repo_name stringlengths 9 75 | topic stringclasses 30
values | issue_number int64 1 203k | title stringlengths 1 976 | body stringlengths 0 254k | state stringclasses 2
values | created_at stringlengths 20 20 | updated_at stringlengths 20 20 | url stringlengths 38 105 | labels listlengths 0 9 | user_login stringlengths 1 39 | comments_count int64 0 452 |
|---|---|---|---|---|---|---|---|---|---|---|---|
jupyter/nbgrader | jupyter | 1,417 | nbgrader feedback fail | <!--
Thanks for helping to improve nbgrader!
If you are submitting a bug report or looking for support, please use the below
template so we can efficiently solve the problem.
If you are requesting a new feature, feel free to remove irrelevant pieces of
the issue template.
-->
when click Generate Feedback ... | closed | 2021-03-05T16:39:40Z | 2021-03-25T23:34:29Z | https://github.com/jupyter/nbgrader/issues/1417 | [
"duplicate"
] | 2010hexi | 1 |
Nemo2011/bilibili-api | api | 613 | [漏洞] 有dolby音效时获取最好的流时出错 | **Python 版本:** 3
**模块版本:** main/dev
**运行环境:** MacOS
**模块路径:** `bilibili_api.video.detect_best_streams()`
**解释器:** cpython
**报错信息:**
```python
Traceback (most recent call last):
File "bilibili-api/server.py", line 181, in <module>
sync(test())
File "bilibili-api/bilibili_api/utils/sync.py",... | closed | 2023-12-28T01:23:12Z | 2023-12-31T00:40:08Z | https://github.com/Nemo2011/bilibili-api/issues/613 | [
"bug",
"solved"
] | nooblong | 3 |
ivy-llc/ivy | pytorch | 28,741 | [Bug]: Failed to sign up | ### Bug Explanation
I followed the instructions (https://unify.ai/docs/ivy/overview/get_started.html) to sign up and add the API key to .ivy/key.pem. However, ivy still asks me to sign up.
### Steps to Reproduce Bug
The test code (located at `~/test_ivy.py`) is simple as follows:
```python
import ivy
ivy.trace_... | closed | 2024-04-21T03:56:48Z | 2024-06-17T08:25:01Z | https://github.com/ivy-llc/ivy/issues/28741 | [
"Bug Report"
] | zhangdongkun98 | 2 |
fugue-project/fugue | pandas | 95 | [FEATURE] Fillna as builtin | **Is your feature request related to a problem? Please describe.**
We should include a fillna method in the execution engine.
**Describe the solution you'd like**
It just needs to be simple like the Spark implementation. No need for forward fill and backfill.
**Additional context**
It will have a very similar... | closed | 2020-11-02T07:45:44Z | 2020-11-08T15:01:56Z | https://github.com/fugue-project/fugue/issues/95 | [
"enhancement",
"Fugue SQL",
"programming interface",
"core feature"
] | kvnkho | 0 |
ResidentMario/geoplot | matplotlib | 74 | Marker size | Hello,
Is there a way to specify the marker size in a pointplot ?
keith | closed | 2019-03-20T22:25:25Z | 2019-03-20T22:46:31Z | https://github.com/ResidentMario/geoplot/issues/74 | [] | vool | 1 |
tqdm/tqdm | pandas | 1,494 | buf-size [cs]hould default to bigger size (when using `--bytes`?) | Hi!
Just stumbled upon this today (mpire linked on HN > mpire's readme links to tqdm for progress bar) and noticed the nice stand-alone program.
As someone frequently using `pv` to visualize how much data went through a pipe when moving data around, I figured it'd be a good replacement (possibly more available, p... | open | 2023-08-12T06:55:36Z | 2023-08-12T06:55:36Z | https://github.com/tqdm/tqdm/issues/1494 | [] | martinetd | 0 |
rthalley/dnspython | asyncio | 307 | Benchmarking functionality | I looked through the examples and around on the websites and didn't find anything about this specifically, but please let me know if I missed it.
It would be nice for this library to have several benchmarking features. We would like to use it to keep tabs on the status of DNS servers, and and their performance in va... | closed | 2018-05-03T20:41:36Z | 2020-07-19T13:19:27Z | https://github.com/rthalley/dnspython/issues/307 | [
"Enhancement Request"
] | isaiahtaylor | 1 |
amdegroot/ssd.pytorch | computer-vision | 328 | about coco dataset labels | I have two questions.
1. why in the config.py, the number of classes in COCO is 201? Is that 81 or 82?
2. why coco_labels.py has two kinds of index?
Thanks! | open | 2019-04-25T06:51:02Z | 2019-11-07T03:20:34Z | https://github.com/amdegroot/ssd.pytorch/issues/328 | [] | qiufeng1994 | 4 |
huggingface/datasets | numpy | 6,982 | cannot split dataset when using load_dataset | ### Describe the bug
when I use load_dataset methods to load mozilla-foundation/common_voice_7_0, it can successfully download and extracted the dataset but It cannot generating the arrow document,
This bug happened in my server, my laptop, so as #6906 , but it won't happen in the google colab. I work for it for da... | closed | 2024-06-19T08:07:16Z | 2024-07-08T06:20:16Z | https://github.com/huggingface/datasets/issues/6982 | [] | cybest0608 | 3 |
nerfstudio-project/nerfstudio | computer-vision | 2,983 | NeRF-W | Hi, nice work! I am wondering if there is any chance I can run NeRF-W by ns-train directly? | open | 2024-03-05T02:53:32Z | 2024-03-14T11:06:08Z | https://github.com/nerfstudio-project/nerfstudio/issues/2983 | [] | cfeng16 | 1 |
Lightning-AI/pytorch-lightning | deep-learning | 20,562 | `BatchSizeFinder` with method `fit` and separate batch_size attributes for train and val (e.g., `self.train_batch_size` and `self.val_batch_size`) | ### Outline & Motivation
I suggest allowing `batch_arg_name` to accept a list of arg names. E.g. `tuner.scale_batch_size(..., batch_arg_name=["train_batch_size", "val_batch_size").
### Pitch
Fit uses both train and val dataloaders. They can have their own batch sizes.
### Additional context
_No response_
cc @lant... | open | 2025-01-24T18:21:04Z | 2025-01-24T18:21:26Z | https://github.com/Lightning-AI/pytorch-lightning/issues/20562 | [
"refactor",
"needs triage"
] | ibro45 | 0 |
PokeAPI/pokeapi | graphql | 1,119 | Little problem with some pokemons | Hi, thank you for the api!
I'm using it to some of my profile projects.
I'm using react and im having problems with 3 pokemons:
- gible
- ralts
- slakoth
error message: SyntaxError: Unexpected end of JSON input
Thank you very much!
| closed | 2024-08-12T16:16:13Z | 2024-10-05T14:30:14Z | https://github.com/PokeAPI/pokeapi/issues/1119 | [] | jrdelrio | 3 |
miguelgrinberg/python-socketio | asyncio | 477 | Client Randomly Disconnecting | Hey Guys,
I recently observe following error when using Python-Socketio as Server with four connected clients.
