url
stringlengths 63
66
| repository_url
stringclasses 1
value | labels_url
stringlengths 77
80
| comments_url
stringlengths 72
75
| events_url
stringlengths 70
73
| html_url
stringlengths 52
56
| id
int64 3.39M
3.04B
| node_id
stringlengths 18
32
| number
int64 1
4.39k
| title
stringlengths 3
367
| user
dict | labels
listlengths 0
5
| state
stringclasses 2
values | locked
bool 2
classes | assignee
dict | assignees
listlengths 0
2
| milestone
null | comments
listlengths 0
30
| created_at
timestamp[s]date 2012-02-26 12:10:46
2025-05-02 18:11:41
| updated_at
timestamp[s]date 2012-02-26 14:52:41
2025-05-02 18:16:27
| closed_at
stringlengths 0
20
| author_association
stringclasses 3
values | type
stringclasses 1
value | active_lock_reason
stringclasses 2
values | sub_issues_summary
dict | body
stringlengths 0
119k
| closed_by
dict | reactions
dict | timeline_url
stringlengths 72
75
| performed_via_github_app
null | state_reason
stringclasses 4
values | is_pull_request
bool 2
classes | draft
bool 2
classes | pull_request
dict |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
https://api.github.com/repos/materialsproject/pymatgen/issues/2201
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2201/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2201/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2201/events
|
https://github.com/materialsproject/pymatgen/issues/2201
| 942,046,369
|
MDU6SXNzdWU5NDIwNDYzNjk=
| 2,201
|
`SpacegroupAnalyzer.get_symmetrized_structure()` weird results on structures with magnetic moments
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"would an `inc_magmoms` arg to `SpacegroupAnalyzer` be an acceptable addition `True` by default but can be turned off if all you care about is the structural spacegroup? ",
"> would an inc_magmoms arg to SpacegroupAnalyzer be an acceptable addition True by default but can be turned off if all you care about is the structural spacegroup?\r\n\r\nYes, I think so.\r\n\r\nI haven't had chance to look at this specific example yet, but this is an issue.\r\n\r\nIt's difficult to know what the \"correct\" action is here; without vector magnetic moments we're in an unfortunate in-between where we cannot properly apply magnetic space groups, but using crystallographic space groups where, essentially, atoms with different moments are treated as non-equivalent are also not appropriate (for example, if you use `get_symmetrized_structure()` on a structure with a specific magnetic ordering, you might find it undesirable for this ordering to be stripped).\r\n\r\nThis problem is exacerbated by the fact that, due to numerical noise, often magnetic moments on atoms are not going to be exactly equal. If I have to handle structures like this myself, the first step I take is usually to round off the magmoms."
] | 2021-07-12T13:10:06
| 2021-07-14T04:46:48
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
If I query a structure with magnetic moments defined i.e. `mp-1228485` and run it through `SpacegroupAnalyzer` then call `get_symmetrized_structure()` it returns
```
SymmetrizedStructure
Full Formula (Al12 Cr4 O24)
Reduced Formula: Al3CrO6
Spacegroup: R3 (146)
abc : 7.165227 7.166083 8.451375
angles: 113.162862 113.161083 85.839799
Sites (40)
# SP a b c Wyckoff magmom
--- ---- -------- -------- -------- --------- --------
0 Al 0.973155 0.473169 0.648738 4a 0.002
1 Al 0.975235 0.975237 0.650128 4a -0.002
2 Al 0.52897 0.028961 0.352616 4a 0.002
3 Cr 0.52741 0.527402 0.35158 4a -0.002
4 O 0.044653 0.704117 0.594498 4b 0.001
5 O 0.780519 0.968294 0.747281 4b -0.001
6 O 0.798074 0.950804 0.406879 4b -0.001
7 O 0.729983 0.528205 0.244689 4b 0.001
8 O 0.709869 0.548396 0.607886 4b -0
9 O 0.447642 0.810549 0.405891 4b 0
```
If I go to bilbao crystallographic server and look up spacegroup 146's wyckoff sets we see that a sites and b sites have multiplicities 3 and 9 respectively (https://www.cryst.ehu.es/cgi-bin/cryst/programs/nph-normsets?from=wycksets&gnum=146).
Given only the above if I applied the symmetry operations I would end up with very large numbers of duplicated sites and wouldn't know which magmoms should be where when merging sites.
Instead what I feel `SymmetrizedStructure` should be can be obtained as `SpacegroupAnalyzer(SpacegroupAnalyzer(struct).get_refined_structure()).get_symmetrized_structure()`
```
SymmetrizedStructure
Full Formula (Al9 Cr3 O18)
Reduced Formula: Al3CrO6
Spacegroup: R3 (146)
abc : 4.879461 4.879461 13.278842
angles: 90.000000 90.000000 120.000000
Sites (30)
# SP a b c Wyckoff
--- ---- -------- -------- -------- ---------
0 Al 0 0 0.148822 3a
1 Al 0.333333 0.666667 0.316863 3a
2 Al 0.666667 0.333333 0.186033 3a
3 Cr 0.333333 0.666667 0.018331 3a
4 O 0.685458 0.995592 0.249595 9b
5 O 0.360234 0.043126 0.086106 9b
```
However doing this discards the magmom information (not important for me but probably important for pymatgen).
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2201/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2201/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2202
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2202/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2202/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2202/events
|
https://github.com/materialsproject/pymatgen/pull/2202
| 944,258,023
|
MDExOlB1bGxSZXF1ZXN0Njg5NzY2MTQ5
| 2,202
|
[WIP - Needs Test] add include magmom toggle to `SpacegroupAnalyzer` #2201
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/41358873)\n\nCoverage decreased (-0.6%) to 83.065% when pulling **8a05fcea91d96cea91a76e81ac8cd8e89d895f92 on CompRhys:magmom-toggle** into **5d600ca610d1a4d3e1532293e5f87062332b9a26 on materialsproject:master**.\n",
"@mkhorton If I need to include a new data file for a test of a `Structure` with magmoms that behaves \"weirdly\" in the way outlined in #2201 what's the current best practise to storing those files within the codebase (#2197)? or this could be merged without an extra test as no bug as such is being fixed that really needs testing",
"A test would definitely be desired. There are three options, (1) pull a test structure from the test framework and add magmoms to it, (2) add a .json to test_files, (3) construct a Structure object within the test itself (either directly or via a minimal from_dict).",
"@mkhorton This is actually more of an issue than I initially thought and it's not as simple as adding a toggle to get the multiplicities right due to the way the code deals with the site properties. When I use the code as is the correct magmoms are passed to the analyzer and this causes wyckoff sites to be split up by spglib (which is what I wanted to avoid) however in the called to `get_symmetrised_structure` the code just accesses the magmoms as a list as stands and then miss-assigns them to elements (see below).\r\n\r\n```\r\nFull Formula (Al12 Cr4 O24)\r\nReduced Formula: Al3CrO6\r\nabc : 7.165227 7.166083 8.451375\r\nangles: 113.162862 113.161083 85.839799\r\nSites (40)\r\n # SP a b c magmom\r\n--- ---- -------- -------- -------- --------\r\n 0 Al 0.973155 0.473169 0.648738 0.002\r\n 1 Al 0.473143 0.973159 0.648728 -0.002\r\n 2 Al 0.223212 0.223202 0.148823 0.002\r\n 3 Al 0.723181 0.723173 0.148783 -0.002\r\n 4 Al 0.975235 0.975237 0.650128 0.001\r\n 5 Al 0.47525 0.47523 0.650127 -0.001\r\n 6 Al 0.225225 0.725244 0.150164 -0.001\r\n 7 Al 0.725226 0.225237 0.150168 0.001\r\n 8 Al 0.52897 0.028961 0.352616 -0\r\n 9 Al 0.028977 0.528957 0.352623 0\r\n 10 Al 0.779009 0.779016 0.852669 -0\r\n 11 Al 0.279008 0.279024 0.852677 0\r\n 12 Cr 0.52741 0.527402 0.35158 -2.949\r\n 13 Cr 0.277385 0.77739 0.851557 -2.949\r\n 14 Cr 0.027476 0.027493 0.351636 2.949\r\n 15 Cr 0.777495 0.27745 0.851649 2.949\r\n 16 O 0.044653 0.704117 0.594498 0.013\r\n 17 O 0.544639 0.204079 0.594503 -0.013\r\n 18 O 0.294562 0.454044 0.094441 0.013\r\n 19 O 0.794545 0.954029 0.094434 -0.013\r\n 20 O 0.780519 0.968294 0.747281 -0.013\r\n 21 O 0.280508 0.468296 0.747287 0.013\r\n 22 O 0.030439 0.718202 0.247303 -0.013\r\n 23 O 0.530433 0.218227 0.247322 0.013\r\n 24 O 0.798074 0.950804 0.406879 -0.013\r\n 25 O 0.298077 0.450856 0.406914 0.013\r\n 26 O 0.047946 0.700753 0.906898 0.013\r\n 27 O 0.547857 0.20076 0.906931 -0.013\r\n 28 O 0.729983 0.528205 0.244689 0.008\r\n 29 O 0.229964 0.028174 0.244634 -0.008\r\n 30 O 0.979915 0.27833 0.74455 -0.004\r\n 31 O 0.479916 0.778318 0.744534 0.004\r\n 32 O 0.709869 0.548396 0.607886 0.006\r\n 33 O 0.20987 0.048404 0.607914 -0.006\r\n 34 O 0.959754 0.298432 0.107708 -0.007\r\n 35 O 0.459708 0.798444 0.107638 0.007\r\n 36 O 0.447642 0.810549 0.405891 0.007\r\n 37 O 0.947641 0.310582 0.405913 -0.007\r\n 38 O 0.697657 0.560783 0.906042 -0.004\r\n 39 O 0.197676 0.060776 0.906051 0.004\r\n```\r\n\r\n```\r\nFull Formula (Al12 Cr4 O24)\r\nReduced Formula: Al3CrO6\r\nSpacegroup: R3 (146)\r\nabc : 7.165227 7.166083 8.451375\r\nangles: 113.162862 113.161083 85.839799\r\nSites (40)\r\n # SP a b c Wyckoff magmom\r\n--- ---- -------- -------- -------- --------- --------\r\n 0 Al 0.973155 0.473169 0.648738 4a 0.002\r\n 1 Al 0.975235 0.975237 0.650128 4a -0.002\r\n 2 Al 0.52897 0.028961 0.352616 4a 0.002\r\n 3 Cr 0.52741 0.527402 0.35158 4a -0.002\r\n 4 O 0.044653 0.704117 0.594498 4b 0.001\r\n 5 O 0.780519 0.968294 0.747281 4b -0.001\r\n 6 O 0.798074 0.950804 0.406879 4b -0.001\r\n 7 O 0.729983 0.528205 0.244689 4b 0.001\r\n 8 O 0.709869 0.548396 0.607886 4b -0\r\n 9 O 0.447642 0.810549 0.405891 4b 0\r\n```",
"closing this PR but leaving #2201 open as this fix doesn't actually do what we need it to do for robustness in the sense I wanted."
] | 2021-07-14T10:02:08
| 2021-09-01T14:10:58
|
2021-09-01T14:10:32Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
re #2201
add toggle on whether to include magmoms in spglib input in order to has equivalent sites correctly map to wyckoff positions without duplicates.
- [ ] Tests added for checking correct Wyckoff position multiplicities
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2202/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2202/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2202",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2202",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2202.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2202.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2203
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2203/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2203/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2203/events
|
https://github.com/materialsproject/pymatgen/issues/2203
| 946,548,760
|
MDU6SXNzdWU5NDY1NDg3NjA=
| 2,203
|
`LocalStructOrderParams` is not translationally invariant when specifying neighbor indices
|
{
"login": "qai222",
"id": 20326044,
"node_id": "MDQ6VXNlcjIwMzI2MDQ0",
"avatar_url": "https://avatars.githubusercontent.com/u/20326044?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/qai222",
"html_url": "https://github.com/qai222",
"followers_url": "https://api.github.com/users/qai222/followers",
"following_url": "https://api.github.com/users/qai222/following{/other_user}",
"gists_url": "https://api.github.com/users/qai222/gists{/gist_id}",
"starred_url": "https://api.github.com/users/qai222/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/qai222/subscriptions",
"organizations_url": "https://api.github.com/users/qai222/orgs",
"repos_url": "https://api.github.com/users/qai222/repos",
"events_url": "https://api.github.com/users/qai222/events{/privacy}",
"received_events_url": "https://api.github.com/users/qai222/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@nisse3000 do you have any thoughts on this issue?",
"@mkhorton, sure. That looks like a neighbor-finding/image-using \"issue\". \r\n\r\nIf you just print out the distances between the locations of the O neighbors from the V central atom using the \"low-fi\" index mechanism, you realize that 2 neighbors don't seem to be in the closest image to the V atom because they exhibit distances way larger than 2:\r\n\r\n-------------------------------------------------------------------\r\n\r\nfrom pymatgen.core.structure import Structure\r\ncifstring = \"\"\"data_primitive\r\n_cell_length_a 8.548\r\n_cell_length_b 9.168\r\n_cell_length_c 12.58\r\n_cell_angle_alpha 89.467\r\n_cell_angle_beta 72.849\r\n_cell_angle_gamma 65.478\r\n\r\n_symmetry_space_group_name_H-M 'P 1'\r\nloop_\r\n_space_group_symop_operation_xyz\r\n'x,y,z'\r\n\r\nloop_\r\n_atom_site_label\r\n_atom_site_fract_x\r\n_atom_site_fract_y\r\n_atom_site_fract_z\r\nV 0.77756 0.13214 0.17138 \r\nO 0.74544 -0.02747 0.16792 \r\nO 1.04270 0.06785 0.12723 \r\nO 0.78324 0.18577 0.01576 \r\nO 0.77380 0.14078 0.33290 \r\nO 0.51586 0.30547 0.22633 \r\nO 0.74690 0.40761 0.19451\"\"\"\r\ns = Structure.from_str(cifstring, \"cif\")\r\nprint(s)\r\n\r\nFull Formula (V1 O6)\r\nReduced Formula: VO6\r\nabc : 8.548000 9.168000 12.580000\r\nangles: 89.467000 72.849000 65.478000\r\nSites (7)\r\n # SP a b c\r\n--- ---- ------- ------- -------\r\n 0 V 0.77756 0.13214 0.17138\r\n 1 O 0.74544 0.97253 0.16792\r\n 2 O 0.0427 0.06785 0.12723\r\n 3 O 0.78324 0.18577 0.01576\r\n 4 O 0.7738 0.14078 0.3329\r\n 5 O 0.51586 0.30547 0.22633\r\n 6 O 0.7469 0.40761 0.19451\r\n\r\nimport numpy as np\r\nfor i in range(1, 7):\r\n print(np.linalg.norm(s.sites[0].coords - s.sites[i].coords))\r\n\r\n7.595027732481124\r\n6.726904760009047\r\n2.005627629286983\r\n2.0244449105916953\r\n2.0408074010602935\r\n2.4394302343682424\r\n\r\n-------------------------------------------------------------------\r\n\r\nIf you, on the other hand, use the standard mechanism for neighbor finding in LocalStructOrderParams, you obtain:\r\n\r\n-------------------------------------------------------------------\r\n\r\nfrom pymatgen.analysis.local_env import VoronoiNN\r\nvnn = VoronoiNN(tol=0.0, targets=None)\r\nneighsites = vnn.get_nn(s, 0)\r\nfor neighsite in neighsites:\r\n print(np.linalg.norm(s.sites[0].coords - neighsite.coords))\r\n\r\n1.600085415591695\r\n2.4394302343682424\r\n2.0408074010602935\r\n2.005627629286983\r\n1.9867913140898044\r\n2.0244449105916953\r\n\r\n-------------------------------------------------------------------\r\n\r\nA different image of a neighboring site surrounds a central atom (possibly) in a different manner than the closest image - especially if the cell angles are not 90 degrees. So, that's why @qai222 obtains different OP values between the two \"los.get_order_parameters(...)\" calls\r\n\r\np1 = los.get_order_parameters(s, 0, indices_neighs=[1, 2, 3, 4, 5, 6])\r\n\r\nand\r\n\r\np2 = los.get_order_parameters(s, 0)\r\n\r\nDoes that make sense?",
"Quick addition: the documentation of the \"get_order_parameters\"-function underscores that \"no nearest images of neighbors are determined when providing an index list.\"",
"Thanks, @nisse3000. If I understand correctly, when `indices_neighs` is provided, the neighbors may not be the image closest to the center atom. Sounds like this can be solved by using `pbc_shortest_vectors`.\r\n\r\nA follow-up question: This VO6 unit has \"pent_pyr\" (0.549) > \"oct\" (0.519), does this suggest it's more like a pentagonal pyramid than an octahedron? To me, it looks just like a distorted octahedron.",
"Welcome, @qai222. Yes, if you want to have it be the closest image, you first have to take care of that manually; that is, before you provide \"get_order_parameters\" the list of indices. \r\n\r\nNumerically, it should be interpreted that way. However, when you think a bit more about the \"rough estimate\" character of the order parameters as likelihoods of pattern resemblance scores, you will probably end up concluding that those 2 values are pretty close to each other and that clear cut conclusions are tricky in this area of science. In the end, we are talking about what individual researchers think/perceive as \"looking rather like an octahedron than a pentagonal pyramid\". To bring that a bit more into perspective: if q_oct would have been 0.9 and q_pent_pyr = 0.5, I believe that most researchers would have interpreted the coordination environment as being a (clear) octahedron rather than a pentagonal pyramid. ",
"@mkhorton, I get the feeling that the \"purported\" implementation problems/issues are addressed/do not exist. I believe that we can close the issue.\r\n\r\nIs that OK with you, @qai222?",
"Sure, you can close the issue. \r\n\r\nFor what it is worth, I end up creating a `Structure` object for the local structure when I need to specify neighbor indices."
] | 2021-07-16T19:36:04
| 2021-07-21T17:30:09
|
2021-07-21T17:30:09Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Values of `LocalStructOrderParams` change after translating sites when specifying neighbor indices.
Also, IIUC, LSOP thinks this distorted octahedron is similar to a pentagonal pyramid.
**To Reproduce**
Steps to reproduce the behavior:
```
from pymatgen.analysis.local_env import LocalStructOrderParams
from pymatgen.core.structure import Structure
# a distorted octahedron, there are fractional coords with value <0 or >1
cifstring = """data_primitive
_cell_length_a 8.548
_cell_length_b 9.168
_cell_length_c 12.58
_cell_angle_alpha 89.467
_cell_angle_beta 72.849
_cell_angle_gamma 65.478
_symmetry_space_group_name_H-M 'P 1'
loop_
_space_group_symop_operation_xyz
'x,y,z'
loop_
_atom_site_label
_atom_site_fract_x
_atom_site_fract_y
_atom_site_fract_z
V 0.77756 0.13214 0.17138
O 0.74544 -0.02747 0.16792
O 1.04270 0.06785 0.12723
O 0.78324 0.18577 0.01576
O 0.77380 0.14078 0.33290
O 0.51586 0.30547 0.22633
O 0.74690 0.40761 0.19451"""
# before translation
s = Structure.from_str(cifstring, "cif")
los = LocalStructOrderParams(types=["hex_plan_max", "pent_pyr", "oct"])
# if specify neighbor indices
p1 = los.get_order_parameters(s, 0, indices_neighs=[1, 2, 3, 4, 5, 6])
print(p1)
# [0.4477591572605329, 0.5664395961928602, 0.23852499675295133]
# if do not specify
p2 = los.get_order_parameters(s, 0)
print(p2)
# [0.19179211488913345, 0.549512347792701, 0.5197264306316182]
# specify neighbor indices and translate the cluster s.t. it is inside the box
s.translate_sites(list(range(len(s))), vector=(-0.2, 0.2, 0.2))
# los = LocalStructOrderParams(types=["hex_plan_max","pent_pyr", "oct"])
p3 = los.get_order_parameters(s, 0, indices_neighs=[1, 2, 3, 4, 5, 6])
print(p3)
# [0.19179211488913417, 0.5495123477927011, 0.5197264306316184]
```
**Expected behavior**
p1 close to p2 close to p3
**Desktop (please complete the following information):**
- OS: Ubuntu 20.04.2
|
{
"login": "qai222",
"id": 20326044,
"node_id": "MDQ6VXNlcjIwMzI2MDQ0",
"avatar_url": "https://avatars.githubusercontent.com/u/20326044?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/qai222",
"html_url": "https://github.com/qai222",
"followers_url": "https://api.github.com/users/qai222/followers",
"following_url": "https://api.github.com/users/qai222/following{/other_user}",
"gists_url": "https://api.github.com/users/qai222/gists{/gist_id}",
"starred_url": "https://api.github.com/users/qai222/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/qai222/subscriptions",
"organizations_url": "https://api.github.com/users/qai222/orgs",
"repos_url": "https://api.github.com/users/qai222/repos",
"events_url": "https://api.github.com/users/qai222/events{/privacy}",
"received_events_url": "https://api.github.com/users/qai222/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2203/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2203/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2204
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2204/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2204/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2204/events
|
https://github.com/materialsproject/pymatgen/issues/2204
| 951,806,585
|
MDU6SXNzdWU5NTE4MDY1ODU=
| 2,204
|
Inconsistency between `formula` and `reduced_formula` for diatomic molecules and Ions
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Overloading `Ion.get_reduced_formula_and_factor` would have the additional advantage of enabling special handling of important aqueous species like `OH[-]` (which otherwise shows up as `HO[-]`) and differentiating between `OH[-]` and hydrogen peroxide, `H2O2`. The fact that we can't distinguish these two species (or other O-H species) causes some problems for certain Pourbaix diagrams."
] | 2021-07-23T18:24:41
| 2023-08-13T16:35:00
|
2023-08-13T16:35:00Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
The output of `Composition.formula` and `Composition.reduced_formula` is counterintuitive for formulas listed in [`special_formulas`](https://github.com/materialsproject/pymatgen/blob/937eff5cc83febdea57258c0488a5b81cb550506/pymatgen/core/composition.py#L78), especially diatomic gases. In addition, the output of `reduced_formula` is not correct for `Ion` objects whose formulas appear in `special_formulas` (`Ion` inherits from `Composition`).
Specifically, for single atom `Composition`, in these cases the `formula` indicates fewer atoms than the `reduced_formula`. The same comment applies to the `composition` and `reduced_composition` attributes.
For Ions in `special_formulas`, the formula returned by `reduced_formula` is generally not the way the ion is canonically represented. This is problematic because the `reduced_formula`, with the charge delineated in brackets, is the clearest way to uniquely identify an ionic species in, e.g. Pourbaix diagrams.
**To Reproduce**
```
# chlorine
>>> from pymatgen.core import Composition
>>> Composition('Cl').formula
'Cl1'
>>> from pymatgen.core import Composition
>>> Composition('Cl').reduced_formula
'Cl2'
# chloride ion
>>> from pymatgen.core.ion import Ion
>>> Ion.from_formula('Cl-').formula
'Cl1 -1'
>>> from pymatgen.core.ion import Ion
>>> Ion.from_formula('Cl-').reduced_formula
'Cl2[2-]'
# hydroxide ion
from pymatgen.core.ion import Ion
Ion.from_formula('OH-').formula
'H1 O1 -1'
from pymatgen.core.ion import Ion
Ion.from_formula('OH-').reduced_formula
'H2O2[2-]'
```
**Expected behavior**
`formula` should not have more atoms than `reduced_formula`. For ions, special formulas should be ignored and `reduced_formula` should return, e.g.`'Cl[-]` or `OH[-]`.
**Suggested fixes**
A minimal fix would be to overload `Ion.reduced_formula` and patch it such that it ignores the `special_formulas`. However, I also think it would be appropriate to change `Composition` to recognize when the composition has only a single atom, e.g. at [this line ](https://github.com/materialsproject/pymatgen/blob/937eff5cc83febdea57258c0488a5b81cb550506/pymatgen/core/composition.py#L388), change to
```
if self.num_atoms > 1 and formula in Composition.special_formulas:
formula = Composition.special_formulas[formula]
factor /= 2
```
My proposed change to `Composition` passes all unit tests in `pymatgen.core` without modification.
Since this would change a core class I thought it best to seek comments from @mkhorton and @shyuep before opening a PR. Please let me know what you think the best resolution would be here. Thanks!
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2204/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2204/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2205
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2205/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2205/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2205/events
|
https://github.com/materialsproject/pymatgen/issues/2205
| 954,089,396
|
MDU6SXNzdWU5NTQwODkzOTY=
| 2,205
|
Problems with saving figures
|
{
"login": "AnshulyadavUnofficial",
"id": 71870780,
"node_id": "MDQ6VXNlcjcxODcwNzgw",
"avatar_url": "https://avatars.githubusercontent.com/u/71870780?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/AnshulyadavUnofficial",
"html_url": "https://github.com/AnshulyadavUnofficial",
"followers_url": "https://api.github.com/users/AnshulyadavUnofficial/followers",
"following_url": "https://api.github.com/users/AnshulyadavUnofficial/following{/other_user}",
"gists_url": "https://api.github.com/users/AnshulyadavUnofficial/gists{/gist_id}",
"starred_url": "https://api.github.com/users/AnshulyadavUnofficial/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/AnshulyadavUnofficial/subscriptions",
"organizations_url": "https://api.github.com/users/AnshulyadavUnofficial/orgs",
"repos_url": "https://api.github.com/users/AnshulyadavUnofficial/repos",
"events_url": "https://api.github.com/users/AnshulyadavUnofficial/events{/privacy}",
"received_events_url": "https://api.github.com/users/AnshulyadavUnofficial/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 4705117170,
"node_id": "LA_kwDOACgets8AAAABGHJj8g",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/question",
"name": "question",
"color": "DB336A",
"default": true,
"description": "Questions about functionality and design choices"
},
{
"id": 5322879170,
"node_id": "LA_kwDOACgets8AAAABPUSwwg",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/help",
"name": "help",
"color": "5ED16E",
"default": false,
"description": "Q&A support-style issues"
}
] |
closed
| true
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-07-27T17:11:48
| 2023-05-24T19:23:31
|
2023-05-24T19:23:31Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I have been trying to save the output of plotter.get_pourbaix_plot. Earlier I could simply putthe pyplot object in a PDFPages objects like below to save the figure. However, due to changes in pymatgen get_pourbaix_plot returns a pyplot, which in theory should be the same as matplotlib.pyplot. The code below tries to save the figure produced by the returned pyplot, however the pdf generated contains a blank image. I am able to save the image manually after it is displayed, however that is undesirable as we would be generating thousands of such images.
```py
# Import necessary tools from pymatgen
from matplotlib import pyplot
from matplotlib.backends.backend_pdf import PdfPages
from pymatgen.ext.matproj import MPRester
from pymatgen.analysis.pourbaix_diagram import PourbaixDiagram
from pymatgen.analysis.pourbaix_diagram import PourbaixPlotter
import os
class ToMakePourbaix(object):
def __init__(self, ele, resultsdir):
self.ele = ele
self.mpr = MPRester("Your ID") #Establishes connection to the MPRester API
self.resultdir = resultsdir
# Create result directory if it does not exist
if not os.path.exists(self.resultdir):
os.mkdir(self.resultdir)
self.plotpdf = PdfPages(os.path.join(self.resultdir, 'stability.pdf')) #To store figures in a pdf file
def drawPourbaix(self):
entries = self.mpr.get_pourbaix_entries(self.ele)
pbd = PourbaixDiagram(entries)
plotter = PourbaixPlotter(pbd)
plt2 = plotter.get_pourbaix_plot() # Output is a pyplot. Equivalent to matplotlib.pyplot in theory
fig = plt2.figure()
fig.savefig("foo.pdf", bbox_inches='tight') #Should save the figure
self.plotpdf.savefig(plt2) #This used to work earlier. However, it does not now.
# savefig() requires a figure object
self.plotpdf.savefig(fig) #This should work but does not
entry = ToMakePourbaix(ele = ["Cu"], resultsdir='Enter your directory')
en = entry.drawPourbaix()
```
In order to use the above code entry the name of your directory where you want to save the figure.
This is the expected output.

This is the output of foo.pdf
[foo.pdf](https://github.com/materialsproject/pymatgen/files/6887452/foo.pdf)
The output pf stability.pdf is empty.
Any suggestions would be much appreciated
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2205/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2205/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2206
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2206/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2206/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2206/events
|
https://github.com/materialsproject/pymatgen/issues/2206
| 956,056,926
|
MDU6SXNzdWU5NTYwNTY5MjY=
| 2,206
|
BruteForceOrderMatching class in molecule_matcher module - very basic example - error
|
{
"login": "mhsiron",
"id": 49957831,
"node_id": "MDQ6VXNlcjQ5OTU3ODMx",
"avatar_url": "https://avatars.githubusercontent.com/u/49957831?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mhsiron",
"html_url": "https://github.com/mhsiron",
"followers_url": "https://api.github.com/users/mhsiron/followers",
"following_url": "https://api.github.com/users/mhsiron/following{/other_user}",
"gists_url": "https://api.github.com/users/mhsiron/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mhsiron/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mhsiron/subscriptions",
"organizations_url": "https://api.github.com/users/mhsiron/orgs",
"repos_url": "https://api.github.com/users/mhsiron/repos",
"events_url": "https://api.github.com/users/mhsiron/events{/privacy}",
"received_events_url": "https://api.github.com/users/mhsiron/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks for the report @mhsiron.\r\n\r\n@fekad, do you have any thoughts on this one?",
"Thanks for the report @mhsiron.\r\n\r\nI have some notes:\r\n- Be careful to use `BruteForceOrderMatcher` with `SymmOp.from_rotation_and_translation`. This method also matches the indices of atoms so if you ignore them you can easily get weird results. You can use `KabschMatcher` insteed.\r\n- The order of rotation and translation are matter. You can use always use `fit` method to do the actual fitting:\r\n```\r\nfrom pymatgen.core import Molecule\r\nfrom pymatgen.analysis.molecule_matcher import BruteForceOrderMatcher\r\nfrom pymatgen.core import SymmOp\r\n\r\nc_cl_x = Molecule(['C', 'Cl'], [[0,0,0], [1.14,0,0]])\r\nc_cl_y = Molecule(['C', 'Cl'], [[0,0,0], [0,1.14,0]])\r\nbfom = BruteForceOrderMatcher(c_cl_x)\r\nc_cl_x, bfom.fit(c_cl_y)[0]\r\n```\r\n```\r\nc_cl_x = Molecule(['C', 'Cl'], [[0,0,-1], [1.14,0,-1]])\r\nc_cl_y = Molecule(['C', 'Cl'], [[1,0,1], [1,1.14,1]])\r\n\r\nbfom = BruteForceOrderMatcher(c_cl_x)\r\nc_cl_x, bfom.fit(c_cl_y)[0]\r\n```\r\n- if you still want to use `SymmOp.from_rotation_and_translation` please check the \"direction\" of the rotation:\r\n```\r\nimport numpy as np\r\nfrom pymatgen.core import Molecule\r\nfrom pymatgen.analysis.molecule_matcher import BruteForceOrderMatcher\r\nfrom pymatgen.core import SymmOp\r\n\r\nc_cl_x = Molecule(['C', 'Cl'], [[0,0,0], [1.14,0,0]])\r\nc_cl_y = Molecule(['C', 'Cl'], [[0,0,0], [0,1.14,0]])\r\nbfom = BruteForceOrderMatcher(c_cl_x)\r\ninds, U, V, rmsd = bfom.match(c_cl_y)\r\n\r\n\r\nc_cl_y_tr = c_cl_y.copy()\r\nc_cl_y_tr.apply_operation(SymmOp.from_rotation_and_translation(np.linalg.inv(U), V))\r\nc_cl_x, c_cl_y_tr\r\n```\r\n```\r\nc_cl_x = Molecule(['C', 'Cl'], [[0,0,-1], [1.14,0,-1]])\r\nc_cl_y = Molecule(['C', 'Cl'], [[1,0,1], [1,1.14,1]])\r\n\r\nbfom = BruteForceOrderMatcher(c_cl_x)\r\ninds, U, V, rmsd = bfom.match(c_cl_y)\r\n\r\nc_cl_y_tr = c_cl_y.copy()\r\nc_cl_y_tr.apply_operation(SymmOp.from_rotation_and_translation(np.linalg.inv(U), V))\r\nc_cl_x, c_cl_y_tr\r\n```\r\n",
"it can work with the other direction namely rotating \"x\" into \"y\":\r\n```\r\nc_cl_x = Molecule(['C', 'Cl'], [[0,0,-1], [1.14,0,-1]])\r\nc_cl_y = Molecule(['C', 'Cl'], [[1,0,1], [1,1.14,1]])\r\n\r\nbfom = BruteForceOrderMatcher(c_cl_x)\r\ninds, U, V, rmsd = bfom.match(c_cl_y)\r\n\r\nc_cl_x_tr = c_cl_x.copy()\r\nc_cl_x_tr.apply_operation(SymmOp.from_rotation_and_translation(U, -np.dot(U, V)))\r\nc_cl_y, c_cl_x_tr\r\n```\r\nHere we also need to replace the order of translation and rotation.\r\n\r\nAlthough the choice of directions/order was determined based on the previous implementations and it is just a convention, it would make sense to the same one as in `SymmOp.from_rotation_and_translation`."
] | 2021-07-29T17:27:54
| 2021-07-30T01:53:11
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
The BruteForceOrderMatching class in the molecule_matcher module appears to return erroneous results for simple molecule matching examples (ie with 2 molecule, with some simple transformations between them).
**To Reproduce**
Example 1:
```
from pymatgen.core import Molecule
from pymatgen.analysis.molecule_matcher import BruteForceOrderMatcher
from pymatgen.core import SymmOp
c_cl_x = Molecule(['C', 'Cl'], [[0,0,0], [1.14,0,0]])
c_cl_y = Molecule(['C', 'Cl'], [[0,0,0], [0,1.14,0]])
bfom = BruteForceOrderMatcher(c_cl_x)
c_cl_y_tr = c_cl_y.copy()
c_cl_y_tr.apply_operation(SymmOp.from_rotation_and_translation(bfom.match(c_cl_y)[1], bfom.match(c_cl_y)[2]))
```
The resulting rotation matrix leads to a c_cl_y_tr which is incorrectly flipped.
Here one could apply the inverse of the rotation matrix, but this can lead to worse results for other simple roations
Example 2:
```
c_cl_x = Molecule(['C', 'Cl'], [[0,0,-1], [1.14,0,-1]])
c_cl_y = Molecule(['C', 'Cl'], [[1,0,1], [1,1.14,1]])
bfom = BruteForceOrderMatcher(c_cl_x)
c_cl_y_tr = c_cl_y.copy()
c_cl_y_tr.apply_operation(SymmOp.from_rotation_and_translation(bfom.match(c_cl_y)[1], bfom.match(c_cl_y)[2]))
```
This leads to an incorrect translation vector for the y component, here, taking the negative of the rotation matrix does not help, like in the previous example. The issue this time is with the translation vector and rotation matrix.
**Expected behavior**
The following code:
```
c_cl_y_tr = c_cl_y.copy()
c_cl_y_tr.apply_operation(SymmOp.from_rotation_and_translation(bfom.match(c_cl_y)[1], bfom.match(c_cl_y)[2]))
```
Should reproduce c_cl_x
**Desktop (please complete the following information):**
- OS: Windows
- Version 2022.0.8
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2206/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2206/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2207
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2207/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2207/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2207/events
|
https://github.com/materialsproject/pymatgen/issues/2207
| 958,569,714
|
MDU6SXNzdWU5NTg1Njk3MTQ=
| 2,207
|
KeyError when accessing PDOS via sites
|
{
"login": "tawe141",
"id": 13124532,
"node_id": "MDQ6VXNlcjEzMTI0NTMy",
"avatar_url": "https://avatars.githubusercontent.com/u/13124532?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/tawe141",
"html_url": "https://github.com/tawe141",
"followers_url": "https://api.github.com/users/tawe141/followers",
"following_url": "https://api.github.com/users/tawe141/following{/other_user}",
"gists_url": "https://api.github.com/users/tawe141/gists{/gist_id}",
"starred_url": "https://api.github.com/users/tawe141/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/tawe141/subscriptions",
"organizations_url": "https://api.github.com/users/tawe141/orgs",
"repos_url": "https://api.github.com/users/tawe141/repos",
"events_url": "https://api.github.com/users/tawe141/events{/privacy}",
"received_events_url": "https://api.github.com/users/tawe141/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This is not really the way this is meant to be used. The structure associated with the complete dos object is stored in the dos.structure attribute. So all you need to do is to replace vr.structures[-1] with dos.structure. vr.structures[-1] may not be exactly the same as the final structure since that is just the structure of the final relaxation step.",
"Got it, thanks Shyue!"
] | 2021-08-02T23:00:56
| 2021-08-02T23:49:46
|
2021-08-02T23:49:46Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
I cannot use `vasprun.complete_dos.get_site_orbital_dos()` in the snippet below because the site I pass in cannot be matched to any of the sites despite passing it a `PeriodicSite` directly from the structure.
My naive suspicion is that there is something going on with how the sites are compared. The hash of a Periodic site is just its atomic number, assuming no partial occupancies. For a structure that has more than one site with the same hash, Python's dict probably does an iterative comparison between the given site and all sites in its keys with the same hash. Somehow, the comparison between two of the same sites is failing.
**To Reproduce**
```
from pymatgen.io.vasp.outputs import Vasprun
vr = Vasprun('vasprun.xml')
print(vr.structures[-1][10] in vr.complete_dos.pdos.keys()) # will print False
vr.complete_dos.get_site_dos(vr.structures[-1][10]) # will throw KeyError
```
```
---------------------------------------------------------------------------
KeyError Traceback (most recent call last)
<ipython-input-7-666f345603d6> in <module>
----> 1 vr.complete_dos.get_site_dos(vr.structures[-1][10])
~/miniconda3/lib/python3.7/site-packages/pymatgen/electronic_structure/dos.py in get_site_dos(self, site)
685 Dos containing summed orbital densities for site.
686 """
--> 687 site_dos = functools.reduce(add_densities, self.pdos[site].values())
688 return Dos(self.efermi, self.energies, site_dos)
689
KeyError: PeriodicSite: Cr (2.9902, 8.4576, 5.1792) [0.0000, 0.5546, 0.0000]
```
See `vasprun.xml` file here: https://www.dropbox.com/s/3qss6jus4uvgjxw/vasprun.xml?dl=0
**Expected behavior**
This should index the pdos just fine, considering the site that pdos is indexing is directly from the structure.
**Desktop (please complete the following information):**
- OS: Mac OSX Big Sur 11.5.1
- Version 2021.2.8.1
|
{
"login": "tawe141",
"id": 13124532,
"node_id": "MDQ6VXNlcjEzMTI0NTMy",
"avatar_url": "https://avatars.githubusercontent.com/u/13124532?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/tawe141",
"html_url": "https://github.com/tawe141",
"followers_url": "https://api.github.com/users/tawe141/followers",
"following_url": "https://api.github.com/users/tawe141/following{/other_user}",
"gists_url": "https://api.github.com/users/tawe141/gists{/gist_id}",
"starred_url": "https://api.github.com/users/tawe141/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/tawe141/subscriptions",
"organizations_url": "https://api.github.com/users/tawe141/orgs",
"repos_url": "https://api.github.com/users/tawe141/repos",
"events_url": "https://api.github.com/users/tawe141/events{/privacy}",
"received_events_url": "https://api.github.com/users/tawe141/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2207/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2207/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2208
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2208/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2208/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2208/events
|
https://github.com/materialsproject/pymatgen/pull/2208
| 959,252,379
|
MDExOlB1bGxSZXF1ZXN0NzAyMzk1Njgy
| 2,208
|
update MP2020 publication details
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/41870449)\n\nCoverage decreased (-0.6%) to 83.063% when pulling **c52889a41d0dfb23efe2cddefee3220435032654 on rkingsbury:master** into **3683e218cc512d15090927faf52794dbcea12322 on materialsproject:master**.\n"
] | 2021-08-03T15:45:26
| 2021-08-03T16:28:09
|
2021-08-03T15:48:41Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Update docstring of `MaterialsProject2020Compatibility` with final published version of manuscript
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2208/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2208/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2208",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2208",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2208.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2208.patch",
"merged_at": "2021-08-03T15:48:41Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2209
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2209/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2209/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2209/events
|
https://github.com/materialsproject/pymatgen/issues/2209
| 961,582,858
|
MDU6SXNzdWU5NjE1ODI4NTg=
| 2,209
|
import Lattice error in Anaconda
|
{
"login": "mersadkhan",
"id": 4376091,
"node_id": "MDQ6VXNlcjQzNzYwOTE=",
"avatar_url": "https://avatars.githubusercontent.com/u/4376091?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mersadkhan",
"html_url": "https://github.com/mersadkhan",
"followers_url": "https://api.github.com/users/mersadkhan/followers",
"following_url": "https://api.github.com/users/mersadkhan/following{/other_user}",
"gists_url": "https://api.github.com/users/mersadkhan/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mersadkhan/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mersadkhan/subscriptions",
"organizations_url": "https://api.github.com/users/mersadkhan/orgs",
"repos_url": "https://api.github.com/users/mersadkhan/repos",
"events_url": "https://api.github.com/users/mersadkhan/events{/privacy}",
"received_events_url": "https://api.github.com/users/mersadkhan/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Lattice needs to be imported from either pymatgen.core or pymatgen.core.lattice. Convenient imports have been removed from v2022.0."
] | 2021-08-05T08:34:58
| 2021-08-05T14:26:46
|
2021-08-05T14:26:46Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Steps to reproduce the behavior:
1. Install Anaconda3-2020.11-Linux-x86_64
2. install pymatgen by :
```
conda install --yes numpy scipy matplotlib
pip install pymatgen
```
3. run python and
`from pymatgen import Lattice `
4. See error
```
Python 3.8.8 (default, Apr 13 2021, 19:58:26)
[GCC 7.3.0] :: Anaconda, Inc. on linux
Type "help", "copyright", "credits" or "license" for more information.
>>> from pymatgen import Lattice
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
ImportError: cannot import name 'Lattice' from 'pymatgen' (unknown location)
```
**Desktop (please complete the following information):**
- OS: [Linux]
- Version [mint 20]
**Additional context**
the below command works and I check the file is all installed in the right place
import pymatgen
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2209/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2209/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2210
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2210/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2210/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2210/events
|
https://github.com/materialsproject/pymatgen/issues/2210
| 962,371,374
|
MDU6SXNzdWU5NjIzNzEzNzQ=
| 2,210
|
Value ERROR: No Voronoi neighbours found for site - try increasing cutoff
|
{
"login": "hasan-sayeed",
"id": 47003757,
"node_id": "MDQ6VXNlcjQ3MDAzNzU3",
"avatar_url": "https://avatars.githubusercontent.com/u/47003757?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/hasan-sayeed",
"html_url": "https://github.com/hasan-sayeed",
"followers_url": "https://api.github.com/users/hasan-sayeed/followers",
"following_url": "https://api.github.com/users/hasan-sayeed/following{/other_user}",
"gists_url": "https://api.github.com/users/hasan-sayeed/gists{/gist_id}",
"starred_url": "https://api.github.com/users/hasan-sayeed/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/hasan-sayeed/subscriptions",
"organizations_url": "https://api.github.com/users/hasan-sayeed/orgs",
"repos_url": "https://api.github.com/users/hasan-sayeed/repos",
"events_url": "https://api.github.com/users/hasan-sayeed/events{/privacy}",
"received_events_url": "https://api.github.com/users/hasan-sayeed/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"> I'm trying to get condensed structure of stable materials from Materials Project using Robocrystellographer. But I'm getting this error `Value ERROR: No Voronoi neighbours found for site - try increasing cutoff` from line 892 of `C:\\Users\\...\\site-packages\\pymatgen\\analysis\\local_env.py`. Seems like it's coming from the `class VoronoiNN`. The default cutoff was 13 and I raised this up to 20 but getting the same error every time at the same point! Can't figure out what's the problem.\r\n\r\nHello, I also met this probelm, have you fixed it ?"
] | 2021-08-06T03:54:24
| 2023-10-29T05:53:30
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I'm trying to get condensed structure of stable materials from Materials Project using Robocrystellographer. But I'm getting this error `Value ERROR: No Voronoi neighbours found for site - try increasing cutoff` from line 892 of `C:\Users\...\site-packages\pymatgen\analysis\local_env.py`. Seems like it's coming from the `class VoronoiNN`. The default cutoff was 13 and I raised this up to 20 but getting the same error every time at the same point! Can't figure out what's the problem.
