url
stringlengths 63
66
| repository_url
stringclasses 1
value | labels_url
stringlengths 77
80
| comments_url
stringlengths 72
75
| events_url
stringlengths 70
73
| html_url
stringlengths 52
56
| id
int64 3.39M
3.04B
| node_id
stringlengths 18
32
| number
int64 1
4.39k
| title
stringlengths 3
367
| user
dict | labels
listlengths 0
5
| state
stringclasses 2
values | locked
bool 2
classes | assignee
dict | assignees
listlengths 0
2
| milestone
null | comments
listlengths 0
30
| created_at
timestamp[s]date 2012-02-26 12:10:46
2025-05-02 18:11:41
| updated_at
timestamp[s]date 2012-02-26 14:52:41
2025-05-02 18:16:27
| closed_at
stringlengths 0
20
| author_association
stringclasses 3
values | type
stringclasses 1
value | active_lock_reason
stringclasses 2
values | sub_issues_summary
dict | body
stringlengths 0
119k
| closed_by
dict | reactions
dict | timeline_url
stringlengths 72
75
| performed_via_github_app
null | state_reason
stringclasses 4
values | is_pull_request
bool 2
classes | draft
bool 2
classes | pull_request
dict |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
https://api.github.com/repos/materialsproject/pymatgen/issues/2101
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2101/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2101/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2101/events
|
https://github.com/materialsproject/pymatgen/pull/2101
| 839,937,107
|
MDExOlB1bGxSZXF1ZXN0NTk5ODcwODQ4
| 2,101
|
small fix for insertion electrode
|
{
"login": "jmmshn",
"id": 14003693,
"node_id": "MDQ6VXNlcjE0MDAzNjkz",
"avatar_url": "https://avatars.githubusercontent.com/u/14003693?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jmmshn",
"html_url": "https://github.com/jmmshn",
"followers_url": "https://api.github.com/users/jmmshn/followers",
"following_url": "https://api.github.com/users/jmmshn/following{/other_user}",
"gists_url": "https://api.github.com/users/jmmshn/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jmmshn/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jmmshn/subscriptions",
"organizations_url": "https://api.github.com/users/jmmshn/orgs",
"repos_url": "https://api.github.com/users/jmmshn/repos",
"events_url": "https://api.github.com/users/jmmshn/events{/privacy}",
"received_events_url": "https://api.github.com/users/jmmshn/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @jmmshn "
] | 2021-03-24T17:03:50
| 2021-03-25T23:18:43
|
2021-03-24T18:56:30Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
When the decomposition energy are missing the `get_max_instability` and `get_min_instability` functions all error out.
Now they will just return None
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2101/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2101/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2101",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2101",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2101.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2101.patch",
"merged_at": "2021-03-24T18:56:30Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2102
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2102/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2102/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2102/events
|
https://github.com/materialsproject/pymatgen/issues/2102
| 840,062,529
|
MDU6SXNzdWU4NDAwNjI1Mjk=
| 2,102
|
Speed up SpacegroupAnalyzer().get_crystal_system() with bisect module
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"For reference, this is the result I'm getting with both functions:\r\n\r\n```py\r\nlog_gvrh.sg_number.apply(get_crystal_system).value_counts()\r\nlog_gvrh.sg_number.apply(new_get_crystal_system).value_counts()\r\n```\r\n\r\n```\r\ncubic 3847\r\ntetragonal 2056\r\northorhombic 1682\r\nhexagonal 1430\r\ntrigonal 978\r\nmonoclinic 795\r\ntriclinic 199\r\nName: sg_number, dtype: int64\r\n```",
"While I have no doubt using bisect might yield some small improvements, I doubt it is necessary. The problem was that the earlier code was badly written with unnecessary complexity. I just modified it. Can you pull the latest pymatgen and try the code again? I am pretty sure it is not that bad.\r\n\r\nFurther, I doubt this is a performance bottleneck for most applications.",
"Not a bottleneck but faster never hurts and like you said, it wasn't good code.",
"My version is still slightly faster but I like yours for readability. :)\r\n```\r\nold took 0.396 sec\r\nnew took 0.292 sec\r\nspeed-up: 1.356\r\n```",
"Quite some variance in those timings...",
"Thanks. Yeah, bisect is worth it mainly when we are trying to find something in a large list. With only 7 crystal systems, the benefit is not that large."
] | 2021-03-24T19:01:17
| 2021-03-24T19:43:50
|
2021-03-24T19:43:50Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Is your feature request related to a problem? Please describe.**
`SpacegroupAnalyzer().get_crystal_system()` can be sped up with the [native `bisect` module](https://docs.python.org/3/library/bisect.html).
**Describe the solution you'd like**
I suggest to replace the current implementation of `get_crystal_system()`
<details>
<summary>Current <code>get_crystal_system()</code></summary>
```py
def get_crystal_system(spg_number):
"""
Get the crystal system for the structure, e.g., (triclinic,
orthorhombic, cubic, etc.).
Returns:
(str): Crystal system for structure or None if system cannot be detected.
"""
def f(i, j):
return i <= spg_number <= j
cs = {
"triclinic": (1, 2),
"monoclinic": (3, 15),
"orthorhombic": (16, 74),
"tetragonal": (75, 142),
"trigonal": (143, 167),
"hexagonal": (168, 194),
"cubic": (195, 230),
}
crystal_sytem = None
for k, v in cs.items():
if f(*v):
crystal_sytem = k
break
return crystal_sytem
```
</details>
with
```py
def new_get_crystal_system(spg_number: int) -> Union[str, None]:
boundaries = (1, 3, 16, 75, 143, 168, 195, 231)
crystal_systems = (
None,
"triclinic",
"monoclinic",
"orthorhombic",
"tetragonal",
"trigonal",
"hexagonal",
"cubic",
None,
)
return crystal_systems[bisect(boundaries, spg_number)]
```
**Additional context**
In this initial testing script, both versions give the same result while the new one is 4.4x faster:
```py
from bisect import bisect
from time import perf_counter
from typing import Union
from matminer.datasets import load_dataset
from tqdm import tqdm
tqdm.pandas()
log_gvrh = load_dataset("matbench_log_gvrh")
log_gvrh[["sg_symbol", "sg_number"]] = log_gvrh.progress_apply(
lambda row: row.structure.get_space_group_info(), axis=1, result_type="expand"
)
# copied from Pymatgen (https://git.io/JYTeF)
def get_crystal_system(spg_number):
"""
Get the crystal system for the structure, e.g., (triclinic,
orthorhombic, cubic, etc.).
Returns:
(str): Crystal system for structure or None if system cannot be detected.
"""
def f(i, j):
return i <= spg_number <= j
cs = {
"triclinic": (1, 2),
"monoclinic": (3, 15),
"orthorhombic": (16, 74),
"tetragonal": (75, 142),
"trigonal": (143, 167),
"hexagonal": (168, 194),
"cubic": (195, 230),
}
crystal_sytem = None
for k, v in cs.items():
if f(*v):
crystal_sytem = k
break
return crystal_sytem
def new_get_crystal_system(spg_number: int) -> Union[str, None]:
boundaries = (1, 3, 16, 75, 143, 168, 195, 231)
crystal_systems = (
None,
"triclinic",
"monoclinic",
"orthorhombic",
"tetragonal",
"trigonal",
"hexagonal",
"cubic",
None,
)
return crystal_systems[bisect(boundaries, spg_number)]
start = perf_counter()
for _ in range(100):
log_gvrh.sg_number.apply(get_crystal_system)
print(f"new took {(old_time := perf_counter() - start):.3f} sec")
start = perf_counter()
for _ in range(100):
log_gvrh.sg_number.apply(new_get_crystal_system)
print(f"new took {(new_time := perf_counter() - start):.3f} sec")
print(f"speed-up: {old_time / new_time:.3f}")
```
I get
```
old took 1.613 sec
new took 0.369 sec
speed-up: 4.366
```
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2102/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2102/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2103
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2103/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2103/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2103/events
|
https://github.com/materialsproject/pymatgen/issues/2103
| 840,946,446
|
MDU6SXNzdWU4NDA5NDY0NDY=
| 2,103
|
get_symmetrically_distinct_miller_indices gives wrong values for primitive fcc cell
|
{
"login": "bernstei",
"id": 1202357,
"node_id": "MDQ6VXNlcjEyMDIzNTc=",
"avatar_url": "https://avatars.githubusercontent.com/u/1202357?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/bernstei",
"html_url": "https://github.com/bernstei",
"followers_url": "https://api.github.com/users/bernstei/followers",
"following_url": "https://api.github.com/users/bernstei/following{/other_user}",
"gists_url": "https://api.github.com/users/bernstei/gists{/gist_id}",
"starred_url": "https://api.github.com/users/bernstei/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/bernstei/subscriptions",
"organizations_url": "https://api.github.com/users/bernstei/orgs",
"repos_url": "https://api.github.com/users/bernstei/repos",
"events_url": "https://api.github.com/users/bernstei/events{/privacy}",
"received_events_url": "https://api.github.com/users/bernstei/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I just realized that this is probably explained by the code not magically knowing the convention that for fcc people use conventional cell Miller indices instead of ones based on the reciprocal of the primitive cell lattice (combined with the fact that conventional 211 can be expressed with max index 1 in primitive reciprocal indices, so even the count of inequivalent indices is different). \r\n\r\nFeel free to close this issue, although I might suggest making this point more explicit in the docs.",
"Yes. The code works primarily for conventional cells. If you supply a conventional fcc crystal, that will yield the correct result. That is usually how people define miller indices. ",
"Also, in general, that method is not called on its own but rather used as part of SlabGenerator. In SlabGenerator, the doc says \"Note that to ensure that the miller indices correspond to usual crystallographic definitions, you should supply a conventional unit cell structure.\""
] | 2021-03-25T13:44:31
| 2021-03-25T15:33:07
|
2021-03-25T15:33:07Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
When passed a primitive fcc cell and max index of 1, `get_symmetrically_distinct_miller_indices()` fails to find (1 0 0), but reports multiple equivalent copies of (110) and (111) differing by negative signs
**To Reproduce**
run the following python script
```
from ase.atoms import Atoms
from pymatgen.io.ase import AseAtomsAdaptor
from pymatgen.core.surface import get_symmetrically_distinct_miller_indices
struct = AseAtomsAdaptor.get_structure( Atoms('Al', cell=[[1,1,0],[1,0,1],[0,1,1]], pbc=[True]*3))
print(struct)
print(get_symmetrically_distinct_miller_indices(struct, 1))
```
**Expected behavior**
Last line should be some permutation of `[(1, 1, 1), (1, 1, 0), (1, 0, 0)]`
**Screenshots**
```
> python tt.py
Full Formula (Al1)
Reduced Formula: Al
abc : 1.414214 1.414214 1.414214
angles: 60.000000 60.000000 60.000000
Sites (1)
# SP a b c
--- ---- --- --- ---
0 Al 0 0 0
[(1, 1, 1), (1, 1, 0), (1, 1, -1), (1, 0, -1)]
```
**Desktop (please complete the following information):**
OS X 10.15, using macports python3 3.8.7, pymatgen version
```
> pip show pymatgen
Name: pymatgen
Version: 2022.0.5
```
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2103/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2103/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2104
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2104/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2104/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2104/events
|
https://github.com/materialsproject/pymatgen/pull/2104
| 841,272,371
|
MDExOlB1bGxSZXF1ZXN0NjAxMDI4OTQ3
| 2,104
|
Update mp2020 to v2021.03.22 DB release, without activating by default
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @rkingsbury \r\n\r\n> There is one test in test_matproj.py that is failing, and I think this is a result of the new DB release rather than any changes in this PR.\r\n\r\nI'll handle this ..."
] | 2021-03-25T20:13:03
| 2021-03-26T04:30:12
|
2021-03-25T20:16:11Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Refit MP2020 corrections to the 2021.03.22 DB release using updated experimental data (see #2006 )
* Add the ability to point `MaterialsProject2020Compatibility` to an alternate configuration file
* Various docstring changes and updates ported from #2006
The purpose of this PR is to enable test builds of the MP database using the new corrections without requiring a new pymatgen release. With this PR, we will be able to iterate on correction values in an alternate configuration file, separate from the one distributed with pymatgen.
## TODO (if any)
* There is one test in `test_matproj.py` that is failing, and I think this is a result of the new DB release rather than any changes in this PR.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2104/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2104/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2104",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2104",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2104.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2104.patch",
"merged_at": "2021-03-25T20:16:10Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2105
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2105/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2105/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2105/events
|
https://github.com/materialsproject/pymatgen/pull/2105
| 844,874,669
|
MDExOlB1bGxSZXF1ZXN0NjA0MDc4NTI5
| 2,105
|
Fix for correct bonds_broken in slab generation
|
{
"login": "fyalcin",
"id": 20908329,
"node_id": "MDQ6VXNlcjIwOTA4MzI5",
"avatar_url": "https://avatars.githubusercontent.com/u/20908329?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/fyalcin",
"html_url": "https://github.com/fyalcin",
"followers_url": "https://api.github.com/users/fyalcin/followers",
"following_url": "https://api.github.com/users/fyalcin/following{/other_user}",
"gists_url": "https://api.github.com/users/fyalcin/gists{/gist_id}",
"starred_url": "https://api.github.com/users/fyalcin/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/fyalcin/subscriptions",
"organizations_url": "https://api.github.com/users/fyalcin/orgs",
"repos_url": "https://api.github.com/users/fyalcin/repos",
"events_url": "https://api.github.com/users/fyalcin/events{/privacy}",
"received_events_url": "https://api.github.com/users/fyalcin/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks for submitting this PR. Do you mind adding a unittest for the bug? You just need to add it to pymatgen/core/tests/test_surface.py. ",
"Just added a unit test. It is the first unit test I ever wrote so I hope I was able to pull it off :)",
"Looks good to me. Thanks!\r\n"
] | 2021-03-30T16:51:27
| 2021-04-01T23:21:59
|
2021-04-01T16:44:15Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
`SlabGenerator._get_c_ranges()` takes in a `bonds` dictionary and returns a set of tuples of fractional z-coordinates of neighboring (nearest) atoms if they have distinct z-coordinates. This set is then used in the `SlabGenerator.get_slabs()` method in order to calculate the total number of broken bonds. The issue with the current version is that the tuples are stored in a `set()` which can only have unique elements so all but one of the bonds between say z_coord1 and z_coord2 are ignored.
I tested this by simply generating Si(111) slabs, calculating the number of bonds broken reported by pymatgen, and comparing them by what I expect from looking at the slabs in VESTA.
Here's the code I used to generate the slabs;
```
from pymatgen.core.surface import SlabGenerator
from pymatgen.symmetry.analyzer import SpacegroupAnalyzer
from pymatgen import MPRester
struct = MPRester().get_structure_by_material_id('mp-149')
bulk_conv = SpacegroupAnalyzer(struct).get_conventional_standard_structure()
SG = SlabGenerator(initial_structure = bulk_conv,
miller_index = [1,1,1],
center_slab = True,
primitive = True,
in_unit_planes = True,
lll_reduce = True,
max_normal_search= 1,
min_slab_size = 7,
min_vacuum_size = 12)
slabs = SG.get_slabs(bonds = {('Si','Si'):2.60}, ftol = 0.1, tol = 0.1, max_broken_bonds = 20,
symmetrize = False, repair = False)
for slab in slabs:
print(slab.energy)
```
This prints 0.5 for both terminations whereas looking at the slabs in VESTA, one expects 2 and 6.
By changing c_ranges from a set() to an empty list instead and appending to it, tuples for all the bonds are preserved and the above script prints 2 and 6.
For consistency, I also changed c_ranges to an empty list if `bonds` is `None` in `get_slabs()`.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2105/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2105/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2105",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2105",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2105.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2105.patch",
"merged_at": "2021-04-01T16:44:15Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2106
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2106/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2106/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2106/events
|
https://github.com/materialsproject/pymatgen/pull/2106
| 859,111,254
|
MDExOlB1bGxSZXF1ZXN0NjE2MjQ0NjEw
| 2,106
|
finalize MP2020 correction values
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@mkhorton there seem to be some problems with the test environment - it complains about enumlib and openbabel not being installed / available",
"That message is just saying those tests have been skipped -- the actual error is that PMG_MAPI_KEY isn't set. I can look into it.",
"Thanks @rkingsbury :)"
] | 2021-04-15T17:49:05
| 2021-04-20T17:45:14
|
2021-04-20T17:44:53Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Final correction fitting for `MaterialsProject2020Compatibility`
* Updates the fitted values to be fully consistent with the new `expt_formation_energy_kingsbury` dataset (see https://github.com/hackingmaterials/matminer/pull/602)
* Use `MaterialsProject2020Compatibility` by default in `get_pourbaix_entries`
* Fixes `DeprecationWarning` in `pourbaix_diagram.py` related to `latexify_ion` (fixes #2056)
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2106/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2106/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2106",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2106",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2106.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2106.patch",
"merged_at": "2021-04-20T17:44:52Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2107
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2107/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2107/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2107/events
|
https://github.com/materialsproject/pymatgen/issues/2107
| 860,052,386
|
MDU6SXNzdWU4NjAwNTIzODY=
| 2,107
|
NaN should be removed from any init functions
|
{
"login": "jmmshn",
"id": 14003693,
"node_id": "MDQ6VXNlcjE0MDAzNjkz",
"avatar_url": "https://avatars.githubusercontent.com/u/14003693?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jmmshn",
"html_url": "https://github.com/jmmshn",
"followers_url": "https://api.github.com/users/jmmshn/followers",
"following_url": "https://api.github.com/users/jmmshn/following{/other_user}",
"gists_url": "https://api.github.com/users/jmmshn/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jmmshn/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jmmshn/subscriptions",
"organizations_url": "https://api.github.com/users/jmmshn/orgs",
"repos_url": "https://api.github.com/users/jmmshn/repos",
"events_url": "https://api.github.com/users/jmmshn/events{/privacy}",
"received_events_url": "https://api.github.com/users/jmmshn/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 4735709485,
"node_id": "LA_kwDOACgets8AAAABGkUxLQ",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/compatability",
"name": "compatability",
"color": "E25438",
"default": false,
"description": "Concerning pymatgen compatibility with different OS, Python versions, numpy versions, etc."
}
] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Using `None` instead of `np.nan` seems like a safe-enough change. PR incoming.",
"Looks like `uncertainty` isn't able to handle `None`:\r\n\r\n```py\r\n> if std_dev < 0 and not isinfinite(std_dev):\r\nE TypeError: '<' not supported between instances of 'NoneType' and 'int'\r\n```\r\n\r\n@jmmshn Would `float(\"nan\")` play better with database validation than `np.nan`? I suspect it might be the same.",
"So looks like this happened in the mean time?\r\nhttps://github.com/materialsvirtuallab/monty/pull/379",
"I think this might be fixed now automatically by monty chanages in the past year. Although if the entries were serializing properly now then this must have been fixed at some point.\r\n\r\nBut either way I think it's still to good idea to get rid of NaN's in any function arguments, even somthing that `math.inf` will be better.",
"@jmmshn Thanks for the monty link. That's good context to have!\r\n\r\n@rkingsbury Do you anticipate any issues with using `math.inf` instead of `np.nan` as default uncertainty value? `math.inf` seems less semantic for a missing value.",
"None, inf and Nan have different meanings. None tells me the uncertainty is not determined. Inf means a huge uncertainty was computed. NaN means a divide by 0 occurred, i.e., there are no datapoints to determine uncertainty. I would use None when uncertainty was not computed at all.",
"> I would use None when uncertainty was not computed at all.\r\n\r\nSame. That's what I did yesterday in https://github.com/materialsproject/pymatgen/pull/3021 but doesn't work with the [`uncertainties` lib](https://pypi.org/project/uncertainties).\r\n\r\n\r\n```py\r\nstd_dev = None\r\n\r\n @std_dev.setter\r\n def std_dev(self, std_dev):\r\n \r\n # We force the error to be float-like. Since it is considered\r\n # as a standard deviation, it must be either positive or NaN:\r\n # (Note: if NaN < 0 is False, there is no need to test\r\n # separately for NaN. But this is not guaranteed, even if it\r\n # should work on most platforms.)\r\n> if std_dev < 0 and not isinfinite(std_dev):\r\nE TypeError: '<' not supported between instances of 'NoneType' and 'int'\r\n\r\n/opt/hostedtoolcache/Python/3.11.3/x64/lib/python3.11/site-packages/uncertainties/core.py:2798: TypeError\r\n```",
"IIRC when implementing the uncertainties we made a deliberate choice that 'np.nan' means \"unknown uncertainty\". That was important when fitting energy corrections because 'np.nan' can be used in mathematical operations whereas 'None' cannot.\r\n\r\n",
"I guess I don't understand what mathematical operations can be performed logically on an unknown uncertainty. It is like saying the uncertainty is the word \"nothing\" and I am performing a mathematical operation on \"nothing\" and a number. For logical uncertainties, either all values in the chain have well-defined uncertainties or the whole operation has no output.",
"> I guess I don't understand what mathematical operations can be performed logically on an unknown uncertainty.\r\n\r\nSome aggregation methods can handle `NaN` but not `None`. E.g.\r\n\r\n```py\r\nmax([1,2, float('nan')])\r\n>>> 2\r\n\r\nmax([1,2, None])\r\nTraceback (most recent call last):\r\n File \"<stdin>\", line 1, in <module>\r\nTypeError: '>' not supported between instances of 'NoneType' and 'int'\r\n```",
"Sorry but what is the max of 1, 2 and NaN in logical terms?",
"I believe our `CorrectionCalculator` and `EnergyAdjustment` classes use the `ufloat` class from `uncertainties`, and this class allows you to do multiplication with error propagation, so if you have\r\n\r\n```\r\n>>> ufloat(2, 0.2) * ufloat(3, np.nan)\r\nufloat(6, np.nan)\r\n```\r\n\r\nWhereas this could not be done with `None`.\r\n\r\n@awvio did most of that development and would know better than me. @awvio, do you have any thoughts on `np.nan` vs. `None` here?",
"I think the main reason we went with `np.nan` instead of `None` is because of the practical reason that `None` doesn't work with the ufloat class. In general, I believe `None` doesn't work well with mathematical operations - as shown in by Janosh's example with the `max()` function. Another example of where `np.nan` works better for us than `None` is when we calculate the average uncertainty of a set of experimental energies. We can use `np.nanmean()` to straightforwardly calculate the average uncertainties, ignoring the unknown uncertainties that have been set to `np.nan`.",
"@awvio Makes sense to me.\r\n\r\nAll things considered, esp. that https://github.com/materialsvirtuallab/monty/pull/379 now handles NaN more gracefully as @jmmshn pointed out, I think the status quo is actually fine. Let's close this without any changes. Happy to reopen if anyone feels strongly otherwise."
] | 2021-04-16T17:58:38
| 2023-05-30T22:55:47
|
2023-05-30T22:55:47Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
An example is shown here:
https://github.com/materialsproject/pymatgen/blob/04beca9af8e575005a40117f1779f1d464c0a697/pymatgen/entries/computed_entries.py#L49
I think these should be removed from any MSONable init function since they don't play well with database and model validation.
I'm currently just replacing these with 0's in any emmet model that contains `ComputedEntry`s
https://github.com/materialsproject/emmet/blob/ebcc50bc47a8b65d661877f9edeb7f4087a7b4e5/emmet-core/emmet/core/utils.py#L117
but if the NaN's are physically meaningful I think we need to replace them with `None`.
@mkhorton @rkingsbury @shyamd
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2107/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2107/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2108
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2108/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2108/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2108/events
|
https://github.com/materialsproject/pymatgen/pull/2108
| 860,116,382
|
MDExOlB1bGxSZXF1ZXN0NjE3MDc2MzM3
| 2,108
|
Radial site perturbation and local_env patch
|
{
"login": "nwinner",
"id": 8825901,
"node_id": "MDQ6VXNlcjg4MjU5MDE=",
"avatar_url": "https://avatars.githubusercontent.com/u/8825901?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/nwinner",
"html_url": "https://github.com/nwinner",
"followers_url": "https://api.github.com/users/nwinner/followers",
"following_url": "https://api.github.com/users/nwinner/following{/other_user}",
"gists_url": "https://api.github.com/users/nwinner/gists{/gist_id}",
"starred_url": "https://api.github.com/users/nwinner/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/nwinner/subscriptions",
"organizations_url": "https://api.github.com/users/nwinner/orgs",
"repos_url": "https://api.github.com/users/nwinner/repos",
"events_url": "https://api.github.com/users/nwinner/events{/privacy}",
"received_events_url": "https://api.github.com/users/nwinner/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39910127)\n\nCoverage decreased (-0.6%) to 83.014% when pulling **82c54470ab958cce369dd70f6cce4893a66f1f31 on nwinner:site-perturb-patch-2** into **5ba1d857cfe1867913e1d7981d2285e79a9233e0 on materialsproject:master**.\n",
"Thanks @nwinner, this looks useful! Apologies this was left to sit for a while"
] | 2021-04-16T19:46:35
| 2021-06-17T00:33:01
|
2021-06-17T00:33:01Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Feature 1: I've added a new site transformation called the RadialSiteDistortionTransformation which takes a selected site, and perturbs the atoms around it. The way I've written is that the first nearest neighbors are perturbed by the specified distance, and then subsequent sites follow a 1/r decay, which seems like the most intuitive way to me.
* Feature 2: A rudimentary unit test has been added
* Fix 1: While writing this transformation, I discovered a bug in MinimumDistanceNN from the local_env module. While this is tagged as being applicable to both structures and molecules, it failed for Molecules. This is because it assumed it could find a periodic image of sites. I added a check to ensure that that when its not a structure, this is not needed.
## Additional dependencies introduced (if any)
* None
## TODO (if any)
Nominally not a WIP, unless some recommendations exist for the new transformation.
## Checklist
- [x ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [ ] All linting and tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2108/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2108/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2108",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2108",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2108.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2108.patch",
"merged_at": "2021-06-17T00:33:00Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2109
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2109/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2109/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2109/events
|
https://github.com/materialsproject/pymatgen/pull/2109
| 860,120,329
|
MDExOlB1bGxSZXF1ZXN0NjE3MDc5NTg1
| 2,109
|
FermiDos patch
|
{
"login": "nwinner",
"id": 8825901,
"node_id": "MDQ6VXNlcjg4MjU5MDE=",
"avatar_url": "https://avatars.githubusercontent.com/u/8825901?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/nwinner",
"html_url": "https://github.com/nwinner",
"followers_url": "https://api.github.com/users/nwinner/followers",
"following_url": "https://api.github.com/users/nwinner/following{/other_user}",
"gists_url": "https://api.github.com/users/nwinner/gists{/gist_id}",
"starred_url": "https://api.github.com/users/nwinner/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/nwinner/subscriptions",
"organizations_url": "https://api.github.com/users/nwinner/orgs",
"repos_url": "https://api.github.com/users/nwinner/repos",
"events_url": "https://api.github.com/users/nwinner/events{/privacy}",
"received_events_url": "https://api.github.com/users/nwinner/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-04-16T19:53:38
| 2021-05-27T16:56:27
|
2021-05-27T13:11:17Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
A simple fix. FermiDos at present uses 1 - f0(), which for very small numbers, causes python to collapse the result to 0. While such small numbers are within the noise of the dos, the function should be more stable. I moved the minus sign into the f0 function, which is equivalent, and has numpy handle all the precision, thereby keeping the small numbers.
Details:
func_old <-> 1-f0(E, fermi,T)
func_new <-> f0(-E, -fermi, T)
A basic example of comparing the two functions
```
from pymatgen.electronic_structure.dos import f0
energies = np.linspace(-1,1,10)
old_f = 1 - f0(energies, 0, 300)
new_f = f0(-energies, -0, 300)
diff = np.subtract(new_f, old_f)
print(diff)
>>> array([ 1.58759376e-17, -4.80244672e-17, -3.25951158e-17, -5.97598685e-17,
7.11236625e-17, 1.11022302e-16, -1.11022302e-16, 0.00000000e+00,
0.00000000e+00, 0.00000000e+00])
```
The array shows that the results are slightly different. This type of rounding error seems like it shouldn't cause issue, but in fact it does. Here are the cb_integral and vb_integrals calculated for efermi=2.522630877151964:
```
func_new -> 1.0281301354247369e-35 1.5922015812226065e-23
func_old -> 1.0281301354247369e-35 0.0
```
So you can see that even though the differences seem small initially, we end up having the vb_integral go to zero even when it's larger than the cb_integral because of the difference in precision thresholds, which leads to errors in the calculated values whenever the vb_integral is small.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2109/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2109/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2109",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2109",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2109.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2109.patch",
"merged_at": "2021-05-27T13:11:17Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2110
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2110/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2110/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2110/events
|
https://github.com/materialsproject/pymatgen/pull/2110
| 862,866,300
|
MDExOlB1bGxSZXF1ZXN0NjE5MzY1NjE4
| 2,110
|
Xps
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi Shyue, \r\n\r\nIs there a reason you didn't add galore as a dependency? Galore only has numpy, Matplotlib and SciPy as dependencies (all of which are dependencies of pymatgen already), so it is not much bloat.\r\n\r\nIf there is something stopping you from using galore as a library (such as the issue you raised here: https://github.com/SMTG-UCL/galore/issues/28) I'd be happy to get that resolved. I think it is better for both packages if there isn't the duplication of data. For example, we occasionally find mistakes in the tabulated data and we are planning on adding more datasets in the future.\r\n\r\n",
"I guess the main reasons are:\r\n1. Smearing was already supported in the DOS object (although only Gaussian and not Lorentzian, though that is not difficult to add).\r\n2. pymatgen already supports other spectra (XANES for instance)\r\n3. There are certain code design choices I prefer, e.g., OOP, static constructors, etc. \r\n\r\nThe issue I reported in galore was just one. In the end, I found it more difficult than necessary to generate XPS from galore. I also feel that smearing should be part of the pymatgen default package since it can be used in many instances, rather than requiring a dependency to do smearing. I am not asking for any credit for this - in fact, the documentation explicitly points to the Galore citation. ",
"Ok. It seems to me that all of those issues do not prevent pymatgen from using galore as a library? Your comments are instead reasons to use the pymatgen interface to galore over galore itself (which is a reasonable preference).\r\n\r\nIf pymatgen were to use galore as a library, I think there are two options:\r\n1. The simplest would be to just import the galore datasets rather than copying the data into pymatgen. \r\n2. Better, I think, would be to replace the entirety of the contents of `XPS.from_dos` to just use the galore api which does all of those things, and just return the XPS object at the end.\r\n\r\nEither way, both of the options are preferable to copying the data into pymatgen, which has the downsides that I mentioned before. Namely, the data might have bugs and will be updated in the future.",
"Hi @shyuep, another factor I noticed is that `galore` is a GPL licensed code, so we really can't import the data into pymatgen directly.\r\n\r\nI'd definitely vote for using it as an optional dependency given that galore has minimal and overlapping dependencies with pymatgen. So something like what @utf suggested in point 2 above.",
"The dataset I used from galore is from a 1980s work. If needed, I can easily parse the data from the original work. If the license is an issue, that's what I will do. I do not wish to introduce another dependency on a function that already exists in pymatgen and a dataset that belongs in the public domain. ",
"Parsing, proofing and formatting a dataset is still an effort, I'm not sure it's in our best interests to re-invent the wheel here when a clean dependency exists that we could use. I think it's to our benefit to use solutions from the community when they exist and are well maintained.",
"To echo Matt's point. It was a lot of effort to parse those datasets (note that Galore includes XPS, UPS, and HAXPES weights with more in the pipeline) and several of those datasets are actually fitted and not just parsing of raw tabulated data.\r\n\r\nI can imagine that a future version of pymatgen would want to include the UPS and HAXPES functionality of galore. So adding galore as an (optional) dependency will give a lot of features for a very small additional footprint. Note, the entire galore installation is only ~500 kb.",
"@utf At this juncture, I *only* need XPS. Further, smearing was already implemented in the DOS object since v1 of pymatgen. As of now, I don't see a reason to add a dependency, even an optional one, for this capability. If and when it is deemed that we need a lot more functionality from galore, I will add it as a dependency. \r\n\r\nPls note that adding any dependency in general requires a consideration of the benefits vs costs. For instance, even though galore only depends on numpy, scipy and matplotlib, there is maintenance there too. E.g., numpy and scipy may deprecate stuff and galore needs to be kept up. Galore may depend on a different version of numpy from pymatgen. Future python versions may also break stuff in either pymatgen or galore that needs to be fixed. Based on the last commit date, galore was last updated in Jul 2019. We have gone through many numpy/scipy versions since then. Further, I generally do not add dependencies for data, but rather only code functionality. Right now, I am not using any code functionality from galore.\r\n\r\nNow, the only things I need are the cross-sections. If it bothers everyone so much that I am using the data file from galore (even though I attributed the entire piece of code to the galore publication), I will delete it and just get another student to parse it out. I just spent 5 mins opening up that 1985 Yeh paper in Acrobat and \"Export to Plain Text\" already yields a semi-usable Table 1. "
] | 2021-04-20T14:16:13
| 2021-04-21T14:37:05
|
2021-04-20T14:16:18Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Port of Galore code for XPS generation from DOS.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2110/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2110/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2110",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2110",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2110.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2110.patch",
"merged_at": "2021-04-20T14:16:18Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2111
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2111/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2111/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2111/events
|
https://github.com/materialsproject/pymatgen/pull/2111
| 863,244,083
|
MDExOlB1bGxSZXF1ZXN0NjE5Njg3OTg2
| 2,111
|
rm spurious warnings filter from GibbsComputedStructureEntryTest
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks Ryan"
] | 2021-04-20T21:09:12
| 2021-04-20T21:35:49
|
2021-04-20T21:35:43Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Remove a no-longer-needed warnings filter from a test in `GibbsComputedStructureEntryTest`. This had been removed previously but was somehow reinstated amidst the shuffle of MP2020-related merges, and was causing tests to fail.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2111/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2111/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2111",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2111",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2111.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2111.patch",
"merged_at": "2021-04-20T21:35:43Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2112
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2112/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2112/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2112/events
|
https://github.com/materialsproject/pymatgen/issues/2112
| 864,645,303
|
MDU6SXNzdWU4NjQ2NDUzMDM=
| 2,112
|
Numpy numerical types in electronic_structure.bandstructure.Kpoint.as_dict result
|
{
"login": "zooks97",
"id": 8887252,
"node_id": "MDQ6VXNlcjg4ODcyNTI=",
"avatar_url": "https://avatars.githubusercontent.com/u/8887252?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/zooks97",
"html_url": "https://github.com/zooks97",
"followers_url": "https://api.github.com/users/zooks97/followers",
"following_url": "https://api.github.com/users/zooks97/following{/other_user}",
"gists_url": "https://api.github.com/users/zooks97/gists{/gist_id}",
"starred_url": "https://api.github.com/users/zooks97/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/zooks97/subscriptions",
"organizations_url": "https://api.github.com/users/zooks97/orgs",
"repos_url": "https://api.github.com/users/zooks97/repos",
"events_url": "https://api.github.com/users/zooks97/events{/privacy}",
"received_events_url": "https://api.github.com/users/zooks97/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-04-22T07:41:42
| 2021-04-22T14:15:32
|
2021-04-22T14:15:32Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
In the [`electronic_structure.bandstructure.Kpoint.as_dict` method](https://github.com/materialsproject/pymatgen/blob/6737f23caaf646cc58425561cf08339acf77b396/pymatgen/electronic_structure/bandstructure.py#L127), the numpy arrays `frac_coords` and `cart_coords` are converted to lists using `list()`, which leaves the coordinates as numpy data types.
This causes issues when using the `as_dict` method to convert the object into a dictionary for storing in mongodb using [mincepy_sci](https://github.com/muhrin/mincepy_sci).
This issue is inherited by objects which contain `Kpoint`s and use their `as_dict` method in their own `as_dict` methods, such as `BandStructure`.
A possible solution is to use the numpy array method `tolist()` which also converts the list contents to python types.
|
{
"login": "zooks97",
"id": 8887252,
"node_id": "MDQ6VXNlcjg4ODcyNTI=",
"avatar_url": "https://avatars.githubusercontent.com/u/8887252?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/zooks97",
"html_url": "https://github.com/zooks97",
"followers_url": "https://api.github.com/users/zooks97/followers",
"following_url": "https://api.github.com/users/zooks97/following{/other_user}",
"gists_url": "https://api.github.com/users/zooks97/gists{/gist_id}",
"starred_url": "https://api.github.com/users/zooks97/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/zooks97/subscriptions",
"organizations_url": "https://api.github.com/users/zooks97/orgs",
"repos_url": "https://api.github.com/users/zooks97/repos",
"events_url": "https://api.github.com/users/zooks97/events{/privacy}",
"received_events_url": "https://api.github.com/users/zooks97/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2112/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2112/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2113
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2113/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2113/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2113/events
|
https://github.com/materialsproject/pymatgen/pull/2113
| 864,723,042
|
MDExOlB1bGxSZXF1ZXN0NjIwOTA3MjYx
| 2,113
|
Use numpy tolist method in electronic_structure.bandstructure.Kpoint.as_dict
|
{
"login": "zooks97",
"id": 8887252,
"node_id": "MDQ6VXNlcjg4ODcyNTI=",
"avatar_url": "https://avatars.githubusercontent.com/u/8887252?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/zooks97",
"html_url": "https://github.com/zooks97",
"followers_url": "https://api.github.com/users/zooks97/followers",
"following_url": "https://api.github.com/users/zooks97/following{/other_user}",
"gists_url": "https://api.github.com/users/zooks97/gists{/gist_id}",
"starred_url": "https://api.github.com/users/zooks97/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/zooks97/subscriptions",
"organizations_url": "https://api.github.com/users/zooks97/orgs",
"repos_url": "https://api.github.com/users/zooks97/repos",
"events_url": "https://api.github.com/users/zooks97/events{/privacy}",
"received_events_url": "https://api.github.com/users/zooks97/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Pls add a unittest to check for this pls. Thanks.",
"> Pls add a unittest to check for this pls. Thanks.\r\n\r\nDone!",
"\n[](https://coveralls.io/builds/39046363)\n\nCoverage decreased (-0.7%) to 83.021% when pulling **49ac26104f98496915d4f2ee94543b444db89099 on zooks97:kpoint-asdict-patch** into **68e4ebd6cbe0b6f71413d9689cd7d64a650a92fb on materialsproject:master**.\n",
"Thanks!"
] | 2021-04-22T09:10:05
| 2021-04-22T14:12:20
|
2021-04-22T14:12:14Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Use `np.ndarray.tolist()` method instead of python `list()` to convert k-point coordinates in `pymatgen.electronic_structure.bandstructure.Kpoint.as_dict()`.
This fixes issue #2112, where numpy numerical types (e.g. numpy.float64) exist in the result of `Kpoint.as_dict()`.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2113/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2113/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2113",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2113",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2113.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2113.patch",
"merged_at": "2021-04-22T14:12:14Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2114
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2114/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2114/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2114/events
|
https://github.com/materialsproject/pymatgen/issues/2114
| 864,783,432
|
MDU6SXNzdWU4NjQ3ODM0MzI=
| 2,114
|
Outdated BoltztrapRunner?
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"There is already a pymatgen.electronic_structure.boltztrap2 module (https://github.com/materialsproject/pymatgen/blob/master/pymatgen/electronic_structure/boltztrap2.py).",
"Oops. But then how come `atomate` doesn't use it?\r\n\r\nhttps://github.com/hackingmaterials/atomate/blob/7bd29a52e5b6f965b73ec035e65fc6c556f4164f/atomate/vasp/firetasks/run_calc.py#L20\r\n\r\nAlso, shouldn't the unhelpful link be updated regardless?",
"Well, the link can be updated if you wish, but the likelihood is that the module will be junked soon anyway. As for atomate, you can pose that question to @computron ."
] | 2021-04-22T10:18:18
| 2021-04-22T13:27:59
|
2021-04-22T13:19:56Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Is your feature request related to a problem? Please describe.**
`BoltztrapRunner` requires `x_trans` to be in path (which appears to be outdated) and links to an [empty page](www.icams.de/content/research/software-development/boltztrap) to get it.
**Describe the solution you'd like**
https://github.com/materialsproject/pymatgen/blob/04beca9af8e575005a40117f1779f1d464c0a697/pymatgen/electronic_structure/boltztrap.py#L54-L66
Perhaps the best solution would be to update `BoltztrapRunner` to use BoltzTraP2 whose official page is https://www.imc.tuwien.ac.at/index.php?id=21094 and which can installed easily with `pip install BoltzTraP2`.
