url
stringlengths 63
66
| repository_url
stringclasses 1
value | labels_url
stringlengths 77
80
| comments_url
stringlengths 72
75
| events_url
stringlengths 70
73
| html_url
stringlengths 52
56
| id
int64 3.39M
3.04B
| node_id
stringlengths 18
32
| number
int64 1
4.39k
| title
stringlengths 3
367
| user
dict | labels
listlengths 0
5
| state
stringclasses 2
values | locked
bool 2
classes | assignee
dict | assignees
listlengths 0
2
| milestone
null | comments
listlengths 0
30
| created_at
timestamp[s]date 2012-02-26 12:10:46
2025-05-02 18:11:41
| updated_at
timestamp[s]date 2012-02-26 14:52:41
2025-05-02 18:16:27
| closed_at
stringlengths 0
20
| author_association
stringclasses 3
values | type
stringclasses 1
value | active_lock_reason
stringclasses 2
values | sub_issues_summary
dict | body
stringlengths 0
119k
| closed_by
dict | reactions
dict | timeline_url
stringlengths 72
75
| performed_via_github_app
null | state_reason
stringclasses 4
values | is_pull_request
bool 2
classes | draft
bool 2
classes | pull_request
dict |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
https://api.github.com/repos/materialsproject/pymatgen/issues/1901
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1901/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1901/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1901/events
|
https://github.com/materialsproject/pymatgen/pull/1901
| 654,364,094
|
MDExOlB1bGxSZXF1ZXN0NDQ3MTA3MDM3
| 1,901
|
## fix openbabel 3.0 compatibility in lammps submodule
|
{
"login": "orionarcher",
"id": 27712051,
"node_id": "MDQ6VXNlcjI3NzEyMDUx",
"avatar_url": "https://avatars.githubusercontent.com/u/27712051?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/orionarcher",
"html_url": "https://github.com/orionarcher",
"followers_url": "https://api.github.com/users/orionarcher/followers",
"following_url": "https://api.github.com/users/orionarcher/following{/other_user}",
"gists_url": "https://api.github.com/users/orionarcher/gists{/gist_id}",
"starred_url": "https://api.github.com/users/orionarcher/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/orionarcher/subscriptions",
"organizations_url": "https://api.github.com/users/orionarcher/orgs",
"repos_url": "https://api.github.com/users/orionarcher/repos",
"events_url": "https://api.github.com/users/orionarcher/events{/privacy}",
"received_events_url": "https://api.github.com/users/orionarcher/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks."
] | 2020-07-09T21:38:50
| 2020-07-09T22:14:43
|
2020-07-09T22:14:39Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Fix: small changes to important statements to align with openbabel 3.0 syntax
* Fix: removed casting to ASCII bytecode that was causing errors
* Feature: added as_dataframe method for XYZ and associated unit tests
## Additional dependencies introduced
* added imports for numpy and pandas to XYZ
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [ ] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1901/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1901/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1901",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1901",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1901.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1901.patch",
"merged_at": "2020-07-09T22:14:39Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1902
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1902/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1902/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1902/events
|
https://github.com/materialsproject/pymatgen/pull/1902
| 654,835,748
|
MDExOlB1bGxSZXF1ZXN0NDQ3NDgzODI5
| 1,902
|
Update the download link and citations for enumlib
|
{
"login": "wsyxbcl",
"id": 13074950,
"node_id": "MDQ6VXNlcjEzMDc0OTUw",
"avatar_url": "https://avatars.githubusercontent.com/u/13074950?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/wsyxbcl",
"html_url": "https://github.com/wsyxbcl",
"followers_url": "https://api.github.com/users/wsyxbcl/followers",
"following_url": "https://api.github.com/users/wsyxbcl/following{/other_user}",
"gists_url": "https://api.github.com/users/wsyxbcl/gists{/gist_id}",
"starred_url": "https://api.github.com/users/wsyxbcl/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/wsyxbcl/subscriptions",
"organizations_url": "https://api.github.com/users/wsyxbcl/orgs",
"repos_url": "https://api.github.com/users/wsyxbcl/repos",
"events_url": "https://api.github.com/users/wsyxbcl/events{/privacy}",
"received_events_url": "https://api.github.com/users/wsyxbcl/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks."
] | 2020-07-10T14:50:07
| 2020-07-10T15:08:30
|
2020-07-10T15:05:37Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Enumlib has been migrated to github.
1. Update link of enumlib from Sourceforge to Github
2. Update the citations according to https://github.com/msg-byu/enumlib#brief-description
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1902/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1902/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1902",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1902",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1902.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1902.patch",
"merged_at": "2020-07-10T15:05:37Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1903
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1903/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1903/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1903/events
|
https://github.com/materialsproject/pymatgen/issues/1903
| 654,841,474
|
MDU6SXNzdWU2NTQ4NDE0NzQ=
| 1,903
|
undefined symbol: _intel_fast_memcpy
|
{
"login": "bbphysics",
"id": 47345012,
"node_id": "MDQ6VXNlcjQ3MzQ1MDEy",
"avatar_url": "https://avatars.githubusercontent.com/u/47345012?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/bbphysics",
"html_url": "https://github.com/bbphysics",
"followers_url": "https://api.github.com/users/bbphysics/followers",
"following_url": "https://api.github.com/users/bbphysics/following{/other_user}",
"gists_url": "https://api.github.com/users/bbphysics/gists{/gist_id}",
"starred_url": "https://api.github.com/users/bbphysics/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/bbphysics/subscriptions",
"organizations_url": "https://api.github.com/users/bbphysics/orgs",
"repos_url": "https://api.github.com/users/bbphysics/repos",
"events_url": "https://api.github.com/users/bbphysics/events{/privacy}",
"received_events_url": "https://api.github.com/users/bbphysics/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This is a problem with the gcc mismatch on Linux. Do a `pip install --user --upgrade pymatgen` and it should disappear.",
"Since you're running conda, you can also try `conda install -c conda-forge pytmagen` too",
"Thanks, Matthew. \r\nconda install -c conda-forge pymatgen worked for me !!\r\nCentOS release 6.5"
] | 2020-07-10T14:58:48
| 2020-07-11T17:08:13
|
2020-07-10T15:07:09Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I have tried loading Pymatgen both using intel and gcc at local cluster however the issue still persist.
Python 3.7.7 (default, May 7 2020, 21:25:33)
[GCC 7.3.0] :: Anaconda, Inc. on linux
Type "help", "copyright", "credits" or "license" for more information.
>>> import pymatgen
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
File "/home/bb6fq/miniconda3/lib/python3.7/site-packages/pymatgen/__init__.py", line 47, in <module>
from .core.periodic_table import Element, Specie, DummySpecie
File "/home/bb6fq/miniconda3/lib/python3.7/site-packages/pymatgen/core/__init__.py", line 12, in <module>
from .structure import Structure, IStructure, Molecule, IMolecule
File "/home/bb6fq/miniconda3/lib/python3.7/site-packages/pymatgen/core/structure.py", line 31, in <module>
from pymatgen.core.lattice import Lattice, get_points_in_spheres
File "/home/bb6fq/miniconda3/lib/python3.7/site-packages/pymatgen/core/lattice.py", line 25, in <module>
from pymatgen.util.coord import pbc_shortest_vectors
File "/home/bb6fq/miniconda3/lib/python3.7/site-packages/pymatgen/util/coord.py", line 15, in <module>
from . import coord_cython as cuc
ImportError: /home/bb6fq/miniconda3/lib/python3.7/site-packages/pymatgen/util/coord_cython.cpython-37m-x86_64-linux-gnu.so: undefined symbol: _intel_fast_memcpy
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1903/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1903/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1904
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1904/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1904/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1904/events
|
https://github.com/materialsproject/pymatgen/pull/1904
| 654,897,671
|
MDExOlB1bGxSZXF1ZXN0NDQ3NTMzMzk2
| 1,904
|
Fix serialisation of slabs with list scale_factor
|
{
"login": "utf",
"id": 1330638,
"node_id": "MDQ6VXNlcjEzMzA2Mzg=",
"avatar_url": "https://avatars.githubusercontent.com/u/1330638?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/utf",
"html_url": "https://github.com/utf",
"followers_url": "https://api.github.com/users/utf/followers",
"following_url": "https://api.github.com/users/utf/following{/other_user}",
"gists_url": "https://api.github.com/users/utf/gists{/gist_id}",
"starred_url": "https://api.github.com/users/utf/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/utf/subscriptions",
"organizations_url": "https://api.github.com/users/utf/orgs",
"repos_url": "https://api.github.com/users/utf/repos",
"events_url": "https://api.github.com/users/utf/events{/privacy}",
"received_events_url": "https://api.github.com/users/utf/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @utf "
] | 2020-07-10T16:30:46
| 2020-07-10T16:32:33
|
2020-07-10T16:32:33Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
A recent bug fix (1a3ce601c932ed80db14930da6e2c11a8a246771) introduced an incompatibility of the `Slab` class with the atomate `TransformerFW` and therefore broke surface workflows. Specifically, the `Slab` class now assumes the `scale_factor` input is a numpy array, whereas the `TransformerFW` supplies a list `scale_factor`. This caused an error during serialisation of the Slab when `tolist()` was called on a list.
This PR casts the `scale_factor` to a numpy array during initialisation of the Slab.
I've added a test.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1904/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1904/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1904",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1904",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1904.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1904.patch",
"merged_at": "2020-07-10T16:32:33Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1905
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1905/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1905/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1905/events
|
https://github.com/materialsproject/pymatgen/issues/1905
| 655,014,984
|
MDU6SXNzdWU2NTUwMTQ5ODQ=
| 1,905
|
QChem Frequency Calculation Import Fails
|
{
"login": "asmith97",
"id": 3400569,
"node_id": "MDQ6VXNlcjM0MDA1Njk=",
"avatar_url": "https://avatars.githubusercontent.com/u/3400569?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/asmith97",
"html_url": "https://github.com/asmith97",
"followers_url": "https://api.github.com/users/asmith97/followers",
"following_url": "https://api.github.com/users/asmith97/following{/other_user}",
"gists_url": "https://api.github.com/users/asmith97/gists{/gist_id}",
"starred_url": "https://api.github.com/users/asmith97/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/asmith97/subscriptions",
"organizations_url": "https://api.github.com/users/asmith97/orgs",
"repos_url": "https://api.github.com/users/asmith97/repos",
"events_url": "https://api.github.com/users/asmith97/events{/privacy}",
"received_events_url": "https://api.github.com/users/asmith97/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @asmith97 for the report, would it be possible to share your output file to diagnose? @samblau could you assign someone to this, or know someone who could take a look?",
"I'm happy to take a look and diagnose. @asmith97, please provide your output, and I'll get right on it!",
"Thank you! I redid a frequency calculation with benzene to see if I got the same error, and it threw the same error. I was using `QCOutput.multiple_outputs_from_file(QCOutput, 'benzene.txt')`\r\n[benzene.txt](https://github.com/materialsproject/pymatgen/files/4906403/benzene.txt)\r\n\r\n",
"Hi @asmith97 - I see the problem. We never calculate Raman intensities, so the regex parsing the frequency modes assumes that they will be immediately preceded by the \"Raman Active: ....\" line rather than the \"Depolar: ...\" line like they are once you set doraman = true. I can work on a fix on Monday that will resolve the issue you're seeing and also parse the Raman intensities and Depolar values. ",
"Thanks!"
] | 2020-07-10T20:20:12
| 2020-07-14T02:17:33
|
2020-07-14T02:17:33Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I'm unable to use QCOutput to import data from a QChem frequency calculation. An error is thrown while loading the eigenvectors of the vibrational modes. I get an `IndexError: list index out of range` from `shape=(len(freqs), len(temp_freq_mode_vecs[0]), 3))` in line 558 of _read_frequency_data because temp_freq_mode_vecs is an array with no elements. I printed out the contents of temp_freq_mode_vecs to confirm this. It seems like either there's an issue with the QChem output file that I have, which comes directly from a QChem frequency calculation, or the regex pattern given to read_table_pattern isn't working. I'm not sure if other people have ran into this issue or if this is the right place for such a message.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1905/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1905/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1906
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1906/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1906/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1906/events
|
https://github.com/materialsproject/pymatgen/issues/1906
| 655,068,051
|
MDU6SXNzdWU2NTUwNjgwNTE=
| 1,906
|
spglib bugs
|
{
"login": "acrutt",
"id": 42151899,
"node_id": "MDQ6VXNlcjQyMTUxODk5",
"avatar_url": "https://avatars.githubusercontent.com/u/42151899?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/acrutt",
"html_url": "https://github.com/acrutt",
"followers_url": "https://api.github.com/users/acrutt/followers",
"following_url": "https://api.github.com/users/acrutt/following{/other_user}",
"gists_url": "https://api.github.com/users/acrutt/gists{/gist_id}",
"starred_url": "https://api.github.com/users/acrutt/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/acrutt/subscriptions",
"organizations_url": "https://api.github.com/users/acrutt/orgs",
"repos_url": "https://api.github.com/users/acrutt/repos",
"events_url": "https://api.github.com/users/acrutt/events{/privacy}",
"received_events_url": "https://api.github.com/users/acrutt/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
] | null |
[
"I don't consider 1 as a bug. An atom existing at the same space is a bad structure.\r\n2 should be addressed and @mkhorton can deal with it.",
"For 1, we should at least make clear the None type is possible to be returned. Will address."
] | 2020-07-10T22:28:15
| 2023-08-13T16:33:43
|
2023-08-13T16:33:43Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
spglib returns None causing errors with SpaceGroupAnalyzer in two cases:
1. Duplicate atom in the structure
2. None magnetic moment assigned to sites in structure
[pymatgen_issue.zip](https://github.com/materialsproject/pymatgen/files/4905854/pymatgen_issue.zip)
<details><summary>Repro in zip archive</summary>
```py
# Get Structures
# %%
from pymatgen import Structure
from pymatgen.symmetry.analyzer import SpacegroupAnalyzer
# %%
has_magmom = Structure.from_file("has_magmom.json")
print(has_magmom)
# %%
duplicate_atoms_no_magmom = Structure.from_file("no_magmom_duplicate_atoms.json")
print(duplicate_atoms_no_magmom)
# SpaceGroupAnalyzer Bug from Duplicate Atoms
# %%
print(has_magmom.get_space_group_info())
# %%
print(duplicate_atoms_no_magmom.get_space_group_info())
# Fix 1: Remove duplicate atoms
# %%
print(len(duplicate_atoms_no_magmom))
duplicate_atoms_no_magmom.merge_sites(mode="delete")
print(len(duplicate_atoms_no_magmom))
print(duplicate_atoms_no_magmom.get_space_group_info())
# SpaceGroupAnalyzer Bug from magmom=None
# %%
combined_sites = list()
combined_sites.extend(duplicate_atoms_no_magmom.sites)
combined_sites.extend(has_magmom.sites)
print(len(combined_sites))
combined_struct = Structure.from_sites(combined_sites)
# %%
for p in combined_struct.site_properties:
print(p)
for s in combined_struct:
print(s.properties)
# %%
sga = SpacegroupAnalyzer(combined_struct, 0.01)
symm_struct = sga.get_symmetrized_structure()
# %%
combined_struct.remove_site_property("magmom")
for p in combined_struct.site_properties:
print(p)
# %%
sga = SpacegroupAnalyzer(combined_struct, 0.01)
symm_struct = sga.get_symmetrized_structure()
```
</details>
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1906/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1906/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1907
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1907/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1907/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1907/events
|
https://github.com/materialsproject/pymatgen/issues/1907
| 655,832,565
|
MDU6SXNzdWU2NTU4MzI1NjU=
| 1,907
|
Build is failing
|
{
"login": "ramav87",
"id": 14301205,
"node_id": "MDQ6VXNlcjE0MzAxMjA1",
"avatar_url": "https://avatars.githubusercontent.com/u/14301205?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ramav87",
"html_url": "https://github.com/ramav87",
"followers_url": "https://api.github.com/users/ramav87/followers",
"following_url": "https://api.github.com/users/ramav87/following{/other_user}",
"gists_url": "https://api.github.com/users/ramav87/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ramav87/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ramav87/subscriptions",
"organizations_url": "https://api.github.com/users/ramav87/orgs",
"repos_url": "https://api.github.com/users/ramav87/repos",
"events_url": "https://api.github.com/users/ramav87/events{/privacy}",
"received_events_url": "https://api.github.com/users/ramav87/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Please use conda install. The instructions are on pymatgen.org."
] | 2020-07-13T13:01:07
| 2020-07-13T15:06:12
|
2020-07-13T15:06:12Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Doing a pip install fails during wheel building, and TravisCI also shows the build failing.
```
Building wheels for collected packages: pymatgen, retrying
Building wheel for pymatgen (setup.py) ... error
ERROR: Complete output from command //anaconda3/bin/python -u -c 'import setuptools, tokenize;__file__='"'"'/private/var/folders/r8/pgjpxss144164gvc52xvk_cj12sy6v/T/pip-install-h86cvy80/pymatgen/setup.py'"'"';f=getattr(tokenize, '"'"'open'"'"', open)(__file__);code=f.read().replace('"'"'\r\n'"'"', '"'"'\n'"'"');f.close();exec(compile(code, __file__, '"'"'exec'"'"'))' bdist_wheel -d /private/var/folders/r8/pgjpxss144164gvc52xvk_cj12sy6v/T/pip-wheel-6s_vb55t --python-tag cp37:
ERROR: running bdist_wheel
running build
running build_py
creating build
creating build/lib.macosx-10.7-x86_64-3.7
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen
copying pymatgen/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen
copying pymatgen/dao.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/entries
copying pymatgen/entries/compatibility.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/entries
copying pymatgen/entries/computed_entries.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/entries
copying pymatgen/entries/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/entries
copying pymatgen/entries/entry_tools.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/entries
copying pymatgen/entries/exp_entries.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/entries
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/xcfunc.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/periodic_table.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/units.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/ion.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/libxcfunc.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/tensors.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/trajectory.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/operations.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/molecular_orbitals.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/spectrum.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/bonds.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/sites.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/lattice.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/structure.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/composition.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/surface.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/transformations
copying pymatgen/transformations/defect_transformations.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/transformations
copying pymatgen/transformations/standard_transformations.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/transformations
copying pymatgen/transformations/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/transformations
copying pymatgen/transformations/transformation_abc.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/transformations
copying pymatgen/transformations/advanced_transformations.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/transformations
copying pymatgen/transformations/site_transformations.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/transformations
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/fragmenter.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/eos.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/energy_models.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/path_finder.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/reaction_calculator.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/find_dimension.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/diffusion_analyzer.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/surface_analysis.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/structure_matcher.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/pourbaix_diagram.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/wulff.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/local_env.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/phase_diagram.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/molecule_matcher.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/structure_analyzer.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/bond_dissociation.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/prototypes.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/piezo.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/ewald.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/nmr.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/interface.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/molecule_structure_comparator.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/thermochemistry.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/bond_valence.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/excitation.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/cost.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/dimensionality.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/adsorption.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/graphs.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/hhi.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/transition_state.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/piezo_sensitivity.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/quasiharmonic.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/substrate_analyzer.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/functional_groups.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/interface_reactions.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/util
copying pymatgen/util/convergence.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util
copying pymatgen/util/num.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util
copying pymatgen/util/sequence.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util
copying pymatgen/util/plotting.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util
copying pymatgen/util/coord.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util
copying pymatgen/util/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util
copying pymatgen/util/io_utils.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util
copying pymatgen/util/string.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util
copying pymatgen/util/provenance.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util
copying pymatgen/util/typing.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util
copying pymatgen/util/testing.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util
copying pymatgen/util/serialization.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/plugins
copying pymatgen/plugins/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/plugins
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/optimization
copying pymatgen/optimization/linear_assignment_numpy.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/optimization
copying pymatgen/optimization/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/optimization
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/ext
copying pymatgen/ext/jhu.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/ext
copying pymatgen/ext/crystalsai.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/ext
copying pymatgen/ext/cod.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/ext
copying pymatgen/ext/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/ext
copying pymatgen/ext/matproj.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/ext
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/fiesta.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/ase.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/xyz.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/atat.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/aiida.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/nwchem.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/xr.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/lmto.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/zeopp.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/pwscf.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/babel.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/shengbte.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/cssr.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/phonopy.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/wannier90.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/cif.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/prismatic.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/gaussian.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/xcrysden.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
copying pymatgen/io/adf.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/phonon
copying pymatgen/phonon/dos.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/phonon
copying pymatgen/phonon/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/phonon
copying pymatgen/phonon/ir_spectra.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/phonon
copying pymatgen/phonon/plotter.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/phonon
copying pymatgen/phonon/bandstructure.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/phonon
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/cli
copying pymatgen/cli/feff_plot_dos.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/cli
copying pymatgen/cli/pmg_structure.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/cli
copying pymatgen/cli/pmg_query.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/cli
copying pymatgen/cli/pmg_plot.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/cli
copying pymatgen/cli/feff_plot_cross_section.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/cli
copying pymatgen/cli/pmg_potcar.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/cli
copying pymatgen/cli/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/cli
copying pymatgen/cli/pmg_analyze.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/cli
copying pymatgen/cli/pmg_config.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/cli
copying pymatgen/cli/gaussian_analyzer.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/cli
copying pymatgen/cli/get_environment.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/cli
copying pymatgen/cli/pmg.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/cli
copying pymatgen/cli/feff_input_generation.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/cli
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/symmetry
copying pymatgen/symmetry/analyzer.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/symmetry
copying pymatgen/symmetry/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/symmetry
copying pymatgen/symmetry/kpath.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/symmetry
copying pymatgen/symmetry/site_symmetries.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/symmetry
copying pymatgen/symmetry/groups.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/symmetry
copying pymatgen/symmetry/structure.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/symmetry
copying pymatgen/symmetry/settings.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/symmetry
copying pymatgen/symmetry/maggroups.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/symmetry
copying pymatgen/symmetry/bandstructure.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/symmetry
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/vis
copying pymatgen/vis/structure_vtk.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/vis
copying pymatgen/vis/plotters.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/vis
copying pymatgen/vis/structure_chemview.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/vis
copying pymatgen/vis/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/vis
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/electronic_structure
copying pymatgen/electronic_structure/cohp.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/electronic_structure
copying pymatgen/electronic_structure/dos.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/electronic_structure
copying pymatgen/electronic_structure/boltztrap.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/electronic_structure
copying pymatgen/electronic_structure/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/electronic_structure
copying pymatgen/electronic_structure/core.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/electronic_structure
copying pymatgen/electronic_structure/plotter.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/electronic_structure
copying pymatgen/electronic_structure/boltztrap2.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/electronic_structure
copying pymatgen/electronic_structure/bandstructure.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/electronic_structure
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/command_line
copying pymatgen/command_line/vampire_caller.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/command_line
copying pymatgen/command_line/bader_caller.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/command_line
copying pymatgen/command_line/critic2_caller.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/command_line
copying pymatgen/command_line/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/command_line
copying pymatgen/command_line/enumlib_caller.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/command_line
copying pymatgen/command_line/aconvasp_caller.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/command_line
copying pymatgen/command_line/gulp_caller.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/command_line
copying pymatgen/command_line/mcsqs_caller.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/command_line
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/apps
copying pymatgen/apps/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/apps
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/alchemy
copying pymatgen/alchemy/materials.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/alchemy
copying pymatgen/alchemy/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/alchemy
copying pymatgen/alchemy/transmuters.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/alchemy
copying pymatgen/alchemy/filters.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/alchemy
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/elasticity
copying pymatgen/analysis/elasticity/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/elasticity
copying pymatgen/analysis/elasticity/stress.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/elasticity
copying pymatgen/analysis/elasticity/elastic.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/elasticity
copying pymatgen/analysis/elasticity/strain.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/elasticity
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/structure_prediction
copying pymatgen/analysis/structure_prediction/volume_predictor.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/structure_prediction
copying pymatgen/analysis/structure_prediction/substitution_probability.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/structure_prediction
copying pymatgen/analysis/structure_prediction/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/structure_prediction
copying pymatgen/analysis/structure_prediction/substitutor.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/structure_prediction
copying pymatgen/analysis/structure_prediction/dopant_predictor.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/structure_prediction
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/xas
copying pymatgen/analysis/xas/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/xas
copying pymatgen/analysis/xas/spectrum.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/xas
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/defects
copying pymatgen/analysis/defects/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/defects
copying pymatgen/analysis/defects/core.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/defects
copying pymatgen/analysis/defects/dilute_solution_model.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/defects
copying pymatgen/analysis/defects/generators.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/defects
copying pymatgen/analysis/defects/thermodynamics.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/defects
copying pymatgen/analysis/defects/utils.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/defects
copying pymatgen/analysis/defects/corrections.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/defects
copying pymatgen/analysis/defects/defect_compatibility.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/defects
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv
copying pymatgen/analysis/chemenv/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/gb
copying pymatgen/analysis/gb/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/gb
copying pymatgen/analysis/gb/grain.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/gb
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/diffraction
copying pymatgen/analysis/diffraction/neutron.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/diffraction
copying pymatgen/analysis/diffraction/xrd.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/diffraction
copying pymatgen/analysis/diffraction/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/diffraction
copying pymatgen/analysis/diffraction/core.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/diffraction
copying pymatgen/analysis/diffraction/tem.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/diffraction
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/ferroelectricity
copying pymatgen/analysis/ferroelectricity/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/ferroelectricity
copying pymatgen/analysis/ferroelectricity/polarization.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/ferroelectricity
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/topological
copying pymatgen/analysis/topological/spillage.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/topological
copying pymatgen/analysis/topological/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/topological
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/magnetism
copying pymatgen/analysis/magnetism/analyzer.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/magnetism
copying pymatgen/analysis/magnetism/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/magnetism
copying pymatgen/analysis/magnetism/heisenberg.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/magnetism
copying pymatgen/analysis/magnetism/jahnteller.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/magnetism
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/connectivity
copying pymatgen/analysis/chemenv/connectivity/connected_components.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/connectivity
copying pymatgen/analysis/chemenv/connectivity/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/connectivity
copying pymatgen/analysis/chemenv/connectivity/structure_connectivity.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/connectivity
copying pymatgen/analysis/chemenv/connectivity/connectivity_finder.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/connectivity
copying pymatgen/analysis/chemenv/connectivity/environment_nodes.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/connectivity
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometry_finder.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments
copying pymatgen/analysis/chemenv/coordination_environments/structure_environments.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments
copying pymatgen/analysis/chemenv/coordination_environments/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments
copying pymatgen/analysis/chemenv/coordination_environments/voronoi.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments
copying pymatgen/analysis/chemenv/coordination_environments/chemenv_strategies.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/utils
copying pymatgen/analysis/chemenv/utils/chemenv_config.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/utils
copying pymatgen/analysis/chemenv/utils/coordination_geometry_utils.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/utils
copying pymatgen/analysis/chemenv/utils/func_utils.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/utils
copying pymatgen/analysis/chemenv/utils/math_utils.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/utils
copying pymatgen/analysis/chemenv/utils/defs_utils.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/utils
copying pymatgen/analysis/chemenv/utils/graph_utils.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/utils
copying pymatgen/analysis/chemenv/utils/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/utils
copying pymatgen/analysis/chemenv/utils/scripts_utils.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/utils
copying pymatgen/analysis/chemenv/utils/chemenv_errors.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/utils
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/strategy_files
copying pymatgen/analysis/chemenv/coordination_environments/strategy_files/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/strategy_files
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/feff
copying pymatgen/io/feff/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/feff
copying pymatgen/io/feff/inputs.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/feff
copying pymatgen/io/feff/sets.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/feff
copying pymatgen/io/feff/outputs.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/feff
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/lobster
copying pymatgen/io/lobster/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/lobster
copying pymatgen/io/lobster/inputs.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/lobster
copying pymatgen/io/lobster/outputs.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/lobster
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/abinit
copying pymatgen/io/abinit/abiobjects.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/abinit
copying pymatgen/io/abinit/abiinspect.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/abinit
copying pymatgen/io/abinit/abitimer.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/abinit
copying pymatgen/io/abinit/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/abinit
copying pymatgen/io/abinit/variable.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/abinit
copying pymatgen/io/abinit/netcdf.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/abinit
copying pymatgen/io/abinit/inputs.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/abinit
copying pymatgen/io/abinit/pseudos.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/abinit
copying pymatgen/io/abinit/helpers.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/abinit
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/vasp
copying pymatgen/io/vasp/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/vasp
copying pymatgen/io/vasp/inputs.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/vasp
copying pymatgen/io/vasp/sets.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/vasp
copying pymatgen/io/vasp/help.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/vasp
copying pymatgen/io/vasp/outputs.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/vasp
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/qchem
copying pymatgen/io/qchem/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/qchem
copying pymatgen/io/qchem/inputs.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/qchem
copying pymatgen/io/qchem/utils.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/qchem
copying pymatgen/io/qchem/sets.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/qchem
copying pymatgen/io/qchem/outputs.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/qchem
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/exciting
copying pymatgen/io/exciting/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/exciting
copying pymatgen/io/exciting/inputs.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/exciting
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/lammps
copying pymatgen/io/lammps/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/lammps
copying pymatgen/io/lammps/inputs.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/lammps
copying pymatgen/io/lammps/utils.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/lammps
copying pymatgen/io/lammps/data.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/lammps
copying pymatgen/io/lammps/outputs.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/lammps
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/apps/borg
copying pymatgen/apps/borg/queen.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/apps/borg
copying pymatgen/apps/borg/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/apps/borg
copying pymatgen/apps/borg/hive.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/apps/borg
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/apps/battery
copying pymatgen/apps/battery/conversion_battery.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/apps/battery
copying pymatgen/apps/battery/analyzer.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/apps/battery
copying pymatgen/apps/battery/__init__.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/apps/battery
copying pymatgen/apps/battery/insertion_battery.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/apps/battery
copying pymatgen/apps/battery/plotter.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/apps/battery
copying pymatgen/apps/battery/battery_abc.py -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/apps/battery
copying pymatgen/entries/MPCompatibility.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/entries
copying pymatgen/entries/MITCompatibility.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/entries
copying pymatgen/core/reconstructions_archive.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/bond_lengths.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/func_groups.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/periodic_table.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/libxc_docs.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/quad_data.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/core/py.typed -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/core
copying pymatgen/analysis/icsd_bv.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/bonds_jmol_ob.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/vesta_cutoffs.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/bvparam_1991.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/op_params.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/cn_opt_params.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/ionic_radii.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/aflow_prototypes.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/costdb_elements.csv -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
copying pymatgen/analysis/hhi_data.csv -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/SiO2.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/Li2O.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/LiFePO4.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/Li10GeP2S12.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/TlBiSe2.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/K2O2.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/Li3V2(PO4)3.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/Graphite.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/He_BCC.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/TiO2.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/CsCl.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/BaNiO3.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/Li2O2.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/Si.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/NaFePO4.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/VO2.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/Sn.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/Pb2TiZrO6.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/util/structures/SrTiO3.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/structures
copying pymatgen/symmetry/symm_data.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/symmetry
copying pymatgen/symmetry/symm_ops.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/symmetry
copying pymatgen/symmetry/symm_data.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/symmetry
copying pymatgen/symmetry/symm_ops.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/symmetry
copying pymatgen/symmetry/symm_data_magnetic.sqlite -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/symmetry
copying pymatgen/vis/ElementColorSchemes.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/vis
copying pymatgen/command_line/OxideTersoffPotentials -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/command_line
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/structure_prediction/data
copying pymatgen/analysis/structure_prediction/data/pair_correlation.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/structure_prediction/data
copying pymatgen/analysis/structure_prediction/data/lambda.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/structure_prediction/data
copying pymatgen/analysis/structure_prediction/DLS_bond_params.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/structure_prediction
copying pymatgen/analysis/diffraction/neutron_scattering_length.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/diffraction
copying pymatgen/analysis/diffraction/atomic_scattering_params.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/diffraction
copying pymatgen/analysis/magnetism/default_magmoms.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/magnetism
copying pymatgen/analysis/chemenv/coordination_environments/strategy_files/ImprovedConfidenceCutoffDefaultParameters.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/strategy_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/allcg.txt -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/BO_2#8.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/H#10.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/TY#3.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/PCPA#11.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/T#4.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/TT_2#9.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/TBT#8.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/TI#9.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/T#5.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/AC#12.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/SBSA#10.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/SH#13.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/HP#12.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/FO#7.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/H#11.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/S#12.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/TBSA#10.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/L#2.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/DI#11.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/TO_1#9.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/DDPN#8.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/TS#3.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/SMA#9.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/PA#10.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/TO_3#9.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/PP#5.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/S#1.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/SBT#8.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/SC#12.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/TC#9.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/DD#8.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/TT#12.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/PBP#12.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/O#6.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/SS#9.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/TT_3#9.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/S#10.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/TL#3.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/PP#6.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/CO#11.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/BO_3#8.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/ST#7.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/C#8.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/HD#9.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/BS_2#10.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/PP#10.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/SS#4.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/BO_1#8.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/C#12.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/TT_1#9.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/MI#10.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/HB#8.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/ET#7.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/A#2.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/S#4.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/O#6_explicit.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/T#6.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/SA#8.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/S#5.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/SY#4.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/HA#12.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/PB#7.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/SH#11.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/TO_2#9.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/BS_1#10.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/I#12.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files/DD#20.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/analysis/chemenv/coordination_environments/coordination_geometries_files
copying pymatgen/io/feff/MPEXAFSSet.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/feff
copying pymatgen/io/feff/MPELNESSet.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/feff
copying pymatgen/io/feff/MPXANESSet.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/feff
copying pymatgen/io/feff/MPEXELFSSet.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/feff
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/lobster/lobster_basis
copying pymatgen/io/lobster/lobster_basis/BASIS_PBE_54_min.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/lobster/lobster_basis
copying pymatgen/io/lobster/lobster_basis/BASIS_PBE_54_max.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/lobster/lobster_basis
copying pymatgen/io/lobster/lobster_basis/BASIS_PBE_54_standard.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/lobster/lobster_basis
copying pymatgen/io/vasp/MVLGWSet.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/vasp
copying pymatgen/io/vasp/MVLRelax52Set.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/vasp
copying pymatgen/io/vasp/MPHSERelaxSet.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/vasp
copying pymatgen/io/vasp/VASPIncarBase.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/vasp
copying pymatgen/io/vasp/MPSCANRelaxSet.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/vasp
copying pymatgen/io/vasp/MPRelaxSet.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/vasp
copying pymatgen/io/vasp/MITRelaxSet.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/vasp
copying pymatgen/io/vasp/vdW_parameters.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/vasp
copying pymatgen/io/vasp/vasp_potcar_pymatgen_hashes.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/vasp
copying pymatgen/io/vasp/incar_parameters.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/vasp
copying pymatgen/io/vasp/vasp_potcar_file_hashes.json -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/vasp
creating build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/lammps/templates
copying pymatgen/io/lammps/templates/md.txt -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/lammps/templates
copying pymatgen/io/lammps/CoeffsDataType.yaml -> build/lib.macosx-10.7-x86_64-3.7/pymatgen/io/lammps
running build_ext
building 'pymatgen.optimization.linear_assignment' extension
creating build/temp.macosx-10.7-x86_64-3.7
creating build/temp.macosx-10.7-x86_64-3.7/pymatgen
creating build/temp.macosx-10.7-x86_64-3.7/pymatgen/optimization
gcc -Wno-unused-result -Wsign-compare -Wunreachable-code -DNDEBUG -g -fwrapv -O3 -Wall -Wstrict-prototypes -I//anaconda3/include -arch x86_64 -I//anaconda3/include -arch x86_64 -I//anaconda3/include/python3.7m -I//anaconda3/lib/python3.7/site-packages/numpy/core/include -c pymatgen/optimization/linear_assignment.c -o build/temp.macosx-10.7-x86_64-3.7/pymatgen/optimization/linear_assignment.o
In file included from //anaconda3/lib/python3.7/site-packages/numpy/core/include/numpy/ndarraytypes.h:1832:0,
from //anaconda3/lib/python3.7/site-packages/numpy/core/include/numpy/ndarrayobject.h:12,
from //anaconda3/lib/python3.7/site-packages/numpy/core/include/numpy/arrayobject.h:4,
from pymatgen/optimization/linear_assignment.c:569:
//anaconda3/lib/python3.7/site-packages/numpy/core/include/numpy/npy_1_7_deprecated_api.h:17:2: warning: #warning "Using deprecated NumPy API, disable it with " "#define NPY_NO_DEPRECATED_API NPY_1_7_API_VERSION" [-Wcpp]
#warning "Using deprecated NumPy API, disable it with " \
^
pymatgen/optimization/linear_assignment.c: In function '__pyx_pf_8pymatgen_12optimization_17linear_assignment_16LinearAssignment___init__.isra.45':
pymatgen/optimization/linear_assignment.c:4673:42: warning: '__pyx_v_last' may be used uninitialized in this function [-Wmaybe-uninitialized]
__pyx_t_45 = (__pyx_v_last + 1);
^
pymatgen/optimization/linear_assignment.c:3115:7: note: '__pyx_v_last' was declared here
int __pyx_v_last;
^
pymatgen/optimization/linear_assignment.c:3959:20: warning: '__pyx_v_j2' may be used uninitialized in this function [-Wmaybe-uninitialized]
__pyx_t_37 = __pyx_v_j1;
^
pymatgen/optimization/linear_assignment.c:3129:7: note: '__pyx_v_j2' was declared here
int __pyx_v_j2;
^
pymatgen/optimization/linear_assignment.c:4652:41: warning: '__pyx_v_m' may be used uninitialized in this function [-Wmaybe-uninitialized]
__pyx_t_9 = ((fabs((__pyx_v_h - __pyx_v_m)) < __pyx_v_eps) != 0);
^
pymatgen/optimization/linear_assignment.c:3125:26: note: '__pyx_v_m' was declared here
__pyx_t_5numpy_float_t __pyx_v_m;
^
gcc -bundle -undefined dynamic_lookup -L//anaconda3/lib -arch x86_64 -L//anaconda3/lib -arch x86_64 -arch x86_64 build/temp.macosx-10.7-x86_64-3.7/pymatgen/optimization/linear_assignment.o -o build/lib.macosx-10.7-x86_64-3.7/pymatgen/optimization/linear_assignment.cpython-37m-darwin.so
building 'pymatgen.util.coord_cython' extension
creating build/temp.macosx-10.7-x86_64-3.7/pymatgen/util
gcc -Wno-unused-result -Wsign-compare -Wunreachable-code -DNDEBUG -g -fwrapv -O3 -Wall -Wstrict-prototypes -I//anaconda3/include -arch x86_64 -I//anaconda3/include -arch x86_64 -I//anaconda3/include/python3.7m -I//anaconda3/lib/python3.7/site-packages/numpy/core/include -c pymatgen/util/coord_cython.c -o build/temp.macosx-10.7-x86_64-3.7/pymatgen/util/coord_cython.o
In file included from //anaconda3/lib/python3.7/site-packages/numpy/core/include/numpy/ndarraytypes.h:1832:0,
from //anaconda3/lib/python3.7/site-packages/numpy/core/include/numpy/ndarrayobject.h:12,
from //anaconda3/lib/python3.7/site-packages/numpy/core/include/numpy/arrayobject.h:4,
from pymatgen/util/coord_cython.c:569:
//anaconda3/lib/python3.7/site-packages/numpy/core/include/numpy/npy_1_7_deprecated_api.h:17:2: warning: #warning "Using deprecated NumPy API, disable it with " "#define NPY_NO_DEPRECATED_API NPY_1_7_API_VERSION" [-Wcpp]
#warning "Using deprecated NumPy API, disable it with " \
^
pymatgen/util/coord_cython.c: In function '__pyx_pf_8pymatgen_4util_12coord_cython_2is_coord_subset_pbc.isra.36':
pymatgen/util/coord_cython.c:28271:36: warning: '__pyx_v_ok' may be used uninitialized in this function [-Wmaybe-uninitialized]
return b ? __Pyx_NewRef(Py_True) : __Pyx_NewRef(Py_False);
^
pymatgen/util/coord_cython.c:4566:7: note: '__pyx_v_ok' was declared here
int __pyx_v_ok;
^
pymatgen/util/coord_cython.c: In function '__pyx_pf_8pymatgen_4util_12coord_cython_4coord_list_mapping_pbc.isra.35':
pymatgen/util/coord_cython.c:5494:6: warning: '__pyx_v_ok_outer' may be used uninitialized in this function [-Wmaybe-uninitialized]
if (unlikely(__pyx_t_21)) {
^
pymatgen/util/coord_cython.c: In function '__pyx_pf_8pymatgen_4util_12coord_cython_pbc_shortest_vectors.isra.37':
pymatgen/util/coord_cython.c:3102:7: warning: '__pyx_v_bestk' may be used uninitialized in this function [-Wmaybe-uninitialized]
int __pyx_v_bestk;
^
gcc -bundle -undefined dynamic_lookup -L//anaconda3/lib -arch x86_64 -L//anaconda3/lib -arch x86_64 -arch x86_64 build/temp.macosx-10.7-x86_64-3.7/pymatgen/util/coord_cython.o -o build/lib.macosx-10.7-x86_64-3.7/pymatgen/util/coord_cython.cpython-37m-darwin.so
building 'pymatgen.optimization.neighbors' extension
gcc -Wno-unused-result -Wsign-compare -Wunreachable-code -DNDEBUG -g -fwrapv -O3 -Wall -Wstrict-prototypes -I//anaconda3/include -arch x86_64 -I//anaconda3/include -arch x86_64 -I//anaconda3/include/python3.7m -I//anaconda3/lib/python3.7/site-packages/numpy/core/include -c pymatgen/optimization/neighbors.cpp -o build/temp.macosx-10.7-x86_64-3.7/pymatgen/optimization/neighbors.o -Wno-cpp -Wno-unused-function -O2 -march=native -std=c++0x -stdlib=libc++
gcc: error: unrecognized command line option '-stdlib=libc++'
error: command 'gcc' failed with exit status 1
----------------------------------------
ERROR: Failed building wheel for pymatgen
Running setup.py clean for pymatgen
```
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1907/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1907/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1908
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1908/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1908/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1908/events
|
https://github.com/materialsproject/pymatgen/pull/1908
| 656,258,628
|
MDExOlB1bGxSZXF1ZXN0NDQ4NTkxMTE0
| 1,908
|
Fixing QChem frequency parsing when calculating Raman intensities
|
{
"login": "samblau",
"id": 7431223,
"node_id": "MDQ6VXNlcjc0MzEyMjM=",
"avatar_url": "https://avatars.githubusercontent.com/u/7431223?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/samblau",
"html_url": "https://github.com/samblau",
"followers_url": "https://api.github.com/users/samblau/followers",
"following_url": "https://api.github.com/users/samblau/following{/other_user}",
"gists_url": "https://api.github.com/users/samblau/gists{/gist_id}",
"starred_url": "https://api.github.com/users/samblau/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/samblau/subscriptions",
"organizations_url": "https://api.github.com/users/samblau/orgs",
"repos_url": "https://api.github.com/users/samblau/repos",
"events_url": "https://api.github.com/users/samblau/events{/privacy}",
"received_events_url": "https://api.github.com/users/samblau/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks."
] | 2020-07-14T01:37:07
| 2020-07-14T02:13:13
|
2020-07-14T02:13:07Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
* Fixes issue #1905
* All frequency information can now be correctly parsed even when Raman intensities are being calculated
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1908/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1908/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1908",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1908",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1908.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1908.patch",
"merged_at": "2020-07-14T02:13:07Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1909
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1909/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1909/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1909/events
|
https://github.com/materialsproject/pymatgen/issues/1909
| 656,908,469
|
MDU6SXNzdWU2NTY5MDg0Njk=
| 1,909
|
Labelling for COHP bonds
|
{
"login": "CifLord",
"id": 8036054,
"node_id": "MDQ6VXNlcjgwMzYwNTQ=",
"avatar_url": "https://avatars.githubusercontent.com/u/8036054?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CifLord",
"html_url": "https://github.com/CifLord",
"followers_url": "https://api.github.com/users/CifLord/followers",
"following_url": "https://api.github.com/users/CifLord/following{/other_user}",
"gists_url": "https://api.github.com/users/CifLord/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CifLord/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CifLord/subscriptions",
"organizations_url": "https://api.github.com/users/CifLord/orgs",
"repos_url": "https://api.github.com/users/CifLord/repos",
"events_url": "https://api.github.com/users/CifLord/events{/privacy}",
"received_events_url": "https://api.github.com/users/CifLord/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi!\r\nThe label corresponds to the label from the ICOHPLIST. We could discuss alternatives but I preferred this as there can be multiple COHPs for a site combination.\r\n(Let me know if you have a better suggestion that is also comparably easy to use. I don't really want to think about which kind of image of the original site I am currently looking at just to identify a relevant COHP. Best strategy for me: identify relevant bonds from ICOHPLIST and then plot those with the labels from there.)",
"Ah OK then, thank you clearing this up. I agree its a hassle to have to think about what site we're dealing with to get a specific COHP. I'll try to think up alternate scheme to organize this since as you said there can be more than one COHP for a pair. ",
"Just to add: I think still think using the labels from the ICOHPLIST makes it very user-friendly (I would also prefer to keep it this way because you can easily identify relevant bonds by checking the ICOHPLIST) but we might be able to add additional ways to identify the relevant COHPs. I agree that there might be potential for improvement."
] | 2020-07-14T21:25:05
| 2023-08-13T16:33:43
|
2023-08-13T16:33:43Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Is there any reason for how an individual bond's COHP gets its label in the CompleteCOHP class (pymatgen.electronic_structure.cohp.py)? It seems to me that the labels are just distinct integers arbitrarily assigned to each bond. If that is the case, I suggest that we might switch to a labeling system where the label is a tuple of the two site indices that make up a COHP bond. This would make searching for a specific COHP via the get_cohp_by_label() method much easier.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1909/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1909/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1910
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1910/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1910/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1910/events
|
https://github.com/materialsproject/pymatgen/pull/1910
| 657,514,418
|
MDExOlB1bGxSZXF1ZXN0NDQ5NjE4MDAw
| 1,910
|
bug and typo fix for boltztrap2
|
{
"login": "fraricci",
"id": 17427517,
"node_id": "MDQ6VXNlcjE3NDI3NTE3",
"avatar_url": "https://avatars.githubusercontent.com/u/17427517?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/fraricci",
"html_url": "https://github.com/fraricci",
"followers_url": "https://api.github.com/users/fraricci/followers",
"following_url": "https://api.github.com/users/fraricci/following{/other_user}",
"gists_url": "https://api.github.com/users/fraricci/gists{/gist_id}",
"starred_url": "https://api.github.com/users/fraricci/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/fraricci/subscriptions",
"organizations_url": "https://api.github.com/users/fraricci/orgs",
"repos_url": "https://api.github.com/users/fraricci/repos",
"events_url": "https://api.github.com/users/fraricci/events{/privacy}",
"received_events_url": "https://api.github.com/users/fraricci/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Pls fix the pylint errors related to boltztrap2.py. The rest of the errors you can ignore for now.",
"Thanks.",
"@fraricci It seems that your changes caused the tests to fail. Can you fix the tests pls? Thanks.",
"it looks like it's not me:\r\n************* Module pymatgen/cli/feff_input_generation.py\r\npymatgen/cli/feff_input_generation.py:1:0: F0001: No module named pymatgen/cli/feff_input_generation.py (fatal)\r\n\r\nAm I right?"
] | 2020-07-15T17:07:43
| 2020-07-16T14:00:58
|
2020-07-15T20:02:10Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1910/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1910/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1910",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1910",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1910.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1910.patch",
"merged_at": "2020-07-15T20:02:10Z"
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/1911
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1911/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1911/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1911/events
|
https://github.com/materialsproject/pymatgen/pull/1911
| 659,723,883
|
MDExOlB1bGxSZXF1ZXN0NDUxNTM4MjM4
| 1,911
|
Add validation and extrapolation for XAS stitching
|
{
"login": "yimingchen-eng",
"id": 27871258,
"node_id": "MDQ6VXNlcjI3ODcxMjU4",
"avatar_url": "https://avatars.githubusercontent.com/u/27871258?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/yimingchen-eng",
"html_url": "https://github.com/yimingchen-eng",
"followers_url": "https://api.github.com/users/yimingchen-eng/followers",
"following_url": "https://api.github.com/users/yimingchen-eng/following{/other_user}",
"gists_url": "https://api.github.com/users/yimingchen-eng/gists{/gist_id}",
"starred_url": "https://api.github.com/users/yimingchen-eng/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/yimingchen-eng/subscriptions",
"organizations_url": "https://api.github.com/users/yimingchen-eng/orgs",
"repos_url": "https://api.github.com/users/yimingchen-eng/repos",
"events_url": "https://api.github.com/users/yimingchen-eng/events{/privacy}",
"received_events_url": "https://api.github.com/users/yimingchen-eng/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"There are failing pylint tests. Pls fix before I merge."