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2210/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2210/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2211
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2211/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2211/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2211/events
|
https://github.com/materialsproject/pymatgen/pull/2211
| 962,971,001
|
MDExOlB1bGxSZXF1ZXN0NzA1NjgyODk0
| 2,211
|
Abstract interface for Input classes
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I am in favor of a more concrete API definition, but I am not sure about this implementation. \r\n1. Input conflicts with the Python built-in input method.\r\n2. I fail to see what the current Input offers beyond the VaspInputSet abstract class. Certainly we can abstract out parts of the VaspInputSet class to make it general. But there doesn't seem to be any further generalizability beyond VASP. It seems that the main definitions are from_files and write_inputs (which I would prefer to rename completely), which are already pretty standard throughout pymatgen.\r\n3. Finally, I think we should be clear about what input/output means. In the context of pymatgne, input/output generally means between Python objects and files/other sources. It does not mean VASP input files or output files. Both of those are actually inputs as far as pymatgen is concerned. ",
"> I am in favor of a more concrete API definition, but I am not sure about this implementation.\r\n\r\nThanks for your thoughts @shyuep! Let me ask some clarifying questions.\r\n\r\n> Input conflicts with the Python built-in input method.\r\n\r\nExcellent point. Since we are trying to separate the data containers from the input settings I was trying to find a pair of names that were clear and evocative for this distinction. I worry that `InputSet` and `InputSettings` would be confusing. What would you think about `InputSet` (the container) and `InputGenerator` (for the settings)?\r\n\r\n> I fail to see what the current Input offers beyond the VaspInputSet abstract class. Certainly we can abstract out parts of the VaspInputSet class to make it general. But there doesn't seem to be any further generalizability beyond VASP. It seems that the main definitions are from_files and write_inputs (which I would prefer to rename completely), which are already pretty standard throughout pymatgen.\r\n\r\nYes, to be honest the number one thing I'm hoping to achieve with this is to standardize the method name and call signature of `from_files` and `write_inputs`. There are analogous methods throughout pymatgen but some go by different names and have different call signatures. How do you want to rename `write_inputs`?\r\n\r\nThe other big benefit beyond the current implementation of `VaspInputSet` (which @utf can probably elaborate on more) is to enforce a clear separation between the data container and the generator. This should remove many points of confusion w.r.t inheritance and streamline workflow development.\r\n\r\n> Finally, I think we should be clear about what input/output means. In the context of pymatgne, input/output generally means between Python objects and files/other sources. It does not mean VASP input files or output files. Both of those are actually inputs as far as pymatgen is concerned.\r\n\r\nI see your point. Would the renaming convention in Item 1 address? Do you have other ideas about how we would clarify this distinction? We could go further and define the base class as `CodeInputSet` or something.\r\n\r\n",
"> I fail to see what the current Input offers beyond the VaspInputSet abstract class. Certainly we can abstract out parts of the VaspInputSet class to make it general. But there doesn't seem to be any further generalizability beyond VASP. It seems that the main definitions are from_files and write_inputs (which I would prefer to rename completely), which are already pretty standard throughout pymatgen.\r\n\r\nI think part of the confusion with VaspInputSet is that it is used as both a generator class and as a container for VASP inputs. This means initialising the class requires all the data needed to generate the inputs (structure, etc) but the actual inputs are generated dynamically (i.e., when accessing the `incar` attribute).\r\n\r\nThe pain points are then:\r\n1. If you have a VaspInputSet it is not obvious how to edit the INCAR or KPOINTS. I.e., you can't just do `vis.incar[\"NSW\"] = 10` because the inputs are generated on the fly.\r\n2. On the flip side, the structure is needed on class creation even though the inputs are generated dynamically, which means that you can't use the input set as a template for delayed input creation.\r\n3. It can be very hard to track down why a certain setting is in your INCAR/KPOINTS because every VaspInputSet subclass implements the Incar, kpoints, etc generation in slightly different ways. E.g., some subclasses choose to edit the k-points in the init method, some choose to do it in the `kpoints` attribute, some in the `write_inputs` function, some in a mixture of all them.\r\n\r\nThis proposal is to have:\r\n1. A fixed input set class that isn’t subclassed for specific calculation types. The init, write_input, and validate functions always do exactly the same thing regardless of whether your calculation is a GGA relaxation, hybrid line mode band structure, or were generated outside of pymatgen.\r\n2. Input set generators for generating the input set class. These contain all the crazy logic for generating the input sets. You can subclass the generators but these only have one function `get_input_set`, so the tracking down their behaviour should be much simpler.\r\n\r\nAs @computron wrote in an email:\r\n\r\n> the \"VaspInput\" class of pymatgen is almost like the Input class proposed. And the VaspInputSet *would* be like the generators proposed if it just ditched all the internal representations of kpoints, poscar, etc. as well as functions for writing out inputs itself, and basically just had the existing method get_vasp_input() retained.\r\n\r\nI think this model can be used beyond the VASP module. The typical pattern used in pymatgen for VASP, QCHEM, CP2K, FEFF, etc is: 1) Make an abstract input set to represent input files, 2) create subclasses of this input set for different calculation types. But in each case, the input sets suffer from the generator/container intermixing and related issues.",
"\r\n> I think part of the confusion with VaspInputSet is that it is used as both a generator class and as a container for VASP inputs. This means initialising the class requires all the data needed to generate the inputs (structure, etc) but the actual inputs are generated dynamically (i.e., when accessing the `incar` attribute).\r\n\r\nThere is a separate VaspInput class in pymatgen.vasp.inputs.\r\n \r\n> The pain points are then:\r\n> \r\n> 1. If you have a VaspInputSet it is not obvious how to edit the INCAR or KPOINTS. I.e., you can't just do `vis.incar[\"NSW\"] = 10` because the inputs are generated on the fly.\r\n\r\nThe idea of an InputSet is to have a relatively rigid way of defining rules for how an input is generated. There is absolutely no reason why you cannot do what you just wrote. `incar = vis.incar; incar[\"NSW\"] = 10` works perfectly fine. So does `user_incar_settings={\"NSW\": 10}`.\r\n\r\n> 2. On the flip side, the structure is needed on class creation even though the inputs are generated dynamically, which means that you can't use the input set as a template for delayed input creation.\r\n\r\nThis has been debated ad nauseum. Previously the input sets did not require structure and generated on the fly. But that led to circular dependencies. E.g., if you are generating inputs for a Bandstructure calculation, you sometimes transform to standard symmetry directions. Basically, that means that if vis.get_incar, vis.get_kpoints, etc. will all call the same structure transformation - 4x for your standard set of inputs. Further, sometimes the incar setting or the kpoint setting can depend on each other. This is why an input set is tied specifically to a structure today. If you have a better idea of how to solve this problem, by all means propose it.\r\n\r\n> 3. It can be very hard to track down why a certain setting is in your INCAR/KPOINTS because every VaspInputSet subclass implements the Incar, kpoints, etc generation in slightly different ways. E.g., some subclasses choose to edit the k-points in the init method, some choose to do it in the kpoints property function, some in the write_inputs function, some in a mixture of all them.\r\n\r\n> \r\n> This proposal is to have:\r\n> \r\n> 1. A fixed input set class that isn’t subclassed for specific calculation types. The init, write_input, and validate functions always do exactly the same thing regardless of whether your calculation is a GGA relaxation, hybrid line mode band structure, or were generated outside of pymatgen.\r\n> 2. Input set generators for generating the input set class. These contain all the crazy logic for generating the input sets. You can subclass the generators but these only have one function `get_input_set`, so the tracking down their behaviour should be much simpler.\r\n\r\nAgain, this is exactly what the situation it is today. Your so-called fix input set class is basically what the VaspInput class is. Your generator is basically what the VaspInputSet is. We can of course do a rename and call it VaspInputGenerator instead of VaspInputSet, but that is semantics, not implementation.\r\n\r\n> \r\n> As @computron wrote in an email:\r\n> \r\n> > the \"VaspInput\" class of pymatgen is almost like the Input class proposed. And the VaspInputSet _would_ be like the generators proposed if it just ditched all the internal representations of kpoints, poscar, etc. as well as functions for writing out inputs itself, and basically just had the existing method get_vasp_input() retained.\r\n> \r\n> I think this model can be used beyond the VASP module. The typical pattern used in pymatgen for VASP, QCHEM, CP2K, FEFF, etc is: 1) Make an abstract input set to represent input files, 2) create subclasses of this input set for different calculation types. But in each case, the input sets suffer from the generator/container intermixing and related issues.\r\n\r\n@computron should be well-aware of the reasons why structure was included in the VaspInputSet in the first place. Solve the issue of the bandstructure input set generation and let me know when you have a good solution. You will find that both types of solutions have their flaws and pain points for different types of calculations.",
"> The idea of an InputSet is to have a relatively rigid way of defining rules for how an input is generated. There is absolutely no reason why you cannot do what you just wrote. `incar = vis.incar; incar[\"NSW\"] = 10` works perfectly fine.\r\n\r\nBut now you no longer have the nice container class for writing all the inputs to file and validating that they are correct. Note that validating an INCAR is reasonable requires having access to the poscar and kpoints files too. Hence, the proposed api allows for the user to write `vis.incar[\"NSW\"] = 10; vis.validate()` which is natural. The `vis.user_incar_settings={\"NSW\": 10}` example is messy and difficult to discover imo.\r\n\r\n> Previously the input sets did not require structure and generated on the fly. But that led to circular dependencies. E.g., if you are generating inputs for a Bandstructure calculation, you sometimes transform to standard symmetry directions. Basically, that means that if vis.get_incar, vis.get_kpoints, etc. will all call the same structure transformation - 4x for your standard set of inputs. Further, sometimes the incar setting or the kpoint setting can depend on each other. This is why an input set is tied specifically to a structure today. If you have a better idea of how to solve this problem, by all means propose it.\r\n\r\nI think this example showcases the strength of current proposal nicely. You have an `InputSetGenerator` with a single function `get_input_set`. You generate the input set with `InputSetGenerator.get_input_set(structure)` and you get back an `InputSet` set that has fully consistent incar, kpoints, poscar and potcar objects. If the structure needs to be transformed, this is done in `get_input_set` and the kpoints, incar etc are all consistent with the transformed structure. \r\n\r\nThe reason why circular dependencies arose was because the old generators allowed you to return individual incar/poscar/kpoints objects. That is not possible with this proposal. You get a full input set back that is fully consistent.",
"> But now you no longer have the nice container class for writing all the inputs to file and validating that they are correct. Note that validating an INCAR is reasonable requires having access to the poscar and kpoints files too. Hence, the proposed api allows for the user to write `vis.incar[\"NSW\"] = 10; vis.validate()` which is natural. The `vis.user_incar_settings={\"NSW\": 10}` example is messy and difficult to discover imo.\r\n\r\nAgain, I would point out, as @computron has pointed out, that there is a VaspInput class. By all means add a validate() method in VaspInput if you wish. I don't even see the point of a validate method. A VaspInputSet/Generator is basically a rigid set of rules to generate *consistent* inputs. Why would you ever generate a consistent set of inputs and then go back to validate that your inputs generated are consistent? If proper unittests are written, the inputs generated must be consistent. Feel free to demonstrate an example \"validate()\" method that actually increases utility. \r\n \r\n> I think this example showcases the strength of current proposal nicely. You have an input set generator with a single function `get_input_set`. You generate the input set with `generator.get_input_set(structure)` and you get back a vasp input set that has fully consistent incar, kpoints, poscar and potcar objects. If the structure needs to be transformed, this is done in `get_input_set` and the kpoints, incar etc are all consistent with the transformed structure.\r\n> \r\n> The reason why circular dependencies arose was because the old generators allowed you to return individual incar/poscar/kpoints objects. That is not possible with this proposal. You get a full input set back that is fully consistent.\r\n\r\nSure, if you want to go this route. The whole idea was to allow people to just generate individual components, rather than shoving the entire VaspInput down their throats whether they want it or not. There are situations where I just need one of the inputs and not all of them. The current implementation generates the inputs *as needed*.\r\n\r\nNote that this is independent from the validate question. I personally think the validate() method is overengineering for no reason. \r\n",
"> Feel free to demonstrate an example \"validate()\" method that actually increases utility.\r\n\r\nThere are two use cases that I have in mind:\r\n1. It is common to customise an input set generator with additional settings, e.g., the `user_incar_settings={\"NSW\": 10}` example mentioned above. Due to the complexity of some of the generators it is nice to have a quick way to check that the customised input is valid. \r\n2. I've had several conversations with VASP users who use pymatgen for managing VASP inputs but do not use the VaspInputSets and who wanted a way to validate their inputs. I see this feature (and the input set container in general) as making pymatgen more useful for these users.\r\n\r\n> Sure, if you want to go this route. The whole idea was to allow people to just generate individual components, rather than shoving the entire VaspInput down their throats whether they want it or not. There are situations where I just need one of the inputs and not all of them. The current implementation generates the inputs as needed.\r\n\r\nTo me, the naming and most common usage of the vasp input sets (at least in atomate) is to generate full input \"sets\" rather than individual inputs. As you mentioned, generating individual inputs can be dangerous as sometimes the inputs are not consistent. Regardless, in this proposal it is easy enough to get a single input using `vis = InputSetGenerator.get_input_set(structure); vis.incar`.\r\n\r\nI think the fact that there is a lot of overlap between VaspInput/VaspInputSet and this proposal is a good thing, as ultimately we don't want to do anything radical and just improve the API for consistency/easier use.\r\n\r\nPerhaps @rkingsbury can comment but I presume the idea is that these new input sets will be developed along side the existing sets, at least at first?\r\n\r\n",
"1. I would like to see how you implement the validate method. If it is merely checking list of species in POTCAR and POSCAR and INCAR are consistent and U values listing, then I would say it is not worth doing at all or we can implement this simple method in the VaspInput method that works generically for all sets of vasp inputs. If it is to check structure consistency etc. like what the bandstructure input sets require, I would say people should just use the VaspInputSets rather than trying to be clever and to do something in a way that it is not intended. There are limits to what kind of protection we want to give users.\r\n2. Atomate is not the only \"customer\" of pymatgen. Generating individual inputs is dangerous only if you do not include the structure in the generation process. If the input set is tied to a structure (like the current VaspInputSets), then the individual inputs are always consistent to the extent that the individual generators use the same processed structure. Your suggested \"solutions\" means that all four inputs are generated, even if I only want the INCAR file.\r\n",
"Going back to the original proposal, my thoughts are:\r\n\r\n1. What about `InputSet` and `InputSetGenerator` as names for the container and generator abstract classes. The generator method would then be called `InputSetGenerator.get_input_set()`.\r\n2. `Input.from_files` is misleading as the function accepts a directory as input. What about `Input.from_directory` instead?\r\n3. In `write_inputs`, some of the argument names are quite long and some override python builtins (e.g., `dir`). My recommendations would be:\r\n - `dir` -> `directory` (and also in `from_files`).\r\n - `make_dir_if_not_present` -> `make_dir`.\r\n - `overwrite_if_present` -> `overwrite`.\r\n4. What happens if a code doesn't require a structure/atomic coordinates as input or if it doesn't make sense to override from a previous directory. I don't think there should be any required/optional arguments for the `get_input_set` method. \r\n\r\nOther than that, I think this proposal is on the right track. I especially like the template code. Obviously the VASP specific code is very rough. If we are happy to move forward with this proposal then I will add some more detailed comments on that section.",
"Thanks everyone for the discussion here so far. I agree with @shyuep in the first post that I am also in favor of a more concrete API definition, so thank you all for your input here and I hope we can agree upon a solution and get this merged.\r\n\r\n### Note on language\r\n\r\nI want to request that we keep threads civil, discussion and disagreements are great if worded politely and constructively. I want to avoid language like \"shoving [...] down their throats\" as not appropriate for this venue.\r\n\r\n### Making individual files/objects accessible like `.incar`\r\n\r\nFrom @shyuep:\r\n\r\n> Your suggested \"solutions\" means that all four inputs are generated, even if I only want the INCAR file.\r\n\r\nAnd context from @utf:\r\n\r\n> The reason why circular dependencies arose was because the old generators allowed you to return individual incar/poscar/kpoints objects. That is not possible with this proposal. You get a full input set back that is fully consistent.\r\n\r\nI do see the benefit in being able to access the individual classes as necessary, e.g. I might sometimes create an input set just to see which POTCARs it uses, and then access the `Potcar` object for additional information.\r\n\r\n@utf, can you clarify why this restriction would be useful? I see your note that about confusion because \"some subclasses choose to edit the k-points in the init method, some choose to do it in the kpoints attribute, some in the write_inputs function, some in a mixture of all them.\" I agree this is an issue. Perhaps we can retain `.incar`, `.kpoints` etc., but maybe there's a way to enforce that these simply return the objects from the input set and so cannot contain any logic within the property itself?\r\n\r\n### Inclusion of `validate` method\r\n\r\n> I would like to see how you implement the validate method [...] There are limits to what kind of protection we want to give users.\r\n\r\nAgreed, I actually also think that the logic _within_ the input set should ensure that inputs are sensible and internally-consistent. However, given the amount of customization possible, I can imagine scenarios where this might be useful. At worst, this is harmless. At best, it might catch some dangerous side cases.\r\n\r\n### Naming of classes and methods\r\n\r\nFrom @utf:\r\n\r\n> 1. What about InputSet and InputSetGenerator as names for the container and generator abstract classes. The generator method would then be called InputSetGenerator.get_input_set(). \r\n\r\nOk.\r\n\r\n> 2. Input.from_files is misleading as the function accepts a directory as input. What about Input.from_directory instead?\r\n\r\nAgreed.\r\n\r\n> 3. In write_inputs, some of the argument names are quite long and some override python builtins (e.g., dir). My recommendations would be:\r\n> dir -> directory (and also in from_files).\r\n> make_dir_if_not_present -> make_dir.\r\n> overwrite_if_present -> overwrite.\r\n\r\nAgreed.\r\n\r\n> What happens if a code doesn't require a structure/atomic coordinates as input or if it doesn't make sense to override from a previous directory. I don't think there should be any required/optional arguments for the get_input_set method.\r\n\r\nI think this makes sense too, it makes it very general but that's what a good base class should be.",
"> Perhaps we can retain .incar, .kpoints etc., but maybe there's a way to enforce that these simply retrn the objects from the input set and so cannot contain any logic within the property itself?\r\n\r\nThe `.incar` etc attributes only work if the generator has already been initialised with the structure. In this proposal, the structure is not needed to initialise the generator, and the generator can be serialized/deserialized and used to generate many sets of input files, not just one set belonging to a certain structure. The `.incar`, `.kpoints` attributes are on the `InputSet` class instead. If you look at how these input sets will be used there is little difference between `vis = InputSetGenerator().get_input_set(structure, prev_dir); vis.incar` (proposed usage) instead of `vis = VaspInputSet.from_prev(structure, prev_dir); vis.incar` (current usage).\r\n\r\nBased on this discussion there are two approaches for the (VASP) input sets. I've tried to summarise the pros and cons of each approach.\r\n\r\n### Structure on initialisation (current approach)\r\n\r\nUsage: \r\n```python\r\nvis = VaspInputSet(structure, user_incar_settings=uis)\r\n# or, for some input sets that support overriding from previous calculations\r\nvis = VaspInputSet.from_prev_calc(prev_dir, structure, user_incar_settings=uis)\r\nvis.incar\r\nvis.write_inputs()\r\n```\r\n\r\n**Pros:**\r\n- Individual inputs accessible from generator.\r\n- Single object.\r\n\r\n**Cons:**\r\n- Hybrid generator/container approach.\r\n- Structure needed to initialise input set. This means, i) the generator can only generate inputs for a single structure; and ii) you cannot use a customised input set as a generic generator for multiple structures.\r\n- Messy overrides of `.kpoints`, etc in subclasses, with code to generate inputs split between init, `from_prev` and `.incar`, `.kpoints`, etc attributes.\r\n- No generic container for VASP inputs, i.e., cannot be used by users who just want to manage VASP inputs but not generate them using pymatgen.\r\n\r\n### Pure generator approach (this proposal)\r\n\r\nUsage: \r\n```python\r\nvis = InputSetGenerator(user_incar_settings=uis).get_input_set(structure, prev_dir)\r\nvis.incar\r\nvis.write_inputs()\r\n```\r\n\r\n**Pros:**\r\n- Generator can be initialised with custom options, serialised and passed around, allowing multiple input sets to be generated for different structures. Very useful for atomate/passing input sets to collaborators.\r\n- All code to generate input set, with/without a previous directory, lives in one place (`get_input_set`).\r\n- Clean separation between generator code (i.e., logic to generate inputs) and the representation of the input set itself (which works without the generator).\r\n\r\n**Cons:**\r\n- Two objects required (generator and container).\r\n\r\nNote that the difference between the approaches is **not** a question of computational efficiency. While in the current approach you can type `.incar` to access just the `Incar` object, calling this attribute also calls `.kpoints` and `.potcar` behind the scenes, even though the `KPOINTS` or `POTCAR` objects are not returned.\r\n\r\n### Validate method\r\n\r\n> If it is merely checking list of species in POTCAR and POSCAR and INCAR are consistent and U values listing ... we can implement this simple method in the VaspInput method that works generically for all sets of vasp inputs. \r\n\r\nThis is exactly what this proposal is suggesting. I.e., the validate function should be generic enough to work on all sets of VASP inputs. From the top of my head there are many other things we can check for:\r\n- KSPACING and `Kpoints` files conflicts.\r\n- Num k-points compatible with ISMEAR (i.e., > 4 k-points if using tetrahedron method).\r\n- NKRED compatible with KPOINTS file\r\n- MAGMOM settings reasonable (also w.r.t. SOC)\r\n- POTCAR functional matches GGA tag in INCAR.\r\n- etc\r\n\r\n> I actually also think that the logic within the input set should ensure that inputs are sensible and internally-consistent. \r\n\r\nThis could be achieved by calling `.validate()` before returning the input set.",
"Thanks @utf, so with the generator approach, the `.incar` attribute remains accessible (once the input set has been generated), in this case I don't see the problem? The generator approach certainly seems cleaner overall.",
"> so with the generator approach, the .incar attribute remains accessible (once the input set has been generated)\r\n\r\nYes, exactly.",
"Thanks all for the robust discussion and especially @utf for the thorough explanations.\r\n\r\nI've pushed some changes implementing @utf and @mkhorton 's suggestions on naming conventions and updated `TemplateInputSet` to more closely follow the generator / container paradigm we're articulating here.\r\n\r\n### On the vision\r\n\r\nFrom @utf \r\n\r\n> I think the fact that there is a lot of overlap between VaspInput/VaspInputSet and this proposal is a good thing, as ultimately we don't want to do anything radical and just improve the API for consistency/easier use.\r\n\r\n> Perhaps @rkingsbury can comment but I presume the idea is that these new input sets will be developed along side the existing sets, at least at first?\r\n\r\nExactly. I don't think we necessarily need to adopt this interface in the VASP input classes right away (or maybe not ever). Those classes are the oldest, most established, and most complex in pymatgen. While there might be advantages to moving toward this framework as @utf has said, it doesn't need to happen right away. I don't have anything to add that hasn't already been said re: advantages or disadvantages over the existing VASP infrastructure.\r\n\r\nI think the biggest value to this interface at this time is to ensure greater consistency among input classes as pymatgen continues to grow. We are seeing continued interest in supporting new codes in pymatgen (xtb, CREST, orca, HOOMD just to name a few I am personally aware of) so I think defining this sooner rather than later will lower the barrier to additional contributions while also streamlining downstream workflows (including bespoke python scripts as well as mainstream atomate workflows).\r\n\r\nMoreover, it seems to me that molecular codes are considerably more \"interchangeable\" than solid state codes, which provides strong motivation for a consistent interface. I feel that if someone uses pymatgen to set up a basic calculation in QChem, it should be relatively trivial to port it to Gaussian or NWChem using pymatgen tools (i.e., the InputSet arguments should be similar). Right now each of these has a different interface even though the underlying input file structure and arguments are not that different. This PR lays a foundation for making these more consistent.\r\n\r\n### On the validate method\r\n\r\nI agree that the internals of the `InputSetGenerator` should ensure that the input is valid. However, there are some limits and edge cases I have run into where this breaks down. I'll give 2 examples I have in mind\r\n\r\n1. QChem: Suppose I have a list of small molecules I want to calculate in DFT. I loop through the list and create a `OptSet` for each structure. If one of my molecules happens to be a single atom, I will still get a valid `InputSet`, but the calculation will fail because QChem doesn't like \"optimizations\" to run on single atoms (they are really single points). Instead of waiting until my calculation fails, this could be caught, either on creation or on the backend, by a `validate` method.\r\n\r\n2. It would be very easy for a future developer to subclass`TemplateInputSetGenerator` to support a new code (in fact, the existing `write_lammps_inputs` method is basically equivalent to this). I could imagine someone wanting not to change the variable substitution logic that's provided in `TemplateInputSet`, but maybe they want to add some rules to the `validate` method as a way to catch non-sensical variable substitutions.\r\n\r\nI personally think having a separate `validate` method to contain these checks and then calling it on creation of a new `InputSet` would provide a clear separation of concerns, but if the consensus is that this is overly complex, we can certainly just leave it to `InputSetGenerator` developers to integrate that logic into the respective classes.\r\n\r\n### on `get_input_set`\r\nFrom @utf and @mkhorton :\r\n\r\n> What happens if a code doesn't require a structure/atomic coordinates as input or if it doesn't make sense to override from a previous directory. I don't think there should be any required/optional arguments for the get_input_set method.\r\n\r\n> I think this makes sense too, it makes it very general but that's what a good base class should be.\r\n\r\nOK, I removed the required arguments. Actually this was needed even to make my `TemplateInputSetGenerator` example work within the paradigm we have here. Outside of that esoteric example though, how likely is it that we will have a code that doesn't require some kind of coordinates? Would a reasonable alternative be to make the `coordinates` arg `Optional` and let developers set it to `None` in the relatively rare cases when a structure isn't required?\r\n\r\nBecause I think `get_input_set` will be used extensively in scripts and atomate workflows, I just feel a little attached to the idea that enforcing that the first arg has to be coordinates, and specifying the types that are allowed, could have a lot of value in terms of code consistency. Maybe that can be saved for later refinement though.",
"Let me suggest a path forward based on the comments.\r\n\r\n1. Build up VaspInput in pymatgen.vasp.inputs to support validation etc. as @utf has suggested.\r\n2. Develop *Mix-in classes* specifying what is expected of an input/output class. This can be in a pymatgen/io/core module (we cannot use `__init__.py` since pymatgen/io is now a namespace package). I think this is better than using a strict hierarchy. This can be based on @rkingsbury initial implementation in this PR.\r\n3. Since everyone has such strong feelings about not including structure, we will develop a parallel equivalent of VaspInputGenerators that does what VaspInputSets do today, but in a no-structure format. This can be in a pymatgne/io/vasp/generators.py module. Once we are satisifed that the generators are a complete replacement for VaspInputSets and have demonstratable benefits, we will deprecate VaspInputSets.\r\n\r\nA final note that I always favor a simple implementation over a complex one. Most other codes are far too serpentine and complex and difficult to use. Do not overengineer something because a 0.01% of people want to do this specialized thing, even if that 0.01% happens to be someone in the Materials Project. If the 0.01% wnat to do something special, they can write their own specialized method. Pymatgen caters to the 80% use case with 20% effort.",
"> Let me suggest a path forward based on the comments.\r\n\r\nThis sounds very reasonable to me.\r\n\r\n@rkingsbury If you're happy going this route and can update this PR to address point 2, I can submit a PR for points 1 and 3.",
"> Let me suggest a path forward based on the comments.\r\n\r\nThank you very much for the proposal @shyuep; apologies for my delayed response.\r\n\r\n> @rkingsbury If you're happy going this route and can update this PR to address point 2, I can submit a PR for points 1 and 3.\r\n\r\nThank you @utf ! I am definitely supportive of some type of stateless generator approach, although after reflecting on this discussion for a while I do have some concerns that we haven't yet converged to the right solution. I'll elaborate below.\r\n\r\n### Reviewing our motivation\r\n\r\nI really appreciate all the thoughtful comments so far. As this discussion has evolved, it's clear that our two objectives are to\r\n\r\n1. Define a more consistent interface for pymatgen `Input` classes\r\n2. Make it possible to serialize and pass around instructions for generating input sets without having to include structures and specific customizations.\r\n\r\nBefore we proceed with anything, I want to make sure that we're honoring those objectives. Although we're getting there, I don't feel the paradigm currently under discussion accomplishes either of them particularly well, but I think it can with a few modifications. Let me elaborate below (apologies for the long post!)\r\n\r\n\r\n### Concern that the Generator / Container paradigm does not accomplish our objectives\r\n\r\nI'm supportive of a stateless input generator class. However, as I've reflected on our discussion, tried to explain the Generator / Container paradigm to others in our group, and developed the `TemplateInputSet` classes, I am concerned that the Generator | Container duality is going to be difficult for the average user or volunteer developer to understand. This relates to @shyuep comment about keeping the code as simple as possible. I do see the value in the two class approach for our operations within MP, and I'm concerned that having two classes is going to create _more_ ambiguity about where to put input construction code rather than less.\r\n\r\nWith the current paradigm under discussion, we could easily wind up with Input creation code in two places if we do not do more to enforce the separation of concerns we intend here. For example, based only on what is required by respective abstract classes, I could do something like this:\r\n\r\n```\r\nclass VaspInputSetGenerator(InputSetGenerator):\r\n\r\n def get_input_set(structure):\r\n return VaspInputSet(structure)\r\n\r\nclass VaspInputSet(InputSet):\r\n\r\n def __init__(structure):\r\n self.structure = structure\r\n # code for creating POTCAR, INCAR, etc. here\r\n```\r\n\r\nBut this would defeat the purpose of having the two classes, since all the input construction code still resides in `VaspInputSet`. Instead, according to our discussion, the proper way to go about this would be:\r\n\r\n```\r\nclass VaspInputSetGenerator(InputSetGenerator):\r\n\r\n def get_input_set(structure):\r\n # code that creates POSCAR, POTCAR, INCAR, KPOINTS here\r\n return VaspInputSet(poscar, potcar, incar, kpoints)\r\n\r\nclass VaspInputSet(InputSet):\r\n\r\n def __init__(poscar, potcar, incar, kpoints):\r\n self.poscar = poscar\r\n self.incar = incar\r\n etc.\r\n```\r\n\r\nI'm not sure that this PR or the proposed path forward do enough to enforce Approach No. 2, and what I think could happen is that we wind up with Input creation code split arbitrarily across the Generator and Container classes in different ways for different codes, and then we would have introduced additional complexity with limited benefit. In fact, if you look at my `TemplateInputSetGenerator` / `TemplateInputSet` implementation as it stands right now, I have some code in the InputSet `__init__` method that really should be in the Generator class because I've had a hard time wrapping my head around the \"right\" way to divide things. \r\n\r\nApproach No. 2 also makes `InputSet` less natural to use by themselves, because more user-friendly input arguments have to be passed to the `InputSetGenerator` which constructs less user-friendly pymatgen objects to pass to `InputSet`. So we effectively force all users to understand the Generator / Container duality. \r\n\r\nIf I put on the \"hat\" of a user that's experienced running calculations in some code but relatively new to pymatgen and to high throughput, that user is going to think about calculation inputs first and foremost as **sets of files**. If that user wants to use pymatgen to generate input files, our paradigm will require navigating two different classes. To take VASP as an example:\r\n\r\n```\r\nVaspInputSetGenerator(user_incar_settings) -> VaspInputSet(structure) -> write_inputs()\r\n```\r\n\r\ninstead of the current one-step process:\r\n\r\n```\r\nVaspInputSet -> write_inputs()\r\n```\r\n\r\nThis creates a potential source of confusion about where to pass different kwargs - do they go to the generator or the container or the write_inputs method? Thinking beyond VASP, in some codes like LAMMPS there are kwargs that are more natural to customize when specific input files are created (like timestep, run time, etc.) rather than on class creation, whereas others like force field settings should definitely be specified at generation. With the paradigm under discussion these might be spread out among 3 places, e.g.:\r\n\r\n```\r\nLammpsInputGenerator(force_field_settings) -> LammpsInputSet(structure) -> write_inputs(timestep, run_time, etc.)\r\n```\r\n\r\nSo again there is much potential for confusion about what arguments go where; although possibly one that could be mitigated with excellent documentation. Still, I think minimizing this kind of confusion was a major objective of @utf original suggestion for the Generator / Container paradigm, so I think we to refine it a bit to keep things as simple as possible.\r\n\r\n### Proposed modified paradigm that accomplishes both objectives\r\n\r\nGoal: Accomplish easy serialization / stateless generation without requiring two classes AND facilitate a consistent interface\r\n\r\nI propose that we make the Container class _optional_ and enforce an optional `get_input_data` method in the `InputSetGenerator` that will yield a container, if defined. Although Container classes are critical for high-throughput work, parsing to databases, etc., they also create a lot of additional development overhead and are not necessary for users that don't use atomate and only want to automate job setup using pymatgen.\r\n\r\nThe key change here is that `write_inputs` would move into the generator class so that an intermediate Container class is not necessary (but still allowed). `validate` and `from_directory` would only make sense in the context of a Container class and would stay where they are.\r\n\r\nFor VASP, this would look like:\r\n\r\n```\r\nclass VaspInputSetGenerator(InputSetGenerator):\r\n \r\n def __init__(user_incar_settings, etc.):\r\n ....\r\n \r\n # optional, not needed by every class\r\n def get_input_data(structure) -> VaspInputSet:\r\n # code that creates POSCAR, POTCAR, INCAR, KPOINTS here\r\n return VaspInputSet(poscar, potcar, incar, kpoints)\r\n \r\n # optional, would require definition of a container class\r\n def from_directory(dir) -> VaspInputSet:\r\n ...\r\n\r\n # required\r\n def write_inputs(structure):\r\n # code that writes files to disk\r\n # can make use of get_input_data if it's defined, but does not have to\r\n vis = self.get_input_data()\r\n for d in [vis.incar, vis.poscar, vis.kpoints, etc.]:\r\n vis.write_file()\r\n\r\n```\r\n\r\nand for LAMMPS, with no container class:\r\n\r\n```\r\nclass LammpsInputGenerator(InputSetGenerator):\r\n\r\n def __init__(force_field_settings, etc.):\r\n ...\r\n\r\n # required\r\n def write_inputs(structure, timestep, run_time, etc.):\r\n # code that creates necessary inputs from structure and kwargs and also writes files to disk\r\n```\r\n\r\nIf we adopt this approach, we need to think carefully about class names because **the generator class would be the primary user-facing class** while the container class would primarily exist to facilitate internal work, databasing, atomate workflow, etc. In that case, I'd propose that we use the name `InputSet` for the generator class (currently `InputSetGenerator`) and come up with something new like `InputContainer` for container classes (currently `InputSet`). \r\n\r\nWe could also leave `InputContainer` (currently `InputSet`) out of the abstract interface entirely and leave it to specific code developers to implement those if they wish.\r\n\r\nThis approach retains more flexibility by not being overly prescriptive about the structure of the Container classes, and also allows us to enforce a minimal interface to `InputSetGenerator` (or whatever we call it). By locating the input generation logic in `write_inputs`(), we make it possible to serialize the class independently from structure or customizations without requiring two user-facing classes for every code.\r\n\r\nRegarding implementation:\r\n\r\n> Develop Mix-in classes specifying what is expected of an input/output class. This can be in a pymatgen/io/core module (we cannot use __init__.py since pymatgen/io is now a namespace package). I think this is better than using a strict hierarchy. This can be based on @rkingsbury initial implementation in this PR.\r\n\r\n@shyuep I'm not sure I understand Mix-in classes very well. Based on `Stringify`, it looks like these can define additional class methods but _not_ define interfaces (i.e., can't use `@abc.abstractmethod` inside a Mix-in class). Is that right, or am I misunderstanding? Here, for example `InputSet.write_inputs` is necessary for every Input class, but there is no way we could define that in a mix-in class because its contents will vary from code to code. Hence, it would need to be an abstract method.\r\n\r\nI believe this proposed modification will accomplish both of our stated objectives. @utf @shyuep please let me know what you think, and I'll move forward.\r\n\r\n\r\n",
"Hi @rkingsbury, thanks for the detailed response. I have some concerns with your proposal and still feel that having the container/generator split will provide the most clarity down the line.\r\n\r\nTo start with, most users are used to the idea of their inputs as files. Having a pymatgen class that closely maps the input files to objects requires minimal cognitive load. \r\n\r\n> I think could happen is that we wind up with Input creation code split arbitrarily across the Generator and Container classes in different ways for different codes\r\n\r\nThe separation between input set container and generator shouldn't be so difficult to decide in practice. E.g., the VaspInputSet shouldn't take a structure as input, only Incar, Kpoints, Potcar and Poscar objects. To put it another way, *you should be able to generate the container class from a directory containing the input files*, there is no ambiguity in this case over what the input set contains and so the init method shouldn't contain any logic for customising the settings.\r\n\r\n> In fact, if you look at my TemplateInputSetGenerator / TemplateInputSet implementation as it stands right now, I have some code in the InputSet __init__ method that really should be in the Generator class because I've had a hard time wrapping my head around the \"right\" way to divide things.\r\n\r\nI think part of the confusion is that the TemplateInputSet you wrote doesn't need a generator. The generator is only to create an input set with a certain set of defaults. In this case, I would just have the TemplateInputSet class and no generator.\r\n\r\n> Thinking beyond VASP, in some codes like LAMMPS there are kwargs that are more natural to customize when specific input files are created (like timestep, run time, etc.) rather than on class creation\r\n\r\nI am a little concerned that some settings are only controlled when writing the files:\r\n- I imagine it will get messy deciding how to split up the settings between the generator init and the write_inputs function. And later on down the line when someone else wants easier control of a certain setting, they will add it to the write_inputs function leading to kwarg bloat.\r\n- What happens if a user specifies one of the settings in the class init as well, how do you decide which one to use (note, respecting user_incar_settings is currently broken in several of the VASP input sets. I think this is a tricky problem to handle and having custom write_input functions will make this harder).\r\n- I can also imagine this leading to a divergence between subclasses of the generators. I.e., some generators accept some kwargs and other generators accept others.\r\n- This would break the recipe for a generic write input set fire task that you suggested in your original message.\r\n\r\nThis is one of the issues avoided by having a single container object with a single write_inputs function. There is only one way to write the inputs and it is the same whichever input generator you're using.\r\n",
"Thanks @utf ; your points are well taken. Based on what you said about the `TemplateInputSet`, what would you think about turning my proposal on its head and saying that `InputSet` (the container) is always required, whereas the generator class is optional?\r\n\r\n>I think part of the confusion is that the TemplateInputSet you wrote doesn't need a generator. The generator is only to create an input set with a certain set of defaults. In this case, I would just have the TemplateInputSet class and no generator.\r\n\r\nIf a generator class is defined, all the customization code should go in there. If it is not defined, it will just go in the `__init__` for the Container. The container class can then have the previously-discussed `from_directory` and `write_inputs` methods with no customziation.\r\n\r\nI'll update my examples to illustrate:\r\n\r\n```\r\nclass VaspInputSetGenerator(InputSetGenerator):\r\n \r\n def __init__(user_incar_settings, etc.):\r\n # all code to construct intermediate objects (e.g. incar) goes here\r\n # all customization is done with this method\r\n ....\r\n \r\n def get_input_set(structure) -> VaspInputSet:\r\n # code that generates incar, poscar, kpoints, etc.\r\n # based on user_incar_settings\r\n return VaspInputSet(incar, potcar, kpoints, structure)\r\n\r\n\r\nclass VaspInputSet(InputSet):\r\n def __init__(incar, potcar, kpoints, structure):\r\n # since we have a generator, the init is based on pre-built objects\r\n self.incar = incar\r\n self.potcar = potcar\r\n etc.\r\n\r\n def write_inputs(directory, make_dir, overwrite):\r\n ...\r\n\r\n def from_directory(directory) -> VaspInputSet:\r\n ...\r\n\r\n ...\r\n\r\n```\r\n\r\nAnd for LAMMPS, if we omit the Generator class:\r\n```\r\nclass LammpsInputSet(InputSet):\r\n\r\n def __init__(structure, timestep, run_time, force_field_settings, etc.):\r\n # In this case all the code that customizes and generates intermediate objects (if needed) is here\r\n # could be moved to a Generator class in the future if we adopt one\r\n ...\r\n\r\n def write_inputs(directory, make_dir, overwrite):\r\n ...\r\n\r\n def from_directory(directory) -> LammpsInputSet:\r\n ...\r\n\r\n```\r\n\r\nSo to summarize, if we adopted what I'm proposing in this post, what would change relative to the current state of affairs is:\r\n1. We standardize the interface for `write_inputs` and `from_directory` for all `InputSet` classes\r\n2. We have the option to make `InputSetGenerator` classes, in which case a lot of `__init__` code for customization would move from the `InputSet` to the `InputSetGenerator`. This is probably an option that will be most appropriate for the older, more complex, and more established `Input` classes and less appropriate for the newer / simpler ones.\r\n\r\nI personally like this approach a lot; I think it achieves both of our stated objectives and should also be intelligible to new users. What do you think?",
"I think that makes sense. I think you only start needing generators when you have multiple default input settings. I.e., let's say you had different input settings for a tight LAMMPS run vs coarse LAMMPS run, these should go into separate generators.",
"\n[](https://coveralls.io/builds/47143838)\n\nCoverage decreased (-0.6%) to 83.511% when pulling **eefa87963dd75c2c66a8d603f43d4d88f05f5f25 on rkingsbury:abstract_input** into **3376f27a9c5a4cac1590169f2212361ab6c94388 on materialsproject:master**.\n",
"Great! I've cleaned up the implementation per our discussion. Noteworthy changes from before include:\r\n\r\n1. Made `write_inputs()` a concrete implementation within `InputSet` so that we can standardize the approach to things like overwriting, making directory if not present, etc. To enable this I added an abstract `_generate_input_data()` method that passes data to `write_inputs()` for writing. So developers of inherited classes can focus on the logic of generating the files they want to write and not get bogged down with writing and testing basic File I/O operations.\r\n3. Removed the previous non-functional VASP demo and made `VaspInputSet` actually inherit from `InputSet`. This entailed adding `_generate_input_data()`, 'write_inputs()', and a dummy `from_directory()`. The new `write_inputs` could be used as a drop-in replacement for the existing `write_input` with a `DeprecationWarning` but I haven't done that yet. I'll leave it up to you guys whether or not you want to adopt this now, but I wanted to show what it would look like in practice.\r\n2. Revised `TemplateInputSet` to not use a generator\r\n\r\n@shyuep , let me know if you want me to move the base classes to a `pymatgen.io.core.py` module. We can also talk about adopting a mix-in class approach if you think that's preferable to the ABCs I have here; though I'll need to better understand how the two approaches differ (I'm not too familiar with mix-in classes)\r\n\r\nPlease let me know what you all think. I feel this should lay a good foundation for more standardization of our workflow components.",
"Thanks for making those edits. A couple of points:\r\n1. Based on the discussion above, `pymatgen.io.vasp.inputs.VaspInput` (rather than `pymatgen.io.vasp.sets.VaspInputSet`, which is the generator) should inherit from `InputSet` .\r\n2. What do you think about renaming `InputSet.write_inputs` -> `InputSet.write_input` as this will match the existing pymatgen input set io in the pymatgen, feff, & boltztrap modules.",
"> Thanks for making those edits. A couple of points:\r\n\r\n> 1. Based on the discussion above, `pymatgen.io.vasp.inputs.VaspInput` (rather than `pymatgen.io.vasp.sets.VaspInputSet`, which is the generator) should inherit from `InputSet` .\r\n\r\nI see what you mean. `InputSet` is supposed to be the container, whereas `InputSetGenerator` is supposed to be the generator. Right now `VaspInputSet` functions as both although `VaspInput` is the lower-level container. Ultimately I agree with your proposal but I think this will take some work to get right and is outside the scope of this PR. Is that something you'd be willing to tackle after we adopt the interface?\r\n\r\n> \r\n> 2. What do you think about renaming `InputSet.write_inputs` -> `InputSet.write_input` as this will match the existing pymatgen input set io in the pymatgen, feff, & boltztrap modules.\r\n\r\nNo objections. It's probably more intuitive without the 's' anyway, as long as it's clear that `write_input` may (depending on the code) generate multiple input files. I went ahead and made that change and commented out the legacy VASP `write_input` method for reference. For now, I've just adopted the existing arg names in the VASP classes (e.g., 'output_dir' instead of 'directory'), but in time it would be good to migrate to the ones we've defined in `InputSet`.\r\n\r\n",
"Happy Monday all! In the interest of forward progress I've gone ahead and pulled the latest master branch and converted this from WIP to ready status. The `test_get_structures_mcloud_2dstructures` is still failing for reasons that appear unrelated to any changes made here.\r\n\r\nDoes anyone have remaining, significant concerns before we make an up or down decision on this interface? I'd like to get it settled soon so that the MD group I'm working with can adopt it as we develop LAMMPS `InputSet`.\r\n\r\nObviously in this PR I have taken baby steps toward a concrete implementation for VASP, but I think it might be best to leave that for a separate PR so that @utf can add `VaspInputSetGenerator` (as desired) and we can test more thoroughly. If that makes sense to everyone, I can split out the VASP parts so that this PR contains only `InputSet`, `InputSetGenerator`, and `TemplateInputSet`.",
"A brief comment on `test_get_structures_mcloud_2dstructures`: this has started failing in CI for reasons unknown to me even though it works locally, I imagine maybe we're getting blocked by an automatic firewall somewhere. This is an _external resource_ test (located in `pymatgen.ext`) so I may disable it. It's also somewhat redundant.",
"On the PR itself, I've been deferring to @shyuep, @utf and @rkingsbury here (I wanted to avoid \"too many cooks in the kitchen\").\r\n\r\nI have referred the PR as it stands however and have no major concerns, aside from the commented-out code block.\r\n\r\nI think the VASP edits here are good and pretty minimal but if @utf or someone else would prefer to tackle VASP (and the generator) wholly in a separate PR, I'd be fine with that too.",
"Revised implementation based on the discussion around `Set`, etc. @utf @shyuep please let me know what you think.\r\n\r\nWe now have 3 classes. As before, `InputSet` is the only one strictly required for supporting a new code and is the primary user-facing one. This implementation maps more closely to the existing behavior of `VaspInput` vs. `VaspInputSet`.\r\n\r\n`InputFile`\r\n\r\n- Class to represent a single input file\r\n- Not named `Input` in order to avoid a name conflict with python built-in type as previously noted\r\n- Must implement `__str__()` and `from_string` methods which are accessed by the `write_file` and `from_file` methods, respectively\r\n- Standardizes file I/O operations in one place\r\n- OPTIONAL to implement\r\n\r\n`InputSet`\r\n\r\n- Container for one or more input files, which I refer to here as 'inputs'\r\n- Instead of inheriting from `Set`, I changed to inherit from `Mapping`. This still gives us `__len__` , `__contains__` , and `__iter__` but makes the class more \"dict-like\" and allows it to behave more like the existing `VaspInput`, e.g. `self.assertEqual(vi[\"INCAR\"][\"ALGO\"], \"Damped\")`. This also makes intuitive sense because the core data structure of `InputSet` is a mapping of filenames and file contents that is accessed by `write_input`.\r\n- Must implement `_inputs` property and `from_directory` method\r\n- The `_inputs` property is a simple mapping of filenames (strings) to either file contents (strings) or `InputFile` objects\r\n- Standardizes file IO via `write_input`. This method loops through `_inputs` and either writes the file contents string directly or calls `write_file` of the underlying `Inputfile` object.\r\n- Note that I've chosen to inherit from `Mapping` rather than `MutableMapping` because our intention with the generators is to make `InputSet` immutable (right?). This change does cause a couple of test failures at present, where tests try to mutate a `VaspInputSet` in place.\r\n\r\n`InputSetGenerator`\r\n\r\n- Class to generate `InputSet` with specific settings, as previously discussed\r\n- Must implement `get_input_set`\r\n\r\nTo illustrate how this all works, I went ahead and inherited `InputFile` in the VASP object classes (`Incar`, `Poscar`, etc.) and updated `VaspInputSet` accordingly.\r\n\r\nI also added some documentation to the root level 'Usage' section of the pymatgen docs, because I feel IO is a big part of the way people use pymatgen. The docs are written to represent a future state in which we've implemented a generator for VASP input sets.",
"Thanks for the feedback @shyuep ; I believe I've addressed all your concerns. I think the interface is basically done. \r\n\r\nI've put the PR back in WIP status pending a couple of cleanup items I want to address:\r\n\r\n- [x] Implement an `InputSetGenerator` for the SCAN input set to verify the class works as expected and to make the documentation consistent with what's actually in pymatgen\r\n- [x] Adopt `TemplateInputSet` in LAMMPS to replace existing `write_lammps_inputs`\r\n- [x] Attempt preliminary `Q-Chem` integration to identify any other unforeseen problems with the interface\r\n- [x] Make another editing pass through all new docstrings and documentation",
"One unresolved issue is what to do about mutable `VaspInputSet`. In this PR we've discussed that `InputSet` should not be mutable; you should use a generator to create a new one rather than mutate in-place. However, in the past we have sometimes mutated VASP input sets (maybe others). Right now this leads to a few test failures like:\r\n\r\n```\r\n def get_vasp_input(self, vasp_input_set=MPRelaxSet, **kwargs):\r\n \"\"\"\r\n Returns VASP input as a dict of vasp objects.\r\n \r\n Args:\r\n vasp_input_set (pymatgen.io.vaspio_set.VaspInputSet): input set\r\n to create vasp input files from structures\r\n \"\"\"\r\n d = vasp_input_set(self.final_structure, **kwargs).get_vasp_input()\r\n> d[\"transformations.json\"] = json.dumps(self.as_dict())\r\nE TypeError: 'VaspInput' object does not support item assignment\r\n```\r\n\r\nWhat I propose to do _for now_ is to inherit from `MutableMapping` instead of `Mapping` and implement a `__setitem__` method so that mutation in place is possible. However, doing, so will raise a warning notifying the user that this type of usage is not supported and will be retired in the future."
] | 2021-08-06T18:52:58
| 2022-08-10T14:01:40
|
2022-03-14T14:05:10Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
After much discussion and feedback, here is a draft abstract interface for pymatgen I/O classes.
Thanks @utf @shyamd @arepstein @mkhorton and @computron for your thoughts so far, and feel free to add more here!
## Motivation
By defining a minimal interface for *all* `Input` classes in pymatgen, we can
1. Make it easier for users to learn pymatgen IO for different codes by providing more consistent interface
2. Make it easier for users to adapt scripts and workflows for different codes
3. Make `FireTask` or `FireWork` call signatures more consistent
4. Make `FireTasks` and `FireWorks` more portable among different codes, lowering barriers to new workflow development
## Objective
The vision here is to define a minimal common interface for *all* `Input` classes in pymatgen, whether for solid DFT, molecular DFT, classical MD, AIMD, etc.
There are two classes. `Input` is a data container holding all input data for a specific calculation. It defines `write_inputs` (required) and `from_files` (optional) methods , and an `is_valid` (optional) property to perform basic input validation.
`InputSettings` is a generator class that holds _settings_ or _instructions_ for how to create an `Input`, given a structure. It's only requirement is a `get_input()` method.
Each code will subclass `Input` one time, e.g. `VaspInput(Input)`, and subclass `InputSettings` multiple times for each type of calculation, e.g. `VaspRelaxation(InputSettings)`, `VaspStatic(InputSettings)`, etc.
For barebones support of any code, it is possible to use`Input` directly without writing an associated `InputSettings` class (see the `TemplateInput` example).
## Implementation
This PR is a vehicle for iterating on the interface. Refactoring existing I/O classes to use the interface is not in scope for this PR. However, I have included a pseudo-functional example VASP implementation to illustrate how this interface could be used in practice. I may add other pseudo-examples, but will leave functional implementations for later.
I also provide a functional concrete implementation of `Input` (`TemplateInput`) that gives a low-barrier way for new users to transition from bash scripts to pymatgen I/O classes.
## Implications for `atomate`
Once we have a consistent call signature for `Input`, it should be possible to write a generic `FireTask` for writing inputs that will work with many different codes, e.g.
```python
class WriteInput(FireTaskBase):
"""
Args:
input_set: Input class
calc_dir: Union[str, Path]
"""
required_params = ["input_set", "calc_dir"]
def run_task(self, fw_spec):
input_set = self["input_set"]
calc_dir = self["calc_dir"]
input_set.write_inputs(calc_dir)
```
This would dramatically lower the barrier to supporting new codes in `atomate`. As long as a code has the associated `Input` class defined, the `FireTask` could be used without modification. Of course customized and more complex `FireTasks` may still be needed and can be built just as they are today.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2211/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2211/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2211",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2211",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2211.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2211.patch",
"merged_at": "2022-03-14T14:05:10Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2212
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2212/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2212/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2212/events
|
https://github.com/materialsproject/pymatgen/issues/2212
| 963,306,640
|
MDU6SXNzdWU5NjMzMDY2NDA=
| 2,212
|
E-above-convex-hull equals 0 for structures with different energy
|
{
"login": "Evmoerman",
"id": 76110846,
"node_id": "MDQ6VXNlcjc2MTEwODQ2",
"avatar_url": "https://avatars.githubusercontent.com/u/76110846?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Evmoerman",
"html_url": "https://github.com/Evmoerman",
"followers_url": "https://api.github.com/users/Evmoerman/followers",
"following_url": "https://api.github.com/users/Evmoerman/following{/other_user}",
"gists_url": "https://api.github.com/users/Evmoerman/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Evmoerman/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Evmoerman/subscriptions",
"organizations_url": "https://api.github.com/users/Evmoerman/orgs",
"repos_url": "https://api.github.com/users/Evmoerman/repos",
"events_url": "https://api.github.com/users/Evmoerman/events{/privacy}",
"received_events_url": "https://api.github.com/users/Evmoerman/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I am not sure what you mean. I ran your code and my output is:\r\n```\r\n##############################################################\r\nenergy Difference (eV): 0.19\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.0010000000002037268\r\n(mavrl) sp@Shyues-Mac-mini py % python ehull.py\r\n##############################################################\r\nenergy Difference (eV): 0.19\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.01900000000023283\r\n(mavrl) sp@Shyues-Mac-mini py % python ehull.py\r\n##############################################################\r\nenergy Difference (eV): 0.0\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0\r\n##############################################################\r\nenergy Difference (eV): 0.01\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.0010000000002037268\r\n##############################################################\r\nenergy Difference (eV): 0.02\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.00200000000018008\r\n##############################################################\r\nenergy Difference (eV): 0.03\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.003000000000156433\r\n##############################################################\r\nenergy Difference (eV): 0.04\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.004000000000132786\r\n##############################################################\r\nenergy Difference (eV): 0.05\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.005000000000109139\r\n##############################################################\r\nenergy Difference (eV): 0.06\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.0060000000000854925\r\n##############################################################\r\nenergy Difference (eV): 0.07\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.007000000000061846\r\n##############################################################\r\nenergy Difference (eV): 0.08\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.008000000000038199\r\n##############################################################\r\nenergy Difference (eV): 0.09\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.009000000000014552\r\n##############################################################\r\nenergy Difference (eV): 0.1\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.010000000000218279\r\n##############################################################\r\nenergy Difference (eV): 0.11\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.011000000000194632\r\n##############################################################\r\nenergy Difference (eV): 0.12\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.012000000000170985\r\n##############################################################\r\nenergy Difference (eV): 0.13\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.012999999999919964\r\n##############################################################\r\nenergy Difference (eV): 0.14\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.014000000000123691\r\n##############################################################\r\nenergy Difference (eV): 0.15\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.015000000000100044\r\n##############################################################\r\nenergy Difference (eV): 0.16\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.016000000000076398\r\n##############################################################\r\nenergy Difference (eV): 0.17\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.01700000000005275\r\n##############################################################\r\nenergy Difference (eV): 0.18\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.018000000000029104\r\n##############################################################\r\nenergy Difference (eV): 0.19\r\nE-above-hull for structure Ca8As2 151: 0\r\nE-above-hull for structure Ca8As2 727: 0.01900000000023283\r\n```\r\n\r\nThis is perfectly in line with the results. Note that ehull is always a energy per atom quantity. ",
"Huh, that's interesting. In my case it was displaying an e-eabove-hull of 0 for both compounds for _energy Difference (eV): 0.0_\r\nup to _energy Difference (eV): 0.13_ and only for higher energy differences one of the compounds stayed on the convex hull and the other had a finite value.\r\n\r\nPerhaps it's just a classical case of not using the most up-to-date version.\r\nI'll check.\r\n\r\nThanks for having a look.",
"Indeed, I was using version 2022.0.6. With 2022.0.11 it returns the correct (that is, your) result.\r\nThanks again."
] | 2021-08-07T21:25:47
| 2021-08-24T15:34:41
|
2021-08-24T15:34:41Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Dear ladies and gentlemen,
In my calculations it happens regularly that two `ComputedEntry`-instances with identical composition and sufficiently different total energies are both given an **energy-above-hull of 0** by `get_decomp_and_e_above_hull()`.
The total energy difference of the two compounds in question **can be as high as 0.12 eV** and still both are assigned e_above_hull = 0.
I'm not sure if this is a bug or if I use the `PhaseDiagram` class of pymatgen incorrectly.
Here's a minimal example script where this is demonstrated:
###########################################################
```
import pymatgen.core as mg
from pymatgen.entries.computed_entries import ComputedEntry
from pymatgen.analysis.phase_diagram import PhaseDiagram, PDPlotter
composition151 = mg.Composition('As8Ca2')
composition727 = mg.Composition('As8Ca2')
#Elements
#As8
composition810 = mg.Composition('As8')
#Ca16
composition243 = mg.Composition('Ca16')
for i in range(20):
ComputedEntry1 = ComputedEntry(composition151, -12106.046992071064)
ComputedEntry2 = ComputedEntry(composition727, -12106.046992071064 + i*0.01)
ComputedEntry3 = ComputedEntry(composition810, -659.3308195375063*8)
ComputedEntry4 = ComputedEntry(composition243, -3406.144052584896*16)
pdEntries = [ComputedEntry1, ComputedEntry2, ComputedEntry3, ComputedEntry4]
pd = PhaseDiagram(pdEntries)
decomp, eAboveHull_151 = pd.get_decomp_and_e_above_hull(ComputedEntry1)
decomp, eAboveHull_727 = pd.get_decomp_and_e_above_hull(ComputedEntry2)
print("##############################################################")
print("energy Difference (eV): ", i*0.01)
print("E-above-hull for structure Ca8As2 151: ", eAboveHull_151)
print("E-above-hull for structure Ca8As2 727: ", eAboveHull_727)
```
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2212/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2212/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2213
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2213/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2213/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2213/events
|
https://github.com/materialsproject/pymatgen/pull/2213
| 963,897,193
|
MDExOlB1bGxSZXF1ZXN0NzA2NDM1NjI2
| 2,213
|
Fix minor bug when reading Structure from a netcdf4 file with hdf5 groups
|
{
"login": "gmatteo",
"id": 3811747,
"node_id": "MDQ6VXNlcjM4MTE3NDc=",
"avatar_url": "https://avatars.githubusercontent.com/u/3811747?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/gmatteo",
"html_url": "https://github.com/gmatteo",
"followers_url": "https://api.github.com/users/gmatteo/followers",
"following_url": "https://api.github.com/users/gmatteo/following{/other_user}",
"gists_url": "https://api.github.com/users/gmatteo/gists{/gist_id}",
"starred_url": "https://api.github.com/users/gmatteo/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/gmatteo/subscriptions",
"organizations_url": "https://api.github.com/users/gmatteo/orgs",
"repos_url": "https://api.github.com/users/gmatteo/repos",
"events_url": "https://api.github.com/users/gmatteo/events{/privacy}",
"received_events_url": "https://api.github.com/users/gmatteo/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/41998310)\n\nCoverage decreased (-0.6%) to 83.13% when pulling **fa70861ca4ce4bae19aa87f8103ba26063b9119b on gmatteo:master** into **e5fd7f5d460025a3f9311659eae722128aa43e67 on materialsproject:master**.\n",
"The [pylint step](https://github.com/materialsproject/pymatgen/runs/3279758523?check_suite_focus=true) fails due to the following error.\r\n\r\n```\r\nRun pylint pymatgen\r\n pylint pymatgen\r\n shell: /usr/bin/bash -e {0}\r\n env:\r\n pythonLocation: /opt/hostedtoolcache/Python/3.9.6/x64\r\n LD_LIBRARY_PATH: /opt/hostedtoolcache/Python/3.9.6/x64/lib\r\n************* Module pymatgen.entries.computed_entries\r\npymatgen/entries/computed_entries.py:704:8: E1101: Instance of 'ComputedStructureEntry' has no '_composition' member (no-member)\r\n```\r\n\r\nbut this PR does not affect the implementation of *computed_entries*."
