repo_name stringlengths 9 75 | topic stringclasses 30
values | issue_number int64 1 203k | title stringlengths 1 976 | body stringlengths 0 254k | state stringclasses 2
values | created_at stringlengths 20 20 | updated_at stringlengths 20 20 | url stringlengths 38 105 | labels listlengths 0 9 | user_login stringlengths 1 39 | comments_count int64 0 452 |
|---|---|---|---|---|---|---|---|---|---|---|---|
liangliangyy/DjangoBlog | django | 647 | Cfe | closed | 2023-03-30T12:33:13Z | 2023-03-31T03:01:43Z | https://github.com/liangliangyy/DjangoBlog/issues/647 | [] | networknice | 0 | |
iperov/DeepFaceLab | deep-learning | 5,228 | "data_src faceset extract" Failing | Choose one or several GPU idxs (separated by comma).
[CPU] : CPU
[0] : GeForce GTX 960M
[0] Which GPU indexes to choose? :
0
[wf] Face type ( f/wf/head ?:help ) :
wf
[0] Max number of faces from image ( ?:help ) :
0
[512] Image size ( 256-2048 ?:help ) :
512
[90] Jpeg quality ( 1-100 ?:help ) :
90
... | open | 2021-01-02T11:18:19Z | 2023-06-08T21:53:08Z | https://github.com/iperov/DeepFaceLab/issues/5228 | [] | adam-eme | 2 |
polakowo/vectorbt | data-visualization | 40 | Returning None from order_func_nb causes numba TypingError | Error:
```
TypingError: Failed in nopython mode pipeline (step: nopython frontend)
Failed in nopython mode pipeline (step: nopython frontend)
Unknown attribute 'size' of type none
```
Code (taken from example with modified order_func_nb)
```python
import vectorbt as vbt
from vectorbt.portfolio.enums impor... | closed | 2020-08-26T03:04:49Z | 2020-09-16T22:31:56Z | https://github.com/polakowo/vectorbt/issues/40 | [] | Ziink | 5 |
keras-team/keras | data-science | 20,259 | ValueError: File not found: filepath=Backend\C4__256g_000040000.keras. Please ensure the file is an accessible `.keras` zip file. | Error:
Please ensure the file is an accessible '.keras' zip file
Keras Version: 3.5.0
tensorflow Version: 2.16.1
I don't have GPU please I need solution
Code:
import numpy as np
# from keras.models import load_model
# from keras.models import load_model
import matplotlib.pyplot as plt
from numpy import ... | open | 2024-09-15T07:32:16Z | 2024-09-20T18:02:08Z | https://github.com/keras-team/keras/issues/20259 | [
"type:Bug"
] | ananthanarayanan431 | 6 |
ivy-llc/ivy | tensorflow | 28,621 | Fix Frontend Failing Test: torch - search.paddle.argsort | To-do List: https://github.com/unifyai/ivy/issues/27498 | closed | 2024-03-17T14:16:17Z | 2024-03-25T12:44:27Z | https://github.com/ivy-llc/ivy/issues/28621 | [
"Sub Task"
] | ZJay07 | 0 |
kizniche/Mycodo | automation | 797 | Error setup -> data after removing pi default admin user | Changed raspberrypi admin user
Traceback (most recent call last):
File "/home/[user]/Mycodo/env/lib/python3.7/site-packages/flask/app.py", line 2447, in wsgi_app
response = self.full_dispatch_request()
File "/home/[user]/Mycodo/env/lib/python3.7/site-packages/flask/app.py", line 1952, in full_dispatch_req... | closed | 2020-07-25T01:11:46Z | 2020-07-27T18:29:22Z | https://github.com/kizniche/Mycodo/issues/797 | [] | stardawg | 1 |
chezou/tabula-py | pandas | 237 | Read_PDF generating single item list | <!--- Provide a general summary of your changes in the Title above -->
When using the tabula.read_pdf for an online pdf files, I expected a dataframe output but seemed to receive a list with the entire table data as one item in the list.
<!-- Write the summary of your issue here -->
# Check list before submit
... | closed | 2020-04-27T15:56:10Z | 2020-06-04T12:08:09Z | https://github.com/chezou/tabula-py/issues/237 | [
"not a bug"
] | clarakheinz | 6 |
pallets-eco/flask-sqlalchemy | flask | 384 | Provide a max_per_page parameter for pagination | Right now the default for per_page is 20 unless otherwise specified. I also see that the code makes sure it is no larger than the total items.
To prevent a request asking for too many items at once, it would be convenient if Pagination could be initialized with a maximum items per page (max_per_page). Then the supplie... | closed | 2016-03-25T20:30:51Z | 2020-12-05T20:55:31Z | https://github.com/pallets-eco/flask-sqlalchemy/issues/384 | [
"pagination"
] | dmulter | 1 |
stanfordnlp/stanza | nlp | 784 | [QUESTION] How to evaluate pre-trained NER model on my domain specific text? | I am trying to get F1 scores for the pre-trained english model on my specific text domain without doing any training.
The docs mention the following command:
`
python -m stanza.utils.training.run_ete ${corpus} --score_${split}
`
However as I dont want to do any training, how can I evaluate the model as is?
I've... | closed | 2021-08-08T13:18:37Z | 2021-08-11T12:09:16Z | https://github.com/stanfordnlp/stanza/issues/784 | [
"question"
] | Pwgeorge-Py | 2 |
pytest-dev/pytest-django | pytest | 696 | DB fails to clean up after tests, says db is used by another user | This is basically a repost of my question from stackoverflow. I hope it will be more useful here.
I have a Django application, and I'm trying to test it using pytest and pytest-django. However, quite often, when the tests finish running, I get the error that the database failed to be deleted: DETAIL: There is 1 oth... | open | 2019-01-28T08:00:40Z | 2024-03-27T15:21:28Z | https://github.com/pytest-dev/pytest-django/issues/696 | [] | ibolit | 1 |
allenai/allennlp | data-science | 4,816 | Make image sets easily accessible | We don't want to automatically download large image sets with `cached_path()`, but we at least want to make it very easy to download them manually (download one archive, extract it, set the image path).
- [x] VQA
- [x] GQA
- [x] SNLI-VE | closed | 2020-11-24T00:27:16Z | 2021-01-04T21:39:11Z | https://github.com/allenai/allennlp/issues/4816 | [] | dirkgr | 2 |
explosion/spaCy | deep-learning | 13,707 | numpy requirements/compatibility | I noticed that conda-forge is struggling with some of the current numpy specifications to the point that they're having to patch the requirements (https://github.com/conda-forge/thinc-feedstock/pull/123).
To improve this and restore numpy v1 compatibility, could you consider using numpy's suggested build+install req... | open | 2024-12-05T09:13:23Z | 2024-12-16T06:31:18Z | https://github.com/explosion/spaCy/issues/13707 | [] | adrianeboyd | 2 |
JaidedAI/EasyOCR | pytorch | 1,166 | Bug in craft training in make_char_box.py | I am hitting a divide by zero exception in `make_char_box.py` when training the craft recognition model where it tries to crop an image by the bounding box, but the bounding box has an area of zero.