] | 2020-07-17T23:14:28
| 2020-07-18T13:29:20
|
2020-07-18T13:29:20Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Add check for empty spectrum and negative intensities
* Fix the jump in intensities at the L2 and L3-edge junction by extrapolating L2-edge XANES
* Add tests
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1911/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1911/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1911",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1911",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1911.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1911.patch",
"merged_at": "2020-07-18T13:29:20Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1912
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1912/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1912/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1912/events
|
https://github.com/materialsproject/pymatgen/pull/1912
| 661,002,878
|
MDExOlB1bGxSZXF1ZXN0NDUyNzE0NjU2
| 1,912
|
Adding JARVIS-Tools adaptor module.
|
{
"login": "knc6",
"id": 16902896,
"node_id": "MDQ6VXNlcjE2OTAyODk2",
"avatar_url": "https://avatars.githubusercontent.com/u/16902896?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/knc6",
"html_url": "https://github.com/knc6",
"followers_url": "https://api.github.com/users/knc6/followers",
"following_url": "https://api.github.com/users/knc6/following{/other_user}",
"gists_url": "https://api.github.com/users/knc6/gists{/gist_id}",
"starred_url": "https://api.github.com/users/knc6/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/knc6/subscriptions",
"organizations_url": "https://api.github.com/users/knc6/orgs",
"repos_url": "https://api.github.com/users/knc6/repos",
"events_url": "https://api.github.com/users/knc6/events{/privacy}",
"received_events_url": "https://api.github.com/users/knc6/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks. \r\n\r\n1. Can you rename jarvisio to just Jarvis. The idea is that it is pymatgen.io.jarvis. For example, pymatgen.io.vasp, not pymatgen.io.vaspio (this was what we started with, but since IO is already in the earlier package, we don't need io again).\r\n2. Pls pin jarvis-tools to a specific version in requirements-optional.txt."
] | 2020-07-19T18:53:17
| 2020-07-19T22:00:58
|
2020-07-19T22:00:58Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
JARVIS-Tools module integration in Pymatgen.
* Creation of pymatgen.io.jarvis
* Fixed setup.cfg for jarvis modules
* Fixed requirements-optional.txt
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1912/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1912/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1912",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1912",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1912.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1912.patch",
"merged_at": "2020-07-19T22:00:58Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1913
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1913/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1913/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1913/events
|
https://github.com/materialsproject/pymatgen/issues/1913
| 661,081,298
|
MDU6SXNzdWU2NjEwODEyOTg=
| 1,913
|
Running SpacegroupAnalyzer on enumerated structure returns wrong space group
|
{
"login": "DoubleKx",
"id": 65990344,
"node_id": "MDQ6VXNlcjY1OTkwMzQ0",
"avatar_url": "https://avatars.githubusercontent.com/u/65990344?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/DoubleKx",
"html_url": "https://github.com/DoubleKx",
"followers_url": "https://api.github.com/users/DoubleKx/followers",
"following_url": "https://api.github.com/users/DoubleKx/following{/other_user}",
"gists_url": "https://api.github.com/users/DoubleKx/gists{/gist_id}",
"starred_url": "https://api.github.com/users/DoubleKx/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/DoubleKx/subscriptions",
"organizations_url": "https://api.github.com/users/DoubleKx/orgs",
"repos_url": "https://api.github.com/users/DoubleKx/repos",
"events_url": "https://api.github.com/users/DoubleKx/events{/privacy}",
"received_events_url": "https://api.github.com/users/DoubleKx/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"The enumerated structure (ordered approximation) will necessarily be lower symmetry than the input disordered structure, so it's reasonable that it would no longer read as F-43m.",
"Ah, I see. So what about after relaxing the structure with DFT? Is there a function or way to \"rebuild\" the relaxed enumerated primitive structure to the conventional cell in F-43m? ",
"You can substitute back disordered species for your ordered species (e.g. if you started with {A: 0.5, B: 0.5} on a site, created an ordered approximation, after relaxation you could replace A with {A: 0.5, B: 0.5} and B with {A: 0.5, B: 0.5}). You could then do a structure refinement with a loose symmetry tolerance. This is obviously more difficult if A or B is a vacancy rather than another species. In general though, \"going back\" is difficult, precisely because you're lowering the symmetry and there'll be local relaxations around your various species, bond lengths will be changing, etc. so it's not a trivial thing. \r\n\r\nWhat you might find useful is storing the mapping (eg lattice transformation) between primitive and conventional cells and then re-applying that mapping after your DFT relaxation.",
"Thanks! That was very useful. I did this using find_mapping in the lattice module and applied the mapping ",
"Great! Glad it helped."
] | 2020-07-19T21:28:25
| 2020-07-21T16:55:32
|
2020-07-21T16:55:32Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I'm trying to take my enumerated primitive cell and return it back to the conventional cell but it's not finding the original space group.
Steps I took:
1. Input conventional cell (s.g. F-43m) and find the primitive cell using SpacegroupAnalyzer.find_primitive()
2. Enumerated using enumlib (i.e. EnumerateStructureTransformation) OR OrderDisorderedStructureTransformation()
3. Generate POSCAR files of enumerated structures
3. Run enumerated cells through SpacegroupAnalyzer.get_refined_structure() and get_space_group_symbol()
4. Result is always P1 or the incorrect space group. It does not find F-43m for some reason even after adjusting tolerances in SpacegroupAnalyzer
[Input_Files.zip](https://github.com/materialsproject/pymatgen/files/4944706/Input_Files.zip)
See the attached CIF file for the conventional cell and the POSCAR after enumeration in step 3
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1913/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1913/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1914
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1914/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1914/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1914/events
|
https://github.com/materialsproject/pymatgen/issues/1914
| 666,540,593
|
MDU6SXNzdWU2NjY1NDA1OTM=
| 1,914
|
enumlib error
|
{
"login": "haesunpark87",
"id": 43765756,
"node_id": "MDQ6VXNlcjQzNzY1NzU2",
"avatar_url": "https://avatars.githubusercontent.com/u/43765756?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/haesunpark87",
"html_url": "https://github.com/haesunpark87",
"followers_url": "https://api.github.com/users/haesunpark87/followers",
"following_url": "https://api.github.com/users/haesunpark87/following{/other_user}",
"gists_url": "https://api.github.com/users/haesunpark87/gists{/gist_id}",
"starred_url": "https://api.github.com/users/haesunpark87/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/haesunpark87/subscriptions",
"organizations_url": "https://api.github.com/users/haesunpark87/orgs",
"repos_url": "https://api.github.com/users/haesunpark87/repos",
"events_url": "https://api.github.com/users/haesunpark87/events{/privacy}",
"received_events_url": "https://api.github.com/users/haesunpark87/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @haesunpark87, this code works for me, a core dump is likely some serious issue with your installation -- have you tried installing `pymatgen` using conda? Does `enumlib` work outside of pymatgen? (e.g. if you call `enum.x` directly?)",
"Dear Matthew, \n\nIt seems you are right. \nI ran the following code,\n\n#!/usr/bin/env python3\n# -*- coding: utf-8 -*-\nfrom pymatgen.core import Lattice, Structure\nfrom pymatgen.transformations.advanced_transformations import EnumerateStructureTransformation\n\nspecie = {\"Cu\": 0.5, \"Au\": 0.5}\ncuau = Structure.from_spacegroup(\"Fm-3m\", Lattice.cubic(3.677), [specie], [[0, 0, 0]])\n\ntrans = EnumerateStructureTransformation(max_cell_size=3)\nss = trans.apply_transformation(cuau, return_ranked_list=1000)\n\nAnd the error message saying \"RuntimeError: EnumlibAdaptor requires the executables 'enum.x' or 'multienum.x' and 'makestr.x' or 'makeStr.py' to be in the path. Please download the library at http://enum.sourceforge.net/ and follow the instructions in the README to compile these two executables accordingly. “\n\nI have the same error in my desktop and the question is where is the path to put the executables. \n\nBest,\nHaesun\n\n\n> On Jul 27, 2020, at 7:53 PM, Matthew Horton <notifications@github.com> wrote:\n> \n> \n> Hi @haesunpark87 <https://github.com/haesunpark87>, this code works for me, a core dump is likely some serious issue with your installation -- have you tried installing pymatgen using conda? Does enumlib work outside of pymatgen? (e.g. if you call enum.x directly?)\n> \n> —\n> You are receiving this because you were mentioned.\n> Reply to this email directly, view it on GitHub <https://github.com/materialsproject/pymatgen/issues/1914#issuecomment-664713216>, or unsubscribe <https://github.com/notifications/unsubscribe-auth/AKN477FDIDHYT2XQKEHJ47TR5YOQNANCNFSM4PJFZ4FA>.\n> \n\n",
"You can install enumlib following the instructions here: https://github.com/msg-byu/enumlib\r\n\r\nYou put it into your PATH meaning that it's available when you type a command in a terminal; the specific instructions to do this vary from Linux, Mac, Windows etc.\r\n\r\nWill close this issue since it's not a pymatgen bug but good luck!"
] | 2020-07-27T19:45:48
| 2020-07-29T18:11:04
|
2020-07-29T18:11:04Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Error when modules from pymatgen.analysis.magnetism.analyzer are used.
**To Reproduce**
Steps to reproduce the behavior:
1. execute the following python script
!/usr/bin/env python3
-*- coding: utf-8 -*-
from pymatgen.core import Lattice, Structure
from pymatgen.analysis.magnetism.analyzer import *
latt = Lattice.cubic(4.17)
species = ["Ni", "O"]
coords = [[0.00000, 0.00000, 0.00000], [0.50000, 0.50000, 0.50000]]
NiO = Structure.from_spacegroup(225, latt, species, coords)
print(NiO)
#enumerator = CollinearMagneticStructureAnalyzer(NiO)
enumerator = MagneticStructureEnumerator(NiO,strategies='ferromagnetic')
2. Then it seems the structure is generated but magnetic analyzing module dose not work
Full Formula (Ni4 O4)
Reduced Formula: NiO
abc : 4.170000 4.170000 4.170000
angles: 90.000000 90.000000 90.000000
Sites (8)
SP a b c
0 Ni 0 0 0
1 Ni 0 0.5 0.5
2 Ni 0.5 0 0.5
3 Ni 0.5 0.5 0
4 O 0.5 0.5 0.5
5 O 0.5 0 0
6 O 0 0.5 0
7 O 0 0 0.5
Illegal instruction (core dumped)
*Expected behavior**
A clear and concise description of what you expected to happen.
**Screenshots**

**Desktop (please complete the following information):**
- The code is ran on the cluster in Argonne National Lab bebop.
- Linux 3.10.0-957.21.3.el7.x86_64 x86_64
**Additional context**
Add any other context about the problem here.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1914/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1914/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1915
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1915/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1915/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1915/events
|
https://github.com/materialsproject/pymatgen/issues/1915
| 668,621,802
|
MDU6SXNzdWU2Njg2MjE4MDI=
| 1,915
|
How to generate vacancy structure and calculate formation energy
|
{
"login": "thienbinh92",
"id": 56110825,
"node_id": "MDQ6VXNlcjU2MTEwODI1",
"avatar_url": "https://avatars.githubusercontent.com/u/56110825?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/thienbinh92",
"html_url": "https://github.com/thienbinh92",
"followers_url": "https://api.github.com/users/thienbinh92/followers",
"following_url": "https://api.github.com/users/thienbinh92/following{/other_user}",
"gists_url": "https://api.github.com/users/thienbinh92/gists{/gist_id}",
"starred_url": "https://api.github.com/users/thienbinh92/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/thienbinh92/subscriptions",
"organizations_url": "https://api.github.com/users/thienbinh92/orgs",
"repos_url": "https://api.github.com/users/thienbinh92/repos",
"events_url": "https://api.github.com/users/thienbinh92/events{/privacy}",
"received_events_url": "https://api.github.com/users/thienbinh92/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @thienbinh92, this issues page is for reporting bugs, if you have a question could you ask it over at our user forum on https://matsci.org/pymatgen ? thanks!"
] | 2020-07-30T11:17:19
| 2020-07-30T16:47:28
|
2020-07-30T16:47:28Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hello, I read this paper "Vacancy Ordering in O3-Type Layered Metal Oxide Sodium-Ion Battery Cathodes", Toumar described that pymatgen was used to generate potential structures and the formation energy vs Na concentration was plotted in figure 3. If I am right, the pymatgen's tutorial does not describe how to generate potential structures. Can anyone show me step by step how to generate vacancy structures using pymatgen? Please take O3 layered oxide as an example.
Thank you so much!
https://escholarship.org/content/qt9648d7wm/qt9648d7wm.pdf
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1915/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1915/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1916
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1916/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1916/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1916/events
|
https://github.com/materialsproject/pymatgen/pull/1916
| 669,040,321
|
MDExOlB1bGxSZXF1ZXN0NDU5NDYwNzc4
| 1,916
|
Remove CPP and use only C in get neighbors optmization
|
{
"login": "chc273",
"id": 22353204,
"node_id": "MDQ6VXNlcjIyMzUzMjA0",
"avatar_url": "https://avatars.githubusercontent.com/u/22353204?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/chc273",
"html_url": "https://github.com/chc273",
"followers_url": "https://api.github.com/users/chc273/followers",
"following_url": "https://api.github.com/users/chc273/following{/other_user}",
"gists_url": "https://api.github.com/users/chc273/gists{/gist_id}",
"starred_url": "https://api.github.com/users/chc273/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/chc273/subscriptions",
"organizations_url": "https://api.github.com/users/chc273/orgs",
"repos_url": "https://api.github.com/users/chc273/repos",
"events_url": "https://api.github.com/users/chc273/events{/privacy}",
"received_events_url": "https://api.github.com/users/chc273/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Many thanks for addressing this @chc273.\r\n\r\nFor documentary purposes, I'm just noting we also noticed this issue on conda installs via Binder (which runs Ubuntu), and previously we also had some issues with Linux wheels via PyPI on #1868 that I think might be related.",
"> Many thanks for addressing this @chc273.\r\n> \r\n> For documentary purposes, I'm just noting we also noticed this issue on conda installs via Binder (which runs Ubuntu), and previously we also had some issues with Linux wheels via PyPI on #1868 that I think might be related.\r\n\r\nAh, I missed that issue. Thanks for bringing that to my attention... I hope this fix will do the job, otherwise, it could take much more time, since it is not very obvious where the problem is and the problem seems to be very dependent on the installation methods..",
"I'm pretty confident that a switch to pure-C will fix this.\r\n\r\nIn that other issue, we were trying a new method of building PyPI wheels, so I'd assumed there was an issue with the build process rather than an issue in the code itself. However, since we're seeing it on conda too, it's clear there must just be a compatibility issue across Linux distributions with the current code."
] | 2020-07-30T18:04:59
| 2020-07-30T19:15:01
|
2020-07-30T19:15:01Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Recently, we have seen some platform-dependent errors caused by this cython extension.
The known error is "illegal instructions (core dumped)" which cause crash of python. This error was found on conda installations. In particular, I have encountered issues on `ubuntu 18.04.4 LTS` (Matt raised this issue to me), and the github action VM `macOS-latest, python3.7`. Such errors however do not show up if pymatgen is installed using `pip` or from source. Interestingly, it also does not show up on github action VM `macOS-latest, python3.8`.
This PR is an attempt to fix the issue. My suspicion is that this is related to C++ compilation since it requires the platform-dependent flags. In this PR, I have removed all CPP-dependent codes and replace them with C only codes.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1916/reactions",
"total_count": 1,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 1,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1916/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1916",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1916",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1916.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1916.patch",
"merged_at": "2020-07-30T19:15:01Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1917
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1917/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1917/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1917/events
|
https://github.com/materialsproject/pymatgen/pull/1917
| 670,100,988
|
MDExOlB1bGxSZXF1ZXN0NDYwMzk0NjU0
| 1,917
|
Scan InputSet revisions
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @rkingsbury ",
"Tests pls ",
"@mkhorton Do you know where to find which pseudopotentials are currently recommended by VASP? There is [this](https://cms.mpi.univie.ac.at/vasp/vasp/Recommended_PAW_potentials_DFT_calculations_using_vasp_5_2.html) old page which is no longer maintained, but the current VASP wiki does not have any such page.",
"No idea, the most recent one I can find is that link -- the basic principles for the recommendations will be pretty much the same between _52 vs _54, with the exception of tungsten (W_pv) which was removed. You can see a discussion of the changes here: https://www.vasp.at/post/highlight-release-paw-datasets-v054/ Also take a look at the discussion of the GW potcars.",
"> No idea, the most recent one I can find is that link -- the basic principles for the recommendations will be pretty much the same between _52 vs _54, with the exception of tungsten (W_pv) which was removed. You can see a discussion of the changes here: https://www.vasp.at/post/highlight-release-paw-datasets-v054/ Also take a look at the discussion of the GW potcars.\r\n\r\nFor reference, here is how our current settings compare to the recommended PSPs (bolded) at the link you shared. All commented lines either differ from VASP recommendations or were just missing from our .yaml file before.\r\n\r\n```\r\n Ac: Ac\r\n Ag: Ag\r\n Al: Al\r\n Am: Am # was not present in the .yaml before\r\n Ar: Ar\r\n As: As\r\n At: At_d_GW # At_d removed in the PBE_54 set\r\n Au: Au\r\n B: B\r\n Ba: Ba_sv\r\n Be: Be_sv # VASP recommends Be\r\n Bi: Bi # VASP recommends Bi_d\r\n Br: Br\r\n C: C\r\n Ca: Ca_sv\r\n Cd: Cd\r\n Ce: Ce\r\n Cl: Cl\r\n Cm: Cm # was not present in the .yaml before\r\n Co: Co\r\n Cr: Cr_pv\r\n Cs: Cs_sv\r\n Cu: Cu_pv # VASP recommends Cu\r\n Dy: Dy_3\r\n Er: Er_3\r\n Eu: Eu # VASP recommends Eu_2\r\n F: F\r\n Fe: Fe_pv # VASP recommends Fe\r\n Fr: Fr_sv # was not present in the .yaml before\r\n Ga: Ga_d\r\n Gd: Gd # VASP recommends Gd_3\r\n Ge: Ge_d\r\n H: H\r\n He: He\r\n Hf: Hf_pv\r\n Hg: Hg\r\n Ho: Ho_3\r\n I: I\r\n In: In_d\r\n Ir: Ir\r\n K: K_sv\r\n Kr: Kr\r\n La: La\r\n Li: Li_sv\r\n Lu: Lu_3\r\n Mg: Mg_pv # VASP recommends Mg\r\n Mn: Mn_pv\r\n Mo: Mo_pv # VASP recommends Mo_sv\r\n N: N\r\n Na: Na_pv\r\n Nb: Nb_pv # VASP recommends Nb_sv\r\n Nd: Nd_3\r\n Ne: Ne\r\n Ni: Ni_pv # VASP recommends Ni\r\n Np: Np\r\n O: O\r\n Os: Os_pv # VASP recommends Os\r\n P: P\r\n Pa: Pa\r\n Pb: Pb_d\r\n Pd: Pd\r\n Pm: Pm_3\r\n Po: Po_d # VASP recommends Po_d Why was this not here before?\r\n Pr: Pr_3\r\n Pt: Pt\r\n Pu: Pu\r\n Ra: Ra_sv # was not present in the .yaml before\r\n Rb: Rb_sv\r\n Re: Re_pv # VASP recommends Re\r\n Rh: Rh_pv\r\n Rn: Rn # was not present in the .yaml before\r\n Ru: Ru_pv\r\n S: S\r\n Sb: Sb\r\n Sc: Sc_sv\r\n Se: Se\r\n Si: Si\r\n Sm: Sm_3\r\n Sn: Sn_d\r\n Sr: Sr_sv\r\n Ta: Ta_pv\r\n Tb: Tb_3\r\n Tc: Tc_pv\r\n Te: Te\r\n Th: Th\r\n Ti: Ti_pv # VASP recommends Ti_sv\r\n Tl: Tl_d\r\n Tm: Tm_3\r\n U: U\r\n V: V_pv # VASP recommends V_sv\r\n W: W_sv\r\n Xe: Xe\r\n Y: Y_sv\r\n Yb: Yb_2\r\n Zn: Zn\r\n Zr: Zr_sv\r\n```",
"> No idea, the most recent one I can find is that link -- the basic principles for the recommendations will be pretty much the same between _52 vs _54, with the exception of tungsten (W_pv) which was removed. You can see a discussion of the changes here: https://www.vasp.at/post/highlight-release-paw-datasets-v054/ Also take a look at the discussion of the GW potcars.\r\n\r\nWhere can I find the \"discussion of the GW potcars\"? I've looked in the VASP manual and read https://wiki.materialsproject.org/Pseudopotentials_Choice but don't see that discussion anywhere.",
"See: https://cms.mpi.univie.ac.at/vasp/vasp/Recommended_GW_PAW_potentials_vasp_5_2.html\r\n",
"> See: https://cms.mpi.univie.ac.at/vasp/vasp/Recommended_GW_PAW_potentials_vasp_5_2.html\r\n\r\nThanks, the GW potentials do sound promising, and I don't believe they were available when MP did the testing to determine the current POTCAR choices. Let's discuss more offline.",
"Sure, happy to discuss -- I've never used the GW POTCARs myself for conventional DFT, but worth testing. As far as I know they're essentially equivalent for ground-state properties. The only reason to prefer them is simply that I think they've seen more active recent development, so might have issues fixed that have not been fixed in the regular POTCARs.",
"Thanks @rkingsbury !"
] | 2020-07-31T18:38:39
| 2020-09-07T15:05:27
|
2020-08-13T20:18:40Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Revise `MPScanRelaxSet` per recent discussions
* LWAVE = False
* LREAL = Auto
* ENAUG = 2 * ENCUT
* remove ICHARG
* Add `MPScanStaticSet` in which we set LREAL=FALSE
## TODO
* Address question about MAGMOM in `MPStaticSet`
* Review POTCAR choices against VASP recommendations
* Update the hash for `MPScanRelaxSet.yaml` so that `SetChangeCheckTest` passes
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1917/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1917/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1917",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1917",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1917.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1917.patch",
"merged_at": "2020-08-13T20:18:40Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1918
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1918/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1918/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1918/events
|
https://github.com/materialsproject/pymatgen/pull/1918
| 672,374,105
|
MDExOlB1bGxSZXF1ZXN0NDYyNDIyNjQw
| 1,918
|
gaussian std orientation bug fix
|
{
"login": "eimrek",
"id": 8721264,
"node_id": "MDQ6VXNlcjg3MjEyNjQ=",
"avatar_url": "https://avatars.githubusercontent.com/u/8721264?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/eimrek",
"html_url": "https://github.com/eimrek",
"followers_url": "https://api.github.com/users/eimrek/followers",
"following_url": "https://api.github.com/users/eimrek/following{/other_user}",
"gists_url": "https://api.github.com/users/eimrek/gists{/gist_id}",
"starred_url": "https://api.github.com/users/eimrek/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/eimrek/subscriptions",
"organizations_url": "https://api.github.com/users/eimrek/orgs",
"repos_url": "https://api.github.com/users/eimrek/repos",
"events_url": "https://api.github.com/users/eimrek/events{/privacy}",
"received_events_url": "https://api.github.com/users/eimrek/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thank you. Can you pls add a unit test for the new functionality pls? Thanks.",
"hi @shyuep, there is no new functionality, just a small bug fix",
"The test would be to catch the bug you've encountered (e.g. with g.txt) so that future code modifications don't re-introduce the bug.",
"Yes, I understand that. but the fact is that the bug fix is fixing something that is not caught by any tests at the moment (otherwise, the tests would be failing since the bug exists). The idea is to write a simple test that demonstrates that the fix actually solves the bug,",
"Also, the new code seems to cause the existing tests to fail. So that needs to be fixed too.",
"Ah yep, I just realized this as well. The new convention (that standard orientation is preferred by default) is incompatible with one of the test, which compares against the \"input orientation\". Should I change that test as well? Other option would be to prefer the \"input orientation\" for the opt_structure but that would be weird as the standard orientation is already preferred for the `structures` attribute",
"Hi @shyuep, @mkhorton \r\n\r\nI added the test and modified an existing one. But now I'm failing `pybtex` import in some completely unrelated file: `pymatgen\\util\\provenance.py`. Any ideas why this could be?",
"Ignore that for now. I have merged it.",
"Thanks @eimrek for this bug fix."
] | 2020-08-03T22:14:02
| 2020-08-10T08:51:29
|
2020-08-04T15:06:18Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hi pymatgen community!
In gaussian log, there are geometries outputted either in "standard" or "input" orientation or both together. The current implementation assumed that "input orientation" is always outputted but that is not the case (try e.g. [g.txt](https://github.com/materialsproject/pymatgen/files/5019019/g.txt)) and I was getting an error.
I modified the code such that by default, if it's there, the `opt_structure` will be of "standard orientation" and otherwise "input orientation". This is reflected by the `standard_orientation` boolean attribute and the same convention was already used for the `structures` attribute.
Previous version was mainly modified by @gVallverdu, so ping to you in case you're curious.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1918/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1918/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1918",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1918",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1918.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1918.patch",
"merged_at": "2020-08-04T15:06:18Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1919
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1919/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1919/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1919/events
|
https://github.com/materialsproject/pymatgen/pull/1919
| 674,485,426
|
MDExOlB1bGxSZXF1ZXN0NDY0MTY5MzU0
| 1,919
|
Make clean=True default in Compatibility.process_entries()
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-08-06T17:32:56
| 2020-08-25T18:01:48
|
2020-08-07T17:23:06Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
The new Compatibility interface (see #1826) provides a boolean `clean` argument to `process_entries` that determines whether or not previously-applied corrections are discarded when `process_entries` is called. We set this to `False` by default, but this has proven to be a bad choice. Myself, @shyamd and @awvio have all made errors using the new `process_entries()` because repeated calls to `process_entries` resulted in corrections being applied multiple times (contrary to our expectations).
This PR changes the default to `True` so that corrections will only be applied one time by default, even if `process_entries` is called multiple times on the same entry.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1919/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1919/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1919",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1919",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1919.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1919.patch",
"merged_at": "2020-08-07T17:23:06Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1920
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1920/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1920/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1920/events
|
https://github.com/materialsproject/pymatgen/pull/1920
| 674,715,954
|
MDExOlB1bGxSZXF1ZXN0NDY0MzU3MzI4
| 1,920
|
Fix JMolNN tolerance
|
{
"login": "utf",
"id": 1330638,
"node_id": "MDQ6VXNlcjEzMzA2Mzg=",
"avatar_url": "https://avatars.githubusercontent.com/u/1330638?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/utf",
"html_url": "https://github.com/utf",
"followers_url": "https://api.github.com/users/utf/followers",
"following_url": "https://api.github.com/users/utf/following{/other_user}",
"gists_url": "https://api.github.com/users/utf/gists{/gist_id}",
"starred_url": "https://api.github.com/users/utf/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/utf/subscriptions",
"organizations_url": "https://api.github.com/users/utf/orgs",
"repos_url": "https://api.github.com/users/utf/repos",
"events_url": "https://api.github.com/users/utf/events{/privacy}",
"received_events_url": "https://api.github.com/users/utf/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"The actual change of this PR was changing a single value (0876de1) but to get linting to pass I had to make many changes in the local_env module.",
"Thanks for fixing everything."
] | 2020-08-07T02:43:55
| 2020-08-07T17:13:08
|
2020-08-07T17:12:59Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Changed the JMolNN tolerance parameter to match the that used by the JMol source code.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1920/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1920/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1920",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1920",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1920.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1920.patch",
"merged_at": "2020-08-07T17:12:59Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1921
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1921/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1921/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1921/events
|
https://github.com/materialsproject/pymatgen/pull/1921
| 676,495,566
|
MDExOlB1bGxSZXF1ZXN0NDY1Nzk3MDY2
| 1,921
|
Added GibbsComputedStructureEntry for predicting ΔGf(T)
|
{
"login": "mattmcdermott",
"id": 40469694,
"node_id": "MDQ6VXNlcjQwNDY5Njk0",
"avatar_url": "https://avatars.githubusercontent.com/u/40469694?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mattmcdermott",
"html_url": "https://github.com/mattmcdermott",
"followers_url": "https://api.github.com/users/mattmcdermott/followers",
"following_url": "https://api.github.com/users/mattmcdermott/following{/other_user}",
"gists_url": "https://api.github.com/users/mattmcdermott/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mattmcdermott/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mattmcdermott/subscriptions",
"organizations_url": "https://api.github.com/users/mattmcdermott/orgs",
"repos_url": "https://api.github.com/users/mattmcdermott/repos",
"events_url": "https://api.github.com/users/mattmcdermott/events{/privacy}",
"received_events_url": "https://api.github.com/users/mattmcdermott/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
] | null |
[
"Thanks @mattmcdermott! Is there any reason this functionality couldn't just be added to the existing `ComputedStructureEntry` rather than made its own class? In general I feel we have too many `Entry` classes already. Interested to hear other's thoughts on this.\r\n\r\nSeparately, I'm working on creating an MPContribs project for experimental thermochemical data that might be a logical home for the NIST gas data you've included. I can upload the data using the file here, but we'll have to discuss whether it makes sense to modify your code to talk to the MPContribs project now, or leave it as-is until we streamline that project more. (Talking to MPContribs requires another API key that not everyone has)",
"I've also been thinking that separate attributes for `enthalpy`, `entropy`, and `free_energy` in `ComputedEntry` might be a good idea to better clarify how some of our analysis works, and your new class takes a step in that direction.",
"Sub-classing is appropriate here, having many classes is not a problem in itself as long as the inheritance makes sense and it's well documented. This Gibbs model, though good, is still its own thing with its own set of caveats and shouldn't be added onto `ComputedStructureEntry` which is more general. Separately, making the distinction between energy/enthalpy might be a good conversation to have, we often interchange these terms inappropriately.\r\n\r\n@mattmcdermott once tests are added this looks about ready to merge, I am thinking a little about `from_pd` though and if this is the best place for it.",
"@rkingsbury I am always ambivalent about furthering an already long chain of subclasses, but I agree with @mkhorton 's reasoning that there is enough distinct functionality here to not integrate it into existing classes... at least not yet.\r\n\r\nI will also add that Chris's model we are using here can only predict this \"G_δ\" term, aka the difference between ΔHf (0K) and ΔGf (desired temp), so it can't quite be separated into enthalpy and entropy terms, although we know from his paper that H changes much less with temperature than the TS term does.\r\n\r\n@mkhorton I always thought that having a method which constructs a list of objects was a bit strange (not sure if this is common practice), but the class is basically useless to the average user without this method, since you have to make a 0 K phase diagram to get MP formation energies in the first place. Were you thinking about putting it into the phase diagram class?",
"@mattmcdermott Yeah, I actually think this is fine after thinking about it more. I was thinking PhaseDiagram, was also thinking Compatibility, but it's a small method and only there for convenience, and it doesn't preclude additional integration elsewhere later.",
"@mkhorton @rkingsbury Just added tests! I also added the `from_entries()` method (another convenient initialization) and cleaned up a problem with saving/loading dicts.",
"Thanks @mattmcdermott !"
] | 2020-08-11T00:29:34
| 2020-08-13T17:23:22
|
2020-08-13T17:23:13Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
- Added a new entry class called **GibbsComputedStructureEntry**. This is an extension to ComputedStructureEntry which
allows for the estimation of Gibbs free energy of formation, ΔGf(T), via a machine-learned model. This can be used to estimate temperature-dependent phase diagrams using MP data.
- Currently incorporates the Gibbs SISSO descriptor for solid compounds as created and described by [Bartel et al. (2018) ](https://www.nature.com/articles/s41467-018-06682-4). This class should be extendable to future models based on pymatgen structural properties if/when they are created.
- As explained in the paper, the calculation of ΔGf(T) requires the use of the temperature-dependent chemical potentials of the elements, which are derived from FactSage and supplied in the _g_els.json_ file.
- Since the SISSO model only applies to inorganic solid compounds, functionality has also been added to use NIST-JANAF data for gases where possible. This data is supplied in the _nist_gas_gf.json_ file.
- The `from_pd()` method constructs and returns a list of GibbsComputedStructureEntry objects given an existing MP phase diagram -- this is typically the easiest way to use the class.
## Checklist
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1921/reactions",
"total_count": 2,
"+1": 2,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1921/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1921",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1921",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1921.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1921.patch",
"merged_at": "2020-08-13T17:23:13Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1922
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1922/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1922/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1922/events
|
https://github.com/materialsproject/pymatgen/issues/1922
| 677,078,833
|
MDU6SXNzdWU2NzcwNzg4MzM=
| 1,922
|
looks like compatibility.process_entries() is not scrubbing previous energy_adjustments
|
{
"login": "computron",
"id": 986759,
"node_id": "MDQ6VXNlcjk4Njc1OQ==",
"avatar_url": "https://avatars.githubusercontent.com/u/986759?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/computron",
"html_url": "https://github.com/computron",
"followers_url": "https://api.github.com/users/computron/followers",
"following_url": "https://api.github.com/users/computron/following{/other_user}",
"gists_url": "https://api.github.com/users/computron/gists{/gist_id}",
"starred_url": "https://api.github.com/users/computron/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/computron/subscriptions",
"organizations_url": "https://api.github.com/users/computron/orgs",
"repos_url": "https://api.github.com/users/computron/repos",
"events_url": "https://api.github.com/users/computron/events{/privacy}",
"received_events_url": "https://api.github.com/users/computron/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Actually closing for now, this looks to be something to do with the way an entry had been serialized in a database and then deserialized. I can't reproduce using more straightforward code. Although I would suggest that the code above is probably more robust than what's there as it works for the example below as well as what I was working with. Feel free to close whenever desired.\r\n\r\nHere is the code that confirms that things are actually working as intended:\r\n\r\n```\r\nfrom pymatgen import MPRester\r\nfrom pymatgen.entries.compatibility import MaterialsProjectCompatibility\r\n\r\nmpr = MPRester()\r\nmpc = MaterialsProjectCompatibility()\r\n\r\nentry = mpr.get_entries(\"mp-19042\")[0]\r\nprint(len(entry.energy_adjustments), entry.correction)\r\n\r\nentry2 = mpc.process_entry(entry) # should clean the previous one\r\nprint(len(entry2.energy_adjustments), entry2.correction)\r\n\r\n# It seems to do so successfully.\r\n```",
"Not against simplifying the `clean` method regardless, I'm not sure what the current logic is. @rkingsbury, thoughts?",
"Just to make sure - @computron did you pull in the latest pymatgen in which I changed the default behavior of `clean`? We had it set to `False` which did indeed cause a lot of unexpected behavior. See https://github.com/materialsproject/pymatgen/pull/1919\r\n\r\nNo objection to simplifying the code a bit; I think the for loop might be a leftover from a previous iteration on this idea in which we were trying to make certain adjustments 'immune' to cleaning.\r\n\r\nI know that there can be some non-obvious behavior with entries that were serialized before we implemented energy adjustments. I tried to handle all of those cases (and have tests in pymatgen that should catch them) but if you find a reproducible case please reopen.",
"To follow up - I did have the most recent version of the code with the default being True.\r\n\r\nI think what happened in my case is as follows:\r\n- A ComputedEntry was exported to MongoDB\r\n- ComputedEntry.from_dict() was used to load it back from MongoDB\r\n- For some reason, the deserialization of energy adjustments probably did not occur in a great manner. Perhaps it got deserialized to some object that didn't have a great equals() method or something.\r\n\r\nSo then the ``clean()`` implementation, which calls list.remove(), was clearly running (I tested this) but failing to actually remove anything. Again, probably because list.remove() requires a proper equals() method to work properly.\r\n\r\nStyle-wise, I also find it problematic to be removing entries from a list while looping over those same entries:\r\n(current code)\r\n```\r\n if clean:\r\n for ea in entry.energy_adjustments:\r\n entry.energy_adjustments.remove(ea)\r\n```\r\n(proposed)\r\n```\r\n if clean:\r\n entry.energy_adjustments = []\r\n```"
] | 2020-08-11T17:58:38
| 2020-08-11T21:01:09
|
2020-08-11T18:21:27Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
pymatgen.entries.compatibility.Compatibility.process_entries() is supposed to be clearing out the previous energy adjustments on an entry (the "clean" parameter is by default True). Doesn't look like it's doing it.
**To Reproduce**
Steps to reproduce the behavior:
- Load a ComputedEntry with existing energy corrections
- Run MaterialsProjectCompatibility.process_entries() on that entry - which should in principle change nothing
- watch the energy adjustments change
**Can fix by**
Making the following patch ~line 474 of compatibility.py
```
if clean:
entry.energy_adjustments = []
```
rather than the weird convoluted code that tries to individually remove each adjustment (why?)
|
{
"login": "computron",
"id": 986759,
"node_id": "MDQ6VXNlcjk4Njc1OQ==",
"avatar_url": "https://avatars.githubusercontent.com/u/986759?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/computron",
"html_url": "https://github.com/computron",
"followers_url": "https://api.github.com/users/computron/followers",
"following_url": "https://api.github.com/users/computron/following{/other_user}",
"gists_url": "https://api.github.com/users/computron/gists{/gist_id}",
"starred_url": "https://api.github.com/users/computron/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/computron/subscriptions",
"organizations_url": "https://api.github.com/users/computron/orgs",
"repos_url": "https://api.github.com/users/computron/repos",
"events_url": "https://api.github.com/users/computron/events{/privacy}",
"received_events_url": "https://api.github.com/users/computron/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1922/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1922/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1923
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1923/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1923/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1923/events
|
https://github.com/materialsproject/pymatgen/issues/1923
| 677,181,200
|
MDU6SXNzdWU2NzcxODEyMDA=
| 1,923
|
BabelMolAdaptor doesn't work with IMolecule, only Molecule
|
{
"login": "smheidrich",
"id": 3827982,
"node_id": "MDQ6VXNlcjM4Mjc5ODI=",
"avatar_url": "https://avatars.githubusercontent.com/u/3827982?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/smheidrich",
"html_url": "https://github.com/smheidrich",
"followers_url": "https://api.github.com/users/smheidrich/followers",
"following_url": "https://api.github.com/users/smheidrich/following{/other_user}",
"gists_url": "https://api.github.com/users/smheidrich/gists{/gist_id}",
"starred_url": "https://api.github.com/users/smheidrich/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/smheidrich/subscriptions",
"organizations_url": "https://api.github.com/users/smheidrich/orgs",
"repos_url": "https://api.github.com/users/smheidrich/repos",
"events_url": "https://api.github.com/users/smheidrich/events{/privacy}",
"received_events_url": "https://api.github.com/users/smheidrich/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-08-11T20:37:53
| 2020-08-11T20:57:13
|
2020-08-11T20:57:13Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
The ``BabelMolAdaptor`` class fails when given ``IMolecule`` instead of ``Molecule`` objects.
**To Reproduce**
```python
from pymatgen import IMolecule
from pymatgen.io.babel import BabelMolAdaptor
m = IMolecule(["H","H"], [[1.,2.,3.],[4.,5.,6.]])
a = BabelMolAdaptor(m)
a.openbabel_mol
```
raises
```
Traceback (most recent call last):
File "babel_no_imolecule.py", line 6, in <module>
a.openbabel_mol
File "/home/smheidrich/pymatgen/pymatgen/io/babel.py", line 97, in openbabel_mol
return self._obmol
AttributeError: 'BabelMolAdaptor' object has no attribute '_obmol'
```
**Expected behavior**
Both classes should be supported.
**Additional context**
This is important because some pymatgen features like [MoleculeMatcher.group_molecules](https://pymatgen.org/pymatgen.analysis.molecule_matcher.html#pymatgen.analysis.molecule_matcher.MoleculeMatcher.group_molecules) perform the ``BabelMolAdaptor`` conversion internally but return data structures containing the originally supplied molecules, which the user might have specifically created as ``IMolecule`` instances to be able to use them as dictionary keys - requiring the user to convert them to ``Molecule`` objects beforehand would render any such dictionaries useless, so that's not an option.
Tested on current revision on master (93e518abfbae96e8d39d972a6b11ad7a9fb39cad).
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1923/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1923/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1924
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1924/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1924/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1924/events
|
https://github.com/materialsproject/pymatgen/pull/1924
| 677,185,839
|
MDExOlB1bGxSZXF1ZXN0NDY2MzQ2NDMw
| 1,924
|
Allow BabelMolAdaptor to convert IMolecules
|
{
"login": "smheidrich",
"id": 3827982,
"node_id": "MDQ6VXNlcjM4Mjc5ODI=",
"avatar_url": "https://avatars.githubusercontent.com/u/3827982?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/smheidrich",
"html_url": "https://github.com/smheidrich",
"followers_url": "https://api.github.com/users/smheidrich/followers",
"following_url": "https://api.github.com/users/smheidrich/following{/other_user}",
"gists_url": "https://api.github.com/users/smheidrich/gists{/gist_id}",
"starred_url": "https://api.github.com/users/smheidrich/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/smheidrich/subscriptions",
"organizations_url": "https://api.github.com/users/smheidrich/orgs",
"repos_url": "https://api.github.com/users/smheidrich/repos",
"events_url": "https://api.github.com/users/smheidrich/events{/privacy}",
"received_events_url": "https://api.github.com/users/smheidrich/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-08-11T20:46:03
| 2020-08-11T20:57:13
|
2020-08-11T20:57:13Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Instead of checking whether supplied objects are instances of ``Molecule``, ``BabelMolAdaptor`` now checks whether they are ``IMolecule``s instead (which is a superclass of ``Molecule`` so the old check still works). This fixes #1923.
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [ ] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1924/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1924/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1924",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1924",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1924.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1924.patch",
"merged_at": "2020-08-11T20:57:13Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1925
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1925/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1925/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1925/events
|
https://github.com/materialsproject/pymatgen/pull/1925
| 677,202,883
|
MDExOlB1bGxSZXF1ZXN0NDY2MzU5NjQw
| 1,925
|
simplify cleaning of EnergyAdjustment
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-08-11T21:16:23
| 2020-08-11T21:20:18
|
2020-08-11T21:20:18Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Replaces the code that removes previous `EnergyAdjustment` from `ComputedEntry` with a more robust alternative. See #1922
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1925/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1925/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1925",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1925",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1925.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1925.patch",
"merged_at": "2020-08-11T21:20:18Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1926
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1926/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1926/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1926/events
|
https://github.com/materialsproject/pymatgen/pull/1926
| 678,013,490
|
MDExOlB1bGxSZXF1ZXN0NDY3MDM2ODQ5
| 1,926
|
fix valence bug, add test
|
{
"login": "rkurchin",
"id": 11233777,
"node_id": "MDQ6VXNlcjExMjMzNzc3",
"avatar_url": "https://avatars.githubusercontent.com/u/11233777?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkurchin",
"html_url": "https://github.com/rkurchin",
"followers_url": "https://api.github.com/users/rkurchin/followers",
"following_url": "https://api.github.com/users/rkurchin/following{/other_user}",
"gists_url": "https://api.github.com/users/rkurchin/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkurchin/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkurchin/subscriptions",
"organizations_url": "https://api.github.com/users/rkurchin/orgs",
"repos_url": "https://api.github.com/users/rkurchin/repos",
"events_url": "https://api.github.com/users/rkurchin/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkurchin/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Many thanks @rkurchin !",
"On the test note, I agree this is unlikely to fail but I'll let the tests pass on CI first before merging. Your issue might be related to using `nose` which is now deprecated, `pytest` should be sufficient to run tests.\r\n\r\nWe're also asking new contributors to [fill out this form](https://forms.gle/JnisFb38QDR8QTFTA) so that everyone can be acknowledged appropriately in the [pymatgen documentation](https://pymatgen.org/team.html).",
"Got it – might want to update the instructions [here](https://pymatgen.org/contributing) then, that's why I used `nose`, even though I saw on its docs page that it was deprecated.\r\n\r\nFor my own stuff (since I need this to work properly for something I'm doing), I imagine once this is merged in I can expect it in the version I get from `pip install` within a couple days? In the meantime I can just use my local version obviously.\r\n",
"Thanks for the catch! Yes that page is definitely out of date, I've pushed some minor edits to the documentation for now but we're over-due for a more thorough documentation update.\r\n\r\nWe don't currently have a fixed schedule for pymatgen releases, we usually make one or two per month as pull requests come in or on an as-needed basis. I'll see if I can get this released by the end of the week.",
"FYI, this was released as part of 2020.8.13"
] | 2020-08-12T22:20:01
| 2020-08-14T22:51:39
|
2020-08-12T23:56:36Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Previously, asking for the valence property on an Element with full shells that wasn't noble (e.g. column 2 or 12) would cause an error. This resolves that.
## Summary
Include a summary of major changes in bullet points:
* Added a check for if the last orbital listed is fully-occupied and to choose that number as the valency
* Added a case to the tests to trigger this
## Additional dependencies introduced (if any)
none
## Checklist
Before a pull request can be merged, the following items must be checked:
- [ x ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [ x ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ x ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [ x ] Tests have been added for any new functionality or bug fixes.
- [ ? ] All existing tests pass. --> I ran the test_periodic_table.py manually, but nosetests continually hangs after 1089 tests. I installed all dependencies (including optional and dev ones) in my environment so I'm not sure what the issue is with that, but I can't imagine what this would break that's not covered by the test I checked.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1926/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1926/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1926",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1926",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1926.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1926.patch",
"merged_at": "2020-08-12T23:56:35Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1927
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1927/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1927/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1927/events
|
https://github.com/materialsproject/pymatgen/pull/1927
| 678,229,020
|
MDExOlB1bGxSZXF1ZXN0NDY3MjExNTk4
| 1,927
|
Updates to LobsterSet
|
{
"login": "JaGeo",
"id": 22094846,
"node_id": "MDQ6VXNlcjIyMDk0ODQ2",
"avatar_url": "https://avatars.githubusercontent.com/u/22094846?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/JaGeo",
"html_url": "https://github.com/JaGeo",
"followers_url": "https://api.github.com/users/JaGeo/followers",
"following_url": "https://api.github.com/users/JaGeo/following{/other_user}",
"gists_url": "https://api.github.com/users/JaGeo/gists{/gist_id}",
"starred_url": "https://api.github.com/users/JaGeo/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/JaGeo/subscriptions",
"organizations_url": "https://api.github.com/users/JaGeo/orgs",
"repos_url": "https://api.github.com/users/JaGeo/repos",
"events_url": "https://api.github.com/users/JaGeo/events{/privacy}",
"received_events_url": "https://api.github.com/users/JaGeo/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I will close this one here and open up a new one. The automatic testing asks me to update the linting of a lot of code.... "
] | 2020-08-13T07:47:46
| 2020-08-13T08:31:42
|
2020-08-13T08:25:23Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
I have just updated some values in the LobsterSet. Let me know if I should include more tests.
Best,
JG
|
{
"login": "JaGeo",
"id": 22094846,
"node_id": "MDQ6VXNlcjIyMDk0ODQ2",
"avatar_url": "https://avatars.githubusercontent.com/u/22094846?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/JaGeo",
"html_url": "https://github.com/JaGeo",
"followers_url": "https://api.github.com/users/JaGeo/followers",
"following_url": "https://api.github.com/users/JaGeo/following{/other_user}",
"gists_url": "https://api.github.com/users/JaGeo/gists{/gist_id}",
"starred_url": "https://api.github.com/users/JaGeo/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/JaGeo/subscriptions",
"organizations_url": "https://api.github.com/users/JaGeo/orgs",
"repos_url": "https://api.github.com/users/JaGeo/repos",
"events_url": "https://api.github.com/users/JaGeo/events{/privacy}",
"received_events_url": "https://api.github.com/users/JaGeo/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1927/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1927/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1927",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1927",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1927.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1927.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1928
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1928/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1928/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1928/events
|
https://github.com/materialsproject/pymatgen/pull/1928
| 678,257,246
|
MDExOlB1bGxSZXF1ZXN0NDY3MjM0ODQ5
| 1,928
|
Updates to LobsterSet
|
{
"login": "JaGeo",
"id": 22094846,
"node_id": "MDQ6VXNlcjIyMDk0ODQ2",
"avatar_url": "https://avatars.githubusercontent.com/u/22094846?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/JaGeo",
"html_url": "https://github.com/JaGeo",
"followers_url": "https://api.github.com/users/JaGeo/followers",
"following_url": "https://api.github.com/users/JaGeo/following{/other_user}",
"gists_url": "https://api.github.com/users/JaGeo/gists{/gist_id}",
"starred_url": "https://api.github.com/users/JaGeo/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/JaGeo/subscriptions",
"organizations_url": "https://api.github.com/users/JaGeo/orgs",
"repos_url": "https://api.github.com/users/JaGeo/repos",
"events_url": "https://api.github.com/users/JaGeo/events{/privacy}",
"received_events_url": "https://api.github.com/users/JaGeo/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Sorry about that, adding an additional rule to ignore in the linter, and accidentally over-rode the default ignore list thereby triggering a lot of errors.\r\n\r\nIt should be fixed now if you merge in the latest master.",
"No worries - should have looked at the latest testss before starting with this pull request.\r\n\r\nI hope all tests run through now.\r\n\r\nBest,\r\nJG",
"Thanks @JaGeo !"
] | 2020-08-13T08:33:24
| 2020-08-13T20:19:07
|
2020-08-13T20:19:03Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
I have just updated some values in the LobsterSet. Let me know if I should include more tests.
There seem to be problems with the linting in the current main branch. I don't really want to fix all of them - there seem to be hundreds.