After some time (about ten minutes) one client disconnects with following log output:
```
INFO:werkzeug:127.0.0.1 - - [05/May/2020 10:23:28] "GET /socket.io/?transport=polling&EIO=3&sid=b91e51d9d9b043ab982... | closed | 2020-05-05T10:30:34Z | 2020-05-05T18:08:40Z | https://github.com/miguelgrinberg/python-socketio/issues/477 | [] | anon767 | 7 |
great-expectations/great_expectations | data-science | 10,603 | add_or_update_expectation suite is gone. will there be an update added to the SuiteFactory? | It looks in v 1.2.0 like suite factory only has CRUD methods: delete, add, and all. Will there be an update so that a suite of a particular name can be updated as was possible in previous versions where you could run add_or_update_expectation_suite on the great expectations context like was possible in version 0.18.17? | closed | 2024-10-30T22:45:22Z | 2025-02-14T16:53:55Z | https://github.com/great-expectations/great_expectations/issues/10603 | [
"feature-request"
] | isaacmartin1 | 5 |
comfyanonymous/ComfyUI | pytorch | 7,347 | [Errno 2] No such file or directory: 'none' is:issue | ### Your question
I am trying to run Live Portrait Lip Sync and I am getting the error message: [Errno 2] No such file or directory: 'none' is:issue Not sure what this means or how to address it. Thanks
### Logs
```powershell
```
### Other
_No response_ | closed | 2025-03-22T02:53:56Z | 2025-03-23T01:29:33Z | https://github.com/comfyanonymous/ComfyUI/issues/7347 | [
"User Support",
"Custom Nodes Bug"
] | mb0021 | 3 |
tensorflow/tensor2tensor | deep-learning | 1,121 | [Question] ASR Transformer performance vs. Google Speech-to-Text | ### Description
We used the ["ASR with Transformer" colab notebook](https://colab.research.google.com/github/tensorflow/tensor2tensor/blob/master/tensor2tensor/notebooks/asr_transformer.ipynb) which let us load the pre-trained checkpoints of the ASR Problems trained on librispeech and Common Voice datasets. We tried o... | open | 2018-10-09T13:09:11Z | 2019-12-15T16:46:50Z | https://github.com/tensorflow/tensor2tensor/issues/1121 | [] | mabergerx | 4 |
ets-labs/python-dependency-injector | flask | 556 | [delete] Core container as singletone for entire app | [App Package (Container) Diagramm](https://online.visual-paradigm.com/community/share/example-app-ub4sde1um)
The `CoreContainer` container contains:
```python
class CoreContainer( containers.DeclarativeContainer ):
arguments = providers.Resource( parse_arguments )
config = providers.Resource( parse... | closed | 2022-02-01T16:14:43Z | 2022-02-02T11:27:48Z | https://github.com/ets-labs/python-dependency-injector/issues/556 | [] | VasyaGaikin | 0 |
jupyterlab/jupyter-ai | jupyter | 289 | Configurable vector store |
### Problem
Users would like to use alternative retrievers for RAG instead of being restricted to using FAISS locally.
### Proposed Solution
1. Offer alternative retrievers, such as Pinecone, Kendra, OpenSearch, etc.
2. Offer retriever configurability to allow operators to switch between them.
### Additi... | open | 2023-07-24T16:23:14Z | 2023-07-27T23:45:33Z | https://github.com/jupyterlab/jupyter-ai/issues/289 | [
"enhancement",
"scope:RAG"
] | dlqqq | 0 |
RobertCraigie/prisma-client-py | pydantic | 811 | Generated Includes for Partial Types | ## Problem
When you use partial types as return values for functions that fetch from your database, you need to include all the relations of the corresponding partial type each time. That is very repetitive, and it would be nice if there were partial_types_includes generated, when generating the partial types.
For ex... | open | 2023-08-29T14:07:19Z | 2023-08-29T14:13:06Z | https://github.com/RobertCraigie/prisma-client-py/issues/811 | [] | TimonGaertner | 0 |
gunthercox/ChatterBot | machine-learning | 1,912 | how to delete one of the learned data training by list or corpus | hello, I want to delete just one of the learned data not all. But I got a problem,I cant't delete it.
it reported an error like this
| open | 2020-02-06T11:19:22Z | 2020-02-06T11:19:22Z | https://github.com/gunthercox/ChatterBot/issues/1912 | [] | qibinaoe | 0 |
dhaitz/mplcyberpunk | matplotlib | 36 | ModuleNotFoundError: No module named 'pkg_resources' | ```
Traceback (most recent call last):
File "C:\Users\Raj Dave\Desktop\New folder\main.py", line 5, in <module>
import mplcyberpunk
File "C:\Users\Raj Dave\AppData\Local\Programs\Python\Python312\Lib\site-packages\mplcyberpunk\__init__.py", line 4, in <module>
import pkg_resources
ModuleNotFoundError:... | closed | 2024-01-30T19:52:37Z | 2024-11-26T21:20:50Z | https://github.com/dhaitz/mplcyberpunk/issues/36 | [] | Rajdave69 | 3 |
chatopera/Synonyms | nlp | 126 | Title: nearby方法输出结果并不是最近的词 | <!-- Sponsor this project / 开源项目支持方 -->
<!-- Chatopera 云服务:低代码、无代码方式定制智能对话机器人,查看 https://bot.chatopera.com/ -->
<!-- 春松客服:快速获得好用的开源客服系统,查看 https://www.cskefu.com/ -->
##

## 预期行为
输出意思近,得分高的词语
#... | open | 2021-08-19T11:36:29Z | 2022-04-24T08:32:41Z | https://github.com/chatopera/Synonyms/issues/126 | [
"bug"
] | Miaotxy | 2 |
pytest-dev/pytest-django | pytest | 682 | pytest.ini directory is not set to rootdir in 3.4.4 | if I set a pytest.ini file and then run pytest in a subfolder it looks like the rootdir is not longer set to the directory that pytest.ini is in. This causes the path to DJANGO_SETTINGS_MODULE to no longer be found