Stack trace below:
```
> /home/connoourke/bin/src/EasyOCRDev/EasyOCR/trainer/craft/data/pseudo_label/make_charbox.... | open | 2023-11-16T12:02:51Z | 2024-01-31T18:31:01Z | https://github.com/JaidedAI/EasyOCR/issues/1166 | [] | connorourke | 1 |
awesto/django-shop | django | 193 | Error in docs | Hi! I'm new to django shop, and I think I've found an error in the docs.
At http://django-shop.readthedocs.org/en/latest/getting-started.html#adding-taxes in the code says def add_extra_cart_price_field(self, cart): and it should say def get_extra_cart_price_field(self, cart):
If it isnt an error i would love to hear... | closed | 2012-11-03T00:44:19Z | 2012-12-06T13:07:26Z | https://github.com/awesto/django-shop/issues/193 | [] | singold | 0 |
mwaskom/seaborn | data-visualization | 2,886 | Area mark raises with log x scale | ```python
so.Plot([1, 10, 100, 1000], [1, 2, 4, 3]).add(so.Area()).scale(x="log")
```
Raises:
<details>
```python-traceback
/Users/mwaskom/miniconda3/envs/seaborn-py39-latest/lib/python3.9/site-packages/matplotlib/transforms.py:2664: RuntimeWarning: invalid value encountered in double_scalars
self._mtx = n... | closed | 2022-07-05T00:28:58Z | 2022-07-14T00:15:25Z | https://github.com/mwaskom/seaborn/issues/2886 | [
"bug",
"objects-mark"
] | mwaskom | 1 |
zappa/Zappa | django | 473 | [Migrated] Add documentation to the README for Cognito triggers | Originally from: https://github.com/Miserlou/Zappa/issues/1268 by [millarm](https://github.com/millarm)
<!--
Before you submit this PR, please make sure that you meet these criteria:
* Did you read the [contributing guide](https://github.com/Miserlou/Zappa/#contributing)?
* If this is a non-trivial commit, di... | closed | 2021-02-20T08:35:20Z | 2024-04-13T16:18:40Z | https://github.com/zappa/Zappa/issues/473 | [
"needs-user-testing",
"no-activity",
"auto-closed"
] | jneves | 2 |
tensorflow/tensor2tensor | deep-learning | 1,310 | How to evaluate the transformer on the summarization task by using the tensor2tensor? | ### Description
e.g. CNN/dailymail dataset. I know how to train the transformer on the dataset, but I don't know how to evaluate the performance of the transformer on the task.
| open | 2018-12-18T05:46:58Z | 2019-07-25T08:57:18Z | https://github.com/tensorflow/tensor2tensor/issues/1310 | [] | zhaoguangxiang | 5 |
sloria/TextBlob | nlp | 75 | pip install fails with latest setuptools version | A recent change in `setuptools` causes `nltk` install to fail (see https://github.com/nltk/nltk/issues/824).
Temporary workaround for users who have `setuptools>=10.0` installed in their python environment (default for all newly created environments using `get_pip.py`):
```
pip install setuptools==9.1
# this will au... | closed | 2015-01-01T15:08:55Z | 2015-01-13T18:41:38Z | https://github.com/sloria/TextBlob/issues/75 | [] | markuskiller | 1 |
PokeAPI/pokeapi | api | 188 | evolution-chain uses inconsistent schema for evolution_details | Best example to compare is evolution-chain/1 and evolution-chain/34.
The evolution_details property for evolves_to can be null, a single value, or an array. I propose wrapping all values in an array so that the expected type remains consistent.
| closed | 2016-05-12T13:29:12Z | 2016-05-24T13:56:16Z | https://github.com/PokeAPI/pokeapi/issues/188 | [] | zberk | 4 |
netbox-community/netbox | django | 18,318 | Mentoning of manufacturer is double in "Module Types/Interface" | ### Deployment Type
NetBox Cloud
### Triage priority
N/A
### NetBox Version
v4.2.0
### Python Version
3.10
### Steps to Reproduce
1. Open A module type that contains interfaces. ex: [https://demo.netbox.dev/dcim/module-types/1/](https://demo.netbox.dev/dcim/module-types/1/)
2. Click on the interface tab: [htt... | closed | 2025-01-07T10:39:29Z | 2025-01-07T15:28:28Z | https://github.com/netbox-community/netbox/issues/18318 | [
"type: bug",
"status: accepted",
"severity: low"
] | elixdreamer | 0 |
litestar-org/litestar | asyncio | 3,481 | Bug: test failures | ### Description
The tests ran right around midnight UTC time.
```
________________ test_default_serializer[v1-condate-2024-05-08] ________________
[gw0] linux -- Python 3.8.18 /home/runner/work/litestar/litestar/.venv/bin/python
model = ModelV1(custom_str='', custom_int=0, custom_float=0.0, custom_list=[], custo... | closed | 2024-05-09T00:21:59Z | 2025-03-20T15:54:42Z | https://github.com/litestar-org/litestar/issues/3481 | [
"Bug :bug:"
] | peterschutt | 0 |
Neoteroi/BlackSheep | asyncio | 128 | Typo in the documentation(middleware) | Hey @RobertoPrevato in the middlewares documentation we should fix code-example
https://www.neoteroi.dev/blacksheep/middlewares/
Handler `home` should receive `request` argument. | closed | 2021-05-11T08:13:07Z | 2021-05-11T08:35:39Z | https://github.com/Neoteroi/BlackSheep/issues/128 | [] | myusko | 2 |
marshmallow-code/marshmallow-sqlalchemy | sqlalchemy | 159 | SQLAlchemy Sessions Not Passed to Nested Fields | https://marshmallow-sqlalchemy.readthedocs.io/en/latest/recipes.html#smart-nested-field
Using this recipe as a base, I was writing a function to assist with serialization/de-serialization that looks somewhat like the below snippet:
```python
class MySchema(ModelSchema):
children = SmartNested(ChildModel)
... | closed | 2018-11-25T00:06:43Z | 2019-08-15T14:24:43Z | https://github.com/marshmallow-code/marshmallow-sqlalchemy/issues/159 | [] | ducharmemp | 2 |
flairNLP/flair | nlp | 2,690 | [Feature/enhancement] Documentation | **Is your feature/enhancement request related to a problem? Please describe.**
After reading the FLERT paper I attempted to implement it on my own data/labels. The quick start guide & tutorials provide a good starting point but lack significant documentation. Very few classes/functions have any documentation at all,... | closed | 2022-03-28T10:51:52Z | 2022-09-09T02:02:40Z | https://github.com/flairNLP/flair/issues/2690 | [
"wontfix"
] | torbenal | 1 |
polakowo/vectorbt | data-visualization | 399 | How to plot indicator? | 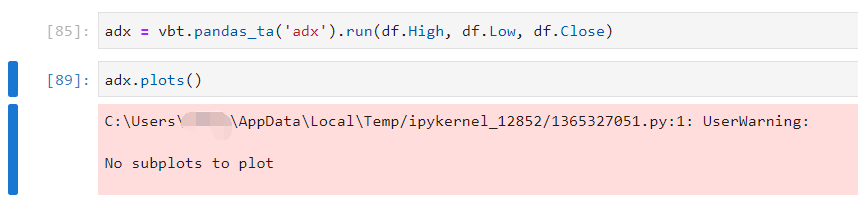
| closed | 2022-03-01T11:58:56Z | 2023-04-28T06:32:46Z | https://github.com/polakowo/vectorbt/issues/399 | [] | GF-Huang | 3 |
robotframework/robotframework | automation | 5,121 | Request to set implicit wait as a basic parameter to a keyword | Our team uses 1.5 min as a basic implicit wait. Sometimes, we need to get results immediately (e.g., is there an element on the page). Since implicit wait is set globally we are failing to structurize scripts nicely and are forced to proceed with the following workaround where we change the global wait, run the keyword... | closed | 2024-04-25T14:19:15Z | 2024-04-26T10:54:36Z | https://github.com/robotframework/robotframework/issues/5121 | [] | vyvy3 | 1 |
vllm-project/vllm | pytorch | 15,144 | [Bug] Mismatch between `get_multimodal_embedding` output and `PlaceholderRange` | In V1, we expect the output of `get_multimodal_embedding` to correspond to the `PlaceholderRange`, which is in turn constructed based on `PromptUpdateDetails.features`. However, the current V1 code doesn't validate this, causing the model to crash during inference when under high load (e.g. #14897, #14963).