I will fix every failing test that seems to be related to my changes.
Best,
JG
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1928/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1928/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1928",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1928",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1928.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1928.patch",
"merged_at": "2020-08-13T20:19:03Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1929
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1929/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1929/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1929/events
|
https://github.com/materialsproject/pymatgen/issues/1929
| 679,126,086
|
MDU6SXNzdWU2NzkxMjYwODY=
| 1,929
|
SpacegroupAnalyzer.get_conventional_standard_structure returns a wrong structure
|
{
"login": "lan496",
"id": 7272505,
"node_id": "MDQ6VXNlcjcyNzI1MDU=",
"avatar_url": "https://avatars.githubusercontent.com/u/7272505?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/lan496",
"html_url": "https://github.com/lan496",
"followers_url": "https://api.github.com/users/lan496/followers",
"following_url": "https://api.github.com/users/lan496/following{/other_user}",
"gists_url": "https://api.github.com/users/lan496/gists{/gist_id}",
"starred_url": "https://api.github.com/users/lan496/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/lan496/subscriptions",
"organizations_url": "https://api.github.com/users/lan496/orgs",
"repos_url": "https://api.github.com/users/lan496/repos",
"events_url": "https://api.github.com/users/lan496/events{/privacy}",
"received_events_url": "https://api.github.com/users/lan496/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Agreed this looks like a serious bug. I'm looking at these lines in particular:\r\n\r\nhttps://github.com/materialsproject/pymatgen/blob/v2020.8.13/pymatgen/symmetry/analyzer.py#L547-L597\r\n\r\nI didn't write this code so it'll take me a while to understand what's meant to be happening here.",
"Interestingly, I'm not sure I can reproduce this on pymatgen 2020.7.14, still investigating ...",
"@mkhorton Check if any of the cleanup that I did caused this. But if it did, seriously the person who wrote the code in the first place should have implemented unittests.\r\n",
"@shyuep I don't think it was those changes, I was able to reproduce on 2020.7.14. Agreed on unit tests, we can use this one + generate some more. Structures don't even `match` in this case! I can imagine a fairly extensive unit test for a set of structures + checking they match before and after getting them in their conventional setting.\r\n\r\n@lan496, haven't got a fix yet, but thank you for such a nice reproducible example ... helps a lot",
"It seems odd that in the `conventional_refined = sga.get_conventional_standard_structure()` all the z co-ordinates have been zeroed out. The transform matrix `transf` is:\r\n\r\n```\r\n[[1. 0. 0.]\r\n [0. 1. 0.]\r\n [0. 0. 0.]]\r\n```\r\n\r\nSo regardless of what the original z fractional co-ordinate is, it would always become zero.\r\n\r\nThis gets set to `1` explicitly here:\r\n\r\nhttps://github.com/materialsproject/pymatgen/blob/185cb592cc0710e6882b3f57177112e3d86414d2/pymatgen/symmetry/analyzer.py#L549\r\n\r\nbut then gets set back to `0` here:\r\n\r\nhttps://github.com/materialsproject/pymatgen/blob/185cb592cc0710e6882b3f57177112e3d86414d2/pymatgen/symmetry/analyzer.py#L595",
"Resetting it back to `transf[2] = [0, 0, 1]` gives the correct result and fixes the issue here, but without code comments or a clear understanding of what the original code is doing, I can't say if this is an appropriate fix.",
"It seems the bug in `get_conventional_standard_structure` for monoclinic also relates to https://github.com/materialsproject/pymatgen/issues/1543. \r\n`SpacegroupAnalyzer` with default symprec judges a structure in this issue as C2/m.\r\n",
"Just adding a note here for test coverage: https://coveralls.io/builds/27639791/source?filename=pymatgen%2Fsymmetry%2Fanalyzer.py#L465\r\n\r\nThere are a few important branches not covered by existing tests.",
"I committed the fix here: https://github.com/materialsproject/pymatgen/commit/f9226b79b8f9ccd0b305e557c9e4878e096bfc77 but I'm not satisfied this issue is closed until we can add some additional tests to make sure the entire function is covered."
] | 2020-08-14T12:31:49
| 2023-08-13T16:33:43
|
2023-08-13T16:33:43Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**To Reproduce**
If I understand correctly, a structure and a symmetry-refined one have the same minimum distance between sites.
So, three structures `structure`, `conventional_refined`, and `spglib_refined` in the following script should have the same minimum distance.
```python
import numpy as np
from pymatgen.core import Structure
from pymatgen.analysis.structure_analyzer import SpacegroupAnalyzer
def min_dist(structure):
all_dists = structure.distance_matrix[np.triu_indices(structure.num_sites, 1)]
return np.min(all_dists)
if __name__ == '__main__':
# spacegroup: C2/m
matrix = [[3.945, 0.0, 3.945], [0.0, 3.945, 3.945], [0.0, 0.0, 11.834999999999999]]
species = ['O'] * 7
frac_coords = np.array([
[0., 0.5, 0.66666667],
[0.5, 0., 0.],
[0.5, 0., 0.33333333],
[0.5, 0., 0.66666667],
[0.5, 0.5, -0.33333333],
[0.5, 0.5, 0.],
[0.5, 0.5, 0.33333333],
])
structure = Structure(matrix, species, frac_coords)
print("original:", min_dist(structure))
sga = SpacegroupAnalyzer(structure)
conventional_refined = sga.get_conventional_standard_structure()
print("conventional_standard:", min_dist(conventional_refined))
spglib_refined = sga.get_refined_structure()
print("refine_cell:", min_dist(spglib_refined))
conventional_refined.to(fmt='cif', filename='conventional_standard.cif', symprec=1e-2)
spglib_refined.to(fmt='cif', filename='refined.cif', symprec=1e-2)
```
But I got the following output. The structure created by `SpacegroupAnalyzer.get_conventional_standard_structure` has a different distance from that of the original structure.
```
original: 2.789536223885567
conventional_standard: 1.610539489774544
refine_cell: 2.7895361959902023
```
**Screenshots**
The structure created by `SpacegroupAnalyzer.get_conventional_standard_structure` is as follows:

The structure created by `SpacegroupAnalyzer.get_refined_structure` is as follows:

**Desktop (please complete the following information):**
- OS: Ubuntu16.04
- Version: 2020.8.13
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1929/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1929/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1930
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1930/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1930/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1930/events
|
https://github.com/materialsproject/pymatgen/pull/1930
| 679,155,700
|
MDExOlB1bGxSZXF1ZXN0NDY3OTgzMTg0
| 1,930
|
Fix type of reciprocal lattice in KPathSeek
|
{
"login": "lan496",
"id": 7272505,
"node_id": "MDQ6VXNlcjcyNzI1MDU=",
"avatar_url": "https://avatars.githubusercontent.com/u/7272505?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/lan496",
"html_url": "https://github.com/lan496",
"followers_url": "https://api.github.com/users/lan496/followers",
"following_url": "https://api.github.com/users/lan496/following{/other_user}",
"gists_url": "https://api.github.com/users/lan496/gists{/gist_id}",
"starred_url": "https://api.github.com/users/lan496/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/lan496/subscriptions",
"organizations_url": "https://api.github.com/users/lan496/orgs",
"repos_url": "https://api.github.com/users/lan496/repos",
"events_url": "https://api.github.com/users/lan496/events{/privacy}",
"received_events_url": "https://api.github.com/users/lan496/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @Ian496!"
] | 2020-08-14T13:25:52
| 2020-08-14T18:19:01
|
2020-08-14T18:19:01Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
The abstract class of KPathSeek, KPathBase, expects a type of a reciprocal lattice as `pymatgen.core.lattice.Lattice`. This tiny PR fixes to cast a reciprocal lattice returned by `seekpath` into pymatgen's lattice class.
## Checklist
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [ ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [ ] All existing tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1930/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1930/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1930",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1930",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1930.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1930.patch",
"merged_at": "2020-08-14T18:19:01Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1931
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1931/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1931/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1931/events
|
https://github.com/materialsproject/pymatgen/issues/1931
| 679,571,406
|
MDU6SXNzdWU2Nzk1NzE0MDY=
| 1,931
|
Problems with Vasprun parsing dielectric vasprun.xml file
|
{
"login": "pjf295",
"id": 69720428,
"node_id": "MDQ6VXNlcjY5NzIwNDI4",
"avatar_url": "https://avatars.githubusercontent.com/u/69720428?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/pjf295",
"html_url": "https://github.com/pjf295",
"followers_url": "https://api.github.com/users/pjf295/followers",
"following_url": "https://api.github.com/users/pjf295/following{/other_user}",
"gists_url": "https://api.github.com/users/pjf295/gists{/gist_id}",
"starred_url": "https://api.github.com/users/pjf295/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/pjf295/subscriptions",
"organizations_url": "https://api.github.com/users/pjf295/orgs",
"repos_url": "https://api.github.com/users/pjf295/repos",
"events_url": "https://api.github.com/users/pjf295/events{/privacy}",
"received_events_url": "https://api.github.com/users/pjf295/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-08-15T13:13:13
| 2023-08-13T16:33:44
|
2023-08-13T16:33:44Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I am trying to use pymatgen to read in the local field (RPA level) frequency dependent dielectric function.
I am using the input files from the SiC example on the vaspwiki page. The structure is SiC
```
<POSCAR>
system SiC
4.35
0.5 0.5 0.0
0.0 0.5 0.5
0.5 0.0 0.5
1 1
cart
0.00 0.00 0.00
0.25 0.25 0.25
```
Initially I calculate the dielectric function in the independent particle approximation with a 8x8x8 Gamma centered grid.
The WAVECAR file from a self-consistent run was used as input.
The INCAR file was as follows.
```
<INCAR>
ALGO = Exact
NBANDS = 64
LOPTICS = .TRUE. ; CSHIFT = 0.100
NEDOS = 2000
LPEAD = .TRUE.
ISMEAR = 0
SIGMA = 0.01
EDIFF = 1.E-8
```
I can read in the frequency dependent dielectric function in pymatgen without error.
Using the WAVECAR and WAVEDIR files from the run above, I then calculated the frequency dependent dielectric
function including the effects of local fields within the RPA approximation.
```
<INCAR>
Frequency dependent dielectric tensor with
# local field effects in RPA
# N.B.: beware one first has to have done a
# calculation with ALGO=Exact, LOPTICS=.TRUE.
# and a reasonable number of virtual states (see above)
ALGO = CHI
# be sure to take the same number of bands as for
# the LOPTICS=.TRUE. calculation, otherwise the
# WAVEDER file is not read correctly
NBANDS = 64
ISMEAR = 0
SIGMA = 0.01
EDIFF = 1.E-8
LWAVE = .FALSE.
LCHARG= .FALSE.
```
Vasp completes normally and I can see the dielectric constant in the OUTCAR file. Attempting to use Vasprun on the
vasprun.xml file, however, results in the following error.
```
IndexError Traceback (most recent call last)
<ipython-input-1-6cd881495936> in <module>
----> 1 run = Vasprun('vasprun.xml')
~/opt/anaconda3/lib/python3.7/site-packages/pymatgen/io/vasp/outputs.py in __init__(self, filename, ionic_step_skip, ionic_step_offset, parse_dos, parse_eigen, parse_projected_eigen, parse_potcar_file, occu_tol, exception_on_bad_xml)
353 self.update_charge_from_potcar(parse_potcar_file)
354
--> 355 if self.incar.get("ALGO", "") != "BSE" and (not self.converged):
356 msg = "%s is an unconverged VASP run.\n" % filename
357 msg += "Electronic convergence reached: %s.\n" % \
~/opt/anaconda3/lib/python3.7/site-packages/pymatgen/io/vasp/outputs.py in converged(self)
582 electronically.
583 """
--> 584 return self.converged_electronic and self.converged_ionic
585
586 @property # type: ignore
~/opt/anaconda3/lib/python3.7/site-packages/pymatgen/io/vasp/outputs.py in converged_electronic(self)
556 ionic step
557 """
--> 558 final_esteps = self.ionic_steps[-1]["electronic_steps"]
559 if 'LEPSILON' in self.incar and self.incar['LEPSILON']:
560 i = 1
IndexError: list index out of range
```
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1931/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1931/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1932
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1932/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1932/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1932/events
|
https://github.com/materialsproject/pymatgen/pull/1932
| 680,523,799
|
MDExOlB1bGxSZXF1ZXN0NDY5MDY2MDY5
| 1,932
|
fix saving _acc_factor instead of _accf
|
{
"login": "lbluque",
"id": 9788715,
"node_id": "MDQ6VXNlcjk3ODg3MTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/9788715?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/lbluque",
"html_url": "https://github.com/lbluque",
"followers_url": "https://api.github.com/users/lbluque/followers",
"following_url": "https://api.github.com/users/lbluque/following{/other_user}",
"gists_url": "https://api.github.com/users/lbluque/gists{/gist_id}",
"starred_url": "https://api.github.com/users/lbluque/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/lbluque/subscriptions",
"organizations_url": "https://api.github.com/users/lbluque/orgs",
"repos_url": "https://api.github.com/users/lbluque/repos",
"events_url": "https://api.github.com/users/lbluque/events{/privacy}",
"received_events_url": "https://api.github.com/users/lbluque/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-08-17T21:02:21
| 2020-08-17T21:22:56
|
2020-08-17T21:22:56Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Fix `as_dict` in EwaldSummation to save `_acc_factor` and not `_accf`, to recreate the exact object as before. Previous tests did not catch this since the actual Ewald summation values remained within numerical tolerance (my bad for not catching this before).
## Checklist
Before a pull request can be merged, the following items must be checked:
- [X] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [X] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [X] Tests have been added for any new functionality or bug fixes.
- [ ] All existing tests pass. pylint is failing, not sure why.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1932/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1932/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1932",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1932",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1932.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1932.patch",
"merged_at": "2020-08-17T21:22:56Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1933
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1933/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1933/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1933/events
|
https://github.com/materialsproject/pymatgen/pull/1933
| 680,589,086
|
MDExOlB1bGxSZXF1ZXN0NDY5MTIwODgw
| 1,933
|
MP2020 Compatibility refinements
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"FYI @awvio. When you are ready to update the .json.gz, the easiest way would be to open a pull request against [my branch](https://github.com/rkingsbury/pymatgen/tree/mp2020-refactor), which I can then merge so that it shows up here. Thanks!",
"@mkhorton @shyamd @shyuep I've reformatted the .yaml file to consolidate uncertainties and corrections into one place, and also to make the keys more descriptive and consistent with the way we describe the corrections in the forthcoming manuscript. Please let me know if you have any feedback.",
"Ready for review per recent discussions @mkhorton. Let me know any further suggestions. Thanks!"
] | 2020-08-17T23:42:20
| 2020-09-15T18:59:18
|
2020-09-15T18:36:40Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
* Refactor `MaterialsProject2020Compatibility` to utilize the new abstract compatibility interface (see #1826 ) instead of the legacy `CorrectionsList` class. Note that the `PotcarCorrection` class is still utilized, but not any other `Correction` classes.
* `MaterialsProject2020Compatibility()` now exposes all keyword arguments that were previously passed to `Correction` classes
* Consolidate .yaml files into a single file and change the keys to more closely match the way each correction type is described in the forthcoming manuscript. Update `make_yaml` to generate this format.
* Refit energy corrections against the `v2020.08.21` database release, using experimental ground-state structures instead of DFT ground states for selected compounds where the DFT ground state is not in the ICSD
* update contents of `calc_compounds.json.gz` to include `ComputedStructureEntry` instead of `ComputedEntry`
* Update docstrings
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1933/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1933/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1933",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1933",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1933.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1933.patch",
"merged_at": "2020-09-15T18:36:40Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1934
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1934/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1934/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1934/events
|
https://github.com/materialsproject/pymatgen/issues/1934
| 680,634,811
|
MDU6SXNzdWU2ODA2MzQ4MTE=
| 1,934
|
Something wrong with the result of DosPlotter
|
{
"login": "Shan19961030",
"id": 69703663,
"node_id": "MDQ6VXNlcjY5NzAzNjYz",
"avatar_url": "https://avatars.githubusercontent.com/u/69703663?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Shan19961030",
"html_url": "https://github.com/Shan19961030",
"followers_url": "https://api.github.com/users/Shan19961030/followers",
"following_url": "https://api.github.com/users/Shan19961030/following{/other_user}",
"gists_url": "https://api.github.com/users/Shan19961030/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Shan19961030/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Shan19961030/subscriptions",
"organizations_url": "https://api.github.com/users/Shan19961030/orgs",
"repos_url": "https://api.github.com/users/Shan19961030/repos",
"events_url": "https://api.github.com/users/Shan19961030/events{/privacy}",
"received_events_url": "https://api.github.com/users/Shan19961030/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I think the problem comes from stack=True. I had a similar result when trying to fill the a DOS plotted horizontally, while was ok if plotted vertically (as by BSDOSPlotter). Please, try with stack=False.\r\nI edited the code to solve the bug but I forgot to push it. I'll do it ASAP.",
"I have tried to solve this problems with stack = False,but the result also seems strange,here is the result\r\n[image](http://m.qpic.cn/psc?/V11CjYqJ0aw8nE/bqQfVz5yrrGYSXMvKr.cqYKcrh4XcC2c7vMugZy9f3ZtSwJy5k2VezXdpGQQ3PIHu1*7g5bKnT5u.SCNKsvgHhQI9qmDgM8gDgs8isQLBiI!/b&bo=sAQgAwAAAAADB7U!&rf=viewer_4) ",
"> I think the problem comes from stack=True. I had a similar result when trying to fill the a DOS plotted horizontally, while was ok if plotted vertically (as by BSDOSPlotter). Please, try with stack=False.\r\n> I edited the code to solve the bug but I forgot to push it. I'll do it ASAP.\r\n\r\nI have tried to solve this problems with stack = False,but the result also seems strange,here is the result\r\n[image](http://m.qpic.cn/psc?/V11CjYqJ0aw8nE/bqQfVz5yrrGYSXMvKr.cqYKcrh4XcC2c7vMugZy9f3ZtSwJy5k2VezXdpGQQ3PIHu1*7g5bKnT5u.SCNKsvgHhQI9qmDgM8gDgs8isQLBiI!/b&bo=sAQgAwAAAAADB7U!&rf=viewer_4)",
"Can you supply the vasprun.xml so that we can debug it?",
"> Can you supply the vasprun.xml so that we can debug it?\r\n\r\nThank you so much for your reply.Here is my vasprun.xml、POSCAR and POTCAR file. If you need more files, please feel free to contact me anytime and I will provide you with them soon.\r\n\r\n[files](https://github.com/Shan19961030/VaspCalculationFile)\r\n\r\nIf you debug it or find the reasons,I hope you can notify me.\r\n\r\nSincere thanks!",
"I need the KPOINTS file for the band structure. But I see what you did wrong. You used a HUGE sigma value. That is the value that is used to smear the DOS plot. A smearing of 0.5 eV is way too big. Try setting it to 0 or some small number. The following code works:\r\n\r\n```python\r\nimport matplotlib.pyplot as plt\r\nfrom pymatgen.io.vasp.outputs import Vasprun\r\nfrom pymatgen.electronic_structure.plotter import BSDOSPlotter, BSPlotter, BSPlotterProjected, DosPlotter\r\ndos_vasprun = Vasprun(\"vasprun.xml\")\r\ndos_data = dos_vasprun.complete_dos\r\nplt1 = DosPlotter(stack=False, sigma=0)\r\nplt1.add_dos('total dos', dos=dos_data)\r\nplt = plt1.get_plot()\r\nplt.xlim(((-4, 6)))\r\nplt.ylim((0, 1000))\r\n```\r\n\r\nNote that you have some strange states at low energies. So I set the xlims and ylims to a reasonable value. Stack does not seem to work for this case right now. This is because this is a no spin DOS I believe. ",
"> I need the KPOINTS file for the band structure. But I see what you did wrong. You used a HUGE sigma value. That is the value that is used to smear the DOS plot. A smearing of 0.5 eV is way too big. Try setting it to 0 or some small number. The following code works:\r\n> \r\n> ```python\r\n> import matplotlib.pyplot as plt\r\n> from pymatgen.io.vasp.outputs import Vasprun\r\n> from pymatgen.electronic_structure.plotter import BSDOSPlotter, BSPlotter, BSPlotterProjected, DosPlotter\r\n> dos_vasprun = Vasprun(\"vasprun.xml\")\r\n> dos_data = dos_vasprun.complete_dos\r\n> plt1 = DosPlotter(stack=False, sigma=0)\r\n> plt1.add_dos('total dos', dos=dos_data)\r\n> plt = plt1.get_plot()\r\n> plt.xlim(((-4, 6)))\r\n> plt.ylim((0, 1000))\r\n> ```\r\n> \r\n> Note that you have some strange states at low energies. So I set the xlims and ylims to a reasonable value. Stack does not seem to work for this case right now. This is because this is a no spin DOS I believe.\r\n\r\nThank you so much ,I will try it with you code again."
] | 2020-08-18T02:11:47
| 2020-08-20T01:49:31
|
2020-08-20T01:45:16Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
when I use the BSDosPlotter model to draw BAND and DOS figure,I got a perfect result comparing to the one drawn by P4V and Origin.But,when I use the same data to draw single DOS picture with DosPlotter ,I got a very strange result .
**To Reproduce**
Follows are my script of DosPlotter:
import matplotlib.pyplot as plt
from pymatgen.io.vasp.outputs import Vasprun
from pymatgen.electronic_structure.plotter import BSDOSPlotter, BSPlotter, BSPlotterProjected, DosPlotter
dos_vasprun = Vasprun("./dos/vasprun.xml")
dos_data = dos_vasprun.complete_dos
bs_vasprun = Vasprun("./band/vasprun.xml", parse_projected_eigen=True)
bs_data = bs_vasprun.get_band_structure(line_mode=True)
plt1 = DosPlotter(stack=True, sigma=0.5)
plt1.add_dos('total dos', dos=dos_data)
plt1.save_plot('DOS.png', img_format=u'png')
**Screenshots**
This is the result of BSDosPlotter
[image](http://m.qpic.cn/psc?/V11CjYqJ0aw8nE/bqQfVz5yrrGYSXMvKr.cqQyJain0wopA8.W9M02nUSSc47hSbSvgwoC*0l6UpJqNaPVGdlAFDgMr3if5O2ECX97msz5kz9M5WBM4KykTlkE!/b&bo=TARSAwAAAAADBzs!&rf=viewer_4)
This is the result of DosPlotter
[image](http://m.qpic.cn/psc?/V11CjYqJ0aw8nE/TmEUgtj9EK6.7V8ajmQrEHrPeDaGjBTlR*k.D5XsIvPoWd31h6XhVEVTwdXkil5.KVPt.nR8HIXuHbv3miCqDcy6xkqY2DhIsyyKEd36z*8!/b&bo=sAQgAwAAAAADF6U!&rf=viewer_4)
**Desktop (please complete the following information):**
- OS: [Windows]
-python Version [3.8]
**Additional context**
Is it a problem of the version of Python?
Thank you for your attention!
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1934/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1934/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1935
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1935/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1935/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1935/events
|
https://github.com/materialsproject/pymatgen/issues/1935
| 680,920,725
|
MDU6SXNzdWU2ODA5MjA3MjU=
| 1,935
|
MoleculeMatcher is failing too easily due to small perturbations
|
{
"login": "fekad",
"id": 6748541,
"node_id": "MDQ6VXNlcjY3NDg1NDE=",
"avatar_url": "https://avatars.githubusercontent.com/u/6748541?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/fekad",
"html_url": "https://github.com/fekad",
"followers_url": "https://api.github.com/users/fekad/followers",
"following_url": "https://api.github.com/users/fekad/following{/other_user}",
"gists_url": "https://api.github.com/users/fekad/gists{/gist_id}",
"starred_url": "https://api.github.com/users/fekad/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/fekad/subscriptions",
"organizations_url": "https://api.github.com/users/fekad/orgs",
"repos_url": "https://api.github.com/users/fekad/repos",
"events_url": "https://api.github.com/users/fekad/events{/privacy}",
"received_events_url": "https://api.github.com/users/fekad/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Not sure if @samblau or @espottesmith want to comment here? (or want to recommend someone who could)"
] | 2020-08-18T10:43:10
| 2023-08-13T16:33:44
|
2023-08-13T16:33:44Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
The current version of MoleculeMatcher does not seem to be robust enough for practical applications.
**To Reproduce**
You can find a simplified code snippet below:
```python
import random
from pymatgen import Lattice, Structure, Molecule
from pymatgen.analysis.molecule_matcher import MoleculeMatcher, IsomorphismMolAtomMapper, InchiMolAtomMapper
coords = [[0, 0, 0], [0.75,0.5,0.75]]
lattice = Lattice.from_parameters(a=3.84, b=3.84, c=3.84, alpha=120, beta=90, gamma=60)
struct = Structure(lattice, ["Si", "Si"], coords)
struct.make_supercell([2,2,2])
# Creating molecule for testing
struct = Molecule.from_sites(struct)
```
Using IsomorphismMolAtomMapper:
```python
N = 10
mm = MoleculeMatcher(tolerance=2., mapper=IsomorphismMolAtomMapper())
# perturbing the atoms' position
for i in range(N):
struct2 = struct.copy()
struct_perturbed = struct.copy()
struct_perturbed.perturb(0.3)
print(mm.get_rmsd(struct2, struct_perturbed))
# perturbing the atoms' order
for i in range(N):
struct2 = struct.copy()
struct_perturbed = struct.copy()
random.shuffle(struct_perturbed)
print(mm.get_rmsd(struct2, struct_perturbed))
```
The output:
```
inf
inf
inf
inf
0.29595839352301634
inf
inf
inf
0.2830535234793642
inf
1.606558315893547e-15
1.606558315893547e-15
1.606558315893547e-15
1.606558315893547e-15
1.606558315893547e-15
1.606558315893547e-15
1.606558315893547e-15
1.606558315893547e-15
1.606558315893547e-15
1.606558315893547e-15
```
Using InchiMolAtomMapper:
```python
N = 10
mm = MoleculeMatcher(tolerance=2., mapper=InchiMolAtomMapper())
# perturbing the atoms' position
for i in range(N):
struct2 = struct.copy()
struct_perturbed = struct.copy()
struct_perturbed.perturb(0.3)
print(mm.get_rmsd(struct2, struct_perturbed))
# perturbing the atoms' order
for i in range(N):
struct2 = struct.copy()
struct_perturbed = struct.copy()
random.shuffle(struct_perturbed)
print(mm.get_rmsd(struct2, struct_perturbed))
```
The output:
```
0.2924931714214089
0.26981844192962556
inf
0.28688401574184996
inf
inf
inf
inf
inf
inf
3.9161327362484313
3.457316914376839
3.5354691763693302
3.9678366662615496
2.944084494876412
3.977348259145049
3.575265287043952
3.5128593012595
3.6150131865387447
3.9656370958396563
```
**Expected behavior**
For perturbing the positions of the atoms by `0.3` you would expect `0.3` rmsd for all the cases.
As far as I can see in the case of `InchiMolAtomMapper` tries to find the good ordering but it is not capable to do that.
**Additional context**
For me, it seems to be the `uniform_labels` method of the mappers which returning with a wrong order (I mean when returns with something at all). Please feel free to add the following line to the loops above to get more details:
```python
print(mm._mapper.uniform_labels(struct2, struct_perturbed))
```
**Desktop (please complete the following information):**
- OS: Mac
- Version: 2020.8.13
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1935/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1935/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1936
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1936/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1936/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1936/events
|
https://github.com/materialsproject/pymatgen/pull/1936
| 681,380,664
|
MDExOlB1bGxSZXF1ZXN0NDY5Nzc4ODcz
| 1,936
|
Overhaul of PhaseDiagram plotting: adding Plotly backend
|
{
"login": "mattmcdermott",
"id": 40469694,
"node_id": "MDQ6VXNlcjQwNDY5Njk0",
"avatar_url": "https://avatars.githubusercontent.com/u/40469694?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mattmcdermott",
"html_url": "https://github.com/mattmcdermott",
"followers_url": "https://api.github.com/users/mattmcdermott/followers",
"following_url": "https://api.github.com/users/mattmcdermott/following{/other_user}",
"gists_url": "https://api.github.com/users/mattmcdermott/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mattmcdermott/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mattmcdermott/subscriptions",
"organizations_url": "https://api.github.com/users/mattmcdermott/orgs",
"repos_url": "https://api.github.com/users/mattmcdermott/repos",
"events_url": "https://api.github.com/users/mattmcdermott/events{/privacy}",
"received_events_url": "https://api.github.com/users/mattmcdermott/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Looks fantastic, very exciting. Will need tests of course but otherwise will seems well structured.",
"Thanks @mattmcdermott ! This is going to be a really nice enhancement. This also includes the uncertainties support (for 2D plots at least), correct?",
"> Thanks @mattmcdermott ! This is going to be a really nice enhancement. This also includes the uncertainties support (for 2D plots at least), correct?\r\n\r\n@rkingsbury Yep, actually I am finishing this up at this exact moment. There will be uncertainty error bars for all plots, plus uncertainty shading for binaries.",
"Ok @mkhorton @rkingsbury -- should be pretty much ready to go! It would be nice if you could test it out on your side to make sure things work (I am sure there are more things layout wise we can play around with as well)",
"Thanks @mattmcdermott! This is great. If there are any additional improvements I think they can be put into a separate PR, since this looks feature complete as-is."
] | 2020-08-18T22:30:23
| 2020-09-11T01:14:38
|
2020-09-11T01:13:46Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
- Added `backend="plotly"` as a default argument to `PDPlotter`. Option `backend="matplotlib"` returns to existing plotting functionality.
- New features includes 2D scatter/line plot for binaries, and 3D scatter/mesh plots for ternaries and quaternaries. The latter include convex hull shading via Plotly's mesh plotting capabilities.
- Benefits of a Plotly backend: **highly increased user interactivity** -- ability to see and rotate around hull in three dimensions, hide traces, see hover annotations for phase info, etc.
- Plotting seems to be pretty fast: ternaries take about ~ 100 ms on my computer, especially with added caching of `pd_plot_data`
### EDITS:
- Improved color scheme with added colorbars
- Default layouts now in json file.
- Added uncertainty error bars for ComputedEntry's new uncertainty property.
- Added uncertainty window shading along the hull for binaries.
- Separated stable labels into a separate trace (to allow hiding).
- Added some functionality for smarter placement of labels (should make this even better in the future).
- Improved some camera angle / rotation capability (should also make this even better in the future).
## Examples (apologies for low resolution):
### Binaries:
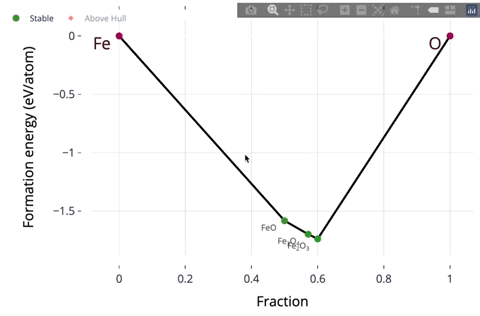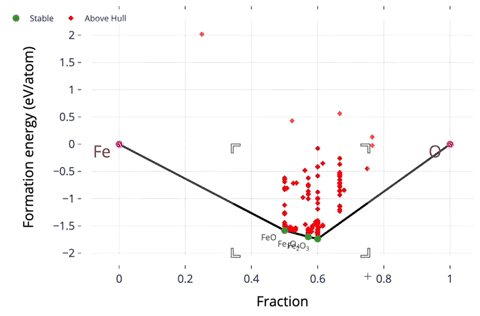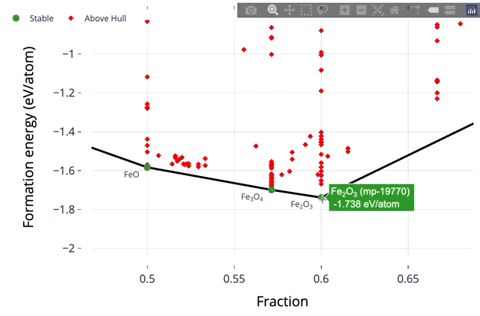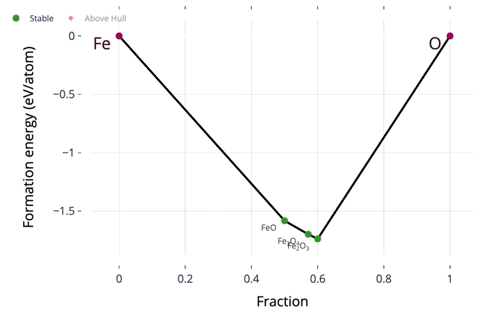
### Ternaries:
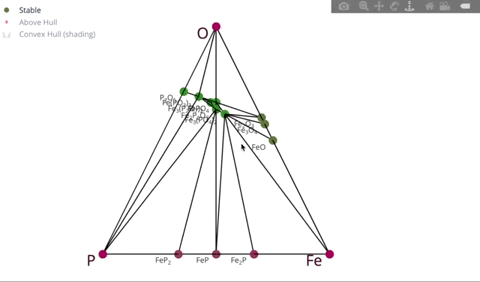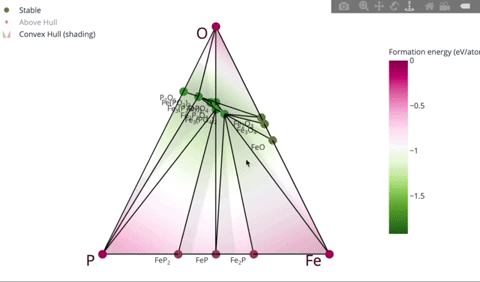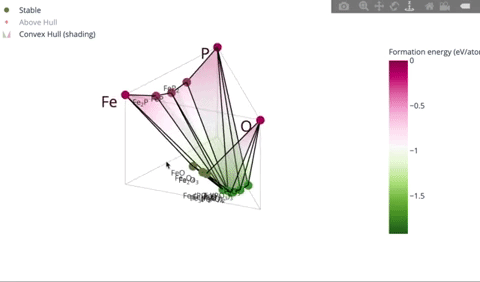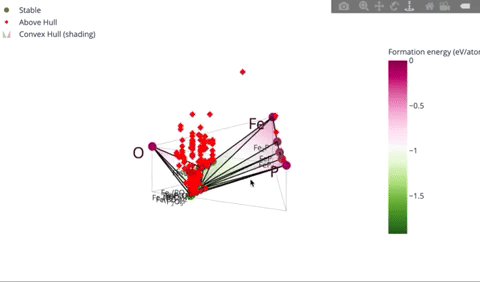
### Quaternaries:
## TODO (if any)
- Would be nice to add Plotly backend for chemical potential range map plots (i.e. predominance diagrams) **EDIT**: This has been done, but will be added in a future PR.
- Would be nice to add rotation support for ternaries; no option for choosing which element is plotted on top
- Phase diagram annotations are not always sensibly placed -- I can't seem to find a good fix for this, other than hiding annotations in general or writing some pretty arbitrary geometrical shifting functions **EDIT**: A simple first pass of this has been implemented, and will likely be edited in the future.
- A couple minor bugs that I've noticed but had trouble fixing: 1) Color of "Stable" trace in legend is not always green (as I would like) which also becomes an annoyance when working with ternary/quaternary hovertext, and 2) Shading of convex hull in quaternaries is obscured by the faces of tetrahedron which are also colored **EDIT**: Both problems fixed. Deleted shading of quaternary hull (too strange).
## Checklist
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
## Projects
This addresses Project #1288.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1936/reactions",
"total_count": 2,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 1,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1936/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1936",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1936",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1936.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1936.patch",
"merged_at": "2020-09-11T01:13:46Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1937
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1937/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1937/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1937/events
|
https://github.com/materialsproject/pymatgen/pull/1937
| 683,154,971
|
MDExOlB1bGxSZXF1ZXN0NDcxMjkwNjk4
| 1,937
|
bugfix in surface and composition
|
{
"login": "gpetretto",
"id": 8665734,
"node_id": "MDQ6VXNlcjg2NjU3MzQ=",
"avatar_url": "https://avatars.githubusercontent.com/u/8665734?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/gpetretto",
"html_url": "https://github.com/gpetretto",
"followers_url": "https://api.github.com/users/gpetretto/followers",
"following_url": "https://api.github.com/users/gpetretto/following{/other_user}",
"gists_url": "https://api.github.com/users/gpetretto/gists{/gist_id}",
"starred_url": "https://api.github.com/users/gpetretto/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/gpetretto/subscriptions",
"organizations_url": "https://api.github.com/users/gpetretto/orgs",
"repos_url": "https://api.github.com/users/gpetretto/repos",
"events_url": "https://api.github.com/users/gpetretto/events{/privacy}",
"received_events_url": "https://api.github.com/users/gpetretto/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Can you implement some tests to catch the bugs fixed pls? Thanks.",
"I implemented the tests. I also slightly changed the docstrings.",
"Thanks."
] | 2020-08-20T23:28:39
| 2020-08-21T15:10:12
|
2020-08-21T15:10:05Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Two bug fixes:
* with the current implementation the Surface object can get weird results when instantiating with `reorient_lattice=True `and `coords_are_cartesian=True`. Fixed by switching to frac coords when needed.
* after dec6ecb6a9a80dbd4bfdfb924040cdfc7fe4ebb3 `Composition.to_reduced_dict` is not returning a dict but a Composition instance.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1937/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1937/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1937",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1937",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1937.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1937.patch",
"merged_at": "2020-08-21T15:10:05Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1938
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1938/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1938/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1938/events
|
https://github.com/materialsproject/pymatgen/pull/1938
| 683,756,962
|
MDExOlB1bGxSZXF1ZXN0NDcxNzk3ODEz
| 1,938
|
[WIP] Implemenation of a robust permutation invariant molecule matcher
|
{
"login": "fekad",
"id": 6748541,
"node_id": "MDQ6VXNlcjY3NDg1NDE=",
"avatar_url": "https://avatars.githubusercontent.com/u/6748541?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/fekad",
"html_url": "https://github.com/fekad",
"followers_url": "https://api.github.com/users/fekad/followers",
"following_url": "https://api.github.com/users/fekad/following{/other_user}",
"gists_url": "https://api.github.com/users/fekad/gists{/gist_id}",
"starred_url": "https://api.github.com/users/fekad/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/fekad/subscriptions",
"organizations_url": "https://api.github.com/users/fekad/orgs",
"repos_url": "https://api.github.com/users/fekad/repos",
"events_url": "https://api.github.com/users/fekad/events{/privacy}",
"received_events_url": "https://api.github.com/users/fekad/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"The results look perfect when I run the same test that was mentioned in issue #1935:\r\n```python\r\nN = 10\r\nmm = PermInvMatcher()\r\n\r\n# perturbing the atoms' position\r\nfor i in range(N):\r\n struct2 = struct.copy()\r\n struct_perturbed = struct.copy()\r\n struct_perturbed.perturb(0.3)\r\n _,_,_,rmsd = mm.match(struct2, struct_perturbed)\r\n print(rmsd)\r\n \r\n\r\n# perturbing the atoms' order\r\nfor i in range(N):\r\n struct2 = struct.copy()\r\n struct_perturbed = struct.copy()\r\n random.shuffle(struct_perturbed)\r\n\r\n _,_,_,rmsd = mm.match(struct2, struct_perturbed)\r\n\r\n print(rmsd)\r\n```\r\noutput:\r\n```\r\n0.1558403374164826\r\n0.15653898357207896\r\n0.16509417765476064\r\n0.15996366581570087\r\n0.15717228359651142\r\n0.160348617463949\r\n0.16101400312730157\r\n0.15850843779432255\r\n0.16499070127970572\r\n0.16268112989375508\r\n7.008175309304189e-16\r\n6.811558627955245e-16\r\n9.037610622348878e-16\r\n7.72009337997319e-16\r\n7.380861825310775e-16\r\n3.5444346724928873e-16\r\n1.032864319121941e-15\r\n2.0355271010290208e-16\r\n7.356775432634027e-16\r\n8.756485695103038e-16\r\n```",
"@mkhorton I'm a quite new user of `pymatgen`. As far as I can see all of the instances can be represented as json (the class is a subclass of MSONable). In the previous implementation, it totally makes sense so you can store/restore the parameters. \r\nthere was a threshold and when you used 2 input structure it returned with a true or false value...\r\n\r\nIn this new implementation, you can fit a structure to a target structure and the function returns with rotation matrix, translation vector rmse etc. I would say it is the user's task to decide what to do with this information. Eg. based on the rmse value he/she can decide was it a good fit or not. \r\n\r\nFollowing this concept, I would use the target structure as an argument of the init function so it would be serialised as a parameter of an instance. Personally I have no problem with it but is there any disadvantage to using a nested structure?\r\n\r\nSo I would use an interface like:\r\n```python\r\nclass AbstractMatcher(MSONable):\r\n\r\n def __init__(self, target: Molecule):\r\n pass\r\n\r\n def match(m: Molecule) -> Tuple[np.array, np.array, float]:\r\n # return rotation, translation, rmse\r\n pass\r\n\r\n def fit(m:Molecule) -> Tuple[Molecule, float]:\r\n # return rotated/translated molecule, rmse\r\n pass\r\n```\r\n",
"Hi @fekad, welcome and many thanks for tackling this issue!\r\n\r\nTo clarify your question, why specifically are you proposing the following:\r\n\r\n```\r\nclass AbstractMatcher(MSONable):\r\n\r\n def __init__(self, target: Molecule):\r\n pass\r\n\r\n def match(m: Molecule) -> Tuple[np.array, np.array, float]:\r\n # return rotation, translation, rmse\r\n pass\r\n```\r\n\r\nvs.\r\n\r\n```\r\nclass AbstractMatcher(MSONable):\r\n\r\n def __init__(self, ...):\r\n pass\r\n\r\n def match(a: Molecule, b: Molecule) -> Tuple[np.array, np.array, float]:\r\n # return rotation, translation, rmse\r\n pass\r\n```\r\n\r\nBoth seem equivalent to me, but the latter seems a bit easier to work with, since you only have to instantiate AbstractMatcher once even if you want to work with multiple molecules.\r\n\r\nIn terms of your other point:\r\n\r\n> there was a threshold and when you used 2 input structure it returned with a true or false value...\r\n> In this new implementation, you can fit a structure to a target structure and the function returns with rotation matrix, translation vector rmse etc. I would say it is the user's task to decide what to do with this information.\r\n\r\nIt might be helpful to take a look at the `StructureMatcher` class for ideas here. I agree that the returning the RMSE is important, and the user can decide based on that, but having a default behavior that returns True/False (possibly via a tolerance parameter set in the init) is also useful. These can simply be two separate methods, e.g. a `match_rmse` method returns the RMSE, and a `match` method uses `match_rmse` but returns a boolean based on the tolerance.",
"@mkhorton Do any of you have experience / best practice to use random numbers for testing? I was trying to use seed method and also creating a random generator object with a specific seed value but it doesn't seem to work...\r\n\r\nObviously, I can just use some predetermined random numbers for testing, but in theory, using random generators with seed method should give the same results and you can easily do more test if you want ...\r\n",
"@fekad I haven't had to do this previously, but using `seed` should work to create a deterministic random number generator, both via the in-built `random` library and `np.random`. I see you're initializing an rng object in some of your kwargs, perhaps that's responsible?\r\n\r\nAs an FYI, we don't currently use this in pymatgen but [hypothesis](https://hypothesis.readthedocs.io/en/latest/) is a nice library for creating unit tests that incorporates random data (following a specific requirement) as the test input, and asserts something about the output.",
"@mkhorton The initialization of the rng object was not an issue because every time I explicitly constructed one and passed that one to the function.\r\n\r\nI also tried to use the - old-fashion - seed method but the result is the same...\r\nI use multiple macs without any issue but It looks like the seed methods are not cross-platform yet.\r\n\r\nIn conclusion, I will use pre-generated structures for the tests unless somebody has a better solution.\r\n",
"Hi @fekad, my apologies for the delay in reviewing this, I will try to review this week. From your end, do you consider this PR ready to merge?\r\n\r\nLooking at the test failures, I'm seeing a minor linter failure, but also some perhaps more serious test failures:\r\n\r\n```\r\nFAILED pymatgen/analysis/tests/test_molecule_matcher.py::KabschMatcherSiTest::test_perturbed_atom_position\r\n129\r\nFAILED pymatgen/analysis/tests/test_molecule_matcher.py::HungarianOrderMatcherSiTest::test_perturbed_atom_position\r\n130\r\nFAILED pymatgen/analysis/tests/test_molecule_matcher.py::GeneticOrderMatcherSiTest::test_all\r\n131\r\nFAILED pymatgen/analysis/tests/test_molecule_matcher.py::GeneticOrderMatcherSiTest::test_perturbed_atom_position\r\n```\r\n\r\nThese are all of the form:\r\n\r\n```\r\nE AssertionError: 0.2680448948923893 != 0.2628450748567651 within 8 places\r\n77\r\n```\r\n\r\nIt's possible these tests just need to be updated for the new method.",
"@mkhorton No worries. I didn't fix the tests yet but I will do in the next days.",
"@mkhorton From my side, this PR is ready for review. Please let me know if something is not understandable or if you have better names for the classes :) ",
"This is a really substantial contribution and nicely tested, thank you! Class names look good to me. Please make sure to include your details in [this form](https://forms.gle/JnisFb38QDR8QTFTA) so that you can be credited appropriately in the [pymatgen documentation](https://pymatgen.org/team.html) too."
] | 2020-08-21T18:44:04
| 2020-10-05T12:28:39
|
2020-10-03T03:03:15Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
The main goal to implement a robust matching method between molecules. This implementation is based on [Kabsch algorithm]( https://en.m.wikipedia.org/wiki/Kabsch_algorithm) and on an excellent python package called [rmsd](https://github.com/charnley/rmsd). The code has been basically rewritten from scratch and adapted to pymatgen. Also, some small bugs in the referred solution have been fixed.
There are three new methods available:
* `KabschMatcher`: this methods works when the order of the atoms is the same in the two structures. Returns with the optimal rotation and translation which belong to the minimal distance between the molecules.
* `BruteForceOrderMatcher`: This method compares all the possible permutations and it also returns with the optimal order of atoms. Works perfectly with small molecules but not feasible larger systems due to it's factorial complexity.
* `HungarianOrderMatcher`: This method tries to identify the atom pairs by aligning the molecules based on their principle axis and sorted by the (Hungarian algorithm)[https://en.m.wikipedia.org/wiki/Hungarian_algorithm]. Please note that it is not guaranteed that this method always returns with the "best" solution.
* (NEW) `GeneticOrderMatcher`: This method was inspired by genetic algorithms and tries to match molecules based on their already matched fragments.
Key features:
* A new method to match `p` onto `q` molecule
* Returns with the rotation matrix, translation vector, proper order of atoms (in case of permutation invariant method) and root-mean-square deviation.
* The implementation is pure python and it is completely independent of `openbabel`.