so if I have a directory structure like this
root
--apps
--test
--project
----settings
------... | closed | 2018-12-18T16:09:08Z | 2019-02-26T15:18:44Z | https://github.com/pytest-dev/pytest-django/issues/682 | [
"bug"
] | danlittlejohn | 2 |
PaddlePaddle/PaddleHub | nlp | 1,700 | paddlehub1.8.1 训练模型出现报错 float division by zore | paddlepaddle版本试过1.7.2、1.8.0、1.8.4,在训练模型时,cls_task.finetune_and_eval() 此步出现报错 float division by zore。float division by zore,strategy,config均能跑通。
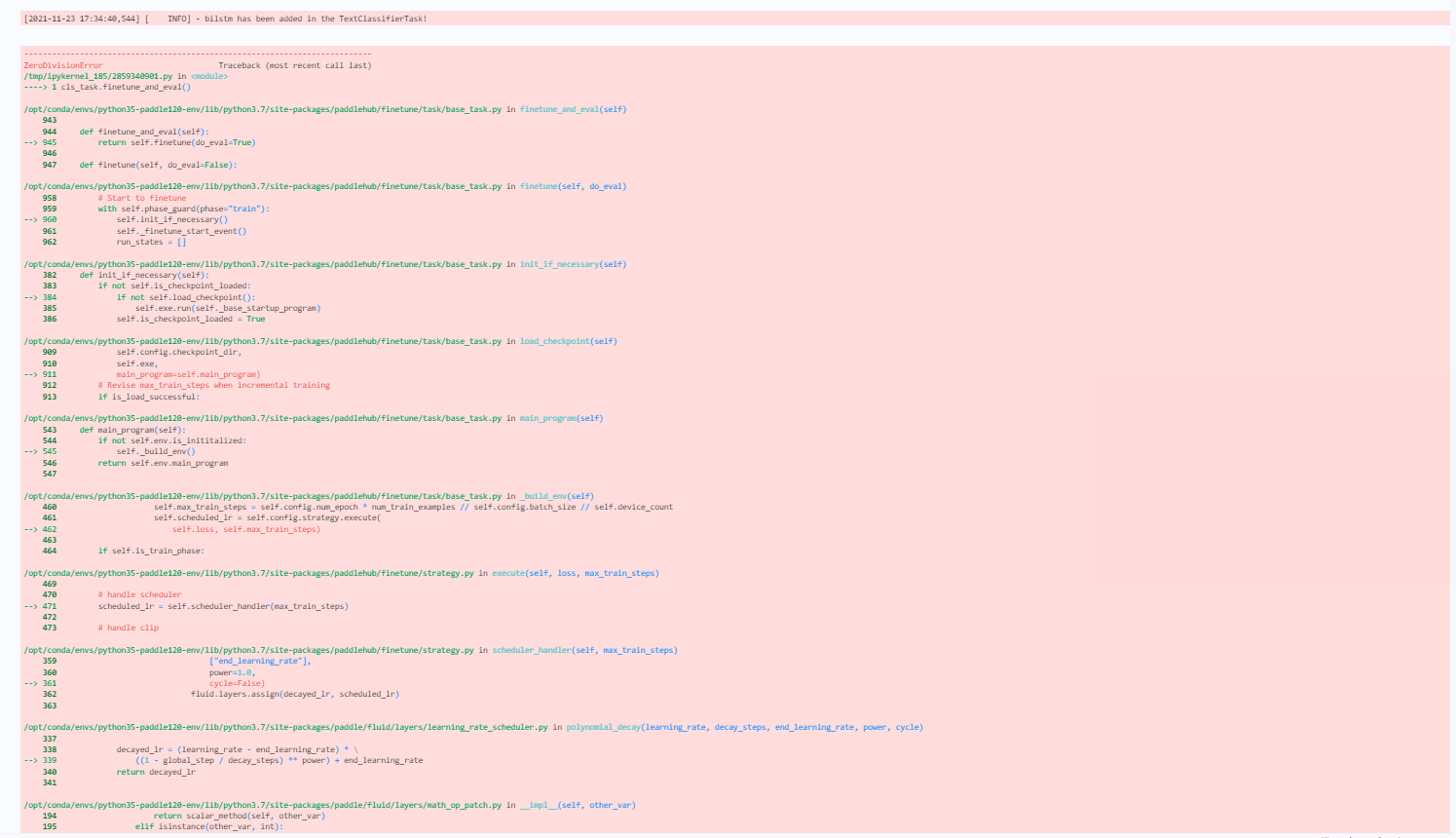
:
for t in res["itemList"]:
response.append(t)
```
"maxCursor" change to "cursor"
```maxCursor = res["cursor"]``` | closed | 2021-03-08T11:43:07Z | 2021-03-08T20:12:42Z | https://github.com/davidteather/TikTok-Api/issues/520 | [
"bug"
] | 11348 | 2 |
Evil0ctal/Douyin_TikTok_Download_API | fastapi | 54 | tiktok请求完数据为空 |
请求结果:
目标链接: https://www.tiktok.com/@jelenew_official/video/7108906863097367851
获取到的TikTok视频ID是7108906863097367851
正在请求API链接:https://api.tiktokv.com/aweme/v1/multi/aweme/detail/?aweme_ids=%5B7108906863097367851%5D
视频ID为: 7108906863097367851
正在请求API链接:https://api.tiktokv.com/aweme/v1/multi/aweme/detail/?aweme_ids... | closed | 2022-07-25T08:29:15Z | 2022-08-01T05:34:22Z | https://github.com/Evil0ctal/Douyin_TikTok_Download_API/issues/54 | [
"API Down",
"Fixed"
] | holmes1849082248 | 10 |
ultralytics/ultralytics | python | 19,770 | mac os M系列芯片怎么跑yolo11 | ### Search before asking
- [x] I have searched the Ultralytics YOLO [issues](https://github.com/ultralytics/ultralytics/issues) and [discussions](https://github.com/orgs/ultralytics/discussions) and found no similar questions.
### Question
/Users/tianlong/anaconda3/envs/yoloenv/bin/python /Users/tianlong/Desktop/de... | open | 2025-03-19T03:46:27Z | 2025-03-19T06:13:41Z | https://github.com/ultralytics/ultralytics/issues/19770 | [
"question",
"dependencies"
] | tianlong19910404 | 2 |
d2l-ai/d2l-en | computer-vision | 2,626 | Numpy version mismatch in d2l installation JAX | I am trying to install d2l for jax but
[Installation Link](https://d2l.ai/chapter_installation/index.html)
and I am getting Error in numpy package version when i install `d2l` by
```
pip install d2l==1.0.3
```
error:
```
ERROR: pip's dependency resolver does not currently take into account all the packages ... | open | 2024-11-22T17:42:22Z | 2024-11-22T17:42:22Z | https://github.com/d2l-ai/d2l-en/issues/2626 | [] | itahang | 0 |
autogluon/autogluon | computer-vision | 4,702 | [BUG] RaySystemError: System error: Failed to unpickle serialized exception | **Bug Report Checklist**
**Describe the bug**
<!-- A clear and concise description of what the bug is. -->
Using many columns ex: 11000 the ray reproduces an error related to catboost...
The error is confusing, because the root of the problem is the number of columns (many)... if you use, for example, only 1,00... | closed | 2024-11-28T14:11:29Z | 2025-01-13T11:37:56Z | https://github.com/autogluon/autogluon/issues/4702 | [
"bug",
"module: tabular",
"Needs Triage",
"priority: 0"
] | celestinoxp | 4 |
nalepae/pandarallel | pandas | 57 | Can't not call progress_apply(lambda x: function(x), axis=0) | The api onl work when axis=1. | closed | 2019-12-04T07:57:47Z | 2019-12-04T08:06:47Z | https://github.com/nalepae/pandarallel/issues/57 | [] | lhduc94 | 0 |
huggingface/datasets | computer-vision | 6,506 | Incorrect test set labels for RTE and CoLA datasets via load_dataset | ### Describe the bug
The test set labels for the RTE and CoLA datasets when loading via datasets load_dataset are all -1.
Edit: It appears this is also the case for every other dataset except for MRPC (stsb, sst2, qqp, mnli (both matched and mismatched), qnli, wnli, ax). Is this intended behavior to safeguard the t... | closed | 2023-12-16T22:06:08Z | 2023-12-21T09:57:57Z | https://github.com/huggingface/datasets/issues/6506 | [] | emreonal11 | 1 |
pytorch/pytorch | deep-learning | 148,937 | [AOTInductor]Only support one model instance when use AOTIModelPackageLoader load aot model? | when i use aoti model in cpp, i try to infer parallel by using multi threads and multi streams, like this:
```cpp
torch::inductor::AOTIModelPackageLoader loader("model.pt2");
torch::inductor::AOTIModelContainerRunner* runner = loader.get_runner();
for thread_id in threads:
// in different threads
auto outputs = r... | closed | 2025-03-11T01:59:53Z | 2025-03-19T02:18:33Z | https://github.com/pytorch/pytorch/issues/148937 | [
"triaged",
"oncall: pt2",
"oncall: export",
"module: aotinductor"
] | zzq96 | 3 |
aiogram/aiogram | asyncio | 1,477 | wrong TypeHint for ReplyKeyboardBuilder button | ### Checklist
- [X] I am sure the error is coming from aiogram code
- [X] I have searched in the issue tracker for similar bug reports, including closed ones
### Operating system
ubuntu 20
### Python version
3.11
### aiogram version
3.5
### Expected behavior
`request_user` and `request_chat` in `ReplyKeyboardB... | closed | 2024-05-03T11:41:23Z | 2024-05-03T12:10:08Z | https://github.com/aiogram/aiogram/issues/1477 | [
"bug"
] | HadiH2o | 0 |
aiortc/aiortc | asyncio | 303 | Strange `trackId` value in stats | I call `pc.addTransceiver()` with a `track` (generated by a `MediaPlayer`) whose `id` is "296ff180-3e3a-44de-b9f9-e0440d686e84".