From a qui... | open | 2025-03-19T16:53:23Z | 2025-03-24T13:56:02Z | https://github.com/vllm-project/vllm/issues/15144 | [
"bug",
"help wanted",
"v1",
"multi-modality"
] | DarkLight1337 | 0 |
zalandoresearch/fashion-mnist | computer-vision | 59 | Benchmark: Conv Net - Accuracy: 92.56% | Tried this network topology that can be summarized as follows:
- Convolutional layer with 32 feature maps of size 5×5.
- Pooling layer taking the max over 2*2 patches.
- Convolutional layer with 64 feature maps of size 5×5.
- Pooling layer taking the max over 2*2 patches.
- Convolutional layer with 128 feature m... | closed | 2017-09-07T09:28:48Z | 2017-09-07T10:50:17Z | https://github.com/zalandoresearch/fashion-mnist/issues/59 | [
"benchmark"
] | umbertogriffo | 2 |
kaliiiiiiiiii/Selenium-Driverless | web-scraping | 85 | cdp_socket.exceptions.CDPError: {'code': -32000, 'message': 'Could not find node with given id'} | > ```python
> from selenium_driverless import webdriver
> from selenium_driverless.types.by import By
> import asyncio
> async def main():
> options = webdriver.ChromeOptions()
> async with webdriver.Chrome(options=options) as driver:
> await driver.get('https://www.google.com/', wait_load=True)
... | closed | 2023-10-07T08:28:35Z | 2023-10-12T03:19:47Z | https://github.com/kaliiiiiiiiii/Selenium-Driverless/issues/85 | [] | samyeid | 4 |
AntonOsika/gpt-engineer | python | 530 | Using gpt-engineer with Azure OpenAI |
Hi, I am trying to test gpt-engineer by using Azure OpenAI but I am getting authentication error. I have added all the additional details that are required for the Azure OpenAI like api_base url, model, etc. in the python file ai.py in the gpt_engineer folder. Am I missing out something can you please help me out wi... | closed | 2023-07-13T11:01:49Z | 2023-10-05T07:44:45Z | https://github.com/AntonOsika/gpt-engineer/issues/530 | [] | RenukaMane | 6 |
huggingface/datasets | pytorch | 7,473 | Webdataset data format problem | ### Describe the bug
Please see https://huggingface.co/datasets/ejschwartz/idioms/discussions/1
Error code: FileFormatMismatchBetweenSplitsError
All three splits, train, test, and validation, use webdataset. But only the train split has more than one file. How can I force the other two splits to also be interpreted ... | closed | 2025-03-21T17:23:52Z | 2025-03-21T19:19:58Z | https://github.com/huggingface/datasets/issues/7473 | [] | edmcman | 1 |
onnx/onnx | scikit-learn | 6,131 | Model zoo test failures | https://github.com/onnx/onnx/actions/runs/8962117051/job/24610534787#step:7:943
```
--------------Time used: 0.31854987144470215 secs-------------
In all 184 models, 4 models failed, 25 models were skipped
ResNet-preproc failed because: Field 'type' of 'value_info' is required but missing.
VGG 16-bn failed becau... | open | 2024-05-06T16:23:02Z | 2024-05-07T23:43:57Z | https://github.com/onnx/onnx/issues/6131 | [] | justinchuby | 1 |
OpenGeoscience/geonotebook | jupyter | 129 | Annotation bounds passed to index are always in WGS84 | These need to be reprojected to the native SRS of the image before being passed to Rasterio's index function. | closed | 2017-07-19T19:19:31Z | 2017-07-21T14:35:10Z | https://github.com/OpenGeoscience/geonotebook/issues/129 | [] | dorukozturk | 0 |
pytorch/vision | machine-learning | 8,956 | `all_ops` argument of `AugMix()` accepts many types of values | ### 🐛 Describe the bug
[The doc](https://pytorch.org/vision/main/generated/torchvision.transforms.v2.AugMix.html) of `AugMix()` says that `all_ops` parameter is `bool` as shown below:
> Parameters:
> ...
> - all_ops ([bool](https://docs.python.org/3/library/functions.html#bool), optional) – Use all operations (inclu... | open | 2025-03-07T14:28:44Z | 2025-03-07T14:28:44Z | https://github.com/pytorch/vision/issues/8956 | [] | hyperkai | 0 |
stanford-oval/storm | nlp | 99 | Fixed Timeout in WebPageHelper Could Lead to Incomplete Data Retrieval | # Fixed Timeout in WebPageHelper Could Lead to Incomplete Data Retrieval
## Description
In the file `utils.py`, the `WebPageHelper` class uses a fixed timeout of 4 seconds for all HTTP requests:
```python
res = self.httpx_client.get(url, timeout=4)
```
This fixed timeout can lead to issues with data retriev... | closed | 2024-07-22T06:03:30Z | 2025-03-08T09:09:48Z | https://github.com/stanford-oval/storm/issues/99 | [] | rmcc3 | 1 |
tensorflow/tensor2tensor | machine-learning | 1,770 | Multiple GPUs are visible but not allocated and used | ### Description
I am trying to start the quick start imagenet run with resnet_101 on 3 GPUs. However, the second and third GPU are not used - the model is not allocated to them. All GPUs are visible -> `$CUDA_VISIBLE_DEVICES=0,1,2` as seen in the logs.