**Describe alternatives you've considered**
More detailed installation instructions for `x_trans`.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2114/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2114/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2115
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2115/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2115/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2115/events
|
https://github.com/materialsproject/pymatgen/pull/2115
| 866,440,887
|
MDExOlB1bGxSZXF1ZXN0NjIyMzM2ODE2
| 2,115
|
Xdatcar fix
|
{
"login": "nkeilbart",
"id": 68927593,
"node_id": "MDQ6VXNlcjY4OTI3NTkz",
"avatar_url": "https://avatars.githubusercontent.com/u/68927593?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/nkeilbart",
"html_url": "https://github.com/nkeilbart",
"followers_url": "https://api.github.com/users/nkeilbart/followers",
"following_url": "https://api.github.com/users/nkeilbart/following{/other_user}",
"gists_url": "https://api.github.com/users/nkeilbart/gists{/gist_id}",
"starred_url": "https://api.github.com/users/nkeilbart/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/nkeilbart/subscriptions",
"organizations_url": "https://api.github.com/users/nkeilbart/orgs",
"repos_url": "https://api.github.com/users/nkeilbart/repos",
"events_url": "https://api.github.com/users/nkeilbart/events{/privacy}",
"received_events_url": "https://api.github.com/users/nkeilbart/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39099092)\n\nCoverage decreased (-0.6%) to 83.032% when pulling **0f65f7b33e512f260fd29b75ff5deb5582974351 on nkeilbart:xdatcar_fix** into **8bb2dd5252909ee8bb54d045b7ab71b4648592fc on materialsproject:master**.\n",
"Thanks."
] | 2021-04-23T21:24:41
| 2021-04-24T12:27:52
|
2021-04-24T12:27:48Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Updated Xdatcar class from pymatgen.io.vasp.outputs to update the lattice constants with each new step. Previously, only a single preamble was read in and used for the rest of the structures.
* Created a new XDATCAR test file named XDATCAR_6 to stay consistent with other filenames
* Added a test for Xdatcar that will assure that the lattice constants are different for the first and last image
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2115/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2115/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2115",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2115",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2115.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2115.patch",
"merged_at": "2021-04-24T12:27:48Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2116
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2116/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2116/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2116/events
|
https://github.com/materialsproject/pymatgen/issues/2116
| 869,231,920
|
MDU6SXNzdWU4NjkyMzE5MjA=
| 2,116
|
MPRestError: REST query returned with error status code 403
|
{
"login": "jrmc502",
"id": 67526255,
"node_id": "MDQ6VXNlcjY3NTI2MjU1",
"avatar_url": "https://avatars.githubusercontent.com/u/67526255?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jrmc502",
"html_url": "https://github.com/jrmc502",
"followers_url": "https://api.github.com/users/jrmc502/followers",
"following_url": "https://api.github.com/users/jrmc502/following{/other_user}",
"gists_url": "https://api.github.com/users/jrmc502/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jrmc502/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jrmc502/subscriptions",
"organizations_url": "https://api.github.com/users/jrmc502/orgs",
"repos_url": "https://api.github.com/users/jrmc502/repos",
"events_url": "https://api.github.com/users/jrmc502/events{/privacy}",
"received_events_url": "https://api.github.com/users/jrmc502/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-04-27T20:20:58
| 2021-05-06T18:56:25
|
2021-05-06T18:56:05Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
MPRestError: REST query returned with error status code 403
**To Reproduce**
Steps to reproduce the behavior:
from pymatgen.ext.matproj import MPRester
from pymatgen.analysis.pourbaix_diagram import PourbaixDiagram, PourbaixPlotter
#here I'm not using pymatgen.core since pourbaix_diagram isn't in core (rather analysis)
%matplotlib inline
mpr = MPRester()
entries = mpr.get_pourbaix_entries(["Cu"])
**Expected behavior**
Use API to create Pourbaix diagram
**Desktop (please complete the following information):**
- OS: Windows 10
- Version 10.0.19042
- Pymatgen: 2020.12.31
**Additional context**
Had issues with creating wheel for spglib for pip install pymatgen
tqdm should also be a required library but is currently not specified.
|
{
"login": "jrmc502",
"id": 67526255,
"node_id": "MDQ6VXNlcjY3NTI2MjU1",
"avatar_url": "https://avatars.githubusercontent.com/u/67526255?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jrmc502",
"html_url": "https://github.com/jrmc502",
"followers_url": "https://api.github.com/users/jrmc502/followers",
"following_url": "https://api.github.com/users/jrmc502/following{/other_user}",
"gists_url": "https://api.github.com/users/jrmc502/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jrmc502/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jrmc502/subscriptions",
"organizations_url": "https://api.github.com/users/jrmc502/orgs",
"repos_url": "https://api.github.com/users/jrmc502/repos",
"events_url": "https://api.github.com/users/jrmc502/events{/privacy}",
"received_events_url": "https://api.github.com/users/jrmc502/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2116/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2116/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2117
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2117/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2117/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2117/events
|
https://github.com/materialsproject/pymatgen/pull/2117
| 869,241,071
|
MDExOlB1bGxSZXF1ZXN0NjI0NjM4ODAw
| 2,117
|
Update bader_caller.py
|
{
"login": "nwinner",
"id": 8825901,
"node_id": "MDQ6VXNlcjg4MjU5MDE=",
"avatar_url": "https://avatars.githubusercontent.com/u/8825901?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/nwinner",
"html_url": "https://github.com/nwinner",
"followers_url": "https://api.github.com/users/nwinner/followers",
"following_url": "https://api.github.com/users/nwinner/following{/other_user}",
"gists_url": "https://api.github.com/users/nwinner/gists{/gist_id}",
"starred_url": "https://api.github.com/users/nwinner/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/nwinner/subscriptions",
"organizations_url": "https://api.github.com/users/nwinner/orgs",
"repos_url": "https://api.github.com/users/nwinner/repos",
"events_url": "https://api.github.com/users/nwinner/events{/privacy}",
"received_events_url": "https://api.github.com/users/nwinner/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @nwinner. For point 2, doesn’t the Bader binary already support this? (Ie getting volumes from the charge density, then integrating the spin density using those volumes) I seem to recall this feature was available in the Bader caller to do this automatically.",
"I hadn't seen that before, but after googling it looks like you can use \r\n\r\nbader CHGCAR_spin -ref CHGCAR_sum.\r\n\r\nIf you generated ACF using that method, then you would get a spin decorated structure by calling get_charge_decorated_structure(), but you would have to flip the sign... \r\n\r\nI'm not sure how to best make that a part of this module though. Thoughts?",
"For the Chgcar interface, we automatically do that analysis when a spin\nchannel is present (I believe it gets added to the summary dictionary). I’m\nnot sure how that could be done for cube. One option would be to something\nlike, get_decorated_structure with two args: the field to integrate and,\noptionally, a reference field?\n\nOn Tue, Apr 27, 2021 at 13:51, nwinner ***@***.***> wrote:\n\n> I hadn't seen that before, but after googling it looks like you can use\n>\n> bader CHGCAR_spin -ref CHGCAR_sum.\n>\n> If you generated ACF using that method, then you would get a spin\n> decorated structure by calling get_charge_decorated_structure(), but you\n> would have to flip the sign...\n>\n> I'm not sure how to best make that a part of this module though. Thoughts?\n>\n> —\n> You are receiving this because you commented.\n>\n>\n> Reply to this email directly, view it on GitHub\n> <https://github.com/materialsproject/pymatgen/pull/2117#issuecomment-827921556>,\n> or unsubscribe\n> <https://github.com/notifications/unsubscribe-auth/AAWWWRATSIBKD532LXLVM7TTK4PU3ANCNFSM43VW37OA>\n> .\n>\n",
"\n[](https://coveralls.io/builds/39319274)\n\nCoverage decreased (-0.7%) to 83.007% when pulling **0b7955ace91b0cddbe4523af0761803c6c20d7cb on nwinner:patch-1** into **df0f9ba74f9b1ba54bb6ede88badb31b7cc1cdb4 on materialsproject:master**.\n",
"Okay so I don't think we need a reference field in decorated structure. It does look like just specifying cghcar_ref when initializing BaderAnalysis, will let you parse a spin density file using the electron density as a reference. So it works as is. \r\n\r\nThe name is not great, as chgcar_ref does seem vasp-specific, but its really a \"general\" reference as far as bader is concerned. \r\n\r\nAs for how to change get_decorated_structure, should I put \"spin decorated structure\" back in, with with the knowledge that you have to set the reference chgcar appropriately, or maybe, should it get_charge_decorated-structure be changed to allow for this? I'm hesitant to rename things because of compatibility. ",
"Okay the newest commit should has a function called get_decorated_structure(). What do you think @mkhorton ?",
"Thanks @nwinner, I clarified the docstring a bit otherwise this looks good to me -- ok to merge?",
"Looks okay to me. Hopefully this doesn't need to be updated again anytime soon."
] | 2021-04-27T20:34:00
| 2021-05-03T21:56:33
|
2021-05-03T21:06:13Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
My last PR for the bader caller had some issues, which this PR resolves. (edited after discussion below)
* Fix 1: Previously, I had "self.nelects" for cube analysis created by looking for the site property with that name. It didn't occur to me that the self.structure object is not user-defined, but created from the cube file, and this property will never exist. I removed this, and changed the functions to have an optional nelect(s) argument for when parsing cube files.
* Feature 1: I have added get_decorated_structure method. Which is like the previous "get_spin_decorated_structure", but more general. Assuming you ran bader with an appropriate reference using `chgref_filename`, this lets you assign a generic property to a structure using the reference file for the partitioning and the main file for the integrating. This can be, for example, be used to parse a spin density cube file to get spin decorated structure or a hartree cube file to get a electrostatic potential decorated structure.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2117/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2117/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2117",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2117",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2117.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2117.patch",
"merged_at": "2021-05-03T21:06:13Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2118
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2118/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2118/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2118/events
|
https://github.com/materialsproject/pymatgen/issues/2118
| 869,673,036
|
MDU6SXNzdWU4Njk2NzMwMzY=
| 2,118
|
Missing Hard-Coded path when using MontyEncoder/Decoder for MP2020Compat
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @CompRhys thanks for reporting this. The file `MP2020Compatibility.yaml` should be distributed with pymatgen and should be at the location listed in your error message. Is it possible that you serialized the entries using a newer version and are trying to open them with an old version? Basically, it's not clear to me why the file is not there.",
"So what I think happens is the config file is there and on computer A when I saved the entry it took the pymatgen path from computer A. When I tried to load on computer B it looked for the path on computer A and failed as it had saved that path as the config file. I have worked around this by just commenting out the `if-else`\nI highlighted and just loaded the distributed file.\n\nThe branch I was using is level with master. I was thinking if you want to keep the option of having a custom config file then add in something like `os.path.isfile()` to the if clause to check if the custom file is available?",
"```python\r\nimport json\r\nimport time\r\n\r\nfrom monty.json import MontyEncoder\r\nfrom pymatgen.ext.matproj import MPRester\r\n\r\nMP_API_KEY = \"very_nice_key\"\r\n\r\nm = MPRester(MP_API_KEY)\r\n# # Materials project provides a python API via pymatgen\r\n# # API key obtained from https://materialsproject.org/open\r\n\r\ndata = m.get_entries(\r\n chemsys_formula_id_criteria={\"material_id\": {\"$in\": [\"mp-10064\"]}},\r\n inc_structure=True,\r\n property_data=[\"formation_energy_per_atom\",\r\n \"e_above_hull\"]\r\n)\r\n\r\ndate = time.strftime(\"%d_%m_%Y\")\r\nwith open(f\"test-bug-mp-query_{date}.json\", 'w') as outfile:\r\n [outfile.write(json.dumps(e, cls=MontyEncoder)+\"/n\") for e in data]\r\n```\r\n\r\n```json\r\n{\"@module\": \"pymatgen.entries.computed_entries\", \"@class\": \"ComputedStructureEntry\", \"energy\": -19.93190347, \"composition\": {\"Si\": 1.0, \"O\": 2.0}, \"entry_id\": \"mp-10064\", \"correction\": -1.376, \"energy_adjustments\": [{\"@module\": \"pymatgen.entries.computed_entries\", \"@class\": \"CompositionEnergyAdjustment\", \"@version\": null, \"adj_per_atom\": -0.688, \"n_atoms\": 2.0, \"uncertainty_per_atom\": 0.002, \"name\": \"MP2020 anion correction (oxide)\", \"cls\": {\"@module\": \"pymatgen.entries.compatibility\", \"@class\": \"MaterialsProject2020Compatibility\", \"@version\": null, \"compat_type\": \"Advanced\", \"correct_peroxide\": true, \"check_potcar_hash\": false, \"config_file\": \"/home/reag2/PhD/pymatgen/pymatgen/entries/MP2020Compatibility.yaml\"}, \"description\": \"Composition-based energy adjustment\"}], \"parameters\": {\"run_type\": \"GGA\", \"is_hubbard\": false, \"pseudo_potential\": {\"functional\": \"PBE\", \"labels\": [\"Si\", \"O\"], \"pot_type\": \"paw\"}, \"hubbards\": {}, \"potcar_symbols\": [\"PBE Si\", \"PBE O\"], \"oxide_type\": \"oxide\"}, \"data\": {\"oxide_type\": \"oxide\", \"formation_energy_per_atom\": -2.0050824800000004, \"e_above_hull\": 1.272290046666666}, \"structure\": {\"@module\": \"pymatgen.core.structure\", \"@class\": \"Structure\", \"charge\": null, \"lattice\": {\"matrix\": [[0.0, 2.291416, 2.291416], [2.291416, 0.0, 2.291416], [2.291416, 2.291416, 0.0]], \"a\": 3.240551584238708, \"b\": 3.240551584238708, \"c\": 3.240551584238708, \"alpha\": 60.00000000000001, \"beta\": 60.00000000000001, \"gamma\": 60.00000000000001, \"volume\": 24.062559428747754}, \"sites\": [{\"species\": [{\"element\": \"Si\", \"occu\": 1}], \"abc\": [0.0, 0.0, 0.0], \"xyz\": [0.0, 0.0, 0.0], \"label\": \"Si\", \"properties\": {\"magmom\": 0.0}}, {\"species\": [{\"element\": \"O\", \"occu\": 1}], \"abc\": [0.25, 0.25, 0.25], \"xyz\": [1.145708, 1.145708, 1.145708], \"label\": \"O\", \"properties\": {\"magmom\": -0.0}}, {\"species\": [{\"element\": \"O\", \"occu\": 1}], \"abc\": [0.75, 0.75, 0.75], \"xyz\": [3.437124, 3.437124, 3.437124], \"label\": \"O\", \"properties\": {\"magmom\": -0.0}}]}, \"@version\": null}/n\r\n```\r\n\r\nI just made this minimal code to replicate and see that it is definitely storing a path particular to my machine, hope this helps debug!",
"Thanks @CompRhys this makes sense. How about the fix below? This way we avoid storing hard-coded paths when serializing entries.\r\n\r\n```\r\n # load corrections and uncertainties\r\n if config_file:\r\n self.config_file = config_file\r\n c = loadfn(self.config_file)\r\n else:\r\n self.config_file = None\r\n c = loadfn(os.path.join(MODULE_DIR, \"MP2020Compatibility.yaml\"))\r\n```\r\n\r\nI'm not real familiar with `MontyDecoder` so if you see a problem with this approach please say so. If not, would you mind opening a PR?",
"Also fwiw, I have used `loadfn` / `dumpfn` to serialize and deserialize `ComputedEntry` across multiple machines and have not run into this problem, so I wonder if it's specific to `MontyEncoder`?",
"That change solves the problem when I pull new entries from the API and save them locally as it now records the config path as \"null\" but it still leaves an issue with the old files where I have the hardcoded path. With the old setup `loadfn`/`dumpfn` failed in the same way for me.\r\n\r\nWith it being fixed for new data requests probably if someone has chosen to use a custom config file that isn't available you probably want to code to crash as otherwise a user might think they're using their custom settings if we let the code default to the pymatgen settings so would suggest adding raising a ValueError and then I will just `crtl+h` replace the old path in the data I downloaded and saved already.\r\n\r\nSo something like this as a final change?\r\n```python\r\n if config_file:\r\n if os.path.isfile(config_file):\r\n self.config_file = config_file\r\n c = loadfn(self.config_file)\r\n else:\r\n raise ValueError(\r\n f\"Custom MaterialsProject2020Compatibility config_file ({config_file}) \"\r\n \"does not exist.\"\r\n )\r\n else:\r\n self.config_file = None\r\n c = loadfn(os.path.join(MODULE_DIR, \"MP2020Compatibility.yaml\"))\r\n```",
"Looks good to me! You make a great point about loading data on a different system that is missing the custom config file.",
"shall I make a PR with the above then?",
"Yes, that would be great. Thanks @CompRhys !"
] | 2021-04-28T08:29:00
| 2021-05-03T00:53:58
|
2021-05-03T00:53:58Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Yesterday I pulled some ComputedStructureEntries using MPRester and saved them to a json file using the `MonthEncoder`, today I am trying to load that json on a google colab instance to do some EDA work. I used the MontyDecoder and I got a path missing error corresponding to my directory setup on the other machine.
```python
from monty.json import MontyDecoder
with open(f"/content/drive/MyDrive/PhD/datasets/mp-query_27_04_2021.json", 'r') as outfile:
mp_data = json.load(outfile, cls=MontyDecoder)
```
```
---------------------------------------------------------------------------
FileNotFoundError Traceback (most recent call last)
<ipython-input-5-ea74e40bc2de> in <module>()
2
3 with open(f"/content/drive/MyDrive/PhD/datasets/mp-query_27_04_2021.json", 'r') as outfile:
----> 4 mp_data = json.load(outfile, cls=MontyDecoder)
5
15 frames
/usr/lib/python3.7/json/__init__.py in load(fp, cls, object_hook, parse_float, parse_int, parse_constant, object_pairs_hook, **kw)
294 cls=cls, object_hook=object_hook,
295 parse_float=parse_float, parse_int=parse_int,
--> 296 parse_constant=parse_constant, object_pairs_hook=object_pairs_hook, **kw)
297
298
/usr/lib/python3.7/json/__init__.py in loads(s, encoding, cls, object_hook, parse_float, parse_int, parse_constant, object_pairs_hook, **kw)
359 if parse_constant is not None:
360 kw['parse_constant'] = parse_constant
--> 361 return cls(**kw).decode(s)
/usr/local/lib/python3.7/dist-packages/monty/json.py in decode(self, s)
381 """
382 d = json.JSONDecoder.decode(self, s)
--> 383 return self.process_decoded(d)
384
385
/usr/local/lib/python3.7/dist-packages/monty/json.py in process_decoded(self, d)
369
370 if isinstance(d, list):
--> 371 return [self.process_decoded(x) for x in d]
372
373 return d
/usr/local/lib/python3.7/dist-packages/monty/json.py in <listcomp>(.0)
369
370 if isinstance(d, list):
--> 371 return [self.process_decoded(x) for x in d]
372
373 return d
/usr/local/lib/python3.7/dist-packages/monty/json.py in process_decoded(self, d)
354 data = {k: v for k, v in d.items() if not k.startswith("@")}
355 if hasattr(cls_, "from_dict"):
--> 356 return cls_.from_dict(data)
357 elif np is not None and modname == "numpy" and classname == "array":
358 if d["dtype"].startswith("complex"):
/usr/local/lib/python3.7/dist-packages/pymatgen/entries/computed_entries.py in from_dict(cls, d)
669 composition=d.get("composition", None),
670 correction=0,
--> 671 energy_adjustments=[dec.process_decoded(e) for e in d.get("energy_adjustments", {})],
672 parameters={k: dec.process_decoded(v) for k, v in d.get("parameters", {}).items()},
673 data={k: dec.process_decoded(v) for k, v in d.get("data", {}).items()},
/usr/local/lib/python3.7/dist-packages/pymatgen/entries/computed_entries.py in <listcomp>(.0)
669 composition=d.get("composition", None),
670 correction=0,
--> 671 energy_adjustments=[dec.process_decoded(e) for e in d.get("energy_adjustments", {})],
672 parameters={k: dec.process_decoded(v) for k, v in d.get("parameters", {}).items()},
673 data={k: dec.process_decoded(v) for k, v in d.get("data", {}).items()},
/usr/local/lib/python3.7/dist-packages/monty/json.py in process_decoded(self, d)
354 data = {k: v for k, v in d.items() if not k.startswith("@")}
355 if hasattr(cls_, "from_dict"):
--> 356 return cls_.from_dict(data)
357 elif np is not None and modname == "numpy" and classname == "array":
358 if d["dtype"].startswith("complex"):
/usr/local/lib/python3.7/dist-packages/monty/json.py in from_dict(cls, d)
167 :return: MSONable class.
168 """
--> 169 decoded = {k: MontyDecoder().process_decoded(v) for k, v in d.items() if not k.startswith("@")}
170 return cls(**decoded)
171
/usr/local/lib/python3.7/dist-packages/monty/json.py in <dictcomp>(.0)
167 :return: MSONable class.
168 """
--> 169 decoded = {k: MontyDecoder().process_decoded(v) for k, v in d.items() if not k.startswith("@")}
170 return cls(**decoded)
171
/usr/local/lib/python3.7/dist-packages/monty/json.py in process_decoded(self, d)
354 data = {k: v for k, v in d.items() if not k.startswith("@")}
355 if hasattr(cls_, "from_dict"):
--> 356 return cls_.from_dict(data)
357 elif np is not None and modname == "numpy" and classname == "array":
358 if d["dtype"].startswith("complex"):
/usr/local/lib/python3.7/dist-packages/monty/json.py in from_dict(cls, d)
168 """
169 decoded = {k: MontyDecoder().process_decoded(v) for k, v in d.items() if not k.startswith("@")}
--> 170 return cls(**decoded)
171
172 def to_json(self) -> str:
/usr/local/lib/python3.7/dist-packages/pymatgen/entries/compatibility.py in __init__(self, compat_type, correct_peroxide, check_potcar_hash, config_file)
867 else:
868 self.config_file = os.path.join(MODULE_DIR, "MP2020Compatibility.yaml")
--> 869 c = loadfn(self.config_file)
870 self.name = c["Name"]
871 self.comp_correction = c["Corrections"].get("CompositionCorrections", defaultdict(float))
/usr/local/lib/python3.7/dist-packages/monty/serialization.py in loadfn(fn, *args, **kwargs)
73 return msgpack.load(fp, *args, **kwargs) # pylint: disable=E1101
74 else:
---> 75 with zopen(fn, "rt") as fp:
76 if fmt == "yaml":
77 if yaml is None:
/usr/local/lib/python3.7/dist-packages/monty/io.py in zopen(filename, *args, **kwargs)
41 if ext in (".GZ", ".Z"):
42 return gzip.open(filename, *args, **kwargs)
---> 43 return io.open(filename, *args, **kwargs)
44
45
FileNotFoundError: [Errno 2] No such file or directory: '/home/reag2/pymatgen/pymatgen/entries/MP2020Compatibility.yaml'
```
the `/home/reag2/...` path is my development path on the cluster we have access to.
I think this can be fixed by changing line 864 in `pymatgen/entries/compatibility.py`
```python
if config_file:
self.config_file = config_file
else:
self.config_file = os.path.join(MODULE_DIR, "MP2020Compatibility.yaml")
```
but am not sure what works best for you setup wise so won't make a PR unless asked. Probably best to default and warn if the file doesn't exist?
@rkingsbury
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2118/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2118/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2119
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2119/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2119/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2119/events
|
https://github.com/materialsproject/pymatgen/pull/2119
| 869,749,463
|
MDExOlB1bGxSZXF1ZXN0NjI1MDYzMDAx
| 2,119
|
Use numpy tolist() in DOS-family object as_dict() methods
|
{
"login": "zooks97",
"id": 8887252,
"node_id": "MDQ6VXNlcjg4ODcyNTI=",
"avatar_url": "https://avatars.githubusercontent.com/u/8887252?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/zooks97",
"html_url": "https://github.com/zooks97",
"followers_url": "https://api.github.com/users/zooks97/followers",
"following_url": "https://api.github.com/users/zooks97/following{/other_user}",
"gists_url": "https://api.github.com/users/zooks97/gists{/gist_id}",
"starred_url": "https://api.github.com/users/zooks97/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/zooks97/subscriptions",
"organizations_url": "https://api.github.com/users/zooks97/orgs",
"repos_url": "https://api.github.com/users/zooks97/repos",
"events_url": "https://api.github.com/users/zooks97/events{/privacy}",
"received_events_url": "https://api.github.com/users/zooks97/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39184219)\n\nCoverage decreased (-0.6%) to 83.036% when pulling **ce0a5f14823fbd0d6a05a8e772dcef88a55c696c on zooks97:dos-asdict-patch** into **df0f9ba74f9b1ba54bb6ede88badb31b7cc1cdb4 on materialsproject:master**.\n",
"Thanks."
] | 2021-04-28T09:41:33
| 2021-04-28T19:56:41
|
2021-04-28T19:56:35Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Use `np.ndarray.tolist()` method instead of python `list()` to convert density of state (`Dos`, `FermiDos`, `CompleteDos`) energies and densities in their `as_dict()` methods.
This fixes an issue where numpy numerical types (e.g. numpy.float64) exist in the results of {`Dos`, `FermiDos`, `CompleteDos`}`.as_dict()`.
This allows these objects to be saved and stored more easily by [mincePy Sci](https://github.com/muhrin/mincepy_sci).
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2119/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2119/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2119",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2119",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2119.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2119.patch",
"merged_at": "2021-04-28T19:56:35Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2120
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2120/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2120/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2120/events
|
https://github.com/materialsproject/pymatgen/issues/2120
| 869,771,333
|
MDU6SXNzdWU4Njk3NzEzMzM=
| 2,120
|
Computed entry `__eq__` doesn't check class instance
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"closing as fixed by 56fa39fcd16e90489c24252e6b6a58520ea427ae "
] | 2021-04-28T10:04:43
| 2021-06-24T14:31:24
|
2021-06-24T14:31:24Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I have a `PhaseDiagram` made with `PDEntry`'s and wanted to get some results for a `ComputedEntry` relative to that phase diagram. As `PDEntry` doesn't have an `entry_id` this causes an error.
```
cse.data["E_ps_per_at"] = phad.get_phase_separation_energy(cse)
File "/home/reag2/pymatgen/pymatgen/analysis/phase_diagram.py", line 818, in get_phase_separation_energy
return self.get_decomp_and_phase_separation_energy(entry, **kwargs)[1]
File "/home/reag2/pymatgen/pymatgen/analysis/phase_diagram.py", line 728, in get_decomp_and_phase_separation_energy
return self.get_decomp_and_e_above_hull(entry, allow_negative=True)
File "/home/reag2/pymatgen/pymatgen/analysis/phase_diagram.py", line 628, in get_decomp_and_e_above_hull
if entry in list(self.stable_entries):
File "/home/reag2/pymatgen/pymatgen/entries/computed_entries.py", line 509, in __eq__
if self.entry_id and other.entry_id and self.entry_id == other.entry_id:
```
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2120/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2120/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2121
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2121/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2121/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2121/events
|
https://github.com/materialsproject/pymatgen/pull/2121
| 870,228,093
|
MDExOlB1bGxSZXF1ZXN0NjI1NDU3ODE1
| 2,121
|
Update cube.py
|
{
"login": "nwinner",
"id": 8825901,
"node_id": "MDQ6VXNlcjg4MjU5MDE=",
"avatar_url": "https://avatars.githubusercontent.com/u/8825901?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/nwinner",
"html_url": "https://github.com/nwinner",
"followers_url": "https://api.github.com/users/nwinner/followers",
"following_url": "https://api.github.com/users/nwinner/following{/other_user}",
"gists_url": "https://api.github.com/users/nwinner/gists{/gist_id}",
"starred_url": "https://api.github.com/users/nwinner/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/nwinner/subscriptions",
"organizations_url": "https://api.github.com/users/nwinner/orgs",
"repos_url": "https://api.github.com/users/nwinner/repos",
"events_url": "https://api.github.com/users/nwinner/events{/privacy}",
"received_events_url": "https://api.github.com/users/nwinner/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39200231)\n\nCoverage decreased (-0.7%) to 82.995% when pulling **270e3d9d6ad1f77902861842866a3cfc5945a3bb on nwinner:patch-2** into **df0f9ba74f9b1ba54bb6ede88badb31b7cc1cdb4 on materialsproject:master**.\n",
"Thanks!"
] | 2021-04-28T18:26:19
| 2021-04-28T20:05:35
|
2021-04-28T19:55:59Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Feature 1: Spherical masks of the cube data is now supported
* Feature 2: Using spherical masks, atomic site averages and totals are now supported natively. These can be used, respectively, for getting the average electrostatic potential around a site using a V_HARTREE file from CP2K or the total spin on a site using a SPIN_DENSITY file from CP2K. These features are useful as an alternative to Bader partitioning for these properties when the bader executable is not available.
* Fix 1: Previously, cube files were loaded using for loops, which was prohibitively slow for files >1Gb. Now it is done with numpy and is much faster.
## TODO (if any)
* While not necessary, it would be nice to support multiprocessing for the atomic site functions. These can be pretty expensive when the cube files are >1Gb. I previously used the multiprocessing module for this, but ran into some errors while trying it out. It was exceeding the memory available for the Drone, and since the Drone is supposed to be automatic/not require user modifications, I removed multiprocessing for robustness, but it would be nice to have it back at some point.
* New tests for the class methods have not been added yet.
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [x ] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2121/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2121/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2121",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2121",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2121.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2121.patch",
"merged_at": "2021-04-28T19:55:59Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2122
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2122/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2122/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2122/events
|
https://github.com/materialsproject/pymatgen/pull/2122
| 870,417,983
|
MDExOlB1bGxSZXF1ZXN0NjI1NjIyMTQ1
| 2,122
|
Update vdW radii to CRC handbook values
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Git does not seem well-equipped to diff `.json` files, so you can use this online tool to better see the changes:\r\n\r\nhttps://jsoncompare.com/#!/diff/id=6924b719277a2a2b7a630352828d155c&fullscreen/\r\n\r\nclick the 'lint' button on both panes and you will see each change nicely highlighted",
"\n[](https://coveralls.io/builds/39206217)\n\nCoverage decreased (-0.6%) to 83.007% when pulling **1c23fb56a667f93724a6abc6df3ebd7f7648edcb on rkingsbury:vdw_data** into **ad5cb53f6255f8af8428bbc6fdff74c50be342e8 on materialsproject:master**.\n",
"Thank you @rkingsbury , definitely happy to have data sourced from somewhere that isn't wikipedia here :-)"
] | 2021-04-28T21:51:44
| 2021-04-29T00:23:50
|
2021-04-29T00:07:17Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This PR updates the `van_der_waals_radius` data used by the `Element` class from Wikipedia data to the latest values in the in the CRC Handbook of Chemistry and Physics. Important changes from the previous values include:
- Revised the value for `H` down by 0.1 angstrom as recommended in a critical review by Rowland and Taylor
- Added values for many lanthanides, actinides, and transition metals that were previously absent
- Updated values or many transition metals
All data, and the justification for which values were chosen, can be found in
"Atomic Radii of the Elements" in CRC Handbook of Chemistry and Physics, 91st Ed.; Haynes, W.M., Ed.; CRC Press: Boca Raton, FL, 2010.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2122/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2122/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2122",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2122",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2122.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2122.patch",
"merged_at": "2021-04-29T00:07:17Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2123
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2123/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2123/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2123/events
|
https://github.com/materialsproject/pymatgen/pull/2123
| 870,442,941
|
MDExOlB1bGxSZXF1ZXN0NjI1NjQzOTk4
| 2,123
|
bug fix in SpacegroupAnalyzer
|
{
"login": "gpetretto",
"id": 8665734,
"node_id": "MDQ6VXNlcjg2NjU3MzQ=",
"avatar_url": "https://avatars.githubusercontent.com/u/8665734?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/gpetretto",
"html_url": "https://github.com/gpetretto",
"followers_url": "https://api.github.com/users/gpetretto/followers",
"following_url": "https://api.github.com/users/gpetretto/following{/other_user}",
"gists_url": "https://api.github.com/users/gpetretto/gists{/gist_id}",
"starred_url": "https://api.github.com/users/gpetretto/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/gpetretto/subscriptions",
"organizations_url": "https://api.github.com/users/gpetretto/orgs",
"repos_url": "https://api.github.com/users/gpetretto/repos",
"events_url": "https://api.github.com/users/gpetretto/events{/privacy}",
"received_events_url": "https://api.github.com/users/gpetretto/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39322138)\n\nCoverage decreased (-0.6%) to 83.011% when pulling **381b843cc02ee839391f259066a1056586c0f3c5 on gpetretto:bugfix** into **ce2fb4ee48a864aa766742caeec4b99c74e54610 on materialsproject:master**.\n",
"Thanks for fixing @gpetretto -- would it be possible to add a quick test case to catch this too? Thank you"
] | 2021-04-28T22:20:43
| 2021-05-06T14:22:33
|
2021-05-06T14:22:32Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
There is a problem in the `get_conventional_standard_structure` method of the `SpacegroupAnalyzer`.
The code below demonstrates the issue:
```python
from pymatgen.symmetry.analyzer import SpacegroupAnalyzer
from pymatgen.analysis.structure_matcher import StructureMatcher
from pymatgen.ext.matproj import MPRester
s = MPRester().get_structure_by_material_id("mp-761986")
sm = StructureMatcher(ltol=0.0001, stol=0.0001, angle_tol=0.1, primitive_cell=False, scale=False)
spga = SpacegroupAnalyzer(s)
sc = spga.get_conventional_standard_structure()
s_ref = spga.get_refined_structure()
print(sm.fit(s, s_ref))
print(sm.fit(s, sc))
```
Output:
```
True
False
```
This comes from the fact that in the triclinic case the reference structure can be modified by the call to `struct.get_reduced_structure("LLL")`, but the lattice object is not updated accordingly.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2123/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2123/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2123",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2123",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2123.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2123.patch",
"merged_at": "2021-05-06T14:22:32Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2124
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2124/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2124/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2124/events
|
https://github.com/materialsproject/pymatgen/pull/2124
| 871,232,128
|
MDExOlB1bGxSZXF1ZXN0NjI2Mjg1MTA4
| 2,124
|
#2118 Stop MP2020Corrections saving hardcoded path
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39232927)\n\nCoverage decreased (-0.6%) to 83.013% when pulling **ce45ed36a71a9e27e825ff84f7d3868ff28f9974 on CompRhys:master** into **cb9854d5ba11f216aafd3c9a49c9fceca485f218 on materialsproject:master**.\n",
"Thanks for the fix @CompRhys!"
] | 2021-04-29T16:43:11
| 2021-05-03T00:53:59
|
2021-05-03T00:53:58Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
resolves #2118 by leaving config path as None if custom path not given. Raises ValueError is a custom path given but the file does not exist.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2124/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2124/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2124",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2124",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2124.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2124.patch",
"merged_at": "2021-05-03T00:53:58Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2125
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2125/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2125/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2125/events
|
https://github.com/materialsproject/pymatgen/pull/2125
| 871,665,967
|
MDExOlB1bGxSZXF1ZXN0NjI2NjcwNDIy
| 2,125
|
Small tweaks / fixes to Q-Chem parsing
|
{
"login": "samblau",
"id": 7431223,
"node_id": "MDQ6VXNlcjc0MzEyMjM=",
"avatar_url": "https://avatars.githubusercontent.com/u/7431223?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/samblau",
"html_url": "https://github.com/samblau",
"followers_url": "https://api.github.com/users/samblau/followers",
"following_url": "https://api.github.com/users/samblau/following{/other_user}",
"gists_url": "https://api.github.com/users/samblau/gists{/gist_id}",
"starred_url": "https://api.github.com/users/samblau/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/samblau/subscriptions",
"organizations_url": "https://api.github.com/users/samblau/orgs",
"repos_url": "https://api.github.com/users/samblau/repos",
"events_url": "https://api.github.com/users/samblau/events{/privacy}",
"received_events_url": "https://api.github.com/users/samblau/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39245138)\n\nCoverage decreased (-0.6%) to 83.012% when pulling **9760bc5ff8e350d49690502c164934db087481c4 on samblau:qchem** into **cb9854d5ba11f216aafd3c9a49c9fceca485f218 on materialsproject:master**.\n",
"Ready to review at your convenience @shyuep ",
"This looks good to me, thanks @samblau"
] | 2021-04-29T23:19:25
| 2021-05-03T00:55:55
|
2021-05-03T00:55:47Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
* Modified encoding to avoid errors when opening Q-Chem output files
* Fixed bug for reading force job outputs
* Removed two outdated warnings
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2125/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2125/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2125",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2125",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2125.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2125.patch",
"merged_at": "2021-05-03T00:55:47Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2126
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2126/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2126/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2126/events
|
https://github.com/materialsproject/pymatgen/issues/2126
| 872,209,877
|
MDU6SXNzdWU4NzIyMDk4Nzc=
| 2,126
|
Source code link in documentation of surface module leads to 404
|
{
"login": "peterschindler",
"id": 45672907,
"node_id": "MDQ6VXNlcjQ1NjcyOTA3",
"avatar_url": "https://avatars.githubusercontent.com/u/45672907?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/peterschindler",
"html_url": "https://github.com/peterschindler",
"followers_url": "https://api.github.com/users/peterschindler/followers",
"following_url": "https://api.github.com/users/peterschindler/following{/other_user}",
"gists_url": "https://api.github.com/users/peterschindler/gists{/gist_id}",
"starred_url": "https://api.github.com/users/peterschindler/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/peterschindler/subscriptions",
"organizations_url": "https://api.github.com/users/peterschindler/orgs",
"repos_url": "https://api.github.com/users/peterschindler/repos",
"events_url": "https://api.github.com/users/peterschindler/events{/privacy}",
"received_events_url": "https://api.github.com/users/peterschindler/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I think this applies to every module in pymatgen.core as they appear to be linking to .../pymatgen/pymatgen/module.py rather than .../pymatgen/core/module.py"
] | 2021-04-30T09:20:15
| 2021-04-30T16:09:14
|
2021-04-30T16:09:14Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Clicking on any of the [source] links on the surface module documentation page leads to a 404 page: https://pymatgen.org/pymatgen.core.surface.html
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2126/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2126/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2127
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2127/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2127/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2127/events
|
https://github.com/materialsproject/pymatgen/pull/2127
| 872,663,388
|
MDExOlB1bGxSZXF1ZXN0NjI3NTQ2Nzkw
| 2,127
|
[WIP] change the relative location of the filename for source code lookup. Fixes #2126
|
{
"login": "adam-kerrigan",
"id": 38280242,
"node_id": "MDQ6VXNlcjM4MjgwMjQy",
"avatar_url": "https://avatars.githubusercontent.com/u/38280242?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/adam-kerrigan",
"html_url": "https://github.com/adam-kerrigan",
"followers_url": "https://api.github.com/users/adam-kerrigan/followers",
"following_url": "https://api.github.com/users/adam-kerrigan/following{/other_user}",
"gists_url": "https://api.github.com/users/adam-kerrigan/gists{/gist_id}",
"starred_url": "https://api.github.com/users/adam-kerrigan/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/adam-kerrigan/subscriptions",
"organizations_url": "https://api.github.com/users/adam-kerrigan/orgs",
"repos_url": "https://api.github.com/users/adam-kerrigan/repos",
"events_url": "https://api.github.com/users/adam-kerrigan/events{/privacy}",
"received_events_url": "https://api.github.com/users/adam-kerrigan/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks.",
"\n[](https://coveralls.io/builds/39267023)\n\nCoverage decreased (-0.6%) to 83.013% when pulling **e2158fc553a238c73e9c61abc9192f46e81daa66 on adam-kerrigan:doc_conf** into **cb9854d5ba11f216aafd3c9a49c9fceca485f218 on materialsproject:master**.\n"
] | 2021-04-30T14:57:58
| 2021-04-30T16:31:48
|
2021-04-30T16:09:15Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Generation of links to source code in the pymatgen.core have been fixed.
* Generation of links to source code where the line numbers cannot be found have been fixed.
This fixes #2126. I was unsure on the difference between conf-docset.py, conf-normal.py and conf.py so I only updated the ones with the linkcode_resolve function in.
## TODO (if any)
Generation of the Documentation, I couldn't find a guide to building the documentation and when I do I get far more changes than I expect as well as 608 warnings from sphinx some of which are listed below:
* Adds '()' to end of property names in genindex.html
* Adds a space to the end of the meta tags
* Removes alt="Documentation Home" from every page.
Also I have noticed that the documentation for BoltzTraP2 is missing on the [doc site](https://pymatgen.org/pymatgen.electronic_structure.boltztrap2.html) however when I build it is present.