However when I print the `transceiver.sender.getStats()` I see a strange `trackId` value into it:
```json
{
"bytesSent": 8643,
"id": "outbound-rtp_4575985168",
"ki... | closed | 2020-02-25T23:44:38Z | 2022-08-07T03:09:36Z | https://github.com/aiortc/aiortc/issues/303 | [
"stale"
] | ibc | 9 |
globaleaks/globaleaks-whistleblowing-software | sqlalchemy | 3,392 | Notification on password expiration | ### Proposal
A nice feature would be notification to users on password expiration - An idea?
/soren
### Motivation and context
We have had multiple requests from recipients, that would like to get a notification on expiration of passwords. This feature will actually make the recipients log in to the platform - ev... | open | 2023-03-20T13:21:28Z | 2023-03-20T15:17:46Z | https://github.com/globaleaks/globaleaks-whistleblowing-software/issues/3392 | [
"C: Backend",
"F: Notification",
"T: Feature"
] | schris-dk | 1 |
onnx/onnx | pytorch | 6,406 | ONNX Windows build relies on MS Store python, fails with official python launcher | # Bug Report
### Description
I'm using a Windows Arm-based PC, and the protobuf compilation stage of the pip installation fails with a fatal error. The error indicates that the mypy plugin is failing because the `python` command isn't found. I believe this is because I'm using the official download from Python do... | closed | 2024-09-30T21:11:35Z | 2024-11-04T19:19:47Z | https://github.com/onnx/onnx/issues/6406 | [
"bug"
] | petewarden | 2 |
fastapi/fastapi | api | 12,554 | Update docs include syntax for source examples | ### Privileged issue
- [X] I'm @tiangolo or he asked me directly to create an issue here.
### Issue Content
This is a good first contribution. :nerd_face:
The code examples shown in the docs are actual Python files. They are even tested in CI, that's why you can always copy paste an example and it will alw... | closed | 2024-10-26T13:06:48Z | 2024-11-18T02:45:54Z | https://github.com/fastapi/fastapi/issues/12554 | [
"good first issue"
] | tiangolo | 30 |
modelscope/data-juicer | streamlit | 375 | Efficient processing OPs for scanned images and pdf | ### Search before continuing 先搜索,再继续
- [X] I have searched the Data-Juicer issues and found no similar feature requests. 我已经搜索了 Data-Juicer 的 issue 列表但是没有发现类似的功能需求。
### Description 描述
There is a large amount of valuable data in the format of scanned images and PDFs. We can continuously discuss and list related proc... | closed | 2024-07-30T03:03:15Z | 2024-09-29T09:32:18Z | https://github.com/modelscope/data-juicer/issues/375 | [
"enhancement",
"dj:multimodal",
"stale-issue",
"dj:op"
] | yxdyc | 5 |
omar2535/GraphQLer | graphql | 11 | Add logging | Implement logging instead of using prints | closed | 2021-11-30T07:56:28Z | 2023-10-15T03:47:12Z | https://github.com/omar2535/GraphQLer/issues/11 | [
"➕enhancement"
] | omar2535 | 1 |
graphql-python/graphene | graphql | 1,226 | Proposal: Alternative API for mutations and subscriptions | # The problem
Currently the mutation api is quite clunky and not very intuitive: https://docs.graphene-python.org/en/latest/types/mutations/
Often users get confused about the `mutate` function being a static method and that you need to define a separate `Mutation` class that extends `ObjectType` with each mutati... | closed | 2020-07-12T12:40:24Z | 2020-11-09T17:05:07Z | https://github.com/graphql-python/graphene/issues/1226 | [
"✨ enhancement"
] | jkimbo | 13 |
nerfstudio-project/nerfstudio | computer-vision | 2,822 | how scale reularizition (use_scale_regularization) used in splatfacto? | how scale reularizition (use_scale_regularization) used in splatfacto?
https://github.com/nerfstudio-project/nerfstudio/blob/3bade3a9dcee19bf0bf28f40c3615891b9b9b444/nerfstudio/models/splatfacto.py#L853 | closed | 2024-01-25T13:37:23Z | 2024-01-25T19:12:36Z | https://github.com/nerfstudio-project/nerfstudio/issues/2822 | [] | pknmax | 1 |
coqui-ai/TTS | pytorch | 2,785 | [Bug] KeyError: 'use_phonemes' | ### Describe the bug
I tried to train a vocoder model and I tried to utilize it with tts-server but this throws KeyError: 'use_phonemes'.
This is the full error that I get:
(venv) PS D:\GitHub\Coqui> tts-server --config_path D:\GitHub\Coqui\models\LJSpeech-1.1\config.json --model_path D:\GitHub\Coqui\models\LJSpee... | closed | 2023-07-19T20:04:17Z | 2023-10-26T19:24:24Z | https://github.com/coqui-ai/TTS/issues/2785 | [
"bug",
"wontfix"
] | MVR3S | 4 |
NullArray/AutoSploit | automation | 438 | Unhandled Exception (6ee01e0a5) | Autosploit version: `2.2.3`
OS information: `Linux-4.19.0-kali1-amd64-x86_64-with-Kali-kali-rolling-kali-rolling`
Running context: `autosploit.py`
Error meesage: `[Errno 17] File exists: '/root/Downloads/AutoSploit/autosploit_out/2019-02-05_05h10m21s/'`
Error traceback:
```
Traceback (most recent call):
File "/root/Do... | closed | 2019-02-05T10:10:25Z | 2019-02-19T04:22:44Z | https://github.com/NullArray/AutoSploit/issues/438 | [] | AutosploitReporter | 0 |
pydata/xarray | numpy | 9,702 | inconsistent 1D interp on 1D and ND DataArrays with NaNs | ### What happened?
May be a duplicate of #5852, and as stated there, interp with nans isn't really supported. But I noticed that in some cases `xr.DataArray.interp` drops valid data if there are NaNs in the array, and that this behavior is different depending on the number of dimensions, even if only a single dimensio... | closed | 2024-10-31T12:41:26Z | 2024-12-02T15:31:58Z | https://github.com/pydata/xarray/issues/9702 | [
"bug",
"topic-interpolation"
] | delgadom | 6 |
InstaPy/InstaPy | automation | 6,038 | InstaPy installation on digital ocean | Hi community,
I am a big fan of instapy and would like to run it on a server.
I tried to follow the instruction: https://github.com/InstaPy/instapy-docs/blob/master/How_Tos/How_To_DO_Ubuntu_on_Digital_Ocean.md
I tried installing both browsers and also just Firefox.
I followed the instructions step by step. They... | closed | 2021-01-18T10:40:31Z | 2021-01-19T10:27:34Z | https://github.com/InstaPy/InstaPy/issues/6038 | [] | tomnewg | 1 |
httpie/cli | python | 1,401 | Bad filename when having non ASCII characters | ## Checklist
- [x] I've searched for similar issues.
- [x] I'm using the latest version of HTTPie.