] | 2021-08-09T11:24:21
| 2021-08-09T14:04:57
|
2021-08-09T14:04:57Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Fix minor bug when reading Structure from a netcdf4 file with hdf5 groups
* Move abiinspect module to abipy to facilitate new developments
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2213/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2213/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2213",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2213",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2213.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2213.patch",
"merged_at": "2021-08-09T14:04:57Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2214
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2214/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2214/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2214/events
|
https://github.com/materialsproject/pymatgen/issues/2214
| 965,955,010
|
MDU6SXNzdWU5NjU5NTUwMTA=
| 2,214
|
get_entries_in_chemsys reports TypeError in the latest version (2022.0.11)
|
{
"login": "jx-fan",
"id": 54628903,
"node_id": "MDQ6VXNlcjU0NjI4OTAz",
"avatar_url": "https://avatars.githubusercontent.com/u/54628903?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jx-fan",
"html_url": "https://github.com/jx-fan",
"followers_url": "https://api.github.com/users/jx-fan/followers",
"following_url": "https://api.github.com/users/jx-fan/following{/other_user}",
"gists_url": "https://api.github.com/users/jx-fan/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jx-fan/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jx-fan/subscriptions",
"organizations_url": "https://api.github.com/users/jx-fan/orgs",
"repos_url": "https://api.github.com/users/jx-fan/repos",
"events_url": "https://api.github.com/users/jx-fan/events{/privacy}",
"received_events_url": "https://api.github.com/users/jx-fan/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I used the exact code you paste and did not get any error. I suspect your install may be out of date. Can you remove pymatgen and reinstall and try again?",
"> I used the exact code you paste and did not get any error. I suspect your install may be out of date. Can you remove pymatgen and reinstall and try again?\r\n\r\nHi Shyue Ping,\r\n\r\nThe error was found when running a pip-installed pymatgen on our HPC cluster:\r\n```\r\npython-3.8.5\r\npip-21.2.4\r\npymatgen-2022.0.11\r\nGadi supercomputer, NCI Australia, as you may know\r\n```\r\nLater on I tried to test it out on my local pc using the recommended conda-forge, the problem was gone. I feel this error might have something to do with the pip install but I don't have a clue yet. I'll just stick to the local conda version by now and thank you very much for your valuable advice. Best wishes, Jiaxin",
"On your HPC, I would suggest you do a local conda install, followed by a conda virtual env and then a pip or conda install in that virutal env. On public systems, there is no way to verify what has actually been installed in the main system or whether it has been properly kept up to date. pymatgen.org provides instructions on computing clusters."
] | 2021-08-11T05:29:40
| 2021-08-15T14:39:47
|
2021-08-15T14:39:47Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
After upgrading to version 2022.0.11, I was trying to use `get_entries_in_chemsys` to obtain entries in a high dimentional chemical space. However a TypeError occurred, which pointed to the compatibility module under entries.
**To Reproduce**
The chemical system I demonstrated here is from pymatgen user manual (https://pymatgen.org/usage.html#:~:text=Finally%2C%20the%20MPRester,can%20be%20determined%3A). Below is the test code:
```
from pymatgen.ext.matproj import MPRester
with MPRester() as m:
mp_entries = m.get_entries_in_chemsys(["Li", "Fe", "O"])
print(len(mp_entries))
```
After running this simple snippet, the error message looks like this:
```
Traceback (most recent call last):
File "sample.py", line 4, in <module>
mp_entries = m.get_entries_in_chemsys(["Li", "Fe", "O"])
File "/.local/lib/python3.8/site-packages/pymatgen/ext/matproj.py", line 872, in get_entries_in_chemsys
entries = self.get_entries(
File "/.local/lib/python3.8/site-packages/pymatgen/ext/matproj.py", line 565, in get_entries
entries = MaterialsProject2020Compatibility().process_entries(entries, clean=True)
File "/.local/lib/python3.8/site-packages/pymatgen/entries/compatibility.py", line 559, in process_entries
for entry in PBar(entries, disable=(not verbose)):
TypeError: __init__() got an unexpected keyword argument 'disable'
```
**Expected behavior**
When I tested with an older version 2022.0.5, I can get the normal output, with the size of chemical space as 362:
```
python3 sample.py
/.local/lib/python3.8/site-packages/pymatgen/ext/matproj.py:567: FutureWarning: __init__ is deprecated
MaterialsProjectCompatibility will be updated with new correction classes as well as new values of corrections and uncertainties in 2020
entries = MaterialsProjectCompatibility().process_entries(entries)
362
```
Do you guys have any thoughts if there's any compatibility issue with the recently updated version?
Cheers
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2214/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2214/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2215
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2215/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2215/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2215/events
|
https://github.com/materialsproject/pymatgen/pull/2215
| 967,375,170
|
MDExOlB1bGxSZXF1ZXN0NzA5NTUyODE1
| 2,215
|
QChem: add CMIRS solvent model support
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/54037541)\n\nCoverage remained the same at 0.0% when pulling **df3a87952d3303699f05292ee04c3d6486b670dd on rkingsbury:cmirs** into **03691a8fdb8bc68db1f23a95930f71a07ffef4df on materialsproject:master**.\n",
"@samblau @espottesmith I want to flag this for you guys to start reviewing as it's 95% done. I'm leaving in WIP status b/c I want to add output parsing before merging, but the input setup parts are solid. I particularly want to call attention to the change in `lower_and_check_unique` that guarantees all numerical values get stored as `str`. It appears innocuous but you might know of situations where that would cause problems.",
"OK, I have the basics of output parsing in place now. However I'm having difficulty understanding how `test_outputs.py` works. I'd also like some guidance on how to structure the output keys. I'll await feedback from @espottesmith and @samblau before moving ahead.\r\n\r\nSpecifically:\r\n\r\n1. I added 4 test files (currently commented out) to `multi_job_out_names` in `test_outputs.py`, and then in a terminal ran `TestQCOutput.generate_multi_job_dict()` to update the .json file. But even after doing this I got a `KeyError` on the new filenames when I tried to run the tests. Note that I have not added any of the new properties to the test yet; I just wanted to make sure they ran cleanly after adding the new files.\r\n\r\n3. Currently I only parse the _energies_ (and dielectric constant) that come out of ISOSVP / CMIRS calculations. This appears to be the approach taken for SMD as well. However unlike SMD, in CMIRS the solvents are not selected by name, but rather by specifying parameters in the `$pcm_nonels section`. Moreover, the only place these parameters show up in the output file is in the `$pcm_nonels` section of the input block at the beginning; they aren't listed elsewhere. So there is no obvious way to determine what the solvent is. I could try construct a `QCInput` from the input block in the output file, but that seems a bit convoluted considering that ultimately we will have a `TaskDocument` with a `calcs_reversed[0]` that will do that for us, and perhaps we can use a builder to map the solvent parameters back to named solvent?\r\n\r\n4. data field names. From what I can tell, PCM and SMD populate completely different keys in the `solvent_data` block, with no overlap. I've taken the same approach here; although there's an argument that we should populate a single `dielectric` field whether we're using PCM, CMIRS, or ISOSVP. I made the field names similar to what is listed in the output file, but feedback there is welcome. For example, it might make sense to prefix the field names with `isosvp` or `cmirs` to make clear what's what in the final task doc, because there are going to be a lot of fields with similar names now that we have four possible solvent models *e.g. `solute_internal_energy` for SMD vs. `change_solute_internal_e` for ISOSVP",
"@samblau ready for another review. Per our discussion, I went ahead and nested the solvent data under `cmirs` and`isosvp` keys in the `solvent_data` dict. I held off doing so for PCM and SMD b/c we think that should be in a separate PR.\r\n\r\nI will test this with atomate soon and report back if I encounter any problems.",
"UPDATE: I've done some testing of this with `atomate` and can confirm the following:\r\n- Task docs are still populated with `float` (e.g. `solvent_data.pcm_dielectric` is still a float) despite the change in `lower_and_check_unique`.\r\n- PCM and SMD calculations parse correctly and now include `None` for keys associated with other solvents (i.e., all `solvent_data` keys are always present for any solvent; those that aren't relevant are `None`).\r\n- ISOSVP and CMIRS Calculations set up as intended and run successfully\r\n\r\n**However**\r\nI've also found one or more \"fun\" undocumented behaviors / bugs in Q-Chem that cause some calculations to completely skip the solvation phase! So although it looks like an ISOSVP calculation has completed successfully, the `solvent_data.isosvp` information will be blank because that phase of the calculation was actually skipped, and the associated information isn't in the output file.\r\n\r\nI'm working on pinning down what's going on here, and that might warrant some small modifications to the IO. See [forum post](https://talk.q-chem.com/t/cmirs-isosvp-unexpected-behavior/835)",
"Alright, I think all issues have been resolved and this is working as intended. I learned (the hard way) that you have to disable `gen_scfman` in order for ISOSVP or CMIRS to work, and add a variable `IPNRF` to the $SVP section in order for CMIRS to work (neither of which is documented right now in the Q-Chem manual - see [forum post](https://talk.q-chem.com/t/cmirs-isosvp-unexpected-behavior/835)).\r\n\r\nAfter adding those changes to our IO, I have successfully set up and run single point calcs using all four solvent models (PCM, SMD, ISOSVP, and CMIRS) and inspected the `solvent_data` key of the resulting task docs. See an example below for a CMIRS calculation. All keys are present always, and assigned values of `None` if they don't apply.\r\n\r\nUnless @samblau or @espottesmith spot anything problematic, I think this is ready to merge.\r\n\r\n<img width=\"785\" alt=\"image\" src=\"https://user-images.githubusercontent.com/1908695/200708577-21141c2b-a022-4eb8-b3f7-ea9c0a078981.png\">",
"I'd like to go ahead and merge this -- @samblau, @espottesmith unless you have any objections, will go ahead.",
"@mkhorton looks good to me! merge away",
"Thanks @rkingsbury, merged :) "
] | 2021-08-11T20:07:42
| 2024-10-10T01:54:53
|
2022-11-22T19:57:08Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Additions to `QCInput` and `QChemDictSet` needed to support the new CMIRS implicit solvent model and the isodensity implementation of SS(V)PE.
See relevant Q-Chem manual pages:
- https://manual.q-chem.com/latest/subsec_PCM_job_control.html
- https://manual.q-chem.com/latest/subsec_SS(V)PE.html
- https://manual.q-chem.com/latest/example_CMIRS-water.html
### Specific changes include:
* Support `$svp` and `$pcm_nonels` sections in `QCInput`
* Add `isosvp_dielectric` and `cmirs_solvent` kwargs and documentation to `QChemDictSet`and all inherited `InputSet` classes
* Add warnings or errors as appropriate to alert users to combinations of inputs with non-obvious results
* Small bugfix to remedy the fact that Q-Chem does not recognize 'dimethyl sulfoxide' as a valid solvent even though that is the keyword given in the manual. `DMSO` works.
* **Noteworthy change** - modify `lower_and_check_unique` utility such that it converts all values (including numeric values) to strings. This was necessary to facilitate reading/writing numerical parameters to and from files. the already-existing `pcm_defaults` dictionary used `str` to store its numerical parameters, so this change appears to be consistent with the original design intent and it appears not to break anything. **However**, as a result of that change, the `dielectric` attribute is now a `str` and not a `float`.
* Fix a bug regarding duplicate key removal and add tests for `lower_and_check_unique`
* Update header info (authors, copyright date, etc.) in all files. The `__version__=0.1` didn't seem meaningful and it didn't make sense to have Brandon's stale email in there
* Add output parsing. CMIRS and ISOSVP solvent data are nested under `solvent_data.cmirs` and `solvent_data.isosvp`, respectively. Per discussion with @samblau , this is in preparation for nesting PCM and SMD data in a similar way, which we plan to do in a separate PR.
* Add tests for all new functionality, including units tests and multi-job output files.
### TODO
* Manually test within atomate workflow
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2215/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2215/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2215",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2215",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2215.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2215.patch",
"merged_at": "2022-11-22T19:57:08Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2216
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2216/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2216/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2216/events
|
https://github.com/materialsproject/pymatgen/issues/2216
| 971,057,757
|
MDU6SXNzdWU5NzEwNTc3NTc=
| 2,216
|
Wrong POSCAR and .cif file generated when using `Structure.to` method
|
{
"login": "755452800",
"id": 18474386,
"node_id": "MDQ6VXNlcjE4NDc0Mzg2",
"avatar_url": "https://avatars.githubusercontent.com/u/18474386?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/755452800",
"html_url": "https://github.com/755452800",
"followers_url": "https://api.github.com/users/755452800/followers",
"following_url": "https://api.github.com/users/755452800/following{/other_user}",
"gists_url": "https://api.github.com/users/755452800/gists{/gist_id}",
"starred_url": "https://api.github.com/users/755452800/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/755452800/subscriptions",
"organizations_url": "https://api.github.com/users/755452800/orgs",
"repos_url": "https://api.github.com/users/755452800/repos",
"events_url": "https://api.github.com/users/755452800/events{/privacy}",
"received_events_url": "https://api.github.com/users/755452800/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Sorry it's my mistake, we can set `coords_are_cartesian` when construct the `Structure` object, so this is not a bug.\r\n"
] | 2021-08-15T05:52:40
| 2021-08-15T12:55:12
|
2021-08-15T12:55:12Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Hi, I was using Pymatgen `Structure` object to write a POSCAR and cif file using `to` method, but the files generated were wrong when using cartesian coordinates.
**To Reproduce**
Steps to reproduce the behavior:
1. generate a pymatgen `Structure` object using cartesian coordinates.
```
from pymatgen.core import Structure
species = [13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 13.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 45.0, 7.0, 1.0]
coords = [[7.025555610656738, 0.0, 12.734589576721191], [6.965488433837891, 10.959248542785645, 17.195571899414062], [1.079264521598816, 5.479624271392822, 14.978279113769531], [-1.9239145517349243, 13.699060440063477, 18.330615997314453], [4.022376537322998, 8.219436645507812, 16.086925506591797], [9.968667030334473, 2.739812135696411, 13.84323501586914], [3.7433207035064697, 12.487018585205078, 14.127511978149414], [9.689611434936523, 7.007394790649414, 11.883822441101074], [0.8452595472335815, 1.5277702808380127, 17.519445419311523], [-2.1738016605377197, 12.644883155822754, 13.752876281738281], [3.772489309310913, 7.165258884429932, 11.509185791015625], [6.760651588439941, 1.6856346130371094, 17.118412017822266], [8.422892570495605, 14.911102294921875, 11.406659126281738], [-0.4214596450328827, 9.431478500366211, 17.042282104492188], [5.524831295013428, 3.9518539905548096, 14.798592567443848], [2.5199215412139893, 12.487018585205078, 17.37525749206543], [8.466212272644043, 7.007394790649414, 15.13156795501709], [2.579988956451416, 1.5277702808380127, 12.914276123046875], [1.2841013669967651, 14.7532377243042, 15.055439949035645], [7.230391979217529, 9.273613929748535, 12.811749458312988], [10.218554496765137, 3.793989658355713, 18.420974731445312], [7.199493408203125, 14.911102294921875, 14.65440559387207], [1.3132697343826294, 9.431478500366211, 12.437112808227539], [4.301432132720947, 3.9518539905548096, 18.04633903503418], [9.023781776428223, 10.220545768737793, 15.377436637878418], [3.1375579833984375, 4.740921974182129, 13.160144805908203], [3.0774905681610107, 15.700170516967773, 17.621126174926758], [0.20023702085018158, 0.9445738196372986, 14.647156715393066], [3.0982983112335205, 11.90382194519043, 11.255224227905273], [6.086460590362549, 6.424198150634766, 16.86444854736328], [-1.5882883071899414, 15.073429107666016, 15.863566398620605], [4.358002662658691, 9.593805313110352, 13.61987590789795], [7.346164703369141, 4.114181041717529, 19.229101181030273], [-1.561689853668213, 10.220545768737793, 12.093321800231934], [1.426472544670105, 4.740921974182129, 17.702547073364258], [4.324533939361572, 15.700170516967773, 14.310613632202148], [-0.30198556184768677, 7.910528659820557, 14.45797348022461], [5.644305229187012, 2.4309046268463135, 12.21428394317627], [5.584238052368164, 13.390152931213379, 16.67526626586914], [4.967262268066406, 0.7387025952339172, 14.55272388458252], [7.865323543548584, 11.697951316833496, 11.16079044342041], [-0.9790287613868713, 6.218327045440674, 16.79641342163086], [2.460514783859253, 3.048719644546509, 15.498584747314453], [5.35857629776001, 14.007967948913574, 12.106651306152344], [8.346738815307617, 8.52834415435791, 17.715877532958984], [-0.34124520421028137, 15.073429107666016, 12.55305290222168], [2.6469171047210693, 9.593805313110352, 18.162277221679688], [8.593208312988281, 4.114181041717529, 15.918587684631348], [3.720219135284424, 0.7387025952339172, 17.863237380981445], [6.618280410766602, 11.697951316833496, 14.471303939819336], [0.732056736946106, 6.218327045440674, 12.254012107849121], [0.6985880136489868, 12.324691772460938, 12.94474983215332], [3.6867501735687256, 6.845067501068115, 18.553974151611328], [9.633041381835938, 1.365443229675293, 16.310285568237305], [-0.5484552979469299, 12.324691772460938, 16.25526237487793], [5.397835731506348, 6.845067501068115, 14.01157283782959], [11.34412670135498, 1.365443229675293, 11.767882347106934], [1.9582922458648682, 10.014674186706543, 15.309401512145996], [7.904582977294922, 4.535050392150879, 13.065711975097656], [7.844515800476074, 15.494298934936523, 17.52669334411621], [-0.4273305833339691, 10.502613067626953, 14.410757064819336], [5.518960475921631, 5.022988796234131, 12.167067527770996], [5.458893299102783, 15.982237815856934, 16.628049850463867], [2.585859775543213, 0.45663538575172424, 15.545801162719727], [5.483921051025391, 11.415884017944336, 12.153867721557617], [8.47208309173584, 5.936259746551514, 17.763093948364258], [0.7029937505722046, 14.357982635498047, 17.574899673461914], [6.649284839630127, 8.878358840942383, 15.331209182739258], [0.7630611062049866, 3.3987345695495605, 13.113917350769043], [6.608130931854248, 16.386812210083008, 12.577348709106445], [9.596293449401855, 10.907187461853027, 18.186574935913086], [3.7100698947906494, 5.427562713623047, 15.969282150268555], [8.433147430419922, 13.040138244628906, 16.003175735473633], [2.5469236373901367, 7.560513973236084, 13.785883903503418], [5.535085678100586, 2.0808897018432617, 19.395109176635742], [2.509666919708252, 14.357982635498047, 12.778741836547852], [5.497829437255859, 8.878358840942383, 18.387968063354492], [-0.38839447498321533, 3.3987345695495605, 16.17067527770996], [4.334682941436768, 11.011309623718262, 16.20456886291504], [10.280974388122559, 5.531685829162598, 13.960878372192383], [1.4366217851638794, 0.05206125229597092, 19.59650230407715], [-1.5926942825317383, 13.040138244628906, 11.233416557312012], [1.3954681158065796, 7.560513973236084, 16.842641830444336], [7.341759204864502, 2.0808897018432617, 14.59895133972168], [10.074357986450195, 0.6700782179832458, 18.050220489501953], [9.816859245300293, 1.2978516817092896, 18.81179428100586]]
laticce = [[ 1.1833e+01, 0.0000e+00, -2.6398e-02],
[-3.0482e+00, 1.6439e+01, -1.1482e+00],
[ 0.0000e+00, 0.0000e+00, 3.1385e+01]]
struc = Structure(laticce, species, coords)
print(struc)
struc.to(fmt='poscar', filename='POSCAR')
```
2. Check the POSCAR generated mentioned above (see attached fig 1), we can see the cartesian coordinates but with a wrong indication showing `direct`, which is wrong and contradictory. The file will be wrongly displayed.
**Expected behavior**
The `to` method should wisely recognize whether the coordinates are cartesian coordinates or direct coordinates and write the corresponding identifier (direct or cartesian in line 8 in POSCAR).
**Screenshots**

Fig .1
**Desktop (please complete the following information):**
- OS: Linux
- Version 2020.8.13 (haven't check the latest version but checked the code and it seems the latest version also has this problem)
**Additional context**
I think the problem occurred because the default setting of `coords_are_cartesian` in `pymatgen/pymatgen/core/structure.py` is not properly set when feeding the `Structure` object cartesian coordinates
|
{
"login": "755452800",
"id": 18474386,
"node_id": "MDQ6VXNlcjE4NDc0Mzg2",
"avatar_url": "https://avatars.githubusercontent.com/u/18474386?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/755452800",
"html_url": "https://github.com/755452800",
"followers_url": "https://api.github.com/users/755452800/followers",
"following_url": "https://api.github.com/users/755452800/following{/other_user}",
"gists_url": "https://api.github.com/users/755452800/gists{/gist_id}",
"starred_url": "https://api.github.com/users/755452800/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/755452800/subscriptions",
"organizations_url": "https://api.github.com/users/755452800/orgs",
"repos_url": "https://api.github.com/users/755452800/repos",
"events_url": "https://api.github.com/users/755452800/events{/privacy}",
"received_events_url": "https://api.github.com/users/755452800/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2216/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2216/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2217
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2217/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2217/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2217/events
|
https://github.com/materialsproject/pymatgen/pull/2217
| 971,120,475
|
MDExOlB1bGxSZXF1ZXN0NzEyOTI2MDk4
| 2,217
|
fix issue #2216
|
{
"login": "755452800",
"id": 18474386,
"node_id": "MDQ6VXNlcjE4NDc0Mzg2",
"avatar_url": "https://avatars.githubusercontent.com/u/18474386?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/755452800",
"html_url": "https://github.com/755452800",
"followers_url": "https://api.github.com/users/755452800/followers",
"following_url": "https://api.github.com/users/755452800/following{/other_user}",
"gists_url": "https://api.github.com/users/755452800/gists{/gist_id}",
"starred_url": "https://api.github.com/users/755452800/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/755452800/subscriptions",
"organizations_url": "https://api.github.com/users/755452800/orgs",
"repos_url": "https://api.github.com/users/755452800/repos",
"events_url": "https://api.github.com/users/755452800/events{/privacy}",
"received_events_url": "https://api.github.com/users/755452800/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This is not a valid check. Fractional coordinations can also be less than 0 and greater than 1, even if they map to the unit cell. This is frequently used when you are working with trajectories. The reason why the coordinates are defaulted to fractional for crystals is because this is by far the most common way coordinates are specified. ",
"Oh, really sorry for the mistake and the wrong pull request.",
"There is no way to check this. A set of coordinates can be cartesian or fractional. Of course, the likelihood is that small numbers mean fractional and large numbers are more likely to be cartesian. But this is not always the case. It is the responsibility of the user to be awre what coordinates they are entering.",
"Thanks for answering!"
] | 2021-08-15T12:16:38
| 2021-08-15T15:59:02
|
2021-08-15T12:20:31Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Add a few lines code to exam the coords fed to `Structure` object are cartesian or not.
The setting of `coords_are_cartesian` was defaulted to `False` and problem may occur when feeding cartesian coords to `Structure`.
* Fix issue #2216
## Additional dependencies introduced (if any)
None
## TODO (if any)
I noticed there are several codes set `coords_are_cartesian` to `False` by default, but I'm not sure if they are properly set when cartesian coords are fed and may be examed first like this case.
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2217/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2217/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2217",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2217",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2217.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2217.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2218
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2218/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2218/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2218/events
|
https://github.com/materialsproject/pymatgen/pull/2218
| 972,105,830
|
MDExOlB1bGxSZXF1ZXN0NzEzNzUwMzA0
| 2,218
|
Add `chempot_diagram` module for creation/plotting of chemical potential diagrams
|
{
"login": "mattmcdermott",
"id": 40469694,
"node_id": "MDQ6VXNlcjQwNDY5Njk0",
"avatar_url": "https://avatars.githubusercontent.com/u/40469694?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mattmcdermott",
"html_url": "https://github.com/mattmcdermott",
"followers_url": "https://api.github.com/users/mattmcdermott/followers",
"following_url": "https://api.github.com/users/mattmcdermott/following{/other_user}",
"gists_url": "https://api.github.com/users/mattmcdermott/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mattmcdermott/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mattmcdermott/subscriptions",
"organizations_url": "https://api.github.com/users/mattmcdermott/orgs",
"repos_url": "https://api.github.com/users/mattmcdermott/repos",
"events_url": "https://api.github.com/users/mattmcdermott/events{/privacy}",
"received_events_url": "https://api.github.com/users/mattmcdermott/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/42531927)\n\nCoverage decreased (-0.6%) to 83.156% when pulling **09d0397c323e3ef624c471a9721a2c986a5ad44e on mattmcdermott:chempot_diagrams** into **13d7c12a822fb64e47a44cc09bdf3b9dcaa0e76d on materialsproject:master**.\n",
"@mkhorton Any thoughts on the content of this PR before I finish docstrings and add tests? \r\n\r\nOne of the things that may feel weird for normal pymatgen users is that I included a `get_plot()` method within the diagram class itself (as opposed to implementing a new `CPDPlotter` or something of the sort). I am not a huge fan of `PDPlotter` in general as I largely think it is a useless class.... wasn't sure what your thoughts were on this?",
"I also have issues with `PDPlotter` (and have a fork somewhere, actually, where I'm working on it) but it needs a large refactor I'm not sure I have time for. I don't feel weird about the `get_plot()` method here. To comment further I'll have to test the PR on my own machine, but I think it looks good.",
"Added docstrings and tests. I believe this is ready to look over before pulling in? @mkhorton \r\n\r\nBtw -- I added support for binary (2D) diagrams!",
"Looks fantastic @mattmcdermott, thank you!"
] | 2021-08-16T21:14:02
| 2021-09-01T00:33:12
|
2021-09-01T00:33:12Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Added new module with class,`ChemicalPotentialDiagram`, for creating and plotting **chemical potential diagrams** for a list of entries with 2 or more elements. Chemical potential diagrams are like normal predominance diagrams (which are already implemented in pymatgen), but they also show the "extra" chemical potential axis -- e.g., a 3D plot for a ternary system.
* Includes ability to plot slices of domains of phases which contain other elements (see reference paper)
This code uses the scipy `HalfspaceIntersection` approach to solve for all domain vertices. For more information, please reference the paper below:
_Todd, Paul K., McDermott, M.J., et al. “Selectivity in yttrium manganese oxide
synthesis via local chemical potentials in hyperdimensional phase space.”
ArXiv:2104.05986 [Cond-Mat], Apr. 2021. arXiv.org, http://arxiv.org/abs/2104.05986._
## TODO (if any)
* Add tests
* Complete remaining docstrings
## Checklist
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/)
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2218/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2218/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2218",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2218",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2218.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2218.patch",
"merged_at": "2021-09-01T00:33:12Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2219
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2219/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2219/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2219/events
|
https://github.com/materialsproject/pymatgen/issues/2219
| 972,141,270
|
MDU6SXNzdWU5NzIxNDEyNzA=
| 2,219
|
Entry ID does not display on hovertext for plots of `CompoundPhaseDiagram` and `GrandPotentialPhaseDiagram`
|
{
"login": "mattmcdermott",
"id": 40469694,
"node_id": "MDQ6VXNlcjQwNDY5Njk0",
"avatar_url": "https://avatars.githubusercontent.com/u/40469694?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mattmcdermott",
"html_url": "https://github.com/mattmcdermott",
"followers_url": "https://api.github.com/users/mattmcdermott/followers",
"following_url": "https://api.github.com/users/mattmcdermott/following{/other_user}",
"gists_url": "https://api.github.com/users/mattmcdermott/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mattmcdermott/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mattmcdermott/subscriptions",
"organizations_url": "https://api.github.com/users/mattmcdermott/orgs",
"repos_url": "https://api.github.com/users/mattmcdermott/repos",
"events_url": "https://api.github.com/users/mattmcdermott/events{/privacy}",
"received_events_url": "https://api.github.com/users/mattmcdermott/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-08-16T22:17:03
| 2021-08-17T00:38:48
|
2021-08-17T00:38:48Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
A phase's Entry ID does not display on the hovertext for Plotly-based plots of `CompoundPhaseDiagram` objects and `GrandPotentialPhaseDiagram` objects.
This has been a known issue and I will address it in a new PR.
Steps to reproduce the behavior:
1. Acquire any set of entry objects which contain an entry ID (e.g., mp-19835, mp-25025, etc.)
2. Create a `GrandPotentialPhaseDiagram` or `CompoundPhaseDiagram`
3. Plot it using `PDPlotter` and observe bug, as below:
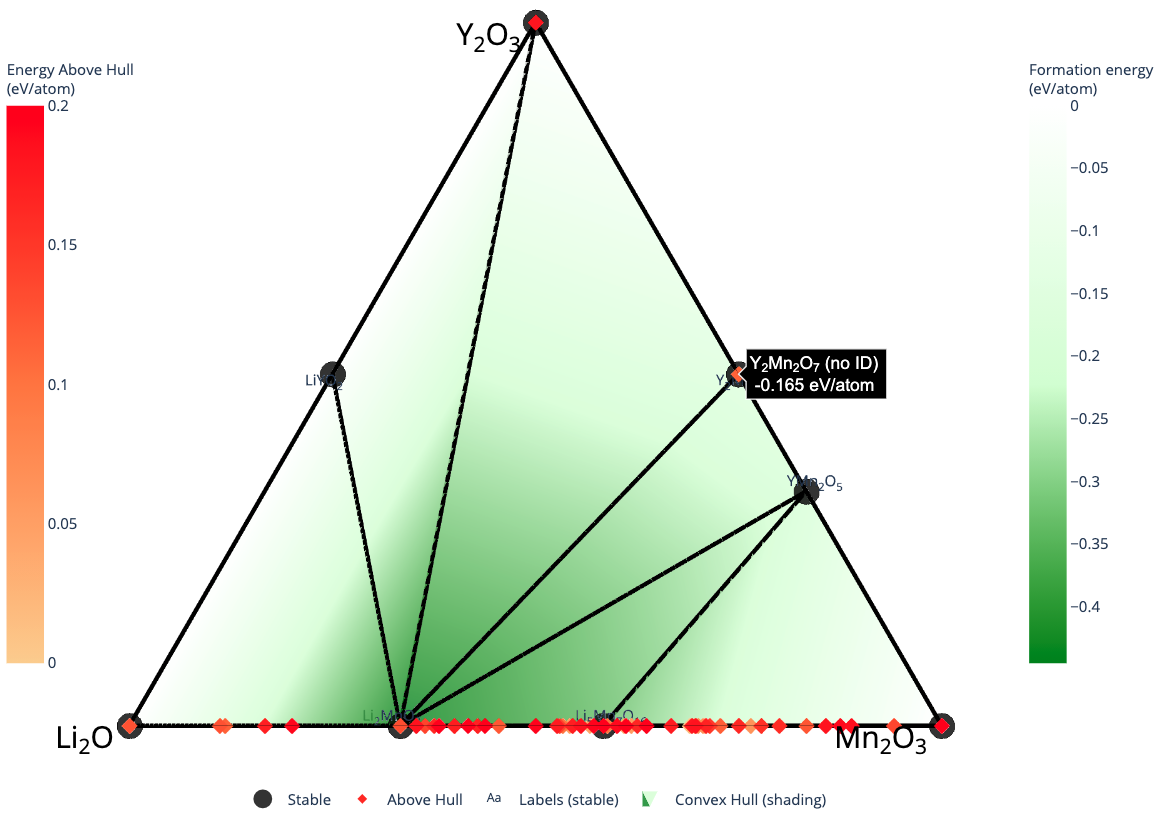
**Desktop:**
- MacOS 11.4
- Version 2022.0.11
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2219/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2219/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2220
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2220/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2220/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2220/events
|
https://github.com/materialsproject/pymatgen/pull/2220
| 972,156,908
|
MDExOlB1bGxSZXF1ZXN0NzEzNzk0MTgy
| 2,220
|
Fix entry_id not displaying on hovertext for plots of CompoundPhaseDiagram and GrandPotentialPhaseDiagram
|
{
"login": "mattmcdermott",
"id": 40469694,
"node_id": "MDQ6VXNlcjQwNDY5Njk0",
"avatar_url": "https://avatars.githubusercontent.com/u/40469694?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mattmcdermott",
"html_url": "https://github.com/mattmcdermott",
"followers_url": "https://api.github.com/users/mattmcdermott/followers",
"following_url": "https://api.github.com/users/mattmcdermott/following{/other_user}",
"gists_url": "https://api.github.com/users/mattmcdermott/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mattmcdermott/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mattmcdermott/subscriptions",
"organizations_url": "https://api.github.com/users/mattmcdermott/orgs",
"repos_url": "https://api.github.com/users/mattmcdermott/repos",
"events_url": "https://api.github.com/users/mattmcdermott/events{/privacy}",
"received_events_url": "https://api.github.com/users/mattmcdermott/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/42187230)\n\nCoverage decreased (-0.6%) to 83.132% when pulling **234a4c1ca19797090ce11b235d62e14eceecf3ef on mattmcdermott:grand_pd_bug_fix** into **0f357e704c7c1a7ff98edba8c21c537114c61b29 on materialsproject:master**.\n",
"Thanks Matt!"
] | 2021-08-16T22:53:39
| 2021-08-17T00:38:16
|
2021-08-17T00:38:16Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Bug fix for entry_ID phase diagram plotting bug described in this Issue: #2219
## Checklist
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2220/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2220/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2220",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2220",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2220.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2220.patch",
"merged_at": "2021-08-17T00:38:16Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2221
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2221/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2221/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2221/events
|
https://github.com/materialsproject/pymatgen/issues/2221
| 973,913,814
|
MDU6SXNzdWU5NzM5MTM4MTQ=
| 2,221
|
Feature request: Interactive plotly viewer for interface reactions
|
{
"login": "mattmcdermott",
"id": 40469694,
"node_id": "MDQ6VXNlcjQwNDY5Njk0",
"avatar_url": "https://avatars.githubusercontent.com/u/40469694?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mattmcdermott",
"html_url": "https://github.com/mattmcdermott",
"followers_url": "https://api.github.com/users/mattmcdermott/followers",
"following_url": "https://api.github.com/users/mattmcdermott/following{/other_user}",
"gists_url": "https://api.github.com/users/mattmcdermott/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mattmcdermott/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mattmcdermott/subscriptions",
"organizations_url": "https://api.github.com/users/mattmcdermott/orgs",
"repos_url": "https://api.github.com/users/mattmcdermott/repos",
"events_url": "https://api.github.com/users/mattmcdermott/events{/privacy}",
"received_events_url": "https://api.github.com/users/mattmcdermott/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I plan to address this feature request in a new PR in the next week(s). \r\n\r\nThe Plotly viewer can also be used on the new MP website.",
"Closing this as this feature was added in #2233 "
] | 2021-08-18T17:46:39
| 2021-11-16T00:27:07
|
2021-11-16T00:27:07Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
The current `InterfacialReactivity` module in `pymatgen.analysis.interface_reactions` is not user-friendly. To view the predicted reactions between two compositions, a user can call `InterfacialReactivity.plot()`, but they are unable to clearly tell which reactions are predicted, as there are no labels on the plot.
To get the reactions themselves, a user has to call `InterfacialReactivity.get_kinks()` and dissect a _zip_ object that has an arbitrary order. This means every user of the class must essentially write their own custom API to use it.
At the very least, we should implement a Plotly-based plotter for the predicted reactions so that a user can easily see which reactions are predicted. If possible, however, we should refactor the module such that reactions can more easily be obtained.
Previously, I have written my own API for the class, where it is possible to get`ComputedReaction` objects by calling `react_interface()`. (Currently, the class only returns basic `Reaction` objects.)
https://github.com/GENESIS-EFRC/reaction-network/blob/main/src/rxn_network/enumerators/minimize.py#L127:L172
|
{
"login": "mattmcdermott",
"id": 40469694,
"node_id": "MDQ6VXNlcjQwNDY5Njk0",
"avatar_url": "https://avatars.githubusercontent.com/u/40469694?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mattmcdermott",
"html_url": "https://github.com/mattmcdermott",
"followers_url": "https://api.github.com/users/mattmcdermott/followers",
"following_url": "https://api.github.com/users/mattmcdermott/following{/other_user}",
"gists_url": "https://api.github.com/users/mattmcdermott/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mattmcdermott/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mattmcdermott/subscriptions",
"organizations_url": "https://api.github.com/users/mattmcdermott/orgs",
"repos_url": "https://api.github.com/users/mattmcdermott/repos",
"events_url": "https://api.github.com/users/mattmcdermott/events{/privacy}",
"received_events_url": "https://api.github.com/users/mattmcdermott/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2221/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2221/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2222
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2222/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2222/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2222/events
|
https://github.com/materialsproject/pymatgen/issues/2222
| 975,948,977
|
MDU6SXNzdWU5NzU5NDg5Nzc=
| 2,222
|
Small bug of pymatgen.apps.borg.hive
|
{
"login": "Jeff-oakley",
"id": 5863275,
"node_id": "MDQ6VXNlcjU4NjMyNzU=",
"avatar_url": "https://avatars.githubusercontent.com/u/5863275?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Jeff-oakley",
"html_url": "https://github.com/Jeff-oakley",
"followers_url": "https://api.github.com/users/Jeff-oakley/followers",
"following_url": "https://api.github.com/users/Jeff-oakley/following{/other_user}",
"gists_url": "https://api.github.com/users/Jeff-oakley/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Jeff-oakley/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Jeff-oakley/subscriptions",
"organizations_url": "https://api.github.com/users/Jeff-oakley/orgs",
"repos_url": "https://api.github.com/users/Jeff-oakley/repos",
"events_url": "https://api.github.com/users/Jeff-oakley/events{/privacy}",
"received_events_url": "https://api.github.com/users/Jeff-oakley/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi Bin, just a note that at Materials Project we mostly now use the drone in [`atomate`](https://github.com/hackingmaterials/atomate/blob/main/atomate/vasp/drones.py) rather than the simple drone here. This is not directly relevant to your bug report but thought I would let you know in case you were looking for this functionality.",
"@mkhorton As I have said several times, the drone in atomate should be moved to pymatgen. It is unclear to me why the concept should be ported to atomate and there is a refusal to move it back to pymatgen proper so that people do not have to do it using atomate. ",
"> Hi Bin, just a note that at Materials Project we mostly now use the drone in [`atomate`](https://github.com/hackingmaterials/atomate/blob/main/atomate/vasp/drones.py) rather than the simple drone here. This is not directly relevant to your bug report but thought I would let you know in case you were looking for this functionality.\r\n\r\nGot it. Thanks for the note."
] | 2021-08-20T22:04:25
| 2021-08-24T15:34:24
|
2021-08-24T15:34:24Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
At the assimilate method of SimpleVaspToComputedEntryDrone, it checks for existence of multiple INCAR POTCAR and DYNMAT with the following commands:
if len(files) == 1 or filename in ("INCAR", "POTCAR", "DYNMAT"):
files_to_parse[filename] = files[0]
If there is no DYNMAT file in the calculation directory, this will raise up an error as files list is empty so the indexing can not be done.
**To Reproduce**
Steps to reproduce the behavior:
This can be easily reproduce if you start a DFT calculation directory that does not have DYNMAT file
**Expected behavior**
It should continue even without DYNMAT file
**Screenshots**
N/A
**Desktop (please complete the following information):**
All platforms
**Additional context**
Can be easily fixed. Such as using try except instead and continue even without DYNMAT file
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2222/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2222/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2223
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2223/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2223/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2223/events
|
https://github.com/materialsproject/pymatgen/issues/2223
| 976,915,784
|
MDU6SXNzdWU5NzY5MTU3ODQ=
| 2,223
|
Inconsistent value from "composition.__len__()"
|
{
"login": "frankNiessen",
"id": 41304940,
"node_id": "MDQ6VXNlcjQxMzA0OTQw",
"avatar_url": "https://avatars.githubusercontent.com/u/41304940?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/frankNiessen",
"html_url": "https://github.com/frankNiessen",
"followers_url": "https://api.github.com/users/frankNiessen/followers",
"following_url": "https://api.github.com/users/frankNiessen/following{/other_user}",
"gists_url": "https://api.github.com/users/frankNiessen/gists{/gist_id}",
"starred_url": "https://api.github.com/users/frankNiessen/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/frankNiessen/subscriptions",
"organizations_url": "https://api.github.com/users/frankNiessen/orgs",
"repos_url": "https://api.github.com/users/frankNiessen/repos",
"events_url": "https://api.github.com/users/frankNiessen/events{/privacy}",
"received_events_url": "https://api.github.com/users/frankNiessen/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"My bad, I have been looping over a pandas dataframe of compositions when talking about multiple compositions, so the iteration over the Composition class should by default always go over the elements."
] | 2021-08-23T11:12:56
| 2021-08-23T11:17:49
|
2021-08-23T11:16:27Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hi,
I would like to point out a problem concerning the inconsistent return value of the __len__() method of the Composition class.
When an object contains one composition, the __len__() command returns the number of elements, whereas an object with multiple compositions returns the number of compositions. This is problematic when using the zip() function to loop over compositions - in the case where the composition object only contains one composition it loops over the elements instead ...
I am not sure if this behavior is desired, if so somebody might be able to suggest an approach to me that works both for single and multiple composition sets. Otherwise I would just like to report it as a bug.
Best wishes
Frank
|
{
"login": "frankNiessen",
"id": 41304940,
"node_id": "MDQ6VXNlcjQxMzA0OTQw",
"avatar_url": "https://avatars.githubusercontent.com/u/41304940?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/frankNiessen",
"html_url": "https://github.com/frankNiessen",
"followers_url": "https://api.github.com/users/frankNiessen/followers",
"following_url": "https://api.github.com/users/frankNiessen/following{/other_user}",
"gists_url": "https://api.github.com/users/frankNiessen/gists{/gist_id}",
"starred_url": "https://api.github.com/users/frankNiessen/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/frankNiessen/subscriptions",
"organizations_url": "https://api.github.com/users/frankNiessen/orgs",
"repos_url": "https://api.github.com/users/frankNiessen/repos",
"events_url": "https://api.github.com/users/frankNiessen/events{/privacy}",
"received_events_url": "https://api.github.com/users/frankNiessen/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2223/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2223/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2224
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2224/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2224/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2224/events
|
https://github.com/materialsproject/pymatgen/pull/2224
| 977,556,706
|
MDExOlB1bGxSZXF1ZXN0NzE4Mjg1OTA1
| 2,224
|
Configured test files dir
|
{
"login": "drew-parsons",
"id": 26508288,
"node_id": "MDQ6VXNlcjI2NTA4Mjg4",
"avatar_url": "https://avatars.githubusercontent.com/u/26508288?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/drew-parsons",
"html_url": "https://github.com/drew-parsons",
"followers_url": "https://api.github.com/users/drew-parsons/followers",
"following_url": "https://api.github.com/users/drew-parsons/following{/other_user}",
"gists_url": "https://api.github.com/users/drew-parsons/gists{/gist_id}",
"starred_url": "https://api.github.com/users/drew-parsons/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/drew-parsons/subscriptions",
"organizations_url": "https://api.github.com/users/drew-parsons/orgs",
"repos_url": "https://api.github.com/users/drew-parsons/repos",
"events_url": "https://api.github.com/users/drew-parsons/events{/privacy}",
"received_events_url": "https://api.github.com/users/drew-parsons/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/42354378)\n\nCoverage decreased (-0.6%) to 83.136% when pulling **86f8981d6b1e38ed83c4dee5060a6536bef2faa6 on drew-parsons:configured_test_files_dir** into **e002969c12ca7e06cacb6984503d69114cb442ef on materialsproject:master**.\n"
] | 2021-08-23T23:15:49
| 2021-08-24T15:36:31
|
2021-08-23T23:39:56Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* use PymatgenTest.TEST_FILES_DIR with all tests
Required to enable tests to run separately from the source, for instance for CI testing of packaging for Linux distributions,
see https://github.com/materialsproject/pymatgen/issues/2025
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). Black is applied.
- [x] All linting and tests pass (testing against release v2022.0.11
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2224/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2224/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2224",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2224",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2224.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2224.patch",
"merged_at": "2021-08-23T23:39:55Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2225
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2225/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2225/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2225/events
|
https://github.com/materialsproject/pymatgen/pull/2225
| 977,560,625
|
MDExOlB1bGxSZXF1ZXN0NzE4Mjg5MzAx
| 2,225
|
update list of data files required for installation
|
{
"login": "drew-parsons",
"id": 26508288,
"node_id": "MDQ6VXNlcjI2NTA4Mjg4",
"avatar_url": "https://avatars.githubusercontent.com/u/26508288?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/drew-parsons",
"html_url": "https://github.com/drew-parsons",
"followers_url": "https://api.github.com/users/drew-parsons/followers",
"following_url": "https://api.github.com/users/drew-parsons/following{/other_user}",
"gists_url": "https://api.github.com/users/drew-parsons/gists{/gist_id}",
"starred_url": "https://api.github.com/users/drew-parsons/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/drew-parsons/subscriptions",
"organizations_url": "https://api.github.com/users/drew-parsons/orgs",
"repos_url": "https://api.github.com/users/drew-parsons/repos",
"events_url": "https://api.github.com/users/drew-parsons/events{/privacy}",
"received_events_url": "https://api.github.com/users/drew-parsons/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/42363078)\n\nCoverage decreased (-0.6%) to 83.134% when pulling **fd9004ddd600c6141a22aae485162720cbc4477e on drew-parsons:install_data_files** into **405a0b22c506077732ed0cabee82d2503ae9d0ce on materialsproject:master**.\n",
"Thanks!"
] | 2021-08-23T23:25:13
| 2021-08-25T16:52:11
|
2021-08-24T15:29:24Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* update list of data files to be installed
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). Checked with black, pycodestyle, flake8.
- [x] All linting and tests pass. (tested against release v2022.0.11)
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2225/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2225/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2225",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2225",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2225.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2225.patch",
"merged_at": "2021-08-24T15:29:24Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2226
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2226/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2226/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2226/events
|
https://github.com/materialsproject/pymatgen/issues/2226
| 978,572,671
|
MDU6SXNzdWU5Nzg1NzI2NzE=
| 2,226
|
A bug in get_neighbors function?
|
{
"login": "qzhu2017",
"id": 29445366,
"node_id": "MDQ6VXNlcjI5NDQ1MzY2",
"avatar_url": "https://avatars.githubusercontent.com/u/29445366?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/qzhu2017",
"html_url": "https://github.com/qzhu2017",
"followers_url": "https://api.github.com/users/qzhu2017/followers",
"following_url": "https://api.github.com/users/qzhu2017/following{/other_user}",
"gists_url": "https://api.github.com/users/qzhu2017/gists{/gist_id}",
"starred_url": "https://api.github.com/users/qzhu2017/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/qzhu2017/subscriptions",
"organizations_url": "https://api.github.com/users/qzhu2017/orgs",
"repos_url": "https://api.github.com/users/qzhu2017/repos",
"events_url": "https://api.github.com/users/qzhu2017/events{/privacy}",
"received_events_url": "https://api.github.com/users/qzhu2017/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@chc273 @shyuep I have tested the new fix. It worked."
] | 2021-08-25T00:00:00
| 2021-08-25T05:03:58
|
2021-08-25T05:03:58Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
I am trying to use the get_neighbors function to process some structures. However, it failed in some cases
**To Reproduce**
```Python
from pymatgen.core import Structure
poscar = '''POSCAR
1.0000000000000000
6.9082208665474800 0.0000000000000005 0.0000000000000011
-0.0000000000000008 14.1745610988433715 0.0000000000000004
0.0000000000000005 -0.0000000000000019 20.0189293405157045
C
14
Direct
1.2615805559557376 1.4846778919841908 0.3162565598606580
-0.7615805559557360 -0.9846778919841882 -0.1837434401393425
1.2615805559557380 -0.4846778919841886 -0.1837434401393425
-0.7615805559557358 0.9846778919841882 0.3162565598606581
-0.2615805559557363 -0.4846778919841892 0.6837434401393432
0.7615805559557360 0.9846778919841881 0.1837434401393430
-0.2653510291469959 0.5275483828607898 0.1211255106369795
0.7653510291469972 -0.0275483828607906 0.6211255106369804
1.2653510291469956 0.5275483828607898 0.3788744893630209
-0.7653510291469972 -0.0275483828607906 -0.1211255106369797
1.2653510291469956 0.4724516171392097 -0.1211255106369793
-0.7653510291469972 0.0275483828607905 0.3788744893630209
-0.2653510291469959 0.4724516171392097 0.6211255106369801
-0.2705230397846415 1.4621722452479102 0.0625618775773844
'''
s=Structure.from_str(poscar, fmt='poscar')
pairs = [(0, 8), (1, 9), (2, 10), (3, 11), (4, 12), (6, 13)]
for pair in pairs:
print('\n Pair', pair)
(id1, id2) = pair
site0=s.sites[id1]
neigh_sites = s.get_neighbors(site0, 2.0)
print("get_neighbors", id1, site0.specie, site0.frac_coords, len(neigh_sites))
site1=s.sites[id2]
neigh_sites = s.get_neighbors(site1, 2.0)
print("get_neighbors", id2, site1.specie, site1.frac_coords, len(neigh_sites))
print("distance_and_image", site0.distance_and_image(site1))
print("distance_and_image", site1.distance_and_image(site0))
```
Executing the above script will generate the following output.
```
Pair (0, 8)
get_neighbors 0 C [1.26158056 1.48467789 0.31625656] 1
get_neighbors 8 C [1.26535103 0.52754838 0.37887449] 1
distance_and_image (1.3933104787298265, array([0, 1, 0]))
distance_and_image (1.3933104787298265, array([ 0, -1, 0]))
Pair (1, 9)
get_neighbors 1 C [-0.76158056 -0.98467789 -0.18374344] 1
get_neighbors 9 C [-0.76535103 -0.02754838 -0.12112551] 0
distance_and_image (1.3933104787298447, array([ 0, -1, 0]))
distance_and_image (1.3933104787298447, array([0, 1, 0]))
Pair (2, 10)
get_neighbors 2 C [ 1.26158056 -0.48467789 -0.18374344] 1
get_neighbors 10 C [ 1.26535103 0.47245162 -0.12112551] 1
distance_and_image (1.3933104787298471, array([ 0, -1, 0]))
distance_and_image (1.3933104787298471, array([0, 1, 0]))
Pair (3, 11)
get_neighbors 3 C [-0.76158056 0.98467789 0.31625656] 1
get_neighbors 11 C [-0.76535103 0.02754838 0.37887449] 1
distance_and_image (1.3933104787298447, array([0, 1, 0]))
distance_and_image (1.3933104787298447, array([ 0, -1, 0]))
Pair (4, 12)
get_neighbors 4 C [-0.26158056 -0.48467789 0.68374344] 1
get_neighbors 12 C [-0.26535103 0.47245162 0.62112551] 1
distance_and_image (1.3933104787298416, array([ 0, -1, 0]))
distance_and_image (1.3933104787298416, array([0, 1, 0]))
Pair (6, 13)
get_neighbors 6 C [-0.26535103 0.52754838 0.12112551] 1
get_neighbors 13 C [-0.27052304 1.46217225 0.06256188] 1
distance_and_image (1.4948199103062294, array([ 0, -1, 0]))
distance_and_image (1.4948199103062294, array([0, 1, 0]))
```
**Expected behavior**
The above script tried to search for the neighboring atoms for each site. In total, there should exist 6 atomic pairs, i.e.,
`(0, 8), (1, 9), (2, 10), (3, 11), (4, 12), (6, 13)`.
It works fine for most of the atoms except atom 9, which cannot find it neighbor atom 1.
**Desktop (please complete the following information):**
- OS: [Mac]
- Version [2022.0.9]
Can anyone help on this issue?
|
{
"login": "qzhu2017",
"id": 29445366,
"node_id": "MDQ6VXNlcjI5NDQ1MzY2",
"avatar_url": "https://avatars.githubusercontent.com/u/29445366?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/qzhu2017",
"html_url": "https://github.com/qzhu2017",
"followers_url": "https://api.github.com/users/qzhu2017/followers",
"following_url": "https://api.github.com/users/qzhu2017/following{/other_user}",
"gists_url": "https://api.github.com/users/qzhu2017/gists{/gist_id}",
"starred_url": "https://api.github.com/users/qzhu2017/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/qzhu2017/subscriptions",
"organizations_url": "https://api.github.com/users/qzhu2017/orgs",
"repos_url": "https://api.github.com/users/qzhu2017/repos",
"events_url": "https://api.github.com/users/qzhu2017/events{/privacy}",
"received_events_url": "https://api.github.com/users/qzhu2017/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2226/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2226/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2227
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2227/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2227/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2227/events
|
https://github.com/materialsproject/pymatgen/pull/2227
| 978,660,013
|
MDExOlB1bGxSZXF1ZXN0NzE5MjEyODgy
| 2,227
|
Fix c mod in neighobrs
|
{
"login": "chc273",
"id": 22353204,
"node_id": "MDQ6VXNlcjIyMzUzMjA0",
"avatar_url": "https://avatars.githubusercontent.com/u/22353204?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/chc273",
"html_url": "https://github.com/chc273",
"followers_url": "https://api.github.com/users/chc273/followers",
"following_url": "https://api.github.com/users/chc273/following{/other_user}",
"gists_url": "https://api.github.com/users/chc273/gists{/gist_id}",
"starred_url": "https://api.github.com/users/chc273/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/chc273/subscriptions",
"organizations_url": "https://api.github.com/users/chc273/orgs",
"repos_url": "https://api.github.com/users/chc273/repos",
"events_url": "https://api.github.com/users/chc273/events{/privacy}",
"received_events_url": "https://api.github.com/users/chc273/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/42390860)\n\nCoverage decreased (-0.6%) to 83.132% when pulling **c42bee17c8ee8cbd5d8fcbea0d96455d29563504 on chc273:fix_c_mod_in_neighobrs** into **40817cc97fe25bc027523a65aab528b59e22e78e on materialsproject:master**.\n"
] | 2021-08-25T03:08:39
| 2021-08-25T03:31:53
|
2021-08-25T03:30:08Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
This PR fixes bug reported by #2226
In the previous implementation, I have turned on the c behavior for division in cython. In this case `-0.1 % 1 = -0.1` while I expected the result to be 0.9 as in python. This causes the atoms out of the unit cell in the negative direction not properly wrapped into the unit cell. I have turned off the cdivision tag in cython. Now it works as expected for the issue #2226.
