url
stringlengths 63
66
| repository_url
stringclasses 1
value | labels_url
stringlengths 77
80
| comments_url
stringlengths 72
75
| events_url
stringlengths 70
73
| html_url
stringlengths 52
56
| id
int64 3.39M
3.04B
| node_id
stringlengths 18
32
| number
int64 1
4.39k
| title
stringlengths 3
367
| user
dict | labels
listlengths 0
5
| state
stringclasses 2
values | locked
bool 2
classes | assignee
dict | assignees
listlengths 0
2
| milestone
null | comments
listlengths 0
30
| created_at
timestamp[s]date 2012-02-26 12:10:46
2025-05-02 18:11:41
| updated_at
timestamp[s]date 2012-02-26 14:52:41
2025-05-02 18:16:27
| closed_at
stringlengths 0
20
| author_association
stringclasses 3
values | type
stringclasses 1
value | active_lock_reason
stringclasses 2
values | sub_issues_summary
dict | body
stringlengths 0
119k
| closed_by
dict | reactions
dict | timeline_url
stringlengths 72
75
| performed_via_github_app
null | state_reason
stringclasses 4
values | is_pull_request
bool 2
classes | draft
bool 2
classes | pull_request
dict |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
https://api.github.com/repos/materialsproject/pymatgen/issues/2001
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2001/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2001/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2001/events
|
https://github.com/materialsproject/pymatgen/issues/2001
| 747,850,839
|
MDU6SXNzdWU3NDc4NTA4Mzk=
| 2,001
|
Error in .from_dict() for elfcar, aeccar0, aeccar2
|
{
"login": "ayushsgupta",
"id": 16586331,
"node_id": "MDQ6VXNlcjE2NTg2MzMx",
"avatar_url": "https://avatars.githubusercontent.com/u/16586331?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ayushsgupta",
"html_url": "https://github.com/ayushsgupta",
"followers_url": "https://api.github.com/users/ayushsgupta/followers",
"following_url": "https://api.github.com/users/ayushsgupta/following{/other_user}",
"gists_url": "https://api.github.com/users/ayushsgupta/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ayushsgupta/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ayushsgupta/subscriptions",
"organizations_url": "https://api.github.com/users/ayushsgupta/orgs",
"repos_url": "https://api.github.com/users/ayushsgupta/repos",
"events_url": "https://api.github.com/users/ayushsgupta/events{/privacy}",
"received_events_url": "https://api.github.com/users/ayushsgupta/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-11-20T23:04:45
| 2023-08-13T16:33:48
|
2023-08-13T16:33:48Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
When trying to call the `.from_dict()` method for elfcar, aeccar0, and aeccar2 files, the following error appears:
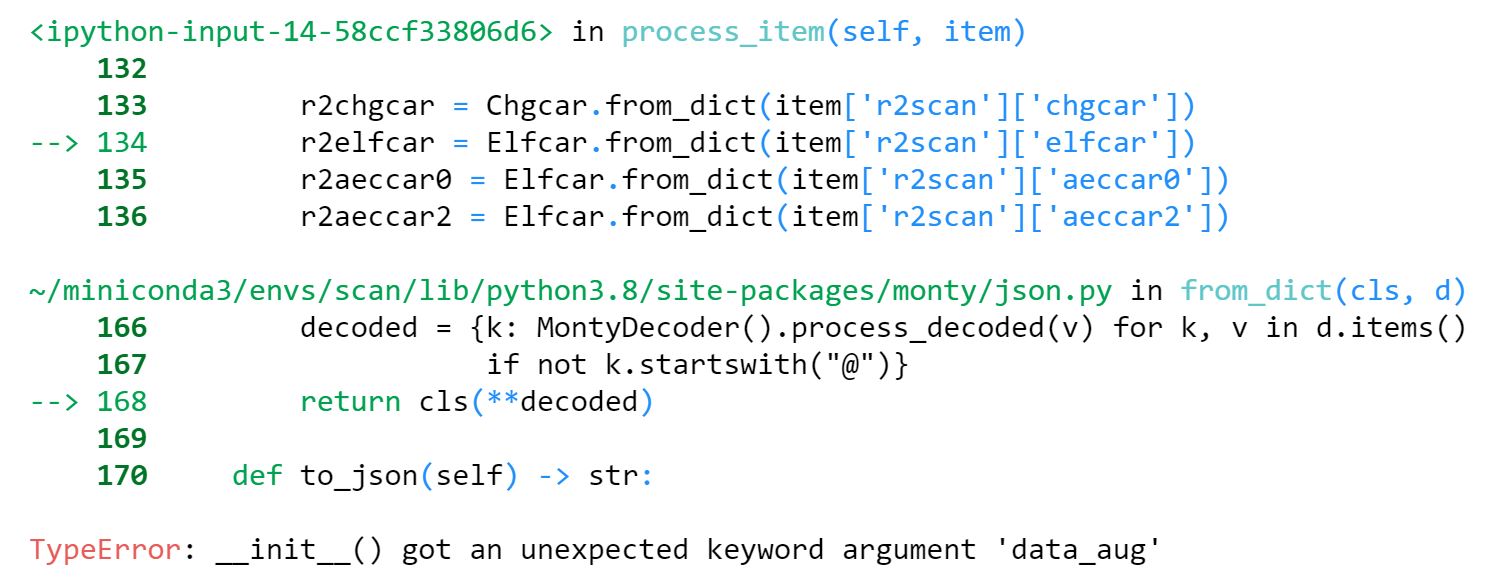
It seems like only the chgcar constructor accepts the `data_aug` argument, yet the dict forms of the other vasp file types contain this argument as well. The current fix is to remove the `data_aug` key manually before calling `.from_dict()`:

Not a very clean solution, so it's probably a good idea to get this fixed. Unfortunately I'm not too familiar with the workings of vasp so I'm not sure whether it's a good idea to prevent the `data_aug` key from being placed in the dict representation of an elfcar/aeccar0/aeccar2 in the first place, or to just accept it as an argument in the elfcar constructor and then ignore it.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2001/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2001/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2002
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2002/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2002/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2002/events
|
https://github.com/materialsproject/pymatgen/issues/2002
| 748,300,006
|
MDU6SXNzdWU3NDgzMDAwMDY=
| 2,002
|
Starting magnetization in quantum espresso (pwscf) file generator
|
{
"login": "bonfus",
"id": 15189887,
"node_id": "MDQ6VXNlcjE1MTg5ODg3",
"avatar_url": "https://avatars.githubusercontent.com/u/15189887?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/bonfus",
"html_url": "https://github.com/bonfus",
"followers_url": "https://api.github.com/users/bonfus/followers",
"following_url": "https://api.github.com/users/bonfus/following{/other_user}",
"gists_url": "https://api.github.com/users/bonfus/gists{/gist_id}",
"starred_url": "https://api.github.com/users/bonfus/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/bonfus/subscriptions",
"organizations_url": "https://api.github.com/users/bonfus/orgs",
"repos_url": "https://api.github.com/users/bonfus/repos",
"events_url": "https://api.github.com/users/bonfus/events{/privacy}",
"received_events_url": "https://api.github.com/users/bonfus/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-11-22T17:40:48
| 2023-08-13T16:33:48
|
2023-08-13T16:33:48Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I didn't really understand how `starting_magnetization` in PWInp is supposed to be specified other than in site_properties, but I eventually did not find a way to let the input generator fill the input with the data I specify in site properties.
This is the original code
https://github.com/materialsproject/pymatgen/blob/25a476e101abbd59affa6ef6b1ad0dfdc02a5e93/pymatgen/io/pwscf.py#L118-L120
and this is how I hacked it to make it work as (I) expected:
for k, v in sorted(site_descriptions.items(), key=lambda i: i[0]):
sub.append(" %s(%d) = %s" % (k2, n, v[k2]))
n += 1
Hope this helps!
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2002/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2002/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2003
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2003/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2003/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2003/events
|
https://github.com/materialsproject/pymatgen/issues/2003
| 751,480,351
|
MDU6SXNzdWU3NTE0ODAzNTE=
| 2,003
|
[Feature Request] Can Contributing.md be updated?
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-11-26T10:36:42
| 2020-12-07T16:45:22
|
2020-12-07T16:45:22Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I have been having issues with the `pylint` linting failing `(no-else-return)` and was trying to find out if I have configured the pre-commit hook incorrectly or something. I stumbled across the `.github/Contributing.md` file and noted it was very out of date (the obvious thing being that the code has now dropped support for python 2.7).
I also think it might be helpful to have `Contributing.md` in the top level directory so that it is more visible to new contributors.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2003/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2003/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2004
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2004/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2004/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2004/events
|
https://github.com/materialsproject/pymatgen/issues/2004
| 753,721,690
|
MDU6SXNzdWU3NTM3MjE2OTA=
| 2,004
|
Incompatibility of mcif and cif file
|
{
"login": "jazmaryphy",
"id": 49759945,
"node_id": "MDQ6VXNlcjQ5NzU5OTQ1",
"avatar_url": "https://avatars.githubusercontent.com/u/49759945?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jazmaryphy",
"html_url": "https://github.com/jazmaryphy",
"followers_url": "https://api.github.com/users/jazmaryphy/followers",
"following_url": "https://api.github.com/users/jazmaryphy/following{/other_user}",
"gists_url": "https://api.github.com/users/jazmaryphy/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jazmaryphy/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jazmaryphy/subscriptions",
"organizations_url": "https://api.github.com/users/jazmaryphy/orgs",
"repos_url": "https://api.github.com/users/jazmaryphy/repos",
"events_url": "https://api.github.com/users/jazmaryphy/events{/privacy}",
"received_events_url": "https://api.github.com/users/jazmaryphy/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-11-30T18:57:07
| 2023-08-13T16:33:49
|
2023-08-13T16:33:49Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hi,
Is there a way to remove the magnetic symmetry operation inside a magnetic mcif file to correspond to symmetry operations in a cif?
For example: check the code using both structure obtain from MAGNDATA (no: 1.16 [http://webbdcrista1.ehu.es/magndata/index.php?this_label=1.16](url) ) and COD (no: 4123009 [http://www.crystallography.net/cod/4123009.html](url) ).
Both file has the same symmetry number.
The function return a more symmetry operation for a mcif than the cif which is understandable because mcif file uses both magnetic and crystallographic symmetry operations.
In my one case, I always want to use data structure from MAGNDATA for my work and the symmetry operations are important in what I am doing.
Is there a way to make both the file (mcif and cif) compatible to each other with the number of symmetry operations generate using the supercell?
def sym(s):
"""
"""
from pymatgen import Structure
from pymatgen.symmetry.analyzer import SpacegroupAnalyzer, SpacegroupOperations
s = Structure.from_file(s)
s.make_supercell([1,2,2])
SA = SpacegroupAnalyzer(s)
no = SA.get_space_group_number()
symbol = SA.get_space_group_symbol()
operations = SA.get_space_group_operations()
return SpacegroupOperations(no, symbol, operations)
[mcif.txt](https://github.com/materialsproject/pymatgen/files/5618126/mcif.txt)
[cif.txt](https://github.com/materialsproject/pymatgen/files/5618128/cif.txt)
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2004/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2004/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2005
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2005/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2005/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2005/events
|
https://github.com/materialsproject/pymatgen/pull/2005
| 754,903,433
|
MDExOlB1bGxSZXF1ZXN0NTMwNzE2OTA0
| 2,005
|
Magmom check
|
{
"login": "jmmshn",
"id": 14003693,
"node_id": "MDQ6VXNlcjE0MDAzNjkz",
"avatar_url": "https://avatars.githubusercontent.com/u/14003693?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jmmshn",
"html_url": "https://github.com/jmmshn",
"followers_url": "https://api.github.com/users/jmmshn/followers",
"following_url": "https://api.github.com/users/jmmshn/following{/other_user}",
"gists_url": "https://api.github.com/users/jmmshn/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jmmshn/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jmmshn/subscriptions",
"organizations_url": "https://api.github.com/users/jmmshn/orgs",
"repos_url": "https://api.github.com/users/jmmshn/repos",
"events_url": "https://api.github.com/users/jmmshn/events{/privacy}",
"received_events_url": "https://api.github.com/users/jmmshn/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @jmmshn. Renamed `DbPhaseDiagram` to be more clear and to match `BaseElement`, also expanded the docstring."
] | 2020-12-02T03:25:38
| 2020-12-12T02:28:58
|
2020-12-12T02:28:16Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Added automatic magmom validation during set creation.
Currently, if there is missing or incompatible magmom data you just get an error when the INCAR is written.
This is not ideal since this often happens on the remote cluster.
So adding an additional validation function should make the behavior smoother.
## Added DBPhaseDiagram
New class for that reading and writing PhaseDiagram to DB
## Added ElementBase
Similar to the case above Element should be a subclass of ElementBase so that other subclasses can be created with a subset of the periodic table (Userful for automatic parsing by the API)
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2005/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2005/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2005",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2005",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2005.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2005.patch",
"merged_at": "2020-12-12T02:28:16Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2006
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2006/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2006/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2006/events
|
https://github.com/materialsproject/pymatgen/pull/2006
| 754,940,764
|
MDExOlB1bGxSZXF1ZXN0NTMwNzQ2NzM1
| 2,006
|
[WIP] enable MaterialsProject2020Compatibility by default
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/37832596)\n\nCoverage decreased (-0.7%) to 82.997% when pulling **65805e23e055377544126e3d936e81e4e2ae0552 on rkingsbury:default_mp2020** into **89e37f004bb1bfc9335552ee4fd3d08c57e67f22 on materialsproject:master**.\n",
"@shyuep , can you help me understand what is happening with these 'Black would make changes' linting errors? They are affecting multiple PRs and I can't figure out how to resolve them. I run `black` on the files in question from the pymatgen root directory, but still see the error during the flake8 test. Thanks!",
"To be honest, I don’t know. As far as I am concerned, black runs fine on my machine. If that is the only problem in your code, I will merge it.",
"@mkhorton the correction values in this PR are now up to date with the new DB release, based on the set of experimental data Amanda compiled and that we've been using all along.\r\n\r\nThe text below summarizes how various energies and magnetic orderings changed compared to the values we fit to at the end of August 2020. Can you please let me know if any of these changes (mainly the magnetic orderings) concern you?\r\n\r\nI'll be evaluating alternative refitting strategies using the new matbench dataset next.\r\n\r\n```\r\nEnergy for entry Na2O (mp-2352) changed from -3.74035 to -3.74051 eV/atom\r\nEnergy for entry MgO (mp-1265) changed from -5.98371 to -5.98399 eV/atom\r\nEnergy for entry CaO (mp-2605) changed from -6.43897 to -6.43930 eV/atom\r\nEnergy for entry NaF (mp-682) changed from -4.32427 to -4.32451 eV/atom\r\nEnergy for entry NaBr (mp-22916) changed from -3.05725 to -3.05739 eV/atom\r\nEnergy for entry MgI2 (mp-23205) changed from -2.57969 to -2.61059 eV/atom\r\nEnergy for entry Al2S3 (mp-684638) changed from -5.03119 to -5.03139 eV/atom\r\nEnergy for entry Si (mp-149) changed from -5.42339 to -5.42531 eV/atom\r\nEnergy for entry V (mp-146) changed from -9.08236 to -9.08391 eV/atom\r\nEnergy for entry Mn (mp-35) changed from -9.16171 to -9.16202 eV/atom\r\nEnergy for entry Fe (mp-13) changed from -8.46930 to -8.47001 eV/atom\r\nEnergy for entry Fe3O4 (mp-19306) changed from -6.74319 to -6.65127 eV/atom\r\nMagnetic type for entry Fe3O4 (mp-19306) changed from FiM to FM\r\nEnergy for entry Co3O4 (mp-18748) changed from -6.00289 to -6.06987 eV/atom\r\nMagnetic type for entry Co3O4 (mp-18748) changed from AFM to FM\r\nEnergy for entry Ni (mp-23) changed from -5.77982 to -5.78014 eV/atom\r\nEnergy for entry MoO2 (mp-510536) changed from -7.40561 to -7.41002 eV/atom\r\nMagnetic type for entry MoO2 (mp-510536) changed from AFM to FM\r\nEnergy for entry WO2 (mp-19372) changed from -7.62360 to -7.71713 eV/atom\r\nMagnetic type for entry MnMoO4 (mp-19081) changed from Unknown to FM\r\nEnergy for entry CaV2O6 (mp-27624) changed from -7.40282 to -7.40419 eV/atom\r\nEnergy for entry Zn (mp-79) changed from -1.25946 to -1.25974 eV/atom\r\nEnergy for entry Ti (mp-72) changed from -7.89505 to -7.89549 eV/atom\r\nEnergy for entry CuSO4 (mp-20525) changed from -5.70722 to -5.72252 eV/atom\r\nMagnetic type for entry CuSO4 (mp-20525) changed from NM to FM\r\nEntry Ba3Al2O6 (mp-1196869) has disappeared from the database\r\nEnergy for entry GaSb (mp-1156) changed from -3.73512 to -3.73532 eV/atom\r\nEnergy for entry Ga2Se3 (mp-1340) changed from -3.89320 to -3.89373 eV/atom\r\nEnergy for entry Ga2Te3 (mp-38970) changed from -3.41813 to -3.41923 eV/atom\r\n```\r\n",
"Related to the above, the selection of fitting compounds we have been using in MP2020 differs in a few cases from the \"likely mpids\" nominated in the `matbench_expt_formation_enthalpy` dataset. In every case, the matbench value is on the hull and experimental, whereas the MP2020 value is off the hull and/or theoretical.\r\n\r\nThe particular one that I wanted to ask about is Fe3O4. The MP2020 nominated material `mp-19306` is FM ordered and and 89 meV off the hull, whereas the matbench value `mp-31770` is on the hull and AFM ordered. Should we specifically be choosing the FM structure here, even though it's so far off the hull?\r\n\r\nAs for the other materials, I think it makes sense to update the mpids to the matbench values.\r\n\r\n```\r\nmpid for K3AlF6 differs: MP2020: mp-1095079 matbench: mp-1201513\r\nmpid for V2O3 differs: MP2020: mp-18937 matbench: mp-21579\r\nmpid for Fe3O4 differs: MP2020: mp-19306 matbench: mp-31770\r\nmpid for LiFeO2 differs: MP2020: mp-19419 matbench: mp-18782\r\nmpid for FeMoO4 differs: MP2020: mp-556667 matbench: mp-505526\r\nmpid for KFeO2 differs: MP2020: mp-759749 matbench: mp-559464\r\nmpid for MoF5 differs: MP2020: mvc-3781 matbench: mp-504759\r\n```",
"> The particular one that I wanted to ask about is Fe3O4. The MP2020 nominated material mp-19306 is FM ordered and and 89 meV off the hull, whereas the matbench value mp-31770 is on the hull and AFM ordered. Should we specifically be choosing the FM structure here, even though it's so far off the hull?\r\n\r\nFe3O4 is a classic ferrimagnet, so should be neither FM nor AFM. I'll have to take a look at this specific case to see if the correct ground state was deprecated somehow. I'm pretty sure the correct mpid is mp-19306 however. (Edited to clarify wording.)",
"> \r\n> \r\n> > The particular one that I wanted to ask about is Fe3O4. The MP2020 nominated material mp-19306 is FM ordered and and 89 meV off the hull, whereas the matbench value mp-31770 is on the hull and AFM ordered. Should we specifically be choosing the FM structure here, even though it's so far off the hull?\r\n> \r\n> Fe3O4 is a classic ferrimagnet, so should be neither FM nor AFM. I'll have to take a look at this specific case to see if the correct ground state was deprecated somehow. I'm pretty sure the correct mpid is mp-19306 however. (Edited to clarify wording.)\r\n\r\nOK. mp-19306 was previously FiM, so it sounds like we should keep that mp-id in anticipation of future DB updates that might put it back on the hull.",
"> > > The particular one that I wanted to ask about is Fe3O4. The MP2020 nominated material mp-19306 is FM ordered and and 89 meV off the hull, whereas the matbench value mp-31770 is on the hull and AFM ordered. Should we specifically be choosing the FM structure here, even though it's so far off the hull?\r\n> > \r\n> > \r\n> > Fe3O4 is a classic ferrimagnet, so should be neither FM nor AFM. I'll have to take a look at this specific case to see if the correct ground state was deprecated somehow. I'm pretty sure the correct mpid is mp-19306 however. (Edited to clarify wording.)\r\n> \r\n> OK. mp-19306 was previously FiM, so it sounds like we should keep that mp-id in anticipation of future DB updates that might put it back on the hull.\r\n\r\nOK, after further investigation I see that ALL but one of the cases in which the new dataset mpid differs from the MP2020 mpid are magnetic. It appears that due to deprecation of some of the magnetic tasks, different mpids are now stable for these six materials. In light of that, I'm going to KEEP the previously-assigned mpids for all these 6 since they are presumably the real ground states (and may be stabilized in a future database release). I will change the mpid for K3AlF6 because in that case, we simply added a new polymorph to the database that is more stable than the previous one.\r\n\r\n```\r\nmpid for K3AlF6 differs: MP2020: mp-1095079 matbench: mp-1201513 NM\r\nmpid for V2O3 differs: MP2020: mp-18937 matbench: mp-21579 AFM\r\nmpid for Fe3O4 differs: MP2020: mp-19306 matbench: mp-31770 FM\r\nmpid for LiFeO2 differs: MP2020: mp-19419 matbench: mp-18782 FM\r\nmpid for FeMoO4 differs: MP2020: mp-556667 matbench: mp-505526 FiM\r\nmpid for KFeO2 differs: MP2020: mp-759749 matbench: mp-559464 AFM\r\nmpid for MoF5 differs: MP2020: mvc-3781 matbench: mp-504759 FM\r\n```",
"@rkingsbury @mkhorton I already cautioned about this a long time back. Things like Fe3O4 (and other mixed valence oxides especially) are extremely sensitive to **charge and magnetic** ordering. Make sure they are really correct before we do this kind of large changes.",
"I've refit the corrections to the latest DB release and updated the experimental data to use higher quality data from the recently-compiled dataset described [here](https://github.com/hackingmaterials/matminer/pull/602). This entailed:\r\n\r\n- Replacing older Kubashewski data with other data for many of the fitting compounds. In particular, this newer data caused non-trivial changes to the experimental enthalpies of halide compounds\r\n- Removing 10 compounds from the fitting set that do not match any ICSD entries\r\n- Updating the mpid of one fitting compound because our latest DB has a new, lower energy polymorph\r\n\r\nIn the end, both the experimental data AND the mpids of fitting compounds are fully consistent with the new dataset, with the exception of the six magnetic compounds described above, for which I retained the older mpids. 115 out of the 263 fitting compounds have uncertainties associated with the formation energies.\r\n\r\nAfter updating the fitting data, our oxygen and halide corrections are more in line with previous work and with the legacy correction scheme.",
"> \r\n> \r\n> Related to the above, the selection of fitting compounds we have been using in MP2020 differs in a few cases from the \"likely mpids\" nominated in the `matbench_expt_formation_enthalpy` dataset. In every case, the matbench value is on the hull and experimental, whereas the MP2020 value is off the hull and/or theoretical.\r\n> \r\n> The particular one that I wanted to ask about is Fe3O4. The MP2020 nominated material `mp-19306` is FM ordered and and 89 meV off the hull, whereas the matbench value `mp-31770` is on the hull and AFM ordered. Should we specifically be choosing the FM structure here, even though it's so far off the hull?\r\n> \r\n> As for the other materials, I think it makes sense to update the mpids to the matbench values.\r\n> \r\n> ```\r\n> mpid for K3AlF6 differs: MP2020: mp-1095079 matbench: mp-1201513\r\n> mpid for V2O3 differs: MP2020: mp-18937 matbench: mp-21579\r\n> mpid for Fe3O4 differs: MP2020: mp-19306 matbench: mp-31770\r\n> mpid for LiFeO2 differs: MP2020: mp-19419 matbench: mp-18782\r\n> mpid for FeMoO4 differs: MP2020: mp-556667 matbench: mp-505526\r\n> mpid for KFeO2 differs: MP2020: mp-759749 matbench: mp-559464\r\n> mpid for MoF5 differs: MP2020: mvc-3781 matbench: mp-504759\r\n> ```\r\n\r\n@mkhorton these are the 7 compounds that should be re-run per today's discussion",
"Hi all - can I get clarification is happening with the magnetic compounds (e.g. Fe3O4, etc) for the new fitting scheme?\r\n\r\n- Will the new fitting scheme be deployed in production prior to these magnetic compounds being completed / re-run?\r\n- Are we now confident on the structure / magnetic state used for these compounds?",
"Also, in case it's helpful, I think we previously tried to follow the example of Wang et al (https://doi.org/10.1103/PhysRevB.73.195107) in picking crystal structures:\r\n\r\n\r\n\r\n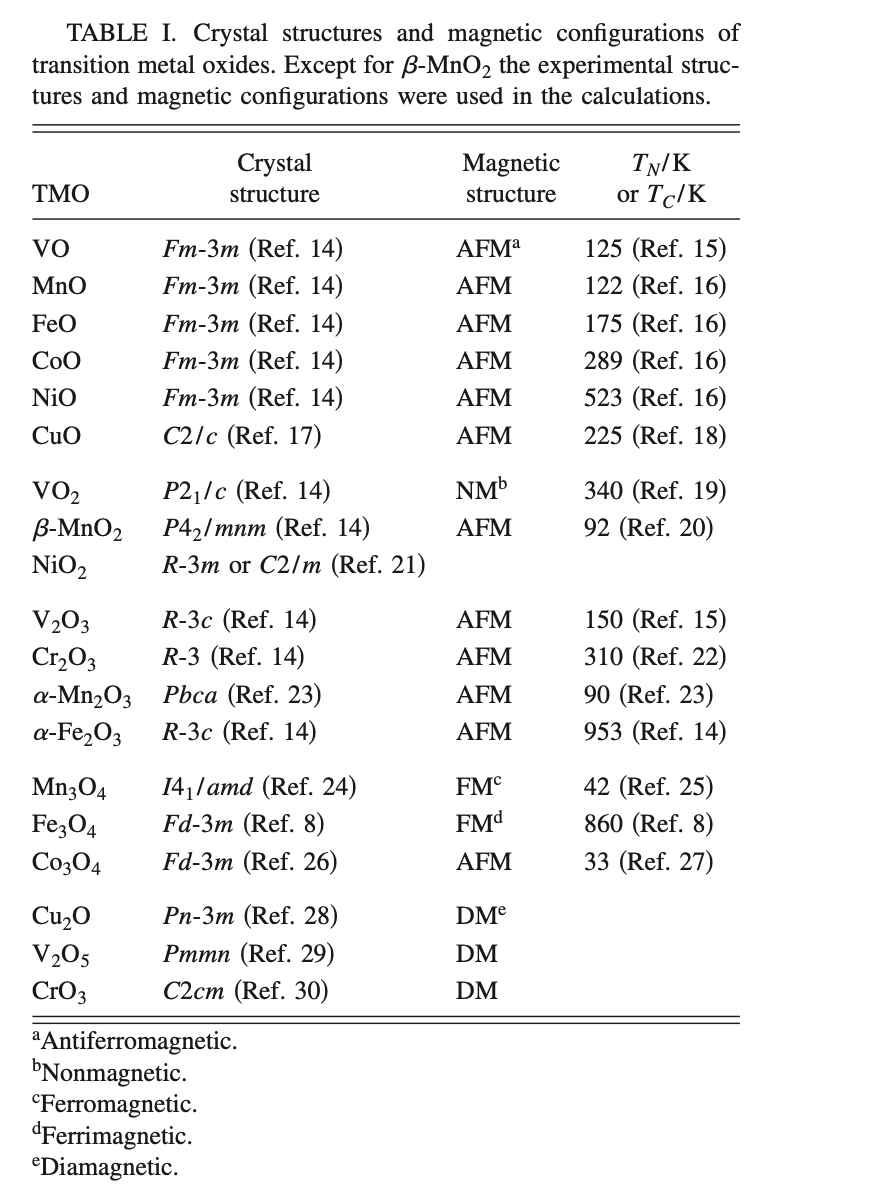\r\n",
"> Hi all - can I get clarification is happening with the magnetic compounds (e.g. Fe3O4, etc) for the new fitting scheme?\r\n> \r\n> * Will the new fitting scheme be deployed in production prior to these magnetic compounds being completed / re-run?\r\n> \r\n> * Are we now confident on the structure / magnetic state used for these compounds?\r\n\r\nHi @computron , I believe we are confident in the magnetic state for the compounds in our fitting set _except_ for the 7 compounds listed above (particularly Fe3O4 which is currently FM but should be FiM). At present our plan is for @mkhorton to rerun this handful of compounds before we use MP2020 in production.\r\n\r\nI can also check what we have against the table you posted @computron. If either of you has an expanded list of correct ground state magnetic orderings please send it my way. \r\n\r\nDidn't we integrate this information into the build pipeline validators? If not, I think we should. \r\n\r\n@mkhorton - I noticed that you went ahead and merged this PR activating MP2020 by default in pymatgen. What is your plan w.r.t. releases? Will MP2020 be the default in the `v2022.xxx` releases but not backported to `v2021.xxx`?",
"As long as the data is live, we can backport to 2021.",
"It is trivial to do backporting. ",
"> \r\n> \r\n> Also, in case it's helpful, I think we previously tried to follow the example of Wang et al (https://doi.org/10.1103/PhysRevB.73.195107) in picking crystal structures:\r\n> \r\n> \r\n\r\n@mkhorton here is a complete list of magnetic materials currently used when fitting the MP2020 corrections, with their magnetic orderings. I've also included any materials whose formulas are listed in the Wang et al. table Anubhav just posted. Any cases where the orderings don't match are marked with MISMATCH\r\n\r\n```\r\nCr2FeO4 (mp-1104680) : FiM \r\nMn2O3 (mp-1172875) : FM MISMATCH (Wang et al. lists AFM)\r\nCa3Al2O6 (mp-12147 ) : Unknown\r\nCuF2 (mp-1229 ) : FM \r\nCa(FeO2)2 (mp-1271532) : FiM \r\nFe (mp-13 ) : FM \r\nCsO2 (mp-1441 ) : FM \r\nCuO (mp-1692 ) : NM MISMATCH (Wang et al. lists AFM)\r\nKO2 (mp-1866 ) : FM \r\nTiNiO3 (mp-18732 ) : AFM \r\nCo3O4 (mp-18748 ) : FM MISMATCH (Wang et al. lists AFM)\r\nNiSO4 (mp-18749 ) : AFM \r\nMn(FeO2)2 (mp-18750 ) : FiM \r\nMn3O4 (mp-18759 ) : FiM MISMATCH (Wang et al. lists FM)\r\nMnCO3 (mp-18814 ) : AFM \r\nMn2SiO4 (mp-18928 ) : AFM \r\nV2O3 (mp-18937 ) : AFM \r\nFeCO3 (mp-18969 ) : AFM \r\nMnO (mp-19006 ) : AFM \r\nNiO (mp-19009 ) : AFM \r\nNaO2 (mp-1901 ) : FM \r\nSi(NiO2)2 (mp-19072 ) : FiM \r\nMnMoO4 (mp-19081 ) : FM \r\nTiMnO3 (mp-19082 ) : AFM \r\nCoWO4 (mp-19092 ) : AFM \r\nVO2 (mp-19094 ) : FM MISMATCH (Wang et al. lists NM)\r\nNiCO3 (mp-19147 ) : AFM \r\nVO (mp-19184 ) : AFM \r\nFeSO4 (mp-19254 ) : FM \r\nFe3O4 (mp-19306 ) : FM MISMATCH (Wang et al. lists FiM)\r\nZn(FeO2)2 (mp-19313 ) : AFM \r\nCr2NiO4 (mp-19344 ) : FiM \r\nNaFeO2 (mp-19359 ) : AFM \r\nCr2(SO4)3 (mp-19361 ) : AFM \r\nWO2 (mp-19372 ) : FM \r\nCoSO4 (mp-19379 ) : AFM \r\nMnO2 (mp-19395 ) : FM MISMATCH (Wang et al. lists AFM)\r\nCr2O3 (mp-19399 ) : AFM \r\nMnWO4 (mp-19407 ) : AFM \r\nZnCr2O4 (mp-19410 ) : AFM \r\nTiFeO3 (mp-19417 ) : AFM \r\nLiFeO2 (mp-19419 ) : FM \r\nFeWO4 (mp-19421 ) : FM \r\nTiCoO3 (mp-19424 ) : AFM \r\nFe2O3 (mp-19770 ) : AFM \r\nFe2SiO4 (mp-20313 ) : FiM \r\nNiSeO3 (mp-20460 ) : AFM \r\nCuSO4 (mp-20525 ) : FM \r\nCr2CoO4 (mp-20758 ) : FiM \r\nNiWO4 (mp-21179 ) : AFM \r\nCoCO3 (mp-21434 ) : AFM \r\nCo2SiO4 (mp-21856 ) : FiM \r\nCa2Fe2O5 (mp-22113 ) : FiM \r\nCoO (mp-22408 ) : AFM \r\nMnSO4 (mp-22554 ) : AFM \r\nNi (mp-23 ) : FM \r\nV2O5 (mp-25279 ) : NM MISMATCH (Wang et al. lists DM)\r\nAl2CuO4 (mp-27719 ) : FM \r\nAl2FeO4 (mp-30084 ) : AFM \r\nMn (mp-35 ) : FiM \r\nCu2O (mp-361 ) : NM MISMATCH (Wang et al. lists DM)\r\nAl2CoO4 (mp-36447 ) : AFM \r\nCr2CuO4 (mp-504573 ) : FiM \r\nFeCuO2 (mp-510281 ) : AFM \r\nCrO3 (mp-510421 ) : NM MISMATCH (Wang et al. lists DM)\r\nMoO2 (mp-510536 ) : FM \r\nCo (mp-54 ) : FM \r\nCaCr2O4 (mp-540582 ) : FiM \r\nTiMn2O4 (mp-540763 ) : AFM \r\nFe2(SO4)3 (mp-540767 ) : AFM \r\nFeClO (mp-540828 ) : AFM \r\nCrF2 (mp-554340 ) : AFM \r\nVF4 (mp-554799 ) : FM \r\nCoF2 (mp-555908 ) : AFM \r\nFeMoO4 (mp-556667 ) : FiM \r\nFeF2 (mp-556911 ) : AFM \r\nCoF3 (mp-559473 ) : FM \r\nNiF2 (mp-559798 ) : FM \r\nCrF3 (mp-560338 ) : AFM \r\nMnF2 (mp-560902 ) : AFM \r\nCoSeO3 (mp-562826 ) : FiM \r\nNaCrO2 (mp-578604 ) : AFM \r\nAl2NiO4 (mp-688785 ) : AFM \r\nFe2CoO4 (mp-753222 ) : FM \r\nMnAl2O4 (mp-755882 ) : AFM \r\nKFeO2 (mp-759749 ) : AFM \r\nTi(FeO2)2 (mp-780585 ) : FiM \r\nCr (mp-90 ) : AFM \r\nMoF5 (mvc-3781 ) : FM \r\n``` "
] | 2020-12-02T04:58:39
| 2021-04-01T21:10:44
|
2021-03-25T20:16:12Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Make `MaterialsProject2020Compatibility` the default energy correction scheme in pymatgen.
* `MPRester().get_entries()` and `MPRester().get_pourbaix_entries()` now use MP2020 to correct energies
* `MaterialsProjectAqueousCompatibility` now uses MP2020 instead of legacy MP as the `solid_compat`
* Updated Pourbaix tests to reflect energy shifts due to above change
* removed deprecation warning for legacy `MaterialsProjectCompatibility`
## TODO (if any)
- [x] Refit energy corrections after upcoming data release
- [ ] Update publication detail after resubmission
- [ ] validate output of emmet `ThermoBuilder` when using MP2020 corrections to build all phase diagrams
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2006/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2006/timeline
| null | true
| true
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2006",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2006",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2006.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2006.patch",
"merged_at": "2021-03-25T20:16:12Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2007
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2007/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2007/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2007/events
|
https://github.com/materialsproject/pymatgen/pull/2007
| 754,950,307
|
MDExOlB1bGxSZXF1ZXN0NTMwNzU0Njcy
| 2,007
|
fix bug in GGA run_type when METAGGA=None
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @rkingsbury "
] | 2020-12-02T05:20:50
| 2020-12-02T19:52:49
|
2020-12-02T19:20:25Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
This PR fixes a recently discovered bug in which the `Vasprun.run_type` of a calculation in which `METAGGA=None` would not be set correctly.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2007/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2007/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2007",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2007",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2007.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2007.patch",
"merged_at": "2020-12-02T19:20:25Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2008
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2008/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2008/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2008/events
|
https://github.com/materialsproject/pymatgen/pull/2008
| 755,677,027
|
MDExOlB1bGxSZXF1ZXN0NTMxMzQ0ODA1
| 2,008
|
Module-wide linting
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
] | null |
[
"Note, this will be merged without `pylint` modified files test passing since, by definition, this PR touches many files. Examination of logs does reveal some areas for improvement, notably `chemenv` (missing docstrings), `boltztrap` (lingering Python 2 code)."
] | 2020-12-02T23:05:26
| 2020-12-03T01:10:47
|
2020-12-03T01:10:40Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
To improve legibility of pull requests, this runs linting tools including formatting with `black` and import sorting with `isort` throughout pymatgen. Will merge after tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2008/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2008/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2008",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2008",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2008.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2008.patch",
"merged_at": "2020-12-03T01:10:40Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2009
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2009/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2009/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2009/events
|
https://github.com/materialsproject/pymatgen/issues/2009
| 756,353,551
|
MDU6SXNzdWU3NTYzNTM1NTE=
| 2,009
|
"Incorrect value of vasp_type given" when reading spin-unpolarized WAVECAR
|
{
"login": "boyoungzheng",
"id": 21117824,
"node_id": "MDQ6VXNlcjIxMTE3ODI0",
"avatar_url": "https://avatars.githubusercontent.com/u/21117824?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/boyoungzheng",
"html_url": "https://github.com/boyoungzheng",
"followers_url": "https://api.github.com/users/boyoungzheng/followers",
"following_url": "https://api.github.com/users/boyoungzheng/following{/other_user}",
"gists_url": "https://api.github.com/users/boyoungzheng/gists{/gist_id}",
"starred_url": "https://api.github.com/users/boyoungzheng/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/boyoungzheng/subscriptions",
"organizations_url": "https://api.github.com/users/boyoungzheng/orgs",
"repos_url": "https://api.github.com/users/boyoungzheng/repos",
"events_url": "https://api.github.com/users/boyoungzheng/events{/privacy}",
"received_events_url": "https://api.github.com/users/boyoungzheng/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I tried your input files with vasp 5.4.4 and pymatgen 2020.7.3. I did the scf first followed by the band structure calculation. I read in the wavecar with the following code:\r\n```python\r\nfrom pymatgen.io.vasp.outputs import Wavecar\r\nw = Wavecar('WAVECAR')\r\n```\r\nThis worked with no problem. I then updated to pymatgen 2020.11.11 and ran it again with no problems. How are you calling Wavecar? Could you try the above code?",
"Thanks for your reply. Weird enough, I tried a clean run and it works (with WAVECAR size 2.2G). I didn't find anything special in OUTCARs. Anyway, here I share the WAVECAR (2.3G) I have problem with.\r\n\r\nhttps://pennstateoffice365-my.sharepoint.com/:f:/g/personal/buz118_psu_edu/Ek8qBSX2661Cn-2eNkZSVPsBbxS2Xn3FdLdm-Z9KO-_8Jg?e=xmZNfo\r\n",
"Well, I'm glad it works now. However, I tried the WAVECAR that you sent with the little code I listed above, and it read with no problem. Python 3.8.6 and pymatgen 2020.12.3. Are you sure you linked the right WAVECAR? :smile: \r\n\r\nWhich python version are you using? I wonder if that has anything to do with it (I would think not, but who knows).",
"MAGICALLY, the same WAVECAR can go through without any problem (on the cluster). I absolutely have no idea what happened. When I copied the WAVECAR to my local mac, I also tested it with no problem. Probably, the original error was due to something in the cluster which may have been secretly fixed by administrators. \r\n\r\nThank you very much for your help. This code helped me a loooooot. 👍",
"No problem, I'm glad it worked out. It could be a memory issue; the entire WAVECAR is read into memory. So if you were using a login node and someone else was using most of the memory, it would throw an error, and then the error would go away once some memory is freed up."
] | 2020-12-03T16:28:13
| 2020-12-08T05:32:30
|
2020-12-08T04:26:25Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Same as the title.
**To Reproduce**
Trying to do a band unfolding process with the attached inputs. Of course, a scf calculation is done first.