---
## Minimal reproduction code and steps
1. save a dummy image as `天狗.png`
2. issue a multipart/form-data, such as: `https --offline --multipart https://example.org name='John Doe' file_field@/home/john/天狗... | open | 2022-05-16T03:55:37Z | 2024-10-30T10:53:32Z | https://github.com/httpie/cli/issues/1401 | [
"bug",
"new"
] | jackdeguest | 4 |
nonebot/nonebot2 | fastapi | 3,209 | Plugin: 他们在聊什么 | ### PyPI 项目名
nonebot-plugin-whats-talk-gemini
### 插件 import 包名
nonebot_plugin_whats_talk_gemini
### 标签
[{"label":"群聊总结","color":"#03f744"},{"label":"AI","color":"#feee06"},{"label":"Gemini","color":"#0609fe"}]
### 插件配置项
```dotenv
WT_AI_KEYS=["xxxxxx"]
```
### 插件测试
- [ ] 如需重新运行插件测试,请勾选左侧勾选框 | closed | 2024-12-26T13:11:00Z | 2024-12-28T15:38:32Z | https://github.com/nonebot/nonebot2/issues/3209 | [
"Plugin",
"Publish"
] | hakunomiko | 1 |
CTFd/CTFd | flask | 2,006 | API documentation is broken | I'm trying to access the API documentation page of CTFd, https://docs.ctfd.io/docs/api/Swagger%20UI, but it's broken and doesn't show Swagger UI.
I didn't find any other contacts, so I reported here. | closed | 2021-10-13T17:35:09Z | 2021-10-14T08:09:56Z | https://github.com/CTFd/CTFd/issues/2006 | [] | MrSuicideParrot | 2 |
SciTools/cartopy | matplotlib | 2,315 | Cannot import cartopy.feature (missing 'config') | ### Description
I'm a first time cartopy user. It installed just fine, but when I try to import cartopy.feature I get an import error related to the config file. As far as I can tell, this _should_ be made automatically by the `__init__` file, but this doesn't seem to be the case.
#### Code to reproduce
```
... | closed | 2024-01-18T17:47:19Z | 2024-01-18T18:04:58Z | https://github.com/SciTools/cartopy/issues/2315 | [] | Elizabethcase | 1 |
521xueweihan/HelloGitHub | python | 2,671 | 【开源自荐】cmd-wrapped,一个 Rust 编写的命令行记录分析 CLI | ## 推荐项目
- 项目地址:https://github.com/YiNNx/cmd-wrapped
- 类别:Rust
- 项目标题:一个 Rust 编写的命令行历史记录分析总结 CLI
- 项目描述:
一个 Rust 编写的命令行历史记录分析 CLI。它可以读取你的命令行操作历史记录,生成一份分析总结:
- 统计任意一年中的命令行活跃分布,如每日最活跃时段,以及常用命令等。
- 类 Github 的年度命令分布图
- 支持 Zsh, Bash, Fish 和 atuin
- 亮点:
通过 cmd-wrapped,你可以获得一份有趣的数据... | closed | 2024-01-08T09:32:50Z | 2024-01-26T02:26:46Z | https://github.com/521xueweihan/HelloGitHub/issues/2671 | [
"已发布",
"Rust 项目"
] | YiNNx | 2 |
Lightning-AI/pytorch-lightning | data-science | 20,152 | Typing for `_restricted_classmethod` (e.g. for `LightningModule.load_from_checkpoint`) has stopped working for mypy 1.11 | ### Bug description
The current release of mypy 1.11 (1.11.0 & 1.11.1) raises an error for methods decorated with `_restricted_classmethod` (e.g. for `LightningModule.load_from_checkpoint`). From what I have found, there used to be a workaround for tricking type checkers into expected behaviour (from `pytorch_lightn... | closed | 2024-08-02T09:05:10Z | 2024-08-03T13:50:03Z | https://github.com/Lightning-AI/pytorch-lightning/issues/20152 | [
"bug",
"help wanted",
"code quality",
"ver: 2.2.x"
] | maciejzj | 1 |
tableau/server-client-python | rest-api | 1,364 | Retrieving subscriptions failed for frequency = Daily, hour (interval) = 24 | tableau_server.subscriptions fails if any subscriptions have schedule frequency="Daily" & interval hours="24"
**Versions**
- Tableau Cloud: 2024.1.0
- Python version: 3.11.4
- TSC library version: 0.30
**To Reproduce**
tableau_server_obj.subscriptions
**Results**
ValueError: Invalid interval 24.0 not ... | closed | 2024-03-15T16:38:14Z | 2024-05-09T23:06:20Z | https://github.com/tableau/server-client-python/issues/1364 | [] | ryanstryker | 2 |
aeon-toolkit/aeon | scikit-learn | 2,660 | [ENH] Implement U-Shapelets clusterer | ### Describe the feature or idea you want to propose
It would be nice to have an implementation of the Unsupervised Shapelets (U-Shapelets)
https://ieeexplore.ieee.org/document/6413851
https://epubs.siam.org/doi/pdf/10.1137/1.9781611974010.101
https://link.springer.com/article/10.1007/s10618-015-0411-4
### Describe ... | open | 2025-03-20T13:14:14Z | 2025-03-20T13:59:05Z | https://github.com/aeon-toolkit/aeon/issues/2660 | [
"enhancement",
"clustering",
"implementing algorithms"
] | MatthewMiddlehurst | 1 |
sloria/TextBlob | nlp | 270 | Python 3.5: no mudule named textblob | When i start tihs program in python 3.5 there came an no module error. But in python 2 it works.
Why | closed | 2019-06-12T12:07:26Z | 2019-09-16T12:51:55Z | https://github.com/sloria/TextBlob/issues/270 | [] | Schokolino1 | 1 |
graphql-python/graphene | graphql | 637 | Docs link to 'Babel Relay Plugin' 404s | Hi!
On this page:
http://docs.graphene-python.org/projects/django/en/latest/introspection/
...the `Babel Relay Plugin` link (pointing at https://facebook.github.io/relay/docs/guides-babel-plugin.html) 404s.
I had a rummage to try and find the new location but didn't have much luck. | closed | 2017-12-29T18:00:13Z | 2017-12-29T18:10:12Z | https://github.com/graphql-python/graphene/issues/637 | [] | edmorley | 1 |
gunthercox/ChatterBot | machine-learning | 1,697 | ChatterBot can handle 10 million database entries? | I want to be able to save as much data as possible, so what is the maximum size that Chatterbot can handle? From what size does it get too slow? can handle 10 million entries? | closed | 2019-04-06T23:52:58Z | 2020-08-22T19:28:29Z | https://github.com/gunthercox/ChatterBot/issues/1697 | [
"answered"
] | EdgarAI | 2 |
kensho-technologies/graphql-compiler | graphql | 260 | NotImplementedError when calling toGremlin in FoldedContextField | Hey guys!
First of all, thanks a lot for this awesome work. I was testing the compiler in combination with Gremlin. The following GraphQL is mentioned in your Readme, but causes a NotImplementedError when trying to generate a Gremlin statement out of it:
```
Animal {
name @output(out_name: "name")
... | open | 2019-05-07T12:36:42Z | 2019-05-08T19:47:14Z | https://github.com/kensho-technologies/graphql-compiler/issues/260 | [
"enhancement",
"help wanted",
"good first issue"
] | dirkkolb | 5 |
iterative/dvc | machine-learning | 9,757 | dvc exp show: External s3 address not properly shown | # Bug Report
<!--
## dvc exp show: External s3 address not properly shown
-->
## Description
Hello,
I extended the example from https://github.com/iterative/dvc/issues/9713. Thank you so much for addressing that so quickly! This is much appreciated!