* closing: #1935
## TODO/Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x] Creating a common interface for the `match` and `fit` methods
- [x] Refactoring the code to have a more object-oriented code
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1938/reactions",
"total_count": 1,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 1,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1938/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1938",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1938",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1938.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1938.patch",
"merged_at": "2020-10-03T03:03:15Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1939
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1939/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1939/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1939/events
|
https://github.com/materialsproject/pymatgen/pull/1939
| 683,855,830
|
MDExOlB1bGxSZXF1ZXN0NDcxODc4MzYy
| 1,939
|
[WIP] Jsmol viewer for jupyter notebook
|
{
"login": "fekad",
"id": 6748541,
"node_id": "MDQ6VXNlcjY3NDg1NDE=",
"avatar_url": "https://avatars.githubusercontent.com/u/6748541?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/fekad",
"html_url": "https://github.com/fekad",
"followers_url": "https://api.github.com/users/fekad/followers",
"following_url": "https://api.github.com/users/fekad/following{/other_user}",
"gists_url": "https://api.github.com/users/fekad/gists{/gist_id}",
"starred_url": "https://api.github.com/users/fekad/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/fekad/subscriptions",
"organizations_url": "https://api.github.com/users/fekad/orgs",
"repos_url": "https://api.github.com/users/fekad/repos",
"events_url": "https://api.github.com/users/fekad/events{/privacy}",
"received_events_url": "https://api.github.com/users/fekad/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"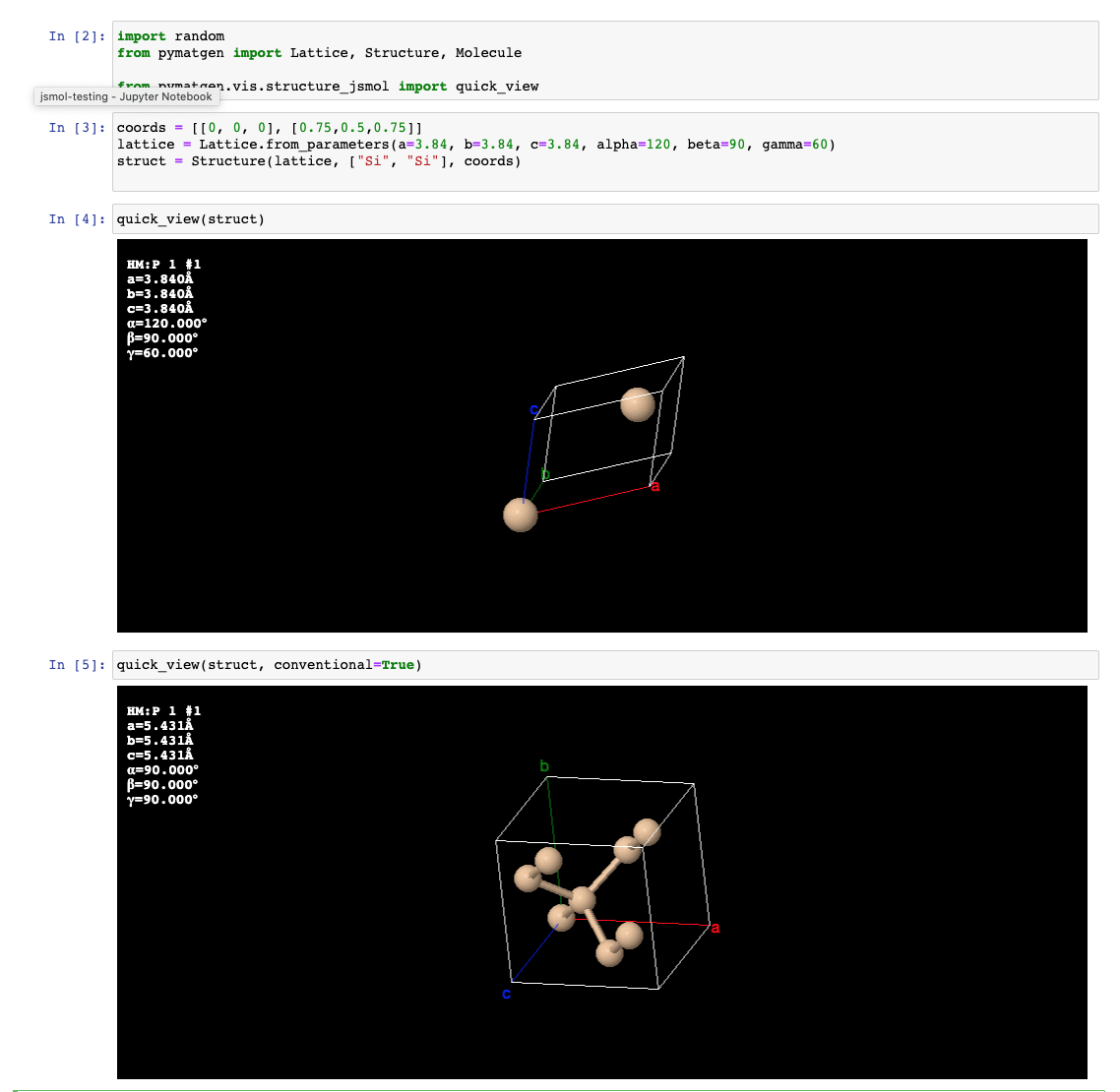\r\n",
"Hi @fekad, this looks useful but probably not something we'd want to merge into pymatgen. We're working on our own library for Jupyter visualization of crystal structures (not released yet, but see [here](https://docs.crystaltoolkit.org/jupyter.html)).",
"For some additional information for why we're making our own solution, it's so we can have 1-to-1 fidelity with the pymatgen objects themselves, e.g. show only the sites that are present and defined, or show only the bonds that are defined by pymatgen (e.g. in the StructureGraph classes). Jmol makes a lot of its own decisions about what bonds to place and what symmetrization to perform.",
"The `crystal_toolkit` looks very interesting and I completely understand why the full control over your viewer is very important.\r\n\r\nI also think that having multiple options for visualising a structure is always a good thing. There are use cases when one can be more useful than the other. These cases can be like drawing tetrahedrons on the atoms, animating vibrational modes by using finite displacement or visualising two molecules and their matching etc...\r\n\r\nAlthough I think these separate approaches have a different purpose I can easily move this function into the jupter_jsmol package so please feel free to close this PR anytime.",
"> I also think that having multiple options for visualising a structure is always a good thing. \r\n\r\nAgreed, I think a good place to share this would be as a public Gist https://gist.github.com, it's a good place to store re-usable useful code snippets even if they don't end up in the library, and they show up in Google search etc."
] | 2020-08-21T22:20:28
| 2020-08-25T01:24:15
|
2020-08-22T10:11:23Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
The tiny interface for visualising structures in jupyter notebook/lab using JSmol. The [`jupyter_jsmol`](https://github.com/fekad/jupyter-jsmol) package provides a widget (fully compatible with any other ipywidgets widgets) which is using the JSmol.
More information about the features of the package can be found [here](https://github.com/fekad/jupyter-jsmol/tree/master/examples).
Features:
* Although it is not the fastest it is full of features. Any custom [command/script](https://chemapps.stolaf.edu/jmol/docs/) can be sent to the JSmol "applet"
* it is easy to integrate the viewer with other ipywidgets (Layout, slider etc...)
* works offline and support Jupyter lab as well
## Checklist
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [ ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
|
{
"login": "fekad",
"id": 6748541,
"node_id": "MDQ6VXNlcjY3NDg1NDE=",
"avatar_url": "https://avatars.githubusercontent.com/u/6748541?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/fekad",
"html_url": "https://github.com/fekad",
"followers_url": "https://api.github.com/users/fekad/followers",
"following_url": "https://api.github.com/users/fekad/following{/other_user}",
"gists_url": "https://api.github.com/users/fekad/gists{/gist_id}",
"starred_url": "https://api.github.com/users/fekad/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/fekad/subscriptions",
"organizations_url": "https://api.github.com/users/fekad/orgs",
"repos_url": "https://api.github.com/users/fekad/repos",
"events_url": "https://api.github.com/users/fekad/events{/privacy}",
"received_events_url": "https://api.github.com/users/fekad/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1939/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1939/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1939",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1939",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1939.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1939.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1940
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1940/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1940/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1940/events
|
https://github.com/materialsproject/pymatgen/pull/1940
| 683,893,719
|
MDExOlB1bGxSZXF1ZXN0NDcxOTA5OTM1
| 1,940
|
SCAN InputSet changes - ADDGRID, ISMEAR for ScanStaticSet
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @rkingsbury "
] | 2020-08-22T00:53:08
| 2020-08-26T19:23:44
|
2020-08-26T19:22:32Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Minor updates to SCAN InputSets per recent communications with Perdew, Kresse, et al.
- Remove ADDGRID
- Enforce `ISMEAR=-5` in all static calculations with `MPScanStaticSet`
Note:
We have learned that when `PREC=Accurate`, setting `ENAUG` has no effect. Although we considered removing ENAUG from the InputSet, we decided to keep it explicitly defined in case a user (or custodian) change the `PREC` setting
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1940/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1940/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1940",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1940",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1940.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1940.patch",
"merged_at": "2020-08-26T19:22:32Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1941
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1941/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1941/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1941/events
|
https://github.com/materialsproject/pymatgen/issues/1941
| 686,253,750
|
MDU6SXNzdWU2ODYyNTM3NTA=
| 1,941
|
Error in fetching band structure for specific materials id
|
{
"login": "goodwilling",
"id": 56999195,
"node_id": "MDQ6VXNlcjU2OTk5MTk1",
"avatar_url": "https://avatars.githubusercontent.com/u/56999195?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/goodwilling",
"html_url": "https://github.com/goodwilling",
"followers_url": "https://api.github.com/users/goodwilling/followers",
"following_url": "https://api.github.com/users/goodwilling/following{/other_user}",
"gists_url": "https://api.github.com/users/goodwilling/gists{/gist_id}",
"starred_url": "https://api.github.com/users/goodwilling/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/goodwilling/subscriptions",
"organizations_url": "https://api.github.com/users/goodwilling/orgs",
"repos_url": "https://api.github.com/users/goodwilling/repos",
"events_url": "https://api.github.com/users/goodwilling/events{/privacy}",
"received_events_url": "https://api.github.com/users/goodwilling/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @goodwilling, this is a Materials Project issue so would be better discussed at [matsci.org/materials-project](matsci.org/materials-project)\r\n\r\nIn brief, we're going through a series of large database updates, including this week expanding our band structures (from 54k -> 76k) and drastically improving the quality of our densities of states (see the release notes on materialsproject.org). In the process we've introduce a bug, this bug is being fixed currently but it might take a day or two before this will work."
] | 2020-08-26T11:47:39
| 2020-09-03T20:53:50
|
2020-09-03T20:53:50Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
The following script/URL, either
mpr.get_bandstructure_by_material_id("mp-867624")
or
https://www.materialsproject.org/rest/v2/materials/mp-867624/vasp/bandstructure?API_KEY=************
gives the error:
{
"valid_response": false,
"error": "too many indices for array: array is 0-dimensional, but 2 were indexed",
"version": {
"db": "2020.08.21",
"pymatgen": "2020.8.3",
"rest": "2.0"
},
"created_at": "2020-08-26T11:00:09.393429",
"traceback": "Traceback (most recent call last):\n File \"/var/www/python/matgen_prod/materials_django/rest/rest.py\", line 94, in wrapped\n d = func(*args, **kwargs)\n File \"/var/www/python/matgen_prod/materials_django/materials/rest.py\", line 114, in get_vasp_property\n loadee=d[\"response\"])\n File \"/var/www/python/matgen_prod/materials_django/materials/rest.py\", line 56, in mid_ETL\n value = transform(s)\n File \"/var/www/python/matgen_prod/materials_django/materials/rest.py\", line 68, in es_transform\n return extracted.as_dict()\n File \"/opt/miniconda3/envs/mpprod3/lib/python3.6/site-packages/pymatgen/electronic_structure/bandstructure.py\", line 858, in as_dict\n d[\"is_metal\"] = self.is_metal()\n File \"/opt/miniconda3/envs/mpprod3/lib/python3.6/site-packages/pymatgen/electronic_structure/bandstructure.py\", line 307, in is_metal\n if np.any(values[i, :] - self.efermi < -efermi_tol) and \\\nIndexError: too many indices for array: array is 0-dimensional, but 2 were indexed\n"
}
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1941/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1941/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1942
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1942/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1942/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1942/events
|
https://github.com/materialsproject/pymatgen/pull/1942
| 687,607,381
|
MDExOlB1bGxSZXF1ZXN0NDc1MDA0MjY2
| 1,942
|
Update SQS integration
|
{
"login": "rwoodsrobinson",
"id": 7270689,
"node_id": "MDQ6VXNlcjcyNzA2ODk=",
"avatar_url": "https://avatars.githubusercontent.com/u/7270689?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rwoodsrobinson",
"html_url": "https://github.com/rwoodsrobinson",
"followers_url": "https://api.github.com/users/rwoodsrobinson/followers",
"following_url": "https://api.github.com/users/rwoodsrobinson/following{/other_user}",
"gists_url": "https://api.github.com/users/rwoodsrobinson/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rwoodsrobinson/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rwoodsrobinson/subscriptions",
"organizations_url": "https://api.github.com/users/rwoodsrobinson/orgs",
"repos_url": "https://api.github.com/users/rwoodsrobinson/repos",
"events_url": "https://api.github.com/users/rwoodsrobinson/events{/privacy}",
"received_events_url": "https://api.github.com/users/rwoodsrobinson/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
] | null |
[
"Thanks @rwoodsrobinson !"
] | 2020-08-27T23:34:46
| 2020-08-29T02:00:51
|
2020-08-29T02:00:43Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
* Added spin parsing to SQSTransformation
* Added cluster parsing to SQSTransformation
* Added tests
* Fix .out file parsing bug
* Fix oxidation state and spin bug in periodic_table
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1942/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1942/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1942",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1942",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1942.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1942.patch",
"merged_at": "2020-08-29T02:00:43Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1943
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1943/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1943/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1943/events
|
https://github.com/materialsproject/pymatgen/issues/1943
| 688,615,834
|
MDU6SXNzdWU2ODg2MTU4MzQ=
| 1,943
|
a problem of import on win10
|
{
"login": "Yuxinya",
"id": 56122867,
"node_id": "MDQ6VXNlcjU2MTIyODY3",
"avatar_url": "https://avatars.githubusercontent.com/u/56122867?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Yuxinya",
"html_url": "https://github.com/Yuxinya",
"followers_url": "https://api.github.com/users/Yuxinya/followers",
"following_url": "https://api.github.com/users/Yuxinya/following{/other_user}",
"gists_url": "https://api.github.com/users/Yuxinya/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Yuxinya/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Yuxinya/subscriptions",
"organizations_url": "https://api.github.com/users/Yuxinya/orgs",
"repos_url": "https://api.github.com/users/Yuxinya/repos",
"events_url": "https://api.github.com/users/Yuxinya/events{/privacy}",
"received_events_url": "https://api.github.com/users/Yuxinya/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-08-29T22:45:23
| 2020-08-30T09:12:50
|
2020-08-30T09:12:50Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
it was install,but it can only' import pymatgen'. if you 'import pymatgen.core' or 'import pymatgen.analysis' ,it would be that:
No module named 'pymatgen.analysis'; 'pymatgen' is not a package
on wind10
|
{
"login": "Yuxinya",
"id": 56122867,
"node_id": "MDQ6VXNlcjU2MTIyODY3",
"avatar_url": "https://avatars.githubusercontent.com/u/56122867?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Yuxinya",
"html_url": "https://github.com/Yuxinya",
"followers_url": "https://api.github.com/users/Yuxinya/followers",
"following_url": "https://api.github.com/users/Yuxinya/following{/other_user}",
"gists_url": "https://api.github.com/users/Yuxinya/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Yuxinya/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Yuxinya/subscriptions",
"organizations_url": "https://api.github.com/users/Yuxinya/orgs",
"repos_url": "https://api.github.com/users/Yuxinya/repos",
"events_url": "https://api.github.com/users/Yuxinya/events{/privacy}",
"received_events_url": "https://api.github.com/users/Yuxinya/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1943/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1943/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1944
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1944/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1944/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1944/events
|
https://github.com/materialsproject/pymatgen/issues/1944
| 691,460,623
|
MDU6SXNzdWU2OTE0NjA2MjM=
| 1,944
|
pymatgen.io.gaussian write_file() gives wrong output when cart_coords=False
|
{
"login": "wudihua",
"id": 4005299,
"node_id": "MDQ6VXNlcjQwMDUyOTk=",
"avatar_url": "https://avatars.githubusercontent.com/u/4005299?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/wudihua",
"html_url": "https://github.com/wudihua",
"followers_url": "https://api.github.com/users/wudihua/followers",
"following_url": "https://api.github.com/users/wudihua/following{/other_user}",
"gists_url": "https://api.github.com/users/wudihua/gists{/gist_id}",
"starred_url": "https://api.github.com/users/wudihua/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/wudihua/subscriptions",
"organizations_url": "https://api.github.com/users/wudihua/orgs",
"repos_url": "https://api.github.com/users/wudihua/repos",
"events_url": "https://api.github.com/users/wudihua/events{/privacy}",
"received_events_url": "https://api.github.com/users/wudihua/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I am confused. How do you know that the cart=False version is wrong? That is in the Zmatrix format. \r\n",
"> \r\n> \r\n> I am confused. How do you know that the cart=False version is wrong? That is in the Zmatrix format.\r\n\r\nI use Gaussview to visualize the two files and they are different.\r\n\r\ncart_output.gjf\r\n\r\n\r\noutput.gjf\r\n\r\n\r\n\r\n",
"Hi\r\n\r\nFor me the issue comes from the conversion into z-matrix and not from the Gaussian class. I cannot exactly say why but my guess is that your atoms are not well sorted and the conversion algorithm from cartesian coords to z-matrix fails. In your case successive atoms may be far away from each other. When the program tries to build the z-matrix it will consider distances, angles and dihedrals that do not correspond to \"real\" internal coordinate. As you have a lot of atoms, at some point, maybe you consider aligned atoms (or closed to be aligned), or another kind of wrong geometry for a z-matrix and from this point the geometry is wrong.\r\n\r\nI sorted the atom of your nanotube according to the axis of the nanotube and at the end the final z-matrix is ok. Both gjf input files (in cartesian coordinates or z-matrix) are correctly written (see the picture).\r\n\r\n\r\n\r\n\r\nHere is what I did to sort the atoms. It is not so elegant but to be quick I used pandas sorting facilities.\r\n\r\n```py\r\nimport pymatgen as mg\r\nimport numpy as np\r\nimport pandas as pd\r\n\r\n# input.xyz is the xyz file you put in your message\r\nmol = mg.Molecule.from_file(\"input.xyz\")\r\n\r\n# I rotated the nanotube around y axes to aligned the nanotube axis and the z-axis\r\n# this is not exactly 30 degrees but it is not so far\r\nop = mg.SymmOp.from_axis_angle_and_translation(axis=[0, 1, 0], angle=-30)\r\nmol.apply_operation(op)\r\nmol.to(fmt=\"xyz\", filename=\"input2.xyz\")\r\n\r\nmol2 = mg.Molecule.from_file(\"input2.xyz\")\r\ncoords = mol2.cart_coords\r\n\r\n# geometric barycenter of the nanotube (without masses)\r\nG = coords.sum(axis=0) / len(coords)\r\ncoords -= G\r\n\r\ndf = pd.DataFrame({\"species\": mol2.species, \"x\": coords[:, 0], \"y\": coords[:, 1], \"z\": coords[:, 2]})\r\n\r\n# compute the radius of the nanotube\r\ndf[\"r\"] = np.sqrt(df[\"x\"]**2 + df[\"y\"]**2)\r\n\r\n# compute the angle theta with x axis.\r\ndf[\"theta\"] = np.degrees(np.arctan2(df.y, df.x))\r\ndf[\"theta\"] = np.where(df[\"theta\"] < 0, 360 + df[\"theta\"], df[\"theta\"])\r\ndf.describe()\r\ndf.head()\r\n\r\n# from now, you can use r, theta, z as cylindrical coordinates\r\n# now the key part, I sort according first to z and second theta (maybe theta is not necessary\r\ndf.sort_values(by=[\"z\", \"theta\"], ascending=True, inplace=True)\r\n\r\n# set up the sorted molecule\r\ncoords = df[[\"x\", \"y\", \"z\"]].values\r\nm_sorted = mg.Molecule(species=df.species.values, coords=coords)\r\n\r\ngau = mg.io.gaussian.GaussianInput(m_sorted)\r\ngau.write_file(\"gau_sorted.gjf\", cart_coords=False)\r\nm_sorted.to(\"xyz\", \"sorted.xyz\")\r\n```\r\n\r\nMay be you do not need to compute radius and angle. Sorting according to z may be enough.\r\n",
"Thank you for your thorough response, it's very helpful. It also inspired me to recheck this issue.\r\n\r\nAfter I looked at this strange problem closely, I think it's the atom 165 that causes the problem, the line is:\r\n`N 164 B164 163 A164 144 D164`\r\nHowever, 164, 163 and 144 is almost on a line, so it causes numerial precision issue when calculating dihedral angle and then the error accumulate.\r\n\r\nThe get_zmatix function (in pymatgen.io.gaussian.GaussianInput class) uses a function called _find_nn_pos_before_site(i) function, this function does not guarantee that the three closest atoms are not (approximatly) collinear. \r\n\r\nI actually make the program work properly by use the following code (line 400 in pymatgen.io.gaussian):\r\n```python\r\n #nn = self._find_nn_pos_before_site(i)\r\n nn = [i-1,i-2,i-3]\r\n```\r\n\r\nBut this would also not always work. I would suggest the program to check the nn[0] - nn[1] - nn[2] angle, if it's close to +/-180 or 0 degree then del nn[2] and recheck. If len(nn)<3 then raise error.\r\n\r\nAnother solution is to tackle the numerical precision issue in dihedral angle calculation, but I think use dihedral angle in dangerous region might not be an approprate practice.",
"Thanks for the debug. I think I know what is happening. But I need to think how best to solve this. There is no unique way of defining the z-matrix and in fact, for something like a nanotube, it is probably even clearer to define it relative to the center axis using virtual atoms since that allows symmetric angles to be defined precisely. "
] | 2020-09-02T22:05:06
| 2023-08-13T16:33:45
|
2023-08-13T16:33:44Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
In some cases pymatgen.io.gaussian write_file() gives wrong output when cart_coords=False
**To Reproduce**
Here is the code to reproduce this issue.
```python
#!/usr/bin/env python
import pymatgen.io.xyz as xyzio
import pymatgen.io.gaussian as gaussianio
xyz_filename = "input.xyz"
xyz = xyzio.Molecule.from_file(xyz_filename)
gjf = gaussianio.GaussianInput(xyz)
gjf_filename = "output.gjf"
gjf.write_file(gjf_filename,cart_coords=False) # Bad structure
gjf.write_file("cart_"+gjf_filename,cart_coords=True) # Good structure
```
The input.xyz is the following:
```
368
N 0.00000000 0.00000000 0.00000000
B 0.00000000 0.00000000 1.46015992
N 1.25732289 0.00000000 2.19736263
B 1.25261991 0.03549742 3.65365385
N 2.51549326 0.02806676 4.38132181
B 2.50997160 0.05650466 5.83894909
N 3.77191199 0.04215517 6.56692842
B 3.76815411 0.06965821 8.02550727
N 5.03084433 0.06452678 8.75174618
B 5.02796191 0.08708214 10.21180961
N 6.29541849 0.10821986 10.93564957
B 6.28247833 0.08613305 12.39412455
N 7.54559625 0.05381271 13.12803913
B 7.53859372 0.06111702 14.58548129
N 8.80105702 0.02888990 15.31550304
B 8.79422011 0.03328746 16.77313675
N 10.05390966 -0.01015904 17.50354864
B 10.05848155 -0.00720206 18.96076520
N 11.32066296 -0.00507892 19.69651073
B 12.50823722 -0.49521155 18.99936572
N 12.42784733 -0.84171398 17.58874183
B 11.23326591 -0.51355407 16.82113136
N 11.18595667 -0.79470248 15.39253350
B 9.98296323 -0.47528008 14.63326139
N 9.92873699 -0.77653922 13.20758313
B 8.72572982 -0.45592657 12.44766477
N 8.66426132 -0.77010942 11.02535181
B 7.46679841 -0.43204174 10.26058297
N 7.41426708 -0.75811093 8.83858839
B 6.21076930 -0.44889080 8.07512200
N 6.15572850 -0.77496332 6.65447245
B 4.95489366 -0.46575924 5.88889983
N 4.90501635 -0.78018609 4.46570478
B 3.70004121 -0.47834236 3.70327870
N 3.64509580 -0.80329604 2.28397418
B 2.43908193 -0.51079315 1.52301176
N 2.37663618 -0.83942908 0.10474481
B 1.20081757 -0.48645210 -0.67534843
N 1.22875767 -0.74705963 -2.11447842
H 0.53113171 -0.34190091 -2.70899625
B 2.15835612 -1.69649261 -2.63987571
N 3.13052271 -2.33492346 -1.78536904
B 3.30956812 -1.81919518 -0.43121460
N 4.36428491 -2.34687670 0.42456895
B 4.56623515 -1.79868914 1.75811355
N 5.61926523 -2.33739901 2.60951705
B 5.81745733 -1.78763845 3.94418212
N 6.86296302 -2.33553573 4.79948324
B 7.06246323 -1.78540046 6.13371688
N 8.09985094 -2.33972811 6.99384643
B 8.30954971 -1.78143039 8.32310709
N 9.34090811 -2.34692278 9.18213336
B 9.56855522 -1.78236097 10.50524829
N 10.61670239 -2.33707801 11.35128121
B 10.83576163 -1.78656858 12.68308433
N 11.88989680 -2.33652718 13.52474454
B 12.09739434 -1.79764370 14.86294576
N 13.14785801 -2.34760033 15.70808360
B 13.35896265 -1.82201432 17.05087326
N 14.44338760 -2.33576343 17.88344134
B 14.97390583 -3.65981947 17.56194895
N 15.89826717 -4.36187136 18.45105491
H 16.38854770 -3.85425336 19.16329573
B 15.97921696 -5.78763522 18.40120182
N 15.23760025 -6.54177996 17.41978309
B 14.58074949 -5.81423585 16.33732109
N 14.49646037 -4.36256788 16.38167510
B 13.66111877 -3.67326770 15.40813357
N 13.25541043 -4.35432476 14.18665968
B 12.40420627 -3.66530386 13.22660885
N 11.97985672 -4.35513094 12.01574734
B 11.13137224 -3.66559552 11.05430665
N 10.70457908 -4.35880481 9.84731483
B 9.86580439 -3.66912986 8.87973099
N 9.44550270 -4.36049304 7.67053874
B 8.61466649 -3.66732964 6.69646725
N 8.20630316 -4.35756927 5.48233758
B 7.37839660 -3.66474755 4.50485910
N 6.97565847 -4.35427115 3.28676130
B 6.13969052 -3.66424149 2.31291592
N 5.73252451 -4.35395400 1.09682725
B 4.88343778 -3.67177516 0.13063051
N 4.46357019 -4.35976118 -1.08283998
B 3.67548758 -3.65765463 -2.08415536
N 3.36173997 -4.36271266 -3.32684330
H 3.00264322 -3.85468526 -4.11263531
B 3.44733728 -5.78827440 -3.37149594
N 3.91577765 -6.54049805 -2.23223635
B 4.53060924 -5.81299530 -1.12604741
N 5.13616935 -6.53484222 -0.01490971
B 5.79024668 -5.80753041 1.06250132
N 6.39699997 -6.54081361 2.16588065
B 7.03687711 -5.80874421 3.24970467
N 7.63660688 -6.54245797 4.35667817
B 8.27490771 -5.81084556 5.44106734
N 8.87774519 -6.54372119 6.54540391
B 9.52104423 -5.81319065 7.62654587
N 10.12930522 -6.54375920 8.72777042
B 10.77875598 -5.81199678 9.80583430
N 11.39383451 -6.54285969 10.90340089
B 12.04608607 -5.80915740 11.97930944
N 12.66149846 -6.54034088 13.07714876
B 13.31557762 -5.80839202 14.15322073
N 13.93059353 -6.53697309 15.25197311
B 13.52017657 -7.91058664 15.49186440
N 13.88999372 -8.56481271 16.73941721
B 14.82521454 -7.92549620 17.65197408
N 15.23662860 -8.66696975 18.84294054
H 16.00777971 -8.33964884 19.39405359
B 14.42944986 -9.74940991 19.31077419
N 13.27718017 -10.19613210 18.56605478
B 13.07289221 -9.66900337 17.21952841
N 11.99365867 -10.17824777 16.38375816
B 11.82380348 -9.68853796 15.02407436
N 12.65000788 -8.59091289 14.54295781
B 12.26768540 -7.92256159 13.30741522
N 11.39294436 -8.60236926 12.36043952
B 11.00402350 -7.92785309 11.13024243
N 10.12468016 -8.60193167 10.18459935
B 9.74125221 -7.92922587 8.95228256
N 8.86566177 -8.60233952 8.00350861
B 8.48657940 -7.92829804 6.76965379
N 7.61285745 -8.60030691 5.81940333
B 7.23848771 -7.92503681 4.58401248
N 6.36382062 -8.59588722 3.63278903
B 5.98980271 -7.91984223 2.39708373
N 5.10812427 -8.58657840 1.44984275
B 4.72615144 -7.90865962 0.21912240
N 3.83351329 -8.56531340 -0.72633981
B 3.50968962 -7.92389719 -1.99140546
N 2.68061839 -8.66281699 -2.94301291
H 2.60388163 -8.34553590 -3.89054968
B 1.88128741 -9.75461196 -2.48360252
N 1.94314120 -10.19364292 -1.11019617
B 3.00869694 -9.67044317 -0.26053792
N 3.19395165 -10.18459723 1.09067416
B 4.28019602 -9.68649530 1.92157379
N 4.45476132 -10.20365745 3.27288611
B 5.53898872 -9.70031340 4.10423275
N 5.71367101 -10.21862500 5.45555715
B 6.79443936 -9.71002565 6.28904691
N 6.97202888 -10.22817532 7.63929805
B 8.05071061 -9.71483848 8.47303827
N 8.23039097 -10.22984228 9.82356039
B 9.31000269 -9.71379289 10.65580521
N 9.49154699 -10.22737507 12.00712228
B 10.57049722 -9.70649348 12.83799927
N 10.74524635 -10.20960689 14.19385952
B 9.61586943 -10.85693611 14.84680604
N 9.65506800 -11.11777353 16.27997567
B 10.86582935 -10.82190587 17.03475556
N 10.90617094 -11.05838577 18.47183207
B 12.13760589 -10.84282565 19.21586436
N 12.15067946 -11.18603379 20.63693686
H 13.02016013 -11.21808311 21.13587892
B 10.91345445 -11.27842448 21.34653134
N 9.65039216 -11.15709956 20.65976768
B 9.64951309 -11.15346994 19.19909560
N 8.39253330 -11.15008223 18.46164727
B 8.39494231 -11.19142462 17.00627149
N 7.13166254 -11.18820529 16.27807768
B 7.13719008 -11.21389795 14.82063435
N 8.39752067 -11.12989862 14.09447815
B 8.35900520 -10.87150937 12.66056390
N 7.14038351 -11.14148944 11.90799550
B 7.10059799 -10.88022662 10.47505320
N 5.88269139 -11.15045936 9.72131008
B 5.84290100 -10.88121032 8.28940915
N 4.62504528 -11.14888160 7.53563592
B 4.58537054 -10.87348000 6.10497678
N 3.36705502 -11.13770919 5.35053527
B 3.32748949 -10.85968718 3.91994628
N 2.10844084 -11.12121251 3.16611439
B 2.06384865 -10.82774545 1.74003964
N 0.84279827 -11.07214516 0.98405567
B 0.81655605 -10.85367998 -0.45424683
N -0.40470552 -11.20382549 -1.17896077
H -0.39477524 -11.25498834 -2.17991969
B -1.63817734 -11.31309847 -0.46579726
N -1.67912519 -11.17887246 0.97056085
B -0.41716411 -11.17388619 1.70498886
N -0.41227627 -11.17359552 3.16266586
B 0.84876864 -11.20407178 3.89083559
N 0.84411039 -11.19194744 5.34836603
B 2.10626429 -11.21981437 6.07609470
N 2.10186652 -11.19860570 7.53350640
B 3.36403679 -11.22814361 8.26215439
N 3.35931725 -11.20681425 9.71922403
B 4.62156436 -11.22929426 10.44779880
N 4.61741855 -11.20295975 11.90463908
B 5.87963116 -11.22383206 12.63362858
N 5.87481733 -11.19738685 14.09132103
B 4.68898696 -10.69106746 14.77063362
N 4.74206996 -10.37763818 16.19347424
B 5.94639795 -10.68001403 16.95594756
N 5.99887638 -10.36290674 18.37771286
B 7.20819487 -10.64688342 19.13753239
N 7.27114642 -10.32115753 20.55671819
B 8.44963756 -10.66762421 21.33566073
N 8.42513940 -10.40474199 22.77388754
H 9.13091225 -10.79957234 23.36659589
B 7.49445972 -9.45680985 23.29926690
N 6.51103987 -8.83016694 22.44935696
B 6.33591440 -9.34321131 21.09293122
N 5.27962931 -8.81914463 20.23767427
B 5.07664260 -9.36645803 18.90381974
N 4.02080406 -8.83159715 18.05533621
B 3.82134326 -9.37835433 16.71960205
N 2.76987365 -8.83754399 15.86924455
B 2.56704714 -9.38360165 14.53356689
N 3.48792874 -10.38230027 14.00680524
B 3.43217733 -10.69755158 12.58554634
N 2.22963968 -10.38869010 11.82331328
B 2.17469320 -10.70121988 10.40155998
N 0.97110252 -10.39182096 9.64053825
B 0.91619045 -10.69984110 8.21799141
N -0.28933535 -10.39304242 7.45770539
B -0.34159919 -10.69497009 6.03282738
N -1.54250598 -10.37814020 5.27124170
B -1.59226735 -10.66965942 3.84564390
N -2.78898912 -10.34816945 3.07882678
B -2.86913219 -10.69843291 1.66909074
N -4.12743905 -10.44142454 0.96915499
H -4.28851463 -10.84590485 0.06644884
B -5.04917982 -9.49091790 1.50659015
N -4.79993438 -8.85076481 2.77590715
B -3.72100656 -9.36663143 3.61381655
N -3.51230555 -8.83684157 4.95498617
B -2.46230698 -9.38358715 5.80253766
N -2.25988004 -8.84552324 7.14141366
B -1.20530740 -9.39057178 7.98610461
N -0.99750311 -8.84425626 9.32029769
B 0.05424619 -9.38996972 10.16845657
N 0.25549007 -8.84685615 11.50444645
B 1.31058295 -9.38771501 12.35060976
N 1.51454784 -8.84225310 13.68516889
B 1.00003867 -7.51056113 13.97819460
N 1.41710280 -6.81957993 15.18994400
B 2.24982811 -7.50887591 16.16351764
N 2.65702354 -6.81917708 17.38086916
B 3.49632907 -7.50577545 18.35237831
N 3.89963496 -6.81689409 19.57118301
B 4.75764098 -7.49516409 20.53174671
N 5.18215398 -6.80510217 21.74224059
B 5.96693139 -7.50569394 22.74589731
N 6.27662948 -6.80134341 23.98936431
H 6.64874232 -7.30714260 24.77088502
B 6.19463234 -5.37541574 24.03117823
N 5.71558917 -4.62387344 22.89666387
B 5.10622626 -5.35273968 21.78739871
N 4.49636641 -4.63231202 20.67836235
B 3.84745138 -5.36229039 19.59913386
N 3.24464862 -4.63124766 18.49338875
B 2.60123409 -5.36417370 17.41122702
N 2.00289335 -4.63370756 16.30387076
B 1.35646654 -5.36500512 15.22178715
N 0.74792318 -4.63336174 14.12125508
B 0.10493095 -5.36663469 13.04072998
N 0.15752770 -6.82266381 13.01089996
B -0.25371173 -7.51292959 11.79775293
N -1.08993396 -6.82178290 10.82603563
B -1.51134952 -7.51392933 9.61775604
N -2.35573743 -6.82512777 8.65024781
B -2.77360134 -7.51566553 7.43809939
N -3.61322953 -6.82517813 6.46934433
B -4.02386620 -7.50966307 5.25206113
N -4.85990045 -6.82277430 4.27715027
B -5.32944235 -7.52609392 3.09315142
N -6.24344871 -6.82013102 2.19555898
H -6.74470838 -7.32794491 1.49184615
B -6.32054884 -5.39405090 2.24449534
N -5.56625015 -4.64258953 3.21820053
B -4.92420748 -5.36930597 4.30922536
N -4.26849738 -4.64583454 5.39052383
B -3.66781147 -5.37173004 6.49935382
N -3.02071043 -4.63870783 7.57932062
B -2.41125516 -5.37079122 8.68115878
N -1.75587327 -4.63779573 9.75502187
B -1.14921600 -5.36775486 10.86183824
N -0.51120262 -4.63348169 11.94283779
B -0.09293379 -3.25942320 11.70579703
N 0.76997725 -2.57452940 12.65622339
B 1.15873433 -3.25683938 13.88329746
N 2.04022915 -2.58631647 14.83053741
B 2.41093693 -3.25632139 16.06763892
N 3.28767297 -2.58390518 17.01809904
B 3.65758395 -3.25538532 18.25598013
N 4.53612364 -2.58587838 19.20560638
B 4.91464372 -3.26175134 20.43777337
N 5.80680323 -2.60531358 21.38426252
B 6.12308940 -3.24088296 22.65363009
N 6.94257187 -2.49625696 23.60864024
H 7.02944191 -2.82166307 24.55319544
B 7.75038154 -1.41239787 23.14638313
N 7.68711471 -0.96789630 21.77516356
B 6.62815436 -1.49793699 20.92025144
N 6.44589399 -0.98419244 19.56880906
B 5.36392673 -1.48582312 18.73357008
N 5.18992881 -0.96875749 17.38264760
B 4.11182179 -1.48034142 16.54575729
N 3.93978732 -0.96677070 15.19366762
B 2.86258134 -1.48310981 14.35707589
N 2.68665397 -0.96687277 13.00687920
B 1.60667041 -1.48292634 12.17749820
N 1.43244338 -0.96751320 10.82373238
B 0.35243905 -1.47944702 9.99545216
N -0.45326259 -2.59821119 10.45634711
B -1.34962777 -3.26080197 9.52110891
N -1.73270709 -2.58661959 8.28801452
B -2.61300187 -3.26153654 7.34276355
N -2.98930009 -2.59395106 6.10478726
B -3.85910406 -3.27241099 5.15427681
N -4.23180091 -2.61343976 3.90981892
B -5.15958910 -3.25712070 2.99229947
N -5.56744629 -2.51770967 1.79807460
H -6.34646146 -2.83672263 1.25417309
B -4.77284263 -1.42171207 1.34115456
N -3.61867163 -0.97972793 2.08641918
B -3.41661992 -1.50579177 3.43316053
N -2.34225058 -0.99105525 4.27267262
B -2.16770549 -1.49182724 5.62854570
N -1.09012425 -0.97111214 6.46071665
B -0.91041625 -1.48333441 7.81197218
N 0.17784137 -0.97814128 8.63867123
B 1.29571836 -0.30050393 7.99553899
N 2.50768004 -0.01689388 8.75023859
B 2.55475625 -0.29622781 10.18084044
N 3.76645440 -0.00894387 10.93452331
B 3.81108764 -0.30345527 12.36102319
N 5.02469976 -0.02199885 13.11919405
B 5.06354804 -0.30314612 14.54776996
N 6.27807862 -0.02787978 15.30843332
B 6.31607528 -0.31008184 16.73788161
N 7.53344545 -0.04484877 17.49571357
B 7.57650541 -0.34163399 18.92092206
N 8.79769800 -0.10287234 19.67999227
B 8.82112831 -0.32010882 21.11804391
N 10.04321734 0.02072243 21.84528732
H 10.03651820 0.04936395 22.84785559
B 11.27878121 0.11309705 21.13396190
H 12.28815840 0.26892673 21.75421617
N 1.24999766 -0.03072387 6.56407890
B 0.03427020 -0.31158005 5.81163413
N -0.00638278 -0.04628751 4.37919206
B -1.21549447 -0.34251288 3.62335575
N -1.25675372 -0.09544106 2.18769673
B -2.48579535 -0.31980854 1.44175037
N -2.49846789 0.02491368 0.02007666
H -3.36874994 0.07438847 -0.47444194
B -1.26205552 0.13159492 -0.68764563
H -1.28863944 0.30140216 -1.86971352
H -5.06165947 -0.89185186 0.31034230
H 8.50231439 -0.88903735 23.91350298
H -6.99962888 -4.82762687 1.44129617
H 6.55425422 -4.80863221 25.01969780
H -6.02118869 -9.21048235 0.87167950
H 7.55877182 -9.16971642 24.45727188
H -2.64580463 -11.48819410 -1.08304845
H 10.94174107 -11.43738954 22.53046121
H 1.13387797 -10.28266112 -3.25120851
H 14.71270342 -10.26917121 20.34882092
H 3.09548669 -6.35465812 -4.36271212
H 16.65790372 -6.35256731 19.20617874
B 14.68428695 -1.70930919 19.16112728
N 13.76352579 -0.76028547 19.70120501
H 13.92075178 -0.36613478 20.60968486
H 15.65250496 -1.99782391 19.79913888
H 2.09920511 -1.97800276 -3.79924276
```
**Expected behavior**
output.gjf and cart_output.gjf should be the same, but only cart_output.gjf gives the right answer.
**Desktop**
OS: Mac
Version: 2020.8.13
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1944/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1944/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1945
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1945/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1945/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1945/events
|
https://github.com/materialsproject/pymatgen/pull/1945
| 692,471,979
|
MDExOlB1bGxSZXF1ZXN0NDc5MDY3NTcw
| 1,945
|
MPRester database version checking and logging
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-09-03T22:51:27
| 2020-09-10T06:36:12
|
2020-09-10T06:36:08Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Merge pending changes to materialsproject.org and subsequent checking.
The intent of this PR is to better notify users of `MPRester` what the Materials Project database version is, and to notify the user if this version changes.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1945/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1945/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1945",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1945",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1945.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1945.patch",
"merged_at": "2020-09-10T06:36:08Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1946
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1946/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1946/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1946/events
|
https://github.com/materialsproject/pymatgen/issues/1946
| 692,711,434
|
MDU6SXNzdWU2OTI3MTE0MzQ=
| 1,946
|
[feature request] embedding structure in another
|
{
"login": "mturiansky",
"id": 8866816,
"node_id": "MDQ6VXNlcjg4NjY4MTY=",
"avatar_url": "https://avatars.githubusercontent.com/u/8866816?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mturiansky",
"html_url": "https://github.com/mturiansky",
"followers_url": "https://api.github.com/users/mturiansky/followers",
"following_url": "https://api.github.com/users/mturiansky/following{/other_user}",
"gists_url": "https://api.github.com/users/mturiansky/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mturiansky/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mturiansky/subscriptions",
"organizations_url": "https://api.github.com/users/mturiansky/orgs",
"repos_url": "https://api.github.com/users/mturiansky/repos",
"events_url": "https://api.github.com/users/mturiansky/events{/privacy}",
"received_events_url": "https://api.github.com/users/mturiansky/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I don't think this functionality is implemented as a method anywhere, but I am unclear whether this requires its own method at all. Thinking about it in my head, this should not require more than 10 lines of code to do?",
"For this specific case, the `merge_sites` docstring claims `mode=='delete'` exists, does this definitely not work? An omission if so.",
"I have not tested, but it seems that it should work. Delete is basically the default option where you don't even \"merge\" sites per se. That is why in the code it says \"if mode == \"sum\". That is when you need to combine site species. For delete, the overlapping site is just ignored.",
"For `merge_sites`, I would expect that for `mode == 'delete'`, the coordinates would not be averaged over each of the sites being merged.\r\n\r\nI agree that a naive implementation making certain assumptions might take 10 lines of code (definitely if `merge_sites` behaves how I expect), but if you want to be more robust, it could get long. The naive implementation might assume the lattices are similar, but this doesn't have to be the case. For example, you might start with the a 3x3x3 cubic primitive cell and want to embed this in a conventional standard supercell. Also, you may only need to determine a translation to align the atoms in the cell, but there could be rotations also, which will get complex.",
"Hi @mturiansky,\r\nAs I understand you want to implant a smaller SC in a large (non-diagonal) SC.\r\nthere might be a way of doing this with a ghost unit cell.\r\n\r\nThe setup will be:\r\n1. Get a reference base unit cell used for both SCs.\r\n2. Use the StructureMatcher module to get the mapping between the pure UC and the SC for both supercells. <-- these is gonna be nice and invertible.\r\n3. The you can get put those two transformations together to implant the smaller SC into the larger SC.\r\n\r\nI'm adding a simpler version of this that only deals with just mapping positions from SC to UC to pymatgen-diffusion but it should also be portable for this case provided the defect is center-cell and you add some logic to lock down the atoms at the boundary during the mapping. ",
"Thanks for the suggestion. Since no one seemed to be interested in a more general version, I just kept it simple for what I needed. I ended up slightly modifying the [merge_sites function on my fork](https://github.com/mturiansky/pymatgen/blob/65977c24727e9d9779dc23ae160da1ceaa43a510/pymatgen/core/structure.py#L3877-L3933) to check for a \"keep\" site property. This way I can just tag the sites that I want to keep before adding them to the supercell and running merge_sites. I just figured out the translation vector to line the sites up by hand. This won't work for vacancies of course, but it won't really care if your defect is in the center of the cell."
] | 2020-09-04T04:49:07
| 2023-08-13T16:33:45
|
2023-08-13T16:33:45Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I'd like to be able to embed one structure inside another. Roughly speaking, some of the sites should match exactly, while others may be quite different. The application that I have in mind is embedding a defect from a small supercell inside a larger supercell to cut down on the computational cost of the relaxation.
It seems like [this](https://github.com/materialsproject/pymatgen/blob/1a31cda59f4d7cb8b1b486d77928cc2ff7037e25/pymatgen/core/structure.py#L3540) almost does part of what I need. Although, it looks like mode == 'delete' (which I need) is not implemented.
Is this functionality implemented anywhere in pymatgen? I can work on it if not, but no sense in duplicating work.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1946/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1946/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1947
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1947/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1947/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1947/events
|
https://github.com/materialsproject/pymatgen/pull/1947
| 693,773,210
|
MDExOlB1bGxSZXF1ZXN0NDgwMjM5OTYw
| 1,947
|
PackmolRunner enhancements for safe looping
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-09-04T22:57:48
| 2020-09-05T17:54:33
|
2020-09-05T16:48:11Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Enhance `PackmolRunner` to work more robustly when called within for loops.
* Use context managers for opening the input file and running the subprocess to ensure that both are properly closed
* Replace `ScratchDir` with `TemporaryDirectory` to eliminate non-obvious file deletion
Re: bullet point 2 - when the `ScratchDir` class is called with `copy_to_current_on_exit=True`, it not only copies output files from the temporary to the current directory, but also DELETES any file in the current working directory that is not present in the temporary directory. This behavior is undocumented and made it impossible to use `PackmolRunner` to loop through a series of structures and place output files in a single directory (everything except the latest output would be deleted on each iteration).
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1947/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1947/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1947",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1947",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1947.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1947.patch",
"merged_at": "2020-09-05T16:48:11Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1948
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1948/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1948/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1948/events
|
https://github.com/materialsproject/pymatgen/pull/1948
| 694,145,555
|
MDExOlB1bGxSZXF1ZXN0NDgwNTY1ODQ1
| 1,948
|
add per_atom attributes to ComputedEntry
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"FYI @mattmcdermott @shyamd @awvio ",
"For certain, this requires new unittests.",
"> For certain, this requires new unittests.\r\n\r\nAbsolutely; tests added"
] | 2020-09-05T18:48:25
| 2021-07-15T14:31:38
|
2020-09-06T00:40:21Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
By analogy with `energy` and `energy_per_atom`, add `per_atom` attributes for other energies of a `ComputedEntry`:
* `correction_per_atom`
* `correction_uncertainty_per_atom`
* `uncorrected_energy_per_atom`
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1948/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1948/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1948",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1948",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1948.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1948.patch",
"merged_at": "2020-09-06T00:40:21Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1949
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1949/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1949/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1949/events
|
https://github.com/materialsproject/pymatgen/issues/1949
| 695,618,382
|
MDU6SXNzdWU2OTU2MTgzODI=
| 1,949
|
How I can get all the ternary metal oxides using MPRester.query without redundant element appear
|
{
"login": "finthon",
"id": 17920490,
"node_id": "MDQ6VXNlcjE3OTIwNDkw",
"avatar_url": "https://avatars.githubusercontent.com/u/17920490?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/finthon",
"html_url": "https://github.com/finthon",
"followers_url": "https://api.github.com/users/finthon/followers",
"following_url": "https://api.github.com/users/finthon/following{/other_user}",
"gists_url": "https://api.github.com/users/finthon/gists{/gist_id}",
"starred_url": "https://api.github.com/users/finthon/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/finthon/subscriptions",
"organizations_url": "https://api.github.com/users/finthon/orgs",
"repos_url": "https://api.github.com/users/finthon/repos",
"events_url": "https://api.github.com/users/finthon/events{/privacy}",
"received_events_url": "https://api.github.com/users/finthon/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-09-08T07:14:52
| 2020-09-08T10:15:05
|
2020-09-08T10:15:05Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I try to search all ternary metal oxides that include specific metal elements in the list:
TM = ['Sc', 'Ti', 'V', 'Cr', 'Mn', 'Fe', 'Co', 'Ni', 'Cu', 'Zn',
'Y', 'Zr', 'Nb', 'Mo', 'Tc', 'Ru', 'Rh', 'Pd', 'Ag', 'Cd',
'La', 'Ce', 'Hf', 'Ta', 'W', 'Re', 'Os', 'Ir', 'Pt', 'Au']
I am getting different results by running this codes:
####
with MPRester("My_API_key") as m:
data = m.query(criteria={"elements": {"$in": TM, "$all": ["O"]}, "nelements": 3}, properties=["pretty_formula", "material_id"])
for i in data:
print(i)
####
some materials, like "NaMnO2", "LiAgO2", are not what I want, because "Na", "Li" are not appeared in the TM list.
In addition, "nelements=2" can get all the binary metal oxides within TM list according to the explanation of the MPRester. query()
Does anybody know the proper mongo-style dict or solution?
I would appreciate any help on this.
