repo_name stringlengths 9 75 | topic stringclasses 30
values | issue_number int64 1 203k | title stringlengths 1 976 | body stringlengths 0 254k | state stringclasses 2
values | created_at stringlengths 20 20 | updated_at stringlengths 20 20 | url stringlengths 38 105 | labels listlengths 0 9 | user_login stringlengths 1 39 | comments_count int64 0 452 |
|---|---|---|---|---|---|---|---|---|---|---|---|
sunscrapers/djoser | rest-api | 468 | SAML metadata page | While following the python-social-auth's [tutorial](https://python-social-auth.readthedocs.io/en/latest/backends/saml.html#basic-usage) for configuring the SAML authentication backend, I've hit a problem. I'm unable to configure the metadata serving page. Even after setting the social urls with the ```social``` namespa... | open | 2020-02-27T18:34:00Z | 2020-02-27T18:34:00Z | https://github.com/sunscrapers/djoser/issues/468 | [] | lm-sousa | 0 |
piccolo-orm/piccolo | fastapi | 865 | name 'Serial' is not defined | When creating a new migration, I think this bug(error) only occurs with ForeignKeys, details below:
```
my_app/tables.py
```
```python
class Order(Table, tablename="orders"):
user = ForeignKey(
LazyTableReference(
table_class_name="Users",
module_path="users.tables"
... | open | 2023-07-14T14:54:22Z | 2023-07-22T19:37:58Z | https://github.com/piccolo-orm/piccolo/issues/865 | [
"bug"
] | hoosnick | 11 |
huggingface/datasets | machine-learning | 7,377 | Support for sparse arrays with the Arrow Sparse Tensor format? | ### Feature request
AI in biology is becoming a big thing. One thing that would be a huge benefit to the field that Huggingface Datasets doesn't currently have is native support for **sparse arrays**.
Arrow has support for sparse tensors.
https://arrow.apache.org/docs/format/Other.html#sparse-tensor
It would be ... | open | 2025-01-21T20:14:35Z | 2025-01-30T14:06:45Z | https://github.com/huggingface/datasets/issues/7377 | [
"enhancement"
] | JulesGM | 1 |
widgetti/solara | jupyter | 36 | Quickstart example does not render correctly with solara run | I have a fresh conda environment with python 3.9 on windows. I've used poetry to install solara.
I have a `myapp.py` file with the following code:
```python
import solara
@solara.component
def Page():
clicks, set_clicks = solara.use_state(0)
def increase_clicks():
set_clicks(clicks+1)
s... | closed | 2023-03-09T10:25:21Z | 2023-03-09T14:07:56Z | https://github.com/widgetti/solara/issues/36 | [] | Jhsmit | 11 |
mwouts/itables | jupyter | 56 | Use Jupyter Book to build the ITables documentation | We should split the README into multiple chapters of a Jupyter Book
TODO
- [x] Fix the table width and font
- [x] Fix the [issue](https://github.com/executablebooks/jupyter-book/issues/1610) with the sample dataframe notebook
- [x] Automatize the publication on GitHub pages | closed | 2022-01-22T00:41:51Z | 2022-01-24T23:29:44Z | https://github.com/mwouts/itables/issues/56 | [] | mwouts | 1 |
pytest-dev/pytest-django | pytest | 306 | Migrations fail silently | If I run `py.test --create-db` with migrations that don't work in dev, py.test just swallows them up and a lot of tests fail due to missing columns etc.
| closed | 2016-01-11T09:19:44Z | 2020-10-16T19:06:52Z | https://github.com/pytest-dev/pytest-django/issues/306 | [
"bug"
] | lee-kagiso | 4 |
jupyterlab/jupyter-ai | jupyter | 510 | Leverage `ExtensionHandlerMixin` from Jupyter Server |
### Problem
Jupyter Server provides an `ExtensionHandlerMixin` class for more complex extension handlers. We should probably take advantage of this.
See discussion in: https://github.com/jupyter-server/jupyter_server_fileid/pull/72#discussion_r1420726422
| open | 2023-12-08T16:34:49Z | 2023-12-08T16:34:49Z | https://github.com/jupyterlab/jupyter-ai/issues/510 | [
"enhancement"
] | dlqqq | 0 |
mage-ai/mage-ai | data-science | 5,618 | Instance-wide API key rotation | **Is your feature request related to a problem? Please describe.**
Our organization values strong security practices, but the application currently lacks a method to rotate the instance-wide API key. This creates a potential risk if the key is ever compromised, as there is no way to retire the compromised key or intro... | closed | 2024-12-16T18:50:27Z | 2024-12-22T21:00:21Z | https://github.com/mage-ai/mage-ai/issues/5618 | [] | the-archbishop | 1 |
vaexio/vaex | data-science | 1,387 | Join with interval | Hi,
is there any way to perform this function in vaex?
trim2 = df1.join(df2, on=[df1['CODMUNRES']==df2['mun_geocod'] ,df2['Date'] >= df1['Trim2_start'], df2['Date'] <= df1['Trim2_stop']],how='left').groupBy('ID','CODMUNRES')
| open | 2021-06-05T06:21:52Z | 2021-06-05T06:21:52Z | https://github.com/vaexio/vaex/issues/1387 | [] | erickkill | 0 |
scikit-learn/scikit-learn | machine-learning | 30,621 | Add links to examples from the docstrings and user guide | _TLDR: Meta-issue for new contributors to add links to the examples in helpful places of the rest of the docs._
## Description
This meta-issue is a good place to start with your first contributions to scikit-learn.