This problem was not captured in previous tests. Hence I have also added a new test for this case.
@qzhu2017 please see if this PR fixes the issues. Thanks!
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2227/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2227/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2227",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2227",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2227.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2227.patch",
"merged_at": "2021-08-25T03:30:08Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2228
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2228/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2228/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2228/events
|
https://github.com/materialsproject/pymatgen/pull/2228
| 980,116,570
|
MDExOlB1bGxSZXF1ZXN0NzIwNDEyOTI1
| 2,228
|
Changes due to new lobster version and changes to lobsterenv
|
{
"login": "JaGeo",
"id": 22094846,
"node_id": "MDQ6VXNlcjIyMDk0ODQ2",
"avatar_url": "https://avatars.githubusercontent.com/u/22094846?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/JaGeo",
"html_url": "https://github.com/JaGeo",
"followers_url": "https://api.github.com/users/JaGeo/followers",
"following_url": "https://api.github.com/users/JaGeo/following{/other_user}",
"gists_url": "https://api.github.com/users/JaGeo/gists{/gist_id}",
"starred_url": "https://api.github.com/users/JaGeo/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/JaGeo/subscriptions",
"organizations_url": "https://api.github.com/users/JaGeo/orgs",
"repos_url": "https://api.github.com/users/JaGeo/repos",
"events_url": "https://api.github.com/users/JaGeo/events{/privacy}",
"received_events_url": "https://api.github.com/users/JaGeo/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I am working on it further. I will let you know once it is finished ... Found another inconsistency and had to make a few changes.\r\n\r\nJG",
"\n[](https://coveralls.io/builds/42448769)\n\nCoverage decreased (-0.7%) to 83.122% when pulling **f786a63458d6e17fc5a9774cb9f0249b07644304 on JaGeo:master** into **a68137eb74c1453642c4067e3f0909df634c4de0 on materialsproject:master**.\n",
"Linting is passing now. Tests should be passing as well. \r\n\r\nThis was faster than expected.\r\n\r\nHappy to hear suggestions/feedback.\r\n\r\nBest, \r\nJG",
"Hi again, \r\nI just saw that there is a problem with file permissions.. Any tip how to resolve the issue? ",
"Thanks @JaGeo ! Happy to merge the new features but seems like something iffy with the PR, says 2864 files changed?",
"Sorry our comments crossed, I'm not sure I see the permissions error, I was just looking at the \"Files changed\" tab.",
"Yes, .. I think so many files \"changed\" because of a change in permission. For some reason all files are executable and I cannot revert it back at the moment. \r\n\r\nI will try to clone it somewhere else and try again . Not sure where this is coming from. Should have probably changed the git config in advance.... ",
"Ah, solved most of it. Sorry for this. There was some weird permission problem on my hard drive. :) Now, only 96 files changed. \r\n\r\nI will take care of the last permission problems.",
"Now the permissions are fixed. Sorry again. @mkhorton .",
"No worries at all, I'll just wait for the CI to pass and then we can merge.",
"Thank you "
] | 2021-08-26T11:06:05
| 2021-08-26T21:11:24
|
2021-08-26T21:02:25Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Hi all,
I have now fixed some problems due to the new Lobster versions. The Icohplist class can now at least handle orbitalwise ICOHPLIST.lobster files -it will ignore the orbitalwise information for now as it will probably take several days of work to think about a new data format for this additional information.
COBICARs can now be handled and plotted. SitePotentials and MadelungEnergies files as well. Also, I made changes to the class for lobsterout since there are several new potential outputs. Lobsterin classes have also be adapted to put out warnings for current lobster bugs and to handle the new input keywords.
And, I made changes to lobsterenv to be more flexible.
I hope that the tests will pass. I will try to fix them as soon as possible if not.
Best,
JG
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2228/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2228/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2228",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2228",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2228.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2228.patch",
"merged_at": "2021-08-26T21:02:25Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2230
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2230/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2230/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2230/events
|
https://github.com/materialsproject/pymatgen/pull/2230
| 984,061,702
|
MDExOlB1bGxSZXF1ZXN0NzIzNTQyNTYx
| 2,230
|
Update bader_caller.py
|
{
"login": "nwinner",
"id": 8825901,
"node_id": "MDQ6VXNlcjg4MjU5MDE=",
"avatar_url": "https://avatars.githubusercontent.com/u/8825901?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/nwinner",
"html_url": "https://github.com/nwinner",
"followers_url": "https://api.github.com/users/nwinner/followers",
"following_url": "https://api.github.com/users/nwinner/following{/other_user}",
"gists_url": "https://api.github.com/users/nwinner/gists{/gist_id}",
"starred_url": "https://api.github.com/users/nwinner/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/nwinner/subscriptions",
"organizations_url": "https://api.github.com/users/nwinner/orgs",
"repos_url": "https://api.github.com/users/nwinner/repos",
"events_url": "https://api.github.com/users/nwinner/events{/privacy}",
"received_events_url": "https://api.github.com/users/nwinner/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @nwinner",
"\n[](https://coveralls.io/builds/42550434)\n\nCoverage decreased (-0.6%) to 83.125% when pulling **aea737ffc405750d5a264fffb78edca1260d259b on nwinner:patch-1** into **48b031a44494b6452b9ca14619448e450bda7af9 on materialsproject:master**.\n"
] | 2021-08-31T16:35:14
| 2021-09-01T01:10:46
|
2021-09-01T00:32:11Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Patch. The chgref option to the bader caller should be applicable to parsing cube files as well.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2230/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2230/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2230",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2230",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2230.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2230.patch",
"merged_at": "2021-09-01T00:32:11Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2231
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2231/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2231/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2231/events
|
https://github.com/materialsproject/pymatgen/pull/2231
| 984,079,361
|
MDExOlB1bGxSZXF1ZXN0NzIzNTU3ODc4
| 2,231
|
indent list for `Composition.contains_element_type` documentation
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @CompRhys !",
"\n[](https://coveralls.io/builds/42551120)\n\nCoverage decreased (-0.6%) to 83.125% when pulling **a88d4069e70048ef248ba91b013e0077687ce236 on CompRhys:master** into **48b031a44494b6452b9ca14619448e450bda7af9 on materialsproject:master**.\n"
] | 2021-08-31T16:55:03
| 2021-09-01T01:15:08
|
2021-09-01T00:31:01Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
should fix a bug in the automatic documentation markup for `Composition.contains_element_type`
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2231/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2231/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2231",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2231",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2231.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2231.patch",
"merged_at": "2021-09-01T00:31:01Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2232
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2232/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2232/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2232/events
|
https://github.com/materialsproject/pymatgen/pull/2232
| 984,923,219
|
MDExOlB1bGxSZXF1ZXN0NzI0MzAwOTky
| 2,232
|
Add `pymatgen-io-fleur` addon to addons page
|
{
"login": "janssenhenning",
"id": 11249470,
"node_id": "MDQ6VXNlcjExMjQ5NDcw",
"avatar_url": "https://avatars.githubusercontent.com/u/11249470?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janssenhenning",
"html_url": "https://github.com/janssenhenning",
"followers_url": "https://api.github.com/users/janssenhenning/followers",
"following_url": "https://api.github.com/users/janssenhenning/following{/other_user}",
"gists_url": "https://api.github.com/users/janssenhenning/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janssenhenning/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janssenhenning/subscriptions",
"organizations_url": "https://api.github.com/users/janssenhenning/orgs",
"repos_url": "https://api.github.com/users/janssenhenning/repos",
"events_url": "https://api.github.com/users/janssenhenning/events{/privacy}",
"received_events_url": "https://api.github.com/users/janssenhenning/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Is there something I forgot to do for this PR to be checked? I notice that the workflows were never run here",
"Thanks. The reason why the CI did not run is because of new security features from Github. They need approval from a maintainer for first time contributors.\r\n\r\nThis looks good. Since it is an add-on, there aren't really any tests per se. So I will merge it once the linting is done. "
] | 2021-09-01T09:27:54
| 2021-09-24T15:27:05
|
2021-09-24T15:27:05Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
This add-on (www.pypi.org/project/pymatgen-io-fleur) provides functionality for reading writing files used
by the FLEUR code (www.flapw.de)
## Summary
Include a summary of major changes in bullet points:
* Added entry for IO namespace package for the fleur code to the addons page
* Added the IO functionality of the addon to the ``Structure`` methods ``to``, ``from_file`` and ``from_str``
* Adds a format for the XML input file of Fleur (read-only), which is chosen for files named ``inp*.xml``
* Adds a format for the input file for the input generator of Fleur which can also be written out
## Additional dependencies introduced (if any)
None
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
> No new functions or methods added to the main package
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
> Tests for the ``Structure`` methods with the new fleur formats will be added to the ``pymatgen-io-fleur`` addon
- [x] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply cp pre-commit .git/hooks and a check will be run prior to allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2232/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2232/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2232",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2232",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2232.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2232.patch",
"merged_at": "2021-09-24T15:27:05Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2233
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2233/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2233/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2233/events
|
https://github.com/materialsproject/pymatgen/pull/2233
| 987,213,079
|
MDExOlB1bGxSZXF1ZXN0NzI2MzMwNzg2
| 2,233
|
Refactor `interface_reactions` module, add support for Plotly
|
{
"login": "mattmcdermott",
"id": 40469694,
"node_id": "MDQ6VXNlcjQwNDY5Njk0",
"avatar_url": "https://avatars.githubusercontent.com/u/40469694?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mattmcdermott",
"html_url": "https://github.com/mattmcdermott",
"followers_url": "https://api.github.com/users/mattmcdermott/followers",
"following_url": "https://api.github.com/users/mattmcdermott/following{/other_user}",
"gists_url": "https://api.github.com/users/mattmcdermott/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mattmcdermott/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mattmcdermott/subscriptions",
"organizations_url": "https://api.github.com/users/mattmcdermott/orgs",
"repos_url": "https://api.github.com/users/mattmcdermott/repos",
"events_url": "https://api.github.com/users/mattmcdermott/events{/privacy}",
"received_events_url": "https://api.github.com/users/mattmcdermott/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@mkhorton My plan is to have this PR done by Wed, September 8, based on the previous timeline goal we set",
"Fantastic, thank you for the update @mattmcdermott !",
"@mkhorton Should be pretty much done. Seems like linting and tests are failing for recent changes related to Optimade.\r\n\r\nNote: you'll see I changed something with `PDPlotter` -- it was an old method I had written called `htmlize_formula()`. Turns out that method already existed in `pymatgen.util.string`, so I removed it from `PDPlotter` and replaced where it was used.",
"\n[](https://coveralls.io/builds/42714836)\n\nCoverage decreased (-0.6%) to 83.153% when pulling **84925584f663e7ece47bdcdec786c6b1db00c52e on mattmcdermott:interface_rxns** into **2c6ac61a557782efee3bceaa6ac39f7aab02eb8e on materialsproject:master**.\n",
"Thanks @mattmcdermott! I think that method is also available on `Composition` etc. directly (via the `Stringify` mixin), just an fyi.",
"> Thanks @mattmcdermott! I think that method is also available on `Composition` etc. directly (via the `Stringify` mixin), just an fyi.\r\n\r\nAh true. The only downside is for compositions like `Composition(\"FeO2\")`, it automatically puts \"1\" as a subscript:\r\ne.g.,`Fe_1 O_2`",
"Ah, I guess it acts on the Composition directly rather than formula, fair enough.",
"Merged. Thanks again @mattmcdermott, this is great!"
] | 2021-09-02T22:16:18
| 2021-09-08T22:23:19
|
2021-09-08T22:22:56Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Add: support for Plotly: calling `plot()` now provides a `backend` option for choosing matplotlib or Plotly.
* Add:`GrandPotentialInterfacialReactivity` class to separate code based on the use of grand potential phase diagrams (this greatly decreases the complexity of the `__init__` args) and is a more inherited approach in general
* Add: `get_dataframe()` method for plotting possible reactions in a Pandas dataframe. Users would typically have to write/copy boilerplate code to do this each time
* Fix: Better matplotlib plotting, included annotations
* Fix: Cleaner, more readable code
## Checklist
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2233/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2233/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2233",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2233",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2233.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2233.patch",
"merged_at": "2021-09-08T22:22:56Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2234
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2234/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2234/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2234/events
|
https://github.com/materialsproject/pymatgen/issues/2234
| 987,733,105
|
MDU6SXNzdWU5ODc3MzMxMDU=
| 2,234
|
Add defined tolerances for `Lattice.__eq__`
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Honestly, even the current version of `__eq__` is not technically correct. There is no fixed reference direction in 3D space. E.g., if I rotate the basis vectors by a fixed angle, the lattices are technically still equivalent. The correct definition of `lattice.__eq__` should be that the lattice parameters (lengths and angles) are all within a certain tolerance. ",
"I don't think that can be changed however because of how the sites positions are specified in terms of Cartesian co-ordinates? the use of Cartesian co-ordinates implies that a choice of axes has been made?",
"It can. Equality of PeriodicSites is based on fractional coordinates and setting the lattice automatically changes the Cartesian coordinates. The problem areas are whether the structure matching algorithms may have issues with the redefinition. ",
"Are you going to change `__eq__` or can I add the tolerance?"
] | 2021-09-03T12:42:04
| 2021-09-16T20:55:46
|
2021-09-16T15:39:35Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
In `Lattice.__eq__` the code uses the default tolerances for np.allclose which affects the strict equality of structures. Elsewhere the code pattern in `pymatgen` appears to be to have a `cls.blah_tol` for different objects i.e. `Site.position_atol = 1e-4`. The np.allclose defaults are quite strict for my use case and think the code pattern in other classes would be a better way for users to vary the tolerances in a consistent manner rather than me just hacking the `__eq__` function directly.
```python
def __eq__(self, other):
"""
A lattice is considered to be equal to another if the internal matrix
representation satisfies np.allclose(matrix1, matrix2) to be True.
"""
if other is None:
return False
# shortcut the np.allclose if the memory addresses are the same
# (very common in Structure.from_sites)
return self is other or np.allclose(self.matrix, other.matrix)
```
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2234/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2234/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2235
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2235/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2235/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2235/events
|
https://github.com/materialsproject/pymatgen/issues/2235
| 988,025,876
|
MDU6SXNzdWU5ODgwMjU4NzY=
| 2,235
|
hash of sites is not discriminate
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Also a thought re: #2077 currently `Composition({\"Fe3+\": 2, \"O2-\":3})` and `Composition({\"Fe\": 2, \"O\":3})` have different hashes. I think this is okay but just want to check, I will make a PR to improve the docstring alongside and possible changes to `Site`'s hash",
"Differing hash methods for different objects is ok, since they're never used across different classes.\r\n\r\nThat being said, it would be good to update the site hash to be salted. All the hashes should be salted. ",
"https://github.com/materialsproject/pymatgen/blob/b8b27c6fa3236f032581aa82f668b25a78aa3b57/pymatgen/core/sites.py#L225-L230\r\n\r\nThe reason I raised it was the potential for the hash collisions we had before where `hash(Composition({\"O\": 1})) = `hash(Composition({\"N\": 0.5, \"F\": 0.5}))` this is much less likely to happen for sites but it **_could_** in principle happen. \r\n\r\nI think I can use the same hash as for `Composition` and then I will sort out the tests so that they don't fail for salted hashes. Any thoughts on the charge assigned species edge case?\r\n\r\nhttps://github.com/materialsproject/pymatgen/blob/b8b27c6fa3236f032581aa82f668b25a78aa3b57/pymatgen/core/composition.py#L236-L240",
"Hash clashes aren't necessarily a bad thing. Remember hashes are not required to be different for the code to work correctly. That is the job of `__eq__`. Good hashes merely improve performance. If you think improving `Site`'s hash will enable something you need to do, then go for it. \r\n\r\nThe salting is important for hashes since they can be a vehicle for security attacks. \r\n\r\nAs for charged vs uncharged composition, it's not an issue that the hashes are different for cases where `__eq__` also says they're different. The issue would be if there was ever a situation in which `__eq__` returns `True`, but we got different hashes. ",
"I am fine with making the Composition Hash use the Site Hash. But @shyamd is correct in that hashes are not meant to be uniquely discriminate. They are meant to be *cheap* and *sufficiently discriminate* to speed up searches. A more expensive hash that is more discriminate can be worse than a less expensive and less discriminate hash. The rule to remember is that equal objects must have equal hashes but equal hashes do not mean equal objects.",
"I will investigate a timings test then and only make the change to site's hash being the same if it's comparable or faster!\r\n\r\nedit ~ it's about 10% slower so unless salting is a big issue I think it might be best to leave it."
] | 2021-09-03T18:54:12
| 2021-09-13T16:26:59
|
2021-09-13T14:40:05Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
In #2077 we added a more discriminate hash from `Composition` individual `Site` are effectively `Composition` but they still use the old composition hash. Shall I update the `Site` hash to match composition? I haven't actually traced a slow-down to site matching like I did with compositions but was looking at `Site` code re: #2234 and I feel that if it helps there it will help here also?
only issue is that the composition hash is salted (as default python hash is salted) whilst the site hash is unsalted therefore would need to remove/change the test on the hash which asserts a value.
@shyamd
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2235/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2235/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2236
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2236/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2236/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2236/events
|
https://github.com/materialsproject/pymatgen/issues/2236
| 988,256,942
|
MDU6SXNzdWU5ODgyNTY5NDI=
| 2,236
|
🐞 Bug in parsing `bandOverlaps.lobster` file
|
{
"login": "ezpzbz",
"id": 13301031,
"node_id": "MDQ6VXNlcjEzMzAxMDMx",
"avatar_url": "https://avatars.githubusercontent.com/u/13301031?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ezpzbz",
"html_url": "https://github.com/ezpzbz",
"followers_url": "https://api.github.com/users/ezpzbz/followers",
"following_url": "https://api.github.com/users/ezpzbz/following{/other_user}",
"gists_url": "https://api.github.com/users/ezpzbz/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ezpzbz/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ezpzbz/subscriptions",
"organizations_url": "https://api.github.com/users/ezpzbz/orgs",
"repos_url": "https://api.github.com/users/ezpzbz/repos",
"events_url": "https://api.github.com/users/ezpzbz/events{/privacy}",
"received_events_url": "https://api.github.com/users/ezpzbz/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-09-04T09:52:48
| 2021-09-10T16:11:54
|
2021-09-10T16:11:54Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Generated `bandOverlaps.lobster` file by `Lobster` (tested with `v4.0.0` and `v4.1.0`), writes spin as `spin 1` and `spin 2`. In the `self._read` method here, spins are parsed as `spin 0` and `spin 1` and therefore, it misses `spin 2` and fails.
**To Reproduce**
I have attached two following files generated by different versions of `Lobster`:
[bandOverlaps.lobster_v4.0.0.txt](https://github.com/materialsproject/pymatgen/files/7109557/bandOverlaps.lobster_v4.0.0.txt)
[bandOverlaps.lobster_v4.1.0.txt](https://github.com/materialsproject/pymatgen/files/7109558/bandOverlaps.lobster_v4.1.0.txt)
The following snippet:
```python
from pymatgen.io.lobster.outputs import Bandoverlaps
Bandoverlaps(filename='./bandOverlaps.lobster_v4.0.0.txt')
Bandoverlaps(filename='./bandOverlaps.lobster_v4.1.0.txt')
```
would results in following exception:
```console
ValueError: could not convert string to float: 'Overlap'
---------------------------------------------------------------------------
ValueError Traceback (most recent call last)
/var/folders/gv/d3m49bs95kl97vw2w2b6k78m0000gp/T/ipykernel_94202/4051178229.py in <module>
----> 1 Bandoverlaps(filename='./bandOverlaps.lobster_v4.0.0.txt')
~/.local/software/miniforge3/envs/aiida/lib/python3.9/site-packages/pymatgen/io/lobster/outputs.py in __init__(self, filename)
1304 contents = f.read().split("\n")
1305
-> 1306 self._read(contents)
1307
1308 def _read(self, contents: list):
~/.local/software/miniforge3/envs/aiida/lib/python3.9/site-packages/pymatgen/io/lobster/outputs.py in _read(self, contents)
1341 for el in line.split(" "):
1342 if el not in [""]:
-> 1343 overlaps.append(float(el))
1344 self.bandoverlapsdict[spin][" ".join(kpoint_array)]["matrix"].append(overlaps)
1345
ValueError: could not convert string to float: 'Overlap'
```
**Expected behavior**
It should not fail and we should get `bandoverlapsdict` and `max_deviation` populated with data.
I'll open a PR shortly to address the issue.
**Desktop (please complete the following information):**
- OS: Mac,Linux
- Version: 2020.0.13
- Lobster: v4.0.0 and v4.1.0
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2236/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2236/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2237
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2237/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2237/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2237/events
|
https://github.com/materialsproject/pymatgen/pull/2237
| 988,258,021
|
MDExOlB1bGxSZXF1ZXN0NzI3MjE0Mjc2
| 2,237
|
🛠 fixes the issue in parsing `bandOverlaps.lobster` file
|
{
"login": "ezpzbz",
"id": 13301031,
"node_id": "MDQ6VXNlcjEzMzAxMDMx",
"avatar_url": "https://avatars.githubusercontent.com/u/13301031?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ezpzbz",
"html_url": "https://github.com/ezpzbz",
"followers_url": "https://api.github.com/users/ezpzbz/followers",
"following_url": "https://api.github.com/users/ezpzbz/following{/other_user}",
"gists_url": "https://api.github.com/users/ezpzbz/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ezpzbz/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ezpzbz/subscriptions",
"organizations_url": "https://api.github.com/users/ezpzbz/orgs",
"repos_url": "https://api.github.com/users/ezpzbz/repos",
"events_url": "https://api.github.com/users/ezpzbz/events{/privacy}",
"received_events_url": "https://api.github.com/users/ezpzbz/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thank you for finding this. ",
"Thanks @pzarabadip! Ideally there would have a minimal test added with the fix, could you add one?",
"I'll do it @mkhorton. I'm having a little bit of issue running tests on M1 Mac but should be fixed soon. I'll updated the available `BandoverlapsTest` test class to reflect this PR. ",
"Thank you. (Let me know if I can help.)",
"One suggestion from my side:\r\n\r\nAdding an additional flag/parameter to the init of this class to still have compatibility with old versions of Lobster (something like old_spin_numbers=False )\r\nThere, spin 0 was used (see test files). If it's too much work for you, I am happy to do this later on. \r\n\r\n(Btw, if you have issues with bandoverlaps, you might need to check your POTCAR version. There are currently issues with the lastest POTCARs in Lobster. You can find more about it in the Lobsterin class.)",
"Thanks a lot @JaGeo, much appreciated. \r\n\r\nIndeed, I've started getting issues with large band overlaps recently. Thanks for pointing out to the solution. \r\n\r\nThat's a great suggestion about having the code backward compatible. I could manage to run tests locally. I'll consider backward compatibility. \r\n\r\nThanks also for your efforts and works for Lobster interface. ",
"Hi @JaGeo and @mkhorton \r\n\r\nRe backward compatibility: As @JaGeo suggested, I have added an argument to the class which can be used to switch between different version of `Lobster`. \r\n\r\nI just pushed the latest changes. I have added two new `bandOverlaps.lobster` files and consequently extended the tests in `test_lobster.py` to check the code against these files too. \r\n\r\nTests were passed locally and I also formatted the code following the guidelines to pass the tests in `lint` workflow. \r\n\r\nCheers,\r\nPezhman\r\n",
"From my side, this looks fine :). Let's see what @mkhorton thinks :).",
"Thanks @JaGeo \r\n\r\nI've been thinking that maybe I should do the following changes before merge:\r\n* Changing the `old_spin_numbers` argument to something like `lobster4` and set it to `True`\r\n* Updating the docstring to explicitly mention something like: Set to `False` if calculations were performed using `Lobster v3.x` ",
"Sounds good, @pzarabadip ",
"Or, you could check for Spin 0 in the file. If it's not there, use Spin 1 and Spin 2. If it's there, use Spin 0 and Spin 1. This would avoid all problems, wouldn't it? Sorry that I haven't thought about this before.",
"Brilliant @JaGeo \r\nThis is a much better solution for this problem. I'll update and push the change at some point today. ",
"Hi @JaGeo @mkhorton \r\nThe latest commit implements the above suggestion. Hope it would be ready to merge. \r\nCheers, \r\nPezhman",
"Looks good from my side :). Let's see what @mkhorton thinks.",
"With regards to Lobster, if you're happy I'm happy @JaGeo ! PR looks great, thank you @pzarabadip. Will merge once the CI checks have completed.\r\n\r\nAlso, for @pzarabadip, since you're a first time contributor please make sure to fill out [this form](https://forms.gle/JnisFb38QDR8QTFTA) so we can acknowledge you appropriately on our [Development Team page](https://pymatgen.org/team.html).\r\n",
"\n[](https://coveralls.io/builds/42791951)\n\nCoverage decreased (-0.6%) to 83.163% when pulling **adf6e058f9117f5209aa35a989e341e16e51c950 on pzarabadip:fix_lobster_parsing** into **e9760e2799a1d05805bbdb9971faa39fd541c1ea on materialsproject:master**.\n",
"Thanks @mkhorton and @JaGeo \r\nI also filled in the form. "
] | 2021-09-04T09:59:21
| 2021-09-10T16:35:49
|
2021-09-10T16:11:54Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Fix #2236
## Additional dependencies introduced (if any)
None.
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2237/reactions",
"total_count": 1,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 1,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2237/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2237",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2237",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2237.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2237.patch",
"merged_at": "2021-09-10T16:11:54Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2238
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2238/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2238/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2238/events
|
https://github.com/materialsproject/pymatgen/pull/2238
| 988,381,016
|
MDExOlB1bGxSZXF1ZXN0NzI3MzA2ODA3
| 2,238
|
Updates to OPTIMADE REST interface
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
] | null |
[
"\n[](https://coveralls.io/builds/42662827)\n\nCoverage decreased (-0.01%) to 83.773% when pulling **f94e8d5c32dc3671ca5aeee3066ac5a2401ecd73 on optimade-updates** into **e9760e2799a1d05805bbdb9971faa39fd541c1ea on master**.\n"
] | 2021-09-04T22:40:57
| 2022-10-10T17:54:17
|
2021-09-05T20:44:01Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Preparing new features in advance of OPTIMADE lesson on Tuesday.
The main feature added in this PR is for `pymatgen.ext.optimade.OptimadeRester` to allow querying multiple providers simultaneously. It also adds more logging and reporting/user experience improvements.
## TODO (if any)
There are several TODOs added inside the code, mainly related to more robust error handling and more closely following (or requiring) conformance with the OPTIMADE specification.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2238/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2238/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2238",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2238",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2238.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2238.patch",
"merged_at": "2021-09-05T20:44:01Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2239
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2239/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2239/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2239/events
|
https://github.com/materialsproject/pymatgen/pull/2239
| 989,467,819
|
MDExOlB1bGxSZXF1ZXN0NzI4MjA1ODEx
| 2,239
|
Speed up nearest-neighbor routines & structure graph generation
|
{
"login": "ltalirz",
"id": 1053245,
"node_id": "MDQ6VXNlcjEwNTMyNDU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1053245?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ltalirz",
"html_url": "https://github.com/ltalirz",
"followers_url": "https://api.github.com/users/ltalirz/followers",
"following_url": "https://api.github.com/users/ltalirz/following{/other_user}",
"gists_url": "https://api.github.com/users/ltalirz/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ltalirz/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ltalirz/subscriptions",
"organizations_url": "https://api.github.com/users/ltalirz/orgs",
"repos_url": "https://api.github.com/users/ltalirz/repos",
"events_url": "https://api.github.com/users/ltalirz/events{/privacy}",
"received_events_url": "https://api.github.com/users/ltalirz/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/42699463)\n\nCoverage decreased (-0.6%) to 83.143% when pulling **30a1b6f1198aa42bb413bad686c9412c10db6aef on ltalirz:speedup-cutoff-dict** into **1ea090680ce34ca944cd7671032fa274bf035192 on materialsproject:master**.\n",
"I don't think the linting and test failures are related to this PR.\r\nHappy to correct issues if there are any",
"Thanks. Can you please merge the latest pymatgen master to yours? Those linting errors were fixed recently.",
"Hi @shyuep , I just rebased my PR but the linting errors persist.\r\nNote that they also occur in [the latest commit to `master`](https://github.com/materialsproject/pymatgen/commit/1ea090680ce34ca944cd7671032fa274bf035192).",
"Thanks.",
"Apologies @ltalirz, I think we're going to have to temporarily revert this commit.\r\n\r\nIt's causing a significant slowdown in production code -- we have code that previously took 0.1 seconds to run that now takes upwards of 20 seconds.\r\n\r\nI suspect this PR is mostly still good but it's going to take some time to find exactly where the slow down is happening and address it.",
"I don't have a minimum reproducible example yet, but I think it might be re-created with a large `Structure`, `CutOffDictNN` with `cut_off_dict=None` (i.e. this should be a very cheap operation). Do you have any thoughts where the bottleneck should be?",
"Hey @mkhorton , sorry about the performance regression - as you can see from the PR description for me the result was quite the opposite.\r\nLet me first check whether I somehow messed something up in the PR (I'll run my tests locally again)\r\n\r\nAnother thing you can check is whether you see the problem already with https://github.com/materialsproject/pymatgen/pull/2239/commits/0d5cb9f1712530cfdc09f12103c85031a7087ac3\r\nor whether you need the second commit of the PR",
"Thanks for looking into this @ltalirz. It's possible this PR has just identified another, different slow part of existing code. I'm sure we can get this resolved.",
"> it might be re-created with a large `Structure`, `CutOffDictNN` with `cut_off_dict=None` (i.e. this should be a very cheap operation\r\n\r\nThanks for pointing this out - I suspect that https://github.com/materialsproject/pymatgen/commit/0d5cb9f1712530cfdc09f12103c85031a7087ac3 will be fine (I still get fast timings for large structures with `cut_off_dict=None` for that commit); will now look into the second commit to see where the problem lies.\r\n\r\nJust to make sure I understand correctly, though: `cut_off_dict=None` means \"no cutoff\", i.e. one needs to create an all-to-all graph, right? I.e. in terms of graph creation this would be an expensive operation (but of course it should not be excessive).",
"Ok, I see now - one does not need to check for neighbors.\r\nPerhaps there is a special case for `None` missing, let's see.",
"So, as you anticipated, the issue seems to be that for `cut_off_dict=None` (there not being any special case) the call to the cythonized `IStructure.get_neighbor_list(max_dist)` gets very expensive; even more expensive than iterating over all atoms and computing their neighbors individually.\r\n\r\nFrom the outside I can't think of a good reason why one should expect this behavior (note that for reasonable cutoffs, calling `get_neighbor_list` once is actually much faster), i.e. I'll need to look at the implementation of `IStructure.get_neighbor_list` which might need to wait until tomorrow. \r\n\r\nUntil this is addressed, however, a quick fix would be to use just https://github.com/materialsproject/pymatgen/commit/0d5cb9f1712530cfdc09f12103c85031a7087ac3 from this PR which does not yet touch `get_all_nn_info` but already gets rid of the scaling issue (O(N^2) => O(N) ).",
"Ok, that helps. Here's what I'm going to do:\r\n\r\n1. Release `v2022.0.16` today that reverts this commit, to address requirements for people who want to use features from `v2022.0.15` without the slow down\r\n2. Suggest you keep using `v2022.0.15` for the time being to enjoy these optimizations\r\n3. Next week, release `v2022.0.17` either including just 0d5cb9f or including the full PR with a fix for the special `None` case",
"Thanks @mkhorton \r\n\r\nFor the fix it would be good to know what your use case for `cut_off_dict=None` is (and thus the expected graph).\r\n\r\nMathematically speaking, in a periodic structure without any cutoff the neighbor list of each atom should be infinite.",
"The way that `CutOffDictNN` works is to only consider connections between specific pairs of atoms, so implicitly if no pairs are requested then there are no bonds.\r\n\r\ne.g. I think there's a semantic distinction between, say, `cut_off_dict={(\"Cl\", \"Cl\"): 0.0}`, `cut_off_dict={(\"Cl\", \"Cl\"): None}` and `cut_off_dict={}` or `cut_off_dict=None`.\r\n\r\nThe current behavior is that if `cut_off_dict=None`, no pairs of atoms are considered and no bonds are drawn. For any atoms present and _not_ specified in the `cut_off_dict`, they are ignored.\r\n\r\nIf you think this behavior should be changed I think that's ok, but since it would be a change in the current behavior, so it'll have to be communicated appropriately.",
"@chc273 Might be an issue with the NN routinues for individual atoms? In any case, I am confused why this issue exists. It should be faster to compute all the atoms in one shot and then getting the neighbors of each one, rather than looping over atoms?",
"@shyuep I think the issue might be using the get neighbor algorithm with a very small cutoff. \r\n\r\nThe current cell list algorithm divides the space into small cubes, with lengths equal to the cutoff. If the cutoff is very small, then the number of cubes (~O((a/cutoff)^3)) will explode. \r\n\r\nI could threshold the cutoff to be greater than 1 A, if people request a smaller cutoff, I can take the 1A neighbor list and then discard those with the neighbor distance greater than `cutoff`? ",
"@shyuep I submitted #2277 to deal with the issue in the previous reply. ",
"Dear all, thanks for the quick follow-ups on your side!\r\n\r\n\r\n> The way that `CutOffDictNN` works is to only consider connections between specific pairs of atoms, so implicitly if no pairs are requested then there are no bonds.\r\n> The current behavior is that if `cut_off_dict=None`, no pairs of atoms are considered and no bonds are drawn. For any atoms present and _not_ specified in the `cut_off_dict`, they are ignored.\r\n\r\nOk, I see. I was certainly not aware of this logic and since I adapted the `get_all_nn_info` from `core.structure.IStructure.get_all_neighbors` (which probably does not have this logic), this may well be the origin of the issue.\r\n\r\nI'm not sure I'll get to it today but I should find time tomorrow.\r\n\r\n> If you think this behavior should be changed I think that's ok, but since it would be a change in the current behavior, so it'll have to be communicated appropriately.\r\n\r\nNo need to change, just opened https://github.com/materialsproject/pymatgen/pull/2278 to make this explicit in the docstring for the future.\r\n\r\n> @shyuep I think the issue might be using the get neighbor algorithm with a very small cutoff.\r\n\r\nThanks @chc273 , I see - `cut_off_dict=None` does indeed result in `CutOffDictNN._max_dist=0.0`, which would be good to speed up as well. \r\n\r\nAt the same time, I now realize that I should also filter the sites passed into the `get_neighbor_list` by the pairs defined in the cutoff dict - an element that does not appear in any cutoff pair should not be passed into `get_neighbor_list` (which would also have fixed the problem observed by @mkhorton ).\r\n\r\nWill work on it tomorrow."
] | 2021-09-06T22:50:14
| 2021-10-23T22:21:00
|
2021-09-07T13:59:43Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
The `get_nn_info` routines of many nearest-neighbor flavors in `pymatgen.analysis.local_env` recomputed the original periodic image of sites although this information was already stored on the `PeriodicNeighbor` objects (at least it seems so to me - correct me if I'm wrong).
This resulted in O(N^2) instead of O(N) computational complexity (for structures beyond a certain size, almost all time was spent there).
This PR
* Changes the `get_nn_info` methods to reuse the information from the `PeriodicNeighbor` objects (1st commit)
* Introduces a faster implementation of `get_all_nn_info` for the `CutoffDictNN` (2nd commit)
The 2nd commit involves some code duplication and is perhaps not absolutely necessary.
It results in further substantial speedup, however.
Example to demonstrate the speedup (adapted from pymatgen test suite)
```python
from pymatgen.analysis.graphs import *
from pymatgen.analysis.local_env import (
CovalentBondNN,
CutOffDictNN,
MinimumDistanceNN,
MinimumOKeeffeNN,
OpenBabelNN,
VoronoiNN,
)
nacl_lattice = Lattice(
[
[3.48543625, 0.0, 2.01231756],
[1.16181208, 3.28610081, 2.01231756],
[0.0, 0.0, 4.02463512],
]
)
nacl = Structure(nacl_lattice, ["Na", "Cl"], [[0, 0, 0], [0.5, 0.5, 0.5]])
nacl.make_supercell([10,10,10])
print(len(nacl)) # 2000 atoms
```
```
# old
In [2]: %timeit -n1 -r1 nacl_graph = Struc
...: tureGraph.with_local_env_strategy(
...: nacl, CutOffDictNN({("Cl", "Cl"):
...: 5.0}))
6min 56s ± 0 ns per loop (mean ± std. dev. of 1 run, 1 loop each)
# improved get_nn_info
In [6]: %timeit -n1 -r1 nacl_graph = Struc
...: tureGraph.with_local_env_strategy(
...: nacl, CutOffDictNN({("Cl", "Cl"):
...: 5.0}))
4.42 s ± 0 ns per loop (mean ± std. dev. of 1 run, 1 loop each)
# improved get_all_nn_info
In [2]: %timeit -n1 -r1 nacl_graph = Struc
...: tureGraph.with_local_env_strategy(
...: nacl, CutOffDictNN({("Cl", "Cl"):
...: 5.0}))
444 ms ± 0 ns per loop (mean ± std. dev. of 1 run, 1 loop each)
```
I picked 2000 atoms since the original implementation was so slow.
On larger structures, the difference between commit 1 and commit 1+2 becomes larger than 10x, e.g. for 8192 atoms (16x16x16 supercell):
```
In [1]: %run test.py
8192
# improved get_nn_info
In [2]: %timeit -n1 -r1 nacl_graph = Struc
...: tureGraph.with_local_env_strategy(
...: nacl, CutOffDictNN({("Cl", "Cl"):
...: 5.0}))
1min 3s ± 0 ns per loop (mean ± std. dev. of 1 run, 1 loop each)
# improved get_all_nn_info
In [2]: %timeit -n1 -r1 nacl_graph = Struc
...: tureGraph.with_local_env_strategy(
...: nacl, CutOffDictNN({("Cl", "Cl"):
...: 5.0}))
1.8 s ± 0 ns per loop (mean ± std. dev. of 1 run, 1 loop each)
```
## Checklist
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [ ] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2239/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2239/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2239",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2239",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2239.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2239.patch",
"merged_at": "2021-09-07T13:59:43Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2240
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2240/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2240/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2240/events
|
https://github.com/materialsproject/pymatgen/issues/2240
| 989,491,820
|
MDU6SXNzdWU5ODk0OTE4MjA=
| 2,240
|
io.lammps.data.LammpsData does not read charge from structure
|
{
"login": "chc273",
"id": 22353204,
"node_id": "MDQ6VXNlcjIyMzUzMjA0",
"avatar_url": "https://avatars.githubusercontent.com/u/22353204?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/chc273",
"html_url": "https://github.com/chc273",
"followers_url": "https://api.github.com/users/chc273/followers",
"following_url": "https://api.github.com/users/chc273/following{/other_user}",
"gists_url": "https://api.github.com/users/chc273/gists{/gist_id}",
"starred_url": "https://api.github.com/users/chc273/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/chc273/subscriptions",
"organizations_url": "https://api.github.com/users/chc273/orgs",
"repos_url": "https://api.github.com/users/chc273/repos",
"events_url": "https://api.github.com/users/chc273/events{/privacy}",
"received_events_url": "https://api.github.com/users/chc273/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"It seems that the `oxidation_states` should have been considered as \"charge\" in the `site_properties` of a structure."
] | 2021-09-07T00:13:00
| 2021-09-07T00:46:33
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
For a `structure` with decorated atomic charges, I would expect `LammpsData.from_structure(structure, atom_style='charge')` to read the atom charges from the `structure`. However, this is not the case and the charges are all assigned zero.
**To Reproduce**
```
from pymatgen.core import Structure
from pymatgen.io.lammps.data import LammpsData
cif = "# generated using pymatgen\ndata_NaCl\n_symmetry_space_group_name_H-M 'P 1'\n_cell_length_a 2.66700000\n_cell_length_b 2.66700000\n_cell_length_c 2.66700000\n_cell_angle_alpha 90.00000000\n_cell_angle_beta 90.00000000\n_cell_angle_gamma 90.00000000\n_symmetry_Int_Tables_number 1\n_chemical_formula_structural NaCl\n_chemical_formula_sum 'Na1 Cl1'\n_cell_volume 18.97007496\n_cell_formula_units_Z 1\nloop_\n _symmetry_equiv_pos_site_id\n _symmetry_equiv_pos_as_xyz\n 1 'x, y, z'\nloop_\n _atom_type_symbol\n _atom_type_oxidation_number\n Na+ 1.0\n Cl- -1.0\nloop_\n _atom_site_type_symbol\n _atom_site_label\n _atom_site_symmetry_multiplicity\n _atom_site_fract_x\n _atom_site_fract_y\n _atom_site_fract_z\n _atom_site_occupancy\n Na+ Na0 1 0.00000000 0.00000000 0.00000000 1.0\n Cl- Cl1 1 0.50000000 0.50000000 0.50000000 1.0\n"
s = Structure.from_str(cif, fmt='cif')
print(s) ## each atom has charges
ld = LammpsData.from_structure(s, atom_style='charge')
ld.write_file('data.static')
```
The `data.static` file is as follows
```
Generated by pymatgen.io.lammps.data.LammpsData
2 atoms
2 atom types
0.000000 2.667000 xlo xhi
0.000000 2.667000 ylo yhi
0.000000 2.667000 zlo zhi
Masses
1 22.989769
2 35.453000
Atoms
1 1 0.0000 0.000000 0.000000 0.000000
2 2 0.0000 1.333500 1.333500 1.333500
```
you can see that the atom charges (the third column in `Atoms`) are all zeros.
**Expected behavior**
Atom charges should not be zero if `structure` is decorated with atom charges.
**Desktop (please complete the following information):**
- OS: [e.g. Mac,Windows,Linux]
- Version [v2022.0.14]
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2240/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2240/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2241
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2241/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2241/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2241/events
|
https://github.com/materialsproject/pymatgen/pull/2241
| 989,754,153
|
MDExOlB1bGxSZXF1ZXN0NzI4NDQ0NzA2
| 2,241
|
improve Structure.from_str fmt doc string and type hint
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Sorry about this hasty PR. I'd never looked into [contravariance](https://www.python.org/dev/peps/pep-0483/#covariance-and-contravariance) before. I still think there would be some value in having a more restrictive type hint (same for docs) but seems that would require use of `# type: ignore` on [line 2338](https://github.com/materialsproject/pymatgen/pull/2241/files#diff-5ea334b067925ce7079b569683cf7cc7ee088d0b19d62e4257f79cbb6aa73914R2338). If you disagree, feel free to close.",
"I think type hints are useful but we should not introduce them and then ignore them? ",
"The type hint would be for IDEs and users reading the code. It would only be ignored by linters, i.e. `mypy`. And I think the doc change in this PR is an improvement in any case.",
"I like this change, I think we should re-open the PR.\r\n\r\nHowever, *I agree type hints should not be ignored* -- can we fix this? We shouldn't merge it with `ignore` present. I'm not familiar with the contravariance issue, can you explain what the problem is here?",
"Paraphrasing from [Wikipedia](https://en.wikipedia.org/wiki/Covariance_and_contravariance_(computer_science)), a typing rule is covariant if it preserves the type ordering from more specific to more generic and contravariant if it reverses this ordering.\r\n\r\nThat wasn't very intelligible to me. But it becomes easier to grasp, if you break it down for our specific case of inheritance and overriding a class method. A method is covariant in its _output_ (i.e. a more restrictive output type means the subclass method is a subtype of the parent method) but contravariant in its _input_, i.e. a more restrictive type rule for inputs means the subclass method becomes more generic and is no longer a subtype of the parent method.\r\n\r\nIn this specific case, `fmt: Literal[\"cif\", \"poscar\", ...]` is a more specific type than `fmt: str` in the parent method so the subclass method is not a proper subtype of the base method and that's why `mypy` is complaining. Tbh, this seems like a face palm for OOP to me.",
"Personally, I think the type problem would be solved if we add an \"Any\" type to some of the type hints?",
"> Personally, I think the type problem would be solved if we add an \"Any\" type to some of the type hints?\r\n\r\nYou mean set the type of `fmt` to `Any` in the parent method, i.e. `SiteCollection.from_str(fmt: Any)`? I think that would work.",
"I mean you can add proper type hints as what you have done, but allow an \"Any\" as a final option. That would still give people lots of information of the normal expected types (e.g., the literals) but at the same time provide enough flexibility if someone introduces a strange new type.",
"Personally, I'd go with adding the `Any` to the parent method. Having the `Any` in the `Structure` would be an incorrect type and it looks bad in type hints:\r\n\r\n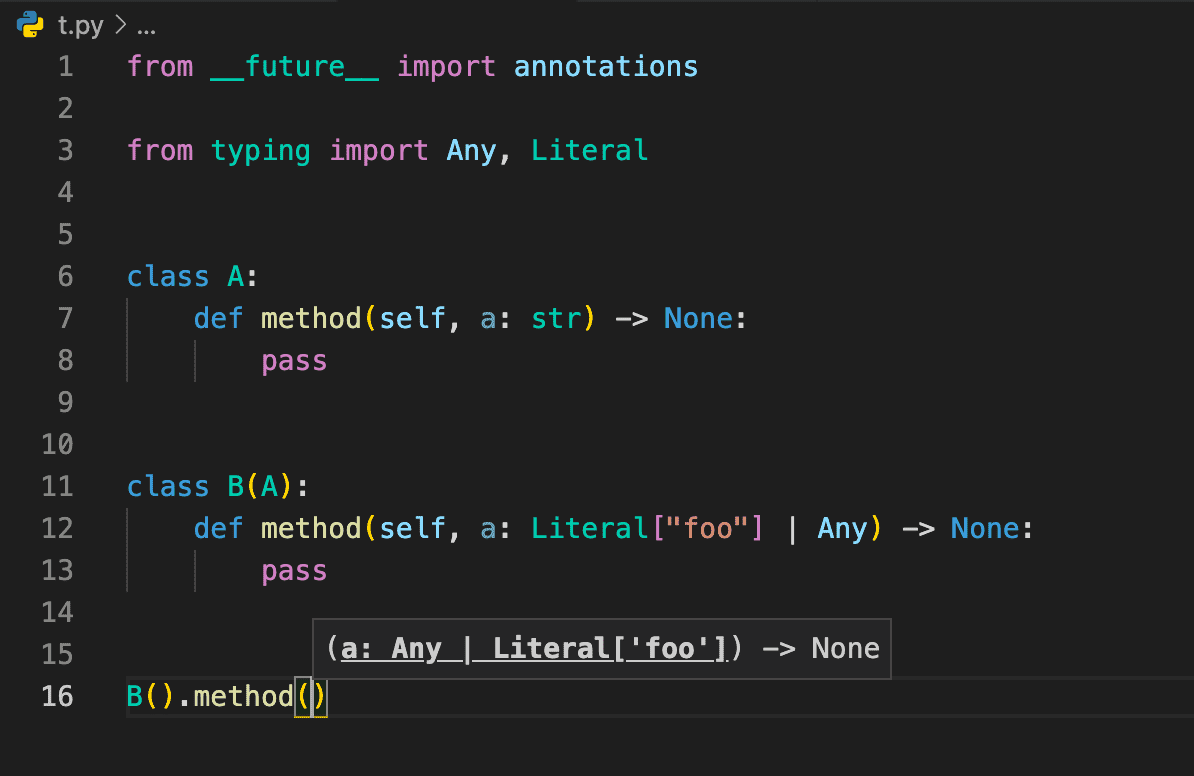\r\n\r\nCompared to this which `mypy` is also fine with:\r\n\r\n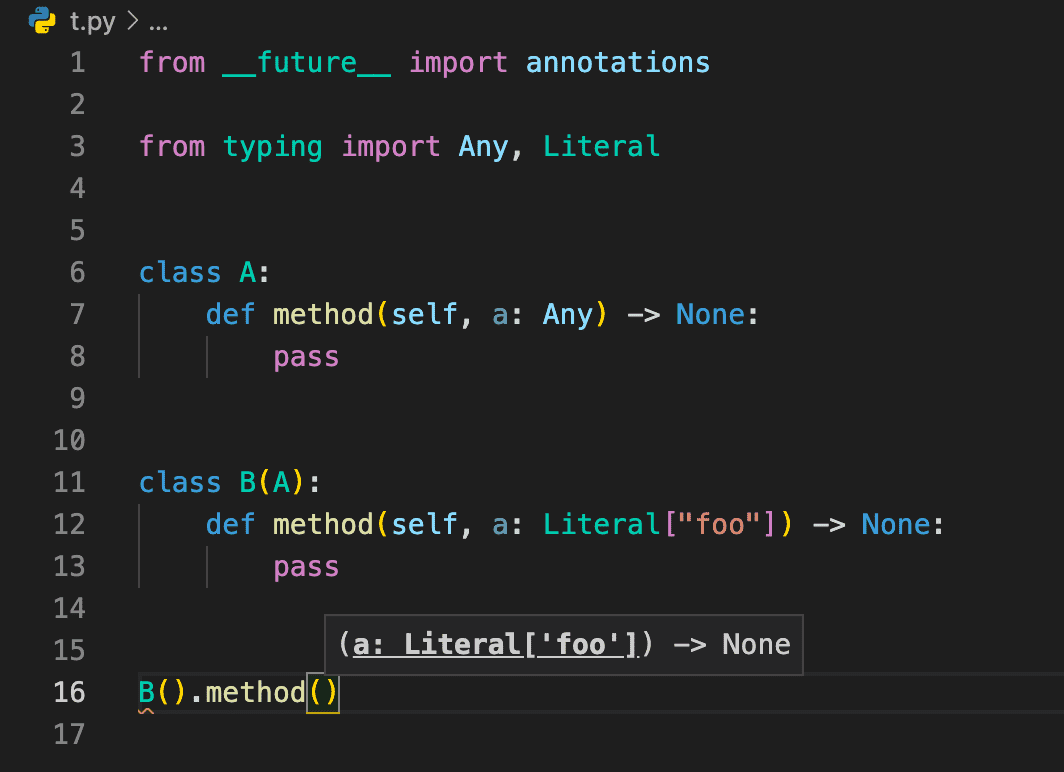\r\n",
"This is fine.",
"@shyuep Made the change. You prob need to reopen for it to sho up here.",
"\n[](https://coveralls.io/builds/42796535)\n\nCoverage decreased (-0.6%) to 83.161% when pulling **6b8019438d726dd1a133265381aee6d2e5867f6c on janosh:patch-1** into **1ea090680ce34ca944cd7671032fa274bf035192 on materialsproject:master**.\n",
"Unrelated question: Why import all of\r\n\r\n```py\r\nfrom pymatgen.io.atat import Mcsqs\r\nfrom pymatgen.io.cif import CifParser\r\nfrom pymatgen.io.cssr import Cssr\r\nfrom pymatgen.io.vasp import Poscar\r\nfrom pymatgen.io.xcrysden import XSF\r\n```\r\n\r\nin `from_str()` if you're only going to use 1 (or none) of them? Why not import inside the matching `if` statement?",
"The remaining CI failures seem unrealted to this PR.",
"The linting failure should have been fixed in the main branch, so please disregard.\r\n\r\nRegarding the imports, I'm not sure it matters too much -- generally it's bad form to import within a method anyway instead of the top of the file (but I see why it's done here), I think they could get moved inside the respective `if` statement, so by all means make the change.\r\n\r\nPerhaps the bigger issue with `Structure.from_str` is how it differs from `Structure.from_file` in non-obvious ways. I think this might be improved by abstracting a `_detect_fmt_from_filename()` function and then having `from_file` call `from_str` directly.",
"I would suggest deferring the changes other than the type hints. My preference is to make these `from_str` or `from_filename` into proper plugins. That way, if a person adds a new io class, `Structure.from_str` would automatically support the new fmt. This requires some inspection methods and probably an abstract class.\r\n\r\nThe reason why the imports are in the methods in this case is due to circular dependencies. E.g., Poscar imports Structure, so Structure cannot import Poscar. You can move them to the respective if statements and I have no objections to that.",
"Ok, sounds sensible, let's defer any additional changes (aside from the `if`) for a later date.",
"While were on the topic of types, how about adding type checking as a pre-commit hook. I noticed you're already using `pre-commit`. I can recommend the [`mypy mirror`](https://github.com/pre-commit/mirrors-mypy), use it on all my projects.",
"Anything blocking this PR?",
"@mkhorton The linting and test errors both appear unrelated to this PR."