Thanks
|
{
"login": "finthon",
"id": 17920490,
"node_id": "MDQ6VXNlcjE3OTIwNDkw",
"avatar_url": "https://avatars.githubusercontent.com/u/17920490?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/finthon",
"html_url": "https://github.com/finthon",
"followers_url": "https://api.github.com/users/finthon/followers",
"following_url": "https://api.github.com/users/finthon/following{/other_user}",
"gists_url": "https://api.github.com/users/finthon/gists{/gist_id}",
"starred_url": "https://api.github.com/users/finthon/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/finthon/subscriptions",
"organizations_url": "https://api.github.com/users/finthon/orgs",
"repos_url": "https://api.github.com/users/finthon/repos",
"events_url": "https://api.github.com/users/finthon/events{/privacy}",
"received_events_url": "https://api.github.com/users/finthon/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1949/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1949/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1950
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1950/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1950/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1950/events
|
https://github.com/materialsproject/pymatgen/issues/1950
| 697,769,058
|
MDU6SXNzdWU2OTc3NjkwNTg=
| 1,950
|
Faster analysis.local_env.VoronoiNN
|
{
"login": "googhgoo",
"id": 16490166,
"node_id": "MDQ6VXNlcjE2NDkwMTY2",
"avatar_url": "https://avatars.githubusercontent.com/u/16490166?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/googhgoo",
"html_url": "https://github.com/googhgoo",
"followers_url": "https://api.github.com/users/googhgoo/followers",
"following_url": "https://api.github.com/users/googhgoo/following{/other_user}",
"gists_url": "https://api.github.com/users/googhgoo/gists{/gist_id}",
"starred_url": "https://api.github.com/users/googhgoo/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/googhgoo/subscriptions",
"organizations_url": "https://api.github.com/users/googhgoo/orgs",
"repos_url": "https://api.github.com/users/googhgoo/repos",
"events_url": "https://api.github.com/users/googhgoo/events{/privacy}",
"received_events_url": "https://api.github.com/users/googhgoo/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks. I have pushed the changes this time. For future reference, the instructions for how to contribute to pymatgen are at https://pymatgen.org/contributing . For a small change such as this, one way is to directly edit within Github itself, which will create a PR.",
"Thank you, I will do that next time.."
] | 2020-09-10T10:56:27
| 2020-09-10T23:48:17
|
2020-09-10T16:31:28Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hello, I have been trying to use VornoiNN at large scale and noticed it does not scale well with large data set.
I noticed that trivial step in line 805 in pymatgen.analysis.local_env.VoronoiNN takes significant amount of time:
`sites = np.array(sites)[uniq_inds]`
The performance improves by about 10 fold over the MP data set if this is changed to:
`sites = [sites[i] for i in uniq_inds]`
I'm not sure how to make a pull request, so I just posted it here.
Thank you very much.. I hope this helps..
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1950/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1950/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1951
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1951/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1951/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1951/events
|
https://github.com/materialsproject/pymatgen/pull/1951
| 698,340,951
|
MDExOlB1bGxSZXF1ZXN0NDg0MTk0MjMx
| 1,951
|
address DeprecationWarning triggered by get_pourbaix_entries
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @rkingsbury. I noticed you commented out a test? Btw, NERSC is having load issues this morning so there are intermittent rester timeouts.",
"> Thanks @rkingsbury. I noticed you commented out a test? Btw, NERSC is having load issues this morning so there are intermittent rester timeouts.\r\n\r\nYes, that was a linting issue. Previously, the entire test was commented out except for the line that assigned a `Structure` to the variable s. The linter was complaining about the variable s being assigned to but never used. Since it seemed that the remainder of the test had already been disabled, I just commented out the whole thing.\r\n",
"There is another warning raised by `get_pourbaix_entries` that needs to be suppressed. I had previously merged a warnings filter to catch it, but it no longer appears to be working. It's not clear to me why not. Still working on that. Here is the relevant code in `matproj.py`\r\n\r\n```\r\n # suppress the warning about supplying the required energies; they will be calculated from the\r\n # entries we get from MPRester\r\n with warnings.catch_warnings():\r\n warnings.filterwarnings(\r\n \"ignore\",\r\n message=\"You did not provide the required O2 and H2O energies.\",\r\n )\r\n compat = MaterialsProjectAqueousCompatibility(solid_compat=solid_compat)\r\n ion_ref_entries = compat.process_entries(ion_ref_entries)\r\n ion_ref_pd = PhaseDiagram(ion_ref_entries)\r\n```\r\n\r\n",
"> There is another warning raised by `get_pourbaix_entries` that needs to be suppressed. I had previously merged a warnings filter to catch it, but it no longer appears to be working. It's not clear to me why not. Still working on that. Here is the relevant code in `matproj.py`\r\n> \r\n> ```\r\n> # suppress the warning about supplying the required energies; they will be calculated from the\r\n> # entries we get from MPRester\r\n> with warnings.catch_warnings():\r\n> warnings.filterwarnings(\r\n> \"ignore\",\r\n> message=\"You did not provide the required O2 and H2O energies.\",\r\n> )\r\n> compat = MaterialsProjectAqueousCompatibility(solid_compat=solid_compat)\r\n> ion_ref_entries = compat.process_entries(ion_ref_entries)\r\n> ion_ref_pd = PhaseDiagram(ion_ref_entries)\r\n> ```\r\n\r\nI can't for the life of me figure out why this warnings filter doesn't work. I've tried several varations (adding `category`, removing the context manager, etc.), but python always emits the warning regardless. I think this should be fixed but perhaps as part of another PR (my suspicion is there's a bigger issue here regarding how `warnings` behaves within pymatgen.\r\n\r\nIf you agree @mkhorton , this should be ready to go.",
"This is only raised if the user calls `get_pourbaix_entries` not on import? So I think it may be less of an issue anyway.",
"Did you try `warnings.simplefilter(\"ignore\")`within the `catch_warnings` block btw?",
"> This is only raised if the user calls `get_pourbaix_entries` not on import? So I think it may be less of an issue anyway.\r\n\r\nCorrect, it's only raised when calling `get_pourbaix_entries`",
"> Did you try `warnings.simplefilter(\"ignore\")`within the `catch_warnings` block btw?\r\n\r\nYes I did, but it has no effect.",
"Ok, I'll merge this as-is. Thanks for addressing this issue @rkingsbury "
] | 2020-09-10T18:33:19
| 2020-09-11T18:36:25
|
2020-09-11T18:34:35Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
The recent introduction of the `solid_compat` kwarg to `get_pourbaix_entries` began triggering a `DeprecationWarning` when the module was imported, because the kwarg defaulted to an instance of `MaterialsProjectCompatibility()`. This commit changes the interface slightly to avoid instantiating the class on import and therefore avoid triggering the `DeprecationWarning` until `get_pourbaix_entries` is actually called.
* `MaterialsProjectAqueousCompatibility` now accepts `None`, `Compatibility()`, or `Compatibility` as the `solid_compat`
* `MaterialsProjectAqueousCompatibility` now defaults to `solid_compat=MaterialsProjectCompatibility` (was previously `None`)
* allow `get_pourbaix_entries` to accept str as the `chemsys` argument (e.g. "Fe-Cr") for consistency with `get_entries_in_chemsys`
* pylint fixes and updates for `pourbaix_diagram.py`
TODO
* Fix the warnings suppression when `get_pourbaix_entries` calls `MaterialsProjectAqueousCompatibility` (can be addressed in a future PR; not changed by this one).
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1951/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1951/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1951",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1951",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1951.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1951.patch",
"merged_at": "2020-09-11T18:34:35Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1952
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1952/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1952/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1952/events
|
https://github.com/materialsproject/pymatgen/pull/1952
| 698,387,162
|
MDExOlB1bGxSZXF1ZXN0NDg0MjM1NzM3
| 1,952
|
Update MPScanRelaxSet with R2Scan
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @rkingsbury !"
] | 2020-09-10T19:15:38
| 2020-09-20T19:14:27
|
2020-09-19T21:42:37Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Migrate `MPScanRelaxSet` to the R2Scan metaGGA functional of Furness et al ([see paper](https://pubs.acs.org/doi/abs/10.1021/acs.jpclett.0c02405)).
* use `R2SCAN` variant of SCAN
* Update test files and tests with R2Scan results
* Update docstrings with description and citation for R2Scan
Iterative changes to specific INCAR parameters (e.g., ENCUT) may be made in the future, pending new benchmarking studies using R2Scan. The purpose of this PR is to allow devlopment tests in the atomate workflow to pass (see https://github.com/hackingmaterials/atomate/pull/454)
Note that R2Scan is not officially supported in VASP. Docstrings have been updated to explain this fact to users and instruct them to contact the manuscript's authors for source code if desired. The docstring also has instructions for substituting the original SCAN functional, which is officially supported.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1952/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1952/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1952",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1952",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1952.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1952.patch",
"merged_at": "2020-09-19T21:42:37Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1953
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1953/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1953/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1953/events
|
https://github.com/materialsproject/pymatgen/pull/1953
| 701,844,268
|
MDExOlB1bGxSZXF1ZXN0NDg3MjI1OTg4
| 1,953
|
Bandstructure bug fix and review of from_dict/as_dict
|
{
"login": "fraricci",
"id": 17427517,
"node_id": "MDQ6VXNlcjE3NDI3NTE3",
"avatar_url": "https://avatars.githubusercontent.com/u/17427517?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/fraricci",
"html_url": "https://github.com/fraricci",
"followers_url": "https://api.github.com/users/fraricci/followers",
"following_url": "https://api.github.com/users/fraricci/following{/other_user}",
"gists_url": "https://api.github.com/users/fraricci/gists{/gist_id}",
"starred_url": "https://api.github.com/users/fraricci/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/fraricci/subscriptions",
"organizations_url": "https://api.github.com/users/fraricci/orgs",
"repos_url": "https://api.github.com/users/fraricci/repos",
"events_url": "https://api.github.com/users/fraricci/events{/privacy}",
"received_events_url": "https://api.github.com/users/fraricci/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Many thanks @fraricci. For some context on the MPRester issue, some particularly large band structures fail due to a timeout on Cloudflare (which we cannot change). We use Cloudflare since it provides caching to reduce load on our servers and firewalls to prevent abuse. However, we're going to establish a second endpoint that bypasses Cloudflare for the purposes of these large files, but we need to purchase a new certificate first so this may take a week or so.",
"Hey @mkhorton, is that related with the issue I'm trying to solve with this issue?\r\n\r\n",
"It is not, separate issue",
"To add additional information, large band structures that may take more than 30 seconds to pull or process will cause a gateway timeout error. So if you're seeing that, that's why.",
"Hi @mkhorton, it seems that this issue #1941 is not solved yet.\r\nFetching bands for mp-867624 still returns the same error:\r\nsbs = rester.get_bandstructure_by_material_id('mp-867624')\r\nMPRestError: too many indices for array: array is 0-dimensional, but 2 were indexed.....\r\n\r\nIf I well remember, to solve this, the server-side version of pymatgen has to contain the patch implemented in this pull request.\r\nDo you have any updates?\r\nThanks!",
"Hi @fraricci, this could simply be because the MP website is still running an older pymatgen (currently, it's `2020.8.13`). We have a release pending so we can update the version of pymatgen then. Note that we're spending most of our development effort on the new API and will be looking for beta testers soon, you might prefer to use that."
] | 2020-09-15T11:19:33
| 2021-01-27T02:13:58
|
2020-10-10T01:17:02Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
This bug fix should solve the problem in issue #1941,
Example: mp-15077
Problem: for some reason some bandstructures on symmetry line in the MP
database contain the bands as dict(array). The BandStructureSymmLine.from_dict()
cannot handle this format. It creates the object but then the other methods
(e.g. is_metal()) fail because they expect an array.
Solution:
The BandStructure.from_dict() actually can handle this case.
So I took this opportunity to review the BandStructure.from_dict()/as_dict()
and make them work for the subclass BandStructureSymmLine as well.
So those methods were removed from the subclass.
Tests are running fine, but I'd like to do some other using data from the database.
I tried to get the json file for the problematic cases from the rester without success.
It seems there's a check of the data server side that fails (because there the bus is
not fixed) and it does not allow the download. I get the usual error instead.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1953/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1953/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1953",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1953",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1953.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1953.patch",
"merged_at": "2020-10-10T01:17:02Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1954
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1954/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1954/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1954/events
|
https://github.com/materialsproject/pymatgen/issues/1954
| 702,956,220
|
MDU6SXNzdWU3MDI5NTYyMjA=
| 1,954
|
pymatgen warnings are immune to filtering
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"The DeprecationWarning message can have unexpected side effects.\r\n\r\non Windows 10, I encountered a case where thousands of DeprecationWarning come out if executing a python library on a terminal: \r\n\r\nDeprecationWarning: species_and_occu is deprecated\r\nUse site.species instead. This will be deprecated with effect from pymatgen 2020.\r\n\r\nHowever, when the stdout/stderr of python is redirected to an input stream reader (buffered stream reader), \r\nno DeprecationWarning comes out but the execution of the python script just hangs there.\r\n(Manual modification of the library by using site.species instead of site.species_and_occu \r\nsolved the problem.)",
"The problem was not in pymatgen, but in monty. The `@deprecated` wrapper has a warnings.simplefilter(\"default\"), which overrides all other warnings filter calls. The reason this was done was because DeprecationWarning is ignored by default, and that is not the intent since we want users to change code to fix deprecated methods. \r\n\r\nIn the end, I discovered that DeprecationWarning is not the correct class to use. It should be FutureWarning. I have updated monty to 4.0.1 and the following code works:\r\n\r\n```python\r\nfrom pymatgen import MPRester\r\nimport warnings\r\nwarnings.filterwarnings(\"ignore\", category=FutureWarning)\r\nmpr = MPRester()\r\nprint(mpr.get_entries(\"mp-19017\"))\r\n```",
"Thank you @shyuep . So if I understand correctly we should still flag soon-to-be-deprecated code with `@deprecated`, but now it raises `FutureWarning` (which is filterable) and not `DeprecationWarning`?",
"Yes use @deprecated. FutureWarning is meant for end users and is always emitted unless filtered. DeprecationMessage is meant for developers and is never emitted unless Python is run in developmental mode. So the correct one to use is FutureWarning, which is now used in the wrapper. To force DeprecationWarg to show, I overrode the filter in the wrapper, which was a bad idea. ",
"Just a small update. I updated the `@deprecated` method in monty v4.0.2 to allow a developer to specify the category of the warning. It defaults to FutureWarning, which I believe would be the most common use case. But in case you want to deprecate methods meant only for developers, this is something to consider."
] | 2020-09-16T17:50:39
| 2020-10-12T17:23:36
|
2020-10-12T14:44:03Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Various warnings emitted by pymatgen appear to be immune to filtering by the normal python approaches like `warnings.simplefilter("ignore")` or the `catch_warnings` context manager.
**To Reproduce**
For example, instantiating `MPRester()` raises a `DeprecationWarning` due to the upcoming change in corrections classes. In some cases I want to suppress this warning; however the warning is still emitted by the following code blocks
```
import warnings
warnings.simplefilter("ignore")
with MPRester() as m:
m.get_entries("Al2O3")
```
```
import warnings
with warnings.catch_warnings():
with MPRester() as m:
m.get_entries("Al2O3")
```
```
import warnings
with warnings.catch_warnings():
warnings.filterwarnings(
"ignore",
message="MaterialsProjectCompatibility will be updated",
)
with MPRester() as m:
m.get_entries("Al2O3")
```
```
import warnings
with warnings.catch_warnings():
warnings.filterwarnings(
"ignore",
category=DeprecationWarning
)
with MPRester() as m:
m.get_entries("Al2O3")
```
**Expected behavior**
Standard Python warning filtering mechanisms should work with pymatgen warnings.
**Desktop (please complete the following information):**
- OS: Mac
- Version 2020.09.14
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1954/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1954/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1955
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1955/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1955/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1955/events
|
https://github.com/materialsproject/pymatgen/issues/1955
| 703,374,951
|
MDU6SXNzdWU3MDMzNzQ5NTE=
| 1,955
|
TransformedStructure.append_tranformation() breaks its history
|
{
"login": "koyamayukinori",
"id": 71375269,
"node_id": "MDQ6VXNlcjcxMzc1MjY5",
"avatar_url": "https://avatars.githubusercontent.com/u/71375269?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/koyamayukinori",
"html_url": "https://github.com/koyamayukinori",
"followers_url": "https://api.github.com/users/koyamayukinori/followers",
"following_url": "https://api.github.com/users/koyamayukinori/following{/other_user}",
"gists_url": "https://api.github.com/users/koyamayukinori/gists{/gist_id}",
"starred_url": "https://api.github.com/users/koyamayukinori/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/koyamayukinori/subscriptions",
"organizations_url": "https://api.github.com/users/koyamayukinori/orgs",
"repos_url": "https://api.github.com/users/koyamayukinori/repos",
"events_url": "https://api.github.com/users/koyamayukinori/events{/privacy}",
"received_events_url": "https://api.github.com/users/koyamayukinori/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-09-17T08:21:11
| 2023-08-13T16:33:45
|
2023-08-13T16:33:45Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
`TransformedStructure.append_transformation()` with `return_alternatives` breaks the structure one before the final.
**To Reproduce**
Here is an example code.
```
#!/usr/bin/env python
from pymatgen import Structure
from pymatgen.alchemy.materials import TransformedStructure
from pymatgen.transformations.advanced_transformations import EnumerateStructureTransformation
from pymatgen.transformations.standard_transformations import SupercellTransformation
a = 3.0
unitcell = Structure(
lattice=[[a, 0, 0], [0, a, 0], [0, 0, a]],
species=[{'Li': 0.25, 'Be': 0.75}],
coords=[[0, 0, 0]],
)
supercell = SupercellTransformation(2).apply_transformation(unitcell)
print('==== Before append_transformation(return_alternatives=...) ====')
transformed = TransformedStructure(supercell)
print(transformed.structures[0])
print()
print('==== After append_transformation(return_alternatives=...) ====')
alternatives = transformed.append_transformation(
EnumerateStructureTransformation(), return_alternatives=10)
print(transformed.structures[0])
print()
print('==== Final structure of the last alternatives ====')
print(alternatives[-1].final_structure)
print()
```
and the output is
```
==== Before append_transformation(return_alternatives=...) ====
Full Formula (Li2 Be6)
Reduced Formula: LiBe3
abc : 6.000000 6.000000 6.000000
angles: 90.000000 90.000000 90.000000
Sites (8)
# SP a b c
--- ------------------ --- --- ---
0 Li:0.250, Be:0.750 0 0 0
1 Li:0.250, Be:0.750 0 0 0.5
2 Li:0.250, Be:0.750 0 0.5 0
3 Li:0.250, Be:0.750 0 0.5 0.5
4 Li:0.250, Be:0.750 0.5 0 0
5 Li:0.250, Be:0.750 0.5 0 0.5
6 Li:0.250, Be:0.750 0.5 0.5 0
7 Li:0.250, Be:0.750 0.5 0.5 0.5
==== After append_transformation(return_alternatives=...) ====
Full Formula (Li2 Be6)
Reduced Formula: LiBe3
abc : 6.000000 6.000000 6.000000
angles: 90.000000 90.000000 90.000000
Sites (8)
# SP a b c
--- ---- --- --- ---
0 Li 0 0 0
1 Li 0.5 0.5 0.5
2 Be 0 0.5 0
3 Be 0.5 0 0
4 Be 0.5 0.5 0
5 Be 0 0 0.5
6 Be 0 0.5 0.5
7 Be 0.5 0 0.5
==== Final structure of the last alternatives ====
Full Formula (Li2 Be6)
Reduced Formula: LiBe3
abc : 6.000000 6.000000 6.000000
angles: 90.000000 90.000000 90.000000
Sites (8)
# SP a b c
--- ---- --- --- ---
0 Li 0 0 0
1 Li 0.5 0.5 0.5
2 Be 0 0.5 0
3 Be 0.5 0 0
4 Be 0.5 0.5 0
5 Be 0 0 0.5
6 Be 0 0.5 0.5
7 Be 0.5 0 0.5
```
`transformed.structures[0]` was overwritten by `alternatives[-1].final_structure`.
pymatgen/alchemy/materials.py:
```
127: hdict = actual_transformation.as_dict()
128: hdict["input_structure"] = input_structure
129: hdict["output_parameters"] = x
130: self.final_structure = s
131: d = self.as_dict()
132: d['history'].append(hdict)
133: d['final_structure'] = s.as_dict()
```
The line 130 looks unnecessary.
**Expected behavior**
Past structures should be unchanged by calling append_transformation().
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1955/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1955/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1956
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1956/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1956/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1956/events
|
https://github.com/materialsproject/pymatgen/issues/1956
| 703,383,818
|
MDU6SXNzdWU3MDMzODM4MTg=
| 1,956
|
TransformedStructure.__str__() destroys its history
|
{
"login": "koyamayukinori",
"id": 71375269,
"node_id": "MDQ6VXNlcjcxMzc1MjY5",
"avatar_url": "https://avatars.githubusercontent.com/u/71375269?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/koyamayukinori",
"html_url": "https://github.com/koyamayukinori",
"followers_url": "https://api.github.com/users/koyamayukinori/followers",
"following_url": "https://api.github.com/users/koyamayukinori/following{/other_user}",
"gists_url": "https://api.github.com/users/koyamayukinori/gists{/gist_id}",
"starred_url": "https://api.github.com/users/koyamayukinori/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/koyamayukinori/subscriptions",
"organizations_url": "https://api.github.com/users/koyamayukinori/orgs",
"repos_url": "https://api.github.com/users/koyamayukinori/repos",
"events_url": "https://api.github.com/users/koyamayukinori/events{/privacy}",
"received_events_url": "https://api.github.com/users/koyamayukinori/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-09-17T08:34:41
| 2023-08-13T16:33:46
|
2023-08-13T16:33:46Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
`TransformedStructure.__str__()` removes `input_structure` from its history.
**To Reproduce**
Here is an example code.
```
#!/usr/bin/env python
from pymatgen import Structure
from pymatgen.alchemy.materials import TransformedStructure
from pymatgen.transformations.standard_transformations import SupercellTransformation
a = 3.0
unitcell = Structure(
lattice=[[a, 0, 0], [0, a, 0], [0, 0, a]],
species=[{'Li': 0.25, 'Be': 0.75}],
coords=[[0, 0, 0]],
)
transformed = TransformedStructure(unitcell, [SupercellTransformation(2)])
print('Before str()')
print(transformed.history[0].keys())
_ = str(transformed)
print('After str()')
print(transformed.history[0].keys())
```
The output is
```
Before str()
dict_keys(['@module', '@class', '@version', 'scaling_matrix', 'input_structure', 'output_parameters'])
After str()
dict_keys(['@module', '@class', '@version', 'scaling_matrix', 'output_parameters'])
```
**Expected behavior**
`__str__()` should not change its attributes.
pymatgen/alchemy/materials.py:
```
219: for h in self.history:
220: h.pop('input_structure', None)
221: output.append(str(h))
```
At the line 220, `pop` should be `get`, shouldn't it?
**Desktop (please complete the following information):**
- OS: Linux
- Version: 2020.8.13
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1956/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1956/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1957
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1957/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1957/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1957/events
|
https://github.com/materialsproject/pymatgen/pull/1957
| 705,078,811
|
MDExOlB1bGxSZXF1ZXN0NDg5ODgxMTkz
| 1,957
|
Updates to diffusion analyzer
|
{
"login": "cajfisher",
"id": 66772456,
"node_id": "MDQ6VXNlcjY2NzcyNDU2",
"avatar_url": "https://avatars.githubusercontent.com/u/66772456?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/cajfisher",
"html_url": "https://github.com/cajfisher",
"followers_url": "https://api.github.com/users/cajfisher/followers",
"following_url": "https://api.github.com/users/cajfisher/following{/other_user}",
"gists_url": "https://api.github.com/users/cajfisher/gists{/gist_id}",
"starred_url": "https://api.github.com/users/cajfisher/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/cajfisher/subscriptions",
"organizations_url": "https://api.github.com/users/cajfisher/orgs",
"repos_url": "https://api.github.com/users/cajfisher/repos",
"events_url": "https://api.github.com/users/cajfisher/events{/privacy}",
"received_events_url": "https://api.github.com/users/cajfisher/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 2414169365,
"node_id": "MDU6TGFiZWwyNDE0MTY5MzY1",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/needs%20testing",
"name": "needs testing",
"color": "66ddab",
"default": false,
"description": "PRs that are not ready to merge due to lacking tests"
}
] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @cajfisher, this looks good but there are some linter errors (see failed tests):\r\n\r\n```\r\npymatgen/analysis/diffusion_analyzer.py:221:39: E261 at least two spaces before inline comment\r\npymatgen/analysis/diffusion_analyzer.py:222:18: E261 at least two spaces before inline comment\r\npymatgen/core/trajectory.py:65:121: E501 line too long (122 > 120 characters)\r\npymatgen/core/trajectory.py:69:121: E501 line too long (124 > 120 characters)\r\nError: Process completed with exit code 1.\r\n```",
"Many thanks @cajfisher this is great, documentation improvements much appreciated too.\r\n\r\nCould you add a quick test for the no framework atoms case? then it looks ready to merge.",
"Hi @cajfisher, I'm just checking in on the status of this PR. If a test is added and the merge conflict resolved, it looks ready to merge.",
"Dear Matthew,\r\n\r\nSorry for the delay – I fell ill for a while and have been desperately trying to catch up on work since then. I’ll sort out the remaining conflicts as soon as I have a chance.\r\n\r\nBest regards,\r\n\r\nCraig\r\n\r\nFrom: Matthew Horton <notifications@github.com>\r\nSent: Wednesday, December 2, 2020 1:39 AM\r\nTo: materialsproject/pymatgen <pymatgen@noreply.github.com>\r\nCc: クレイグ・フィッシャー <c_fisher@jfcc.or.jp>; Mention <mention@noreply.github.com>\r\nSubject: Re: [materialsproject/pymatgen] Updates to diffusion analyzer (#1957)\r\n\r\n\r\nHi @cajfisher<https://github.com/cajfisher>, I'm just checking in on the status of this PR. If a test is added and the merge conflict resolved, it looks ready to merge.\r\n\r\n—\r\nYou are receiving this because you were mentioned.\r\nReply to this email directly, view it on GitHub<https://github.com/materialsproject/pymatgen/pull/1957#issuecomment-736670077>, or unsubscribe<https://github.com/notifications/unsubscribe-auth/AP5N32GX3MWXD4CPAQXXUF3SSUL25ANCNFSM4RTRJWKQ>.\r\n",
"No worries at all, there's no harm if the PR sits here for a while."
] | 2020-09-20T09:44:26
| 2022-01-09T14:51:25
|
2022-01-09T14:51:25Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Fixes and changes to diffusion_analyzer.py:
* Now handles single-species systems correctly (specifically, when no framework atoms are present, calculation of framework drift is skipped)
* Various corrections to function descriptions (typo and grammar fixes, formatting and phrasing made consistent, etc.)
Nb. The original analysis tool assumes all species are ions, but in liquid and gaseous systems, for example, this is not necessarily the case. With a few minor modifications the functions can be made to handle more general msd calculations (i.e., neutral or charged species; obviously mscds are still only useful for charged species). This is the purpose of the proposed changes.
## No additional dependencies introduced
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1957/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1957/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1957",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1957",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1957.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1957.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1958
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1958/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1958/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1958/events
|
https://github.com/materialsproject/pymatgen/pull/1958
| 705,160,250
|
MDExOlB1bGxSZXF1ZXN0NDg5OTM5ODA1
| 1,958
|
Update nomad Information
|
{
"login": "wuxiaohua1011",
"id": 17909837,
"node_id": "MDQ6VXNlcjE3OTA5ODM3",
"avatar_url": "https://avatars.githubusercontent.com/u/17909837?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/wuxiaohua1011",
"html_url": "https://github.com/wuxiaohua1011",
"followers_url": "https://api.github.com/users/wuxiaohua1011/followers",
"following_url": "https://api.github.com/users/wuxiaohua1011/following{/other_user}",
"gists_url": "https://api.github.com/users/wuxiaohua1011/gists{/gist_id}",
"starred_url": "https://api.github.com/users/wuxiaohua1011/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/wuxiaohua1011/subscriptions",
"organizations_url": "https://api.github.com/users/wuxiaohua1011/orgs",
"repos_url": "https://api.github.com/users/wuxiaohua1011/repos",
"events_url": "https://api.github.com/users/wuxiaohua1011/events{/privacy}",
"received_events_url": "https://api.github.com/users/wuxiaohua1011/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@tschaume , please take a look. the MPRester should now be able to get a url like this:\r\n\r\nhttps://nomad-lab.eu/prod/rae/api/repo/?file_pattern=vasprun*&external_id=mp-739635\r\n\r\n\r\nthe runner script is as follows:\r\n```\r\nimport os\r\nmaterial_ids = [\"mp-32800\"]\r\ntask_types = [TaskType.GGA_OPT, TaskType.GGA_UNIFORM]\r\nfile_patterns = [\"vasprun*\"]\r\nwith MPRester(MP_API_KEY) as mpr:\r\n meta, urls = mpr.get_download_info(\r\n material_ids, task_types=task_types, file_patterns=file_patterns\r\n )\r\n\r\nprint(meta)\r\nprint(urls)\r\n```",
"Here's the python script to test it\r\n\r\n```\r\nfrom pymatgen.ext.matproj import MPRester, TaskType\r\n\r\nmaterial_ids = [\"mvc-2970\"]\r\ntask_types = [TaskType.GGA_OPT, TaskType.GGAU_UNIFORM]\r\nfile_patterns = [\"vasprun*\", \"OUTCAR*\"]\r\nwith MPRester(\"SECRET KEY HERE\") as mpr:\r\n meta, urls = mpr.get_download_info(material_ids, task_types, file_patterns)\r\n\r\nprint(urls)\r\n```",
"Hi @wuxiaohua1011, could you double-check this PR, there are two stray test files (one deleted, one added) that I think are unrelated to your changes.\r\n\r\nOtherwise, @tschaume, @wuxiaohua1011, is this ready to merge?",
"> Hi @wuxiaohua1011, could you double-check this PR, there are two stray test files (one deleted, one added) that I think are unrelated to your changes.\r\n> \r\n> Otherwise, @tschaume, @wuxiaohua1011, is this ready to merge?\r\n\r\nHi\r\nFor the test files, I am having a bit of problem with getting pass code style checkking...\r\nfor the file in `pymatgen/io/cp2k/__init__.py`, it doesn't have docstring, and pycode style is complaining.\r\n\r\n\r\n\r\n\r\n",
"I think I was able to solve that issue :) Please review it again @mkhorton ",
"Looks ok to me, thanks @wuxiaohua1011! @tschaume ?",
"\n[](https://coveralls.io/builds/40680635)\n\nCoverage decreased (-0.7%) to 83.086% when pulling **8dd90e407e9d658a159740c4e05d34b6dcc69ad0 on wuxiaohua1011:master** into **df22c1a523eee198ad6381dad585ac9b851dc9b3 on materialsproject:master**.\n",
"Thanks @wuxiaohua1011! Apologies for the slow merge, I was under the mistaken impression this was waiting on something."
] | 2020-09-20T18:29:45
| 2021-09-23T18:12:13
|
2021-09-23T18:12:13Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Update Nomad information
* Relink `get_download_info` in `ext/matproj.py` to the new URL: `https://nomad-lab.eu/prod/rae/api/repo/query?`
* Relink the `test_get_download_info` in the `ext/tests/matproj.py` to new URL
## TODO (if any)
This is a work in progress, I am still not able to run all the tests locally, requesting @tschaume 's help in that
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1958/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1958/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1958",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1958",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1958.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1958.patch",
"merged_at": "2021-09-23T18:12:13Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1959
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1959/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1959/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1959/events
|
https://github.com/materialsproject/pymatgen/pull/1959
| 705,302,634
|
MDExOlB1bGxSZXF1ZXN0NDkwMDUyNTE2
| 1,959
|
Added code to calculate NGX/NGY/NGZ NGXF/NGYF/NGZF
|
{
"login": "jmmshn",
"id": 14003693,
"node_id": "MDQ6VXNlcjE0MDAzNjkz",
"avatar_url": "https://avatars.githubusercontent.com/u/14003693?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jmmshn",
"html_url": "https://github.com/jmmshn",
"followers_url": "https://api.github.com/users/jmmshn/followers",
"following_url": "https://api.github.com/users/jmmshn/following{/other_user}",
"gists_url": "https://api.github.com/users/jmmshn/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jmmshn/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jmmshn/subscriptions",
"organizations_url": "https://api.github.com/users/jmmshn/orgs",
"repos_url": "https://api.github.com/users/jmmshn/repos",
"events_url": "https://api.github.com/users/jmmshn/events{/privacy}",
"received_events_url": "https://api.github.com/users/jmmshn/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@mkhorton for the tests I just grabbed a bunch of random calculations and check that the grids matched.\r\nI actually check vs all 100K + calculations in my DB but only uploaded 300 to keep the file size small.",
"Thanks @jmmshn! Interesting you have to do a prime factorization with this too",
"This was amusing for me as well. It's because of how VASP interacts with FFTs.",
"Thanks @jmmshn !"
] | 2020-09-21T05:46:06
| 2020-09-25T16:32:37
|
2020-09-25T16:32:36Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Calculate the grid sizes from vasp inputs
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1959/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1959/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1959",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1959",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1959.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1959.patch",
"merged_at": "2020-09-25T16:32:36Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1960
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1960/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1960/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1960/events
|
https://github.com/materialsproject/pymatgen/pull/1960
| 705,983,821
|
MDExOlB1bGxSZXF1ZXN0NDkwNjE0OTEz
| 1,960
|
populate "Generated by" in EnergyAdjustment generated by MP2020 and fix serialization bug
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Unless you want to communicate that people should be able to change `adj_per_atom` after instatiation, you probably want to set it with `self._adj_per_atom` (similar to eg `self._value` previously), likewise with `uncertainty`. Otherwise this looks good."
] | 2020-09-21T23:41:05
| 2020-09-29T17:49:45
|
2020-09-29T15:33:01Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Populate the `cls` kwarg when creating `EnergyAdjustment` within `MaterialsProject2020Compatibility`. This information will be helpful in tracking the provenance of energy adjustments.
Now that the `cls` kwarg is populated as intended, executing `ComputedEntry.energy_adjustments` on an entry that has been processed with the MP2020 correction scheme should generate output similar to
```
[CompositionEnergyAdjustment:
Name: MP2020 anion correction (oxide)
Value: -5.920 eV
Uncertainty: 0.014 eV
Description: Composition-based energy adjustment (-0.740 eV/atom x 8.0 atoms)
Generated by: MaterialsProject2020Compatibility,
CompositionEnergyAdjustment:
Name: MP2020 GGA/GGA+U mixing correction (Fe)
Value: -8.728 eV
Uncertainty: 0.036 eV
Description: Composition-based energy adjustment (-2.182 eV/atom x 4.0 atoms)
Generated by: MaterialsProject2020Compatibility]
```
Whereas before, the "Generated by:" line read `None`
In addition, I discovered that several classes inherited from `EnergyAdjustment` (including `CompositionEnergyAdjustment`, which is used by MP2020) were not serializing properly. This was because the `description` attribute was being modified during init and because some kwargs were not stored as attributes. This PR also fixes those problems.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1960/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1960/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1960",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1960",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1960.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1960.patch",
"merged_at": "2020-09-29T15:33:01Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1961
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1961/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1961/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1961/events
|
https://github.com/materialsproject/pymatgen/issues/1961
| 707,051,301
|
MDU6SXNzdWU3MDcwNTEzMDE=
| 1,961
|
diffusion_analyzer assumes more than one species is present in model
|
{
"login": "cajfisher",
"id": 66772456,
"node_id": "MDQ6VXNlcjY2NzcyNDU2",
"avatar_url": "https://avatars.githubusercontent.com/u/66772456?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/cajfisher",
"html_url": "https://github.com/cajfisher",
"followers_url": "https://api.github.com/users/cajfisher/followers",
"following_url": "https://api.github.com/users/cajfisher/following{/other_user}",
"gists_url": "https://api.github.com/users/cajfisher/gists{/gist_id}",
"starred_url": "https://api.github.com/users/cajfisher/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/cajfisher/subscriptions",
"organizations_url": "https://api.github.com/users/cajfisher/orgs",
"repos_url": "https://api.github.com/users/cajfisher/repos",
"events_url": "https://api.github.com/users/cajfisher/events{/privacy}",
"received_events_url": "https://api.github.com/users/cajfisher/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-09-23T05:07:23
| 2023-08-13T16:33:46
|
2023-08-13T16:33:46Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
MSD calculations fail when only one species is present in the system, as it is assumed all systems are ionic with framework atoms present in addition to the species of interest.
**To Reproduce**
Steps to reproduce the behavior:
1. Run DiffusionAnalzyer.from_structures using structures containing only one species.
2. See error
**Expected behavior**
The MSD should be calculated for the single species with zero drift correction from the (non-existent) framework.
**Desktop (please complete the following information):**
- OS: Fedora 31 (Linux)
**Additional context**
Simple liquid and melt systems often contain only one species. It would be good to generalize DiffusionAnalyzer to work equally well for non-charged atoms and charged atoms (ions) as well as single component and multicomponent systems.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1961/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1961/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1962
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1962/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1962/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1962/events
|
https://github.com/materialsproject/pymatgen/issues/1962
| 708,606,338
|
MDU6SXNzdWU3MDg2MDYzMzg=
| 1,962
|
Problems with terminology
|
{
"login": "cajfisher",
"id": 66772456,
"node_id": "MDQ6VXNlcjY2NzcyNDU2",
"avatar_url": "https://avatars.githubusercontent.com/u/66772456?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/cajfisher",
"html_url": "https://github.com/cajfisher",
"followers_url": "https://api.github.com/users/cajfisher/followers",
"following_url": "https://api.github.com/users/cajfisher/following{/other_user}",
"gists_url": "https://api.github.com/users/cajfisher/gists{/gist_id}",
"starred_url": "https://api.github.com/users/cajfisher/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/cajfisher/subscriptions",
"organizations_url": "https://api.github.com/users/cajfisher/orgs",
"repos_url": "https://api.github.com/users/cajfisher/repos",
"events_url": "https://api.github.com/users/cajfisher/events{/privacy}",
"received_events_url": "https://api.github.com/users/cajfisher/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @cajfisher, your comment is well-received but this is not likely to change at this point. The decision to use the term `Specie` is not made for grammatical reasons but for programmatical reasons: it is to unambiguously declare that this refers to a *single* species, vs., say, a list of multiple species. In terms of a programming language, only a single definition can be used (e.g., we could not use the term `Species` to refer to both a single `Species` object and an object that represents multiple `Species`), and it is very important that we unambiguously declare that this refers to the singular form. I appreciate this is grammatically frustrating however so my apologies for that :-)",
"Thanks for your quick response and explanation. I understand the need to distinguish between single and multiple species in the coding. The main problem (as I see it) is that this has bled over into the \"natural\" English used in the manual and module/function descriptions. For example, in the pymatgen.core.sites module under \"species\" it says \"ii. An element / specie specified either as a string...\", even though \"species\" is meant to contain multiple species; in the pymatgen.analysis.diffusion_analyzer module: \"Specie to calculate diffusivity for as a String\" (why am I calculating the diffusivity of a python variable?); and in the pymatgen.alchemy.materials module: \"when the specie to replace...\", but elsewhere \"with a single species\". In other words, there appears to be a lack of consistency. Tidying this up (although tedious!) should improve readability and reduce confusion, thereby speeding up both the uptake of pymatgen and future code development (easier bug checking, and so on). That's my (non-inflationary) two-cents worth, anyway.\r\nJust for reference, in other codes I've worked on, variable names such as \"species_type\" and \"atom_type\" are commonly used to overcome/avoid this tricky linguistic problem (although you have probably already considered these). Cheers!",
"I think this can be resolved easily. Let's rename it as Ion, and put Specie as a trivial subclass to maintain backwards compatibility. I do not think it is going to cause any issues?\n\nSpecies_type or Atom_type is far too long a name for a very commonly used class.",
"Looking at the definition of the Specie class in pymatgen.core.periodic_table, this seems to be a sensible choice to me (as an outsider looking in).",
"Ok, there is a complication (slight one). There is actually an ion class that is used defined in pymatgen.core.ion. The difference is that the current Ion is Composition(which can mean 1 or many atoms)+Oxidation State. This Ion is used just in the Pourbaix diagram construction.\r\n\r\nThere are a few potential solutions in order of effort:\r\n1. Do nothing.\r\n2. Rename Specie to Species and have Specie be a backwards compatible name.\r\n3. Rename Specie to Ion and Ion to PolyIon. This will break backwards compatibility for sure for the current PourbaixDiagrams but will Specie can still be made backwards compatible by mapping to Ion.\r\n\r\n@mkhorton I am leaning towards option 2. It is ~ the first option in terms of effort. It solves the grammar issue. And for existing users, it is invisible. What do you think?",
"I have implemented the change in https://github.com/materialsproject/pymatgen/tree/species_rename branch. Seems to be working fine. There is one slight complication in that Site.specie is used to refer to a single species in ordered sites. For that, I don't think we can do a rename since the programmatic choice that Matt mentioned (single vs plural variable names) unfortunately applies here. I have added a statement on the deliberate non-grammatical naming choice in the documentation for that property.",
"I don't think `Ion` is a good fit for `Specie`, because `Specie` can contain `spin` instead of `oxidation_state`, i.e. it's not always an ion, and as you say ion is already used in Pourbaix regardless",
"My primary motivation *against* changing it is the backwards compatibility argument, since `Specie` has been in use for a long time at this point. If we wanted to break backwards compatibility, I would have it subclass `Element` instead, and allow a more uniform interface between `Element` and `Specie`",
"Another alternative label is \"Particle\". A few more characters than \"Ion\" and a bit more general in meaning, but it seems to cover the properties entailed by \"Specie\" reasonably well, assuming it isn't used elsewhere. Also, if you did want to break backwards compatibility, Matthew's suggestion sounds good from an OOP point of view. ",
"Thanks for the feedback, it is now `Species` 🎉"
] | 2020-09-25T03:10:08
| 2020-09-30T03:52:48
|
2020-09-25T03:31:41Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
A number of functions and their descriptions (as well as the online manual) use the incorrect term "specie" to describe a single species. The singular and plural forms are both "species". ("Specie" refers to the physical form of money, usually coins.)
See, e.g., https://brians.wsu.edu/2016/05/31/specie/
https://thetrcompany.com/en/difference-specie-species/
https://grammarist.com/usage/species/
Although the above explanations are concerned primarily with species as living organisms, the same applies for other usages:
https://www.merriam-webster.com/dictionary/species (1e)
It would be good if this error could be fixed at some stage to minimize confusion and improve readability and consistency across the various modules.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1962/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1962/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1963
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1963/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1963/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1963/events
|
https://github.com/materialsproject/pymatgen/pull/1963
| 709,074,577
|
MDExOlB1bGxSZXF1ZXN0NDkzMTkwOTYz
| 1,963
|
Species rename
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@mkhorton Pls review.",
"I'm fine merging this since the original `Specie` and `DummySpecie` are not deprecated but will remain as simple subclasses, so net disruption to the end user should be minimal.\r\n\r\nHowever, I'm mixed overall since it seems like the change doesn't produce net benefit, but will result in e.g. schemas becoming more complicated, teaching materials becoming out of date. I'm more of a [lingustic descriptivist](https://dictionary.cambridge.org/dictionary/english/descriptivism) than a prescriptivist, so although I concede the original usage has always been technically incorrect, that doesn't necessarily mean it's *wrong*."
] | 2020-09-25T16:21:23
| 2020-09-29T15:32:17
|
2020-09-29T15:32:17Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
* Rename Specie and DummySpecie to Species and DummySpecies
* Specie and DummySpecie is allowed as a mapping for backwards compatibility.
* Corresponding changes to doc where possible.
* Pylint does not pass because of refactoring in chemenv and defects and abinit packages. These are unrelated to the changes made.
## Additional dependencies introduced (if any)
## TODO (if any)
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [X ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [ X] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [X] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [X] Tests have been added for any new functionality or bug fixes.
- [X] All existing tests pass.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1963/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1963/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1963",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1963",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1963.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1963.patch",
"merged_at": "2020-09-29T15:32:17Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1964
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1964/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1964/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1964/events
|
https://github.com/materialsproject/pymatgen/issues/1964
| 709,673,489
|
MDU6SXNzdWU3MDk2NzM0ODk=
| 1,964
|
Faile to run: pip install -r requirements-optional.txt
|
{
"login": "hongyi-zhao",
"id": 11155854,
"node_id": "MDQ6VXNlcjExMTU1ODU0",
"avatar_url": "https://avatars.githubusercontent.com/u/11155854?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/hongyi-zhao",
"html_url": "https://github.com/hongyi-zhao",
"followers_url": "https://api.github.com/users/hongyi-zhao/followers",
"following_url": "https://api.github.com/users/hongyi-zhao/following{/other_user}",
"gists_url": "https://api.github.com/users/hongyi-zhao/gists{/gist_id}",
"starred_url": "https://api.github.com/users/hongyi-zhao/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/hongyi-zhao/subscriptions",
"organizations_url": "https://api.github.com/users/hongyi-zhao/orgs",
"repos_url": "https://api.github.com/users/hongyi-zhao/repos",
"events_url": "https://api.github.com/users/hongyi-zhao/events{/privacy}",
"received_events_url": "https://api.github.com/users/hongyi-zhao/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @hongyi-zhao, this is a dependency for the optional requirement `BoltzTrap2`, if you don't want to use BoltzTrap2 functionality you don't need to install this. To get it to work, you will want to install `cmake`, e.g. `snap install cmake` or `apt-get install cmake`",
"Thanks a lot. The following command solves the problem:\r\n\r\n`$ sudo apt-get install cmake`\r\n"
] | 2020-09-27T05:41:24
| 2020-09-27T05:45:21
|
2020-09-27T05:45:21Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hi,
The environment is Ubuntu 20.04, python 3.8.3 managed by pyenv, and latest git master version of pymatgen. See following for more info:
```
$ python --version
Python 3.8.3
$ pip install -r requirements-optional.txt
[...]
ERROR: Command errored out with exit status 1:
command: /home/werner/.pyenv/versions/3.8.3/envs/datasci/bin/python3.8 -u -c 'import sys, setuptools, tokenize; sys.argv[0] = '"'"'/tmp/pip-install-elz3n1in/boltztrap2/setup.py'"'"'; __file__='"'"'/tmp/pip-install-elz3n1in/boltztrap2/setup.py'"'"';f=getattr(tokenize, '"'"'open'"'"', open)(__file__);code=f.read().replace('"'"'\r\n'"'"', '"'"'\n'"'"');f.close();exec(compile(code, __file__, '"'"'exec'"'"'))' install --record /tmp/pip-record-ed6pmgp6/install-record.txt --single-version-externally-managed --compile --install-headers /home/werner/.pyenv/versions/3.8.3/envs/datasci/include/site/python3.8/BoltzTraP2
cwd: /tmp/pip-install-elz3n1in/boltztrap2/
Complete output (25 lines):
running install
running build
running build_py
creating build
creating build/lib.linux-x86_64-3.8
creating build/lib.linux-x86_64-3.8/BoltzTraP2
copying BoltzTraP2/fermisurface.py -> build/lib.linux-x86_64-3.8/BoltzTraP2
copying BoltzTraP2/__init__.py -> build/lib.linux-x86_64-3.8/BoltzTraP2
copying BoltzTraP2/io.py -> build/lib.linux-x86_64-3.8/BoltzTraP2
copying BoltzTraP2/bandlib.py -> build/lib.linux-x86_64-3.8/BoltzTraP2
copying BoltzTraP2/misc.py -> build/lib.linux-x86_64-3.8/BoltzTraP2
copying BoltzTraP2/units.py -> build/lib.linux-x86_64-3.8/BoltzTraP2
copying BoltzTraP2/dft.py -> build/lib.linux-x86_64-3.8/BoltzTraP2
copying BoltzTraP2/interface.py -> build/lib.linux-x86_64-3.8/BoltzTraP2
copying BoltzTraP2/serialization.py -> build/lib.linux-x86_64-3.8/BoltzTraP2
copying BoltzTraP2/fite.py -> build/lib.linux-x86_64-3.8/BoltzTraP2
copying BoltzTraP2/version.py -> build/lib.linux-x86_64-3.8/BoltzTraP2
copying BoltzTraP2/fd.py -> build/lib.linux-x86_64-3.8/BoltzTraP2
creating build/lib.linux-x86_64-3.8/BoltzTraP2/sphere
copying BoltzTraP2/sphere/__init__.py -> build/lib.linux-x86_64-3.8/BoltzTraP2/sphere
running build_ext
running build_spglib
About to create a new build directory for spglib
About to run 'cmake' for spglib
error: [Errno 2] No such file or directory: 'cmake'
----------------------------------------
ERROR: Command errored out with exit status 1: /home/werner/.pyenv/versions/3.8.3/envs/datasci/bin/python3.8 -u -c 'import sys, setuptools, tokenize; sys.argv[0] = '"'"'/tmp/pip-install-elz3n1in/boltztrap2/setup.py'"'"'; __file__='"'"'/tmp/pip-install-elz3n1in/boltztrap2/setup.py'"'"';f=getattr(tokenize, '"'"'open'"'"', open)(__file__);code=f.read().replace('"'"'\r\n'"'"', '"'"'\n'"'"');f.close();exec(compile(code, __file__, '"'"'exec'"'"'))' install --record /tmp/pip-record-ed6pmgp6/install-record.txt --single-version-externally-managed --compile --install-headers /home/werner/.pyenv/versions/3.8.3/envs/datasci/include/site/python3.8/BoltzTraP2 Check the logs for full command output.