### Environment information
OS: Ubuntu 18.04
$ pip free... | closed | 2019-12-13T17:51:11Z | 2019-12-13T18:30:23Z | https://github.com/tensorflow/tensor2tensor/issues/1770 | [] | cgebe | 1 |
voxel51/fiftyone | computer-vision | 5,220 | [BUG] AttributeError: 'FiftyOneRTDETRModelConfig' object has no attribute 'model' for models rtdetr-l-coco-torch and rtdetr-x-coco-torch | ### Describe the problem
The `rtdetr-l-coco-torch` and `rtdetr-x-coco-torch` models (https://docs.voxel51.com/model_zoo/models.html#rtdetr-l-coco-torch) do not load anymore. Running the provided notebook from the docs gives
```
Traceback (most recent call last):
File "<ipython-input-10-c73e2e7b9249>", line 11... | closed | 2024-12-05T15:14:13Z | 2024-12-09T16:38:51Z | https://github.com/voxel51/fiftyone/issues/5220 | [
"bug"
] | daniel-bogdoll | 0 |
sqlalchemy/alembic | sqlalchemy | 491 | drop python 2.6 support and bump version to 1.0 | **Migrated issue, originally created by Michael Bayer ([@zzzeek](https://github.com/zzzeek))**
the argparse requirement in setup.py can be removed once we support 2.7 and above. That in turn would make a universal wheel package on pypi possible.
| closed | 2018-04-21T13:14:57Z | 2018-06-30T01:09:24Z | https://github.com/sqlalchemy/alembic/issues/491 | [
"bug",
"installation"
] | sqlalchemy-bot | 3 |
JaidedAI/EasyOCR | deep-learning | 1,131 | Letter 'o' is wrongly interpreted as 0 (zero) | Portuguese language uses letter 'o' a lot as a word in sentences. They are almost every time recognized as 0 (zero). How can I overcome this issue?

| open | 2023-09-04T13:48:19Z | 2023-09-30T02:13:30Z | https://github.com/JaidedAI/EasyOCR/issues/1131 | [] | bilalsattar | 4 |
Anjok07/ultimatevocalremovergui | pytorch | 1,642 | resources to understand which model to pick ? | hi, are there any resources to understood which models to pick, and which are the most advanced ? I use hq 5 but idk if thats' the best at this time | open | 2024-11-29T22:01:49Z | 2024-11-29T22:01:49Z | https://github.com/Anjok07/ultimatevocalremovergui/issues/1642 | [] | avichou | 0 |
waditu/tushare | pandas | 1,701 | 利润表下载数据有重复 | 例如:000001.sz pro.income(ts_code=‘000001.sz ’, start_date=‘20000101’, end_date=‘20230418’, repot_type=1) 得到的数据,多处重复
ID:280603 | open | 2023-04-19T08:41:39Z | 2023-04-19T08:43:38Z | https://github.com/waditu/tushare/issues/1701 | [] | YellowStarr | 0 |
ydataai/ydata-profiling | pandas | 825 | Phi K correlation variable order | For me all correlation plots show variables in the (domain-specific sensible) order of the columns in my data frame.
Only Phi K shows them in some other order.
Is this a bug or a feature?
Is there a setting to get the "good" order?
This is with pandas 1.3 and pandas-profiling 3.0.0
<img width="879" alt="... | closed | 2021-09-05T19:46:25Z | 2021-09-16T08:31:52Z | https://github.com/ydataai/ydata-profiling/issues/825 | [
"bug 🐛",
"help wanted 🙋"
] | cdeil | 5 |
huggingface/datasets | pytorch | 6,595 | Loading big dataset raises pyarrow.lib.ArrowNotImplementedError 2 | ### Describe the bug
I'm aware of the issue #5695 .
I'm using a modified SDXL trainer: https://github.com/kopyl/diffusers/blob/5e70f604155aeecee254a5c63c5e4236ad4a0d3d/examples/text_to_image/train_text_to_image_sdxl.py#L1027C16-L1027C16
So i
1. Map dataset
2. Save to disk
3. Try to upload:
```
import data... | closed | 2024-01-16T02:03:09Z | 2024-01-27T18:26:33Z | https://github.com/huggingface/datasets/issues/6595 | [] | kopyl | 14 |
napari/napari | numpy | 7,226 | layers.events.changed not calling | ### 🐛 Bug Report
The callback for layers.events.changed doesn't seem to be working, although this may be due to confusion over how it's meant to work thanks to the ongoing issues with events documentation in Napari.
At least to me, the name implies that this event is called every time any change to the layerlist o... | closed | 2024-08-29T15:42:48Z | 2025-02-19T06:59:03Z | https://github.com/napari/napari/issues/7226 | [
"documentation"
] | Joseph-Garvey | 7 |
junyanz/pytorch-CycleGAN-and-pix2pix | deep-learning | 1,676 | Repeated Pattern in generated images | I'm using pix2pix model to generate images with some datasets I collected but the model keeps generate images with repeated pattern in the middle.
I'm using scale_width_and_crop on preprocess option.
Is there anyone know why?
This is a sample image i generated.
 being similar in clear and FHE predictions. Howev... | closed | 2023-10-29T07:17:12Z | 2023-10-31T14:19:30Z | https://github.com/zama-ai/concrete-ml/issues/380 | [
"bug"
] | bhuvneshchaturvedi2512 | 4 |
Johnserf-Seed/TikTokDownload | api | 482 | [BUG] | 大佬你好,我这边发现tiktok的视频合集下载每次都报错,可能是因为tiktok的主页链接和抖音不同。我试了下载抖音的个人合集没有问题。tiktok的主页链接是https://www.tiktok.com/@soon_ne,也试过您主页提供的链接也不行。能麻烦解答一下吗,感谢
Traceback (most recent call last):
File "C:\Users\XTY\Desktop\test\TikTokTool.py", line 32, in <module>
profile.getProfile(cmd.setting())
File "C:\Users\XTY\Desktop\test... | open | 2023-07-29T03:49:08Z | 2023-08-03T15:01:13Z | https://github.com/Johnserf-Seed/TikTokDownload/issues/482 | [
"故障(bug)",
"额外求助(help wanted)",
"无效(invalid)"
] | xty8623 | 2 |
CorentinJ/Real-Time-Voice-Cloning | deep-learning | 428 | Windows 10 - DLL Error | I've been stuck on this step for 2 days and I know people have posted this problem before, but the answers have not been too helpful to me considering I am retarded, technologically speaking. Anyway, when I type:
> python demo_cli.py
I am returned with:
> Traceback (most recent call last):
> File "C:\Users\... | closed | 2020-07-17T16:32:39Z | 2020-07-27T00:40:13Z | https://github.com/CorentinJ/Real-Time-Voice-Cloning/issues/428 | [] | gfhsrtras | 3 |
CorentinJ/Real-Time-Voice-Cloning | python | 285 | AssertionError | while sythesizer_preprocess_audio.py running it intrupted by an assetion error at this point every time . can anyone tell me what is happening?
AssertionError
LibriSpeech: 21%|███▊ | 251/1172 [48:17<2:57:10, 11.54s/speakers] | closed | 2020-02-19T06:37:12Z | 2020-07-04T22:38:57Z | https://github.com/CorentinJ/Real-Time-Voice-Cloning/issues/285 | [] | AmitBDA | 1 |
numba/numba | numpy | 9,398 | CUDA: Providing a signature for a function reference when eagerly compiling does not work | ## Reporting a bug
- [x] I have tried using the latest released version of Numba (most recent is
visible in the release notes
(https://numba.readthedocs.io/en/stable/release-notes-overview.html).