[inputs.zip](https://github.com/materialsproject/pymatgen/files/5637393/inputs.zip)
**Expected behavior**
Successful reading of WAVECAR
**Screenshots**
(None)
**Versions**
- pymatgen: 2020.11.11
- vasp: 5.4.1
**Additional context**
(None)
|
{
"login": "boyoungzheng",
"id": 21117824,
"node_id": "MDQ6VXNlcjIxMTE3ODI0",
"avatar_url": "https://avatars.githubusercontent.com/u/21117824?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/boyoungzheng",
"html_url": "https://github.com/boyoungzheng",
"followers_url": "https://api.github.com/users/boyoungzheng/followers",
"following_url": "https://api.github.com/users/boyoungzheng/following{/other_user}",
"gists_url": "https://api.github.com/users/boyoungzheng/gists{/gist_id}",
"starred_url": "https://api.github.com/users/boyoungzheng/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/boyoungzheng/subscriptions",
"organizations_url": "https://api.github.com/users/boyoungzheng/orgs",
"repos_url": "https://api.github.com/users/boyoungzheng/repos",
"events_url": "https://api.github.com/users/boyoungzheng/events{/privacy}",
"received_events_url": "https://api.github.com/users/boyoungzheng/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2009/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2009/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2010
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2010/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2010/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2010/events
|
https://github.com/materialsproject/pymatgen/issues/2010
| 758,449,490
|
MDU6SXNzdWU3NTg0NDk0OTA=
| 2,010
|
Cannot install pymatgen==2019.7.2 with python3.6
|
{
"login": "kpace",
"id": 5939689,
"node_id": "MDQ6VXNlcjU5Mzk2ODk=",
"avatar_url": "https://avatars.githubusercontent.com/u/5939689?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/kpace",
"html_url": "https://github.com/kpace",
"followers_url": "https://api.github.com/users/kpace/followers",
"following_url": "https://api.github.com/users/kpace/following{/other_user}",
"gists_url": "https://api.github.com/users/kpace/gists{/gist_id}",
"starred_url": "https://api.github.com/users/kpace/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/kpace/subscriptions",
"organizations_url": "https://api.github.com/users/kpace/orgs",
"repos_url": "https://api.github.com/users/kpace/repos",
"events_url": "https://api.github.com/users/kpace/events{/privacy}",
"received_events_url": "https://api.github.com/users/kpace/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I think this was a problem with past versions. Try the newest version of pymatgen instead. (Or if that fails, you should use a py37 environment. Py36 is pretty old and a lot of packages already dropped support for it)",
"Yes, to expand a little here, `numpy` has now officially dropped support for Python 3.6 in their latest version. While `pymatgen` should still work with 3.6 (all our tests still pass on 3.6) by using an older version of numpy such as 1.19.2, I concur with @shyuep that it would probably be easier to use Python 3.7+ if that's an option.",
"Hi! I would appreciate if you could re-open the issue as the underlying bug breaks installation flows for all packages that depend on pymatgen, but still support Python 3.6. Concretely, this issue is currently causing a break of our standard installation path for AiiDA, see issue https://github.com/aiidateam/aiida-core/issues/4614 and related pull request https://github.com/aiidateam/aiida-core/pull/4615 . Important to mention that this issue has occurred in the past before, and was previously reported in https://github.com/materialsproject/pymatgen/issues/1520 .\r\n\r\nI understand that the pymatgen maintainer perspective is to simply recommend the use of a newer Python version, but this is unfortunately not possible for some users and packages that have long-term support schedules. But even if that was an acceptable solution, this bug would still cause an issue, because it causes *all constraints that dependent packages have placed on their numpy dependency to be ignored.*\r\n\r\nThis problem would be much less severe if at least older versions of pymatgen would still behave properly, but the root cause makes it basically impossible to have any pymatgen dependency, because -- as was demonstrated by the OP -- even older versions of pymatgen will break. To put it plainly: unless the build setup is fixed, pymatgen will *always* cause issues when numpy drops support for older Python versions even for older versions of pymatgen, making it basically impossible to declare pymatgen as a dependency without giving up all control over numpy constraints.",
"@csadorf I am unclear what is needed from pymatgen side. Certainly older versions of pymatgen will work for python 3.6. We do not specify a numpy version in the setup.py beyond \">=1.14.x\". That means even relatively ancient numpy versions should work. The requirements.txt pinning for older versions of pymatgen are based on the numpy versions at those times and is in line with python best practices.\r\n\r\nI did a test and even as recent as pymatgen 2020.8.3, install on py36 works.\r\n\r\n```\r\nconda create -n py36 python=3.6\r\nconda activate py36\r\nconda install numpy\r\npip install pymatgen==2020.8.3\r\n```\r\n\r\ngives me\r\n\r\n```\r\nCollecting pymatgen==2020.8.3\r\n Downloading pymatgen-2020.8.3-cp36-cp36m-macosx_10_14_x86_64.whl (3.3 MB)\r\n |████████████████████████████████| 3.3 MB 3.4 MB/s\r\nRequirement already satisfied: numpy>=1.14.3 in ./opt/miniconda3/envs/py36/lib/python3.6/site-packages (from pymatgen==2020.8.3) (1.19.2)\r\nCollecting dataclasses>=0.6\r\n Downloading dataclasses-0.8-py3-none-any.whl (19 kB)\r\nCollecting matplotlib>=1.5\r\n Downloading matplotlib-3.3.3-cp36-cp36m-macosx_10_9_x86_64.whl (8.5 MB)\r\n |████████████████████████████████| 8.5 MB 29.1 MB/s\r\n....\r\n```\r\n\r\nSo it does seem the numpy setup.py is respected. I also tried it with your pymatgen==2019.7.2 and it works as well.\r\n\r\nIn short, I have no idea why your install is failing. I suspect it si an issue with your test-env caching. I suggest you use a clean environment and re-test.\r\n\r\nNow, that said, you will have to pin to an older version of pymatgen. It seems anything before Sep 2020 should work fine. But newer features will not be available. Unfortunately, we cannot keep retaining compatibility with old versions. Once major packages such as numpy move on, newer versions of pymatgen will have to move on.",
"Thanks for taking the time to investigate this @shyuep . The problem seems to be when installing `pymatgen` from the tarball distribution and so the wheel has to be built. In that case, something (I am not sure which package is responsible for this) is installing `numpy==1.20.0rc1`. Since this doesn't support Python 3.6, the build will crash. For an example, you can check [this log of a run on Github Actions](https://github.com/sphuber/aiida-core/runs/1566282532). Some of the relevant lines are:\r\n```\r\nCollecting pymatgen==2020.3.2\r\n Downloading pymatgen-2020.3.2.tar.gz (2.6 MB)\r\n ERROR: Command errored out with exit status 1:\r\n command: /opt/hostedtoolcache/Python/3.6.12/x64/bin/python -c 'import sys, setuptools, tokenize; sys.argv[0] = '\"'\"'/tmp/pip-install-s526ezws/pymatgen_06d20c37b3a749d187e77525e2902b22/setup.py'\"'\"'; __file__='\"'\"'/tmp/pip-install-s526ezws/pymatgen_06d20c37b3a749d187e77525e2902b22/setup.py'\"'\"';f=getattr(tokenize, '\"'\"'open'\"'\"', open)(__file__);code=f.read().replace('\"'\"'\\r\\n'\"'\"', '\"'\"'\\n'\"'\"');f.close();exec(compile(code, __file__, '\"'\"'exec'\"'\"'))' egg_info --egg-base /tmp/pip-pip-egg-info-1m_tqngz\r\n cwd: /tmp/pip-install-s526ezws/pymatgen_06d20c37b3a749d187e77525e2902b22/\r\n Complete output (68 lines):\r\n Traceback (most recent call last):\r\n File \"/opt/hostedtoolcache/Python/3.6.12/x64/lib/python3.6/site-packages/setuptools/sandbox.py\", line 154, in save_modules\r\n yield saved\r\n File \"/opt/hostedtoolcache/Python/3.6.12/x64/lib/python3.6/site-packages/setuptools/sandbox.py\", line 195, in setup_context\r\n yield\r\n File \"/opt/hostedtoolcache/Python/3.6.12/x64/lib/python3.6/site-packages/setuptools/sandbox.py\", line 250, in run_setup\r\n _execfile(setup_script, ns)\r\n File \"/opt/hostedtoolcache/Python/3.6.12/x64/lib/python3.6/site-packages/setuptools/sandbox.py\", line 45, in _execfile\r\n exec(code, globals, locals)\r\n File \"/tmp/easy_install-q6kydy_z/numpy-1.20.0rc1/setup.py\", line 30, in <module>\r\n if sys.platform.startswith('win') and platform.machine().endswith('64'):\r\n RuntimeError: Python version >= 3.7 required.\r\n```\r\nNow what is remarkable is the conditional which is included in the last line `if sys.platform.startswith('win') and platform.machine().endswith('64'):`. The tests are actually running on an Ubuntu 18.04 host. Now I am not sure if this is just a virtual machine that is running on a host runner that is actually using windows. Given that Github is now owned by MIcrosoft is not wholly unlikely.\r\n\r\nI have been trying to reproduce this on my own machine which also runs Ubuntu 18.04, but without success. There, even in a clean Python 3.6 virtual environment, the `pymatgen` tarball is downloaded and built some problems. I am now wondering if the version of `wheel` or `setuptools` has anything to do with it. I am investigating this further and will update here.\r\n",
"@sphuber I just retried on a Ubuntu server the exact same series of commands (the previous test was done on a Mac). There is still no error and if a numpy > 1.14 was installed, pymatgen installs without error. This was done on both a supercomputer as well as a virtual machine. So in short, I cannot reproduce the error you mention. I suspect it is something percuilar to the CI environment.",
"@shyuep did you `pip install --upgrade pip`, I was concerned maybe the new [pip dependency resolver](http://pyfound.blogspot.com/2020/11/pip-20-3-new-resolver.html) might be responsible (just a hunch)",
"Note that if we pre-install a version of `numpy` that is compatible with the Python version of the environment, then the installation of `pymatgen` goes without problem (this is the current workaround that we use). So at least one condition for the problem to appear is for the environment to have no version of `numpy` installed whatsoever when `pymatgen` gets installed.",
"@mkhorton we experienced the same problem over a year ago when Python 2.7 and 3.5 support was dropped by `numpy` and this was well before the new dependency resolver, so I don't think this is necessarily at fault.",
"Interesting. Then I think this is an issue at either the versioning end or pip end. Basically, pip install should choose the latest version of numpy compatible with that python version (3.6 in your case). It is strange that it is attempting to install 1.20 which is clearly incompatible with py36. ",
"> Interesting. Then I think this is an issue at either the versioning end or pip end. Basically, pip install should choose the latest version of numpy compatible with that python version (3.6 in your case). It is strange that it is attempting to install 1.20 which is clearly incompatible with py36.\r\n\r\nI think that is exactly where the crux of the problem lies: if I understand things correctly it _is not `pip`_ that installs `numpy==1.20.0rc1`. At the point when that occurs, I think it is another package manager or installation tool that interprets the requirements of `pymatgen`. My suspicion is that when the `pymatgen` wheel is getting built, it is either `wheel` or `setuptools` that is responsible for installing the dependencies and there the requirements imposed do not get respected.\r\n\r\nIf we look again at the failing build, it is the following command that fails:\r\n```\r\n/opt/hostedtoolcache/Python/3.6.12/x64/bin/python -c 'import sys, setuptools, tokenize; sys.argv[0] = '\"'\"'/tmp/pip-install-s526ezws/pymatgen_06d20c37b3a749d187e77525e2902b22/setup.py'\"'\"'; __file__='\"'\"'/tmp/pip-install-s526ezws/pymatgen_06d20c37b3a749d187e77525e2902b22/setup.py'\"'\"';f=getattr(tokenize, '\"'\"'open'\"'\"', open)(__file__);code=f.read().replace('\"'\"'\\r\\n'\"'\"', '\"'\"'\\n'\"'\"');f.close();exec(compile(code, __file__, '\"'\"'exec'\"'\"'))' egg_info --egg-base /tmp/pip-pip-egg-info-1m_tqngz\r\n```\r\nI have no idea who calls this command, but it seems likely that `pip` at this point is no longer involved. Since again `setuptools` is involved here as it gets imported, it seems to me the problem has to do with that.\r\n",
"More evidence that `setuptools` is the culprit. I noticed that the minimum version you require is very low:\r\nhttps://github.com/materialsproject/pymatgen/blob/bb6ca9eefeda4f400537c7530212d3b18a8d6bb4/setup.py#L111\r\n\r\nI tried to update `setuptools` before installing `pymatgen` and that got rid of the problem. I guess the question now would be to figure out starting from which version of `setuptools` the problem was solved. If we find it, would it be a possible solution for you to increase the minimum required version of `setuptools` in your `setup_requires`? The current minimum [version dates back to 2015](https://pypi.org/project/setuptools/18.0/).",
"@sphuber I tried doing this from scratch in a clean ubuntu env and a clean conda, i.e., no numpy installed. I even let the system auto-find the correct version of pymatgen. I still cannot reproduce this error. This is the output.\r\n\r\n```\r\n(base) shyuep@ubuntu:~$ conda activate py36\r\n(py36) shyuep@ubuntu:~$ pip install pymatgen\r\nCollecting pymatgen\r\n Downloading pymatgen-2020.12.3.tar.gz (2.8 MB)\r\n |████████████████████████████████| 2.8 MB 3.8 MB/s\r\nCollecting dataclasses>=0.6\r\n Downloading dataclasses-0.8-py3-none-any.whl (19 kB)\r\nCollecting matplotlib>=1.5\r\n Downloading matplotlib-3.3.3-cp36-cp36m-manylinux1_x86_64.whl (11.6 MB)\r\n |████████████████████████████████| 11.6 MB 7.9 kB/s\r\nCollecting cycler>=0.10\r\n Downloading cycler-0.10.0-py2.py3-none-any.whl (6.5 kB)\r\nCollecting kiwisolver>=1.0.1\r\n Downloading kiwisolver-1.3.1-cp36-cp36m-manylinux1_x86_64.whl (1.1 MB)\r\n |████████████████████████████████| 1.1 MB 8.4 MB/s\r\nCollecting monty>=3.0.2\r\n Downloading monty-4.0.2-py3-none-any.whl (62 kB)\r\n |████████████████████████████████| 62 kB 37 kB/s\r\nCollecting networkx>=2.2\r\n Downloading networkx-2.5-py3-none-any.whl (1.6 MB)\r\n |████████████████████████████████| 1.6 MB 8.8 MB/s\r\nCollecting decorator>=4.3.0\r\n Downloading decorator-4.4.2-py2.py3-none-any.whl (9.2 kB)\r\nCollecting numpy>=1.14.3\r\n Using cached numpy-1.19.4-cp36-cp36m-manylinux2010_x86_64.whl (14.5 MB)\r\nCollecting palettable>=3.1.1\r\n Downloading palettable-3.3.0-py2.py3-none-any.whl (111 kB)\r\n |████████████████████████████████| 111 kB 11.3 MB/s\r\nCollecting pillow>=6.2.0\r\n Downloading Pillow-8.0.1-cp36-cp36m-manylinux1_x86_64.whl (2.2 MB)\r\n |████████████████████████████████| 2.2 MB 7.6 MB/s\r\nCollecting plotly>=4.5.0\r\n Downloading plotly-4.14.1-py2.py3-none-any.whl (13.2 MB)\r\n |████████████████████████████████| 13.2 MB 56 kB/s\r\nCollecting pyparsing!=2.0.4,!=2.1.2,!=2.1.6,>=2.0.3\r\n Downloading pyparsing-2.4.7-py2.py3-none-any.whl (67 kB)\r\n |████████████████████████████████| 67 kB 682 kB/s\r\nCollecting python-dateutil>=2.1\r\n Downloading python_dateutil-2.8.1-py2.py3-none-any.whl (227 kB)\r\n |████████████████████████████████| 227 kB 8.5 MB/s\r\nCollecting retrying>=1.3.3\r\n Downloading retrying-1.3.3.tar.gz (10 kB)\r\nCollecting ruamel.yaml>=0.15.6\r\n Downloading ruamel.yaml-0.16.12-py2.py3-none-any.whl (111 kB)\r\n |████████████████████████████████| 111 kB 8.8 MB/s\r\nCollecting ruamel.yaml.clib>=0.1.2\r\n Downloading ruamel.yaml.clib-0.2.2-cp36-cp36m-manylinux1_x86_64.whl (549 kB)\r\n |████████████████████████████████| 549 kB 9.7 MB/s\r\nCollecting scipy>=1.5.0\r\n Downloading scipy-1.5.4-cp36-cp36m-manylinux1_x86_64.whl (25.9 MB)\r\n |████████████████████████████████| 25.9 MB 1.3 MB/s\r\n...\r\n```\r\n\r\nThe bottom line is that it is successfully installed and numpy 1.19.4 (which is compatible with py36) was automatically installed. \r\n\r\nOne other thing you might want to look into is whether any existing packages in your env might be causing problems. Usually it is best to have the virtual env be isolated and not use any site-packages.",
"@sphuber Thanks. Ok. I will make setuptools>=50 as the requirement. That should fix some of this for now. In general though, I think these things typically are updated by themselves....",
"> One other thing you might want to look into is whether any existing packages in your env might be causing problems. Usually it is best to have the virtual env be isolated and not use any site-packages.\r\n\r\nWe do use virtual environments. This is what we recommend to all users for installation and is definitely what we use in our CI. Each run gets a completely new and clean virtual environment. As described above, I really think it is due to the outdated version of `setuptools`. I suspect that in all your tests you had a more recent version where the problem was fixed.",
"@sphuber Ok. I have increased the min setuptools version. That will not fix the old pymatgen releases, but hopefully it will prevent future releases from having issues. ",
"@shyuep Thank you very much for the help and sorry that I was not able to respond earlier. But I'm glad that @sphuber was available to clear up some of the questions. It seems that it is mostly figured out that `setuptools` is the culprit here.\r\n\r\nHere is a minimal example that demonstrates that the bug was apparently resolved in setuptools version 42:\r\n```\r\n# fails:\r\n$ docker run --env SETUPTOOLS_VERSION=41 --rm -it python:3.6 /bin/bash -c 'pip install -I setuptools==${SETUPTOOLS_VERSION} && pip install pymatgen==2019.7.2'\r\n# succeeds:\r\n$ docker run --env SETUPTOOLS_VERSION=42 --rm -it python:3.6 /bin/bash -c 'pip install -I setuptools==${SETUPTOOLS_VERSION} && pip install pymatgen==2019.7.2'\r\n```\r\nSame behavior when installed as dependency (example with aiida-core):\r\n```\r\n# fails:\r\n docker run --env SETUPTOOLS_VERSION=41 --rm -it python:3.6 /bin/bash -c 'pip install -I setuptools==${SETUPTOOLS_VERSION} && pip install aiida-core[atomic_tools]'\r\n# succeeds:\r\n docker run --env SETUPTOOLS_VERSION=42 --rm -it python:3.6 /bin/bash -c 'pip install -I setuptools==${SETUPTOOLS_VERSION} && pip install aiida-core[atomic_tools]'\r\n```\r\nI therefore assume that it might be related to this change: https://setuptools.readthedocs.io/en/latest/history.html#id170",
"Sorry to add to a long thread, but while some of the issues may have been fixed, one root problem remains unaddressed:\r\n\r\npymatgen is using the `setup_requires` in the `setup.py` which is [discouraged by setuptools](https://setuptools.readthedocs.io/en/latest/references/keywords.html).\r\nThe problem is that this results in using `easy_install` to fetch the build dependencies. [`easy_install` is outdated and should not be used](https://setuptools.readthedocs.io/en/latest/deprecated/easy_install.html) - it is significantly dumber than pip and can lead to all sorts of issues and incompatibilities [1].\r\n\r\nThe solution is to follow [PEP-518](http://www.python.org/dev/peps/pep-0518/) and use the `pyproject.toml` to specify the build dependencies.\r\nI see that @shyuep just added a `pyproject.toml` an hour ago in a different context. \r\nI will prepare a PR to use it also for the build dependencies.\r\n\r\n\r\n[1] To take a recent example, our builds of the AiiDA documentation on readthedocs (using python 3.8) have been broken since December 4th due to an [issue with `numpy.ndarray size changed` introduced by the pymatgen dependency](https://readthedocs.org/projects/aiida-core/builds/12491657/) - in this particular case I haven't checked that `easy_install` is the perpetrator but I strongly suspect it (we had a similar case before where it was)."
] | 2020-12-07T12:04:55
| 2021-01-30T20:26:07
|
2020-12-07T16:37:28Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Cannot install pymatgen==2019.7.2 on ubuntu 16.04 in a clean virtualenv with python3.6.11.
**To Reproduce**
```
(test-venv) jenkins@ip:~/workspace/PR-212$ python --version
Python 3.6.11
(test-venv) jenkins@ip:~/workspace/PR-212$ pip --version
pip 20.3.1 from /usr/src/test-venv/lib/python3.6/site-packages/pip (python 3.6)
(test-venv) jenkins@ip-:~/workspace/PR-212$ pip3 install pymatgen==2019.7.2
Collecting pymatgen==2019.7.2
Using cached pymatgen-2019.7.2.tar.gz (2.4 MB)
ERROR: Command errored out with exit status 1:
command: /usr/src/test-venv/bin/python3.6 -c 'import sys, setuptools, tokenize; sys.argv[0] = '"'"'/tmp/pip-install-gcjf5s0s/pymatgen_0d09ccb94359449695eafa5c38d65f5c/setup.py'"'"'; __file__='"'"'/tmp/pip-install-gcjf5s0s/pymatgen_0d09ccb94359449695eafa5c38d65f5c/setup.py'"'"';f=getattr(tokenize, '"'"'open'"'"', open)(__file__);code=f.read().replace('"'"'\r\n'"'"', '"'"'\n'"'"');f.close();exec(compile(code, __file__, '"'"'exec'"'"'))' egg_info --egg-base /tmp/pip-pip-egg-info-98jjh4w1
cwd: /tmp/pip-install-gcjf5s0s/pymatgen_0d09ccb94359449695eafa5c38d65f5c/
Complete output (68 lines):
Traceback (most recent call last):
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/sandbox.py", line 154, in save_modules
yield saved
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/sandbox.py", line 195, in setup_context
yield
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/sandbox.py", line 250, in run_setup
_execfile(setup_script, ns)
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/sandbox.py", line 45, in _execfile
exec(code, globals, locals)
File "/tmp/easy_install-exzgb8j1/numpy-1.20.0rc1/setup.py", line 30, in <module>
Official docs: [http://pymatgen.org](http://pymatgen.org/)
RuntimeError: Python version >= 3.7 required.
During handling of the above exception, another exception occurred:
Traceback (most recent call last):
File "<string>", line 1, in <module>
File "/tmp/pip-install-gcjf5s0s/pymatgen_0d09ccb94359449695eafa5c38d65f5c/setup.py", line 164, in <module>
'get_environment = pymatgen.cli.get_environment:main',
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/__init__.py", line 142, in setup
_install_setup_requires(attrs)
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/__init__.py", line 137, in _install_setup_requires
dist.fetch_build_eggs(dist.setup_requires)
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/dist.py", line 586, in fetch_build_eggs
replace_conflicting=True,
File "/usr/src/test-venv/lib/python3.6/site-packages/pkg_resources/__init__.py", line 780, in resolve
replace_conflicting=replace_conflicting
File "/usr/src/test-venv/lib/python3.6/site-packages/pkg_resources/__init__.py", line 1063, in best_match
return self.obtain(req, installer)
File "/usr/src/test-venv/lib/python3.6/site-packages/pkg_resources/__init__.py", line 1075, in obtain
return installer(requirement)
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/dist.py", line 653, in fetch_build_egg
return cmd.easy_install(req)
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/command/easy_install.py", line 679, in easy_install
return self.install_item(spec, dist.location, tmpdir, deps)
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/command/easy_install.py", line 705, in install_item
dists = self.install_eggs(spec, download, tmpdir)
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/command/easy_install.py", line 890, in install_eggs
return self.build_and_install(setup_script, setup_base)
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/command/easy_install.py", line 1158, in build_and_install
self.run_setup(setup_script, setup_base, args)
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/command/easy_install.py", line 1144, in run_setup
run_setup(setup_script, args)
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/sandbox.py", line 253, in run_setup
raise
File "/usr/lib/python3.6/contextlib.py", line 99, in __exit__
self.gen.throw(type, value, traceback)
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/sandbox.py", line 195, in setup_context
yield
File "/usr/lib/python3.6/contextlib.py", line 99, in __exit__
self.gen.throw(type, value, traceback)
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/sandbox.py", line 166, in save_modules
saved_exc.resume()
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/sandbox.py", line 141, in resume
six.reraise(type, exc, self._tb)
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/_vendor/six.py", line 685, in reraise
raise value.with_traceback(tb)
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/sandbox.py", line 154, in save_modules
yield saved
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/sandbox.py", line 195, in setup_context
yield
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/sandbox.py", line 250, in run_setup
_execfile(setup_script, ns)
File "/usr/src/test-venv/lib/python3.6/site-packages/setuptools/sandbox.py", line 45, in _execfile
exec(code, globals, locals)
File "/tmp/easy_install-exzgb8j1/numpy-1.20.0rc1/setup.py", line 30, in <module>
Official docs: [http://pymatgen.org](http://pymatgen.org/)
RuntimeError: Python version >= 3.7 required.
----------------------------------------
ERROR: Command errored out with exit status 1: python setup.py egg_info Check the logs for full command output.
(test-venv) jenkins@ip:~/workspace/PR-212$
```
**Desktop:**
- OS: Ubuntu 16.04.5 LTS
**Additional context**
It seems it tries to download `numpy-1.20.0rc1` which requires python >= 3.7. I also tried installing a lower version of `numpy` (1.15.1) and then installing `pymatgen` but it didn't work as well.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2010/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2010/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2011
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2011/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2011/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2011/events
|
https://github.com/materialsproject/pymatgen/pull/2011
| 760,549,908
|
MDExOlB1bGxSZXF1ZXN0NTM1MzYyOTA2
| 2,011
|
Add IsayevNN near neighbor class
|
{
"login": "utf",
"id": 1330638,
"node_id": "MDQ6VXNlcjEzMzA2Mzg=",
"avatar_url": "https://avatars.githubusercontent.com/u/1330638?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/utf",
"html_url": "https://github.com/utf",
"followers_url": "https://api.github.com/users/utf/followers",
"following_url": "https://api.github.com/users/utf/following{/other_user}",
"gists_url": "https://api.github.com/users/utf/gists{/gist_id}",
"starred_url": "https://api.github.com/users/utf/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/utf/subscriptions",
"organizations_url": "https://api.github.com/users/utf/orgs",
"repos_url": "https://api.github.com/users/utf/repos",
"events_url": "https://api.github.com/users/utf/events{/privacy}",
"received_events_url": "https://api.github.com/users/utf/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @utf!"
] | 2020-12-09T18:04:16
| 2020-12-09T21:05:29
|
2020-12-09T21:05:29Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Added `IsayevNN`, a class for finding near neighbors based on: [10.1038/ncomms15679](http://www.nature.com/articles/ncomms15679).
Two sites are considered coordinated if: (i) they share a Voronoi face and (ii) the near neighbor distance is less than the sum of the covalent radii + 0.25 Å.
Also added tests.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2011/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2011/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2011",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2011",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2011.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2011.patch",
"merged_at": "2020-12-09T21:05:29Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2012
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2012/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2012/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2012/events
|
https://github.com/materialsproject/pymatgen/issues/2012
| 760,746,782
|
MDU6SXNzdWU3NjA3NDY3ODI=
| 2,012
|
Proposed change to how electrode are defined
|
{
"login": "jmmshn",
"id": 14003693,
"node_id": "MDQ6VXNlcjE0MDAzNjkz",
"avatar_url": "https://avatars.githubusercontent.com/u/14003693?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jmmshn",
"html_url": "https://github.com/jmmshn",
"followers_url": "https://api.github.com/users/jmmshn/followers",
"following_url": "https://api.github.com/users/jmmshn/following{/other_user}",
"gists_url": "https://api.github.com/users/jmmshn/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jmmshn/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jmmshn/subscriptions",
"organizations_url": "https://api.github.com/users/jmmshn/orgs",
"repos_url": "https://api.github.com/users/jmmshn/repos",
"events_url": "https://api.github.com/users/jmmshn/events{/privacy}",
"received_events_url": "https://api.github.com/users/jmmshn/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"NOte that db \"friendliness\" is a distant consideration to actual user-friendliness.\r\nIn this case, I do agree using dataclasses is better. The only reason why it wasn't done in the past is to maintain backwards compatibility. Since most other packages have abandoned py36, there is nothing stopping us from doing dataclasses anymore.",
"@shyuep Thanks for the quick reply! I will make these changes and create a PR."
] | 2020-12-09T23:17:22
| 2020-12-12T20:30:14
|
2020-12-12T20:30:14Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Is your feature request related to a problem? Please describe.**
The current implementation of the insertion and conversion electrodes are not very db friendly.
The information that users will interface with is essentially the data defined in the abstract properties of each class in addition to the `_vpair` parameter.
However, the current implementation has custom `as_dict` and `from_dict` using the input entries.
**Describe the solution you'd like**
Convert the Abstract classes to `dataclass` that subclass `MSONables`.
This automatically creates the optimal `__init__`, `as_dict`, `from_dict` functions for the electrodes and voltages pairs.
Additionally, this will make the code significantly smaller
I tested the interaction between `MSONable` and `dataclass` and it seems to behave as expected.
```
from dataclasses import dataclass
from monty.json import MSONable
@dataclass
class TestClass(MSONable):
"""Class for keeping track of an item in inventory."""
name: str
unit_price: float
quantity_on_hand: int = 0
tmp = TestClass(name="J", unit_price=12)
dd = tmp.as_dict()
print(A.from_dict(dd))
```
@mkhorton if you are ok with this I will implement ASAP.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2012/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2012/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2013
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2013/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2013/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2013/events
|
https://github.com/materialsproject/pymatgen/pull/2013
| 760,862,209
|
MDExOlB1bGxSZXF1ZXN0NTM1NjE3MDU4
| 2,013
|
Changed electrode document to dataclasses
|
{
"login": "jmmshn",
"id": 14003693,
"node_id": "MDQ6VXNlcjE0MDAzNjkz",
"avatar_url": "https://avatars.githubusercontent.com/u/14003693?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jmmshn",
"html_url": "https://github.com/jmmshn",
"followers_url": "https://api.github.com/users/jmmshn/followers",
"following_url": "https://api.github.com/users/jmmshn/following{/other_user}",
"gists_url": "https://api.github.com/users/jmmshn/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jmmshn/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jmmshn/subscriptions",
"organizations_url": "https://api.github.com/users/jmmshn/orgs",
"repos_url": "https://api.github.com/users/jmmshn/repos",
"events_url": "https://api.github.com/users/jmmshn/events{/privacy}",
"received_events_url": "https://api.github.com/users/jmmshn/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @jmmshn, this is great"
] | 2020-12-10T03:50:11
| 2020-12-11T18:18:44
|
2020-12-11T18:18:41Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
PR to address #2012
- Changed the `AbstractVoltagePair` and `AbstractElectrode` into `dataclass`s
- Changed the `InsertionVoltagePair` and `InsertionElectrode` into `dataclass`s
- For `InsertionVoltagePair` and `InsertionElectrode`, moved original entries based init into `from_entires` function
- For `InsertionElectrode`, moved original `as_dict` `from_dict` to `*_legacy`
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2013/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2013/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2013",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2013",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2013.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2013.patch",
"merged_at": "2020-12-11T18:18:41Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2014
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2014/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2014/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2014/events
|
https://github.com/materialsproject/pymatgen/pull/2014
| 762,949,388
|
MDExOlB1bGxSZXF1ZXN0NTM3NDQ3MzY3
| 2,014
|
feature: more standard approach to __hash__ and __eq__ for Entry and children
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thank you @CompRhys ! Looks like you're missing a newline character in pymatgen.entries.__init__.py. Otherwise looks good to me. I tested with the MP2020 corrections enabled as well and saw no issues.",
"> What is this new TransformedPDEntry logic for? What is `sp_mapping`? Can you add docstrings and maybe rename `sp_mapping` to something that is a bit more easily readable and understandable? There's a lot of changes to PD. Can you summarize what is going on and why? \r\n\r\nI missed that I hadn't updated the docstring here, I will do it now! I haven't changed the logic behind `TransformedPDEntry` just moved in from inside `CompoundPhaseDiagram`, the reason for this was that previously `TransformedPDEntry` didn't have any information about what the terminal compositions were. \r\n\r\n`sp_mapping` is the species mapping between dummy species and the terminal compositions. I inherited the name from `CompoundPhaseDiagram` but can change it to something clearer? \r\n\r\nThe other changes to PD are only cosmetic/for clarity \r\n\r\n1. several are due to swapping older `.. attribute:: energy` style docstrings to the google docstring format.\r\n2. some are to use list comprehensions rather than for loops that append to a set/list (not that this speed difference is significant here)\r\n3. `chempot_range_map` had some anti-patterns of using `pd=self` then using `pd.method()` than I took out\r\n4. `finalfacets` became `final_facets` to be more readable also `allfacets` to `all_facets`\r\n5. removing strange auto-formatting edge cases e.g. assigning `test_open` in line 1187 because it was previously being computed multiple times and each time it was being spread over 5 lines by the auto-formatting used.\r\n6. I also removed the duplicate checks from `get_equilibirum_reaction_energy` that caused issues with `ComputedEntry` as it was indicated that this degree of hand holding wasn't typical for `pymatgen` (although the changes to `__hash__` alone would fix the bug)\r\n7. I have disabled my `get_quasi_e_to_hull` for `ComputedEntry` until I can check all the edge cases about whether the duplicate checking can be removed there. This seemed to be the best option last week when @rkingsbury asked me to split out these changes from the other PR.",
"> Some modifications i've suggested. @mattmcdermott has said he no longer needs the hash consistency and in general, it's not a good idea to override salted hashes for `__hash__`. This was done on purpose early on in python 3 to prevent insecurities in dictionaries and sets.\r\n\r\nshall I remove the `hashlib` dependency and go back to hashing the strings? It depends on whether it's likely that other people encounter the same consistency issues I can't think of any security issues but that doesn't mean they don't exist",
"I think unless we have a compelling use case for using unsalted hashes, we should continue to use salted hashes. Basically, the problem has to do with inserting objects into dictionaries, but it also is an issue for sets. This was solved in 2012 by salting the builtin hash function: https://bugs.python.org/issue13703\r\n\r\n> > Some modifications i've suggested. @mattmcdermott has said he no longer needs the hash consistency and in general, it's not a good idea to override salted hashes for `__hash__`. This was done on purpose early on in python 3 to prevent insecurities in dictionaries and sets.\r\n> \r\n> shall I remove the `hashlib` dependency and go back to hashing the strings? It depends on whether it's likely that other people encounter the same consistency issues I can't think of any security issues but that doesn't mean they don't exist\r\n\r\n",
"I ended up not adding temperature to the `GibbsComputedStructureEntry` to try make this PR as simple as possible. If it appears that this affects performance then it could be easily added at a later date.\r\n\r\nI removed `hashlib` and now just use the default python hash. \r\n\r\nI removed my `per_atom` kwarg due to the potential issues highlighted by @shyamd. ",
"I checked all the edge cases I could think of with `get_quasi_e_to_hull` and `ComputedEntry` and tweaked the normalizations to eliminate the few that did cause issues.\r\n\r\nI added a note to `__eq__` in the `Entry` class highlighting that the `e.as_dict()` comparison isn't robust to floating point changes in the energy which can arise following repeated normalizations. I tweaked `get_quasi_e_to_hull` to avoid this and doubt that it would come up anywhere else. ",
"The lack of robustness in equality using the dict format is a problem. That's why I suggest explicitly breaking out what is being compared and using the appropriate comparator such as `np.allclose` ",
"I'm going to unapprove this PR @CompRhys because `__eq__` should be 100% reliable. I think the only viable way is to explicitly check each parameter introduced by that class and then doing a super `__eq__` rather than relying on the dict equals. ",
"A potential problem with that is that lots of the `Entry` children have there own dictionaries optional data arguments i.e. `data` and `parameters` in `ComputedEntry` or `chempots` for `GrandPotPDEntry` and I want to avoid going down that rabbit hole.\r\n\r\nOne thing to note is that even if the dictionary comparison isn't 100% robust for a lot of use cases it is more robust than \r\n```\r\n def __eq__(self, other):\r\n return self is other\r\n \r\n def __hash__(self):\r\n return hash(id(self)/16)\r\n``` \r\nfor which is currently the case for `ComputedEntry` or\r\n```\r\n def __eq__(self, other):\r\n if isinstance(other, self.__class__):\r\n return self.as_dict() == other.as_dict()\r\n else:\r\n return False\r\n\r\n def __hash__(self):\r\n return id(self)\r\n```\r\nwhich was the original `__hash__` and `__eq__` for `PDEntry`. \r\n\r\nAs I was told it was \"urgent\" to fix this issue and given that this is equivalent or better in terms of robustness than what is currently in the code I think that it would probably be better for someone on the core team to take over this PR as this now seems to be more of a core issue where a wider knowledge of how robust is robust enough in `pymatgen` would be needed -- it's certainly not the small issue that was holding back my `PatchedPhaseDiagram` contribution I initially thought i was trying to fix.",
"We have a project that would be much easier to proceed with if this hashing issue and #1974 were resolved. Would be great if, as @CompRhys suggested, @mkhorton or someone else on the core team could have another look at this PR to help get it merged.",
"Let's leave it as-is then and merge this in as there isn't a \"core\" team. Pymatgen is very much a community-driven effort. I will say that while this works out now, it's almost a guarantee that anyone that relies on Entries will hit a bug they can't diagnose because of the floating-point issue. ",
"Thanks! After finishing #1974 I will come back and have a look at whether we can further improve the robustness here and what equality actually means in terms of `Entries` and how to deal with the edge cases thrown up by the optional `parameters`, `attributes`, `data` attributes.",
"Thanks @CompRhys! If we ever hand out annual PRs-with-most-comments-responded-to award I'll make sure you're on the list :-) "
] | 2020-12-11T21:47:46
| 2021-01-27T13:18:50
|
2021-01-27T01:56:36Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Split out the Entry changes as cleanly as possible from #1974 @rkingsbury
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2014/reactions",
"total_count": 3,
"+1": 3,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2014/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2014",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2014",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2014.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2014.patch",
"merged_at": "2021-01-27T01:56:36Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2015
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2015/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2015/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2015/events
|
https://github.com/materialsproject/pymatgen/issues/2015
| 762,985,844
|
MDU6SXNzdWU3NjI5ODU4NDQ=
| 2,015
|
LammpsData.from_file does not read in bond types, angle types, etc.
|
{
"login": "tawe141",
"id": 13124532,
"node_id": "MDQ6VXNlcjEzMTI0NTMy",
"avatar_url": "https://avatars.githubusercontent.com/u/13124532?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/tawe141",
"html_url": "https://github.com/tawe141",
"followers_url": "https://api.github.com/users/tawe141/followers",
"following_url": "https://api.github.com/users/tawe141/following{/other_user}",
"gists_url": "https://api.github.com/users/tawe141/gists{/gist_id}",
"starred_url": "https://api.github.com/users/tawe141/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/tawe141/subscriptions",
"organizations_url": "https://api.github.com/users/tawe141/orgs",
"repos_url": "https://api.github.com/users/tawe141/repos",
"events_url": "https://api.github.com/users/tawe141/events{/privacy}",
"received_events_url": "https://api.github.com/users/tawe141/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-12-11T22:21:17
| 2023-08-13T16:34:12
|
2023-08-13T16:34:12Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Reading a LAMMPS data file via `LammpsData.from_file()` does not read in the bond types, angle types, dihedral types, and improper types.
**To Reproduce**
```
from pymatgen.io.lammps.data import LammpsData
ld = LammpsData.from_file(filepath)
print(ld)
```
gives the output:
```
Generated by pymatgen.io.lammps.data.LammpsData
174 atoms
201 bonds
420 angles
438 dihedrals
54 impropers
107 atom types # bond types, angle types, etc. should be below this line
0.000000 21.657000 xlo xhi
0.000000 18.755512 ylo yhi
0.000000 6.766600 zlo zhi
-10.828500 0.000000 0.000000 xy xz yz
...
```
Here's the data file to reproduce this: https://www.dropbox.com/s/8rezwjb5rs9hddn/system.data?dl=0
**Desktop (please complete the following information):**
- OS: Mac OSX 10.15.17
- Version: 2020.12.3
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2015/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2015/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2016
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2016/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2016/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2016/events
|
https://github.com/materialsproject/pymatgen/pull/2016
| 765,860,392
|
MDExOlB1bGxSZXF1ZXN0NTM5MTEyODA2
| 2,016
|
Further cleanup of Electrode objects
|
{
"login": "jmmshn",
"id": 14003693,
"node_id": "MDQ6VXNlcjE0MDAzNjkz",
"avatar_url": "https://avatars.githubusercontent.com/u/14003693?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jmmshn",
"html_url": "https://github.com/jmmshn",
"followers_url": "https://api.github.com/users/jmmshn/followers",
"following_url": "https://api.github.com/users/jmmshn/following{/other_user}",
"gists_url": "https://api.github.com/users/jmmshn/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jmmshn/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jmmshn/subscriptions",
"organizations_url": "https://api.github.com/users/jmmshn/orgs",
"repos_url": "https://api.github.com/users/jmmshn/repos",
"events_url": "https://api.github.com/users/jmmshn/events{/privacy}",
"received_events_url": "https://api.github.com/users/jmmshn/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Yes, please add a deprecation decorator to `as_dict_summary()`, and we can remove at a later date. Thanks!",
"@mkhorton I made some additional (independent) changes to the plotter.",
"Can you make `capacity` synonymous with `capacity_grav`? e.g. just add an `or` in line 73, for backwards compatibility (the docstring can be left as-is), I'm not sure if features in the current materials project website rely on the existing behavior",
"OK done. Also maybe we should consider raising the version number up from 0.1.",
"I'm not sure the version numbers are useful, we don't have a policy for what they mean or when they're incremented",
"Thanks @jmmshn!"
] | 2020-12-14T04:08:43
| 2020-12-17T22:21:57
|
2020-12-17T22:21:54Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
- Change `ConversionElectrode` to type `dataclass` similar to the change to `InsertionElectrode` in #2013
- Added `x_charge` and `x_discharge` properties to report the number of working ion ver formula unit
- Standardized how the summary dictionary is created, now the base Electrode class has a `get_summary_dict` function which can be improved upon by the Insertion and Conversion objects.
- Added some more tests of the `summary_dict`
The old `as_dict_summary` methods are still around but maybe should be deprecated?
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2016/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2016/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2016",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2016",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2016.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2016.patch",
"merged_at": "2020-12-17T22:21:54Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2017
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2017/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2017/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2017/events
|
https://github.com/materialsproject/pymatgen/pull/2017
| 768,120,140
|
MDExOlB1bGxSZXF1ZXN0NTQwNjQ5MTk5
| 2,017
|
[WIP] add pretty_parity_plot function
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This looks useful but I'm trying not to merge new matplotlib plotting into pymatgen in favor of plotly-based plots which allow for more interactive data exploration. If you are using matplotlib, the [seaborn](https://seaborn.pydata.org) wrapper has some nice options for this kind of plot, including marginal histograms and 2D histograms, which can help deal with with data overlap issues.",
"> This looks useful but I'm trying not to merge new matplotlib plotting into pymatgen in favor of plotly-based plots which allow for more interactive data exploration. If you are using matplotlib, the [seaborn](https://seaborn.pydata.org) wrapper has some nice options for this kind of plot, including marginal histograms and 2D histograms, which can help deal with with data overlap issues.\r\n\r\nOK, it sounds like the method in this PR needs to find a different home then. I do like plotly but personally find it very difficult to generate polished / publication-ready figures with it. I think matplotlib is still superior in that regard and use the existing `pretty_plot` class regularly for that purpose.",
"I personally have found no significant difference between them for publication, plotly's default template seems heavily inspired by ggplot2 so can look a bit different, but it has a white-background template if that's preferred. I find their Jupyter extension useful for quick edits of the chart too and being able to save the chart to JSON and make edits after the fact, independently of the code used to generate the chart, has been useful. But not trying to sell anyone on this; use in pymatgen is purely because it allows easier data exploration, and makes it easier to transfer plots to web apps (so they're 1-to-1 equivalent between publication and website)."
] | 2020-12-15T19:58:27
| 2021-07-15T14:31:43
|
2020-12-18T01:13:47Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Adds a new plotting function `pretty_parity_plot` to facilitate creating customizable and nicely-formatted plots that compare two quantities. An example is shown below.
* Most common plot customizations are exposed through explicitly documented kwargs
* Automatically plot or hide linear regression line, regression statistics, and 1:1 line
* 'units' are automatically appended to both axes and error metrics (if visible)
* Robust enough to plot data that includes `np.nan` / `np.inf`
* Optional `ax` kwarg allows plotting directly to a subplot axis of another figure
## Example
```
parityplot(x,
y,
"$\Delta H_f^{R2SCAN}$",
"$\Delta H_f^{SCAN}$",
units="eV/atom",
show_regression_line=False,
show_diagonal=True,
regression_ls='k-'
)
```
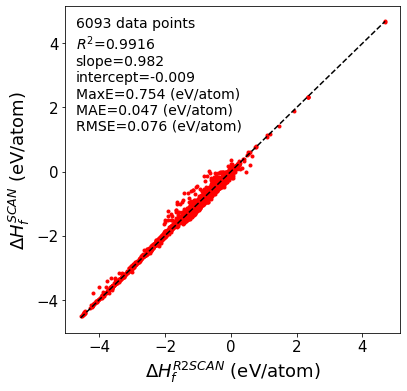
## Additional dependencies introduced (if any)
* `scikit-learn` for calculation of error metrics (see TODO)
## TODO (if any)
- [ ] Find an alternative to `sklearn.metrics` to avoid introducing a new dependency (probably will just hand-code the calculations of RMSE, max error, mean absolute error)
- [ ] Implement the `ax` kwarg
- [ ] Determine what kind of tests are needed
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2017/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2017/timeline
| null | true
| true
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2017",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2017",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2017.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2017.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2018
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2018/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2018/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2018/events
|
https://github.com/materialsproject/pymatgen/pull/2018
| 772,338,911
|
MDExOlB1bGxSZXF1ZXN0NTQzNjIyNjY1
| 2,018
|
Minor change to definition of InsertionVoltagePair
|
{
"login": "jmmshn",
"id": 14003693,
"node_id": "MDQ6VXNlcjE0MDAzNjkz",
"avatar_url": "https://avatars.githubusercontent.com/u/14003693?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jmmshn",
"html_url": "https://github.com/jmmshn",
"followers_url": "https://api.github.com/users/jmmshn/followers",
"following_url": "https://api.github.com/users/jmmshn/following{/other_user}",
"gists_url": "https://api.github.com/users/jmmshn/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jmmshn/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jmmshn/subscriptions",
"organizations_url": "https://api.github.com/users/jmmshn/orgs",
"repos_url": "https://api.github.com/users/jmmshn/repos",
"events_url": "https://api.github.com/users/jmmshn/events{/privacy}",
"received_events_url": "https://api.github.com/users/jmmshn/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @jmmshn!"
] | 2020-12-21T17:22:56
| 2021-01-04T19:53:45
|
2021-01-04T19:53:38Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Moved the entry_charge/discharge into the data model.
This allows the from_dict to recover the entries after serialization.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2018/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2018/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2018",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2018",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2018.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2018.patch",
"merged_at": "2021-01-04T19:53:38Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2019
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2019/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2019/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2019/events
|
https://github.com/materialsproject/pymatgen/pull/2019
| 772,537,792
|
MDExOlB1bGxSZXF1ZXN0NTQzNzg2NjM2
| 2,019
|
Improvement to continuous band structure transformation method
|
{
"login": "munrojm",
"id": 6877334,
"node_id": "MDQ6VXNlcjY4NzczMzQ=",
"avatar_url": "https://avatars.githubusercontent.com/u/6877334?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/munrojm",
"html_url": "https://github.com/munrojm",
"followers_url": "https://api.github.com/users/munrojm/followers",
"following_url": "https://api.github.com/users/munrojm/following{/other_user}",
"gists_url": "https://api.github.com/users/munrojm/gists{/gist_id}",
"starred_url": "https://api.github.com/users/munrojm/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/munrojm/subscriptions",
"organizations_url": "https://api.github.com/users/munrojm/orgs",
"repos_url": "https://api.github.com/users/munrojm/repos",
"events_url": "https://api.github.com/users/munrojm/events{/privacy}",
"received_events_url": "https://api.github.com/users/munrojm/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This is great, thanks for adding @munrojm !"
] | 2020-12-21T23:53:56
| 2021-01-04T19:54:29
|
2021-01-04T19:54:28Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This PR improves the `get_continuous_path` method of `HighSymmKpath` by returning a new complete `BandStructureSymmLine` object instead of just the new path labels and distance segment mappings.
Additionally, the docstring for `KPathLatimerMunro` has been updated to list the appropriate publication.
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2019/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2019/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2019",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2019",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2019.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2019.patch",
"merged_at": "2021-01-04T19:54:28Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2020
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2020/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2020/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2020/events
|
https://github.com/materialsproject/pymatgen/pull/2020
| 774,530,952
|
MDExOlB1bGxSZXF1ZXN0NTQ1NDQxMDY0
| 2,020
|
CREST Basic I/O
|
{
"login": "arepstein",
"id": 17616763,
"node_id": "MDQ6VXNlcjE3NjE2NzYz",
"avatar_url": "https://avatars.githubusercontent.com/u/17616763?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/arepstein",
"html_url": "https://github.com/arepstein",
"followers_url": "https://api.github.com/users/arepstein/followers",
"following_url": "https://api.github.com/users/arepstein/following{/other_user}",
"gists_url": "https://api.github.com/users/arepstein/gists{/gist_id}",
"starred_url": "https://api.github.com/users/arepstein/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/arepstein/subscriptions",
"organizations_url": "https://api.github.com/users/arepstein/orgs",
"repos_url": "https://api.github.com/users/arepstein/repos",
"events_url": "https://api.github.com/users/arepstein/events{/privacy}",
"received_events_url": "https://api.github.com/users/arepstein/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/40638027)\n\nCoverage decreased (-0.6%) to 83.093% when pulling **ba54f14de5d865f3ae941a0344ee3c4ff716f537 on arepstein:ae_Crest** into **14150a9d631996686cdc386c36272f96cf8d7e99 on materialsproject:master**.\n",
"I'm going to merge this since the only issue is the same listing issue affecting the main branch and otherwise this PR looks excellent -- thank you @arepstein !"
] | 2020-12-24T18:18:41
| 2021-06-17T00:26:51
|
2021-06-17T00:26:51Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
- Added CRESTOutput to parse CREST output directories in pymatgen.io.xtb.outputs
- Added CRESTInput to write CREST input files in pymatgen.io.xtb.inputs
- Added tests and test files for CRESTOutput and CRESTInput
## Additional dependencies introduced (if any)
- None
## TODO (if any)
- XTB i/o still missing
- CRESTOutput supports the default conformer searching method, but needs tests for non-default methods.
- CRESTInput supports writing a coordinate file and a .constrains file, but other files might be valuable to add
- Add parsing of .xcontrol
- Add improved error handling methods
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [ x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [ x] Tests have been added for any new functionality or bug fixes.
- [ x] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2020/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2020/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2020",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2020",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2020.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2020.patch",
"merged_at": "2021-06-17T00:26:51Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2021
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2021/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2021/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2021/events
|
https://github.com/materialsproject/pymatgen/issues/2021
| 774,904,119
|
MDU6SXNzdWU3NzQ5MDQxMTk=
| 2,021
|
Pymatgen does not support reading EIGENVAL files from VASP < 5.4
|
{
"login": "mdforti",
"id": 30444792,
"node_id": "MDQ6VXNlcjMwNDQ0Nzky",
"avatar_url": "https://avatars.githubusercontent.com/u/30444792?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mdforti",
"html_url": "https://github.com/mdforti",
"followers_url": "https://api.github.com/users/mdforti/followers",
"following_url": "https://api.github.com/users/mdforti/following{/other_user}",
"gists_url": "https://api.github.com/users/mdforti/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mdforti/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mdforti/subscriptions",
"organizations_url": "https://api.github.com/users/mdforti/orgs",
"repos_url": "https://api.github.com/users/mdforti/repos",
"events_url": "https://api.github.com/users/mdforti/events{/privacy}",
"received_events_url": "https://api.github.com/users/mdforti/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 4812645414,
"node_id": "LA_kwDOACgets8AAAABHtskJg",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/outdated",
"name": "outdated",
"color": "2A890C",
"default": false,
"description": "Issues resulting from using outdated versions of pymatgen or dependencies"
},
{
"id": 5304335471,
"node_id": "LA_kwDOACgets8AAAABPCm8bw",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/io",
"name": "io",
"color": "c2e0c6",
"default": false,
"description": "Input/output functionality"
},
{
"id": 5457739150,
"node_id": "LA_kwDOACgets8AAAABRU59jg",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/vasp",
"name": "vasp",
"color": "BF4B01",
"default": false,
"description": "Vienna Ab initio Simulation Package"
},
{
"id": 5756346769,
"node_id": "LA_kwDOACgets8AAAABVxrhkQ",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/wont%20fix",
"name": "wont fix",
"color": "0B4643",
"default": false,
"description": "Won't be worked on due to lack of priority"
}
] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @mdforti, thanks for reporting this. Would you willing to prepare a PR to fix?",
"yes, of course. hope I do it correctly.",
"Note to @janosh: The correct description here should be that Pymatgen does not support reading EIGENVAL files from pre-VASP 5.4, as noted in #2403."