This issue builds on top of #26927 and is introduced for easier maintainability. The goal is exactly th... | open | 2025-01-10T12:29:04Z | 2025-03-24T15:21:08Z | https://github.com/scikit-learn/scikit-learn/issues/30621 | [
"Documentation",
"Sprint",
"good first issue",
"Meta-issue"
] | StefanieSenger | 101 |
pydantic/pydantic-core | pydantic | 853 | manylinux aarch64 wheels aren't manylinux2014 compliant | This could be due to an underlying issue with maturin, but the published `aarch64` wheels for `pydantic_core` do not appear to be `manylinux2014` compliant. This doesn't look like a huge surprise given that they appear to be built in a `manylinux_2_24` container, but somehow the final wheels haven't ended up with the c... | closed | 2023-08-04T15:58:44Z | 2023-08-16T16:57:27Z | https://github.com/pydantic/pydantic-core/issues/853 | [
"unconfirmed"
] | adrianeboyd | 2 |
adap/flower | tensorflow | 4,919 | Add a SizePartitioner with IID Distribution | ### Describe the type of feature and its functionality.
The current implementation of Flower’s partitioners provides two separate functionalities: the IidPartitioner and SizePartitioner. This proposal suggests introducing a new partitioner "SizePartitionerIID" that merges both functionalities. It will allow users to s... | closed | 2025-02-07T10:49:29Z | 2025-02-20T10:25:28Z | https://github.com/adap/flower/issues/4919 | [
"feature request"
] | RedPandaY | 0 |
autogluon/autogluon | data-science | 4,218 | [tabular] Add log-scaling to regression for appropriate metrics | Reference: [Discord Thread](https://discord.com/channels/1043248669505368144/1241688613725536296)
Kudos to @giladrubin1 for starting the thread and @LennartPurucker for helping with brainstorming
## The Idea
In regression tasks for metrics such as RMLSE (root mean squared log error), it is beneficial to first lo... | open | 2024-05-22T19:02:57Z | 2025-02-05T15:05:56Z | https://github.com/autogluon/autogluon/issues/4218 | [
"enhancement",
"module: tabular",
"priority: 1"
] | Innixma | 3 |
home-assistant/core | python | 141,257 | Roborock Connection | ### The problem
Hello,
ich try to connect my Roborock but i get an error:
Traceback (most recent call last):
File "/usr/src/homeassistant/homeassistant/config_entries.py", line 640, in __async_setup_with_context
result = await component.async_setup_entry(hass, self)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^... | open | 2025-03-24T06:37:32Z | 2025-03-24T16:31:18Z | https://github.com/home-assistant/core/issues/141257 | [
"needs-more-information",
"integration: roborock"
] | Dengo91 | 3 |
freqtrade/freqtrade | python | 11,511 | How to Enable the Backtesting Option in FreqUI When Running Freqtrade via Docker on Windows 10? | Hello Freqtrade community,
Could I kindly ask a beginner question? I’m running Freqtrade using Docker on Windows 10 and noticed in the documentation that FreqUI includes backtesting functionality. However, when I access FreqUI, I only see the options "Trade," "Dashboard," and "Chart Logs." Could you please guide me on... | closed | 2025-03-15T19:34:16Z | 2025-03-15T23:57:40Z | https://github.com/freqtrade/freqtrade/issues/11511 | [
"Question"
] | ray147291617 | 2 |
holoviz/panel | matplotlib | 7,771 | Feature: A lighter weight pn.Card | With the advent of LLMs, I see a lot of collapsible divs, e.g.
Claude:
<img width="748" alt="Image" src="https://github.com/user-attachments/assets/4721460a-6c04-4fe4-a7a2-cf8547848009" />
Cursor:
<img width="395" alt="Image" src="https://github.com/user-attachments/assets/965cd583-eedb-49a9-bbcd-0603c3d3ee34" />
Th... | open | 2025-03-11T18:38:03Z | 2025-03-13T11:24:36Z | https://github.com/holoviz/panel/issues/7771 | [] | ahuang11 | 0 |
localstack/localstack | python | 12,062 | bug: EventBridge event target pointing on API destination is not expanding header_parameters | ### Is there an existing issue for this?