When now using an external s3 address `s3://<BUCKET>/... | open | 2023-07-25T00:01:48Z | 2024-10-23T08:06:33Z | https://github.com/iterative/dvc/issues/9757 | [
"bug",
"p2-medium",
"ui",
"A: experiments"
] | kpetersen-hf | 1 |
autogluon/autogluon | data-science | 4,457 | [BUG] refit_every_n_windows doesn't reduce training time for long time series | **Bug Report Checklist**
<!-- Please ensure at least one of the following to help the developers troubleshoot the problem: -->
- [x] I provided code that demonstrates a minimal reproducible example. <!-- Ideal, especially via source install -->
- [x] I confirmed bug exists on the latest mainline of AutoGluon via s... | open | 2024-09-06T13:37:42Z | 2025-01-06T10:32:15Z | https://github.com/autogluon/autogluon/issues/4457 | [
"bug: unconfirmed",
"Needs Triage",
"module: timeseries"
] | loek-scholt | 1 |
benbusby/whoogle-search | flask | 1,136 | [unable to start] <errno 13 Permission denied: whoogle.key> | **Describe the bug**
After a server reboot whoogle refuses to start. Docker logs indicate a permissions issue for the file whoogle.key which to the best of my knowledge has never been present in my deployment.
**To Reproduce**
Steps to reproduce the behavior:
1. Running Ubuntu 22.04 (latest patches as of Monday ... | closed | 2024-04-08T12:11:29Z | 2024-04-19T18:51:00Z | https://github.com/benbusby/whoogle-search/issues/1136 | [
"bug"
] | laoistom | 0 |
deeppavlov/DeepPavlov | nlp | 1,667 | Repl: KeyboardInterrupt, EOFError | - При выходе из интерактивного режима через нажатие ctrl-c или ctrl-d происходит падение скрипта с ошибкой.
- Пустая строка защитывается за контекст или вопрос. А по хорошему просто по новой запрашивать данные.
**DeepPavlov version** (you can look it up by running `pip show deeppavlov`):
1.4.0
**Python versi... | open | 2023-10-26T14:20:00Z | 2023-10-26T14:20:00Z | https://github.com/deeppavlov/DeepPavlov/issues/1667 | [
"bug",
"enhancement"
] | optk2k | 0 |
marimo-team/marimo | data-science | 3,531 | Feature request: Support collapsible sections in app mode | ### Description
Hi there,
first things first: Thank you _so much_ for conceiving marimo. It is really a pleasure to work with, full of so many details expanding upon but re-thinking Jupyter and other Notebook ideas and technologies. @WalBeh and me love it.
### Status quo
We are very pleased with the "collapsible sec... | open | 2025-01-21T23:10:28Z | 2025-01-24T02:06:00Z | https://github.com/marimo-team/marimo/issues/3531 | [
"enhancement"
] | amotl | 7 |
tensorpack/tensorpack | tensorflow | 918 | Add examples to tensorpack import | I vote for adding the examples to the tensorpack (at least add a `__init__.py`). I end up with copying or symlinking these files. But something along
```python
from tensorpack.examples.GAN import GanTrainer
from tensorpack.examples.OpticalFlow.FlowNet import Model
```
would be super useful. I know, these examp... | closed | 2018-10-04T13:13:16Z | 2019-06-26T00:03:50Z | https://github.com/tensorpack/tensorpack/issues/918 | [
"enhancement"
] | PatWie | 6 |
graphql-python/graphene | graphql | 1,496 | @include and @skip directives lead to irrelevant errors | I am unsure whether to put this under bugs or feature requests, because I would consider this to be both.
The directives `@include` and `@skip` are awesome concepts, but they miss the mark when a certain field is not queryable. That is, if the boolean variable on which the `@include` directive is conditioned evaluat... | closed | 2023-02-22T13:10:26Z | 2023-02-27T16:29:52Z | https://github.com/graphql-python/graphene/issues/1496 | [
"question"
] | DrumsnChocolate | 2 |
nteract/testbook | pytest | 97 | tb contains method | > Also, it would be nice if the following check is supported:
> 'happy_fraction' in tb
Implement a `__contains__` method which returns whether the variable/function is present in the notebook or not (in memory).
_Originally posted by @rohitsanj in https://github.com/nteract/testbook/issues/96#issuecomment-819540... | open | 2021-04-14T13:59:33Z | 2021-05-20T09:34:19Z | https://github.com/nteract/testbook/issues/97 | [
"enhancement",
"good first issue",
"sprint-friendly"
] | rohitsanj | 0 |
allenai/allennlp | pytorch | 5,064 | RuntimeError: CUDA error: CUBLAS_STATUS_EXECUTION_FAILED when calling cublasSgemm | ## Checklist
<!-- To check an item on the list replace [ ] with [x]. -->
- [x] I have verified that the issue exists against the `master` branch of AllenNLP.
- [x] I have read the relevant section in the [contribution guide](https://github.com/allenai/allennlp/blob/master/CONTRIBUTING.md#bug-fixes-and-new-featur... | closed | 2021-03-19T20:25:55Z | 2021-06-03T18:29:55Z | https://github.com/allenai/allennlp/issues/5064 | [
"bug"
] | nelson-liu | 17 |
unionai-oss/pandera | pandas | 1,071 | `validate` is slow with when coercing several hundreds columns. | **Describe the bug**
Validating against a `SchemaModel` with several hundred is used with coerce takes a lot of time, even if the dataframe is already valid. It doesn’t occur when there is no `coerce`.
- [x] I have checked that this issue has not already been reported.
- [x] I have confirmed this bug exists on... | open | 2023-01-20T13:05:07Z | 2025-02-14T22:11:55Z | https://github.com/unionai-oss/pandera/issues/1071 | [
"bug"
] | koalp | 4 |
dfki-ric/pytransform3d | matplotlib | 178 | Photogrammetry Example? | In the Readme, there is an example of a tree trunk with the computed position of the camera being displayed with the reconstructed 3D mesh. Does an example do this at the current moment? | closed | 2022-01-12T23:49:44Z | 2022-01-13T08:27:58Z | https://github.com/dfki-ric/pytransform3d/issues/178 | [] | stanleyshly | 0 |
jina-ai/clip-as-service | pytorch | 608 | word embedding for emojis | Hi, I'm using bc.encode() to generate the word vector, but the text contains some emojis. Does BertClient encode this emojis properly? Or should I remove the emojis before the word encoding? If they can be encoded properly, can you explain how briefly? Thank you! | open | 2020-12-03T13:29:25Z | 2020-12-03T13:29:25Z | https://github.com/jina-ai/clip-as-service/issues/608 | [] | jianGuoQiang | 0 |
BeanieODM/beanie | asyncio | 572 | [BUG] Fetch_Links & Inheritance dont work if Link not on Parent Class | Hi,
To avoid a circular import error with Links, I put my "Base_House" in a separate module than houses.py and house_parts.py. [In reality my modules are much larger]. This is now causing an issue whereby, when I use .find() on Base_House (which really is just an abstract class in a sense, it just has the common feat... | closed | 2023-05-24T13:29:56Z | 2023-07-10T02:10:34Z | https://github.com/BeanieODM/beanie/issues/572 | [
"Stale"
] | mg3146 | 3 |
fastapi/sqlmodel | fastapi | 337 | select specific columns returning Row objects instead of ORM class, load_only() doesn't work | ### First Check
- [X] I added a very descriptive title to this issue.
- [X] I used the GitHub search to find a similar issue and didn't find it.
- [X] I searched the SQLModel documentation, with the integrated search.
- [X] I already searched in Google "How to X in SQLModel" and didn't find any information.
- [X] I al... | open | 2022-05-11T20:40:49Z | 2024-08-07T23:42:11Z | https://github.com/fastapi/sqlmodel/issues/337 | [
"question"
] | johnatr | 2 |
scikit-optimize/scikit-optimize | scikit-learn | 489 | random_state not working? | Greetings!