] | 2020-12-26T17:02:25
| 2023-08-13T16:34:12
|
2023-08-13T16:34:12Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
https://github.com/materialsproject/pymatgen/blob/f6aa22ae178f3ea1e0332e8ec2a7731d6b260548/pymatgen/io/vasp/outputs.py#L5526
the line referenced skips if the eigenvalue table for a kpoint has only two columns. an option should be added for non spin polarized calculations where only two columns are present, one for band number and another for corresponding eigenvalue.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2021/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2021/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2022
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2022/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2022/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2022/events
|
https://github.com/materialsproject/pymatgen/pull/2022
| 775,252,064
|
MDExOlB1bGxSZXF1ZXN0NTQ1OTcxNjAz
| 2,022
|
fix MoS test filename in test_graph.py
|
{
"login": "drew-parsons",
"id": 26508288,
"node_id": "MDQ6VXNlcjI2NTA4Mjg4",
"avatar_url": "https://avatars.githubusercontent.com/u/26508288?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/drew-parsons",
"html_url": "https://github.com/drew-parsons",
"followers_url": "https://api.github.com/users/drew-parsons/followers",
"following_url": "https://api.github.com/users/drew-parsons/following{/other_user}",
"gists_url": "https://api.github.com/users/drew-parsons/gists{/gist_id}",
"starred_url": "https://api.github.com/users/drew-parsons/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/drew-parsons/subscriptions",
"organizations_url": "https://api.github.com/users/drew-parsons/orgs",
"repos_url": "https://api.github.com/users/drew-parsons/repos",
"events_url": "https://api.github.com/users/drew-parsons/events{/privacy}",
"received_events_url": "https://api.github.com/users/drew-parsons/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks for the fix @drew-parsons!\r\n\r\nSince this is your first contribution, please do fill out [the form here](https://forms.gle/JnisFb38QDR8QTFTA) so that we can [acknowledge you appropriately](https://pymatgen.org/team.html). I know this is only a quick fix, but we want to make sure we acknowledge all contributions :-)"
] | 2020-12-28T07:30:13
| 2021-01-04T19:56:23
|
2021-01-04T19:56:23Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
test file is MoS2_single.pdf not MOS2_single.pdf
"MOS2_single.pdf" triggers an error: "No such file or directory: 'MOS2_single.pdf'" from `os.remove(test_file)`
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2022/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2022/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2022",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2022",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2022.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2022.patch",
"merged_at": "2021-01-04T19:56:22Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2023
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2023/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2023/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2023/events
|
https://github.com/materialsproject/pymatgen/issues/2023
| 775,261,368
|
MDU6SXNzdWU3NzUyNjEzNjg=
| 2,023
|
test_wulff index error in WulffShapeTest (test_get_plotly)
|
{
"login": "drew-parsons",
"id": 26508288,
"node_id": "MDQ6VXNlcjI2NTA4Mjg4",
"avatar_url": "https://avatars.githubusercontent.com/u/26508288?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/drew-parsons",
"html_url": "https://github.com/drew-parsons",
"followers_url": "https://api.github.com/users/drew-parsons/followers",
"following_url": "https://api.github.com/users/drew-parsons/following{/other_user}",
"gists_url": "https://api.github.com/users/drew-parsons/gists{/gist_id}",
"starred_url": "https://api.github.com/users/drew-parsons/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/drew-parsons/subscriptions",
"organizations_url": "https://api.github.com/users/drew-parsons/orgs",
"repos_url": "https://api.github.com/users/drew-parsons/repos",
"events_url": "https://api.github.com/users/drew-parsons/events{/privacy}",
"received_events_url": "https://api.github.com/users/drew-parsons/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-12-28T07:52:24
| 2023-08-13T16:34:13
|
2023-08-13T16:34:13Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
WulffShapeTest (via test_get_plotly) from test_wulff.py is failing on Debian systems (python3.9, numpy 1.19.4)
The error message is
```
ERROR: test_get_plotly (pymatgen.analysis.tests.test_wulff.WulffShapeTest)
----------------------------------------------------------------------
Traceback (most recent call last):
File "/projects/pymatgen/.pybuild/cpython3_3.9_pymatgen/build/pymatgen/analysis/tests/test_wulff.py", line 90, in test_get_plotly
self.wulff_Ti.get_plotly()
File "/projects/pymatgen/.pybuild/cpython3_3.9_pymatgen/build/pymatgen/analysis/wulff.py", line 624, in get_plotly
i=tri_indices[0],
IndexError: too many indices for array: array is 0-dimensional, but 1 were indexed
```
The error is reproducible from the toplevel source dir running test_wullf.py directly, `python3 pymatgen/analysis/tests/test_wulff.py`
More precisely, with binary installation ( including cython .so extensions) located in .pybuild/cpython3_3.9_pymatgen/build/ ,
```
$ PYTHONPATH=/projects/pymatgen/.pybuild/cpython3_3.9_pymatgen/build/ python3 /projects/pymatgen/.pybuild/cpython3_3.9_pymatgen/build/pymatgen/analysis/tests/test_wulff.py -v
test_corner_and_edges (__main__.WulffShapeTest) ... ok
test_get_azimuth_elev (__main__.WulffShapeTest) ... ok
test_get_plot (__main__.WulffShapeTest) ... ok
test_get_plotly (__main__.WulffShapeTest) ... ERROR
test_properties (__main__.WulffShapeTest) ... ok
======================================================================
ERROR: test_get_plotly (__main__.WulffShapeTest)
----------------------------------------------------------------------
Traceback (most recent call last):
File "/projects/pymatgen/.pybuild/cpython3_3.9_pymatgen/build/pymatgen/analysis/tests/test_wulff.py", line 90, in test_get_plotly
self.wulff_Ti.get_plotly()
File "/projects/pymatgen/.pybuild/cpython3_3.9_pymatgen/build/pymatgen/analysis/wulff.py", line 624, in get_plotly
i=tri_indices[0],
IndexError: too many indices for array: array is 0-dimensional, but 1 were indexed
----------------------------------------------------------------------
Ran 5 tests in 1.403s
FAILED (errors=1)
```
**Desktop:**
- OS: Debian GNU/Linux (debian unstable)
- python 3.9
- numpy 1.19.4
- plotly 4.12.0
- cython3 0.29.21
- gcc 10.2.1
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2023/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2023/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2024
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2024/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2024/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2024/events
|
https://github.com/materialsproject/pymatgen/issues/2024
| 775,468,761
|
MDU6SXNzdWU3NzU0Njg3NjE=
| 2,024
|
feff_input_generation executable is broken: No module named 'pymatgen.cli.feff_input_generation'
|
{
"login": "drew-parsons",
"id": 26508288,
"node_id": "MDQ6VXNlcjI2NTA4Mjg4",
"avatar_url": "https://avatars.githubusercontent.com/u/26508288?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/drew-parsons",
"html_url": "https://github.com/drew-parsons",
"followers_url": "https://api.github.com/users/drew-parsons/followers",
"following_url": "https://api.github.com/users/drew-parsons/following{/other_user}",
"gists_url": "https://api.github.com/users/drew-parsons/gists{/gist_id}",
"starred_url": "https://api.github.com/users/drew-parsons/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/drew-parsons/subscriptions",
"organizations_url": "https://api.github.com/users/drew-parsons/orgs",
"repos_url": "https://api.github.com/users/drew-parsons/repos",
"events_url": "https://api.github.com/users/drew-parsons/events{/privacy}",
"received_events_url": "https://api.github.com/users/drew-parsons/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Yes, this has been deprecated and removed. Pls use the modules directly to generate the input files.",
"But the executable scripts are still being generated. They are listed in the entry_points list in setup.py. Hence this bug is still open.",
"Yes, I just removed them in a new commit. This will be fixed in the next release.",
"I see it, thanks @shyuep ."
] | 2020-12-28T15:59:26
| 2020-12-29T01:15:03
|
2020-12-28T17:01:45Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
pymatgen generates a handful of executable files invoking various pymatgen functionality, including pmg,
gaussian_analyzer, feff_input_generation
feff_input_generation appears to be broken and fails on attempting to load self-referential python submodule:
```
$ feff_input_generation
Traceback (most recent call last):
File "/usr/bin/feff_input_generation", line 33, in <module>
sys.exit(load_entry_point('pymatgen==2020.12.18', 'console_scripts', 'feff_input_generation')())
File "/usr/bin/feff_input_generation", line 25, in importlib_load_entry_point
return next(matches).load()
File "/usr/lib/python3.9/importlib/metadata.py", line 77, in load
module = import_module(match.group('module'))
File "/usr/lib/python3.9/importlib/__init__.py", line 127, in import_module
return _bootstrap._gcd_import(name[level:], package, level)
File "<frozen importlib._bootstrap>", line 1030, in _gcd_import
File "<frozen importlib._bootstrap>", line 1007, in _find_and_load
File "<frozen importlib._bootstrap>", line 984, in _find_and_load_unlocked
ModuleNotFoundError: No module named 'pymatgen.cli.feff_input_generation'
```
feff_input_generation is the calling script. As far as I'm aware it should not be a submodule of pymatgen.cli as implied in this error message.
All the other executable scripts run fine.
**Desktop:**
- OS: Debian GNU/Linux
- Version debian/unstable
- Python 3.9
- cython3 0.29.21
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2024/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2024/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2025
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2025/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2025/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2025/events
|
https://github.com/materialsproject/pymatgen/issues/2025
| 775,669,834
|
MDU6SXNzdWU3NzU2Njk4MzQ=
| 2,025
|
enable installation CI testing
|
{
"login": "drew-parsons",
"id": 26508288,
"node_id": "MDQ6VXNlcjI2NTA4Mjg4",
"avatar_url": "https://avatars.githubusercontent.com/u/26508288?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/drew-parsons",
"html_url": "https://github.com/drew-parsons",
"followers_url": "https://api.github.com/users/drew-parsons/followers",
"following_url": "https://api.github.com/users/drew-parsons/following{/other_user}",
"gists_url": "https://api.github.com/users/drew-parsons/gists{/gist_id}",
"starred_url": "https://api.github.com/users/drew-parsons/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/drew-parsons/subscriptions",
"organizations_url": "https://api.github.com/users/drew-parsons/orgs",
"repos_url": "https://api.github.com/users/drew-parsons/repos",
"events_url": "https://api.github.com/users/drew-parsons/events{/privacy}",
"received_events_url": "https://api.github.com/users/drew-parsons/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Generally the tests are meant to be operated on a cloned repository of pymatgen, rather than being run on a pip install version. None of the test files are provided in the standard wheels or tar.gz distributions because they are hundreds of Mb. So these are mainly provided just in the repository for cloning by developers.\r\n\r\nNote that pymatgen already runs CI on the repo. These can be seen from the Github Actions.",
"Your response does not address the issue raised.\r\n\r\nIt needs to be possible for distributions to be able to test the module as installed. Testing the repo does not achieve that.",
"As I said, the test files are very large. Neither the tests nor the input files needed for testing are distributed with the installation. This is the reality with the kind of files we have. It is theoretically possible to do the tests such that those that do not require input files can be tested using some script. But that would test a minuscule fraction of what pymatgen does. If you have a good suggestion on a way to design the tests, I am all ears. As it is, pymatgen is tested on both Mac and Linux platforms. ",
"github CI testing does not achieve CI testing of distribution installations. We need to confirm, for instance, if new versions of numpy or other modules updated in Debian will break client packages such as pymatgen.\r\n\r\nThe pypi source tarball does not include the test files, but the github source tarball does include them.\r\n\r\nCurrently the test files are embedded in the github source directories alongside the python module files. I recommend placing test files in a separate directory with hierarchical structure mirroring the module files which they are testing. Something like\r\n\r\n```\r\npymatgen/\r\n __init__.py\r\n core/\r\n __init__.py\r\n ...\r\n ...\r\ntest_files/\r\n ATOMS\r\n AgO_kpath_test.cif\r\n ...\r\ntests/\r\n __init__.py\r\n test_init.py\r\n core/\r\n __init__.py\r\n slab.cif\r\n test_bonds.py\r\n ...\r\n ...\r\n```\r\nrather than\r\n```\r\npymatgen/\r\n __init__.py\r\n tests/\r\n __init__.py\r\n test_init.py\r\n core/\r\n __init__.py\r\n ...\r\n tests/\r\n __init__.py\r\n slab.cif\r\n test_bonds.py\r\n ...\r\n ...\r\ntest_files/\r\n ATOMS\r\n AgO_kpath_test.cif\r\n ...\r\ntests/\r\n __init__.py\r\n test_init.py\r\n core/\r\n slab.cif\r\n test_bonds.py\r\n ...\r\n ```\r\n\r\nIn that case `cd tests; pytest` could run the tests for all types of installation, whether in-source, pypi or distribution installation.\r\n\r\nThe relationship between the tests and the input files in the `test_files` dir can also be made more robust by allowing for a different location of the test_files dir, possibly checking a hierarchy of alternatives, possibly with an override that can be set at runtime, e.g. via environment variable PYMATGEN_TEST_FILES or via pytest_addoption in conf.py ",
"Ok, I have implemented your suggestion to use an environmental variable. You can specify the root location of test files using PMG_TEST_DIR with the code in the master branch. This will be available in the next release. I will not be making the rest of the changes, e.g., the structure of the tests hirerarchy since that will require too much maintenance. Note that having tests directories in the modules is standard practice (e.g., see numpy own tests).",
"Thanks @shyuep , I'll give PMG_TEST_DIR a try.",
"Use of PMG_TEST_FILES_DIR works well enough. Some tests were missed in the patch. I'll prepare a PR."
] | 2020-12-29T02:54:15
| 2021-08-23T21:55:37
|
2020-12-31T01:16:20Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
For Linux (and other) distributions, it would be good to be able to regularly test the pymatgen installation is working correctly (configuration integration CI testing).
pymatgen already provides an extensive amount of tests. They can't really be used with CI testing however. The location of input files is hardcoded to an outside directory, e.g. core/tests/test_surface.py uses `path = os.path.join(cwd, "..", "..", "..", "test_files", "surface_tests", path_str)`
If a distribution installs pymatgen into /usr/lib/python3/dist-packages/pymatgen, then it means input files would be expected in /usr/lib/python3/dist-packages/test_files, which is not the right place for this kind of data. /usr/share/pymatgen/test_files would be more appropriate.
Even in a private user installation, it means test files would be expected in ~/.local/lib/python3.9/site-packages/test_files, which again is not a good path for such data.
The pymatgen tests should allow for a different location of the test_files dir, possibly checking a hierarchy of alternatives, possibly with an override that can be set at runtime, e.g. via environment variable PYMATGEN_TEST_FILES or via pytest_addoption in conf.py (cf. https://docs.pytest.org/en/stable/example/simple.html#dynamically-adding-command-line-options)
Alternatively, tests could be placed in a separate tests dir, separate from the actual module dirs.
I'm preparing packaging for the Debian GNU/Linux distribution, but the point applies to other distributions.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2025/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2025/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2026
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2026/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2026/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2026/events
|
https://github.com/materialsproject/pymatgen/issues/2026
| 780,674,968
|
MDU6SXNzdWU3ODA2NzQ5Njg=
| 2,026
|
Calling specie from a site gives two different object classes
|
{
"login": "mm04926412",
"id": 21348397,
"node_id": "MDQ6VXNlcjIxMzQ4Mzk3",
"avatar_url": "https://avatars.githubusercontent.com/u/21348397?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mm04926412",
"html_url": "https://github.com/mm04926412",
"followers_url": "https://api.github.com/users/mm04926412/followers",
"following_url": "https://api.github.com/users/mm04926412/following{/other_user}",
"gists_url": "https://api.github.com/users/mm04926412/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mm04926412/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mm04926412/subscriptions",
"organizations_url": "https://api.github.com/users/mm04926412/orgs",
"repos_url": "https://api.github.com/users/mm04926412/repos",
"events_url": "https://api.github.com/users/mm04926412/events{/privacy}",
"received_events_url": "https://api.github.com/users/mm04926412/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I would need a bit more context the inconsistent behavior you are referring to. Python uses duck typing. Objects that basically implement a common subset of methods/properties can substitute for each other.\r\n\r\nThere are instances where people want to just use Element to represent the site, e.g., when they are not interested or do not know the oxidation state. For example, an alloy like CuAu. There are other instances where they want to specify a formal oxidation state. Say Li2O with Li+ and O2-. \r\n\r\nIn all cases, specie.Z and specie.element.Z should return exactly the same number. If it doesn't, something is seriously wrong. But I would need you to provide an example code demonstrating such a problem.",
"Using specie.element.Z where only an element is defined results in the error\r\n\r\n\"Object of type Element has no member element\" \r\n\r\nThe specific line where I get different results is \r\n\r\ntoken_list = torch.tensor(\r\n [int(i.specie.Z) for i in structure], dtype=torch.long\r\n )\r\n\r\nwhere\r\n\r\ntoken_list = torch.tensor(\r\n [int(i.specie.element.Z) for i in structure], dtype=torch.long\r\n )\r\n\r\nis stable for compounds with a defined oxidation state but gives \"Object of type Element has no member element\" otherwise.\r\n\r\nthe version I'm using is '2020.12.18'",
"Well, in general, you should not be using specie.element.Z since specie.Z basically works for both Element and Specie.\r\n",
"Ah ok that makes sense, I'm stil confused about the strange behaviour though. For context I have structure objects for the ICSD database and every now and again some of tensors generated by the list comprehension above have arbitary huge values in them with it furthermore being arbitrary if they are positive or negative.\r\n\r\nThis behaviour only happens with the i.specie.Z syntax and not i.specie.element.Z",
"The only way that happens is that your ICSD structure contains a specie notation that is not actually a valid element. Then a DummySpecie is generated, for which the Z is a nonsensical value.",
"Would it be possible to have this raise an Exception or default to nan? Casting to a seemingly random int - a behaviour not shared by species.Element.Z - seems troublesome.",
"That is not possible. There are many instances where we want the DummySpecie (e.g., to represent vacancies). You can catch this by checking whether the Z is < 100.",
"I understand that but you don't want DummySpecie().Z to ever be called right? It seems there is already precedent to return NaN when a request has no physical meaning?",
"We cannot use NaN. Otherwise a DummySpecie(X) and DummySpecie(Z) will have the same Z and that can result in issues when you want to treat multiple dummy specie."
] | 2021-01-06T16:19:29
| 2021-01-07T18:42:52
|
2021-01-07T18:42:51Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Depending on the source of the structure the return value for "Site.specie" is either an specie object or an element object. When I'm working with a database of pymatgen structures this seems to vary depending on the data source. This causes inconsistent behaviour with attempting to access the atomic number with .Z as specie.Z and specie.element.Z can report different values for some structures.
It seems like a very strange design choice to me that two different classes can be returned from the same method?
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2026/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2026/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2027
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2027/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2027/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2027/events
|
https://github.com/materialsproject/pymatgen/pull/2027
| 780,854,369
|
MDExOlB1bGxSZXF1ZXN0NTUwNjc4NzI4
| 2,027
|
QuasiRRHO Thermochemistry Analysis
|
{
"login": "arepstein",
"id": 17616763,
"node_id": "MDQ6VXNlcjE3NjE2NzYz",
"avatar_url": "https://avatars.githubusercontent.com/u/17616763?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/arepstein",
"html_url": "https://github.com/arepstein",
"followers_url": "https://api.github.com/users/arepstein/followers",
"following_url": "https://api.github.com/users/arepstein/following{/other_user}",
"gists_url": "https://api.github.com/users/arepstein/gists{/gist_id}",
"starred_url": "https://api.github.com/users/arepstein/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/arepstein/subscriptions",
"organizations_url": "https://api.github.com/users/arepstein/orgs",
"repos_url": "https://api.github.com/users/arepstein/repos",
"events_url": "https://api.github.com/users/arepstein/events{/privacy}",
"received_events_url": "https://api.github.com/users/arepstein/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-01-06T21:22:42
| 2021-01-06T21:55:41
|
2021-01-06T21:55:41Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Added quasirrho.py to the analysis subpackage to calculate the Quasi-RRHO free energy from a Gaussian or QChem frequency calculation.
* Calculates Grimme's Quasi-RRHO free energy
* Option for also correcting for solvent concentration
## Additional dependencies introduced (if any)
*None
## TODO (if any)
* The rotational symmetry number is required as an input. To calculate this on the fly, will consider updates to PointGroupAnalyzer
Before a pull request can be merged, the following items must be checked:
- [ x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x ] Tests have been added for any new functionality or bug fixes.
- [ x] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "arepstein",
"id": 17616763,
"node_id": "MDQ6VXNlcjE3NjE2NzYz",
"avatar_url": "https://avatars.githubusercontent.com/u/17616763?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/arepstein",
"html_url": "https://github.com/arepstein",
"followers_url": "https://api.github.com/users/arepstein/followers",
"following_url": "https://api.github.com/users/arepstein/following{/other_user}",
"gists_url": "https://api.github.com/users/arepstein/gists{/gist_id}",
"starred_url": "https://api.github.com/users/arepstein/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/arepstein/subscriptions",
"organizations_url": "https://api.github.com/users/arepstein/orgs",
"repos_url": "https://api.github.com/users/arepstein/repos",
"events_url": "https://api.github.com/users/arepstein/events{/privacy}",
"received_events_url": "https://api.github.com/users/arepstein/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2027/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2027/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2027",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2027",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2027.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2027.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2028
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2028/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2028/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2028/events
|
https://github.com/materialsproject/pymatgen/pull/2028
| 780,935,131
|
MDExOlB1bGxSZXF1ZXN0NTUwNzQ2MTQx
| 2,028
|
Quasi-RRHO Thermochemistry Analysis Module
|
{
"login": "arepstein",
"id": 17616763,
"node_id": "MDQ6VXNlcjE3NjE2NzYz",
"avatar_url": "https://avatars.githubusercontent.com/u/17616763?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/arepstein",
"html_url": "https://github.com/arepstein",
"followers_url": "https://api.github.com/users/arepstein/followers",
"following_url": "https://api.github.com/users/arepstein/following{/other_user}",
"gists_url": "https://api.github.com/users/arepstein/gists{/gist_id}",
"starred_url": "https://api.github.com/users/arepstein/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/arepstein/subscriptions",
"organizations_url": "https://api.github.com/users/arepstein/orgs",
"repos_url": "https://api.github.com/users/arepstein/repos",
"events_url": "https://api.github.com/users/arepstein/events{/privacy}",
"received_events_url": "https://api.github.com/users/arepstein/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 1427867026,
"node_id": "MDU6TGFiZWwxNDI3ODY3MDI2",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/enhancement",
"name": "enhancement",
"color": "c5e06d",
"default": true,
"description": "A new feature or improvement to an existing one"
},
{
"id": 5525608888,
"node_id": "LA_kwDOACgets8AAAABSVoZuA",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/analysis",
"name": "analysis",
"color": "7E50AA",
"default": false,
"description": "Concerning pymatgen.analysis"
},
{
"id": 5653124555,
"node_id": "LA_kwDOACgets8AAAABUPPVyw",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/molecules",
"name": "molecules",
"color": "EF81DB",
"default": false,
"description": "Molecule stuff"
}
] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@arepstein is this PR ready for review to be merged?",
"I believe so. The only pending update in my mind is automated detection of the rotational symmetry number, but I think it's okay as an input parameter for now.",
"Hey @arepstein , I get to know this PR in today's subgroup meeting. I am just wondering how the functionalities here comparing to those in the Goodvibes package https://github.com/bobbypaton/GoodVibes?",
"> Hey @arepstein , I get to know this PR in today's subgroup meeting. I am just wondering how the functionalities here comparing to those in the Goodvibes package https://github.com/bobbypaton/GoodVibes?\r\n\r\nThanks for pointing this out, I didn't know about Goodvibes when I put in this PR. This PR should be identical to the Grimme Quasi-RRHO approximation for the entropy that's implemented in GoodVibes, but does not include any other methods that GoodVibes implements. One big difference I see in terms of implementation into pymatgen infrastructure is that this PR can calculate Quasi-RRHO entropeis for Q-Chem output files, Gaussian output files, or manual input parameters, which is useful for Atomate integration. It would be a good idea to check that this agrees with GoodVibes.\r\n",
"\n[](https://coveralls.io/builds/47139417)\n\nCoverage decreased (-0.7%) to 83.428% when pulling **d08a1ca5c4a2d5e3f29703767642b4bfa5fd7ef0 on arepstein:readytoPR** into **3376f27a9c5a4cac1590169f2212361ab6c94388 on materialsproject:master**.\n",
"See also some small edits I made when I used the code, in a PR against your fork: https://github.com/arepstein/pymatgen/pull/1",
"I will also note for other reviewers that although the GoodVibes package contains similar functionality, it _only_ accepts Gaussian output files, and hence is not useful with our high-throughput Q-Chem infrastructure. So this PR is valuable because it gives us those capabilities.",
"@arepstein Is this still WIP?",
"@janosh Yes, still WIP, I haven't incorporated Ryan's comments yet. Thanks @rkingsbury for the comments!",
"> @janosh Yes, still WIP, I haven't incorporated Ryan's comments yet. Thanks @rkingsbury for the comments!\r\n\r\nNo problem! Recall that I also opened a [small PR](https://github.com/arepstein/pymatgen/pull/1/files) against your branch with some other edits, notably exposing `h_corrected` as an attribute.\r\n\r\nI had another thought about the call signature which I think would be better than what I initially proposed in that PR. Currently you have\r\n\r\n```\r\ndef __init__(self, output: GaussianOutput | QCOutput | dict, sigma_r=1, temp=298.15, press=101317, conc=1, v0=100):\r\n```\r\n\r\nI think it would be cleaner to make the regular `__init__` method generic by using explicit kwargs, e.g.\r\n```\r\ndef __init__(self, mol: Molecule, frequencies: List, energy: float, sigma_r=1, temp=298.15, press=101317, conc=1, v0=100):\r\n```\r\nWhere `mol`, `frequencies`, and `energy` are the items currently extracted from the `GaussianOutput` / `QCOutput` / `dict`. Then, you would add two `@classmethod`: `.from_gaussian(self, output: GaussianOutput)` and `.from_qchem(self, output: QCOutput)` to allow instantiating from the respective classes.\r\n\r\nThis structure allows for 1) use of the module with output from arbitrary codes (i.e., any molecule and frequency combo) and 2) leaves the door open for additional `.from_xxxx` methods to cleanly expand support to new codes in the future. You could even add some error handling that ensure the number of `frequencies` is consistent with the size of the `molecule`",
"Funny timing - I now would very much like to use this capability! One thing I want to note is that it would be very nice if, in addition to an actual Q-Chem output file, we could pass it a summary document that comes out of the new Molecules Emmet infrastructure. Happy to provide an example JSON.",
"> \r\nHey @samblau , I'm getting to this now. Could you please provide an example JSON? Right now it can take a manual dictionary input, so it should be easy to incorporate.\r\n",
"Hey @rkingsbury and @janosh, I've incorporated Ryan's edits and recommendations. While making these edits it became clear that a Class might not be the best choice for QuasiRRHO. As it stands now, it might be better as a simple function. Ideally, I think it could be nice to have a `Thermochemistry` class that is a `Molecule` plus some extra information like frequencies. We could then have different treatments of thermochemistry, like GoodVibes implements, but in a way works well with our ecosystem. \r\n\r\nThat being said, hopefully QuasiRRHO is still useful for now and can be a starting point for later changes if desired.",
"## [Codecov](https://app.codecov.io/gh/materialsproject/pymatgen/pull/2028?src=pr&el=h1&utm_medium=referral&utm_source=github&utm_content=comment&utm_campaign=pr+comments&utm_term=materialsproject) Report\n> :exclamation: No coverage uploaded for pull request base (`master@57d8a2f`). [Click here to learn what that means](https://docs.codecov.io/docs/error-reference?utm_medium=referral&utm_source=github&utm_content=comment&utm_campaign=pr+comments&utm_term=materialsproject#section-missing-base-commit).\n> Patch has no changes to coverable lines.\n\n<details><summary>Additional details and impacted files</summary>\n\n\n```diff\n@@ Coverage Diff @@\n## master #2028 +/- ##\n=========================================\n Coverage ? 74.06% \n=========================================\n Files ? 230 \n Lines ? 69403 \n Branches ? 16161 \n=========================================\n Hits ? 51403 \n Misses ? 14957 \n Partials ? 3043 \n```\n\n\n\n\n</details>\n\n[:umbrella: View full report in Codecov by Sentry](https://app.codecov.io/gh/materialsproject/pymatgen/pull/2028?src=pr&el=continue&utm_medium=referral&utm_source=github&utm_content=comment&utm_campaign=pr+comments&utm_term=materialsproject). \n:loudspeaker: Have feedback on the report? [Share it here](https://about.codecov.io/codecov-pr-comment-feedback/?utm_medium=referral&utm_source=github&utm_content=comment&utm_campaign=pr+comments&utm_term=materialsproject).\n",
"I think this is ready to merge, right @arepstein ? @janosh can you remove the 'stale' label?",
"Yes, I believe ready to merge!",
"Thanks @janosh and @rkingsbury for all the help, edits, and revisions!"
] | 2021-01-07T00:19:44
| 2023-08-22T14:58:05
|
2023-08-21T22:02:27Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Added quasirrho.py to the analysis subpackage to calculate the Quasi-RRHO free energy from a Gaussian or QChem frequency calculation.
* Calculates Grimme's Quasi-RRHO free energy
* Option for also correcting for solvent concentration
## Additional dependencies introduced (if any)
*None
## TODO (if any)
* The rotational symmetry number is required as an input. To calculate this on the fly, will consider updates to PointGroupAnalyzer
Before a pull request can be merged, the following items must be checked:
- [ x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x ] Tests have been added for any new functionality or bug fixes.
- [ x] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2028/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2028/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2028",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2028",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2028.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2028.patch",
"merged_at": "2023-08-21T22:02:26Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2029
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2029/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2029/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2029/events
|
https://github.com/materialsproject/pymatgen/pull/2029
| 782,165,995
|
MDExOlB1bGxSZXF1ZXN0NTUxNzcxMTk5
| 2,029
|
fix: typo in get_cbm docstring
|
{
"login": "kjappelbaum",
"id": 32935233,
"node_id": "MDQ6VXNlcjMyOTM1MjMz",
"avatar_url": "https://avatars.githubusercontent.com/u/32935233?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/kjappelbaum",
"html_url": "https://github.com/kjappelbaum",
"followers_url": "https://api.github.com/users/kjappelbaum/followers",
"following_url": "https://api.github.com/users/kjappelbaum/following{/other_user}",
"gists_url": "https://api.github.com/users/kjappelbaum/gists{/gist_id}",
"starred_url": "https://api.github.com/users/kjappelbaum/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/kjappelbaum/subscriptions",
"organizations_url": "https://api.github.com/users/kjappelbaum/orgs",
"repos_url": "https://api.github.com/users/kjappelbaum/repos",
"events_url": "https://api.github.com/users/kjappelbaum/events{/privacy}",
"received_events_url": "https://api.github.com/users/kjappelbaum/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @kjappelbaum, thanks for this! Even for small fixes or edits, please do fill out our [developer attribution form](https://forms.gle/JnisFb38QDR8QTFTA) so we can [acknowledge you appropriately on our website](https://pymatgen.org/team.html), improvements to pymatgen are always appreciated."
] | 2021-01-08T14:42:16
| 2021-01-11T20:04:24
|
2021-01-08T15:44:48Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Just a tiny typo fix.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2029/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2029/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2029",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2029",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2029.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2029.patch",
"merged_at": "2021-01-08T15:44:48Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2030
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2030/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2030/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2030/events
|
https://github.com/materialsproject/pymatgen/issues/2030
| 782,380,061
|
MDU6SXNzdWU3ODIzODAwNjE=
| 2,030
|
Test coverage for cmd line stuff
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I can deal with `mcsqs` and `critic2` since I am familiar with both and compilations are straight-forward. I don't know who's responsible for the `aconvasp` and `zeo` interfaces?",
"Sure, we can start with those 2. The other two are not that important for now.",
"Rather than committing compiled versions in the repository, I think we might instead have a custom Dockerfile with the utilities pre-installed, and launch from this Dockerfile in the test workflows. This is supported by GitHub Actions. Thoughts?",
"Sounds good to me.\r\n"
] | 2021-01-08T20:33:49
| 2023-08-13T16:34:58
|
2023-08-13T16:34:58Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Right now, the unit tests are not being run for a number of those requiring certain cmd line utilities. (see recent tests).
1. mcsqs
2. critic2
3. aconvasp
4. zeo
I would like to address at least a number of those, esp 1-3. We do not need executables for all platforms, but at the minimum, the Ubuntu tests should aim for 100% coverage (I have managed to push it up quite significantly). @mkhorton can you make sure at least 1-3 are addressed and we will think of adding zeo? All we need is to have compiled and executable versions in the pymatgen/cmd_line directory under appropriate structures.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2030/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2030/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2031
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2031/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2031/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2031/events
|
https://github.com/materialsproject/pymatgen/pull/2031
| 784,762,507
|
MDExOlB1bGxSZXF1ZXN0NTUzODk5NzIx
| 2,031
|
Optimizing pymatgen start-up time
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This should speed up import by 2x to 548,074 µs.\r\n\r\nRemoving all top-level imports from pymatgen would be the more forward-looking approach, and would maybe add an extra 10x speed-up. This would break backwards compatibility for anyone doing a root import (e.g. `from pymatgen import Structure` would have to become `from pymatgen.core.structure import Structure`). To leave the door open for this, I've made sure all tests etc. are using the canonical import.\r\n\r\nI tried lazy importing these top level imports, but the user experience wasn't there."
] | 2021-01-13T03:47:10
| 2021-02-13T17:29:22
|
2021-01-28T22:15:21Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This PR will be to explore strategies for optimizing pymatgen's start-up and import times.
Current root-level import is ~ 1 second (specifically, 1,100,534 µs as profiled by `python -X importtime`). Reducing this will also have consequences for speed of test suite.
Initial investigations suggest this could be sped up by at least 2x.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2031/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2031/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2031",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2031",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2031.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2031.patch",
"merged_at": "2021-01-28T22:15:21Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2032
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2032/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2032/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2032/events
|
https://github.com/materialsproject/pymatgen/pull/2032
| 786,394,494
|
MDExOlB1bGxSZXF1ZXN0NTU1MjcxNTQ3
| 2,032
|
Adding Q-Chem cube file plotting and small tweaks to the output parser
|
{
"login": "samblau",
"id": 7431223,
"node_id": "MDQ6VXNlcjc0MzEyMjM=",
"avatar_url": "https://avatars.githubusercontent.com/u/7431223?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/samblau",
"html_url": "https://github.com/samblau",
"followers_url": "https://api.github.com/users/samblau/followers",
"following_url": "https://api.github.com/users/samblau/following{/other_user}",
"gists_url": "https://api.github.com/users/samblau/gists{/gist_id}",
"starred_url": "https://api.github.com/users/samblau/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/samblau/subscriptions",
"organizations_url": "https://api.github.com/users/samblau/orgs",
"repos_url": "https://api.github.com/users/samblau/repos",
"events_url": "https://api.github.com/users/samblau/events{/privacy}",
"received_events_url": "https://api.github.com/users/samblau/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @samblau !"
] | 2021-01-14T22:43:44
| 2021-01-16T00:33:18
|
2021-01-16T00:33:14Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Adding Q-Chem cube file plotting and small tweaks to the output parser
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2032/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2032/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2032",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2032",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2032.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2032.patch",
"merged_at": "2021-01-16T00:33:14Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2033
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2033/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2033/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2033/events
|
https://github.com/materialsproject/pymatgen/pull/2033
| 786,460,565
|
MDExOlB1bGxSZXF1ZXN0NTU1MzI2NjE4
| 2,033
|
Parse OUTCAR core potentials in fixed width format
|
{
"login": "utf",
"id": 1330638,
"node_id": "MDQ6VXNlcjEzMzA2Mzg=",
"avatar_url": "https://avatars.githubusercontent.com/u/1330638?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/utf",
"html_url": "https://github.com/utf",
"followers_url": "https://api.github.com/users/utf/followers",
"following_url": "https://api.github.com/users/utf/following{/other_user}",
"gists_url": "https://api.github.com/users/utf/gists{/gist_id}",
"starred_url": "https://api.github.com/users/utf/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/utf/subscriptions",
"organizations_url": "https://api.github.com/users/utf/orgs",
"repos_url": "https://api.github.com/users/utf/repos",
"events_url": "https://api.github.com/users/utf/events{/privacy}",
"received_events_url": "https://api.github.com/users/utf/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks."
] | 2021-01-15T01:27:05
| 2021-01-15T04:16:29
|
2021-01-15T04:16:23Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
The electrostatic core potentials in the OUTCAR file are written based on a fix width format. The potentials are numbered from 1 to natoms, and sometimes the fixed width means there is no space between the index and the potential. For example:
```
average (electrostatic) potential at core
the test charge radii are 1.0894 0.9698
(the norm of the test charge is 1.0000)
1-101.5055 2-101.5055 3-101.5055 4-101.5055 5 -85.7708
6 -85.7708 7 -85.7708 8 -85.7708
```
This PR changes the parsing of the potentials to be based on a fix width format (rather than `str.split()`). I've added a new test.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2033/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2033/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2033",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2033",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2033.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2033.patch",
"merged_at": "2021-01-15T04:16:23Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2034
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2034/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2034/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2034/events
|
https://github.com/materialsproject/pymatgen/pull/2034
| 786,756,678
|
MDExOlB1bGxSZXF1ZXN0NTU1NTgzMzgw
| 2,034
|
Wavefunctions from Lobster
|
{
"login": "JaGeo",
"id": 22094846,
"node_id": "MDQ6VXNlcjIyMDk0ODQ2",
"avatar_url": "https://avatars.githubusercontent.com/u/22094846?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/JaGeo",
"html_url": "https://github.com/JaGeo",
"followers_url": "https://api.github.com/users/JaGeo/followers",
"following_url": "https://api.github.com/users/JaGeo/following{/other_user}",
"gists_url": "https://api.github.com/users/JaGeo/gists{/gist_id}",
"starred_url": "https://api.github.com/users/JaGeo/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/JaGeo/subscriptions",
"organizations_url": "https://api.github.com/users/JaGeo/orgs",
"repos_url": "https://api.github.com/users/JaGeo/repos",
"events_url": "https://api.github.com/users/JaGeo/events{/privacy}",
"received_events_url": "https://api.github.com/users/JaGeo/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"From my side, this code would now be ready to merge. Let me know, if there is anything else to do.",
"Thanks @JaGeo, this looks great!\r\n\r\nI did have a question on the `set_volumetric_data`, wasn't sure what the `runner` variable and the following block was doing.",
"I need to filter out some values from the Lobster output (at x=1.0, y=1.0 or z=1.0). These values are written out twice (also at x=0.0, y=0.0 and z=0.0) if I use a box output of the wavefunction between the fractional coordinates 0.00 0.00 0.00 and 1.00 1.00 1.00. As far as I understood the volumeticdata object, they should not be included. \r\n\r\nI hope my explanation is understandable.",
"Yes, that makes sense, thanks. And yes, VolumetricData wouldn't want values for both."
] | 2021-01-15T09:47:21
| 2021-01-19T21:16:06
|
2021-01-16T00:29:31Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This is a implementation of the new class Wavefunction for the Lobster module that can read in Lobster wavefunctions and transfer them to a VolumetricData object (This allows plotting with VESTA and compability with other VASP files). It can also calculate densities based on the Lobster output.
Thanks to @rnels12 for helpful comments on the Lobster implementation.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2034/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2034/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2034",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2034",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2034.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2034.patch",
"merged_at": "2021-01-16T00:29:31Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2035
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2035/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2035/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2035/events
|
https://github.com/materialsproject/pymatgen/pull/2035
| 788,239,263
|
MDExOlB1bGxSZXF1ZXN0NTU2Nzg0MzI0
| 2,035
|
Fixed coordination_geometry_finder.
|
{
"login": "davidwaroquiers",
"id": 3625296,
"node_id": "MDQ6VXNlcjM2MjUyOTY=",
"avatar_url": "https://avatars.githubusercontent.com/u/3625296?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/davidwaroquiers",
"html_url": "https://github.com/davidwaroquiers",
"followers_url": "https://api.github.com/users/davidwaroquiers/followers",
"following_url": "https://api.github.com/users/davidwaroquiers/following{/other_user}",
"gists_url": "https://api.github.com/users/davidwaroquiers/gists{/gist_id}",
"starred_url": "https://api.github.com/users/davidwaroquiers/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/davidwaroquiers/subscriptions",
"organizations_url": "https://api.github.com/users/davidwaroquiers/orgs",
"repos_url": "https://api.github.com/users/davidwaroquiers/repos",
"events_url": "https://api.github.com/users/davidwaroquiers/events{/privacy}",
"received_events_url": "https://api.github.com/users/davidwaroquiers/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hello,\r\nis there anything left to do for this pull request? It would help me with another implementation if it could be merged? Thanks.\r\n\r\nBest, \r\nJG\r\n\r\n",
"Pls fix all missing documentation. I already put #TODO in all the relevant places in the chemenv package. We would like to make sure that all pymatgen is fully tested and documented.",
"Thanks."
] | 2021-01-18T12:43:23
| 2021-02-05T15:39:13
|
2021-02-05T15:39:12Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Fixed small bug in coordination_geometry_finder
* Fixed mutable argument in method for separation plane algorithm
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2035/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2035/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2035",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2035",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2035.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2035.patch",
"merged_at": "2021-02-05T15:39:12Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2036
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2036/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2036/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2036/events
|
https://github.com/materialsproject/pymatgen/issues/2036
| 788,984,013
|
MDU6SXNzdWU3ODg5ODQwMTM=
| 2,036
|
Issue in `Structure.get_neighbor_list()` returning sites as their own neighbours
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"tolerence is already in pymatgen"
] | 2021-01-19T12:31:12
| 2021-01-19T14:45:25
|
2021-01-19T13:50:49Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
According to the OCP codebase `Structure.get_neighbor_list()` can sometimes return self neighbours in some edge cases.
https://github.com/Open-Catalyst-Project/ocp/blob/899de1c1f46f44bf99e3ccd190b9a37795b60b67/ocpmodels/preprocessing/atoms_to_graphs.py#L99-L114
They resolve this by setting a minimum distance between neighbours of 1e-8. Flagging as there could potentially be some benefit to including this tolerance in `pymatgen` directly.
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2036/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2036/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2037
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2037/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2037/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2037/events
|
https://github.com/materialsproject/pymatgen/pull/2037
| 789,492,578
|
MDExOlB1bGxSZXF1ZXN0NTU3ODMwMzk2
| 2,037
|
HighSymmKpath Hinuma setting bug fix
|
{
"login": "munrojm",
"id": 6877334,
"node_id": "MDQ6VXNlcjY4NzczMzQ=",
"avatar_url": "https://avatars.githubusercontent.com/u/6877334?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/munrojm",
"html_url": "https://github.com/munrojm",
"followers_url": "https://api.github.com/users/munrojm/followers",
"following_url": "https://api.github.com/users/munrojm/following{/other_user}",
"gists_url": "https://api.github.com/users/munrojm/gists{/gist_id}",
"starred_url": "https://api.github.com/users/munrojm/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/munrojm/subscriptions",
"organizations_url": "https://api.github.com/users/munrojm/orgs",
"repos_url": "https://api.github.com/users/munrojm/repos",
"events_url": "https://api.github.com/users/munrojm/events{/privacy}",
"received_events_url": "https://api.github.com/users/munrojm/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @munrojm "
] | 2021-01-20T00:08:38
| 2021-01-21T20:00:25
|
2021-01-21T20:00:23Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This PR contains a small bug fix for the `HighSymmKpath` class. Currently, the k-path from this class produced by the Hinuma et al. convention is not properly transformed into the basis of the reciprocal lattice of the input structure in all cases. This has been fixed, and a warning has been added to tell the user to use `KPathSeek` if they would like the path in the original author-intended basis.
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2037/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2037/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2037",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2037",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2037.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2037.patch",
"merged_at": "2021-01-21T20:00:23Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2038
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2038/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2038/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2038/events
|
https://github.com/materialsproject/pymatgen/issues/2038
| 790,363,805
|
MDU6SXNzdWU3OTAzNjM4MDU=
| 2,038
|
Composition shows weird things when reading in unknown element
|
{
"login": "TimoSommer",
"id": 67800320,
"node_id": "MDQ6VXNlcjY3ODAwMzIw",
"avatar_url": "https://avatars.githubusercontent.com/u/67800320?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/TimoSommer",
"html_url": "https://github.com/TimoSommer",
"followers_url": "https://api.github.com/users/TimoSommer/followers",
"following_url": "https://api.github.com/users/TimoSommer/following{/other_user}",
"gists_url": "https://api.github.com/users/TimoSommer/gists{/gist_id}",
"starred_url": "https://api.github.com/users/TimoSommer/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/TimoSommer/subscriptions",
"organizations_url": "https://api.github.com/users/TimoSommer/orgs",
"repos_url": "https://api.github.com/users/TimoSommer/repos",
"events_url": "https://api.github.com/users/TimoSommer/events{/privacy}",
"received_events_url": "https://api.github.com/users/TimoSommer/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This is a feature, not a bug. It is common practice to use symbols to represent abstract species, e.g., vacancies or just some group of elements. ",
"For more context on this, read the documentation on `DummySpecies`. It is possible to initialize a DummySpecies without oxidation state, e.g. `DummySpecies('M', oxidation_state=None)` but this has to be done manually. When initializing a `Composition` from a string, the string is parsed and a best guess is taken (historically, a 0 oxidation state is assigned because Species used to require an oxidation state, but this is now optional -- the old behavior was retained so as not to break backwards compatibility).",
"Alright, so this is historic behaviour. Thank you very much for clearing this up!"
] | 2021-01-20T21:58:51
| 2021-01-22T07:35:21
|
2021-01-20T22:01:02Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
When I make a new Composition from a string that contains a chemical formula with non valid (but still often e.g. as placeholder used symbols) like "M", this pseudo element gets represented with the string "M0+" which is not the expected behaviour.