- [X] I have searched the existing issues
### Current Behavior
I have EventBus event target configured similarly to the below terraform:
```
resource "aws_cloudwatch_event_api_destination" "http_destination" {
# .......
}
resource "aws_cloudwatch_event_target" "tes... | open | 2024-12-22T19:31:12Z | 2025-01-03T15:26:06Z | https://github.com/localstack/localstack/issues/12062 | [
"type: bug",
"aws:events",
"status: backlog"
] | gemyago | 1 |
google-deepmind/graph_nets | tensorflow | 59 | Cannot interpret feed_dict as Tensor | Hi! I'm trying to run a model that predicts node attributes based on global and edge inputs.
I've been largely following the shortest_path.ipynb demo to write my code, and my code at the moment looks as follows (happy to include more if need be!):
```python
# train_input, train_target, test_input etc. are all list... | closed | 2019-04-04T15:33:53Z | 2019-04-05T13:28:22Z | https://github.com/google-deepmind/graph_nets/issues/59 | [] | jmcs100 | 6 |
mitmproxy/pdoc | api | 268 | Get line number of classes and functions... | I implement a README generate using pdoc here: https://github.com/boxine/bx_py_utils/pull/76
Example result is currently: https://github.com/boxine/bx_py_utils/blob/auto-doc/README.md
The idea is to add links to the github code view page, e.g.:
https://github.com/boxine/bx_py_utils/blob/auto-doc/bx_py_utils/auto_... | closed | 2021-05-31T10:18:28Z | 2021-06-05T10:46:45Z | https://github.com/mitmproxy/pdoc/issues/268 | [
"enhancement"
] | jedie | 10 |
matplotlib/mplfinance | matplotlib | 203 | Bug Report: Can't display Chinese character even matplotlib can work with Chinese | Respect,
Here is a user from China. I met a problem when I updated to latest version of mapfinance. I can't show Chinese characters anymore even the matplotlib can work with Chinese characters well.
Thank you very much | closed | 2020-07-02T08:03:38Z | 2020-08-09T18:42:43Z | https://github.com/matplotlib/mplfinance/issues/203 | [
"bug"
] | xuelangqingkong | 8 |
AUTOMATIC1111/stable-diffusion-webui | deep-learning | 16,878 | solved | ### Checklist
- [ ] The issue exists after disabling all extensions
- [ ] The issue exists on a clean installation of webui
- [ ] The issue is caused by an extension, but I believe it is caused by a bug in the webui
- [x] The issue exists in the current version of the webui
- [ ] The issue has not been reported before... | closed | 2025-03-03T15:38:47Z | 2025-03-14T16:30:52Z | https://github.com/AUTOMATIC1111/stable-diffusion-webui/issues/16878 | [
"bug-report"
] | Sensanko52123 | 2 |
aimhubio/aim | data-visualization | 3,272 | The API reference spec seems missing | ## 📚 Documentation
The documentation sometimes contains links to the full API reference specs, like for instance on the [Manage Runs](https://aimstack.readthedocs.io/en/latest/using/manage_runs.html) page:
> Run class full [spec](https://aimstack.readthedocs.io/en/latest/refs/sdk.html#aim.sdk.run.Run).
Unfort... | open | 2024-12-17T12:09:52Z | 2024-12-17T12:09:52Z | https://github.com/aimhubio/aim/issues/3272 | [
"area / docs"
] | bluenote10 | 0 |
opengeos/leafmap | streamlit | 164 | Add GUI for opening COG and STAC | This feature allows loading raster datasets onto the map without coding. | closed | 2021-12-30T15:03:51Z | 2022-01-11T05:38:21Z | https://github.com/opengeos/leafmap/issues/164 | [
"Feature Request"
] | giswqs | 1 |
wandb/wandb | data-science | 8,937 | 'wandb.tensorboard.unpatch()' missing in documentation | Hey everyone,
I log my experiments with Tensorboard and have multiple experiments per run. Thus, I need to run:
```
wandb.tensorboard.patch(root_logdir=log_directory)
wand.init()
writer = SummaryWriter(log_dir=log_directory)
...
My experiment
...
wandb.finish()
writer.close()
wandb.tensorboard.unpatch()
```
Howeve... | open | 2024-11-22T22:43:09Z | 2024-11-28T14:46:49Z | https://github.com/wandb/wandb/issues/8937 | [
"c:docs"
] | daniel-bogdoll | 5 |
lexiforest/curl_cffi | web-scraping | 464 | Unable to download libcurl-impersonate when installing package (build.py) | Please check the following items before reporting a bug, otherwise it may be closed immediately.
- [ + ] **This is NOT a site-related "bugs"**, e.g. some site blocks me when using curl_cffi,
UNLESS it has been verified that the reason is missing pieces in the impersonation.