] | 2021-09-07T08:56:45
| 2021-10-06T19:15:01
|
2021-10-06T19:12:59Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2241/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2241/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2241",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2241",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2241.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2241.patch",
"merged_at": "2021-10-06T19:12:59Z"
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2242
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2242/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2242/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2242/events
|
https://github.com/materialsproject/pymatgen/pull/2242
| 990,007,621
|
MDExOlB1bGxSZXF1ZXN0NzI4NjYyNTA4
| 2,242
|
Add two more pre-defined Optimade aliases
|
{
"login": "blokhin",
"id": 177388,
"node_id": "MDQ6VXNlcjE3NzM4OA==",
"avatar_url": "https://avatars.githubusercontent.com/u/177388?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/blokhin",
"html_url": "https://github.com/blokhin",
"followers_url": "https://api.github.com/users/blokhin/followers",
"following_url": "https://api.github.com/users/blokhin/following{/other_user}",
"gists_url": "https://api.github.com/users/blokhin/gists{/gist_id}",
"starred_url": "https://api.github.com/users/blokhin/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/blokhin/subscriptions",
"organizations_url": "https://api.github.com/users/blokhin/orgs",
"repos_url": "https://api.github.com/users/blokhin/repos",
"events_url": "https://api.github.com/users/blokhin/events{/privacy}",
"received_events_url": "https://api.github.com/users/blokhin/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @blokhin!"
] | 2021-09-07T13:50:23
| 2021-09-07T16:06:59
|
2021-09-07T15:57:01Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2242/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2242/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2242",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2242",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2242.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2242.patch",
"merged_at": "2021-09-07T15:57:01Z"
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2243
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2243/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2243/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2243/events
|
https://github.com/materialsproject/pymatgen/issues/2243
| 990,049,615
|
MDU6SXNzdWU5OTAwNDk2MTU=
| 2,243
|
Extraneous parameter in OPTIMADE extension when using raw filters (plus a missing dependency)
|
{
"login": "ml-evs",
"id": 7916000,
"node_id": "MDQ6VXNlcjc5MTYwMDA=",
"avatar_url": "https://avatars.githubusercontent.com/u/7916000?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ml-evs",
"html_url": "https://github.com/ml-evs",
"followers_url": "https://api.github.com/users/ml-evs/followers",
"following_url": "https://api.github.com/users/ml-evs/following{/other_user}",
"gists_url": "https://api.github.com/users/ml-evs/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ml-evs/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ml-evs/subscriptions",
"organizations_url": "https://api.github.com/users/ml-evs/orgs",
"repos_url": "https://api.github.com/users/ml-evs/repos",
"events_url": "https://api.github.com/users/ml-evs/events{/privacy}",
"received_events_url": "https://api.github.com/users/ml-evs/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @ml-evs,\r\n\r\nThanks for the write-up. I will address 1. myself, but if you're willing to make a PR for the latter, that would be of course welcome!",
"Great!\r\n\r\nre: `retrying`, I ran into it in a fresh binder env that I used for your tutorial: https://mybinder.org/v2/gh/Materials-Consortia/optimade-tutorial-exercises/HEAD?filepath=notebooks%2Fdemonstration-pymatgen-for-optimade-queries.ipynb\r\n\r\nIf you remove it from the first `pip install` line then the import of `OptimadeRester` fails, so not sure it is pulled in by plotly.",
"Looks like binder's default env is Python 3.7, if that helps, and some plotly subpackage has some kind of dependency on it... \r\n\r\nhttps://github.com/plotly/plotly.py/blob/03979d105c65dda3df3a155322eaff18f203b03f/packages/python/chart-studio/setup.py#L44",
"Yes, I think I mis-remembered, or maybe it changed. I disabled it for now at least pending a re-evaluation."
] | 2021-09-07T14:30:57
| 2021-09-15T21:06:49
|
2021-09-15T21:06:49Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Some minor bugs arising from today's OPTIMADE NOMAD tutorial:
1. The library [`retrying`](https://pypi.org/project/retrying/) is used in the [OPTIMADE extension](https://github.com/materialsproject/pymatgen/blob/f72963b038be33c058c844f93ec2392f1613fffe/pymatgen/ext/optimade.py#L13) but it is not listed as a pymatgen dependency (this was not a problem for the Colab examples as Colab already includes `retrying` in its default environment).
2. When providing a "raw" OPTIMADE filter to `OptimadeRester`, the response fields are set incorrectly, with an extraneous `fields=` prefix. I'm a bit worried that this is actually somewhere where OPTIMADE diverged from [JSON:API Fields](https://jsonapi.org/format/1.1/#document-resource-object-fields) that we could maybe address in the future (by e.g., allowing both).
The fix is just changing:
https://github.com/materialsproject/pymatgen/blob/f72963b038be33c058c844f93ec2392f1613fffe/pymatgen/ext/optimade.py#L282
to
```diff
+ url = join(resource, f"v1/structures?filter={optimade_filter}&fields={fields}")
- url = join(resource, f"v1/structures?filter={optimade_filter}&{fields}")
```
(reported by Daniel Massote)
**Desktop (please complete the following information):**
- Version 2020.0.14
I'm happy to make the PR to address no. 2, but I assume a pymatgen maintainer will want to deal with no. 1 as they see fit.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2243/reactions",
"total_count": 2,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 1,
"eyes": 1
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2243/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2244
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2244/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2244/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2244/events
|
https://github.com/materialsproject/pymatgen/pull/2244
| 990,143,321
|
MDExOlB1bGxSZXF1ZXN0NzI4Nzc2Mzk1
| 2,244
|
Remove bad param when setting OPTIMADE response fields
|
{
"login": "ml-evs",
"id": 7916000,
"node_id": "MDQ6VXNlcjc5MTYwMDA=",
"avatar_url": "https://avatars.githubusercontent.com/u/7916000?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ml-evs",
"html_url": "https://github.com/ml-evs",
"followers_url": "https://api.github.com/users/ml-evs/followers",
"following_url": "https://api.github.com/users/ml-evs/following{/other_user}",
"gists_url": "https://api.github.com/users/ml-evs/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ml-evs/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ml-evs/subscriptions",
"organizations_url": "https://api.github.com/users/ml-evs/orgs",
"repos_url": "https://api.github.com/users/ml-evs/repos",
"events_url": "https://api.github.com/users/ml-evs/events{/privacy}",
"received_events_url": "https://api.github.com/users/ml-evs/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @ml-evs! Since you're a first time contributor, I had to manually approve the CI, but will merge once it's run.\r\n\r\nBtw, (also since you're a first time contributor), please make sure to fill out [this form](https://forms.gle/JnisFb38QDR8QTFTA) so we can acknowledge you appropriately on our [development team page](https://pymatgen.org/team.html).",
"Hi @mkhorton, looks like I need approval again for the workflows.\r\n\r\nI've rebased my PR on master so now the `retrying` dep failures should not occur. I also added a test for the raw filter.",
"\n[](https://coveralls.io/builds/42905434)\n\nCoverage decreased (-0.6%) to 83.162% when pulling **b6f24189c6161097899419620a32e0afbf11fc0f on ml-evs:patch-1** into **b8b27c6fa3236f032581aa82f668b25a78aa3b57 on materialsproject:master**.\n",
"Much obliged @ml-evs !"
] | 2021-09-07T16:12:56
| 2021-09-15T21:06:49
|
2021-09-15T21:06:49Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
OPTIMADE uses the `response_fields` URL parameter to select the response fields, this PR removes an extraneous URL parameter that is added when using a raw OPTIMADE filter through `OptimadeRester`.
Addresses and closes #2243 (cc @mkhorton)
~Haven't added any tests as I don't think this extension is being tested at the moment~
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2244/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2244/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2244",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2244",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2244.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2244.patch",
"merged_at": "2021-09-15T21:06:49Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2245
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2245/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2245/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2245/events
|
https://github.com/materialsproject/pymatgen/issues/2245
| 993,147,749
|
MDU6SXNzdWU5OTMxNDc3NDk=
| 2,245
|
cannot import name 'coord_cython' from 'pymatgen.util'
|
{
"login": "courage0412",
"id": 90034730,
"node_id": "MDQ6VXNlcjkwMDM0NzMw",
"avatar_url": "https://avatars.githubusercontent.com/u/90034730?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/courage0412",
"html_url": "https://github.com/courage0412",
"followers_url": "https://api.github.com/users/courage0412/followers",
"following_url": "https://api.github.com/users/courage0412/following{/other_user}",
"gists_url": "https://api.github.com/users/courage0412/gists{/gist_id}",
"starred_url": "https://api.github.com/users/courage0412/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/courage0412/subscriptions",
"organizations_url": "https://api.github.com/users/courage0412/orgs",
"repos_url": "https://api.github.com/users/courage0412/repos",
"events_url": "https://api.github.com/users/courage0412/events{/privacy}",
"received_events_url": "https://api.github.com/users/courage0412/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-09-10T11:31:22
| 2021-09-10T11:31:57
|
2021-09-10T11:31:57Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
When I run 'import pymatgen.core.structure import Molecule' to check whether it works well, something wrong happens like this:
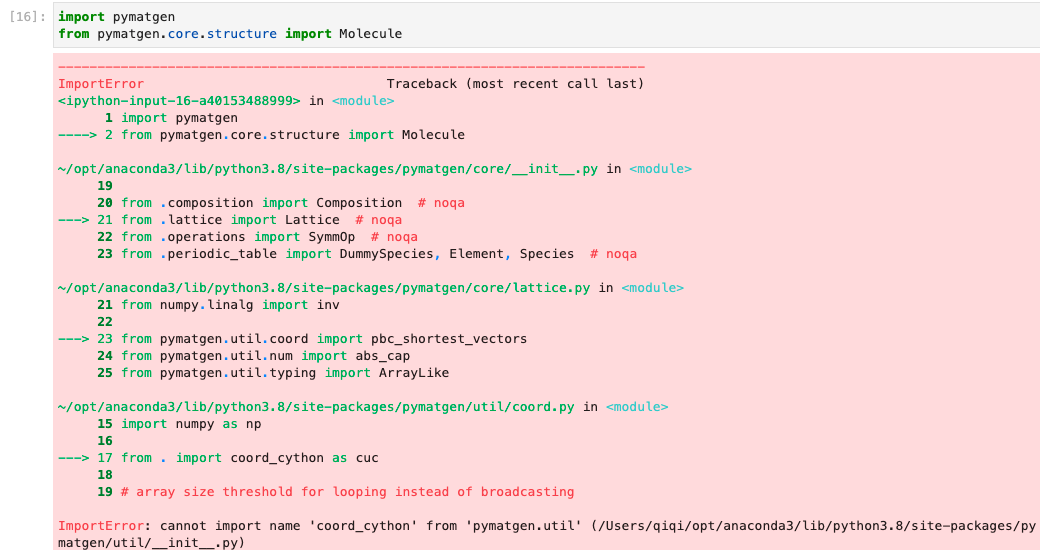
Is there any reason or solution for this?
**To Reproduce**
Steps to reproduce the behavior:
1. Go to '...'
2. Click on '....'
3. Scroll down to '....'
4. See error
Provide any example files that are needed to reproduce the error,
especially if the bug pertains to parsing of a file.
**Expected behavior**
A clear and concise description of what you expected to happen.
**Screenshots**
If applicable, add screenshots to help explain your problem.
**Desktop (please complete the following information):**
- OS: [e.g. Mac,Windows,Linux]
- Version [e.g. 2019.9.16]
**Additional context**
Add any other context about the problem here.
|
{
"login": "courage0412",
"id": 90034730,
"node_id": "MDQ6VXNlcjkwMDM0NzMw",
"avatar_url": "https://avatars.githubusercontent.com/u/90034730?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/courage0412",
"html_url": "https://github.com/courage0412",
"followers_url": "https://api.github.com/users/courage0412/followers",
"following_url": "https://api.github.com/users/courage0412/following{/other_user}",
"gists_url": "https://api.github.com/users/courage0412/gists{/gist_id}",
"starred_url": "https://api.github.com/users/courage0412/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/courage0412/subscriptions",
"organizations_url": "https://api.github.com/users/courage0412/orgs",
"repos_url": "https://api.github.com/users/courage0412/repos",
"events_url": "https://api.github.com/users/courage0412/events{/privacy}",
"received_events_url": "https://api.github.com/users/courage0412/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2245/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2245/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2246
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2246/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2246/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2246/events
|
https://github.com/materialsproject/pymatgen/issues/2246
| 993,149,388
|
MDU6SXNzdWU5OTMxNDkzODg=
| 2,246
|
cannot import name 'coord_cython' from 'pymatgen.util'
|
{
"login": "courage0412",
"id": 90034730,
"node_id": "MDQ6VXNlcjkwMDM0NzMw",
"avatar_url": "https://avatars.githubusercontent.com/u/90034730?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/courage0412",
"html_url": "https://github.com/courage0412",
"followers_url": "https://api.github.com/users/courage0412/followers",
"following_url": "https://api.github.com/users/courage0412/following{/other_user}",
"gists_url": "https://api.github.com/users/courage0412/gists{/gist_id}",
"starred_url": "https://api.github.com/users/courage0412/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/courage0412/subscriptions",
"organizations_url": "https://api.github.com/users/courage0412/orgs",
"repos_url": "https://api.github.com/users/courage0412/repos",
"events_url": "https://api.github.com/users/courage0412/events{/privacy}",
"received_events_url": "https://api.github.com/users/courage0412/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Pls reinstall pymatgen. Try pip install --update pymatgen.",
"> Pls reinstall pymatgen. Try pip install --update pymatgen.\r\n\r\nIt works, thank you so much for your help!",
"I've tried upgrade it but it didn't work. There is no coord_cython.py but a coord_cython.cp39-win_amd64.pyd in pymatgen/util/.\r\nAnd seems the error just happen with jupyter notebook (it runs well on pycharm...).",
"This just means that your Jupyter is in a different environment from your pymatgen install. Use a dedicated environment. "
] | 2021-09-10T11:33:53
| 2023-09-21T03:26:43
|
2021-09-16T15:38:52Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hello everyone,
When I run 'from pymatgen.core.structure import Molecule' to check whether it works well, something wrong happens like this:
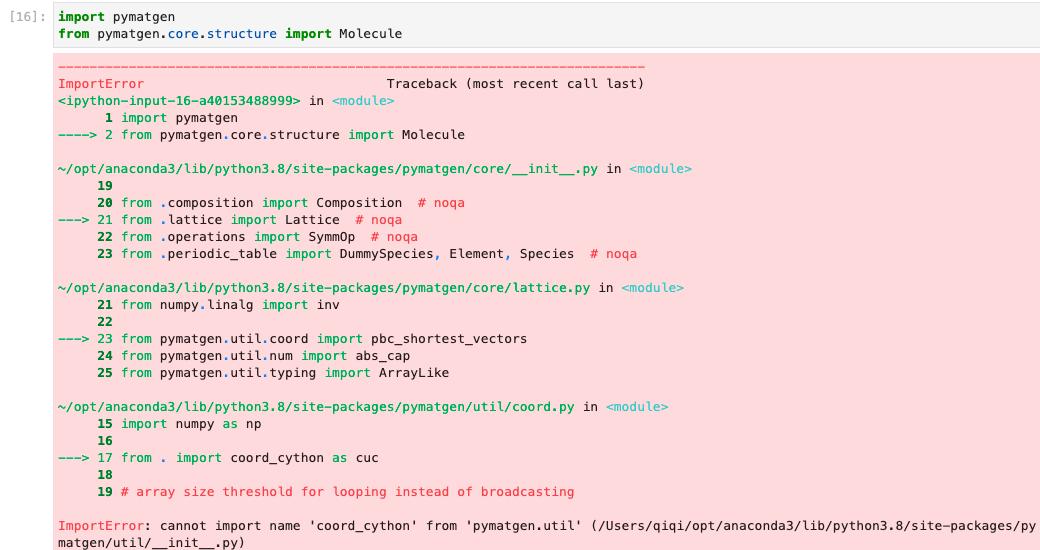
Is there any reason or solution for this issue?
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2246/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2246/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2247
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2247/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2247/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2247/events
|
https://github.com/materialsproject/pymatgen/pull/2247
| 993,915,879
|
MDExOlB1bGxSZXF1ZXN0NzMxOTg0MTY4
| 2,247
|
Type hints for literal string kwargs
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/43286494)\n\nCoverage decreased (-0.6%) to 83.134% when pulling **069a01ae5d36ef04776d09b92a933195baf2dca2 on janosh:literal-types** into **2926a20ccdf5c41a17eb4bd79fe76f1d6ae0afc8 on materialsproject:master**.\n",
"I think that would be great, no objections from me as long as any mypy failures are also dealt with, thank you!",
"This is great, thank you!"
] | 2021-09-11T18:58:52
| 2021-10-06T19:41:31
|
2021-10-06T19:41:03Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Follow up to #2241. Here are a few more examples where `Literal` type hints make sense. I think there are ~80 more places throughout pymatgen. Happy to address more of them in this PR if you guys will accept.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2247/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2247/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2247",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2247",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2247.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2247.patch",
"merged_at": "2021-10-06T19:41:02Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2248
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2248/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2248/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2248/events
|
https://github.com/materialsproject/pymatgen/issues/2248
| 994,798,750
|
MDU6SXNzdWU5OTQ3OTg3NTA=
| 2,248
|
Chemical system depends on assigned charges
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Definitely not desired behaviour!\r\n\r\nThat was my code so mea culpa. Was there a reason you closed this issue?",
"It was related to much hash concerns about `Site` and there the takeaway for me was that because two compositions are not equal if they have assigned charges then it doesn't matter that the hashes aren't equal #2235. The chemical system is related to this because a few months back I needed a hash with less collisions for my `PatchedPhaseDiagram` and so the hash for `Composition` was changed to `hash(frozenset(Composition._data.keys()))` as it was barely slower than the sum of species charges but a lot more discriminating. In the end because so few sites are disordered and the discriminative hash is ~10% slower I decided against the PR for `Site`.\r\n\r\nI was going to submit a PR later to add this example to the docstring:\r\n\r\n```python\r\n>>> Composition({\"Si4+\": 1, \"O2-\":2}).chemical_system\r\n'O2--Si4+'\r\n>>> Composition({\"Si\": 1, \"O\":2}).chemical_system\r\n'O-Si'\r\n``` ",
"A docstring improvement would be welcome (given that this is the current behaviour), but I do think we should change this to just return the elements. I'm less concerned with the hashing and more with expected/correct behaviour.",
"```python\r\n @property\r\n def chemical_system(self) -> str:\r\n \"\"\"\r\n Get the chemical system of a Composition, for example \"O-Si\" for\r\n SiO2. Chemical system is a string of a list of elements\r\n sorted alphabetically and joined by dashes, by convention for use\r\n in database keys.\r\n \"\"\"\r\n return \"-\".join(sorted([el.symbol for el in self.elements]))\r\n```\r\n\r\ngives\r\n\r\n```python\r\nPython 3.7.10 (default, Jun 4 2021, 14:48:32) \r\n[GCC 7.5.0] :: Anaconda, Inc. on linux\r\nType \"help\", \"copyright\", \"credits\" or \"license\" for more information.\r\n>>> from pymatgen.core.composition import Composition\r\n>>> Composition({\"Si\": 1, \"O\":2}).chemical_system\r\n'O-Si'\r\n>>> Composition({\"Si4+\": 1, \"O2-\":2}).chemical_system\r\n'O-Si'\r\n```\r\n\r\nhaven't tested speed but `Species.__str__` accesses symbol so it shouldn't be slower. \r\n\r\nIf happy with this I can make PR and add a test for charged composition chemical systems?",
"I think we should keep this definition of chemsys. We either treat Species as distinct or not by default. Pymatgen treats them as different. And there are instances where we need to define Fe2+-Fe3+-O as opposed to just Fe-O system. If we want to ignore oxidation states, we can make it a different property or method.\r\n",
"would it not make more sense to update `chemical_system` to match it's current docstring and then add another option for the `species system` @mkhorton has already said this definition was a bug? \r\n\r\nalso in the species system just as a matter to ease of manipulation using regex the use of dash as a deliminator seems like a poor choice as we have things like `\"O2--Cu2+\".split(deliminator) = [\"O2\", \"\", \"Cu2+\"]` which could cause hidden errors if like me you think the current behaviour is unexpected."
] | 2021-09-13T11:55:21
| 2021-10-07T07:28:22
|
2021-10-07T07:28:22Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
```python
>>> from pymatgen.core.composition import Composition
>>> Composition({"Fe3+": 2, "O2-":3}).chemical_system
'Fe3+-O2-'
>>> Composition({"Fe": 2, "O":3}).chemical_system
'Fe-O'
>>> Composition({"Fe3+": 2, "Fe2+": 1, "O2-":4}).chemical_system
'Fe2+-Fe3+-O2-'
```
Not sure if this is a bug or not, it not the behaviour i would have expected so if planned I will look at doing a docstring improvement.
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2248/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2248/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2249
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2249/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2249/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2249/events
|
https://github.com/materialsproject/pymatgen/pull/2249
| 995,904,699
|
PR_kwDOACgets4ruYz_
| 2,249
|
Update functionality of composition.chemical_system to match expected behaviour
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/42858314)\n\nCoverage decreased (-0.6%) to 83.161% when pulling **6c571fc272554b0e8f07e77099f92e90a7085a02 on CompRhys:chemsystem** into **b8b27c6fa3236f032581aa82f668b25a78aa3b57 on materialsproject:master**.\n",
"Thanks!"
] | 2021-09-14T11:26:10
| 2021-10-13T08:59:44
|
2021-10-06T19:39:26Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
fix as suggested in #2248. Added additional test.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2249/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2249/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2249",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2249",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2249.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2249.patch",
"merged_at": "2021-10-06T19:39:26Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2250
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2250/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2250/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2250/events
|
https://github.com/materialsproject/pymatgen/issues/2250
| 996,205,727
|
I_kwDOACgets47YOSf
| 2,250
|
A bug in pointgroupanalyzer?
|
{
"login": "qzhu2017",
"id": 29445366,
"node_id": "MDQ6VXNlcjI5NDQ1MzY2",
"avatar_url": "https://avatars.githubusercontent.com/u/29445366?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/qzhu2017",
"html_url": "https://github.com/qzhu2017",
"followers_url": "https://api.github.com/users/qzhu2017/followers",
"following_url": "https://api.github.com/users/qzhu2017/following{/other_user}",
"gists_url": "https://api.github.com/users/qzhu2017/gists{/gist_id}",
"starred_url": "https://api.github.com/users/qzhu2017/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/qzhu2017/subscriptions",
"organizations_url": "https://api.github.com/users/qzhu2017/orgs",
"repos_url": "https://api.github.com/users/qzhu2017/repos",
"events_url": "https://api.github.com/users/qzhu2017/events{/privacy}",
"received_events_url": "https://api.github.com/users/qzhu2017/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"The relevant tolerance in this case is the `eigen_tolerance`, which determines whether it is classified as a symmetric top or not. Setting `eigen_tolerance=1e-3` gives C2 as expected. "
] | 2021-09-14T16:22:50
| 2021-09-14T16:32:15
|
2021-09-14T16:32:15Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
I am trying to use the `PointGroupAnalyzer` to perform some symmetry analysis on some cluster. However, it does not work as I expected for some cases.
**To Reproduce**
```python
from pymatgen.core import Molecule
from pymatgen.symmetry.analyzer import PointGroupAnalyzer
xyz = """15
C7 N8
C -0.000000 -0.000002 1.048832
C 1.008447 0.765201 0.191851
C -1.008448 -0.765196 0.191849
C 2.019657 1.482290 1.088066
C -2.019653 -1.482295 1.088058
N 2.932883 2.279851 0.284256
N -2.932877 -2.279853 0.284246
C 3.030291 3.615607 0.223535
C -3.030270 -3.615608 0.223507
N 3.994395 3.952771 -0.650098
N -3.994383 -3.952767 -0.650120
N 4.468392 2.802027 -1.112496
N -4.468410 -2.802025 -1.112481
N 3.829808 1.789427 -0.549517
N -3.829830 -1.789428 -0.549487
"""
m = Molecule.from_str(xyz, fmt='xyz')
for tol in [0.01, 0.1, 0.5]:
pga = PointGroupAnalyzer(m, 0.01);
print(tol, pga.sch_symbol)
```
```
$ python bug.py
0.01 C1
0.1 C1
0.5 C1
```
**Expected behavior**
The above example clearly has a C2 symmetry along z-axis, i.e., (x, y, z) -> (-x, -y, z),
However, I tried to used different tolerance, it always returns `C1` symmetry.
**Desktop (please complete the following information):**
I am using pymatgen-2022.0.12 on Mac-os.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2250/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2250/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2251
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2251/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2251/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2251/events
|
https://github.com/materialsproject/pymatgen/issues/2251
| 996,392,593
|
I_kwDOACgets47Y76R
| 2,251
|
Outcar parsing error from pymatgen
|
{
"login": "yurisanspeur",
"id": 72325364,
"node_id": "MDQ6VXNlcjcyMzI1MzY0",
"avatar_url": "https://avatars.githubusercontent.com/u/72325364?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/yurisanspeur",
"html_url": "https://github.com/yurisanspeur",
"followers_url": "https://api.github.com/users/yurisanspeur/followers",
"following_url": "https://api.github.com/users/yurisanspeur/following{/other_user}",
"gists_url": "https://api.github.com/users/yurisanspeur/gists{/gist_id}",
"starred_url": "https://api.github.com/users/yurisanspeur/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/yurisanspeur/subscriptions",
"organizations_url": "https://api.github.com/users/yurisanspeur/orgs",
"repos_url": "https://api.github.com/users/yurisanspeur/repos",
"events_url": "https://api.github.com/users/yurisanspeur/events{/privacy}",
"received_events_url": "https://api.github.com/users/yurisanspeur/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-09-14T20:11:33
| 2021-09-14T20:11:33
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
I am relaxing an oxide slab e.g. IrO2 with ISPIN=2 LORBIT=11 and with the WalltimeHandler from Custodian switched on. The vasp command that I am using is mpirun --map-by core -np 32 /opt/vasp.5.4.4.pl2/bin/vasp_std. Sometimes I will receive a parsing error from pymatgen when it is trying to parse the OUTCAR with examples of the stacktraces below:
"Traceback (most recent call last):\n File \"/opt/conda/lib/python3.7/site-packages/fireworks/core/rocket.py\", line 262, in run\n m_action = t.run_task(my_spec)\n File \"/opt/conda/lib/python3.7/site-packages/atomate/vasp/firetasks/run_calc.py\", line 210, in run_task\n c.run()\n File \"/opt/conda/lib/python3.7/site-packages/custodian/custodian.py\", line 368, in run\n self._run_job(job_n, job)\n File \"/opt/conda/lib/python3.7/site-packages/custodian/custodian.py\", line 461, in _run_job\n has_error = self._do_check(self.monitors, terminate)\n File \"/opt/conda/lib/python3.7/site-packages/custodian/custodian.py\", line 634, in _do_check\n if h.check():\n File \"/opt/conda/lib/python3.7/site-packages/custodian/vasp/handlers.py\", line 1417, in check\n outcar = Outcar(\"OUTCAR\")\n File \"/opt/conda/lib/python3.7/site-packages/pymatgen/io/vasp/outputs.py\", line 1762, in __init__\n toks = [float(i) for i in re.findall(r\"[\\d\\.\\-]+\", clean)]\n File \"/opt/conda/lib/python3.7/site-packages/pymatgen/io/vasp/outputs.py\", line 1762, in <listcomp>\n toks = [float(i) for i in re.findall(r\"[\\d\\.\\-]+\", clean)]\nValueError: could not convert string to float: '-'\n"
"Traceback (most recent call last):\n File \"/opt/conda/lib/python3.7/site-packages/fireworks/core/rocket.py\", line 262, in run\n m_action = t.run_task(my_spec)\n File \"/opt/conda/lib/python3.7/site-packages/atomate/vasp/firetasks/run_calc.py\", line 210, in run_task\n c.run()\n File \"/opt/conda/lib/python3.7/site-packages/custodian/custodian.py\", line 368, in run\n self._run_job(job_n, job)\n File \"/opt/conda/lib/python3.7/site-packages/custodian/custodian.py\", line 479, in _run_job\n has_error = self._do_check(self.handlers)\n File \"/opt/conda/lib/python3.7/site-packages/custodian/custodian.py\", line 634, in _do_check\n if h.check():\n File \"/opt/conda/lib/python3.7/site-packages/custodian/vasp/handlers.py\", line 1417, in check\n outcar = Outcar(\"OUTCAR\")\n File \"/opt/conda/lib/python3.7/site-packages/pymatgen/io/vasp/outputs.py\", line 1844, in __init__\n r\"maximum and minimum number of plane-waves\",\n File \"/opt/conda/lib/python3.7/site-packages/pymatgen/io/vasp/outputs.py\", line 2056, in read_table_pattern\n retained_data = tables[-1]\nIndexError: list index out of range\n"
"Traceback (most recent call last):\n File \"/opt/conda/lib/python3.8/site-packages/fireworks/core/rocket.py\", line 262, in run\n m_action = t.run_task(my_spec)\n File \"/opt/conda/lib/python3.8/site-packages/atomate/vasp/firetasks/run_calc.py\", line 299, in run_task\n c.run()\n File \"/opt/conda/lib/python3.8/site-packages/custodian/custodian.py\", line 368, in run\n self._run_job(job_n, job)\n File \"/opt/conda/lib/python3.8/site-packages/custodian/custodian.py\", line 461, in _run_job\n has_error = self._do_check(self.monitors, terminate)\n File \"/opt/conda/lib/python3.8/site-packages/custodian/custodian.py\", line 634, in _do_check\n if h.check():\n File \"/opt/conda/lib/python3.8/site-packages/custodian/vasp/handlers.py\", line 1417, in check\n outcar = Outcar(\"OUTCAR\")\n File \"/opt/conda/lib/python3.8/site-packages/pymatgen/io/vasp/outputs.py\", line 1876, in __init__\n self.data[\"nplwv\"] = [[int(self.data[\"nplwv\"][0][0])]]\nIndexError: list index out of range\n"
Provide any example files that are needed to reproduce the error,
especially if the bug pertains to parsing of a file.
[OUTCAR.gz](https://github.com/materialsproject/pymatgen/files/7164858/OUTCAR.gz)
Notice also that the parsing error seems to happen when the magnetic moments are being written to the OUTCAR file.
**Expected behavior**
There should not be a parsing error from pymatgen/custodian
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2251/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2251/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2252
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2252/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2252/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2252/events
|
https://github.com/materialsproject/pymatgen/issues/2252
| 997,719,116
|
I_kwDOACgets47d_xM
| 2,252
|
convert from pymatgen Molecule/Structure into SMILES/SELFIES and vice versa
|
{
"login": "turmeric-blend",
"id": 62788745,
"node_id": "MDQ6VXNlcjYyNzg4NzQ1",
"avatar_url": "https://avatars.githubusercontent.com/u/62788745?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/turmeric-blend",
"html_url": "https://github.com/turmeric-blend",
"followers_url": "https://api.github.com/users/turmeric-blend/followers",
"following_url": "https://api.github.com/users/turmeric-blend/following{/other_user}",
"gists_url": "https://api.github.com/users/turmeric-blend/gists{/gist_id}",
"starred_url": "https://api.github.com/users/turmeric-blend/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/turmeric-blend/subscriptions",
"organizations_url": "https://api.github.com/users/turmeric-blend/orgs",
"repos_url": "https://api.github.com/users/turmeric-blend/repos",
"events_url": "https://api.github.com/users/turmeric-blend/events{/privacy}",
"received_events_url": "https://api.github.com/users/turmeric-blend/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Converting to SMILES is already possible. That can be done via the babel interface in pymatgen.io.babel. As for SELFIES, I am happy for it to be included if someone implements it.",
"Hi, could you give a small example of how to use pymatgen.io.babel to convert `pymatgen` Structure/Molecule to SMILES?\r\n\r\nI am trying to initiate `BabelMolAdaptor()`, but I am getting this error:\r\n\r\n> RuntimeError: BabelMolAdaptor requires openbabel to be installed with Python bindings. Please get it at http://openbabel.org (version >=3.0.0).\r\n\r\nI am not sure how to properly install this.",
"openbabel is available on the conda-forge channel via `conda install -c conda-forge openbabel`. This is documented in pymatgen.org. Pls refer to that if you have other questions.",
"Once openbabel is installed, the smiles is as easy as \r\n\r\n```python\r\nfrom pymatgen.core import Molecule\r\nm = Molecule.from_file(\"molecules/Carbon_Disulfide.xyz\")\r\nprint(m.to(fmt=\"SMILES\"))\r\n```",
"I see, I guess that SMILES string can then be converted into SELFIES using:\r\n\r\n```\r\nimport selfies\r\nselfies_string = selfies.encoder(smiles_string)\r\nprint(selfies_string)\r\n```\r\n\r\nThanks for your help!"
] | 2021-09-16T03:28:41
| 2021-09-16T15:36:01
|
2021-09-16T15:36:01Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Is your feature request related to a problem? Please describe.**
It would be a useful feature to have, especially if this could be done for [SELFIES](https://github.com/aspuru-guzik-group/selfies). SELFIES is a new more robust version of SMILES. It is useful because it might be good to explore machine learning on string-based structures as they have different ease of computation or might learn different compared to structure-based molecules such as pymatgen.
**Describe the solution you'd like**
convert from pymatgen's `Structure`/`Molecule` to SELFIES (ideally)
otherwise
convert from pymatgen's `Structure`/`Molecule` to SMILES
**Additional context**
note the SELFIES github page has a feature to convert SMILES TO SELFIES and vice versa
|
{
"login": "turmeric-blend",
"id": 62788745,
"node_id": "MDQ6VXNlcjYyNzg4NzQ1",
"avatar_url": "https://avatars.githubusercontent.com/u/62788745?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/turmeric-blend",
"html_url": "https://github.com/turmeric-blend",
"followers_url": "https://api.github.com/users/turmeric-blend/followers",
"following_url": "https://api.github.com/users/turmeric-blend/following{/other_user}",
"gists_url": "https://api.github.com/users/turmeric-blend/gists{/gist_id}",
"starred_url": "https://api.github.com/users/turmeric-blend/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/turmeric-blend/subscriptions",
"organizations_url": "https://api.github.com/users/turmeric-blend/orgs",
"repos_url": "https://api.github.com/users/turmeric-blend/repos",
"events_url": "https://api.github.com/users/turmeric-blend/events{/privacy}",
"received_events_url": "https://api.github.com/users/turmeric-blend/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2252/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2252/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2253
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2253/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2253/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2253/events
|
https://github.com/materialsproject/pymatgen/issues/2253
| 998,194,624
|
I_kwDOACgets47fz3A
| 2,253
|
get_pointgroup function takes long time?
|
{
"login": "qzhu2017",
"id": 29445366,
"node_id": "MDQ6VXNlcjI5NDQ1MzY2",
"avatar_url": "https://avatars.githubusercontent.com/u/29445366?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/qzhu2017",
"html_url": "https://github.com/qzhu2017",
"followers_url": "https://api.github.com/users/qzhu2017/followers",
"following_url": "https://api.github.com/users/qzhu2017/following{/other_user}",
"gists_url": "https://api.github.com/users/qzhu2017/gists{/gist_id}",
"starred_url": "https://api.github.com/users/qzhu2017/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/qzhu2017/subscriptions",
"organizations_url": "https://api.github.com/users/qzhu2017/orgs",
"repos_url": "https://api.github.com/users/qzhu2017/repos",
"events_url": "https://api.github.com/users/qzhu2017/events{/privacy}",
"received_events_url": "https://api.github.com/users/qzhu2017/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This can be resolved by setting matrix_tolerance=0.2 in your PointGroupAnalyzer. Unfortunately, if too small a tolerance is set, the code just goes on... I will add a warning message."
] | 2021-09-16T13:08:42
| 2021-10-27T15:52:19
|
2021-10-27T15:52:19Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
I am trying to use the PointGroupAnalyzer to parse the molecular symmetry. It works well in general. However, I sometimes encounter some cases in which the get_pointgroup function takes forever.
**To Reproduce**
```python
from pymatgen.symmetry.analyzer import PointGroupAnalyzer
from pymatgen.core import Molecule
xyz="""10
C4 N6
C 1.887551 0.000000 -0.075956
N 2.605011 0.000000 -1.145031
N 1.722945 0.000000 -2.225207
C 0.522780 0.000000 -1.755613
N 0.567925 0.000000 -0.391239
N -0.515557 0.000000 0.463576
C -1.184399 1.083249 0.935390
N -2.210929 0.702391 1.629345
N -2.210929 -0.702391 1.629345
C -1.184399 -1.083249 0.935390
"""
m = Molecule.from_str(xyz, fmt='xyz')
pga = PointGroupAnalyzer(m, 0.25, eigen_tolerance=1e-3)
print(pga.sch_symbol)
pg = pga.get_pointgroup() #This step sometime will take forever
```
**Expected behavior**
If you execute the above script, you will find that the code will get stuck in the last step.
**Desktop (please complete the following information):**
- OS: Mac
- Version: 2022.0.13
**Additional context**
Add any other context about the problem here.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2253/reactions",
"total_count": 1,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 1
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2253/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2254
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2254/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2254/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2254/events
|
https://github.com/materialsproject/pymatgen/issues/2254
| 1,011,029,712
|
I_kwDOACgets48QxbQ
| 2,254
|
Update AFLOW prototype library
|
{
"login": "jacksund",
"id": 47992949,
"node_id": "MDQ6VXNlcjQ3OTkyOTQ5",
"avatar_url": "https://avatars.githubusercontent.com/u/47992949?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jacksund",
"html_url": "https://github.com/jacksund",
"followers_url": "https://api.github.com/users/jacksund/followers",
"following_url": "https://api.github.com/users/jacksund/following{/other_user}",
"gists_url": "https://api.github.com/users/jacksund/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jacksund/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jacksund/subscriptions",
"organizations_url": "https://api.github.com/users/jacksund/orgs",
"repos_url": "https://api.github.com/users/jacksund/repos",
"events_url": "https://api.github.com/users/jacksund/events{/privacy}",
"received_events_url": "https://api.github.com/users/jacksund/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I reached out to the AFLOW team, and they don't want archives of the full library unless you purchase through them ([here's the thread](https://groups.io/g/aflow/topic/85959008)). \r\n\r\nI'll close this request because these prototypes cannot be updated to the full library😢",
"Hahahaha"
] | 2021-09-29T14:21:51
| 2021-09-29T22:25:00
|
2021-09-29T22:22:26Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Would someone be able to update the `aflow_prototypes.json` in the `pymatgen.analysis` module?
Currently it only includes 288 prototypes (so Part 1 of their library), but they are now at 1,100 prototypes (thanks to Parts 2 and 3).
I don't use the aflow software, so I'm not sure how to download these structures and update them myself. I was hoping someone here could update it with a quick script!
|
{
"login": "jacksund",
"id": 47992949,
"node_id": "MDQ6VXNlcjQ3OTkyOTQ5",
"avatar_url": "https://avatars.githubusercontent.com/u/47992949?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jacksund",
"html_url": "https://github.com/jacksund",
"followers_url": "https://api.github.com/users/jacksund/followers",
"following_url": "https://api.github.com/users/jacksund/following{/other_user}",
"gists_url": "https://api.github.com/users/jacksund/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jacksund/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jacksund/subscriptions",
"organizations_url": "https://api.github.com/users/jacksund/orgs",
"repos_url": "https://api.github.com/users/jacksund/repos",
"events_url": "https://api.github.com/users/jacksund/events{/privacy}",
"received_events_url": "https://api.github.com/users/jacksund/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2254/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2254/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2255
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2255/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2255/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2255/events
|
https://github.com/materialsproject/pymatgen/issues/2255
| 1,013,531,871
|
I_kwDOACgets48aUTf
| 2,255
|
Pymatgen Zeo++ installation help
|
{
"login": "cwaitt",
"id": 36715383,
"node_id": "MDQ6VXNlcjM2NzE1Mzgz",
"avatar_url": "https://avatars.githubusercontent.com/u/36715383?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/cwaitt",
"html_url": "https://github.com/cwaitt",
"followers_url": "https://api.github.com/users/cwaitt/followers",
"following_url": "https://api.github.com/users/cwaitt/following{/other_user}",
"gists_url": "https://api.github.com/users/cwaitt/gists{/gist_id}",
"starred_url": "https://api.github.com/users/cwaitt/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/cwaitt/subscriptions",
"organizations_url": "https://api.github.com/users/cwaitt/orgs",
"repos_url": "https://api.github.com/users/cwaitt/repos",
"events_url": "https://api.github.com/users/cwaitt/events{/privacy}",
"received_events_url": "https://api.github.com/users/cwaitt/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 5322879170,
"node_id": "LA_kwDOACgets8AAAABPUSwwg",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/help",
"name": "help",
"color": "5ED16E",
"default": false,
"description": "Q&A support-style issues"
},
{
"id": 5456916296,
"node_id": "LA_kwDOACgets8AAAABRUHvSA",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/docs",
"name": "docs",
"color": "35B067",
"default": false,
"description": "Documentation, examples, user guides"
}
] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Zeo++ and its related tools have moved around a bit over time. The current version of Zeo++ can be found [here](http://www.zeoplusplus.org/download.html) and Voro++ [here](https://github.com/chr1shr/voro.git). However, I'm pretty sure that Zeo++ ships with everything needed for installation, including a compatible version of Voro++.",
"@mkhorton -- looks like I closed this one in https://github.com/materialsproject/pymatgen/pull/2400.",
"@janosh: This can be closed based on my prior PR."
] | 2021-10-01T16:18:08
| 2023-06-03T02:25:52
|
2023-06-03T02:25:52Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I am following the directions on the pymatgen documentation site on installing and setting up Zeo++ and Voro++ (https://pymatgen.org/pymatgen.io.zeopp.html#zeo-installation-steps). When I do the svn chackout command I get an "Access denied" error. I found an older Bug back from 2016 that offered a solution but when I tried that it still did not work. any guidance will be much appreciated!
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2255/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2255/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2256
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2256/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2256/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2256/events
|
https://github.com/materialsproject/pymatgen/pull/2256
| 1,015,808,197
|
PR_kwDOACgets4sq6qF
| 2,256
|
Type hint and correct documentation of Structure.remove_site_properties
|
{
"login": "kmu",
"id": 3815357,
"node_id": "MDQ6VXNlcjM4MTUzNTc=",
"avatar_url": "https://avatars.githubusercontent.com/u/3815357?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/kmu",
"html_url": "https://github.com/kmu",
"followers_url": "https://api.github.com/users/kmu/followers",
"following_url": "https://api.github.com/users/kmu/following{/other_user}",
"gists_url": "https://api.github.com/users/kmu/gists{/gist_id}",
"starred_url": "https://api.github.com/users/kmu/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/kmu/subscriptions",
"organizations_url": "https://api.github.com/users/kmu/orgs",
"repos_url": "https://api.github.com/users/kmu/repos",
"events_url": "https://api.github.com/users/kmu/events{/privacy}",
"received_events_url": "https://api.github.com/users/kmu/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/43268100)\n\nCoverage decreased (-0.6%) to 83.134% when pulling **27ce06b72049a2e76e3e206a3c4e99c5d0029a34 on kmu:master** into **5011c0a6e1dc48380ed2d7a29ec6b0561f60be3b on materialsproject:master**.\n",
"Thanks."
] | 2021-10-05T02:12:18
| 2021-10-05T13:39:43
|
2021-10-05T13:39:39Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Type hint and correct documentation of `Structure.remove_site_properties`
## Additional dependencies introduced (if any)
None
## TODO (if any)
None
* Feature 1 supports A, but not B.
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [ ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [ ] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2256/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2256/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2256",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2256",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2256.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2256.patch",
"merged_at": "2021-10-05T13:39:39Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2257
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2257/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2257/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2257/events
|
https://github.com/materialsproject/pymatgen/pull/2257
| 1,021,509,739
|
PR_kwDOACgets4s9tpW
| 2,257
|
InsertionElectrode bug fix & doc clarification
|
{
"login": "acrutt",
"id": 42151899,
"node_id": "MDQ6VXNlcjQyMTUxODk5",
"avatar_url": "https://avatars.githubusercontent.com/u/42151899?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/acrutt",
"html_url": "https://github.com/acrutt",
"followers_url": "https://api.github.com/users/acrutt/followers",
"following_url": "https://api.github.com/users/acrutt/following{/other_user}",
"gists_url": "https://api.github.com/users/acrutt/gists{/gist_id}",
"starred_url": "https://api.github.com/users/acrutt/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/acrutt/subscriptions",
"organizations_url": "https://api.github.com/users/acrutt/orgs",
"repos_url": "https://api.github.com/users/acrutt/repos",
"events_url": "https://api.github.com/users/acrutt/events{/privacy}",
"received_events_url": "https://api.github.com/users/acrutt/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/44306257)\n\nCoverage decreased (-0.6%) to 83.111% when pulling **59f1b2e7439f20588bee501aa96c711ef59d1c30 on acrutt:master** into **9276567df1bf01eea473933688975003449dc5e5 on materialsproject:master**.\n",
"Note for future reference on the appropriate value for the terminal element entries when constructing the phase diagram. This value cannot be set infinitely high otherwise there will be precision errors in constructing the ConvexHull. \r\n\r\n[http://www.qhull.org/html/qh-impre.html](http://www.qhull.org/html/qh-impre.html)\r\n\r\n> Your input coordinates start with the same five or more digits (i.e., it is shifted relative to the origin). This reduces the available precision.\r\n\r\nTherefore the terminal elements need to have an energy comparable to the provided battery material energies. 10 can be an appropriate value while 1e9 is too big. If the value is set too big (values greater than 1e6), pymatgen's tests will fail due to incorrect convex hull construction that results in misidentification of the stable battery entries.",
"Thanks @acrutt and for providing additional context here",
"@mkhorton I believe this PR is ready to merge. Ultimately the changes are now limited to some documentation clarifications. Let me know if any other work is needed!",
"Thanks!"
] | 2021-10-08T22:24:26
| 2021-11-16T17:37:14
|
2021-11-16T17:36:50Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
The InsertionElectrode.from_entries() method can use either ComputedEntries or ComputedStructureEntries - this has been added to the documentation. Setting strip_structures=True only works with ComputedStructureEntries so this has also been clarified.
In constructing the PhaseDiagram to compare the energies of the provided battery entries, the element energies were not always set high enough if there was a large energy difference between the empty host material and intercalated material. As a result, no intercalated entry was found to be stable and voltage pairs could not be formed. The factor for increasing the element_energy was set higher to 1e9 to avoid this problem.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2257/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2257/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2257",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2257",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2257.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2257.patch",
"merged_at": "2021-11-16T17:36:50Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2258
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2258/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2258/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2258/events
|
https://github.com/materialsproject/pymatgen/issues/2258
| 1,021,527,822
|
I_kwDOACgets4840cO
| 2,258
|
LammpsData.from_file changes data type for some dihedral coeffs from int to float
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Just trying to give some insight here:\r\nI believe this is because in Pandas, NaN (in `coeff6` here) will automatically upscale the rest of the data in that column to float. \r\nThe `NaN` is from when we don't have a coefficient (the first 3 rows here)\r\n\r\nA proposed solution to this issue would be to use `fillna` here or to use `pd.convert_dtypes()` which should automatically choose the best datatype to support `Na` in the dataframe ",
"@rkingsbury Are you using a hybrid Dihedral Coeffs set where line 409-411 and line 412-417 have different Dihedral style? If so, I think you might need to specify that style instead of `1 `and `2`. LammpsData will help make sure certain coeff has the correct type according to the style.",
"> @rkingsbury Are you using a hybrid Dihedral Coeffs set where line 409-411 and line 412-417 have different Dihedral style? If so, I think you might need to specify that style instead of `1 `and `2`. LammpsData will help make sure certain coeff has the correct type according to the style.\r\n\r\nNo, as best I can tell all dihedrals are supposed to have the style `fourier` according to the output files generated by antechamber and moltemplate. However, not all dihedrals have the same number of coefficients. This is the raw output from the data file generated by moltemplate for one of the molecules, which corresponds to the `LammpsData` shown above.\r\n\r\n```\r\nDihedral Coeffs\r\n\r\n1 1 0.155555555556 3 0.0 # JCC,7,(1986),230\r\n2 1 0.166666666667 3 0.0 # JCC,7,(1986),230\r\n3 1 0.383333333333 3 0.0 # JCC,7,(1986),230\r\n4 2 0.16 3 0.0 0.25 1 0.0 # Junmei et al, 1999\r\n5 2 0.383 3 0.0 0.1 2 180.0 #\r\n6 2 0.144 3 0.0 1.175 2 0.0 # parm98, TC,PC,PAK\r\n7 2 0.144 3 0.0 1.175 2 0.0 # parm98, TC,PC,PAK\r\n8 2 0.0 3 0.0 0.25 1 0.0 # Junmei et al, 1999\r\n9 2 0.0 3 0.0 0.25 1 0.0 # Junmei et al, 1999\r\n```\r\n\r\nThat said, the input files generated by moltemplate do list the style as `hybrid fourier` even though every individual coefficient is assigned the fourier style",
"OK, after further investigation, the issues i that the [fourier style](https://docs.lammps.org/dihedral_fourier.html) can take a variable number of parameters. The integer in the second column above indicates how many groups of 3 parameters there are. So in the first 3 rows, the 2nd column is one and I need only 3 parameters. For subsequent rows, the second column is 2 and I need 2 groups of 3 parameters. In each group of parameters, the middle number is an integer.\r\n\r\nI tried to work around this bug by explicitly supplying zeros for the missing parameters in the first 3 rows in my data file, but LAMMPS then fails with\r\n\r\n```\r\nERROR: Incorrect number of arguments for dihedral coefficients (src/USER-MISC/dihedral_fourier.cpp:315)\r\n```\r\n\r\nSo I would say this is a bug, but I don't see an elegant way to fix it.",
"@rkingsbury Thanks for the update. OK, in that case, this line will remove all the `nan` in the output str representation. https://github.com/materialsproject/pymatgen/blob/d9d342b69dc838aeb046aaefbca6ace6de985e70/pymatgen/io/lammps/data.py#L479 So I don't think storing an empty cell as NaN will cause a very big problem. \r\n\r\nThe only issue here is that, without explicitly feeding the `fourier` keyword, pymatgen won't know what is the correct data type to output. But if you add that as in this modified file [PEG400_with_FF.data1.txt](https://github.com/materialsproject/pymatgen/files/7428355/PEG400_with_FF.data1.txt), pymatgen will be able to tell the correct data type as I mentioned. Adding `fourier` explicitly will also make sure that the output datatype strictly complies with the style requirement even if your input file has the wrong data type.\r\n\r\nThis is indeed a corner case where coeffs have different numbers of columns even if they are the same style. The automatic datatype correction as mentioned by @druyang does not work here because it also converts `180.0` to `180`. So I think the safest way here is to just add that style column and let pymatgen determine the correct data type post hoc.\r\n\r\n\r\n",
"> @rkingsbury Thanks for the update. OK, in that case, this line will remove all the `nan` in the output str representation.\r\n> \r\n> https://github.com/materialsproject/pymatgen/blob/d9d342b69dc838aeb046aaefbca6ace6de985e70/pymatgen/io/lammps/data.py#L479\r\n> So I don't think storing an empty cell as NaN will cause a very big problem.\r\n> \r\n> The only issue here is that, without explicitly feeding the `fourier` keyword, pymatgen won't know what is the correct data type to output. But if you add that as in this modified file [PEG400_with_FF.data1.txt](https://github.com/materialsproject/pymatgen/files/7428355/PEG400_with_FF.data1.txt), pymatgen will be able to tell the correct data type as I mentioned. Adding `fourier` explicitly will also make sure that the output datatype strictly complies with the style requirement even if your input file has the wrong data type.\r\n> \r\n> This is indeed a corner case where coeffs have different numbers of columns even if they are the same style. The automatic datatype correction as mentioned by @druyang does not work here because it also converts `180.0` to `180`. So I think the safest way here is to just add that style column and let pymatgen determine the correct data type post hoc.\r\n\r\nThanks @htz1992213 ; your suggestion worked well."
] | 2021-10-08T23:11:12
| 2021-11-15T22:25:35
|
2021-11-15T22:25:35Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
When reading a file containing a `Dihedral Coeffs` section, `LammpsData.from_file` sometimes change the datatype of the `coeff6` column from `int` to `float`. This is a problem because LAMMPS is very particular about the data types for dihedral coefficients. Having a float instead of an int can result in an error like
```
ERROR: Expected integer parameter instead of '1.0' in input script or data file
```
**To Reproduce**
Steps to reproduce the behavior:
1. Download the data file [PEG400_with_FF.data.txt](https://github.com/materialsproject/pymatgen/files/7314582/PEG400_with_FF.data.txt)
2. Read in with `LammpsData.from_file(<file>)`
3. Inspect the `.force_field["Dihedral Coeffs"]` attribute. Notice that the coeff6 column is populated with `float` even though the corresponding values are given as `int` in the data file.