- [x] I have included a self contained code sample to reproduce the problem.
i.e. it's possible to run as 'pyth... | open | 2024-01-19T07:51:29Z | 2024-02-27T07:07:54Z | https://github.com/numba/numba/issues/9398 | [
"feature_request",
"CUDA"
] | s-m-e | 3 |
TracecatHQ/tracecat | pydantic | 15 | Readme: add definition of SOAR | **Is your feature request related to a problem? Please describe.**
When someone stumbles across this repo and is not familiar with the acronym, why should he care and read any further.
**Describe the solution you'd like**
A clear and concise description of SOAR, e.g:
**SOAR** ([Security Orchestration, Automatio... | closed | 2024-03-28T10:48:48Z | 2024-03-28T17:37:40Z | https://github.com/TracecatHQ/tracecat/issues/15 | [
"documentation"
] | cleder | 0 |
pyeve/eve | flask | 1,348 | Fix simple typo: wether -> whether | There is a small typo in eve/io/base.py.
Should read `whether` rather than `wether`.
| closed | 2020-01-26T11:28:58Z | 2020-02-01T08:31:49Z | https://github.com/pyeve/eve/issues/1348 | [] | timgates42 | 1 |
pykaldi/pykaldi | numpy | 303 | Install kaldi script fails with missing folder | The `./install_kaldi.sh` script mentioned in the README fails with the following error:
All done OK.
Configuring KALDI to use MKL.
Checking compiler g++ ...
Checking OpenFst library in /home/chris/pykaldi/tools/kaldi/tools/openfst-1.6.7 ...
Performing OS specific configuration ...
On Lin... | closed | 2022-05-28T23:08:39Z | 2023-09-14T21:43:16Z | https://github.com/pykaldi/pykaldi/issues/303 | [] | chrisspen | 10 |
modin-project/modin | pandas | 6,729 | Use custom pytest mark instead of `--extra-test-parameters` option | closed | 2023-11-09T01:28:33Z | 2023-12-08T14:42:27Z | https://github.com/modin-project/modin/issues/6729 | [
"Code Quality 💯",
"Testing 📈"
] | anmyachev | 0 | |
neuml/txtai | nlp | 692 | Create temporary tables once per database session | Currently, databases check to create temporary tables before every `createbatch` and `createscores` call. Temporary database tables should be created when the database connection is created, as they have a lifespan of the session. | closed | 2024-04-17T12:12:08Z | 2024-04-18T17:18:16Z | https://github.com/neuml/txtai/issues/692 | [] | davidmezzetti | 0 |
oegedijk/explainerdashboard | plotly | 289 | skorch models raising The SHAP explanations do not sum up to the model's output | Seems related to this bug: https://github.com/shap/shap/issues/3363
Can be avoided with passing `shap_kwargs=dict(check_additivity=False)` to the explainer, but then you might get inaccurate shap values.
Added check_additivity=False param to the skorch tests for now.
| open | 2023-12-17T12:20:51Z | 2023-12-17T12:20:51Z | https://github.com/oegedijk/explainerdashboard/issues/289 | [] | oegedijk | 0 |
amisadmin/fastapi-amis-admin | sqlalchemy | 110 | 使用postgresql asyncpg 时报错 |
The above exception was the direct cause of the following exception:
Traceback (most recent call last):
File "/Users/yu/.conda/envs/turingmodel/lib/python3.8/site-packages/uvicorn/protocols/http/httptools_impl.py", line 371, in run_asgi
result = await app(self.scope, self.receive, self.send)
File "/User... | open | 2023-07-08T15:40:36Z | 2023-07-08T15:40:36Z | https://github.com/amisadmin/fastapi-amis-admin/issues/110 | [] | yxlwfds | 0 |
JaidedAI/EasyOCR | deep-learning | 1,361 | Fine-Tune training for new and rare Chinese characters in Traditional Chinese | Hello
Can I ask a question?
How to fine-tune training for new and rare Chinese characters in Traditional Chinese?
For rare words that are not established in the vocabulary, the official teaching is currently used, and the parameters of config.yaml are
FT: True, new_prediction: True
A small number of training set... | open | 2025-01-08T10:36:00Z | 2025-01-08T10:41:00Z | https://github.com/JaidedAI/EasyOCR/issues/1361 | [] | 6692a | 0 |
amfoss/cms | graphql | 23 | Broken link for amfoss linkedIn account. | **Describe the bug**
LinkedIn link is broken.
**Screenshots**

| closed | 2019-03-12T08:22:56Z | 2019-03-13T12:24:53Z | https://github.com/amfoss/cms/issues/23 | [] | harshithpabbati | 0 |
jupyter-book/jupyter-book | jupyter | 1,782 | Make Pyppeteer configurable for pdfhtml builder | ### Context
SImilar to sphinx, pyppeteer also has a bunch of configs as described [here](https://miyakogi.github.io/pyppeteer/reference.html#launcher). However, JB does not allow default configs to be overridden while converting HTML to PDF.
For example, pyppeteer attempts to download and install chromium in the d... | open | 2022-07-17T17:51:42Z | 2022-09-14T18:49:19Z | https://github.com/jupyter-book/jupyter-book/issues/1782 | [
"enhancement"
] | SimplyOm | 2 |
tensorflow/tensor2tensor | machine-learning | 1,096 | AttributeError: 'RunConfig' object has no attribute 'data_parallelism' t2t=1.9, TPU TF version 1.9 | ### Description
Hi
I am getting this error while running translation workloads on TPU.
AttributeError: 'RunConfig' object has no attribute 'data_parallelism'
...
### Environment information
Tensor2Tensor =1.9
TPU tensorflow=1.9
```
OS: <your answer here>
$ pip freeze | grep tensor
tensor2tensor==1.... | open | 2018-09-25T21:51:40Z | 2018-09-25T21:51:40Z | https://github.com/tensorflow/tensor2tensor/issues/1096 | [] | nicks165 | 0 |
pyro-ppl/numpyro | numpy | 1,467 | Leak when running examples with JAX_CHECK_TRACER_LEAKS=1 | Hi
when I run ar2.py from yours examples (or bnn.py did not try with others) with the environment variable JAX_CHECK_TRACER_LEAKS=1 they fail. (I had to use it to try to find an issue with a function I had written)
the exception raised is
Exception: Leaked level MainTrace(1,DynamicJaxprTrace). Leaked tracer(s):... | closed | 2022-08-12T08:54:00Z | 2022-08-23T13:34:29Z | https://github.com/pyro-ppl/numpyro/issues/1467 | [
"bug"
] | hyperfra | 4 |
yunjey/pytorch-tutorial | deep-learning | 37 | [image captioning] training /test/validation data? | training /test/validation data? I saw in image captioning and the other examples that there is no training/test/validation splits. Should the examples have this to promote best practices? If I wrote code to add it, would it be merged in? | closed | 2017-05-26T17:28:12Z | 2017-05-28T11:21:12Z | https://github.com/yunjey/pytorch-tutorial/issues/37 | [] | jtoy | 1 |
junyanz/pytorch-CycleGAN-and-pix2pix | pytorch | 1,594 | pix2pix training doesn't go past "create web directory ./checkpoints\cartoonify_pix2pix\web..." | I am training a pix2pix model on my own dataset. I trained for 170 epochs at first and then another 20 epochs. Now when I try to continue training, it loads the model and does everything normally but stops execution without giving any error. This issue was there before, but doing random things like restarting the PC or... | closed | 2023-09-01T06:16:56Z | 2024-06-25T02:19:59Z | https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/issues/1594 | [] | TehreemFarooqi | 2 |
vimalloc/flask-jwt-extended | flask | 418 | AssertionError: View function mapping is overwriting an existing endpoint function: wrapper | Hello, Im coding a REST API for a website I got his problem when I use @jwt_required. Couldn't find any solutions on the internet so I thought asking here would be the only option.