Before a pull request can be merged, the following items must be checked:
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [ ] All linting and tests pass.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2127/reactions",
"total_count": 1,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 1,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2127/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2127",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2127",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2127.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2127.patch",
"merged_at": "2021-04-30T16:09:14Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2128
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2128/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2128/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2128/events
|
https://github.com/materialsproject/pymatgen/pull/2128
| 872,814,585
|
MDExOlB1bGxSZXF1ZXN0NjI3NjgzNTE4
| 2,128
|
Add auto oxidation state detection to MP2020 ; incremental update to correction values
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39270514)\n\nCoverage decreased (-0.6%) to 83.011% when pulling **faed4eff372fe402613e4427715e66cc3c9882ef on rkingsbury:default_mp2020** into **af726e3b4f6c93b3faf78671e4a3908eb2ae3932 on materialsproject:master**.\n",
"Thanks @rkingsbury "
] | 2021-04-30T16:24:37
| 2021-05-03T00:53:20
|
2021-05-03T00:53:17Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
- Automatically try to guess the oxidation state of compounds in `process_entries` if it is not already provided
- Small update to the MP2020 correction values as we finalize for the upcoming database release.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2128/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2128/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2128",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2128",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2128.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2128.patch",
"merged_at": "2021-05-03T00:53:17Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2129
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2129/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2129/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2129/events
|
https://github.com/materialsproject/pymatgen/pull/2129
| 873,502,538
|
MDExOlB1bGxSZXF1ZXN0NjI4MzA3Mjgz
| 2,129
|
Symmetrically equivalent surfaces in slabs
|
{
"login": "CifLord",
"id": 8036054,
"node_id": "MDQ6VXNlcjgwMzYwNTQ=",
"avatar_url": "https://avatars.githubusercontent.com/u/8036054?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CifLord",
"html_url": "https://github.com/CifLord",
"followers_url": "https://api.github.com/users/CifLord/followers",
"following_url": "https://api.github.com/users/CifLord/following{/other_user}",
"gists_url": "https://api.github.com/users/CifLord/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CifLord/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CifLord/subscriptions",
"organizations_url": "https://api.github.com/users/CifLord/orgs",
"repos_url": "https://api.github.com/users/CifLord/repos",
"events_url": "https://api.github.com/users/CifLord/events{/privacy}",
"received_events_url": "https://api.github.com/users/CifLord/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39319301)\n\nCoverage decreased (-0.7%) to 82.996% when pulling **f1182e02fd0990268d2ad969202da9063cb4e6d3 on richardtran415:surfaces** into **af726e3b4f6c93b3faf78671e4a3908eb2ae3932 on materialsproject:master**.\n",
"Thanks richard."
] | 2021-05-01T02:15:34
| 2021-05-05T22:26:12
|
2021-05-04T12:18:33Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
This pull request contains an improvement for analyzing surface symmetry of slabs. A slab now has symmetrically equivalent surfaces if (hkl) is a mirror plane, if [hkl] is a screw axis or if the slab has inversion symmetry.
Include a summary of major changes in bullet points:
* Modifications to is_symmetric() Slab class method. If slab contains inversion symmetry, the (hkl) plane is a mirror plane, or there is a screw axis about [hkl], the surfaces of the slab are symmetrically equivalent.
* Test for generating slabs that don't need inversion
* Remove really inefficient Slab class method for checking equivalent surfaces have_equivalent_surfaces()
* Modify SlabGenerator class method nonstoichiometric_symmetrized_slab() to use is_symmetric() when generating slabs
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2129/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2129/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2129",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2129",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2129.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2129.patch",
"merged_at": "2021-05-04T12:18:32Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2130
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2130/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2130/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2130/events
|
https://github.com/materialsproject/pymatgen/pull/2130
| 874,224,843
|
MDExOlB1bGxSZXF1ZXN0NjI4ODI3NDIy
| 2,130
|
Ensure all MP2020 corrections have unique names; tweak oxi state detection
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39341111)\n\nCoverage decreased (-0.7%) to 83.008% when pulling **c63779000d0ee50b6b0ec863628e96bfd76be64f on rkingsbury:default_mp2020** into **12d6e2b1b6b49f7c6524dec2f834b959205f459b on materialsproject:master**.\n"
] | 2021-05-03T04:56:40
| 2021-07-15T14:32:32
|
2021-05-06T14:24:58Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Ensures that energy corrections applied to each anion have unique names (e.g., N vs. Cl vs. Br).
If this is not the case, then in a compound with multiple corrected anions like `C2NCl`, `Compatibility` will apply a correction to the first anion, then think the second is a duplicate, warning
```
warnings.warn(
2021-05-01T11:22:13.8878461Z /users/home/ghrunner/actions-runner/_work/_tool/Python/3.8.7/x64/lib/python3.8/site-packages/pymatgen/entries/compatibility.py:579: UserWarning: Entry mp-567221 already has an energy adjustment called MP2020 anion correction, but its value differs from the value of -0.718 calculated here. This Entry will be discarded.
```
Also makes some tweaks to oxidation state detection to work better with the Builders.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2130/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2130/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2130",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2130",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2130.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2130.patch",
"merged_at": "2021-05-06T14:24:58Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2131
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2131/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2131/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2131/events
|
https://github.com/materialsproject/pymatgen/pull/2131
| 874,934,263
|
MDExOlB1bGxSZXF1ZXN0NjI5Mzg1OTkw
| 2,131
|
Fix QChem gzipped parsing and add gzipped test
|
{
"login": "samblau",
"id": 7431223,
"node_id": "MDQ6VXNlcjc0MzEyMjM=",
"avatar_url": "https://avatars.githubusercontent.com/u/7431223?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/samblau",
"html_url": "https://github.com/samblau",
"followers_url": "https://api.github.com/users/samblau/followers",
"following_url": "https://api.github.com/users/samblau/following{/other_user}",
"gists_url": "https://api.github.com/users/samblau/gists{/gist_id}",
"starred_url": "https://api.github.com/users/samblau/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/samblau/subscriptions",
"organizations_url": "https://api.github.com/users/samblau/orgs",
"repos_url": "https://api.github.com/users/samblau/repos",
"events_url": "https://api.github.com/users/samblau/events{/privacy}",
"received_events_url": "https://api.github.com/users/samblau/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39320108)\n\nCoverage decreased (-0.6%) to 83.01% when pulling **03372b1b25a8d2eaf8091bc0bda11f12311c2bdc on samblau:qchem** into **ce2fb4ee48a864aa766742caeec4b99c74e54610 on materialsproject:master**.\n",
"@mkhorton ready for review. Thanks!",
"Thanks @samblau "
] | 2021-05-03T21:44:47
| 2021-05-04T01:03:54
|
2021-05-04T01:03:41Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
My last PR fixed one QChem parsing problem, but introduced a new one - the zopen call could no longer accommodate gzipped output files. I have fixed this problem and also gzipped one of my already present test output files to ensure that the ability to parse gzipped outputs is now tested.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2131/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2131/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2131",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2131",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2131.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2131.patch",
"merged_at": "2021-05-04T01:03:41Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2132
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2132/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2132/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2132/events
|
https://github.com/materialsproject/pymatgen/pull/2132
| 874,986,728
|
MDExOlB1bGxSZXF1ZXN0NjI5NDI3NjA0
| 2,132
|
Speedup LammpsData.as_string; add as_lammpsdata method to CombinedData
|
{
"login": "htz1992213",
"id": 25351437,
"node_id": "MDQ6VXNlcjI1MzUxNDM3",
"avatar_url": "https://avatars.githubusercontent.com/u/25351437?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/htz1992213",
"html_url": "https://github.com/htz1992213",
"followers_url": "https://api.github.com/users/htz1992213/followers",
"following_url": "https://api.github.com/users/htz1992213/following{/other_user}",
"gists_url": "https://api.github.com/users/htz1992213/gists{/gist_id}",
"starred_url": "https://api.github.com/users/htz1992213/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/htz1992213/subscriptions",
"organizations_url": "https://api.github.com/users/htz1992213/orgs",
"repos_url": "https://api.github.com/users/htz1992213/repos",
"events_url": "https://api.github.com/users/htz1992213/events{/privacy}",
"received_events_url": "https://api.github.com/users/htz1992213/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39352112)\n\nCoverage decreased (-0.6%) to 83.018% when pulling **e11261e75ec5bfeef67d5afab6ddea0349197149 on htz1992213:master** into **ce2fb4ee48a864aa766742caeec4b99c74e54610 on materialsproject:master**.\n",
"Thanks. Can you add a unittest for the new method pls?",
"> Thanks. Can you add a unittest for the new method pls?\r\n\r\nI have added the tests accordingly. Thanks!",
"THanks!"
] | 2021-05-03T23:36:02
| 2021-05-05T12:24:44
|
2021-05-05T12:24:44Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
* Speedup LammpsData.as_string for non-hybrid data with large coeff sections
* Add as_lammpsdata method to CombinedData which allows CombinedData object to be converted to its superclass
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2132/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2132/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2132",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2132",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2132.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2132.patch",
"merged_at": "2021-05-05T12:24:44Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2133
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2133/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2133/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2133/events
|
https://github.com/materialsproject/pymatgen/pull/2133
| 875,634,105
|
MDExOlB1bGxSZXF1ZXN0NjI5OTM3ODI4
| 2,133
|
Add $van_der_waals input block to QCInput
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I'm fine with it as-is. It's a little awkward, but it would also be a little awkward to shoehorn mode into the van_der_waals dict.",
"\n[](https://coveralls.io/builds/39345618)\n\nCoverage decreased (-0.6%) to 83.004% when pulling **c04c01c306e202c5e3ebf736eefff6c6406e2da2 on rkingsbury:qcinput-vdw** into **1269f9cff8fd3e1988913907cef495b9e5a8bf26 on materialsproject:master**.\n"
] | 2021-05-04T16:32:45
| 2021-05-06T15:45:18
|
2021-05-06T14:23:10Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Adds support for custom vdW radii to `QCInput` and `QChemDictSet`. These radii are used in the construction of PCM cavities and when calculating charges.
## TODO
- [x] Feedback requested from @samblau regarding the `vdw_mode` kwarg that I had to add. This construction is a little bit awkward. An alternative would be to include a special 'mode' key in the `van_der_waals` dict. Please let me know if you have a preference.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2133/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2133/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2133",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2133",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2133.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2133.patch",
"merged_at": "2021-05-06T14:23:10Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2134
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2134/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2134/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2134/events
|
https://github.com/materialsproject/pymatgen/pull/2134
| 877,500,722
|
MDExOlB1bGxSZXF1ZXN0NjMxNDQzNDE2
| 2,134
|
Allow compressed log files for parse_lammps_log
|
{
"login": "ab5424",
"id": 57258530,
"node_id": "MDQ6VXNlcjU3MjU4NTMw",
"avatar_url": "https://avatars.githubusercontent.com/u/57258530?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ab5424",
"html_url": "https://github.com/ab5424",
"followers_url": "https://api.github.com/users/ab5424/followers",
"following_url": "https://api.github.com/users/ab5424/following{/other_user}",
"gists_url": "https://api.github.com/users/ab5424/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ab5424/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ab5424/subscriptions",
"organizations_url": "https://api.github.com/users/ab5424/orgs",
"repos_url": "https://api.github.com/users/ab5424/repos",
"events_url": "https://api.github.com/users/ab5424/events{/privacy}",
"received_events_url": "https://api.github.com/users/ab5424/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I didn't add any new tests, but changed the existing tests to compressed files. Hope that's ok.",
"\n[](https://coveralls.io/builds/39413703)\n\nCoverage decreased (-0.7%) to 83.008% when pulling **419d9111910cea9f520dfddc29f1873024bbf0d4 on ab5424:lammps** into **2ef73af08fea7d7d04d0376239f9ff1bfc3ff015 on materialsproject:master**.\n",
"Thanks."
] | 2021-05-06T13:30:41
| 2021-05-10T08:22:01
|
2021-05-06T14:21:44Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Change parsing lammps logs to zopen
## Checklist
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2134/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2134/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2134",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2134",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2134.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2134.patch",
"merged_at": "2021-05-06T14:21:44Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2135
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2135/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2135/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2135/events
|
https://github.com/materialsproject/pymatgen/pull/2135
| 878,250,164
|
MDExOlB1bGxSZXF1ZXN0NjMyMTAxMjQ0
| 2,135
|
Add van_der_waals support to QCInput.from_file
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/40653175)\n\nCoverage decreased (-0.6%) to 83.103% when pulling **c0b5d4e23ac289d005ffcc2b24cc914a1d823fca on rkingsbury:qcinput-vdw** into **41d82c498900b6dffdd5c3c0db6de2564bdf75be on materialsproject:master**.\n",
"Many thanks @rkingsbury!"
] | 2021-05-07T00:43:11
| 2021-07-15T14:33:03
|
2021-06-17T00:35:05Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Follow up to #2133 , adds support for reading the `van_der_waals` block from a QChem output file.
This PR also refactors `test_write_file`. Previously, this test used `QCinput.from_file` to construct a dict, and then compared the contents of the dicts. The problem with this approach is that if `QCInput.from_file` has a bug in it that prevents parsing a section (as was the case when I was developing this code), the test will still pass.
The refactored test performs a direct comparison of the text file written by `write_input` and the reference text file.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2135/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2135/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2135",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2135",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2135.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2135.patch",
"merged_at": "2021-06-17T00:35:05Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2136
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2136/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2136/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2136/events
|
https://github.com/materialsproject/pymatgen/pull/2136
| 879,109,568
|
MDExOlB1bGxSZXF1ZXN0NjMyODU2MzM3
| 2,136
|
Add progress bar to applying compatibility settings
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39899802)\n\nCoverage decreased (-0.6%) to 83.023% when pulling **111f3348360b11d3765f61f8ca4de5e028332566 on CompRhys:master** into **adee29710a382a2438a84109c4994d0a2f394620 on materialsproject:master**.\n",
"Thanks @CompRhys !"
] | 2021-05-07T14:29:23
| 2021-05-23T23:11:02
|
2021-05-23T23:05:53Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Add pymatgen `PBar` to applying compatibility settings to help impatient users like myself. I have by default suppressed the progress bar and added a boolean verbose flag to turn it on.
Changes made are superficial therefore no new tests added.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2136/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2136/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2136",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2136",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2136.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2136.patch",
"merged_at": "2021-05-23T23:05:53Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2137
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2137/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2137/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2137/events
|
https://github.com/materialsproject/pymatgen/issues/2137
| 879,285,388
|
MDU6SXNzdWU4NzkyODUzODg=
| 2,137
|
Inconsistent dependencies for Python 3.6
|
{
"login": "ajjackson",
"id": 2511383,
"node_id": "MDQ6VXNlcjI1MTEzODM=",
"avatar_url": "https://avatars.githubusercontent.com/u/2511383?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ajjackson",
"html_url": "https://github.com/ajjackson",
"followers_url": "https://api.github.com/users/ajjackson/followers",
"following_url": "https://api.github.com/users/ajjackson/following{/other_user}",
"gists_url": "https://api.github.com/users/ajjackson/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ajjackson/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ajjackson/subscriptions",
"organizations_url": "https://api.github.com/users/ajjackson/orgs",
"repos_url": "https://api.github.com/users/ajjackson/repos",
"events_url": "https://api.github.com/users/ajjackson/events{/privacy}",
"received_events_url": "https://api.github.com/users/ajjackson/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"We have abandoned py36 support given that most of the underlying scientific python packages have abandoned py36. "
] | 2021-05-07T16:10:46
| 2021-05-07T16:33:38
|
2021-05-07T16:33:38Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Installation with Python 3.6 using Pip initially fails due to the lack of a compatible Numpy version.
The setup.py of Pymatgen reports that Python >= 3.6 is required, and Numpy >= 1.20 is required. However, Numpy version 1.20 [only supports Python 3.7-3.9](https://numpy.org/devdocs/release/1.20.0-notes.html).
When I run `pip install pymatgen` interactively, it is able to recover by figuring out that the previous Pymatgen version needed an acceptable Numpy version and rolling back. But in the Travis-CI for Galore, a package that depends on Pymatgen, it raises an error and gives up. So I expect the reliability of this depends a bit on your pip version and/or install flags.
**To Reproduce**
```
conda create -n pymatgen-py3.6 --python=3.6
pip install pymatgen
```
**Expected behavior**
Install without tripping over the incompatible Numpy version.
**Desktop (please complete the following information):**
- OS: Fedora Workstation 33
- Version 2021.3.3
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2137/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2137/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2138
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2138/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2138/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2138/events
|
https://github.com/materialsproject/pymatgen/issues/2138
| 883,265,004
|
MDU6SXNzdWU4ODMyNjUwMDQ=
| 2,138
|
Structure.from_str reads from str(Structure)
|
{
"login": "goodwilling",
"id": 56999195,
"node_id": "MDQ6VXNlcjU2OTk5MTk1",
"avatar_url": "https://avatars.githubusercontent.com/u/56999195?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/goodwilling",
"html_url": "https://github.com/goodwilling",
"followers_url": "https://api.github.com/users/goodwilling/followers",
"following_url": "https://api.github.com/users/goodwilling/following{/other_user}",
"gists_url": "https://api.github.com/users/goodwilling/gists{/gist_id}",
"starred_url": "https://api.github.com/users/goodwilling/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/goodwilling/subscriptions",
"organizations_url": "https://api.github.com/users/goodwilling/orgs",
"repos_url": "https://api.github.com/users/goodwilling/repos",
"events_url": "https://api.github.com/users/goodwilling/events{/privacy}",
"received_events_url": "https://api.github.com/users/goodwilling/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @goodwilling, we've had some requests for a Structure.from_str before, however I don't think this is in general a good approach. The string representation is lossy. It would be better if we could handle Pandas conversion natively. I know Pandas support extensions and custom data types but I'm not familiar with the details.",
"Are you able to use `MontyEncoder`?\r\n\r\ne.g.\r\n\r\n```\r\nfrom monty.json import MontyEncoder\r\n\r\ndf.to_json(default_handler=MontyEncoder().encode)\r\n```",
"Thank you very much. MontyEncoder works well for DataFrame containing Structure data. "
] | 2021-05-10T03:23:26
| 2021-05-14T02:53:03
|
2021-05-14T02:53:03Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Is your feature request related to a problem? Please describe.**
It is necessary to convert str(Structure) to a Structure when a DataFrame contains column of structures and default_handler=str is set in DataFrame.to_json() in order to get rid of "OverflowError: Maximum recursion level reached".
**Describe the solution you'd like**
Strcture.from_str(str(struct), fmt="str") reads from str(struct), where struct is a Structure.
**Additional context**
The following code:
from pymatgen.core.structure import Structure
struct = Structure.from_file("Si2.cif")
print(str(struct))
gives the structure information:
Full Formula (Si2)
Reduced Formula: Si
abc : 3.839136 3.839136 3.839136
angles: 60.000000 60.000000 60.000000
Sites (2)
# SP a b c
--- ---- ----- ----- -----
0 Si 0.125 0.125 0.125
1 Si 0.875 0.875 0.875
However, there is no method to convert str(struct) to a Structure,
like
Structure.from_str(str(struct), fmt="str")
The feature should be necessary when a DataFrame contains column of structures
and default_handler=str is set in DataFrame.to_json() in order to get rid of
"OverflowError: Maximum recursion level reached".
|
{
"login": "goodwilling",
"id": 56999195,
"node_id": "MDQ6VXNlcjU2OTk5MTk1",
"avatar_url": "https://avatars.githubusercontent.com/u/56999195?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/goodwilling",
"html_url": "https://github.com/goodwilling",
"followers_url": "https://api.github.com/users/goodwilling/followers",
"following_url": "https://api.github.com/users/goodwilling/following{/other_user}",
"gists_url": "https://api.github.com/users/goodwilling/gists{/gist_id}",
"starred_url": "https://api.github.com/users/goodwilling/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/goodwilling/subscriptions",
"organizations_url": "https://api.github.com/users/goodwilling/orgs",
"repos_url": "https://api.github.com/users/goodwilling/repos",
"events_url": "https://api.github.com/users/goodwilling/events{/privacy}",
"received_events_url": "https://api.github.com/users/goodwilling/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2138/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2138/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2139
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2139/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2139/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2139/events
|
https://github.com/materialsproject/pymatgen/pull/2139
| 883,808,871
|
MDExOlB1bGxSZXF1ZXN0NjM3MjQ5OTMx
| 2,139
|
Add spacegroup details to SymmetrizedStructure string
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Great, thanks @CompRhys!"
] | 2021-05-10T09:22:19
| 2021-06-28T13:53:39
|
2021-05-11T20:04:53Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Wyckoff letters are meaningless without the context of the spacegroup, current SymmetrizedStructure __str__ doesn't include the spacegroup so I've added it in - this is purely a cosmetic change.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2139/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2139/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2139",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2139",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2139.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2139.patch",
"merged_at": "2021-05-11T20:04:53Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2140
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2140/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2140/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2140/events
|
https://github.com/materialsproject/pymatgen/pull/2140
| 887,586,357
|
MDExOlB1bGxSZXF1ZXN0NjQwODA4ODQx
| 2,140
|
Update MPScanRelaxSet doc string
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39579302)\n\nCoverage decreased (-0.6%) to 82.992% when pulling **4d3be5af32fed1edac61ae89a58ba22d3a3b41e1 on janosh:master** into **08aa5214a1b1a5fc6872de76b12cf97f5ceb03c9 on materialsproject:master**.\n"
] | 2021-05-11T15:11:45
| 2021-05-11T15:49:25
|
2021-05-11T15:49:25Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
[VASP officially supports r2SCAN in v6](https://www.vasp.at/forum/viewtopic.php?f=4&t=18058&p=20113&hilit=r2scan#p20113).
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2140/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2140/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2140",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2140",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2140.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2140.patch",
"merged_at": "2021-05-11T15:49:25Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2141
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2141/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2141/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2141/events
|
https://github.com/materialsproject/pymatgen/pull/2141
| 891,453,763
|
MDExOlB1bGxSZXF1ZXN0NjQ0MzIzMTE4
| 2,141
|
MP2020 corrections post-release housekeeping
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39820330)\n\nCoverage decreased (-0.6%) to 83.0% when pulling **daa95f3cdfaf131ef641614510f51f236c59ddba on rkingsbury:production_mp2020** into **9953bbe3431865bc637d7aff8a5199b21a3e7912 on materialsproject:master**.\n",
"@shyuep as part of this PR I'd like to address your request to minimize the dependency of Pourbaix Diagram tests on specific energy values from the database. However, I noticed that the test in question ([link](https://github.com/materialsproject/pymatgen/blob/9953bbe3431865bc637d7aff8a5199b21a3e7912/pymatgen/analysis/tests/test_pourbaix_diagram.py#L275)) appears to be designed specifically to test the pipeline from `MPRester` to `PourbaixDiagram`.\r\n\r\nSo, I left the the test intact, but I changed [this line](https://github.com/materialsproject/pymatgen/blob/9953bbe3431865bc637d7aff8a5199b21a3e7912/pymatgen/analysis/tests/test_pourbaix_diagram.py#L287) to test the identity of the stable entry rather than its energy. I also moved the custom ion test in [this line](https://github.com/materialsproject/pymatgen/blob/9953bbe3431865bc637d7aff8a5199b21a3e7912/pymatgen/analysis/tests/test_pourbaix_diagram.py#L298) to a different part of the code, and integrated that data into the local test file because this test doesn't have anything to do with MPRester at all.\r\n\r\nLet me know if you think this will accomplish what you were looking for.",
"Pls remove the MPRester dependency. The pipeline to MP should be tested in the ext.matproj so that we can isolate it. I don't think we should have external dependencies in test in the rest of pymatgen.",
"> \r\n> \r\n> Pls remove the MPRester dependency. The pipeline to MP should be tested in the ext.matproj so that we can isolate it. I don't think we should have external dependencies in test in the rest of pymatgen.\r\n\r\nDone. All `MPRester` related tests have been moved to `test_matproj.py`\r\n",
"Thanks."
] | 2021-05-13T23:16:18
| 2021-07-15T14:32:31
|
2021-05-21T17:12:38Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Some housekeeping changes following up the MP2020 database release v2021.05.13
* Silence the oxidation states warning when using `MPRester`
* Update the Pourbaix Diagram tests to be less dependent on specific energy values from the database
* Add ChemRxiv link for MP2020 manuscript to the `MaterialsProject2020Compatibility` docstring
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2141/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2141/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2141",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2141",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2141.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2141.patch",
"merged_at": "2021-05-21T17:12:38Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2142
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2142/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2142/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2142/events
|
https://github.com/materialsproject/pymatgen/issues/2142
| 891,853,336
|
MDU6SXNzdWU4OTE4NTMzMzY=
| 2,142
|
black changes all quotes to double, flake8 complains about them
|
{
"login": "flaviu-gostin",
"id": 37553536,
"node_id": "MDQ6VXNlcjM3NTUzNTM2",
"avatar_url": "https://avatars.githubusercontent.com/u/37553536?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/flaviu-gostin",
"html_url": "https://github.com/flaviu-gostin",
"followers_url": "https://api.github.com/users/flaviu-gostin/followers",
"following_url": "https://api.github.com/users/flaviu-gostin/following{/other_user}",
"gists_url": "https://api.github.com/users/flaviu-gostin/gists{/gist_id}",
"starred_url": "https://api.github.com/users/flaviu-gostin/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/flaviu-gostin/subscriptions",
"organizations_url": "https://api.github.com/users/flaviu-gostin/orgs",
"repos_url": "https://api.github.com/users/flaviu-gostin/repos",
"events_url": "https://api.github.com/users/flaviu-gostin/events{/privacy}",
"received_events_url": "https://api.github.com/users/flaviu-gostin/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 4812645414,
"node_id": "LA_kwDOACgets8AAAABHtskJg",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/outdated",
"name": "outdated",
"color": "2A890C",
"default": false,
"description": "Issues resulting from using outdated versions of pymatgen or dependencies"
},
{
"id": 5318918309,
"node_id": "LA_kwDOACgets8AAAABPQhApQ",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/linting",
"name": "linting",
"color": "5DBC83",
"default": false,
"description": "Linting and quality assurance"
}
] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@janosh: This can be closed as out of scope.",
"`flake8` was replaced with `ruff`."
] | 2021-05-14T11:41:27
| 2023-06-03T02:23:27
|
2023-06-03T02:23:16Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Conflict between blake and flake8 settings.
**To Reproduce**
Steps to reproduce the behavior:
1. Commit changes with `git commit`
2. pre-commit is triggered
3. black (I think) changes all quotes to double "
4. flake8 complains about the double quotes with: Q000 Remove bad quotes
Recommend adding in setup.cfg (according to [https://github.com/zheller/flake8-quotes])
[flake8]
inline-quotes = double
This stops the Q000 ...
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2142/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2142/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2143
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2143/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2143/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2143/events
|
https://github.com/materialsproject/pymatgen/pull/2143
| 891,969,192
|
MDExOlB1bGxSZXF1ZXN0NjQ0NzUyMDI3
| 2,143
|
Add new compact style for hkl annotations to peaks in diffraction patterns
|
{
"login": "flaviu-gostin",
"id": 37553536,
"node_id": "MDQ6VXNlcjM3NTUzNTM2",
"avatar_url": "https://avatars.githubusercontent.com/u/37553536?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/flaviu-gostin",
"html_url": "https://github.com/flaviu-gostin",
"followers_url": "https://api.github.com/users/flaviu-gostin/followers",
"following_url": "https://api.github.com/users/flaviu-gostin/following{/other_user}",
"gists_url": "https://api.github.com/users/flaviu-gostin/gists{/gist_id}",
"starred_url": "https://api.github.com/users/flaviu-gostin/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/flaviu-gostin/subscriptions",
"organizations_url": "https://api.github.com/users/flaviu-gostin/orgs",
"repos_url": "https://api.github.com/users/flaviu-gostin/repos",
"events_url": "https://api.github.com/users/flaviu-gostin/events{/privacy}",
"received_events_url": "https://api.github.com/users/flaviu-gostin/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks. Feel free to make it the default!",
"Thanks!",
"Thank you for accepting this change!",
"We should be thanking you! The XRDs now look much better!"
] | 2021-05-14T14:27:53
| 2021-05-14T16:19:32
|
2021-05-14T15:12:31Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Introducing a new optional style for the hkl labels in diffraction patterns (see example figure below).
The default style was not changed. However, I would recommend making this new more-compact style the default because it prevents overlap of labels as in https://matgenb.materialsvirtuallab.org/2013/01/01/Calculating-XRD-patterns.html (a similar image can be seen here at the bottom)
```python
from pymatgen.core import Lattice, Structure
from pymatgen.analysis.diffraction.xrd import XRDCalculator
import matplotlib.pyplot as plt
a = 4.209
latt = Lattice.cubic(a)
struct = Structure(latt, ["Cs", "Cl"], [[0, 0, 0], [0.5, 0.5, 0.5]])
c = XRDCalculator()
```
1. The new more-compact style:
```python
c.show_plot(struct, annotate_peaks='compact')
```
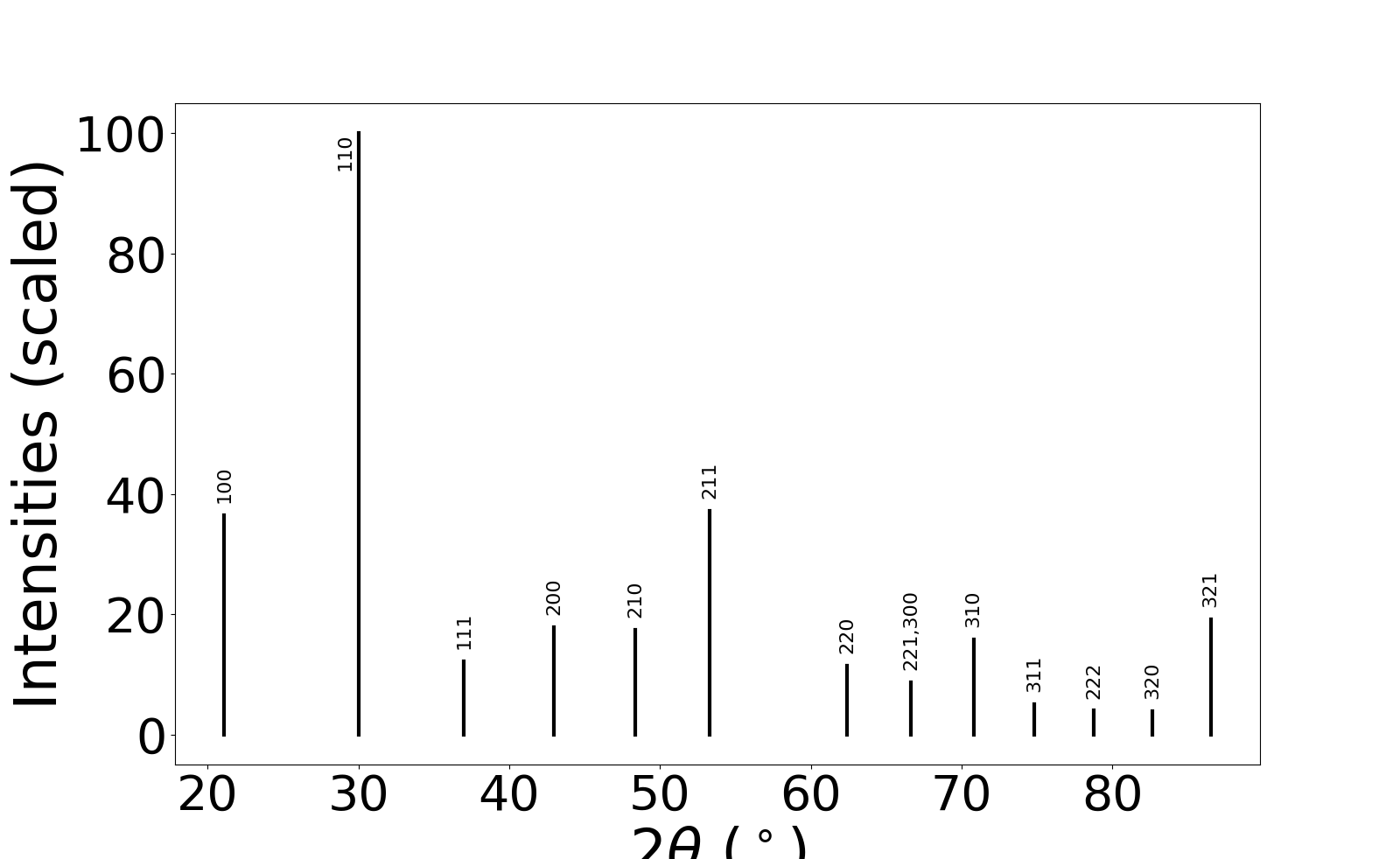
2. The current default style (previously: True, now: 'full'):
```python
c.show_plot(struct, annotate_peaks='full')
```
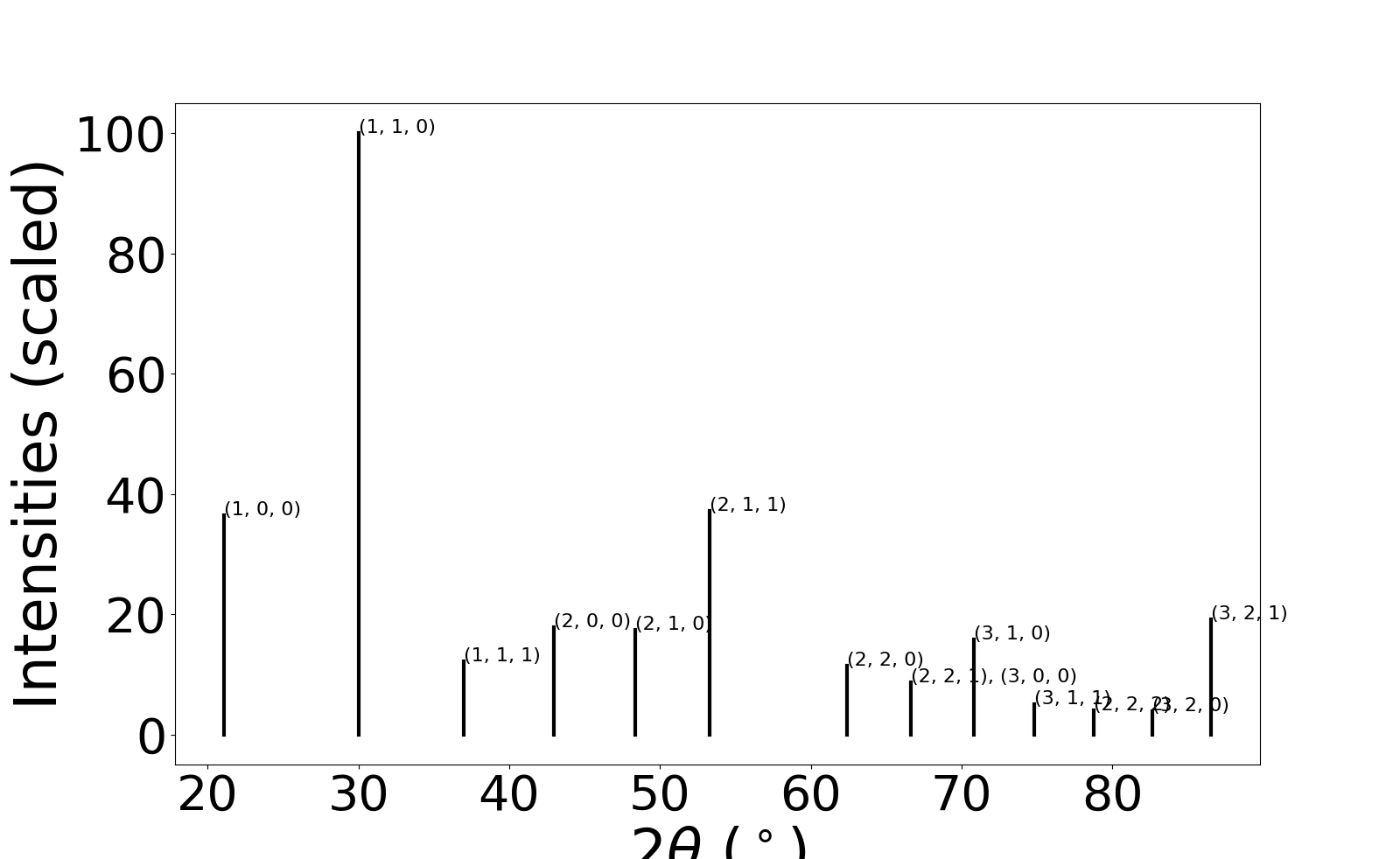
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2143/reactions",
"total_count": 1,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 1,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2143/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2143",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2143",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2143.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2143.patch",
"merged_at": "2021-05-14T15:12:31Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2144
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2144/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2144/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2144/events
|
https://github.com/materialsproject/pymatgen/pull/2144
| 893,646,960
|
MDExOlB1bGxSZXF1ZXN0NjQ2MTI3ODky
| 2,144
|
Bandstructure path name changes and Kpoint from_dict
|
{
"login": "munrojm",
"id": 6877334,
"node_id": "MDQ6VXNlcjY4NzczMzQ=",
"avatar_url": "https://avatars.githubusercontent.com/u/6877334?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/munrojm",
"html_url": "https://github.com/munrojm",
"followers_url": "https://api.github.com/users/munrojm/followers",
"following_url": "https://api.github.com/users/munrojm/following{/other_user}",
"gists_url": "https://api.github.com/users/munrojm/gists{/gist_id}",
"starred_url": "https://api.github.com/users/munrojm/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/munrojm/subscriptions",
"organizations_url": "https://api.github.com/users/munrojm/orgs",
"repos_url": "https://api.github.com/users/munrojm/repos",
"events_url": "https://api.github.com/users/munrojm/events{/privacy}",
"received_events_url": "https://api.github.com/users/munrojm/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39793806)\n\nCoverage decreased (-0.6%) to 82.996% when pulling **85583920bae3e045ca2ff593f3d21acb4621b72e on munrojm:bandstructure_fixes** into **37d8c1e4a94e79948bf2104d9b9caf61c05a603c on materialsproject:master**.\n",
"Thanks."
] | 2021-05-17T19:34:27
| 2021-05-21T17:12:21
|
2021-05-21T17:12:18Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Path type names in `HighSymmKpath` changed from short form ('sc', 'hin', and 'lm') to long form ('setyawan_curtarolo', 'hinuma', 'latimer_munro').
* Proper `from_dict` method added to `Kpoint` class in `from pymatgen.electronic_structure.bandstructure`.
## Checklist
- [X] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [X] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [X] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2144/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2144/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2144",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2144",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2144.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2144.patch",
"merged_at": "2021-05-21T17:12:18Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2145
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2145/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2145/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2145/events
|
https://github.com/materialsproject/pymatgen/issues/2145
| 893,717,264
|
MDU6SXNzdWU4OTM3MTcyNjQ=
| 2,145
|
This looks like dead code since the class is redefined immediately after
|
{
"login": "jmmshn",
"id": 14003693,
"node_id": "MDQ6VXNlcjE0MDAzNjkz",
"avatar_url": "https://avatars.githubusercontent.com/u/14003693?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jmmshn",
"html_url": "https://github.com/jmmshn",
"followers_url": "https://api.github.com/users/jmmshn/followers",
"following_url": "https://api.github.com/users/jmmshn/following{/other_user}",
"gists_url": "https://api.github.com/users/jmmshn/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jmmshn/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jmmshn/subscriptions",
"organizations_url": "https://api.github.com/users/jmmshn/orgs",
"repos_url": "https://api.github.com/users/jmmshn/repos",
"events_url": "https://api.github.com/users/jmmshn/events{/privacy}",
"received_events_url": "https://api.github.com/users/jmmshn/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"nvm `Dos` not `DOS`",
"I think it is strange to have both though... I'm not sure that a work-in-progress replacement like this should be in the main branch since it causes confusion when a user goes to import the class. Maybe it's better to rename it `_DOS` or remove it until it's ready to be used?",
"Yes this can be removed. "
] | 2021-05-17T21:10:45
| 2021-05-23T23:00:54
|
2021-05-17T21:11:07Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
https://github.com/materialsproject/pymatgen/blob/ad5cb53f6255f8af8428bbc6fdff74c50be342e8/pymatgen/electronic_structure/dos.py#L26
|
{
"login": "jmmshn",
"id": 14003693,
"node_id": "MDQ6VXNlcjE0MDAzNjkz",
"avatar_url": "https://avatars.githubusercontent.com/u/14003693?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jmmshn",
"html_url": "https://github.com/jmmshn",
"followers_url": "https://api.github.com/users/jmmshn/followers",
"following_url": "https://api.github.com/users/jmmshn/following{/other_user}",
"gists_url": "https://api.github.com/users/jmmshn/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jmmshn/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jmmshn/subscriptions",
"organizations_url": "https://api.github.com/users/jmmshn/orgs",
"repos_url": "https://api.github.com/users/jmmshn/repos",
"events_url": "https://api.github.com/users/jmmshn/events{/privacy}",
"received_events_url": "https://api.github.com/users/jmmshn/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2145/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2145/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2146
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2146/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2146/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2146/events
|
https://github.com/materialsproject/pymatgen/issues/2146
| 893,803,258
|
MDU6SXNzdWU4OTM4MDMyNTg=
| 2,146
|
from_dict in PourbaixEntry fails to read pbx entries with structure information
|
{
"login": "zbwang",
"id": 8037294,
"node_id": "MDQ6VXNlcjgwMzcyOTQ=",
"avatar_url": "https://avatars.githubusercontent.com/u/8037294?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/zbwang",
"html_url": "https://github.com/zbwang",
"followers_url": "https://api.github.com/users/zbwang/followers",
"following_url": "https://api.github.com/users/zbwang/following{/other_user}",
"gists_url": "https://api.github.com/users/zbwang/gists{/gist_id}",
"starred_url": "https://api.github.com/users/zbwang/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/zbwang/subscriptions",
"organizations_url": "https://api.github.com/users/zbwang/orgs",
"repos_url": "https://api.github.com/users/zbwang/repos",
"events_url": "https://api.github.com/users/zbwang/events{/privacy}",
"received_events_url": "https://api.github.com/users/zbwang/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-05-17T23:50:01
| 2021-05-21T19:23:32
|
2021-05-21T19:23:32Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
The from_dict method in PourbaixEntry fails to read a computed entry with structure information
**Expected behavior**
When reloading saved (dumpfn) pbx_entries using loadfn, it is expected pbx_entries still contain the structure information.
|
{
"login": "zbwang",
"id": 8037294,
"node_id": "MDQ6VXNlcjgwMzcyOTQ=",
"avatar_url": "https://avatars.githubusercontent.com/u/8037294?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/zbwang",
"html_url": "https://github.com/zbwang",
"followers_url": "https://api.github.com/users/zbwang/followers",
"following_url": "https://api.github.com/users/zbwang/following{/other_user}",
"gists_url": "https://api.github.com/users/zbwang/gists{/gist_id}",
"starred_url": "https://api.github.com/users/zbwang/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/zbwang/subscriptions",
"organizations_url": "https://api.github.com/users/zbwang/orgs",
"repos_url": "https://api.github.com/users/zbwang/repos",
"events_url": "https://api.github.com/users/zbwang/events{/privacy}",
"received_events_url": "https://api.github.com/users/zbwang/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2146/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2146/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2147
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2147/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2147/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2147/events
|
https://github.com/materialsproject/pymatgen/pull/2147
| 895,886,071
|
MDExOlB1bGxSZXF1ZXN0NjQ4MDQ2ODYy
| 2,147
|
add basis_not_supported error to QCOutput
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-05-19T20:59:01
| 2021-07-15T14:33:10
|
2021-05-21T17:11:42Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Adds detection of basis set not supported error to `QCOutput`. This error will cause the calculation to fail before it even starts, so it is treated similarly to the existing `input_file_error`.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2147/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2147/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2147",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2147",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2147.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2147.patch",
"merged_at": "2021-05-21T17:11:42Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2148
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2148/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2148/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2148/events
|
https://github.com/materialsproject/pymatgen/pull/2148
| 896,923,257
|
MDExOlB1bGxSZXF1ZXN0NjQ4OTc3OTU3
| 2,148
|
add test and fix for computed entries
|
{
"login": "JosephMontoya-TRI",
"id": 45643466,
"node_id": "MDQ6VXNlcjQ1NjQzNDY2",
"avatar_url": "https://avatars.githubusercontent.com/u/45643466?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/JosephMontoya-TRI",
"html_url": "https://github.com/JosephMontoya-TRI",
"followers_url": "https://api.github.com/users/JosephMontoya-TRI/followers",
"following_url": "https://api.github.com/users/JosephMontoya-TRI/following{/other_user}",
"gists_url": "https://api.github.com/users/JosephMontoya-TRI/gists{/gist_id}",
"starred_url": "https://api.github.com/users/JosephMontoya-TRI/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/JosephMontoya-TRI/subscriptions",
"organizations_url": "https://api.github.com/users/JosephMontoya-TRI/orgs",
"repos_url": "https://api.github.com/users/JosephMontoya-TRI/repos",
"events_url": "https://api.github.com/users/JosephMontoya-TRI/events{/privacy}",
"received_events_url": "https://api.github.com/users/JosephMontoya-TRI/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks"
] | 2021-05-20T14:43:14
| 2021-05-21T17:11:05
|
2021-05-21T17:11:00Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Updates PourbaixEntry.from_dict to invoke entries other than IonEntries using MontyDecoder, rather than hard-coding PDEntry.