**To Reproduce**
Steps to reproduce the behavior:
import pymatgen
string = M2 # Notice M is no valid chemical element
comp = pymatgen.Composition(string)
If one inspects comp then you see that the element "M" somehow is represented as "M0+". You can for example easily see this by getting the dictionary
comp_dict = comp.as_dict() # comp_dict = defaultdict(<class 'float'>, {'M0+': 2.0})
**Expected behavior**
I would expect that though this element is not valid, it still shows up as "M" and not as "M0+". For example, when using the method Composition.as_dict() there should be a key "M" and not a key "M0+".
**Desktop (please complete the following information):**
- OS: Linux Mint 19.1 Cinnamon
- Version: 2020.12.31
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2038/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2038/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2039
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2039/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2039/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2039/events
|
https://github.com/materialsproject/pymatgen/pull/2039
| 790,855,410
|
MDExOlB1bGxSZXF1ZXN0NTU5MDAyMjQw
| 2,039
|
[WIP] add kwarg to pass in a plt ax to plot_brillouin
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I noticed that `get_ax3d_fig_plt` still prevents Brillouin zones from being plotted into a subplot because matplotlib appears unable to find the current figure and instead creates a new one:\r\n\r\n```py\r\ndef get_ax3d_fig_plt(ax=None, **kwargs):\r\n \"\"\"\r\n Helper function used in plot functions supporting an optional Axes3D\r\n argument. If ax is None, we build the `matplotlib` figure and create the\r\n Axes3D else we return the current active figure.\r\n\r\n Args:\r\n kwargs: keyword arguments are passed to plt.figure if ax is not None.\r\n\r\n Returns:\r\n ax: :class:`Axes` object\r\n figure: matplotlib figure\r\n plt: matplotlib pyplot module.\r\n \"\"\"\r\n import matplotlib.pyplot as plt\r\n from mpl_toolkits.mplot3d import axes3d\r\n if ax is None:\r\n fig = plt.figure(**kwargs)\r\n ax = axes3d.Axes3D(fig)\r\n else:\r\n fig = plt.gcf()\r\n # preventing the call to plt.gcf() avoids creating new figures\r\n # fig = None\r\n```\r\n\r\n```py\r\ndef gcf():\r\n \"\"\"\r\n Get the current figure.\r\n\r\n If no current figure exists, a new one is created using\r\n `~.pyplot.figure()`.\r\n \"\"\"\r\n figManager = _pylab_helpers.Gcf.get_active()\r\n if figManager is not None:\r\n return figManager.canvas.figure\r\n else:\r\n return figure()\r\n```\r\n\r\nCurrently: producing lots of empty figures\r\n\r\n\r\n\r\n\r\nGoal: subplots\r\n\r\n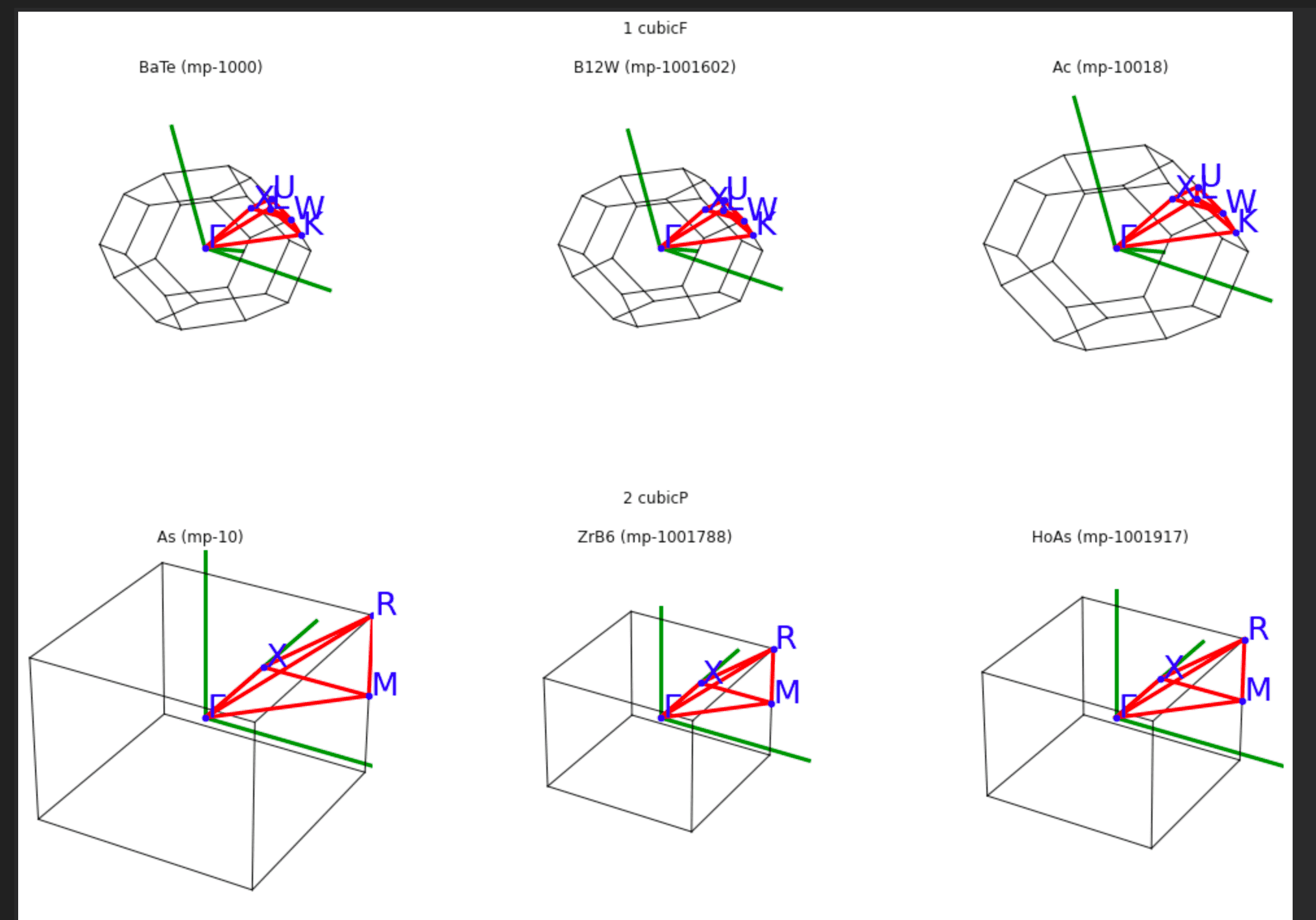\r\n\r\nIf the user passes in an `plt.axis` instance, presumably, they don't need `BSPlotter.plot_brillouin()` to return the current `fig`. Would it be alright if we get rid of the call to `get_ax3d_fig_plt` altogether if the user provided their own axis?\r\n\r\n@mkhorton Pinging you as this is related to https://github.com/materialsproject/api/issues/181#issuecomment-763514303.",
"@shyuep What's your take on this?",
"It isn't clear to me why `gcf` is creating a new figure. Is `plt.show()` being called before the subplots are added?",
"> It isn't clear to me why gcf is creating a new figure.\r\n\r\n@utf That confuses me as well.\r\n\r\n> Is plt.show() being called before the subplots are added?\r\n\r\nI looked for that as well. Not in my code and not (as far as I can tell) by `pymatgen` either. `BSPlotter` _does_ have a `show` method (with an unused argument `smooth_tol` btw):\r\n\r\n```py\r\n def show(self, zero_to_efermi=True, ylim=None, smooth=False, smooth_tol=None):\r\n \"\"\"\r\n Show the plot using matplotlib.\r\n\r\n Args:\r\n zero_to_efermi: Automatically subtract off the Fermi energy from\r\n the eigenvalues and plot (E-Ef).\r\n ylim: Specify the y-axis (energy) limits; by default None let\r\n the code choose. It is vbm-4 and cbm+4 if insulator\r\n efermi-10 and efermi+10 if metal\r\n smooth: interpolates the bands by a spline cubic\r\n smooth_tol (float) : tolerance for fitting spline to band data.\r\n Default is None such that no tolerance will be used.\r\n \"\"\"\r\n plt = self.get_plot(zero_to_efermi, ylim, smooth)\r\n plt.show()\r\n```\r\n\r\nbut that's not being called by any of the other methods.",
"Hello @janosh,\r\njust to understand the problem you're point out.\r\nI managed to create subplots of brillouin zones with not change in the code already there.\r\nHere my simple approach:\r\nfrom pymatgen.electronic_structure.plotter import plot_brillouin_zone_from_kpath, plot_brillouin\r\nkpath = HighSymmKpath(structure)\r\nplt.subplot(121,projection='3d')\r\nplot_brillouin_zone_from_kpath(kpath,plt.gca())\r\nplt.subplot(122,projection='3d')\r\nplot_brillouin_zone_from_kpath(kpath,plt.gca())\r\n\r\nhere the result:\r\n\r\n",
"@fraricci You're not using the code I'm suggesting to modify. I'm aware that `plot_brillouin_zone_from_kpath` already has an `ax` kwarg. But the most user-facing method `BSPlotter.plot_brillouin()` doesn't, hence my PR."
] | 2021-01-21T09:18:37
| 2021-03-09T16:22:52
|
2021-03-09T16:22:52Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Adds the ability to plot Brillouin zones into a `plt.subplot`
## TODO (if any)
- Does this need a test?
## Checklist
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [ ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2039/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2039/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2039",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2039",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2039.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2039.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2040
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2040/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2040/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2040/events
|
https://github.com/materialsproject/pymatgen/pull/2040
| 792,849,527
|
MDExOlB1bGxSZXF1ZXN0NTYwNjQyMTQ3
| 2,040
|
fix: #1985 and add test to stop computed entry being used for get_quasi_e_to_hull
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @CompRhys ! I appreciate you doing this and I know that #2014 has become more involved than originally expected. This fixes #1985 at least insofar as it affects my use cases and looks like a very reasonable stopgap solution. @mkhorton , do you have any concerns about merging this as a temporary fix?",
"@rkingsbury I plan to merge both asap, stay tuned... I think #2014 is in good shape, though it did indeed end up longer than originally intended :-)",
"Given that #2014 is merged, shall we close this?",
"OK with me @mkhorton !",
"not needed due to #2014!"
] | 2021-01-24T17:29:07
| 2021-01-27T09:32:02
|
2021-01-27T09:00:31Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
@rkingsbury
As the proper fix in #2014 seems like it might still take a while to get merged this is the temp fix I should have done for #1985 when I first saw it. It avoids the bug that I introduced by not (at that point) being aware that `ComputedEntry` was supposed to be interoperable with `PDEntry` and adds a catch to stop another bug in `get_quasi_e_to_hull` that would be caused by the same issues with `__eq__` and `__hash__` for `ComputedEntry` that caused this bug.
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2040/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2040/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2040",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2040",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2040.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2040.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2041
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2041/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2041/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2041/events
|
https://github.com/materialsproject/pymatgen/pull/2041
| 793,457,302
|
MDExOlB1bGxSZXF1ZXN0NTYxMTQyMTE0
| 2,041
|
add documentation for QChemDictSet
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@samblau @espottesmith would love to get your feedback on this. As I started using the QChem workflows I found myself having to do a lot of code diving and searching the QChem manual to understand how to set things properly. That's what motivated me to do this; hoping to provide some clarification for new users about what typical settings are.",
"@samblau @espottesmith friendly reminder - any comments on this revised docstring? ",
"My apologies for the delay, Ryan. Hoping to get to this later today.",
"Hi @rkingsbury, thanks for your patience. These are very helpful additions. A few suggested tweaks:\r\n\r\n- I haven't checked recently, but not that long ago the new GenSCFman module was limited in what SCF algorithms it supported. It's muuuuuuch better than the old SCF code across the board, so I'd say it's basically always better to use it even if it has limitations. Thus, I'd recommend changing some of the documentation for scf_algorithm to note that DIIS, DIIS_GDM, and GDM are definitely supported by GenSCFman, and that if the user wants to use a different SCF algorithm, they should consult the manual to see if it is supported by GenSCFman. If it isn't, they might have to set genscfman to False in rem - I'm not sure what is default at this point or if requesting an SCF alg that is only in the old module will automatically force the switch. \r\n- I would perhaps not use tetrahydrofuran as an example SMD solvent input. There is this super awkward thing (which I hope has been fixed, but I'm not sure) where you actually have to request \"thf\" instead of \"tetrahydrofuran\" in the QChem input file. There are two stupid lines of code in QCInputs to handle this, but yeah, just maybe not the best example to avoid any potential confusion.\r\n- Beyond rem, pcm, and solvent, overwrite_inputs will also work with smx and plots as per the lines following \"if self.overwrite_inputs:\".\r\n\r\nThanks for these additions!",
"@samblau let me know what you think of my latest edits per your comments. FYI, this is what the current Qchem manual recommends re: SCF_ALGORITHM. I could not find an explicit list of algorithms that are supported by GEN_SCFMAN, but it seems like almost all (except perhaps RCA) are at this point. RCA_DIIS and DIIS_GDM *might* use the old SCF algorithm as well but it isn't clear.\r\n\r\n> Use DIIS unless performing a restricted open-shell calculation, in which case GDM is rec-ommended. If DIIS fails to find a reasonable approximate solution in the initial iterations,RCA_DIIS is the recommended fallback option. If DIIS approaches the correct solution but failsto finally converge, DIIS_GDM is the recommended fallback. For systems with small HOMO-LUMO gaps and DIIS fails to converge, LS_DIIS could help",
"@rkingsbury The SCF docstring looks good now. I'm 100% confident that DIIS_GDM uses the new algorithm. I'm hoping that RCA_DIIS either already does or will soon. At some point I'll need to go back and check because if RCA_DIIS is now compatible with GenSCFman then it almost certainly has a place in our SCF error handling procedure. All the other minor stuff looks good as well. Thanks!",
"@shyuep This is ready for review / merge at your convenience. For some reason the flake8 linter is failing with\r\n\r\n```\r\nBLK100 Black would make changes\r\n```\r\neven though I have run `black` on the file in question. If you want me to take any further action on the formatting, please let me know.",
"@samblau I just realized that changing `pcm_dielectric` from string to int is causing a test failure. I can fix it, but want to confirm you agree that we should change the type of this variable first.",
"\n[](https://coveralls.io/builds/37290915)\n\nCoverage decreased (-0.7%) to 79.239% when pulling **ebef17dfa5485d7addd3b220d4fb4f4cc7863d99 on rkingsbury:qchem** into **19d7f56ba07850ce7cef743cc60272057a289c43 on materialsproject:master**.\n",
"@shyuep This is ready for review / merge at your convenience. All tests are passing, except that for some reason the flake8 linter is failing with\r\n\r\nBLK100 Black would make changes\r\n\r\neven though I have run black on the file in question. If you want me to take any further action on the formatting, please let me know.",
"This usually means you are not running black from pymatgen root directory.",
"@shyuep Thanks for the guidance. I ran `black pymatgen/io/qchem` from the root directory, pushed the changes, but am still getting the same linting error. Any other ideas?",
"@shyuep following up your comment about `Black would make changes` warnings in #2006 , this PR is ready to merge. Thanks.",
"@rkingsbury For the record, when I ran black from my pymatgen root, I get\r\n\r\n```\r\nreformatted /Users/shyue/repos/pymatgen/pymatgen/io/qchem/utils.py\r\nreformatted /Users/shyue/repos/pymatgen/pymatgen/io/qchem/sets.py\r\nreformatted /Users/shyue/repos/pymatgen/pymatgen/io/qchem/inputs.py\r\nreformatted /Users/shyue/repos/pymatgen/pymatgen/io/qchem/tests/test_sets.py\r\nreformatted /Users/shyue/repos/pymatgen/pymatgen/io/tests/test_gaussian.py\r\nreformatted /Users/shyue/repos/pymatgen/pymatgen/io/qchem/outputs.py\r\nreformatted /Users/shyue/repos/pymatgen/pymatgen/io/vasp/tests/test_outputs.py\r\nAll done! ✨ 🍰 ✨\r\n7 files reformatted, 485 files left unchanged.\r\n```\r\n\r\nFurther, there were pylint errors.\r\n```\r\n************* Module pymatgen.io.qchem.outputs\r\npymatgen/io/qchem/outputs.py:700:15: R1726: Boolean condition 'parsed_geometry == [] or None' may be simplified to 'parsed_geometry == []' (simplifiable-condition)\r\npymatgen/io/qchem/outputs.py:721:11: R1726: Boolean condition 'parsed_optimized_geometries == [] or None' may be simplified to 'parsed_optimized_geometries == []' (simplifiable-condition)\r\n-----------------------------------\r\nYour code has been rated at 9.98/10\r\n```\r\n\r\nI really suggest you use the pre-commit hook included in pymatgen. That will avoid a lot of issues in future. I have no idea why black made changes in my case when it didn't in your case. But pylint errors should be dealt with regardless.\r\n",
"@rkingsbury See this commit for the actual changes made by black: https://github.com/materialsproject/pymatgen/commit/d67a637825647cb8ecae95f07021be37f2bd8d7c",
"Thanks @shyuep! I'll have to examine my local configuration and figure out what's going on. Can you tell me what version of black you're using? Somehow it seems that my local linter configurations have gone out of sync with upstream.",
"I am using black==20.8b1. It turns out the flake8-black was using an old version on the Github actions. I just added a pin to black==20.8b1. Hopefully that solves the issue on Github Actions being inconsistent with our local machines. I also rearranged the linters such that the other lints are done first and flake8 is the last one to run."
] | 2021-01-25T15:02:53
| 2021-02-24T15:40:06
|
2021-02-24T15:24:08Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Add documentation to `QChemDictSet` and potentially other QChem `InputSet`
TODO:
- [x] Clarify type of `pcm_dielectric`. The original docstring specified a `str` but I believe a `float` is more appropriate and intuitive
- [x] Copy docstring to child sets
- [x] complete documentation of `opt_variables` and `scan_variables`
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2041/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2041/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2041",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2041",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2041.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2041.patch",
"merged_at": "2021-02-24T15:24:08Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2042
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2042/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2042/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2042/events
|
https://github.com/materialsproject/pymatgen/pull/2042
| 795,123,898
|
MDExOlB1bGxSZXF1ZXN0NTYyNTI0NTQ3
| 2,042
|
[WIP] Add PatchedPhaseDiagram to accelerate handling of large approximately disjoint data sets
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"> @CompRhys I was just working on filtering large chemical spaces and tested out the PatchedPhaseDiagram class.\r\n> \r\n> I ran it on all entries in the \"Y-Mn-O-Li-Cl-C-Na-Fe-W-Bi-Ti-Be-S\" chemical system (as acquired from MP), which has 13 elements and about 3,595 entries. The PatchedPhaseDiagram code performed equivalently fast to the `expand_pd()` method I linked to a while back (about 7 seconds) on a single core, but actually took about 20x longer (~155 seconds) with multiprocessing enabled. I think that the current setup of doing a map to each chemical system and supplying `all_entries` as an argument each time is causing a great deal of overhead, as it tries to copy the list of all entries into the memory of each process.\r\n> \r\n> Wasn't sure if you already had planned on addressing this, but to help speed this up, you might try to do some batching so that there are only a few processes created (maybe total # processes = # cores), making sure only to enable multiprocessing when each process takes a substantial amount of compute time (maybe 10+ seconds?)... this system seemed to work well for me in similar multiprocessing applications :)\r\n\r\n@mattmcdermott I have changed the multiprocessing a bit now do you still encounter this slow-down? I made the `_get_pd_for_space` function into a method which means it shouldn't need to copy `all_entries` each time? However, I am not sure about that as I didn't manage to replicate the slow down with the older setup",
"\n[](https://coveralls.io/builds/39149998)\n\nCoverage decreased (-0.6%) to 83.024% when pulling **a938e9f65d82d154f47a775b847a97e67756395e on CompRhys:ppd** into **df0f9ba74f9b1ba54bb6ede88badb31b7cc1cdb4 on materialsproject:master**.\n",
"@mkhorton I've been using this code for a while now and I think this is probably now stable enough to think about reviewing/merging. I refactored get `phase_separation_energy` a bit on this branch and added a fix to an edge case I found today, I fixed it here as this is the code I have been using but can make a separate PR for the fix in prefered ",
"a small note is that I did some testing when swapping `tqdm` for the `pymatgen` `PBar` and am pretty sure that `PBarSafe` isn't safe or at least not how I was using `tqdm` so could be worth making `tqdm` a requirement?",
"Actually tried applying to whole of MP today and hit some issues speed issues that I am not sure can be fixed to make this generally useful, will close for the time being and maybe open again if I can speed it up",
"Could you clarify what the speed issues were, and what the unsafe PBar behavior was? I'm not against merging this if it's useful for intermediate cases, even if it can't handle very large data sets.\r\n\r\n",
"When I purposefully removed tqdm from my environment PBar should default for PBarSafe but PBarsafe threw an error. I didn't really explore further than that as PBar worked fine as a direct substitution for tqdm.\r\n\r\nI can get the `ppd` for the whole of MP in <40minutes using 20 workers if I first convert `ComputedStructureEntry`/`ComputuedEntry` to `PDEntry` to reduce memory and function calls (hence why I encountered #2120). I don't know how it compares to the equivalent stages in your database builds for getting the pds for the get e_above_hull calcs. The main bottlenecks are having to copy the object to different processes and then selecting the entries to build the relevant patches. Selecting entries scales badly to larger entry sets due to it being done in serial (calls to elements and frozenset is superset in profile are due to this step).\r\n\r\nThe other speed issues of concern are a mix-match quite a lot due to me not knowing what was slow in pymatgen and trying to write pretty code that kept walking into speed-traps. These effect calls to `get_e_above_hull` and `get_phase_separation_energy`. Mostly the problem was that I was using a lot of calls to fractional_composition which ends up being very slow and running checks to avoid computation that are slower than the computation being avoided. Tweaking this I think i've got a 4x speed-up on `get_phase_separation_energy` but still haven't worked out how to make use of multi-processing with the ppd to evaluate lots of these in parallel. The issue is that the object for the whole of MP ppd can be v large (esp if using ComputedStructureEntries) so don't necessarily want to make too many copies.\r\n\r\n[ppd-profile.txt](https://github.com/materialsproject/pymatgen/files/6400552/ppd-profile.txt)\r\n\r\n\r\n"
] | 2021-01-27T13:57:16
| 2021-04-30T15:21:47
|
2021-04-28T12:15:31Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This PR takes over from #1974.
Introduce a new `PatchedPhaseDiagram` based class to handle large disjoint data sets. Calculation of the convex hull of a data set scales with the dimensions and amount of data. Breaking up disjoint datasets into smaller patches reduces the complexity in both these regards and would allow for parallel computation of different patches.
The code has been adapted such that as much as possible can be inherited from `PhaseDiagram`. However, not all the functionality of `PhaseDiagram` will be available in `PatchedPhaseDiagram` as some of these functions can't be calculated without access to the facets. My primary aim is to be able to extract `get_decomp_and_quasi_e_to_hull` for large high dimensional phase diagrams in a timely manner to allow for more robust testing workflows of ML models extending on ideas in doi.org/10.1038/s41524-020-00362-y .
Where functionality is not able to be replicated I have attempted to modularise the code such that if approximate methods can be devised then only smaller modular methods need to be re-written for functionality to be inherited correctly. e.g. `_get_simplex_intersections` and `_get_all_facets_and_simplexes`
More of the code has been converted into google docstring format.
#### Speed
The code has been tested for speed using the following `ppd_test.py` script:
```python
import pymatgen
from pymatgen.analysis.phase_diagram import (
PhaseDiagram,
PatchedPhaseDiagram,
)
with pymatgen.MPRester("API_KEY") as m:
entries = m.get_entries_in_chemsys("Y-Mn-O-Li-Cl-C-Na-Fe-W-Bi-Ti-Be-S")
def foo():
pd = PatchedPhaseDiagram(entries, workers=0)
def bar():
pd = PatchedPhaseDiagram(entries, workers=1)
def flob():
pd = PatchedPhaseDiagram(entries, workers=3)
def fum():
pd = PatchedPhaseDiagram(entries, workers=-1) # i.e. 4 workers
def fifo():
pd = PhaseDiagram(entries)
```
The options were tested on a laptop with 4 threads
```
% python -mtimeit -s'import ppd_test as test' 'test.foo()'
1 loop, best of 5: 14.6 sec per loop
```
```
% python -mtimeit -s'import ppd_test as test' 'test.bar()'
1 loop, best of 5: 15.3 sec per loop
```
```
% python -mtimeit -s'import ppd_test as test' 'test.flob()'
1 loop, best of 5: 10 sec per loop
```
```
% python -mtimeit -s'import ppd_test as test' 'test.fum()'
1 loop, best of 5: 10.2 sec per loop
```
I gave up testing `fifo` as it took too long.
On a dataset of 56788 entries drawn from MP it took ~1hr40m to construct a ppd on my laptop. CPU: Intel i5-5200U (4) @ 2.700GHz
With 20 cores on the cds3 cluster it takes ~5m. CPU: Intel(R) Xeon(R) Gold 6142 CPU @ 2.60GHz
## Checklist
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
- [x] Tests have been added for any new functionality or bug fixes.
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2042/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2042/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2042",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2042",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2042.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2042.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2043
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2043/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2043/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2043/events
|
https://github.com/materialsproject/pymatgen/pull/2043
| 795,156,952
|
MDExOlB1bGxSZXF1ZXN0NTYyNTUyMzQ1
| 2,043
|
Read Eigenval for Vasp non spin polarized Calculations
|
{
"login": "mdforti",
"id": 30444792,
"node_id": "MDQ6VXNlcjMwNDQ0Nzky",
"avatar_url": "https://avatars.githubusercontent.com/u/30444792?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mdforti",
"html_url": "https://github.com/mdforti",
"followers_url": "https://api.github.com/users/mdforti/followers",
"following_url": "https://api.github.com/users/mdforti/following{/other_user}",
"gists_url": "https://api.github.com/users/mdforti/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mdforti/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mdforti/subscriptions",
"organizations_url": "https://api.github.com/users/mdforti/orgs",
"repos_url": "https://api.github.com/users/mdforti/repos",
"events_url": "https://api.github.com/users/mdforti/events{/privacy}",
"received_events_url": "https://api.github.com/users/mdforti/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 2414169365,
"node_id": "MDU6TGFiZWwyNDE0MTY5MzY1",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/needs%20testing",
"name": "needs testing",
"color": "66ddab",
"default": false,
"description": "PRs that are not ready to merge due to lacking tests"
},
{
"id": 3093102097,
"node_id": "MDU6TGFiZWwzMDkzMTAyMDk3",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/stale",
"name": "stale",
"color": "A99B62",
"default": false,
"description": "Abandoned or conflicting PRs and outdated issues"
}
] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @mdforti! Do you have an example non-spin-polarized EIGENVAL file to hand too? We can include it as a test case",
"(Note: I've edited my comment now that I understand what is going on.)\r\n\r\nRegarding this PR and #2021 by @mdforti, I believe VASP might have changed how the EIGENVAL file is formatted several years ago. At least in VASP 5.4.x, I can confirm that Pymatgen's EIGENVAL reader works appropriately for spin-unpolarized calculations. Such calculations have three columns in the EIGENVAL file, describing the index, energy, and occupation. You can find an example EIGENVAL file here ([EIGENVAL.txt](https://github.com/materialsproject/pymatgen/files/6730391/EIGENVAL.txt)). No issues occur upon parsing with the code below:\r\n\r\n```python\r\nfrom pymatgen.io.vasp.outputs import Eigenval\r\neig = Eigenval('EIGENVAL')\r\nprint(eig.eigenvalue_band_properties)\r\n````\r\n\r\nHowever, traversing the web leads to instances where it is clear the EIGENVAL file _can_ contain only two columns (in which case a fix like that proposed by @mdforti would be necessary). For instance, see the EIGENVAL file supplied on [this website](http://grandcentral.apam.columbia.edu:5555/tutorials/dft_procedures/bands/srvo3/vasp/index.html), attached here ([EIGENVAL_old.txt](https://github.com/materialsproject/pymatgen/files/6730436/EIGENVAL_old.txt)). I used their VASP input files to generate my EIGENVAL and despite that, I have a three-column EIGENVAL file whereas they have a two-column EIGENVAL file.\r\n\r\nSo, I do think it could be worthwhile to provided support for what I presume is an older two-column format. That being said, I can only imagine that a spin-polarized calculation would then have three columns in the old VASP version. That could cause problems because in the current versions of VASP, the three-column format is for spin-unpolarized calculations (and five columns for spin-polarized). Either way, this is all to say that test cases should definitely be provided.",
"Thank you for taking this. It has been a long time since I was working on it. The version of vasp I was working was older and the EIGENVAL looks exactly by the EIGENVAL_old provided by @arosen93 , where the occupations are missing. In that case, also for spin polarized there are no occupations listed, so the reading would not be correct either. ",
"From @arosen93's comment, sounds like even with this merged, spin-polarized calcs (3 cols) run with pre 5.4.x Vasp would still be parsed incorrectly as spin-unpolarized by `pymatgen`? I assume the unspoken conclusion here was `pymatgen` EIGENVAL reader doesn't support parsing pre 5.4.x Vasp?",
"Closing this as abandoned."
] | 2021-01-27T14:37:14
| 2023-05-08T19:51:49
|
2023-05-08T19:51:49Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
* add a ine to include the case where the eigenval corresponds to non spin polarized
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2043/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2043/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2043",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2043",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2043.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2043.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2044
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2044/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2044/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2044/events
|
https://github.com/materialsproject/pymatgen/pull/2044
| 795,641,394
|
MDExOlB1bGxSZXF1ZXN0NTYyOTU0OTE3
| 2,044
|
method to estimate NBANDS + tests
|
{
"login": "rwoodsrobinson",
"id": 7270689,
"node_id": "MDQ6VXNlcjcyNzA2ODk=",
"avatar_url": "https://avatars.githubusercontent.com/u/7270689?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rwoodsrobinson",
"html_url": "https://github.com/rwoodsrobinson",
"followers_url": "https://api.github.com/users/rwoodsrobinson/followers",
"following_url": "https://api.github.com/users/rwoodsrobinson/following{/other_user}",
"gists_url": "https://api.github.com/users/rwoodsrobinson/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rwoodsrobinson/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rwoodsrobinson/subscriptions",
"organizations_url": "https://api.github.com/users/rwoodsrobinson/orgs",
"repos_url": "https://api.github.com/users/rwoodsrobinson/repos",
"events_url": "https://api.github.com/users/rwoodsrobinson/events{/privacy}",
"received_events_url": "https://api.github.com/users/rwoodsrobinson/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @rwoodsrobinson !",
"If possible, I'd also take into account whether spin-orbit coupling is included in the calculation (`LSORBIT = True`), as that will approximately double NBANDS.",
"Thanks for mentioning @utf, I'll add it. Know of any other cases where VASP changes the NBANDS value, or how they are estimated as a function of # of cores?",
"I'm not sure of any other settings that will change the number of bands drastically. I think my rough understanding is that NBANDS has to be divisible by `NCORE` (or equivalently by #cores / NPAR) and if neither are set then by the number of cores per node.",
"@utf it can definitely change drastically in some cases, e.g. imagine a small cell with few bands on something like KNL (64 cores) with default parallelization (I think; I would have to check specific circumstances, but I've definitely seen it change significantly, perhaps this was with OMP threads too)",
"Apologies, I meant changing due to a specific INCAR setting (like LSORBIT) rather than the number of cores.",
"Good point about SOC btw, VASP manual doesn't mention that but makes sense"
] | 2021-01-28T03:46:45
| 2021-01-28T20:45:34
|
2021-01-28T04:23:02Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* created method to estimate NBANDS for VASP input set
* simplified `nelect` property
* added tests
## Checklist
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2044/reactions",
"total_count": 2,
"+1": 2,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2044/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2044",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2044",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2044.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2044.patch",
"merged_at": "2021-01-28T04:23:02Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2045
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2045/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2045/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2045/events
|
https://github.com/materialsproject/pymatgen/issues/2045
| 797,344,594
|
MDU6SXNzdWU3OTczNDQ1OTQ=
| 2,045
|
"Failed to build spglib" while installing pymatgem
|
{
"login": "pratapslair",
"id": 41544177,
"node_id": "MDQ6VXNlcjQxNTQ0MTc3",
"avatar_url": "https://avatars.githubusercontent.com/u/41544177?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/pratapslair",
"html_url": "https://github.com/pratapslair",
"followers_url": "https://api.github.com/users/pratapslair/followers",
"following_url": "https://api.github.com/users/pratapslair/following{/other_user}",
"gists_url": "https://api.github.com/users/pratapslair/gists{/gist_id}",
"starred_url": "https://api.github.com/users/pratapslair/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/pratapslair/subscriptions",
"organizations_url": "https://api.github.com/users/pratapslair/orgs",
"repos_url": "https://api.github.com/users/pratapslair/repos",
"events_url": "https://api.github.com/users/pratapslair/events{/privacy}",
"received_events_url": "https://api.github.com/users/pratapslair/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Pls report your OS when reporting issues. For this, I would recommend you try using conda installation as recommended in the official pymatgen documentation first. ",
"OS is Windows 10. ",
"The issue was resolved after i updated Microsoft Visual C++ greater than 14.0. Thank you."
] | 2021-01-30T05:54:06
| 2021-01-30T11:13:20
|
2021-01-30T11:13:20Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I tried to install 'pymatgem' using 'pip install pymatgem' command. It threw an error saying
"ERROR: Failed building wheel for spglib
Failed to build spglib
ERROR: Could not build wheels for spglib which use PEP 517 and cannot be installed directly"
The python version i am using is, python 3.8
Can you please let me know what I should do to troubleshoot this?
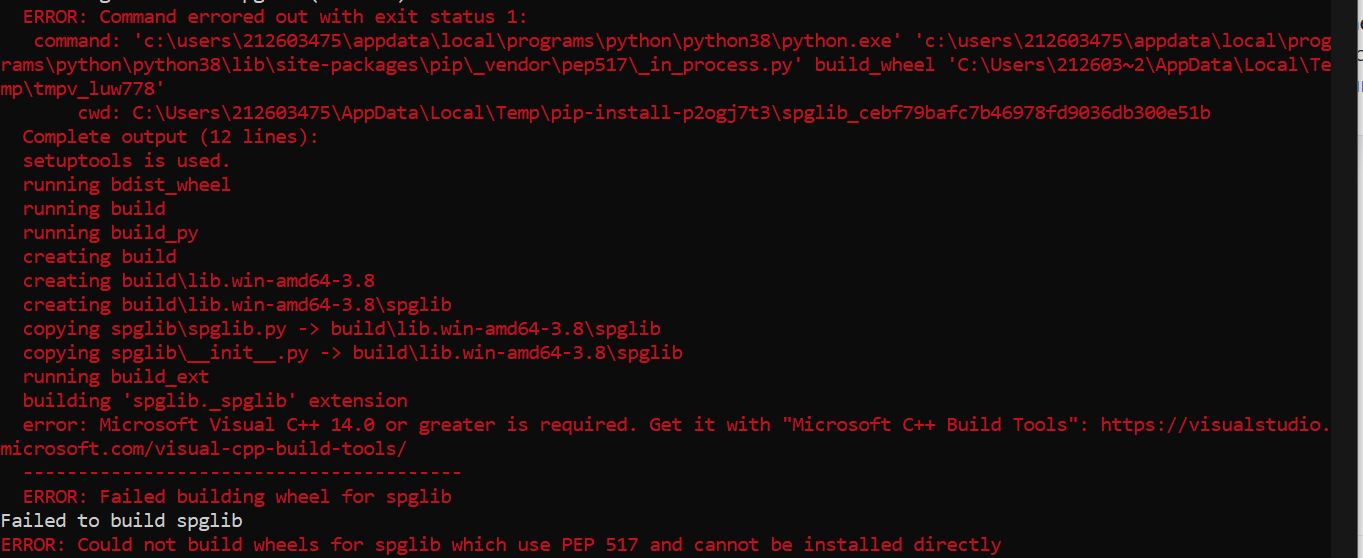
|
{
"login": "pratapslair",
"id": 41544177,
"node_id": "MDQ6VXNlcjQxNTQ0MTc3",
"avatar_url": "https://avatars.githubusercontent.com/u/41544177?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/pratapslair",
"html_url": "https://github.com/pratapslair",
"followers_url": "https://api.github.com/users/pratapslair/followers",
"following_url": "https://api.github.com/users/pratapslair/following{/other_user}",
"gists_url": "https://api.github.com/users/pratapslair/gists{/gist_id}",
"starred_url": "https://api.github.com/users/pratapslair/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/pratapslair/subscriptions",
"organizations_url": "https://api.github.com/users/pratapslair/orgs",
"repos_url": "https://api.github.com/users/pratapslair/repos",
"events_url": "https://api.github.com/users/pratapslair/events{/privacy}",
"received_events_url": "https://api.github.com/users/pratapslair/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2045/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2045/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2046
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2046/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2046/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2046/events
|
https://github.com/materialsproject/pymatgen/pull/2046
| 797,523,093
|
MDExOlB1bGxSZXF1ZXN0NTY0NTA0OTQ5
| 2,046
|
[WIP] build system: adopt PEP-518
|
{
"login": "ltalirz",
"id": 1053245,
"node_id": "MDQ6VXNlcjEwNTMyNDU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1053245?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ltalirz",
"html_url": "https://github.com/ltalirz",
"followers_url": "https://api.github.com/users/ltalirz/followers",
"following_url": "https://api.github.com/users/ltalirz/following{/other_user}",
"gists_url": "https://api.github.com/users/ltalirz/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ltalirz/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ltalirz/subscriptions",
"organizations_url": "https://api.github.com/users/ltalirz/orgs",
"repos_url": "https://api.github.com/users/ltalirz/repos",
"events_url": "https://api.github.com/users/ltalirz/events{/privacy}",
"received_events_url": "https://api.github.com/users/ltalirz/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Ironic that the switch to the `pyproject.toml` surfaces the issue described in https://github.com/materialsproject/pymatgen/issues/2010#issuecomment-770274405 ;-)\r\nI'll try to have a look what's going on",
"@shyuep perhaps one of your recent commits broke the macos build; if I remember correctly they were working just before (at least the issue doesn't look related to the `pyproject.toml`) - but this `pyproject.toml` now seems to fix the linux builds.\r\n\r\nnot sure whether you saw this PR previously or were just working on the same thing by chance.",
"However, it is not an ideal solution - unfortunately, build isolation was introduced in pip without making it possible to ensure that the version of a package (`numpy`) is the same both in the build environment and in the target environment (without hardcoding one specific version). See also https://github.com/pypa/pip/issues/6144\r\n\r\nIf two numpy versions result in incompatible binaries (as now seems to be the case with numpy >= 1.20 vs numpy < 1.20), the only \"safe\" way I see is to limit the numpy version within a boundary (e.g. >=1.18, <1.20) inside which such incompatibility does not exist (and to do it both in the `pyproject.toml` and in the `setup.py` `install_requires`).\r\n\r\nOne can still make this window depend on the python version if desired.\r\n",
"Just to reiterate - while this `pyproject.toml` makes the tests pass, I believe this is because on CI you install numpy 1.19.2 *before* installing pymatgen.\r\n\r\nSomeone installing pymatgen into an empty environment (therefore pulling in numpy 1.20.0 on python >=3.7) will still have issues.\r\n\r\nThe question is: which numpy version range do you prefer for which python version?\r\n\r\nI guess for the moment one can decide to skip numpy 1.20 across the board, and also use `\"numpy>=1.18,<1.20; python_version<='3.9'\"` in the `install_requires`, but at some point people will probably want to use it since it [brings wider use of simd](https://numpy.org/devdocs/release/1.20.0-notes.html).",
"Thanks. I already incorporated your changes manually since I had to fix my CI build. I basically just copied your pyproject. For now, let's pin to bumpy 1.19. I couldn't get 1.20 to work. ",
"Yes, but please don't forget to limit it also in the `setup.py` (as I did in the last commit) or I believe users installing pymatgen in to a new environment may face issues.\r\n\r\nI would also suggest to add a CI test where you *don't* install your `requirements.txt` first, but directly `pip install -e .` (since this is what many users will do).\r\n\r\nUnless you already have such a test - then it's fine already.",
"Thanks. Yes, I did change the setup.py as well. I did try the pip install -e . method and it seems to be fine for now. "
] | 2021-01-30T20:22:45
| 2021-01-31T16:52:44
|
2021-01-31T01:01:20Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
fix #2010
## Summary
The setup.py was using the deprecated `setup_requires` keyword, which
can result in dependencies being installed via `easy_install`.
`easy_install` is outdated and should no longer be used to install
packages.
This commit adopts PEP 518 and uses the `pyproject.toml` file to specify
the dependencies of the build system.
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [ ] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2046/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2046/timeline
| null | true
| true
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2046",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2046",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2046.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2046.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2047
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2047/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2047/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2047/events
|
https://github.com/materialsproject/pymatgen/issues/2047
| 797,542,729
|
MDU6SXNzdWU3OTc1NDI3Mjk=
| 2,047
|
Possible bug with Gaussian smearing
|
{
"login": "jmmshn",
"id": 14003693,
"node_id": "MDQ6VXNlcjE0MDAzNjkz",
"avatar_url": "https://avatars.githubusercontent.com/u/14003693?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jmmshn",
"html_url": "https://github.com/jmmshn",
"followers_url": "https://api.github.com/users/jmmshn/followers",
"following_url": "https://api.github.com/users/jmmshn/following{/other_user}",
"gists_url": "https://api.github.com/users/jmmshn/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jmmshn/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jmmshn/subscriptions",
"organizations_url": "https://api.github.com/users/jmmshn/orgs",
"repos_url": "https://api.github.com/users/jmmshn/repos",
"events_url": "https://api.github.com/users/jmmshn/events{/privacy}",
"received_events_url": "https://api.github.com/users/jmmshn/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"> But maybe since testing is lacking we should just add a deprecation warning for now?\r\n\r\nDeprecation warning is only for if we're going to remove the function. A general warning that it is untested would be useful; we should either add tests asap or remove though, untested code shouldn't be in pymatgen.",
"Thanks for reporting/noticing this.",
"I think the code has been migrated to pymatgen-diffusion. Can you check? If it is already there, I will just delete this code.",
"The `pathfinder` in pymatgen diffusion is different from this `pathfinder` and the code that I added to pymatgen-diffusion uses both `pathfinders` but I cannot find anywhere in our codebase that uses this `StaticPotential` that contains this incorrect Gaussian smearing implementation."
] | 2021-01-30T21:47:06
| 2023-08-13T16:34:58
|
2023-08-13T16:34:58Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
The code might have an issue.
https://github.com/materialsproject/pymatgen/blob/4f906e3578cc20372ff4f12fcda9db3aaebec975/pymatgen/analysis/path_finder.py#L388
I used a Si CHGCAR and ran
```
from pymatgen.analysis.path_finder import StaticPotential
sp = StaticPotential(chgcar.structure, pot=chgcar.data['total'])
sp.gaussian_smear(r=2)
sp.get_v().shape
```
This should return an `sp` where potential has the same dimensions as the original but instead give a smaller array.
Also just looking at the code I think something is wrong with how the number of required images are computed for skewed cells.
(There's also no tests for these classes)
I have a working solution for this kind of smearing in PyRho but might take a while until things are migrated here.
I just wanna create this issue for future reference. But maybe since testing is lacking we should just add a deprecation warning for now?
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2047/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2047/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2048
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2048/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2048/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2048/events
|
https://github.com/materialsproject/pymatgen/pull/2048
| 798,966,795
|
MDExOlB1bGxSZXF1ZXN0NTY1NjkxNzg2
| 2,048
|
[WIP] Input/output for CASTEP
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 3093102097,
"node_id": "MDU6TGFiZWwzMDkzMTAyMDk3",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/stale",
"name": "stale",
"color": "A99B62",
"default": false,
"description": "Abandoned or conflicting PRs and outdated issues"
}
] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-02-02T06:12:32
| 2024-08-03T19:01:57
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Uses some code from the MIT-licensed [sumo](https://github.com/SMTG-UCL/sumo/blob/master/sumo/io/castep.py) implementation by @ajjackson, [used with permission](https://github.com/SMTG-UCL/sumo/pull/61#issuecomment-567632927) (thank you!).
I've re-worked the .cell implementation to make it more consistent with other parts of pymatgen (`write_file` instead of `to_file`, adding a `__str__` method), added type hints. I also added a `CellKeyword` and `ParamKeyword` Enum nominally for avoiding bugs, though in practice likely just to make things more awkward. This interface also adds magnetic moment writing and parsing.
Opening the PR now in case there are any comments that might affect the implementation before I finish working on it.
## Additional dependencies introduced (if any)
* None
## TODO (if any)
- [x] `.cell` I/O
- [x] `.param` I/O
- [ ] A more general `KPoints` class, perhaps a unified interface for both VASP and CASTEP
- [x] Basic input set for static energy calculation and relaxation
Later, not in this PR, will be band structure parsing.