If you run the following code:
``python
import numpy as np
from skopt import gp_minimize
def f(x):
#print(x)
return (np.sin(5 * x[0]) * (1 - np.tanh(x[0] ** 2)) *
np.random.randn() * 0.1)
res1 = forest_minimize(f, [(-2.0, 2.0)], random_state=np.random.RandomState(1))
r... | closed | 2017-08-29T11:04:02Z | 2017-08-29T11:55:43Z | https://github.com/scikit-optimize/scikit-optimize/issues/489 | [] | echo66 | 1 |
scrapy/scrapy | web-scraping | 5,842 | Values sent from api do not work in custom_settings | ### Description
I send some `custom_settings` values through api. While these values are successfully received in kwargs, they are not applied in crawler settings.
I wrote two different codes for the same thing, but I didn't succeed, and maybe scrapy has a bug in this part or I made a mistake.
### First code:
<... | closed | 2023-03-03T07:08:56Z | 2023-03-03T16:48:09Z | https://github.com/scrapy/scrapy/issues/5842 | [] | SardarDelha | 3 |
scrapy/scrapy | python | 5,763 | Cookie Injection won't work to all allowed url in LinkExtractor of CrawlSpider class | <!--
Thanks for taking an interest in Scrapy!
If you have a question that starts with "How to...", please see the Scrapy Community page: https://scrapy.org/community/.
The GitHub issue tracker's purpose is to deal with bug reports and feature requests for the project itself.
Keep in mind that by filing an iss... | closed | 2022-12-18T16:13:46Z | 2022-12-18T16:26:20Z | https://github.com/scrapy/scrapy/issues/5763 | [] | arcans1998 | 0 |
nolar/kopf | asyncio | 432 | [archival placeholder] | This is a placeholder for later issues/prs archival.
It is needed now to reserve the initial issue numbers before going with actual development (PRs), so that later these placeholders could be populated with actual archived issues & prs with proper intra-repo cross-linking preserved... | closed | 2020-08-18T20:06:28Z | 2020-08-18T20:06:29Z | https://github.com/nolar/kopf/issues/432 | [
"archive"
] | kopf-archiver[bot] | 0 |
noirbizarre/flask-restplus | flask | 613 | The name of the body can only be called payload? | In the generated document, the name of the body can only be called payload? I want change the name to make more sense.
```
@au.route('/authenticate')
@au.response(400, 'params error')
class Authenticate(Resource):
@au.doc('Get a accessT... | open | 2019-03-27T07:42:34Z | 2020-01-16T02:58:23Z | https://github.com/noirbizarre/flask-restplus/issues/613 | [
"Needed: Feedback"
] | ELvisZxc | 3 |
ploomber/ploomber | jupyter | 331 | Notebook/script templates generated by "ploomber scaffold" should contain a cell with the autoreload magic | If `ploomber scaffold` finds a `pipeline.yaml` if checks all `tasks[*].sources` and creates files for all tasks whose source is missing. e.g.,
```yaml
tasks:
- source: scripts/some-script.py
product: output.ipynb
```
If `scripts/some-script.py` does not exist, it creates one.
The created script ... | closed | 2021-09-04T15:19:46Z | 2021-10-08T18:49:53Z | https://github.com/ploomber/ploomber/issues/331 | [
"good first issue"
] | edublancas | 0 |
google-deepmind/graph_nets | tensorflow | 80 | Placeholders from data dicts with no edges | Hi,
I am currently trying out graph_nets and came across a problem, that I wasn't able to solve.
My graphs have node and global properties as well as senders/receivers, but use no edge properties at all. I tried to create a placeholder from such a data dict and provide it as input for my model, however, session.run... | closed | 2019-06-11T14:37:04Z | 2019-06-24T09:15:13Z | https://github.com/google-deepmind/graph_nets/issues/80 | [] | felbecker | 2 |
plotly/dash-core-components | dash | 443 | Upgrade to Plotly.js 1.44.1 | https://github.com/plotly/plotly.js/releases/tag/v1.44.1 | closed | 2019-01-23T20:24:42Z | 2019-01-25T03:02:34Z | https://github.com/plotly/dash-core-components/issues/443 | [
"dash-type-maintenance"
] | Marc-Andre-Rivet | 0 |
mirumee/ariadne | graphql | 1,188 | make_federated_schema is unable to parse repeatable directives | Hello, I'm extending our application to be integrated in GraphQL federation and I encountered a weird bug.
We are heavily utilizing directives in our schema, to extend resolving logic of our fields and we used to consume schema with ariadne function `make_executable_schema`. During integration of our service to Grap... | closed | 2024-07-08T12:26:19Z | 2024-11-19T16:46:17Z | https://github.com/mirumee/ariadne/issues/1188 | [] | ezopezo | 1 |
mage-ai/mage-ai | data-science | 5,264 | [BUG] MAGE_BASE_PATH issue with fonts (.ttf) | ### Mage version
0.9.72
### Describe the bug
Hello Mage team,
after upgrading from 0.9.71 to 0.9.72, MAGE_BASE_PATH is not working with fonts.
ex. MAGE_BASE_PATH=test
font URLs should be like https://mageurl-example.com/test/fonts/~~~, but frontend is loading fonts from https://mageurl-example.com/fonts/~~~
Tha... | closed | 2024-07-13T01:37:06Z | 2024-07-15T08:35:52Z | https://github.com/mage-ai/mage-ai/issues/5264 | [
"bug"
] | farmboy-dev | 0 |
akfamily/akshare | data-science | 5,355 | option_dce_daily输出有误 | 调用d1, d2 = akshare.option_dce_daily("玉米期权", trade_date="20241121"),d2输出如下,包含了多余玉米淀粉期权的信息
合约系列 隐含波动率(%)
0 c2501 13.18
1 c2503 10.13
2 c2505 14.28
3 c2507 15.57
4 c2509 13.44
5 c2511 10.9
6 cs2501 13.24
7 cs2503 10.99
8 cs2505 13.69
9 cs2507 13.6... | closed | 2024-11-21T11:42:16Z | 2024-11-21T12:21:50Z | https://github.com/akfamily/akshare/issues/5355 | [
"bug"
] | akkezhu | 1 |
pytorch/vision | computer-vision | 8,967 | Failed to install vision based on python 3.13t(free-threaded) on Windows OS | ### 🐛 Describe the bug
**Reproduce steps:**
[Windows 11 OS]
conda create -n nogil2 --override-channels -c conda-forge python-freethreading
conda activate nogil2
pip install torch torchvision torchaudio --pre --index-url https://download.pytorch.org/whl/nightly/cu128
ERROR: Cannot install torchvision==0.22.0.dev20250... | open | 2025-03-13T07:28:42Z | 2025-03-14T09:06:47Z | https://github.com/pytorch/vision/issues/8967 | [] | jameszhouyi | 2 |
plotly/jupyter-dash | jupyter | 54 | call-backs from Dash-plotly to JupyterLab | 
Hi,
I am trying to incorporate ArcGIS WebScenes in my Dash-plotly workflow.
I think I have two main options
1. To use iFrame and I louse the call back (As far as I know), wh... | open | 2021-02-12T06:16:52Z | 2021-02-12T06:16:52Z | https://github.com/plotly/jupyter-dash/issues/54 | [] | Shai2u | 0 |
OpenInterpreter/open-interpreter | python | 1,018 | Function calling broken with Azure OpenAI | ### Describe the bug
To confirm the model works, I asked it to write a haiku:

Next, to test the project, I asked it for _python script for hello world_
, where providing multiple properties at build/create will have the system check for those and fail if they don't match what is in the DB, unless non... | open | 2015-10-28T18:49:41Z | 2020-10-13T07:51:45Z | https://github.com/FactoryBoy/factory_boy/issues/241 | [] | stephenross | 1 |
ageitgey/face_recognition | python | 791 | Comparing new encoding with saved encodings from .dat file | * face_recognition version: probably 1.2.2
* Python version: 3.6.3
* Operating System: CentOS 7
### Description
Hello, I have access to lots of pictures and 64 cores CPU. I am not that great at python.