* I initially had a bigger refactor which changed the ways Ions/IonEntries are invoked, but I'm a little worried about backwards compatibility of a core refactor, so I'm issuing this one for now.
* Addresses #2146
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [ ] All linting and tests pass.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2148/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2148/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2148",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2148",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2148.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2148.patch",
"merged_at": "2021-05-21T17:11:00Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2149
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2149/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2149/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2149/events
|
https://github.com/materialsproject/pymatgen/pull/2149
| 897,508,923
|
MDExOlB1bGxSZXF1ZXN0NjQ5NTEwNzc4
| 2,149
|
Overhaul interfaces
|
{
"login": "shyamd",
"id": 16827130,
"node_id": "MDQ6VXNlcjE2ODI3MTMw",
"avatar_url": "https://avatars.githubusercontent.com/u/16827130?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyamd",
"html_url": "https://github.com/shyamd",
"followers_url": "https://api.github.com/users/shyamd/followers",
"following_url": "https://api.github.com/users/shyamd/following{/other_user}",
"gists_url": "https://api.github.com/users/shyamd/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyamd/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyamd/subscriptions",
"organizations_url": "https://api.github.com/users/shyamd/orgs",
"repos_url": "https://api.github.com/users/shyamd/repos",
"events_url": "https://api.github.com/users/shyamd/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyamd/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39901075)\n\nCoverage decreased (-0.6%) to 83.075% when pulling **0aa785d900801e9caa9cc782adfcec729e62eb98 on shyamd:interfaces** into **37d8c1e4a94e79948bf2104d9b9caf61c05a603c on materialsproject:master**.\n",
"Nice PR!"
] | 2021-05-20T22:15:47
| 2021-05-23T22:40:52
|
2021-05-22T06:29:01Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
This PR overhauls all the interface code including the ZSL Generator and the substrate matcher:
- Creates a new `Interface` data class in core
- Moves interface algorithm to `analysis.interfaces`; imports and warnings in the old modules ensure they should still work
- Cleans up `ZSLGenerator` and `SubstrateAnalyzer` by implementing functional dataclasses for the matches making it easier to understand how to extra important quantities like the strain and transformation matrix
- Implements a `CoherentInterfaceBuilder` which builds `Interface` objects using the lattice matching of the Zur and McGill algorithm
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2149/reactions",
"total_count": 1,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 1,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2149/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2149",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2149",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2149.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2149.patch",
"merged_at": "2021-05-22T06:29:01Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2150
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2150/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2150/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2150/events
|
https://github.com/materialsproject/pymatgen/pull/2150
| 897,644,914
|
MDExOlB1bGxSZXF1ZXN0NjQ5NjI1NTkw
| 2,150
|
Python 3.6+ behavior of Exception.message
|
{
"login": "kmu",
"id": 3815357,
"node_id": "MDQ6VXNlcjM4MTUzNTc=",
"avatar_url": "https://avatars.githubusercontent.com/u/3815357?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/kmu",
"html_url": "https://github.com/kmu",
"followers_url": "https://api.github.com/users/kmu/followers",
"following_url": "https://api.github.com/users/kmu/following{/other_user}",
"gists_url": "https://api.github.com/users/kmu/gists{/gist_id}",
"starred_url": "https://api.github.com/users/kmu/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/kmu/subscriptions",
"organizations_url": "https://api.github.com/users/kmu/orgs",
"repos_url": "https://api.github.com/users/kmu/repos",
"events_url": "https://api.github.com/users/kmu/events{/privacy}",
"received_events_url": "https://api.github.com/users/kmu/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/39878391)\n\nCoverage decreased (-0.7%) to 82.984% when pulling **83537c312cd4d0dd65beb8564cdc4937ea3d3fd4 on kmu:MPResterError-message** into **37d8c1e4a94e79948bf2104d9b9caf61c05a603c on materialsproject:master**.\n",
"Thanks."
] | 2021-05-21T03:33:39
| 2021-05-21T16:56:49
|
2021-05-21T16:56:43Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Hey guys,
Python 3.6+ seems to abolish `Exception.message`.
This PR addresses this problem.
It does not seem to be easy to write a unittest to reproduce this without imposing a large load on MP.
Here is a working example.
- Current implementation
```python
import re
from pymatgen.ext.matproj import MPRestError
try:
raise MPRestError("error status code 000")
except MPRestError as e:
# pylint: disable=E1101
match = re.search(r"error status code (\d+)", e.message)
if match:
print(e)
```
```
MPRestError Traceback (most recent call last)
<ipython-input-3-83f89fe43bab> in <module>
2 try:
----> 3 raise MPRestError("error status code 000")
4 except MPRestError as e:
MPRestError:
During handling of the above exception, another exception occurred:
AttributeError Traceback (most recent call last)
<ipython-input-3-83f89fe43bab> in <module>
4 except MPRestError as e:
5 # pylint: disable=E1101
----> 6 match = re.search(r"error status code (\d+)", e.message)
7 if match:
8 print(e)
AttributeError: 'MPRestError' object has no attribute 'message'
```
- This PR
```python
import re
from pymatgen.ext.matproj import MPRestError
try:
raise MPRestError("error status code 000")
except MPRestError as e:
# pylint: disable=E1101
match = re.search(r"error status code (\d+)", str(e))
if match:
print(e)
```
```
error status code 000
```
## Additional dependencies introduced (if any)
None
## TODO (if any)
None
## Checklist
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2150/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2150/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2150",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2150",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2150.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2150.patch",
"merged_at": "2021-05-21T16:56:43Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2151
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2151/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2151/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2151/events
|
https://github.com/materialsproject/pymatgen/issues/2151
| 897,758,428
|
MDU6SXNzdWU4OTc3NTg0Mjg=
| 2,151
|
how to handle properties of tritium and deuterium?
|
{
"login": "kjappelbaum",
"id": 32935233,
"node_id": "MDQ6VXNlcjMyOTM1MjMz",
"avatar_url": "https://avatars.githubusercontent.com/u/32935233?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/kjappelbaum",
"html_url": "https://github.com/kjappelbaum",
"followers_url": "https://api.github.com/users/kjappelbaum/followers",
"following_url": "https://api.github.com/users/kjappelbaum/following{/other_user}",
"gists_url": "https://api.github.com/users/kjappelbaum/gists{/gist_id}",
"starred_url": "https://api.github.com/users/kjappelbaum/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/kjappelbaum/subscriptions",
"organizations_url": "https://api.github.com/users/kjappelbaum/orgs",
"repos_url": "https://api.github.com/users/kjappelbaum/repos",
"events_url": "https://api.github.com/users/kjappelbaum/events{/privacy}",
"received_events_url": "https://api.github.com/users/kjappelbaum/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I would actually go for option 1. I don't think it is that difficult to make it such that isotopes can be handled. The main problem, though, is that I don't have a source of compilation for all the other properties that most elements have (e.g., melting point, etc.) for isotopes.\r\n\r\nIf it is just D and T, I think it is a simple matter to just add those as additional element symbolsm, like what ASE does.\r\n\r\nAlternatively, we can just translate D/T to H unless there is a very good reason why we want D or T to be present. \r\n\r\nI don't think most of the analysis we do differentiates between D or T or H (except maybe diffusion where the mass differences can be large). For most other isotopes, the properties are similar enough. ",
"I think this is a great question.\r\n\r\nI agree that symbols `D` and `T` should be handled automatically, which would involve coding a special case somewhere.\r\n\r\nIf we did want to add isotope support (one might imagine this useful for other elements too, e.g. in the diffusion case mentioned, I've seen MD runs where specific isotopes are used for example), then one option is to add it as a supported property on `Species`. Currently, `Species` only formally supports `spin` and `oxidation_state`, but we could add `mass_number`. The string representation of this might be `Fe,A=56` for example.\r\n\r\nI agree that adding D, T to periodic_table.json is going to be the easiest/lowest effort route.",
"Actually, the advantage of the `Species` approach is that these cases, e.g. D, T, would still be handled properly as H by, for example, VASP I/O without any additional custom logic.",
"This may be worth re-opening in the context of MP's current ingestion of new (since 2019) ICSD structures, about 10k of which have undetermined light atom positions. Mostly hydrogen but some deuterium as well."
] | 2021-05-21T07:11:24
| 2023-11-02T22:37:44
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Is your feature request related to a problem? Please describe.**
In `cif`s (e.g., from the Cambridge Crystallographic Database) one sometimes finds symbols for common isotopes such as Tritium (`T`) or Deuterium (`D`). `pymatgen` parses them without issues, but it complains when one likes to access some properties like the atomic mass.
**Describe alternatives you've considered**
- A "quick and dirty" solution I tried is to just add `D` to `periodic_table.json` with the `Atomic mass` and some other physical properties. It is somehow ugly because adding the `Atomic no` will lead to some issues when one wants to create an `Element.from_z` because then there is ambiguity. For this reason, one cannot simply add the `Atomic no` which might lead to unexpected issues, when users want to use this property.
- ASE simply replaces `Atom` with the symbol `D`. with an atom with symbol `H` and mass 2. But this behavior can sometimes be quite unexpected I feel.
- Explicitly handle isotopes, https://github.com/pkienzle/periodictable does this. This seems like a major change.
- One could also simply raise an error if there is a symbol `pymatgen` does not know. But this is not really pythonic and there are many functionalities for which one does not need to know any physical properties.
**Additional context**
I would be happy to contribute a PR to fix this but I'd like to get some feedback on the design you would use before I go for it.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2151/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2151/timeline
| null |
reopened
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2152
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2152/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2152/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2152/events
|
https://github.com/materialsproject/pymatgen/pull/2152
| 899,733,183
|
MDExOlB1bGxSZXF1ZXN0NjUxMzkzMTE5
| 2,152
|
Ignore typical virtual environment file/directory names
|
{
"login": "flaviu-gostin",
"id": 37553536,
"node_id": "MDQ6VXNlcjM3NTUzNTM2",
"avatar_url": "https://avatars.githubusercontent.com/u/37553536?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/flaviu-gostin",
"html_url": "https://github.com/flaviu-gostin",
"followers_url": "https://api.github.com/users/flaviu-gostin/followers",
"following_url": "https://api.github.com/users/flaviu-gostin/following{/other_user}",
"gists_url": "https://api.github.com/users/flaviu-gostin/gists{/gist_id}",
"starred_url": "https://api.github.com/users/flaviu-gostin/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/flaviu-gostin/subscriptions",
"organizations_url": "https://api.github.com/users/flaviu-gostin/orgs",
"repos_url": "https://api.github.com/users/flaviu-gostin/repos",
"events_url": "https://api.github.com/users/flaviu-gostin/events{/privacy}",
"received_events_url": "https://api.github.com/users/flaviu-gostin/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-05-24T14:50:31
| 2021-05-24T14:56:48
|
2021-05-24T14:56:48Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Taken from https://github.com/github/gitignore
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2152/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2152/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2152",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2152",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2152.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2152.patch",
"merged_at": "2021-05-24T14:56:48Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2153
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2153/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2153/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2153/events
|
https://github.com/materialsproject/pymatgen/issues/2153
| 899,951,145
|
MDU6SXNzdWU4OTk5NTExNDU=
| 2,153
|
Question on TEM diffraction calculator
|
{
"login": "YuLin999",
"id": 44076136,
"node_id": "MDQ6VXNlcjQ0MDc2MTM2",
"avatar_url": "https://avatars.githubusercontent.com/u/44076136?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/YuLin999",
"html_url": "https://github.com/YuLin999",
"followers_url": "https://api.github.com/users/YuLin999/followers",
"following_url": "https://api.github.com/users/YuLin999/following{/other_user}",
"gists_url": "https://api.github.com/users/YuLin999/gists{/gist_id}",
"starred_url": "https://api.github.com/users/YuLin999/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/YuLin999/subscriptions",
"organizations_url": "https://api.github.com/users/YuLin999/orgs",
"repos_url": "https://api.github.com/users/YuLin999/repos",
"events_url": "https://api.github.com/users/YuLin999/events{/privacy}",
"received_events_url": "https://api.github.com/users/YuLin999/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"The issue is that `mpr.get_structure_by_material_id` will download the primitive cell structure by default. This means that the [1, 0, 0] direction in your structure does not correspond to the [1, 0, 0] of the conventional cell but instead the [1, 1, 1] of the conventional cell.\r\n\r\nAsking for the conventional cell gives the correct 4-fold symmetry.\r\n\r\n```python\r\nfrom pymatgen.ext.matproj import MPRester\r\nfrom pymatgen.analysis.diffraction.tem import TEMCalculator\r\n\r\nstructure = mpr.get_structure_by_material_id('mp-13', conventional_unit_cell=True)\r\ncalculator = TEMCalculator(beam_direction=[1,0,0])\r\nplt = calculator.get_plot_2d(structure=structure)\r\nplt.show()\r\n```\r\n\r\n<img width=\"556\" alt=\"Screenshot 2021-05-24 at 13 53 37\" src=\"https://user-images.githubusercontent.com/1330638/119406543-d1bf7080-bc97-11eb-9383-3bdc1b6d1348.png\">\r\n",
"@utf [100] in primitive cell is not [111] in the conventional either. In cubic system, the [111] remains [111] in both conventional and primitive.",
"Actually wait.. for bcc, the primitive [100] is the [111]. You are right. It is for fcc that the [111] remains the same.",
"Okay I see. Thanks a lot for the quick response!!"
] | 2021-05-24T19:42:14
| 2021-05-24T22:01:32
|
2021-05-24T21:02:26Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I am trying to generate some simulation diffraction pattern data using and got some patterns inconsistent with the real patterns. Wondering if I was doing this correctly. Here is the codes that I used (and the generated pattern is attached). mp-13 is Fe with cubic bcc structure so I am expecting the simulated pattern will has 4-fold symmetry.
structure = mpr.get_structure_by_material_id('mp-13')
calculator = tem.TEMCalculator(beam_direction=[1,0,0])
fig = calculator.get_plot_2d(structure=structure)
fig.show()

|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2153/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2153/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2154
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2154/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2154/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2154/events
|
https://github.com/materialsproject/pymatgen/issues/2154
| 900,154,093
|
MDU6SXNzdWU5MDAxNTQwOTM=
| 2,154
|
Surface slab visualization KeyError with oxidation states
|
{
"login": "peterschindler",
"id": 45672907,
"node_id": "MDQ6VXNlcjQ1NjcyOTA3",
"avatar_url": "https://avatars.githubusercontent.com/u/45672907?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/peterschindler",
"html_url": "https://github.com/peterschindler",
"followers_url": "https://api.github.com/users/peterschindler/followers",
"following_url": "https://api.github.com/users/peterschindler/following{/other_user}",
"gists_url": "https://api.github.com/users/peterschindler/gists{/gist_id}",
"starred_url": "https://api.github.com/users/peterschindler/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/peterschindler/subscriptions",
"organizations_url": "https://api.github.com/users/peterschindler/orgs",
"repos_url": "https://api.github.com/users/peterschindler/repos",
"events_url": "https://api.github.com/users/peterschindler/events{/privacy}",
"received_events_url": "https://api.github.com/users/peterschindler/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I tried ```color = color_dict[sites[n].element.symbol]```\r\n\r\nbut I'm getting: ```[AttributeError: element]()```",
"Mhmm, I see. How about \r\n`color = color_dict[sites[n].species.elements[0].symbol]`\r\nor\r\n`color = color_dict[sites[n].species.remove_charges().elements[0].symbol]` ?",
"Thanks. I just pushed a fix.",
"Thanks @peterschindler. With both of them I’m getting errors below. I also retried by removing oxidations states. It seem that something is wrong with the radius. \r\n\r\n\r\n```[---> ]()[23]()[ plot_slab(slab, ax, adsorption_sites=False)\r\n ]()[24]()[ ax.set_title(\"Si (1, 1, 1) surface %i\" %(n+1))\r\n ]()[25]()[ ax.set_xticks([])\r\n\r\nFile ~/xraylarch/envs/my_pymatgen/lib/python3.10/site-packages/pymatgen/analysis/adsorption.py:710, in plot_slab(slab, ax, scale, repeat, window, draw_unit_cell, decay, adsorption_sites, inverse)\r\n ]()[708]()[ # Draw circles at sites and stack them accordingly\r\n ]()[709]()[ for n, coord in enumerate(coords):\r\n--> ]()[710]()[ r = sites[n].specie.atomic_radius * scale\r\n ]()[711]()[ ax.add_patch(patches.Circle(coord[:2] - lattsum * (repeat // 2), r, color=\"w\", zorder=2 * n))\r\n ]()[712]()[ color = color_dict[sites[n].species.elements[0].symbol]\r\n\r\nFile ~/xraylarch/envs/my_pymatgen/lib/python3.10/site-packages/pymatgen/core/sites.py:78, in Site.__getattr__(self, a)\r\n ]()[76]()[ if a in p:\r\n ]()[77]()[ return p[a]\r\n---> ]()[78]()[ raise AttributeError(a)\r\n\r\nAttributeError: specie]() ```\r\n\r\n****\r\n\r\n```[21]()[ for n, slab in enumerate(slabs):\r\n ]()[22]()[ ax = fig.add_subplot(1, len(slabs), n+1)\r\n---> ]()[23]()[ plot_slab(slab, ax, adsorption_sites=False)\r\n ]()[24]()[ ax.set_title(\"Si (1, 1, 1) surface %i\" %(n+1))\r\n ]()[25]()[ ax.set_xticks([])\r\n\r\nFile ~/xraylarch/envs/my_pymatgen/lib/python3.10/site-packages/pymatgen/analysis/adsorption.py:710, in plot_slab(slab, ax, scale, repeat, window, draw_unit_cell, decay, adsorption_sites, inverse)\r\n ]()[708]()[ # Draw circles at sites and stack them accordingly\r\n ]()[709]()[ for n, coord in enumerate(coords):\r\n--> ]()[710]()[ r = sites[n].specie.atomic_radius * scale\r\n ]()[711]()[ ax.add_patch(patches.Circle(coord[:2] - lattsum * (repeat // 2), r, color=\"w\", zorder=2 * n))\r\n ]()[712]()[ color = color_dict[sites[n].species.remove_charges().elements[0].symbol]\r\n\r\nFile ~/xraylarch/envs/my_pymatgen/lib/python3.10/site-packages/pymatgen/core/sites.py:78, in Site.__getattr__(self, a)\r\n ]()[76]()[ if a in p:\r\n ]()[77]()[ return p[a]\r\n---> ]()[78]()[ raise AttributeError(a)\r\n\r\nAttributeError: specie]()```",
"That's a different error from what I originally referred to but actually related. The same solution I mentioned above should work here as well:\r\n`r = sites[n].species.elements[0].atomic_radius * scale`\r\nSeems like `site.species_string` is also used in line 607 which should potentially also be replaced?",
"@peterschindler Thanks Peter. In ```adsorption.py```, This fixed it:\r\n\r\n``` # Draw circles at sites and stack them accordingly\r\n for n, coord in enumerate(coords):\r\n r = sites[n].species.elements[0].atomic_radius * scale\r\n ax.add_patch(patches.Circle(coord[:2] - lattsum * (repeat // 2), r, color=\"w\", zorder=2 * n))\r\n color = color_dict[sites[n].species.remove_charges().elements[0].symbol]\r\n ax.add_patch(\r\n patches.Circle(\r\n coord[:2] - lattsum * (repeat // 2),\r\n r,\r\n facecolor=color,\r\n alpha=alphas[n],\r\n edgecolor=\"k\",\r\n lw=0.3,\r\n zorder=2 * n + 1,```",
"Great, glad it worked @Ardalanhayatifar!\r\nPlease, could you push the change in line 710 as well, @shyuep?",
"I fixed line 710. But I think line 607 should remain since it is based on target species. In general, I have always counseled that all code should avoid assuming an ordered structure or a non-oxidation state structure. Quite clearly it is violated in this code."
] | 2021-05-25T01:14:51
| 2022-04-26T02:48:46
|
2022-04-19T14:20:06Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
There seems to be a bug in displaying surface slabs when sites are decorated with oxidation states. Mentioned [here](https://matsci.org/t/is-it-a-bug-or-just-something-i-need-to-do-when-use-plot-slab-function/36521) and [here](https://matsci.org/t/surface-slab-visualization-key-error-o2/34557). I didn't have the time to test this but I think replacing line 716 in analysis/adsorption.py with
`color = color_dict[sites[n].element.symbol]`
should fix the issue, I believe.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2154/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2154/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2155
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2155/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2155/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2155/events
|
https://github.com/materialsproject/pymatgen/issues/2155
| 903,389,596
|
MDU6SXNzdWU5MDMzODk1OTY=
| 2,155
|
unexpected result of get_conventional_standard_structure() to mp-644693
|
{
"login": "lwk8891",
"id": 81547431,
"node_id": "MDQ6VXNlcjgxNTQ3NDMx",
"avatar_url": "https://avatars.githubusercontent.com/u/81547431?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/lwk8891",
"html_url": "https://github.com/lwk8891",
"followers_url": "https://api.github.com/users/lwk8891/followers",
"following_url": "https://api.github.com/users/lwk8891/following{/other_user}",
"gists_url": "https://api.github.com/users/lwk8891/gists{/gist_id}",
"starred_url": "https://api.github.com/users/lwk8891/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/lwk8891/subscriptions",
"organizations_url": "https://api.github.com/users/lwk8891/orgs",
"repos_url": "https://api.github.com/users/lwk8891/repos",
"events_url": "https://api.github.com/users/lwk8891/events{/privacy}",
"received_events_url": "https://api.github.com/users/lwk8891/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @lwk8891, this is almost certainly a serious bug. I will investigate.\r\n\r\nFor now, I suggest you use `.get_refined_structure()` which might work for your purposes.",
"Has this issue been resolved? I need to transform to the conventional setting and want to ensure I'm getting the correct results. Currently, I'm getting the same unit cell parameters using both ```.get_conventional_standard_structure()``` and ```.get_refined_structure()```.\r\n\r\nExample:\r\n```\r\nentry = \"mp-28243\"\r\nwith MPRester(api key) as mpr:\r\n structure = mpr.get_structure_by_material_id(entry)\r\n sga = `SpacegroupAnalyzer(structure)` \r\n print('Prim',structure.lattice.abc, structure.lattice.angles)\r\n conventional_structure = sga.get_conventional_standard_structure()\r\n print('Conventional', conventional_structure.lattice.abc, conventional_structure.lattice.angles)\r\n refined_structure = sga.get_refined_structure()\r\n print('Refined', refined_structure.lattice.abc, refined_structure.lattice.angles)\r\n```\r\n\r\nOutput:\r\nPrim (4.121743287390061, 7.1843702, 7.499651169364684) (90.0, 105.94984245397592, 90.0)\r\nConventional (4.121743287390061, 14.421868563834614, 7.1843702) (90.0, 90.0, 90.0)\r\nRefined (4.121743287390061, 14.421868563834614, 7.1843702) (90.0, 90.0, 90.0)\r\n\r\nThese parameters do not match the associated entry in the ICSD (https://www.ccdc.cam.ac.uk/structures/Search?Ccdcid=36216&DatabaseToSearch=ICSD) even though ```.get_refined_structure()``` is said to have the same parameters as the ICSD. ",
"Hi!\r\n\r\nI quickly checked the original issue and could not reproduce it.\r\nI get the same structure for mp-644693 after using the get_conventional_standard_structure(). There is just a swap of the a and b axes, \r\nbut the structure is the same. This swap is due to the \"convention\" used in the definition\r\nof the cell as reported in the doc of the function:\r\n```\r\nGives a structure with a conventional cell according to certain\r\nstandards. The standards are defined in Setyawan, W., & Curtarolo,\r\nS. (2010). High-throughput electronic band structure calculations:\r\nChallenges and tools. Computational Materials Science,\r\n49(2), 299-312. doi:10.1016/j.commatsci.2010.05.010\r\nThey basically enforce as much as possible\r\nnorm(a1)<norm(a2)<norm(a3). NB This is not necessarily the same as the\r\nstandard settings within the International Tables of Crystallography,\r\nfor which get_refined_structure should be used instead.\r\n```\r\n\r\n\r\nIn your case, there is a similar swap of the a and b axes, but I'm guessing the structure\r\nis the same. The small difference in the decimal digits could come from the fact that\r\nthe ICSD is the exp. structure while the stucture from MP is relaxed.\r\n\r\nHope this helps.",
"Hi! Thanks for the help! What is the difference between ```get_conventional_standard_structure()``` and ```get_refined_structure()``` then? In the documentation, it says that ```get_refined_structure()``` should be used to get the standard settings used within the International Tables of Crystallography, but it seems to return the same results as ```get_conventional_standard_structure()``` in the example above. ",
"well, `get_conventional_standard_structure()` follows the convention in Setyawan, W., & Curtarolo,\r\nS. (2010); whereas the `get_refined_structure()` follows the International Tables of Crystallography ;-)\r\nI'm not sure about which convention is used in ICSD, it might be neither of the above.",
"@janosh: Looks like this can be closed based on @fraricci's analysis."
] | 2021-05-27T08:18:34
| 2023-09-03T19:43:23
|
2023-09-03T19:43:23Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
The get_conventional_standard_structure() method of pymatgen.symmetry.analyzer.SpacegroupAnalyzer class changes the atomic connection of mp-644693.
**To Reproduce**
1. get the structure of mp-644693 by calling s=get_structure_by_materials_id('mp-644693') of pymatgen.ext.matproj.MPRestrer class
2. Create the SpacegroupAnalyzer class for structure s by anal=SpacegroupAnalyzer(s)
3. Get the conventional standard structure by conv_struct=anal.get_conventional_standard_structure()
4. Visualize the structures s and conv_struct by making POSCAR format of VASP, then the atomic connections are different
**Expected behavior**
The atomic connection is maintained after getting conventional standard structure.
**Screenshots**

**Additional context**
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2155/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2155/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2156
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2156/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2156/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2156/events
|
https://github.com/materialsproject/pymatgen/pull/2156
| 906,452,691
|
MDExOlB1bGxSZXF1ZXN0NjU3NDUzODU3
| 2,156
|
Update usage.rst
|
{
"login": "755452800",
"id": 18474386,
"node_id": "MDQ6VXNlcjE4NDc0Mzg2",
"avatar_url": "https://avatars.githubusercontent.com/u/18474386?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/755452800",
"html_url": "https://github.com/755452800",
"followers_url": "https://api.github.com/users/755452800/followers",
"following_url": "https://api.github.com/users/755452800/following{/other_user}",
"gists_url": "https://api.github.com/users/755452800/gists{/gist_id}",
"starred_url": "https://api.github.com/users/755452800/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/755452800/subscriptions",
"organizations_url": "https://api.github.com/users/755452800/orgs",
"repos_url": "https://api.github.com/users/755452800/repos",
"events_url": "https://api.github.com/users/755452800/events{/privacy}",
"received_events_url": "https://api.github.com/users/755452800/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks.",
"\n[](https://coveralls.io/builds/40140538)\n\nCoverage decreased (-0.6%) to 83.078% when pulling **3183ae8fbb036c116c3c909c98e3d306c054b967 on 755452800:master** into **1d0471d5232f19dfb3eff2d988d0f82460d0a832 on materialsproject:master**.\n"
] | 2021-05-29T11:48:53
| 2021-05-29T12:12:03
|
2021-05-29T11:50:18Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
I think there is a typo in line 189:
`w = CifWriter(p.struct)`
should be:
`w = CifWriter(p.structure)`
or else error **AttributeError: 'Poscar' object has no attribute 'struct'** will occur.
Include a summary of major changes in bullet points:
* fix a typo in usage.rst line 189
## Additional dependencies introduced (if any)
* None
## TODO (if any)
* None
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All linting and tests pass.
Note that the CI system will run all the above checks. But it will be much more efficient if you already fix most
errors prior to submitting the PR. It is highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2156/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2156/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2156",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2156",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2156.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2156.patch",
"merged_at": "2021-05-29T11:50:18Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2157
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2157/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2157/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2157/events
|
https://github.com/materialsproject/pymatgen/pull/2157
| 906,554,635
|
MDExOlB1bGxSZXF1ZXN0NjU3NTMxOTcy
| 2,157
|
Support combining data with multiple mol-id
|
{
"login": "htz1992213",
"id": 25351437,
"node_id": "MDQ6VXNlcjI1MzUxNDM3",
"avatar_url": "https://avatars.githubusercontent.com/u/25351437?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/htz1992213",
"html_url": "https://github.com/htz1992213",
"followers_url": "https://api.github.com/users/htz1992213/followers",
"following_url": "https://api.github.com/users/htz1992213/following{/other_user}",
"gists_url": "https://api.github.com/users/htz1992213/gists{/gist_id}",
"starred_url": "https://api.github.com/users/htz1992213/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/htz1992213/subscriptions",
"organizations_url": "https://api.github.com/users/htz1992213/orgs",
"repos_url": "https://api.github.com/users/htz1992213/repos",
"events_url": "https://api.github.com/users/htz1992213/events{/privacy}",
"received_events_url": "https://api.github.com/users/htz1992213/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/40215177)\n\nCoverage decreased (-0.6%) to 83.078% when pulling **3a90f50895f128fabcdeb30687552f531decf612 on htz1992213:master** into **37da2710fc3e23c901f6e6a275adf4ee540cd1f3 on materialsproject:master**.\n",
"Thanks @htz1992213!"
] | 2021-05-29T20:36:53
| 2021-06-01T20:56:45
|
2021-06-01T20:56:45Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
* Feature 1: Support combining data files that have multiple molecules (mol id) in a single data object
* Feature 2: Added restriction to LAMMPS data name list for easier parsing.
* Fix 1: Changed maintainer
* Fix 2: Linting errors in other modules
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2157/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2157/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2157",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2157",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2157.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2157.patch",
"merged_at": "2021-06-01T20:56:45Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2158
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2158/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2158/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2158/events
|
https://github.com/materialsproject/pymatgen/issues/2158
| 909,518,842
|
MDU6SXNzdWU5MDk1MTg4NDI=
| 2,158
|
typo
|
{
"login": "ddopplereffekt",
"id": 77629108,
"node_id": "MDQ6VXNlcjc3NjI5MTA4",
"avatar_url": "https://avatars.githubusercontent.com/u/77629108?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ddopplereffekt",
"html_url": "https://github.com/ddopplereffekt",
"followers_url": "https://api.github.com/users/ddopplereffekt/followers",
"following_url": "https://api.github.com/users/ddopplereffekt/following{/other_user}",
"gists_url": "https://api.github.com/users/ddopplereffekt/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ddopplereffekt/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ddopplereffekt/subscriptions",
"organizations_url": "https://api.github.com/users/ddopplereffekt/orgs",
"repos_url": "https://api.github.com/users/ddopplereffekt/repos",
"events_url": "https://api.github.com/users/ddopplereffekt/events{/privacy}",
"received_events_url": "https://api.github.com/users/ddopplereffekt/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @ddopplereffekt, thanks for reporting, I've fixed this now. In future, you can also suggest edits to files directly by clicking the \"Edit\" button on GitHub for small typos like this, and then we can merge the change in with one-click."
] | 2021-06-02T13:43:49
| 2021-06-02T19:24:10
|
2021-06-02T19:24:10Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
in pymatgen/entries/compatebility.py line 1031 is a type: the enclosing bracket for the formular is missing
**Expected behavior**
f"({entry.composition.reduced_formula}). Assigning anion correction to "
instead of
f"({entry.composition.reduced_formula}. Assigning anion correction to "
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2158/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2158/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2159
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2159/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2159/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2159/events
|
https://github.com/materialsproject/pymatgen/issues/2159
| 909,691,268
|
MDU6SXNzdWU5MDk2OTEyNjg=
| 2,159
|
Support for ASE-style "extended XYZ" with Lattice parameters
|
{
"login": "ghutchis",
"id": 41128,
"node_id": "MDQ6VXNlcjQxMTI4",
"avatar_url": "https://avatars.githubusercontent.com/u/41128?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ghutchis",
"html_url": "https://github.com/ghutchis",
"followers_url": "https://api.github.com/users/ghutchis/followers",
"following_url": "https://api.github.com/users/ghutchis/following{/other_user}",
"gists_url": "https://api.github.com/users/ghutchis/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ghutchis/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ghutchis/subscriptions",
"organizations_url": "https://api.github.com/users/ghutchis/orgs",
"repos_url": "https://api.github.com/users/ghutchis/repos",
"events_url": "https://api.github.com/users/ghutchis/events{/privacy}",
"received_events_url": "https://api.github.com/users/ghutchis/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I think the would be a welcome inclusion, I know this format (or a similar one) is used in AtomEye too. Would you be interested in preparing a PR?",
"I can't promise I can support everything - since it seems \"Properties\" can include a lot. But the Lattice extension IMHO is easy and IMHO the big first piece.\r\n\r\nLet me get Avogadro 1.94 tagged today and I'll send a PR soon.",
"Thanks, I think the Lattice support is sufficient, as far as I know it's an informal spec anyway, we would definitely encourage people to prefer CIF for archival purposes."
] | 2021-06-02T16:42:50
| 2021-06-02T19:57:17
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
As far as I can tell, the current versions of `pymatgen` support the traditional non-lattice XYZ from Open Babel.
I just implemented initial support for "extended XYZ" in Avogadro2 - the first layer is pretty easy. The second line (for comments / title) now have the lattice vectors and optionally additional properties.
e.g. Lattice="H11 H21 H31 H12 H22 H32 H13 H23 H33"
https://atomsk.univ-lille.fr/doc/en/format_xyz.html
https://gitlab.com/ase/ase/-/merge_requests/62
It seems like a useful interchange format, and wondered if `pymatgen` would support it too, either transparently in the .xyz reader or in an "extxyz" format.
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2159/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2159/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2160
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2160/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2160/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2160/events
|
https://github.com/materialsproject/pymatgen/pull/2160
| 909,801,582
|
MDExOlB1bGxSZXF1ZXN0NjYwMzIxNzIx
| 2,160
|
fix another CombinedData bug
|
{
"login": "htz1992213",
"id": 25351437,
"node_id": "MDQ6VXNlcjI1MzUxNDM3",
"avatar_url": "https://avatars.githubusercontent.com/u/25351437?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/htz1992213",
"html_url": "https://github.com/htz1992213",
"followers_url": "https://api.github.com/users/htz1992213/followers",
"following_url": "https://api.github.com/users/htz1992213/following{/other_user}",
"gists_url": "https://api.github.com/users/htz1992213/gists{/gist_id}",
"starred_url": "https://api.github.com/users/htz1992213/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/htz1992213/subscriptions",
"organizations_url": "https://api.github.com/users/htz1992213/orgs",
"repos_url": "https://api.github.com/users/htz1992213/repos",
"events_url": "https://api.github.com/users/htz1992213/events{/privacy}",
"received_events_url": "https://api.github.com/users/htz1992213/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/40429423)\n\nCoverage decreased (-0.6%) to 83.082% when pulling **8a30eab02d056fc51b6af18475a0c0ddc3cfca6b on htz1992213:master** into **c724e7b679581783dcdb4690a83c374a57437ae6 on materialsproject:master**.\n",
"Thanks @htz1992213. Rather than edit the pylintrc with this specific case, I think you can add an in-line comment (something like `# ignore`).\r\n\r\nCan you comment on what the issue was with `ComputedStructureEntry._composition`? This is a protected member anyway, so shouldn't be access outside the class.",
"Thanks for the advice @mkhorton! The access comes from the following line, I am not familiar with these classes so I will change the code with the # ignore workaround.\r\n\r\nhttps://github.com/materialsproject/pymatgen/blob/463e4f1b577287c593e493b2b1138320418f0bc7/pymatgen/entries/computed_entries.py#L700",
"> \r\n> \r\n> Thanks @htz1992213. Rather than edit the pylintrc with this specific case, I think you can add an in-line comment (something like `# ignore`).\r\n> \r\n> Can you comment on what the issue was with `ComputedStructureEntry._composition`? This is a protected member anyway, so shouldn't be access outside the class.\r\n\r\n@mkhorton I will note that `ComputedStructureEntry._composition` seems to be causing a `1pylint` failure in another PR as well, in which the changes are completely unrelated. Does this test pass in master?\r\nhttps://github.com/materialsproject/pymatgen/pull/2135/checks?check_run_id=2754058779",
"Hey @mkhorton, could you please let me know if there are other things that I should address for the PR to be merged? Thanks!",
"This is all good, thank you Tingzheng!"
] | 2021-06-02T19:02:49
| 2021-06-09T20:07:31
|
2021-06-09T20:07:30Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
* Fix 1: The CombinedData constructor previously creates new ff dict only according to the keys in the first mol in the mols llist, which may cause the ff information in the remaining mols lost. The new implementation create the ff dict using a set of keys containing all ff keys in mols
* Fix 2: The CombinedData constructor previously traverses the topo and ff info of mol in mols list without checking None type, which may cause error. The new implementation checks for None before iteration, and set ff/topo of the CombinedData to be None if all mol in mols list has None ff/topo.
Tests have been created accordingly.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2160/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2160/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2160",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2160",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2160.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2160.patch",
"merged_at": "2021-06-09T20:07:30Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2161
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2161/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2161/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2161/events
|
https://github.com/materialsproject/pymatgen/issues/2161
| 910,202,173
|
MDU6SXNzdWU5MTAyMDIxNzM=
| 2,161
|
Cleave a (111) surface that alpha=beta=90 degree
|
{
"login": "Rasic2",
"id": 44538197,
"node_id": "MDQ6VXNlcjQ0NTM4MTk3",
"avatar_url": "https://avatars.githubusercontent.com/u/44538197?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Rasic2",
"html_url": "https://github.com/Rasic2",
"followers_url": "https://api.github.com/users/Rasic2/followers",
"following_url": "https://api.github.com/users/Rasic2/following{/other_user}",
"gists_url": "https://api.github.com/users/Rasic2/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Rasic2/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Rasic2/subscriptions",
"organizations_url": "https://api.github.com/users/Rasic2/orgs",
"repos_url": "https://api.github.com/users/Rasic2/repos",
"events_url": "https://api.github.com/users/Rasic2/events{/privacy}",
"received_events_url": "https://api.github.com/users/Rasic2/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"set the max_normal_search=1 can resolve the problem. "
] | 2021-06-03T07:32:38
| 2021-06-03T07:46:03
|
2021-06-03T07:43:31Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
When I try to cleave a CeO2(111) surface, I found that the lattice matrix is not similar with the Material Studio produced, that is the alpha=85 degree not the 90 degree,How can I get the same result with the Material studio.