```
'Dihedral Coeffs': coeff1 coeff2 coeff3 coeff4 coeff5 coeff6 coeff7
1 1 0.155556 3 0.0 NaN NaN NaN
2 1 0.166667 3 0.0 NaN NaN NaN
3 1 0.383333 3 0.0 NaN NaN NaN
4 2 0.160000 3 0.0 0.250 1.0 0.0
5 2 0.383000 3 0.0 0.100 2.0 180.0
6 2 0.144000 3 0.0 1.175 2.0 0.0
7 2 0.144000 3 0.0 1.175 2.0 0.0
8 2 0.000000 3 0.0 0.250 1.0 0.0
9 2 0.000000 3 0.0 0.250 1.0 0.0}
```
**Additional context**
I believe the relevant line is [here](https://github.com/materialsproject/pymatgen/blob/84561a916b984de0ff927508101340bf5e871ce0/pymatgen/io/lammps/data.py#L711). It's unclear to me how or where the data types get set or why this issue only seems to affect the `coeff6` column in my example (`coeff1` and `coeff3` are also provided as `int` and their types are preserved).
Flagging @htz1992213 who may have some perspective on this.
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2258/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2258/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2259
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2259/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2259/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2259/events
|
https://github.com/materialsproject/pymatgen/issues/2259
| 1,023,357,412
|
I_kwDOACgets48_zHk
| 2,259
|
Parsing error in vasprun.xml using Outcar method (Vasp 6)
|
{
"login": "pjf295",
"id": 69720428,
"node_id": "MDQ6VXNlcjY5NzIwNDI4",
"avatar_url": "https://avatars.githubusercontent.com/u/69720428?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/pjf295",
"html_url": "https://github.com/pjf295",
"followers_url": "https://api.github.com/users/pjf295/followers",
"following_url": "https://api.github.com/users/pjf295/following{/other_user}",
"gists_url": "https://api.github.com/users/pjf295/gists{/gist_id}",
"starred_url": "https://api.github.com/users/pjf295/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/pjf295/subscriptions",
"organizations_url": "https://api.github.com/users/pjf295/orgs",
"repos_url": "https://api.github.com/users/pjf295/repos",
"events_url": "https://api.github.com/users/pjf295/events{/privacy}",
"received_events_url": "https://api.github.com/users/pjf295/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 3093102097,
"node_id": "MDU6TGFiZWwzMDkzMTAyMDk3",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/stale",
"name": "stale",
"color": "A99B62",
"default": false,
"description": "Abandoned or conflicting PRs and outdated issues"
},
{
"id": 5304335471,
"node_id": "LA_kwDOACgets8AAAABPCm8bw",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/io",
"name": "io",
"color": "c2e0c6",
"default": false,
"description": "Input/output functionality"
},
{
"id": 5457739150,
"node_id": "LA_kwDOACgets8AAAABRU59jg",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/vasp",
"name": "vasp",
"color": "BF4B01",
"default": false,
"description": "Vienna Ab initio Simulation Package"
}
] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@yang-ruoxi: You've done XAS calculations before, right? Do you think this issue is still relevant?\r\n\r\n@janosh ",
"Hi @pjf295, VAPS can't do XANES calculation as far as I know. the IO for XANES in pymatgen is for the code FEFF. \r\nThanks, @arosen93 for keeping track of the issue! ",
"Oops, sorry 'bout that! Of course! ",
"Sorry, let me correct myself here: it appears VASP **CAN** do XANES from 6.0 onwards. It would be great to read that output in pymatgen. Would you be interested in opening a PR for that @pjf295? ",
"I am interested in this functionality as well. Has there been any progress on it?"
] | 2021-10-12T04:44:01
| 2024-03-18T14:08:47
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
The current version of Pymatgen (2022.0.14) does not recognize XANES output from VASP 6.
**Proposed Solution:** Enable Pymatgen to read VASP 6 XANES output from either the `vasprun.xml` file or the `OUTCAR` file. Additionally, consider supporting electron-phonon coupling results.
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2259/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2259/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2260
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2260/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2260/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2260/events
|
https://github.com/materialsproject/pymatgen/issues/2260
| 1,024,889,793
|
I_kwDOACgets49FpPB
| 2,260
|
defects test_generators fails on non-amd64 architecture
|
{
"login": "drew-parsons",
"id": 26508288,
"node_id": "MDQ6VXNlcjI2NTA4Mjg4",
"avatar_url": "https://avatars.githubusercontent.com/u/26508288?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/drew-parsons",
"html_url": "https://github.com/drew-parsons",
"followers_url": "https://api.github.com/users/drew-parsons/followers",
"following_url": "https://api.github.com/users/drew-parsons/following{/other_user}",
"gists_url": "https://api.github.com/users/drew-parsons/gists{/gist_id}",
"starred_url": "https://api.github.com/users/drew-parsons/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/drew-parsons/subscriptions",
"organizations_url": "https://api.github.com/users/drew-parsons/orgs",
"repos_url": "https://api.github.com/users/drew-parsons/repos",
"events_url": "https://api.github.com/users/drew-parsons/events{/privacy}",
"received_events_url": "https://api.github.com/users/drew-parsons/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-10-13T07:28:00
| 2021-10-13T07:33:20
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
The test suite includes VoronoiInterstitialGeneratorTest.test_int_gen in pymatgen/analysis/defects/tests/test_generators.py,
https://github.com/materialsproject/pymatgen/blob/fffcc6d3edf95132fbf910ddc520d00309f1b448/pymatgen/analysis/defects/tests/test_generators.py#L107
This test run at build-time is failing any many non-amd64 architectures, e.g. for arm64,
https://buildd.debian.org/status/fetch.php?pkg=pymatgen&arch=arm64&ver=2022.0.11%2Bdfsg1-1&stamp=1633948064&raw=0
```
________________ VoronoiInterstitialGeneratorTest.test_int_gen _________________
self = <defects.tests.test_generators.VoronoiInterstitialGeneratorTest testMethod=test_int_gen>
def test_int_gen(self):
struc = PymatgenTest.get_structure("VO2")
int_gen = VoronoiInterstitialGenerator(struc, "Li")
ints = list(int_gen)
self.assertEqual(len(ints), 3)
multiplicities = [i.multiplicity for i in ints]
self.assertEqual(multiplicities, [8, 8, 4])
self.assertEqual(str(ints[0].site.specie), "Li")
self.assertEqual(str(ints[1].site.specie), "Li")
self.assertEqual(str(ints[2].site.specie), "Li")
# self.assertEqual(str(ints[3].site.specie), "Li")
> self.assertArrayAlmostEqual(ints[0].site.coords, (1.5177146, 2.6784354, 3.9481299))
.pybuild/test_python3.9/pymatgen/analysis/defects/tests/test_generators.py:107:
_ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _
actual = array([1.5177146 , 3.94812992, 2.67843543])
desired = (1.5177146, 2.6784354, 3.9481299), decimal = 7, err_msg = ''
verbose = True
@staticmethod
def assertArrayAlmostEqual(actual, desired, decimal=7, err_msg="", verbose=True):
"""
Tests if two arrays are almost equal to a tolerance. The CamelCase
naming is so that it is consistent with standard unittest methods.
"""
> return nptu.assert_almost_equal(actual, desired, decimal, err_msg, verbose)
E AssertionError:
E Arrays are not almost equal to 7 decimals
E
E Mismatched elements: 2 / 3 (66.7%)
E Max absolute difference: 1.26969452
E Max relative difference: 0.47404336
E x: array([1.5177146, 3.9481299, 2.6784354])
E y: array([1.5177146, 2.6784354, 3.9481299])
.pybuild/test_python3.9/pymatgen/util/testing.py:82: AssertionError
```
Not all non-amd64 architectures fail. mipsel and armel pass. arm64 and ppc64el fail.
A list of failing builds and build logs can be found at https://buildd.debian.org/status/package.php?p=pymatgen
**To Reproduce**
Steps to reproduce the behavior:
1. Build pymatgen on non-amd64 hardware, e.g. arm64
2. Prepare `test_files` dir e.g. from the source dir, with the compiled module installed to pybuilddir
```
testdir=build/test_py
mkdir -p $testdir
cp -a $pybuilddir $testdir
for t in `find pymatgen -name tests`; do
d=`dirname $t`;
cp -a $t $testdir/$d;
done
PMG_TEST_FILES_DIR=`pwd`/test_files PYTHONPATH=$testdir pytest-3 -v $testdir;
```
3. Launch tests
```
PMG_TEST_FILES_DIR=`pwd`/test_files pytest-3 -v $TESTDIR
```
4. See error
**Expected behavior**
The tests should pass on all architectures.
**Desktop (please complete the following information):**
- OS: Debian GNU/Linux (debian unstable)
- Version 2022.0.11
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2260/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2260/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2261
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2261/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2261/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2261/events
|
https://github.com/materialsproject/pymatgen/pull/2261
| 1,025,094,184
|
PR_kwDOACgets4tIxba
| 2,261
|
Literal-types
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"You can install `typing_extensions`, which is actually already a requirement of pymatgen if you're running in Python 3.7.\r\n\r\nThe correct way of resolving this is following this example:\r\n\r\nhttps://github.com/materialsproject/pymatgen/blob/f97868791cb8f01aadf1eb3620d1df63fb0ff49a/pymatgen/io/qchem/inputs.py#L19-L22\r\n\r\nApologies, I should have mentioned this in your earlier PR but forgot.",
"@mkhorton Thanks for that advice. Changed all the `Literal` imports from #2247.",
"pulled this branch everything seems to work",
"Thanks both!"
] | 2021-10-13T10:53:42
| 2021-10-14T19:51:45
|
2021-10-14T16:57:29Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Follow up to #2247.
@CompRhys informed me that due to #2247, Pymatgen is no longer installable in Google Colab which is locked to Python 3.7 while `typing.Literal` was only added in Python 3.8.
There's a relatively recent but quiet issue tracking the ability to select different Python versions in Colab: https://github.com/googlecolab/colabtools/issues/2165
In the meantime though, I assume Pymatgen wants to keep supporting Google Colab. I think the best solution for that would be to protect `Literal` imports with
```py
from typing import TYPE_CHECKING
if TYPE_CHECKING:
from typing import Literal
```
and then use the string form `"Literal"` in type hints. Happy to add those changes to this PR if that's acceptable.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2261/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2261/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2261",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2261",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2261.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2261.patch",
"merged_at": "2021-10-14T16:57:29Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2262
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2262/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2262/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2262/events
|
https://github.com/materialsproject/pymatgen/pull/2262
| 1,027,949,105
|
PR_kwDOACgets4tRnQ-
| 2,262
|
removed _ from beginning of arguments
|
{
"login": "jmmshn",
"id": 14003693,
"node_id": "MDQ6VXNlcjE0MDAzNjkz",
"avatar_url": "https://avatars.githubusercontent.com/u/14003693?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jmmshn",
"html_url": "https://github.com/jmmshn",
"followers_url": "https://api.github.com/users/jmmshn/followers",
"following_url": "https://api.github.com/users/jmmshn/following{/other_user}",
"gists_url": "https://api.github.com/users/jmmshn/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jmmshn/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jmmshn/subscriptions",
"organizations_url": "https://api.github.com/users/jmmshn/orgs",
"repos_url": "https://api.github.com/users/jmmshn/repos",
"events_url": "https://api.github.com/users/jmmshn/events{/privacy}",
"received_events_url": "https://api.github.com/users/jmmshn/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @jmmshn!"
] | 2021-10-16T04:30:45
| 2021-10-18T22:15:55
|
2021-10-18T22:15:49Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
variables used in constructor should not be "privated" with underscore
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2262/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2262/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2262",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2262",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2262.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2262.patch",
"merged_at": "2021-10-18T22:15:49Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2263
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2263/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2263/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2263/events
|
https://github.com/materialsproject/pymatgen/pull/2263
| 1,027,996,985
|
PR_kwDOACgets4tRu-a
| 2,263
|
Use all short-circuit rather than loop to make code more pythonic
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"> I have several small QOL changes like this on that branch and think it would be easier if they are merged separately first \r\n\r\nThanks @CompRhys -- are you planning on submitting these as separate PRs? i.e. should I merge this one, or wait?",
"These two PRs are the only two not in phase diagram relevant files\n\nFor the others the question is whether you would prefer a PR for the changes to phase diagram docstrings/Qol separately to the PR for the new class or all in a bigger PR? I think breaking it out should be helpful but whatever is easier for you",
"I'm fine with either, with a mild preference to breaking them out."
] | 2021-10-16T08:56:10
| 2021-10-19T09:15:18
|
2021-10-18T22:14:38Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
in-built all short circuits when checking a boolean list which means a for loop isn't needed here.
I am making this PR as I am looking at trying to get my `PatchedPhaseDiagram` ready to be merged in near future, I have several small QOL changes like this on that branch and think it would be easier if they are merged separately first to making reviewing the `PatchedPhaseDiagram` PR easier (or I can revert on ppd branch if there are issues).
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2263/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2263/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2263",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2263",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2263.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2263.patch",
"merged_at": "2021-10-18T22:14:38Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2264
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2264/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2264/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2264/events
|
https://github.com/materialsproject/pymatgen/pull/2264
| 1,028,001,066
|
PR_kwDOACgets4tRvn3
| 2,264
|
Remove method and function that are no longer used for entry equality
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks!"
] | 2021-10-16T09:17:09
| 2021-10-19T09:14:57
|
2021-10-18T22:15:29Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Previously when dealing with entry equality we wrote these function to do a robust eq check. After these changes didn't fix the results @mkhorton and @mattmcdermott came up with a better solution for the `__hash__` and `__eq__` but these functions were not removed. They are not referenced in `pymatgen` beyond their definitions and so I removed them in my `ppd` branch for code hygiene and think that it might also be sensible to remove on `main`.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2264/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2264/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2264",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2264",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2264.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2264.patch",
"merged_at": "2021-10-18T22:15:28Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2265
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2265/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2265/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2265/events
|
https://github.com/materialsproject/pymatgen/pull/2265
| 1,028,756,801
|
PR_kwDOACgets4tT0IU
| 2,265
|
[WIP] New Structure.get_element_distance() method
|
{
"login": "rpw199912j",
"id": 53155036,
"node_id": "MDQ6VXNlcjUzMTU1MDM2",
"avatar_url": "https://avatars.githubusercontent.com/u/53155036?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rpw199912j",
"html_url": "https://github.com/rpw199912j",
"followers_url": "https://api.github.com/users/rpw199912j/followers",
"following_url": "https://api.github.com/users/rpw199912j/following{/other_user}",
"gists_url": "https://api.github.com/users/rpw199912j/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rpw199912j/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rpw199912j/subscriptions",
"organizations_url": "https://api.github.com/users/rpw199912j/orgs",
"repos_url": "https://api.github.com/users/rpw199912j/repos",
"events_url": "https://api.github.com/users/rpw199912j/events{/privacy}",
"received_events_url": "https://api.github.com/users/rpw199912j/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 3093102097,
"node_id": "MDU6TGFiZWwzMDkzMTAyMDk3",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/stale",
"name": "stale",
"color": "A99B62",
"default": false,
"description": "Abandoned or conflicting PRs and outdated issues"
}
] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks for the contribution. However, I think this is too specific an analysis to be added to pymatgen and it can be done with the existing get_distance_matrix coupled with the elemental information. Can you provide more details on where such an analysis would be useful that it justifies the creation of this method? Thanks.",
"The original motivation is that I was writing several machine learning featurizers where you need to know the metal-metal and metal-nonmental elemental distance to calculate [approximation values for the Hubbard U and charge-transfer gap](https://www.sciencedirect.com/science/article/pii/0921453491905346), as seen in the Zaanen-Sawatzky-Allen framework. These two energy features can be used to differentiate metals from insulators.\r\n\r\nAs you mentioned, this is a convenience method to quickly get the distance between elements without having to manually look up elemental information beforehand, and it serves as the basis for the machine learning featurizers I used in my own research. I totally agree with you that people can achieve the same result using existing methods, but before writing this method, I often find myself finding the element-site mapping by printing out the structure, or loading the structure in a visualization software like VESTA to figure out which sites belong to which elements and are the closest to one another. The biggest utility of this method is that it does all of the above functionalities for you, and can accommodate disordered structures with fractional occupancy, and automatically make a supercell when the input structure does not reflect global symmetry. It would be especially useful if you need to process hundreds if not thousands of structures in a high-throughput study.\r\n\r\nI could just include this method within the Hubbard U and charge transfer gap featurizers and submit a pull request to matminer, but I think this method would benefit a more general audience who are not just looking to classify metals and insulators for material informatics research.",
"Hi @rpw199912j, thank you for the thoughtful PR and explanation.\r\n\r\nI agree with @shyuep that this is a somewhat niche functionality for general use but I can see the usefulness for the featurizer. The code is also very well-documented.",
"@rpw199912j we can surely include this in matminer, please open a WIP PR on the matminer repo and tag me!\r\n",
"Closing this as not planned since the functionality seems better placed in `matminer`."
] | 2021-10-18T07:58:54
| 2023-02-03T17:52:36
|
2023-02-03T17:52:35Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Added a new get_element_distance() method to the pymatgen.core.structure.Structure class that:
* Calculates the minimum element-element distance for one element or between two different elements.
* Automatically calculates with a supercell if any element in the input structure occupies only one site or needs additional sites to better reflect global symmetry.
## Algorithm Overview
Using Fe2O3 as an example (assuming Fe occupies site 0 and 1; O occupies site 2, 3, 4).
1. For each element in the structure, figure out the site(s) it occupies. For example, Fe2O3 would produce {Fe: [0, 1], O: [2, 3, 4]}. When an element in the input structure occupies only one site, by default, make a supercell of [2a, 2b, 2c] to avoid getting a distance of 0.
2. Check if the specified element(s) can be found in the structure. If not, raise a ValueError.
3. If two different elements are specified, create a cartesian product of the indices. For example, finding the Fe-O element distance in Fe2O3 would produce [(0, 2), (0, 3), (0, 4), (1, 2), (1, 3), (1, 4)]. If only one unique element is specified, create a list of pairwise combinations from the indices of that particular element. For example, finding the O-O element distance would produce [(2, 3), (2, 4), (3, 4)].
4. For each pair of site indices, calculate the distance between the sites with the `Structure.get_distance(i, j, jimage=None)` method, resulting in a list of distances.
5. Find the minimum of the distance list and determine a maximum cutoff distance by multiplying the minimum distance with some arbitrary value. The arbitrary value is calculated as (1 + a threshold value). Find all the values in the range of [minimum_distance, minimum_distance * (1 + threshold)] and return the average of this minimum distance list, with the option to return this list itself.
## Additional dependencies introduced (if any)
* None
## TODO (if any)
This is my first time making a pull request. Any comment and advice would be much appreciated!
So far, I've added several tests for the following categories:
- [x] The Si-Si distance in the default structure that comes with the StructureTest class.
- [x] Raising a ValueError if elements that are not in the input structure are passed in as parameters.
- [x] The Mg-O distance in a disordered structure with fractional occupancy modified from the default StructureTest structure.
- [x] The Si-Si distance in a custom-defined simple cubic structure with the lengths of all cell edges set to be 1 angstrom. This structure only has one silicon site specified at the [0, 0, 0] corner, so a supercell of [2a, 2b, 2c] should be made in order to obtain the correct distance of 1 instead of 0.
- [x] Make sure the input structure does not get changed inplace whenever a supercell is made.
- [x] Still with the same structure from 4., the Si-Si distance if the `threshold` value is set to be 1, which would mean all the distances that are less than or equal to 2 angstroms are included. As a result, the calculated distance should be greater than 1 as the distances of the face diagonals and body diagonals are now accepted.
- [x] Whether or not the original minimum distances are returned in a NumPy array if `return_lst` is set to be True.
- [x] Whether or not only unique values are returned if `only_unique` is set to be True.
However, there are several tests concerning the use of `jimage` in the Structure.get_distance() methods that I am not quite sure about, especially when it is used along with the scaling matrix.
- [x] If `jimage` is set to be `None`, meaning choosing the closest periodic image in the get_distance() method, in some cases, using different scaling matrices would produce the same element distance. When `jimage` is set to be `0`, the behavior is changed, and it seems now the get_distance() method only calculates distance for sites that strictly fall inside the cell. The following code segment can be used to reproduce this issue. (**Note**: current implementation defaults to `jimage=None` so that the get_distance() method always uses the nearest periodic image.)
```python
from pymatgen.core.lattice import Lattice
from pymatgen.core.structure import Structure
struct_primitive_3 = Structure(
Lattice([[6, 0, 3], [4, 1, 3], [2, 0, 2]]), ["Fe", "O"], [[0, 0, 0], [0.5, 0, 0]]
)
struct_primitive_3_222 = struct_primitive_3.copy()
struct_primitive_3_222.make_supercell(scaling_matrix=[2, 2, 2])
struct_primitive_3_121 = struct_primitive_3.copy()
struct_primitive_3_121.make_supercell(scaling_matrix=[1, 2, 1])
print("Make sure we are using the same sites for the two different scaled structures")
for site_indices, scaling_matrix, struct_scaled in zip([[0, 2], [0, 1]], [[2, 2, 2], [1, 2, 1]], [struct_primitive_3_222, struct_primitive_3_121]):
print(f"For the structure scaled by {scaling_matrix}")
for site_index in site_indices:
print(f"Site {site_index}")
print(struct_scaled[site_index])
print(struct_primitive_3_222.get_distance(0, 2, jimage=None))
print(struct_primitive_3_121.get_distance(0, 1, jimage=None))
print(struct_primitive_3_121.get_distance(0, 1, jimage=0))
```
Please let me know if I should be adding more tests for additional edge cases, or if the current implementation of `jimage` should be changed.
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2265/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2265/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2265",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2265",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2265.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2265.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2266
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2266/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2266/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2266/events
|
https://github.com/materialsproject/pymatgen/issues/2266
| 1,029,871,315
|
I_kwDOACgets49YpbT
| 2,266
|
Should the called processes be terminated automatically when the execution of pymatgen script is stopped?
|
{
"login": "goodwilling",
"id": 56999195,
"node_id": "MDQ6VXNlcjU2OTk5MTk1",
"avatar_url": "https://avatars.githubusercontent.com/u/56999195?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/goodwilling",
"html_url": "https://github.com/goodwilling",
"followers_url": "https://api.github.com/users/goodwilling/followers",
"following_url": "https://api.github.com/users/goodwilling/following{/other_user}",
"gists_url": "https://api.github.com/users/goodwilling/gists{/gist_id}",
"starred_url": "https://api.github.com/users/goodwilling/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/goodwilling/subscriptions",
"organizations_url": "https://api.github.com/users/goodwilling/orgs",
"repos_url": "https://api.github.com/users/goodwilling/repos",
"events_url": "https://api.github.com/users/goodwilling/events{/privacy}",
"received_events_url": "https://api.github.com/users/goodwilling/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"For general purposes, it is desired to stop the execution of mcsqs when the python script calling \r\nSQSTransformation is terminated. Therefore, I added the following lines (in bold) to \r\npymatgen/command_line/mcsqs_caller.py and all mcsqs processes can be stopped when the execution of \r\npython script is terminated.\r\n\r\nimport signal\r\n\r\n......\r\n\r\n\r\ndef run_mcsqs(\r\n......\r\n\r\n mcsqs_find_sqs_processes = []\r\n if instances and instances > 1:\r\n # if multiple instances, run a range of commands using \"-ip\"\r\n for i in range(instances):\r\n instance_cmd = [\"-ip {}\".format(i + 1)]\r\n cmd = mcsqs_find_sqs_cmd + add_ons + instance_cmd\r\n p = Popen(cmd, shell=True) # pylint: disable=R1732\r\n mcsqs_find_sqs_processes.append(p)\r\n else:\r\n # run normal mcsqs command\r\n cmd = mcsqs_find_sqs_cmd + add_ons\r\n p = Popen(cmd, shell=True) # pylint: disable=R1732\r\n mcsqs_find_sqs_processes.append(p)\r\n\r\n **def signal_handler(sig, frame):\r\n for p in mcsqs_find_sqs_processes:\r\n p.kill()\r\n p.communicate()\r\n\r\n signal.signal(signal.SIGTERM, signal_handler)** \r\n\r\n try:\r\n for idx, p in enumerate(mcsqs_find_sqs_processes):\r\n p.communicate(timeout=search_time * 60)\r\n\r\n if instances and instances > 1:\r\n p = Popen([\"wsl\", \"mcsqs\", \"-best\"]) # pylint: disable=R1732\r\n p.communicate()\r\n ...",
"Thanks. Pls provide a PR."
] | 2021-10-19T05:17:53
| 2021-10-26T16:01:10
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I am using SQSTransformation to generate SQS in RHEL 8.3 by using the command
python test_sqs.py &
where test_sqs.py is just a copy from pymatgen/transformation/tests.
Since it takes time, the execution of the python script is stopped by
pkill -f test_sqs.py
However, many mcsqs processes still keep running after that. In the other hand, if I use the command
python test_sqs.py
and press Ctrl+C to stop the execution, all mcsqs processes will be killed immediately.
I wonder what is the correct behavior for the called processes when the caller is stopped, especially for
the mcsqs processes called by SQSTransformation.
Thank you very much.
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2266/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2266/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2267
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2267/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2267/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2267/events
|
https://github.com/materialsproject/pymatgen/issues/2267
| 1,030,528,298
|
I_kwDOACgets49bJ0q
| 2,267
|
Round trip potcar writing/reading
|
{
"login": "JosephMontoya-TRI",
"id": 45643466,
"node_id": "MDQ6VXNlcjQ1NjQzNDY2",
"avatar_url": "https://avatars.githubusercontent.com/u/45643466?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/JosephMontoya-TRI",
"html_url": "https://github.com/JosephMontoya-TRI",
"followers_url": "https://api.github.com/users/JosephMontoya-TRI/followers",
"following_url": "https://api.github.com/users/JosephMontoya-TRI/following{/other_user}",
"gists_url": "https://api.github.com/users/JosephMontoya-TRI/gists{/gist_id}",
"starred_url": "https://api.github.com/users/JosephMontoya-TRI/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/JosephMontoya-TRI/subscriptions",
"organizations_url": "https://api.github.com/users/JosephMontoya-TRI/orgs",
"repos_url": "https://api.github.com/users/JosephMontoya-TRI/repos",
"events_url": "https://api.github.com/users/JosephMontoya-TRI/events{/privacy}",
"received_events_url": "https://api.github.com/users/JosephMontoya-TRI/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I think the reason for this is simply that the hash database is incomplete.\r\n\r\nThere are two sets of hashes, a \"file\" hash (checks the POTCAR is byte-to-byte identical) and a \"data\" hash (which is a bit more permissive, e.g. if there are changes in comment strings etc.)\r\n\r\nThe data hash matches, but the file match does not, hence it is in error. The warning message should probably be less extreme, i.e. acknowledge the possibility that the pymatgen database is incomplete.\r\n\r\nThe warning does not appear on set construction since only the \"data\" hash is checked.\r\n\r\nSuggested remedy here is:\r\n\r\n1. Only check the data hash in both places to avoid inconsistency.\r\n2. Supplement the hash file with additional hashes.\r\n\r\nA separate issue here is that `PotcarSingle.identify_potcar()` returns `(['PBE'], ['runelements', 'C'])` for reasons that are not clear to me.",
"Hm, I think the `runelements` is simply because there was an older VASP POTCAR that identified itself as such (or perhaps a parsing bug).",
"Ok, I mis-spoke previously. I think this is a round-trip issue due to some logic in `Potcar.from_file`. The file itself is correct.\r\n\r\nI have a fix which is to hash the `str(PotcarSingle)` rather than `PotcarSingle.data`. The failing file hash is because `PotcarSingle.data` removes a trailing newline."
] | 2021-10-19T16:42:05
| 2021-10-20T23:32:09
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Round trip Potcar parsing results in a warning that's not present when writing input.
**To Reproduce**
```
import unittest
from pymatgen.io.vasp.sets import MPRelaxSet
from pymatgen.io.vasp.inputs import VaspInput
from pymatgen.core.lattice import Lattice
from pymatgen.core.structure import Structure
from pymatgen.core.periodic_table import Element
from monty.tempfile import ScratchDir
class PotcarCompatTest(unittest.TestCase):
def test_round_trip(self):
with ScratchDir('.'):
structure = Structure(Lattice.cubic(1.5), ['C'], [[0, 0, 0,]])
relax_set = MPRelaxSet(structure)
# This should complain if md5 hashes are wrong
print("Write set")
relax_set.write_input('.')
print("Read")
vasp_input = VaspInput.from_directory('.') # This results in a warning
if __name__ == "__main__":
unittest.main()
```
**Expected behavior**
No warning if input set is written without a warning.
**Desktop (please complete the following information):**
- OS: Linux
**Additional context**
Believe this is related to the regex parsing of PotcarSingle from a multiple potcar file, or perhaps on the write itself, which may leave out a trailing line break present in the original file.
On the write - the PotcarData contains
a trailing line break, but not on the read.
```
test_potcar.py Write set
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>> PDB set_trace >>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>
> /root/miniconda3/lib/python3.9/site-packages/pymatgen/io/vasp/inputs.py(1780)__init__()
-> self.hash = self.get_potcar_hash()
(Pdb) self.data[-20:]
'+01\n End of Dataset\n'
(Pdb) c
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>> PDB continue >>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>
Read
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>> PDB set_trace >>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>
> /root/miniconda3/lib/python3.9/site-packages/pymatgen/io/vasp/inputs.py(1780)__init__()
-> self.hash = self.get_potcar_hash()
(Pdb) self.data[-20:]
'E+01\n End of Dataset'
(Pdb)
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2267/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2267/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2268
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2268/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2268/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2268/events
|
https://github.com/materialsproject/pymatgen/pull/2268
| 1,030,745,426
|
PR_kwDOACgets4tZ_-9
| 2,268
|
QOL changes to phase_diagram.py
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks."
] | 2021-10-19T21:07:44
| 2021-10-20T13:59:59
|
2021-10-20T13:54:46Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Added google docstrings to fit `pymatgen` style guide and whitespace line changes for better readability. Several typos/whitespaces. Slightly adjusted the function of `_get_slsqp_decomp()` to make it more useful as a stand alone function. Added a few ease of use methods for intermediary calculations that help in the `PatchedPhaseDiagram` implementation to come.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2268/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2268/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2268",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2268",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2268.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2268.patch",
"merged_at": "2021-10-20T13:54:46Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2269
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2269/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2269/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2269/events
|
https://github.com/materialsproject/pymatgen/pull/2269
| 1,030,916,998
|
PR_kwDOACgets4tajIP
| 2,269
|
Moladaptor fix
|
{
"login": "orionarcher",
"id": 27712051,
"node_id": "MDQ6VXNlcjI3NzEyMDUx",
"avatar_url": "https://avatars.githubusercontent.com/u/27712051?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/orionarcher",
"html_url": "https://github.com/orionarcher",
"followers_url": "https://api.github.com/users/orionarcher/followers",
"following_url": "https://api.github.com/users/orionarcher/following{/other_user}",
"gists_url": "https://api.github.com/users/orionarcher/gists{/gist_id}",
"starred_url": "https://api.github.com/users/orionarcher/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/orionarcher/subscriptions",
"organizations_url": "https://api.github.com/users/orionarcher/orgs",
"repos_url": "https://api.github.com/users/orionarcher/repos",
"events_url": "https://api.github.com/users/orionarcher/events{/privacy}",
"received_events_url": "https://api.github.com/users/orionarcher/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"thanks\r\n"
] | 2021-10-20T02:46:04
| 2021-10-20T13:38:39
|
2021-10-20T13:38:32Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
* Fix 1: allowed pmg.io.babel.BabelMolAdaptor to accept pybel.Molecule
## Checklist
Before a pull request can be merged, the following items must be checked:
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code. (not needed)
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [ ] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2269/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2269/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2269",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2269",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2269.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2269.patch",
"merged_at": "2021-10-20T13:38:32Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2270
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2270/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2270/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2270/events
|
https://github.com/materialsproject/pymatgen/issues/2270
| 1,031,462,294
|
I_kwDOACgets49et2W
| 2,270
|
BasePhaseDiagram should be renamed
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
] | null |
[
"I have no problem changing the name, and agree re. the private method call, but I believe the class itself exists for a good reason.\r\n\r\nIf I remember correctly, the _reason_ for this class was to make PhaseDiagram itself MSONable without breaking backwards compatibility, but I would have to check.",
"No. The purpose is merely to store the results of the convex hull to avoid recomputation. Theoretically, just serializing the list of entries alone would suffice for serializing the PD. ",
"Ok, but that's the point I was making -- being able to serialize the entries is different from being able to serialize the phase diagram itself. Since recomputing the convex hull is expensive, reconstructing the PhaseDiagram from entries alone would be a very slow operation, which is prohibitive when dealing with a large number of phase diagrams.",
"I understand that point. Just that this is not the right way to do this. Let me suggest another approach:\r\n\r\n```python\r\nclass PhaseDiagram(MSONAble):\r\n\r\n def __init__(entries, *, computed_data:dict=None):\r\n if computed_data is None:\r\n <same old code>\r\n else:\r\n <load computed data and skip computation>\r\n```\r\n\r\n`as_dict` and `from_dict` will store and get the computed data.\r\n",
"That seems reasonable, I'm fine with that change (with the caveat that it would break backwards compatibility, but I think that would affect a small number of people, probably mostly ourselves).",
"Why would it break backwards compatibility? Just the people who are using BasePD at the moment? As far as normal users go, all method signatures are still the same?\r\n",
"> Just the people who are using BasePD at the moment?\r\n\r\nYes, exactly.",
"In the first place, the current doc for PhaseDiagram is completely wrong because of this change. Ridiculous.",
"I pushed my implementation. In any case, the old implementation makes no sense. While it allows for pre-computed data, it does not actually obey the MSONable protocol. That is because the underlying objects like Simplex are actually not json serializable and do not obey the MSONable protocol as well. The only use case is actually just to dump it to a dict and reconstitute the object immediately but if you go through any form of actual serialization, it doesn't work. \r\n",
"Interesting, I'll let the person know who implemented it. I imagine they only added tests for as_dict/from_dict rather than checking the full round trip to JSON string.",
"@mkhorton I am going to close this issue since the MSONable part of phase diagram is a separate one from naming. I see you have some recent fixes attempting to fix this. ",
"Thanks, yes, this should all be addressed now."
] | 2021-10-20T14:12:11
| 2021-10-27T15:58:19
|
2021-10-27T15:42:17Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I am not sure when this class was introduced. But whatever it is, the naming is not good. A new user coming to pymatgen will be wondering what the heck is a BasePhaseDiagram. It has the connotations of an abstract base class but is actually not one. Then the original PhaseDiagram class inherits from BasePhaseDiagram but calls a PRIVATE method in the init. That is all kinds of ludicrous in design.
Pls revert such that PhaseDiagram is the actual base class and if you want something to just store the computed PD results, write a dataclass or a "CalculatedPhaseDiagram" (similar to "BalancedReaction") that stores the results.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2270/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2270/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2271
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2271/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2271/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2271/events
|
https://github.com/materialsproject/pymatgen/issues/2271
| 1,031,591,987
|
I_kwDOACgets49fNgz
| 2,271
|
Logger calls that appear to be historic and could potentially be removed in `ReactionDiagram`
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"They're debug calls anyway, presumably were useful to whoever was writing that class.",
"happy for me to remove? the git blame it 4 years ago so I am sure that they're finished debugging by now?",
"It's more a philosophical point -- I think `logger.debug()` statements are useful for whoever develops the code next and harmless otherwise? (unlike, say, a stray print statement which is nearly-always bad)\r\n\r\nHappy to reconsider my position though, is this considered bad practice?",
"Loggers should be kept. These do not affect normal usage. When something bad happens, we need these to figure out what went wrong."
] | 2021-10-20T16:12:55
| 2021-10-21T17:54:14
|
2021-10-21T17:54:14Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
https://github.com/materialsproject/pymatgen/blob/b40ae2d962936a2ad8c2a024a3c0b2efc503e8e8/pymatgen/analysis/phase_diagram.py#L1487-L1489
@shyuep are these logging calls still needed in the code? I can't trace anywhere where they might be used and was wondering if I should clean them up?
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2271/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2271/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2272
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2272/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2272/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2272/events
|
https://github.com/materialsproject/pymatgen/pull/2272
| 1,031,664,689
|
PR_kwDOACgets4tc-Nq
| 2,272
|
Make the appearance of periodic_table_heatmap changeable.
|
{
"login": "penicillin0",
"id": 56141035,
"node_id": "MDQ6VXNlcjU2MTQxMDM1",
"avatar_url": "https://avatars.githubusercontent.com/u/56141035?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/penicillin0",
"html_url": "https://github.com/penicillin0",
"followers_url": "https://api.github.com/users/penicillin0/followers",
"following_url": "https://api.github.com/users/penicillin0/following{/other_user}",
"gists_url": "https://api.github.com/users/penicillin0/gists{/gist_id}",
"starred_url": "https://api.github.com/users/penicillin0/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/penicillin0/subscriptions",
"organizations_url": "https://api.github.com/users/penicillin0/orgs",
"repos_url": "https://api.github.com/users/penicillin0/repos",
"events_url": "https://api.github.com/users/penicillin0/events{/privacy}",
"received_events_url": "https://api.github.com/users/penicillin0/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/43671060)\n\nCoverage decreased (-0.6%) to 83.131% when pulling **30af0330ecdd2d6c70bc979ce771058269d13268 on penicillin0:add-option-periodic_table_heatmap** into **b40ae2d962936a2ad8c2a024a3c0b2efc503e8e8 on materialsproject:master**.\n",
"thanks\r\n"
] | 2021-10-20T17:37:49
| 2021-10-21T15:26:28
|
2021-10-21T15:26:22Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
I found the current periodic table difficult to read because of the color and the size of the letters, so I made it partially changeable.
* The color of the periodic table border can be changed.
* The font size of value and symbol on the periodic table can be changed.
* Added the option to change the font color to a color that is easier to read depending on the background color.
## Additional dependencies introduced (if any)
- None
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
## Demo
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2272/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2272/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2272",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2272",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2272.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2272.patch",
"merged_at": "2021-10-21T15:26:22Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2273
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2273/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2273/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2273/events
|
https://github.com/materialsproject/pymatgen/pull/2273
| 1,031,913,827
|
PR_kwDOACgets4tdyZw
| 2,273
|
Correct POTCAR hash warnings
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/43657330)\n\nCoverage decreased (-0.02%) to 83.745% when pulling **21f23fbb471aa4f5249978a71fe3b255623273a7 on potcar-warning** into **6f73449fd8cc7da094505a7bceaacda24386875c on master**.\n"
] | 2021-10-20T23:35:15
| 2022-10-10T17:54:05
|
2021-10-21T16:41:29Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Fix for #2267
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2273/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2273/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2273",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2273",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2273.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2273.patch",
"merged_at": "2021-10-21T16:41:29Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2274
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2274/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2274/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2274/events
|
https://github.com/materialsproject/pymatgen/issues/2274
| 1,032,329,114
|
I_kwDOACgets49iBea
| 2,274
|
Implicit hydrogen parsing should not be default for doped cifs.
|
{
"login": "TimoSommer",
"id": 67800320,
"node_id": "MDQ6VXNlcjY3ODAwMzIw",
"avatar_url": "https://avatars.githubusercontent.com/u/67800320?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/TimoSommer",
"html_url": "https://github.com/TimoSommer",
"followers_url": "https://api.github.com/users/TimoSommer/followers",
"following_url": "https://api.github.com/users/TimoSommer/following{/other_user}",
"gists_url": "https://api.github.com/users/TimoSommer/gists{/gist_id}",
"starred_url": "https://api.github.com/users/TimoSommer/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/TimoSommer/subscriptions",
"organizations_url": "https://api.github.com/users/TimoSommer/orgs",
"repos_url": "https://api.github.com/users/TimoSommer/repos",
"events_url": "https://api.github.com/users/TimoSommer/events{/privacy}",
"received_events_url": "https://api.github.com/users/TimoSommer/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-10-21T10:33:51
| 2021-10-21T10:50:04
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
I have found that when reading in a cif file of a doped structure, where hydrogen and another element share the same lattice site, the hydrogen is not shown in the structure, except as 'implicit_hydrogens' in structure.site_properties. If however the lattice site of the hydrogen is not shared with another element, the hydrogen is shown and treated like every other element. I think this is very confusing and should not be the default way of parsing (I think this was added in #692). I have not found any mention of how to turn off this behaviour in the documentation. Neither have I gotten a warning like I think I should get according to #1287.
**To Reproduce**
1. Save the following cif as variable:
```
cif = """# generated using pymatgen
data_Ce2Fe2As2H1.8O0.2
_symmetry_space_group_name_H-M P4/nmm
_cell_length_a 3.97371900
_cell_length_b 3.97371900
_cell_length_c 8.96776000
_cell_angle_alpha 90.00000000
_cell_angle_beta 90.00000000
_cell_angle_gamma 90.00000000
_symmetry_Int_Tables_number 129
_chemical_formula_structural Ce2Fe2As2H1.8O0.2
_chemical_formula_sum 'Ce2 Fe2 As2 H1.8 O0.2'
_cell_volume 141.60490035
_cell_formula_units_Z 1
loop_
_symmetry_equiv_pos_site_id
_symmetry_equiv_pos_as_xyz
1 'x, y, z'
2 '-y+1/2, x+1/2, z'
3 '-x, -y, z'
4 'y+1/2, -x+1/2, z'
5 'x+1/2, -y+1/2, -z'
6 '-y, -x, -z'
7 '-x+1/2, y+1/2, -z'
8 'y, x, -z'
9 '-x+1/2, -y+1/2, -z'
10 'y, -x, -z'
11 'x+1/2, y+1/2, -z'
12 '-y, x, -z'
13 '-x, y, z'
14 'y+1/2, x+1/2, z'
15 'x, -y, z'
16 '-y+1/2, -x+1/2, z'
loop_
_atom_site_type_symbol
_atom_site_label
_atom_site_symmetry_multiplicity
_atom_site_fract_x
_atom_site_fract_y
_atom_site_fract_z
_atom_site_occupancy
Ce Ce0 2 0.00000000 0.50000000 0.86157000 1.0
Fe Fe1 2 0.00000000 0.00000000 0.50000000 1.0
As As2 2 0.00000000 0.50000000 0.31128600 1.0
O O3 2 0.00000000 0.00000000 0.00000000 0.1
H H4 2 0.00000000 0.00000000 0.00000000 0.9
"""
```
2. Read in the cif file as structure:
```
from pymatgen.core.structure import Structure
structure = Structure.from_str(cif, fmt='cif')
```
3. The structure shows (except for structure.site_properties) no sign of the hydrogen!
`structure.formula == 'Ce2 Fe2 As2 O0.2'
`
**Expected behavior**
The doped hydrogen should be treated like every other element and shown in the formula and the structure. Implicit hydrogens make sense for molecules, but it doesn't make sense that the lattice site (H:0.99 O:0.01) is parsed completely different from the lattice site (H:1.0). On the other side, if this is truly the wanted default, I would wish for a simple option to turn off this behaviour and a clear warning when this happens.