```
Any hints for this problem?
Regards,
HY
|
{
"login": "hongyi-zhao",
"id": 11155854,
"node_id": "MDQ6VXNlcjExMTU1ODU0",
"avatar_url": "https://avatars.githubusercontent.com/u/11155854?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/hongyi-zhao",
"html_url": "https://github.com/hongyi-zhao",
"followers_url": "https://api.github.com/users/hongyi-zhao/followers",
"following_url": "https://api.github.com/users/hongyi-zhao/following{/other_user}",
"gists_url": "https://api.github.com/users/hongyi-zhao/gists{/gist_id}",
"starred_url": "https://api.github.com/users/hongyi-zhao/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/hongyi-zhao/subscriptions",
"organizations_url": "https://api.github.com/users/hongyi-zhao/orgs",
"repos_url": "https://api.github.com/users/hongyi-zhao/repos",
"events_url": "https://api.github.com/users/hongyi-zhao/events{/privacy}",
"received_events_url": "https://api.github.com/users/hongyi-zhao/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1964/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1964/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1965
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1965/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1965/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1965/events
|
https://github.com/materialsproject/pymatgen/issues/1965
| 709,674,867
|
MDU6SXNzdWU3MDk2NzQ4Njc=
| 1,965
|
The error description in the document for installation: pip install pymatgen[extra]
|
{
"login": "hongyi-zhao",
"id": 11155854,
"node_id": "MDQ6VXNlcjExMTU1ODU0",
"avatar_url": "https://avatars.githubusercontent.com/u/11155854?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/hongyi-zhao",
"html_url": "https://github.com/hongyi-zhao",
"followers_url": "https://api.github.com/users/hongyi-zhao/followers",
"following_url": "https://api.github.com/users/hongyi-zhao/following{/other_user}",
"gists_url": "https://api.github.com/users/hongyi-zhao/gists{/gist_id}",
"starred_url": "https://api.github.com/users/hongyi-zhao/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/hongyi-zhao/subscriptions",
"organizations_url": "https://api.github.com/users/hongyi-zhao/orgs",
"repos_url": "https://api.github.com/users/hongyi-zhao/repos",
"events_url": "https://api.github.com/users/hongyi-zhao/events{/privacy}",
"received_events_url": "https://api.github.com/users/hongyi-zhao/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Here, \"extra\" is a placeholder for the specific \"extra\" that you require.\r\n\r\nCurrently, you can choose from \"provenance\", \"ase\", \"vis\" or \"abinit\" (see [here](https://github.com/materialsproject/pymatgen/blob/4b406abb589b9df1b7fbc3ba52e375f6fd3ed37e/setup.py#L115)).\r\n\r\nAs with the BoltzTrap2 dependencies, extras are optional. Most users do not need them.",
"> Here, \"extra\" is a placeholder for the specific \"extra\" that you require.\r\n> \r\n> Currently, you can choose from \"provenance\", \"ase\", \"vis\" or \"abinit\" (see [here](https://github.com/materialsproject/pymatgen/blob/4b406abb589b9df1b7fbc3ba52e375f6fd3ed37e/setup.py#L115)).\r\n\r\nDo you mean any of them will do the trick, i.e., running one of the following:\r\n\r\n```\r\n$ pip install .[provenance]\r\n$ pip install .[ase]\r\n$ pip install .[vis]\r\n$ pip install .[abinit]\r\n```\r\n> \r\n> As with the BoltzTrap2 dependencies, extras are optional. Most users do not need them.\r\n\r\nI use the following command to install all possible dependencies:\r\n \r\n`$ pip install -r requirements.txt -r requirements-dev.txt -r requirements-optional.txt`\r\n\r\n",
"The recommended way to install pymatgen is either:\r\n\r\n`pip install pymatgen` (or, if you have a desired extra, `pip install pymatgen[provenance]` etc.)\r\n\r\nor\r\n\r\n`conda install -c conda-forge pymatgen`\r\n\r\nIf you are doing development, then you can manually install those dependencies yourself, such as doing `pip install -r requirements.txt`, but in general you only need to install dependencies for functionality you want to use. If you're not familiar with this additional functionality or you're not doing development using it, it's safe not to install it.",
"> `pip install pymatgen` (or, if you have a desired extra, `pip install pymatgen[provenance]` etc.)\r\n\r\nFor my case, I run the command immediately under the local git repo root of pymatgen, so I think it should be invoked like the forms I posted previously.\r\n\r\n> If you are doing development, then you can manually install those dependencies yourself, such as doing `pip install -r requirements.txt`, \r\n\r\nBut the **requirements.txt** doesn't include any of these [packages](https://github.com/materialsproject/pymatgen/blob/4b406abb589b9df1b7fbc3ba52e375f6fd3ed37e/setup.py#L115).",
"Yes, that's correct, because these are optional requirements (hence \"extras\"), they are in `requirements-optional.txt`\r\n\r\nPerhaps it would be useful to ask what you're intending to use pymatgen for, or why you're installing it? Typically, if you're new to pymatgen, I would recommend following the `pip install` or `conda install` instructions above, and not performing `git clone` etc.\r\n\r\nThe important thing is to understand that the `setup.py` and `requirements*.txt` files both describe similar things but with different purposes. The `setup.py` is generally used for installing, while the `requirements*.txt` give you specific known versions of dependencies, and can be useful for development. This structure is fairly typical for a Python code, and not specific to pymatgen itself.",
"> Yes, that's correct, because these are optional requirements (hence \"extras\"), they are in `requirements-optional.txt`\r\n\r\nBut even the **requirements-optional.txt** doesn't include all the dependencies we discussed here. \r\n",
"I'm sorry, I'm not sure I understand the problem you're having. Pymatgen will work fine even if optional dependencies are not installed. If you find that you want to use a feature for which you don't have the requisite dependency, you will get a message and you can install it.\r\n\r\nThe `requirements-optional.txt` *does* have these optional dependencies, e.g. it has `pybtex` (the \"provenance\" extra), it has `ase` (the \"ase\" extra), it has `netcdf4` (the \"abinit\" extra), it does not have `vtk` but this is because vtk can be difficult to install on certain systems and is only used for the visualization package which is being deprecated.",
"> The `requirements-optional.txt` _does_ have these optional dependencies, e.g. it has `pybtex` (the \"provenance\" extra), it has `ase` (the \"ase\" extra), it has `netcdf4` (the \"abinit\" extra), it does not have `vtk` but this is because vtk can be difficult to install on certain systems and is only used for the visualization package which is being deprecated.\r\n\r\nThanks a lot, got it. Another question: do you mean **vtk** is being deprecated by pymatgen?",
"> The recommended way to install pymatgen is either:\r\n> \r\n> `pip install pymatgen` (or, if you have a desired extra, `pip install pymatgen[provenance]` etc.)\r\n\r\nThere is still another question I can't figure out, see following for more info.\r\n\r\nThe following codes are the content of [extras_require section of setup.py](https://github.com/materialsproject/pymatgen/blob/4b406abb589b9df1b7fbc3ba52e375f6fd3ed37e/setup.py#L115). \r\n```\r\n extras_require={\r\n \"provenance\": [\"pybtex\"],\r\n \"ase\": [\"ase>=3.3\"],\r\n \"vis\": [\"vtk>=6.0.0\"],\r\n \"abinit\": [\"netcdf4\"],\r\n ':python_version < \"3.7\"': [\r\n \"dataclasses>=0.6\",\r\n ]},\r\n```\r\n So, if I want to install all of these extra dependencies with one run of pip, I find that only the following command can do the trick:\r\n\r\n```\r\n$ pip install .[provenance,ase,vis,abinit] |grep -i warning\r\n$\r\n```\r\nAnd all the following commands failed:\r\n```\r\n$ pip install .[extras_require] |grep -i warning\r\n WARNING: pymatgen 2020.9.14 does not provide the extra 'extras_require'\r\n$ pip install .[extras] |grep -i warning\r\n WARNING: pymatgen 2020.9.14 does not provide the extra 'extras'\r\n$ pip install .[extra] |grep -i warning\r\n WARNING: pymatgen 2020.9.14 does not provide the extra 'extra'\r\n```\r\n\r\n\r\n\r\n",
"The correct pip install command would be `pip install pymatgen[provenance]` etc.",
"> The correct pip install command would be `pip install pymatgen[provenance]` etc.\r\n\r\nI'm still confused why can't I combine them into one like the following, considering that the usage abides with the pip's specification:\r\n\r\n`$ pip install pymatgen[provenance,ase,vis,abinit] `\r\n\r\nor run as following if we are directly in the pymatgen's top level subdirectory:\r\n\r\n`$ pip install .[provenance,ase,vis,abinit] `\r\n\r\nSee [here](https://github.com/pypa/pip/issues/8940) for the discussion relevant to this problem."
] | 2020-09-27T05:54:40
| 2020-09-30T09:31:09
|
2020-09-30T03:51:21Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hi,
The document told the following:
```
$ egrep -inR 'pymatgen\[extra\]' .
./docs_rst/introduction.rst:182: pip install pymatgen[extra]
```
But the output of the following installation command is not coherent with the above description:
```
$ pip install .[extra]
Looking in indexes: https://mirrors.aliyun.com/pypi/simple/
Processing /home/werner/Public/hpc/tools/materialsproject/pymatgen.git
WARNING: pymatgen 2020.9.14 does not provide the extra 'extra'
[...]
```
Any hints for this problem?
Regards,
HY
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1965/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1965/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1966
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1966/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1966/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1966/events
|
https://github.com/materialsproject/pymatgen/issues/1966
| 710,262,553
|
MDU6SXNzdWU3MTAyNjI1NTM=
| 1,966
|
pymatgen==2020.9.14: PDPlotter cannot write image (AttributeError)
|
{
"login": "LuciusV",
"id": 11925386,
"node_id": "MDQ6VXNlcjExOTI1Mzg2",
"avatar_url": "https://avatars.githubusercontent.com/u/11925386?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/LuciusV",
"html_url": "https://github.com/LuciusV",
"followers_url": "https://api.github.com/users/LuciusV/followers",
"following_url": "https://api.github.com/users/LuciusV/following{/other_user}",
"gists_url": "https://api.github.com/users/LuciusV/gists{/gist_id}",
"starred_url": "https://api.github.com/users/LuciusV/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/LuciusV/subscriptions",
"organizations_url": "https://api.github.com/users/LuciusV/orgs",
"repos_url": "https://api.github.com/users/LuciusV/repos",
"events_url": "https://api.github.com/users/LuciusV/events{/privacy}",
"received_events_url": "https://api.github.com/users/LuciusV/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@LuciusV My apologies for just seeing this now! Yes, the new PDPlotter (with Plotly backend) does not currently work with the `write_image()` function; that's my fault for missing that implementation when I wrote the code!\r\n\r\nUnfortunately, Plotly is quite difficult when it comes to saving high-quality figures, but I've identified a workaround for now. (using some PhaseDiagram object named `pd`):\r\n\r\n```\r\nfig = PDPlotter(pd).get_plot()\r\nconfig = {\r\n 'toImageButtonOptions': {\r\n 'format': 'svg', \r\n }\r\n}\r\n\r\nfig.write_html(\"my_img.html\", config=config)\r\n```\r\nThis changes the image save button to be SVG, then creates an HTML file that you can open in your browser and adjust to your liking before clicking save. I find this to be the most convenient method as of now.\r\n\r\nI'll look into fixing the `write_image()` function next week... I know I've had some issues with some of the Plotly saving methods in the past.",
"Yes, to write images you need to have `orca` installed (typically) and use `plotly.io`, writing to html is another way to do it. The former solution is probably the better one, though that will require the extra dependency for people who want to write images out with the plotly backend.",
"This is very good workaround for my case, thank you very much!",
"@mattmcdermott For me the error still exists, but I found a way that works for me.\r\nI modified the PDPlotter.write_image function to:\r\n\r\n```\r\ndef write_image(self, stream, image_format=\"svg\", **kwargs):\r\n \r\n plot = self.get_plot(**kwargs)\r\n plot.write_image(stream, format=image_format, width=1073, height=700, scale=1)\r\n ```\r\nIt uses the function plotly.io.write_image() instead of plt.savefig().",
"@TholeoR This has been addressed with #3032. Apologies for the delay!"
] | 2020-09-28T13:24:26
| 2023-06-02T23:37:07
|
2020-10-30T19:55:25Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
If I try to .write_image() of ternary phase diagram with default backend, it fails.
**To Reproduce**
Steps to reproduce the behavior:
In jupyter notebook:
1. Make some 3-component PhaseDiagram
2. Make PDPlotter on this Phasediagram
3. Call write_image('something.svg'). PNG also fails.
4. it fails (AttributeError: 'Figure' object has no attribute 'gcf') apparently expecting matplotlib object...
**Expected behavior**
SVG Image saved normally
**Desktop (please complete the following information):**
- OS: Linux
- Version: 2020.9.14
**Additional context**
I know that I can save png from resulting plotly plot, but I need svg :(
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1966/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1966/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1967
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1967/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1967/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1967/events
|
https://github.com/materialsproject/pymatgen/pull/1967
| 710,791,348
|
MDExOlB1bGxSZXF1ZXN0NDk0NTc4NzIw
| 1,967
|
Cube parsing for Bader Analysis
|
{
"login": "nwinner",
"id": 8825901,
"node_id": "MDQ6VXNlcjg4MjU5MDE=",
"avatar_url": "https://avatars.githubusercontent.com/u/8825901?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/nwinner",
"html_url": "https://github.com/nwinner",
"followers_url": "https://api.github.com/users/nwinner/followers",
"following_url": "https://api.github.com/users/nwinner/following{/other_user}",
"gists_url": "https://api.github.com/users/nwinner/gists{/gist_id}",
"starred_url": "https://api.github.com/users/nwinner/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/nwinner/subscriptions",
"organizations_url": "https://api.github.com/users/nwinner/orgs",
"repos_url": "https://api.github.com/users/nwinner/repos",
"events_url": "https://api.github.com/users/nwinner/events{/privacy}",
"received_events_url": "https://api.github.com/users/nwinner/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Many thanks @nwinner, this is great! Thanks for breaking it off into a smaller PR too from the CP2k PR too, makes review easier.\r\n\r\nPlease make sure to include your details in [this form](https://forms.gle/JnisFb38QDR8QTFTA) so that you can be credited appropriately in the [pymatgen documentation](https://pymatgen.org/team.html) too."
] | 2020-09-29T06:04:53
| 2021-04-27T19:46:50
|
2020-10-03T03:14:37Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Bader caller has been amended to allow for parsing VASP outputs (CHGCAR) as well as the Gaussian Cube format.
* Feature 1: pymatgen.io.cube module created for Cube object, which reads and stores volumetric data from cube file just as Chgcar does for CHGCAR files.
* Feature 2: BaderAnalysis now supports either VASP or Cubes. Unit tests were overridden to explicitly reference the keyword argument desired. Because "from_path" and related methods in the bader_caller.py file are used to assimilate several VASP files, they can remain as they are, and so bader analysis done by drones in atomate will still function the same.
## Note
Passes tests and lints _except_ for one that was recently added, but the master branch also does not pass this so I believe it is out of my hands.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1967/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1967/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1967",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1967",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1967.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1967.patch",
"merged_at": "2020-10-03T03:14:37Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1968
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1968/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1968/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1968/events
|
https://github.com/materialsproject/pymatgen/pull/1968
| 711,184,779
|
MDExOlB1bGxSZXF1ZXN0NDk0ODg3OTU0
| 1,968
|
Better BSPlotter
|
{
"login": "fraricci",
"id": 17427517,
"node_id": "MDQ6VXNlcjE3NDI3NTE3",
"avatar_url": "https://avatars.githubusercontent.com/u/17427517?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/fraricci",
"html_url": "https://github.com/fraricci",
"followers_url": "https://api.github.com/users/fraricci/followers",
"following_url": "https://api.github.com/users/fraricci/following{/other_user}",
"gists_url": "https://api.github.com/users/fraricci/gists{/gist_id}",
"starred_url": "https://api.github.com/users/fraricci/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/fraricci/subscriptions",
"organizations_url": "https://api.github.com/users/fraricci/orgs",
"repos_url": "https://api.github.com/users/fraricci/repos",
"events_url": "https://api.github.com/users/fraricci/events{/privacy}",
"received_events_url": "https://api.github.com/users/fraricci/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Many thanks @fraricci ! This is a lot of very nice work.\r\n\r\nFor this point:\r\n\r\n> TODO: might by useful to implement a plotly version\r\n\r\nWe have plotly versions already (in another code, to be merged into pymatgen when they're fairly mature). We've been taking this strategy with the phase diagram plotting too, which was recently merged in. These use `bs_plot_data` fairly heavily. To make sure I understand your changes, if `split_branches=True`, `bs_plot_data` returns the same information that it did previously?\r\n\r\nIf you would like to address the plotly issue, I can share the current plotly code we have to build off that, but I don't think that's necessary to merge in this PR as-is.",
"Exactly, when split_branches=True bs_plot_data returns the usual data.\r\n(I think False case could be convenient in plotly too, since it reduces \r\nthe number of line to plot)\r\n\r\nBUT, I also swapped the order of distances and Spin in the energy key.\r\nThis means that this:\r\nbs_data[\"energy\"][d][str(Spin.up)]\r\nshould be changed in:\r\nbs_data[\"energy\"][str(Spin.up)][d]\r\n\r\nI found easier to code the rest if having the data in this way.\r\nI adapted easily the code in BSPlotterProjected, \r\nI guess it can be done as well in crystaltoolkit bs component.\r\nBut, let me know what you think.\r\n",
"Thanks for clarifying, that change makes sense. Tagging @utf @ajjackson here too since I'm not sure if this change will affect [sumo](https://sumo.readthedocs.io) (if it doesn't, please disregard)\r\n\r\nApologies for the final linter failure, I'm not sure what change was made that means the linter is checking all files.",
"So one way to replicate this issue with out the specific failing Bandstructure data is to do this on a working bandstructure object `bs`. Note that while this is a bit roundabout, it's a way to force the flaw to the front, which shouldn't happen in the first place. \r\n\r\n```python\r\nimport json\r\nfrom monty.json import MontyEncoder()\r\n\r\ndb_string = MontyEncoder().encode(bs)\r\ndb_doc = json.loads(db_string)\r\nnew_bs = BandStructureSymmLine.from_dict(db_doc)\r\nnew_bs.is_metal()\r\n```",
"<s>Thanks @shyamd, I will try this trick to test the fix impemented on some mp-id (many people are reporting mp-ids with downloading bandstructure failure). </s>\r\nOk, I misunderstood what you meant.\r\nSure, this replicate the issue: \r\nthe BandStructure obj contains numpy array that are converted by monty in a special dict.\r\nWhile BandStructure.from_dict() can recognize that dict and convert back the bands to numpy array, \r\nthe BandStructureSymmLine.from_dict() CANNOT do the same. It actually expects to get a list.\r\nFor this reason, BandStructureSymmLine.as_dict() convert bands from numpy to list.\r\n\r\nSo my question is why there are jsons of BandStructureSymmLine which contains the monty converted dict of the bands \r\ninstead of a list, given that when a BandStructureSymmLine is saved to json its as_dict is called.\r\n\r\nDo we know why this issue showed up? Is this due to the new set of bandstructure data? \r\nor is pmg that has changed at some point?",
"The last test failing is due to a \"Unnecessary \"elif\" after \"raise\" (no-else-raise)\".\r\nThe logic there is a little complex, so I'd tend to let it like it is, if you do not mind.\r\nAlso it is not in the new BSPlotter.",
"Many thanks @fraricci; I've merged this in now, for any remaining issues with serialization we can save them for another PR."
] | 2020-09-29T14:45:46
| 2020-10-10T01:18:16
|
2020-10-10T01:17:00Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
- BSPlotter now handle multiple bandstructures when they are on the same symmetry kpath.
- Usage: bsp = BSPlotter(sbs) or bsp = BSPlotter([sbs1,sbs2]); also bsp.add_bs(sbs2) or bsp.add_bs([sbs2,sbs3])
- Old plot_compare method still works but a warning will be raised.
- Differences in the distances scale are managed automatically.
- To have faster and lighter plot, it plots only a line for all the branches that are continuous.
- bs_plot_data method handles both splitting (each branch and continuous branches) for compatibility.
- bs_plot_data method handles the scaling if a bandstructure is passed as reference.
- interpolation code with B-Splines has been polished.
- interpolation can be applied selectively to some bandstructures.
- Automatic setting of the ylim according to the bandstructure plotted.
- Auto tight_layout when resizing the window ot by pressing the "T" key (thinking to add this to all mpl plots...)
- Overall improvement of the code.
- Docs updated.
- Tests added.
- TODO: might by useful to implement a plotly version
- TODO: update ipynb on matgenb
- Long standing requests of adding tests for BoltztrapPlotter has also been addressed here
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1968/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1968/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1968",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1968",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1968.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1968.patch",
"merged_at": "2020-10-10T01:16:59Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1969
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1969/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1969/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1969/events
|
https://github.com/materialsproject/pymatgen/issues/1969
| 711,608,426
|
MDU6SXNzdWU3MTE2MDg0MjY=
| 1,969
|
Issue with inheriting magmoms
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-09-30T03:44:39
| 2023-08-13T16:33:47
|
2023-08-13T16:33:46Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
See the following:
> I'm not sure if related, but I had similar problem with MPSOCSet/MPStaticSet and MAGMOM. It might be worth having a look at this issue #1397. We solved it but we let it open for further checks.
_Originally posted by @fraricci in https://github.com/materialsproject/pymatgen/pull/1917#discussion_r484482863_
Creating this issue not to lose track.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1969/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1969/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1970
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1970/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1970/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1970/events
|
https://github.com/materialsproject/pymatgen/issues/1970
| 712,611,443
|
MDU6SXNzdWU3MTI2MTE0NDM=
| 1,970
|
The "too close" atoms are not removed in the grain boundary structure for bcc [001] twist GB with primitive cell
|
{
"login": "goodwilling",
"id": 56999195,
"node_id": "MDQ6VXNlcjU2OTk5MTk1",
"avatar_url": "https://avatars.githubusercontent.com/u/56999195?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/goodwilling",
"html_url": "https://github.com/goodwilling",
"followers_url": "https://api.github.com/users/goodwilling/followers",
"following_url": "https://api.github.com/users/goodwilling/following{/other_user}",
"gists_url": "https://api.github.com/users/goodwilling/gists{/gist_id}",
"starred_url": "https://api.github.com/users/goodwilling/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/goodwilling/subscriptions",
"organizations_url": "https://api.github.com/users/goodwilling/orgs",
"repos_url": "https://api.github.com/users/goodwilling/repos",
"events_url": "https://api.github.com/users/goodwilling/events{/privacy}",
"received_events_url": "https://api.github.com/users/goodwilling/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@ucsdlxg Can you take a look?",
"Sure. I will take a look.\n\n> On Oct 10, 2020, at 9:45 PM, Shyue Ping Ong <notifications@github.com> wrote:\n> \n> \n> @ucsdlxg <https://github.com/ucsdlxg> Can you take a look?\n> \n> —\n> You are receiving this because you were mentioned.\n> Reply to this email directly, view it on GitHub <https://github.com/materialsproject/pymatgen/issues/1970#issuecomment-706641049>, or unsubscribe <https://github.com/notifications/unsubscribe-auth/AHCWHSMXMXDJ2KCJOGI2PZLSKEL3HANCNFSM4SADCIQA>.\n> \n\n",
"It seems to me that the problem is due to the structure.merge_sites() function. The temporary solution is to do one more merge_sites(tol = tol, mode='d') with your obtained GB object. Or you can add a vacuum layer, e.g. set vacuum_thickness=1.0."
] | 2020-10-01T08:16:39
| 2023-08-13T16:33:47
|
2023-08-13T16:33:47Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
In the GB structure, some atoms are too close to each other (distance < rm_ratio*bond_length_in_bulk).
Similar results are also obtained for other bcc systems, with [100], [010], [001] as the rotation axis for the
twist GB and the primitive cell as the crystal bulk structure, together with normal = False.
**To Reproduce**
Using the following python script with pymatgen 2020.9.4:
-----------------------------------
from pymatgen import Structure
from pymatgen.analysis.gb.grain import GrainBoundary, GrainBoundaryGenerator
angle = GrainBoundaryGenerator.get_rotation_angle_from_sigma(5, [0, 0, 1], lat_type='c')
struc = Structure.from_file("Fe_mp-13_primitive.cif")
generator = GrainBoundaryGenerator(struc)
gb = generator.gb_from_parameters([0, 0, 1], angle[0], expand_times=8)
gb.to(filename="Fe_mp-13_gb001.cif")
-----------------------------------
**Expected behavior**
0.7*bond_length of bulk system is the criteria of bond length,
below which the atom will be removed.
**Desktop (please complete the following information):**
- OS: Windows 10 Pro
- Version [2004]
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1970/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1970/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1971
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1971/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1971/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1971/events
|
https://github.com/materialsproject/pymatgen/issues/1971
| 712,729,413
|
MDU6SXNzdWU3MTI3Mjk0MTM=
| 1,971
|
ComputedEntry.as_dict() doesn't work with energy_adjustments
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thank you for reporting this @CompRhys ! Have you tested against the latest master branch? I believe what you are describing is the same bug that we caught and fixed in #1960 , which was just merged a few days ago. ",
"For me, using the your code snippet with the latest upstream pymatgen works properly, with the slight modification that the `uncertainty_per_degK` kwarg has been renamed to `uncertainty_per_deg`. I get the following output:\r\n\r\n```{'@module': 'pymatgen.entries.computed_entries',\r\n '@class': 'ComputedEntry',\r\n 'energy': 6.9,\r\n 'composition': defaultdict(float, {'Na': 5.0, 'Cl': 5.0}),\r\n 'energy_adjustments': [{'@module': 'pymatgen.entries.computed_entries',\r\n '@class': 'ManualEnergyAdjustment',\r\n '@version': '2020.9.14',\r\n 'value': 5},\r\n {'@module': 'pymatgen.entries.computed_entries',\r\n '@class': 'ConstantEnergyAdjustment',\r\n '@version': '2020.9.14',\r\n 'value': 5,\r\n 'uncertainty': nan,\r\n 'name': 'Constant energy adjustment',\r\n 'cls': {},\r\n 'description': 'Constant energy adjustment'},\r\n {'@module': 'pymatgen.entries.computed_entries',\r\n '@class': 'CompositionEnergyAdjustment',\r\n '@version': '2020.9.14',\r\n 'adj_per_atom': 1,\r\n 'n_atoms': 5,\r\n 'uncertainty_per_atom': 0,\r\n 'name': 'Na',\r\n 'cls': {},\r\n 'description': 'Composition-based energy adjustment'},\r\n {'@module': 'pymatgen.entries.computed_entries',\r\n '@class': 'TemperatureEnergyAdjustment',\r\n '@version': '2020.9.14',\r\n 'adj_per_deg': 0.005,\r\n 'temp': 100,\r\n 'n_atoms': 10,\r\n 'uncertainty_per_deg': 0,\r\n 'name': '',\r\n 'cls': {},\r\n 'description': 'Temperature-based energy adjustment'}],\r\n 'parameters': {},\r\n 'data': {},\r\n 'entry_id': None,\r\n 'correction': 20.0}\r\n```",
"@rkingsbury you're totally right I wasn't quite up to date and it now does work! apologies"
] | 2020-10-01T10:53:55
| 2020-10-01T17:14:36
|
2020-10-01T17:14:36Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
`ComputedEntry.as_dict()` doesn't work with `energy_adjustments`
**To Reproduce**
```python
from pymatgen.entries.computed_entries import (
ComputedEntry,
ConstantEnergyAdjustment,
CompositionEnergyAdjustment,
TemperatureEnergyAdjustment,
ManualEnergyAdjustment,
)
ealist = [
ManualEnergyAdjustment(5),
ConstantEnergyAdjustment(5),
CompositionEnergyAdjustment(1, 5, uncertainty_per_atom=0, name="Na"),
TemperatureEnergyAdjustment(0.005, 100, 10, uncertainty_per_degK=0)
]
entry = ComputedEntry("Na5Cl5", 6.9, energy_adjustments=ealist)
entry.as_dict()
```
this gives the error
```bash
Traceback (most recent call last):
File "/home/reag2/PhD/second-year/pymatgen/dupl_test.py", line 72, in <module>
entry.as_dict()
File "/home/reag2/PhD/second-year/pymatgen/pymatgen/entries/computed_entries.py", line 517, in as_dict
json.dumps(self.energy_adjustments, cls=MontyEncoder)
File "/home/reag2/anaconda3/lib/python3.7/json/__init__.py", line 238, in dumps
**kw).encode(obj)
File "/home/reag2/anaconda3/lib/python3.7/json/encoder.py", line 199, in encode
chunks = self.iterencode(o, _one_shot=True)
File "/home/reag2/anaconda3/lib/python3.7/json/encoder.py", line 257, in iterencode
return _iterencode(o, 0)
File "/home/reag2/anaconda3/lib/python3.7/site-packages/monty/json.py", line 255, in default
d = o.as_dict()
File "/home/reag2/anaconda3/lib/python3.7/site-packages/monty/json.py", line 143, in as_dict
"Unable to automatically determine as_dict "
NotImplementedError: Unable to automatically determine as_dict format from class. MSONAble requires all args to be present as either self.argname or self._argname, and kwargs to be present undera self.kwargs variable to automatically determine the dict format. Alternatively, you can implement both as_dict and from_dict.
```
Tagging authors of relevant code @rkingsbury @mattmcdermott
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1971/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1971/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1972
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1972/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1972/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1972/events
|
https://github.com/materialsproject/pymatgen/pull/1972
| 713,269,906
|
MDExOlB1bGxSZXF1ZXN0NDk2NjE4MDY5
| 1,972
|
Ignore POTCAR.spec in Vasprun
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @rkingsbury "
] | 2020-10-02T00:36:43
| 2020-10-02T03:10:37
|
2020-10-02T02:03:27Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
To facilitate testing in the absence of proprietary POTCAR files, `InputSet` have the option to write a `POTCAR.spec` file instead of an actual POTCAR. However, when a `POTCAR.spec` file is present, `Vasprun.get_potcars()` tries to read it, resulting in an error.
This PR causes Vasprun.get_potcars() to ignore POTCAR.spec when present. See also a related atomate PR which requires this change: https://github.com/hackingmaterials/atomate/pull/454
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1972/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1972/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1972",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1972",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1972.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1972.patch",
"merged_at": "2020-10-02T02:03:27Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1973
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1973/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1973/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1973/events
|
https://github.com/materialsproject/pymatgen/pull/1973
| 713,969,871
|
MDExOlB1bGxSZXF1ZXN0NDk3MTg5MTkz
| 1,973
|
Bug fixes for PDPlotter (labeling in grand + compound PDs)
|
{
"login": "mattmcdermott",
"id": 40469694,
"node_id": "MDQ6VXNlcjQwNDY5Njk0",
"avatar_url": "https://avatars.githubusercontent.com/u/40469694?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mattmcdermott",
"html_url": "https://github.com/mattmcdermott",
"followers_url": "https://api.github.com/users/mattmcdermott/followers",
"following_url": "https://api.github.com/users/mattmcdermott/following{/other_user}",
"gists_url": "https://api.github.com/users/mattmcdermott/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mattmcdermott/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mattmcdermott/subscriptions",
"organizations_url": "https://api.github.com/users/mattmcdermott/orgs",
"repos_url": "https://api.github.com/users/mattmcdermott/repos",
"events_url": "https://api.github.com/users/mattmcdermott/events{/privacy}",
"received_events_url": "https://api.github.com/users/mattmcdermott/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @mattmcdermott!"
] | 2020-10-03T00:49:16
| 2020-10-03T03:04:12
|
2020-10-03T03:04:12Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Fix 1: phase labels now appear correctly for instances of CompoundPhaseDiagram and GrandPotentialPhaseDiagram
* Fix 2: shaded uncertainty window now appears correctly for terminal compounds in instances of CompoundPhaseDiagram and GrandPotentialPhaseDiagram
* Feature: uncertainty value appears in hover label for each phase
Tagging @rkingsbury who first observed these bugs.
## Checklist
Before a pull request can be merged, the following items must be checked:
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [ ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [ ] All existing tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1973/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1973/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1973",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1973",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1973.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1973.patch",
"merged_at": "2020-10-03T03:04:12Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1974
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1974/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1974/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1974/events
|
https://github.com/materialsproject/pymatgen/pull/1974
| 714,139,625
|
MDExOlB1bGxSZXF1ZXN0NDk3MzEyNjkw
| 1,974
|
[WIP] Add PatchedPhaseDiagram to accelerate handling of large approximately disjoint data sets.
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Nice -- very timely as I've actually been wanting to solve this issue as well! I frequently need to make phase diagrams of 10+ elements, which is essentially impossible (that's around when it starts taking hours). I came up with a \"phase diagram expansion\" solution (not actually in pymatgen). It's the `expand_pd()` method here -- is this similar to your approach?\r\n\r\nhttps://github.com/GENESIS-EFRC/reaction-network/blob/96850b013538f36ee310d37c028b9000575d20f7/rxn_network/helpers.py#L611\r\n\r\nMy thought at the time was to sort the entries in reverse order by size (number of elements), and then check that each next entry is found within one of the existing larger PDs",
"The problem I'm working on now is figuring out how to do this expansion method, but preserve the information about the simplicial facets (i.e. not just the formation energies), and then combine these facets in each subspace back into one gigantic phase diagram... ",
"I think what I have is pretty similar to your function. All it does it just break up the data into the different (smaller) chemical spaces, I haven't (yet?) excluded the subsets so I would have Li-O and Li-Fe-O in my dictionary. \n\nIn terms of stitching the patches back together I haven't really thought about it at all. I would need to get more familiar with the qhull output to even give it a go\n",
"As a simple example the problem you're looking at is something like: I have NaCl, MgO, Na2O and MgCl2 - Can I get the Na-Mg-O-Cl quaternary pd from the 4 binary pds? The use would then be that I could get a decomposition for something like Mg2OCl2?",
"Yep, that is spot on. In my particular case, I'd also like to get predicted reaction products when mixing together two compositions on the phase diagram (which we already have code for via the `InterfacialReactivity` class), but you need to know the facets of the energy hull to be able to get the products. Furthermore, it turns out that it's quite possible (although not super common) to get reactions between two pentanary compounds, meaning that you will occasionally be required to create phase diagrams with 5+5=10 (or more) dimensions.",
"Sounds like an interesting problem which if it can be done would improve the functionality of this `PatchedPhaseDiagram` class.\r\n\r\nI think the thing to do is introduce a new `PhaseDiagram` child class called `StitchedPhaseDiagram` (working name) that we initialise from a list of pds and stray entries. Then if we can reconstruct all the attributes of `PhaseDiagram` in the `StitchedPhaseDiagram.__init__` we can let it inherit all the functions from `PhaseDiagram` which make the code nice and clean. Although honestly it looks pretty hard to do the simplicies and facets.\r\n\r\nI think there's two hard stages to stitching them back together firstly defining a new co-ordinate system in the new space and then secondly you'd need to iteratively check all the possible triangular facets that can be added and take the ones that are furthest away. The one thing I'm not sure about is whether you would need to handle cases where you have facets that bisect as I can visualise it beyond 3D.\r\n",
"@mattmcdermott Looking at your particular problem I don't think you need to go the whole way with `StitchedPhaseDiagram` straight away (although long term I think it would be good to add if the stitching can be achieved in a timely manner^). The `InterfacialReactivity` code only needs the facets to get the decomposition/hull_energy and it's possible to get these by solving a constrained optimisation problem like I implemented in order to add `PhaseDiagram.get_quasi_e_to_hull`** . You could probably even reuse the `_slsqp_decomp_solution` function.\r\n\r\n^ I haven't found any particularly useful resources/examples that makes me think this is possible most stuff is to do with non overlapping polygons in 2/3D geometries not non-overlapping in different dimensions\r\n**provided you don't have too many competing stable entries in the space which might not be the case in such large chemical spaces",
"@mattmcdermott I implemented `get_hull_energy` for `PatchedPhaseDiagram` using the slsqp method if we have examples that don't fit in a given patch. This might help with your problem or at least offer a partial speed up - better (i.e. more narrow) selection of the competing entries would help as the spaces get larger.\r\n\r\nHere's an example showing it gets the same results:\r\n\r\n```python\r\nfrom pymatgen import Composition\r\nfrom pymatgen.entries.entry_tools import EntrySet\r\nfrom pymatgen.analysis.tests.test_phase_diagram import module_dir\r\nfrom pymatgen.analysis.phase_diagram import (\r\n PhaseDiagram,\r\n PatchedPhaseDiagram,\r\n)\r\n\r\nentries = EntrySet.from_csv(str(module_dir / \"reaction_entries_test.csv\"))\r\n\r\npd = PhaseDiagram(entries=entries)\r\nppd = PatchedPhaseDiagram(entries=entries)\r\n\r\nprint(ppd.spaces)\r\n\r\n# novel entries not in any of the patches\r\nc1 = Composition(\"H5C2OP\")\r\nc2 = Composition(\"V2PH4C\")\r\n\r\nassert c1.chemical_system not in ppd.spaces\r\nassert c2.chemical_system not in ppd.spaces\r\n\r\nprint(\"Full Pd\")\r\nprint(pd.get_hull_energy(c1))\r\nprint(pd.get_hull_energy(c2))\r\n\r\nprint(\"Patched Pd\")\r\nprint(ppd.get_hull_energy(c1))\r\nprint(ppd.get_hull_energy(c2))\r\n```\r\n\r\nI think this should be suitable as a drop-in replacement in `InterfacialReactivity` for non-grand phase diagrams",
"First of all, thank you so much. Your PR and phase diagram updates are literally a godsend for me.\r\n\r\nThe code seems to be performing pretty well! Was able to make some gigantic (P)PDs no problem. Unfortunately, I still need to play around with `InterfacialReactivity` to get it working with the new ppds, since it actually relies on using the `get_critical_compositions()` method to get the kinks in tieline across the hull. I'm pretty confident I can get something working though.\r\n\r\nThanks again for all your additions!\r\n\r\n",
"No problem, the fact that the slsqp method can be used for more than the decomposition energy (`quasi_e_to_hull`) justifies merging my last PR despite skepticism about the usefulness of the energy in question!\n\nI have no idea how `get_critical_compositions()` works but it looks like it uses the simiplicies so that may be a problem.\n\nI probably have most of the functionality I want from ppd but it would be cool to try get as much of the functionality from pd as possible.\n\nDo you envisage there being a need for a `GrandPotentialPatchedPhaseDiagram` in your use case?",
"I don't frequently do grand potential PDs for high-throughput reaction analysis, but I do use them a lot for looking at single reactions. It would be nice eventually to add, but the reality is I can often get away with the time it takes to create several large (i.e. 9-10) element grand PDs at different chemical potential values.",
"@CompRhys so I thought a bit more about this idea of \"stitching together\" a phase diagram, and it may be an impossible effort. Consider this example ternary phase diagram which contains no ternary compounds. The question is -- how would we, given only knowledge of the hull along the binary edges of the triangle, be able to predict which binary compounds are stable against each other? For example, how would we ever determine that there is a tie-line between TiFe-TiCl3, but not one between TiFe-TiCl4?\r\n<img width=\"568\" alt=\"Screen Shot 2020-10-05 at 1 16 17 PM\" src=\"https://user-images.githubusercontent.com/40469694/95135404-3635bb80-0719-11eb-9b16-cc17ed0c2fa6.png\">\r\n\r\nMy naive idea was to do a triangulation scheme to come up with facets, but it turns out that triangulation (at least Delaunay) is done by computing the convex hull anyway, so you're going to run into the same problem.\r\n\r\nThe reason why I think it might be truly impossible to \"beat\" the qhull algorithm is that despite being our best algorithm, it also fails due to large combinatorics, i.e. that is why it stops working after 9-10 dimensions... combinatorial explosion in the number of possible facets. So it appears that if we really want to get EVERY possible simplex we will never get there even if the compositional \"middle\" of the phase diagram is sparse.\r\n\r\nInstead, I think the better idea might be to stick to using your decomposition method (with SLSQP) to get several decomposition product sets along the hull between two compositions (i.e. don't use precomputed facets to find kinks, just grab import ones intelligently -- TBD on how to do this). Effectively what I'd like to do is to say \"hey, we don't actually need EVERY facet on the hull\", and just find good guesses at what it looks like between certain compounds. Does that make any sense?\r\n\r\nAlso tagging @shyamd because we've talked about this issue in the past.",
"Posting this in case it's useful: approximate convex hull algorithm for high-dimensional data, https://arxiv.org/abs/1603.04422 ",
"I believe (?) that we already have the points that define the convex hull as the union of the stable points from the patch pds so as @mattmcdermott says the issue is that patching it together we don't get the facets/simplicies which is what we need for the other functions in code.\r\n\r\nI have done a little refactoring to add this `_get_simplex_intersections` method which if we can implement for the ppd would allow us to inherit `get_critical_compositions` from pd. In terms of implementing it I still don't have a firm grasp of what the `Simplex.line_intersections` returns so haven't made any progress.",
"You might consider using the methodology in the `CompoundPhaseDiagram` to stitch together lower dimension PDs. ",
"@shyamd The compound methodology is definitely helpful, but in this particular case since there are no entries in the \"middle\" of the gigantic phase diagram (e.g. you only have binary compounds on a ternary diagram like the pic above), you would never find the facets of the hull. In other words, the compound methodology looks for entries which *directly* exist on the tie-line between two compounds on a PD and in this case there are 0 of those compounds. Compare this to the InterfacialReactivity model, which is a true slice of the bigger convex hull and thus uses knowledge about the facet structure to find *combinations* of product compounds which minimize energy (not just single compounds).",
"Doing some reading it's the triangulation that brings in the exponential scaling. For a set of n points in d dimensions the expected number of simplicies is <img src=\"https://render.githubusercontent.com/render/math?math=\\large O(n^{d / 2})\">. The only way to avoid this will be to not generate all the simplicies. I would propose that I aim to refactor more of the code to add in private methods that could later be approximated in the ppd if someone (@mattmcdermott) has a good idea about a smart approach ",
"@mkhorton After deciding that the task of stitching back the patches isn't feasible are you still in principle happy to accept this PR based on the functionality of calculating the `e_above_hull`, `equilibrium_reaction_energy` and `quasi_e_from_hull` more quickly when starting from a high dimensional set of entries?",
"I have changed the hash for `PDEntry` to the following:\r\n\r\n```python\r\n def __hash__(self):\r\n return hash(json.dumps(self.as_dict(), sort_keys=True, cls=MontyEncoder))\r\n```\r\n\r\nAs a consequence I needed to change two tests that were set based on the hash not removing duplicates. \r\n\r\nAs `Entry` is and ABC i think it would be good to move the `__eq__` and `__hash__` from `PDEntry` into `Entry` such that they can be inherited by other classes that use the ABC e.g. `ComputedEntry`. Both these functions are written using generic ideas that should be extensible to newer `Entry` types as everything should be `MSONable`. \r\n\r\n```python\r\n def __eq__(self, other):\r\n # NOTE Scaled duplicates are not equal unless normalized separately\r\n if isinstance(other, self.__class__):\r\n return self.as_dict() == other.as_dict()\r\n return False\r\n\r\n def __hash__(self):\r\n return hash(json.dumps(self.as_dict(), sort_keys=True, cls=MontyEncoder))\r\n```\r\n\r\nIf all the tests apart from the two mentioned above still pass would this be an acceptable thing to do? I imagine that it may impact some users workflows. ",
"It looks like the test used to assert 492 unique entries and now it only has 490 in the hull data file. Can you double check that there should actually only be 490 unique entries? That is my only major concern.\r\n\r\nIn general, though, the pattern of putting `__eq__` and `__hash__` in an `ABC` interface class is not a good idea. We can get away with it because of `MSONable`. ",
"> It looks like the test used to assert 492 unique entries and now it only has 490 in the hull data file. Can you double check that there should actually only be 490 unique entries? That is my only major concern.\r\n\r\n@shyamd Yes there are literal duplicates in the `pymatgen/analysis/tests/pdentries_test.csv` file. `Li2O,2,0,1,-14.31361175` appears in rows 8 and 14 and `Fe,0,1,0,-6.59394176` appears in rows 11 and 19.\r\n\r\n> In general, though, the pattern of putting `__eq__` and `__hash__` in an `ABC` interface class is not a good idea. We can get away with it because of `MSONable`.\r\n\r\nOkay I will leave it in PDEntry!\r\n\r\n",
"@CompRhys do you have an idea how the new `__hash__` function impacts the speed of operations with `PDEntry` objects? \r\n\r\nBringing this up because I had to implement a new `__hash__` function for `GibbsComputedStructureEntry`to deal with a multiprocessing issue in my reaction network code. I originally tried that exact same code with `json.dumps` and it really slowed down everything drastically (not sure exactly which lines but I think anything involving sets/dictionaries of entries). ",
"@mattmcdermott I don't know how it might affect anything that isn't implemented for ppd so that should explored before this gets merged. \r\n\r\nI would guess the issue wouldn't be so pronounced here as `GibbsComputedStructureEntry` seems to be a much more complicated object with many more nested layers to encode and dump. What hash operation did you end up using?\r\n\r\nThe other hash I started with was\r\n\r\n```python\r\n def __hash__(self):\r\n return hash(str(self))\r\n```\r\nbut that hash seemed as though it might to be less discriminative",
"I ended up doing this:\r\n\r\n```python\r\ndef __hash__(self):\r\n data_md5 = hashlib.md5(f\"{self.composition}_\"\r\n f\"{self.formation_enthalpy}_{self.entry_id}_\"\r\n f\"{self.temp}\".encode('utf-8')).hexdigest()\r\n return int(data_md5, 16)\r\n\r\n```\r\n\r\nNot saying this is the best solution by any means but it seemed to work decently for me :) FYI I was having trouble creating entry objects on different CPUs that would hash equivalently so I could use them as keys in a shared dictionary.\r\n\r\nAlso yeah I think part of the reason is that the structure is stored with the entry, and structures can get massive -- this code removes that structure object before computing the hash (which is fine for me). So I guess now that I'm thinking about it the speed issue might only be for ComputedStructureEntry-like objects...",
"@mattmcdermott to try and make the `__eq__` and `__hash__` consistent for `Entry` type classes I have been doing a bit of refactoring. One thing I can't quite get my head around is what is supposed to/should happen to the structure when we call `normalize` on a `ComputedStructureEntry`. After calling `normalize` `self.composition` and `self.structure.composition` are not related anymore. \r\n\r\nOne option would be to just allow this disconnect (current solution) or the other I can think of would be to disable the `normalize` method for `ComputedStructureEntry` and then have `self.compositon` always return `self.structure.composition`. Do you have any thoughts on this?\r\n\r\nIt would also be really great if you could check for me whether adding this `__eq__` to your version of `GibbsComputedStructureEnergy` slows your workflow down:\r\n\r\n```python\r\n def __eq__(self, other: object) -> bool:\r\n # NOTE Scaled duplicates i.e. physically equivalent materials\r\n # are not equal unless normalized separately\r\n if id(self) == id(other):\r\n return True\r\n\r\n if isinstance(other, self.__class__):\r\n if self.entry_id != other.entry_id:\r\n return False\r\n if self._energy != other._energy:\r\n return False\r\n if self._composition != other._composition:\r\n return False\r\n # NOTE this is not performant and should be preceeded\r\n # if faster rigorous checks are available.\r\n return self.as_dict() == other.as_dict()\r\n return False\r\n```\r\n\r\nFor `GibbsComputedStructureEntry` do you ever have entries with the same `entry_id` but different temperatures? should I add another check in for that?",
"Due the issues with `__hash__` and `__eq__` for `Entry` and it's child classes (`PDEntry`, `ComputedEntry`, `GrandPotPDentry`, etc.) this PR has ballooned in scope slightly. I have refactored all the `Entry` type classes to have a nominally private/immutable `_composition` as well as `_energy` such that the `__hash__` and `__eq__` can be defined consistently across all of them. \r\n\r\nI have also moved the reaction used to calculate the dummy species for `TransformedPDEntry` inside the class such that it can be normalized (previously you couldn't normalize based on the formulae unit as there are not electronegativities for dummy species). \r\n\r\nWhilst this PR passes all the tests I am not sure about whether it can be merged yet as I imagine that there could be edge cases not covered by the tests that this might create issues for. The speed of `__eq__` may potentially be problematic for some workflows that contain lots of similar entries such that simple checks don't work and the dictionaries need to be compared.",
"I appreciate the question was directed to @mattmcdermott, but regarding:\r\n\r\n> One option would be to just allow this disconnect (current solution) or the other I can think of would be to disable the normalize method for ComputedStructureEntry and then have self.compositon always return self.structure.composition. Do you have any thoughts on this?\r\n\r\nI imagine that the structure object is retained essentially for provenance (or to calculate, say, what bonding is present so that an appropriate correction can be applied). Normalization to the entry's composition would therefore be fine, even if it results in a disconnect/disagreement. That said, a disagreement between the two certainly might be surprising so it could be clarified in the docstring.\r\n\r\nMore broadly, my only feeling here is that having Entry's be mutable at all is unfortunate, since it complicates things (rather than normalizing always returning a new Entry). I'm not sure the refactoring that would be necessary to change it at this point however. It's a shame we have to spend time thinking about `__hash__` `__eq__` etc. when something like Entry seems ideal for `@dataclass` where these features would come \"for free.\"\r\n\r\nI haven't followed this PR as closely though so will defer to @mattmcdermott on this.",
"Sorry for such a late response! @CompRhys I checked out your fork of pymatgen and some of the entry operations are ~30% slower, but that is because I put an extremely basic `__eq__()` method on my version of GibbsComputedStructureEntry, which deviated from pymatgen a while back:\r\n\r\n```python\r\ndef __eq__(self, other):\r\n if isinstance(other, self.__class__):\r\n return (self.composition == other.composition) and \\\r\n (self.formation_enthalpy == other.formation_enthalpy) and \\\r\n (self.entry_id == other.entry_id) and (self.temp == other.temp)\r\n else:\r\n return False\r\n```\r\nOf course, this doesn't check the structure, nor is it a great idea to write a slightly different version of a pymatgen class in my code, but since checking structure is a huge cost that's why I excluded it. Not sure this should be the way it functions in pymatgen though. Now that I've been thinking some more, I wouldn't worry too much about small performance differences -- just do what you think is logical and if something major comes up in my code that I think will help others, I will raise an issue :)\r\n\r\nRegarding the hashing... the current methods you added look good to me. It's really very unfortunate that Entry is mutable as @mkhorton mentioned. IMO I would refactor to make immutable, but I also agree that it's not necessary to do in this PR. Nice work on all this btw!",
"@mkhorton \r\n> Normalization to the entry's composition would therefore be fine, even if it results in a disconnect/disagreement. That said, a disagreement between the two certainly might be surprising so it could be clarified in the docstring.\r\n\r\nI have it so that it raises a warning if you normalize a computed structure entry but there are definitely edge cases that exist that I haven't looked at i.e. if I make a `GrandPotPDEntry` from a `ComputedStructureEntry` what happens if I normalize.\r\n\r\n> when something like Entry seems ideal for `@dataclass` where these features would come \"for free.\"\r\n\r\ntwo points here, `@dataclass` is py3.7+ which would mean pymatgen would need to drop support for <py3.6 and apparently most people still use 3.6** and in #1985 @rkingsbury suggested that there was some benefit to `ComputedEntry` being mutable in regards to adding corrections. I am not sure if `@dataclass` allows for some things to be frozen i.e. the composition, structure etc and others to be mutable i.e. the corrections applied. This may be something that the dev team wants to think about but as we can mostly solve the `__hash__` and `__eq__` issues with getting the correct sets of stable entries for `PatchedPhaseDiagram` without making any breaking changes I think I should try do the minimum required amount such that this PR works. \r\n\r\nThat said I am not in a huge rush to merge this PR as I think more consideration of edge cases (like #1985 for my last PR) are needed as we're changing `ComputedEntry`a little.\r\n\r\n@mattmcdermott \r\n> checking structure is a huge cost that's why I excluded it.\r\n\r\nI also don't really think we need to but there's balance to this, if a user is using the code as intended they will be using an unique id to distinguish things in which case it could be faster for everyone to not check structures. I am not sure where on the balance between **robust** and **(fairly) robust** to target. \r\n\r\n\r\n**although dropping <py3.6 would make the tests pass as it appears to have a floating point error that the other test actions didn't encounter.",
"Brief note on the `@dataclass` point, there's an official back port to 3.6 and it's already in the pymatgen requirements. Nevertheless, it was more a general comment than a strong recommendation."