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2048/reactions",
"total_count": 3,
"+1": 2,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 1
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2048/timeline
| null | true
| true
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2048",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2048",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2048.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2048.patch",
"merged_at": null
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2049
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2049/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2049/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2049/events
|
https://github.com/materialsproject/pymatgen/pull/2049
| 799,186,387
|
MDExOlB1bGxSZXF1ZXN0NTY1ODcyMTM4
| 2,049
|
[WIP] magnetic spacegroup analyzer
|
{
"login": "mt-huebsch",
"id": 42400095,
"node_id": "MDQ6VXNlcjQyNDAwMDk1",
"avatar_url": "https://avatars.githubusercontent.com/u/42400095?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mt-huebsch",
"html_url": "https://github.com/mt-huebsch",
"followers_url": "https://api.github.com/users/mt-huebsch/followers",
"following_url": "https://api.github.com/users/mt-huebsch/following{/other_user}",
"gists_url": "https://api.github.com/users/mt-huebsch/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mt-huebsch/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mt-huebsch/subscriptions",
"organizations_url": "https://api.github.com/users/mt-huebsch/orgs",
"repos_url": "https://api.github.com/users/mt-huebsch/repos",
"events_url": "https://api.github.com/users/mt-huebsch/events{/privacy}",
"received_events_url": "https://api.github.com/users/mt-huebsch/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 3093102097,
"node_id": "MDU6TGFiZWwzMDkzMTAyMDk3",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/stale",
"name": "stale",
"color": "A99B62",
"default": false,
"description": "Abandoned or conflicting PRs and outdated issues"
}
] |
open
| false
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
] | null |
[
"Thank you @mt-huebsch! This looks like an incredibly useful contribution, I would be happy to see this merged.\r\n\r\nI will take some time to review the PR carefully to provide comments and a response to your question on testing.\r\n\r\n> I noticed that this file does not get installed when installing pymatgen from my repository in a new enviroment. Maybe I should have chosen a different extension? Or does this need to be configured?\r\n\r\nBy default, only `.json`, `.csv` and `.yaml` files are packaged as specified by [this line in the setup.py](https://github.com/materialsproject/pymatgen/blob/5a7830b0a833e35ca99eb637c82d5679086e8675/setup.py#L52). We can either choose to include .txt files too (either all .txt files, or a specific file), or we can change the file to be one of these other formats instead.",
"Dear @mkhorton, thank you for your quick response!\r\n\r\nRegarding the .txt file, I will change the format. No problem.\r\nI will wait for some more comments from you and then make the changes. \r\n\r\nThank you for showing interest in my contribution.",
"This is fantastic. Thank you @mt-huebsch!\r\n\r\nI have a couple of questions. The major roadblock that I encountered while trying to develop a magnetic symmetry analyzer was for cases in which the \"setting\" of the msg operations is non-standard. From doing a search on the MAGNDATA server, you can see that the majority of magnetic structures (822 out of the 1217 structures), the msg operations are defined in a non-standard setting (defined by a similarity transform from the standard lattice or vector space).\r\n\r\nWhile I get the same very nice result for Ca3Co2-xMnxO6, I get an error: \r\n`Error in the list of symmetry operations.`\r\nif I try a structure in which the msg is in a non-standard setting, such as [FePO4](http://webbdcrista1.ehu.es/magndata/index.php?this_label=0.17), which has an msg P212121 (\\#19.25) within the non-standard setting (a,b,c;0,1/2,3/4).\r\n\r\nI'm curious if I'm doing something wrong? Here is how I got the error:\r\n\r\n```\r\n>>> filename_a = \"0.13_Ca3Co2-xMnxO6.mcif\"\r\n>>> struct_a = Structure.from_file( filename_a )\r\n>>> ma_a = MagneticSpacegroupAnalyzer( struct_a )\r\n>>> print( \"OG info: \", ma_a.get_og() )\r\nOG info: ('R3c', '161.1.1300')\r\n>>> \r\n>>> filename_b = \"0.17_FePO4.mcif\"\r\n>>> struct_b = Structure.from_file( filename_b )\r\n>>> ma_b = MagneticSpacegroupAnalyzer( struct_b )\r\nError in the list of symmetry operations.\r\n```\r\n\r\nIt wasn't trivial to me how to find the similarity transform for a msg. I'm curious if that is what's happening here. I'll dive deeper into your code to see if you account for this. Did you encounter a similar issue? It's possible to check this setting transformation with the symmetry finder in the ISOTROPY suite: https://stokes.byu.edu/iso/findsym.php. \r\n\r\nI believe that there are ways of identifying if two sg representations are \"similar\" using [matrix normal forms](https://en.wikipedia.org/wiki/Canonical_form), which appears to be the basis for many of the symmetry finding algorithms developed by Stokes et. al. and in `spglib`. I would be happy to talk about this in more depth if you also explored this. \r\n\r\nIf you found a work-around for finding the transformation to another setting, I am very curious to learn what your approach was. I don't see where you account for this in the code but I'll look more closely. If this is an issue, maybe this feature can be implemented in the future.\r\n\r\nThank you again!",
"Dear @guymoore13,\r\n\r\nThank you for running tests on the code!!\r\n\r\nIndeed, you have discovered a bug. I need to transform to the standardized conventional cell. As far as I understand the same ambiguity is already present in the spacegroup operations, so spglib can handle it. I am already working on the solution and can perform all necessary transformations, but at the moment I haven't figured out how \"standardized\" is defined exactly. Maybe I am wrong but spglib and [Stokes et al](https://stokes.byu.edu/iso/magneticspacegroups.php) seem to have a different definition of standardized.\r\n\r\nPlease let me know in case you have any thoughts on that.",
"Dear @mt-huebsch, thank you for your response!\r\n\r\nYou are correct that spglib accounts for this, although I'm also still trying to identify how they go about addressing the setting/basis issue. Due to the ambiguity in the \"standard setting\" in which to compare operations, I was trying a different approach - because I encountered this question as well.\r\n\r\nFor this reason, I was trying to find a way of comparing \"reduced representations\" of the msg operations - which should be independent from the settings of each representation (and therefore the similarity transform to the standard setting). I haven't made much progress on this front, but I will let you know if I do - it may be nice to compare notes at some point. I was having issues with the \"uniqueness\" of the diagonalization scheme that I was using.\r\n\r\nYour approach to find the standard setting may be more robust. I look forward to hearing how it goes!",
"Great work and a very important contribution! I want to report just a couple of minor things I have noticed so far\r\n\r\n1. In _paramag_symmetry_operations of MagneticSpacegroupAnalyzer class:\r\n`\r\n rotation = self._spacegroupAnalyzer._space_group_data['rotations']\r\n`\r\n`\r\n translation = self._spacegroupAnalyzer._space_group_data['translations']\r\n`\r\n\r\nYour rotations and translations come directly from spglib. However, it often returns small translation vectors that are duplicated i,e. [0, 0, 0], [1e-17,1e-17,1e-17] etc You can reproduce it on 1.0.1_Ag2CrO2.mcif from MAGNDATA. As a result, the number of symmetry operations is larger than it should be (40 symmetries instead of 4 in case of Ag2CrO2). A similar issue is described in _get_symmetry of SpacegroupAnalyzer. One can avoid it by using \r\n\r\n`\r\n rotation = self._spacegroupAnalyzer._get_symmetry()[0]\r\n`\r\n`\r\n translation = self._spacegroupAnalyzer._get_symmetry()[1]\r\n`\r\n\r\n2. You got the following line twice:\r\n`\r\n if np.any(np.abs(self._structure._lattice.matrix - self._spacegroupAnalyzer._cell[0]) > 1e-12):\r\n exit(\"Error in the definitions of the unit cell.\")\r\n`\r\n\r\nFirst, in `__init__` and then in `_paramag_symmetry_operations`.",
"@maxmarkov thanks for the testing. Could you clarify how:\r\n\r\n```\r\nrotation = self._spacegroupAnalyzer._get_symmetry()[0]\r\ntranslation = self._spacegroupAnalyzer._get_symmetry()[1]\r\n```\r\n\r\navoids the duplicated `SymmOp` issue? Related to this, we use `np.allclose()` to define whether two `SymmOps` are equal but the tolerance is currently very tight, do you think this should be relaxed to make it easier to filter out duplicates?",
"Dear @maxmarkov! Thank you for testing. Honestly, I haven't had a look at q>0 magnetic structures, but that is just a side remark. I will add it to my to do list.\r\n\r\nRegarding _spacegroupAnalyzer._get_symmetry(), it is always good to make the code more robust. But rather than hardcoding a rounding, maybe it should be implemented using pos_tolerance with a good fall back default value. What is pymatgen's policy on that, @mkhorton? \r\n\r\nBdw, previously I used _spacegroupAnalyzer._get_symmetry_operations() at that point, but this function always returns the conventional symmetries. This is unexpected behavior to me, if the input structure is primitive. That is why I wrote _paramag_symmetry_operations() in the first place. Although it is probably not good if the SpacegroupAnalyzer and the MagneticSpacegroupAnalyzer behave differently. Harmonizing the behavior should maybe be added to the to do list, too.\r\n\r\n@guymoore13, regarding the setting issue, I have a small update:\r\n\r\nEvery MSPG symbol is derived from a non-magnetic SPG symbol. That is why fortunately the same alternative settings apply. But the standard setting for MSPGs is not the one closest to the standard setting of the parent SPG [1]. \r\nFurthermore, Hall symbols [2] represent an explicit-origin spacegroup notation. There is a method to decode Hall symbols into matrix representations described in ITC volume B. The resulting matrices are precomputed and stored in spglib [3].\r\nSpglib determines matrix representations for SPG operations and then compares to that dataset. Both tasks, determining the symmetries and comparing representations, are quite involved and currently not available individually.\r\nIt should be possible to overload the initializer of spglib to accept a set of symmetry operations as an input instead of a structure. Then, the transformation matrix to a chosen setting (by selecting the Hall number) can be returned. I will contact Togo-sensei and look into spglib's repo to check how difficult this will be.\r\n\r\n[1] https://www.annualreviews.org/doi/10.1146/annurev-matsci-070214-021008\r\n[2] https://doi.org/10.1107/S0567739481001228\r\n[3] https://arxiv.org/pdf/1808.01590.pdf\r\n\r\nI will try to carve out more time to get this functionality ready for distribution. Thank you all for your feedback and patience. \r\n\r\nBest regards!",
"Thank you for the detailed response @mt-huebsch. Yes, I think we are saying more or less the same thing. You mention the Hall symbol, and my approach has been to store the matrix normal forms required to identify the Hall symbols (on [page 7 of the spglib paper that you shared](https://arxiv.org/pdf/1808.01590.pdf#page=7)). They use the [Smith canonical form](https://en.wikipedia.org/wiki/Smith_normal_form). I saw this a bit later - I was using a different matrix diagonalization. However, I recently discovered that the Smith canonical form has some very nice properties, especially for integer matrices. They use `SageMath` to compute the Smith normal forms. I have been experimenting with other Python packages to compute normal forms.\r\n\r\nTake care, Guy"
] | 2021-02-02T11:16:46
| 2024-08-03T19:01:57
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Added MagneticSpacegroupAnalyzer that given a magnetic structure
* finds all magnetic spacegroup operations
* determines the magnetic spacegroup
* can return the magnetic pointgroup
* can compute all magnetic domains
Added ApplyMagSymmOpTransformation to standard transformations
* apply MagSymmOp to a magnetic structure
## Additional dependencies introduced
None
## To Do
* add a test
I was not sure how/where to add a test. This snipped shows how to run the main functionality of the code, i.e. finding the magnetic spacegroup:
`path_to_s = "0.13_Ca3Co2-xMnxO6.mcif"`
`s = mg.Structure.from_file( path_to_s)`
`ma = MagneticSpacegroupAnalyzer( s )`
`print( "OG info: ", ma.get_og() )`
0.13_Ca3Co2-xMnxO6.mcif can be downloaded from MAGNDATA
[http://webbdcrista1.ehu.es/magndata/index.php?index=0.13](http://webbdcrista1.ehu.es/magndata/index.php?index=0.13)
The OG info for this material is ('R3c', '161.1.1300')
To obtain the magnetic domains, run
`domain_dict = ma.get_domains()`
`print( "Magnetic domains: ", len(domain_dict), [mop.as_xyzt_string() for mop in domain_dict.keys()] )`
This retruns
Magnetic domains: 4 ['x, y, z, +1', 'x, y, z, -1', '-x, -y, -z, +1', '-x, -y, -z, -1']
I'd be happy to provide more extensive tests, if you point me in the right direction.
* enable pip to get pymatgen/symmetry/mspg2mpg_map.txt
I noticed that this file does not get installed when installing pymatgen from my repository in a new enviroment. Maybe I should have chosen a different extension? Or does this need to be configured?
* code review
This is my first contribution, so I would appreciate any comment.
## Comment
In the past I have submitted a contribution, where the search for the magnetic spacegroup was dependent on tools of the Bilbao Crystallographic Server. This has now been replaced by a search independent of any external sources. The code is robust enough to handle uncertainties of a typical ab initio run and can return all magnetic domains. Thus, if two ab initio results correspond to different magnetic domains, this feature will help identify the correspondence.
Looking forwards to your feedback!
Cheers,
Marie-Therese
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2049/reactions",
"total_count": 3,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 3,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2049/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2049",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2049",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2049.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2049.patch",
"merged_at": null
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2050
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2050/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2050/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2050/events
|
https://github.com/materialsproject/pymatgen/issues/2050
| 799,451,307
|
MDU6SXNzdWU3OTk0NTEzMDc=
| 2,050
|
Typing for pymatgen
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 5318918309,
"node_id": "LA_kwDOACgets8AAAABPQhApQ",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/linting",
"name": "linting",
"color": "5DBC83",
"default": false,
"description": "Linting and quality assurance"
},
{
"id": 5509830160,
"node_id": "LA_kwDOACgets8AAAABSGlWEA",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/types",
"name": "types",
"color": "7D66D9",
"default": false,
"description": "Type all the things"
}
] |
closed
| false
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
] | null |
[
"1. I've been looking for solutions for this problem too. I found [nptyping](https://github.com/ramonhagenaars/nptyping) which defines a set of types for Numpy arrays of specific shapes. I'd rather not rely on a third-party here though. We can simply alias `Vector` and `Matrix` to `ArrayLike` for now, so at least it's easier to replace with an appropriate type later.\r\n2. Ok, sounds good.\r\n3. Agreed -- I think mixing of species/element in different places already causes some confusion. Avoiding custom types will make things simpler. The type hint I am using a lot however is `Literal` (as a substitute for `str` when only one of a few specific strings are valid), but this is in the standard library.",
"I've been slowly adding types to the most frequently used parts of `pymatgen` over the last 2 years. Still far from complete coverage but `core` is mostly there which is what this issue was aiming for. Closing this as completed."
] | 2021-02-02T16:41:59
| 2023-05-30T16:58:02
|
2023-05-30T16:58:01Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I think we should try to get at least the core aspects of pymatgen typed. Here are some thoughts:
1. Numpy 1.20 has introduced the numpy.typing.ArrayLike type, which we can rely on for many instances. I created one in pymatgen.util.typing that uses ArrayLike where available and falls back to a default creation where it is not. One disadvantage of using the numpy ArrayLike is that it is extremely broad and does not distinguish between vectors or matrices. But the advantage is that we do not need to maintain anything. Not sure what is the best course here.
2. I added the SpeciesLike and CompositionLike types. These are basically things that can be passed into get_el_sp or Composition() to return an Element/Species/DummySpecies or a Composition object respectively. CompositionLike is a superset of SpeciesLike since Species can be converted to just {sp: 1} compositions.
3. We should minimize the types we create. The above + the core python types should cover 90% of use cases. But I welcome any suggestions of addition types we want to create.
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2050/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2050/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2051
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2051/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2051/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2051/events
|
https://github.com/materialsproject/pymatgen/issues/2051
| 800,472,711
|
MDU6SXNzdWU4MDA0NzI3MTE=
| 2,051
|
Issue with EnumlibAdaptor during doping transformation
|
{
"login": "marcoloco23",
"id": 55764616,
"node_id": "MDQ6VXNlcjU1NzY0NjE2",
"avatar_url": "https://avatars.githubusercontent.com/u/55764616?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/marcoloco23",
"html_url": "https://github.com/marcoloco23",
"followers_url": "https://api.github.com/users/marcoloco23/followers",
"following_url": "https://api.github.com/users/marcoloco23/following{/other_user}",
"gists_url": "https://api.github.com/users/marcoloco23/gists{/gist_id}",
"starred_url": "https://api.github.com/users/marcoloco23/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/marcoloco23/subscriptions",
"organizations_url": "https://api.github.com/users/marcoloco23/orgs",
"repos_url": "https://api.github.com/users/marcoloco23/repos",
"events_url": "https://api.github.com/users/marcoloco23/events{/privacy}",
"received_events_url": "https://api.github.com/users/marcoloco23/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This error typically occurs if the enumeration was unsuccessful. Either the cell is too large to be enumerated or the enum.x exited with error. "
] | 2021-02-03T16:07:38
| 2021-02-04T10:31:06
|
2021-02-04T10:31:06Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
While trying to perform a doping transformation, I get the following attribute error:
'EnumlibAdaptor' object has no attribute 'structures'.
The error occurs on line 457 of "https://github.com/materialsproject/pymatgen/blob/v2021.1.28/pymatgen/transformations/advanced_transformations.py#L1095-L1258".
It seems as though 'EnumlibAdaptor' has an attribute "structure" but not "structures" which leads to the error.
|
{
"login": "marcoloco23",
"id": 55764616,
"node_id": "MDQ6VXNlcjU1NzY0NjE2",
"avatar_url": "https://avatars.githubusercontent.com/u/55764616?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/marcoloco23",
"html_url": "https://github.com/marcoloco23",
"followers_url": "https://api.github.com/users/marcoloco23/followers",
"following_url": "https://api.github.com/users/marcoloco23/following{/other_user}",
"gists_url": "https://api.github.com/users/marcoloco23/gists{/gist_id}",
"starred_url": "https://api.github.com/users/marcoloco23/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/marcoloco23/subscriptions",
"organizations_url": "https://api.github.com/users/marcoloco23/orgs",
"repos_url": "https://api.github.com/users/marcoloco23/repos",
"events_url": "https://api.github.com/users/marcoloco23/events{/privacy}",
"received_events_url": "https://api.github.com/users/marcoloco23/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2051/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2051/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2052
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2052/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2052/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2052/events
|
https://github.com/materialsproject/pymatgen/issues/2052
| 800,573,311
|
MDU6SXNzdWU4MDA1NzMzMTE=
| 2,052
|
get_computed_entry() is failing in the Vasprun
|
{
"login": "kshumilov",
"id": 15161636,
"node_id": "MDQ6VXNlcjE1MTYxNjM2",
"avatar_url": "https://avatars.githubusercontent.com/u/15161636?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/kshumilov",
"html_url": "https://github.com/kshumilov",
"followers_url": "https://api.github.com/users/kshumilov/followers",
"following_url": "https://api.github.com/users/kshumilov/following{/other_user}",
"gists_url": "https://api.github.com/users/kshumilov/gists{/gist_id}",
"starred_url": "https://api.github.com/users/kshumilov/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/kshumilov/subscriptions",
"organizations_url": "https://api.github.com/users/kshumilov/orgs",
"repos_url": "https://api.github.com/users/kshumilov/repos",
"events_url": "https://api.github.com/users/kshumilov/events{/privacy}",
"received_events_url": "https://api.github.com/users/kshumilov/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks for reporting this. @rkingsbury I believe this was added by you? Can you fix it pls? Thanks.",
"Thanks for reporting @kshumilov . @shyuep see #2054 for a fix."
] | 2021-02-03T18:09:59
| 2021-02-03T22:53:05
|
2021-02-03T22:53:05Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
If `io.vasp.outputs.Vasprun()` is describing a run with IVDW tag set to `int` the function `get_computed_entry()` is failing.
The bug is hiding in the line [759](https://github.com/materialsproject/pymatgen/blob/v2021.1.28/pymatgen/io/vasp/outputs.py#L759) where the output of `get()` method on the `self.incar` returns an integer, which doesn't have `strip()` method.
**To Reproduce**
1. Have a `vasprun.xml` with IVDW set to `int`
2. Get `Vasprun` object
3. Call `Vasprun.get_computed_entry()`
Unable to provide file exact IVDW file, but technically it's not needed since the `Vasprun()` can be constructed with the custom `Incar` object.
**Expected behavior**
`ComputedStructureEntry` is returned successfully with correctly identified run IVDW variant.
**Screenshots**
If applicable, add screenshots to help explain your problem.
**Desktop (please complete the following information):**
- OS: Mac
- Version 2021.1.28
**Additional context**
Traceback
```python
/usr/local/Caskroom/miniconda/base/envs/quantum/lib/python3.9/site-packages/pymatgen/io/vasp/outputs.py in get_computed_entry(self, inc_structure, parameters, data)
807 if parameters:
808 param_names.update(parameters)
--> 809 params = {p: getattr(self, p) for p in param_names}
810 data = {p: getattr(self, p) for p in data} if data is not None else {}
811
/usr/local/Caskroom/miniconda/base/envs/quantum/lib/python3.9/site-packages/pymatgen/io/vasp/outputs.py in <dictcomp>(.0)
807 if parameters:
808 param_names.update(parameters)
--> 809 params = {p: getattr(self, p) for p in param_names}
810 data = {p: getattr(self, p) for p in data} if data is not None else {}
811
/usr/local/Caskroom/miniconda/base/envs/quantum/lib/python3.9/site-packages/pymatgen/io/vasp/outputs.py in run_type(self)
757 if self.parameters.get("LUSE_VDW", False):
758 rt += "+rVV10"
--> 759 elif self.incar.get("IVDW", "").strip().upper() in IVDW_TYPES:
760 rt += "vdW-" + IVDW_TYPES[self.incar.get("IVDW", "").strip().upper()]
761 elif self.incar.get("IVDW"):
AttributeError: 'int' object has no attribute 'strip'
```
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2052/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2052/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2053
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2053/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2053/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2053/events
|
https://github.com/materialsproject/pymatgen/issues/2053
| 800,700,595
|
MDU6SXNzdWU4MDA3MDA1OTU=
| 2,053
|
Bug: GibbsComputedStructureEntry returning incorrect energies, running slowly
|
{
"login": "mattmcdermott",
"id": 40469694,
"node_id": "MDQ6VXNlcjQwNDY5Njk0",
"avatar_url": "https://avatars.githubusercontent.com/u/40469694?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mattmcdermott",
"html_url": "https://github.com/mattmcdermott",
"followers_url": "https://api.github.com/users/mattmcdermott/followers",
"following_url": "https://api.github.com/users/mattmcdermott/following{/other_user}",
"gists_url": "https://api.github.com/users/mattmcdermott/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mattmcdermott/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mattmcdermott/subscriptions",
"organizations_url": "https://api.github.com/users/mattmcdermott/orgs",
"repos_url": "https://api.github.com/users/mattmcdermott/repos",
"events_url": "https://api.github.com/users/mattmcdermott/events{/privacy}",
"received_events_url": "https://api.github.com/users/mattmcdermott/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"The phase diagram parts running more slowly will be due to the equality now checking the structure in getting stable_entries. It might be worth removing the structure check from the `__eq__` if the slow down in prohibitive? \n\nI can't think of any edge cases with the gibbs energy correction I went through all the edge cases I could think of.\n\n",
"Thanks for making a PR @mattmcdermott -- could you add an explicit test for the CO2 data too? Just so we can catch a bug like this in future.\r\n\r\nRe. equality checking, open to suggestions on the cleanest way to do that.",
"@CompRhys the issue with the refactoring is double-counting the energy for experimental entries. i.e. if the composition appears in one of the expt. tables, that expt Gibbs Free Energy is added as a `gibbs_correction` to the formation enthalpy, when it should really just be the final energy!\r\n",
"@mkhorton cleanest way would probably be to replace\r\nhttps://github.com/materialsproject/pymatgen/blob/7d2d2001e4b9fb38bf2389a09ac0cfa3030b64ef/pymatgen/entries/__init__.py#L163\r\nwith \r\n```python\r\n return all(_is_robust_eq(other_dict[k], v) for k, v in self_dict.items() if k != \"structure\")\r\n```\r\n\r\nAlthough it doesn't show any speed impact for me in the above code snippet with both checking the struct and not checking it taking ~ 1s but I would guess the slow down was observed in a larger system.",
"@CompRhys I did some line profiling and interestingly the vast majority (99%) of compute time is spent just on the `as_dict()` methods for the Entry objects. The line you referenced with `_is_robust_eq()` only takes about 1% of the time, regardless of whether it's checking the structure!\r\n\r\nNot quite sure how to get around that yet... definitely a drawback of computing gigantic dictionaries. Any thoughts @mkhorton ?\r\n\r\nAlso I will say that the slowdown is significant enough to forbid me from using any equality-based method with `Entry`-derived objects. But that's also because I have to generate **many...many** phase diagrams :) ",
"Using as_dict was to keep the code clean as the alternative was to have lots of custom `__eq__`. Inside the calls to as_dict the slow part will be getting the dict of the structure right?\n\nMaybe it would be better to make `__eq__` and abstract method to avoid as_dict or could memoization of as_dict help?",
"> Not quite sure how to get around that yet... definitely a drawback of computing gigantic dictionaries. Any thoughts @mkhorton ?\r\n\r\n@mattmcdermott Yes, I proposed the as_dict() as a way to get approx_equal behavior between disparate classes. Given that you're encountering performance issues, I'm inclined to say this was the wrong call and we should do something else.",
"@CompRhys The memoization idea seems to work pretty well! I saw a 6x speed-up by adding `functools.lru_cache` to the `ComputedStructureEntry.as_dict()` method. I then saw another 10x speedup by removing the requirement of computing `Structure.as_dict()`. So altogether, about a 60x speedup to cache the result and not do structure dict-checking. \r\n\r\n@mkhorton Is there anything wrong with doing caching for the as_dict() method? Since I'd also like to have the option to remove the structure in equality checking, maybe the better fix is to just implement a different `__eq__` method for ComputedStructureEntry alone?",
"@mattmcdermott nothing wrong with caching, it's just a hack (caching) on top of a hack (using as_dict). But it is a fairly minimal one. To rephrase; I'm ok doing the caching, but would prefer a cleaner solution if one exists.",
"Caching via lru_cache might run into problems as the entries are mutable and could be changed without the cache being cleared\r\n\r\nI think custom `__eq__` for all `EntryLike` is probably the best option but it won't be particularly clean. \r\n",
"Honestly, if classes subclassing `Entry` are meant to be hashable, then every subclass should have to define a `__hash__` and `__eq__`. The big thing is to use a combination of `super()` functions and then check what the new subclass added in.\r\n",
"Another hacky speed up would be to adjust the verbosity of the structure `as_dict` call to be zero rather than default. Reading the code looking for bottlenecks it seems like this could help with the current setup but not sure what it would do practically and haven't tried it yet. It would have to be done in the `_is_dict_eq` so might require adding a `**kwargs` to a lot of `as_dict` so they don't break when given a verbosity argument.",
"I did implement a solution like that, but I think it's too hacky. @mkhorton and I are working on a fix",
"Closing the issue since it was fixed in #2058 ",
"I just wanted to update this thread to say that we merged in a change that makes `Entry` immutable (immutability not enforced, but there is no longer a way to mutate Entry in-place). We also swapped out the `__eq__` for a faster version to correct my mis-step of suggesting the dictionary comparisons; this includes the `entry_id` comparison as a quick way to exit the equality check. I suspect that it can still be improved further but hopefully this will address performance issues for now."
] | 2021-02-03T21:06:22
| 2021-02-16T20:32:51
|
2021-02-05T23:32:11Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
`GibbsComputedStructureEntry` is now returning incorrect formation energies and running very slowly, I think due to refactoring in #2014 (tagging @CompRhys for reference, but I should be able to address).
**To Reproduce**
```python
from pymatgen.analysis.phase_diagram import PhaseDiagram
from pymatgen import MPRester
from pymatgen.entries.computed_entries import GibbsComputedStructureEntry
with MPRester() as mpr:
all_entries = mpr.get_entries_in_chemsys("C-O", inc_structure="final")
pd_mp = PhaseDiagram(all_entries)
temp = 900
gibbs_entries = GibbsComputedStructureEntry.from_pd(pd_mp,temp)
pd_gibbs = PhaseDiagram(gibbs_entries)
for e in sorted(list(pd_gibbs.stable_entries),key=lambda x: x.composition.reduced_formula):
print(e.composition.reduced_formula, pd_gibbs.get_form_energy_per_atom(e))
```
**Expected behavior**
The formation energy of CO2 should be around -1.367 eV/atom at T=900 K, according to NIST-JANAF data. Instead, the energy output currently shows -3.147 eV/atom.
**Desktop (please complete the following information):**
- OS: Mac
**Additional context**
I will be making a PR to address ASAP -- will also try to add more tests to catch bugs like this in the future.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2053/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2053/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2054
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2054/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2054/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2054/events
|
https://github.com/materialsproject/pymatgen/pull/2054
| 800,742,918
|
MDExOlB1bGxSZXF1ZXN0NTY3MTc5NDc5
| 2,054
|
fix IVDW bug in Vasprun.run_type
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @rkingsbury "
] | 2021-02-03T22:12:23
| 2021-02-04T00:43:57
|
2021-02-03T22:53:05Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Fix #2052
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2054/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2054/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2054",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2054",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2054.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2054.patch",
"merged_at": "2021-02-03T22:53:05Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2055
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2055/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2055/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2055/events
|
https://github.com/materialsproject/pymatgen/pull/2055
| 801,705,466
|
MDExOlB1bGxSZXF1ZXN0NTY3OTc0NTY4
| 2,055
|
Additions to Q-Chem IO
|
{
"login": "espottesmith",
"id": 17201238,
"node_id": "MDQ6VXNlcjE3MjAxMjM4",
"avatar_url": "https://avatars.githubusercontent.com/u/17201238?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/espottesmith",
"html_url": "https://github.com/espottesmith",
"followers_url": "https://api.github.com/users/espottesmith/followers",
"following_url": "https://api.github.com/users/espottesmith/following{/other_user}",
"gists_url": "https://api.github.com/users/espottesmith/gists{/gist_id}",
"starred_url": "https://api.github.com/users/espottesmith/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/espottesmith/subscriptions",
"organizations_url": "https://api.github.com/users/espottesmith/orgs",
"repos_url": "https://api.github.com/users/espottesmith/repos",
"events_url": "https://api.github.com/users/espottesmith/events{/privacy}",
"received_events_url": "https://api.github.com/users/espottesmith/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Pls just run black through your code first. Also, can you try to add some type annotations? That would be helpful.",
"I'll work on the linting problems.\r\n\r\nAre there any specific areas that you think need type hints? Mostly I've just modified existing functions or made close copies (such as the new Sets classes in Q-Chem IO). I didn't add a lot of new functions in places that already had type hints.",
"Well, type hints are not compulsory, but if you can add them wherever in the qchem package, that would be helpful.",
"Hmm... I am not sure what you need to do with black. Simply running `black <filename>` should fix all the black related lining errors. Then you can focus on the rest of the stuff. That was what I did for the entire pymatgen code base in ~10 seconds.",
"Yeah, I have this problem pretty consistently where running black doesn't completely resolve all issues, so I have to go back and fix by hand.",
"Thanks for all the work on this PR @espottesmith !"
] | 2021-02-04T23:12:35
| 2022-01-31T21:04:43
|
2021-02-08T17:27:56Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Add ability to handle more diverse job types (including potential energy surface scans and transition-state optimizations) to the Q-Chem IO module.
One other small change was moving the metal_edge_extender function from pymatgen.analysis.fragmenter to pymatgen.analysis.local_env. As this function determines bonding and doesn't directly have to do with creating molecular fragments, it belongs with the NearNeighbor classes.
## TODO (if any)
None
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [X ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [ X] Tests have been added for any new functionality or bug fixes.
- [ ] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2055/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2055/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2055",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2055",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2055.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2055.patch",
"merged_at": "2021-02-08T17:27:56Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2056
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2056/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2056/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2056/events
|
https://github.com/materialsproject/pymatgen/issues/2056
| 801,745,014
|
MDU6SXNzdWU4MDE3NDUwMTQ=
| 2,056
|
latexify_ion is broken
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Proposed fix:\r\n\r\nChanging the regex match pattern from\r\n`\r\nre.sub(r\"()\\[([^)]*)\\]\", r\"\\1$^{\\2}$\", formula)\r\n`\r\nto \r\n`\r\nre.sub(r\"([-+][\\d\\.]*)\", r\"$^{\\1}$\", )\r\n`\r\n\r\nSeems to work as expected. See [regex tester](https://regex101.com/r/HKAgL1/1).\r\n\r\nIt seems to me it might be better to integrate this with `pymatgen.util.string.latexify` and deprecate `latexify_ion` rather than have a standalone function to recognize charged formulae. Thoughts?",
"Go ahead and integrate with pymatgen.util.string latexify. Frankly, I don't know why the heck someone implemented a separate one elsewhere.",
"Actually, if we're proposing changes, I think I would prefer a `.latex_string()` option on Species, Composition, etc. -- then people don't have to find or hunt for the appropriate util function.",
"Please don't remove the pymatgen.util.string function though.",
"Pls do not add a method to every single class. That would be a horrendous way to do this. I think we should use a MixIn class that implements to_latex_string that works for most use cases,eg some adaptation of current latexify. ",
"Eg MSONable is the classic example of a mixin. ",
"Why would it be horrendous? We have `__str__` representations on a class, this is a variant of `__str__`",
"We're not talking about a lot of classes here, there's `Composition`, `Species`, `SpaceGroup`, `PointGroup` maybe.",
"There are classes you haven't thought of. Ion. Defects. There are also alternative conversions that would be extremely useful. Eg to_html_string. Even if it is 3-4 classes, having ONE implementation rather than the same code replicated in multiple locations. ",
"> Even if it is 3-4 classes, having ONE implementation rather than the same code replicated in multiple locations.\r\n\r\nYes, this is fair -- I was presuming that it wouldn't be a common implementation, e.g. Species has to deal with superscripts, SpaceGroup handles minus signs as overbars etc. I was indeed neglecting Ion, Defects however. (Edit: actually, `Ion` subclasses `Composition`, so that would be fine)",
"I have added a basic implementation in https://github.com/materialsproject/pymatgen/commit/0d38d3f9e86ad3dfb498d2db28f3f28c253bf1c8 .\r\nThis works on both Specie and compositions.",
"> I have added a basic implementation in [0d38d3f](https://github.com/materialsproject/pymatgen/commit/0d38d3f9e86ad3dfb498d2db28f3f28c253bf1c8) .\r\n> This works on both Specie and compositions.\r\n\r\n@shyuep thanks. This implementation requires one to choose EITHER subscripts or superscripts though, which I don't think is necessary. We can have a single method that will, for example, convert `B(OH)4-` to `B(OH)$_4^-$`. This can be accomplished with a combination of the regex pattern in the existing `latexify` and the one in my post up above.\r\n\r\nThe only caveat is that the charge of multivalent species must be written _after_ the + or - sign (e.g., `Fe+2` not `Fe2+`, which is something we can specify in the docstring. I think this is a good practice to enforce anyway as it prevents ambiguity. What do you think?",
"> The only caveat is that the charge of multivalent species must be written after the + or - sign (e.g., Fe+2 not Fe2+, which is something we can specify in the docstring. I think this is a good practice to enforce anyway as it prevents ambiguity. What do you think?\r\n\r\nThe problem with this is that `latexify` needs to work on, say, the output of a `str(Species)` which would do `Fe2+`, likewise `Species.from_str` has to support this. I'm not sure if this notation is strictly conventional but it's definitely common.",
"> > The only caveat is that the charge of multivalent species must be written after the + or - sign (e.g., Fe+2 not Fe2+, which is something we can specify in the docstring. I think this is a good practice to enforce anyway as it prevents ambiguity. What do you think?\r\n> \r\n> The problem with this is that `latexify` needs to work on, say, the output of a `str(Species)` which would do `Fe2+`, likewise `Species.from_str` has to support this. I'm not sure if this notation is strictly conventional but it's definitely common.\r\n\r\nOK that's a good point. Can / should we consider changing the output of `str(Species)` then? I feel that notation is outdated and ambiguous but I don't know what's common practice in every field.",
"@rkingsbury @mkhorton This is an initial attempt. Feel free to modify if you think there is a better way. The general idea with the mix-in is that you can always override one method. E.g., let's say we want to deal with space groups and it is not clear that a single regex can work for Species, Composition and Spacegroups. So for Spacegroups, we can implement a special to_latex method. It will still inherit the mix-in class because the latex->HTML is much easier and well-defined than basic string -> latex. I am sure there are other markup languages we can think of, e.g., markdown.\r\n\r\nThere is also some code for unicode versions, which are helpful. I think those can be added relatively easily though they don't work in all instances.",
"As for from_str, I don't think there is a standard definition. I would suggest we follow the convention that the `__repr__` is usually the more complete string output and stipulate that from_str should work with the output from `repr(obj)`. `__str__` is usually the more user-friendly output that may not include all information necessary to reconstitute the class.",
"One other option: we define in Stringify a \"to_pretty_string\" method that defaults to `__str__`. All other methods then works on the to_pretty_string output. That will allow us to decouple the actual output of `__str__` and a pretty output when we want to.",
"@rkingsbury @mkhorton I have a mostly working implementation at https://github.com/materialsproject/pymatgen/commit/8455650c5a40a8a5c69f79df015ab0e9580a6429 .\r\n\r\nNow a general method implements latex, html and unicode strings and a `to_pretty_string` is the input (defaulting to just `__str__`).\r\n\r\nI am going to just leave this for now. You can make changes if you desire, including fixing the Ion latex string. ",
"Great, thanks @shyuep!"
] | 2021-02-05T00:42:28
| 2021-02-06T21:18:08
|
2021-02-06T21:18:08Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
The `latexify_ion` method in `analysis.pourbaix_diagram` does not work; it just returns the input formula.
**To Reproduce**
```
from pymatgen.analysis.pourbaix_diagram import latexify_ion
latexify_ion('Na+')
'Na+'
latexify_ion('SO4-2')
'SO4-2'
```
**Expected behavior**
The formula would be reformatted with appropriate LateX sub/superscripts
**Desktop (please complete the following information):**
- OS: Mac
- Version: latest Github master as of Feb 3, 2021
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2056/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2056/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2057
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2057/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2057/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2057/events
|
https://github.com/materialsproject/pymatgen/pull/2057
| 802,157,660
|
MDExOlB1bGxSZXF1ZXN0NTY4MzQ4MTEy
| 2,057
|
Determine local environments based on bonding analysis
|
{
"login": "JaGeo",
"id": 22094846,
"node_id": "MDQ6VXNlcjIyMDk0ODQ2",
"avatar_url": "https://avatars.githubusercontent.com/u/22094846?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/JaGeo",
"html_url": "https://github.com/JaGeo",
"followers_url": "https://api.github.com/users/JaGeo/followers",
"following_url": "https://api.github.com/users/JaGeo/following{/other_user}",
"gists_url": "https://api.github.com/users/JaGeo/gists{/gist_id}",
"starred_url": "https://api.github.com/users/JaGeo/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/JaGeo/subscriptions",
"organizations_url": "https://api.github.com/users/JaGeo/orgs",
"repos_url": "https://api.github.com/users/JaGeo/repos",
"events_url": "https://api.github.com/users/JaGeo/events{/privacy}",
"received_events_url": "https://api.github.com/users/JaGeo/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I would like to have a better organization of all the local environment analysis packages, including local_env and chemenv. Can we create a parent package called pymatgen.analysis.local_environment to house all of them. I would also prefer some common API that unifies all the different kinds of analysis and the various options. Right now, the packages are so big, fragmented and difficult to find and use.",
"I am happy to move my code to another place. \r\n\r\nI am not sure regarding the common interface and moving the localenv and chemenv code (it will probably break a lot of existing code and documentation in publications and so on) but there are other people that might be able to comment better on this. ",
"Well, all this is open for discussion. There is no need to create breaking changes. We can always retain existing code where they are with deprecation messages. @mkhorton what do you think?",
"The main problem is that `chemenv` and `local_env` are developed by different people/teams and have different philosophies.\r\n\r\nThe code @JaGeo is proposing is related to neither, but based on the Lobster code.\r\n\r\nThere has been some attempt at having a common interface, namely the `NearNeighbors` class (in local_env) is an interface to define neighbors and `StructureGraph` in `pymatgen.analysis.graphs` is a container to store the resulting graph. I think both of these are good.\r\n\r\n@JaGeo here is using `NearNeighbors` here, so is conformant with that interface, so I think this is a good PR. I think it should be in its own file too, due to its length, but perhaps `io.lobster` is the better location since this is so closely coupled to the output of the Lobster code.\r\n\r\nChemenv (unrelated to this PR) pre-dates the `NearNeighbors` class, but it would be nice to get a `NearNeighbors` interface to Chemenv as well. It is too large a module to merge with `local_env` however, but a better page on all our local environment algorithms would be good in the docs to explain that we do provide multiple algorithms, and to give the relevant publications for each.",
"I am totally fine with moving the code to `io.lobster`. \r\nBuilding some interface to ChemEnv via the `NearNeighbors` class also seems to be feasible (but I would expect that it would only be able to use a limited number of the capabilities of ChemEnv). ",
"> Building some interface to ChemEnv via the NearNeighbors class also seems to be feasible (but I would expect that it would only be able to use a limited number of the capabilities of ChemEnv).\r\n\r\nYes, perhaps it depends on what kwargs are available for `ChemEnvNeighbors(NearNeighbors)`, I hope it may be possible to include the advanced functionality too. But ChemEnv is outside the scope of this PR so let's not worry about it here.",
"\n[](https://coveralls.io/builds/37447438)\n\nCoverage decreased (-0.4%) to 79.521% when pulling **c13852b068c8e460d7e9446488594d9f682faa16 on JaGeo:master** into **754c2593f069b38b8188b03d064dc52a674d55cb on materialsproject:master**.\n",
"I am now quite happy with the current state of the code. I would be happy if this could be merged for now. (I will probably extend the code within the next months)\r\nI am still having problems with some of the tests though and I am not completely sure what is going on. At least all tests on ubuntu work.",
"I know what the issue with the mac/win tests is. I have made changes. I have merged this PR. Thanks.",
"Thank you. "
] | 2021-02-05T13:18:18
| 2021-02-26T08:20:37
|
2021-02-25T16:26:36Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This is work in progress to add an analysis of the local environments based on bonding analysis. The classes will allow to both use localenv and chemenv in combination with Lobster output. It will also provide additional methods to analyse the ICOHPs/COHPs for sites with neighbor atoms.
@mkhorton , let me know if you already have comments.
I hope it is okay to put it in pymatgen.analysis.lobster_env
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2057/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2057/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2057",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2057",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2057.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2057.patch",
"merged_at": "2021-02-25T16:26:36Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2058
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2058/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2058/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2058/events
|
https://github.com/materialsproject/pymatgen/pull/2058
| 802,393,101
|
MDExOlB1bGxSZXF1ZXN0NTY4NTQzODY5
| 2,058
|
Fix bug in GibbsComputedStructureEntry, add test
|
{
"login": "mattmcdermott",
"id": 40469694,
"node_id": "MDQ6VXNlcjQwNDY5Njk0",
"avatar_url": "https://avatars.githubusercontent.com/u/40469694?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mattmcdermott",
"html_url": "https://github.com/mattmcdermott",
"followers_url": "https://api.github.com/users/mattmcdermott/followers",
"following_url": "https://api.github.com/users/mattmcdermott/following{/other_user}",
"gists_url": "https://api.github.com/users/mattmcdermott/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mattmcdermott/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mattmcdermott/subscriptions",
"organizations_url": "https://api.github.com/users/mattmcdermott/orgs",
"repos_url": "https://api.github.com/users/mattmcdermott/repos",
"events_url": "https://api.github.com/users/mattmcdermott/events{/privacy}",
"received_events_url": "https://api.github.com/users/mattmcdermott/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @mattmcdermott "
] | 2021-02-05T18:46:42
| 2021-02-05T19:27:15
|
2021-02-05T19:17:49Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Addresses bug in #2053, where experimentally-derived Gibbs free energies (for gases like CO2) were being added incorrectly to DFT energies. Adds a test to check that energy of CO2 is correct, even after normalization of entry.
Note: a future PR should address the slow equality checking, as well as the additional computational delay from having to do the Gibbs computation each time.
## Checklist
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2058/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2058/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2058",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2058",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2058.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2058.patch",
"merged_at": "2021-02-05T19:17:49Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2059
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2059/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2059/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2059/events
|
https://github.com/materialsproject/pymatgen/pull/2059
| 802,578,266
|
MDExOlB1bGxSZXF1ZXN0NTY4Njk2NjU3
| 2,059
|
Remove custom entry normalize methods, solve via optional composition
|
{
"login": "mattmcdermott",
"id": 40469694,
"node_id": "MDQ6VXNlcjQwNDY5Njk0",
"avatar_url": "https://avatars.githubusercontent.com/u/40469694?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mattmcdermott",
"html_url": "https://github.com/mattmcdermott",
"followers_url": "https://api.github.com/users/mattmcdermott/followers",
"following_url": "https://api.github.com/users/mattmcdermott/following{/other_user}",
"gists_url": "https://api.github.com/users/mattmcdermott/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mattmcdermott/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mattmcdermott/subscriptions",
"organizations_url": "https://api.github.com/users/mattmcdermott/orgs",
"repos_url": "https://api.github.com/users/mattmcdermott/repos",
"events_url": "https://api.github.com/users/mattmcdermott/events{/privacy}",
"received_events_url": "https://api.github.com/users/mattmcdermott/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-02-06T01:43:27
| 2021-02-06T02:22:26
|
2021-02-06T02:22:26Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
For **entry-equality branch**, to be reviewed by @mkhorton:
Solves the normalization issue by adding an optional composition parameter so that composition is not taken directly from the structure object. Also cleaned up a ton of other things!
This doesn't yet do the major entry refactor.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2059/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2059/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2059",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2059",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2059.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2059.patch",
"merged_at": "2021-02-06T02:22:26Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2060
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2060/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2060/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2060/events
|
https://github.com/materialsproject/pymatgen/pull/2060
| 803,747,604
|
MDExOlB1bGxSZXF1ZXN0NTY5NjIzOTc3
| 2,060
|
Some improvements to `Entry`
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-02-08T16:59:34
| 2021-02-13T17:29:23
|
2021-02-08T17:07:26Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Makes `Entry` immutable, this vastly simplifies debugging
* `Entry.normalize()` always returns a new Entry
* Issues in `GibbsComputedStructureEntry` resolved
Thanks to @mattmcdermott for work on this and @CompRhys for discussion.
## TODO (if any)
* Introduce a new `.normalized_energy()` `.normalized_composition()` on Entry, to allow retrieving these values without having to create a new object
* Remove `ComputedStructureEntry.normalize()` (the fundamental discrepancy between the un-normalized Structure and normalized Composition causes bugs)
* Ensure `EnergyAdjustment` similarly immutable
* Look at serialization issues for `Entry` subclasses, some custom serialization is over-complex
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [ ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [ ] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2060/reactions",
"total_count": 2,
"+1": 2,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2060/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2060",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2060",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2060.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2060.patch",
"merged_at": "2021-02-08T17:07:26Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2061
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2061/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2061/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2061/events
|
https://github.com/materialsproject/pymatgen/pull/2061
| 804,034,195
|
MDExOlB1bGxSZXF1ZXN0NTY5ODcxMzQ0
| 2,061
|
Add tqdm as requirement
|
{
"login": "jan-janssen",
"id": 3854739,
"node_id": "MDQ6VXNlcjM4NTQ3Mzk=",
"avatar_url": "https://avatars.githubusercontent.com/u/3854739?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jan-janssen",
"html_url": "https://github.com/jan-janssen",
"followers_url": "https://api.github.com/users/jan-janssen/followers",
"following_url": "https://api.github.com/users/jan-janssen/following{/other_user}",
"gists_url": "https://api.github.com/users/jan-janssen/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jan-janssen/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jan-janssen/subscriptions",
"organizations_url": "https://api.github.com/users/jan-janssen/orgs",
"repos_url": "https://api.github.com/users/jan-janssen/repos",
"events_url": "https://api.github.com/users/jan-janssen/events{/privacy}",
"received_events_url": "https://api.github.com/users/jan-janssen/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks for bringing this to our attention @jan-janssen, I'll close this PR since I fixed it via https://github.com/materialsproject/pymatgen/commit/5126b5bc73173b911acb7d4c20a65655d2a9fe1c instead.\r\n\r\n`tqdm` should be an optional dependency, we have a wrapper called `PBar` which will fall back gracefully if tqdm is not available."