- [ + ] A code snippet that can repr... | closed | 2024-12-17T12:10:38Z | 2024-12-17T12:42:24Z | https://github.com/lexiforest/curl_cffi/issues/464 | [
"bug"
] | lrdcxdes | 1 |
pytorch/vision | computer-vision | 8,720 | wrap_dataset_for_transforms_v2 with transforms not working as intended | ### 🐛 Describe the bug
When using the `wrap_dataset_for_transforms_v2` wrapper for `torchvision.datasets` classes it seems that the `transform` being passed during instantiation of the dataset is not utilized properly. The issue was observed the `VOCDetection` dataset:
- when using `RandomCrop`, passing it to `VOCDe... | closed | 2024-11-11T12:54:52Z | 2024-12-23T09:16:41Z | https://github.com/pytorch/vision/issues/8720 | [] | liopeer | 3 |
zappa/Zappa | flask | 386 | [Migrated] Deleted Resource resulting in `Invalid REST API identifier` | Originally from: https://github.com/Miserlou/Zappa/issues/967 by [yuric](https://github.com/yuric)
<!--- Provide a general summary of the issue in the Title above -->
## Context
<!--- Provide a more detailed introduction to the issue itself, and why you consider it to be a bug -->This is not an issue or a bug but a ... | closed | 2021-02-20T08:27:46Z | 2022-08-16T00:44:23Z | https://github.com/zappa/Zappa/issues/386 | [] | jneves | 1 |
Teemu/pytest-sugar | pytest | 32 | Possible to convert live video into animated GIF? | I wonder if the cool animated video at http://pivotfinland.com/pytest-sugar/
could be converted to an animated GIF?
Because GitHub, as far as I can tell, won't let us embed a video in a README, but you can embed an animated GIF and that would let the GitHub page have the sexy video.
I think I tried to convert it mys... | closed | 2014-02-13T16:07:09Z | 2014-12-12T22:14:13Z | https://github.com/Teemu/pytest-sugar/issues/32 | [] | msabramo | 2 |
autogluon/autogluon | scikit-learn | 4,832 | [BUG] [timeseries] Covariate regressor creates an empty folder under `AutogluonModels/ag-{TIMESTAMP}CatBoostModel`` | **Bug Report Checklist**
<!-- Please ensure at least one of the following to help the developers troubleshoot the problem: -->
- [x] I provided code that demonstrates a minimal reproducible example. <!-- Ideal, especially via source install -->
- [x] I confirmed bug exists on the latest mainline of AutoGluon via sour... | closed | 2025-01-23T12:55:15Z | 2025-01-28T19:52:14Z | https://github.com/autogluon/autogluon/issues/4832 | [
"bug",
"module: timeseries",
"module: core"
] | shchur | 0 |
fugue-project/fugue | pandas | 259 | [FEATURE] Duckdb support | **Describe the solution you'd like**
We should support DuckDB as a sql backend.
| closed | 2021-10-17T22:02:50Z | 2021-10-18T20:37:34Z | https://github.com/fugue-project/fugue/issues/259 | [
"enhancement",
"Fugue SQL"
] | goodwanghan | 0 |
kizniche/Mycodo | automation | 1,308 | Unable to Generate Camera Timelapses | Attempting to add a time-lapse using the basic function page using a working camera. I'm unable to generate any time-lapse stills now.
1. Images of the time-lapse information with error are shown.
2. No pictures are ever generated in the time-lapse photo. There were time-lapse photos from a different date, indicat... | closed | 2023-05-17T20:26:37Z | 2023-08-21T18:25:01Z | https://github.com/kizniche/Mycodo/issues/1308 | [
"bug",
"Fixed and Committed"
] | robocode-LAB | 1 |
biolab/orange3 | data-visualization | 6,435 | ODBC Support | **What's your use case?**
I would like to connect through oledb to an existing database (MonetDB here)
<!-- Is your request related to a problem, or perhaps a frustration? -->
Well, I can't connect to the said database :)
<!-- Tell us the story that led you to write this request. -->
**What's your proposed solut... | open | 2023-04-26T19:23:05Z | 2024-09-12T11:49:37Z | https://github.com/biolab/orange3/issues/6435 | [] | simonaubertbd | 3 |
jmcnamara/XlsxWriter | pandas | 212 | Content is unreadable in excel | I have the following code
```
workbook = xlsxwriter.Workbook('test.xls')
worksheet = workbook.add_worksheet()
e_opts = ['test5', 'other']
for i in range(10):
worksheet.write('A%d'%i, 'test1')
worksheet.write('B%d'%i, 'test2')
worksheet.write('C%d'%i, 'test3')
worksheet.write('D%d'%i, 'test4')
w... | closed | 2015-01-16T18:24:51Z | 2015-01-16T20:28:04Z | https://github.com/jmcnamara/XlsxWriter/issues/212 | [
"bug"
] | SeanWhipple | 2 |
microsoft/hummingbird | scikit-learn | 287 | Add support for sklearn MLPRegressor | Extend the existing sklearn MLPClassifier to also support MLPRegressor. | closed | 2020-09-03T20:18:56Z | 2020-09-04T04:36:57Z | https://github.com/microsoft/hummingbird/issues/287 | [] | scnakandala | 1 |
Ehco1996/django-sspanel | django | 603 | (1045, "Access denied for user 'root'@'172.18.0.4' (using password: YES)") | **问题的描述**
(1045, "Access denied for user 'root'@'172.18.0.4' (using password: YES)")
**项目的配置文件**
```
#--->服务端口 nginx
port=8080
#--->mysql
# mysql数据库设置 db.py
MYSQL_USER=root
MYSQL_PASSWORD=yourpass
# mysql服务设置
# mysql host默认请保持注释*
# MYSQL_HOS=mysql
# mysql服务密码设置
MYSQL_ROOT_PASSWORD=${MYSQL_PASSWORD}
#设置... | closed | 2021-12-09T07:32:45Z | 2021-12-28T00:34:30Z | https://github.com/Ehco1996/django-sspanel/issues/603 | [
"bug"
] | taotecode | 5 |
strawberry-graphql/strawberry | asyncio | 3,631 | strawberry.ext.mypy_plugin Pydantic 2.9.0 PydanticModelField.to_argument error missing 'model_strict' and 'is_root_model_root' | Hello!