] | 2020-10-03T17:43:17
| 2021-01-19T17:56:31
|
2021-01-19T17:56:31Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Introduce a new `PhaseDiagram` based class to handle large disjoint data sets. Calculation of the convex hull of a data set scales with the dimensions and amount of data. Breaking up disjoint datasets into smaller patches reduces the complexity in both these regards and would allow for parallel computation of different patches.
Not all the functionality of `PhaseDiagram` will be available in `PatchedPhaseDiagram` as some of these functions can't be quickly calculated in very very large chemical spaces without the full phase diagram. Currently the aim will be to keep all functionality that would be helpful for those investigating Machine Learning workflows who may be interested in the relative stability large spaces I.e. the aim is to allow for `get_quasi_e_to_hull` to be evaluated from scratch for the entirety of the Materials Project on a 5 year old laptop in a timely manner.
## TODO
#### Functionality
* ~~Use multiprocessing to accelerate the calculation of the convex hulls within the patches~~ **on a 85 dimensional ppd with 4000 entries we take 90 seconds without mp and 60 seconds using mp across 3 cores**
* ~~Add function to stitch patches together when required~~ **deemed impossible (see discussion below)**
* Standardise `__eq__` and `__hash__` for `Entry` and it's child classes.
#### Speed
* ~~Carry out a speed test to show this is actually faster~~ **Much faster provided we have a high rank chemical space e.g. 140msec vs 10.4 sec for a toy set of just 30 materials containing 10 diverse materials and 20 different elements**
## Checklist
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
- [x] Tests have been added for any new functionality or bug fixes. **Tests expose functionality that was not configured correctly previously - this now needs to be resolved**
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1974/reactions",
"total_count": 2,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 1,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1974/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1974",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1974",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1974.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1974.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1975
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1975/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1975/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1975/events
|
https://github.com/materialsproject/pymatgen/pull/1975
| 714,807,926
|
MDExOlB1bGxSZXF1ZXN0NDk3ODI3NzE3
| 1,975
|
Parameters can be written to exciting input file via nested dictionary
|
{
"login": "vorwerkc",
"id": 2338896,
"node_id": "MDQ6VXNlcjIzMzg4OTY=",
"avatar_url": "https://avatars.githubusercontent.com/u/2338896?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/vorwerkc",
"html_url": "https://github.com/vorwerkc",
"followers_url": "https://api.github.com/users/vorwerkc/followers",
"following_url": "https://api.github.com/users/vorwerkc/following{/other_user}",
"gists_url": "https://api.github.com/users/vorwerkc/gists{/gist_id}",
"starred_url": "https://api.github.com/users/vorwerkc/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/vorwerkc/subscriptions",
"organizations_url": "https://api.github.com/users/vorwerkc/orgs",
"repos_url": "https://api.github.com/users/vorwerkc/repos",
"events_url": "https://api.github.com/users/vorwerkc/events{/privacy}",
"received_events_url": "https://api.github.com/users/vorwerkc/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Many thanks for this @vorwerkc ! Since `paramdict` is freeform (for any settings not otherwise specified), perhaps it would be better to set this at the end as `**kwargs` instead? Otherwise this looks good to merge.",
"Hi @vorwerkc, I made a quick commit to illustrate what I meant by the \"kwargs\" comment, so now the relevant line is simply:\r\n\r\n```\r\n self._dicttoxml(kwargs, root)\r\n```\r\n\r\nand I've updated the test accordingly too. This means, for example, you can still do:\r\n\r\n```\r\n paradir = {'grst': {'do': 'fromscratch', 'ngridk': '8 8 8', \r\n 'xctype': 'GGA_PBE_SOL', 'gmaxvr': '14.0'}, \r\n 'xs': {'xstype': 'BSE', 'ngridk': '4 4 4', \r\n 'ngridq': '4 4 4', 'nempty': '30', 'gqmax': '3.0', \r\n 'broad': '0.07', 'tevout': 'true', 'energywindow':\r\n {'intv': '0.0 1.0', 'points': '1200'},\r\n 'screening': {'screentype': 'full', 'nempty': '100'}, \r\n 'BSE': {'bsetype': 'singlet', 'nstlbse': '1 5 1 4'}}}\r\n\r\n test_input = ExcitingInput(struct)\r\n test_input.write_string('unchanged', **paradir)\r\n```\r\n\r\nbut also, for example,\r\n\r\n```\r\n test_input = ExcitingInput(struct)\r\n test_input.write_string('unchanged', grst={\"xctype\": \"GGA_PBE_SOL\"}, xs=...) # etc.\r\n```\r\n\r\nif you just want to make a smaller change.",
"Hi @mkhorton,\r\nthanks for the review! You are right, your solution is more general and more elegant. I was stuck in a FORTRAN mindset...\r\nFrom my side, the pull request is ready to be merged.",
"Thanks @vorwerkc !"
] | 2020-10-05T12:45:57
| 2020-12-02T21:02:27
|
2020-12-02T21:02:24Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Small additional feature that allows the user to generate an exciting input file with additional parameters. The parameters are provided in the form of a nested dictionary, which allows to generate a full exciting input files with all elements and attributes.
* Additional arguments for io.exciting.ExcitingInput.write_etree, io.exciting.ExcitingInput.write_string, io.exciting.ExcitingInput.write_file to provide additional parameters that are included in the exciting input
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1975/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1975/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1975",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1975",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1975.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1975.patch",
"merged_at": "2020-12-02T21:02:24Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1976
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1976/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1976/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1976/events
|
https://github.com/materialsproject/pymatgen/pull/1976
| 715,181,191
|
MDExOlB1bGxSZXF1ZXN0NDk4MTMzNzQ3
| 1,976
|
Default Co to low spin.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-10-05T21:40:54
| 2020-10-05T21:44:34
|
2020-10-05T21:43:12Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
* Feature 1
* Feature 2
* Fix 1
* Fix 2
## Additional dependencies introduced (if any)
* List all new dependencies needed and justify why. While adding dependencies that bring
significantly useful functionality is perfectly fine, adding ones that
add trivial functionality, e.g., to use one single easily implementable
function, is frowned upon. Provide a justification why that dependency is needed.
Especially frowned upon are circular dependencies, e.g., depending on derivative
modules of pymatgen such as custodian or Fireworks.
## TODO (if any)
If this is a work-in-progress, write something about what else needs
to be done
* Feature 1 supports A, but not B.
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [ ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [ ] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1976/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1976/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1976",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1976",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1976.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1976.patch",
"merged_at": "2020-10-05T21:43:11Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1977
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1977/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1977/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1977/events
|
https://github.com/materialsproject/pymatgen/issues/1977
| 715,677,132
|
MDU6SXNzdWU3MTU2NzcxMzI=
| 1,977
|
XSF atomic symbol as label request
|
{
"login": "seryi70",
"id": 1672249,
"node_id": "MDQ6VXNlcjE2NzIyNDk=",
"avatar_url": "https://avatars.githubusercontent.com/u/1672249?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/seryi70",
"html_url": "https://github.com/seryi70",
"followers_url": "https://api.github.com/users/seryi70/followers",
"following_url": "https://api.github.com/users/seryi70/following{/other_user}",
"gists_url": "https://api.github.com/users/seryi70/gists{/gist_id}",
"starred_url": "https://api.github.com/users/seryi70/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/seryi70/subscriptions",
"organizations_url": "https://api.github.com/users/seryi70/orgs",
"repos_url": "https://api.github.com/users/seryi70/repos",
"events_url": "https://api.github.com/users/seryi70/events{/privacy}",
"received_events_url": "https://api.github.com/users/seryi70/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Can you upload a sample XSF file?",
"Sure thing, thanks a lot.\r\nSergiu\r\n\r\n[pymatgen_example.zip](https://github.com/materialsproject/pymatgen/files/5339451/pymatgen_example.zip)\r\n\r\n\r\n",
"Done. Now the default format uses atom symbols."
] | 2020-10-06T13:25:05
| 2020-10-11T02:44:29
|
2020-10-11T02:44:17Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
We get frustrated that pymatgen XSF implemented only the number as atom label, whereas most software can handle the regular atomic symbol, much easier to find elements of interest in the file.
**Describe the solution you'd like**
Please add the possibility to dump/read XSF files with atomic symbol (H,He,O) instead of atomic number (1,2,8) for atom label.
Thank you.
Sergiu
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1977/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1977/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1978
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1978/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1978/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1978/events
|
https://github.com/materialsproject/pymatgen/issues/1978
| 715,681,781
|
MDU6SXNzdWU3MTU2ODE3ODE=
| 1,978
|
Dump structure to Lammps format
|
{
"login": "seryi70",
"id": 1672249,
"node_id": "MDQ6VXNlcjE2NzIyNDk=",
"avatar_url": "https://avatars.githubusercontent.com/u/1672249?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/seryi70",
"html_url": "https://github.com/seryi70",
"followers_url": "https://api.github.com/users/seryi70/followers",
"following_url": "https://api.github.com/users/seryi70/following{/other_user}",
"gists_url": "https://api.github.com/users/seryi70/gists{/gist_id}",
"starred_url": "https://api.github.com/users/seryi70/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/seryi70/subscriptions",
"organizations_url": "https://api.github.com/users/seryi70/orgs",
"repos_url": "https://api.github.com/users/seryi70/repos",
"events_url": "https://api.github.com/users/seryi70/events{/privacy}",
"received_events_url": "https://api.github.com/users/seryi70/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"You can use the pymatgen.io.lammps to create LAMMPS format. Not all formats are given in fmt because LAMMPS require other parameters to be specified."
] | 2020-10-06T13:30:33
| 2020-10-11T02:37:03
|
2020-10-11T02:37:02Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Dear pymatgen developers, is it possible to have pymatge.core.structure.to(fmt='lammps',...) ?
**Describe the solution you'd like**
A simple format conversion described here: https://atomsk.univ-lille.fr/tutorial_lammps.php
Besr regards,
Sergiu
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1978/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1978/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1979
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1979/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1979/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1979/events
|
https://github.com/materialsproject/pymatgen/pull/1979
| 716,868,916
|
MDExOlB1bGxSZXF1ZXN0NDk5NTMxMjc2
| 1,979
|
Remove self.kwargs.get pattern in MP input sets
|
{
"login": "utf",
"id": 1330638,
"node_id": "MDQ6VXNlcjEzMzA2Mzg=",
"avatar_url": "https://avatars.githubusercontent.com/u/1330638?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/utf",
"html_url": "https://github.com/utf",
"followers_url": "https://api.github.com/users/utf/followers",
"following_url": "https://api.github.com/users/utf/following{/other_user}",
"gists_url": "https://api.github.com/users/utf/gists{/gist_id}",
"starred_url": "https://api.github.com/users/utf/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/utf/subscriptions",
"organizations_url": "https://api.github.com/users/utf/orgs",
"repos_url": "https://api.github.com/users/utf/repos",
"events_url": "https://api.github.com/users/utf/events{/privacy}",
"received_events_url": "https://api.github.com/users/utf/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @utf!"
] | 2020-10-07T21:38:15
| 2020-10-08T06:03:43
|
2020-10-08T06:03:36Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
A common pattern in many of the MP VASP input sets is to store the input kwargs in the constructor and then access them using `self.kwargs.get` in the `incar` and `kpoint` functions. Unfortunately, this can lead to discrepancies between the actual attributes of the input set and the data stored in the `kwargs` variable.
For example, the following causes an error:
```python
static_set = MPStaticSet(structure, user_incar_settings=None)
static_set.incar()
# File ".../pymatgen/io/vasp/sets.py", line 974, in incar
# self.kwargs.get("user_incar_settings", {}).keys()
# AttributeError: 'NoneType' object has no attribute 'keys'
```
This is surprising since the default value of `user_incar_settings` in the `VaspInputSet` constructor is `None`, and the following code works fine.
```python
static_set = MPStaticSet(structure)
static_set.incar()
```
The issue arises because the `incar` and `kpoints` functions call `self.kwargs.get("user_incar_settings", {}).keys()` which evaluates to `None.key()`. This issue can be avoided by just calling `self.user_incar_settings.keys()`. This is because the `VaspInputSet` constructor initialises `user_incar_settings` to as an empty dictionary if `None` is given as the argument.
I've left the `self.kwargs` pattern in the input set constructor as the `kwargs` variable is used by the monty decoder in the `as_dict` function. Note that I only edited MP input sets.
This issue is responsible for a bug in atomate: hackingmaterials/atomate#445.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1979/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1979/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1979",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1979",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1979.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1979.patch",
"merged_at": "2020-10-08T06:03:36Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1980
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1980/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1980/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1980/events
|
https://github.com/materialsproject/pymatgen/issues/1980
| 718,251,011
|
MDU6SXNzdWU3MTgyNTEwMTE=
| 1,980
|
Plot of neutron diffraction is broken
|
{
"login": "goodwilling",
"id": 56999195,
"node_id": "MDQ6VXNlcjU2OTk5MTk1",
"avatar_url": "https://avatars.githubusercontent.com/u/56999195?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/goodwilling",
"html_url": "https://github.com/goodwilling",
"followers_url": "https://api.github.com/users/goodwilling/followers",
"following_url": "https://api.github.com/users/goodwilling/following{/other_user}",
"gists_url": "https://api.github.com/users/goodwilling/gists{/gist_id}",
"starred_url": "https://api.github.com/users/goodwilling/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/goodwilling/subscriptions",
"organizations_url": "https://api.github.com/users/goodwilling/orgs",
"repos_url": "https://api.github.com/users/goodwilling/repos",
"events_url": "https://api.github.com/users/goodwilling/events{/privacy}",
"received_events_url": "https://api.github.com/users/goodwilling/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks. I implemented the fix with a new test to ensure this does not recur.\r\n"
] | 2020-10-09T15:48:22
| 2020-10-11T02:36:07
|
2020-10-11T02:35:52Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Neutron diffraction pattern can not be plot. A bug fix is suggested.
**To Reproduce**
Error always comes out if using the following script:
from pymatgen import Structure
from pymatgen.analysis.diffraction.neutron import NDCalculator
import sys
struc = Structure.from_file("Fe_mp-13_primitive.cif");
plot=NDCalculator().get_plot(struc)
plot.show()
with the following error message
------------------
Traceback (most recent call last):
File "calc_XRD.py", line 6, in <module>
plot=NDCalculator().get_plot(struc)
File "C:\ProgramData\miniconda3\lib\site-packages\pymatgen\analysis\diffraction\core.py", line 108, in get_plot
label = ", ".join([str(hkl["hkl"]) for hkl in hkls])
File "C:\ProgramData\miniconda3\lib\site-packages\pymatgen\analysis\diffraction\core.py", line 108, in <listcomp>
label = ", ".join([str(hkl["hkl"]) for hkl in hkls])
TypeError: tuple indices must be integers or slices, not str
------------------
**Expected behavior**
A matplotlib plot should be shown.
**Suggested bug fix**
With the following correction, the neutron diffraction pattern can be plot as expected:
pymatgen\analysis\diffraction\neutron.py, Line 197:
hkls.append(fam)
-->
#hkls.append(fam)
hkls.append([{"hkl": hkl, "multiplicity": mult}
for hkl, mult in fam.items()])
**Desktop (please complete the following information):**
- OS: Windows 10
- Version: Pymatgen-2020.09.14
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1980/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1980/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1981
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1981/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1981/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1981/events
|
https://github.com/materialsproject/pymatgen/issues/1981
| 718,738,351
|
MDU6SXNzdWU3MTg3MzgzNTE=
| 1,981
|
Confirm the equivalence of different surfaces belong to different spacegroups which are related via the change-of-basis transformation.
|
{
"login": "hongyi-zhao",
"id": 11155854,
"node_id": "MDQ6VXNlcjExMTU1ODU0",
"avatar_url": "https://avatars.githubusercontent.com/u/11155854?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/hongyi-zhao",
"html_url": "https://github.com/hongyi-zhao",
"followers_url": "https://api.github.com/users/hongyi-zhao/followers",
"following_url": "https://api.github.com/users/hongyi-zhao/following{/other_user}",
"gists_url": "https://api.github.com/users/hongyi-zhao/gists{/gist_id}",
"starred_url": "https://api.github.com/users/hongyi-zhao/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/hongyi-zhao/subscriptions",
"organizations_url": "https://api.github.com/users/hongyi-zhao/orgs",
"repos_url": "https://api.github.com/users/hongyi-zhao/repos",
"events_url": "https://api.github.com/users/hongyi-zhao/events{/privacy}",
"received_events_url": "https://api.github.com/users/hongyi-zhao/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Your statement is incorrect. C2/c and I2/b are both monoclinic. They even have the same space group (No. 15). It is just a different choice of axes. All this can be found in the International Tables for Space group No. 15."
] | 2020-10-11T01:54:32
| 2020-10-11T02:31:36
|
2020-10-11T02:31:36Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hi,
Based on the descriptions in [this paper](https://pubs.acs.org/doi/pdf/10.1021/acs.chemmater.9b05047), see top left column on pp.3 for more info, I learned the following fact:
`
The (010) surface in C2/c is equivalent to the (001) surface in I2/b. And these two space groups are related through a change-of-basis transformation. An I2/b cell is with tetragonal lattice while a C2/c cell is related to the conventional monoclinic cell.
`
But The above description is very abstract and difficult to understand, so I want to know whether I can have a better understanding of the problem based on the help of pymatgen?
Regards,
HY
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1981/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1981/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1982
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1982/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1982/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1982/events
|
https://github.com/materialsproject/pymatgen/issues/1982
| 719,461,054
|
MDU6SXNzdWU3MTk0NjEwNTQ=
| 1,982
|
Wrong tag naming causing pylint errors
|
{
"login": "shyamd",
"id": 16827130,
"node_id": "MDQ6VXNlcjE2ODI3MTMw",
"avatar_url": "https://avatars.githubusercontent.com/u/16827130?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyamd",
"html_url": "https://github.com/shyamd",
"followers_url": "https://api.github.com/users/shyamd/followers",
"following_url": "https://api.github.com/users/shyamd/following{/other_user}",
"gists_url": "https://api.github.com/users/shyamd/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyamd/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyamd/subscriptions",
"organizations_url": "https://api.github.com/users/shyamd/orgs",
"repos_url": "https://api.github.com/users/shyamd/repos",
"events_url": "https://api.github.com/users/shyamd/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyamd/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I just pin it to the without v for now."
] | 2020-10-12T15:17:22
| 2020-10-12T16:36:16
|
2020-10-12T16:36:15Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
It looks like the last release didn't have a `v` in front of the version in the git tag. This is causing the pylint script to fail. This could be fixed by just updating that for now, or we could force push update the git tag?
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1982/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1982/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1983
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1983/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1983/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1983/events
|
https://github.com/materialsproject/pymatgen/issues/1983
| 722,403,950
|
MDU6SXNzdWU3MjI0MDM5NTA=
| 1,983
|
Python 3.9 support
|
{
"login": "sheerluck",
"id": 5173768,
"node_id": "MDQ6VXNlcjUxNzM3Njg=",
"avatar_url": "https://avatars.githubusercontent.com/u/5173768?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/sheerluck",
"html_url": "https://github.com/sheerluck",
"followers_url": "https://api.github.com/users/sheerluck/followers",
"following_url": "https://api.github.com/users/sheerluck/following{/other_user}",
"gists_url": "https://api.github.com/users/sheerluck/gists{/gist_id}",
"starred_url": "https://api.github.com/users/sheerluck/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/sheerluck/subscriptions",
"organizations_url": "https://api.github.com/users/sheerluck/orgs",
"repos_url": "https://api.github.com/users/sheerluck/repos",
"events_url": "https://api.github.com/users/sheerluck/events{/privacy}",
"received_events_url": "https://api.github.com/users/sheerluck/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This is related to the fact that numpy and cython does not exist in Python 3.9 yet. See https://github.com/pandas-dev/pandas/issues/32045 . Just keep pymatgen to python3.7 and 3.8 for now.",
"FYI for this, it looks like `tp_print` has been [removed in Python 3.9](https://docs.python.org/3.9/whatsnew/3.9.html#id3), since it hasn't been used for since Python 3.0. I see that the `coord_cython.c` committed in the repo was last built 2 years ago, it's possible just re-building it with any newer Cython might fix it.",
"I recompiled it. Not sure if it will fix it.",
"Thanks, I'm testing now. Looks like we might have the same issue in `linear_assignment.c` (presumably `neighbors.c` too)",
"Ok, I rebuilt those. Perhaps we should add some code in tasks.py to regenerate those C files prior to each release using the latest cython?",
"Ok, pymatgen seems to build and work fine for me now.\r\n\r\nPython 3.9 support in general is still spotty for now though, neither numpy, pandas or matplotlib have pre-built wheels available yet.\r\n\r\n> Perhaps we should add some code in tasks.py to regenerate those C files prior to each release using the latest cython?\r\n\r\nProbably not a bad idea. I'm not that familiar with Cython. It seems like these changes are quite rare."
] | 2020-10-15T14:45:31
| 2020-10-16T23:17:24
|
2020-10-16T12:05:25Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
pymatgen fails to build with Python 3.9.0
```
pymatgen/util/coord_cython.c:22836:25: error: «PyTypeObject» {aka «struct _typeobject»} has no member named «tp_print»
22836 | __pyx_type___pyx_array.tp_print = 0;
```
**To Reproduce**
Steps to reproduce the behavior:
1. build with Python 3.9.0
2. See error
**Desktop (please complete the following information):**
- OS: Gentoo linux with Python 3.9.0
- Version: master branch
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1983/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1983/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1984
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1984/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1984/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1984/events
|
https://github.com/materialsproject/pymatgen/issues/1984
| 723,780,939
|
MDU6SXNzdWU3MjM3ODA5Mzk=
| 1,984
|
[Feature Request] Better compatibility between PDPlotter and matplotlib subplots
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"+1 . I agree this would be very useful. ",
"I think it might be nicer to just change the `plt` argument to `ax` as I feel that is more natural to work with.\r\n\r\nThat said, to use the existing `plt` argument with subplots often the only thing you have to do is set the current axis to the axis you want to plot to. I.e.,\r\n\r\n```python\r\nimport matplotlib.pyplot as plt\r\n\r\nfig, (ax1, ax2) = plt.subplots(2, 1)\r\n\r\n# ... generate PDPlotter object...\r\n\r\nplt.sca(ax2) # set the current axis to ax2\r\nplotter.get_plot(..., plt=plt)\r\n```",
"I don't have a problem with this. Can @utf implement and submit a PR?",
"Yes I can implement this.",
"@utf I submitted a similar PR that adds an `ax` kwarg to `plot_brillouin`. Could you have a look at #2039?",
"I just updated `PDPlotter` in https://github.com/materialsproject/pymatgen/pull/3032 but this is still not addressed. I don't feel particularly good about touching some of the matplotlib code, so if anyone is interested in improving it sometime, please feel free!"
] | 2020-10-17T15:17:38
| 2023-08-13T16:33:47
|
2023-08-13T16:33:47Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Is your feature request related to a problem? Please describe.**
It would be nice to be able to pass `matplotlib` axes objects to `PDPlotter` when using the `matplotlib` backend such that they can be easily incorporated into subplots.
**Describe the solution you'd like**
a `kwarg` called `axes` in PDPlotter that gives the axes to plot the phase diagram on in a subplots
**Additional context**
there is a `kwarg` called `plt` in `PDPlotter` that may have similar functionality but I haven't managed to use it for this use case successfully
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1984/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1984/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1985
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1985/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1985/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1985/events
|
https://github.com/materialsproject/pymatgen/issues/1985
| 725,177,010
|
MDU6SXNzdWU3MjUxNzcwMTA=
| 1,985
|
PhaseDiagram.get_equilibrium_reaction_energy no longer works for ComputedEntry
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@shyamd feel free to add anything I missed\r\n",
"I think this might actually be a bug in `.normalize`. The unittest we have for `get_equilibrium_reaction_energy` passes, and it builds a `PhaseDiagram` from very simple `PDEntry` - they just have an energy and a composition. Using those entries,\r\n```\r\n[e.normalize(inplace=False) for e in pd.stable_entries][0] in [e.normalize(inplace=False) for e in pd.stable_entries]\r\n```\r\nreturns `True`. So somehow, using `ComputedEntry` instead of `PDEntry` causes this to fail.",
"The behavior of `ComputedEntry` did change recently, see https://github.com/materialsproject/pymatgen/commit/513eddb3fe32a43270df1f51cd58b93f9cc1aa9b",
"I think this might be a subtle memory referencing thing. Using `ComputedEntry` (not `PDEntry`), I can do\r\n\r\n```\r\n>>> with MPRester() as m:\r\n entries = m.get_entries_in_chemsys(\"Zn-S\")\r\n\r\n>>> pd = PhaseDiagram(entries)\r\n>>> [e.normalize(inplace=False) for e in pd.stable_entries][0] in pd.get_stable_entries_normed()\r\nFalse\r\n>>> [e.normalize(inplace=False) for e in pd.stable_entries][0].as_dict() == pd.get_stable_entries_normed()[0].as_dict()\r\nTrue\r\n```\r\nand visual inspection of both dicts shows that the entries are identical.\r\n\r\nIn the case of a `PDEntry` that has far fewer attributes than a `ComputedEntry`, is it possible that Python is doing some sort of lazy referencing where it doesn't create a new copy of the entry because an identical one is already in memory? And then somehow it is NOT doing this for `ComputedEntry`?\r\n\r\n",
"That last bit is because you're using Dict equals and i bet normalize isn't changing any of the values so the final dicts do have the same data. This also highlights that there might be an intrinsic issue where there should be a `__eq__` and `__hash__` defined if comparisons are being done on Entries.",
"@rkingsbury @shyamd I implemented `pd.get_stable_entries_normed()` which returns a list and so comparisons against it use `__eq__` rather than hash equality as a way to get around the fact that the `__hash__` for Entries is based on their id in memory rather than the object being considered. I think that if this is causing issues it would be valid just to remove the check in `get_equilibrium_reaction_energy` as it wasn't their before I added `quasi_e_from_hull`.\r\n\r\nHowever, I think that setting a `__hash__` based on the object would be the preferable solution. The question is then do you hash based on some normalised representation or the inputted representation. I wanted to hash based on a normalised representation but that may lead to issues elsewhere in the code which is why @mkhorton advised me to try find a solution that avoided changing the hash when I added `get_quasi_e_from_hull`. ",
"Thanks for investigating this @CompRhys! You are correct that if I comment out the stability check, the bug goes away. However, the code always returns a reaction energy of zero because the next list equality check still fails.\r\n\r\n```\r\n def get_equilibrium_reaction_energy(self, entry):\r\n \"\"\"\r\n Provides the reaction energy of a stable entry from the neighboring\r\n equilibrium stable entries (also known as the inverse distance to\r\n hull).\r\n\r\n Args:\r\n entry (PDEntry): A PDEntry like object\r\n\r\n Returns:\r\n Equilibrium reaction energy of entry. Stable entries should have\r\n equilibrium reaction energy <= 0. The energy is given per atom.\r\n \"\"\"\r\n # if entry.normalize(inplace=False) not in self.get_stable_entries_normed():\r\n # raise ValueError(\r\n # \"{} is unstable, the equilibrium reaction energy is\"\r\n # \"available only for stable entries.\".format(entry)\r\n # )\r\n\r\n if entry.is_element:\r\n return 0\r\n\r\n entries = [e for e in self.stable_entries if e.normalize(inplace=False) != entry.normalize(inplace=False)]\r\n modpd = PhaseDiagram(entries, self.elements)\r\n return modpd.get_decomp_and_e_above_hull(entry, allow_negative=True)[1]\r\n```\r\n\r\nI'm not well acquainted with `__eq__` vs hash equality. Which is normally invoked by Python when testing for membership in a list?",
"So I spent a bit of time digging into this and here is my big picture recommendations:\r\n\r\n1. Read https://hynek.me/articles/hashes-and-equality/\r\n2. Make `Entry` a fully functioning class by having `Entry.energy` just return `Entry._energy`.\r\n3. Thus`Entry` should have a `__hash__` and `__eq__` .\r\na. `__hash__` can just use the hash of the reduced composition\r\nb. `__eq__` can check the composition are equals and the energies are equals within 1E-6\r\n4. `PDEntry` `__eq__` should first check if the other is a PDEntry\r\na. if so, equality should be based on ID no?\r\nb. if both IDs are `None`, then use`Entry.__eq__`\r\n\r\n",
"Wouldn't leaving `Entry` as an `ABC` but setting `__eq__` and `__hash__` as abstractmethods be an alternative? That would force careful consideration of what the ops do. \n\nI think that `PDEntry` needs to hash based on the object as the funky behaviour that arises from the hash for entries being based on mutable properties isn't fixed in terms of code functionality by the current hash based on the id (I can change the energy of something in `pd.stable_entries` such that it is unstable and it will remain in the set of stable entries) and things don't behave as expected (i.e. literal duplicates in the entry \"set\" used to test `PhaseDiagram` here due to the order they're loaded into memory hash(a)!=hash(b) even though a==b which is the condition of a hash).\n\nIn #1974 when stitching the `stable_entries` together in `PatchedPhaseDiagram` I end up with large numbers of duplicates that result in strange nominal decompositions. \n\nCould `Entry`, `PDEntry`, `ComputedEntry`, etc. reasonably be made immutable using `__slots__=[]` and making them child classes of `tuple`? Are there reasons to be able to change their properties once they have been declared?\n",
"The default object/user-defined-class `__hash__` and `__eq__` are \r\n\r\n```python \r\n def __eq__(self, o) -> bool:\r\n # NOTE this is the default __eq__ of a user defined class\r\n # return object.__eq__(self, o)\r\n return id(self) == id(o)\r\n\r\n def __hash__(self) -> int:\r\n # NOTE this is the default __hash__ of a user defined class\r\n return hash(id(self)/16)\r\n```\r\n\r\ntherefore these are the `__eq__` and `__hash__` for `ComputedEntry`. The check based on the normalized entries is based on the `__eq__` of `PDEntry` which leads to the bug above.\r\n\r\n@rkingsbury As I understand list comparisons use `__eq__` and set comparisons use `__hash__` first then `__eq__`. This is why the hashing criteria can be weaker than the equality criteria.\r\n\r\nIt's interesting that it appears that the reason the `__hash__` is explicitly defined in `PDEntry` is that py3+ doesn't allow you to define `__eq__` explicitly without also defining the `__hash__`. The `__eq__` that was defined changes based on the object being mutable but the `__hash__` does not. This appears not to be great code practise and is probably the underlying cause of the issues both here and the ones I am having in getting the correct functionality in #1974 \r\n\r\nLooking a little more if pymatgen were to move to having py3.7+ as it's req we could use https://docs.python.org/3/library/dataclasses.html with `frozen=True` to make the entries immutable (much cleaner than the `__slots__` approach - so much so that I would say it's not worth looking at the `__slots__` method as it's super hacky). Making them immutable would have implications on normalisation in-place and potentially in other parts of the code also.\r\n",
"@rkingsbury This is pretty much what it was before I added in the checks for normalised duplicates of stable entries.\r\n\r\n```python\r\n if entry not in self.stable_entries:\r\n raise ValueError(\r\n \"{} is unstable, the equilibrium reaction energy is\"\r\n \"available only for stable entries.\".format(entry)\r\n )\r\n\r\n if entry.is_element:\r\n return 0\r\n\r\n entries = [e for e in self.stable_entries if e != entry]\r\n modpd = PhaseDiagram(entries, self.elements)\r\n return modpd.get_decomp_and_e_above_hull(entry, allow_negative=True)[1]\r\n```\r\n\r\nThis code has the pathology that say you ran two calculations one with a supercell twice the unit cell of the other such that the energy and composition were scaled by 2. Only one of them would end up in the stable materials and have a non-zero reaction energy whilst the other would have a reaction energy of 0. It was like this for many years so I am not sure if this is an edge case that actually arises in user's workflows.",
"That is sort of expected. PD is not expected to perform duplicate checking. The user is expected to take care of that. Maybe we could provide some sort of warning in that case though?",
"the check is necessary to pass the tests I wrote for `get_quasi_e_from_hull` using the testing entry set as it contains literal duplicates. I didn't necessarily see the problem of duplicates/scaled-duplicates arising in actual use but it does in the test set of entries so I had to work the edge case.\r\n\r\nAs the same edge case exists in `get_equilibrium_reaction_energy` (although it isn't being tested) I added the check into `get_equilibrium_reaction_energy` for parity. \r\n\r\nI wasn't aware at that time that `ComputedEntry` didn't use the same `__eq__` as `PDEntry`. I think it's best to revert the change and remove the check in `get_equilibrium_reaction_energy` but would really like to explicitly resolve the `__eq__` and `__hash__` issue as it also impacts #1974.",
"I'd agree that we should resolve the discrepancy where, as I understand it, `__eq__` is different between `ComputedEntry` and `PDEntry` while `__hash__` is inherited from `PDEntry`.\r\n\r\nI don't think it's practical to make `ComputedEntry` immutable, mainly because we need to be able to process them to apply energy corrections, which are stored in the `energy_adjustments` attribute. That attribute has to be mutable in order for corrections to work.",
"For `ComputedEntry` and `Entry` we have:\r\n\r\n```python\r\n def __eq__(self, o) -> bool:\r\n # NOTE this is the default __eq__ of a user defined class\r\n return id(self) == id(o)\r\n\r\n def __hash__(self) -> int:\r\n # NOTE this is the default __hash__ of a user defined class\r\n return hash(id(self)/16)\r\n```\r\n\r\nFor `PDEntry` we have:\r\n\r\n```python\r\n def __eq__(self, other):\r\n # NOTE Scaled duplicates are not equal unless normalized separately\r\n if isinstance(other, self.__class__):\r\n return self.as_dict() == other.as_dict()\r\n return False\r\n\r\n def __hash__(self) -> int:\r\n # NOTE This hashing operation means that equivalent entries\r\n # hash to different values.\r\n return id(self)\r\n```\r\n\r\nso the hashes are slightly different but both based on the id.\r\n\r\nI would propose for all we have something like:\r\n\r\n```python\r\n def __hash__(self):\r\n data_md5 = hashlib.md5((\r\n f\"{str(self.__class__.__name__)}\"\r\n f\"{self.composition.reduced_formula}\"\r\n f\"{self._energy}\").encode('utf-8')\r\n ).hexdigest()\r\n return int(data_md5, 16)\r\n```\r\n\r\nas the hash.\r\n\r\nThe `__eq__` is less straightforward. I think @shyamd 's idea of comparing on something cheap like the `entry_id` first instead of going straight to comparing the dictionary representations. However, in `PDEntry` there isn't the same separation between `entry_id` and `name`. If `name` isn't given then it's set to the `reduced_formula` so I wouldn't be sure what you would want to do there. Likewise for `ComputedEntry` equality based on the dictionary seems as though it might be quite heavy but I am not familiar with how `ComputedEntry` is used.",
"I realized yesterday that this bug actually affects our ability to run `ThermoBuilder`, so I'd like to elevate it's priority. If we can come to a consensus here about the best fix I'll be happy to implement it.\r\n\r\nFrom my perspective, I think we're looking for a solution that\r\n\r\n- makes minimal changes to what exists now\r\n- Puts as much of the logic in the `Entry` abstract base class as possible, to maximize consistency between different types of entries\r\n- maintains the mutability of `ComputedEntry`, so that energy corrections and `normalize` continue to work the same way.\r\n\r\nWith respect to scaling compositions - I feel that there is a difference between a `ComputedEntry` with one composition (e.g., LiCl) and another with double its composition and energy (e.g., Li2Cl2), so those two should not be considered equal. Arguably, they could be considered equal as `PDEntry` for the purposes of `PhaseDiagram` construction, but from a practical perspective I think many users will build `PhaseDiagram` using `ComputedEntry` rather than `PDEntry`, so it would be best not to have too many inconsistencies in how we treat their `__eq__` and `__hash__`",
"@mattmcdermott do you have any thoughts on this issue?",
"can we link this issue to #1974 and move the discussion there? The critical path there is also `__hash__` and `__eq__`. That PR is maybe mostly in a merge-able state but would appreciate feedback on the wider implications of the proposed changes made to `Entry` classes.\r\n\r\n> With respect to scaling compositions - I feel that there is a difference between a ComputedEntry with one composition (e.g., LiCl) and another with double its composition and energy (e.g., Li2Cl2), so those two should not be considered equal.\r\n\r\nI agree that they shouldn't be equal which was the my intention by adding the out of place normalization checks so they could be functionally equivalent when needed but otherwise not equal (this was how it worked for `PDEntry`. I have reverted the change that was causing this issue originally in `get_equilibrium_reaction_energy`. If users wanted this functionality with the normalization check they could still use `get_quasi_e_from_hull(stable_only=True)` which should also work for `ComputedEntry` given the changes to `__eq__` and `__hash__` - however, I should add a test for this probably.",
"Sure, happy to move the discussion to #1974 ",
"@rkingsbury can this be closed?",
"Fixed by #2014 . Thanks for all the patient work on this @CompRhys !"
] | 2020-10-20T04:10:58
| 2021-01-27T15:26:18
|
2021-01-27T15:26:18Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
`PhaseDiagram.get_equlibrium_reaction_energy()` fails when called on a stable entry from the phase diagram, and raises
```ValueError: None is unstable, the equilibrium reaction energy is available only for stable entries```
**To Reproduce**
```
with MPRester() as m:
entries = m.get_entries_in_chemsys("Zn-S")
pd = PhaseDiagram(entries)
pd.get_equilibrium_reaction_energy([e for e in pd.stable_entries][0])
```
**Desktop (please complete the following information):**
- OS: Mac
- Version 2020.10.9
**Additional context**
The problem is caused by `entry.normalize(inplace=False)`, here:
https://github.com/materialsproject/pymatgen/blob/055805bc5e58a9951add78c441f5cb275da60679/pymatgen/analysis/phase_diagram.py#L665
`self.get_stable_entries_normed()` is populated by calling `entry.normalize(inplace=False)`, which creates copies of each entry. `entry.normalize(inplace=False)` creates _another_ copy of the entry that is not in the list.
I am working on a fix.
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1985/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1985/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1986
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1986/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1986/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1986/events
|
https://github.com/materialsproject/pymatgen/pull/1986
| 725,814,231
|
MDExOlB1bGxSZXF1ZXN0NTA2OTg4ODM1
| 1,986
|
Add f* diagram generator
|
{
"login": "jonathanjdenney",
"id": 48103676,
"node_id": "MDQ6VXNlcjQ4MTAzNjc2",
"avatar_url": "https://avatars.githubusercontent.com/u/48103676?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jonathanjdenney",
"html_url": "https://github.com/jonathanjdenney",
"followers_url": "https://api.github.com/users/jonathanjdenney/followers",
"following_url": "https://api.github.com/users/jonathanjdenney/following{/other_user}",
"gists_url": "https://api.github.com/users/jonathanjdenney/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jonathanjdenney/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jonathanjdenney/subscriptions",
"organizations_url": "https://api.github.com/users/jonathanjdenney/orgs",
"repos_url": "https://api.github.com/users/jonathanjdenney/repos",
"events_url": "https://api.github.com/users/jonathanjdenney/events{/privacy}",
"received_events_url": "https://api.github.com/users/jonathanjdenney/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 3093102097,
"node_id": "MDU6TGFiZWwzMDkzMTAyMDk3",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/stale",
"name": "stale",
"color": "A99B62",
"default": false,
"description": "Abandoned or conflicting PRs and outdated issues"
}
] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @jonathanjdenney, thanks for this.\r\n\r\nA few comments before merging, mostly just to follow Python conventions:\r\n\r\n* The file `pymatgen/analysis/Fstar/__pycache__/fstar.cpython-36.pyc` should be removed\r\n* The folder `pymatgen/analysis/Fstar/` should be renamed to `pymatgen/analysis/fstar/`\r\n* The files `neutron_factors.csv` `xray_factors_2.csv` are duplicated in the `tests` folder, this is unnecessary.\r\n* `FStarDiagram_test` -> `test_fstardiagram` (method names should be lower case)\r\n* `x_ray_scatter_df`, `neutron_scatter_df` should be `X_RAY_SCATTER_DF`, `NEUTRON_SCATTER_DF` (global variables upper case)\r\n* instead of `try` `except AttributeError` you can do an `if` statement, e.g. `if isinstance(sp, Species` or `if hasattr(sp, \"element\")` etc.\r\n\r\nI see changes were made via direct file upload, I highly recommend [GitHub Desktop](https://desktop.github.com) for making easier edits.\r\n\r\nThese are all my suggestions for now, but I'll take another look after the above changes.",
"Also, I would add a reference to the papers using f* (e.g. https://doi.org/10.1063/1.5044555 https://doi.org/10.1021/acs.chemmater.9b03646) to the docstring to point newcomers in the right direction.",
"Hi @mkhorton ,\r\nI will get to work on your suggestions. Thanks for looking at it.",
"Hi @mkhorton ,\r\nI believe I have addressed all of your comments. Please let me know if I missed anything.",
"Thanks @jonathanjdenney ! I'll take another look. Were some tests removed in the most recent iteration?",
"@mkhorton No tests were actually removed. I had previously accidentally added a file that was a non-functioning version of the test file which I have now removed. ",
"@mkhorton \r\nHello Matt,\r\nSorry for not getting to your comments sooner, I've had other deadlines. The purpose of including a way to input cifs directly into the script was to make it more user friendly to our target audience, experimentalists. Experimentalists will typically be more familiar with cifs over structure objects. In addition, this method allows the user to easily input non-physical values for the occupancies which may have resulted from the refinement. As far as I know, the cif parser will round greater than 1 occupancies to 1 regardless of what tolerance you set. This script can take structure objects as its input without any trouble, so is it really unacceptable that I've given the option to use cifs? \r\nIn regards to your last comment I agree it is a bit confusing. I will add to the code description an explanation that you use the labels generated in the site_labels list, which can be viewed by printing the list. It is input as a list of strings. \r\nBest,\r\nJonathan",
"> In addition, this method allows the user to easily input non-physical values for the occupancies which may have resulted from the refinement. As far as I know, the cif parser will round greater than 1 occupancies to 1 regardless of what tolerance you set. \r\n\r\nIf there's a case where unphysical occupancies are useful, then we should change `CifParser` accordingly.\r\n\r\n> The purpose of including a way to input cifs directly into the script was to make it more user friendly to our target audience, experimentalists.\r\n> This script can take structure objects as its input without any trouble, so is it really unacceptable that I've given the option to use cifs?\r\n\r\nMaybe to explain the issue a different way, pymatgen is intended as a library. For someone using it, regardless of their background or expertise (I started off as an experimentalist myself), you have to learn how to use Python, how to install the package, how to import the relevant functionality you want etc.\r\n\r\nTherefore, with the recommendation to people using the script will be to say, \"first you write one line to import your CIFs\" and then \"you run the FStarDiagram generator\", in my own view the extra directive to write the additional line is not the limiting hurdle for using a library like pymatgen.\r\n\r\nThe objection to having the CIF input is that it creates confusion and makes it more difficult to reason about what the code is doing internally. The existence of the Structure object is not to create yet another file format (e.g. on Materials Project, we offer CIF downloads, not Structure downloads), it's to create a standardized interface to crystal structures internally within pymatgen.\r\n\r\nMore importantly however, this discussion has highlighted a limitation (the CIF unphysical occupancies mentioned), and we'd be papering over this limitation rather than addressing it properly. By adding the capability to CifParser, other people can also have the benefit of importing CIF with unphysical occupancies, and the resulting code within FStarDiagram will be a lot simpler too."
] | 2020-10-20T17:51:34
| 2023-02-07T19:13:33
|
2023-02-07T19:13:32Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Added f* diagram generator.
## Additional dependencies introduced (if any)
* List all new dependencies needed and justify why. While adding dependencies that bring
significantly useful functionality is perfectly fine, adding ones that
add trivial functionality, e.g., to use one single easily implementable
function, is frowned upon. Provide a justification why that dependency is needed.
Especially frowned upon are circular dependencies, e.g., depending on derivative
modules of pymatgen such as custodian or Fireworks.
## TODO (if any)
If this is a work-in-progress, write something about what else needs
to be done
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "jonathanjdenney",
"id": 48103676,
"node_id": "MDQ6VXNlcjQ4MTAzNjc2",
"avatar_url": "https://avatars.githubusercontent.com/u/48103676?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jonathanjdenney",
"html_url": "https://github.com/jonathanjdenney",
"followers_url": "https://api.github.com/users/jonathanjdenney/followers",
"following_url": "https://api.github.com/users/jonathanjdenney/following{/other_user}",
"gists_url": "https://api.github.com/users/jonathanjdenney/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jonathanjdenney/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jonathanjdenney/subscriptions",
"organizations_url": "https://api.github.com/users/jonathanjdenney/orgs",
"repos_url": "https://api.github.com/users/jonathanjdenney/repos",
"events_url": "https://api.github.com/users/jonathanjdenney/events{/privacy}",
"received_events_url": "https://api.github.com/users/jonathanjdenney/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1986/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1986/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1986",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1986",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1986.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1986.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1987
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1987/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1987/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1987/events
|
https://github.com/materialsproject/pymatgen/issues/1987
| 727,613,298
|
MDU6SXNzdWU3Mjc2MTMyOTg=
| 1,987
|
BSPlotterProjected not work for d orbital
|
{
"login": "Yang-YM",
"id": 48523891,
"node_id": "MDQ6VXNlcjQ4NTIzODkx",
"avatar_url": "https://avatars.githubusercontent.com/u/48523891?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Yang-YM",
"html_url": "https://github.com/Yang-YM",
"followers_url": "https://api.github.com/users/Yang-YM/followers",
"following_url": "https://api.github.com/users/Yang-YM/following{/other_user}",
"gists_url": "https://api.github.com/users/Yang-YM/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Yang-YM/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Yang-YM/subscriptions",
"organizations_url": "https://api.github.com/users/Yang-YM/orgs",
"repos_url": "https://api.github.com/users/Yang-YM/repos",
"events_url": "https://api.github.com/users/Yang-YM/events{/privacy}",
"received_events_url": "https://api.github.com/users/Yang-YM/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I run that code on a vasprun that contains d orbilals projection without any error.\r\nCould you check that your vasprun has the needed data? In positive case,\r\nsend me that vasprun so that I can reproduce the error. @Yang-YM\r\n"
] | 2020-10-22T18:12:04
| 2023-08-13T16:33:48
|
2023-08-13T16:33:48Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
When I want to plot the projected band structure using `BSPlotterProjected`, I find it not work if the `dictio` has d orbitals.