**Desktop (please complete the following information):**
- OS: Linux Mint 19.1 Cinnamon
- pymatgen: pymatgen-2022.0.14
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2274/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2274/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2275
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2275/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2275/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2275/events
|
https://github.com/materialsproject/pymatgen/issues/2275
| 1,033,725,417
|
I_kwDOACgets49nWXp
| 2,275
|
Tests fail: invalid module name: 'pymatgen.io.vasp.inputs'
|
{
"login": "yurivict",
"id": 271906,
"node_id": "MDQ6VXNlcjI3MTkwNg==",
"avatar_url": "https://avatars.githubusercontent.com/u/271906?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/yurivict",
"html_url": "https://github.com/yurivict",
"followers_url": "https://api.github.com/users/yurivict/followers",
"following_url": "https://api.github.com/users/yurivict/following{/other_user}",
"gists_url": "https://api.github.com/users/yurivict/gists{/gist_id}",
"starred_url": "https://api.github.com/users/yurivict/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/yurivict/subscriptions",
"organizations_url": "https://api.github.com/users/yurivict/orgs",
"repos_url": "https://api.github.com/users/yurivict/repos",
"events_url": "https://api.github.com/users/yurivict/events{/privacy}",
"received_events_url": "https://api.github.com/users/yurivict/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"You need to provide more details. That one liner does not give any information about your installation and there is no way we can diagnose where the problem lies. ",
"Build [log](https://people.freebsd.org/~yuri/py-pymatgen.log).\r\n\r\n[List](https://people.freebsd.org/~yuri/py-pymatgen-plist.txt) of installed files.\r\n",
"I am not sure what is happening, but it would appear your pymatgen install is not complete. Further, you seem to be installing to the OS python, which is highly discoouraged. I would recommend you redo the install within a proper virtual env or even better, use Miniconda to do the install. Please reopen the issue if you still encounter problems. ",
"I've came by this failure and have fixed it partially. The testing is set up properly now, but\r\n1. Some tests fail because data files seem to be missing from the distribution.\r\n2. Other tests fail trying to access the network. Policy prohibits network access during build/testing, as it would hinder tests reproducibility and stability.\r\n3. Because these attempts are made directly from the root of test code\r\nhttps://github.com/materialsproject/pymatgen/blob/da8d0a996584b716e2f9a6f2e3fdf9a94d80454a/pymatgen/vis/tests/test_plotters.py#L17-L18\r\ninstead of being wrapped into fixtures, test setup/teardown functions or skip's, pytest collection fails, and no tests which could probably still be executed are run.\r\n\r\nAt least 1 and 3 should be fixed in pymatgen code, so please reopen.\r\n\r\n> I would recommend you redo the install within a proper virtual env or even better, use Miniconda to do the install\r\n\r\nThis is absolutely not possible for packaging. Additionally, miniconda is not available for FreeBSD.\r\n\r\n```\r\nrunning build_ext\r\ncopying build/lib.freebsd-13.0-RELEASE-amd64-3.8/pymatgen/optimization/linear_assignment.cpython-38.so -> pymatgen/optimization\r\ncopying build/lib.freebsd-13.0-RELEASE-amd64-3.8/pymatgen/util/coord_cython.cpython-38.so -> pymatgen/util\r\ncopying build/lib.freebsd-13.0-RELEASE-amd64-3.8/pymatgen/optimization/neighbors.cpython-38.so -> pymatgen/optimization\r\n============================= test session starts ==============================\r\nplatform freebsd13 -- Python 3.8.13, pytest-7.1.1, pluggy-0.13.1 -- /usr/local/bin/python3.8\r\ncachedir: .pytest_cache\r\nrootdir: /work/usr/ports/science/py-pymatgen/work-py38/pymatgen-2022.0.17, configfile: setup.cfg\r\ncollecting ... collected 1434 items / 7 errors / 2 skipped\r\n\r\n==================================== ERRORS ====================================\r\n___ ERROR collecting pymatgen/electronic_structure/tests/test_boltztrap2.py ____\r\npymatgen/electronic_structure/tests/test_boltztrap2.py:35: in <module>\r\n vrun = Vasprun(vrunfile, parse_projected_eigen=True)\r\npymatgen/io/vasp/outputs.py:345: in __init__\r\n with zopen(filename, \"rt\") as f:\r\n/usr/local/lib/python3.8/site-packages/monty/io.py:43: in zopen\r\n return io.open(filename, *args, **kwargs) # pylint: disable=R1732\r\nE FileNotFoundError: [Errno 2] No such file or directory: '/work/usr/ports/science/py-pymatgen/work-py38/pymatgen-2022.0.17/pymatgen/util/../../test_files/boltztrap2/vasprun.xml'\r\n_______ ERROR collecting pymatgen/entries/tests/test_computed_entries.py _______\r\npymatgen/entries/tests/test_computed_entries.py:29: in <module>\r\n vasprun = Vasprun(filepath)\r\npymatgen/io/vasp/outputs.py:345: in __init__\r\n with zopen(filename, \"rt\") as f:\r\n/usr/local/lib/python3.8/site-packages/monty/io.py:43: in zopen\r\n return io.open(filename, *args, **kwargs) # pylint: disable=R1732\r\nE FileNotFoundError: [Errno 2] No such file or directory: '/work/usr/ports/science/py-pymatgen/work-py38/pymatgen-2022.0.17/pymatgen/util/../../test_files/vasprun.xml'\r\n_______________ ERROR collecting pymatgen/ext/tests/test_cod.py ________________\r\n/usr/local/lib/python3.8/site-packages/urllib3/connection.py:174: in _new_conn\r\n conn = connection.create_connection(\r\n/usr/local/lib/python3.8/site-packages/urllib3/util/connection.py:72: in create_connection\r\n for res in socket.getaddrinfo(host, port, family, socket.SOCK_STREAM):\r\n/usr/local/lib/python3.8/socket.py:918: in getaddrinfo\r\n for res in _socket.getaddrinfo(host, port, family, type, proto, flags):\r\nE socket.gaierror: [Errno 8] Name does not resolve\r\n\r\nDuring handling of the above exception, another exception occurred:\r\n/usr/local/lib/python3.8/site-packages/urllib3/connectionpool.py:703: in urlopen\r\n httplib_response = self._make_request(\r\n/usr/local/lib/python3.8/site-packages/urllib3/connectionpool.py:386: in _make_request\r\n self._validate_conn(conn)\r\n/usr/local/lib/python3.8/site-packages/urllib3/connectionpool.py:1040: in _validate_conn\r\n conn.connect()\r\n/usr/local/lib/python3.8/site-packages/urllib3/connection.py:358: in connect\r\n self.sock = conn = self._new_conn()\r\n/usr/local/lib/python3.8/site-packages/urllib3/connection.py:186: in _new_conn\r\n raise NewConnectionError(\r\nE urllib3.exceptions.NewConnectionError: <urllib3.connection.HTTPSConnection object at 0x8312b2d60>: Failed to establish a new connection: [Errno 8] Name does not resolve\r\n\r\nDuring handling of the above exception, another exception occurred:\r\n/usr/local/lib/python3.8/site-packages/requests/adapters.py:440: in send\r\n resp = conn.urlopen(\r\n/usr/local/lib/python3.8/site-packages/urllib3/connectionpool.py:785: in urlopen\r\n retries = retries.increment(\r\n/usr/local/lib/python3.8/site-packages/urllib3/util/retry.py:592: in increment\r\n raise MaxRetryError(_pool, url, error or ResponseError(cause))\r\nE urllib3.exceptions.MaxRetryError: HTTPSConnectionPool(host='www.crystallography.net', port=443): Max retries exceeded with url: / (Caused by NewConnectionError('<urllib3.connection.HTTPSConnection object at 0x8312b2d60>: Failed to establish a new connection: [Errno 8] Name does not resolve'))\r\n\r\nDuring handling of the above exception, another exception occurred:\r\npymatgen/ext/tests/test_cod.py:14: in <module>\r\n website_is_up = requests.get(\"https://www.crystallography.net\").status_code == 200\r\n/usr/local/lib/python3.8/site-packages/requests/api.py:75: in get\r\n return request('get', url, params=params, **kwargs)\r\n/usr/local/lib/python3.8/site-packages/requests/api.py:61: in request\r\n return session.request(method=method, url=url, **kwargs)\r\n/usr/local/lib/python3.8/site-packages/requests/sessions.py:529: in request\r\n resp = self.send(prep, **send_kwargs)\r\n/usr/local/lib/python3.8/site-packages/requests/sessions.py:645: in send\r\n r = adapter.send(request, **kwargs)\r\n/usr/local/lib/python3.8/site-packages/requests/adapters.py:519: in send\r\n raise ConnectionError(e, request=request)\r\nE requests.exceptions.ConnectionError: HTTPSConnectionPool(host='www.crystallography.net', port=443): Max retries exceeded with url: / (Caused by NewConnectionError('<urllib3.connection.HTTPSConnection object at 0x8312b2d60>: Failed to establish a new connection: [Errno 8] Name does not resolve'))\r\n____________ ERROR collecting pymatgen/ext/tests/test_crystalsai.py ____________\r\n/usr/local/lib/python3.8/site-packages/urllib3/connection.py:174: in _new_conn\r\n conn = connection.create_connection(\r\n/usr/local/lib/python3.8/site-packages/urllib3/util/connection.py:72: in create_connection\r\n for res in socket.getaddrinfo(host, port, family, socket.SOCK_STREAM):\r\n/usr/local/lib/python3.8/socket.py:918: in getaddrinfo\r\n for res in _socket.getaddrinfo(host, port, family, type, proto, flags):\r\nE socket.gaierror: [Errno 8] Name does not resolve\r\n\r\nDuring handling of the above exception, another exception occurred:\r\n/usr/local/lib/python3.8/site-packages/urllib3/connectionpool.py:703: in urlopen\r\n httplib_response = self._make_request(\r\n/usr/local/lib/python3.8/site-packages/urllib3/connectionpool.py:398: in _make_request\r\n conn.request(method, url, **httplib_request_kw)\r\n/usr/local/lib/python3.8/site-packages/urllib3/connection.py:239: in request\r\n super(HTTPConnection, self).request(method, url, body=body, headers=headers)\r\n/usr/local/lib/python3.8/http/client.py:1256: in request\r\n self._send_request(method, url, body, headers, encode_chunked)\r\n/usr/local/lib/python3.8/http/client.py:1302: in _send_request\r\n self.endheaders(body, encode_chunked=encode_chunked)\r\n/usr/local/lib/python3.8/http/client.py:1251: in endheaders\r\n self._send_output(message_body, encode_chunked=encode_chunked)\r\n/usr/local/lib/python3.8/http/client.py:1011: in _send_output\r\n self.send(msg)\r\n/usr/local/lib/python3.8/http/client.py:951: in send\r\n self.connect()\r\n/usr/local/lib/python3.8/site-packages/urllib3/connection.py:205: in connect\r\n conn = self._new_conn()\r\n/usr/local/lib/python3.8/site-packages/urllib3/connection.py:186: in _new_conn\r\n raise NewConnectionError(\r\nE urllib3.exceptions.NewConnectionError: <urllib3.connection.HTTPConnection object at 0x83117f730>: Failed to establish a new connection: [Errno 8] Name does not resolve\r\n\r\nDuring handling of the above exception, another exception occurred:\r\n/usr/local/lib/python3.8/site-packages/requests/adapters.py:440: in send\r\n resp = conn.urlopen(\r\n/usr/local/lib/python3.8/site-packages/urllib3/connectionpool.py:785: in urlopen\r\n retries = retries.increment(\r\n/usr/local/lib/python3.8/site-packages/urllib3/util/retry.py:592: in increment\r\n raise MaxRetryError(_pool, url, error or ResponseError(cause))\r\nE urllib3.exceptions.MaxRetryError: HTTPConnectionPool(host='megnet.crystals.ai', port=80): Max retries exceeded with url: / (Caused by NewConnectionError('<urllib3.connection.HTTPConnection object at 0x83117f730>: Failed to establish a new connection: [Errno 8] Name does not resolve'))\r\n\r\nDuring handling of the above exception, another exception occurred:\r\npymatgen/ext/tests/test_crystalsai.py:12: in <module>\r\n website_is_up = requests.get(\"http://megnet.crystals.ai\").status_code == 200\r\n/usr/local/lib/python3.8/site-packages/requests/api.py:75: in get\r\n return request('get', url, params=params, **kwargs)\r\n/usr/local/lib/python3.8/site-packages/requests/api.py:61: in request\r\n return session.request(method=method, url=url, **kwargs)\r\n/usr/local/lib/python3.8/site-packages/requests/sessions.py:529: in request\r\n resp = self.send(prep, **send_kwargs)\r\n/usr/local/lib/python3.8/site-packages/requests/sessions.py:645: in send\r\n r = adapter.send(request, **kwargs)\r\n/usr/local/lib/python3.8/site-packages/requests/adapters.py:519: in send\r\n raise ConnectionError(e, request=request)\r\nE requests.exceptions.ConnectionError: HTTPConnectionPool(host='megnet.crystals.ai', port=80): Max retries exceeded with url: / (Caused by NewConnectionError('<urllib3.connection.HTTPConnection object at 0x83117f730>: Failed to establish a new connection: [Errno 8] Name does not resolve'))\r\n_______________ ERROR collecting pymatgen/ext/tests/test_jhu.py ________________\r\n/usr/local/lib/python3.8/site-packages/urllib3/connection.py:174: in _new_conn\r\n conn = connection.create_connection(\r\n/usr/local/lib/python3.8/site-packages/urllib3/util/connection.py:72: in create_connection\r\n for res in socket.getaddrinfo(host, port, family, socket.SOCK_STREAM):\r\n/usr/local/lib/python3.8/socket.py:918: in getaddrinfo\r\n for res in _socket.getaddrinfo(host, port, family, type, proto, flags):\r\nE socket.gaierror: [Errno 8] Name does not resolve\r\n\r\nDuring handling of the above exception, another exception occurred:\r\n/usr/local/lib/python3.8/site-packages/urllib3/connectionpool.py:703: in urlopen\r\n httplib_response = self._make_request(\r\n/usr/local/lib/python3.8/site-packages/urllib3/connectionpool.py:398: in _make_request\r\n conn.request(method, url, **httplib_request_kw)\r\n/usr/local/lib/python3.8/site-packages/urllib3/connection.py:239: in request\r\n super(HTTPConnection, self).request(method, url, body=body, headers=headers)\r\n/usr/local/lib/python3.8/http/client.py:1256: in request\r\n self._send_request(method, url, body, headers, encode_chunked)\r\n/usr/local/lib/python3.8/http/client.py:1302: in _send_request\r\n self.endheaders(body, encode_chunked=encode_chunked)\r\n/usr/local/lib/python3.8/http/client.py:1251: in endheaders\r\n self._send_output(message_body, encode_chunked=encode_chunked)\r\n/usr/local/lib/python3.8/http/client.py:1011: in _send_output\r\n self.send(msg)\r\n/usr/local/lib/python3.8/http/client.py:951: in send\r\n self.connect()\r\n/usr/local/lib/python3.8/site-packages/urllib3/connection.py:205: in connect\r\n conn = self._new_conn()\r\n/usr/local/lib/python3.8/site-packages/urllib3/connection.py:186: in _new_conn\r\n raise NewConnectionError(\r\nE urllib3.exceptions.NewConnectionError: <urllib3.connection.HTTPConnection object at 0x830fdd040>: Failed to establish a new connection: [Errno 8] Name does not resolve\r\n\r\nDuring handling of the above exception, another exception occurred:\r\n/usr/local/lib/python3.8/site-packages/requests/adapters.py:440: in send\r\n resp = conn.urlopen(\r\n/usr/local/lib/python3.8/site-packages/urllib3/connectionpool.py:785: in urlopen\r\n retries = retries.increment(\r\n/usr/local/lib/python3.8/site-packages/urllib3/util/retry.py:592: in increment\r\n raise MaxRetryError(_pool, url, error or ResponseError(cause))\r\nE urllib3.exceptions.MaxRetryError: HTTPConnectionPool(host='muellergroup.jhu.edu', port=8080): Max retries exceeded with url: / (Caused by NewConnectionError('<urllib3.connection.HTTPConnection object at 0x830fdd040>: Failed to establish a new connection: [Errno 8] Name does not resolve'))\r\n\r\nDuring handling of the above exception, another exception occurred:\r\npymatgen/ext/tests/test_jhu.py:20: in <module>\r\n website_is_up = requests.get(\"http://muellergroup.jhu.edu:8080\").status_code == 200\r\n/usr/local/lib/python3.8/site-packages/requests/api.py:75: in get\r\n return request('get', url, params=params, **kwargs)\r\n/usr/local/lib/python3.8/site-packages/requests/api.py:61: in request\r\n return session.request(method=method, url=url, **kwargs)\r\n/usr/local/lib/python3.8/site-packages/requests/sessions.py:529: in request\r\n resp = self.send(prep, **send_kwargs)\r\n/usr/local/lib/python3.8/site-packages/requests/sessions.py:645: in send\r\n r = adapter.send(request, **kwargs)\r\n/usr/local/lib/python3.8/site-packages/requests/adapters.py:519: in send\r\n raise ConnectionError(e, request=request)\r\nE requests.exceptions.ConnectionError: HTTPConnectionPool(host='muellergroup.jhu.edu', port=8080): Max retries exceeded with url: / (Caused by NewConnectionError('<urllib3.connection.HTTPConnection object at 0x830fdd040>: Failed to establish a new connection: [Errno 8] Name does not resolve'))\r\n_____________ ERROR collecting pymatgen/ext/tests/test_matproj.py ______________\r\n/usr/local/lib/python3.8/site-packages/urllib3/connection.py:174: in _new_conn\r\n conn = connection.create_connection(\r\n/usr/local/lib/python3.8/site-packages/urllib3/util/connection.py:72: in create_connection\r\n for res in socket.getaddrinfo(host, port, family, socket.SOCK_STREAM):\r\n/usr/local/lib/python3.8/socket.py:918: in getaddrinfo\r\n for res in _socket.getaddrinfo(host, port, family, type, proto, flags):\r\nE socket.gaierror: [Errno 8] Name does not resolve\r\n\r\nDuring handling of the above exception, another exception occurred:\r\n/usr/local/lib/python3.8/site-packages/urllib3/connectionpool.py:703: in urlopen\r\n httplib_response = self._make_request(\r\n/usr/local/lib/python3.8/site-packages/urllib3/connectionpool.py:386: in _make_request\r\n self._validate_conn(conn)\r\n/usr/local/lib/python3.8/site-packages/urllib3/connectionpool.py:1040: in _validate_conn\r\n conn.connect()\r\n/usr/local/lib/python3.8/site-packages/urllib3/connection.py:358: in connect\r\n self.sock = conn = self._new_conn()\r\n/usr/local/lib/python3.8/site-packages/urllib3/connection.py:186: in _new_conn\r\n raise NewConnectionError(\r\nE urllib3.exceptions.NewConnectionError: <urllib3.connection.HTTPSConnection object at 0x830fa1e20>: Failed to establish a new connection: [Errno 8] Name does not resolve\r\n\r\nDuring handling of the above exception, another exception occurred:\r\n/usr/local/lib/python3.8/site-packages/requests/adapters.py:440: in send\r\n resp = conn.urlopen(\r\n/usr/local/lib/python3.8/site-packages/urllib3/connectionpool.py:785: in urlopen\r\n retries = retries.increment(\r\n/usr/local/lib/python3.8/site-packages/urllib3/util/retry.py:592: in increment\r\n raise MaxRetryError(_pool, url, error or ResponseError(cause))\r\nE urllib3.exceptions.MaxRetryError: HTTPSConnectionPool(host='www.materialsproject.org', port=443): Max retries exceeded with url: / (Caused by NewConnectionError('<urllib3.connection.HTTPSConnection object at 0x830fa1e20>: Failed to establish a new connection: [Errno 8] Name does not resolve'))\r\n\r\nDuring handling of the above exception, another exception occurred:\r\npymatgen/ext/tests/test_matproj.py:31: in <module>\r\n website_is_up = requests.get(\"https://www.materialsproject.org\").status_code == 200\r\n/usr/local/lib/python3.8/site-packages/requests/api.py:75: in get\r\n return request('get', url, params=params, **kwargs)\r\n/usr/local/lib/python3.8/site-packages/requests/api.py:61: in request\r\n return session.request(method=method, url=url, **kwargs)\r\n/usr/local/lib/python3.8/site-packages/requests/sessions.py:529: in request\r\n resp = self.send(prep, **send_kwargs)\r\n/usr/local/lib/python3.8/site-packages/requests/sessions.py:645: in send\r\n r = adapter.send(request, **kwargs)\r\n/usr/local/lib/python3.8/site-packages/requests/adapters.py:519: in send\r\n raise ConnectionError(e, request=request)\r\nE requests.exceptions.ConnectionError: HTTPSConnectionPool(host='www.materialsproject.org', port=443): Max retries exceeded with url: / (Caused by NewConnectionError('<urllib3.connection.HTTPSConnection object at 0x830fa1e20>: Failed to establish a new connection: [Errno 8] Name does not resolve'))\r\n_____________ ERROR collecting pymatgen/vis/tests/test_plotters.py _____________\r\npymatgen/vis/tests/test_plotters.py:18: in <module>\r\n with open(os.path.join(test_dir, \"LiCoO2_k_xanes.json\")) as fp:\r\nE FileNotFoundError: [Errno 2] No such file or directory: '/work/usr/ports/science/py-pymatgen/work-py38/pymatgen-2022.0.17/pymatgen/util/../../test_files/spectrum_test/LiCoO2_k_xanes.json'\r\n=============================== warnings summary ===============================\r\npymatgen/electronic_structure/dos.py:14\r\n /work/usr/ports/science/py-pymatgen/work-py38/pymatgen-2022.0.17/pymatgen/electronic_structure/dos.py:14: DeprecationWarning: Please use `value` from the `scipy.constants` namespace, the `scipy.constants.codata` namespace is deprecated.\r\n from scipy.constants.codata import value as _cd\r\n\r\npymatgen/core/spectrum.py:15\r\n /work/usr/ports/science/py-pymatgen/work-py38/pymatgen-2022.0.17/pymatgen/core/spectrum.py:15: DeprecationWarning: Please use `convolve1d` from the `scipy.ndimage` namespace, the `scipy.ndimage.filters` namespace is deprecated.\r\n from scipy.ndimage.filters import convolve1d\r\n\r\n../../../../../../../usr/local/lib/python3.8/site-packages/past/builtins/misc.py:45\r\n /usr/local/lib/python3.8/site-packages/past/builtins/misc.py:45: DeprecationWarning: the imp module is deprecated in favour of importlib; see the module's documentation for alternative uses\r\n from imp import reload\r\n\r\n-- Docs: https://docs.pytest.org/en/stable/how-to/capture-warnings.html\r\n=========================== short test summary info ============================\r\nSKIPPED [1] pymatgen/analysis/tests/test_functional_groups.py:19: OpenBabel not installed\r\nSKIPPED [1] pymatgen/io/tests/test_babel.py:18: OpenBabel not installed\r\n!!!!!!!!!!!!!!!!!!! Interrupted: 7 errors during collection !!!!!!!!!!!!!!!!!!!!\r\n=================== 2 skipped, 3 warnings, 7 errors in 5.77s ===================\r\n```",
"Please be specific about what data is missing from the distribution. Note that the test files are deliberately not included within the usual pymatgen distribution. The test dir alone is several GB because it contains output files from electronic structure software. It is not possible to include this in an install file (and it is not common practice). If you want the full suite, you can download directly from the Github repo. For network access, you can exclude all tests in the pymatgen/ext folder (this is just using the --exclude options. As far as I know, pymatgen/ext is the only one that communicates on the network. ",
"> Please be specific about what data is missing from the distribution\r\n\r\nThe exhaustive log was attached, the files were mentioned right there. For instance,\r\n- test_files/boltztrap2/vasprun.xml\r\n- test_files/vasprun.xml\r\n- test_files/spectrum_test/LiCoO2_k_xanes.json\r\n\r\n> Note that the test files are deliberately not included within the usual pymatgen distribution. The test dir alone is several GB\r\n\r\nThis is understandable, but in this case not skipping tests if data files required for them are missing is the problem.",
"For a code like pymatgen, most of the tests involve an external file of some sort. If we skip all tests that require things in test_files, we will be left with a tiny fraction of the code being tested.\r\n\r\nAs I said before, testing is meant to be done from the Github full repository, which includes the test files. It is not meant to be done from the distributed wheels or the tar.gz source. ",
"> If we skip all tests that require things in test_files, we will be left with a tiny fraction of the code being tested.\r\n\r\nThis is expected, but it is better than nothing, and definitely better than failing for unrelated reason, and requiring investigation from a maintainer like in this case. If introducing proper test skips would be too much work, there probably are other solution, such as failing all the tests early with some informative message, or removing tests from the distribution as well. That would be consistent at the very least.",
"The tests were never included in the distribution. Go ahead and download the tar.gz from https://pypi.org/project/pymatgen/ and see for yourself. Pretty sure there are no tests in there.",
"Ok, that's true for the latest version."
] | 2021-10-22T15:51:38
| 2022-04-27T08:42:50
|
2021-10-22T19:19:19Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
```python-3.8 -m pytest``` fails:
```
===> py38-pymatgen-2022.0.15 depends on file: /usr/local/bin/python3.8 - found
INTERNALERROR> Traceback (most recent call last):
INTERNALERROR> File "/usr/local/lib/python3.8/site-packages/_pytest/main.py", line 206, in wrap_session
INTERNALERROR> session.exitstatus = doit(config, session) or 0
INTERNALERROR> File "/usr/local/lib/python3.8/site-packages/_pytest/main.py", line 249, in _main
INTERNALERROR> config.hook.pytest_collection(session=session)
INTERNALERROR> File "/usr/local/lib/python3.8/site-packages/pluggy/hooks.py", line 286, in __call__
INTERNALERROR> return self._hookexec(self, self.get_hookimpls(), kwargs)
INTERNALERROR> File "/usr/local/lib/python3.8/site-packages/pluggy/manager.py", line 93, in _hookexec
INTERNALERROR> return self._inner_hookexec(hook, methods, kwargs)
INTERNALERROR> File "/usr/local/lib/python3.8/site-packages/pluggy/manager.py", line 84, in <lambda>
INTERNALERROR> self._inner_hookexec = lambda hook, methods, kwargs: hook.multicall(
INTERNALERROR> File "/usr/local/lib/python3.8/site-packages/pluggy/callers.py", line 208, in _multicall
INTERNALERROR> return outcome.get_result()
INTERNALERROR> File "/usr/local/lib/python3.8/site-packages/pluggy/callers.py", line 80, in get_result
INTERNALERROR> raise ex[1].with_traceback(ex[2])
INTERNALERROR> File "/usr/local/lib/python3.8/site-packages/pluggy/callers.py", line 182, in _multicall
INTERNALERROR> next(gen) # first yield
INTERNALERROR> File "/usr/local/lib/python3.8/site-packages/_pytest/warnings.py", line 150, in pytest_collection
INTERNALERROR> with catch_warnings_for_item(
INTERNALERROR> File "/usr/local/lib/python3.8/contextlib.py", line 113, in __enter__
INTERNALERROR> return next(self.gen)
INTERNALERROR> File "/usr/local/lib/python3.8/site-packages/_pytest/warnings.py", line 88, in catch_warnings_for_item
INTERNALERROR> _setoption(warnings, arg)
INTERNALERROR> File "/usr/local/lib/python3.8/site-packages/_pytest/warnings.py", line 27, in _setoption
INTERNALERROR> category = wmod._getcategory(category)
INTERNALERROR> File "/usr/local/lib/python3.8/warnings.py", line 262, in _getcategory
INTERNALERROR> raise _OptionError("invalid module name: %r" % (module,)) from None
INTERNALERROR> warnings._OptionError: invalid module name: 'pymatgen.io.vasp.inputs'
no tests ran in 0.45 seconds
*** Error code 3
```
FreeBSD 13
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2275/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2275/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2276
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2276/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2276/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2276/events
|
https://github.com/materialsproject/pymatgen/pull/2276
| 1,033,862,162
|
PR_kwDOACgets4tkGBS
| 2,276
|
Make `Simplex` `MSONable`
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I think there should be a test for this MSONable for Simplex.",
"Yes, this PR is not complete",
"\n[](https://coveralls.io/builds/43711331)\n\nCoverage increased (+0.001%) to 83.748% when pulling **7899caf4efbd3fc42004444eaa6cd5846f283c15 on simplex** into **2dd3a9febe770de8a6a6e1ebb980f4d04eb4ecce on master**.\n"
] | 2021-10-22T18:52:09
| 2022-10-10T17:53:58
|
2021-10-22T20:30:25Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
To enable `PhaseDiagram` to be JSON-serializable.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2276/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2276/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2276",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2276",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2276.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2276.patch",
"merged_at": "2021-10-22T20:30:25Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2277
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2277/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2277/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2277/events
|
https://github.com/materialsproject/pymatgen/pull/2277
| 1,034,209,991
|
PR_kwDOACgets4tlJ7G
| 2,277
|
Fix small cutoff neighbor
|
{
"login": "chc273",
"id": 22353204,
"node_id": "MDQ6VXNlcjIyMzUzMjA0",
"avatar_url": "https://avatars.githubusercontent.com/u/22353204?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/chc273",
"html_url": "https://github.com/chc273",
"followers_url": "https://api.github.com/users/chc273/followers",
"following_url": "https://api.github.com/users/chc273/following{/other_user}",
"gists_url": "https://api.github.com/users/chc273/gists{/gist_id}",
"starred_url": "https://api.github.com/users/chc273/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/chc273/subscriptions",
"organizations_url": "https://api.github.com/users/chc273/orgs",
"repos_url": "https://api.github.com/users/chc273/repos",
"events_url": "https://api.github.com/users/chc273/events{/privacy}",
"received_events_url": "https://api.github.com/users/chc273/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/43719978)\n\nCoverage decreased (-0.6%) to 83.101% when pulling **00ecc46d9242e4a92cf6438abf8bd41bbc2b5505 on chc273:fix_small_cutoff_neighbor** into **71c8ab8da6369970d5766113d9f688db78cba899 on materialsproject:master**.\n"
] | 2021-10-23T16:59:38
| 2021-10-26T16:01:27
|
2021-10-26T16:01:27Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
In the current cell list implementation for finding neighbors, the space is divided into ~O((cell length / cutoff)^3) cubes for fast neighbor finding. This causes issues when the user chooses unrealistic small or even zero cutoffs.
In this PR, I am doing the following:
1) For reasonable cutoff (default to > 1.0 Angstrom), I am calculating the neighbors directly using the algorithm.
2) For small cutoffs (e.g., < 1.0 Angstrom, including neg cutoffs), I am calculating the 1.0-cutoff neighbor list, and use a boolean mask to keep those neighbors having distances less than the cutoff afterward. This will also make sure that if the users input negative cutoff, the returned neighbors will be empty.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2277/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2277/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2277",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2277",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2277.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2277.patch",
"merged_at": "2021-10-26T16:01:27Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2278
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2278/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2278/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2278/events
|
https://github.com/materialsproject/pymatgen/pull/2278
| 1,034,272,984
|
PR_kwDOACgets4tlV86
| 2,278
|
update docstring of CutOffDictNN
|
{
"login": "ltalirz",
"id": 1053245,
"node_id": "MDQ6VXNlcjEwNTMyNDU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1053245?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ltalirz",
"html_url": "https://github.com/ltalirz",
"followers_url": "https://api.github.com/users/ltalirz/followers",
"following_url": "https://api.github.com/users/ltalirz/following{/other_user}",
"gists_url": "https://api.github.com/users/ltalirz/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ltalirz/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ltalirz/subscriptions",
"organizations_url": "https://api.github.com/users/ltalirz/orgs",
"repos_url": "https://api.github.com/users/ltalirz/repos",
"events_url": "https://api.github.com/users/ltalirz/events{/privacy}",
"received_events_url": "https://api.github.com/users/ltalirz/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/43721197)\n\nCoverage decreased (-0.6%) to 83.101% when pulling **09ac6172c9606ebaa99fa05294ff6d57ba7187e0 on ltalirz:update-cutoff-docs** into **71c8ab8da6369970d5766113d9f688db78cba899 on materialsproject:master**.\n",
"@chc273 Has fixed some of the NN calculations. I think you can retest your old implementation.",
"Thanks @shyuep - sorry for the delay on my side; a couple of other tasks popped into existence, I'll get to it as soon as I can",
"by the way, this PR can be merged without further ado",
"Thanks! Sorry for delay here too"
] | 2021-10-23T22:03:44
| 2021-11-08T23:58:38
|
2021-11-08T23:58:08Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
follow-up on @mkhorton 's comments
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2278/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2278/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2278",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2278",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2278.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2278.patch",
"merged_at": "2021-11-08T23:58:07Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2279
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2279/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2279/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2279/events
|
https://github.com/materialsproject/pymatgen/issues/2279
| 1,035,296,191
|
I_kwDOACgets49tV2_
| 2,279
|
Support for FPLO lattice convention
|
{
"login": "mueslo",
"id": 847751,
"node_id": "MDQ6VXNlcjg0Nzc1MQ==",
"avatar_url": "https://avatars.githubusercontent.com/u/847751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mueslo",
"html_url": "https://github.com/mueslo",
"followers_url": "https://api.github.com/users/mueslo/followers",
"following_url": "https://api.github.com/users/mueslo/following{/other_user}",
"gists_url": "https://api.github.com/users/mueslo/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mueslo/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mueslo/subscriptions",
"organizations_url": "https://api.github.com/users/mueslo/orgs",
"repos_url": "https://api.github.com/users/mueslo/repos",
"events_url": "https://api.github.com/users/mueslo/events{/privacy}",
"received_events_url": "https://api.github.com/users/mueslo/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"The link doesn't work. Not sure what FLPO is. But in general, the lattice vectors should not have any impact on any analysis.",
"[FPLO is a major DFT code](https://www.fplo.de/). Lattice vectors do have impact on analysis, e.g. if you use the cartesian k-vectors output by the DFT code, they will make no sense in the lattice generated by `pymatgen`, since their bases are not identical - but only in certain circumstances (e.g. when gamma is not 90°).\r\n\r\nI've double-checked, and the link works from here, can you be more specific? I think you have to click the \"I am not a spammer\" button, and then reload the original link (unnecessarily complex, but works).\r\n\r\nFor convenience, I will quote it here:\r\n\r\n> Hexagonal/trigonal lattices are fixed as c=z, b=y, a in xy-plane at 30 degree\r\n> clockwise from x.\r\n> Rhomobohedral is the rhombo bravais cell of above mentioned trigonal cell.\r\n> There are two settings, either as simple cell or as centered trigonal cell.\r\n> But\r\n> both have the same orientation with respect to the cartesian axes.\r\n> cubic, ortho and tetra are along cartesian axes (tetra: c=z).\r\n> monoclinic: c=z, b=y, a in in xz=plane\r\n> triclinic are: c=z, b in yz=plane, a free\r\n> \r\n> In general centered lattices are based on conventional cells as described\r\n> above.\r\n> Reciprocial cells are fully determined by the duality relation of both\r\n> lattices.",
"<img width=\"1222\" alt=\"Screen Shot 2021-10-25 at 9 18 12 AM\" src=\"https://user-images.githubusercontent.com/577107/138732809-b3c1724c-7757-4f2d-92a0-8127a389fc14.png\">\r\nInitially the link brought me to this. But after reclicking it several times, it finally brought me to the definition.\r\n\r\nI have no objections if someone wanted to implement the FLPO convention in pymatgen. I don't think it is a lot of effort. Just needs an additional \"convention\" parameter, which will allow other conventions to be specified too. Personally, I prefer if codes output things in formats that are invariant to the definition of the lattice vectors, which are honestly arbitrary. Alternatively, you can always parse the lattice vectors from the FLPO output itself and construct the lattice directly, e.g., Lattice(vectors), rather than using lattice parameters.\r\n\r\n",
"I will either use a workaround (currently writing something similar to your suggestion) or write a new convention argument while keeping the `vesta` argument for the time being for compatibility until it is deprecated, with something like \r\n\r\n if vesta: \r\n convention = 'vesta'\r\n\r\n Either way, I will come back to this issue and delete it or reference the pull request. Thank you for your input!\r\n\r\nI definitely agree that it would be nice to get invariant definitions, but sadly FPLO nowhere outputs the lattice matrix, and I'm currently trying to go through the fortran code to find the exact code used to build the lattice so I'm sure it works in all cases."
] | 2021-10-25T15:45:55
| 2021-10-25T16:31:45
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
https://github.com/materialsproject/pymatgen/blob/fcb84a1e129da3a3010b9f6b9cf18beddf39f245/pymatgen/core/lattice.py#L310-L363
It would be great if pymatgen also supported FPLO conventions on lattice generation from parameters.
They are as follows: https://www.listserv.dfn.de/sympa/arc/fplo-users/2020-01/msg00002.html
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2279/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2279/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2280
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2280/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2280/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2280/events
|
https://github.com/materialsproject/pymatgen/issues/2280
| 1,037,382,411
|
I_kwDOACgets491TML
| 2,280
|
🪛 [RFC] Returning index of vbm and cbm
|
{
"login": "ezpzbz",
"id": 13301031,
"node_id": "MDQ6VXNlcjEzMzAxMDMx",
"avatar_url": "https://avatars.githubusercontent.com/u/13301031?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ezpzbz",
"html_url": "https://github.com/ezpzbz",
"followers_url": "https://api.github.com/users/ezpzbz/followers",
"following_url": "https://api.github.com/users/ezpzbz/following{/other_user}",
"gists_url": "https://api.github.com/users/ezpzbz/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ezpzbz/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ezpzbz/subscriptions",
"organizations_url": "https://api.github.com/users/ezpzbz/orgs",
"repos_url": "https://api.github.com/users/ezpzbz/repos",
"events_url": "https://api.github.com/users/ezpzbz/events{/privacy}",
"received_events_url": "https://api.github.com/users/ezpzbz/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I have no objections to this really, but I would ask that you add a kwarg such as \"include_kpoint_and_index=False\". By default, the return will still be the same. If that kwarg is set to true, the kpoint and index will be returned as per your suggestion. That will ensure that the method itself retains backward compatibility with code already out there using it.",
"I totally agree that this is a better approach @shyuep."
] | 2021-10-27T12:49:44
| 2021-10-27T15:59:30
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Is your feature request related to a problem? Please describe.**
Currently, [`eigenvalue_band_properties`](https://github.com/materialsproject/pymatgen/blob/fcb84a1e129da3a3010b9f6b9cf18beddf39f245/pymatgen/io/vasp/outputs.py#L958) does not return the kpoint and eigenvalue index for `vbm` and `cbm`. The former one is already there and just needs to be returned while the eigenvalue index requires a little tweak to be captured and returned.
**Use-case:** I use [`vaspkit`](https://vaspkit.com/index.html) to visualize the real-part of wavefunction at specific eigenvalues (more specifically `vbm` and `cbm`). The above mentioned indexes makes the life much easier by quickly getting the kpoint and eigenvalue indexes and passing them to `vaspkit`.
**Describe the solution you'd like**
I'm currently using the modified version of mentioned method to get this done:
```python
def eigenvalue_band_properties(vrun, occu_tol):
vbm = -float("inf")
vbm_kpoint = None
vbm_index = None
cbm = float("inf")
cbm_kpoint = None
cbm_index = None
for spin, d in vrun.eigenvalues.items():
for k, val in enumerate(d):
for index, (eigenval, occu) in enumerate(val):
if occu > occu_tol and eigenval > vbm:
vbm = eigenval
vbm_kpoint = k
vbm_index = index
elif occu <= occu_tol and eigenval < cbm:
cbm = eigenval
cbm_kpoint = k
cbm_index = index
return max(cbm - vbm, 0), cbm, vbm, vbm_kpoint, cbm_kpoint, vbm_index, cbm_index
```
,where in the third `for loop`, I enumerate the `val` and record the `index`.
**Describe alternatives you've considered**
I have not.
**Additional context**
I have opened this issue to initiate discussion on this feature and request for comments. If this makes sense, I would be happy to open a PR to address this.
Cheers,
Pezhman
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2280/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2280/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2281
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2281/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2281/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2281/events
|
https://github.com/materialsproject/pymatgen/issues/2281
| 1,037,881,803
|
I_kwDOACgets493NHL
| 2,281
|
Ion.from_formula has unexpected behavior when +/- precede charge number
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-10-27T21:34:48
| 2021-11-09T19:30:58
|
2021-11-09T19:30:58Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Ion.from_formula should correctly recognize formulae in which the `+` or `-` sign precede the charge number. See also #2204
**Describe the bug**
```
>>> ion = Ion.from_formula('La[3+]')
>>> ion.charge
3.0
```
```
>>> ion = Ion.from_formula('La[+3]')
>>> ion.charge
1.0
```
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2281/reactions",
"total_count": 1,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 1
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2281/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2282
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2282/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2282/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2282/events
|
https://github.com/materialsproject/pymatgen/issues/2282
| 1,038,920,381
|
I_kwDOACgets497Kq9
| 2,282
|
Add Composition.replace()
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Composition is meant to be immutable. Which is why it inherits Mapping and Hashable and not dict or MutableMapping. Composition objects are probably the one of the most used keys in dictionaries and sets. So in place modifications are *not* allowed. FYI, the above can be easily done using `Composition({(k if k.symbol != \"Fe\" else \"Mn\"): v for k, v in comp.items()})`. ",
"Just a note that I have no objection if you want to create a method to do substitutions that return a *new* composition object. But in-place modifications are not supported.",
"Probably should have stated explicitly that I didn't want this op to be in-place."
] | 2021-10-28T20:19:45
| 2021-10-28T20:54:30
|
2021-10-28T20:52:54Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Is your feature request related to a problem? Please describe.**
`Composition` does not support elemental substitutions.
**Describe the solution you'd like**
New method `Composition("Fe2O3").replace({"Fe": "Mg"})` similar to [`Structure.replace_species()`](https://github.com/materialsproject/pymatgen/blob/v2022.0.15/pymatgen/core/structure.py#L462-L494).
**Describe alternatives you've considered**
None.
**Additional context**
Useful when working with [data-mined substitution matrices](https://iopscience.iop.org/article/10.1088/1367-2630/18/9/093011).
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2282/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2282/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2283
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2283/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2283/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2283/events
|
https://github.com/materialsproject/pymatgen/pull/2283
| 1,038,964,019
|
PR_kwDOACgets4t0fWD
| 2,283
|
Add PatchedPhaseDiagram to allow for handling of Large Disjoint Chemical systems
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"```\r\ne_h = np.array(e_h)\r\ne_h_mp = np.array(e_h_mp)\r\nidx = np.argwhere(np.invert(np.isclose(e_h, e_h_mp)))\r\nprint(len(idx))\r\nprint(e_h[idx][2:3])\r\nprint(e_h_mp[idx][2:3])\r\nprint([ppd.get_decomp_and_e_above_hull(mp_data[i]) for i in idx[2:3, 0]])\r\nprint([mp_data[i] in ppd.stable_entries for i in idx[2:3, 0]])\r\nprint([mp_data[i] for i in idx[2:3, 0]])\r\n```\r\n```\r\n68\r\n[[2.41988421]]\r\n[[2.68688421]]\r\n[({mp-23154 ComputedEntry - Br4 (Br)\r\nEnergy (Uncorrected) = -6.5477 eV (-1.6369 eV/atom)\r\nCorrection = 0.0000 eV (0.0000 eV/atom)\r\nEnergy (Final) = -6.5477 eV (-1.6369 eV/atom)\r\nEnergy Adjustments:\r\n None\r\nParameters:\r\n run_type = GGA\r\n is_hubbard = False\r\n pseudo_potential = {'functional': 'PBE', 'labels': ['Br'], 'pot_type': 'paw'}\r\n hubbards = {}\r\n potcar_symbols = ['PBE Br']\r\n oxide_type = None\r\nData:\r\n oxide_type = None\r\n e_above_hull = 0: 0.5, mp-25 ComputedEntry - N8 (N2)\r\nEnergy (Uncorrected) = -66.6920 eV (-8.3365 eV/atom)\r\nCorrection = 0.0000 eV (0.0000 eV/atom)\r\nEnergy (Final) = -66.6920 eV (-8.3365 eV/atom)\r\nEnergy Adjustments:\r\n None\r\nParameters:\r\n run_type = GGA\r\n is_hubbard = False\r\n pseudo_potential = {'functional': 'PBE', 'labels': ['N'], 'pot_type': 'paw'}\r\n hubbards = {}\r\n potcar_symbols = ['PBE N']\r\n oxide_type = None\r\nData:\r\n oxide_type = None\r\n e_above_hull = 0: 0.5}, 2.419884208749999)]\r\n[False]\r\n[mp-1059160 ComputedEntry - Br1 N1 (BrN)\r\nEnergy (Uncorrected) = -4.2387 eV (-2.1193 eV/atom)\r\nCorrection = -0.8950 eV (-0.4475 eV/atom)\r\nEnergy (Final) = -5.1337 eV (-2.5668 eV/atom)\r\nEnergy Adjustments:\r\n MP2020 anion correction (Br): -0.5340 eV (-0.2670 eV/atom)\r\n MP2020 anion correction (N): -0.3610 eV (-0.1805 eV/atom)\r\nParameters:\r\n run_type = GGA\r\n is_hubbard = False\r\n pseudo_potential = {'functional': 'PBE', 'labels': ['Br', 'N'], 'pot_type': 'paw'}\r\n hubbards = {}\r\n potcar_symbols = ['PBE Br', 'PBE N']\r\n oxide_type = None\r\nData:\r\n oxide_type = None\r\n e_above_hull = 2.6868842087499996\r\n oxidation_states = {'Br': -1.0, 'N': 1.0}]\r\n/usr/local/lib/python3.7/dist-packages/pymatgen/analysis/phase_diagram.py:1631: UserWarning: No suitable PhaseDiagrams found for Br1 N1. Using SLSQP to find decomposition\r\n warnings.warn(str(e) + \" Using SLSQP to find decomposition\")\r\n```\r\n\r\nFor this material https://materialsproject.org/materials/mp-1059160/#corrections-eqn the final energy per atom on the website seems inconsistent",
"appears to be a bug with the `PhaseDiagram` serialisation changes leading to test errors that are unrelated to this PR",
"Tests are currently passing in the main branch (aside from two or three recent unrelated commits), so the bug is likely in this PR?",
"I see the change now, I removed the `dict()` around `el_refs`, will revert tomorrow ",
"No worries. The conversion to a list happens because you can't serialize a dict with `Element` keys. The change to serialization was to allow the computed information to be included when converting the PD to JSON, so as to speed up the reverse process when you have many thousands of PDs saved.",
"In my workflow I just pickle this big ppd but for all of the data ~400k it does get pretty large.\n\nI am not sure about how to serialise to json atm as then it will make a bunch of duplicates of entries inside the patches whereas building from scratch we don't (as far as I can tell) make hard copies.",
"\n[](https://coveralls.io/builds/50559312)\n\nCoverage decreased (-0.06%) to 63.525% when pulling **2034520d4660812e774f1b450f7ede8f2b647916 on CompRhys:ppd** into **85796d24190aca51a2674a7895430eb7c561b0a9 on materialsproject:master**.\n",
"Just to update this I am pretty sure there is no nice solution to the problem I outline re serialising to json when we have an object containing multiple references to other objects inside itself. Pickle has the correct behaviour in that it serialises a single copy and then keeps track of references for the copies.\r\n\r\nAs such I think this PR is okay to be merged provided you're happy with the serialisation and de-serialisation to/from json having to repeat the calculations of the hulls/patches. ",
"Very excited about this contribution - I've been using OQMD's implementation of GCLP-based phase diagrams to handle chemically diverse datasets, but would love to simplify my stack by using this instead.",
"@montoyjh do you want to give a run-down of the pros/cons of the GCLP approach for the uninitiated?\r\n\r\n@CompRhys only just saw this PR was marked as good-to-go. Will review shortly.",
"> @montoyjh do you want to give a run-down of the pros/cons of the GCLP approach for the uninitiated?\r\n\r\nI haven't looked at GCLP - my approach here could easily be a sub-optimal approach but it was intended to work with the SLSQP approach I previously implemented to get e_above_hull in chemical systems without phase diagrams for supersets of the chemical system.\r\n\r\n> @CompRhys only just saw this PR was marked as good-to-go. Will review shortly.\r\n\r\nNo worries, I was investigating some pickling issues last week unrelated and so came back to this PR as I was thinking about alternatives to pickling and realized that I didn't change the title back in Oct/Jan.\r\n\r\n-- closing was accidental",
"Re: GCLP - it's a linear programming approach to energy minimization that does so for each queried composition, see [this paper from OQMD](https://onlinelibrary.wiley.com/doi/abs/10.1002/adma.200700843) and a [recent paper](https://chemrxiv.org/engage/api-gateway/chemrxiv/assets/orp/resource/item/629135424d8b597d2ea14f76/original/environmental-stability-of-crystals-why-be-greedy-when-you-can-be-exact.pdf) that describes a method that's basically the same a little more explicitly. I think if you want to come up with the stability of *any* composition in a large chemical space (say a 12-element composition in a 96 dimensional space) - GCLP scales better, but on a practical basis we only ever deal with <8 elements and I think it's easier to understand a strategy like the one that's in this PR - plus having the convex hulls of the subspaces is useful.",
"Thanks @montoyjh :)",
"We have an upfront cost to generate the ppd and then getting the decomposition is cheap for entries within a patch and for entries outside a patch I think the SLSQP I mentioned is just an algorithm to solve the GCLP as opposed to using gurobi to solve the LP as in the second paper.\r\n\r\nIn the paper they claim that using gurobi takes ~10 minutes to solve the LP whereas the approach here with the combination of convex hull, simplex method and SLSQP would take a few hours. ",
"Thanks for this, it's excellent! Apologies for the delay, longer PRs always take a moment to review."
] | 2021-10-28T21:22:26
| 2022-08-11T03:46:17
|
2022-07-21T00:54:29Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Handler Class to allow for thermodynamic properties to be calculated across large chemically diverse phase diagrams whilst only generating relevant phase diagrams once.
Designed to inherit as much as possible and be modular to allow additional functionality to be approximated if solutions can be devised.
```python
m = MPRester(MP_API_KEY)
# Materials project provides a python API via pymatgen
# API key obtained from https://materialsproject.org/open
#
mp_data = m.get_entries(
chemsys_formula_id_criteria={"nelements":{"$lt":3}},
inc_structure=False,
)
```
```bash
100%|██████████| 21160/21160 [00:14<00:00, 1504.11it/s]
```
```python
%timeit ppd = PatchedPhaseDiagram(mp_data, verbose=True)
```
```bash
100%|██████████| 2127/2127 [00:12<00:00, 172.51it/s]
100%|██████████| 2127/2127 [00:12<00:00, 174.56it/s]
100%|██████████| 2127/2127 [00:12<00:00, 174.54it/s]
100%|██████████| 2127/2127 [00:12<00:00, 172.13it/s]
100%|██████████| 2127/2127 [00:12<00:00, 169.93it/s]
100%|██████████| 2127/2127 [00:12<00:00, 172.21it/s]
1 loop, best of 5: 26.2 s per loop
```
c.f. the equivalent `PhaseDiagram` is intractable.
When calling thermodynamic properties, e.g. `e_above_hull`, class first looks to see if sufficient pd exists in patch library, if not we use the `_get_slsqp_decomp` method to get the decomp on the minimal set of competing entries.
If I compare the `e_above_hull` I calculate there are 68 materials where they differ from the MP values. I think these are most likely due to manual fixes/rounding in MP (see below)
### TODO
- [ ] ~~Refactor to allow serialisation in a manner that mimics new `PhaseDiagram`~~
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2283/reactions",
"total_count": 2,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 1,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2283/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2283",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2283",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2283.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2283.patch",
"merged_at": "2022-07-21T00:54:29Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2284
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2284/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2284/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2284/events
|
https://github.com/materialsproject/pymatgen/pull/2284
| 1,039,488,884
|
PR_kwDOACgets4t2KcF
| 2,284
|
Add Composition.replace()
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-10-29T11:57:42
| 2021-10-29T18:05:45
|
2021-10-29T17:04:36Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Addresses #2282.
I originally attempted to handle `SpeciesLike` rather than just `str` as input, i.e.
```py
elem_map: Dict[SpeciesLike, Union[SpeciesLike, Dict[SpeciesLike, float]]]
```
instead of the current
```py
elem_map: Dict[str, Union[str, Dict[str, float]]]
```
but it turned into much more of a hassle with perhaps not much extra value. What's your take?
## Checklist
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2284/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2284/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2284",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2284",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2284.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2284.patch",
"merged_at": "2021-10-29T17:04:36Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2285
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2285/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2285/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2285/events
|
https://github.com/materialsproject/pymatgen/pull/2285
| 1,039,579,182
|
PR_kwDOACgets4t2dWa
| 2,285
|
Fix oddly split strings and a few typos
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Dropped some unused variables that `flake8` pre-commit hook was complaining about.",
"\n[](https://coveralls.io/builds/43882122)\n\nCoverage decreased (-0.6%) to 83.104% when pulling **b180cac685a432ed62ea0a80926ce4bb61ef0651 on janosh:str-fixes** into **acfe5899ee50add1e2a0dd6385ee4fba78122e0f on materialsproject:master**.\n",
"Thanks @janosh !"
] | 2021-10-29T13:38:24
| 2021-11-09T19:34:48
|
2021-11-09T19:32:19Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
`"Lots of " "oddly split" " string in pymatgen"`, presumably as a result of `black` formatting.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2285/reactions",
"total_count": 1,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 1,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2285/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2285",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2285",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2285.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2285.patch",
"merged_at": "2021-11-09T19:32:19Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2286
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2286/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2286/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2286/events
|
https://github.com/materialsproject/pymatgen/pull/2286
| 1,040,266,805
|
PR_kwDOACgets4t4cgS
| 2,286
|
ElementBase.X drop NaN warning
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/43983158)\n\nCoverage decreased (-0.6%) to 83.102% when pulling **85898a64447e5103c1e5dc0c256d5b07e0991bbd on janosh:silence-elem-x** into **acfe5899ee50add1e2a0dd6385ee4fba78122e0f on materialsproject:master**.\n",
"No the warning needs to be there. BEcause a Nan is still a float, it can lead to strange errors if you use it in the wrong way. If you wish, you can set a proper warning type and on your end, you can capture all the warnings. I personally found it useful to know if I have elements that do not have electronegativities in my dataset.",
"> BEcause a Nan is still a float, it can lead to strange errors if you use it in the wrong way.\r\n\r\nThen it would be better to return `None` to avoid tripping up inexperienced users.",
"It used to return None. That caused more problems because Noke is a type that needs to be caught. Which means everyone who uses electro negativity needs to use try catch. ",
"Could also use an electronegativity scale that assigns values to all elements:\r\n\r\nhttps://wikipedia.org/wiki/Electronegativity#Allen_electronegativity",
"You can of course add Allen electro negativity as a separate property. But most people's understanding of electronegativity refers to Pauling. "
] | 2021-10-30T18:24:20
| 2021-11-18T19:29:04
|
2021-11-03T13:44:52Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
The warning about missing electronegativities is by far the most prevalent one in my use of pymatgen and it has never once been useful. Would be great to get rid of it.
It comes up all the time and usually repeated a dozen times or so when iterating over a dataframe column involving creation of new `Elements()`.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2286/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2286/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2286",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2286",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2286.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2286.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2287
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2287/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2287/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2287/events
|
https://github.com/materialsproject/pymatgen/pull/2287
| 1,040,348,095
|
PR_kwDOACgets4t4sL3
| 2,287
|
Ion bugfixes and enhancements
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/43894854)\n\nCoverage decreased (-0.6%) to 83.115% when pulling **d378d277e70ef8b9d6b48ebb4ba0b82b9b3f7ee1 on rkingsbury:ion_fixes** into **acfe5899ee50add1e2a0dd6385ee4fba78122e0f on materialsproject:master**.\n",
"Thanks for this PR @rkingsbury ! looks great.\r\n\r\nOne concern is the proliferation of special cases for organics, I wonder if there's a better long-term solution to this that's more general/transferrable."
] | 2021-10-31T01:01:26
| 2022-06-08T22:25:54
|
2021-11-09T19:30:58Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This PR contains some miscellaneous bugfixes and enhancements for the `Ion` class.
* Fixes #2281
* Partially address #2204 by overloading `get_reduced_formula_and_factor` to provide special formula handling for ions. The `reduced_formula` for hydroxide, hydrogen peroxide, acetate, acetic acid, and small alcohols are now more consistent with the way these species are usually written.
* Implement a `utils.string.charge_string` method to make the representation of ion charges more consistent across `formula`, `reduced_formula`, etc.
* Make representation of `Ion` formulae more consistent. By default, charges are now always appended in brackets, with the sign preceding the magnitude, e.g. `Na[+1]` or `SO4[-2]`. This provides unambiguous differentiation between stoichiometric coefficients and charges, and will facilitate using `Ion.reduced_formula` as database keys in, e.g., the Pourbaix ion data. For uncharged `Ion`, the formula is always followed by `(aq)`.
* Change display of ions that may be hydrated metal complexes, e.g. `Zr(OH)4(aq)` becomes `ZrO2.2H2O(aq)`
* Add extra tests for `to_latex_string` and `Ion.from_formula`
* Add type hinting
## Additional dependencies introduced (if any)
None
## Checklist
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2287/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2287/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2287",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2287",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2287.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2287.patch",
"merged_at": "2021-11-09T19:30:58Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2288
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2288/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2288/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2288/events
|
https://github.com/materialsproject/pymatgen/issues/2288
| 1,040,378,938
|
I_kwDOACgets4-Auw6
| 2,288
|
How to install and use zeopp modules
|
{
"login": "ctc-er",
"id": 56148779,
"node_id": "MDQ6VXNlcjU2MTQ4Nzc5",
"avatar_url": "https://avatars.githubusercontent.com/u/56148779?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ctc-er",
"html_url": "https://github.com/ctc-er",
"followers_url": "https://api.github.com/users/ctc-er/followers",
"following_url": "https://api.github.com/users/ctc-er/following{/other_user}",
"gists_url": "https://api.github.com/users/ctc-er/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ctc-er/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ctc-er/subscriptions",
"organizations_url": "https://api.github.com/users/ctc-er/orgs",
"repos_url": "https://api.github.com/users/ctc-er/repos",
"events_url": "https://api.github.com/users/ctc-er/events{/privacy}",
"received_events_url": "https://api.github.com/users/ctc-er/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Please refer to pymatgen.org's instructions. But honestly, these questions should be posed to the Zeo+ developers rather than to pymatgen since we only provide an interface to it."