It seems the Pydantic 2.9.0 version introduced a breaking change on PydanticModelField.to_argument adding two new arguments:
https://github.com/pydantic/pydantic/commit/d6df62aaa34c21272cb5fcbcbe3a8b88474732f8
and
https://github.com/pydantic/pydantic/commit/93ced97b00491da4778e0608f2a3be62e64437a8
##... | open | 2024-09-13T12:02:43Z | 2025-03-20T15:56:52Z | https://github.com/strawberry-graphql/strawberry/issues/3631 | [
"bug"
] | victor-nb | 0 |
timkpaine/lantern | plotly | 40 | plotly - pie | closed | 2017-10-10T01:33:08Z | 2017-10-19T04:53:44Z | https://github.com/timkpaine/lantern/issues/40 | [
"feature",
"plotly/cufflinks"
] | timkpaine | 0 | |
modelscope/modelscope | nlp | 701 | 加载本地数据集时报错NotImplementedError: Loading a dataset cached in a LocalFileSystem is not supported. | ```python
from modelscope.msdatasets import MsDataset
from modelscope.utils.constant import DownloadMode
ms_train_dataset = MsDataset.load(
'./data/garbage265',
subset_name='default',split='train',) # 加载训练集
```
使用以上代码加载本地自定义数据集报错
NotImplementedError: Loading a dataset cached in a Loc... | closed | 2023-12-28T08:52:28Z | 2024-06-08T01:49:39Z | https://github.com/modelscope/modelscope/issues/701 | [
"Stale"
] | 1006076811 | 3 |
qubvel-org/segmentation_models.pytorch | computer-vision | 294 | [Feature request] Add an option BatchNorm => Instance norm | I would like to be able to replace BatchNorm with Instance norm in the network at the initialization, including pre-trained backbones. | closed | 2020-12-06T17:48:37Z | 2020-12-07T15:10:36Z | https://github.com/qubvel-org/segmentation_models.pytorch/issues/294 | [] | ternaus | 1 |
tflearn/tflearn | data-science | 356 | Is there any way to disable printing loss? | when i use "fit" function, loss is printed on console each step
I guess if i can disable printing it, the speed of learning process will be faster
However, i don't know how to disable... (it's possible to disable printing accuracy though)
Is there any way?
| closed | 2016-09-23T15:38:16Z | 2016-09-25T14:28:11Z | https://github.com/tflearn/tflearn/issues/356 | [] | y-rok | 2 |
deepset-ai/haystack | pytorch | 8,692 | Document ID doesn't updated upon metadata update | **Describe the bug**
If you assign the `meta` field post initialization to a `Document`, the id of the document doesn't get updated.
This is e.g. done in the [PyPDFConverter](https://github.com/deepset-ai/haystack/blob/28ad78c73d6c11c9b77089aba42799508178a2fa/haystack/components/converters/pypdf.py#L225).
Document... | closed | 2025-01-09T12:23:59Z | 2025-02-13T09:01:32Z | https://github.com/deepset-ai/haystack/issues/8692 | [
"P3"
] | wochinge | 2 |
NullArray/AutoSploit | automation | 634 | Unhandled Exception (a2bc5a14e) | Autosploit version: `3.0`
OS information: `Linux-4.19.0-parrot1-20t-amd64-x86_64-with-Parrot-4.6-stable`
Running context: `autosploit.py`
Error meesage: `global name 'Except' is not defined`
Error traceback:
```
Traceback (most recent call):
File "/home/lnx-crew/3xploit/AutoSploit-master/autosploit/main.py", line 113,... | closed | 2019-04-07T00:24:37Z | 2019-04-18T17:31:35Z | https://github.com/NullArray/AutoSploit/issues/634 | [] | AutosploitReporter | 0 |
raphaelvallat/pingouin | pandas | 408 | Remove call to sns.despine in paired_plot | ### Discussed in https://github.com/raphaelvallat/pingouin/discussions/407
<div type='discussions-op-text'>
<sup>Originally posted by **timobage** February 17, 2024</sup>
Hi,
how can I change the y-axis limits of the plot_paired() function?