] | 2021-02-08T22:50:56
| 2021-02-08T23:46:13
|
2021-02-08T23:46:12Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Because it is used in pymatgen.analysis.pourbaix_diagram
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2061/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2061/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2061",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2061",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2061.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2061.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2062
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2062/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2062/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2062/events
|
https://github.com/materialsproject/pymatgen/pull/2062
| 805,631,712
|
MDExOlB1bGxSZXF1ZXN0NTcxMjAxMDI0
| 2,062
|
Fix KeyError when initializing XRDCalculator with integer wavelength
|
{
"login": "flaviu-gostin",
"id": 37553536,
"node_id": "MDQ6VXNlcjM3NTUzNTM2",
"avatar_url": "https://avatars.githubusercontent.com/u/37553536?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/flaviu-gostin",
"html_url": "https://github.com/flaviu-gostin",
"followers_url": "https://api.github.com/users/flaviu-gostin/followers",
"following_url": "https://api.github.com/users/flaviu-gostin/following{/other_user}",
"gists_url": "https://api.github.com/users/flaviu-gostin/gists{/gist_id}",
"starred_url": "https://api.github.com/users/flaviu-gostin/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/flaviu-gostin/subscriptions",
"organizations_url": "https://api.github.com/users/flaviu-gostin/orgs",
"repos_url": "https://api.github.com/users/flaviu-gostin/repos",
"events_url": "https://api.github.com/users/flaviu-gostin/events{/privacy}",
"received_events_url": "https://api.github.com/users/flaviu-gostin/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks. Can you merge main master into your branch pls? I believe that's why your linting failed.",
"> Thanks. Can you merge main master into your branch pls? I believe that's why your linting failed.\r\n\r\nI just checked and it seems my master is based on the latest commit on the main master, so that wasn't the issue. I checked the Linting workflow on GitHub and it seems there were not enough blank lines in composition.py. Run black, pycodestyle and flake8 on that file. It should be ok now. I wonder how this file got through in the previous commit when the linting error was introduced (bf9880a).",
"Thanks!"
] | 2021-02-10T15:45:31
| 2021-02-11T16:24:39
|
2021-02-11T16:24:35Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
* Fix 1: attempting to create a XRDCalculator instance with an integer wavelength (e.g. 1, when wavelength is 1 Angstrom) throws a KeyError. Made changes to the __init__ method to accept integers for wavelength. Also raise TypeError if wavelength is neither of float, int or str. Added test to test if TypeError is raised.
Before a pull request can be merged, the following items must be checked:
- [x ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [ ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x ] Tests have been added for any new functionality or bug fixes.
- [x ] All linting and tests pass. --- Some linting does not pass. However, it was inherited. It is not due to this contribution.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2062/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2062/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2062",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2062",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2062.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2062.patch",
"merged_at": "2021-02-11T16:24:35Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2063
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2063/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2063/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2063/events
|
https://github.com/materialsproject/pymatgen/issues/2063
| 807,646,919
|
MDU6SXNzdWU4MDc2NDY5MTk=
| 2,063
|
pymatgen installation
|
{
"login": "frank-lin-liu",
"id": 33406238,
"node_id": "MDQ6VXNlcjMzNDA2MjM4",
"avatar_url": "https://avatars.githubusercontent.com/u/33406238?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/frank-lin-liu",
"html_url": "https://github.com/frank-lin-liu",
"followers_url": "https://api.github.com/users/frank-lin-liu/followers",
"following_url": "https://api.github.com/users/frank-lin-liu/following{/other_user}",
"gists_url": "https://api.github.com/users/frank-lin-liu/gists{/gist_id}",
"starred_url": "https://api.github.com/users/frank-lin-liu/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/frank-lin-liu/subscriptions",
"organizations_url": "https://api.github.com/users/frank-lin-liu/orgs",
"repos_url": "https://api.github.com/users/frank-lin-liu/repos",
"events_url": "https://api.github.com/users/frank-lin-liu/events{/privacy}",
"received_events_url": "https://api.github.com/users/frank-lin-liu/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Pls use the installation using conda as recommended in the pymatgen documentation."
] | 2021-02-13T01:32:33
| 2021-02-13T15:23:15
|
2021-02-13T15:23:15Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
When I install pymatgen in windows 10 using "python setup.py install", I have the following error,
error: command 'C:\\Program Files (x86)\\Microsoft Visual studio\\2019\\Community\\VC\\Tools\\MSVC\\14.28.29333\\bin\\HostX86\\x64\\cl.exe' failed with exit status 2
Please, could you help me out?
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2063/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2063/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2064
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2064/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2064/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2064/events
|
https://github.com/materialsproject/pymatgen/issues/2064
| 808,968,002
|
MDU6SXNzdWU4MDg5NjgwMDI=
| 2,064
|
cannot import pymatgen as mg
|
{
"login": "frank-lin-liu",
"id": 33406238,
"node_id": "MDQ6VXNlcjMzNDA2MjM4",
"avatar_url": "https://avatars.githubusercontent.com/u/33406238?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/frank-lin-liu",
"html_url": "https://github.com/frank-lin-liu",
"followers_url": "https://api.github.com/users/frank-lin-liu/followers",
"following_url": "https://api.github.com/users/frank-lin-liu/following{/other_user}",
"gists_url": "https://api.github.com/users/frank-lin-liu/gists{/gist_id}",
"starred_url": "https://api.github.com/users/frank-lin-liu/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/frank-lin-liu/subscriptions",
"organizations_url": "https://api.github.com/users/frank-lin-liu/orgs",
"repos_url": "https://api.github.com/users/frank-lin-liu/repos",
"events_url": "https://api.github.com/users/frank-lin-liu/events{/privacy}",
"received_events_url": "https://api.github.com/users/frank-lin-liu/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Pls try to install the latest pymatgen released yesterday. I believe it should solve this problem. It has to do with an incompatibility introduced in numpy 1.20.0 but solved in 1.20.1",
"Thank you. Pymatgen can be used now. "
] | 2021-02-16T04:13:36
| 2021-02-16T21:16:11
|
2021-02-16T21:16:11Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
operating system: Windows 10
I use the command "conda install --channel conda-forge pymatgen" to install pymatgen in Conda Prompt as described in https://pymatgen.org/installation.html. After installation, I run a python script using "python pymatgen_sample.py". In the pymatgen_sample.py, I import pymatgen using "import pymatgen as mg". But I have the following error,
_**Traceback (most recent call last):
File "pymatgen_sample.py", line 1, in <module>
import pymatgen as mg
File "C:\Users\frank\Anaconda3\lib\site-packages\pymatgen\__init__.py", line 56, in <module>
from .core.composition import Composition # noqa
File "C:\Users\frank\Anaconda3\lib\site-packages\pymatgen\core\__init__.py", line 11, in <module>
from .lattice import Lattice # noqa
File "C:\Users\frank\Anaconda3\lib\site-packages\pymatgen\core\lattice.py", line 23, in <module>
from pymatgen.util.coord import pbc_shortest_vectors
File "C:\Users\frank\Anaconda3\lib\site-packages\pymatgen\util\coord.py", line 17, in <module>
from . import coord_cython as cuc
File "pymatgen/util/coord_cython.pyx", line 1, in init pymatgen.util.coord_cython
ValueError: numpy.ndarray size changed, may indicate binary incompatibility. Expected 88 from C header, got 80 from PyObject_**
Could you please help me out?
|
{
"login": "frank-lin-liu",
"id": 33406238,
"node_id": "MDQ6VXNlcjMzNDA2MjM4",
"avatar_url": "https://avatars.githubusercontent.com/u/33406238?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/frank-lin-liu",
"html_url": "https://github.com/frank-lin-liu",
"followers_url": "https://api.github.com/users/frank-lin-liu/followers",
"following_url": "https://api.github.com/users/frank-lin-liu/following{/other_user}",
"gists_url": "https://api.github.com/users/frank-lin-liu/gists{/gist_id}",
"starred_url": "https://api.github.com/users/frank-lin-liu/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/frank-lin-liu/subscriptions",
"organizations_url": "https://api.github.com/users/frank-lin-liu/orgs",
"repos_url": "https://api.github.com/users/frank-lin-liu/repos",
"events_url": "https://api.github.com/users/frank-lin-liu/events{/privacy}",
"received_events_url": "https://api.github.com/users/frank-lin-liu/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2064/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2064/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2065
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2065/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2065/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2065/events
|
https://github.com/materialsproject/pymatgen/pull/2065
| 809,578,238
|
MDExOlB1bGxSZXF1ZXN0NTc0NDE3MTM0
| 2,065
|
Parse projected magnetisation from non-collinear calculations
|
{
"login": "utf",
"id": 1330638,
"node_id": "MDQ6VXNlcjEzMzA2Mzg=",
"avatar_url": "https://avatars.githubusercontent.com/u/1330638?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/utf",
"html_url": "https://github.com/utf",
"followers_url": "https://api.github.com/users/utf/followers",
"following_url": "https://api.github.com/users/utf/following{/other_user}",
"gists_url": "https://api.github.com/users/utf/gists{/gist_id}",
"starred_url": "https://api.github.com/users/utf/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/utf/subscriptions",
"organizations_url": "https://api.github.com/users/utf/orgs",
"repos_url": "https://api.github.com/users/utf/repos",
"events_url": "https://api.github.com/users/utf/events{/privacy}",
"received_events_url": "https://api.github.com/users/utf/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I've run black on the files, so I don't know why linting is failing saying \"Black would make changes\". It doesn't...",
"\n[](https://coveralls.io/builds/37179362)\n\nCoverage decreased (-0.7%) to 79.63% when pulling **a71607a151a3b790142ec28f85c9d34282dda280 on utf:parse-magnetisation** into **42f7c765330f609d4dbc1483e766b3f10bb606fb on materialsproject:master**.\n",
"@utf different set of black rules maybe? https://github.com/materialsproject/pymatgen/blob/1e36e835967fb8665a6ede344f4e608af50e93c2/pyproject.toml#L10\r\n\r\n",
"I can run black on outputs.py using a line length of 120 but that will cause lots of unrelated changes. Are you sure you'd like me to do that?",
"Just run black from the root level of pymatgen.",
"> Just run black from the root level of pymatgen.\r\n\r\nI did. There were no changes.",
"I have merged this for now\r\n",
"Thanks!",
"I ran black on my system on your pulled code and it definitely reformatted the file. Pls check your black version. It might be out of date.",
"I am using the latest version of black, so it can't be that. 🤷 \r\n\r\n```\r\n› black --version\r\nblack, version 20.8b1\r\n```",
"Also seeing this issue myself actually."
] | 2021-02-16T19:15:07
| 2021-02-17T01:47:57
|
2021-02-16T21:20:11Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Added the ability to parse the projected magnetisation from VASP calculations that include spin-orbit coupling or non-collinear magnetism.
For calculations that include orbital projections (`LORBIT = 11`), the projections are written to the vasprun.xml file in the same format as in the PROCAR file (https://www.vasp.at/wiki/index.php/PROCAR). The format of the projections depends on the other calculation parameters:
- For spin-paired calculations (`ISPIN = 1`): There is only one spin channel present.
- For spin-polarised calculations (`ISPIN = 2`): There are two spin channels present.
- For non-collinear or SOC calculations: There are 4 spin channels present.
In the last case, the first spin channel is the usual projection of the wavefunctions onto the atomic orbitals. The last 3 spin channels are actually the projected magnetisation of those orbitals in the 3 cartesian directions.
Previously, the projected eigenvalues from SOC calculations incorrectly had two spin channels, while the remaining spin channels were ignored. This PR fixes the parsing so that the projected magnetisation is extracted correctly for SOC calculations.
I've added a unit test.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2065/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2065/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2065",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2065",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2065.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2065.patch",
"merged_at": "2021-02-16T21:20:10Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2066
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2066/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2066/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2066/events
|
https://github.com/materialsproject/pymatgen/pull/2066
| 809,771,195
|
MDExOlB1bGxSZXF1ZXN0NTc0NTc2NzMy
| 2,066
|
Add convenience interface for OPTIMADE-compliant APIs
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/37188701)\n\nCoverage decreased (-0.1%) to 79.514% when pulling **8447c788d514233aa5c060f80392c2c0b863d80b on optimade** into **5c5628265429bf33d1181d58add13a77e405a919 on master**.\n"
] | 2021-02-17T01:14:49
| 2021-03-09T01:04:01
|
2021-02-17T02:34:15Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
An `OptimadeRester` in `pymatgen.ext.optimade`.
This is by no means the final version of this class, but it is functional and a starting point for future improvements.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2066/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2066/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2066",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2066",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2066.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2066.patch",
"merged_at": "2021-02-17T02:34:15Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2067
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2067/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2067/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2067/events
|
https://github.com/materialsproject/pymatgen/issues/2067
| 810,400,378
|
MDU6SXNzdWU4MTA0MDAzNzg=
| 2,067
|
Gcc 8.3 errors when compiling Zeo++
|
{
"login": "feacluster",
"id": 34378182,
"node_id": "MDQ6VXNlcjM0Mzc4MTgy",
"avatar_url": "https://avatars.githubusercontent.com/u/34378182?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/feacluster",
"html_url": "https://github.com/feacluster",
"followers_url": "https://api.github.com/users/feacluster/followers",
"following_url": "https://api.github.com/users/feacluster/following{/other_user}",
"gists_url": "https://api.github.com/users/feacluster/gists{/gist_id}",
"starred_url": "https://api.github.com/users/feacluster/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/feacluster/subscriptions",
"organizations_url": "https://api.github.com/users/feacluster/orgs",
"repos_url": "https://api.github.com/users/feacluster/repos",
"events_url": "https://api.github.com/users/feacluster/events{/privacy}",
"received_events_url": "https://api.github.com/users/feacluster/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I tried on another system with older GCC 4.8.5 but got the same errors. The errors are:\r\n\r\nzeo/netstorage.cpp: In function ‘PyObject* __Pyx_PyCFunction_FastCall(PyObject*, PyObject**, Py_ssize_t)’:\r\nzeo/netstorage.cpp:8776:69: error: too many arguments to function\r\n return (*((__Pyx_PyCFunctionFast)meth)) (self, args, nargs, NULL);\r\n ^\r\nzeo/netstorage.cpp:8766:15: warning: unused variable ‘result’ [-Wunused-variable]\r\n PyObject *result;\r\n ^\r\nzeo/netstorage.cpp:8767:9: warning: unused variable ‘flags’ [-Wunused-variable]\r\n int flags;\r\n ^\r\nzeo/netstorage.cpp: In function ‘void __Pyx__ExceptionSwap(PyThreadState*, PyObject**, PyObject**, PyObject**)’:\r\nzeo/netstorage.cpp:10133:24: error: ‘PyThreadState’ has no member named ‘exc_type’\r\n tmp_type = tstate->exc_type;\r\n ^\r\nzeo/netstorage.cpp:10134:25: error: ‘PyThreadState’ has no member named ‘exc_value’\r\n tmp_value = tstate->exc_value;\r\n ^\r\nzeo/netstorage.cpp:10135:22: error: ‘PyThreadState’ has no member named ‘exc_traceback’\r\n tmp_tb = tstate->exc_traceback;\r\n ^\r\nzeo/netstorage.cpp:10136:13: error: ‘PyThreadState’ has no member named ‘exc_type’\r\n tstate->exc_type = *type;\r\n ^\r\nzeo/netstorage.cpp:10137:13: error: ‘PyThreadState’ has no member named ‘exc_value’\r\n tstate->exc_value = *value;\r\n ^\r\nzeo/netstorage.cpp:10138:13: error: ‘PyThreadState’ has no member named ‘exc_traceback’\r\n tstate->exc_traceback = *tb;\r\n ^\r\n\r\n",
"We can't really help you on this since zeopp is not a pymatgen code per se. Pls contact the Zeo++ authors on its installation and usage."
] | 2021-02-17T17:45:57
| 2021-02-17T18:24:38
|
2021-02-17T18:24:38Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I am following your instructions here for the pymatgen.io.zeopp module:
https://pymatgen.org/pymatgen.io.zeopp.html#zeo-installation-steps
I get gcc errors when attempting step 5(c). The first error is:
zeo/netstorage.cpp: In function ‘PyObject* __Pyx_PyCFunction_FastCall(PyObject*,
zeo/netstorage.cpp:8776:69: error: too many arguments to function
return (*((__Pyx_PyCFunctionFast)meth)) (self, args, nargs, NULL);
This is then followed by a few other errors before the compilation fails.
I suspect it is because I am using the latest gcc 8.3.1
(VOID) [hussaif1@7920 cython_wrapper]$ gcc --version
gcc (GCC) 8.3.1 20191121 (Red Hat 8.3.1-5)
Or could it be something else?
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2067/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2067/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2068
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2068/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2068/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2068/events
|
https://github.com/materialsproject/pymatgen/issues/2068
| 811,048,761
|
MDU6SXNzdWU4MTEwNDg3NjE=
| 2,068
|
Outcar parsing for vasp 6.2.0 is broken
|
{
"login": "MichaelWolloch",
"id": 66372013,
"node_id": "MDQ6VXNlcjY2MzcyMDEz",
"avatar_url": "https://avatars.githubusercontent.com/u/66372013?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/MichaelWolloch",
"html_url": "https://github.com/MichaelWolloch",
"followers_url": "https://api.github.com/users/MichaelWolloch/followers",
"following_url": "https://api.github.com/users/MichaelWolloch/following{/other_user}",
"gists_url": "https://api.github.com/users/MichaelWolloch/gists{/gist_id}",
"starred_url": "https://api.github.com/users/MichaelWolloch/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/MichaelWolloch/subscriptions",
"organizations_url": "https://api.github.com/users/MichaelWolloch/orgs",
"repos_url": "https://api.github.com/users/MichaelWolloch/repos",
"events_url": "https://api.github.com/users/MichaelWolloch/events{/privacy}",
"received_events_url": "https://api.github.com/users/MichaelWolloch/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 4735709485,
"node_id": "LA_kwDOACgets8AAAABGkUxLQ",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/compatability",
"name": "compatability",
"color": "E25438",
"default": false,
"description": "Concerning pymatgen compatibility with different OS, Python versions, numpy versions, etc."
},
{
"id": 5304335471,
"node_id": "LA_kwDOACgets8AAAABPCm8bw",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/io",
"name": "io",
"color": "c2e0c6",
"default": false,
"description": "Input/output functionality"
},
{
"id": 5457739150,
"node_id": "LA_kwDOACgets8AAAABRU59jg",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/vasp",
"name": "vasp",
"color": "BF4B01",
"default": false,
"description": "Vienna Ab initio Simulation Package"
}
] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"> I can create a pull request that does exactly that if needed.\r\n\r\nThat would be great, thank you!\r\n\r\nI would modify it slightly:\r\n\r\n```\r\ntry:\r\n run_stats[tok[0].strip()] = float(tok[1].strip())\r\nexcept ValueError:\r\n run_stats[tok[0].strip()] = None\r\n```\r\n",
"Pull request was made and all checks have passed.",
"@mkhorton -- looks like this one can be closed.",
"@janosh: This can be closed."
] | 2021-02-18T12:19:43
| 2023-06-03T02:22:04
|
2023-06-03T02:22:04Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
I recently used VASP version 6.2.0 (newest official release afaik) and found that the Outcar parser of pymatgen (class `Outcar`, `__init__` method, in pymatgen.io.vasp.outputs.py) is not compatible with it.
The problem is a tiny change in the OUTCAR format concerning memory, especially the line where the average memory consumption is detailed. (I think VASP often actually does not know the average memory used…)
In 5.4.* and 6.1.* I get:
~~~
Average memory used (kb): 0.
~~~
In version 6.2.0 I get:
~~~
Average memory used (kb): N/A
~~~
**To Reproduce**
Steps to reproduce the behavior:
1. Run a small vasp calculation with version 6.2.0 and wait till it is completed (or use the OUTCAR from below that I created)
2. in the execution directory open ipython
3. from pymatgen.io.vasp.output import Outcar
4. outcar_parsed = Outcar('OUTCAR')
5. See `ValueError: could not convert string to float: 'N/A'`
Provide any example files that are needed to reproduce the error,
especially if the bug pertains to parsing of a file:
[OUTCAR.gz](https://github.com/materialsproject/pymatgen/files/6002640/OUTCAR.gz)
**Expected behavior**
No parsing error
**Additional context**
The problematic line is [1715](https://github.com/materialsproject/pymatgen/blob/27d766cae2a62bbf565b4dc28bf2781fd160ee50/pymatgen/io/vasp/outputs.py#L1715) in pymatgen.io.vasp.outputs.py:
~~~
run_stats[tok[0].strip()] = float(tok[1].strip())
~~~
I would simply replace this with the following try/except block:
~~~
try:
run_stats[tok[0].strip()] = float(tok[1].strip())
except:
run_stats[tok[0].strip()] = 0.0
~~~
I can create a pull request that does exactly that if needed.
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2068/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2068/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2069
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2069/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2069/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2069/events
|
https://github.com/materialsproject/pymatgen/pull/2069
| 811,892,866
|
MDExOlB1bGxSZXF1ZXN0NTc2MzQ4NzI3
| 2,069
|
Fix Outcar parsing for VASP 6.2.0
|
{
"login": "MichaelWolloch",
"id": 66372013,
"node_id": "MDQ6VXNlcjY2MzcyMDEz",
"avatar_url": "https://avatars.githubusercontent.com/u/66372013?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/MichaelWolloch",
"html_url": "https://github.com/MichaelWolloch",
"followers_url": "https://api.github.com/users/MichaelWolloch/followers",
"following_url": "https://api.github.com/users/MichaelWolloch/following{/other_user}",
"gists_url": "https://api.github.com/users/MichaelWolloch/gists{/gist_id}",
"starred_url": "https://api.github.com/users/MichaelWolloch/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/MichaelWolloch/subscriptions",
"organizations_url": "https://api.github.com/users/MichaelWolloch/orgs",
"repos_url": "https://api.github.com/users/MichaelWolloch/repos",
"events_url": "https://api.github.com/users/MichaelWolloch/events{/privacy}",
"received_events_url": "https://api.github.com/users/MichaelWolloch/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 5304335471,
"node_id": "LA_kwDOACgets8AAAABPCm8bw",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/io",
"name": "io",
"color": "c2e0c6",
"default": false,
"description": "Input/output functionality"
},
{
"id": 5318590534,
"node_id": "LA_kwDOACgets8AAAABPQNARg",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/fix",
"name": "fix",
"color": "40DE1F",
"default": false,
"description": "Bug fix PRs"
},
{
"id": 5457739150,
"node_id": "LA_kwDOACgets8AAAABRU59jg",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/vasp",
"name": "vasp",
"color": "BF4B01",
"default": false,
"description": "Vienna Ab initio Simulation Package"
}
] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/37280078)\n\nCoverage decreased (-0.7%) to 79.241% when pulling **f7c9e2553668ea2f07224c7220185a89ebcdca21 on MichaelWolloch:Fix_issue_#2068** into **c9a7e9810b539d13398219ff06e333682881184e on materialsproject:master**.\n",
"Pls add a simple unit test to ensure this bug does not recur. Thanks. ",
"I hope the test I added was what you had in mind @shyuep . I have not much experience writing tests...",
"Thanks @MichaelWolloch! That test works fine; simply having a VASP 6.2 OUTCAR parsed as part of the test suite will be useful.",
"Hi @MichaelWolloch, this has now been merged. Please do fill out our [developer attribution form](https://forms.gle/JnisFb38QDR8QTFTA) so we can [acknowledge you appropriately on our website](https://pymatgen.org/team.html), improvements/bug fixes to pymatgen are always appreciated."
] | 2021-02-19T10:16:55
| 2023-06-03T02:22:26
|
2021-02-19T18:29:54Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Close https://github.com/materialsproject/pymatgen/issues/2068 as [discussed](https://github.com/materialsproject/pymatgen/issues/2068#issuecomment-781645771) with @mkhorton.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2069/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2069/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2069",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2069",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2069.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2069.patch",
"merged_at": "2021-02-19T18:29:54Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2070
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2070/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2070/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2070/events
|
https://github.com/materialsproject/pymatgen/issues/2070
| 812,045,333
|
MDU6SXNzdWU4MTIwNDUzMzM=
| 2,070
|
Make CrystalNN work with fractional occupation
|
{
"login": "nisse3000",
"id": 11105218,
"node_id": "MDQ6VXNlcjExMTA1MjE4",
"avatar_url": "https://avatars.githubusercontent.com/u/11105218?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/nisse3000",
"html_url": "https://github.com/nisse3000",
"followers_url": "https://api.github.com/users/nisse3000/followers",
"following_url": "https://api.github.com/users/nisse3000/following{/other_user}",
"gists_url": "https://api.github.com/users/nisse3000/gists{/gist_id}",
"starred_url": "https://api.github.com/users/nisse3000/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/nisse3000/subscriptions",
"organizations_url": "https://api.github.com/users/nisse3000/orgs",
"repos_url": "https://api.github.com/users/nisse3000/repos",
"events_url": "https://api.github.com/users/nisse3000/events{/privacy}",
"received_events_url": "https://api.github.com/users/nisse3000/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"No. That is the wrong approach. The basic premise is that people should code for disordered structures, rather than assume that all structures will always be ordered. In this case, `specie` was the wrong attribute to use in the first place. Feel free to just change the code to work with disordered structures and submit a PR.\r\n\r\n@mkhorton Let's make sure when people submit PRs in the future that we reject anything that assumes a structure is ordered without good reason. The only cases where full occupancies should be enforced are when interfacing with computational codes that require ordered structures. All generic analysis must work with ordered and disordered structures.",
"There's a logical difficulty here, in that edges/bonds are defined with respect to sites, and basically all bonding algorithms require there to be a specific atom at each end of the bond. In disordered structures, there's going to be a fundamental ambiguity present.\r\n\r\nA general \"fix\" for this might be in the `NearNeighbors` class (i.e. not CrystalNN specifically) to calculate bonding only with the majority member of that site. This is still a compromise.\r\n\r\n@shyuep sure, I think enforcing tests with disordered structures where appropriate would be useful -- we shouldn't be seeing an AttributeError here, if the logic doesn't work for disordered structures we should simply be told that.",
"Stumbled into the above error too just now.\r\n\r\n> to calculate bonding only with the majority member of that site. This is still a compromise.\r\n\r\nWhat about using a weighted average?",
"> What about using a weighted average?\r\n\r\nI agree this should be an acceptable workaround. \r\n"
] | 2021-02-19T13:58:24
| 2022-08-30T21:21:30
|
2022-08-30T21:21:30Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
When the CrystalNN neighbor-finding method is applied to a structure with fractional occupation (in the test case from a CIF file), it exits with an attribute error:
File "pymatgen/pymatgen/analysis/local_env.py", line 4010, in get_cn
return super().get_cn(structure, n, use_weights)
File "pymatgen/pymatgen/analysis/local_env.py", line 265, in get_cn
siw = self.get_nn_info(structure, n)
File "pymatgen/pymatgen/analysis/local_env.py", line 3842, in get_nn_info
nndata = self.get_nn_data(structure, n)
File "pymatgen/pymatgen/analysis/local_env.py", line 3905, in get_nn_data
X1 = structure[n].specie.X
File "pymatgen/pymatgen/core/sites.py", line 80, in __getattr__
raise AttributeError(a)
AttributeError: specie
Removing the fractional occupation in the underlying CIF file results in no problems.
Maybe the fractional occupation should simply be ignored for determining the neighbors and coordination numbers. The only issue that remains is then when fractional site a close to each other so that they might be deemed coordinated when the question might be: is either site 1 or site 2 present, not both so that they cannot be coordinated.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2070/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2070/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2071
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2071/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2071/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2071/events
|
https://github.com/materialsproject/pymatgen/issues/2071
| 812,185,988
|
MDU6SXNzdWU4MTIxODU5ODg=
| 2,071
|
Installation problem in Google colab
|
{
"login": "Amadeus-System",
"id": 76824867,
"node_id": "MDQ6VXNlcjc2ODI0ODY3",
"avatar_url": "https://avatars.githubusercontent.com/u/76824867?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Amadeus-System",
"html_url": "https://github.com/Amadeus-System",
"followers_url": "https://api.github.com/users/Amadeus-System/followers",
"following_url": "https://api.github.com/users/Amadeus-System/following{/other_user}",
"gists_url": "https://api.github.com/users/Amadeus-System/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Amadeus-System/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Amadeus-System/subscriptions",
"organizations_url": "https://api.github.com/users/Amadeus-System/orgs",
"repos_url": "https://api.github.com/users/Amadeus-System/repos",
"events_url": "https://api.github.com/users/Amadeus-System/events{/privacy}",
"received_events_url": "https://api.github.com/users/Amadeus-System/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"The issue is that Google won't let you install numpy 1.20.1 (try `!pip install numpy==1.20.1` to verify), I think because it's not supported by Python 3.6 (the default interpreter). For now, I would advice you install an older version of pymatgen e.g. using `pip install pymatgen==` until Google supports this.",
"If google collab allows you to install conda, you can do that instead. \r\n"
] | 2021-02-19T17:03:27
| 2021-02-19T20:17:15
|
2021-02-19T20:17:15Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hi. I was using pymatgen in google colab.
In google colab, pymatgen is not installed , so I was using pymatgen by installing every time when I need to use it.
"!pip3 install pymatgen" or "!pip install pymatgen" command was normally working a few days ago, but now it's not working so I can't use pymatgen in colab.
I need your help. please teach me how to install pymatgen in google colab.
I think it's because some sort of problem happend in pymatgen library.
The following is my capture image for the error and message.
"""
Collecting pymatgen
Downloading https://files.pythonhosted.org/packages/9b/50/a22eb063ca5ceb27c2be7e51e775ee4b5ade664525be143a2b1b77e4cc70/pymatgen-2021.2.16.tar.gz (3.0MB)
|████████████████████████████████| 3.0MB 6.4MB/s
Installing build dependencies ... error
ERROR: Command errored out with exit status 1: /usr/bin/python3 /usr/local/lib/python3.6/dist-packages/pip install --ignore-installed --no-user --prefix /tmp/pip-build-env-u3bnrkjk/overlay --no-warn-script-location -v --no-binary :none: --only-binary :none: -i https://pypi.org/simple -- 'numpy>=1.20.1' 'setuptools>=43.0.0' Check the logs for full command output.
"""
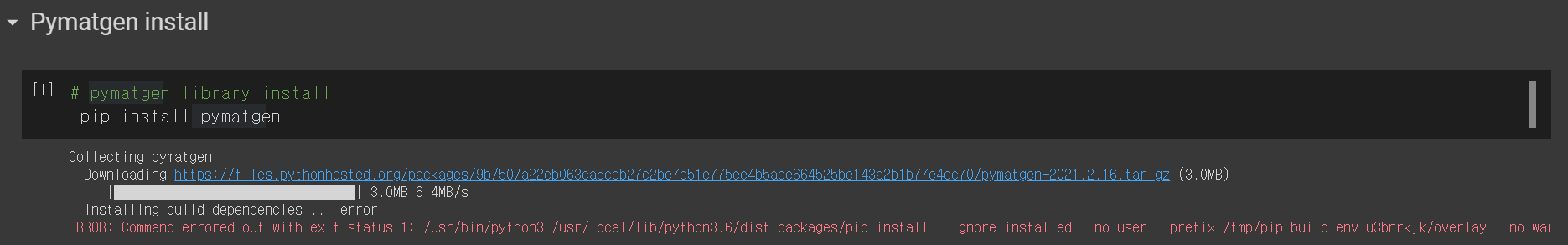
I expect your kind answer.
Thank you.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2071/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2071/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2072
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2072/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2072/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2072/events
|
https://github.com/materialsproject/pymatgen/pull/2072
| 812,859,023
|
MDExOlB1bGxSZXF1ZXN0NTc3MTE0MDM1
| 2,072
|
Gaussian: allow None for functional, bset, charge and multiplicity
|
{
"login": "eimrek",
"id": 8721264,
"node_id": "MDQ6VXNlcjg3MjEyNjQ=",
"avatar_url": "https://avatars.githubusercontent.com/u/8721264?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/eimrek",
"html_url": "https://github.com/eimrek",
"followers_url": "https://api.github.com/users/eimrek/followers",
"following_url": "https://api.github.com/users/eimrek/following{/other_user}",
"gists_url": "https://api.github.com/users/eimrek/gists{/gist_id}",
"starred_url": "https://api.github.com/users/eimrek/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/eimrek/subscriptions",
"organizations_url": "https://api.github.com/users/eimrek/orgs",
"repos_url": "https://api.github.com/users/eimrek/repos",
"events_url": "https://api.github.com/users/eimrek/events{/privacy}",
"received_events_url": "https://api.github.com/users/eimrek/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/37304418)\n\nCoverage decreased (-0.7%) to 79.244% when pulling **9f9ab99653cb6b5e9c86530460ca557d5077b640 on eimrek:master** into **19d7f56ba07850ce7cef743cc60272057a289c43 on materialsproject:master**.\n",
"Thanks!"
] | 2021-02-21T13:44:00
| 2021-02-21T23:41:33
|
2021-02-21T23:41:31Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hi pymatgen community!
I would like to use the Gaussian input generator for post-processing runs, such as this one
~~~
%chk=g.chk
%oldchk=../g.chk
#P chkbasis geom=allcheck guess=(only,read) pop=naturalorbital
test
~~~
This would read the geometry, functional, basis set, charge and multiplicity from `../g.chk` and perform natural orbital analysis.
This PR enables this by making
* functional and basis set are not outputted when set to `None`
* if geometry is `None`, then charge or multiplicity set to `None` will not be displayed
I also added a test for this and I had to modify an existing test.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2072/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2072/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2072",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2072",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2072.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2072.patch",
"merged_at": "2021-02-21T23:41:31Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2073
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2073/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2073/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2073/events
|
https://github.com/materialsproject/pymatgen/issues/2073
| 813,886,152
|
MDU6SXNzdWU4MTM4ODYxNTI=
| 2,073
|
pymatgen.analysis.defects.point_defects doesn't exist
|
{
"login": "yurivict",
"id": 271906,
"node_id": "MDQ6VXNlcjI3MTkwNg==",
"avatar_url": "https://avatars.githubusercontent.com/u/271906?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/yurivict",
"html_url": "https://github.com/yurivict",
"followers_url": "https://api.github.com/users/yurivict/followers",
"following_url": "https://api.github.com/users/yurivict/following{/other_user}",
"gists_url": "https://api.github.com/users/yurivict/gists{/gist_id}",
"starred_url": "https://api.github.com/users/yurivict/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/yurivict/subscriptions",
"organizations_url": "https://api.github.com/users/yurivict/orgs",
"repos_url": "https://api.github.com/users/yurivict/repos",
"events_url": "https://api.github.com/users/yurivict/events{/privacy}",
"received_events_url": "https://api.github.com/users/yurivict/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"It's because you copied it with an extra space at the start :-) This is generally true wherever you copy and paste Python code, be careful of leading tabs or spaces. I would recommend reading some general Python tutorials before getting started with pymatgen specifically.\r\n\r\nThat example is out of date for a separate reason however, it should now read:\r\n\r\n```\r\nfrom pymatgen.analysis.defects.core import Interstitial\r\n```\r\n\r\nYou can check the API docs for more information on specific classes, such as this page for the [`Interstitial`](https://pymatgen.org/pymatgen.analysis.defects.core.html#pymatgen.analysis.defects.core.Interstitial) object."
] | 2021-02-22T21:38:31
| 2021-02-22T21:46:45
|
2021-02-22T21:46:45Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
This command from the manual @ https://pymatgen.org/installation.html fails:
```
$ python3.7
Python 3.7.9 (default, Dec 12 2020, 01:25:17)
[Clang 8.0.1 (tags/RELEASE_801/final 366581)] on freebsd12
Type "help", "copyright", "credits" or "license" for more information.
>>> from pymatgen.analysis.defects.point_defects import Interstitial
File "<stdin>", line 1
from pymatgen.analysis.defects.point_defects import Interstitial
^
IndentationError: unexpected indent
>>>
```
Version 2021.2.16
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2073/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2073/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2074
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2074/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2074/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2074/events
|
https://github.com/materialsproject/pymatgen/pull/2074
| 815,418,754
|
MDExOlB1bGxSZXF1ZXN0NTc5MjUxNzQ3
| 2,074
|
More Informative Message with PhaseDiagramError
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/37416969)\n\nCoverage decreased (-0.7%) to 79.244% when pulling **9b83131e07eb95e6d049e82949e805dcd212199e on CompRhys:pderror** into **47c2bef6d01c6d2fd53e950931b923b94bf6f175 on materialsproject:master**.\n",
"Please fix the linting error. Also, I suggest you change the error to ValueError instead of PhaseDiagramError. I am planning to remove custom errors in favor of the common ValueError, RuntimeError and KeyError unless there is a good reason to create a new error class.",
"I have removed PhaseDiagramError and update the tests.\r\n\r\nI always get hit by the linter even when I auto-format the code... have you considered something like https://pre-commit.ci/ to autoformat as an action? It might stop most of the lining issues that are seen?",
"There is already a pre-commit hook provided. But I prefer not to make it autoformat since the coder might have other ideas. ",
"Thanks."
] | 2021-02-24T12:10:03
| 2021-02-24T17:34:29
|
2021-02-24T16:31:17Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
update `PhaseDiagramError` message to tell users which elements are missing.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2074/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2074/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2074",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2074",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2074.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2074.patch",
"merged_at": "2021-02-24T16:31:17Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2075
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2075/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2075/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2075/events
|
https://github.com/materialsproject/pymatgen/pull/2075
| 815,860,095
|
MDExOlB1bGxSZXF1ZXN0NTc5NjE5NzQ0
| 2,075
|
[WIP] reconfigure `get_quasi_e_to_hull` when polymorphs are present
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/37587648)\n\nCoverage decreased (-0.6%) to 79.523% when pulling **d038832f61d0e70dda4a8953f49777e3124e9f7c on CompRhys:decomp** into **877715f11f72457bafca6346b0378c5cfc7d1131 on materialsproject:master**.\n",
"@mkhorton I think this can be reviewed/merged. I added some notes to the code to explain the small changes that may seem unrelated to the PR title. ",
"Thanks @CompRhys! I'm happy to merge this.\r\n\r\nOne note regarding the method re-name: given that it's a recently added method and the re-name makes sense, I'm ok to do it here, but as a rule I'd prefer not to merge in changes that rename methods so that we don't cause frustration for users when methods go missing. For future PRs, if renaming does make sense, one option is just to add a stub for the old method name to point to the new method name and give a DeprecationWarning."
] | 2021-02-24T21:24:10
| 2021-03-02T20:01:26
|
2021-03-02T19:06:16Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Currently for the stable entry in the toy hull shown below in the presence of a polymorph (blue filled circle) the current code returns the red energy from `get_quasi_e_to_hull` whilst the intended behaviour should be that it gives the orange energy. I have checked this with @CJBartel who agrees that the orange energy is the correct energy. This PR should now return the orange energy.
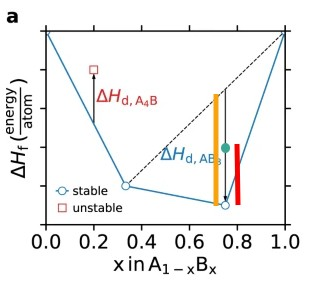
## TODO (if any)
1. Double check edge cases
2. Maybe rename the function to have a more informative name `get_phase_separation_energy` may be appropriate
1. Some materials are not directly tested as the only way to get a test would rely on calling the method being tested for a different material
## Checklist
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2075/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2075/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2075",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2075",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2075.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2075.patch",
"merged_at": "2021-03-02T19:06:16Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2076
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2076/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2076/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2076/events
|
https://github.com/materialsproject/pymatgen/issues/2076
| 816,513,050
|
MDU6SXNzdWU4MTY1MTMwNTA=
| 2,076
|
Adding a more discriminative hash for Composition
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"A good hash function has two qualities:\r\n1) It's fast\r\n2) It hashes very similar but different things to completely different hashes. The more granual that definition of similar is, the better the hash function.\r\n\r\nYour 3rd example is a good test to implement IMHO. Something like:\r\n``` python\r\nassert hash(Composition(f\"O{0.8}Fe{0.2}\")) == hash(Composition(f\"O{0.8}Fe{0.2}Na{1e-9}\"))\r\nassert hash(Composition(f\"O{0.8}Fe{0.2}\")) != hash(Composition(f\"O{0.8}Fe{0.2+Composition.amount_tolerance}\"))\r\n```\r\n\r\nWhy not just do something like\r\n```python\r\ndef __hash__(self)\r\n return hash(self.formula)\r\n```",
"I think without the manipulations the formula alone isn't robust, as a counter-example: \r\n\r\n```python\r\n>>> Composition(f\"O{0.2+Composition.amount_tolerance}Fe{0.8}\") == Composition(f\"O{0.2}Fe{0.8}\")\r\nTrue\r\n>>> hash(Composition(f\"O{0.2+Composition.amount_tolerance}Fe{0.8}\").formula) == hash(Composition(f\"O{0.2}Fe{0.8}\").formula)\r\nFalse\r\n>>> hash(Composition(f\"O{0.2+Composition.amount_tolerance}Fe{0.8}\")) == hash(Composition(f\"O{0.2}Fe{0.8}\"))\r\nTrue\r\n```",
"Ah! Ok, I didn't realize that those two compositions should actually be equal. Then, this looks mostly good. The rounding logic is a bit confusing, so you might want some comments on why you chose the numbers you did. For instance, I don't understand why 1e7 is the right multiplier. It should probably something like divide by tolerance since that could change.\r\n\r\n You might drop the switch to a string and even the `get_el_amt_dict` and use something like:\r\n```python\r\n\r\n def __hash__(self):\r\n \"\"\"\r\n Minimally effective hash function that just distinguishes between\r\n Compositions with different elements.\r\n \"\"\"\r\n sym_amt = {\r\n el: floor(1e7*(amt + Composition.amount_tolerance * 2))\r\n for el, amt in self.items()\r\n if abs(amt) > Composition.amount_tolerance\r\n }\r\n\r\n return hash(frozenset(sym_amt.items()))\r\n```\r\n\r\nEdit: removed unnecessary lines in the code.",
"Using `frozenset` is faster and I split out the 1e7 coefficient into a divide by `tol` and multiply by 10. The 2 is to deal with the case where the delta is equal to the tol, anything larger than 1 would do so I picked 2 because it only takes one character...\r\n\r\n```python\r\n def __hash__(self):\r\n \"\"\"\r\n Minimally effective hash function that just distinguishes between\r\n Compositions with different elements.\r\n \"\"\"\r\n sym_amt = {\r\n el: floor((amt + Composition.amount_tolerance * 2) / (Composition.amount_tolerance * 10))\r\n for el, amt in self.items()\r\n if abs(amt) > Composition.amount_tolerance\r\n }\r\n\r\n return hash(frozenset(sym_amt.items()))\r\n```\r\n```\r\n1 loop, best of 5: 547 msec per loop\r\n```",
"Yeah! I like it. As long as this passes all existing tests, I think it's a good modification. The docstring will need an update to be clear it now distinguishes the full composition, but that can be in the PR :)",
"I don't think something this complicated ius necessary. The problem with the earlier hash is mainly because it just sums the Z. While that is somewhat ok, there are many ways to arrive at the same Z. We just need to use something else. What if you just change `el.Z` to `el.Z * el.X`? My suspicion is that you will get most of the speed savings.",
"Can't use `el.X` due to noble gases but trying `int(el.Z * el.mendeleev_no)` took\r\n```1 loop, best of 5: 1.01 sec per loop```\r\nwithout wrapping in int I got an error",
"Cool. That is already just 2x of the solution with the amounts. Will this be sufficient? I am loath to introduce things like floating point comparisons, floors, etc. It is a lot more prone to errors in edge cases.",
"Why not just `hash({frozentset(self.elements)})` then?\r\n\r\nAlso, the tests on the larger data set of 400K+ might be a more useful comparison here. ",
"on the larger set `hash(frozenset(self.elements))` gave \r\n```\r\n1 loop, best of 5: 21.1 sec per loop (365msec on small)\r\n```\r\n\r\nwhilst the one with the amounts was\r\n```\r\n1 loop, best of 5: 14.9 sec per loop (616msec on small)\r\n```\r\n\r\nas such `hash(frozenset(self.elements))` is performant enough to be useable at the current size of the screening set if I end up screening millions of things it might be slow but in that case I have something workable",
"NIce. Let's implement that then. ",
"BTW, you can probably get a small bit of performance boost from using `self._data.keys()` instead of `self.elements`. The former uses a built in attribute.",
"okay making PR and adding a small test for robustness"
] | 2021-02-25T14:55:50
| 2021-02-25T23:14:12
|
2021-02-25T23:14:12Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
For a project I am doing I have produced a large screening library of compositions, to reduce the computation I need to do at the next stage I want only the unique compositions. Unique compositions will be determined by hashing and then within each hashbin checking the n(n-1) equality options. The current hash ends up hashing very large numbers of compositions to the same hash code
``` python
def __hash__(self):
"""
Minimally effective hash function that just distinguishes between
Compositions with different elements.
"""
hashcode = 0
for el, amt in self.items():
if abs(amt) > Composition.amount_tolerance:
hashcode += el.Z
return hashcode
```
A more discriminative hash might be:
```python
from math import floor
def __hash__(self):
"""
Minimally effective hash function that just distinguishes between
Compositions with different elements.
"""
sym_amt = {
el: floor(1e7*(amt + Composition.amount_tolerance * 2))
for el, amt in self.get_el_amt_dict().items()
if abs(amt) > Composition.amount_tolerance
}
syms = sorted(sym_amt.keys())
formula = [s + formula_double_format(sym_amt[s], False) for s in syms]
return hash(" ".join(formula))
```
### robustness
```
>>> hash(Composition(f"O{0.8}Fe{0.2}"))
-2228561952991124878
>>> hash(Composition(f"O{0.8}Fe{0.2}Na{1e-9}"))
-2228561952991124878
>>> hash(Composition(f"O{0.8}Fe{0.2+Composition.amount_tolerance}"))
-2228561952991124878
>>> hash(Composition(f"O{0.8}Fe{0.2-Composition.amount_tolerance}"))
-2228561952991124878
>>> hash(Composition(f"O{0.8}Fe{0.2-2*Composition.amount_tolerance}"))
-2228561952991124878
>>> hash(Composition(f"O{0.8}Fe{0.2-5*Composition.amount_tolerance}"))
-414002531081527414
>>> hash(Composition(f"O{0.8+Composition.amount_tolerance}Fe{0.2-2*Composition.amount_tolerance}"))
-2228561952991124878
>>> hash(Composition(f"Cl{0.8+Composition.amount_tolerance}Na{0.2-2*Composition.amount_tolerance}"))
1665041687307082964
>>> hash(Composition(f"Fe{0.2}O{0.8-Composition.amount_tolerance}"))
-2228561952991124878
```
@shyamd do you see any issues with this change to the hash? I think it maintains a greater level of robustness than `__eq__`.
an issue might be that the current hash is not salted but the proposed hash would be
### speed
for a set of 18,322 compositions where 14,201 are unique I timed the following `foo()` function
```python
df = pnd.read_csv("data/hash-test-small.csv", na_filter=False)
df = StrToComposition(target_col_id="composition_obj").featurize_dataframe(df, "composition")
df["frac_comp"] = df["composition_obj"].apply(lambda x: x.fractional_composition)
def foo():
print(len(df["frac_comp"].unique()))
```
```
proposed: 1 loop, best of 5: 793 msec per loop
current: 1 loop, best of 5: 4.94 sec per loop
```
for a set of 454,484 where 317,764 are unique timing the new hash gives
```
proposed: 1 loop, best of 5: 23.1 sec per loop
```
The current hash is still running whilst writing this post.