My Code:
```python
@app.route('/login', methods=['POST'])
def login():
email = request.json.get('email', None)
password ... | closed | 2021-04-24T17:20:33Z | 2021-04-25T03:40:07Z | https://github.com/vimalloc/flask-jwt-extended/issues/418 | [] | GoekhanDev | 2 |
exaloop/codon | numpy | 25 | PyObjects in `if`-statements fail | ```python
from python import foo
if foo() > 1:
pass
```
gives
```
Trunc only operates on integer
%262 = trunc i8* %261 to i1, !dbg !24241
Assert failed: module broken
Expression: !broken
Source: /Users/arshajii/Documents/workspace/codon/codon/sir/llvm/optimize.cpp:232
zsh: abort build/codon ... | closed | 2022-04-21T15:03:58Z | 2022-07-22T16:50:44Z | https://github.com/exaloop/codon/issues/25 | [] | arshajii | 1 |
ets-labs/python-dependency-injector | flask | 424 | Erroneous initialization of Resources | After upgrading from `4.10.1` to `4.29.2`, I noticed that some Resources which remained unitialized in prior version unexpectedly started to initialize during `init_resorces()` call. The **Selector** provider initializes all nested resources disregarding selector key:
```
from dependency_injector import containers,... | open | 2021-03-11T11:34:59Z | 2022-04-05T09:41:48Z | https://github.com/ets-labs/python-dependency-injector/issues/424 | [] | atten | 9 |
jonaswinkler/paperless-ng | django | 1,594 | [BUG] Installed from script and Gotenburg and Tika not working? | Hello, thanks for this great work!
I am new to paperless-ng do not normally use docker, so I may be doing something wrong.
My paperless works well, but when I try to import a .docx file for example, it fails with, `Error while converting document to PDF: 404 Client Error: Not Found for url: http://gotenberg:3000... | open | 2022-02-01T11:56:09Z | 2022-04-19T11:13:49Z | https://github.com/jonaswinkler/paperless-ng/issues/1594 | [] | 2600box | 10 |
yzhao062/pyod | data-science | 274 | AutoEncoder.fit() times ever increasing | While working with the AutoEncoder, we noticed that times required to fit a model are increasing a bit every time AutoEncoder.fit() is invoked.
As an example, if the following is executed
```
F1 = list(np.random.normal(0, 0.2, 250))`
F1.extend(list(np.random.normal(1, 0.2, 50)))
F2 = list(np.random.normal(0, 0.2, ... | open | 2021-01-15T13:23:39Z | 2021-01-27T09:11:18Z | https://github.com/yzhao062/pyod/issues/274 | [] | fgabbaninililly | 1 |
microsoft/nni | tensorflow | 5,483 | TypeError: __new__() missing 1 required positional argument: 'task' | **Describe the issue**: Hello, developers. I'm a newbie just getting started learning nas. When I run the tutorial file in the notebook, I get an error, I hope you can help me to solve this problem:
File "search_2.py", line 59, in <module>
max_epochs=5)
File "D:\anaconda3\envs\venv_copy\lib\site-packages\nn... | closed | 2023-03-28T01:18:25Z | 2023-03-29T02:38:04Z | https://github.com/microsoft/nni/issues/5483 | [] | zzfer490 | 3 |
jupyterlab/jupyter-ai | jupyter | 978 | Cannot import name 'RunnableWithMessageHistory' from 'langchain_core.runnables' | ## Description
https://github.com/jupyterlab/jupyter-ai/pull/943 replaced import:
```diff
- from langchain_core.runnables.history import RunnableWithMessageHistory
+ from langchain_core.runnables import ConfigurableFieldSpec, RunnableWithMessageHistory
```
This errors out on specific version of `langchain_c... | closed | 2024-09-07T09:41:39Z | 2024-09-09T20:20:29Z | https://github.com/jupyterlab/jupyter-ai/issues/978 | [
"bug"
] | krassowski | 1 |
microsoft/MMdnn | tensorflow | 834 | [Caffe2Keras] converted keras gives wrong output value | Hi,
I have a Caffe model and I converted to Keras model (Caffe - IR - Keras) but when testing with same input, the Keras model gave different output value although the dimension is correct. What should I do?
The caffe model: https://github.com/CongWeilin/mtcnn-caffe/tree/master/12net
Thanks | open | 2020-05-11T15:41:37Z | 2020-05-16T07:41:46Z | https://github.com/microsoft/MMdnn/issues/834 | [] | nhatuan84 | 1 |
Guovin/iptv-api | api | 479 | open_m3u_result设置成了False依然输出的是m3u | docker-compose.yml:
services:
tv-requests:
image: guovern/tv-requests:latest
ports:
- "8000:8000"
environment:
open_m3u_result: False
volumes:
- /volume1/IPTV/config:/tv-requests/config
- /volume1/IPTV/output:/tv-requests/output
restart: always
访问http://IP:80... | closed | 2024-10-29T13:26:27Z | 2024-10-29T15:46:57Z | https://github.com/Guovin/iptv-api/issues/479 | [
"invalid"
] | iFoxox | 3 |
postmanlabs/httpbin | api | 359 | How to debug issues in URL? | I'm trying to debug an encoding issue in the URL. I want to quickly test an HTTP request language, and I want to see the results of:
```
request.get("http://httpbin.org/get/test");
request.get("http://httpbin.org/get/café");
request.get("http://httpbin.org/get/🐶");
```
I want to see if the library encodes th... | open | 2017-05-30T09:50:39Z | 2018-04-26T17:51:14Z | https://github.com/postmanlabs/httpbin/issues/359 | [] | Flimm | 3 |
Gozargah/Marzban | api | 731 | sni رندم در ریالیتی | دوستان آیا راهی هست بشه برای ریالیتی توی فایل کانفیگ servername / sni رو با * گذاشت مرزبان به صورت رندم یک مقدار بسازه و xray قبول کنه؟
من یه سایت با tls 1.3 آووردم بالا و تنظیم کردم روی همه ساب دامنه هام هم هم سایت میاره بالا اما مشکل اینه وقتی توی فایل xray configuration وقتی مقدار *.domain.com رو میزنم کاربر اگ... | closed | 2024-01-05T22:15:44Z | 2024-01-09T21:58:09Z | https://github.com/Gozargah/Marzban/issues/731 | [
"Bug"
] | lvlrf | 3 |
Sanster/IOPaint | pytorch | 513 | [Feature Request] Add Zits-PlusPlus model | A new version of the Zits model [has been released](https://github.com/ewrfcas/ZITS-PlusPlus)
The inference code and models have been released, so I think it should be possible to add it. Seems to be a fairly strong model that should be able to outperform a lot of the models in IOPaint. | closed | 2024-04-24T19:38:01Z | 2025-03-13T20:43:27Z | https://github.com/Sanster/IOPaint/issues/513 | [
"stale"
] | mn7216 | 3 |
scikit-tda/kepler-mapper | data-visualization | 169 | Ayasdi's Neighborhood Lens 1 and 2? | I feel embarrassed asking this here and if it's not appropriate then please delete, but I have not been able to find the answer anywhere...