If I use ax.set_ylim() it only shrinks/expands the existing plots but ... | closed | 2024-02-20T19:43:00Z | 2024-03-02T12:05:59Z | https://github.com/raphaelvallat/pingouin/issues/408 | [
"bug :boom:"
] | raphaelvallat | 0 |
modin-project/modin | data-science | 7,391 | whats the fastest way to add a new column that already has the same partitions (probably)? | There are a bunch of ways to add a column to a dataframe..
what is the fastest with modin?
say get a new column by applying a function to another one
```py
new_c = df['column'].apply(lambda x: abs(x))
```
the resulting series should have the same partitions as the dataframe right?
we can use...
merge, or... | open | 2024-09-06T16:26:28Z | 2024-09-06T16:58:22Z | https://github.com/modin-project/modin/issues/7391 | [
"question ❓",
"Triage 🩹"
] | Liquidmasl | 1 |
thtrieu/darkflow | tensorflow | 1,197 | How to put a darkflow model into Android Studio | I created an object detection model that I trained through darkflow. and I changed this to the tflite format to put into Android Studio
but I when I try to add metadata to this file, the following error occurs:
`ValueError: The number of output tensors (1) should match the number of output tensor metadata (4)`
F... | open | 2021-05-15T07:27:20Z | 2021-05-15T07:27:20Z | https://github.com/thtrieu/darkflow/issues/1197 | [] | M1nseoPark | 0 |
sinaptik-ai/pandas-ai | data-visualization | 1,343 | Questions about the train function | Thanks for the great work.
I have several questions about the instruct train function
1. May I know that what vectorDB perform during the train? Does it act as a RAG?
2. After the train, is that anyway to save the trained model or stuff? Or it requires to call the train function for the prompt everytime?
3. For the... | closed | 2024-08-30T02:21:56Z | 2025-02-11T16:00:11Z | https://github.com/sinaptik-ai/pandas-ai/issues/1343 | [] | mrgreen3325 | 3 |
WZMIAOMIAO/deep-learning-for-image-processing | deep-learning | 394 | 想问下有没有分割网络读取coco数据集的代码? | 想问下有没有分割网络读取coco数据集的代码? | closed | 2021-11-06T09:25:17Z | 2021-11-13T07:25:25Z | https://github.com/WZMIAOMIAO/deep-learning-for-image-processing/issues/394 | [] | ily666666 | 1 |
StackStorm/st2 | automation | 5,087 | NO_PROXY environment variable is not considered while installing the pack dependencies | ## SUMMARY
I have HTTP_PROXY, HTTPS_PROXY and NO_PROXY environment variables set in my setup and also, I maintain an internal PyPI hosting most of the python dependencies. Now, I want the `st2 pack install file:///path/to/pack/folder` command to download python dependencies from this internal PyPI instead of the offic... | open | 2020-11-20T11:54:24Z | 2025-02-14T07:48:47Z | https://github.com/StackStorm/st2/issues/5087 | [
"bug"
] | RaviTezu | 9 |
plotly/dash | flask | 3,008 | Dash slow pattern-matching performance on many elements | Hello,
First, thank you for your amazing work on Dash.
**Describe your context**
```
dash 2.17.1
dash_ag_grid 31.2.0
dash-bootstrap-components 1.4.1
dash-core-components 2.0.0
dash-html-components 2.0.0
dash-table 5.0.0
jupyter-dash 0... | open | 2024-09-19T08:25:15Z | 2024-12-21T11:10:35Z | https://github.com/plotly/dash/issues/3008 | [
"performance",
"bug",
"P3"
] | Spriteware | 4 |
zappa/Zappa | flask | 708 | [Migrated] `certify` custom domain doesn't take effect until `update` is called | Originally from: https://github.com/Miserlou/Zappa/issues/1796 by [pickledish](https://github.com/pickledish)
<!--- Provide a general summary of the issue in the Title above -->
## Context
<!--- Provide a more detailed introduction to the issue itself, and why you consider it to be a bug -->
<!--- Also, please make... | closed | 2021-02-20T12:40:55Z | 2024-04-13T18:14:20Z | https://github.com/zappa/Zappa/issues/708 | [
"no-activity",
"auto-closed"
] | jneves | 2 |
tensorflow/tensor2tensor | machine-learning | 1,472 | Dependency mismatch for Cloud ML Engine runtime 1.12 | ### Description
TensorFlow Probability has just released 0.6.0 to PyPI, which expects TF 1.13.1. The current Runtime Version on Cloud ML Engine comes with TF 1.12.0. Without either:
1. upgrading the default tensorflow version to 1.13.1 on ML Engine, or
2. downgrading the version of tensorflow-probability to 0.5.0,... | closed | 2019-02-26T23:21:49Z | 2020-12-14T07:46:21Z | https://github.com/tensorflow/tensor2tensor/issues/1472 | [] | jvmncs | 5 |
biolab/orange3 | pandas | 6,250 | File widget: URL is lost when saving and reopening OWS file |