**To Reproduce**
The following code I used in this issue:
#!/usr/bin/env python
from pymatgen.core import Structure, Lattice
from pymatgen.core.surface import generate_all_slabs
CeO2=Structure.from_spacegroup("Fm-3m",Lattice.cubic(5.450),["Ce","O"],[[0,0,0],[0.25,0.25,0.25]])
slabs=generate_all_slabs(CeO2,max_index=1,min_slab_size=8.0,min_vacuum_size=15.0,center_slab=True)
CeO2_111=[slab for slab in slabs if slab.miller_index==(1,1,1)][1]
print(CeO2_111.lattice.angles)
**The output**
(85.6269933583793, 85.6269933583793, 60.00000000000001)
**Expected behavior**
(90, 90, 60.00000000000001)
**Desktop (please complete the following information):**
- OS: Mac
- Version [e.g. 2022.0.8]
|
{
"login": "Rasic2",
"id": 44538197,
"node_id": "MDQ6VXNlcjQ0NTM4MTk3",
"avatar_url": "https://avatars.githubusercontent.com/u/44538197?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Rasic2",
"html_url": "https://github.com/Rasic2",
"followers_url": "https://api.github.com/users/Rasic2/followers",
"following_url": "https://api.github.com/users/Rasic2/following{/other_user}",
"gists_url": "https://api.github.com/users/Rasic2/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Rasic2/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Rasic2/subscriptions",
"organizations_url": "https://api.github.com/users/Rasic2/orgs",
"repos_url": "https://api.github.com/users/Rasic2/repos",
"events_url": "https://api.github.com/users/Rasic2/events{/privacy}",
"received_events_url": "https://api.github.com/users/Rasic2/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2161/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2161/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2162
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2162/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2162/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2162/events
|
https://github.com/materialsproject/pymatgen/issues/2162
| 910,381,028
|
MDU6SXNzdWU5MTAzODEwMjg=
| 2,162
|
Inefficient k-point generation for NiO (and possibly other structures)
|
{
"login": "ab5424",
"id": 57258530,
"node_id": "MDQ6VXNlcjU3MjU4NTMw",
"avatar_url": "https://avatars.githubusercontent.com/u/57258530?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ab5424",
"html_url": "https://github.com/ab5424",
"followers_url": "https://api.github.com/users/ab5424/followers",
"following_url": "https://api.github.com/users/ab5424/following{/other_user}",
"gists_url": "https://api.github.com/users/ab5424/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ab5424/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ab5424/subscriptions",
"organizations_url": "https://api.github.com/users/ab5424/orgs",
"repos_url": "https://api.github.com/users/ab5424/repos",
"events_url": "https://api.github.com/users/ab5424/events{/privacy}",
"received_events_url": "https://api.github.com/users/ab5424/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"You didn't use the default `grid_density`, so you're getting a k-point mesh commensurate with that. If you want a smaller k-point mesh, then set `grid_density` to a smaller number. \r\n\r\nAlso, that is an example calculation from the VASP manual. It is not in any way a recommendation on k-point density. That you have to decide based on what you want to do. ",
"Sorry @shyamd, I probably wasn't precise enough. I did two calculations with **exactly the same settings**, one with the structure from the MP and one with the structure from VASP, like this:\r\n\r\n```\r\nwith MPRester() as a:\r\n struct_MP = a.get_structure_by_material_id('mp-19009')\r\nrelax = MPRelaxSet(structure=struct_MP)\r\nrelax.write_input(\"struct_MP\")\r\n\r\n\r\nstruct_VASP = Structure.from_file('NiO-VASP-POSCAR') # This is the structure from the VASP website\r\nrelax = MPRelaxSet(structure=struct_VASP)\r\nrelax.write_input(\"struct_VASP\")\r\n```\r\n\r\nWith the structure from the MP, I got `Gamma 9 9 5` and with the VASP structure I got `Gamma 7 7 7` in the `KPOINTS` file, with the latter being far more efficient, as I explained in my first post. (In this case, changing the grid setting didn't influence the setup.) \r\n\r\nThe issue seems to be the initial cell vectors. I'm not sure if that affects other compounds with the same structure type but saving ~62% of computing time by changing the orientation of the vectors seems a lot to me. I'm currently calculating phonons where I need very accurate forces, so I have to use strict (and expensive) convergence criteria. I also used `StructureMatcher` to verify that both structures are the same.",
"This is a difficult problem. It's hard to tell apriori which setting will be the fastest. Sometimes you can get lower k-points, but that will result in slower electronic convergence. Depending on what the user is doing it's possible the settings are purposely in a setting that's not ideal. The `InputSet` respects whatever structure the user provides and its up to the user to optimize the settings. \r\n\r\nMP's audience isn't just researchers using DFT. We choose to put structures in a conventional standard-setting to make the structure easily interpretable. ",
"@ab5424 are you saying that both cases have equivalent numbers of overall k-points, but a difference in the number of irreducible k-points due to a difference in crystallographic setting?",
"@mkhorton It's almost the same overall number (9*9*5= 405, 7*7*7=343), due to rounding. But yes, that was what i tried to say, due to the different lattice vectors, the number of irreducible k-points differ significantly.\r\n\r\n@shyamd I assume that it's best for any code to use an input structure that is as symmetric as possible, in any aspect. The \"VASP website\" structure has reciprocal lattice vectors with equal length, the MP structure does not:\r\n\r\nVASP:\r\n```\r\n direct lattice vectors reciprocal lattice vectors\r\n 4.170000000 2.085000000 2.085000000 0.359712230 -0.119904077 -0.119904077\r\n 2.085000000 4.170000000 2.085000000 -0.119904077 0.359712230 -0.119904077\r\n 2.085000000 2.085000000 4.170000000 -0.119904077 -0.119904077 0.359712230\r\n\r\n length of vectors\r\n 5.107186114 5.107186114 5.107186114 0.397676833 0.397676833 0.397676833\r\n ```\r\n\r\nMP:\r\n```\r\n direct lattice vectors reciprocal lattice vectors\r\n 2.109361000 -0.000052000 -2.109347000 0.355613956 -0.118450847 -0.118461130\r\n 0.000054000 -2.109368000 2.109347000 0.118451353 -0.355612888 0.118460906\r\n -2.105111000 -2.105116000 -4.214497000 -0.118699391 -0.118699109 -0.118697252\r\n\r\n length of vectors\r\n 2.983077035 2.983081985 5.159940955 0.393096589 0.393095708 0.205591978\r\n```\r\n\r\nThe volume is the same, the positions are the same etc. Is there any way to construct the structure with equally long (reciprocal) lattice vectors, and would that conflict with the MP's standard settings?",
"I would offer a few general points. This doesn't respond to your issue directly Alexander, but might help elucidate:\r\n\r\n* The MP standard settings (our current defaults) are comparatively old, so don't necessarily reflect best practices _today_\r\n* Newer MP data (not yet released) is using KSPACING as default rather than this method, you can check out the MPScanSet as an example of how we're using this -- our new scheme is using a KSPACING that is band-gap dependent\r\n* For an arbitrary material, it is very difficult to know in advance which setting would give the optimal k-point grid if using the Monkhorst-Pack algorithm\r\n* However, to address this, there are now even better methods available, e.g. see generalized k-point grids in [this paper](https://doi.org/10.1016/j.commatsci.2020.110100) and [this paper](https://doi.org/10.1016/j.commatsci.2019.109340). Code exists for both. In pymatgen, we had an interface to the k-point generation API in `pymatgen.ext.jhu`, but both groups now offer local codes that can be used instead. I believe the former integrates nicely with pymagen already (@shyamd might comment further, given that he's on the former paper).\r\n\r\n> The volume is the same, the positions are the same etc. Is there any way to construct the structure with equally long (reciprocal) lattice vectors, and would that conflict with the MP's standard settings?\r\n\r\nTo this point, what you probably want is the LLL reduced structure. There is a method on the `Structure` object that can give you this. This applies a fast algorithm that gives lattice vectors that likely are as orthogonal as possible. It does not _guarantee_ to fulfill this condition, but algorithms that do this rigorously are that much slower, and the LLL reduced lattice is almost always \"good enough.\"",
"@mkhorton Thanks for the clarification. I guess NiO is a special case due to the AFM structure. For now, `KSPACING` might be a better option when writing new input sets and I will try this for my phonon calculations."
] | 2021-06-03T11:04:37
| 2021-07-06T10:23:57
|
2021-07-06T10:23:57Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
When using the NiO structure (mp-19009) from the MP, the generation of k-points in VASP is inefficient.
**To Reproduce**
Using
```
with MPRester() as a:
struct = a.get_structure_by_material_id('mp-19009')
relax = MPRelaxSet(structure=struct,
user_kpoints_settings={"grid_density": 2000},
force_gamma=True)
```
yields 115 irreducible k-points in the calculation:
`Found 115 irreducible k-points:` (from OUTCAR)
**Expected behavior**
When using the POSCAR provided by the VASP manual (https://www.vasp.at/wiki/index.php/NiO), I get 44 irreducible k-points, with otherwise identical settings.
`Found 44 irreducible k-points:`
This is attributed to the different meshes in the k-point files (`Gamma 9 9 5` for MP structure vs `Gamma 7 7 7` for VASP structure). This isn't really a bug but it seems quite inefficient to me (I stumbled across this when carrying out HSE calculations). I'm also not sure if that only affects this structure or others as well.
|
{
"login": "ab5424",
"id": 57258530,
"node_id": "MDQ6VXNlcjU3MjU4NTMw",
"avatar_url": "https://avatars.githubusercontent.com/u/57258530?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ab5424",
"html_url": "https://github.com/ab5424",
"followers_url": "https://api.github.com/users/ab5424/followers",
"following_url": "https://api.github.com/users/ab5424/following{/other_user}",
"gists_url": "https://api.github.com/users/ab5424/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ab5424/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ab5424/subscriptions",
"organizations_url": "https://api.github.com/users/ab5424/orgs",
"repos_url": "https://api.github.com/users/ab5424/repos",
"events_url": "https://api.github.com/users/ab5424/events{/privacy}",
"received_events_url": "https://api.github.com/users/ab5424/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2162/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2162/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2163
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2163/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2163/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2163/events
|
https://github.com/materialsproject/pymatgen/pull/2163
| 911,825,066
|
MDExOlB1bGxSZXF1ZXN0NjYyMDQyMzA2
| 2,163
|
Fix KSPACING formula in MPScanRelaxSet
|
{
"login": "ab5424",
"id": 57258530,
"node_id": "MDQ6VXNlcjU3MjU4NTMw",
"avatar_url": "https://avatars.githubusercontent.com/u/57258530?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ab5424",
"html_url": "https://github.com/ab5424",
"followers_url": "https://api.github.com/users/ab5424/followers",
"following_url": "https://api.github.com/users/ab5424/following{/other_user}",
"gists_url": "https://api.github.com/users/ab5424/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ab5424/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ab5424/subscriptions",
"organizations_url": "https://api.github.com/users/ab5424/orgs",
"repos_url": "https://api.github.com/users/ab5424/repos",
"events_url": "https://api.github.com/users/ab5424/events{/privacy}",
"received_events_url": "https://api.github.com/users/ab5424/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"FYI @rkingsbury, can I ask for your review?",
"Thanks for the catch @ab5424 ",
"\n[](https://coveralls.io/builds/40323647)\n\nCoverage decreased (-0.6%) to 83.077% when pulling **bb1de613d938ea89e4eecc96227d8a849c5b62fc on ab5424:kspacing** into **463e4f1b577287c593e493b2b1138320418f0bc7 on materialsproject:master**.\n",
"Thanks for catching @ab5424 . @mkhorton I can confirm; I looked at the paper and the workflow again. The difference in KSPACING appears to be small, and the previous (incorrect) equation would have resulted in a tighter / more stringent KSPACING value than the one that will be computed now.",
"Great, will merge. Thanks both!"
] | 2021-06-04T20:46:24
| 2021-06-07T19:25:32
|
2021-06-07T19:25:24Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Change rmin formula to match Eq. 25 from the paper.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2163/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2163/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2163",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2163",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2163.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2163.patch",
"merged_at": "2021-06-07T19:25:24Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2164
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2164/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2164/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2164/events
|
https://github.com/materialsproject/pymatgen/pull/2164
| 912,656,269
|
MDExOlB1bGxSZXF1ZXN0NjYyNzk4NzY5
| 2,164
|
reorder SETTINGS load priority
|
{
"login": "ardunn",
"id": 19936203,
"node_id": "MDQ6VXNlcjE5OTM2MjAz",
"avatar_url": "https://avatars.githubusercontent.com/u/19936203?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ardunn",
"html_url": "https://github.com/ardunn",
"followers_url": "https://api.github.com/users/ardunn/followers",
"following_url": "https://api.github.com/users/ardunn/following{/other_user}",
"gists_url": "https://api.github.com/users/ardunn/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ardunn/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ardunn/subscriptions",
"organizations_url": "https://api.github.com/users/ardunn/orgs",
"repos_url": "https://api.github.com/users/ardunn/repos",
"events_url": "https://api.github.com/users/ardunn/events{/privacy}",
"received_events_url": "https://api.github.com/users/ardunn/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/40337082)\n\nCoverage decreased (-0.6%) to 83.078% when pulling **ea9c9e54431280223f9f2c70435fad256f463245 on ardunn:master** into **463e4f1b577287c593e493b2b1138320418f0bc7 on materialsproject:master**.\n",
"Thanks @ardunn "
] | 2021-06-06T06:13:29
| 2021-06-07T18:21:57
|
2021-06-07T18:21:57Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
When both a .pmgrc.yaml and environment variables are specified, any pmg environment variable will not be read into `pymatgen.core.__init__`'s `SETTINGS`.
E.g., I have a `PMG_MAPI_KEY` as an environment variable and my .pmgrc.yaml does not contain that key - it looks like
```
#.pmgrc.yaml
MAPI_DB_VERSION:
LAST_ACCESSED: '2021_05_13'
LOG: {'2021_05_13': 6}
```
`PMG_MAPI_KEY` will not be found in `SETTINGS`.
This PR assumes this is not the desired behavior. `SETTINGS` here are now loaded as the union of env vars and pmgrc.yaml values, with the priority being pmgrc.yaml in case of a setting being defined in both sources.
If the desired behavior is in fact to void env variables for `SETTINGS` if .pmgrc.yaml exists, please close this PR.
## Checklist
Before a pull request can be merged, the following items must be checked:
- [X] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/). The easiest way to handle this
is to run the following in the **correct sequence** on your local machine. Start with running
[black](https://black.readthedocs.io/en/stable/index.html) on your new code. This will automatically reformat
your code to PEP8 conventions and removes most issues. Then run
[pycodestyle](https://pycodestyle.readthedocs.io/en/latest/), followed by
[flake8](http://flake8.pycqa.org/en/latest/).
- [X] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [X] N/A Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/) to type check your code.
- [X] N/A Tests have been added for any new functionality or bug fixes.
- [X] All linting and tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2164/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2164/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2164",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2164",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2164.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2164.patch",
"merged_at": "2021-06-07T18:21:57Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2165
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2165/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2165/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2165/events
|
https://github.com/materialsproject/pymatgen/pull/2165
| 912,855,849
|
MDExOlB1bGxSZXF1ZXN0NjYyOTc0NjUz
| 2,165
|
QChem: detect NLebdevPts error in QCOutput
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Note that there seems to be a new pylint error causing linting to fail on all PRs\r\n```\r\n************* Module pymatgen.entries.computed_entries\r\npymatgen/entries/computed_entries.py:700:8: E1101: Instance of 'ComputedStructureEntry' has no '_composition' member (no-member)\r\n```",
"\n[](https://coveralls.io/builds/40340146)\n\nCoverage decreased (-0.6%) to 83.079% when pulling **7ff37b4acd7298b05bdaf32ddcec2694745c8788 on rkingsbury:qcoutput_lebdevpts** into **463e4f1b577287c593e493b2b1138320418f0bc7 on materialsproject:master**.\n",
"You can try to fix this error? Technically, there should be a _compositon variable. If not, if you are sure this is purely a linter issue, you can set the ignore.",
"@shyuep this is not related to this PR, this is a pre-existing issue. I've taken a look at it already + will fix soon. I think it's just started appearing in logs because mypy is getting smarter.",
"Thanks @rkingsbury! Have merged since linter issue was unrelated to this PR and otherwise looks good. Will try to get linters passing in the main branch again soon."
] | 2021-06-06T15:01:08
| 2021-07-15T14:33:04
|
2021-06-17T00:37:19Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Detect `need to increase the array of NLebdevPts` error in `QCOutput` that can occur when computing RESP charges. Detecting this error is necessary to enable a handler in custodian. See [custodian #170](https://github.com/materialsproject/custodian/pull/170)
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2165/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2165/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2165",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2165",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2165.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2165.patch",
"merged_at": "2021-06-17T00:37:19Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2166
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2166/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2166/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2166/events
|
https://github.com/materialsproject/pymatgen/issues/2166
| 914,064,642
|
MDU6SXNzdWU5MTQwNjQ2NDI=
| 2,166
|
typing_extensions dependency
|
{
"login": "htz1992213",
"id": 25351437,
"node_id": "MDQ6VXNlcjI1MzUxNDM3",
"avatar_url": "https://avatars.githubusercontent.com/u/25351437?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/htz1992213",
"html_url": "https://github.com/htz1992213",
"followers_url": "https://api.github.com/users/htz1992213/followers",
"following_url": "https://api.github.com/users/htz1992213/following{/other_user}",
"gists_url": "https://api.github.com/users/htz1992213/gists{/gist_id}",
"starred_url": "https://api.github.com/users/htz1992213/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/htz1992213/subscriptions",
"organizations_url": "https://api.github.com/users/htz1992213/orgs",
"repos_url": "https://api.github.com/users/htz1992213/repos",
"events_url": "https://api.github.com/users/htz1992213/events{/privacy}",
"received_events_url": "https://api.github.com/users/htz1992213/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This is an issue in `pymatgen.io.qchem.inputs` not in `setup.py`, typing_extensions is not required for Python >= 3.8 since the relevant type hints are built-in. Fixed with https://github.com/materialsproject/pymatgen/commit/73783005ce1e9030f06d50125abd87e4071183f3.\r\n\r\nThanks @htz1992213 for reporting, fyi @espottesmith "
] | 2021-06-07T23:55:21
| 2021-06-08T00:25:15
|
2021-06-08T00:25:14Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Dependency requirements for `typing-extensions` should apply for any python versions, not just python<3.8. Current version:
https://github.com/materialsproject/pymatgen/blob/dfc90b07d6362848442ce611ebe1c3a3e1ca2926/requirements.txt#L17
https://github.com/materialsproject/pymatgen/blob/dfc90b07d6362848442ce611ebe1c3a3e1ca2926/setup.py#L124-L125
**To Reproduce**
Steps to reproduce the behavior:
```
conda create -n test_pymatgen python=3.8
source activate test_pymatgen
conda install --channel conda-forge pymatgen
python
>>> import pymatgen.io.qchem.inputs
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
File "/global/homes/t/thou/.conda/envs/test_pymatgen/lib/python3.8/site-packages/pymatgen/io/qchem/inputs.py", line 10, in <module>
from typing_extensions import Literal
ModuleNotFoundError: No module named 'typing_extensions'
```
**Expected behavior**
The import should work.
**Additional context**
The `typing-extensions` package is not a built-in package for any python version. It should be explicitly required not only for python<3.8, but also >=3.8, even though the imported class is a built-in `typing` class for python>=3.8.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2166/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2166/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2167
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2167/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2167/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2167/events
|
https://github.com/materialsproject/pymatgen/issues/2167
| 914,977,545
|
MDU6SXNzdWU5MTQ5Nzc1NDU=
| 2,167
|
Could it manually enter high symmetry point information when I draw energy band?
|
{
"login": "Nick12-hub",
"id": 62950321,
"node_id": "MDQ6VXNlcjYyOTUwMzIx",
"avatar_url": "https://avatars.githubusercontent.com/u/62950321?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Nick12-hub",
"html_url": "https://github.com/Nick12-hub",
"followers_url": "https://api.github.com/users/Nick12-hub/followers",
"following_url": "https://api.github.com/users/Nick12-hub/following{/other_user}",
"gists_url": "https://api.github.com/users/Nick12-hub/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Nick12-hub/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Nick12-hub/subscriptions",
"organizations_url": "https://api.github.com/users/Nick12-hub/orgs",
"repos_url": "https://api.github.com/users/Nick12-hub/repos",
"events_url": "https://api.github.com/users/Nick12-hub/events{/privacy}",
"received_events_url": "https://api.github.com/users/Nick12-hub/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-06-08T12:33:39
| 2023-08-13T16:34:59
|
2023-08-13T16:34:59Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Could it manually enter high symmetry point information when I draw energy band?In fact I have Dirac points (k and k') inside the input KPOINTS file, which prevents my energy band diagram from showing the horizontal coordinates.Hope for reply,thanks!
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2167/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2167/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2168
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2168/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2168/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2168/events
|
https://github.com/materialsproject/pymatgen/issues/2168
| 916,127,970
|
MDU6SXNzdWU5MTYxMjc5NzA=
| 2,168
|
[Improvment] duplicated data selection from `exterema_df`
|
{
"login": "ezpzbz",
"id": 13301031,
"node_id": "MDQ6VXNlcjEzMzAxMDMx",
"avatar_url": "https://avatars.githubusercontent.com/u/13301031?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ezpzbz",
"html_url": "https://github.com/ezpzbz",
"followers_url": "https://api.github.com/users/ezpzbz/followers",
"following_url": "https://api.github.com/users/ezpzbz/following{/other_user}",
"gists_url": "https://api.github.com/users/ezpzbz/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ezpzbz/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ezpzbz/subscriptions",
"organizations_url": "https://api.github.com/users/ezpzbz/orgs",
"repos_url": "https://api.github.com/users/ezpzbz/repos",
"events_url": "https://api.github.com/users/ezpzbz/events{/privacy}",
"received_events_url": "https://api.github.com/users/ezpzbz/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-06-09T12:08:40
| 2021-06-09T12:08:40
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hi,
I have been using the `ChargeInsertionAnalyzer` class and realized that within the `get_label` function, we first select the desired data with average charge density below the maximum threshold from `extrema_df`:
https://github.com/materialsproject/pymatgen/blob/c724e7b679581783dcdb4690a83c374a57437ae6/pymatgen/analysis/defects/utils.py#L1344
However, later below where we are constructing the `inserted_strc`, we check again if the site average charge density is below the threshold:
https://github.com/materialsproject/pymatgen/blob/c724e7b679581783dcdb4690a83c374a57437ae6/pymatgen/analysis/defects/utils.py#L1347-L1348
I think we remove the latter condition from the loop.
If you think this makes sense to be done, I'd be happy to open a PR.
Cheers,
Pezhman
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2168/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2168/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2169
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2169/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2169/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2169/events
|
https://github.com/materialsproject/pymatgen/issues/2169
| 916,803,036
|
MDU6SXNzdWU5MTY4MDMwMzY=
| 2,169
|
None radius for He Ne
|
{
"login": "LanceKnight",
"id": 5760199,
"node_id": "MDQ6VXNlcjU3NjAxOTk=",
"avatar_url": "https://avatars.githubusercontent.com/u/5760199?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/LanceKnight",
"html_url": "https://github.com/LanceKnight",
"followers_url": "https://api.github.com/users/LanceKnight/followers",
"following_url": "https://api.github.com/users/LanceKnight/following{/other_user}",
"gists_url": "https://api.github.com/users/LanceKnight/gists{/gist_id}",
"starred_url": "https://api.github.com/users/LanceKnight/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/LanceKnight/subscriptions",
"organizations_url": "https://api.github.com/users/LanceKnight/orgs",
"repos_url": "https://api.github.com/users/LanceKnight/repos",
"events_url": "https://api.github.com/users/LanceKnight/events{/privacy}",
"received_events_url": "https://api.github.com/users/LanceKnight/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 5322879170,
"node_id": "LA_kwDOACgets8AAAABPUSwwg",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/help",
"name": "help",
"color": "5ED16E",
"default": false,
"description": "Q&A support-style issues"
}
] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This is just that there is no provided radius in the data source, https://github.com/materialsproject/pymatgen/blob/master/pymatgen/core/periodic_table.json\r\n\r\ne.g. for He\r\n\r\n```json\r\n \"He\":{\r\n \"Atomic mass\":4.002602,\r\n \"Atomic no\":2,\r\n \"Atomic orbitals\":{\r\n \"1s\":-0.570425\r\n },\r\n \"Atomic radius\":\"no data\",\r\n \"Atomic radius calculated\":0.31,\r\n \"Boiling point\":\"4.22 K\",\r\n \"Brinell hardness\":\"no data MN m<sup>-2</sup>\",\r\n```",
"Thanks!"
] | 2021-06-10T01:38:29
| 2023-05-30T02:31:46
|
2023-05-30T02:31:46Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hello, I'm a new user of pymatgen. When I try to calculate the radius of He and Ne, I got None. I guess this is designed so on purpose. Can someone explain to me why?
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2169/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2169/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2170
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2170/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2170/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2170/events
|
https://github.com/materialsproject/pymatgen/issues/2170
| 917,087,551
|
MDU6SXNzdWU5MTcwODc1NTE=
| 2,170
|
Adding alternative symmetry finding backends to pymatgen
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"It depends on how it's implemented, and how easy it is to switch backends. We would not want any core code that requires installing packages outside pip, and would not want to add clutter within the existing classes. A more general coupling between pymatgen and the symmetry-finding code might result in the pymatgen code being cleaner however, and the feature in general sounds good.\r\n\r\n> I think I have a reasonable idea on how to implement this\r\n\r\nCould you outline this more concretely?",
"I'm not familiar with that paper, but from a quick skim it seems that the only major advance is that they iteratively change the symmetry-finding tolerance until a consistency criterion is satisfied. I suspect spglib could be used in a similar way too. Regardless, I think having the option to switch or try out different methods is a good thing.",
"I am always open to better implementations. But in this case, I would suggest you define what is \"better results\" first. Even I am not entirely clear how to compare different symmetry finding packages.\r\n\r\nI would rank my preference as follows:\r\n1. Built in support in pymatgen for symmetry finding, either as a Python-only or as a Python C extension. The former would need to be nearly as fast as the latter to be worthwhile. But I can live with the latter.\r\n2. Spglib or some other external pip installable package. \r\n3. External command line tools (I am aware of a few).\r\n\r\nThere is nothing preventing us from having multiple of these supported. But it needs to be worthwhile to spend the effort. So far, we don't have the first option since none of us want to make that considerable effort. But that would be an ideal long term goal.",
"> I am always open to better implementations. But in this case, I would suggest you define what is \"better results\" first. Even I am not entirely clear how to compare different symmetry finding packages.\r\n\r\nI think the better I am targeting is robustness but that robustness comes from adapting the tolerances and looking for consistency. In the testing, I've done looking at more \"robust\" symmetry finders `findsym` is about 10x slower than `spglib` and `aflow-sym` maybe 100x slower although both do place many fewer materials in P1 which is the proxy I was using for robustness. \r\n\r\nI think that probably on balance the backend idea using cli tools might end up being unsupported longer term and so maybe it's better if I look at whether it would be possible to do as @mkhorton suggests and mimic the consistency checking of `aflow-sym` using `spglib` as the backend inside pymatgen and then if I can get that to work it could be turned on and off with a `auto_adjust_tol` kwarg or something like that. \r\n\r\n",
"Ok, I think that is not really a definition of better. If I set tolerances to be 0, I am guaranteed to get P1 far more often than a higher symmetry. If I set tolerances really high, I tend to get cubic or other high symmetries more often. I wouldn't even know where to begin to stipulate what the ground truth symmetry is - which is what you need to say something is better.\r\n\r\nAdaptive tolerances is generally a good idea - I have done this in the past. ",
"> mimic the consistency checking of aflow-sym using spglib as the backend inside pymatgen and then if I can get that to work it could be turned on and off with a auto_adjust_tol kwarg or something like that.\r\n\r\nThis would be interesting. Perhaps @atztogo would be interested in implementing directly in spglib too if the benefit is clear. However, I don't want to be too quick to form an opinion at this point since I have not had time to review the paper thoroughly.",
"Tolerance parameter is a complicated parameter in symmetry finding. At least I don't define it mathematically well, so what I could explain was just how it is implemented in spglib, https://arxiv.org/abs/1808.01590 (maybe see Fig. 1.) We can improve spglib. But I want to balance among (1) reasonable symmetry identification (2) speed (3) portability (4) maintainability under the constraint that the fact that the main developer is only me.\r\n\r\nWhat might be wanted to discuss here may be not as simple as you might think. As @shyuep wrote, the definition is unclear. What @mkhorton may expect to spglib is not clear for me. If you could define what you really want, I am happy to consider its implementation. Probably, the difficulty exists not in the implementation, but in the definition and the design.\r\n\r\nBy the way, I've never watched the aflow-sym code, but it seems it's under the GPL. So in a strict license manner, it might invoke license issue.",
"> By the way, I've never watched the aflow-sym code, but it seems it's under the GPL. So in a strict license manner, it might invoke license issue.\r\n\r\nMy understanding is that the code is GPL, but the algorithm described in the paper is freely available to implement.\r\n\r\n> What @mkhorton may expect to spglib is not clear for me. If you could define what you really want, I am happy to consider its implementation.\r\n\r\nYes, I may have a poor understanding here. I was looking at Figure 5 of the [paper here](https://onlinelibrary.wiley.com/doi/pdf/10.1107/S2053273318003066?casa_token=dKAY7wSjt1AAAAAA:AFlijXEKJK-Rb-HqRlapIi9EARUEwLs4UwViPhXsHpbm70ja1Rxk0EISUq5SWSj8RQoBtzDO-GhlN0h96Q). My understanding is that spglib already does most of what this describes except perhaps for the consistency checks specified in §2.6 of the paper. Thank you for the response!",
"> My understanding is that the code is GPL, but the algorithm described in the paper is freely available to implement.\r\n\r\nYes. I wanted to mention to the idea by @CompRhys.\r\n\r\n> Yes, I may have a poor understanding here. I was looking at Figure 5 of the paper here. My understanding is that spglib already does most of what this describes except perhaps for the consistency checks specified in §2.6 of the paper. Thank you for the response!\r\n\r\nIn spglib, those symmetries are check to be consistent. Probably we can have more checks. I am not a specialist of crystallography. So if the materials science community wants to have a better symmetry finder, the first thing to do may be to discuss with crystallographers, such as the cctbx project. If I remember correctly (maybe wrong), sginfo was under GPL or something that I wanted to avoid to use (not now), and so I could not use it when I needed it.\r\n\r\nThe adjustment of tolerance parameter (or the way to tolerate) is done in a primitive way in spglib. Here \"a primitive way\" does not mean with respect to other symmetry finders (because I don't know others), but with respect to my feeling that we can implement a more clever way. We know many things to do for the improvement but this kind of work including maintenance and support is not respected and no chance to get paid."
] | 2021-06-10T08:51:38
| 2021-06-11T00:03:36
|
2021-06-10T17:32:23Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Would a PR be welcomed adding alternative symmetry finding backends to pymatgen?
Currently pymatgen uses `spglib` which should definitely remain the default due to speed and the fact that it's a pip-installable package but alternative finders such as find-sym and aflow-sym trade can produce better results - https://doi.org/10.1107/S2053273318003066.
I think I have a reasonable idea on how to implement this but it would require a config file somewhere to allow users to add paths to their aflow/findsym executables.
Would this be supported if I try to do it?
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2170/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2170/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2171
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2171/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2171/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2171/events
|
https://github.com/materialsproject/pymatgen/pull/2171
| 917,238,212
|
MDExOlB1bGxSZXF1ZXN0NjY2ODAwMTY1
| 2,171
|
Avoid error exit for ALGO=CHI
|
{
"login": "KazMorita",
"id": 22847784,
"node_id": "MDQ6VXNlcjIyODQ3Nzg0",
"avatar_url": "https://avatars.githubusercontent.com/u/22847784?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/KazMorita",
"html_url": "https://github.com/KazMorita",
"followers_url": "https://api.github.com/users/KazMorita/followers",
"following_url": "https://api.github.com/users/KazMorita/following{/other_user}",
"gists_url": "https://api.github.com/users/KazMorita/gists{/gist_id}",
"starred_url": "https://api.github.com/users/KazMorita/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/KazMorita/subscriptions",
"organizations_url": "https://api.github.com/users/KazMorita/orgs",
"repos_url": "https://api.github.com/users/KazMorita/repos",
"events_url": "https://api.github.com/users/KazMorita/events{/privacy}",
"received_events_url": "https://api.github.com/users/KazMorita/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Can you add a simple unittest for this? Thanks.",
"\n[](https://coveralls.io/builds/40537438)\n\nCoverage decreased (-0.6%) to 83.083% when pulling **180153deaf7da2d6bfbd016c20d78c305bb273a2 on KazMorita:master** into **afa2cb9ba4111b4488933692d14f3c4356060199 on materialsproject:master**.\n",
"> Can you add a simple unittest for this? Thanks.\r\n\r\nI added a file, \"test_files/vasprun.xml.chi.gz\", and implemented a test to read it.\r\nPlease check."
] | 2021-06-10T11:32:11
| 2021-06-13T02:31:24
|
2021-06-13T02:27:57Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Added ALGO=CHI in place of ALGO=BSE to avoid the code from error exiting.
* This change is minor, so no change to the document is required.
## Additional dependencies introduced (if any)
* non
## TODO (if any)
* non
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2171/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2171/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2171",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2171",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2171.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2171.patch",
"merged_at": "2021-06-13T02:27:57Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2172
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2172/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2172/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2172/events
|
https://github.com/materialsproject/pymatgen/pull/2172
| 917,544,118
|
MDExOlB1bGxSZXF1ZXN0NjY3MDYzODg0
| 2,172
|
Use tighter EDIFF for DFPT calculations
|
{
"login": "utf",
"id": 1330638,
"node_id": "MDQ6VXNlcjEzMzA2Mzg=",
"avatar_url": "https://avatars.githubusercontent.com/u/1330638?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/utf",
"html_url": "https://github.com/utf",
"followers_url": "https://api.github.com/users/utf/followers",
"following_url": "https://api.github.com/users/utf/following{/other_user}",
"gists_url": "https://api.github.com/users/utf/gists{/gist_id}",
"starred_url": "https://api.github.com/users/utf/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/utf/subscriptions",
"organizations_url": "https://api.github.com/users/utf/orgs",
"repos_url": "https://api.github.com/users/utf/repos",
"events_url": "https://api.github.com/users/utf/events{/privacy}",
"received_events_url": "https://api.github.com/users/utf/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"test pls.",
"Done.",
"\n[](https://coveralls.io/builds/40484767)\n\nCoverage decreased (-0.6%) to 83.082% when pulling **d1ee3c77f84dd68ad30dd93d74cbd63d74f8ec60 on utf:dfpt** into **77c6dd669e159ca5fc5b2605b5a07b0bd3e500f9 on materialsproject:master**.\n",
"Failing linting is unrelated to this PR:\r\n\r\n```\r\npymatgen/cli/pmg.py:14: error: Library stubs not installed for \"tabulate\" (or incompatible with Python 3.9)\r\n... etc\r\n```"
] | 2021-06-10T16:24:36
| 2021-06-11T16:49:47
|
2021-06-11T16:49:47Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Set EDIFF = 1e-5 when `lepsilon = True` in `MPStaticSet`. Note that this is [already used in atomate](https://github.com/hackingmaterials/atomate/blob/128a754000533029ce2b3d1faf3eb06aabe6f88b/atomate/vasp/firetasks/write_inputs.py#L378), I'm just moving the configuration to the input set itself.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2172/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2172/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2172",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2172",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2172.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2172.patch",
"merged_at": "2021-06-11T16:49:47Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2173
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2173/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2173/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2173/events
|
https://github.com/materialsproject/pymatgen/issues/2173
| 919,202,513
|
MDU6SXNzdWU5MTkyMDI1MTM=
| 2,173
|
Significantly different Pourbaix diagrams somewhere between 2020.3.13 and 2022.0.8
|
{
"login": "mwitman1",
"id": 10645657,
"node_id": "MDQ6VXNlcjEwNjQ1NjU3",
"avatar_url": "https://avatars.githubusercontent.com/u/10645657?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mwitman1",
"html_url": "https://github.com/mwitman1",
"followers_url": "https://api.github.com/users/mwitman1/followers",
"following_url": "https://api.github.com/users/mwitman1/following{/other_user}",
"gists_url": "https://api.github.com/users/mwitman1/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mwitman1/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mwitman1/subscriptions",
"organizations_url": "https://api.github.com/users/mwitman1/orgs",
"repos_url": "https://api.github.com/users/mwitman1/repos",
"events_url": "https://api.github.com/users/mwitman1/events{/privacy}",
"received_events_url": "https://api.github.com/users/mwitman1/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @mwitman1, please see the Materials Project database release notes here for more information: https://matsci.org/t/materials-project-database-release-log/1609/18\r\n\r\n@rkingsbury can comment further (though he's not available this week, so may take some time for a response)",
"Hi @rkingsbury thanks for looking into it whenever you have some time, and sorry in the first post also wanted to highlight how the Materials Project web app still seems to be more consistent with the previous 2020.03.13 version at more reducing potentials, if you have any comment on that as well\r\n\r\n<img width=\"857\" alt=\"image\" src=\"https://user-images.githubusercontent.com/10645657/121938710-00fa5800-cd01-11eb-9903-36b69a9db6a1.png\">\r\n",
"Hi @mwitman, thanks for reporting. The discrepancy you found between your code and the website is due to the fact that we set the `filter_solids=True` kwarg when instantiating `PourbaixDiagram`, e.g.\r\n\r\n```\r\npbx = PourbaixDiagram(pourbaix, filter_solids=True)\r\n```\r\n\r\n `filter_solids=True` includes only solid entries that are stable on the compositional phase diagram when it constructs the Pourbaix diagram, whereas with `filter_solids=False`, even unstable solid polymorphs are included in the diagram. That can at times lead to strange results.\r\n\r\n To be honest, `filter_solids=True` should probably be the default because we consider that to be the most correct way to build these diagrams and that is what we use on the website. @montoyjh , do you have any further comments on this?\r\n\r\nI think the differences in energy you saw among some materials between your two versions are an artifact of a transitional period in our database release cycle (i.e., `get_pourbaix_entries` using legacy corrections even though the database was updated with new energies), so I wouldn't worry about that.\r\n\r\nHope this helps!\r\n\r\n",
"Yeah, my opinion is that `filter_solids=True` is the most correct way to represent equilibrium phase stability. The practical consequence of this is that highly oxidized or reduced phases that might show up in experiments due to kinetic limitations on oxygen/hydrogen evolution won't appear, but they're not actually \"stable\" (and are frequently overstabilized from DFT errors). I'd keep the option to not filter, but would be happy to see the default change to be consistent with the Materials Project website."
] | 2021-06-11T20:51:49
| 2021-06-20T23:12:59
|
2021-06-20T23:12:59Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
There seems to significant difference in the Pourbaix diagrams generated in 2022.0.08 and those up to at least v2020.3.13. I’m referencing v2020.3.13 because it’s the last version in the Change Log that makes a mention of “pourbaix”. Ultimately it seems to come down to the computed energies for each species in the diagram between the two versions, some of which are quite large (up to 1 ev/atom).