It gives the error :
```
C:\ProgramData\Anaconda3\lib\site-packages\pymatgen\electronic_structure\plotter.py:1058: MatplotlibDeprecationWarning: Adding an axes using the same arguments as a previous axes currently reuses the earlier instance. In a future version, a new instance will always be created and returned. Meanwhile, this warning can be suppressed, and the future behavior ensured, by passing a unique label to each axes instance.
\plt.subplot(fig_rows+fig_cols+count)
Traceback (most recent call last):
File ".\band-dos.py", line 41, in <module>
pband_fig.get_projected_plots_dots({'S':['d']}, ylim=(-2,4))
File "C:\ProgramData\Anaconda3\lib\site-packages\pymatgen\electronic_structure\plotter.py", line 1088, in get_projected_plots_dots
markersize=proj[b][str(Spin.up)][i][j][str(el)][o]
KeyError: 'd'
```
According to follow the code, I find the function `get_projections_on_elements_and_orbitals(self, el_orb_spec)` in class `BandStructure` can not return the information of d orbitals. I want to know how to solve it
This is my code
```
bs_run = Vasprun("./vasprun.xml", parse_projected_eigen=True)
bs_data = bs_run.get_band_structure("./KPOINTS",line_mode=True)
pband_fig = BSPlotterProjected(bs=bs_data)
pband_fig.get_projected_plots_dots({'Cu':['d']}, ylim=(-2,4))
plt.savefig('pband_orbital_fig.png',img_format='png',dpi=300)
plt.show()
```
- OS: Windows
- Version 2020.10.20
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1987/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1987/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1988
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1988/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1988/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1988/events
|
https://github.com/materialsproject/pymatgen/issues/1988
| 728,978,320
|
MDU6SXNzdWU3Mjg5NzgzMjA=
| 1,988
|
If there any way to get x-y axis non-orthogonal slabs?
|
{
"login": "lixinyuu",
"id": 33710988,
"node_id": "MDQ6VXNlcjMzNzEwOTg4",
"avatar_url": "https://avatars.githubusercontent.com/u/33710988?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/lixinyuu",
"html_url": "https://github.com/lixinyuu",
"followers_url": "https://api.github.com/users/lixinyuu/followers",
"following_url": "https://api.github.com/users/lixinyuu/following{/other_user}",
"gists_url": "https://api.github.com/users/lixinyuu/gists{/gist_id}",
"starred_url": "https://api.github.com/users/lixinyuu/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/lixinyuu/subscriptions",
"organizations_url": "https://api.github.com/users/lixinyuu/orgs",
"repos_url": "https://api.github.com/users/lixinyuu/repos",
"events_url": "https://api.github.com/users/lixinyuu/events{/privacy}",
"received_events_url": "https://api.github.com/users/lixinyuu/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"In pymatgen's slab generator, the surface is in the x-y direction. The only question is whether the z direction is constrained to be orthogonal or not. By default the SlabGenerator makes non-orthgonal slabs that are the smallest possible for that surface. If you use Slab.get_orthogonal_c_slab, you then get the orthogonal version.",
"In future, please use https://matsci.org/ for queries on usage. Github Issues is meant for reporting bugs and requesting new features. Thanks.",
"Thanks, Prof. Ong."
] | 2020-10-25T09:20:20
| 2020-10-25T15:04:33
|
2020-10-25T14:49:46Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hi pymatgen developers,
I have lots of x-y axis orthorhombic slabs which is constructed by SlabGenerator. However, my problem is sensitive to the slab x-y angle (gamma) thus I would like to check if these slabs could be transformed into non-orthorhombic slabs.
The following picture (get from https://wiki.fysik.dtu.dk/ase/ase/build/surface.html) shows what I want to do. So I have lots of orthorhombic slabs like the right slabs, I would like to know if they could be reshaped into non-orthorhombic ones. I have checked both SlabGenerator and SpacegroupAnalyzer, but didn't find way to do this.
Could you please give me some suggestions? Thanks in advance.
![Screensho![Uploading Screenshot 2020-10-25 201005.png…]()t 2020-10-25 201005](https://user-images.githubusercontent.com/33710988/97103188-fe5dda80-16fe-11eb-99bd-b387e69b84b0.png)
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1988/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1988/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1989
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1989/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1989/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1989/events
|
https://github.com/materialsproject/pymatgen/pull/1989
| 730,018,088
|
MDExOlB1bGxSZXF1ZXN0NTEwMzk4MzE3
| 1,989
|
Fix outcar serialization
|
{
"login": "utf",
"id": 1330638,
"node_id": "MDQ6VXNlcjEzMzA2Mzg=",
"avatar_url": "https://avatars.githubusercontent.com/u/1330638?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/utf",
"html_url": "https://github.com/utf",
"followers_url": "https://api.github.com/users/utf/followers",
"following_url": "https://api.github.com/users/utf/following{/other_user}",
"gists_url": "https://api.github.com/users/utf/gists{/gist_id}",
"starred_url": "https://api.github.com/users/utf/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/utf/subscriptions",
"organizations_url": "https://api.github.com/users/utf/orgs",
"repos_url": "https://api.github.com/users/utf/repos",
"events_url": "https://api.github.com/users/utf/events{/privacy}",
"received_events_url": "https://api.github.com/users/utf/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This should be ready to merge btw.",
"Thanks @utf"
] | 2020-10-27T01:12:14
| 2020-10-28T22:59:57
|
2020-10-28T22:27:04Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
`Outcar.as_dict` method fails if the OUTCAR doesn't contain the average electrostatic potential at atomic cores. This can occur if `ICORELEVEL = 1` is set. I've fixed this behaviour and added a test.
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1989/reactions",
"total_count": 2,
"+1": 2,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1989/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1989",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1989",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1989.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1989.patch",
"merged_at": "2020-10-28T22:27:04Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1990
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1990/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1990/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1990/events
|
https://github.com/materialsproject/pymatgen/issues/1990
| 733,472,248
|
MDU6SXNzdWU3MzM0NzIyNDg=
| 1,990
|
No module named 'pymatgen.symmetry.kpath'
|
{
"login": "manishkothakonda",
"id": 32824488,
"node_id": "MDQ6VXNlcjMyODI0NDg4",
"avatar_url": "https://avatars.githubusercontent.com/u/32824488?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/manishkothakonda",
"html_url": "https://github.com/manishkothakonda",
"followers_url": "https://api.github.com/users/manishkothakonda/followers",
"following_url": "https://api.github.com/users/manishkothakonda/following{/other_user}",
"gists_url": "https://api.github.com/users/manishkothakonda/gists{/gist_id}",
"starred_url": "https://api.github.com/users/manishkothakonda/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/manishkothakonda/subscriptions",
"organizations_url": "https://api.github.com/users/manishkothakonda/orgs",
"repos_url": "https://api.github.com/users/manishkothakonda/repos",
"events_url": "https://api.github.com/users/manishkothakonda/events{/privacy}",
"received_events_url": "https://api.github.com/users/manishkothakonda/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This is too brief an error message for us to debug. I don't get this error."
] | 2020-10-30T19:48:10
| 2020-10-30T19:57:08
|
2020-10-30T19:57:08Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
It says the module is not available, I have been using pymatgen for a year and have not came across anything like this before.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1990/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1990/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1991
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1991/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1991/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1991/events
|
https://github.com/materialsproject/pymatgen/pull/1991
| 737,069,446
|
MDExOlB1bGxSZXF1ZXN0NTE2MTc2NDg4
| 1,991
|
Change convention used to generate Abinit typat/znucl variables
|
{
"login": "gmatteo",
"id": 3811747,
"node_id": "MDQ6VXNlcjM4MTE3NDc=",
"avatar_url": "https://avatars.githubusercontent.com/u/3811747?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/gmatteo",
"html_url": "https://github.com/gmatteo",
"followers_url": "https://api.github.com/users/gmatteo/followers",
"following_url": "https://api.github.com/users/gmatteo/following{/other_user}",
"gists_url": "https://api.github.com/users/gmatteo/gists{/gist_id}",
"starred_url": "https://api.github.com/users/gmatteo/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/gmatteo/subscriptions",
"organizations_url": "https://api.github.com/users/gmatteo/orgs",
"repos_url": "https://api.github.com/users/gmatteo/repos",
"events_url": "https://api.github.com/users/gmatteo/events{/privacy}",
"received_events_url": "https://api.github.com/users/gmatteo/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks. Can you uncomment abinit in the setup.cfg and make sure the pydocstyle tests pass too? Thanks.",
"Hi, I've removed abinit from setup.cfg and fixed the doc strings as requested. All tests seem to be OK. ",
"Thanks."
] | 2020-11-05T16:14:30
| 2020-11-06T16:02:55
|
2020-11-06T16:02:55Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Use deterministic algorithm (not based on python set) to obtain list of elements before writing Abinit input file.
This is required to maintain compatibility with the original implementation of structure.types_of_specie and all the DFPT
results produced so far.
* Add refine_struct optional argument to CifWriter
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1991/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1991/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1991",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1991",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1991.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1991.patch",
"merged_at": "2020-11-06T16:02:55Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1992
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1992/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1992/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1992/events
|
https://github.com/materialsproject/pymatgen/pull/1992
| 737,853,250
|
MDExOlB1bGxSZXF1ZXN0NTE2ODIyNTQ2
| 1,992
|
Fix bug in PhononBandStructureSymmLine
|
{
"login": "gpetretto",
"id": 8665734,
"node_id": "MDQ6VXNlcjg2NjU3MzQ=",
"avatar_url": "https://avatars.githubusercontent.com/u/8665734?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/gpetretto",
"html_url": "https://github.com/gpetretto",
"followers_url": "https://api.github.com/users/gpetretto/followers",
"following_url": "https://api.github.com/users/gpetretto/following{/other_user}",
"gists_url": "https://api.github.com/users/gpetretto/gists{/gist_id}",
"starred_url": "https://api.github.com/users/gpetretto/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/gpetretto/subscriptions",
"organizations_url": "https://api.github.com/users/gpetretto/orgs",
"repos_url": "https://api.github.com/users/gpetretto/repos",
"events_url": "https://api.github.com/users/gpetretto/events{/privacy}",
"received_events_url": "https://api.github.com/users/gpetretto/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @gpetretto :-)"
] | 2020-11-06T15:35:43
| 2020-11-07T02:58:47
|
2020-11-07T02:58:45Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Small bug fix in PhononBandStructureSymmLine. Added tests.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1992/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1992/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1992",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1992",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1992.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1992.patch",
"merged_at": "2020-11-07T02:58:45Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1993
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1993/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1993/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1993/events
|
https://github.com/materialsproject/pymatgen/issues/1993
| 742,115,893
|
MDU6SXNzdWU3NDIxMTU4OTM=
| 1,993
|
ValueError when using get_decomposition_energy for computed entries in Pourbaix diagrams
|
{
"login": "CJBR-97",
"id": 69945487,
"node_id": "MDQ6VXNlcjY5OTQ1NDg3",
"avatar_url": "https://avatars.githubusercontent.com/u/69945487?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CJBR-97",
"html_url": "https://github.com/CJBR-97",
"followers_url": "https://api.github.com/users/CJBR-97/followers",
"following_url": "https://api.github.com/users/CJBR-97/following{/other_user}",
"gists_url": "https://api.github.com/users/CJBR-97/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CJBR-97/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CJBR-97/subscriptions",
"organizations_url": "https://api.github.com/users/CJBR-97/orgs",
"repos_url": "https://api.github.com/users/CJBR-97/repos",
"events_url": "https://api.github.com/users/CJBR-97/events{/privacy}",
"received_events_url": "https://api.github.com/users/CJBR-97/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"The phase used in the argument of Pourbaix.get_decomposition_energy must have a ratio of non-OH elements that matches the ratio corresponding to the pourbaix diagram in order to work. When you generate the pourbaix diagram, add a composition dictionary to match the entry whose stability you want to determine, e.g.\r\n\r\n```\r\npbx = PourbaixDiagram(entries, comp_dict={\"Zn\": 27/108, \"V\": 81/108})\r\n```",
"@CJBR-97 does this resolve your issue?",
"Yes this was great! Thank you.\r\n\r\nSorry for the late reply.",
"Hi,\r\n\r\nI get the same error even when I use a composition dictionary. Could you help me understand why? My code is:\r\n\r\n```\r\nels = ['O', 'Na', 'Si', 'Al']\r\nmp_id = \"mp-560334\"\r\ncomp_dict = {'Na': 0.14285714285714285, 'Al': 0.14285714285714285, 'Si': 0.14285714285714285, 'O': 0.5714285714285714}\r\n\r\nwith MPRester(api_key) as m:\r\n pe = m.get_pourbaix_entries(els) \r\n \r\ndia = pb.PourbaixDiagram(pe, comp_dict) \r\nplot = pb.PourbaixPlotter(dia)\r\n\r\nmpid_entry = [entry for entry in pe if entry.entry_id==mp_id][0] \r\ndecomp_e = dia.get_decomposition_energy(mpid_entry, pH, V)\r\n```\r\n\r\n\r\nThis gives the error:\r\n`ValueError: Composition of stability entry does not match Pourbaix Diagram`\r\nEverything works until I use dia.get_decomposition_energy",
"Hi @mfdresser, the `comp_dict` argument is meant to be a non-OH composition that pins the ratio of non-OH elements. Try using:\r\n\r\n```\r\ncomp_dict = {\"Na\": 1/3, \"Al\": 1/3, \"Si\": 1/3}\r\n```",
"As an aside for dev purposes - we could probably do a better job of preprocessing this argument, I don't see any issue removing OH elements from a supplied composition, and it would probably be useful to accept a composition, as opposed to only a dictionary, and to just call the argument that. I think the the holdover is just from legacy behavior.",
"@JosephMontoya-TRI I think it is better to issue an error (or at least a warning) if OH is detected. And yes, I agree a Comp is better. Though technically there shouldn't be mcuh of a difference since Comp is basically a dict-like object.",
"Hi, thanks for your response! So, even if my solid of interest is an oxide, I should still leave the O composition out of my composition dictionary? ",
"That's right - the comp_dict is intended to pin the ratio of non-OH elements, so it shouldn't include O or H.",
"Thank you! This fixed the problem."
] | 2020-11-13T03:59:33
| 2023-09-20T02:07:38
|
2020-11-22T22:05:32Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hello,
I am new to pymatgen and am trying to use the Pourbaix diagram features to obtain a decomposition energy for a computed entry. However I keep encountering the ValueError associated with the get_decomposition_energy() method:
_ValueError: Composition of stability entry does not match Pourbaix Diagram_
Is it possible to resolve this error?
Below is the code:
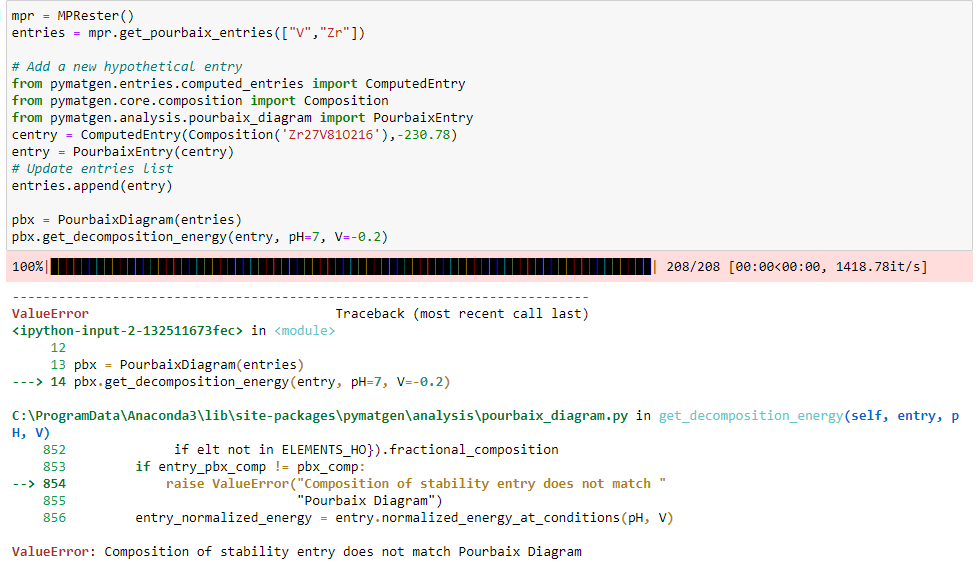
|
{
"login": "CJBR-97",
"id": 69945487,
"node_id": "MDQ6VXNlcjY5OTQ1NDg3",
"avatar_url": "https://avatars.githubusercontent.com/u/69945487?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CJBR-97",
"html_url": "https://github.com/CJBR-97",
"followers_url": "https://api.github.com/users/CJBR-97/followers",
"following_url": "https://api.github.com/users/CJBR-97/following{/other_user}",
"gists_url": "https://api.github.com/users/CJBR-97/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CJBR-97/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CJBR-97/subscriptions",
"organizations_url": "https://api.github.com/users/CJBR-97/orgs",
"repos_url": "https://api.github.com/users/CJBR-97/repos",
"events_url": "https://api.github.com/users/CJBR-97/events{/privacy}",
"received_events_url": "https://api.github.com/users/CJBR-97/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1993/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1993/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1994
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1994/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1994/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1994/events
|
https://github.com/materialsproject/pymatgen/issues/1994
| 743,938,562
|
MDU6SXNzdWU3NDM5Mzg1NjI=
| 1,994
|
Failed pymatgen import in python v3.8
|
{
"login": "deepanshs",
"id": 21365911,
"node_id": "MDQ6VXNlcjIxMzY1OTEx",
"avatar_url": "https://avatars.githubusercontent.com/u/21365911?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/deepanshs",
"html_url": "https://github.com/deepanshs",
"followers_url": "https://api.github.com/users/deepanshs/followers",
"following_url": "https://api.github.com/users/deepanshs/following{/other_user}",
"gists_url": "https://api.github.com/users/deepanshs/gists{/gist_id}",
"starred_url": "https://api.github.com/users/deepanshs/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/deepanshs/subscriptions",
"organizations_url": "https://api.github.com/users/deepanshs/orgs",
"repos_url": "https://api.github.com/users/deepanshs/repos",
"events_url": "https://api.github.com/users/deepanshs/events{/privacy}",
"received_events_url": "https://api.github.com/users/deepanshs/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I've also had this issue.",
"I just tried on my system and I cannot reproduce this. Can you try forcing a reinstall of ruamel?\r\n\r\n`pip install --force ruamel.yaml`",
"Forcing the reinstall gives me the following error messages:\r\n\r\nERROR: Cannot uninstall 'ruamel-yaml'. It is a distutils installed project and thus we cannot accurately determine which files belong to it which would lead to only a partial uninstall.",
"Hmmm.... are you using a virtual env?",
"I am. I have it activated for the pip install.",
"Honestly, I am unsure what is wrong. There are some suggestions that using the pip install --ignore-installed flag would work. Maybe you can try that.",
"Changing the ruamel.yaml import from:\r\n\r\nimport ruamel.yaml as yaml\r\n\r\nto: \r\n\r\nimport ruamel_yaml as yaml\r\n\r\n in the following files:\r\n1. __init__.py\" (line 14)\r\n2. ext/matproj.py (line 27)\r\n3. analysis/local_env.py (line 23)\r\n\r\nSolved it for me. It looks like the naming for ruamel just changed.",
"That's very odd then. Because I did ruamel.yaml and it works. And this is the latest ruamel.yaml version.\r\n\r\n```\r\nPython 3.8.5 (default, Sep 4 2020, 02:22:02)\r\n[Clang 10.0.0 ] :: Anaconda, Inc. on darwin\r\nType \"help\", \"copyright\", \"credits\" or \"license\" for more information.\r\n>>> import ruamel.yaml\r\n>>> ruamel.yaml.__version__\r\n'0.16.12'\r\n```",
"Hmm, that is weird. I just did a fresh install of Anaconda and Python. Same thing still happens.",
"Python 3.8.5 (default, Sep 4 2020, 02:22:02) \r\n[Clang 10.0.0 ] :: Anaconda, Inc. on darwin\r\nType \"help\", \"copyright\", \"credits\" or \"license\" for more information.\r\n>>> import ruamel.yaml\r\nTraceback (most recent call last):\r\n File \"<stdin>\", line 1, in <module>\r\nModuleNotFoundError: No module named 'ruamel.yaml'\r\n>>> import ruamel_yaml\r\n>>> ",
"Version of ruamel?",
">>> ruamel_yaml.__version__\r\n'0.15.87'",
"Ok, unclear what is happening. Apparently someone on anaconda did a patched version that uses ruamel_yaml. The official pip version is ruamel.yaml for sure. Anyway, this is an easy fix. ",
"Fix pushed."
] | 2020-11-16T15:40:17
| 2021-01-16T00:50:07
|
2021-01-16T00:50:00Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Error importing the pymatgen module.
```
>>> import pymatgen
---------------------------------------------------------------------------
ModuleNotFoundError Traceback (most recent call last)
<ipython-input-2-a3a39e60e980> in <module>
----> 1 import pymatgen
~/anaconda3/lib/python3.8/site-packages/pymatgen/__init__.py in <module>
12 from fnmatch import fnmatch
13
---> 14 import ruamel.yaml as yaml
15
16 from monty.json import MontyEncoder, MontyDecoder, MSONable
ModuleNotFoundError: No module named 'ruamel'
```
**To Reproduce**
1. Install pymatgen using pip: ``pip install pymatgen``
2. Single line code: ``import pymatgen``
**Expected behavior**
No error
**Desktop/System info**
- OS: macOS Big Sur, v11.0.1
- Python v3.8.5
**Other details**
1. `ruamel.yaml` is installed on python v3.8.
```
>>> pip show ruamel.yaml
Name: ruamel-yaml
Version: 0.15.87
Summary: ruamel_yaml is a YAML parser/emitter that supports roundtrip preservation of comments, seq/map flow style, and map key order
Home-page: UNKNOWN
Author: Anthon van der Neut
Author-email: a.van.der.neut@ruamel.eu
License: UNKNOWN
Location: ~/anaconda3/lib/python3.8/site-packages
Requires:
Required-by: pymatgen, conda, anaconda-project
```
2. The `import pymatgen` is successful on python v3.7.7
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1994/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1994/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1995
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1995/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1995/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1995/events
|
https://github.com/materialsproject/pymatgen/issues/1995
| 744,178,187
|
MDU6SXNzdWU3NDQxNzgxODc=
| 1,995
|
Reading WAVECAR for SOC is not working
|
{
"login": "aamirshafique2977",
"id": 51540002,
"node_id": "MDQ6VXNlcjUxNTQwMDAy",
"avatar_url": "https://avatars.githubusercontent.com/u/51540002?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/aamirshafique2977",
"html_url": "https://github.com/aamirshafique2977",
"followers_url": "https://api.github.com/users/aamirshafique2977/followers",
"following_url": "https://api.github.com/users/aamirshafique2977/following{/other_user}",
"gists_url": "https://api.github.com/users/aamirshafique2977/gists{/gist_id}",
"starred_url": "https://api.github.com/users/aamirshafique2977/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/aamirshafique2977/subscriptions",
"organizations_url": "https://api.github.com/users/aamirshafique2977/orgs",
"repos_url": "https://api.github.com/users/aamirshafique2977/repos",
"events_url": "https://api.github.com/users/aamirshafique2977/events{/privacy}",
"received_events_url": "https://api.github.com/users/aamirshafique2977/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 5304335471,
"node_id": "LA_kwDOACgets8AAAABPCm8bw",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/io",
"name": "io",
"color": "c2e0c6",
"default": false,
"description": "Input/output functionality"
},
{
"id": 5322879170,
"node_id": "LA_kwDOACgets8AAAABPUSwwg",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/help",
"name": "help",
"color": "5ED16E",
"default": false,
"description": "Q&A support-style issues"
},
{
"id": 5457739150,
"node_id": "LA_kwDOACgets8AAAABRU59jg",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/vasp",
"name": "vasp",
"color": "BF4B01",
"default": false,
"description": "Vienna Ab initio Simulation Package"
}
] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Can you provide your pymatgen version, VASP version, and a copy of the WAVECAR?",
"I have used pymatgen==2020.11.11 and vasp 5.4.4 version. I am unable to attach the WAVECAR file because the size of the file is very large (approx. 155 GB).",
"Do you have enough memory? The entire WAVECAR will be read in to memory. I've tested against a WAVECAR generated from 5.4.4, so it should work. Without the WAVECAR, I'm not really sure how to debug it for you.",
"Hi Mark, \r\n\r\nI have also run into the 'incorrect vasp_type'\r\n\r\n issue trying to convert WAVECAR to wavefunction.h5 file. The WAVECAR is generated by VASP 5.4.4 and is about 5 MB. So, I do not think mine is a memory issue. Could you test if you could convert my WAVECAR file? I have tested this with pymatgen installed on different OS (both linux and windows). Any insights will be much appreciated.\r\n\r\nThank you,\r\nManoj\r\n\r\n[WAVECAR.gz](https://github.com/materialsproject/pymatgen/files/6651818/WAVECAR.gz)\r\n \r\n",
"Hi @ManojSettipalli, I took a look at the file you sent. It seems that the number of plane waves that are read in from the WAVECAR at the first kpoint is 339, which is not evenly divisible by 2, so something is wrong. A noncollinear WAVECAR has spinors, so there should be twice as many plane waves as usual, one set for spin up and one set for down. By any chance, was this WAVECAR from a relaxation where the volume was also relaxed? That's my only guess as to what may be going wrong. Perhaps try doing a fixed geometry calculation from this WAVECAR to generate a new one and see if that fixes it.\r\n\r\nIf that's not the case, then I'm not really sure what might be wrong. I can look into it, but I'm quite busy at the moment, so I may not get back to you for some time.",
"Hi Mark,\r\n\r\nThank you for your response. I should have mentioned that I got the error for an 'std' type calculation so it is not a noncollinear calculation. I still had this issue. As a last attempt, I fully uninstalled Python on my computer and reinstalled a new python on Conda, and then installed pymatgen. Now, the WAVECAR is being read. So, while I am not entirely sure what the issue was, I am just going with this for now. \r\n\r\nThank you,\r\nManoj",
"That's strange. I'm glad it worked out in the end.",
"I guess this problem may come from the precision of calculating `Gpoints` from `_generate_G_points` function.\r\nAs you can find in the code, the error is raised when the deduced `Gpoints` is mismatched with `nplane`.\r\nThe plane waves near the boundary of the cutoff radius (in energy) are critical to include as bases.\r\n\r\n```\r\n if len(self.Gpoints[ink]) != nplane and 2 * len(self.Gpoints[ink]) != nplane:\r\n raise ValueError(\r\n f\"Incorrect value of vasp_type given ({vasp_type}).\"\r\n \" Please open an issue if you are certain this WAVECAR\"\r\n \" was generated with the given vasp_type.\"\r\n )\r\n```\r\n\r\nSo, changing the k-point that encounters the error might solve the problem.\r\n\r\n---\r\n\r\n@mturiansky \r\n\r\nI found that three lines in the `outputs.py` script below could cause the problem.\r\n\r\n```\r\n for i in range(2 * self._nbmax[2] + 1):\r\n i3 = i - 2 * self._nbmax[2] - 1 if i > self._nbmax[2] else i\r\n for j in range(2 * self._nbmax[1] + 1):\r\n j2 = j - 2 * self._nbmax[1] - 1 if j > self._nbmax[1] else j\r\n for k in range(kmax):\r\n k1 = k - 2 * self._nbmax[0] - 1 if k > self._nbmax[0] else k\r\n if gamma and ((k1 == 0 and j2 < 0) or (k1 == 0 and j2 == 0 and i3 < 0)):\r\n continue\r\n G = np.array([k1, j2, i3])\r\n v = kpoint + G\r\n g = np.linalg.norm(np.dot(v, self.b))\r\n E = g**2 / self._C\r\n if E < self.encut:\r\n gpoints.append(G)\r\n if gamma and (k1, j2, i3) != (0, 0, 0):\r\n extra_gpoints.append(-G)\r\n extra_coeff_inds.append(G_ind)\r\n G_ind += 1\r\n```\r\n\r\n`i3 = i - 2 * self._nbmax[2] - 1 if i > self._nbmax[2] else i`\r\n`j2 = j - 2 * self._nbmax[1] - 1 if j > self._nbmax[1] else j`\r\n`k1 = k - 2 * self._nbmax[0] - 1 if k > self._nbmax[0] else k`\r\n\r\nThree lines could be critical with the if conditions. During the calculation of `Gpoints`, the vector G must be in the 1st Brillouin zone. The folding way, `if condition in three lines`, causes the lack of `Gpoints`.\r\n\r\nSome K-points outside the 1st Brillouin zone were set in my WAVECAR.\r\n~~It seems that an even element in `self._nbmax` causes the error.~~\r\n\r\n___\r\n\r\nReplacing `_generate_G_points` with the code below works for my specific WAVECAR by accident, but I could understand how it works.\r\nI confirmed the way that the code reads `Gpoints` matches the way VASP reads `Gpoints` (only for my case).\r\n```\r\n def _generate_G_points(self, kpoint: np.ndarray, gamma: bool = False) -> tuple[list, list, list]:\r\n \"\"\"\r\n Helper function to generate G-points based on nbmax.\r\n This function iterates over possible G-point values and determines\r\n if the energy is less than G_{cut}. Valid values are appended to\r\n the output array. This function should not be called outside of\r\n initialization.\r\n Args:\r\n kpoint (np.array): the array containing the current k-point value\r\n gamma (bool): determines if G points for gamma-point only executable\r\n should be generated\r\n Returns:\r\n a list containing valid G-points\r\n \"\"\"\r\n if gamma:\r\n kmax = self._nbmax[0] + 1\r\n else:\r\n kmax = 2 * self._nbmax[0] + 1\r\n\r\n gpoints = []\r\n extra_gpoints = []\r\n extra_coeff_inds = []\r\n G_ind = 0\r\n for i in range(2 * self._nbmax[2] + 1):\r\n i3 = i - 2 * self._nbmax[2] - 1 if i > self._nbmax[2] else i\r\n for j in range(2 * self._nbmax[1] + 1):\r\n j2 = j - 2 * self._nbmax[1] - 1 if j > self._nbmax[1] else j\r\n for k in range(kmax):\r\n k1 = k - 2 * self._nbmax[0] - 1 if k > self._nbmax[0] else k\r\n if gamma and ((k1 == 0 and j2 < 0) or (k1 == 0 and j2 == 0 and i3 < 0)):\r\n continue\r\n G = np.array([k1, j2, i3])\r\n v = kpoint + G\r\n g = np.linalg.norm(np.dot(v, self.b))\r\n E = g**2 / self._C\r\n if E < self.encut:\r\n gpoints.append(G)\r\n if gamma and (k1, j2, i3) != (0, 0, 0):\r\n extra_gpoints.append(-G)\r\n extra_coeff_inds.append(G_ind)\r\n G_ind += 1\r\n elif i == self._nbmax[2]:\r\n i3 = i - 2 * self._nbmax[2] - 1\r\n if gamma and ((k1 == 0 and j2 < 0) or (k1 == 0 and j2 == 0 and i3 < 0)):\r\n continue\r\n G = np.array([k1, j2, i3])\r\n v = kpoint + G\r\n g = np.linalg.norm(np.dot(v, self.b))\r\n E = g**2 / self._C\r\n if E < self.encut:\r\n gpoints.append(G)\r\n if gamma and (k1, j2, i3) != (0, 0, 0):\r\n extra_gpoints.append(-G)\r\n extra_coeff_inds.append(G_ind)\r\n G_ind += 1\r\n elif j == self._nbmax[1]:\r\n j2 = j - 2 * self._nbmax[1] - 1\r\n if gamma and ((k1 == 0 and j2 < 0) or (k1 == 0 and j2 == 0 and i3 < 0)):\r\n continue\r\n G = np.array([k1, j2, i3])\r\n v = kpoint + G\r\n g = np.linalg.norm(np.dot(v, self.b))\r\n E = g**2 / self._C\r\n if E < self.encut:\r\n gpoints.append(G)\r\n if gamma and (k1, j2, i3) != (0, 0, 0):\r\n extra_gpoints.append(-G)\r\n extra_coeff_inds.append(G_ind)\r\n G_ind += 1\r\n elif k == self._nbmax[0]:\r\n k1 = k - 2 * self._nbmax[0] - 1\r\n if gamma and ((k1 == 0 and j2 < 0) or (k1 == 0 and j2 == 0 and i3 < 0)):\r\n continue\r\n G = np.array([k1, j2, i3])\r\n v = kpoint + G\r\n g = np.linalg.norm(np.dot(v, self.b))\r\n E = g**2 / self._C\r\n if E < self.encut:\r\n gpoints.append(G)\r\n if gamma and (k1, j2, i3) != (0, 0, 0):\r\n extra_gpoints.append(-G)\r\n extra_coeff_inds.append(G_ind)\r\n G_ind += 1\r\n return (gpoints, extra_gpoints, extra_coeff_inds)\r\n```\r\n\r\nI hope to share whether it works for other WAVECARs with the error.",
"@Nokimann Feel free to open a new issue if this is still an issue. I'll close this issue as completed for now.",
"I have encountered a similar problem on line 4345 of the vasp outputs.py module. I have explicitly included the correct precision and vasp binary type (accurate, ncl). The incar that produced it is the following:\r\n\r\n```\r\nALGO = Fast\r\nAMIN = 0.01\r\nEDIFF = 1e-06\r\nENCUT = 600 eV\r\nGGA = PE\r\nISMEAR = 0\r\nLASPH = True\r\nLDAU = False\r\nLMAXMIX = 4\r\nLORBIT = 11\r\nLREAL = Auto\r\nLSORBIT = True\r\nLWAVE = True\r\nMAGMOM = 100*0\r\nNELM = 100\r\nPREC = Accurate\r\nSIGMA = 0.05\r\n```\r\n\r\nDo keep in mind that I produced this WAVECAR using line mode, so perhaps that might be an issue? \r\n\r\nFor now, I have commented out the conditional statement to see if there are any further issues, as all I need is the charge density at a specific single kpoint. ",
"@wladerer feel free to open an issue with a repro if you can confirm there's a bug"
] | 2020-11-16T20:46:46
| 2024-02-22T07:45:41
|
2023-06-23T02:49:22Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I trying to load the spin-orbit coupling WAVCAR but failed. I got the following error message:
```py
Traceback (most recent call last):
File "/lustre/project/k1388/shafiq/CSLD-AMSET/install4/bin/amset", line 33, in <module>
sys.exit(load_entry_point('amset', 'console_scripts', 'amset')())
File "/lustre/project/k1388/shafiq/CSLD-AMSET/install4/lib/python3.7/site-packages/click/core.py", line 829, in __call__
return self.main(*args, **kwargs)
File "/lustre/project/k1388/shafiq/CSLD-AMSET/install4/lib/python3.7/site-packages/click/core.py", line 782, in main
rv = self.invoke(ctx)
File "/lustre/project/k1388/shafiq/CSLD-AMSET/install4/lib/python3.7/site-packages/click/core.py", line 1259, in invoke
return _process_result(sub_ctx.command.invoke(sub_ctx))
File "/lustre/project/k1388/shafiq/CSLD-AMSET/install4/lib/python3.7/site-packages/click/core.py", line 1066, in invoke
return ctx.invoke(self.callback, **ctx.params)
File "/lustre/project/k1388/shafiq/CSLD-AMSET/install4/lib/python3.7/site-packages/click/core.py", line 610, in invoke
return callback(*args, **kwargs)
File "/lustre/project/k1388/shafiq/CSLD-AMSET/install4/amset/amset/tools/wavefunction.py", line 74, in wave
coeffs, gpoints = _wavefunction_vasp(ibands, planewave_cutoff, **kwargs)
File "/lustre/project/k1388/shafiq/CSLD-AMSET/install4/amset/amset/tools/wavefunction.py", line 93, in _wavefunction_vasp
wf = get_wavefunction(wavecar=wavecar, directory=directory, vasp_type=vasp_type)
File "/lustre/project/k1388/shafiq/CSLD-AMSET/install4/amset/amset/wavefunction/vasp.py", line 28, in get_wavefunction
return Wavecar(wavecar, vasp_type=vasp_type)
File "/lustre/project/k1388/shafiq/CSLD-AMSET/install4/lib/python3.7/site-packages/pymatgen/io/vasp/outputs.py", line 4433, in __init__
raise ValueError(f'Incorrect value of vasp_type given ({vasp_type}).'
ValueError: Incorrect value of vasp_type given (ncl). Please open an issue if you are certain this WAVECAR was generated with the given vasp_type.
```
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1995/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1995/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1996
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1996/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1996/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1996/events
|
https://github.com/materialsproject/pymatgen/pull/1996
| 744,245,782
|
MDExOlB1bGxSZXF1ZXN0NTIyMDA0Njk0
| 1,996
|
Add tests and SCAN variant support to Vasprun.run_type
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I think we can make it stricter. Explicitly set the type to \"unknown\" with a warning if it is not within the list of defined types. ",
"Might be more robust to set it to the METAGGA tag if that tag is specified, in terms of if additional meta GGAs are added in future.",
"Specifically this line:\r\n\r\n```\r\n elif self.incar.get(\"METAGGA\", \"\").strip().upper() in METAGGA_TYPES:\r\n```\r\n\r\ncould instead be:\r\n\r\n```\r\n elif self.incar.get(\"METAGGA\"):\r\n rt = self.incar.get(\"METAGGA\").strip().upper()\r\n```",
"I would also break out the +U logic to a separate if statement, so it doesn't have to be duplicated, i.e. at the end of the method:\r\n\r\n```\r\nif self.is_hubbard or self.parameters.get(\"LDAU\", True):\r\n rt += \"+U\"\r\n```\r\n\r\nand maybe also add to the end:\r\n\r\n```\r\nIVDW_TYPES = {1: 'DFT-D2', 10: 'DFT-D2', 11: 'DFT-D3', 12: 'DFT-D3-BJ', 2: 'TS', 20: 'TS', 21: 'TS-H', 202: 'MBD', 4: 'dDsC'}\r\nif self.incar.get(\"IVDW\", \"\").strip().upper() in IVDW_TYPES:\r\n rt += \"vdW-\" + IVDW_TYPES[self.incar.get(\"IVDW\", \"\").strip().upper()]\r\nelif self.incar.get(\"IVDW\"):\r\n rt += \"vdW-unknown\"\r\n```\r\n\r\nsince this is an important case not captured by current logic.",
"(The `TODO: Fix for other functional types like PW91, other vdW types, etc.` could be removed then too)",
"@mkhorton @shyuep do you have any other comments / concerns here?",
"Thanks @rkingsbury! going to go ahead and merge"
] | 2020-11-16T22:34:18
| 2021-07-15T14:31:39
|
2020-12-01T16:32:50Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
At present, `OUTCAR.run_type` incorrectly sets the `run_type` to 'GGA' when METAGGA is equal to `R2SCAN` or `RSCAN`. This commit adds support for these functionals in `run_type` and also adds a test for regular `SCAN` run_type, which was not previously covered by a test.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1996/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1996/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1996",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1996",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1996.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1996.patch",
"merged_at": "2020-12-01T16:32:50Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1997
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1997/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1997/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1997/events
|
https://github.com/materialsproject/pymatgen/pull/1997
| 744,250,607
|
MDExOlB1bGxSZXF1ZXN0NTIyMDA5MjM5
| 1,997
|
added insertion site code and tests
|
{
"login": "jmmshn",
"id": 14003693,
"node_id": "MDQ6VXNlcjE0MDAzNjkz",
"avatar_url": "https://avatars.githubusercontent.com/u/14003693?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jmmshn",
"html_url": "https://github.com/jmmshn",
"followers_url": "https://api.github.com/users/jmmshn/followers",
"following_url": "https://api.github.com/users/jmmshn/following{/other_user}",
"gists_url": "https://api.github.com/users/jmmshn/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jmmshn/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jmmshn/subscriptions",
"organizations_url": "https://api.github.com/users/jmmshn/orgs",
"repos_url": "https://api.github.com/users/jmmshn/repos",
"events_url": "https://api.github.com/users/jmmshn/events{/privacy}",
"received_events_url": "https://api.github.com/users/jmmshn/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
] | null |
[
"Thanks @jmmshn, going to merge this."
] | 2020-11-16T22:38:04
| 2020-12-01T16:31:55
|
2020-12-01T16:31:55Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Added new class for general site-insertion algorithm
- Added ChargeInsertionAnalyzer class and tests
- Reformatted and ignoring .pre-commit-config.yaml
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1997/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1997/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1997",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1997",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1997.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1997.patch",
"merged_at": "2020-12-01T16:31:55Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1998
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1998/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1998/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1998/events
|
https://github.com/materialsproject/pymatgen/issues/1998
| 744,921,025
|
MDU6SXNzdWU3NDQ5MjEwMjU=
| 1,998
|
Calculated energy is not same with MP
|
{
"login": "haesunpark87",
"id": 43765756,
"node_id": "MDQ6VXNlcjQzNzY1NzU2",
"avatar_url": "https://avatars.githubusercontent.com/u/43765756?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/haesunpark87",
"html_url": "https://github.com/haesunpark87",
"followers_url": "https://api.github.com/users/haesunpark87/followers",
"following_url": "https://api.github.com/users/haesunpark87/following{/other_user}",
"gists_url": "https://api.github.com/users/haesunpark87/gists{/gist_id}",
"starred_url": "https://api.github.com/users/haesunpark87/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/haesunpark87/subscriptions",
"organizations_url": "https://api.github.com/users/haesunpark87/orgs",
"repos_url": "https://api.github.com/users/haesunpark87/repos",
"events_url": "https://api.github.com/users/haesunpark87/events{/privacy}",
"received_events_url": "https://api.github.com/users/haesunpark87/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@haesunpark87 I recently noticed the same issue. The problem was due to the different pseudopotentials used for Li_sv. The PAW_PBE Li_sv in MP should be the `23Jan2001` version. Maybe the Li_sv you are using was `10Sep2004`. @mkhorton Is this something significant? I also have friends who had the same issues and spent days debugging. ",
"I'm looking into it, it's difficult to debug this kind of issue, I will say that the MP database is by no means perfect and we have encountered changes in energy in non-deterministic ways which is always troubling, as a result of different VASP compilations, parallelization settings, etc.\r\n\r\nHowever, I re-ran the attached input and got the following as the final step:\r\n\r\n```\r\n 3 F= -.14263627E+02 E0= -.14263627E+02 d E =-.663574E-04 mag= -0.0000\r\n```\r\n\r\nvs the output from @haesunpark87:\r\n\r\n```\r\n 3 F= -.14347976E+02 E0= -.14347976E+02 d E =-.283537E-02 mag= -0.0000\r\n```\r\n\r\n@haesunpark87 My repeat is 80 meV different from your output but only 3 meV different from the MP database so perhaps you're using different pseudo potentials?\r\n\r\nAlso a small note on your script, you should be careful not to paste the API key publicly, and `str` is not a good variable name for a structure since it shadows the built-in Python string type (`str`). You can also use `write_input` on the input set object to write all the inputs (kpoints, incar, poscar, potcar) rather than having to write out each input individually.\r\n\r\n@chc273 please do let us know if anyone is spending days debugging, would definitely like to help if we can to save time.",
"@chc273 @mkhorton I'm sorry for the confusion, but I've been using the wrong version of POTCAR. The calculation which yields 30meV/atom discrepancy was done with POTCAR version of potpaw_PBE.52 which is not compatible with MP data base. After switching the POTCAR version to potpaw_PBE, the result was very similar to MP data base. ",
"Glad to hear you figured it out :)"
] | 2020-11-17T17:07:20
| 2021-03-09T02:23:49
|
2021-03-09T02:23:49Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I have calculated the energy of Li2O using the input setting made by **MPRelaxSet** module from pymatgen.
However, the calculated energy is way off from that in the MP data base(https://materialsproject.org/materials/mp-1960/#corrections-eqn).
The uncorrected energy differs by ~30meV/atom.
I used the POTCAR of Li_sv and O which are specified by pymatgen code and attempts with the different versions of vasp did not fix the issue.
I'm wondering what make this discrepancy and I attach the standard output(stdout_1), vasprun.xml file, and the python script generating the vasp inputs (vasp_input.py) for reference.
[outputs.tar.gz](https://github.com/materialsproject/pymatgen/files/5554990/outputs.tar.gz)
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1998/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1998/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1999
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1999/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1999/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1999/events
|
https://github.com/materialsproject/pymatgen/pull/1999
| 745,949,733
|
MDExOlB1bGxSZXF1ZXN0NTIzNDI1ODI4
| 1,999
|
imporve legibility of 3D phase diagram plot
|
{
"login": "bayesfactor",
"id": 16023856,
"node_id": "MDQ6VXNlcjE2MDIzODU2",
"avatar_url": "https://avatars.githubusercontent.com/u/16023856?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/bayesfactor",
"html_url": "https://github.com/bayesfactor",
"followers_url": "https://api.github.com/users/bayesfactor/followers",
"following_url": "https://api.github.com/users/bayesfactor/following{/other_user}",
"gists_url": "https://api.github.com/users/bayesfactor/gists{/gist_id}",
"starred_url": "https://api.github.com/users/bayesfactor/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/bayesfactor/subscriptions",
"organizations_url": "https://api.github.com/users/bayesfactor/orgs",
"repos_url": "https://api.github.com/users/bayesfactor/repos",
"events_url": "https://api.github.com/users/bayesfactor/events{/privacy}",
"received_events_url": "https://api.github.com/users/bayesfactor/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks for the improvement @bayesfactor, apologies for letting such a brief PR sit for a while.",
"Since you are a first-time contributor, please do complete [this form](https://forms.gle/JnisFb38QDR8QTFTA) so we can credit you appropriately in the [documentation](https://pymatgen.org/team.html). We ask all contributors to do this regardless of the size of the contribution.",
"I did complete the form. Thank you for accepting the PR, and thank you for your work in maintaining materialsproject, which is a fantastic asset to materials science. "
] | 2020-11-18T19:39:29
| 2020-12-01T21:23:30
|
2020-12-01T16:41:59Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
3D phase diagram plots have fonts that are so small they are nearly illegible. This PR has minor changes to aesthetics that take full use of the space in the plot and increase font size.
## Summary
Include a summary of major changes in bullet points:
* Larger fonts for component labels
* Changed 3D plot layout to make full use of the figure
## Additional dependencies introduced (if any)
None
* List all new dependencies needed and justify why. While adding dependencies that bring
significantly useful functionality is perfectly fine, adding ones that
add trivial functionality, e.g., to use one single easily implementable
function, is frowned upon. Provide a justification why that dependency is needed.
Especially frowned upon are circular dependencies, e.g., depending on derivative
modules of pymatgen such as custodian or Fireworks.
## TODO (if any)
None
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [ ] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1999/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1999/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1999",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1999",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1999.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1999.patch",
"merged_at": "2020-12-01T16:41:59Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2000
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2000/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2000/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2000/events
|
https://github.com/materialsproject/pymatgen/pull/2000
| 746,994,223
|
MDExOlB1bGxSZXF1ZXN0NTI0MjkwMTA3
| 2,000
|
improved Fermi level / bandgap detection in Vasprun
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@utf @mkhorton see `smart_efermi` method based on Alex's code. Fermi level seems to be set in the `init` method of `Outcar`, `BSVasprun`, and `Wavecar`. It is also read from the DOS in `Vasprun`.\r\n\r\nWould it make sense to default to this method inside `Vasprun.get_band_structure()`? Where else in outputs.py would we want to use this by default?",
"Aside from the comment above this seems ready to merge.",
"Thanks @rkingsbury "
] | 2020-11-19T23:05:36
| 2021-07-15T14:31:41
|
2020-12-04T01:53:24Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
- Implement a `smart_efermi` property in `Vasprun` to detect cases in which the Fermi level reported by VASP crosses one of the bands, but the bands otherwise indicate insulating behavior. This issue sometimes causes `Vasprun` to report inaccurate bandgaps (for example, insulators may be reported to have zero bandgap)
- Changes `Vasprun.get_bandstructure` to default to using the `smart_efermi` property to determine Fermi energy.
- Add type hinting and update docstrings in affected methods
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2000/reactions",
"total_count": 2,
"+1": 2,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2000/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2000",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2000",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2000.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2000.patch",
"merged_at": "2020-12-04T01:53:24Z"
}
|
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.