**What's wrong?**
If I specify a (specific type of?) URL referring to a data file in the File widget, then save the workflow and reopen it, the URL field is empty and the widget produces an error "No file selected"
**How can we reproduce the problem?**
Place a File widget in the canvas, double-click on it, se... | closed | 2022-12-09T11:34:56Z | 2023-01-20T07:41:15Z | https://github.com/biolab/orange3/issues/6250 | [
"bug"
] | wvdvegte | 1 |
keras-team/keras | pytorch | 20,706 | Loaded Keras Model Throws Error While Predicting (Likely Issues with Masking) | I am currently developing and testing a RNN that relies upon a large amount of data for training, and so have attempted to separate my training and testing files. I have one file where I create, train, and save a tensorflow.keras model to a file 'model.keras' I then load this model in another file and predict some valu... | closed | 2024-12-31T20:46:49Z | 2025-01-23T21:57:52Z | https://github.com/keras-team/keras/issues/20706 | [
"type:Bug"
] | JoeDoyle12 | 4 |
ets-labs/python-dependency-injector | asyncio | 58 | Add docs for @inject decorator | closed | 2015-05-08T14:48:03Z | 2015-08-05T14:48:10Z | https://github.com/ets-labs/python-dependency-injector/issues/58 | [
"docs"
] | rmk135 | 0 | |
geex-arts/django-jet | django | 87 | how to enable multiselect actions in pages that opens in popup windows? | When opening an admin list in a popup window, the column with checkoxes to do multiselection does not appear, and combo with actions is not appearing also...
I've done a search and i«m not finding what exactly how can i enable this... basically it's related with GET var ?_popup=1, but can't find the files i should cha... | closed | 2016-07-11T21:58:02Z | 2016-08-27T12:58:42Z | https://github.com/geex-arts/django-jet/issues/87 | [] | carlosfvieira | 7 |
proplot-dev/proplot | data-visualization | 300 | x axis inverts when using plot with negative y | <!-- Thanks for helping us make proplot a better package! If this is a bug report, please use the template provided below. If this is a feature request, you can delete the template text (just try to be descriptive with your request). -->
### Description
within cartesian grid, using plot, x axis inverts with y set... | closed | 2021-11-01T19:17:22Z | 2021-11-03T14:39:00Z | https://github.com/proplot-dev/proplot/issues/300 | [
"wontfix",
"documentation"
] | dewinkelwaar | 4 |
falconry/falcon | api | 2,314 | Get rid of `setup.cfg` | `setup.cfg` does not really contribute much value anymore, almost everything can be migrated to `pyproject.toml`.
We will have to continue augmenting `pyproject.toml` with `setup.py` though, because it is where we programmatically manage our build process. | closed | 2024-08-31T09:06:16Z | 2024-08-31T19:44:16Z | https://github.com/falconry/falcon/issues/2314 | [
"breaking-change",
"maintenance"
] | vytas7 | 0 |
quantmind/pulsar | asyncio | 158 | test suite does not stop with ctrl-C | The reason being all test classes are loaded in the event loop.
| closed | 2015-09-07T13:14:52Z | 2015-10-02T13:46:45Z | https://github.com/quantmind/pulsar/issues/158 | [
"bug",
"test"
] | lsbardel | 3 |
ultralytics/yolov5 | machine-learning | 13,019 | Issue when try to validate openvino format model | ### Search before asking
- [X] I have searched the YOLOv5 [issues](https://github.com/ultralytics/yolov5/issues) and found no similar bug report.
### YOLOv5 Component
Validation
### Bug
The next script, to validate a trained yolov5 works well:
!python ./yolov5_train100kbdd/yolov5s_originsize_300epochs... | closed | 2024-05-16T17:36:09Z | 2024-06-30T00:24:58Z | https://github.com/ultralytics/yolov5/issues/13019 | [
"bug",
"Stale"
] | FrancoArtale | 4 |
reloadware/reloadium | flask | 98 | Reloadium doesn't work on M1 mac with Python 3.9 | ## Describe the bug*
On some M1 macs, reloadium is working fine with Pyton 3.9. On few other macs, it is failing to run.
## To Reproduce
Not able to reproduce the bug on all M1 macs.
## Expected behavior
Reloadium should run the application.
## Screenshots
<img width="1280" alt="Screenshot 2023-02... | closed | 2023-02-06T03:16:06Z | 2023-02-06T06:36:05Z | https://github.com/reloadware/reloadium/issues/98 | [] | ChillarAnand | 7 |
pandas-dev/pandas | pandas | 60,322 | BUG: Specifying `hour` param, but not year, month, day in pandas.Timestamp() sets hour-value as minutes | ### Pandas version checks
- [X] I have checked that this issue has not already been reported.
- [X] I have confirmed this bug exists on the [latest version](https://pandas.pydata.org/docs/whatsnew/index.html) of pandas.