] | 2021-10-31T05:17:06
| 2021-10-31T15:07:12
|
2021-10-31T15:07:12Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
when I open the address in http://zeoplusplus.org/, it was a cpp source and I can not using it. could you tell me how to install it in the py? thank you for your patience.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2288/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2288/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2289
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2289/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2289/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2289/events
|
https://github.com/materialsproject/pymatgen/issues/2289
| 1,044,308,093
|
I_kwDOACgets4-PuB9
| 2,289
|
Load bonds from .cif to make StructureGraph
|
{
"login": "alex-l-m",
"id": 35587576,
"node_id": "MDQ6VXNlcjM1NTg3NTc2",
"avatar_url": "https://avatars.githubusercontent.com/u/35587576?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/alex-l-m",
"html_url": "https://github.com/alex-l-m",
"followers_url": "https://api.github.com/users/alex-l-m/followers",
"following_url": "https://api.github.com/users/alex-l-m/following{/other_user}",
"gists_url": "https://api.github.com/users/alex-l-m/gists{/gist_id}",
"starred_url": "https://api.github.com/users/alex-l-m/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/alex-l-m/subscriptions",
"organizations_url": "https://api.github.com/users/alex-l-m/orgs",
"repos_url": "https://api.github.com/users/alex-l-m/repos",
"events_url": "https://api.github.com/users/alex-l-m/events{/privacy}",
"received_events_url": "https://api.github.com/users/alex-l-m/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"That would be nice. Can you point to the CIF specification? Last time I looked into it, it was unclear to me how CIF defined bonds that crossed periodic boundaries.\r\n\r\nIf we do add, I think StructureGraph -> CIF export would also be useful.",
"The bond information in CIF files is defined by the [GEOM_BOND](https://www.iucr.org/__data/iucr/cifdic_html/1/cif_core.dic/Cgeom_bond.html) category. It's kind of annoying to set up due to the non-intuitive periodicity flags, but it can be done. Bonding across boundaries is specified using a flag of the format `1_XYZ`. If I recall correctly (it's been a long time...), it's such that any number after the underscore other than 5 indicates a cross over a periodic boundary. For example, I think `1_545` specifies that it crosses the boundary in the -y dimension, whereas `1_556` specifies that it crosses the boundary in the +z dimension. Or something very similar to this.",
"Thanks @arosen93, that info on the periodicity labels `_geom_bond_site_symmetry_1` etc. was exactly what I was missing. More info on that tag here: https://www.iucr.org/__data/iucr/cifdic_html/1/cif_core.dic/Igeom_bond_site_symmetry_.html ",
"@alex-l-m do you have a sample CIF file with bonding information that we could incorporate into pymatgen as a test case?",
"Hi, this would be a great feature to have! I am attaching an example cif with the bond info :)\r\n\r\n```\r\n# CIF file generated by openbabel 2.4.90, see http://openbabel.sf.net\r\ndata_I\r\n_chemical_name_common 'str_m12_o9_o13_f0_bcu.sym.134'\r\n_cell_length_a 15.0575\r\n_cell_length_b 15.1814\r\n_cell_length_c 16.8456\r\n_cell_angle_alpha 108.912\r\n_cell_angle_beta 105.512\r\n_cell_angle_gamma 109.889\r\n_space_group_name_H-M_alt 'P 1'\r\n_space_group_name_Hall 'P 1'\r\nloop_\r\n _symmetry_equiv_pos_as_xyz\r\n x,y,z\r\nloop_\r\n _atom_site_label\r\n _atom_site_type_symbol\r\n _atom_site_fract_x\r\n _atom_site_fract_y\r\n _atom_site_fract_z\r\n _atom_site_occupancy\r\n C0 C 0.08427 0.29217 0.08263 1.000\r\n C1 C 0.91134 0.72437 0.96799 1.000\r\n C2 C 0.02437 0.58531 0.74592 1.000\r\n C3 C 0.96994 0.53523 0.27556 1.000\r\n C4 C 0.76020 0.19775 0.72483 1.000\r\n C5 C 0.33697 0.78219 0.25411 1.000\r\n C6 C 0.30651 0.47935 0.98438 1.000\r\n C7 C 0.63846 0.42759 0.93764 1.000\r\n C8 C 0.91112 0.93227 0.04701 1.000\r\n C9 C 0.09230 0.11281 0.10333 1.000\r\n C10 C 0.00420 0.92694 0.05744 1.000\r\n C11 C 0.99663 0.11391 0.07721 1.000\r\n C12 C 0.09743 0.02056 0.08989 1.000\r\n C13 C 0.90545 0.02428 0.05526 1.000\r\n C14 C 0.71802 0.95340 0.95291 1.000\r\n C15 C 0.28234 0.07865 0.18993 1.000\r\n C16 C 0.60326 0.99837 0.01510 1.000\r\n C17 C 0.40056 0.05270 0.12294 1.000\r\n C18 C 0.48628 0.56359 0.27606 1.000\r\n C19 C 0.50672 0.32366 0.64285 1.000\r\n C20 C 0.48091 0.54814 0.18811 1.000\r\n C21 C 0.49355 0.33012 0.72323 1.000\r\n C22 C 0.93468 0.52380 0.74922 1.000\r\n C23 C 0.06291 0.55049 0.26576 1.000\r\n C24 C 0.45126 0.44758 0.12025 1.000\r\n C25 C 0.47645 0.41134 0.77444 1.000\r\n C26 C 0.42986 0.36334 0.14243 1.000\r\n C27 C 0.47005 0.48446 0.74216 1.000\r\n C28 C 0.43482 0.37876 0.23018 1.000\r\n C29 C 0.48288 0.47782 0.66158 1.000\r\n C30 C 0.46059 0.47795 0.29728 1.000\r\n C31 C 0.50376 0.39938 0.61206 1.000\r\n C32 C 0.51643 0.40195 0.00422 1.000\r\n C33 C 0.39120 0.44291 0.88187 1.000\r\n C34 C 0.53471 0.40743 0.92849 1.000\r\n C35 C 0.38375 0.45256 0.96522 1.000\r\n C36 C 0.46754 0.42052 0.86238 1.000\r\n C37 C 0.44639 0.43058 0.02682 1.000\r\n C38 C 0.61880 0.94325 0.94027 1.000\r\n C39 C 0.38174 0.09070 0.20079 1.000\r\n C40 C 0.95912 0.58124 0.36227 1.000\r\n C41 C 0.02800 0.63229 0.68164 1.000\r\n C42 C 0.12933 0.71851 0.61392 1.000\r\n C43 C 0.85183 0.56103 0.44638 1.000\r\n C44 C 0.77698 0.20321 0.81194 1.000\r\n C45 C 0.28065 0.78425 0.17571 1.000\r\n C46 C 0.12238 0.70603 0.69081 1.000\r\n C47 C 0.86035 0.54880 0.36404 1.000\r\n C48 C 0.93616 0.59493 0.60393 1.000\r\n C49 C 0.04850 0.64882 0.44657 1.000\r\n C50 C 0.94195 0.60944 0.52789 1.000\r\n C51 C 0.04031 0.66253 0.52986 1.000\r\n C52 C 0.71958 0.10268 0.63794 1.000\r\n C53 C 0.44598 0.85643 0.32010 1.000\r\n C54 C 0.59148 0.03144 0.39124 1.000\r\n C55 C 0.61668 0.91305 0.55199 1.000\r\n C56 C 0.49966 0.94972 0.31608 1.000\r\n C57 C 0.68440 0.00306 0.63561 1.000\r\n C58 C 0.49278 0.84030 0.39432 1.000\r\n C59 C 0.70089 0.10975 0.55462 1.000\r\n C60 C 0.58186 0.92296 0.47121 1.000\r\n C61 C 0.62833 0.02099 0.47142 1.000\r\n C62 C 0.68705 0.06412 0.10257 1.000\r\n C63 C 0.32013 0.00309 0.03400 1.000\r\n C64 C 0.78709 0.07671 0.11504 1.000\r\n C65 C 0.22064 0.99138 0.02275 1.000\r\n C66 C 0.27673 0.55014 0.96545 1.000\r\n C67 C 0.69672 0.39596 0.98754 1.000\r\n C68 C 0.80302 0.02105 0.04027 1.000\r\n C69 C 0.20082 0.02771 0.10082 1.000\r\n C70 C 0.78225 0.29834 0.73432 1.000\r\n C71 C 0.27441 0.68998 0.25265 1.000\r\n C72 C 0.25528 0.44334 0.03211 1.000\r\n C73 C 0.70404 0.49802 0.91860 1.000\r\n C74 C 0.10642 0.60045 0.82024 1.000\r\n C75 C 0.89016 0.46115 0.18837 1.000\r\n C76 C 0.91242 0.20510 0.00246 1.000\r\n C77 C 0.08457 0.80538 0.05117 1.000\r\n C78 C 0.99613 0.20051 0.05639 1.000\r\n C79 C 0.99949 0.82293 0.02891 1.000\r\n H80 H 0.16125 0.32392 0.13610 1.000\r\n H81 H 0.83135 0.70787 0.93520 1.000\r\n H82 H 0.72878 0.90860 0.89487 1.000\r\n H83 H 0.26880 0.10912 0.25072 1.000\r\n H84 H 0.51190 0.64245 0.32739 1.000\r\n H85 H 0.51837 0.25909 0.60503 1.000\r\n H86 H 0.50034 0.61456 0.17296 1.000\r\n H87 H 0.49684 0.27154 0.74565 1.000\r\n H88 H 0.40844 0.28548 0.09140 1.000\r\n H89 H 0.45719 0.54803 0.78025 1.000\r\n H90 H 0.41773 0.31292 0.24605 1.000\r\n H91 H 0.47808 0.53524 0.63809 1.000\r\n H92 H 0.56743 0.39101 0.05493 1.000\r\n H93 H 0.33798 0.45340 0.83264 1.000\r\n H94 H 0.12510 0.68214 0.44778 1.000\r\n H95 H 0.86084 0.54460 0.59721 1.000\r\n H96 H 0.78846 0.50779 0.30271 1.000\r\n H97 H 0.19280 0.74752 0.75410 1.000\r\n H98 H 0.20491 0.76536 0.61846 1.000\r\n H99 H 0.77522 0.52547 0.44495 1.000\r\n H100 H 0.85622 0.50749 0.71060 1.000\r\n H101 H 0.14133 0.60185 0.31920 1.000\r\n H102 H 0.55371 0.89165 0.87274 1.000\r\n H103 H 0.44430 0.12993 0.26958 1.000\r\n H104 H 0.73209 0.18526 0.55455 1.000\r\n H105 H 0.45465 0.76821 0.39794 1.000\r\n H106 H 0.70169 0.99438 0.69833 1.000\r\n H107 H 0.46778 0.96307 0.25822 1.000\r\n H108 H 0.62762 0.10574 0.38984 1.000\r\n H109 H 0.58352 0.83769 0.55225 1.000\r\n H110 H 0.52616 0.98946 0.00531 1.000\r\n H111 H 0.47762 0.06221 0.13173 1.000\r\n H112 H 0.67448 0.10546 0.16074 1.000\r\n H113 H 0.33501 0.97423 0.97378 1.000\r\n H114 H 0.85133 0.12743 0.18321 1.000\r\n H115 H 0.15908 0.95383 0.95362 1.000\r\n H116 H 0.76785 0.14020 0.83056 1.000\r\n H117 H 0.31080 0.83562 0.14707 1.000\r\n H118 H 0.31233 0.60136 0.93937 1.000\r\n H119 H 0.66599 0.32839 0.99935 1.000\r\n H120 H 0.77781 0.32416 0.68147 1.000\r\n H121 H 0.29757 0.65422 0.29473 1.000\r\n H122 H 0.25723 0.38609 0.05621 1.000\r\n H123 H 0.68325 0.53284 0.87468 1.000\r\n H124 H 0.83075 0.15379 0.97779 1.000\r\n H125 H 0.16359 0.85952 0.10062 1.000\r\n H126 H 0.84092 0.86321 0.02877 1.000\r\n H127 H 0.16411 0.18090 0.12359 1.000\r\n N128 N 0.05566 0.34855 0.04562 1.000\r\n N129 N 0.94434 0.65145 0.95438 1.000\r\n N130 N 0.96083 0.50785 0.82370 1.000\r\n N131 N 0.03917 0.49215 0.17630 1.000\r\n N132 N 0.81129 0.30341 0.87165 1.000\r\n N133 N 0.18871 0.69659 0.12835 1.000\r\n N134 N 0.20520 0.55299 0.99767 1.000\r\n N135 N 0.79480 0.44701 0.00233 1.000\r\n N136 N 0.81480 0.36183 0.82370 1.000\r\n N137 N 0.18520 0.63817 0.17630 1.000\r\n N138 N 0.20168 0.49457 0.04562 1.000\r\n N139 N 0.79832 0.50543 0.95438 1.000\r\n N140 N 0.06842 0.56053 0.87165 1.000\r\n N141 N 0.93158 0.43947 0.12835 1.000\r\n N142 N 0.94807 0.29586 0.99767 1.000\r\n N143 N 0.05193 0.70414 0.00233 1.000\r\n Ni144 Ni 0.11934 0.49342 0.10996 1.000\r\n Ni145 Ni 0.88066 0.50658 0.89004 1.000\r\n Ni146 Ni 0.87259 0.37259 0.00000 1.000\r\n Ni147 Ni 0.12741 0.62741 0.00000 1.000\r\n O148 O 0.46722 0.48837 0.38403 1.000\r\n C149 C 0.42695 0.55494 0.42131 1.000\r\n H150 H 0.35373 0.54062 0.36841 1.000\r\n H151 H 0.48676 0.63880 0.45517 1.000\r\n H152 H 0.40769 0.53839 0.47575 1.000\r\n O153 O 0.51338 0.39669 0.53148 1.000\r\n C154 C 0.58545 0.36138 0.51562 1.000\r\n H155 H 0.54825 0.27392 0.48248 1.000\r\n H156 H 0.65753 0.39671 0.58014 1.000\r\n H157 H 0.60946 0.38620 0.46637 1.000\r\n O158 O 0.21125 0.65560 0.84420 1.000\r\n C159 C 0.23641 0.59272 0.78035 1.000\r\n H160 H 0.21386 0.51360 0.77769 1.000\r\n H161 H 0.19888 0.58148 0.70908 1.000\r\n H162 H 0.32203 0.63262 0.80288 1.000\r\n O163 O 0.78525 0.41314 0.16592 1.000\r\n C164 C 0.75499 0.30844 0.15067 1.000\r\n H165 H 0.79856 0.30542 0.21376 1.000\r\n H166 H 0.76534 0.26297 0.09023 1.000\r\n H167 H 0.67087 0.26987 0.13475 1.000\r\nloop_\r\n _geom_bond_atom_site_label_1\r\n _geom_bond_atom_site_label_2\r\n _geom_bond_distance\r\n _geom_bond_site_symmetry_2\r\n _ccdc_geom_bond_type\r\n Ni146 N135 1.88315 . S\r\n Ni146 N141 1.85224 . S\r\n Ni146 N132 1.85224 1_554 S\r\n Ni146 N142 1.88287 1_554 S\r\n Ni147 N143 1.88287 . S\r\n Ni147 N133 1.85224 . S\r\n Ni147 N140 1.85224 1_554 S\r\n Ni147 N134 1.88315 1_554 S\r\n N135 N139 1.37737 1_554 S\r\n N135 C67 1.32029 1_554 S\r\n N143 C77 1.31556 . S\r\n N143 N129 1.38433 1_454 S\r\n C76 C78 1.38112 . S\r\n C76 H124 1.07554 1_554 S\r\n C76 N142 1.33612 1_554 S\r\n C32 C37 1.36569 . S\r\n C32 H92 1.08388 . S\r\n C32 C34 1.39695 1_554 S\r\n H110 C16 1.08156 . S\r\n C16 C62 1.39809 1_565 S\r\n C16 C38 1.39680 1_554 S\r\n C65 C63 1.39891 1_565 S\r\n C65 C69 1.40520 1_565 S\r\n C65 H115 1.08314 1_554 S\r\n C37 C24 1.48695 . S\r\n C37 C35 1.41173 1_554 S\r\n H126 C8 1.08367 . S\r\n C79 C77 1.37575 1_655 S\r\n C79 C10 1.46394 1_655 S\r\n C79 C1 1.40665 1_554 S\r\n C72 N138 1.31109 . S\r\n C72 H122 1.07732 . S\r\n C72 C6 1.36747 1_554 S\r\n C63 C17 1.39865 . S\r\n C63 H113 1.08230 1_544 S\r\n C68 C13 1.47560 . S\r\n C68 C64 1.40279 . S\r\n C68 C14 1.40612 1_544 S\r\n N128 C0 1.32672 . S\r\n N128 Ni144 1.84262 . S\r\n N128 N142 1.38433 1_454 S\r\n N138 Ni144 1.85163 . S\r\n N138 N134 1.37737 1_554 S\r\n C8 C13 1.39272 1_565 S\r\n C8 C10 1.39866 1_655 S\r\n C77 H125 1.07050 . S\r\n C13 C11 1.41583 . S\r\n C78 C11 1.46819 . S\r\n C78 C0 1.38697 1_655 S\r\n C10 C12 1.42044 1_565 S\r\n C11 C9 1.39683 1_655 S\r\n C0 H80 1.08042 . S\r\n C12 C69 1.47923 . S\r\n C12 C9 1.37691 . S\r\n H166 C164 1.11344 . S\r\n H88 C26 1.08348 . S\r\n C69 C15 1.40501 . S\r\n C62 C64 1.39941 . S\r\n C62 H112 1.08225 . S\r\n C9 H127 1.08056 . S\r\n Ni144 N131 1.85100 . S\r\n Ni144 N137 1.84227 . S\r\n C64 H114 1.08315 . S\r\n C24 C26 1.40458 . S\r\n C24 C20 1.40304 . S\r\n C17 H111 1.08142 . S\r\n C17 C39 1.39726 . S\r\n N133 C45 1.33846 . S\r\n N133 N137 1.37738 . S\r\n N141 N131 1.38433 1_655 S\r\n N141 C75 1.33446 . S\r\n H167 C164 1.10953 . S\r\n C26 C28 1.39661 . S\r\n H117 C45 1.08049 . S\r\n C164 O163 1.40714 . S\r\n C164 H165 1.11406 . S\r\n O163 C75 1.37402 . S\r\n H86 C20 1.08328 . S\r\n C45 C5 1.38205 . S\r\n N131 C23 1.34710 . S\r\n N137 C71 1.33357 . S\r\n C20 C18 1.39805 . S\r\n C75 C3 1.38984 . S\r\n C15 C39 1.39730 . S\r\n C15 H83 1.08311 . S\r\n C39 H103 1.08257 . S\r\n C28 H90 1.08237 . S\r\n C28 C30 1.40002 . S\r\n C71 C5 1.38413 . S\r\n C71 H121 1.07931 . S\r\n C5 C53 1.46735 . S\r\n H107 C56 1.07955 . S\r\n C23 C3 1.40297 1_455 S\r\n C23 H101 1.07827 . S\r\n C3 C40 1.48026 . S\r\n C18 C30 1.41448 . S\r\n C18 H84 1.07988 . S\r\n C30 O148 1.38993 . S\r\n H96 C47 1.07378 . S\r\n C56 C53 1.40577 . S\r\n C56 C54 1.39002 1_565 S\r\n C53 C58 1.40167 . S\r\n C40 C47 1.41083 . S\r\n C40 C49 1.40510 1_655 S\r\n C47 C43 1.38538 . S\r\n H150 C149 1.11478 . S\r\n O148 C149 1.41160 . S\r\n H108 C54 1.08409 . S\r\n C54 C61 1.39843 . S\r\n C58 H105 1.07960 . S\r\n C58 C60 1.38690 . S\r\n C149 H151 1.10986 . S\r\n C149 H152 1.10922 . S\r\n H99 C43 1.08440 . S\r\n C43 C50 1.39193 . S\r\n C49 H94 1.08184 . S\r\n C49 C51 1.39556 . S\r\n H157 C154 1.10951 . S\r\n C60 C61 1.41109 1_565 S\r\n C60 C55 1.39773 . S\r\n C61 C59 1.38883 . S\r\n H155 C154 1.11119 . S\r\n C154 O153 1.41224 . S\r\n C154 H156 1.11448 . S\r\n C50 C51 1.41012 1_655 S\r\n C50 C48 1.38802 . S\r\n C51 C42 1.39770 . S\r\n O153 C31 1.39235 . S\r\n C55 H109 1.08356 . S\r\n C55 C57 1.38824 1_565 S\r\n H104 C59 1.08136 . S\r\n C59 C52 1.40280 . S\r\n H95 C48 1.08093 . S\r\n C48 C41 1.40159 1_655 S\r\n H85 C19 1.08143 . S\r\n C31 C19 1.41477 . S\r\n C31 C29 1.40139 . S\r\n C42 H98 1.08366 . S\r\n C42 C46 1.39383 . S\r\n C57 C52 1.40538 . S\r\n C57 H106 1.08058 1_545 S\r\n C52 C4 1.46648 . S\r\n H91 C29 1.08282 . S\r\n C19 C21 1.39758 . S\r\n C29 C27 1.39834 . S\r\n H120 C70 1.07828 . S\r\n C41 C46 1.40871 . S\r\n C41 C2 1.47988 . S\r\n C46 H97 1.07860 . S\r\n H161 C159 1.10839 . S\r\n H100 C22 1.07965 . S\r\n C21 H87 1.08326 . S\r\n C21 C25 1.40573 . S\r\n C4 C70 1.39149 . S\r\n C4 C44 1.39025 . S\r\n C70 N136 1.33400 . S\r\n C27 C25 1.40464 . S\r\n C27 H89 1.08299 . S\r\n C2 C22 1.38375 1_455 S\r\n C2 C74 1.39267 . S\r\n C22 N130 1.33623 . S\r\n C25 C36 1.48915 . S\r\n H160 C159 1.11339 . S\r\n C159 H162 1.10950 . S\r\n C159 O158 1.41770 . S\r\n C44 H116 1.07733 . S\r\n C44 N132 1.33718 . S\r\n C74 O158 1.37721 . S\r\n C74 N140 1.34683 . S\r\n N136 N132 1.37738 . S\r\n N136 Ni145 1.84227 . S\r\n N130 N140 1.38433 1_655 S\r\n N130 Ni145 1.85100 . S\r\n H93 C33 1.08274 . S\r\n C36 C33 1.39283 . S\r\n C36 C34 1.40721 . S\r\n H102 C38 1.08247 . S\r\n H123 C73 1.08029 . S\r\n C33 C35 1.40259 . S\r\n Ni145 N139 1.85163 . S\r\n Ni145 N129 1.84262 . S\r\n H82 C14 1.08332 . S\r\n C73 C7 1.37650 . S\r\n C73 N139 1.33457 . S\r\n C34 C7 1.43985 . S\r\n H81 C1 1.08051 . S\r\n C7 C67 1.36396 . S\r\n H118 C66 1.07185 . S\r\n C38 C14 1.39748 . S\r\n N129 C1 1.34007 . S\r\n C35 C6 1.43524 . S\r\n C66 C6 1.38970 . S\r\n C66 N134 1.33637 . S\r\n C67 H119 1.07920 . S\r\n\r\n```"
] | 2021-11-04T02:46:37
| 2022-06-04T19:41:34
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
There is a StructureGraph class, and some .cif files include bonding information. Is there any interest in creating StructureGraph's from the bonding information in a .cif file?
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2289/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2289/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2290
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2290/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2290/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2290/events
|
https://github.com/materialsproject/pymatgen/issues/2290
| 1,045,846,464
|
I_kwDOACgets4-VlnA
| 2,290
|
Error while using MagOrderingTransformation
|
{
"login": "t-reents",
"id": 77727843,
"node_id": "MDQ6VXNlcjc3NzI3ODQz",
"avatar_url": "https://avatars.githubusercontent.com/u/77727843?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/t-reents",
"html_url": "https://github.com/t-reents",
"followers_url": "https://api.github.com/users/t-reents/followers",
"following_url": "https://api.github.com/users/t-reents/following{/other_user}",
"gists_url": "https://api.github.com/users/t-reents/gists{/gist_id}",
"starred_url": "https://api.github.com/users/t-reents/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/t-reents/subscriptions",
"organizations_url": "https://api.github.com/users/t-reents/orgs",
"repos_url": "https://api.github.com/users/t-reents/repos",
"events_url": "https://api.github.com/users/t-reents/events{/privacy}",
"received_events_url": "https://api.github.com/users/t-reents/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This suggests that your enum.x and makeStr.x are incorrectly compiled. I suggest you compile from the enumlib source. ",
"Thanks @shyuep for the quick reply. I followed the description in the readme https://github.com/msg-byu/enumlib and compiled it again. The error still occurs",
"What OS are you using? Also, there shouldn't be a makeStr.py. It should be makeStr.x\r\n",
"I am using Ubuntu WSL. They mention in the readme that the makeStr.x was superceded by the makeStr.py. Moreover, I think this is also considered here:\r\n\r\nhttps://github.com/materialsproject/pymatgen/blob/50f51443b118d1f72dd8c9e3269ad1969eba5fb9/pymatgen/command_line/enumlib_caller.py#L58",
"any updates on this? I'm struggling with the same error on Linux and MacOS11. Thanks ",
"You need to edit the makeStr.py file and add the following at the top:\n\n#! /usr/bin/env python\n\nThis could probably be avoided by modifying pymatgen itself. But I tested this fix and it works for me. ",
"perfect, thanks Alex!"
] | 2021-11-05T13:25:24
| 2023-12-06T12:57:12
|
2023-12-06T12:57:11Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I tried to use the `MagOrderingTransformation` module https://github.com/materialsproject/pymatgen/blob/50f51443b118d1f72dd8c9e3269ad1969eba5fb9/pymatgen/transformations/advanced_transformations.py#L621
Code snippet:
```ruby
magnetic_species = {"Ni": 1.0, "O": 0.0}
MagOrder = MagOrderingTransformation(mag_species_spin=magnetic_species)
ordered = MagOrder.apply_transformation(structure_pymatgen)
```
I encountered the following error:
```ruby
---------------------------------------------------------------------------
OSError Traceback (most recent call last)
/tmp/ipykernel_6126/1801449226.py in <module>
----> 1 MagOrder.apply_transformation(structure_pymatgen)
~/.venvs/aiida/lib/python3.8/site-packages/pymatgen/transformations/advanced_transformations.py in apply_transformation(self, structure, return_ranked_list)
872 t = EnumerateStructureTransformation(**enum_kwargs)
873
--> 874 alls = t.apply_transformation(structure, return_ranked_list=return_ranked_list)
875
876 # handle the fact that EnumerateStructureTransformation can either
~/.venvs/aiida/lib/python3.8/site-packages/pymatgen/transformations/advanced_transformations.py in apply_transformation(self, structure, return_ranked_list)
423 )
424 try:
--> 425 adaptor.run()
426 structures = adaptor.structures
427 if structures:
~/.venvs/aiida/lib/python3.8/site-packages/pymatgen/command_line/enumlib_caller.py in run(self)
146 # Read in the enumeration output as structures.
147 if num_structs > 0:
--> 148 self.structures = self._get_structures(num_structs)
149 else:
150 raise EnumError("Unable to enumerate structure.")
~/.venvs/aiida/lib/python3.8/site-packages/pymatgen/command_line/enumlib_caller.py in _get_structures(self, num_structs)
334 options = ["struct_enum.out", str(0), str(num_structs - 1)]
335
--> 336 with subprocess.Popen(
337 [makestr_cmd] + options,
338 stdout=subprocess.PIPE,
/usr/lib/python3.8/subprocess.py in __init__(self, args, bufsize, executable, stdin, stdout, stderr, preexec_fn, close_fds, shell, cwd, env, universal_newlines, startupinfo, creationflags, restore_signals, start_new_session, pass_fds, encoding, errors, text)
856 encoding=encoding, errors=errors)
857
--> 858 self._execute_child(args, executable, preexec_fn, close_fds,
859 pass_fds, cwd, env,
860 startupinfo, creationflags, shell,
/usr/lib/python3.8/subprocess.py in _execute_child(self, args, executable, preexec_fn, close_fds, pass_fds, cwd, env, startupinfo, creationflags, shell, p2cread, p2cwrite, c2pread, c2pwrite, errread, errwrite, restore_signals, start_new_session)
1702 if errno_num != 0:
1703 err_msg = os.strerror(errno_num)
-> 1704 raise child_exception_type(errno_num, err_msg, err_filename)
1705 raise child_exception_type(err_msg)
1706
OSError: [Errno 8] Exec format error: '/home/treents/.venvs/aiida/bin/makeStr.py'
```
How to solve this? The `enum.x` and `makeStr.py` files are in the path otherwise the error would be different.
|
{
"login": "t-reents",
"id": 77727843,
"node_id": "MDQ6VXNlcjc3NzI3ODQz",
"avatar_url": "https://avatars.githubusercontent.com/u/77727843?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/t-reents",
"html_url": "https://github.com/t-reents",
"followers_url": "https://api.github.com/users/t-reents/followers",
"following_url": "https://api.github.com/users/t-reents/following{/other_user}",
"gists_url": "https://api.github.com/users/t-reents/gists{/gist_id}",
"starred_url": "https://api.github.com/users/t-reents/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/t-reents/subscriptions",
"organizations_url": "https://api.github.com/users/t-reents/orgs",
"repos_url": "https://api.github.com/users/t-reents/repos",
"events_url": "https://api.github.com/users/t-reents/events{/privacy}",
"received_events_url": "https://api.github.com/users/t-reents/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2290/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2290/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2291
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2291/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2291/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2291/events
|
https://github.com/materialsproject/pymatgen/pull/2291
| 1,046,055,990
|
PR_kwDOACgets4uKMpx
| 2,291
|
Replace-species
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/44171744)\n\nCoverage decreased (-0.6%) to 83.115% when pulling **d1d01d6d166191a203dca6ad02642de08bcbdea6 on janosh:replace-species** into **9276567df1bf01eea473933688975003449dc5e5 on materialsproject:master**.\n",
"Thanks @janosh, I see you reverted the other commit so going to merge this"
] | 2021-11-05T17:00:09
| 2021-11-18T18:34:06
|
2021-11-18T18:32:41Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
`Structure().replace_species` doc string should make it clear that original structure is modified in place.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2291/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2291/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2291",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2291",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2291.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2291.patch",
"merged_at": "2021-11-18T18:32:40Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2292
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2292/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2292/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2292/events
|
https://github.com/materialsproject/pymatgen/issues/2292
| 1,046,689,278
|
I_kwDOACgets4-YzX-
| 2,292
|
Broken docstrings on the dos module
|
{
"login": "Andrew-S-Rosen",
"id": 8674072,
"node_id": "MDQ6VXNlcjg2NzQwNzI=",
"avatar_url": "https://avatars.githubusercontent.com/u/8674072?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Andrew-S-Rosen",
"html_url": "https://github.com/Andrew-S-Rosen",
"followers_url": "https://api.github.com/users/Andrew-S-Rosen/followers",
"following_url": "https://api.github.com/users/Andrew-S-Rosen/following{/other_user}",
"gists_url": "https://api.github.com/users/Andrew-S-Rosen/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Andrew-S-Rosen/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Andrew-S-Rosen/subscriptions",
"organizations_url": "https://api.github.com/users/Andrew-S-Rosen/orgs",
"repos_url": "https://api.github.com/users/Andrew-S-Rosen/repos",
"events_url": "https://api.github.com/users/Andrew-S-Rosen/events{/privacy}",
"received_events_url": "https://api.github.com/users/Andrew-S-Rosen/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Fixed in #2293."
] | 2021-11-07T09:26:13
| 2021-11-15T01:49:12
|
2021-11-15T01:49:12Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
In the [docs](https://pymatgen.org/pymatgen.electronic_structure.dos.html) for `pymatgen.electronic_structure.dos`, many of the docstrings are cutoff because of ":" characters not playing well with the doc generator. For instance, see the `Returns` entries [here](https://pymatgen.org/pymatgen.electronic_structure.dos.html#pymatgen.electronic_structure.dos.LobsterCompleteDos.get_site_t2g_eg_resolved_dos) and [here](https://pymatgen.org/pymatgen.electronic_structure.dos.html#pymatgen.electronic_structure.dos.LobsterCompleteDos.get_site_t2g_eg_resolved_dos).
|
{
"login": "Andrew-S-Rosen",
"id": 8674072,
"node_id": "MDQ6VXNlcjg2NzQwNzI=",
"avatar_url": "https://avatars.githubusercontent.com/u/8674072?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Andrew-S-Rosen",
"html_url": "https://github.com/Andrew-S-Rosen",
"followers_url": "https://api.github.com/users/Andrew-S-Rosen/followers",
"following_url": "https://api.github.com/users/Andrew-S-Rosen/following{/other_user}",
"gists_url": "https://api.github.com/users/Andrew-S-Rosen/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Andrew-S-Rosen/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Andrew-S-Rosen/subscriptions",
"organizations_url": "https://api.github.com/users/Andrew-S-Rosen/orgs",
"repos_url": "https://api.github.com/users/Andrew-S-Rosen/repos",
"events_url": "https://api.github.com/users/Andrew-S-Rosen/events{/privacy}",
"received_events_url": "https://api.github.com/users/Andrew-S-Rosen/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2292/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2292/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2293
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2293/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2293/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2293/events
|
https://github.com/materialsproject/pymatgen/pull/2293
| 1,046,689,912
|
PR_kwDOACgets4uMHrJ
| 2,293
|
Docstring updates: Fix missing Outcar attributes and update elemental_dos_dos string
|
{
"login": "Andrew-S-Rosen",
"id": 8674072,
"node_id": "MDQ6VXNlcjg2NzQwNzI=",
"avatar_url": "https://avatars.githubusercontent.com/u/8674072?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Andrew-S-Rosen",
"html_url": "https://github.com/Andrew-S-Rosen",
"followers_url": "https://api.github.com/users/Andrew-S-Rosen/followers",
"following_url": "https://api.github.com/users/Andrew-S-Rosen/following{/other_user}",
"gists_url": "https://api.github.com/users/Andrew-S-Rosen/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Andrew-S-Rosen/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Andrew-S-Rosen/subscriptions",
"organizations_url": "https://api.github.com/users/Andrew-S-Rosen/orgs",
"repos_url": "https://api.github.com/users/Andrew-S-Rosen/repos",
"events_url": "https://api.github.com/users/Andrew-S-Rosen/events{/privacy}",
"received_events_url": "https://api.github.com/users/Andrew-S-Rosen/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/44069430)\n\nCoverage decreased (-0.6%) to 83.103% when pulling **105253d538a972a09f06e5383f41c11a95f2a8ad on arosen93:rosen-docs** into **50f51443b118d1f72dd8c9e3269ad1969eba5fb9 on materialsproject:master**.\n",
"Thanks."
] | 2021-11-07T09:30:12
| 2021-11-15T05:47:51
|
2021-11-08T13:12:27Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* The docs for `pymatgen.io.vasp.outputs.Outcar` are missing many of the attributes. For instance, it wasn't clear to me until I read the code that the `Outcar()` object has a `total_mag` keyword. I've update the docstring to include the missing attributes.
* The docs for `pymatgen.electronic_structure.dos.get_element_spd_dos` have an incorrect description of the `Returns` dictionary. This has been updated. Note: There are still some aesthetic issues with the docs, as mentioned in #2292.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2293/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2293/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2293",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2293",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2293.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2293.patch",
"merged_at": "2021-11-08T13:12:27Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2294
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2294/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2294/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2294/events
|
https://github.com/materialsproject/pymatgen/pull/2294
| 1,050,134,538
|
PR_kwDOACgets4uXSA6
| 2,294
|
Pyupgrade 3.7 plus
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/44425206)\n\nCoverage decreased (-0.6%) to 83.114% when pulling **b587d68a2877f90bb9c051f9628cab52439b119e on janosh:pyupgrade** into **8fd02cf3a7d33059a62200c12dd48a15be21a8a3 on materialsproject:master**.\n",
"Thanks for this PR @janosh. While I don't expect any significant issues, given the size of the diff I'll need some time to review it.",
"(I'll note that similar automatic code tools have been run against pymatgen in the past, and 99% worked but did introduce one or two small issues that needed to be manually fixed)",
"> I'll need some time to review it.\r\n\r\nSure thing. Is there something I can help check? Run or write more tests for anything particular?",
"Nope, not at all. Perhaps being over-cautious, but I just wanted to skim through it myself",
"Actually there is a merge conflict now, so that could be fixed?",
"Let me know when to resolve the merge conflicts here. Best done close to review time.",
"Thanks, will do. Before I get to this I need to figure out what's happening with CI and the current test failures in the main branch.\r\n\r\nWe actually had some code tools introduce incorrect type hints recently (Eg `Tuple` -> `tuple`) which does make me want to review more closely but if you can vouch for it, based on your previous PRs, I would be happy to merge without a too-detailed review.",
"I've been using `pyupgrade` for a while now and have never once known it to create problems if used correctly. E.g. it too would convert `Tuple -> tuple` if run in `--py310-plus` mode but of course that'd be user error if your project needs to support older Pythons.",
"We can close this. I already did pyupgrade on pymatgen a few days back. Thanks for the help.",
"@shyuep I think there are still some things worth merging here like 552e0d9, ab2c75d and b587d68.",
"Can you submit those as a separate PR? Thanks.",
"@shyuep I rebased onto `master` and force pushed to my fork. Only 34 changed files with 84 additions and 94 deletions. So much more manageable PR now. I think easiest to reopen this PR though. Can update the title/rename the branch if that was your reason for suggesting a new PR?",
"It seems that I can't reopen the PR after you force pushed it. I suggest you reopen a new PR.\r\n",
"@shyuep See #2318."
] | 2021-11-10T17:57:04
| 2021-12-04T17:46:09
|
2021-12-03T20:35:15Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Lots of automatic updates to use newer language features by running
```sh
pyupgrade --py37-plus **/*.py
```
across the whole code base.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2294/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2294/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2294",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2294",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2294.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2294.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2295
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2295/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2295/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2295/events
|
https://github.com/materialsproject/pymatgen/pull/2295
| 1,051,188,850
|
PR_kwDOACgets4uauCP
| 2,295
|
Update DOI URLs
|
{
"login": "e-kwsm",
"id": 11627708,
"node_id": "MDQ6VXNlcjExNjI3NzA4",
"avatar_url": "https://avatars.githubusercontent.com/u/11627708?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/e-kwsm",
"html_url": "https://github.com/e-kwsm",
"followers_url": "https://api.github.com/users/e-kwsm/followers",
"following_url": "https://api.github.com/users/e-kwsm/following{/other_user}",
"gists_url": "https://api.github.com/users/e-kwsm/gists{/gist_id}",
"starred_url": "https://api.github.com/users/e-kwsm/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/e-kwsm/subscriptions",
"organizations_url": "https://api.github.com/users/e-kwsm/orgs",
"repos_url": "https://api.github.com/users/e-kwsm/repos",
"events_url": "https://api.github.com/users/e-kwsm/events{/privacy}",
"received_events_url": "https://api.github.com/users/e-kwsm/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thank you @e-kwsm!",
"\n[](https://coveralls.io/builds/44381135)\n\nCoverage decreased (-0.6%) to 83.112% when pulling **d7044eb809f1b8b0c96caab8f52061ee23542bad on e-kwsm:doi** into **9276567df1bf01eea473933688975003449dc5e5 on materialsproject:master**.\n",
"@e-kwsm, since you are a new contributor, please do fill out [this form](https://forms.gle/JnisFb38QDR8QTFTA) so we can acknowledge you appropriately in [pymatgen documentation](https://pymatgen.org/team.html)"
] | 2021-11-11T17:08:07
| 2021-11-19T00:42:42
|
2021-11-18T19:07:00Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
* URLs are changed to https://doi.org/ in accordance with https://www.doi.org/doi_handbook/2_Numbering.html#htmlencoding.
<!--
## Additional dependencies introduced (if any)
* List all new dependencies needed and justify why. While adding dependencies that bring
significantly useful functionality is perfectly fine, adding ones that
add trivial functionality, e.g., to use one single easily implementable
function, is frowned upon. Provide a justification why that dependency is needed.
Especially frowned upon are circular dependencies, e.g., depending on derivative
modules of pymatgen such as custodian or Fireworks.
## TODO (if any)
If this is a work-in-progress, write something about what else needs
to be done
* Feature 1 supports A, but not B.
-->
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [ ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2295/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2295/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2295",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2295",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2295.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2295.patch",
"merged_at": "2021-11-18T19:07:00Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2296
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2296/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2296/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2296/events
|
https://github.com/materialsproject/pymatgen/pull/2296
| 1,052,804,666
|
PR_kwDOACgets4ufa3h
| 2,296
|
Remove accidentally tracked files and unset executable flag
|
{
"login": "e-kwsm",
"id": 11627708,
"node_id": "MDQ6VXNlcjExNjI3NzA4",
"avatar_url": "https://avatars.githubusercontent.com/u/11627708?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/e-kwsm",
"html_url": "https://github.com/e-kwsm",
"followers_url": "https://api.github.com/users/e-kwsm/followers",
"following_url": "https://api.github.com/users/e-kwsm/following{/other_user}",
"gists_url": "https://api.github.com/users/e-kwsm/gists{/gist_id}",
"starred_url": "https://api.github.com/users/e-kwsm/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/e-kwsm/subscriptions",
"organizations_url": "https://api.github.com/users/e-kwsm/orgs",
"repos_url": "https://api.github.com/users/e-kwsm/repos",
"events_url": "https://api.github.com/users/e-kwsm/events{/privacy}",
"received_events_url": "https://api.github.com/users/e-kwsm/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Good catch @e-kwsm, thank you",
"\n[](https://coveralls.io/builds/44289132)\n\nCoverage decreased (-0.6%) to 83.112% when pulling **23c0e07833ea5ef1054186617f4d1e4083722afd on e-kwsm:chmod-x** into **9276567df1bf01eea473933688975003449dc5e5 on materialsproject:master**.\n"
] | 2021-11-13T23:52:39
| 2021-11-19T00:41:35
|
2021-11-18T18:30:59Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
<!--Include a summary of major changes in bullet points:-->
Subject says it all.
<!--
## Additional dependencies introduced (if any)
* List all new dependencies needed and justify why. While adding dependencies that bring
significantly useful functionality is perfectly fine, adding ones that
add trivial functionality, e.g., to use one single easily implementable
function, is frowned upon. Provide a justification why that dependency is needed.
Especially frowned upon are circular dependencies, e.g., depending on derivative
modules of pymatgen such as custodian or Fireworks.
## TODO (if any)
If this is a work-in-progress, write something about what else needs
to be done
* Feature 1 supports A, but not B.
-->
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [ ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2296/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2296/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2296",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2296",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2296.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2296.patch",
"merged_at": "2021-11-18T18:30:59Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2297
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2297/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2297/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2297/events
|
https://github.com/materialsproject/pymatgen/pull/2297
| 1,053,058,962
|
PR_kwDOACgets4ugIpv
| 2,297
|
Add warning if LASPH != True for meta-GGA/hybrid/vdW/+U
|
{
"login": "Andrew-S-Rosen",
"id": 8674072,
"node_id": "MDQ6VXNlcjg2NzQwNzI=",
"avatar_url": "https://avatars.githubusercontent.com/u/8674072?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Andrew-S-Rosen",
"html_url": "https://github.com/Andrew-S-Rosen",
"followers_url": "https://api.github.com/users/Andrew-S-Rosen/followers",
"following_url": "https://api.github.com/users/Andrew-S-Rosen/following{/other_user}",
"gists_url": "https://api.github.com/users/Andrew-S-Rosen/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Andrew-S-Rosen/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Andrew-S-Rosen/subscriptions",
"organizations_url": "https://api.github.com/users/Andrew-S-Rosen/orgs",
"repos_url": "https://api.github.com/users/Andrew-S-Rosen/repos",
"events_url": "https://api.github.com/users/Andrew-S-Rosen/events{/privacy}",
"received_events_url": "https://api.github.com/users/Andrew-S-Rosen/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/44468724)\n\nCoverage decreased (-0.6%) to 83.117% when pulling **debcc753d732d2d0db0bf09c248e6a659ac7b4fe on arosen93:rosen-mixing** into **9276567df1bf01eea473933688975003449dc5e5 on materialsproject:master**.\n",
"Thanks @arosen93!"
] | 2021-11-14T22:31:27
| 2021-11-24T03:30:02
|
2021-11-24T01:56:48Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
This PR implements a simple `BadInputSetWarning` if `LASPH = False` (or is not set) yet the user is running a meta-GGA/hybrid/DFT+U/vdW-DF calculation.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2297/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2297/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2297",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2297",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2297.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2297.patch",
"merged_at": "2021-11-24T01:56:48Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2298
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2298/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2298/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2298/events
|
https://github.com/materialsproject/pymatgen/pull/2298
| 1,053,426,608
|
PR_kwDOACgets4uhUXS
| 2,298
|
Add warning if improper ALGO is used for hybrid calculations
|
{
"login": "Andrew-S-Rosen",
"id": 8674072,
"node_id": "MDQ6VXNlcjg2NzQwNzI=",
"avatar_url": "https://avatars.githubusercontent.com/u/8674072?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Andrew-S-Rosen",
"html_url": "https://github.com/Andrew-S-Rosen",
"followers_url": "https://api.github.com/users/Andrew-S-Rosen/followers",
"following_url": "https://api.github.com/users/Andrew-S-Rosen/following{/other_user}",
"gists_url": "https://api.github.com/users/Andrew-S-Rosen/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Andrew-S-Rosen/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Andrew-S-Rosen/subscriptions",
"organizations_url": "https://api.github.com/users/Andrew-S-Rosen/orgs",
"repos_url": "https://api.github.com/users/Andrew-S-Rosen/repos",
"events_url": "https://api.github.com/users/Andrew-S-Rosen/events{/privacy}",
"received_events_url": "https://api.github.com/users/Andrew-S-Rosen/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/44468633)\n\nCoverage decreased (-0.6%) to 83.116% when pulling **e0f7d28c14b22740ea59ff0986771d83b061490f on arosen93:rosen-lhfcalc** into **9276567df1bf01eea473933688975003449dc5e5 on materialsproject:master**.\n",
"Thanks @arosen93! Could you add a quick test?",
"@mkhorton, apologies about all these failed tests in my recent pymatgen PRs. I'm having trouble figuring out some of the nuances of `pymatgen.vasp.io.sets`. I'll ping you when they're done after I talk to someone in the group to learn more about what's going on in this part of pymatgen.",
"@mkhorton, the PRs are good to go now :) #2298, #2297, #2301 (thanks for the help with debugging the tests, @munrojm!!)",
"Thanks @arosen93 !"
] | 2021-11-15T09:46:33
| 2021-11-23T18:58:06
|
2021-11-23T16:14:00Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Per the VASP manual (https://www.vasp.at/wiki/index.php/LHFCALC), hybrid calculations should only be run with ALGO = All, Damped, or Normal (never Fast or VeryFast despite no clear error written in the output). This PR implements a `BadInputSetWarning` if LHFCALC = True and Algo is not one of the three supported options.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2298/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2298/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2298",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2298",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2298.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2298.patch",
"merged_at": "2021-11-23T16:14:00Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2299
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2299/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2299/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2299/events
|
https://github.com/materialsproject/pymatgen/pull/2299
| 1,056,922,779
|
PR_kwDOACgets4usUGH
| 2,299
|
[WIP] Pourbaix: new methods and enhancements for customizability
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 3093102097,
"node_id": "MDU6TGFiZWwzMDkzMTAyMDk3",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/stale",
"name": "stale",
"color": "A99B62",
"default": false,
"description": "Abandoned or conflicting PRs and outdated issues"
}
] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/44358398)\n\nCoverage decreased (-0.6%) to 83.112% when pulling **a0dcaf93c9325aa2cbf09791af5e69060fb94c98 on rkingsbury:pourbaix_updates** into **e084ec1383aa7006f9e4c9f6781bd9199043e05a on materialsproject:master**.\n",
"@rkingsbury Is this still WIP?",
"@janosh I still think it would add value, but I have not started and it's not something I'll likely have time to work on anytime soon.",
"No worries. Was just looking through the list of open PRs seeing if anything just needed a little push over the finish line.",
"Here are a few ideas for further Pourbaix enhancements:\r\n\r\n1. Can print an output file containing all the phases in their respective stable regions, all considered phases from `pymatgen` database for that Pourbaix Diagram, Equation of lines on the Pourbaix Diagram, and their corresponding equations.\r\n2. Addition of metastable and missing phases in the `pymatgen` database. Better exchange correlation (meta-GGA) values can also be considered.\r\n3. Include one more dimension of concentration to the Pourbaix diagram. "
] | 2021-11-18T04:19:49
| 2024-10-10T01:54:44
|
2024-10-10T01:54:43Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
The eventual goal of this PR is to provide some additional methods that make `PourbaixDiagram` more easily customizable within pymatgen. This is a WIP motivated by requests and challenges faced by several users that I've interacted with.
Part of the goal here is to provide similar functionality to [MPRester.get_pourbaix_entries](https://github.com/materialsproject/api/blob/4f13ce2b905970246b975f83e7effe36f3f401fd/src/mp_api/client.py#L411) and related methods, but accessible for locally-derived entries, rather than just those retrieved from MPRester.
* `PourbaixDiagram.from_phase_diagram`
* `PourbaixDiagram.from_computed_entries`
* TBD!
## Additional dependencies introduced (if any)
None
## TODO (if any)
Seek feedback on mock up docstrings before implementing.
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [ ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [ ] All linting and tests pass.
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2299/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2299/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2299",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2299",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2299.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2299.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2300
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2300/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2300/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2300/events
|
https://github.com/materialsproject/pymatgen/pull/2300
| 1,059,033,545
|
PR_kwDOACgets4uzBSA
| 2,300
|
wrap SC to unit cell
|
{
"login": "jmmshn",
"id": 14003693,
"node_id": "MDQ6VXNlcjE0MDAzNjkz",
"avatar_url": "https://avatars.githubusercontent.com/u/14003693?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jmmshn",
"html_url": "https://github.com/jmmshn",
"followers_url": "https://api.github.com/users/jmmshn/followers",
"following_url": "https://api.github.com/users/jmmshn/following{/other_user}",
"gists_url": "https://api.github.com/users/jmmshn/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jmmshn/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jmmshn/subscriptions",
"organizations_url": "https://api.github.com/users/jmmshn/orgs",
"repos_url": "https://api.github.com/users/jmmshn/repos",
"events_url": "https://api.github.com/users/jmmshn/events{/privacy}",
"received_events_url": "https://api.github.com/users/jmmshn/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This should be an option, but not forced. You can modify the kwargs of the method to provide an option called to_unit_cell.",
"I don't think we can add a kwarg for `__mul__` and I would argue that this kind of folding is the \"expect\" behavior when we construct supercells. \r\n\r\nIf we really can't change it maybe we should just offer a quick way for users to fold the sites back to the UC after a Structure has been initialized? I looked but it seems this kind of folding only occurs during construction.",
"I think Structure.sanitize is what you are looking for.",
"For `__mul__` specifically, on a change of basis, would you not reasonably expect the fractional co-ordinates to be inside the unit cell? It's a semantic question, but I think there's a good case for it.\r\n\r\nTo put it a different way, I think there's a good case that `to_unit_cell` should be True by default, and that people can disable it if they wish. I can imagine situations where people might not fractional co-ordinates wrapped, but it seems like this would be the exception rather than the rule.",
"I will accept this change for now. Personally I don't usually care about wrapping because all analysis should be independent of wrapping just for robustness. Any kind of analysis that assumes the coordinates are 0-1 are prone to hard to debug errors. None of the codes in pymatgen depends on the coordinates being within the unit cell.",
"> Any kind of analysis that assumes the coordinates are 0-1 are prone to hard to debug errors. \r\n\r\nAgreed. Still glad to see this merged though, thank you."
] | 2021-11-20T02:00:04
| 2021-11-23T20:14:56
|
2021-11-23T16:59:46Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
When you create supercell I think the expected behavior should be that the sites in the SC sit in the unit-cell.
Currently, the behavior is:
```
# Si unit cell
Full Formula (Si2)
Reduced Formula: Si
abc : 3.866974 3.866975 3.866975
angles: 60.000006 60.000002 60.000010
Sites (2)
# SP a b c
--- ---- ----- ----- -----
0 Si 0.875 0.875 0.875
1 Si 0.125 0.125 0.125
# Si [[-1,1,1], [1,-1,1], [1,1,-1]] supercell
Full Formula (Si8)
Reduced Formula: Si
abc : 5.468729 5.468729 5.468728
angles: 90.000014 90.000001 90.000001
Sites (8)
# SP a b c
--- ---- ----- ----- -----
0 Si 0.875 0.875 0.875
1 Si 1.375 1.375 0.875
2 Si 1.375 0.875 1.375
3 Si 0.875 1.375 1.375
4 Si 0.125 0.125 0.125
5 Si 0.625 0.625 0.125
6 Si 0.625 0.125 0.625
7 Si 0.125 0.625 0.625
```
The additional flag should fix this problem
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2300/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2300/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2300",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2300",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2300.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2300.patch",
"merged_at": "2021-11-23T16:59:46Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2301
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2301/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2301/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2301/events
|
https://github.com/materialsproject/pymatgen/pull/2301
| 1,059,223,579
|
PR_kwDOACgets4uzk13
| 2,301
|
Clearer handling of the MAGMOM flag in pymatgen.io.vasp.sets
|
{
"login": "Andrew-S-Rosen",
"id": 8674072,
"node_id": "MDQ6VXNlcjg2NzQwNzI=",
"avatar_url": "https://avatars.githubusercontent.com/u/8674072?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Andrew-S-Rosen",
"html_url": "https://github.com/Andrew-S-Rosen",
"followers_url": "https://api.github.com/users/Andrew-S-Rosen/followers",
"following_url": "https://api.github.com/users/Andrew-S-Rosen/following{/other_user}",
"gists_url": "https://api.github.com/users/Andrew-S-Rosen/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Andrew-S-Rosen/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Andrew-S-Rosen/subscriptions",
"organizations_url": "https://api.github.com/users/Andrew-S-Rosen/orgs",
"repos_url": "https://api.github.com/users/Andrew-S-Rosen/repos",
"events_url": "https://api.github.com/users/Andrew-S-Rosen/events{/privacy}",
"received_events_url": "https://api.github.com/users/Andrew-S-Rosen/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @arosen93!\r\n\r\nFor the PR itself, if WIP, please either put [WIP] in the title or you can explicitly \"convert to draft\" in the top-right.\r\n\r\nFor the proposed changes, this looks like an important thing to clarify, and the PR looks good.\r\n\r\nI did also want to make sure you knew about the [`Magmom` class](https://pymatgen.org/pymatgen.electronic_structure.core.html?#pymatgen.electronic_structure.core.Magmom) which was my own old attempt at starting to unify magmom representations, specifically between non-collinear and collinear cases.\r\n\r\nIn terms of handling of magmom, we've ended up in a slightly awkward situation:\r\n\r\n1. It has been more typical to use the `magmom` site property, however this does not allow the representation of disordered magnetic materials.\r\n2. It might be more naturally to represent using a `Species` `spin` property, however this representation is awkward because if symmetrically equivalent sites have _slightly_ different magnetic moments (due to numerical noise, etc.), they will get treated as inequivalent in a lot of places in pymatgen, in symmetry analysis, etc.\r\n3. They are sometimes represented as vectors and sometimes as scalars (hence the `Magmom` class, to hopefully make this easier, and also encode a spin axis)\r\n\r\nThis is broader context not immediately relevant to this PR, but may be good to bear in mind if you're editing this code.\r\n\r\n(Edit: sorry, submitted this comment too soon and missed the second half)",
"Thanks."
] | 2021-11-20T20:29:13
| 2021-11-23T18:57:18
|
2021-11-23T17:00:46Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Currently, the handling of the MAGMOM INCAR flag in `pymatgen.io.vasp.sets` could be made clearer. This became apparent to me when trying to modify MAGMOM in an atomate workflow, which did not at all do what I expected it to do (and led to some confusing errors). This PR does not change how magnetic moments are handled in Pymatgen. It just adds and clarifies some warning messages.
1. An error is now raised if MAGMOM is not a dict, and the user is appropriately notified that this is an element-specific flag, not a site-specific flag like the typical VASP INCAR's MAGMOM argument.
2. The `constrain_total_magmom` flag, when set to `True`, sums up the magnetic moments on each site and sets that as NUPDOWN. If the user specified a custom MAGMOM, then there is no guarantee the summed values are integers. I have therefore added a warning if `round(sum(magmoms)) != sum(magmoms)`.
3. The warning about Co being initialized as low-spin could be made clearer. Currently, Pymatgen prints out this warning whenever MAGMOM has an entry for Co and there is no site-specific `magmom` or `spin`. If one were to change the config to have high-spin Co or modify MAGMOM directly (e.g. `{"MAGMOM": {"Co":5}}`), the warning would still appear despite the fact that it is no longer relevant. I have clarified the wording in the warning.
4. The documentation says that if the symbol is not in the config file, VASP's default of 0.6 is used. The [VASP default is 1.0](https://www.vasp.at/wiki/index.php/MAGMOM); the Pymatgen default is 0.6. I've clarified the phrasing.
Of course, if the user specified the magnetic moments as site properties, much of this wouldn't be needed, but this subtlety surrounding how MAGMOM is treated wasn't initially clear to me.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2301/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2301/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2301",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2301",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2301.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2301.patch",
"merged_at": "2021-11-23T17:00:46Z"
}
|
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.