I noticed that face_recognition can't use multiple core while it is loading images, faces, encodings,it only ... | open | 2019-04-02T07:58:52Z | 2019-06-06T11:52:08Z | https://github.com/ageitgey/face_recognition/issues/791 | [] | franckyz0r | 4 |
httpie/cli | python | 1,331 | Sessions doesn't support multiple headers sharing the same name | ## Checklist
- [x] I've searched for similar issues.
- [x] I'm using the latest version of HTTPie.
---
## Minimal reproduction code and steps
Run `http --offline : hello:world hello:people --session test`
## Current result
The header `hello: world` is not saved in the session file
```
$ cat ~/.http... | closed | 2022-03-19T19:02:31Z | 2022-04-03T13:48:31Z | https://github.com/httpie/cli/issues/1331 | [
"enhancement",
"needs product design"
] | ducaale | 0 |
ultralytics/ultralytics | pytorch | 18,982 | YOLOv11 vs SSD performance on 160x120 infrared images | ### Search before asking
- [x] I have searched the Ultralytics YOLO [issues](https://github.com/ultralytics/ultralytics/issues) and [discussions](https://github.com/orgs/ultralytics/discussions) and found no similar questions.
### Question
Hello,
in our previous project, we successfully implemented object detection... | open | 2025-02-03T20:01:01Z | 2025-02-27T22:16:34Z | https://github.com/ultralytics/ultralytics/issues/18982 | [
"question",
"detect",
"embedded"
] | BigMuscle85 | 32 |
fastapi-users/fastapi-users | asyncio | 188 | user handling with HTML template | hello, how do I pass a ValidationError or 400 error back to the login.html template?
for example if user enters empty username or incorrect password, they just get a text return,
```
1 validation error for Request
body -> password
field required (type=value_error.missing)
Validation Error:
```
How can... | closed | 2020-05-17T18:41:44Z | 2023-05-09T12:46:41Z | https://github.com/fastapi-users/fastapi-users/issues/188 | [
"question"
] | perfecto25 | 4 |
WeblateOrg/weblate | django | 14,174 | Batch triggering of source checks | ### Describe the problem
The source checks like MultipleFailingCheck or LongUntranslatedCheck are currently the slowest paths in the `update_checks` task. These could probably be done using a single SQL query instead of doing a query per string.
### Describe the solution you would like
Similarly to the target checks... | open | 2025-03-12T09:05:37Z | 2025-03-21T12:36:55Z | https://github.com/WeblateOrg/weblate/issues/14174 | [
"enhancement",
"Area: Quality checks"
] | nijel | 3 |
thp/urlwatch | automation | 282 | Xpath filtering doesn't work with text() and node() | Example:
That will gives me the first result in a google's search:
```
name: "Test"
url: "https://www.google.com/search?q=something&tbm=nws&source=lnt&tbs=qdr:h&biw=1920&bih=957&dpr=1"
filter: xpath://div[@class='g'][1]/descendant::a/text()
```
Expected Behavior: Monitoring the changes on the first result`s... | closed | 2018-09-19T18:20:15Z | 2018-10-23T18:24:19Z | https://github.com/thp/urlwatch/issues/282 | [] | eduardohayashi | 0 |
strawberry-graphql/strawberry | graphql | 2,813 | Exception handling with status code . As graphql-strawberry always return status code as 200 OK. | <!---I am new to graphql how to handle status code in graphql-strawberry-->
## Feature Request Type
- [ ] Core functionality
- [ ] Alteration (enhancement/optimization) of existing feature(s)
- [ ] New behavior
## Description
<!-- A few sentences describing what it is. -->
def default_resolver(root, fiel... | closed | 2023-06-06T08:08:09Z | 2025-03-20T15:56:12Z | https://github.com/strawberry-graphql/strawberry/issues/2813 | [] | itsckguru | 1 |
pyg-team/pytorch_geometric | pytorch | 8,894 | Torch 2.2 | ### 😵 Describe the installation problem
Is pyg compatible with torch 2.2 yet? The readme says test passing 2.2, but on conda channel it still requires torch to be highest at 2.1.
### Environment
* PyG version:
* PyTorch version: 2.2
* OS:
* Python version:
* CUDA/cuDNN version:
* How you installed PyTorch and ... | open | 2024-02-10T16:12:27Z | 2024-02-10T17:09:34Z | https://github.com/pyg-team/pytorch_geometric/issues/8894 | [
"installation"
] | Will-Zhao0 | 1 |
Yorko/mlcourse.ai | scikit-learn | 596 | links are broken on mlcourse.ai | example: https://festline.github.io/notebooks/blob/master/jupyter_english/topic02_visual_data_analysis/topic2_visual_data_analysis.ipynb?flush_cache=true | closed | 2019-05-30T02:15:44Z | 2019-06-27T14:54:06Z | https://github.com/Yorko/mlcourse.ai/issues/596 | [] | nickcorona | 4 |
pandas-dev/pandas | data-science | 61,018 | BUG: df.plot() "Subplots" changes behavior of how values are stacked using the "Stacked" property | ### Pandas version checks
- [x] I have checked that this issue has not already been reported.
- [x] I have confirmed this bug exists on the [latest version](https://pandas.pydata.org/docs/whatsnew/index.html) of pandas.
- [x] I have confirmed this bug exists on the [main branch](https://pandas.pydata.org/docs/dev/ge... | open | 2025-02-28T05:41:34Z | 2025-03-02T13:55:10Z | https://github.com/pandas-dev/pandas/issues/61018 | [
"Bug",
"Visualization"
] | eicchen02 | 3 |
microsoft/MMdnn | tensorflow | 281 | Error when converting Pytorch model to IR | Platform (like ubuntu 16.04/win10):
ubuntu 16.04
Python version:
python 3.5.2
Source framework with version (like Tensorflow 1.4.1 with GPU):
Pytorch 0.4.0 with GPU
Destination framework with version (like CNTK 2.3 with GPU):
caffe
Pre-trained model path (webpath or webdisk path)... | closed | 2018-07-03T03:11:27Z | 2019-10-10T00:29:04Z | https://github.com/microsoft/MMdnn/issues/281 | [] | Miranda0920 | 10 |
facebookresearch/fairseq | pytorch | 5,111 | how to finetune a language pair that not in the language dictionary | ## ❓ Questions and Help
### Before asking:
1. search the issues.
2. search the docs.
<!-- If you still can't find what you need: -->
#### What is your question?
hello, I'm using m2m100 model and i want finetune a language pair that not in the language dictionary ,what should i do ?
#### Code
<!-- Ple... | open | 2023-05-22T07:22:40Z | 2023-05-22T07:22:40Z | https://github.com/facebookresearch/fairseq/issues/5111 | [
"question",
"needs triage"
] | ecoli-hit | 0 |
proplot-dev/proplot | data-visualization | 381 | Error when passing keyword arguments like edgecolor and linewidth to Quiver function | <!-- Thanks for helping us make proplot a better package! If this is a bug report, please use the template provided below. If this is a feature request, you can delete the template text (just try to be descriptive with your request). -->
### Description
Absolutely love the project, thank you for all your efforts wi... | open | 2022-08-02T00:26:14Z | 2023-03-29T08:42:43Z | https://github.com/proplot-dev/proplot/issues/381 | [
"bug"
] | jordanbrook | 0 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.