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2076/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2076/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2077
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2077/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2077/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2077/events
|
https://github.com/materialsproject/pymatgen/pull/2077
| 816,691,859
|
MDExOlB1bGxSZXF1ZXN0NTgwMzEwMjcz
| 2,077
|
Add a more discriminative hash for Composition
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/37458509)\n\nCoverage decreased (-0.7%) to 79.517% when pulling **9e09cf9526eca1d064ac98ff0ecd98d22fbb03de on CompRhys:comphash** into **6cb45f5fbb1bde78aef668c48582a92f7a910604 on materialsproject:master**.\n",
"Thanks."
] | 2021-02-25T18:37:07
| 2021-02-25T21:03:31
|
2021-02-25T20:19:21Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
New more discriminative hash see #2076 for discussion
## Checklist
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2077/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2077/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2077",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2077",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2077.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2077.patch",
"merged_at": "2021-02-25T20:19:21Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2078
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2078/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2078/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2078/events
|
https://github.com/materialsproject/pymatgen/pull/2078
| 817,752,237
|
MDExOlB1bGxSZXF1ZXN0NTgxMTk1MzI1
| 2,078
|
QChem Input OPT Section
|
{
"login": "arepstein",
"id": 17616763,
"node_id": "MDQ6VXNlcjE3NjE2NzYz",
"avatar_url": "https://avatars.githubusercontent.com/u/17616763?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/arepstein",
"html_url": "https://github.com/arepstein",
"followers_url": "https://api.github.com/users/arepstein/followers",
"following_url": "https://api.github.com/users/arepstein/following{/other_user}",
"gists_url": "https://api.github.com/users/arepstein/gists{/gist_id}",
"starred_url": "https://api.github.com/users/arepstein/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/arepstein/subscriptions",
"organizations_url": "https://api.github.com/users/arepstein/orgs",
"repos_url": "https://api.github.com/users/arepstein/repos",
"events_url": "https://api.github.com/users/arepstein/events{/privacy}",
"received_events_url": "https://api.github.com/users/arepstein/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/37495101)\n\nCoverage decreased (-0.6%) to 79.521% when pulling **b3e37b460207ec7bb639717bc6f8f27ed00b8d29 on arepstein:ae_qchem** into **ccd04311c957b03cfbe3735348d57842c51cabc6 on materialsproject:master**.\n",
"Thanks.\r\n"
] | 2021-02-26T23:41:41
| 2021-03-04T16:05:27
|
2021-03-04T16:05:21Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
* Allows QChem OPT input file section to be written from OptSet
## Additional dependencies introduced (if any)
*None
## TODO (if any)
*None
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [ x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [ x] Tests have been added for any new functionality or bug fixes.
- [ x] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2078/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2078/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2078",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2078",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2078.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2078.patch",
"merged_at": "2021-03-04T16:05:21Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2079
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2079/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2079/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2079/events
|
https://github.com/materialsproject/pymatgen/issues/2079
| 818,251,122
|
MDU6SXNzdWU4MTgyNTExMjI=
| 2,079
|
pymatgen 2021.2.16 documentation: get_spacegroup_symbol() seems to be typo for get_space_group_symbol()
|
{
"login": "eiichirou",
"id": 16527923,
"node_id": "MDQ6VXNlcjE2NTI3OTIz",
"avatar_url": "https://avatars.githubusercontent.com/u/16527923?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/eiichirou",
"html_url": "https://github.com/eiichirou",
"followers_url": "https://api.github.com/users/eiichirou/followers",
"following_url": "https://api.github.com/users/eiichirou/following{/other_user}",
"gists_url": "https://api.github.com/users/eiichirou/gists{/gist_id}",
"starred_url": "https://api.github.com/users/eiichirou/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/eiichirou/subscriptions",
"organizations_url": "https://api.github.com/users/eiichirou/orgs",
"repos_url": "https://api.github.com/users/eiichirou/repos",
"events_url": "https://api.github.com/users/eiichirou/events{/privacy}",
"received_events_url": "https://api.github.com/users/eiichirou/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-02-28T16:51:00
| 2021-03-01T00:04:03
|
2021-03-01T00:04:03Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
In Section "Usage" of [pymatgen project top page](https://pymatgen.org/),
part of
`# Integrated symmetry analysis tools from spglib.`
`from pymatgen.symmetry.analyzer import SpacegroupAnalyzer`
`finder = SpacegroupAnalyzer(structure)`
`finder.get_spacegroup_symbol()`
returns
> AttributeError: 'SpacegroupAnalyzer' object has no attribute 'get_spacegroup_symbol'
alternatively,
`finder.get_space_group_symbol()`
returns
> 'Pm-3m'
Though it seems to be just a typo, one can confirm installation check using these codes.
Thus I report here above mentioned.
Thanks
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2079/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2079/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2080
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2080/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2080/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2080/events
|
https://github.com/materialsproject/pymatgen/pull/2080
| 821,427,103
|
MDExOlB1bGxSZXF1ZXN0NTg0MjE5MTYw
| 2,080
|
Modify Critic2Caller for more modular use
|
{
"login": "samblau",
"id": 7431223,
"node_id": "MDQ6VXNlcjc0MzEyMjM=",
"avatar_url": "https://avatars.githubusercontent.com/u/7431223?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/samblau",
"html_url": "https://github.com/samblau",
"followers_url": "https://api.github.com/users/samblau/followers",
"following_url": "https://api.github.com/users/samblau/following{/other_user}",
"gists_url": "https://api.github.com/users/samblau/gists{/gist_id}",
"starred_url": "https://api.github.com/users/samblau/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/samblau/subscriptions",
"organizations_url": "https://api.github.com/users/samblau/orgs",
"repos_url": "https://api.github.com/users/samblau/repos",
"events_url": "https://api.github.com/users/samblau/events{/privacy}",
"received_events_url": "https://api.github.com/users/samblau/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/37628931)\n\nCoverage decreased (-0.6%) to 79.5% when pulling **0780b2a4a2f33d9761ad8e50c21980b876ee63fe on samblau:qchem** into **b8d63d20728deed91d02d507e739d05f3d49a61e on materialsproject:master**.\n",
"@mkhorton If this looks good to you, I'd appreciate if a new version could be released following the merge. Thanks!",
"thanks,.",
"Why is a release needed?",
"So that I can use this functionality in my open Atomate PR #598",
"I will release today.",
"Thank you!"
] | 2021-03-03T19:44:31
| 2021-03-03T20:18:05
|
2021-03-03T20:10:22Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Modifications per discussions with @mkhorton in Atomate PR #598. Made `Critic2Caller.__init__` only responsible for actually calling `Critic2`, and put the rest of the chgcar-specific details into a `from_chgcar` class method.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2080/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2080/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2080",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2080",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2080.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2080.patch",
"merged_at": "2021-03-03T20:10:22Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2081
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2081/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2081/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2081/events
|
https://github.com/materialsproject/pymatgen/issues/2081
| 822,064,645
|
MDU6SXNzdWU4MjIwNjQ2NDU=
| 2,081
|
Context managers for perturbing and translating lattice sites
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Based on preliminary testing, the new API would be about twice as fast as the current one, probably due to not [creating new `Site`s to overwrite existing ones in `translate_sites`](https://github.com/materialsproject/pymatgen/blob/33ad121f3ad99ab5e61d4142d4200e7519e12d28/pymatgen/core/structure.py#L3995) but simply reusing them (as the operation is destructive anyway):\r\n\r\n```py\r\nfrom contextlib import contextmanager\r\nfrom time import perf_counter\r\nfrom pymatgen import MPRester\r\n\r\nmpr = MPRester(api_key=\"foobar\")\r\ncystal = mpr.get_structure_by_material_id('mp-1000')\r\n\r\n@contextmanager\r\ndef perturb_crystal(crystal, std=0.03):\r\n try:\r\n old_coords = crystal.cart_coords\r\n\r\n # loop over each site and apply Gaussian noice to position\r\n for site in crystal:\r\n perturbation = np.random.normal(0, std, [3])\r\n site.coords += perturbation\r\n\r\n yield crystal\r\n\r\n finally:\r\n for coords, site in zip(old_coords, crystal):\r\n site.coords = coords\r\n\r\n\r\nstart = perf_counter()\r\n\r\nfor _ in range(10000):\r\n with perturb_crystal(crystal) as cryst:\r\n pass\r\n\r\nprint(f\"context took {perf_counter() - start:.3f} sec\")\r\n\r\n\r\nstart = perf_counter()\r\n\r\nfor _ in range(10000):\r\n old_coords = crystal.cart_coords\r\n crystal.perturb(0.03)\r\n \r\n for site, coords in zip(crystal, old_coords):\r\n site.coords = coords\r\n\r\nprint(f\"existing perturb() took {perf_counter() - start:.3f} sec\")\r\n```\r\n\r\nThis printed\r\n\r\n```txt\r\nConnection established to Materials Project database, version 2021_02_08.\r\ncontext took 0.581 sec\r\nperturb() took 1.032 sec\r\n```",
"I don't think this is a common enough use case to justify a context manager. It is rare that you want to \"go back\" to the original structure after you perturb it. Even when you want to keep the structure, there are simpler ways to do this.\r\n\r\nE..g.\r\n\r\n```python\r\ns2 = structure.copy()\r\ns2.perturb(0.01)\r\n```\r\n\r\nor if you are going to be doing it millions of times,\r\n\r\n```\r\nlattice = structure.lattice\r\nspecies = structure.species\r\ncoords = structure.cart_coords\r\nfor _ in range(1000000):\r\n new_coords = coords + np.random.normal(coords.shape[0], 3)\r\n s2 = Structure(lattice, species, new_coords, coords_are_cartesian=True)\r\n```\r\n\r\n\r\n",
"Thanks for the quick reply!\r\n\r\n> I don't think this is a common enough use case to justify a context manager.\r\n\r\nI think it's likely that data augmentation for structure-based models will become more widely used in the future.\r\n\r\n> Even when you want to keep the structure, there are simpler ways to do this.\r\n> [...]\r\n> \r\n> ```\r\n> lattice = structure.lattice\r\n> species = structure.species\r\n> coords = structure.cart_coords\r\n> for _ in range(1000000):\r\n> new_coords = coords + np.random.normal(coords.shape[0], 3)\r\n> s2 = Structure(lattice, species, new_coords, coords_are_cartesian=True)\r\n> ```\r\n\r\nI don't find that as simple as:\r\n\r\n```py\r\nfor _ in range(10000):\r\n with crystal.perturbed() as cryst:\r\n pass\r\n```\r\n\r\nBut it's slightly faster so good to know. :)",
"I agree data augmentation is useful for ML. But the best way to do that isn't using the structure.perturb method (with or without context managers). The work you were referring to uses MEGNet graph models from my group. The input is a crystal graph, not a structure. Of course, in a typical case where you have unique structures, you construct a graph from a Structure object. But if you are planning to do data augmentation, a much faster way is to directly construct the graph from coordinates rather than doing Structure -> Perturbed Structure -> Graph because the species, i.e., nodes, are not actually changing. The cost in constructing the Structure object is always expensive. While manipulating numpy arrays are cheap.",
"Thanks for this advice! I'm aware that Megnet ultimately uses a graph representation but it still computes that from a `Structure()` in its `get_graphs_within_cutoff()` function.\r\n\r\nhttps://github.com/materialsvirtuallab/megnet/blob/0c75e9cfd0c9b949c6d5d08ef7bcd211718b2138/megnet/utils/data.py#L14-L36\r\n\r\nSo your suggestion is to modify `get_graphs_within_cutoff` to not accept a `Structure` but pass in the `lattice.matrix` and `cart_coords` as bare numpy arrays? And then turn\r\n\r\n```py\r\ncart_coords = np.ascontiguousarray(np.array(structure.cart_coords), dtype=float)\r\n```\r\n\r\ninto\r\n\r\n```py\r\ncart_coords = np.ascontiguousarray(cart_coords + np.random.normal(...), dtype=float)\r\n```",
"Yeah, my advice is to go the the lower level numpy arrays to speed things up, especially if you plan to do lots of augmentation. @chc273 can probably advise you on the best way to proceed. Note that the get_graphs_within_cutoff basically uses the underlying cartesian coordinates (with PBC) anyway. If it is just augmenting a few structures, then probably the cost savings is not really worth it and you might as well use the slower methods I mentioned before.",
"@shyuep @janosh Yes I think working with the numpy arrays directly instead of going from Structure -> Perturbed Structure -> Perturbed Graph will be much faster since constructing python objects is slow. You can basically call the find_points_in_spheres method in pymatgen, which takes numpy arrays as inputs. The get_graphs_within_cutoff is just a thin wrapper of the pymatgen method. "
] | 2021-03-04T11:58:05
| 2021-03-04T17:33:14
|
2021-03-04T17:33:14Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Is your feature request related to a problem? Please describe.**
[`Structure.perturb()`](https://pymatgen.org/pymatgen.core.structure.html#pymatgen.core.structure.Structure.perturb) and [`Structure.translate_sites()`](https://pymatgen.org/pymatgen.core.structure.html#pymatgen.core.structure.Structure.translate_sites) make in-place modifications to the lattice sites of a crystal. This can lead to user errors when people don't consider the implications of permanent changes for downstream tasks or simply forget to revert to the original positions manually.
**Describe the solution you'd like**
A way to avoid these pitfalls would be to provide new methods, say `Structure.perturbed()` and `Structure.translated_sites()`, as context managers which perform temporary lattice site modifications and revert back to the original positions when the context exits. The new API could look like this:
```py
from pymatgen import Structure
crystal = Structure()
print('original')
print(crystal.cart_coords)
with crystal.perturbed(0.05) as cryst:
print('perturbed coordinates')
print(cryst.cart_coords)
print('unchanged original')
print(crystal.cart_coords)
```
which would print:
```txt
original
[[ 3.34134688 0.94786948 1.78078667]
[ 0.69595394 3.04452513 1.31349259]
[ 2.26161323 2.87664712 -0.55981345]
[ 1.34395046 1.61686791 3.70986864]
[ 3.64365627 4.24525709 -3.22423295]]
perturbed coordinates
[[ 3.31185334 0.92091982 1.80484822]
[ 0.64055798 3.06264923 1.26460508]
[ 2.19797793 2.82148173 -0.50081053]
[ 1.33806345 1.61946541 3.75244629]
[ 3.67168166 4.20351193 -3.19816276]]
unchanged original
[[ 3.34134688 0.94786948 1.78078667]
[ 0.69595394 3.04452513 1.31349259]
[ 2.26161323 2.87664712 -0.55981345]
[ 1.34395046 1.61686791 3.70986864]
[ 3.64365627 4.24525709 -3.22423295]]
```
and be implemented like this:
```py
from contextlib import contextmanager
import numpy as np
class Structure:
@contextmanager
def perturbed(self, std=0.03):
try:
old_coords = self.cart_coords
# loop over each site and apply Gaussian noise to its position
for site in self:
perturbation = np.random.normal(0, std, [3])
site.coords += perturbation
yield self
finally:
for site, coords in zip(self, old_coords):
site.coords = coords
```
**Additional context**
Such temporary lattice perturbations are useful for on-the-fly data augmentation while training structure-based ML models, see e.g. ["Crystal Structure Search with Random Relaxations Using Graph Networks"](https://arxiv.org/abs/2012.02920).
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2081/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2081/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2082
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2082/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2082/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2082/events
|
https://github.com/materialsproject/pymatgen/issues/2082
| 822,246,988
|
MDU6SXNzdWU4MjIyNDY5ODg=
| 2,082
|
Out of date example using Interstitial in documentation
|
{
"login": "katerinavr",
"id": 43647831,
"node_id": "MDQ6VXNlcjQzNjQ3ODMx",
"avatar_url": "https://avatars.githubusercontent.com/u/43647831?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/katerinavr",
"html_url": "https://github.com/katerinavr",
"followers_url": "https://api.github.com/users/katerinavr/followers",
"following_url": "https://api.github.com/users/katerinavr/following{/other_user}",
"gists_url": "https://api.github.com/users/katerinavr/gists{/gist_id}",
"starred_url": "https://api.github.com/users/katerinavr/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/katerinavr/subscriptions",
"organizations_url": "https://api.github.com/users/katerinavr/orgs",
"repos_url": "https://api.github.com/users/katerinavr/repos",
"events_url": "https://api.github.com/users/katerinavr/events{/privacy}",
"received_events_url": "https://api.github.com/users/katerinavr/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@mbkumar can you update pls?",
"@shyuep,\r\n\r\nThere are two parts. a) The installation for Zeo++, which could be still valid. b) The validation example which uses the Interstitial class is outdated. The defects subpackage was rewritten by Shyam and Danny. They were using a new algorithm introduced by Nils Zimmerman for interstitials. \r\n\r\nSo my suggestion would be to rewrite the interstitial generation example using the latest defects code and move it to a new subsection titled \"Defects\" if not already present. And leave the rest of the Zeo++ section intact after removing/relocating the example code. \r\n\r\n@shyamd,\r\ncould you please take a look at the non-working example and write a new canonical example for interstitial generation.\r\n\r\nBharat"
] | 2021-03-04T15:37:11
| 2023-08-13T16:34:58
|
2023-08-13T16:34:58Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
The example given in the documentation making use of `pymatgen.analysis.defects.core.Interstitial` seems to be outdated or incorrect. Following the example shown on the installation page yields to the following error message:
```
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
TypeError: __init__() got an unexpected keyword argument 'radii'
```
**To Reproduce**
See the last code snippet at https://pymatgen.org/installation.html.
```
from pymatgen.analysis.defects.core import Interstitial
from pymatgen.core.structure import Structure
structure = Structure.from_file('/path/to/file')
radii, valences = {}, {}
for element in structure.composition.elements:
radii[element.symbol] = element.atomic_radius
valence = element.group # Just a first guess..
if element.group > 12:
valence -= 10
valences[element.symbol] = valence
interstitial = Interstitial(structure, radii=radii, valences=valences)
```
If you could please provide an up-to-date method of using this class that would be really helpful
Regards
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2082/reactions",
"total_count": 1,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 1,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2082/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2083
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2083/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2083/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2083/events
|
https://github.com/materialsproject/pymatgen/issues/2083
| 823,062,398
|
MDU6SXNzdWU4MjMwNjIzOTg=
| 2,083
|
please consider switching back to semantic versioning
|
{
"login": "ltalirz",
"id": 1053245,
"node_id": "MDQ6VXNlcjEwNTMyNDU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1053245?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ltalirz",
"html_url": "https://github.com/ltalirz",
"followers_url": "https://api.github.com/users/ltalirz/followers",
"following_url": "https://api.github.com/users/ltalirz/following{/other_user}",
"gists_url": "https://api.github.com/users/ltalirz/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ltalirz/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ltalirz/subscriptions",
"organizations_url": "https://api.github.com/users/ltalirz/orgs",
"repos_url": "https://api.github.com/users/ltalirz/repos",
"events_url": "https://api.github.com/users/ltalirz/events{/privacy}",
"received_events_url": "https://api.github.com/users/ltalirz/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"> These problems would be solved by switching back to [semantic versioning](https://semver.org/)\r\n\r\nShould note that this solves problems _provided_ that minor and patch versions do not break backwards compatibility. Might be obvious given the definition of semantic versioning but is in practice not always easy nor guaranteed.",
"The reason why calendar versioning is used instead of semantic versioning is because the maintenance costs of semantic versioning far outweigh the benefits for us. It is rare that pymatgen introduces breaking changes (other than the recent v2021.3.4, I can't remember really when was the last time this happened). There are also breaking changes that can be introduced by codes that pymatgen depend on, e.g., the recent numpy v1.20 that dropped support for Python < 3.7 and introduced support for the ArrayLike typing object.\r\n\r\nI am also unclear what the benefit of semantic versioning is. Other than the options you mentioned, you can also specify a range of pymatgen versions that would work (a strategy that a lot of people use for something like numpy) even with calendar versioning. Also, any code that is compatible with a higher version of pymatgen, e.g., 2021.3.4 would usually work with pymatgen<2021.3.4. @mkhorton and I try hard to make sure this is the case. So even if your project decides to make sure that you are compatible with pymatgen>=2021.3.4, it does not have to make it a requirement. In fact, if you set your requirements to say pymatgen>=2020.12.31 or even 2019.12.31, that usually poses no issues at all though you may lose some access to later analysis. It is much rarer that other codes depend on the latest analysis published within pymatgen. Most codes depend on pymatgen for its core functionality, which has rarely changed beyond some minor speed improvements over the past few years.\r\n\r\nNow, I would like to take this opportunity to explain **why** the breaking changes of removing root-level imports was implemented. The reason is in fact to better help other developers. We want to eventually move to making pymatgen and certain subpackages (e.g., analysis and io would be main examples) into [namespace packages](https://packaging.python.org/guides/packaging-namespace-packages/). This will allow us to distribute pymatgen in smaller modular pieces. Developers can also write add-on packages for pymatgen that can be maintained in separate repositories with their own requirements. \r\n\r\nI will use the example of the package from my group called pymatgen-diffusion, which provides for diffusion analysis. Arguably, this could have been implemented under `pymatgen.analysis` in the main repo, but it was decided to do this in a separate repository because I did not deem it of sufficiently broad appeal. However, because it is not a namespace package, pymatgen-diffusion exists as a separate package that has to be imported using the awkward `from pymatgen_diffusion import`. Once we move to a namespace package, we can make pymatgen-diffusion as part of the pymatgen.analysis sub-package (imported via `from pymatgen.analysis.diffusion import`) while still maintaining the flexibility to keep it in a separate install (`pip install pymatgen-diffusion`).\r\n\r\nThe above example is a more trivial one and merely one of optical cleanliness. But there are situations where this would be extremely useful. For instance, if someone wanted to introduce an add-on machine learning package for pymatgen that depends on tensorflow. Tensorflow is a horrific dependency that we would not want to impose on everyone using pymatgen for core functionality and does not care about ML. Namespace packages would allow these functionality to be implemented as add-ons. Another example are of course io packages, e.g., an add-on to support a new quantum mechanics software used by its own small community.\r\n\r\nFinally, I just want to say that while we take pains to minimize such changes, we also try to make sure that the transition is made as painless as possible. I did provide the two commands that if you run them on your repos, it should resolve most of the breaking imports. \r\n\r\n```bash\r\n# for mac\r\nfind . -name '*.py' | xargs sed -i \"\" 's/from pymatgen import MPRester/from pymatgen.ext.matproj import MPRester/g'\r\nfind . -name '*.py' | xargs sed -i \"\" 's/from pymatgen import/from pymatgen.core import/g'\r\n\r\n# for linux\r\nfind . -name '*.py' | xargs sed -i 's/from pymatgen import MPRester/from pymatgen.ext.matproj import MPRester/g'\r\nfind . -name '*.py' | xargs sed -i 's/from pymatgen import/from pymatgen.core import/g'\r\n```\r\n\r\nI tried them on the several codes that I maintain and they work for the most part. There may be a few instances, e.g., if you have something like `from pymatgen import Element, MPRester` somewhere in your code that needs to be manually broken into two separate imports now. Again, I stress that just because you made your code compatible with pymatgen>=2021.3.4 does not mean that you have to make that your minimal pymatgen version. Your code still works with previous pymatgen versions.",
"Thanks for the detailed reply @shyuep !\r\n\r\nLet me just follow up on some of the points you made:\r\n\r\n> I am also unclear what the benefit of semantic versioning is. [...]\r\n> In fact, if you set your requirements to say pymatgen>=2020.12.31 or even 2019.12.31, that usually poses no issues at all though you may lose some access to later analysis.\r\n\r\nOne key benefit of semantic versioning is the ability to say (as a developer of an upstream tool): I am ok with any pymatgen version X or later, as long as it does not introduce breaking changes.\r\n\r\nEverybody understands that breaking changes are unavoidable from time to time.\r\nThe breaking change https://github.com/materialsproject/pymatgen/commit/8f097cf7ea20b8fb42635452b02f520152218394 likely means that a substantial fraction of the 493 projects depending on pymatgen stopped working today for anybody doing a fresh install from PyPI.\r\n\r\nIf we imagine for the moment that you had been using semantic versioning, then you would have increased the major version number together with this change (say, to 6.0).\r\nAny project that would have depended (as they should) on pymatgen up to version 5.x (e.g. via `pymatgen~=5.2` or even `pymatgen>=4.0,<6`) would still be functional today.\r\nInstead of many projects now having to scramble to immediately make their code base compatible with the latest pymatgen version, they could do this in their own time at some point in the future - or never, if they don't require the latest pymatgen functionality.\r\n\r\nI certainly agree that keeping an eye on breaking changes and reflecting this in the version number is more work than just taking the day of the release.\r\nHowever, I would argue the larger the number of projects depending on a given library, the higher is the payoff for doing so.",
"> I am also unclear what the benefit of semantic versioning is. Other than the options you mentioned, you can also specify a range of pymatgen versions that would work (a strategy that a lot of people use for something like numpy) even with calendar versioning. \r\n\r\nI guess the benefit is that with sematic versioning one specify an sensible *upper* limit of the version, which is not possible with calendar versioning. In both cases, there is no problem to specify a range for already released versions. Taking `numpy` as an example, one should specify `numpy>=1.2.0,<2` to avoid picking up `2.x` numpy version when it gets a new release that can potentially causes problems. ",
"@ltalirz @zhubonan You can also do the same with calendar versioning? E.g., pymatgen>=2019.12.31,<2021.3.4? Of course, I recognize it is not as clean as saying <6.0.0 in the case of semantic versioning. \r\n\r\nI wouldn't think that the 493 projects that depend on pymatgen would stop working today. In fact, if you pip-installed say matminer or custodian today from Pypi, they will still install an old version of pymatgen and everything would work as per normal. The main people who would in fact be affected are those that install from the Github source and attempted to install the latest pymatgen.\r\n\r\nI welcome suggestions on how else I can try to make this more painless. We had a lot of discussions internally on how best to move ahead and trust me, we generally try not to implement breaking changes (except those that upstream codes such as numpy cause). ",
"> You can also do the same with calendar versioning? E.g., pymatgen>=2019.12.31,<2021.3.4?\r\n\r\nThat approach does indeed work for \"past\" versions, but it does not work for future versions, since there is no way of knowing whether they will be compatible.\r\n\r\n> if you pip-installed say matminer or custodian today from Pypi, they will still install an old version of pymatgen and everything would work as per normal.\r\n\r\nRight - this is because matminer chose option 1. I mentioned [above](https://github.com/materialsproject/pymatgen/issues/2083#issue-823062398) - with the corresponding downside of clashing with any other tool that would take such an approach (it can work for tools that go into the direction of a \"distribution\", which are never meant to be installed alongside something else).\r\n\r\nhttps://github.com/hackingmaterials/matminer/blob/master/requirements.txt\r\n\r\n> I welcome suggestions on how else I can try to make this more painless. We had a lot of discussions internally on how best to move ahead and trust me, we generally try not to implement breaking changes (except those that upstream codes such as numpy cause).\r\n\r\nTrying to avoid breaking changes is certainly extremely valuable (and thanks for putting so much effort into this!).\r\n\r\nHowever, when the time comes eventually where a breaking change becomes necessary, in my view the most painless way to communicate this is through semantic versioning.",
"@shyuep The problem is that on the day of 2021.1.1 I have no idea that pymatgen version 2021.3.4 will introduce breaking changes - so I cannot put `pymatgen>=2019.12.31,<2021.3.4` in the dependency. \r\n\r\nOn the other hand, if semantic versioning is used, and on 2021.1.1 pymatgen is at 5.3.2 version. I just need to put `pymatgen>=5.3,<6` in my project dependency. Later, on 2021.3.4, a new version of pymatgen released and contains breaking changes, but it has a version number like `6.x`. Install my package will still pick up the old compatible version instead of the new one. ",
"@zhubonan Thanks for clarifying. Good point. Then may I propose a compromise and feel free to tell me this works.\r\n* I will remove v2021.3.4 and release a new pymatgen v2021.3.5 that contains the root-level import. This will reverse the breaking change for most people.\r\n* I will release a new version called 2022.0.0 with the breaking change. This has the effect of \"semantic\" versioning. Once we get into y 2022, any new versions will of course supersede 2022.0.0.\r\n* If there are critical bug fixes that needs to be backported, I can release 2021.x.x versions that can include those fixes.\r\n\r\nThis will help the developers who actually pin to a semantic version. I doubt that a lot of people do that. But I am willing to do this to help everyone along if this is useful.",
"Thanks a lot @shyuep ! I think this would be greatly appreciated by many developers relying on pymatgen.\r\n\r\n> I will remove v2021.3.4 \r\n\r\nIf this turns out to be too difficult (I think it can take time to take a release off pypi), skipping this step is probably also ok, since most people will pick up the latest version v2021.3.5 anyways.",
"Ok, thanks all for the input. I have just released v2021.3.5 and v2022.0.0. These will be live once the release Github Actions workflow are done (~30mins). v2021.x development will be on the \"legacy\" branch while we will continue the main line of development on v2022.0.x for the remainder of 2021.\r\n",
"I saw @shyuep just closed this. I don't have any concrete suggestions, but I did want to offer some support and encouragement.\r\n\r\nThere are a significant number of contributions to pymatgen from people outside of the core development team and it would take significant resources on their part to maintain proper SemVer. The definition of \"breaking change\" is really hard to determine - I recently came across [Hyrum's Law](https://www.hyrumslaw.com) from this [relevant blog post advocating against SemVer](https://hynek.me/articles/semver-will-not-save-you/):\r\n\r\n> With a sufficient number of users of an API,\r\nit does not matter what you promise in the contract:\r\nall observable behaviors of your system\r\nwill be depended on by somebody.\r\n\r\nIf pymatgen strictly followed SemVer under a definition of \"breaking\" similar to Hyrum's Law for the past few years, the major version number increments would be about the same burden on packages downstream of pymatgen as CalVer. While breaking changes like v2021.3.4 can be surprising to downstream packages, I think calendar versioning on the whole has worked well for the pymatgen community. \r\n\r\nThank you, pymatgen team, for your hard work and dedication to this package and community.",
"I just wanted to add that we've added a page on compatibility and versioning issues to our documentation:\r\n\r\nhttps://pymatgen.org/compatibility.html\r\n\r\nThis page is just a first attempt and will be improved, and we do take the issue of backwards compatibility very seriously. I can't remember the last time we renamed or removed a method in pymatgen.core, for example.\r\n\r\nI do agree with @bocklund that the main issue with semantic versioning is drawing the line where the main version should be incremented. The example of root-level imports is a clear case where it would have been useful to have semver, but these cases are rare. We're also not alone in using date versioning: many large packages do so also (e.g. `pip` itself for example). In retrospect, we could have made this change just before 2021.x, as we did when we dropped Python 2.x support, but we missed that opportunity. \r\n\r\nIn any case, I'm also grateful for the discussion and feedback, and sincerely apologize for causing the extra work in this case (as it will for myself as well). Hopefully it will bring sufficient benefits in the long run to be worthwhile.",
"Also just catching up with the above messages too, looks like I should update that doc page 🙃 Hopefully this is a good resolution for everyone though!",
"Just wanted to say that I appreciate it a lot that you guys responded so quickly and willing to discuss these things. I agree with what has been said before that deciding what exactly constitutes as breaking is not always easy. There are very clear cut and dry cases, but there is often a grey area. I personally think that calendar based versioning definitely has a place, but is probably best used for end-applications. I think that for libraries semantic versioning still is better suited as it allows consumers upstream to better prepare for eventually inevitable breaking changes. Anyway, thanks a lot for the compromise solution :+1: and all the work you put in `pymatgen`.",
"> we do take the issue of backwards compatibility very seriously\r\n\r\nRespectfully 😬, I would just point out that after you said this, multiple breaking changes were made in patch versions; ironically to how you can actually retrieve the version of pymatgen:\r\n\r\n`2022.0.0 -> 2022.0.1` it was moved from `pymatgen.__version__` to `pymatgen.settings.__version__` : https://github.com/materialsproject/pymatgen/commit/f462d35cab98d15f6abbbb9d3eec4d0ec53e49a8\r\n\r\n`2022.0.2 -> 2022.0.3` it was moved from `pymatgen.settings.__version__` to `pymatgen.core.__version__`: https://github.com/materialsproject/pymatgen/commit/2d8b9927b0ef3df9ef3821f9e501de01392c0a93\r\n\r\nSo now people have to use a number of try/excepts just to get the version of pymatgen\r\n\r\n> We want to eventually move to making pymatgen and certain subpackages (e.g., analysis and io would be main examples) into namespace packages. This will allow us to distribute pymatgen in smaller modular pieces.\r\n> However, because it is not a namespace package, pymatgen-diffusion exists as a separate package that has to be imported using the awkward from `pymatgen_diffusion` import. \r\n\r\nIt sounds like this decision has already been made, but anyhow I would still advise against using this namespace package structure, in favour of just moving/leaving the packages to different repositories.\r\nSure this is a step towards modularity, but you still end up with one list of issues/PRs for a bunch of packages with different development/release cadences. I'm not sure why `pymatgen_diffusion` is so awkward 😉 \r\n\r\nThis reminds of ansible's recent restructuring, that now is split into ansible-core and community repositories (and also adopted semantic versioning):\r\n\r\n- https://www.ansible.com/blog/thoughts-on-restructuring-the-ansible-project\r\n- https://www.ansible.com/blog/announcing-the-community-ansible-3.0.0-package ",
"@shyuep maybe worth removing the `2022.0.0`, `2022.0.1`, `2022.0.2` releases for this reason?",
"@chrisjsewell I'd only add that the commitment of \"taking backwards compatibility seriously\" is not in conflict with also sometimes making mistakes or mis-steps, so I'd ask for patience while we figure this out. Ultimately, _pymatgen_ is a community-driven non-profit/academic code and we're all trying to do our best here.",
"@chrisjsewell Yes, there were a few moves from 2022.0.0-2022.0.3. I accept culpability for it since this is the first time I am doing a namespace package and mistakes were made for the first few releases. That was fixed in 2022.0.3. Since SETTINGS and __version__ are primarily used internally within pymatgen, I don't think a lot of people were affected.\r\n\r\nAs a compromise, I have \"yanked\" 2022.0.0-2022.0.2, i.e., a form of soft-delete that prevents new users from installing them but allows users who have pinned them to continue to use them with warning messages. Hard deletes are frowned upon.\r\n\r\nI am unclear why you think avoiding namespace packages and simply moving packages to different repos is a better outcome. That would certainly cause more issues for end users. And you are mistaken in thinking this was done merely to break pymatgen into separate components (though we may indeed move certain less frequently used subpackages that are primarily maintained by external collaborators to separate repos). This is a forward move for more decentralized pymatgen functionality development by other developers. It is up to those developers to set their own cadence of release and manage their Issues and PRs. The main pymatgen distribution will focus on core functionality that caters to the majority of use cases by the broader community. This is really no different from the relationship between something like Jupyter or Flask and their various extensions and plugins. No one would think that the flask core developer has any role in setting the cadence of a plugin written by another developer.\r\n\r\nFinally, the 2022 versions are in fact a compromise towards temporary semantic version for the remainder of this year. I do not want to get into philosophical arguments about the pros and cons of semantic vs calendar versioning. ",
"> I am unclear why you think avoiding namespace packages and simply moving packages to different repos is a better outcome. \r\n> This is really no different from the relationship between something like Jupyter or Flask and their various extensions and plugins\r\n\r\nBoth your examples use multiple repositories over namespace packages, jupyter even specifically has `jupyter/jupyter_core`.\r\nI'm just yet to see a project that successfully uses namespace packages\r\n\r\n> No one would think that the flask core developer has any role in setting the cadence of a plugin written by another developer.\r\n\r\nthey would if the plugins were in the same repository as the core package\r\n\r\n> It is up to those developers to set their own cadence of release and manage their Issues and PRs\r\n\r\nwell I'm going to be interested to see how this works in the same repository; e.g. having release tags for multiple packages in the same git history 😬 \r\n\r\nBut anyhow, I'm just giving my personal opinion, its entirely up to you ✌️ ",
"The plugins are *not* going to be in the same repository as the core package. That is the whole point. E.g., pymatgen-diffusion is not in the same repo as pymatgen. If all plugins are in the same repo, there is hardly any point in making namespace packages isn't it?\r\n\r\nPymatgen main repo is now 681 Mb. I don't think anyone fancies it getting bigger. ",
"oh if you are saying that you are planning to move folders out of pymatgen into their own repositories then thats fine 👍 \r\n\r\nPersonally, it feels like namespaces is an over-complication then and the [namespace documentation](https://packaging.python.org/guides/packaging-namespace-packages/) even says:\r\n\r\n> However, namespace packages come with several caveats and are not appropriate in all cases. A simple alternative is to use a prefix on all of your distributions such as import mynamespace_subpackage_a (you could even use import mynamespace_subpackage_a as subpackage_a to keep the import object short).\r\n\r\n(Note also, I say this having made a very similar change in one of the packages I maintain: https://github.com/executablebooks/markdown-it-py/commit/bddc8829c33faa26a89ad1df85bbe8cfe0d80729, so I do put my money where my mouth is 😆)",
"This is a final update on this channel. I just released a v2022.0.4. There are no breaking changes in this one - just new functionality for Element (ionization energies and electron affinities). I also did a backport to a v2021.3.9. Feel free to let me and @mkhorton know if any issues arise. Otherwise, I think v2022.0.3 onwards is reasonably stable to migrate to as and when you are ready to do so. We will refrain from making any other breaking changes. :-)"
] | 2021-03-05T12:58:56
| 2021-03-09T19:34:34
|
2021-03-05T15:21:26Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Is your feature request related to a problem? Please describe.**

As the github landing page nicely illustrates, `pymatgen` is used as a dependency in many other projects, some of which are again used as components of other projects and so on.
pymatgen's [calendar versioning](https://calver.org/) approach essentially leaves developers of such projects with two options:
1. lock the version of the pymatgen dependency completely, e.g. `pymatgen==2021.3.4`.
This has the disadvantage that your tool becomes incompatible with any other tool following the same strategy because their pymatgen dependency versions would clash.
2. provide a minimal required pymatgen version, e.g. `pymatgen>=2020.3.4`.
This means your project can (and likely will) break when pymatgen makes breaking changes (such as the recent https://github.com/materialsproject/pymatgen/commit/8f097cf7ea20b8fb42635452b02f520152218394 )
These problems would be solved by switching back to [semantic versioning](https://semver.org/), which would allow developers of upstream tools to depend on pymatgen without risking to break their projects (e.g. `pymatgen~=5.0`) while still making it possible to work together with other tools that share the pymatgen dependency.
I know that pymatgen originally used semantic versioning and then switched to calendar versioning in https://github.com/materialsproject/pymatgen/commit/51411b19f27906d162de2328214ca3ffed16fdda#diff-60f61ab7a8d1910d86d9fda2261620314edcae5894d5aaa236b821c7256badd7 in 2017.
I would kindly ask the developers to elaborate on the reasons for this decision and, potentially, to reconsider it.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2083/reactions",
"total_count": 13,
"+1": 13,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2083/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2084
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2084/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2084/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2084/events
|
https://github.com/materialsproject/pymatgen/issues/2084
| 823,542,014
|
MDU6SXNzdWU4MjM1NDIwMTQ=
| 2,084
|
Outdated docstrings related to PeriodicNeighbor
|
{
"login": "lan496",
"id": 7272505,
"node_id": "MDQ6VXNlcjcyNzI1MDU=",
"avatar_url": "https://avatars.githubusercontent.com/u/7272505?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/lan496",
"html_url": "https://github.com/lan496",
"followers_url": "https://api.github.com/users/lan496/followers",
"following_url": "https://api.github.com/users/lan496/following{/other_user}",
"gists_url": "https://api.github.com/users/lan496/gists{/gist_id}",
"starred_url": "https://api.github.com/users/lan496/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/lan496/subscriptions",
"organizations_url": "https://api.github.com/users/lan496/orgs",
"repos_url": "https://api.github.com/users/lan496/repos",
"events_url": "https://api.github.com/users/lan496/events{/privacy}",
"received_events_url": "https://api.github.com/users/lan496/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks for reporting this. I have updated the doc.",
"Thank you very much for your quick reply and fix. Also, I appreciate the functions for getting neighbor atoms in pymatgen, which have helped my research a lot."
] | 2021-03-06T02:41:45
| 2021-03-07T05:56:59
|
2021-03-07T05:46:33Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
The docstrings for member functions of `IStructure` to get neighbor atoms are seemed to be outdated.
Related to `get_sites_in_sphere`, `get_neighbors`, `get_neighbors_old`, `get_all_neighbors`, `get_all_neighbors_py`, and `get_neighbors_in_shell`.
The difference between docstrings and codes are as follows:
- `include_index` and `include_image` were deprecated, but some of these functions do not mention so in the docstrings.
- Some docstrings say returned `PeriodicNeighbor`s have `distance` attribute, but `PeriodicNeighbor` has actually `nn_distance` instead.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2084/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2084/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2085
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2085/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2085/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2085/events
|
https://github.com/materialsproject/pymatgen/issues/2085
| 824,212,501
|
MDU6SXNzdWU4MjQyMTI1MDE=
| 2,085
|
Critic2Caller.from_path error
|
{
"login": "pjf295",
"id": 69720428,
"node_id": "MDQ6VXNlcjY5NzIwNDI4",
"avatar_url": "https://avatars.githubusercontent.com/u/69720428?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/pjf295",
"html_url": "https://github.com/pjf295",
"followers_url": "https://api.github.com/users/pjf295/followers",
"following_url": "https://api.github.com/users/pjf295/following{/other_user}",
"gists_url": "https://api.github.com/users/pjf295/gists{/gist_id}",
"starred_url": "https://api.github.com/users/pjf295/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/pjf295/subscriptions",
"organizations_url": "https://api.github.com/users/pjf295/orgs",
"repos_url": "https://api.github.com/users/pjf295/repos",
"events_url": "https://api.github.com/users/pjf295/events{/privacy}",
"received_events_url": "https://api.github.com/users/pjf295/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @pjf295, this is due to a recent change in #2080. The call on line 301 just needs to be changed to `cls.from_chgcar(...)`.",
"Actually, my mistake, this precedes that PR. Let me look into it"
] | 2021-03-08T06:14:08
| 2023-08-13T16:34:59
|
2023-08-13T16:34:59Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Provide any example files that are needed to reproduce the error,
especially if the bug pertains to parsing of a file.
**Expected behavior**
Properly return with a Critic2Caller object.
**Screenshots**
If applicable, add screenshots to help explain your problem.
**Desktop (please complete the following information):**
- OS: CentOS 7.9.2009
- Version '2020.12.31'
**Additional context**
The files in the example are quite large and it would be inappropriate to include them here. This is especially true in that hte error seems to prevent the files from being read at all. Note also that manually running Critic2 on the same files (constructing the sum of AECCAR0 and AECCAR2 within Critic2) works fine.
The Critic2Caller's from_path() method is a convenience method that aggregates CHGCAR AECCAR0 and AECCAR2. The method constructs the sum of the core and valence charge densities in a temporary file and then runs critic2 according to the documentation. When, however, I presented the from_path() method with a directory containing these files the following error occurred. Note that running critic2 manually worked without problem. In addition, the documentation claims that there is another method of Critic2Caller, namely from_chgcar() available, but in fact, it is not defined. Attempting to bring up the source results in an 404 error on github and, of course, a direct attempt to import the method also fails. The source also appears to be missing for Critic2Analysis.
The code was run under python 3.8.5/ipython 7.20.0 of anaconda.
In [1]: from pymatgen.command_line.critic2_caller import Critic2Analysis,Critic2Caller, CriticalPoint, CriticalPointType
In [2]: critic2 = Critic2Caller.from_path('.')
/home/paulfons/miniconda3/lib/python3.8/site-packages/pymatgen/command_line/bader_caller.py:458: UserWarning: Multiple files detected, using .
---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
<ipython-input-2-491bd541980e> in <module>
----> 1 critic2 = Critic2Caller.from_path('.')