**To Reproduce**
```
from pymatgen.ext.matproj import MPRester
from pprint import pprint
from pymatgen.analysis.pourbaix_diagram import PourbaixDiagram, PourbaixPlotter
with MPRester(APIKEY) as mpr:
pourbaix=mpr.get_pourbaix_entries(['Cu'])
def f(lst):
“”” pretty prints the most stable entry for a given compound “””
lst = [(entry.composition.reduced_formula, entry.energy_per_atom) for entry in lst]
d = {}
for tup in lst:
if tup[0] not in d.keys():
d[tup[0]]=tup[1]
elif tup[1] < d[tup[0]]:
d[tup[0]] = tup[1]
return d
pprint(f(pourbaix))
pbx = PourbaixDiagram(pourbaix)
plotter = PourbaixPlotter(pbx)
plotter.get_pourbaix_plot().show()
```
**Expected behavior**
In **v2020.3.13** I get the following, and the pourbaix diagram is pretty close to what appears on the web app, although not quite identical:
{'Fe': -1.622125307056931,
'Fe(HO)2': -0.08565486141138604,
'Fe(HO)3': -0.037704901008132855,
'Fe10O11': -0.272528083333333,
'Fe11O12': -0.27000459956521927,
'Fe12O13': -0.23810561700000094,
'Fe13O14': -0.2507718170370378,
'Fe13O15': -0.22857405517857185,
'Fe13O19': 0.053782962968749404,
'Fe14O15': -0.2649596684482761,
'Fe15O16': -0.24799346483870952,
'Fe17O18': -0.2864528791428575,
'Fe21HO32': -0.04000913377074037,
'Fe21O23': -0.2655534476136363,
'Fe21O32': -0.019029330943396604,
'Fe23O25': -0.2702020153125,
'Fe23O32': -0.1528225829090913,
'Fe25O32': -0.09684627649122858,
'Fe2O3': -0.2420518570000006,
'Fe2O5': 0.9901734950000005,
'Fe32O35': -0.26137829828358267,
'Fe35O36': 0.45011210830985887,
'Fe38O39': -0.262887671623377,
'Fe3H': 1.1950894790949995,
'Fe3O': 1.1023692849999995,
'Fe3O4': -0.2570221514285707,
'Fe41O56': -0.17948183690721772,
'Fe43O64': -0.12777606065420513,
'Fe4H14O13': 1.0941170194941923,
'Fe4O13': 1.5738661873529407,
'Fe4O5': -0.1980533911111119,
'Fe5HO8': 0.05780113652714368,
'Fe5O7': -0.0888776762499992,
'Fe5O8': 0.036853481730769196,
'Fe7O8': -0.2445972323333334,
'Fe7O9': -0.06110978593750005,
'Fe8H10O17': 0.6559708343942865,
'Fe8O17': 0.6088237528000002,
'Fe8O9': -0.26275042058823506,
'Fe9O10': -0.2755539805263164,
'Fe9O13': 0.012218070227270868,
'FeH': 1.1894350456899998,
'FeH3': 2.034037118535,
'FeH4': 2.227225052104,
'FeHO': -0.4067181023523105,
'FeHO2': 0.049041423235767434,
'FeO': -0.3401552675000008,
'FeO2': 0.09847189764768978,
'FeO3': 1.4467245549999985,
'FeO4': 1.1378171385886138}
<img width="992" alt="Pasted Graphic 3" src="https://user-images.githubusercontent.com/10645657/121747165-d075ac80-cabb-11eb-8e77-df79a4b0e1a8.png">
Whereas in **v2022.0.8** I get the following. There is a very large discrepancy in the energies of various compounds including (especially H containing and even Fe itself), while some others seem to be relatively close. But it has a significant impact on the diagram. Would it be possible for the Materials Project team to trouble shoot this issue? Is this something to do with new data on MP or pymatgen processing of the pourbaix entries being incorrect in the newest version?
{'Fe': -1.1221903070569303,
'Fe(HO)2': 0.014332138588614107,
'Fe(HO)3': 0.033714384706152956,
'Fe10O11': -0.03737617857142726,
'Fe11O12': -0.033896773478261984,
'Fe12O13': -0.0011948170000003699,
'Fe13O14': -0.013177002222222607,
'Fe13O15': 0.0010813019642864471,
'Fe13O19': 0.25664265046874957,
'Fe14O15': -0.02677518568965515,
'Fe15O16': -0.009295400322580307,
'Fe17O18': -0.04690373628571385,
'Fe21HO32': 0.1409162460185182,
'Fe21O23': -0.02990185670454518,
'Fe21O32': 0.17920236716981103,
'Fe23O25': -0.03367597364583327,
'Fe23O32': 0.05554614436363517,
'Fe25O32': 0.12094810947368503,
'Fe2O3': -0.04207785700000031,
'Fe2O5': 1.1373806378571427,
'Fe32O35': -0.025570089328357977,
'Fe35O36': 0.6930056294366199,
'Fe38O39': -0.019740788506492496,
'Fe3H': 0.7268496062499992,
'Fe3O': 1.4639417849999998,
'Fe3O4': -0.04385643714285666,
'Fe41O56': 0.03096393628865947,
'Fe43O64': 0.0730609486915891,
'Fe4H14O13': 0.829550475,
'Fe4O13': 1.697793834411764,
'Fe4O5': 0.02244105333333203,
'Fe5HO8': 0.18430188714285656,
'Fe5O7': 0.11879149041666744,
'Fe5O8': 0.22972425096153898,
'Fe7O8': -0.013842565666666692,
'Fe7O9': 0.15617833906249956,
'Fe8H10O17': 0.5620518368571433,
'Fe8O17': 0.7718609528000002,
'Fe8O9': -0.03018512647058813,
'Fe9O10': -0.04155924368421145,
'Fe9O13': 0.2163894338636359,
'FeH': 0.2529553,
'FeH3': 0.6293175000000009,
'FeH4': 0.7288574589999989,
'FeHO': -0.24007310235231025,
'FeHO2': 0.17402517323576738,
'FeO': -0.09401026750000052,
'FeO2': 0.2651168976476897,
'FeO3': 1.5547095549999987,
'FeO4': 1.2378041385886138}
<img width="987" alt="10 12 14" src="https://user-images.githubusercontent.com/10645657/121747138-c6ec4480-cabb-11eb-8fe2-1b0b901328c2.png">
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2173/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2173/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2174
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2174/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2174/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2174/events
|
https://github.com/materialsproject/pymatgen/pull/2174
| 919,313,541
|
MDExOlB1bGxSZXF1ZXN0NjY4NjUxNzky
| 2,174
|
Q-Chem NBO
|
{
"login": "samblau",
"id": 7431223,
"node_id": "MDQ6VXNlcjc0MzEyMjM=",
"avatar_url": "https://avatars.githubusercontent.com/u/7431223?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/samblau",
"html_url": "https://github.com/samblau",
"followers_url": "https://api.github.com/users/samblau/followers",
"following_url": "https://api.github.com/users/samblau/following{/other_user}",
"gists_url": "https://api.github.com/users/samblau/gists{/gist_id}",
"starred_url": "https://api.github.com/users/samblau/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/samblau/subscriptions",
"organizations_url": "https://api.github.com/users/samblau/orgs",
"repos_url": "https://api.github.com/users/samblau/repos",
"events_url": "https://api.github.com/users/samblau/events{/privacy}",
"received_events_url": "https://api.github.com/users/samblau/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/40715205)\n\nCoverage decreased (-0.6%) to 83.141% when pulling **7f07cc4693489915ce953b7546ece9b30bccdd8d on samblau:qchem** into **df22c1a523eee198ad6381dad585ac9b851dc9b3 on materialsproject:master**.\n",
"@mkhorton This is ready for review at your convenience - no rush. Note that I have fixed the outstanding Linting issues currently in main.",
"Much appreciated @samblau !"
] | 2021-06-11T23:51:00
| 2021-06-19T00:06:23
|
2021-06-19T00:06:21Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Add Q-Chem NBO functionality, whereby the NBO package can be called from within Q-Chem
* Input reading/writing in QCInput and corresponding testing
* Output parsing in QCOutput and corresponding testing
* NBO functionality in Q-Chem sets and corresponding testing
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2174/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2174/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2174",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2174",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2174.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2174.patch",
"merged_at": "2021-06-19T00:06:21Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2175
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2175/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2175/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2175/events
|
https://github.com/materialsproject/pymatgen/issues/2175
| 921,836,006
|
MDU6SXNzdWU5MjE4MzYwMDY=
| 2,175
|
SpacegroupAnalyzer setting _space_group_data to None
|
{
"login": "kyledmiller",
"id": 45434005,
"node_id": "MDQ6VXNlcjQ1NDM0MDA1",
"avatar_url": "https://avatars.githubusercontent.com/u/45434005?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/kyledmiller",
"html_url": "https://github.com/kyledmiller",
"followers_url": "https://api.github.com/users/kyledmiller/followers",
"following_url": "https://api.github.com/users/kyledmiller/following{/other_user}",
"gists_url": "https://api.github.com/users/kyledmiller/gists{/gist_id}",
"starred_url": "https://api.github.com/users/kyledmiller/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/kyledmiller/subscriptions",
"organizations_url": "https://api.github.com/users/kyledmiller/orgs",
"repos_url": "https://api.github.com/users/kyledmiller/repos",
"events_url": "https://api.github.com/users/kyledmiller/events{/privacy}",
"received_events_url": "https://api.github.com/users/kyledmiller/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This is not a problem with pymatgen but rather with the CIF file itself. Some of the atoms generated are too near each other, resulting in atomic collisions in the symmetry finding algorithm.\r\n\r\nUsing your second CIF file as an example,\r\n\r\n```python\r\nfrom pymatgen.core import Structure\r\ns = Structure.from_file(\"bad.cif\")\r\nprint(s.get_space_group_info()). # this fails\r\ns.merge_sites(tol=0.01, mode=\"delete\") # this deletes atoms that are too close, i.e., less than 0.01 A apart\r\nprint(s.get_space_group_info()) # this returns ('R-3', 148)\r\n```\r\n\r\nBottom line is that CIF is a horrible format for a file and some CIFs have bad information. Pymatgen errs on the side of preserving as much information as possible, i.e., we do not merge or delete atoms unless they are really super close together and letting the user do any processing themselves.",
"To add some additional context, I think the real issue here is likely the finite precision of the atomic positions stated in the CIF file. When the CIF file is parsed, the full set of atomic positions are constructed with reference to the stated symmetry operations, but if the atomic positions are not supplied with sufficient precision, it can lead to two atoms being generated close to one another rather than the single atom that is desired.",
"Thank you both very much for the rapid response. These were indeed re-symmetrized using Pymatgen/SPGLib with fairly loose tolerances. I'll check out the atom collision functions you've created."
] | 2021-06-15T21:57:37
| 2021-06-16T16:02:03
|
2021-06-16T12:27:33Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
When attempting to fetch the spacegroup info for structures parsed from certain cifs (provided), an exception is thrown about the space group data being None.
**To Reproduce**
Copy in cifs provided at the bottom (sorry, I can't upload .cif files to these issues), run the code below.
```
from pymatgen.core.structure import Structure
struc = Structure.from_file('Zn26S31_42794.cif')
struc.get_space_group_info()
struc = Structure.from_file('Sc8NiCl12_424477.cif')
struc.get_space_group_info()
```
**Expected behavior**
No exception, correct space group assignment.
**Desktop (please complete the following information):**
- Ubuntu (Windows Subsystem for Linux)
- Version 18.04.5 LTS
**CIFs**
---Zn26S31_42794-------------------------------------------------------------------------------------
```
#(C) 2019 by FIZ Karlsruhe - Leibniz Institute for Information Infrastructure. All rights reserved.
data_42794-ICSD
_database_code_ICSD 42794
_audit_creation_date 2000-07-15
_chemical_name_systematic 'Zinc sulfide'
_chemical_formula_structural 'Zn S'
_chemical_formula_sum 'S1 Zn1'
_chemical_name_structure_type ZnS(78R)
_exptl_crystal_density_diffrn 4.09
_publ_section_title 'New Zn S polytypes of the families 10L, 22L and 26L'
loop_
_citation_id
_citation_journal_full
_citation_year
_citation_journal_volume
_citation_page_first
_citation_page_last
_citation_journal_id_ASTM
primary
;
Acta Crystallographica, Section B: Structural Crystallography and Crystal
Chemistry
; 1969 25 1195 1197 ACBCAR
loop_
_publ_author_name
'Kiflawi, I.'
'Mardix, S.'
_cell_length_a 3.823
_cell_length_b 3.823
_cell_length_c 243.672
_cell_angle_alpha 90.
_cell_angle_beta 90.
_cell_angle_gamma 120.
_cell_volume 3084.22
_cell_formula_units_Z 78
_symmetry_space_group_name_H-M 'R 3 m H'
_symmetry_Int_Tables_number 160
loop_
_symmetry_equiv_pos_site_id
_symmetry_equiv_pos_as_xyz
1 '-x+y, y, z'
2 'x, x-y, z'
3 '-y, -x, z'
4 '-x+y, -x, z'
5 '-y, x-y, z'
6 'x, y, z'
7 '-x+y+2/3, y+1/3, z+1/3'
8 'x+2/3, x-y+1/3, z+1/3'
9 '-y+2/3, -x+1/3, z+1/3'
10 '-x+y+2/3, -x+1/3, z+1/3'
11 '-y+2/3, x-y+1/3, z+1/3'
12 'x+2/3, y+1/3, z+1/3'
13 '-x+y+1/3, y+2/3, z+2/3'
14 'x+1/3, x-y+2/3, z+2/3'
15 '-y+1/3, -x+2/3, z+2/3'
16 '-x+y+1/3, -x+2/3, z+2/3'
17 '-y+1/3, x-y+2/3, z+2/3'
18 'x+1/3, y+2/3, z+2/3'
loop_
_atom_type_symbol
_atom_type_oxidation_number
Zn2+ 2
S2- -2
loop_
_atom_site_label
_atom_site_type_symbol
_atom_site_symmetry_multiplicity
_atom_site_Wyckoff_symbol
_atom_site_fract_x
_atom_site_fract_y
_atom_site_fract_z
_atom_site_B_iso_or_equiv
_atom_site_occupancy
_atom_site_attached_hydrogens
Zn1 Zn2+ 3 a 0 0 0 . 1. 0
Zn2 Zn2+ 3 a 0.3333 0.6667 0.01282 . 1. 0
Zn3 Zn2+ 3 a 0.6667 0.3333 0.02564 . 1. 0
Zn4 Zn2+ 3 a 0 0 0.03846 . 1. 0
Zn5 Zn2+ 3 a 0.3333 0.6667 0.05128 . 1. 0
Zn6 Zn2+ 3 a 0.6667 0.3333 0.0641 . 1. 0
Zn7 Zn2+ 3 a 0 0 0.07692 . 1. 0
Zn8 Zn2+ 3 a 0.3333 0.6667 0.08974 . 1. 0
Zn9 Zn2+ 3 a 0.6667 0.3333 0.10256 . 1. 0
Zn10 Zn2+ 3 a 0 0 0.11538 . 1. 0
Zn11 Zn2+ 3 a 0.6667 0.3333 0.12821 . 1. 0
Zn12 Zn2+ 3 a 0.3333 0.6667 0.14103 . 1. 0
Zn13 Zn2+ 3 a 0 0 0.15385 . 1. 0
Zn14 Zn2+ 3 a 0.3333 0.6667 0.16667 . 1. 0
Zn15 Zn2+ 3 a 0.6667 0.3333 0.17949 . 1. 0
Zn16 Zn2+ 3 a 0 0 0.19231 . 1. 0
Zn17 Zn2+ 3 a 0.6667 0.3333 0.20513 . 1. 0
Zn18 Zn2+ 3 a 0.3333 0.6667 0.21795 . 1. 0
Zn19 Zn2+ 3 a 0 0 0.23077 . 1. 0
Zn20 Zn2+ 3 a 0.3333 0.6667 0.24359 . 1. 0
Zn21 Zn2+ 3 a 0.6667 0.3333 0.25641 . 1. 0
Zn22 Zn2+ 3 a 0 0 0.26923 . 1. 0
Zn23 Zn2+ 3 a 0.3333 0.6667 0.28205 . 1. 0
Zn24 Zn2+ 3 a 0.6667 0.3333 0.29487 . 1. 0
Zn25 Zn2+ 3 a 0.3333 0.6667 0.30769 . 1. 0
Zn26 Zn2+ 3 a 0 0 0.32051 . 1. 0
S1 S2- 3 a 0 0 0.00962 . 1. 0
S2 S2- 3 a 0.3333 0.6667 0.02244 . 1. 0
S3 S2- 3 a 0.6667 0.3333 0.03526 . 1. 0
S4 S2- 3 a 0 0 0.04808 . 1. 0
S5 S2- 3 a 0.3333 0.6667 0.0609 . 1. 0
S6 S2- 3 a 0.6667 0.3333 0.07372 . 1. 0
S7 S2- 3 a 0 0 0.08654 . 1. 0
S8 S2- 3 a 0.3333 0.6667 0.09936 . 1. 0
S9 S2- 3 a 0.6667 0.3333 0.11218 . 1. 0
S10 S2- 3 a 0 0 0.125 . 1. 0
S11 S2- 3 a 0.6667 0.3333 0.13782 . 1. 0
S12 S2- 3 a 0.3333 0.6667 0.15064 . 1. 0
S13 S2- 3 a 0 0 0.16346 . 1. 0
S14 S2- 3 a 0.3333 0.6667 0.17628 . 1. 0
S15 S2- 3 a 0.6667 0.3333 0.1891 . 1. 0
S16 S2- 3 a 0 0 0.20192 . 1. 0
S17 S2- 3 a 0.6667 0.3333 0.21474 . 1. 0
S18 S2- 3 a 0.3333 0.6667 0.22756 . 1. 0
S19 S2- 3 a 0 0 0.24038 . 1. 0
S20 S2- 3 a 0.3333 0.6667 0.25321 . 1. 0
S21 S2- 3 a 0.6667 0.3333 0.26603 . 1. 0
S22 S2- 3 a 0 0 0.27885 . 1. 0
S23 S2- 3 a 0.3333 0.66677 0.29167 . 1. 0
S24 S2- 3 a 0.6667 0.3333 0.30449 . 1. 0
S25 S2- 3 a 0.3333 0.6667 0.31731 . 1. 0
S26 S2- 3 a 0 0 0.33013 . 1. 0
#End of TTdata_42794-ICSD
```
---------------------------------------------------------------------------------------------------------
---Sc8NiCl12_424477----------------------------------------------------------------------------------
```
#(C) 2019 by FIZ Karlsruhe - Leibniz Institute for Information Infrastructure. All rights reserved.
data_424477-ICSD
_database_code_ICSD 424477
_audit_creation_date 2013-02-01
_chemical_name_systematic 'Nickel hexascandium dodecachloride scandium'
_chemical_formula_structural '(Ni Sc6) Cl12 Sc'
_chemical_formula_sum 'Cl12 Ni1 Sc7'
_chemical_name_structure_type NaLa6OsI12-Ca0.65Pr0.35Pr6CoI12
_exptl_crystal_density_diffrn 2.91
_cell_measurement_temperature 293.
_publ_section_title
;
The prolific {Z R6} X12 R and {Z R6} X10 structure types with isolated
endohedrally stabilized (Z) rare-earth metal (R) cluster halide (X) complexes
;
loop_
_citation_id
_citation_journal_full
_citation_year
_citation_journal_volume
_citation_page_first
_citation_page_last
_citation_journal_id_ASTM
primary 'Zeitschrift fuer Anorganische und Allgemeine Chemie (1950) (DE)' 2012
638 1922 1931 ZAACAB
loop_
_publ_author_name
'Rustige, C.'
'Bruehmann, M.'
'Steinberg, S.'
'Meyer, E.'
'Daub, K.'
'Zimmermann, S.'
'Wolberg, M.'
'Mudring, A.V.'
'Meyer, G.'
_cell_length_a 13.172(2)
_cell_length_b 13.172(2)
_cell_length_c 9.0896(19)
_cell_angle_alpha 90.
_cell_angle_beta 90.
_cell_angle_gamma 120.
_cell_volume 1365.77
_cell_formula_units_Z 3
_symmetry_space_group_name_H-M 'R -3 H'
_symmetry_Int_Tables_number 148
_refine_ls_R_factor_all 0.0501
loop_
_symmetry_equiv_pos_site_id
_symmetry_equiv_pos_as_xyz
1 'x-y, x, -z'
2 'y, -x+y, -z'
3 '-x, -y, -z'
4 '-x+y, -x, z'
5 '-y, x-y, z'
6 'x, y, z'
7 'x-y+2/3, x+1/3, -z+1/3'
8 'y+2/3, -x+y+1/3, -z+1/3'
9 '-x+2/3, -y+1/3, -z+1/3'
10 '-x+y+2/3, -x+1/3, z+1/3'
11 '-y+2/3, x-y+1/3, z+1/3'
12 'x+2/3, y+1/3, z+1/3'
13 'x-y+1/3, x+2/3, -z+2/3'
14 'y+1/3, -x+y+2/3, -z+2/3'
15 '-x+1/3, -y+2/3, -z+2/3'
16 '-x+y+1/3, -x+2/3, z+2/3'
17 '-y+1/3, x-y+2/3, z+2/3'
18 'x+1/3, y+2/3, z+2/3'
loop_
_atom_type_symbol
_atom_type_oxidation_number
Ni0+ 0
Cl0+ 0
Sc0+ 0
loop_
_atom_site_label
_atom_site_type_symbol
_atom_site_symmetry_multiplicity
_atom_site_Wyckoff_symbol
_atom_site_fract_x
_atom_site_fract_y
_atom_site_fract_z
_atom_site_B_iso_or_equiv
_atom_site_occupancy
_atom_site_attached_hydrogens
Ni1 Ni0+ 3 b 0. 0. 0.5 0.0127(4) 1 0
Cl1 Cl0+ 18 f -0.31170(11) -0.08262(11) 0.49115(15) 0.0169(4) 1 0
Cl2 Cl0+ 18 f -0.48915(11) -0.20474(10) 0.83376(12) 0.0148(4) 1 0
Sc1 Sc0+ 18 f -0.12308(8) 0.04361(8) 0.65125(10) 0.0115(3) 1 0
Sc2 Sc0+ 3 a -0.6667 -0.3333 0.6667 0.0176(6) 1 0
loop_
_atom_site_aniso_label
_atom_site_aniso_type_symbol
_atom_site_aniso_U_11
_atom_site_aniso_U_22
_atom_site_aniso_U_33
_atom_site_aniso_U_12
_atom_site_aniso_U_13
_atom_site_aniso_U_23
Ni1 Ni0+ 0.0104(6) 0.0104(6) 0.0173(8) 0.0052(3) 0. 0.
Cl1 Cl0+ 0.0093(6) 0.0186(7) 0.0227(7) 0.0069(5) -0.0017(4) -0.0060(5)
Cl2 Cl0+ 0.0114(7) 0.0118(7) 0.0197(6) 0.0046(5) 0.0016(5) 0.0017(4)
Sc1 Sc0+ 0.0095(5) 0.0093(5) 0.0163(5) 0.0052(5) 0.0004(4) 0.0000(4)
Sc2 Sc0+ 0.0143(8) 0.0143(8) 0.0243(13) 0.0071(4) 0. 0.
#End of TTdata_424477-ICSD
```
---------------------------------------------------------------------------------------------------------
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2175/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2175/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2176
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2176/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2176/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2176/events
|
https://github.com/materialsproject/pymatgen/issues/2176
| 925,001,239
|
MDU6SXNzdWU5MjUwMDEyMzk=
| 2,176
|
BabelMolAdaptor in pymatgen.io.babel: Converted openbabel_mol is different from pymatgen_mol
|
{
"login": "Hyunseung-Kim",
"id": 31006321,
"node_id": "MDQ6VXNlcjMxMDA2MzIx",
"avatar_url": "https://avatars.githubusercontent.com/u/31006321?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Hyunseung-Kim",
"html_url": "https://github.com/Hyunseung-Kim",
"followers_url": "https://api.github.com/users/Hyunseung-Kim/followers",
"following_url": "https://api.github.com/users/Hyunseung-Kim/following{/other_user}",
"gists_url": "https://api.github.com/users/Hyunseung-Kim/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Hyunseung-Kim/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Hyunseung-Kim/subscriptions",
"organizations_url": "https://api.github.com/users/Hyunseung-Kim/orgs",
"repos_url": "https://api.github.com/users/Hyunseung-Kim/repos",
"events_url": "https://api.github.com/users/Hyunseung-Kim/events{/privacy}",
"received_events_url": "https://api.github.com/users/Hyunseung-Kim/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Babel has its own bonding algorithm. You would use this to get what Babel thinks should be bonded, so it's not surprising that it disagrees with what you put in. If you put in the resulting babel bonded-molecule and it changed again, then there would be a bug. ",
"Your answer was helpful for me. Thank you, shyamd."
] | 2021-06-18T15:15:45
| 2021-06-25T16:54:11
|
2021-06-25T16:54:00Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hello. I'm a newbie of pymatgen.
BabelMolAdaptor(mol) seems that a bridge between pymatgen_mol and openbabel_mol.
But if I try to convert pymatgen_mol to openbabel_mol with the following codes:
`openbabel_mol = BabelMolAdaptor(pymatgen_mol).openbabel_mol`
the number of bonds in openbabel_mol is different from pymatgen_mol. Why it is different from each other?
Here is the failed example for pymatgen_molGraph:
```
Full Formula (H3 C3 O3 F1)
Reduced Formula: H3C3O3F
Charge = -1, Spin Mult = 2
Sites (10)
0 C 0.154383 0.035399 0.408406
1 C 1.574747 0.156457 0.801702
2 O 2.256278 -1.093528 0.687596
3 C 3.193129 -1.246086 -0.372539
4 O 3.417788 -0.253020 -1.084838
5 O 3.667329 -2.391423 -0.414259
6 H -0.628536 -0.403324 1.019965
7 F -0.082731 -0.123535 -0.918526
8 H 1.609127 0.462383 1.852347
9 H 2.083332 0.903659 0.188073
from to to_image weight
---- ---- ------------ ------------------
0 1 (0, 0, 0) 0.000e+00
0 6 (0, 0, 0) 0.000e+00
0 7 (0, 0, 0) 0.000e+00
1 2 (0, 0, 0) 0.000e+00
1 8 (0, 0, 0) 0.000e+00
1 9 (0, 0, 0) 0.000e+00
2 3 (0, 0, 0) 0.000e+00
3 4 (0, 0, 0) 0.000e+00
3 5 (0, 0, 0) 0.000e+00
4 9 (0, 0, 0) 0.000e+00
```
The pymatgen_molGraph has 10 number of bonds, however, openbabel_mol has 9 number of bonds.
The bond [atom4(O) - atom9(H)] is not in openbabel_mol.
Here I attach the test code, easy to copy (Some parts of the code is borrowed from BonDNet, https://github.com/mjwen/bondnet)
```
from pymatgen.analysis.graphs import MoleculeGraph, MolGraphSplitError
from pymatgen import Molecule
from pymatgen.io.babel import BabelMolAdaptor
from openbabel import openbabel as ob
import numpy as np
def pymatgen_2_babel_atom_idx_map(pmg_mol, ob_mol):
"""
borrowed from: BonDNet (https://github.com/mjwen/bondnet)
"""
"""
Create an atom index mapping between pymatgen mol and openbabel mol.
This does not require pymatgen mol and ob mol has the same number of atoms.
But ob_mol can have smaller number of atoms.
Args:
pmg_mol (pymatgen.Molecule): pymatgen molecule
ob_mol (ob.Mol): openbabel molecule
Returns:
dict: with atom index in pymatgen mol as key and atom index in babel mol as
value. Value is `None` if there is not corresponding atom in babel.
"""
pmg_coords = pmg_mol.cart_coords
ob_coords = [[a.GetX(), a.GetY(), a.GetZ()] for a in ob.OBMolAtomIter(ob_mol)]
ob_index = [a.GetIdx() for a in ob.OBMolAtomIter(ob_mol)]
mapping = {i: None for i in range(len(pmg_coords))}
for idx, oc in zip(ob_index, ob_coords):
for i, gc in enumerate(pmg_coords):
if np.allclose(oc, gc):
mapping[i] = idx
break
else:
raise RuntimeError("Cannot create atom index mapping pymatgen and ob mols")
return mapping
pymatgen_mol = Molecule(species=['C', 'C', 'O', 'C', 'O', 'O', 'H', 'F', 'H', 'H'], coords=[[0.1543828012, 0.03539906, 0.40840565500000003], [1.5747466355, 0.156456812, 0.8017021181], [2.2562777668000003, -1.093528376, 0.6875961097000001], [3.1931291901, -1.2460856975, -0.3725393443], [3.4177877228, -0.2530196865, -1.0848382041], [3.6673293558, -2.3914227018, -0.4142593592], [-0.6285355493, -0.4033244492, 1.0199650487], [-0.0827305468, -0.1235353602, -0.9185260584], [1.6091269596000002, 0.4623828547, 1.8523474326], [2.0833316644, 0.9036585446000001, 0.1880726019]], charge=-1, spin_multiplicity=2)
bonds = {i: None for i in tuple(map(tuple, [[0, 7], [0, 1], [0, 6], [1, 9], [1, 2], [1, 8], [2, 3], [3, 4], [3, 5], [4, 9]]))}
pymatgen_molgraph = MoleculeGraph.with_edges(pymatgen_mol, bonds)
openbabel_mol = BabelMolAdaptor(pymatgen_molgraph.molecule).openbabel_mol
ob_bond_order = {}
for bd in ob.OBMolBondIter(openbabel_mol):
k = tuple(sorted([bd.GetBeginAtomIdx(), bd.GetEndAtomIdx()]))
v = bd.GetBondOrder()
ob_bond_order[k] = v
atom_idx_mapping = pymatgen_2_babel_atom_idx_map(pymatgen_mol, openbabel_mol)
print("atom_idx_mapping (key=pymatgen_mol_atom_idx: value=:openbabel_mol_atom_idx)", atom_idx_mapping)
print("pymatgen_bonds:", len(bonds), "ea.", bonds)
print("openbabel_bonds:", len(ob_bond_order), "ea.", ob_bond_order)
```
And, the following is the results:
```
atom_idx_mapping (key=pymatgen_mol_atom_idx: value=:openbabel_mol_atom_idx) {0: 1, 1: 2, 2: 3, 3: 4, 4: 5, 5: 6, 6: 7, 7: 8, 8: 9, 9: 10}
pymatgen_bonds: 10 ea. {(0, 7): None, (0, 1): None, (0, 6): None, (1, 9): None, (1, 2): None, (1, 8): None, (2, 3): None, (3, 4): None, (3, 5): None, (4, 9): None}
openbabel_bonds: 9 ea. {(4, 5): 2, (1, 8): 1, (4, 6): 1, (3, 4): 1, (2, 10): 1, (1, 2): 1, (1, 7): 1, (2, 3): 1, (2, 9): 1}
```
|
{
"login": "Hyunseung-Kim",
"id": 31006321,
"node_id": "MDQ6VXNlcjMxMDA2MzIx",
"avatar_url": "https://avatars.githubusercontent.com/u/31006321?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Hyunseung-Kim",
"html_url": "https://github.com/Hyunseung-Kim",
"followers_url": "https://api.github.com/users/Hyunseung-Kim/followers",
"following_url": "https://api.github.com/users/Hyunseung-Kim/following{/other_user}",
"gists_url": "https://api.github.com/users/Hyunseung-Kim/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Hyunseung-Kim/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Hyunseung-Kim/subscriptions",
"organizations_url": "https://api.github.com/users/Hyunseung-Kim/orgs",
"repos_url": "https://api.github.com/users/Hyunseung-Kim/repos",
"events_url": "https://api.github.com/users/Hyunseung-Kim/events{/privacy}",
"received_events_url": "https://api.github.com/users/Hyunseung-Kim/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2176/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2176/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2177
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2177/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2177/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2177/events
|
https://github.com/materialsproject/pymatgen/pull/2177
| 925,357,915
|
MDExOlB1bGxSZXF1ZXN0NjczODUzMjU5
| 2,177
|
Pourbaix: default to filter_solids=True
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I think it was more intended to include single entries that one might want to inspect and use for analysis post serial/deserialization in the multi-entry case, but I think it's probably overly complex and misnamed, so I'm in support of getting rid of it.",
"Thanks @rkingsbury, @shyamd, including discussion in https://github.com/materialsproject/pymatgen/commit/e8028b0cfe783f8628c9090da07ff6e435244f6c#commitcomment-52383083"
] | 2021-06-19T09:39:59
| 2021-06-20T23:13:58
|
2021-06-20T23:12:59Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Default `PourbaixDiagram` to use `filter_solids=True` for consistency with the MP website and discussion in #2173.
In addition:
* Fix a bug in `MaterialsProjectAqueousCompatibility` introduced by e8028b0cfe783f8628c9090da07ff6e435244f6c
* add type hinting to `PourbaixDiagram` and make a few type hinting fixes to `Entry` classes (necessary to make mypy happy on `PourbaixDiagram`).
* Deprecate the `include_unprocessed_entries` kwarg to `PourbaixDiagram.as_dict`. This was necessary in order to make serialization work properly after switching to `filter_solids=True` by default. To me, it appears this kwarg was a sort of workaround to allow a user to effectively decide whether to filter solids at the serialization / deserialization stage. In my mind, it makes more sense for the user to make that decision via the `filter_solids` kwarg when the diagram is constructed, then serialize the `filter_solids` kwarg along with all unprocessed entries always. I feel this is more consistent with the way I "expect" serialization to work. @montoyjh please comment if you have any concerns or had a specific reason for keeping this is a kwarg to `as_dict`
Closes #2173
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2177/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2177/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2177",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2177",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2177.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2177.patch",
"merged_at": "2021-06-20T23:12:59Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2178
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2178/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2178/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2178/events
|
https://github.com/materialsproject/pymatgen/pull/2178
| 925,515,934
|
MDExOlB1bGxSZXF1ZXN0NjczOTcxMjc3
| 2,178
|
QE IO: allow to distinguish same element with different oxidation state
|
{
"login": "FCMeng",
"id": 82957832,
"node_id": "MDQ6VXNlcjgyOTU3ODMy",
"avatar_url": "https://avatars.githubusercontent.com/u/82957832?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/FCMeng",
"html_url": "https://github.com/FCMeng",
"followers_url": "https://api.github.com/users/FCMeng/followers",
"following_url": "https://api.github.com/users/FCMeng/following{/other_user}",
"gists_url": "https://api.github.com/users/FCMeng/gists{/gist_id}",
"starred_url": "https://api.github.com/users/FCMeng/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/FCMeng/subscriptions",
"organizations_url": "https://api.github.com/users/FCMeng/orgs",
"repos_url": "https://api.github.com/users/FCMeng/repos",
"events_url": "https://api.github.com/users/FCMeng/events{/privacy}",
"received_events_url": "https://api.github.com/users/FCMeng/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Can you add a simple unittest pls?\r\n",
"> Can you add a simple unittest pls?\r\n@shyuep I uploaded a simple unittest here. Thanks!\r\n\r\n",
"Hi there, you do not need to send it as a zip file. All you need to do is to edit pymatgen/io/tests/test_pwscf.py to add this test to it. Then the checks will automatically run to make sure that the code is running as it should.",
"> Hi there, you do not need to send it as a zip file. All you need to do is to edit pymatgen/io/tests/test_pwscf.py to add this test to it. Then the checks will automatically run to make sure that the code is running as it should.\r\n\r\n@shyuep Thank you, Dr. Ong. I have committed a modified test_pwscf.py file, where there are three tests for the oxidation states: one with fully oxidation(like Li+, Li+, O2- in the test), second without any oxidation (like Li, Li, O in the test), and the third with a mixed oxidation (like Li, Li+, O in the test).",
"\n[](https://coveralls.io/builds/42080597)\n\nCoverage decreased (-0.6%) to 83.133% when pulling **4b26d06c8fbc38be37aa14e7025d831df0d92754 on FCMeng:master** into **ef6c0c56a214334fd0f1af26f10b570f3bad1565 on materialsproject:master**.\n",
"Thanks @FCMeng, merging because the listing issue is not related to this PR and otherwise this looks good!"
] | 2021-06-20T04:12:00
| 2021-08-11T22:28:59
|
2021-08-11T22:28:59Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Allow to distinguish same element with different oxidation state, which might have different pseudopotentials.
## Summary
The only change I made is at line 153 of pwscf.py file:
out.append(" %s %.6f %.6f %.6f" % (site.specie.symbol, site.a, site.b, site.c))
-> out.append(" %s %.6f %.6f %.6f" % (site.specie, site.a, site.b, site.c))
For example, if in the structure, there is a "Ti+" state, it will be exported as "Ti" if we use site.specie.symbol to export the pwscf input file. In real QE calculations, it is possible that "Ti" and "Ti+" elements have different pseudopotentials.
By making the change from site.specie.symbol to site.specie, the oxidization state information is preserved.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2178/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2178/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2178",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2178",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2178.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2178.patch",
"merged_at": "2021-08-11T22:28:59Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2179
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2179/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2179/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2179/events
|
https://github.com/materialsproject/pymatgen/pull/2179
| 926,824,218
|
MDExOlB1bGxSZXF1ZXN0Njc1MDcxMTg5
| 2,179
|
enforce KSPACING cap in MPScanRelaxSet
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/40766573)\n\nCoverage decreased (-0.6%) to 83.149% when pulling **9f113be5cfe4156aefe0964dd81f9ed6e0c4adb5 on rkingsbury:kspacing-guard** into **33844780c5f735a18d6d28d788f5fc22f208b646 on materialsproject:master**.\n"
] | 2021-06-22T04:44:45
| 2021-07-15T14:32:57
|
2021-06-22T16:03:14Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Add a guard statement and a test to ensure that the KSPACING value calculated by `MPScanRelaxSet` never exceeds the range 0.22 - 0.44.
This is a follow up to #2163. After that PR was merged I noticed from our atomate tests of the SCAN workflow that if a material's bandgap was sufficiently large >9 eV or so, it could actually reverse the sign of KSPACING, leading to large negative values that broke the logic. This PR fixes that bug and will be needed to keep atomate tests passing.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2179/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2179/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2179",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2179",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2179.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2179.patch",
"merged_at": "2021-06-22T16:03:13Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2180
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2180/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2180/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2180/events
|
https://github.com/materialsproject/pymatgen/issues/2180
| 927,683,522
|
MDU6SXNzdWU5Mjc2ODM1MjI=
| 2,180
|
Convert XYZ to cif format
|
{
"login": "VictorPulzz",
"id": 23406402,
"node_id": "MDQ6VXNlcjIzNDA2NDAy",
"avatar_url": "https://avatars.githubusercontent.com/u/23406402?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/VictorPulzz",
"html_url": "https://github.com/VictorPulzz",
"followers_url": "https://api.github.com/users/VictorPulzz/followers",
"following_url": "https://api.github.com/users/VictorPulzz/following{/other_user}",
"gists_url": "https://api.github.com/users/VictorPulzz/gists{/gist_id}",
"starred_url": "https://api.github.com/users/VictorPulzz/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/VictorPulzz/subscriptions",
"organizations_url": "https://api.github.com/users/VictorPulzz/orgs",
"repos_url": "https://api.github.com/users/VictorPulzz/repos",
"events_url": "https://api.github.com/users/VictorPulzz/events{/privacy}",
"received_events_url": "https://api.github.com/users/VictorPulzz/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @VictorPullzz, since this is not a bug, you would be better off asking for help at our discussion forum: matsci.org/pymatgen\r\n\r\nIn brief, the CIF format is a crystallographic format, meaning it also defines a periodic lattice. XYZ files typically describe molecules, although some have a header with lattice information. To convert to CIF, import the XYZ file (e.g. using `Molecule`), construct a `Structure` object with the appropriate `Lattice`, and then use `Structure.to(filename=\"your_file.cif\")` to export.",
"@mkhorton Hello thanks for you anwser, please can you send me example, how I can do it )))"
] | 2021-06-22T22:31:31
| 2021-06-23T20:36:33
|
2021-06-23T19:16:07Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Hello I have a question. How I can convert XYZ file to cif format?
For ex:
I have some XYZ file
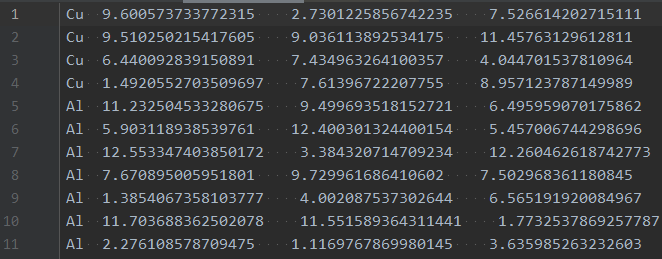
I need convert this to cif format file.
How I can doing this?