(Some of ?) the lenses that Ayasdi uses are listed in [this](https://platform.ayasdi.com/sdkdocs/userdoc/lenses_supp_overview.html) page (scroll to the bottom of the page). The ... | closed | 2019-04-18T10:48:25Z | 2021-04-15T17:12:24Z | https://github.com/scikit-tda/kepler-mapper/issues/169 | [] | karinsasaki | 4 |
erdewit/ib_insync | asyncio | 180 | decode with correct encoding | utils.py --> def decode()
return s.decode(errors='backslashreplace') -->> some problem here.
s.decode(encoding="GBK") # non-english location have to specify the correct one that set in tws.
so another question is how to get the language you set in tws? | closed | 2019-08-25T19:20:33Z | 2019-08-29T10:32:31Z | https://github.com/erdewit/ib_insync/issues/180 | [] | viponedream | 1 |
roboflow/supervision | machine-learning | 1,463 | Notebook not found: Serialise Detections to a CSV File | ### Search before asking
- [X] I have searched the Supervision [issues](https://github.com/roboflow/supervision/issues) and found no similar bug report.
### Bug
The Colab in this [cookbook](https://supervision.roboflow.com/develop/notebooks/serialise-detections-to-csv/) is not found.
<img width="1118" al... | closed | 2024-08-19T20:22:59Z | 2024-08-20T14:08:37Z | https://github.com/roboflow/supervision/issues/1463 | [
"bug"
] | ediardo | 2 |
PaddlePaddle/models | nlp | 5,293 | paddlepaddle2.0.0、paddlehub2.0.0版本ocr接口疑似存在内存泄漏 | ocr接口疑似存在内存泄漏请看一下问题出现在什么地方,谢谢。
1、版本:
paddlepaddle2.0.0、
paddlehub2.0.0
运行的为cpu版本
服务器内存16G
2、问题现象
a、python单进程调用ocr.recognize_text接口读取指定目录下的100张不同图片进行图片文字识别,每识别一张图片,占用内存增加200M以上,随着识别图片数的增加,内存持续增长,直到内存被耗尽被操作系统kill掉
b、如果只是反复识别同一张图片的话,内存增长的一定数后不再增长
3、导致内存泄漏的疑似代码
通过代码调用链的分析最终定位到c++库中的PD_Pre... | closed | 2021-03-20T02:46:24Z | 2021-03-22T02:31:38Z | https://github.com/PaddlePaddle/models/issues/5293 | [] | weihyemail | 0 |
huggingface/datasets | deep-learning | 7,363 | ImportError: To support decoding images, please install 'Pillow'. | ### Describe the bug
Following this tutorial locally using a macboko and VSCode: https://huggingface.co/docs/diffusers/en/tutorials/basic_training
This line of code: for i, image in enumerate(dataset[:4]["image"]):
throws: ImportError: To support decoding images, please install 'Pillow'.
Pillow is installed.
###... | open | 2025-01-08T02:22:57Z | 2025-02-07T07:30:33Z | https://github.com/huggingface/datasets/issues/7363 | [] | jamessdixon | 3 |
sebp/scikit-survival | scikit-learn | 278 | Confusion with dynamic AUC | I am not able to reproduce the AUC that is returned by `cumulative_dynamic_auc()`. I am pretty certain that this is due to my lack of understanding, but would like some clarification if possible. Below is a reproducible example, mostly derived from the documentation example:
```
# Most imports
import numpy as np
... | closed | 2022-07-01T23:43:54Z | 2022-07-03T08:20:10Z | https://github.com/sebp/scikit-survival/issues/278 | [] | Dekermanjian | 1 |
CTFd/CTFd | flask | 2,550 | [Question] Timing's Impact on CTF Ranking | It's often observed that when two teams amass equal points, the team which solves challenges later tends to be ranked higher on the scoreboard.
Why does the timing of challenge solutions become a determining factor in team rankings?
To shed light on this, consider a scenario where two teams complete all challenges ... | closed | 2024-06-02T09:59:09Z | 2024-06-07T13:23:33Z | https://github.com/CTFd/CTFd/issues/2550 | [] | mosheDO | 2 |
rthalley/dnspython | asyncio | 1,028 | 2.5.0 Release | I just looked at the amount of unreleased stuff we have and concluded it's enough for a release. So, I plan to start the release process in early 2024 with the usual RC and release following two weeks or so later if all goes well. | closed | 2023-12-24T22:23:54Z | 2024-01-20T13:32:45Z | https://github.com/rthalley/dnspython/issues/1028 | [] | rthalley | 2 |
opengeos/leafmap | streamlit | 715 | Incorrect data labels with `.add_data()`? | <!-- Please search existing issues to avoid creating duplicates. -->
### Environment Information
- leafmap version: leafmap==0.31.7
- Python version: 3.10.12
- Operating System: Ubuntu Linux
### Description
### What I Did
I use `add_data()` to map color fill to a categorical variable. This se... | closed | 2024-04-12T22:35:14Z | 2024-04-14T18:44:10Z | https://github.com/opengeos/leafmap/issues/715 | [
"bug"
] | cboettig | 2 |
horovod/horovod | pytorch | 3,347 | GPU Head tests for elastic ray are flaky in CI | Elastic ray tests(scale up and down) become flaky in GPU Heads. The following tests fail:
1. test_ray_elastic_v2::test_fault_tolerance_hosts_remove_and_add
2. test_ray_elastic_v2::test_fault_tolerance_hosts_added_and_removed
3. test_ray_elastic_v2::test_fault_tolerance_hosts_remove_and_add_cooldown
| open | 2022-01-06T00:17:49Z | 2022-01-06T00:17:49Z | https://github.com/horovod/horovod/issues/3347 | [
"bug"
] | ashahab | 0 |
scrapy/scrapy | web-scraping | 6,590 | Path to removal of spider middleware iterable downgrading | We added support for async spider middlewares in 2.7 (In October 2022), and [mixing sync and async middlewares](https://docs.scrapy.org/en/latest/topics/coroutines.html#sync-async-spider-middleware) was intended to be temporary and eventually deprecated and removed, after (most?) 3rd-party middlewares are converted. Th... | open | 2024-12-24T18:44:15Z | 2025-02-08T13:17:53Z | https://github.com/scrapy/scrapy/issues/6590 | [
"discuss",
"cleanup",
"asyncio"
] | wRAR | 5 |
modelscope/modelscope | nlp | 1,029 | [Bug] 模型存在重复下载的问题,不能正确识别已经上传至缓存文件夹的权重文件 | Thanks for your error report and we appreciate it a lot.