~/miniconda3/lib/python3.8/site-packages/pymatgen/command_line/critic2_caller.py in from_path(cls, path, suffix, zpsp)
299 chgcar_ref = aeccar0.linear_add(aeccar2) if (aeccar0 and aeccar2) else None
300
--> 301 return cls(chgcar.structure, chgcar, chgcar_ref, zpsp=zpsp)
302
303
~/miniconda3/lib/python3.8/site-packages/monty/dev.py in decorated(*args, **kwargs)
94 if not self.condition:
95 raise RuntimeError(self.message)
---> 96 return _callable(*args, **kwargs)
97
98 return decorated
~/miniconda3/lib/python3.8/site-packages/pymatgen/command_line/critic2_caller.py in __init__(self, structure, chgcar, chgcar_ref, user_input_settings, write_cml, write_json, zpsp)
189
190 if chgcar and isinstance(chgcar, VolumetricData):
--> 191 chgcar.write_file("int.CHGCAR")
192 elif chgcar:
193 os.symlink(chgcar, "int.CHGCAR")
~/miniconda3/lib/python3.8/site-packages/pymatgen/io/vasp/outputs.py in write_file(self, file_name, vasp4_compatible)
3756 f.write("".join(self.data_aug.get(data_type, [])))
3757
-> 3758 write_spin("total")
3759 if self.is_spin_polarized and self.is_soc:
3760 write_spin("diff_x")
~/miniconda3/lib/python3.8/site-packages/pymatgen/io/vasp/outputs.py in write_spin(data_type)
3754 if count % 5 != 0:
3755 f.write(" " + "".join(lines) + " \n")
-> 3756 f.write("".join(self.data_aug.get(data_type, [])))
3757
3758 write_spin("total")
TypeError: can only join an iterable
**To Reproduce**
Steps to reproduce the behavior:
1. Go to '...'
2. Click on '....'
3. Scroll down to '....'
4. See error
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2085/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2085/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2086
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2086/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2086/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2086/events
|
https://github.com/materialsproject/pymatgen/pull/2086
| 824,824,023
|
MDExOlB1bGxSZXF1ZXN0NTg2OTk1NDky
| 2,086
|
docs: update compatibility guidance
|
{
"login": "ltalirz",
"id": 1053245,
"node_id": "MDQ6VXNlcjEwNTMyNDU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1053245?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ltalirz",
"html_url": "https://github.com/ltalirz",
"followers_url": "https://api.github.com/users/ltalirz/followers",
"following_url": "https://api.github.com/users/ltalirz/following{/other_user}",
"gists_url": "https://api.github.com/users/ltalirz/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ltalirz/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ltalirz/subscriptions",
"organizations_url": "https://api.github.com/users/ltalirz/orgs",
"repos_url": "https://api.github.com/users/ltalirz/repos",
"events_url": "https://api.github.com/users/ltalirz/events{/privacy}",
"received_events_url": "https://api.github.com/users/ltalirz/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I don't think the writeup is necessarily true. I don't want to give up calendar version just because we are temporarily doing semantic versioning due to the switch over to a namespace package. In future, it will still be calendar versioned.\r\n\r\nI think a more important statement is that if you happen to just use pymatgen for its core classes, it really doesn't matter in most instances whether you are using v2018, 2019 or 2021 or 2022. None of the core classes have undergone any backwards incompatible changes in the past few years.\r\n\r\nThose who work on the bleeding age of pymatgen will of course have to live with the nature of a research code, i.e., it is always changing!",
"\n[](https://coveralls.io/builds/37749342)\n\nCoverage decreased (-0.7%) to 82.985% when pulling **29e873e83c63c3c21e0213fd61ab1fe04709e300 on ltalirz:issue_2083_semver** into **3786ba9edd101a5418d636d6a89f6f687e0ddbff on materialsproject:master**.\n",
"> I don't think the writeup is necessarily true. I don't want to give up calendar version just because we are temporarily doing semantic versioning due to the switch over to a namespace package. In future, it will still be calendar versioned.\r\n\r\nOh, I see.\r\n\r\nSorry, I had understood you were going to follow the principle of incrementing the major version number upon breaking changes not just until 2022 but also going forward.\r\n\r\nFeel free to disregard this PR then.\r\n\r\n> I think a more important statement is that if you happen to just use pymatgen for its core classes, it really doesn't matter in most instances whether you are using v2018, 2019 or 2021 or 2022. None of the core classes have undergone any backwards incompatible changes in the past few years.\r\n\r\nSince you mention this, I went back through the aiida-core issue tracker looking for pymatgen-related issues from the last two years\r\n\r\nhttps://github.com/aiidateam/aiida-core/issues/3244\r\nhttps://github.com/aiidateam/aiida-core/issues/3297\r\nhttps://github.com/aiidateam/aiida-core/issues/3263\r\nhttps://github.com/aiidateam/aiida-core/issues/4339\r\nhttps://github.com/aiidateam/aiida-core/issues/4614\r\nhttps://github.com/aiidateam/aiida-core/issues/4687\r\nhttps://github.com/aiidateam/aiida-core/issues/4793\r\n\r\nOn closer inspection, you are right that most of these are not related to breaking changes. Most were caused by issues with the `numpy` dependency (which should hopefully be fixed now with the `pyproject.toml`) and other dependencies.\r\nOnly two were breaking changes - the `species_and_occu` and the recent removal of imports from the core.\r\n\r\nAll in all, I still think that for a tool like aiida it would still be useful if pymatgen was using semantic versioning... \r\nbut of course announcing them via warnings well in advance already helps as well.",
"Adding the `species_and_occu` example to the compatibility page may be a good idea though, if at least to give people a better idea of how often these changes have happened in the past.",
"@ltalirz Thanks. I will merge this and update the doc to reflect what Matt suggested.",
"@ltalirz I have clarified the compatibility doc based on your suggested edits. Thanks."
] | 2021-03-08T18:15:37
| 2021-03-09T16:45:49
|
2021-03-09T16:20:08Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Drafting an update of the Compatibility guide as requested by @mkhorton in
https://github.com/materialsproject/pymatgen/issues/2083#issuecomment-791597868
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [ ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [ ] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2086/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2086/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2086",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2086",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2086.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2086.patch",
"merged_at": "2021-03-09T16:20:07Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2087
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2087/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2087/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2087/events
|
https://github.com/materialsproject/pymatgen/issues/2087
| 824,935,904
|
MDU6SXNzdWU4MjQ5MzU5MDQ=
| 2,087
|
Defect code should be aware of MP GGA + U mixing correction
|
{
"login": "sthartman",
"id": 38861184,
"node_id": "MDQ6VXNlcjM4ODYxMTg0",
"avatar_url": "https://avatars.githubusercontent.com/u/38861184?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/sthartman",
"html_url": "https://github.com/sthartman",
"followers_url": "https://api.github.com/users/sthartman/followers",
"following_url": "https://api.github.com/users/sthartman/following{/other_user}",
"gists_url": "https://api.github.com/users/sthartman/gists{/gist_id}",
"starred_url": "https://api.github.com/users/sthartman/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/sthartman/subscriptions",
"organizations_url": "https://api.github.com/users/sthartman/orgs",
"repos_url": "https://api.github.com/users/sthartman/repos",
"events_url": "https://api.github.com/users/sthartman/events{/privacy}",
"received_events_url": "https://api.github.com/users/sthartman/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-03-08T20:43:42
| 2021-03-08T20:45:49
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hello,
The defects package in Pymatgen doesn't seem to be aware of the PBE + U correction terms used in the Materials Project for certain transition metal compounds. Supposing that I'm calculating the formation energy of a Fe vacancy in Fe3O4, pymatgen will calculate the uncorrected energy of a Defect_entry just as the difference between the pristine and defect cells. Various corrections can then be added to this Defect_entry, such as the Freysoldt charge correction, and then this Defect_entry can be used to obtain the formation energy for a certain set of chemical potentials. The chemical potentials are derived from the equilibrium between phases, such as Fe3O4/Fe2O3 or Fe3O4/FeO in this case.
However, these chemical potentials are usually pulled from the Materials Project, which applies a correction term to many metal oxides. So the initial energy of the Fe atom, in the Fe3O4, is not corrected in this way, but its final energy in Fe2O3 is corrected, resulting in an incorrect neutral vacancy formation energy which is negative by several eV. It's not hard to modify the Defect_entry with a custom MP_oxide_correction using my own scripts, but this could trip up less experienced users. (The issue didn't show up in the original PyCDT paper because they didn't test any transition metal oxides that would be subject to a GGA+U mixing correction)
I think it would be nice to have a built-in way of adding the GGA+U mixing correction to a Defect_entry, and a warning to users if they don't correct materials that would be eligible for the correction.
edit: This also applies to the oxide correction
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2087/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2087/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2088
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2088/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2088/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2088/events
|
https://github.com/materialsproject/pymatgen/pull/2088
| 826,154,833
|
MDExOlB1bGxSZXF1ZXN0NTg4MTc4OTA1
| 2,088
|
correct get_neighbor_list speed claim
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks.",
"\n[](https://coveralls.io/builds/37787014)\n\nCoverage decreased (-0.7%) to 82.993% when pulling **ded832a04ab268c1169c3b3a05d70bca6291d7ce on janosh:docs** into **f02c081348e17f0ecbd1d87b109710ade520ba4f on materialsproject:master**.\n"
] | 2021-03-09T16:01:41
| 2021-03-09T17:53:37
|
2021-03-09T16:12:36Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
`Structure.get_neighbor_list` claims to be orders of magnitude faster than `get_all_neighbors` which probably refers to `get_all_neighbors_old`. It's actually only about 2-3x faster than `get_all_neighbors`.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2088/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2088/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2088",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2088",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2088.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2088.patch",
"merged_at": "2021-03-09T16:12:36Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2089
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2089/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2089/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2089/events
|
https://github.com/materialsproject/pymatgen/issues/2089
| 826,534,680
|
MDU6SXNzdWU4MjY1MzQ2ODA=
| 2,089
|
ImportError: cannot import name 'Molecule' from 'pymatgen' (unknown location)
|
{
"login": "javiSYD",
"id": 66537664,
"node_id": "MDQ6VXNlcjY2NTM3NjY0",
"avatar_url": "https://avatars.githubusercontent.com/u/66537664?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/javiSYD",
"html_url": "https://github.com/javiSYD",
"followers_url": "https://api.github.com/users/javiSYD/followers",
"following_url": "https://api.github.com/users/javiSYD/following{/other_user}",
"gists_url": "https://api.github.com/users/javiSYD/gists{/gist_id}",
"starred_url": "https://api.github.com/users/javiSYD/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/javiSYD/subscriptions",
"organizations_url": "https://api.github.com/users/javiSYD/orgs",
"repos_url": "https://api.github.com/users/javiSYD/repos",
"events_url": "https://api.github.com/users/javiSYD/events{/privacy}",
"received_events_url": "https://api.github.com/users/javiSYD/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @jr31na,\r\n\r\nThis is part of a recent breaking change with pymatgen version 2022 onwards.\r\n\r\nThe older import `from pymatgen import Molecule` is what's known as a \"convenience import\", and was short-hand for the canonical import which is `from pymatgen.core.molecule import Molecule`. For technical reasons, these convenience imports have been removed. You can read more about this in our documentation here: \r\n\r\nhttps://pymatgen.org/compatibility.html#v2022-0-0\r\n\r\nYou have two options, you can either install an older pymatgen version, `pip install pymatgen==2021.3.3` for example, or you can start using the canonical imports.",
"Just to add that the workshop materials are from last year and precede this change, we will be updating the workshop materials for this year's workshop but have not done so yet.",
"Thank you very much @mkhorton It was very helpful. \r\n\r\nI find the Material Project very interesting. Although I'm a beginner in the subject and I have been learning a lot from your videos and documentation I want to use it to create something. \r\n\r\nI hope I can share it with you when I am a pro on the subject :)",
"Good luck!"
] | 2021-03-09T19:45:21
| 2021-03-09T20:49:04
|
2021-03-09T20:49:04Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
I am practicing with the Intro to Pymatgen workshop. I create a virtual environment on my computer and download [pip install pymatgen] in it, create a pymatgen.py file and write the first lines of code in Visual Studio Code.
**Code**
import pymatgen
from pymatgen import Molecule
c_monox = Molecule (["C", "O"], [[0., 0., 0.], [0., 0., 1.2]])
print (c_monox)
→ImportError: cannot import name 'Molecule' from 'pymatgen' (unknown location)
Also when I run [print(pymatgen.__version__)], I get the following error
→ AttributeError: module 'pymatgen' has no attribute '__version__'
**Expected behavior**
Full Formula (C1 01)
Reduced Formula: C0
Charge = 0.0, Spin Mult = 1
Sites(2)
0 C 0.000000 0.000000 0.000000
1 0 0.000000 0.000000. 1.200000
**Desktop (please complete the following information):**
- OS: [macOS Big Sur 11.2.1]
- Version [pymatgen==2022.0.3]
**Additional context**
Those are the libraries that are installed in the virtual environment when I run [pip freeze]
certifi==2020.12.5
chardet==4.0.0
cycler==0.10.0
decorator==4.4.2
future==0.18.2
idna==2.10
kiwisolver==1.3.1
matplotlib==3.3.4
monty==2021.3.3
mpmath==1.2.1
networkx==2.5
numpy==1.20.1
palettable==3.3.0
pandas==1.2.3
Pillow==8.1.2
plotly==4.14.3
pymatgen==2022.0.3
pyparsing==2.4.7
python-dateutil==2.8.1
pytz==2021.1
requests==2.25.1
retrying==1.3.3
ruamel.yaml==0.16.13
ruamel.yaml.clib==0.2.2
scipy==1.6.1
six==1.15.0
spglib==1.16.1
sympy==1.7.1
tabulate==0.8.9
uncertainties==3.1.5
urllib3==1.26.3
certifi==2020.12.5
chardet==4.0.0
cycler==0.10.0
decorator==4.4.2
future==0.18.2
idna==2.10
kiwisolver==1.3.1
matplotlib==3.3.4
monty==2021.3.3
mpmath==1.2.1
networkx==2.5
numpy==1.20.1
palettable==3.3.0
pandas==1.2.3
Pillow==8.1.2
plotly==4.14.3
pymatgen==2022.0.3
pyparsing==2.4.7
python-dateutil==2.8.1
pytz==2021.1
requests==2.25.1
retrying==1.3.3
ruamel.yaml==0.16.13
ruamel.yaml.clib==0.2.2
scipy==1.6.1
six==1.15.0
spglib==1.16.1
sympy==1.7.1
tabulate==0.8.9
uncertainties==3.1.5
urllib3==1.26.3
------
Thank you very much and I appreciate your help to continue learning and being able to create things with this library.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2089/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2089/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2090
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2090/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2090/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2090/events
|
https://github.com/materialsproject/pymatgen/issues/2090
| 830,831,260
|
MDU6SXNzdWU4MzA4MzEyNjA=
| 2,090
|
Pourbaix diagram for big supercells
|
{
"login": "hitarth64",
"id": 26575079,
"node_id": "MDQ6VXNlcjI2NTc1MDc5",
"avatar_url": "https://avatars.githubusercontent.com/u/26575079?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/hitarth64",
"html_url": "https://github.com/hitarth64",
"followers_url": "https://api.github.com/users/hitarth64/followers",
"following_url": "https://api.github.com/users/hitarth64/following{/other_user}",
"gists_url": "https://api.github.com/users/hitarth64/gists{/gist_id}",
"starred_url": "https://api.github.com/users/hitarth64/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/hitarth64/subscriptions",
"organizations_url": "https://api.github.com/users/hitarth64/orgs",
"repos_url": "https://api.github.com/users/hitarth64/repos",
"events_url": "https://api.github.com/users/hitarth64/events{/privacy}",
"received_events_url": "https://api.github.com/users/hitarth64/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @hitarth64 , make sure that the energy you use to create your extra `ComputedEntry` in this line\r\n\r\n`centry = ComputedEntry(Composition('Fe11 Co11 O64 Ru10'), -606.313)`\r\n\r\nis a _formation energy_ and not a raw DFT energy. All energies supplied to `PourbaixDiagram` need to be free energies of formation with respect to elements. I suspect that's what is causing this material to be overstabilized relative to the others.",
"Thanks. This helps! "
] | 2021-03-13T07:18:29
| 2021-06-27T15:15:53
|
2021-06-27T15:15:53Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
I am trying to model an alloy with three elements (Co11-Fe11-Ru10-O64) in it. I am working with a 96 atoms supercell. Calculating the Pourbaix diagram only yields that particular compound as the stable entity across the whole pH range which doesn't make sense.
**To Reproduce**
Steps to reproduce the behavior:
```
from pymatgen import MPRester
mpr = MPRester()
entries = mpr.get_pourbaix_entries(['Co','Fe','Ru'])
from pymatgen.entries.computed_entries import ComputedEntry
from pymatgen.core.composition import Composition
from pymatgen.analysis.pourbaix_diagram import PourbaixEntry
from pymatgen.analysis.pourbaix_diagram import PourbaixDiagram, PourbaixPlotter
# Energy is obtained from by performing DFT in consistency with MPRelaxSet parameters
centry = ComputedEntry(Composition('Fe11 Co11 O64 Ru10'), -606.313)
entry = PourbaixEntry(centry)
entries.append(entry)
pbx = PourbaixDiagram(entries,comp_dict={'Fe':11/32,'Co':11/32,'Ru':10/32})
print('Decomposition energy: ',pbx.get_decomposition_energy(entry,pH=0,V=1.23))
plotter = PourbaixPlotter(pbx)
plotter.get_pourbaix_plot().show()
```
**Expected behavior**
Expected behavior would be to see other competing phases too. I guess there is some problem or inconsistency within pymatgen when dealing with bigger supercells. Could you help me figure out if there is a specific parameter to be taken into account?
**Screenshots**

**Desktop (please complete the following information):**
- OS: Windows
- Version: 2021.2.16
|
{
"login": "hitarth64",
"id": 26575079,
"node_id": "MDQ6VXNlcjI2NTc1MDc5",
"avatar_url": "https://avatars.githubusercontent.com/u/26575079?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/hitarth64",
"html_url": "https://github.com/hitarth64",
"followers_url": "https://api.github.com/users/hitarth64/followers",
"following_url": "https://api.github.com/users/hitarth64/following{/other_user}",
"gists_url": "https://api.github.com/users/hitarth64/gists{/gist_id}",
"starred_url": "https://api.github.com/users/hitarth64/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/hitarth64/subscriptions",
"organizations_url": "https://api.github.com/users/hitarth64/orgs",
"repos_url": "https://api.github.com/users/hitarth64/repos",
"events_url": "https://api.github.com/users/hitarth64/events{/privacy}",
"received_events_url": "https://api.github.com/users/hitarth64/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2090/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2090/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2091
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2091/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2091/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2091/events
|
https://github.com/materialsproject/pymatgen/issues/2091
| 831,254,620
|
MDU6SXNzdWU4MzEyNTQ2MjA=
| 2,091
|
Import problems in Google Collab
|
{
"login": "VIGNESHinZONE",
"id": 45672459,
"node_id": "MDQ6VXNlcjQ1NjcyNDU5",
"avatar_url": "https://avatars.githubusercontent.com/u/45672459?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/VIGNESHinZONE",
"html_url": "https://github.com/VIGNESHinZONE",
"followers_url": "https://api.github.com/users/VIGNESHinZONE/followers",
"following_url": "https://api.github.com/users/VIGNESHinZONE/following{/other_user}",
"gists_url": "https://api.github.com/users/VIGNESHinZONE/gists{/gist_id}",
"starred_url": "https://api.github.com/users/VIGNESHinZONE/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/VIGNESHinZONE/subscriptions",
"organizations_url": "https://api.github.com/users/VIGNESHinZONE/orgs",
"repos_url": "https://api.github.com/users/VIGNESHinZONE/repos",
"events_url": "https://api.github.com/users/VIGNESHinZONE/events{/privacy}",
"received_events_url": "https://api.github.com/users/VIGNESHinZONE/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"We removed root level imports in the recent versions of pymatgen as part of a move to a more modular structure. Pls change from pymatgen import Structure to from pymatgen.core.structure import Structure."
] | 2021-03-14T20:41:46
| 2021-03-14T20:49:28
|
2021-03-14T20:49:28Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Unable to install Pymatgen properly on google collab,
```
!pip3 install pymatgen
from pymatgen import Structure
```
Could you please share a working code for a collab,
**To Reproduce**
Please run this [collab file](https://colab.research.google.com/drive/1XAb7q78cKf1n-MA3T9LjNscVgO0SRCsw?usp=sharing) for checking
Screenshot -
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2091/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2091/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2092
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2092/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2092/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2092/events
|
https://github.com/materialsproject/pymatgen/issues/2092
| 832,687,270
|
MDU6SXNzdWU4MzI2ODcyNzA=
| 2,092
|
Construct PointGroup class failed from the result of SpaceGroup for some space groups
|
{
"login": "hitliaomq",
"id": 18226671,
"node_id": "MDQ6VXNlcjE4MjI2Njcx",
"avatar_url": "https://avatars.githubusercontent.com/u/18226671?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/hitliaomq",
"html_url": "https://github.com/hitliaomq",
"followers_url": "https://api.github.com/users/hitliaomq/followers",
"following_url": "https://api.github.com/users/hitliaomq/following{/other_user}",
"gists_url": "https://api.github.com/users/hitliaomq/gists{/gist_id}",
"starred_url": "https://api.github.com/users/hitliaomq/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/hitliaomq/subscriptions",
"organizations_url": "https://api.github.com/users/hitliaomq/orgs",
"repos_url": "https://api.github.com/users/hitliaomq/repos",
"events_url": "https://api.github.com/users/hitliaomq/events{/privacy}",
"received_events_url": "https://api.github.com/users/hitliaomq/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Some of the point groups are just different settings of existing point groups. E.g., -4m2 is similar to -42m. Others are just more lengthy names. E.g., 321 is just 32. ",
"just as a comment, here is a way to convert them:\r\n```\r\npoint_group_conv = {'321' :'32', '312': '32', '3m1' :'3m', '31m': '3m',\r\n '-3m1' : '-3m', '-31m': '-3m', '-4m2': '-42m', '-62m': '-6m2' }\r\n\r\n#Using this dictionary we can correct to the standard point group notation.\r\ncorrected_point_groups = [point_group_conv.get(pg, pg) for pg in point_groups]\r\n```\r\n\r\nThis was used by @blondegeek in developing the FE database.",
"@shyuep @fraricci Thank you very much."
] | 2021-03-16T11:17:09
| 2021-03-21T01:29:35
|
2021-03-17T21:43:43Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Construct PointGroup class failed from the result of SpaceGroup for some space groups.
Failed space groups: 115-120, 149-154, 156-159, 162-165, 189-190
**To Reproduce**
Steps to reproduce the behavior:
```python
from pymatgen.symmetry.groups import SpaceGroup, PointGroup
for spacegroup in range(1, 231):
pointgroup = SpaceGroup.from_int_number(int(spacegroup)).point_group
try:
pointsym = PointGroup(pointgroup)
except Exception as e:
print(spacegroup)
print(pointgroup)
```
It will give the following results
```
[115, 116, 117, 118, 119, 120, 149, 150, 151, 152, 153, 154, 156, 157, 158, 159, 162, 163, 164, 165, 189, 190]
['-4m2', '-4m2', '-4m2', '-4m2', '-4m2', '-4m2', '312', '321', '312', '321', '312', '321', '3m1', '31m', '3m1', '31m', '-31m', '-31m', '-3m1', '-3m1', '-62m', '-62m']
[Finished in 0.7s]
```
**Desktop (please complete the following information):**
- OS: Windows,Linux
- Version 2022.0.4
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2092/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2092/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2093
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2093/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2093/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2093/events
|
https://github.com/materialsproject/pymatgen/issues/2093
| 832,863,302
|
MDU6SXNzdWU4MzI4NjMzMDI=
| 2,093
|
source code links from docs seem to be broken
|
{
"login": "rkurchin",
"id": 11233777,
"node_id": "MDQ6VXNlcjExMjMzNzc3",
"avatar_url": "https://avatars.githubusercontent.com/u/11233777?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkurchin",
"html_url": "https://github.com/rkurchin",
"followers_url": "https://api.github.com/users/rkurchin/followers",
"following_url": "https://api.github.com/users/rkurchin/following{/other_user}",
"gists_url": "https://api.github.com/users/rkurchin/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkurchin/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkurchin/subscriptions",
"organizations_url": "https://api.github.com/users/rkurchin/orgs",
"repos_url": "https://api.github.com/users/rkurchin/repos",
"events_url": "https://api.github.com/users/rkurchin/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkurchin/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Looks like @shyuep has fixed, thanks for reporting!"
] | 2021-03-16T14:31:19
| 2021-03-16T19:18:09
|
2021-03-16T19:18:09Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
The links in the documentation to the source code associated with that release currently give a 404 error, at least in the cases I have tried.
**To Reproduce**
Steps to reproduce the behavior:
1. Go to https://pymatgen.org/pymatgen.analysis.local_env.html
2. Click on any "source" link
4. See "not found" error
**Expected behavior**
I should have been brought to the source code...
**Desktop (please complete the following information):**
- MacOS
- Version 10.15.7
- browser: Vivaldi v3.6.2165.40
**Additional context**
I was going to just make a PR to fix it but I couldn't figure out how to make the actual correct link 😆
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2093/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2093/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2094
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2094/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2094/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2094/events
|
https://github.com/materialsproject/pymatgen/issues/2094
| 834,955,201
|
MDU6SXNzdWU4MzQ5NTUyMDE=
| 2,094
|
pymatgen.__version__ does not work
|
{
"login": "adelaiden",
"id": 29924845,
"node_id": "MDQ6VXNlcjI5OTI0ODQ1",
"avatar_url": "https://avatars.githubusercontent.com/u/29924845?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/adelaiden",
"html_url": "https://github.com/adelaiden",
"followers_url": "https://api.github.com/users/adelaiden/followers",
"following_url": "https://api.github.com/users/adelaiden/following{/other_user}",
"gists_url": "https://api.github.com/users/adelaiden/gists{/gist_id}",
"starred_url": "https://api.github.com/users/adelaiden/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/adelaiden/subscriptions",
"organizations_url": "https://api.github.com/users/adelaiden/orgs",
"repos_url": "https://api.github.com/users/adelaiden/repos",
"events_url": "https://api.github.com/users/adelaiden/events{/privacy}",
"received_events_url": "https://api.github.com/users/adelaiden/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"We do make `pymatgen.core.__version__` to be the correct version number. However, it is not the best way to properly get version numbers for packages since it depends on the developer actually updating it. You should use importlib or pkgutil. See https://stackoverflow.com/questions/20180543/how-to-check-version-of-python-modules",
"Great, thank you! That worked for me:\r\n\r\nPython 3.8.8 (default, Feb 24 2021, 13:46:16) \r\n[Clang 10.0.0 ] :: Anaconda, Inc. on darwin\r\nType \"help\", \"copyright\", \"credits\" or \"license\" for more information.\r\n\\>>> import pymatgen\r\n\\>>> import pkg_resources\r\n\\>>> pkg_resources.get_distribution('pymatgen').version\r\n'2022.0.4'",
"Specifically,\r\n\r\n```\r\nIn [1]: import pkg_resources\r\n\r\nIn [2]: pkg_resources.get_distribution('pymatgen').version\r\nOut[2]: '2022.0.4'\r\n```\r\n\r\n"
] | 2021-03-18T15:35:54
| 2021-03-18T15:48:56
|
2021-03-18T15:46:27Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hi, I have installed the latest version of pymatgen (v2022.0.4) in a fresh conda environment. Previously when checking the version I could open my command line, type `import pymatgen` and then `pymatgen.__version__` and I could obtain the version number. However, with the latest version (since the backwards incompatible transition to namespaces), it doesn't work.
Here is a sample output:
Python 3.8.8 (default, Feb 24 2021, 13:46:16)
[Clang 10.0.0 ] :: Anaconda, Inc. on darwin
Type "help", "copyright", "credits" or "license" for more information.
\>>> import pymatgen
\>>> pymatgen.\_\_version__
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
AttributeError: module 'pymatgen' has no attribute '\_\_version__'
\>>>
I can check the version of modules such as pymatgen.core by importing `import pymatgen.core` and `core.__version__`, but I would still like to be able to check the version of pymatgen, especially when designing packages that have dependencies on specific versions of pymatgen. Previous versions of pymatgen do appear to work.
Sample output for older pymatgen version, in different conda environment with Python 3.7:
Python 3.7.3 | packaged by conda-forge | (default, Dec 6 2019, 08:36:57)
[Clang 9.0.0 (tags/RELEASE_900/final)] :: Anaconda, Inc. on darwin
Type "help", "copyright", "credits" or "license" for more information.
\>>> import pymatgen
\>>> pymatgen.\_\_version__
'2020.9.14'
- OS: Mac, Catalina, 10.14.3
- New test conda environment with python 3.8
|
{
"login": "adelaiden",
"id": 29924845,
"node_id": "MDQ6VXNlcjI5OTI0ODQ1",
"avatar_url": "https://avatars.githubusercontent.com/u/29924845?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/adelaiden",
"html_url": "https://github.com/adelaiden",
"followers_url": "https://api.github.com/users/adelaiden/followers",
"following_url": "https://api.github.com/users/adelaiden/following{/other_user}",
"gists_url": "https://api.github.com/users/adelaiden/gists{/gist_id}",
"starred_url": "https://api.github.com/users/adelaiden/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/adelaiden/subscriptions",
"organizations_url": "https://api.github.com/users/adelaiden/orgs",
"repos_url": "https://api.github.com/users/adelaiden/repos",
"events_url": "https://api.github.com/users/adelaiden/events{/privacy}",
"received_events_url": "https://api.github.com/users/adelaiden/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2094/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2094/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2095
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2095/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2095/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2095/events
|
https://github.com/materialsproject/pymatgen/pull/2095
| 835,136,956
|
MDExOlB1bGxSZXF1ZXN0NTk1ODM3NzU0
| 2,095
|
Add convention to remove duplicate edges when i==j in StructureGraph + test
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/38064815)\n\nCoverage increased (+0.01%) to 83.652% when pulling **528ca078a121f8fced073529f43995ae5a5cdeef on graph-improvements** into **ac8f65e336008b6c219b8043f299cdeb50945c62 on master**.\n"
] | 2021-03-18T18:43:35
| 2021-03-18T20:48:38
|
2021-03-18T20:33:28Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
A fix to the graphs class that enforces a new convention to ensure duplicate edges are not added. This will only happen in rare instances normally (namely, small unit cells with NearNeighbor strategies where the cut-off is >> the length of a chemical bond).
Tagging @jmmshn @kim-jiyoon
While this convention fixes the present issue, I'm not convinced it's sufficient in all cases and will give it some further thought. It includes the mp-683887 6 Å cut-off example as a test-case but not the bow-tie example.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2095/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2095/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2095",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2095",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2095.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2095.patch",
"merged_at": "2021-03-18T20:33:28Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2096
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2096/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2096/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2096/events
|
https://github.com/materialsproject/pymatgen/issues/2096
| 835,760,814
|
MDU6SXNzdWU4MzU3NjA4MTQ=
| 2,096
|
Duplicated atoms in Structure.from_str compared to source string
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"You need to use mode=\"delete\" for merging. It is impossible to know whether some files have duplicate sites because they were wrongly entered, or it was intentional. Further, it is quite often that H atoms are excluded from the atomic positions but included in the composition of a file. So asserting a composition cannot work. Now, we can of course come up with some clever algorithm that will automatically merge sites and choose delete or sum depending on composition, but we'd be trying to divine the intent of the end user and there will be edge cases where something wrong is done and no error will be passed. I'd rather have an error and let the user figure out what they really want to do.\r\n\r\nThe code below will do what you want.\r\n\r\n```python\r\nfrom pymatgen.ext.cod import COD\r\n\r\nstructure = COD().get_structure_by_id('1510206')\r\nstructure.merge_sites(tol=0.003, mode=\"delete\")\r\n```\r\n\r\n\r\n",
"Makes sense, thanks for the detailed reply.",
"Yes. To add to this, COD is usually pretty good, but the amount of out-of-spec or malformed CIF files that are in the wild is staggering. The pymatgen CifParser is very forgiving in this respect (another issue is finite precision of atomic site co-ordinates which can make reconstruction via space group operations tricky), but it's impossible to get a good answer automatically in all cases.",
"Given a set of COD or ICSD IDs, is is possible to query with `MPRester` for corresponding MP IDs where they exist?",
"@janosh Yes, we anticipated this need. The best way is to use https://pymatgen.org/pymatgen.ext.matproj.html#pymatgen.ext.matproj.MPRester.find_structure . \r\n\r\n```\r\nfrom pymatgen.ext.cod import COD\r\n\r\nstructure = COD().get_structure_by_id('1510206')\r\nstructure.merge_sites(tol=0.003, mode=\"delete\")\r\nfrom pymatgen.ext.matproj import MPRester\r\nmpr = MPRester()\r\nmpr.find_structure(structure)\r\n```\r\n\r\nreturns ['mp-29138']\r\n\r\nThis uses StructureMatcher. It is somewhat slower than actually assigning a COD id in the database, but it is completely general and works for a structure from any source. Congrats on being the first person who have ever asked about this method after I coded it years ago!",
"Also to add, for icsd_ids, there is a simpler way since the database does store the icsd ids.\r\n\r\n```python\r\nfrom pymatgen.ext.matproj import MPRester\r\nmpr = MPRester()\r\nmpr.query({\"icsd_ids\": 1}, [\"task_id\"])\r\n```\r\n\r\n"
] | 2021-03-19T09:39:06
| 2021-03-21T20:10:28
|
2021-03-19T15:25:54Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
`Structure.from_str` duplicates atoms.
**To Reproduce**
```py
import requests
resp = requests.get("http://www.crystallography.net/cod/1510206.cif")
with open("1510206.cif", 'w') as file:
file.write(resp.text)
```
The above writes a CIF file with contents
```cif
...
_chemical_formula_sum 'Au K Se2'
_chemical_name_systematic 'K (Au Se2)'
_space_group_IT_number 127
_symmetry_space_group_name_Hall '-P 4 2ab'
_symmetry_space_group_name_H-M 'P 4/m b m'
_cell_angle_alpha 90
_cell_angle_beta 90
_cell_angle_gamma 90
_cell_formula_units_Z 2
_cell_length_a 7.699
_cell_length_b 7.699
_cell_length_c 3.665
_cell_volume 217.241
_citation_journal_id_ASTM JALCEU
_cod_data_source_file gold_264.cif
_cod_data_source_block Au1K1Se2
_cod_original_cell_volume 217.2414
_cod_original_formula_sum 'Au1 K1 Se2'
_cod_database_code 1510206
...
```
On the other hand,
```py
from pymatgen.ext.cod import COD
cod.get_structure_by_id('1510206')
```
returns
```sh
Structure Summary
Lattice
abc : 7.699 7.699 3.665
angles : 90.0 90.0 90.0
volume : 217.24141266499998
A : 7.699 0.0 4.714277853317737e-16
B : -4.714277853317737e-16 7.699 4.714277853317737e-16
C : 0.0 0.0 3.665
PeriodicSite: K (0.0000, 0.0000, 0.0000) [0.0000, 0.0000, 0.0000]
PeriodicSite: K (3.8495, 3.8495, 0.0000) [0.5000, 0.5000, 0.0000]
PeriodicSite: Au (3.8495, 0.0000, 0.0000) [0.5000, 0.0000, 0.0000]
PeriodicSite: Au (-0.0000, 3.8495, 0.0000) [0.0000, 0.5000, 0.0000]
PeriodicSite: Se (2.6716, 1.1764, 1.8325) [0.3470, 0.1528, 0.5000]
PeriodicSite: Se (6.5226, 2.6716, 1.8325) [0.8472, 0.3470, 0.5000]
PeriodicSite: Se (5.0274, 6.5226, 1.8325) [0.6530, 0.8472, 0.5000]
PeriodicSite: Se (1.1764, 5.0274, 1.8325) [0.1528, 0.6530, 0.5000]
PeriodicSite: Se (6.5211, 2.6731, 1.8325) [0.8470, 0.3472, 0.5000]
PeriodicSite: Se (5.0259, 6.5211, 1.8325) [0.6528, 0.8470, 0.5000]
PeriodicSite: Se (1.1779, 5.0259, 1.8325) [0.1530, 0.6528, 0.5000]
PeriodicSite: Se (2.6731, 1.1779, 1.8325) [0.3472, 0.1530, 0.5000]
```
i.e. double the amount of `Se`. Apparently all the `Se` are duplicated with a 0.002 A offset. If I pass `merge_tol`,
```py
cod.get_structure_by_id('1510206', merge_tol=0.003)
```
I get
```sh
ValueError: Species occupancies sum to more than 1!
```
How to proceed in this case?
Also, there should probably be a test for `Structure.from_str` that asserts the reduced compositions of the structure and the source string/file match.
**Desktop (please complete the following information):**
- OS: Mac & Linux
- Version: 2021.2.16
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2096/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2096/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2097
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2097/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2097/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2097/events
|
https://github.com/materialsproject/pymatgen/issues/2097
| 835,915,642
|
MDU6SXNzdWU4MzU5MTU2NDI=
| 2,097
|
SpacegroupAnalyzer.get_conventional_standard_structure additional documentation
|
{
"login": "tom-wood",
"id": 23095965,
"node_id": "MDQ6VXNlcjIzMDk1OTY1",
"avatar_url": "https://avatars.githubusercontent.com/u/23095965?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/tom-wood",
"html_url": "https://github.com/tom-wood",
"followers_url": "https://api.github.com/users/tom-wood/followers",
"following_url": "https://api.github.com/users/tom-wood/following{/other_user}",
"gists_url": "https://api.github.com/users/tom-wood/gists{/gist_id}",
"starred_url": "https://api.github.com/users/tom-wood/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/tom-wood/subscriptions",
"organizations_url": "https://api.github.com/users/tom-wood/orgs",
"repos_url": "https://api.github.com/users/tom-wood/repos",
"events_url": "https://api.github.com/users/tom-wood/events{/privacy}",
"received_events_url": "https://api.github.com/users/tom-wood/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks. Feel free to submit a PR to modify the doc. I don't think the alternative is needed though. There are a lot of different settings for space groups. People who work with space groups understand the differences."
] | 2021-03-19T12:09:15
| 2021-03-22T18:53:44
|
2021-03-22T18:53:44Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Is your feature request related to a problem? Please describe.**
It is a bit confusing (i.e. I spent quite a bit of time under a false impression) that the get_conventional_standard_structure method in the SpacegroupAnalyzer class does not necessarily return a Structure instance that is in the standard settting described by the International Tables of Crystallography, whereas get_refined_structure will. A pathological example would be the following, where the "conventional standard structure" is in fact 'P2_1mn':
```
from pymatgen.core import Structure, Lattice
from pymatgen.symmetry.analyzer import SpacegroupAnalyzer
latt = Lattice.orthorhombic(3, 1, 2)
struc = Structure(latt, ["H"] * 4 + ["Li"] * 4,
[(0.1, 0.2, 0.3), (0.4, 0.8, 0.8), (0.6, 0.8, 0.8), (0.9, 0.2, 0.3),
(0.2, 0.3, 0.4), (0.3, 0.7, 0.9), (0.7, 0.7, 0.9), (0.8, 0.3, 0.4)])
s = SpacegroupAnalyzer(struc)
con = s.get_conventional_standard_structure()
ref = s.get_refined_structure()
print(con.lattice.abc) # (1.0, 2.0, 3.0)
print(con.get_space_group_info()) # ('Pmn2_1', 31)
print(ref.lattice.abc) # (3.0, 1.0, 2.0)
print(ref.get_space_group_info()) # ('Pmn2_1', 31)
```
This _is_ subtly implicit in the documentation for get_conventional_standard_structure but would be even better if it were explicitly warned about.
**Describe the solution you'd like**
An additional line in the docstring for get_conventional_standard_structure with a warning that get_refined_structure should be used if the ITC settings are required. I'm happy to submit a PR to do this if needed.
**Describe alternatives you've considered**
I don't know if there would be any appetite for this, but maybe the get_space_group_info method of the Structure class could print a warning that the current structure is in a non-standard setting if the transformation matrix to standard settings does not match the identity matrix. I'm agnostic as to whether this would be a useful feature, however.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2097/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2097/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2098
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2098/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2098/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2098/events
|
https://github.com/materialsproject/pymatgen/issues/2098
| 836,552,872
|
MDU6SXNzdWU4MzY1NTI4NzI=
| 2,098
|
Please don't deprecate latexify
|
{
"login": "utf",
"id": 1330638,
"node_id": "MDQ6VXNlcjEzMzA2Mzg=",
"avatar_url": "https://avatars.githubusercontent.com/u/1330638?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/utf",
"html_url": "https://github.com/utf",
"followers_url": "https://api.github.com/users/utf/followers",
"following_url": "https://api.github.com/users/utf/following{/other_user}",
"gists_url": "https://api.github.com/users/utf/gists{/gist_id}",
"starred_url": "https://api.github.com/users/utf/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/utf/subscriptions",
"organizations_url": "https://api.github.com/users/utf/orgs",
"repos_url": "https://api.github.com/users/utf/repos",
"events_url": "https://api.github.com/users/utf/events{/privacy}",
"received_events_url": "https://api.github.com/users/utf/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Ok, this sounds fair to me, I can definitely see how these might be useful for people for formula strings that cannot be cast to Composition.\r\n\r\n> If you'd like to remove the duplication, I'd recommend that the Stringify mix-in class use these functions directly rather than duplicating the code in both places.\r\n\r\nThis seems like the best solution. We can retain the benefits of `Stringify` and use this going forwards while keeping these around for use cases like yours. Tagging @shyuep here in case there are objections.\r\n\r\nIf you'd like to submit a PR, please do so, otherwise I'll add it to my list of items next time I'm doing pymatgen maintenance work.",
"Ok, thanks. I will submit a PR.",
"This was done (deprecation notice removed) so closing the issue."
] | 2021-03-20T01:14:17
| 2021-09-30T01:14:18
|
2021-09-30T01:14:18Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
The `latexify`, `unicodeify`, etc have been deprecated in a recent update. We are now recommended to use the `Stringify` mix-in class.
Please can you not remove these functions. I use them fairly extensively in my code and in many cases switching to the `Stringify` mix-in class is not possible. For example, I have a string "n-CH3NH3PbI3". There is no way to convert this to a latex string directly using the mix-in class, whereas `latexify("n-CH3NH3PbI3")` gives the output I'd expect: `"n-CH$_3$NH$_3$PbI$_3$`.
If you'd like to remove the duplication, I'd recommend that the `Stringify` mix-in class use these functions directly rather than duplicating the code in both places.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2098/reactions",
"total_count": 2,
"+1": 2,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2098/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2099
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2099/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2099/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2099/events
|
https://github.com/materialsproject/pymatgen/issues/2099
| 837,058,773
|
MDU6SXNzdWU4MzcwNTg3NzM=
| 2,099
|
[2022+] Structure predictor raises TypeError
|
{
"login": "LuciusV",
"id": 11925386,
"node_id": "MDQ6VXNlcjExOTI1Mzg2",
"avatar_url": "https://avatars.githubusercontent.com/u/11925386?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/LuciusV",
"html_url": "https://github.com/LuciusV",
"followers_url": "https://api.github.com/users/LuciusV/followers",
"following_url": "https://api.github.com/users/LuciusV/following{/other_user}",
"gists_url": "https://api.github.com/users/LuciusV/gists{/gist_id}",
"starred_url": "https://api.github.com/users/LuciusV/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/LuciusV/subscriptions",
"organizations_url": "https://api.github.com/users/LuciusV/orgs",
"repos_url": "https://api.github.com/users/LuciusV/repos",
"events_url": "https://api.github.com/users/LuciusV/events{/privacy}",
"received_events_url": "https://api.github.com/users/LuciusV/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Sorry invalid report "
] | 2021-03-21T12:03:00
| 2021-03-21T12:25:28
|
2021-03-21T12:25:28Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
If I decorate structures with oxidation stater and use them in Substitutor
(from pymatgen.analysis.structure_prediction.substitutor import Substitutor)
then call to
sub = Substitutor(threshold=1E-4)
species = [Species('Li', 1), Species('B', 3), Species('O', -2), Species('F', -1)] # just an example
sub.pred_from_structures(target_species=species, structures_list=decorated_structures)
will give an exception:
`TypeError Traceback (most recent call last)
<ipython-input-8-ae259adef2d0> in <module>
----> 1 sub.pred_from_structures(target_species=species, structures_list=decorated_structures)
~/virtualenvs/science/lib/python3.7/site-packages/pymatgen/analysis/structure_prediction/substitutor.py in pred_from_structures(self, target_species, structures_list, remove_duplicates, remove_existing)
122 # check if: species are in the domain,
123 # and the probability of subst. is above the threshold
--> 124 els = s["structure"].composition.elements
125 if (
126 len(els) == len(permut)
~/virtualenvs/science/lib/python3.7/site-packages/pymatgen/core/structure.py in __getitem__(self, ind)
286
287 def __getitem__(self, ind):
--> 288 return self.sites[ind]
289
290 def __len__(self):
TypeError: list indices must be integers or slices, not str
`
If I use pickled decorated structures from earlier versions of pymatgen, this does not happen and Substitutor works normally
**To Reproduce**
Steps to reproduce the behavior:
1. Make some oxidation states decorated structures
2. Try to use them with Substitutor from pymatgen.analysis.structure_prediction.substitutor
3. See error
**Expected behavior**
Substitutor excected to work.
**Screenshots**
If applicable, add screenshots to help explain your problem.
**Desktop (please complete the following information):**
- OS: any
- Version 2022.0.3
**Additional context**
Add any other context about the problem here.
|
{
"login": "LuciusV",
"id": 11925386,
"node_id": "MDQ6VXNlcjExOTI1Mzg2",
"avatar_url": "https://avatars.githubusercontent.com/u/11925386?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/LuciusV",
"html_url": "https://github.com/LuciusV",
"followers_url": "https://api.github.com/users/LuciusV/followers",
"following_url": "https://api.github.com/users/LuciusV/following{/other_user}",
"gists_url": "https://api.github.com/users/LuciusV/gists{/gist_id}",
"starred_url": "https://api.github.com/users/LuciusV/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/LuciusV/subscriptions",
"organizations_url": "https://api.github.com/users/LuciusV/orgs",
"repos_url": "https://api.github.com/users/LuciusV/repos",
"events_url": "https://api.github.com/users/LuciusV/events{/privacy}",
"received_events_url": "https://api.github.com/users/LuciusV/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2099/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2099/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2100
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2100/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2100/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2100/events
|
https://github.com/materialsproject/pymatgen/pull/2100
| 837,969,344
|
MDExOlB1bGxSZXF1ZXN0NTk4MjI0NDE3
| 2,100
|
Small clarification to get_conventional_standard_structure docstring in symmetry/analyzer.py
|
{
"login": "tom-wood",
"id": 23095965,
"node_id": "MDQ6VXNlcjIzMDk1OTY1",
"avatar_url": "https://avatars.githubusercontent.com/u/23095965?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/tom-wood",
"html_url": "https://github.com/tom-wood",
"followers_url": "https://api.github.com/users/tom-wood/followers",
"following_url": "https://api.github.com/users/tom-wood/following{/other_user}",
"gists_url": "https://api.github.com/users/tom-wood/gists{/gist_id}",
"starred_url": "https://api.github.com/users/tom-wood/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/tom-wood/subscriptions",
"organizations_url": "https://api.github.com/users/tom-wood/orgs",
"repos_url": "https://api.github.com/users/tom-wood/repos",
"events_url": "https://api.github.com/users/tom-wood/events{/privacy}",
"received_events_url": "https://api.github.com/users/tom-wood/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/38148505)\n\nCoverage decreased (-0.6%) to 83.002% when pulling **8aae967617ad4b310f90d4b4140dca8457a8326d on tom-wood:master** into **2e5a4bb06950bf0d54bad07121435531eebc49a9 on materialsproject:master**.\n",
"Thanks for adding this @tom-wood. I agree this is confusing, at minimum being more explicit in the documentation definitely helps."
] | 2021-03-22T17:50:45
| 2021-03-22T18:55:09
|
2021-03-22T18:53:44Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
*Small addition to docstring of get_conventional_standard_structure to clarify that it does not produce the standard ITC setting (this is done by get_refined_structure).
Closes #2097.
## Additional dependencies introduced (if any)
* None
## TODO (if any)
None
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [ ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [ ] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2100/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2100/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2100",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2100",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2100.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2100.patch",
"merged_at": "2021-03-22T18:53:44Z"
}
|
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.