Pelase help me)
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2180/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2180/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2181
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2181/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2181/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2181/events
|
https://github.com/materialsproject/pymatgen/issues/2181
| 929,240,783
|
MDU6SXNzdWU5MjkyNDA3ODM=
| 2,181
|
checking whether an entry is stable makes getting `e_above_hull` O(N) as opposed to O(1)
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks for reporting this. Can you provide an actual profile code showing this is indeed the offending line? I would assume a quick search through stable entries would be faster than doing the actual decomp computation for stable entries. Of course, if you have 100000 unstable entries and only 10 stable entries, that may result in a lot of unnecessary checks.\r\n\r\nA few things we can do:\r\n1. Swap the order of the entry `__eq__` checks - I believe checking entry_id first might be faster than checking energy or composition since that is usually just a string. - I just pushed a change to implement this.\r\n2. We can add a boolean parameter called `skip_stable_check` to bypass that if statement. It is up to the user to then make sure they do the checks themselves.\r\n\r\n@rkingsbury @mkhorton Pls comment.",
"I am currently refactoring code to try before releasing the source and so my workflow is currently broken but I will post come profiling results once I have it working again. \r\n\r\nThe workflow was looking at the stabilities of randomly generated structures (initial screening library ~400k) against a patched phase diagram of MP + the MP compatible data from https://archive.materialscloud.org/record/2021.68 ~320k entries.",
"@rkingsbury One thing to consider is whether we want ot make the entry id check *exclusionary* rather than inclusionary,. Right now, the check says that if the entry ids are equal, we set to True. If not, we check composition and energy. Given that most entries are UNEQUAL, we can say that if entry ids exist and are not equal, we return False without checking composition or energy. I believe that will make the entry check basically many times faster.",
"@shyuep on this front what would your opinion be on adding entry_ids to `PDEntry` and it's children and removing reference to the class in the `__hash__` for entry type objects? \r\n\r\nThis would allow another hack(?) i've done using `PDEntry` in the pd construction but computing the properties of `ComputedStructureEntries` from the second data set to become standard. Using `PDEntry` helps a fair amount with speed as returning the energy becomes a look-up rather than a function call with the corrections.",
"I would generally avoid adding entry ids to PDEntry. I prefer to keep objects to their *minimum required specification*. When you add something like entry ids to an object, you are imposing a requirement for the user to properly designate such ids. If someone were to say, just set `entry_id = 1` for all the entries he creates, that would result in wrong behavior. If you were going to just assign a unique number to each PDENtry you create, that would be no different from the `object._id` that comes part and parcel of every Python object. In essence, you are replicating default functionality already present.\r\n\r\nFor computed entries, this is more justifiable since we have things like MP that assigns unique ids.",
"@CompRhys I had a thought that if you are going to generate a lot of entries from your own vaspruns, it would be helpful if a default entry_id is added to get_computed_entry. I have just pushed a change that implements this. What this means is that all computed entries generated from your own Vaspruns would now have a entry id \"vasprun-{datetime that it was parsed}\". This would at the very least ensure that almost all ComputedEntries have a valid entry_id, which would speed up the checks a lot.",
"I will defer to you guys on this one; this is beyond the extent of my software engineering knowledge.",
"@shyuep have fixed the workflow and an profiling now. Removing the check getting e_above_hull for all ternaries and below from my `PatchedPhaseDiagram` takes ~ 97 seconds whilst with the check it will take ~60 hours. This is very extreme as the `stable_entries` of the `PatchedPhaseDiagram` covers all the patches not just the individual patch (i.e. `PhaseDiagram`) being considered in each chemical system.\r\n\r\nedit: for clarity the `PatchedPhaseDiagram` is essentially a ease of use tool to calculate stabilities over the whole of MP and new materials. It is a dictionary of {chemsys: `PhaseDiagram`} for all the non-overlapping chemical systems. If you have an entry not in any of the chemical systems the decomposition is calculated using the linear algebra code introduced for `get_phase_separation_energy`. ",
"Thanks. Can you paste the output of the profiling here? I am curious which function call is causing that speed difference.",
"Sure, because I don't want to wait 60 hours with check is just for 100 entries whilst without check is for all 84944 tertiary or smaller systems.\r\n\r\nThe vast majority of the time is spent checking the equality of `ComputedEntries`.\r\n\r\n[with_check.txt](https://github.com/materialsproject/pymatgen/files/6727035/with_check.txt)\r\n[without_check.txt](https://github.com/materialsproject/pymatgen/files/6727036/without_check.txt)\r\n",
"Did you update with the latest pymatgen yet? \r\n\r\n```python\r\nfrom pymatgen.ext.matproj import MPRester\r\nfrom pymatgen.analysis.phase_diagram import PhaseDiagram\r\n\r\nmpr = MPRester()\r\nentries = mpr.get_entries_in_chemsys([\"Li\", \"Fe\", \"P\", \"O\"])\r\npd = PhaseDiagram(entries)\r\n\r\ndef main():\r\n return [pd.get_decomp_and_e_above_hull(e) for e in entries]\r\n\r\nif __name__ == \"__main__\":\r\n import cProfile\r\n import os\r\n import pstats\r\n cProfile.run('main()', 'profile_stats')\r\n p = pstats.Stats('profile_stats')\r\n p.sort_stats('cumulative').print_stats(20)\r\n os.remove(\"profile_stats\")\r\n```\r\n\r\nThis is my profiling code. The output is:\r\n\r\n```\r\n 664439 function calls (658491 primitive calls) in 0.325 seconds\r\n\r\n Ordered by: cumulative time\r\n List reduced from 73 to 20 due to restriction <20>\r\n\r\n ncalls tottime percall cumtime percall filename:lineno(function)\r\n 1 0.000 0.000 0.326 0.326 {built-in method builtins.exec}\r\n 1 0.000 0.000 0.326 0.326 <string>:1(<module>)\r\n 1 0.000 0.000 0.326 0.326 /Users/sp/Library/Mobile Documents/com~apple~CloudDocs/Downloads/py/profilepd.py:20(main)\r\n 1 0.000 0.000 0.326 0.326 /Users/sp/Library/Mobile Documents/com~apple~CloudDocs/Downloads/py/profilepd.py:21(<listcomp>)\r\n 943 0.002 0.000 0.326 0.000 /Users/sp/repos/pymatgen/pymatgen/analysis/phase_diagram.py:549(get_decomp_and_e_above_hull)\r\n 943 0.002 0.000 0.255 0.000 /Users/sp/repos/pymatgen/pymatgen/analysis/phase_diagram.py:520(get_decomposition)\r\n 879 0.009 0.000 0.223 0.000 /Users/sp/repos/pymatgen/pymatgen/analysis/phase_diagram.py:488(_get_facet_and_simplex)\r\n 28768 0.038 0.000 0.203 0.000 /Users/sp/repos/pymatgen/pymatgen/util/coord.py:417(in_simplex)\r\n 29711 0.018 0.000 0.118 0.000 /Users/sp/repos/pymatgen/pymatgen/util/coord.py:391(bary_coords)\r\n 59422 0.076 0.000 0.076 0.000 {built-in method numpy.core._multiarray_umath.implement_array_function}\r\n 2727 0.002 0.000 0.068 0.000 /Users/sp/repos/pymatgen/pymatgen/entries/__init__.py:83(energy_per_atom)\r\n 2727 0.002 0.000 0.065 0.000 /Users/sp/repos/pymatgen/pymatgen/entries/computed_entries.py:368(energy)\r\n 2727 0.003 0.000 0.063 0.000 /Users/sp/repos/pymatgen/pymatgen/entries/computed_entries.py:384(correction)\r\n 29711 0.010 0.000 0.061 0.000 <__array_function__ internals>:2(concatenate)\r\n 28768 0.007 0.000 0.051 0.000 {method 'all' of 'numpy.ndarray' objects}\r\n 28768 0.005 0.000 0.044 0.000 /Users/sp/miniconda3/envs/mavrl/lib/python3.9/site-packages/numpy/core/_methods.py:59(_all)\r\n 943 0.001 0.000 0.042 0.000 /Users/sp/repos/pymatgen/pymatgen/analysis/phase_diagram.py:572(<listcomp>)\r\n 29711 0.009 0.000 0.039 0.000 <__array_function__ internals>:2(dot)\r\n 28768 0.039 0.000 0.039 0.000 {method 'reduce' of 'numpy.ufunc' objects}\r\n 7174 0.022 0.000 0.038 0.000 /Users/sp/miniconda3/envs/mavrl/lib/python3.9/site-packages/uncertainties/core.py:633(f_with_affine_o\r\n```\r\n\r\nI enabled and disabled the check, and it made absolutely no difference in the time. ",
"Anyway, I added a check_stable option to get_decomp_and_ehull to the latest commit. If you set that to false, that should allow you to bypass the stability check.",
"Thanks, I think you're right that the difference between it being O(N) and O(1) isn't significant unless you're dealing with very large numbers of entries but happy that there's a supported work around."
] | 2021-06-24T13:38:25
| 2021-06-28T16:22:19
|
2021-06-28T14:46:32Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
https://github.com/materialsproject/pymatgen/blob/f091ddba0a82c586eca91b454ebeeafb45ac80fe/pymatgen/analysis/phase_diagram.py#L566-L569
In `PhaseDiagram.get_decomp_and_e_above_hull` the current code first checks if an entry is in the stable entries. This wastes a large amount of compute to ensure that the decomp is exactly 1 as opposed to 1.0000000000000002 which is what we get from the bary_coord calculation for stable entries.
Removing this check speeds up my workflow by several hours and I think it would be sensible to remove it from the dev branch also.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2181/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2181/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2182
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2182/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2182/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2182/events
|
https://github.com/materialsproject/pymatgen/pull/2182
| 929,432,531
|
MDExOlB1bGxSZXF1ZXN0Njc3Mjg1MzEx
| 2,182
|
fix: update matplotlib 3daxes to use modern api.
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/40858948)\n\nCoverage decreased (-0.6%) to 83.149% when pulling **5420e901364090346f3174e7125ba155eb568bc2 on CompRhys:3daxes** into **f4d2b6a53061074ccebba5bf0e75afa9fed15281 on materialsproject:master**.\n"
] | 2021-06-24T17:00:45
| 2021-06-28T12:42:40
|
2021-06-28T12:41:46Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
matplotlib is deprecating the old way to setup 3d plots used by the "matplotlib" backend in PDPlotter. This was giving me deprecation warnings in the testing scripts so I've replaced the calls with the more up to date api usage. I have tested the backend and it still works as before for 4d gibbs simplicies.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2182/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2182/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2182",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2182",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2182.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2182.patch",
"merged_at": "2021-06-28T12:41:46Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2183
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2183/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2183/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2183/events
|
https://github.com/materialsproject/pymatgen/issues/2183
| 929,679,648
|
MDU6SXNzdWU5Mjk2Nzk2NDg=
| 2,183
|
Inconsistency between function definition and documentation
|
{
"login": "gVallverdu",
"id": 5417018,
"node_id": "MDQ6VXNlcjU0MTcwMTg=",
"avatar_url": "https://avatars.githubusercontent.com/u/5417018?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/gVallverdu",
"html_url": "https://github.com/gVallverdu",
"followers_url": "https://api.github.com/users/gVallverdu/followers",
"following_url": "https://api.github.com/users/gVallverdu/following{/other_user}",
"gists_url": "https://api.github.com/users/gVallverdu/gists{/gist_id}",
"starred_url": "https://api.github.com/users/gVallverdu/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/gVallverdu/subscriptions",
"organizations_url": "https://api.github.com/users/gVallverdu/orgs",
"repos_url": "https://api.github.com/users/gVallverdu/repos",
"events_url": "https://api.github.com/users/gVallverdu/events{/privacy}",
"received_events_url": "https://api.github.com/users/gVallverdu/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @gVallverdu, fixed with https://github.com/materialsproject/pymatgen/commit/8c96e7a395a2c5894d9d7d6d3094873b7e172741"
] | 2021-06-24T23:06:01
| 2021-06-24T23:37:42
|
2021-06-24T23:37:42Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hi,
There is an inconsistency between the definition of the `append()` function of the `Molecule` class and the documentation.
The `validate_proximity` is set to `False` by default but in the docstring of the function it is said that the default value is `True`.
Here is the link to the `append()` method in the `pymatgen.core.structure.Molecule` class:
https://github.com/materialsproject/pymatgen/blob/b789d74639aa851d7e5ee427a765d9fd5a8d1079/pymatgen/core/structure.py#L3861-L3887
I am using the version 2022.0.5.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2183/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2183/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2184
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2184/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2184/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2184/events
|
https://github.com/materialsproject/pymatgen/issues/2184
| 931,466,729
|
MDU6SXNzdWU5MzE0NjY3Mjk=
| 2,184
|
Lattice object should not be hashable
|
{
"login": "Develpy",
"id": 54062404,
"node_id": "MDQ6VXNlcjU0MDYyNDA0",
"avatar_url": "https://avatars.githubusercontent.com/u/54062404?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Develpy",
"html_url": "https://github.com/Develpy",
"followers_url": "https://api.github.com/users/Develpy/followers",
"following_url": "https://api.github.com/users/Develpy/following{/other_user}",
"gists_url": "https://api.github.com/users/Develpy/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Develpy/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Develpy/subscriptions",
"organizations_url": "https://api.github.com/users/Develpy/orgs",
"repos_url": "https://api.github.com/users/Develpy/repos",
"events_url": "https://api.github.com/users/Develpy/events{/privacy}",
"received_events_url": "https://api.github.com/users/Develpy/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Strictly speaking, yes. Practically, this is normal practice and sometimes, there are cases where you want to hash a Lattice or some other mutable object in a dict. The default Python Object is hashable. "
] | 2021-06-28T11:24:14
| 2021-06-28T12:41:32
|
2021-06-28T12:41:32Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
The lattice object (pymatgen/pymatgen/core/lattice.py) implements the hash-function as follows:
```
def __hash__(self):
return 7
```
I think that this is not a good way to implement the hash-function and that lattice should not be hashable because it is mutable.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2184/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2184/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2185
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2185/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2185/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2185/events
|
https://github.com/materialsproject/pymatgen/issues/2185
| 932,225,519
|
MDU6SXNzdWU5MzIyMjU1MTk=
| 2,185
|
eigenvalue_band_properties based on separate spin channels
|
{
"login": "Andrew-S-Rosen",
"id": 8674072,
"node_id": "MDQ6VXNlcjg2NzQwNzI=",
"avatar_url": "https://avatars.githubusercontent.com/u/8674072?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Andrew-S-Rosen",
"html_url": "https://github.com/Andrew-S-Rosen",
"followers_url": "https://api.github.com/users/Andrew-S-Rosen/followers",
"following_url": "https://api.github.com/users/Andrew-S-Rosen/following{/other_user}",
"gists_url": "https://api.github.com/users/Andrew-S-Rosen/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Andrew-S-Rosen/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Andrew-S-Rosen/subscriptions",
"organizations_url": "https://api.github.com/users/Andrew-S-Rosen/orgs",
"repos_url": "https://api.github.com/users/Andrew-S-Rosen/repos",
"events_url": "https://api.github.com/users/Andrew-S-Rosen/events{/privacy}",
"received_events_url": "https://api.github.com/users/Andrew-S-Rosen/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Addressed in #2187."
] | 2021-06-29T05:24:31
| 2021-07-01T04:45:31
|
2021-07-01T04:45:31Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Is your feature request related to a problem? Please describe.**
Currently, the band gaps computed using the `eigenvalue_band_properties` function in `pymatgen.io.vasp.outputs` (e.g. for the `Vasprun` or `Eigenval` classes) are based on the VBM and CBM independent of spin. For some spin-polarized systems, this might result in a band gap that is formally a spin-forbidden transition, which may not be the gap of interest depending on the use-case. Additionally, it is not currently possible to quickly determine if a material might be something like a [half-metal](https://en.wikipedia.org/wiki/Half-metal), where one spin channel has metallic behavior and the other does not. [Disclaimer: I'm by no means an expert on this topic, so apologies if I've butchered it.]
As an example, see the DOS below. Band gap 1 (and its corresponding CBM and VBM) is the current value computed in Pymatgen. Band gap 2, however, may be a desirable property to compute since there is no spin transition. Band gap 3, while larger than band gap 2, may also be of interest. Currently, there is no way to compute band gap 2 or band gap 3.

**Describe the solution you'd like**
I propose to add a flag `separate_spins` that, when set to False (default), does not change the behavior of the eigenvalue band properties. However, when `separate_spins` is set to True, the returned tuple will contain properties for the spin-up and spin-down channels separately. The user would most likely be interested in the minimum value of band gap 2 and 3, but they can take the minimum on their own if desired.
**Describe alternatives you've considered**
Nothing major to note.
**Additional context**
Here is an example of how the band gap can currently be computed.
```python
from pymatgen.io.vasp.outputs import Eigenval
eig = Eigenval('EIGENVAL') # I am proposing to add a new keyword argument here
bg = eig.eigenvalue_band_properties[0]
print(bg)
```
**Note**
I am currently working on the PR. Hopefully I can submit it soon if you think this is desirable.
|
{
"login": "Andrew-S-Rosen",
"id": 8674072,
"node_id": "MDQ6VXNlcjg2NzQwNzI=",
"avatar_url": "https://avatars.githubusercontent.com/u/8674072?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Andrew-S-Rosen",
"html_url": "https://github.com/Andrew-S-Rosen",
"followers_url": "https://api.github.com/users/Andrew-S-Rosen/followers",
"following_url": "https://api.github.com/users/Andrew-S-Rosen/following{/other_user}",
"gists_url": "https://api.github.com/users/Andrew-S-Rosen/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Andrew-S-Rosen/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Andrew-S-Rosen/subscriptions",
"organizations_url": "https://api.github.com/users/Andrew-S-Rosen/orgs",
"repos_url": "https://api.github.com/users/Andrew-S-Rosen/repos",
"events_url": "https://api.github.com/users/Andrew-S-Rosen/events{/privacy}",
"received_events_url": "https://api.github.com/users/Andrew-S-Rosen/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2185/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2185/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2186
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2186/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2186/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2186/events
|
https://github.com/materialsproject/pymatgen/pull/2186
| 932,274,036
|
MDExOlB1bGxSZXF1ZXN0Njc5NjQyMjQ4
| 2,186
|
Revise #2164 - Make SETTINGS loading more robust
|
{
"login": "ardunn",
"id": 19936203,
"node_id": "MDQ6VXNlcjE5OTM2MjAz",
"avatar_url": "https://avatars.githubusercontent.com/u/19936203?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ardunn",
"html_url": "https://github.com/ardunn",
"followers_url": "https://api.github.com/users/ardunn/followers",
"following_url": "https://api.github.com/users/ardunn/following{/other_user}",
"gists_url": "https://api.github.com/users/ardunn/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ardunn/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ardunn/subscriptions",
"organizations_url": "https://api.github.com/users/ardunn/orgs",
"repos_url": "https://api.github.com/users/ardunn/repos",
"events_url": "https://api.github.com/users/ardunn/events{/privacy}",
"received_events_url": "https://api.github.com/users/ardunn/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/40957332)\n\nCoverage decreased (-0.6%) to 83.148% when pulling **be53de0347d21b5fbd55512316dee670fcd6b0e5 on ardunn:master** into **0e3181fe3bd2553267340dcb8b8afd19332387a7 on materialsproject:master**.\n",
"Thanks @ardunn!"
] | 2021-06-29T06:47:59
| 2021-06-30T18:04:51
|
2021-06-30T18:04:50Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Changes in SETTINGS load priority from #2164 can sometimes cause the loaded SETTINGS dictionary to be empty, leading to the following kind of error:
```
File "/home/runner/work/matminer/matminer/test_env/lib/python3.8/site-packages/pymatgen/ext/matproj.py", line 202, in __init__
if "MAPI_DB_VERSION" not in d:
TypeError: argument of type 'NoneType' is not iterable
```
This PR corrects that scenario
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2186/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2186/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2186",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2186",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2186.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2186.patch",
"merged_at": "2021-06-30T18:04:50Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2187
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2187/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2187/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2187/events
|
https://github.com/materialsproject/pymatgen/pull/2187
| 932,820,349
|
MDExOlB1bGxSZXF1ZXN0NjgwMTIwNjcy
| 2,187
|
Add support for spin-dependent eig band props
|
{
"login": "Andrew-S-Rosen",
"id": 8674072,
"node_id": "MDQ6VXNlcjg2NzQwNzI=",
"avatar_url": "https://avatars.githubusercontent.com/u/8674072?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Andrew-S-Rosen",
"html_url": "https://github.com/Andrew-S-Rosen",
"followers_url": "https://api.github.com/users/Andrew-S-Rosen/followers",
"following_url": "https://api.github.com/users/Andrew-S-Rosen/following{/other_user}",
"gists_url": "https://api.github.com/users/Andrew-S-Rosen/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Andrew-S-Rosen/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Andrew-S-Rosen/subscriptions",
"organizations_url": "https://api.github.com/users/Andrew-S-Rosen/orgs",
"repos_url": "https://api.github.com/users/Andrew-S-Rosen/repos",
"events_url": "https://api.github.com/users/Andrew-S-Rosen/events{/privacy}",
"received_events_url": "https://api.github.com/users/Andrew-S-Rosen/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/41009700)\n\nCoverage decreased (-0.6%) to 83.158% when pulling **9c748d3a87c4593b83a9e51eb6c30751d1db1884 on arosen93:master** into **0e3181fe3bd2553267340dcb8b8afd19332387a7 on materialsproject:master**.\n",
"Thanks for this PR @arosen93, it looks like a good addition to me.\r\n\r\nNote there's possibly a more serious logic bug in the current `eigenvalue_band_properties` in that I believe it's broken on metals, in that it will give a very small but non-zero gap.",
"@mkhorton -- thanks for the head's up. I think I may have observed similar behavior before.\r\n\r\nI think the PR is pretty much done. I am going to use it for a day or so more to make sure nothing unexpected pops up. I'll reach out here once I'm done with that.\r\n\r\n~Regarding the TODO I mentioned, do you know the best way for me to check if ISPIN = 2 in the Vasprun and BSVasprun classes so I can raise an error if it's not equal to 2 but `separate_spins` is called?~",
"Okay, @mkhorton -- I'm feeling good about this one whenever you wish to proceed with the PR.",
"Thanks for adding this! Have merged.\r\n\r\nWhile reviewing, I noticed we have two copies of this implementation (one for `Vasprun`, one for `Eigenval`), I'm not sure if we should simplify this at some point to have a common implementation."
] | 2021-06-29T15:19:47
| 2021-07-01T04:46:19
|
2021-07-01T04:43:47Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
PR to address #2185 (please refer to this Issue for additional context). This PR allows for the band gap, CBM, VBM, and is_direct_band_gap to be computed for each individual spin channel in a spin-polarized calculation when using the VASP Eigenval or Vasprun parsers via a new `separate_spins` keyword argument. Test cases are also included.
As an example, one can run:
```python
from pymatgen.io.vasp.outputs import Eigenval
eig = Eigenval('EIGENVAL', separate_spins=True)
bg = eig.eigenvalue_band_properties[0]
print(bg)
```
which would print out a list of length two where index 0 and index 1 are the band gaps for the up- and down-spin channels, respectively. Analogous behavior would occur if the Vasprun parser were used.
## TODO (if any)
~Setting `separate_spins=True` should raise an error if the calculation is not spin-polarized (i.e. if ISPIN != 2). I have done this already for the Eigenval class (as shown [here](https://github.com/materialsproject/pymatgen/blob/42156a757ad9a52cd28d42005f4c02c6289e6286/pymatgen/io/vasp/outputs.py#L5276)) but wasn't sure the best way to introduce this for the Vasprun and BSVasprun classes.~
**Any suggestions?**
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2187/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2187/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2187",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2187",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2187.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2187.patch",
"merged_at": "2021-07-01T04:43:47Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2188
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2188/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2188/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2188/events
|
https://github.com/materialsproject/pymatgen/issues/2188
| 933,073,477
|
MDU6SXNzdWU5MzMwNzM0Nzc=
| 2,188
|
CombinedData serialization with monty is broken
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"FYI @htz1992213 @orioncohen",
"Per a side discussion with @mkhorton , we might want to consider adding `DataFrame` support to `monty` so that we don't have to implement custom `as_dict` / `from_dict`. If we do that, just inheriting from `MSONable` and making sure that all kwargs are stored as class attributes should be enough. But currently `MSONable` doesn't support serializing `DataFrame`. We can discuss. See https://github.com/materialsproject/pymatgen/issues/2138",
"Thanks for bringing this up @rkingsbury. If I am reading this correctly, the method is throwing an error when you call `data = loadfn('some_path.json')`, right?\r\nWhat would adding `DataFrame` support to `monty` involve? Would that be as easy as importing a specialized package? \r\n\r\nI support using monty to serialize without using custom `as_dict` and `from_dict` methods. My one concern is that serialization is not very persistent if we change the underlying object code. If we want to be able to store permanently in MongoDB, it might be wise to create a more flexible schema for storage.",
"@rkingsbury Will make a patch overriding superclass method. I might set some method, e.g. `from_ff_and_topologies`, to be unsupported.",
"> \r\n> \r\n> Thanks for bringing this up @rkingsbury. If I am reading this correctly, the method is throwing an error when you call `data = loadfn('some_path.json')`, right?\r\n> What would adding `DataFrame` support to `monty` involve? Would that be as easy as importing a specialized package?\r\n\r\nI think what would be required is to add a type check to `MontyEncoder`, similar to what is currently done for datetime objects ([here](https://github.com/materialsvirtuallab/monty/blob/9cced458b657ab9aa7ee7dc39f0360c9e5aa6049/monty/json.py#L273)).\r\n\r\nI'm a little out of my depth here, but from #2138 I know that this works:\r\n```\r\ndf.to_json(default_handler=MontyEncoder().encode)\r\n```\r\n\r\nand `loadfn` uses `MontyEncoder` by default (as I understand it), so I think we would just need to add something to `loadfn` and `dumpfn` like:\r\n\r\n```\r\nif isinstance(o, pd.DataFrame):\r\n return o.to_json(default_handler=MontyEncoder().encode)\r\n```\r\n\r\n@mkhorton does that look right to you?",
"Hah, yes! I actually typed out a reply to this saying exactly that:\r\n\r\n> It would mean adding `pandas` to [this block](https://github.com/materialsvirtuallab/monty/blob/9cced458b657ab9aa7ee7dc39f0360c9e5aa6049/monty/json.py#L378) and then encoding the data frame using `df.to_json(default_handler=MontyEncoder().encode)`. In this way, any data frame can be serialized automatically provided it contains only native or MSON data types, could be stored in Mongo, etc.\r\n\r\nbut forgot to submit it 🤦🏻♂️\r\n\r\nNote that this is slightly different to your proposal just in that the code should be in `MontyEncoder`/`MontyDecoder`, rather than needing to touch `loadfn` or `dumpfn`.",
"Thanks @mkhorton ! So something like\r\n\r\n```\r\nif modname and modname not in [\"bson.objectid\", \"numpy\"]:\r\n if modname == \"pandas\" and classname == \"DataFrame\":\r\n return= d.to_json(default_handler=MontyEncoder().encode)\r\n if modname == \"pandas\" and classname == \"Series\":\r\n return= d.to_json(default_handler=MontyEncoder().encode)\r\n\r\n```\r\n?",
"Yes, I believe so, but I haven't tried -- see [here](https://github.com/materialsvirtuallab/monty/blob/9cced458b657ab9aa7ee7dc39f0360c9e5aa6049/monty/json.py#L279) too. Note that including the `@module` and `@class` is important, and then the pandas json could be under a `data` key.\r\n\r\nIt's not clear to me if `Series` is necessary, or if additional pandas data types would also be useful, but sounds reasonable to me.",
"@rkingsbury @mkhorton @orioncohen I have made a PR (https://github.com/materialsproject/pymatgen/pull/2191) for serialization using conventional encoding, but please let me know if the newer way works!",
"@rkingsbury this issue can be closed, right?",
"> @rkingsbury this issue can be closed, right?\r\n\r\nI think so? It looks like #2191 should have taken care of it, although I have not personally tried serializing the returned `dict`"
] | 2021-06-29T20:21:36
| 2023-11-05T22:19:54
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
I am unable to serialize and deserialize a `CombinedData` object using `loadfn` / `dumpfn`. I believe this occurs because `CombinedData` inherits `from_dict` from `LammpsData`, but `CombinedData.init()` takes different kwargs.
**To Reproduce**
Create a `CombinedData` object called `data`
```
from monty.serialization import loadfn, dumpfn
dumpfnn(data, 'some_path.json')
data = loadfn('some_path.json')
```
will fail with
```
------------------------------------------------------------------------
TypeError Traceback (most recent call last)
<ipython-input-26-b8f560ec17df> in <module>
1 # data = LammpsData.from_file(pipeline_dir / '2021-05_prelim_lammps_TMA-Cl' / 'md.data')
----> 2 data = loadfn(pipeline_dir / '2021-05_prelim_lammps_TMA-Cl' / 'combined_data.json')
~/miniconda3/envs/md/lib/python3.9/site-packages/monty/serialization.py in loadfn(fn, *args, **kwargs)
83 if "cls" not in kwargs:
84 kwargs["cls"] = MontyDecoder
---> 85 return json.load(fp, *args, **kwargs)
86
87 raise TypeError("Invalid format: {}".format(fmt))
~/miniconda3/envs/md/lib/python3.9/json/__init__.py in load(fp, cls, object_hook, parse_float, parse_int, parse_constant, object_pairs_hook, **kw)
291 kwarg; otherwise ``JSONDecoder`` is used.
292 """
--> 293 return loads(fp.read(),
294 cls=cls, object_hook=object_hook,
295 parse_float=parse_float, parse_int=parse_int,
~/miniconda3/envs/md/lib/python3.9/json/__init__.py in loads(s, cls, object_hook, parse_float, parse_int, parse_constant, object_pairs_hook, **kw)
357 if parse_constant is not None:
358 kw['parse_constant'] = parse_constant
--> 359 return cls(**kw).decode(s)
~/miniconda3/envs/md/lib/python3.9/site-packages/monty/json.py in decode(self, s)
381 """
382 d = json.JSONDecoder.decode(self, s)
--> 383 return self.process_decoded(d)
384
385
~/miniconda3/envs/md/lib/python3.9/site-packages/monty/json.py in process_decoded(self, d)
354 data = {k: v for k, v in d.items() if not k.startswith("@")}
355 if hasattr(cls_, "from_dict"):
--> 356 return cls_.from_dict(data)
357 elif np is not None and modname == "numpy" and classname == "array":
358 if d["dtype"].startswith("complex"):
~/miniconda3/envs/md/code/pymatgen/pymatgen/io/lammps/data.py in from_dict(cls, d)
917 topology = {k: decode_df(v) for k, v in topology.items()}
918 items["topology"] = topology
--> 919 return cls(**items)
920
921 def as_dict(self):
TypeError: __init__() got an unexpected keyword argument 'box'
```
**Expected behavior**
The `CombinedData` object should be created from the file
**Desktop (please complete the following information):**
- Windows 10 / WSL 2 Ubuntu
- Version 2022.0.9
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2188/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2188/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2189
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2189/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2189/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2189/events
|
https://github.com/materialsproject/pymatgen/issues/2189
| 935,341,982
|
MDU6SXNzdWU5MzUzNDE5ODI=
| 2,189
|
Tensorflow and Pymatgen incompatible
|
{
"login": "arunppsg",
"id": 26398753,
"node_id": "MDQ6VXNlcjI2Mzk4NzUz",
"avatar_url": "https://avatars.githubusercontent.com/u/26398753?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/arunppsg",
"html_url": "https://github.com/arunppsg",
"followers_url": "https://api.github.com/users/arunppsg/followers",
"following_url": "https://api.github.com/users/arunppsg/following{/other_user}",
"gists_url": "https://api.github.com/users/arunppsg/gists{/gist_id}",
"starred_url": "https://api.github.com/users/arunppsg/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/arunppsg/subscriptions",
"organizations_url": "https://api.github.com/users/arunppsg/orgs",
"repos_url": "https://api.github.com/users/arunppsg/repos",
"events_url": "https://api.github.com/users/arunppsg/events{/privacy}",
"received_events_url": "https://api.github.com/users/arunppsg/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Try 2019.12.31",
"Thanks for your suggestion. Pymatgen 2020.12.31 worked for me. In colab, it worked fine. In conda, there was module not found error for ruamel.yaml. But `import ruamel_yaml` worked. It was resolved by following steps:\r\n\r\n1. `cd anaconda3/lib/python3.8/site-packages/pymatgen`\r\n2. `find . -name '*.py' | xargs sed -i 's/ruamel.yaml/ruamel_yaml/g'`\r\n",
"Just to note that older versions of numpy are likely fine with the current pymatgen too if you're willing to manually git clone and run `python setup.py install`.",
"Thanks!"
] | 2021-07-02T01:56:44
| 2021-07-04T02:03:06
|
2021-07-02T14:13:46Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
The latest version of tensorflow and pymatgen are incompatible due to dependency errors. Tensorflow depends on older version of numpy (1.19.1) while pymatgen uses numpy 1.20.1.
When installing pymatgen in a new conda environment or in colab containing tensorflow, installation errors with exit status 1. Maybe it will be helpful if someone could point to the version of pymatgen using numpy 1.19.1 version.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2189/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2189/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2190
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2190/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2190/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2190/events
|
https://github.com/materialsproject/pymatgen/pull/2190
| 935,914,897
|
MDExOlB1bGxSZXF1ZXN0NjgyNzUxMjYx
| 2,190
|
Gruneisen classes
|
{
"login": "JaGeo",
"id": 22094846,
"node_id": "MDQ6VXNlcjIyMDk0ODQ2",
"avatar_url": "https://avatars.githubusercontent.com/u/22094846?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/JaGeo",
"html_url": "https://github.com/JaGeo",
"followers_url": "https://api.github.com/users/JaGeo/followers",
"following_url": "https://api.github.com/users/JaGeo/following{/other_user}",
"gists_url": "https://api.github.com/users/JaGeo/gists{/gist_id}",
"starred_url": "https://api.github.com/users/JaGeo/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/JaGeo/subscriptions",
"organizations_url": "https://api.github.com/users/JaGeo/orgs",
"repos_url": "https://api.github.com/users/JaGeo/repos",
"events_url": "https://api.github.com/users/JaGeo/events{/privacy}",
"received_events_url": "https://api.github.com/users/JaGeo/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Will fix the rest of the problems on Monday... (sorry)",
"Looking at the logs, it's unclear why so many linting issues are coming up for classes that are not touched by this PR -- only linting issues associated with the files modified by this PR need to be addressed.\r\n\r\nOne request is that when a new plotter is added, that it supports a `plotly` backend (see `PDPlotter` for an example). This is not required for the PR to be merged. The reason for this is that plotly plots can be more easily integrated with web apps, and also provide useful interactive features in Jupyter notebooks.\r\n\r\nOtherwise, this looks like a great addition! Thank you @ab5424 @gpetretto @JaGeo.",
"@mkhorton The linting issues are caused by the pylint upgrade, which now detects more instances of .item used,.",
"Ah, thanks @shyuep ",
"@mkhorton Thanks for your comment.\r\n\r\nRegarding the `plotly` backend: the new band structure plotter inherits from [`PhononBSPlotter`](https://github.com/materialsproject/pymatgen/blob/b789d74639aa851d7e5ee427a765d9fd5a8d1079/pymatgen/phonon/plotter.py#L242), which has no `plotly` backend either. Would it be ok to merge this first and add the new backend later? I think all classes in phonon/plotter only have a `matplotlib` backend, so it might be easier to rework everything in a separate PR. ",
"Yes, absolutely. As I said above that’s not required for this to be merged, it was more a comment to share our roadmap.\r\n\r\nWe actually do have plotly code for the BSPlotter that could be merged in, I’m not sure without checking to what extent PhononBSPlotter could make use of this.",
"Ok, thanks. `PhononBSPlotter` definitely uses some code from `BSPlotter` so this seems like a good start. ",
"I have fixed all problems in the linting part and also a problem with the test of the Gruneisen classes. The other errors seem to come from a different part of pymatgen that we haven't touched. Is it okay like this for now or do I need to do something else?\r\n\r\nThanks in advance. \r\n\r\nJG",
"There are still some problems from our side. They result from the fact that phonopy is needed for some of the tests. I hope that we can get back to them in the next days.",
"Has one of you an idea why only one test crashes? https://github.com/materialsproject/pymatgen/pull/2190/checks?check_run_id=3001365472\r\nIs there something different with regard to the requirements? Locally, the test runs correctly with and without phonopy installed.",
"Okay, found the problem... Let me know if we should do something more before this can be merged. \r\n\r\nJG",
"\n[](https://coveralls.io/builds/41239163)\n\nCoverage decreased (-0.8%) to 83.062% when pulling **836caa5b93f08c2c951ecd8e662d397f65b66802 on ab5424:gruneisen-classes-george** into **3855d50c857fc61e70a0a4497e194a819d617a64 on materialsproject:master**.\n",
"@mkhorton, I have now included all suggestions. I hope it is fine now.",
"Thanks all!",
"Thank you as well!"
] | 2021-07-02T16:30:37
| 2021-07-09T17:21:21
|
2021-07-09T16:41:53Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
@ab5424 , @gpetretto and I have worked on classes to handle Grüneisen parametrs in pymatgen. We have added a module phonopy.phonon.gruneisen.py and also plotter classes. The parameters can also be imported from phonopy.
Please let me know if we can do anything further to get this pull request merged soon.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2190/reactions",
"total_count": 2,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 1,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2190/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2190",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2190",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2190.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2190.patch",
"merged_at": "2021-07-09T16:41:53Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2191
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2191/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2191/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2191/events
|
https://github.com/materialsproject/pymatgen/pull/2191
| 936,138,980
|
MDExOlB1bGxSZXF1ZXN0NjgyOTQxODc3
| 2,191
|
CombinedData.as_dict/from_dict and other methods overwrite; CombinedData.as_lammpsdata deepcopied
|
{
"login": "htz1992213",
"id": 25351437,
"node_id": "MDQ6VXNlcjI1MzUxNDM3",
"avatar_url": "https://avatars.githubusercontent.com/u/25351437?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/htz1992213",
"html_url": "https://github.com/htz1992213",
"followers_url": "https://api.github.com/users/htz1992213/followers",
"following_url": "https://api.github.com/users/htz1992213/following{/other_user}",
"gists_url": "https://api.github.com/users/htz1992213/gists{/gist_id}",
"starred_url": "https://api.github.com/users/htz1992213/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/htz1992213/subscriptions",
"organizations_url": "https://api.github.com/users/htz1992213/orgs",
"repos_url": "https://api.github.com/users/htz1992213/repos",
"events_url": "https://api.github.com/users/htz1992213/events{/privacy}",
"received_events_url": "https://api.github.com/users/htz1992213/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Fixes #2188 ",
"\n[](https://coveralls.io/builds/42396886)\n\nCoverage decreased (-0.6%) to 83.143% when pulling **cab0331c70be487ee9b8edbdd8e8d98b888a6cba on htz1992213:master** into **40817cc97fe25bc027523a65aab528b59e22e78e on materialsproject:master**.\n",
"Hey @mkhorton, I have added tests for the new methods. ",
"The serialization of the `CombinedData` class should work as expected with the update @rkingsbury ",
"> The serialization of the `CombinedData` class should work as expected with the update @rkingsbury\r\n\r\nThanks @htz1992213 ! This looks like a great update that will really improve robustness and clarity of `CombinedData`.",
"Longer-term, I still think we should work on getting `DataFrame` supported in `monty` so that we can do away with custom `as_dict` and `from_dict` methods (and this comment applies throughout pymatgen). But for now this is a good solution. `CombinedData` has been inheriting too many methods from `LammpsData` that don't make sense, so I'm glad you added these as well as the `AttributeError` where appropriate.",
"I think we should prefer the monty solution over the custom `as_dict`/`from_dict` here actually; let's try and get the monty solution first, and fall back to this if that's impractical? I think it would be an equivalent number of lines of code.\r\n\r\nI'm trying to actively discourage custom `as_dict`, `from_dict` methods wherever possible, they're creating a lot of technical debt.",
"> I think we should prefer the monty solution over the custom `as_dict`/`from_dict` here actually; let's try and get the monty solution first, and fall back to this if that's impractical? I think it would be an equivalent number of lines of code.\r\n> \r\n> I'm trying to actively discourage custom `as_dict`, `from_dict` methods wherever possible, they're creating a lot of technical debt.\r\n\r\n@mkhorton I agree with you, however this PR fixes several pretty serious flaws in `CombinedData` that are affecting our workflows, and brings it into line with `LammpsData` which also has custom `as_dict` / `from_dict`. I'd encourage you to consider merging this as a stopgap solution until we get monty patched, which may take more time. Once that happens, we can remove `as_dict` / `from_dict` from all the Lammps I/O classes at once.",
"Unfortunately \"stopgap\" solutions are how we accrue technical debt, because we forget to go back and remove them. I am reluctant to merge this when there has been a clear alternative suggested, with specific code proposed, and that could be implemented on a comparable timeframe.",
"Note that this comment is for the `as_dict`, `from_dict` specifically, not for the remainder of the PR which is fine to merge.",
"> Unfortunately \"stopgap\" solutions are how we accrue technical debt, because we forget to go back and remove them. I am reluctant to merge this when there has been a clear alternative suggested, with specific code proposed, and that could be implemented on a comparable timeframe.\r\n\r\n@mkhorton I think modifying `monty` is beyond my expertise because it involves making test cases that I may not familiar. I don't think this is a stopgap because it is for sure built on the latest dependencies of` pymatgen`. But I would be happy to modify both `LammpsData`/`CombinedData` immediately once the patch to `monty` is in place. I am quite actively maintaining this submodule so no worries about forgetting to go back here. And I think the urgent is to make sure the serialization of `CombinedData` works.",
"> @mkhorton I think modifying monty is beyond my expertise because it involves making test cases that I may not familiar. \r\n\r\nOk, that's not a problem -- I'm happy to propose this change as a PR to monty myself but it'll take me a few days until I can get to it.",
"Hey @mkhorton @rkingsbury, Shyue merged the Monty update, so I was able to deprecate the as_dict and from_dict methods for LammpsData and CombinedData. The MontyEncoder/Decoder works fine here. One thing I want to add is that, the pandas.DataFrame.to_dict method will convert the int type \"index\" of the df to str type. This won't cause a fundamental error here because LammpsData will just output that column as str anyway. But I think in general it is something that could be fixed in Monty.",
"Thanks @htz1992213 ! Good catch re: the index type. I don't have a well-informed opinion of whether that type conversion is a problem or not.",
"Thanks @htz1992213! I'll go ahead and merge.\r\n\r\nThe index conversion is interesting to know -- this is done by pandas JSON export itself rather than monty, I believe?",
"@rkingsbury @mkhorton Yes I think pandas export JSON by itself. But I think that default behavior can be changed as described in this [answer](https://stackoverflow.com/a/44127109/15193876)",
"@htz1992213 ah, I see. If we make this change it would be a breaking change since it would change the serialization format, would that be ok?",
"@mkhorton I made the tests in this PR to only check if the actual values in dfs are the same regardless of the type. So it should be ok updating monty according to the link. I don't think it will break the things here.",
"I was asking more if it would cause any problems for you for your own work, if you've already been using the functionality, otherwise I will submit another PR to monty ASAP.",
"@mkhorton Oh I see. No, I don't think that is a problem for me. I was just discussing if it is a better practice to maintain the int type of the index for serializing a df in Monty.",
"I don't think it's a big issue, but it does seem best practice for serialization/deserialization to retain type information, and since it's a simple change I think we should do that before it sees widespread usage.",
"@mkhorton Yeah I totally agree, a simple fix would be great!"