**Checklist**
* I have searched the tutorial on modelscope [doc-site](https://modelscope.cn/docs)
* I have searched related issues but cannot get the expected help.
* The bug has not been fixed in the latest version.
**Describe the bug**
将一个下载容易断线的模型... | closed | 2024-10-18T06:52:07Z | 2024-10-21T09:34:38Z | https://github.com/modelscope/modelscope/issues/1029 | [] | zydmtaichi | 10 |
quantumlib/Cirq | api | 6,727 | Incorrect behaviour when inserting classical control circuit element, causing `ValueError` | **Description of the issue**
Under specific circumstances, it seems that classical controls are inserted wrongly and causing errors when getting measurement keys.
**How to reproduce the issue**
Run the following code:
```py
from cirq import *
from cirq.transformers import *
import numpy as np
all_pas... | closed | 2024-09-13T14:22:33Z | 2025-01-15T17:07:11Z | https://github.com/quantumlib/Cirq/issues/6727 | [
"good first issue",
"kind/bug-report",
"triage/accepted"
] | Bennybenassius | 3 |
neuml/txtai | nlp | 163 | Make adding pipelines to API easier | Currently, the list of available pipelines and endpoints is hard coded in the API. This should be modified to dynamically detect pipelines within the txtai.pipeline package. This will make all txtai.pipelines available via API workflows.
Exposing API endpoints for each pipeline still needs to be explicitly set but ... | closed | 2021-11-28T00:38:46Z | 2021-11-28T00:40:41Z | https://github.com/neuml/txtai/issues/163 | [] | davidmezzetti | 0 |
quantmind/pulsar | asyncio | 172 | HTTPException message is not escaped | When an HTTPException is raised, and [render_error()](https://github.com/quantmind/pulsar/blob/74d1e6e58095679291c14c826ce63f6312a93256/pulsar/apps/wsgi/utils.py#L262) decide to return HTML, the exception message should be escaped to prevent [XSS](https://en.wikipedia.org/wiki/Cross-site_scripting).
[OWASP recommends ... | closed | 2015-10-30T10:56:12Z | 2016-01-06T14:55:06Z | https://github.com/quantmind/pulsar/issues/172 | [
"enhancement",
"security"
] | msornay | 2 |
getsentry/sentry | django | 87,029 | Search functionality for schema hints slideout | same search functionality as breadcrumbs. AKA local filtering logic. | closed | 2025-03-13T20:43:29Z | 2025-03-18T18:16:40Z | https://github.com/getsentry/sentry/issues/87029 | [] | nikkikapadia | 0 |
junyanz/pytorch-CycleGAN-and-pix2pix | pytorch | 890 | No models saved in Colab | I was using the PyTorch Colab notebook of CycleGAN. I trained the network using `!python train.py --dataroot ./datasets/horse2zebra --name horse2zebra --model cycle_gan --display_id -1 --n_epochs 5 --n_epochs_decay 5` but it didn't saved any model (I'm using default settings to save every 5000 iterations, in fact it pr... | closed | 2020-01-03T01:59:05Z | 2022-04-28T21:33:35Z | https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/issues/890 | [] | domef | 6 |
automagica/automagica | automation | 1 | Math functions | Add math functionality to activities.py to support native math operations as described in the documentation | closed | 2018-05-14T14:57:58Z | 2018-10-04T09:45:02Z | https://github.com/automagica/automagica/issues/1 | [
"enhancement"
] | tvturnhout | 0 |
ultralytics/yolov5 | machine-learning | 13,304 | hyp.finetune.yaml missing | ### Search before asking
- [X] I have searched the YOLOv5 [issues](https://github.com/ultralytics/yolov5/issues) and [discussions](https://github.com/ultralytics/yolov5/discussions) and found no similar questions.
### Question
Hi I cannot find hyp.finetune.yaml in this repo as it was mentioned in finetuning ... | closed | 2024-09-11T02:27:47Z | 2024-09-13T05:58:50Z | https://github.com/ultralytics/yolov5/issues/13304 | [
"question"
] | spacewalk01 | 3 |
iperov/DeepFaceLab | machine-learning | 5,598 | 3070TI (laptop) shows better compatibility with DX12 build compared to 3000 build (More memoryerrors) | First of all, I want to ask that, theoretically, should 3000 build provide a better performance or not? If not, maybe I will stop trying to figure out how to run 3000 build properly.
OK, now the description of the problems:
In general, for training, 3000 build always crashes, and usually with a MemoryError or low p... | closed | 2022-12-10T23:50:51Z | 2022-12-18T08:40:35Z | https://github.com/iperov/DeepFaceLab/issues/5598 | [] | JulesLiu | 0 |
ivy-llc/ivy | numpy | 28,090 | Fix Frontend Failing Test: paddle - math.paddle.pow | To-do List: https://github.com/unifyai/ivy/issues/27500 | closed | 2024-01-27T16:36:15Z | 2024-01-29T13:11:44Z | https://github.com/ivy-llc/ivy/issues/28090 | [
"Sub Task"
] | Sai-Suraj-27 | 0 |
plotly/dash | data-visualization | 3,076 | add support for ignoring specific folders under the assets directory | The existing parameter `assets_ignore` can ignore files under the assets folder that match a specified regular expression pattern, but it cannot ignore specific folders under the assets folder as a whole, which would be useful in many scenarios.
| open | 2024-11-15T01:58:40Z | 2024-11-20T13:30:18Z | https://github.com/plotly/dash/issues/3076 | [
"feature",
"P3"
] | CNFeffery | 3 |
cobrateam/splinter | automation | 387 | browser.get_alert() does not return "None" when there is no alert | When I call `browser.get_alert()` on a page without an alert, instead of `None`, it returns `NoAlertPresentException: Message: No alert is present`-- contrary to the documentation.
Ie.,
```
from splinter import Browser
browser = Browser()
browser.visit('https://www.google.com')
browser.get_alert()
```
It'd be nice... | closed | 2015-04-06T20:46:31Z | 2019-10-02T20:48:42Z | https://github.com/cobrateam/splinter/issues/387 | [
"NeedsInvestigation"
] | njbennett | 2 |
dinoperovic/django-salesman | rest-api | 19 | item_validate is called twice per item | validate_basket_item is called twice per item | closed | 2022-04-25T19:03:24Z | 2022-04-25T19:18:55Z | https://github.com/dinoperovic/django-salesman/issues/19 | [] | IncreaseComputers | 1 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.