] | 2021-07-03T01:19:59
| 2021-08-26T18:00:19
|
2021-08-25T16:45:55Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
* Add `as_dict`, `from_dict` for CombinedData to overwrite superclass methods
* Overwrite `structure`, `disassemble`, `from_ff_and_topologie`, `from_structur` methods for CombinedData
* Rewrite `as_lammpsdata` method for CombinedData so that each attribute is deep copied
* Update docs
* Join string frags
* Added tests
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2191/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2191/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2191",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2191",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2191.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2191.patch",
"merged_at": "2021-08-25T16:45:55Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2192
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2192/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2192/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2192/events
|
https://github.com/materialsproject/pymatgen/issues/2192
| 937,354,786
|
MDU6SXNzdWU5MzczNTQ3ODY=
| 2,192
|
SubstrateAnalyzer not returning the substrate matches
|
{
"login": "emily-l-wang",
"id": 54878006,
"node_id": "MDQ6VXNlcjU0ODc4MDA2",
"avatar_url": "https://avatars.githubusercontent.com/u/54878006?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/emily-l-wang",
"html_url": "https://github.com/emily-l-wang",
"followers_url": "https://api.github.com/users/emily-l-wang/followers",
"following_url": "https://api.github.com/users/emily-l-wang/following{/other_user}",
"gists_url": "https://api.github.com/users/emily-l-wang/gists{/gist_id}",
"starred_url": "https://api.github.com/users/emily-l-wang/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/emily-l-wang/subscriptions",
"organizations_url": "https://api.github.com/users/emily-l-wang/orgs",
"repos_url": "https://api.github.com/users/emily-l-wang/repos",
"events_url": "https://api.github.com/users/emily-l-wang/events{/privacy}",
"received_events_url": "https://api.github.com/users/emily-l-wang/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Also tagging @shyamd here. A recent PR #2149 cleaned up a lot of the interface functionality and added new features.",
"Yup. Dumb bug on my part. Already opened a PR to fix it in #2198 ",
"Thanks!"
] | 2021-07-05T20:44:12
| 2021-09-13T08:39:12
|
2021-09-13T08:39:12Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
pymatgen.analysis.interfaces.substrate_analyzer returns None objects rather than SubstrateMatch objects.
**To Reproduce**
Code to run:
from pymatgen.analysis.interfaces.substrate_analyzer import SubstrateAnalyzer
from pymatgen.ext.matproj import MPRester
mpr = MPRester("API_KEY")
substrate = mpr.get_structure_by_material_id("mp-19009")
film = mpr.get_structure_by_material_id("mp-23")
sa = SubstrateAnalyzer()
gener = sa.calculate(film, substrate, lowest = True)
list(gener)
**Expected behavior**
Expect an equal sized list of SubstrateMatches.
**Desktop (please complete the following information):**
- Windows 10
- Version 2022.0.9
**Additional context**
A way to fix this would be adding a return at L69 here: https://github.com/materialsproject/pymatgen/blob/v2022.0.9/pymatgen/analysis/interfaces/substrate_analyzer.py, but I'm not certain if the list of Nones is expected behavior somehow and I might just be calling it incorrectly.
Also, is it just the build rather than the calculations themselves that are different from Version 2020.7.3's substrate_analyzer (https://pymatgen.org/_modules/pymatgen/analysis/substrate_analyzer.html)? The reason I ask is because I find calculating the MCIA much simpler with the dict version of returning the matches. Is there something simple like in this forum post (https://matsci.org/t/how-to-find-the-substrate-for-my-own-structure/967/2) I can use without doing a cross product with the film_sl_vectors field?
|
{
"login": "shyamd",
"id": 16827130,
"node_id": "MDQ6VXNlcjE2ODI3MTMw",
"avatar_url": "https://avatars.githubusercontent.com/u/16827130?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyamd",
"html_url": "https://github.com/shyamd",
"followers_url": "https://api.github.com/users/shyamd/followers",
"following_url": "https://api.github.com/users/shyamd/following{/other_user}",
"gists_url": "https://api.github.com/users/shyamd/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyamd/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyamd/subscriptions",
"organizations_url": "https://api.github.com/users/shyamd/orgs",
"repos_url": "https://api.github.com/users/shyamd/repos",
"events_url": "https://api.github.com/users/shyamd/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyamd/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2192/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2192/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2193
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2193/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2193/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2193/events
|
https://github.com/materialsproject/pymatgen/pull/2193
| 939,111,891
|
MDExOlB1bGxSZXF1ZXN0Njg1MzgzMTAw
| 2,193
|
Bugfix for defect BandFillingCorrection
|
{
"login": "kavanase",
"id": 51478689,
"node_id": "MDQ6VXNlcjUxNDc4Njg5",
"avatar_url": "https://avatars.githubusercontent.com/u/51478689?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/kavanase",
"html_url": "https://github.com/kavanase",
"followers_url": "https://api.github.com/users/kavanase/followers",
"following_url": "https://api.github.com/users/kavanase/following{/other_user}",
"gists_url": "https://api.github.com/users/kavanase/gists{/gist_id}",
"starred_url": "https://api.github.com/users/kavanase/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/kavanase/subscriptions",
"organizations_url": "https://api.github.com/users/kavanase/orgs",
"repos_url": "https://api.github.com/users/kavanase/repos",
"events_url": "https://api.github.com/users/kavanase/events{/privacy}",
"received_events_url": "https://api.github.com/users/kavanase/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/41191123)\n\nCoverage decreased (-0.6%) to 83.191% when pulling **6c69bc4c6b6657b60883ecc892885a1882f9bf4a on kavanase:master** into **3855d50c857fc61e70a0a4497e194a819d617a64 on materialsproject:master**.\n",
"Thanks @kavanase! Both for catching and fixing. Tagging @nwinner in case this is relevant to your work.",
"Thanks @kavanase ! This is something I never would have noticed @mkhorton since we don't use band filling corrections for my work."
] | 2021-07-07T17:33:20
| 2021-07-12T19:11:56
|
2021-07-08T19:51:21Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This is a bugfix for two significant errors in the BandFillingCorrection class for defects.
I have attached a PDF, notebook and corresponding data to demonstrate the errors, and how these are now fixed with this PR. I have changed the value in the BandFillingCorrection test to match the correct value as well, and have attached a PDF & notebook for this, calculating this correction manually to verify that it is the correct result.
* **Fix 1**: Fix the potential alignment shifting of the band extrema.
* **Fix 2**: Fix the eigenstate occupancy spinfactor calculation, so that spin-orbit coupling (SOC) calculations can be parsed correctly (previously giving a spurious doubling of the occupancies in these cases).
I have added a new test for the BandFillingCorrection with a SOC defect calculation, and have attached files here demonstrating that this gives the correct value (by also calculating it manually).
[BandFillingCorrection_Tests.zip](https://github.com/materialsproject/pymatgen/files/6779207/BandFillingCorrection_Tests.zip)
[Te_i_Pymatgen_Bandfilling_Error_Demo.zip](https://github.com/materialsproject/pymatgen/files/6779213/Te_i_Pymatgen_Bandfilling_Error_Demo.zip)
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2193/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2193/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2193",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2193",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2193.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2193.patch",
"merged_at": "2021-07-08T19:51:21Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2194
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2194/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2194/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2194/events
|
https://github.com/materialsproject/pymatgen/issues/2194
| 939,419,843
|
MDU6SXNzdWU5Mzk0MTk4NDM=
| 2,194
|
ZSLGenerator not returning all inequivalent transformations
|
{
"login": "bernstei",
"id": 1202357,
"node_id": "MDQ6VXNlcjEyMDIzNTc=",
"avatar_url": "https://avatars.githubusercontent.com/u/1202357?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/bernstei",
"html_url": "https://github.com/bernstei",
"followers_url": "https://api.github.com/users/bernstei/followers",
"following_url": "https://api.github.com/users/bernstei/following{/other_user}",
"gists_url": "https://api.github.com/users/bernstei/gists{/gist_id}",
"starred_url": "https://api.github.com/users/bernstei/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/bernstei/subscriptions",
"organizations_url": "https://api.github.com/users/bernstei/orgs",
"repos_url": "https://api.github.com/users/bernstei/repos",
"events_url": "https://api.github.com/users/bernstei/events{/privacy}",
"received_events_url": "https://api.github.com/users/bernstei/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I just noticed the undocumented `bidirectional` argument for the `ZSLGenerator` constructor, but it's not clear to me how exactly it relates to this behavior. It would be nice if it could be explained in the docstring.",
"I just updated to 2022.0.10, and I'm running into a new bug. I've modified my code for the change from the match object being a dict to a proper object, but now `match.match_area` gives an error:\r\n```\r\nTraceback (most recent call last):\r\n File \"t.py\", line 7, in <module>\r\n print(match.match_area)\r\n File \"/share/apps/python/lib/python3.8/site-packages/pymatgen/analysis/interfaces/zsl.py\", line 37, in match_area\r\n return vec_area(*self.film_sl_vectors.tolist())\r\nAttributeError: 'list' object has no attribute 'tolist'\r\n```",
"Tagging @shyamd here. A recent PR #2149 cleaned up a lot of the interface functionality and added new features, I'm not sure if this is related.",
"Might be. I'll take a look tomorrow. ",
"The match area problem should be fixed in #2198 with an appropriate test. ",
"On Jul 8, 2021, at 4:29 PM, Shyam Dwaraknath ***@***.******@***.***>> wrote:\n\nThe match area problem should be fixed in #2198<https://github.com/materialsproject/pymatgen/pull/2198> with an appropriate test.\n\nThanks. Any guesses how long before the pypi version has this update, or should I just install directly from the github source if I want to take advantage of it soon?\n\n",
"Probably best to just grab it off github. I don't think you're using the latest pymatgen as-is. The interfaces refactor longer returns just plain python dictionaries, but rather a dataclass object with properly named attributes to reduce ambiguities. Your output would look more like:\r\n``` \r\nZSLMatch(film_sl_vectors=[array([0. , 1.02, 0. ]), array([3., 0., 0.])], substrate_sl_vectors=[array([0. , 1.01, 0. ]), array([2.97, 0. , 0. ])], film_vectors=[(1, 0, 0), (0, 1.02, 0)], substrate_vectors=[(0.99, 0.0, 0.0), (0.0, 1.01, 0.0)], film_transformation=array([[3., 0.],\r\n [0., 1.]]), substrate_transformation=array([[3., 0.],\r\n [0., 1.]]))\r\n```",
"This should be fixed. Let me know if it's not yet. "
] | 2021-07-08T03:00:57
| 2021-09-13T08:39:41
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
`ZSLGenerator` doesn't return some expected mappings between two low symmetry lattices.
**To Reproduce**
run
```
from pymatgen.analysis.substrate_analyzer import ZSLGenerator
# find in-plane supercell lattice pairs that are compatible
zsl = ZSLGenerator(max_area=4.5, max_length_tol = 0.2, max_angle_tol=0.2, max_area_ratio_tol=0.2)
for match in zsl([(1, 0, 0), (0, 1.02, 0)], [(0.99, 0.0, 0.0), (0.0, 1.01, 0.0)]):
print(match)
```
This only outputs one match with area ~1:
```
tin 1097 : python t.py | head -20
{'film_sl_vecs': array([[1. , 0. , 0. ],
[0. , 1.02, 0. ]]), 'sub_sl_vecs': array([[0.99, 0. , 0. ],
[0. , 1.01, 0. ]]), 'match_area': 1.02, 'film_vecs': array([[1. , 0. , 0. ],
[0. , 1.02, 0. ]]), 'sub_vecs': array([[0.99, 0. , 0. ],
[0. , 1.01, 0. ]]), 'film_transformation': array([[1., 0.],
[0., 1.]]), 'substrate_transformation': array([[1., 0.],
[0., 1.]])}
{'film_sl_vecs': array([[1. , 0. , 0. ],
[0. , 2.04, 0. ]]), 'sub_sl_vecs': array([[0.99, 0. , 0. ],
[0. , 2.02, 0. ]]), 'match_area': 2.04, 'film_vecs': array([[1. , 0. , 0. ],
[0. , 1.02, 0. ]]), 'sub_vecs': array([[0.99, 0. , 0. ],
[0. , 1.01, 0. ]]), 'film_transformation': array([[1., 0.],
[0., 2.]]), 'substrate_transformation': array([[1., 0.],
[0., 2.]])}
```
**Expected behavior**
Two matches that have area ~1, one that maps the first material's vector0 to the second material's vector0, and the second that maps the first material's vector0 to the second material's vector1, i.e. rotated by 90 deg. Both do have the same pair of surface cells, so maybe that's why they are not returned separately, but they are two different relative orientations, so I expected they'd each be in the list separately.
**Desktop (please complete the following information):**
CentOS 7, anaconda3 python 3.8, pypi pymatgen 2022.0.8
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2194/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2194/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2195
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2195/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2195/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2195/events
|
https://github.com/materialsproject/pymatgen/issues/2195
| 939,460,311
|
MDU6SXNzdWU5Mzk0NjAzMTE=
| 2,195
|
Inconsistent use of output file path in Structure.to() method
|
{
"login": "yw-choi",
"id": 1888873,
"node_id": "MDQ6VXNlcjE4ODg4NzM=",
"avatar_url": "https://avatars.githubusercontent.com/u/1888873?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/yw-choi",
"html_url": "https://github.com/yw-choi",
"followers_url": "https://api.github.com/users/yw-choi/followers",
"following_url": "https://api.github.com/users/yw-choi/following{/other_user}",
"gists_url": "https://api.github.com/users/yw-choi/gists{/gist_id}",
"starred_url": "https://api.github.com/users/yw-choi/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/yw-choi/subscriptions",
"organizations_url": "https://api.github.com/users/yw-choi/orgs",
"repos_url": "https://api.github.com/users/yw-choi/repos",
"events_url": "https://api.github.com/users/yw-choi/events{/privacy}",
"received_events_url": "https://api.github.com/users/yw-choi/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 19613987,
"node_id": "MDU6TGFiZWwxOTYxMzk4Nw==",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/bug",
"name": "bug",
"color": "9222c1",
"default": true,
"description": ""
},
{
"id": 5304335471,
"node_id": "LA_kwDOACgets8AAAABPCm8bw",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/io",
"name": "io",
"color": "c2e0c6",
"default": false,
"description": "Input/output functionality"
}
] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks for the report. I didn't see your issue at the time but this was fixed in 6a6056cd417f1310d6810cf3ebdd2d2126fe4d41 and released in [v2022.11.1](https://github.com/materialsproject/pymatgen/releases/tag/v2022.11.1)."
] | 2021-07-08T04:36:58
| 2023-05-30T16:53:47
|
2023-05-30T16:53:47Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Structure.to() method ignores the parent path of filename for certain file formats, say, .xsf format.
For example, in the following scenario,
```python
struct = Structure(...)
struct.to(filename="/path/to/struct.ext")
```
the output file is correctly written to /path/to/struct.ext when .ext is .cif, .mcif, .json, .cssr, .poscar. But, for some formats including .xsf, it is written at ./struct.ext (current directory).
This is because, in the source code of to(), fname (which only contains the basename of the given filename) is used instead of filename variable for some extensions.
For example, in core/structure.py:2303-2310,
```python
elif fmt == "xsf" or fnmatch(fname.lower(), "*.xsf*"):
from pymatgen.io.xcrysden import XSF
s = XSF(self).to_string()
if filename:
with zopen(fname, "wt", encoding="utf8") as f:
f.write(s)
return
```
zopen uses fname instead of filename. It is also the case for some other extensions.
**Expected behavior**
I think to() method should always use filename as the output path.
**Desktop (please complete the following information):**
- OS: MacOS Big Sur
|
{
"login": "janosh",
"id": 30958850,
"node_id": "MDQ6VXNlcjMwOTU4ODUw",
"avatar_url": "https://avatars.githubusercontent.com/u/30958850?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/janosh",
"html_url": "https://github.com/janosh",
"followers_url": "https://api.github.com/users/janosh/followers",
"following_url": "https://api.github.com/users/janosh/following{/other_user}",
"gists_url": "https://api.github.com/users/janosh/gists{/gist_id}",
"starred_url": "https://api.github.com/users/janosh/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/janosh/subscriptions",
"organizations_url": "https://api.github.com/users/janosh/orgs",
"repos_url": "https://api.github.com/users/janosh/repos",
"events_url": "https://api.github.com/users/janosh/events{/privacy}",
"received_events_url": "https://api.github.com/users/janosh/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2195/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2195/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2196
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2196/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2196/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2196/events
|
https://github.com/materialsproject/pymatgen/issues/2196
| 939,917,407
|
MDU6SXNzdWU5Mzk5MTc0MDc=
| 2,196
|
Typo in init.py Ruamel import
|
{
"login": "arepstein",
"id": 17616763,
"node_id": "MDQ6VXNlcjE3NjE2NzYz",
"avatar_url": "https://avatars.githubusercontent.com/u/17616763?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/arepstein",
"html_url": "https://github.com/arepstein",
"followers_url": "https://api.github.com/users/arepstein/followers",
"following_url": "https://api.github.com/users/arepstein/following{/other_user}",
"gists_url": "https://api.github.com/users/arepstein/gists{/gist_id}",
"starred_url": "https://api.github.com/users/arepstein/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/arepstein/subscriptions",
"organizations_url": "https://api.github.com/users/arepstein/orgs",
"repos_url": "https://api.github.com/users/arepstein/repos",
"events_url": "https://api.github.com/users/arepstein/events{/privacy}",
"received_events_url": "https://api.github.com/users/arepstein/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-07-08T14:25:18
| 2021-07-08T14:27:09
|
2021-07-08T14:27:09Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
core/__init__.py tries to import ruamal, which results in an error, rather than ruamel, which I think is the proper import statement.
**To Reproduce**
Steps to reproduce the behavior:
import pymatgen.core
Provide any example files that are needed to reproduce the error,
especially if the bug pertains to parsing of a file.
**Expected behavior**
Expect `import pymatgen.core` to execute successfully
**Screenshots**
<img width="1041" alt="Screen Shot 2021-07-08 at 10 22 14 AM" src="https://user-images.githubusercontent.com/17616763/124938779-945c3b80-dfd6-11eb-854f-48871f88dcbc.png">
**Desktop (please complete the following information):**
- OS: Mac
- Version 10.14.6
**Additional context**
` from ruamel import yaml` fixes the issue for me
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2196/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2196/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2197
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2197/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2197/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2197/events
|
https://github.com/materialsproject/pymatgen/issues/2197
| 940,131,243
|
MDU6SXNzdWU5NDAxMzEyNDM=
| 2,197
|
Improve resource structure to work better with systems that package code and data
|
{
"login": "jdagdelen",
"id": 9101793,
"node_id": "MDQ6VXNlcjkxMDE3OTM=",
"avatar_url": "https://avatars.githubusercontent.com/u/9101793?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jdagdelen",
"html_url": "https://github.com/jdagdelen",
"followers_url": "https://api.github.com/users/jdagdelen/followers",
"following_url": "https://api.github.com/users/jdagdelen/following{/other_user}",
"gists_url": "https://api.github.com/users/jdagdelen/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jdagdelen/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jdagdelen/subscriptions",
"organizations_url": "https://api.github.com/users/jdagdelen/orgs",
"repos_url": "https://api.github.com/users/jdagdelen/repos",
"events_url": "https://api.github.com/users/jdagdelen/events{/privacy}",
"received_events_url": "https://api.github.com/users/jdagdelen/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks for the feedback. but I need a bit more context about where this is causing problems. In general, data files that are packaged are meant to be defaults. If alternatives are allowed, those code should be implemented with an argument to allow other files to be supplied. So pls provide a specific scenario where these break and code changes are needed. We use pymatgen on HPC resources too and as far as I know, the code and data reside in the same paths. \r\nI don't remember a scenario where python will install code and data files to different paths, even though the MANIFEST specifically includes the files in the right paths.\r\n\r\nLoading by filename is a feature, not a bug. I got tired of typing multiple lines with lots of words to load files. Of course, I can add an arg to monty to force a choice of format. That can be an additional flexibility.\r\n\r\nMinor thing: I think we should avoid stipulating feedback sources as \"from FAANG engineer\". That is not a credential. A FANNG engineer can be someone like Jeff Dean or a lowly intern. The source can either come with proper name and credentials (e.g., master python programmer) or no source needs to be specified and we will objectively evaluate the feedback regardless of source.",
"I have implemented a fmt override for monty. See https://github.com/materialsvirtuallab/monty/commit/02de6b36dd8a00519a40789bee38e66d513ac48a \r\n\r\nWill wait for discussion of the rest of the feedback.",
"Thanks for getting back to me so quickly. I'm a student in Kristin's group but doing an industry internship this summer, which is where this suggestion comes from. To clarify, I just said \"FAANG\" to clue in Matt/you about the kind of system this is an issue on, not as a credential. I used his quote because I'm not too familiar with monty and I didn't want to mis-translate something.\r\n\r\n> We use pymatgen on HPC resources too and as far as I know, the code and data reside in the same paths.\r\n\r\nThat's a key difference between NERSC and our system. I'm still sort of new to this infrastructure, so I might not have a 100% accurate understanding of the details, but on our system code and data don't reside on the same path and we have to explicitly choose what files/packages get included in the packaged bundle. I'm not sure if it's for security, performance, or something else, but I think other companies/institutions might use similar infrastructure. \r\n\r\nWe can certainly keep all the modifications we have made to pymatgen in our internal version, but if it made sense for other users too maybe it makes sense to enable that behavior. I'm happy to make these changes to how resources are accessed and submit a PR myself, but I just wanted to have the discussion with you and Matt first before writing any code. ",
"Doing a little digging, I think this is a broader topic on how resources are handled in python, not specific to pymatgen. \r\n\r\nFor example, [importlib.resources](https://docs.python.org/3/library/importlib.html#module-importlib.resources) seems to be addressing a similar issue. "
] | 2021-07-08T18:50:52
| 2021-08-24T15:35:27
|
2021-08-24T15:35:27Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Is your feature request related to a problem? Please describe.**
The current implementation of pymatgen is difficult to adapt/use in environments where data files are packaged separately from source code. In these cases, the actual file name of the resource, such as `pymatgen/analysis/op_params.yaml` won't necessarily have the same root path as the code opening it, so things like the following do not work:
```
_directory = os.path.join(os.path.dirname(__file__))
with open(os.path.join(_directory, "op_params.yaml"), "rt") as f:
default_op_params = yaml.safe_load(f)
```
We also use monty to read data files which can introduce non-deterministic bugs. Feedback from engineer at FAANG company using pymatgen regarding monty:
"Monty loads check by filename for a mplk. Since these are packaged files, they have a lot of random identifiers early on
And if you're running 10K jobs, these often yield bugs."
**Describe the solution you'd like**
1. Create a single submodule/function for loading resources so that users on HPC systems that package data files can modify one load function instead of dozens/hundreds across entire library.
2. Where possible, use yaml.safe_load and json.load instead of monty
Open to discussion about best way to/whether to accommodate these use cases @mkhorton @shyuep.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2197/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2197/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/2198
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2198/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2198/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2198/events
|
https://github.com/materialsproject/pymatgen/pull/2198
| 940,181,846
|
MDExOlB1bGxSZXF1ZXN0Njg2MjkyMDA5
| 2,198
|
Fix substrate analyzer not actually returning objects
|
{
"login": "shyamd",
"id": 16827130,
"node_id": "MDQ6VXNlcjE2ODI3MTMw",
"avatar_url": "https://avatars.githubusercontent.com/u/16827130?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyamd",
"html_url": "https://github.com/shyamd",
"followers_url": "https://api.github.com/users/shyamd/followers",
"following_url": "https://api.github.com/users/shyamd/following{/other_user}",
"gists_url": "https://api.github.com/users/shyamd/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyamd/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyamd/subscriptions",
"organizations_url": "https://api.github.com/users/shyamd/orgs",
"repos_url": "https://api.github.com/users/shyamd/repos",
"events_url": "https://api.github.com/users/shyamd/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyamd/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"\n[](https://coveralls.io/builds/41228307)\n\nCoverage decreased (-0.6%) to 83.187% when pulling **33c9032ca48e98c119d13b20b2db96bd9386cc2a on shyamd:fix_sub_analyzer** into **ec611b9f0fc04c49de003132a53ff4df4b2a3f7f on materialsproject:master**.\n",
"> This should fix #2195, which was a bug introduced in the interfaces reorganization where the substrate analyzer was constructing matches to return, but not actually returning them... DOH!\r\n> \r\n> I've added a test to check for this in the future.\r\n\r\nAre you sure it's #2195 ? That seems to be about some file I/O bug.",
"Thanks @shyamd !"
] | 2021-07-08T20:07:19
| 2023-05-30T16:49:34
|
2021-07-09T22:45:15Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
This should fix #2198 , which was a bug introduced in the interfaces reorganization where the substrate analyzer was constructing matches to return, but not actually returning them... DOH!
I've added a test to check for this in the future.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2198/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2198/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/2198",
"html_url": "https://github.com/materialsproject/pymatgen/pull/2198",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/2198.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/2198.patch",
"merged_at": "2021-07-09T22:45:15Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/2199
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2199/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2199/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2199/events
|
https://github.com/materialsproject/pymatgen/issues/2199
| 940,199,792
|
MDU6SXNzdWU5NDAxOTk3OTI=
| 2,199
|
Outdated costdb_elements.csv, active elemental price updates for CostAnalysis?
|
{
"login": "CifLord",
"id": 8036054,
"node_id": "MDQ6VXNlcjgwMzYwNTQ=",
"avatar_url": "https://avatars.githubusercontent.com/u/8036054?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CifLord",
"html_url": "https://github.com/CifLord",
"followers_url": "https://api.github.com/users/CifLord/followers",
"following_url": "https://api.github.com/users/CifLord/following{/other_user}",
"gists_url": "https://api.github.com/users/CifLord/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CifLord/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CifLord/subscriptions",
"organizations_url": "https://api.github.com/users/CifLord/orgs",
"repos_url": "https://api.github.com/users/CifLord/repos",
"events_url": "https://api.github.com/users/CifLord/events{/privacy}",
"received_events_url": "https://api.github.com/users/CifLord/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"If there is an API that you can use to use latest prices, by all means implement an interface to it. I think there is an XML feed that can be used quite easily."
] | 2021-07-08T20:33:10
| 2021-07-08T21:17:53
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
The costdb_elements.csv is vastly outdated, especially for metal prices. If we look at Rh today, its currently at $600000/kg compared to the $20000/kg price obtained from WolframAlpha in 2013, about a 3000% increase. However even if we update all the elements with the most recent costs, there's still a lot of volatility in price points, e.g. Rh was going for nearly 1mil/kg just a few months ago. Is there a way to implement a pymatgen interface with specific websites like lme.com or anywhere that keeps a daily update on prices so a user has the option of using a static price point provided by a more up-to-date csv file in pymatgen or they use an api-like object (maybe using something like beautifulsoup) that does analysis based on currently reported prices.
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2199/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2199/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||||
https://api.github.com/repos/materialsproject/pymatgen/issues/2200
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2200/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2200/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2200/events
|
https://github.com/materialsproject/pymatgen/issues/2200
| 940,234,112
|
MDU6SXNzdWU5NDAyMzQxMTI=
| 2,200
|
V+2 and V+3 do not appear in Pourbaix Diagrams
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
open
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2021-07-08T21:25:41
| 2021-07-08T21:26:59
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
V+2 and V+3 species do not appear on our `PourbaixDiagram` for vanadium, even though several credible sources indicate that they should. This shortcoming was reported by an external user.
I've included below the pymatgen / Materials Project Pourbaix Diagram and several other points of reference. The majority of these show V+3 present in the low pH-pE region. Not all show V+2.
In our diagrams, it appears that we overstabilize V2O3 in this region, and hence do not see the weak acid behavior of V2O3 -> V(OH)2+ -> VOH+2 -> V+3 that several diagrams show.
**Additional context**
V+2 and V+3 (aq) are not in the NBS Tables, which is the source of the majority of the aqueous ion reference data we use. A possible way to improve the accuracy of the Vanadium-containing diagrams is to find experimental data for these species and add it to the ion data. I will investigate this during the next round of improvements to the Pourbaix code.
**Current pymatgen Pourbaix diagram (1e-8 M concentration, via website)**
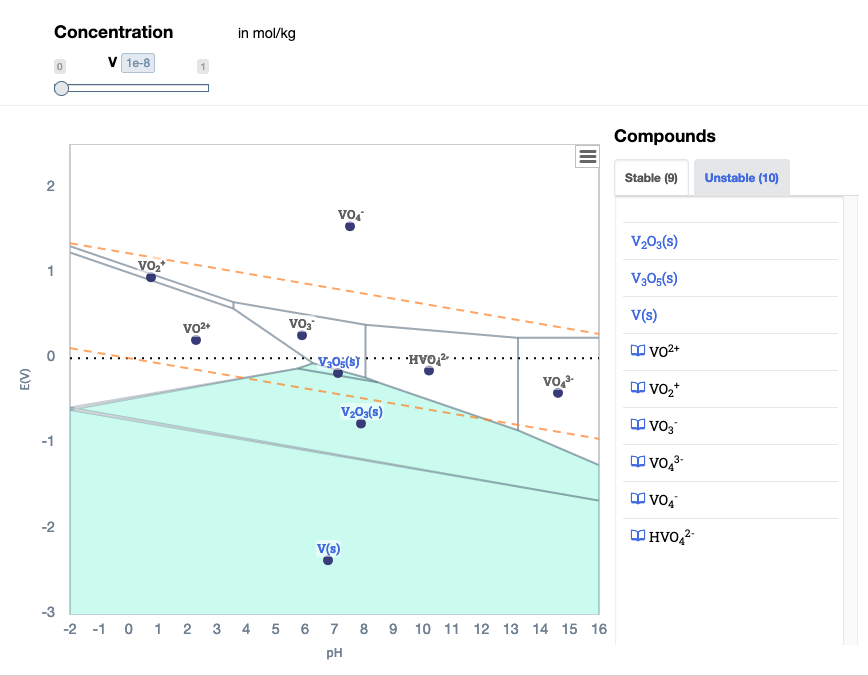
**Diagram constructed by external user (1 M concentration)**
Based on data from _Standard Potentials in Aqueous Solution_, A.J. Bard, R. Parsons and J. Jordan (Eds.), Marcel Dekker, New York, 1985.
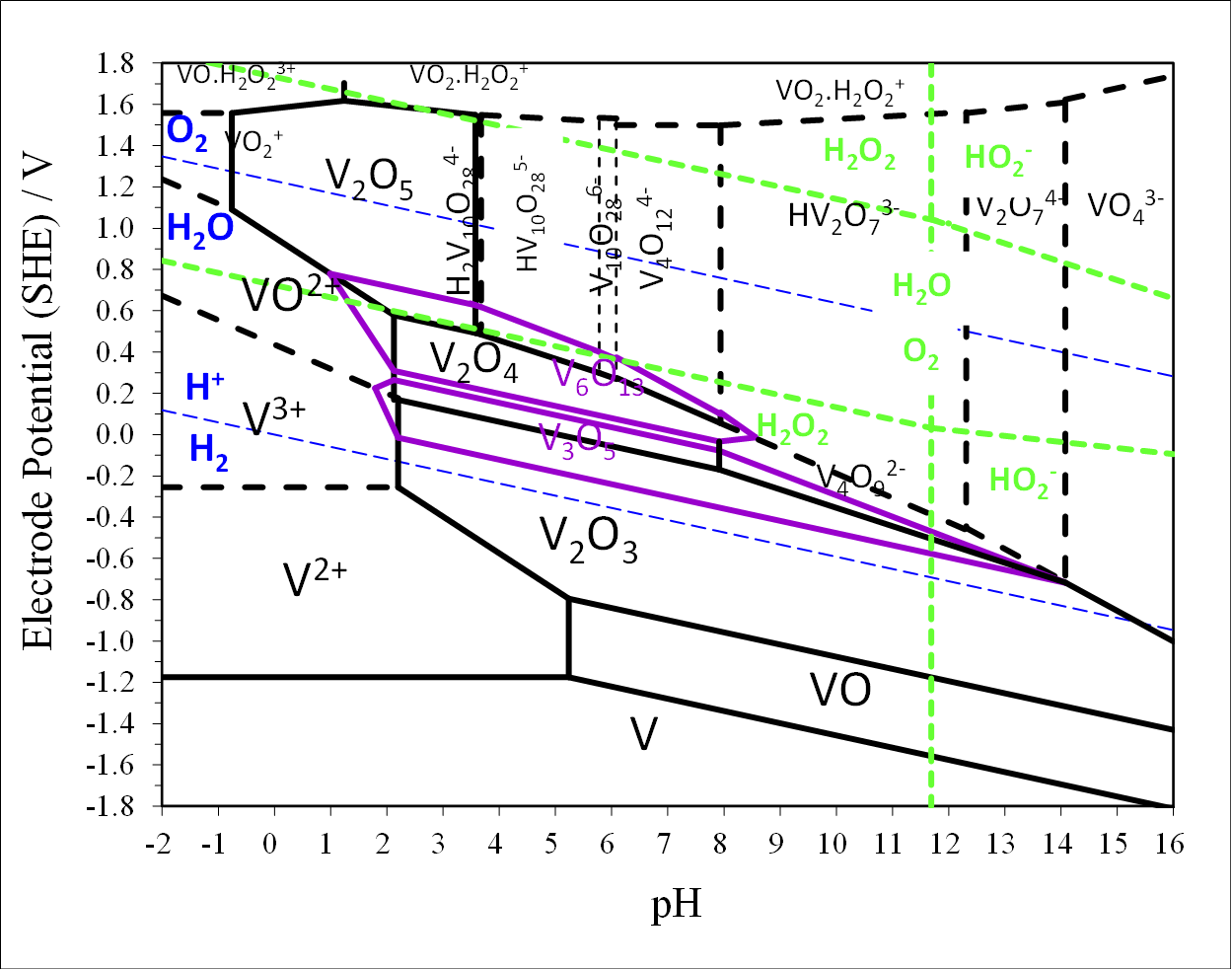
**A comparison of several databases available [here](https://www.nrc.gov/docs/ML1808/ML18089A638.pdf)**

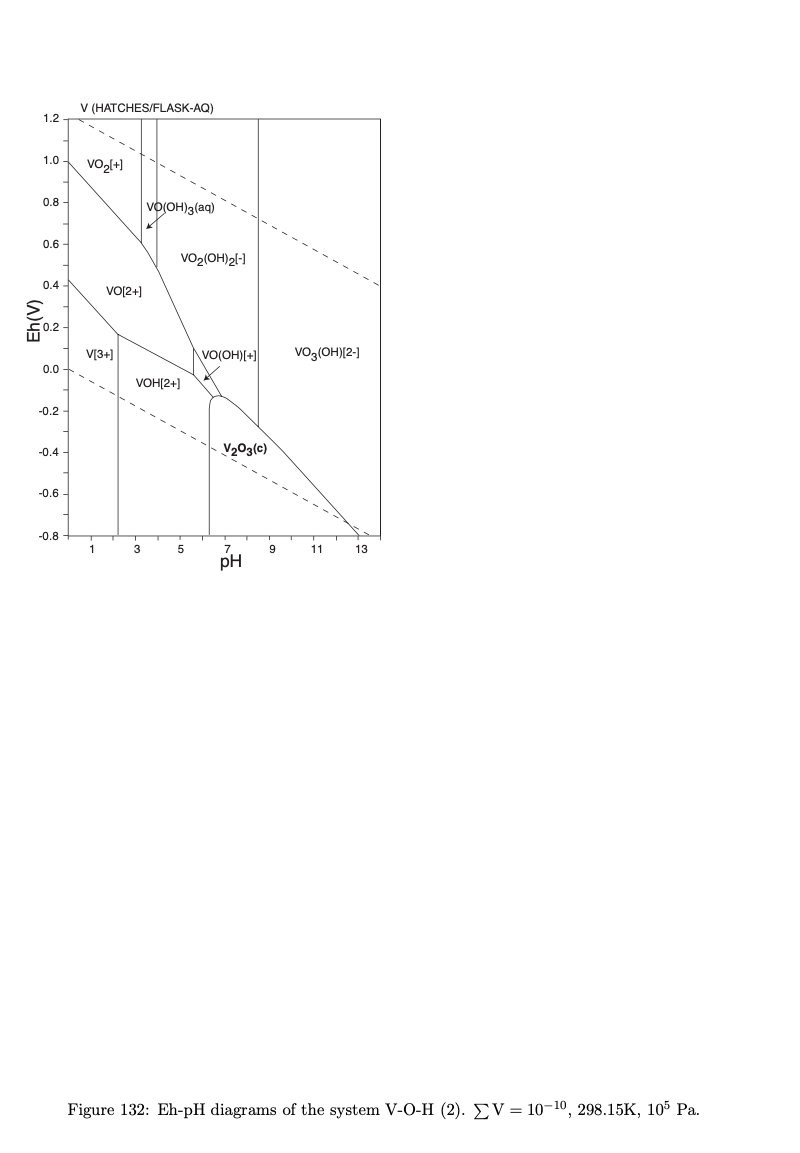
**Diagram from Pourbaix's Atlas**
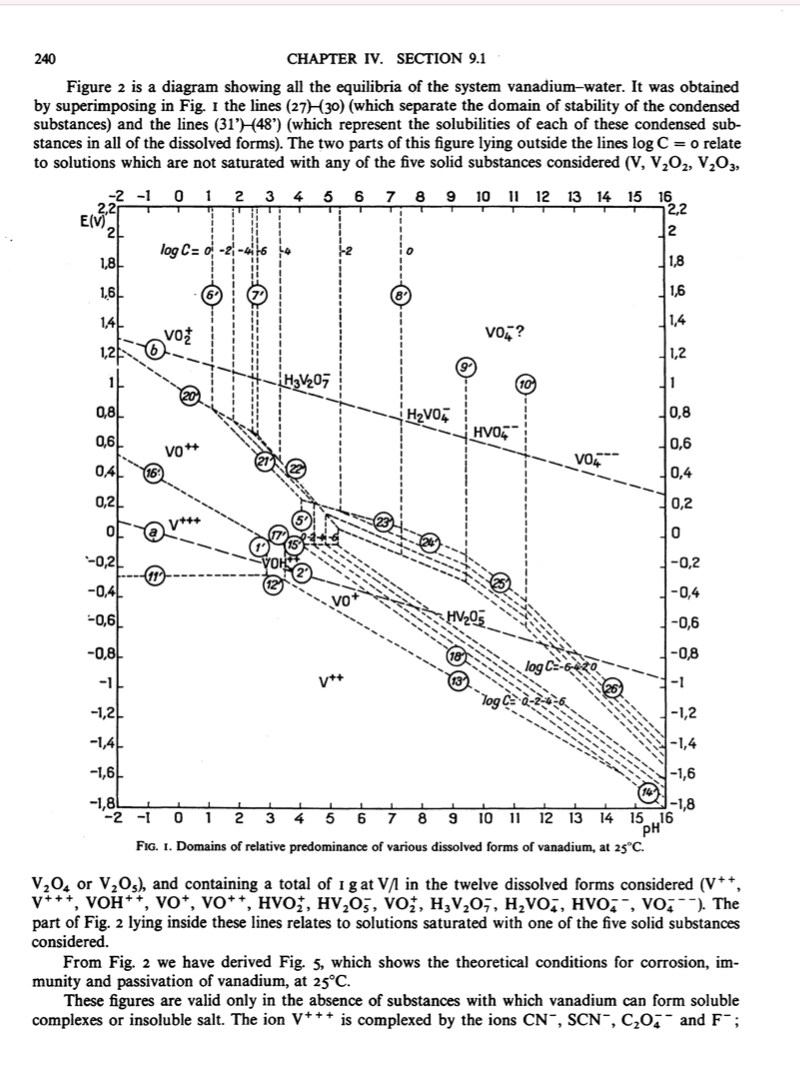
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/2200/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/2200/timeline
| null | false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.