- [ ] I have confirmed this bug exists on the [main branch](https://pandas.pydata.org/docs/dev/ge... | closed | 2024-11-15T11:40:32Z | 2024-11-17T13:41:11Z | https://github.com/pandas-dev/pandas/issues/60322 | [
"Bug",
"Duplicate Report",
"Timestamp"
] | christoffer-hk | 2 |
ml-tooling/opyrator | pydantic | 4 | Finalize docker export capabilities | **Feature description:**
Finalize capabilities to export an opyrator to a Docker image.
The export can be executed via command line:
```bash
opyrator export my_opyrator:hello_world --format=docker my-opyrator-image:latest
```
_💡 The Docker export requires that Docker is installed on your machine._
Af... | closed | 2021-04-19T10:01:47Z | 2023-07-27T14:30:30Z | https://github.com/ml-tooling/opyrator/issues/4 | [
"feature",
"stale"
] | lukasmasuch | 6 |
CorentinJ/Real-Time-Voice-Cloning | tensorflow | 583 | Email | @blue-fish Hey I have some question regarding the repo, would it be possible to get in touch with you through email? | closed | 2020-10-31T10:10:32Z | 2020-10-31T17:47:57Z | https://github.com/CorentinJ/Real-Time-Voice-Cloning/issues/583 | [] | JakubReha | 1 |
flairNLP/flair | nlp | 3,356 | [Question]: How to train an end-to-end Entity Linking model? | ### Question
Hello,
I am interested in implementing an end-to-end Entity Linking model using a custom dataset. I would greatly appreciate any guidance on how to proceed, particularly concerning the required input data format, the training procedure, and the inference process.
Thank you in advance for your time a... | open | 2023-10-25T10:06:57Z | 2024-08-06T09:01:09Z | https://github.com/flairNLP/flair/issues/3356 | [
"question"
] | anna-shopova | 6 |
PeterL1n/BackgroundMattingV2 | computer-vision | 22 | problem running inference_images.py | It seems that there arer multiple threads running at the same time since I got this overrride question many times:
This is what I see when answering no (the output folder does not exist before)
```
(bgm2) C:\ZeroBox\src\BackgroundMattingV2> python inference_images.py --model-type mattingrefine --model-backbone mob... | closed | 2020-12-31T03:51:41Z | 2020-12-31T21:29:15Z | https://github.com/PeterL1n/BackgroundMattingV2/issues/22 | [] | jinzishuai | 5 |
horovod/horovod | pytorch | 2,957 | Large Batch Simulation Breaks with Mixed Precision | **Environment:**
1. Framework: TensorFlow
2. Framework version: 2.3
3. Horovod version: 0.22.0
7. Python version: 3.7
**Checklist:**
1. Did you search issues to find if somebody asked this question before? yes
2. If your question is about hang, did you read [this doc](https://github.com/horovod/horovod/blob/... | open | 2021-06-07T19:43:15Z | 2021-06-21T17:07:35Z | https://github.com/horovod/horovod/issues/2957 | [
"bug"
] | czmrand | 2 |
apachecn/ailearning | python | 432 | 第9章_树回归 - ApacheCN | http://ailearning.apachecn.org/ml/9.TreeRegression/
ApacheCN 专注于优秀项目维护的开源组织 | closed | 2018-08-24T07:10:06Z | 2021-09-07T17:40:30Z | https://github.com/apachecn/ailearning/issues/432 | [
"Gitalk",
"162d4b1d1067f1f6264103c55a777526"
] | jiangzhonglian | 0 |
jonaswinkler/paperless-ng | django | 324 | Quotation marks in PAPERLESS_FILENAME_FORMAT cause malformed filenames | Hi Jonas,
I just tried to change the filename format by setting (in docker-compose.env)
`PAPERLESS_FILENAME_FORMAT="{created_year}/{correspondent}/{created_year}-{created_month}-{created_day} - {title}"`.
Paperless does not complain and starts up as usual. I let paperelss consume a new file, which also works ... | closed | 2021-01-11T18:14:01Z | 2021-02-03T00:20:16Z | https://github.com/jonaswinkler/paperless-ng/issues/324 | [] | niarbx | 4 |
FujiwaraChoki/MoneyPrinter | automation | 142 | [BUG] | **Describe the bug**
A clear and concise description of what the bug is.
**To Reproduce**
Steps to reproduce the behavior:
1. Go to 'front page '
2. Click on ' Generate'
3. Scroll down to '....'
4. See error 'An error occurred. Please try again later.'
**Expected behavior**
A clear and concise description ... | closed | 2024-02-10T10:02:32Z | 2024-02-10T10:45:51Z | https://github.com/FujiwaraChoki/MoneyPrinter/issues/142 | [] | FranklinOP-IND | 3 |
apache/airflow | data-science | 47,521 | Test Deployment via breeze in KinD/K8s need to realx Same Origin Policy | ### Body
When a local K8s cluster (via KinD) is started through `breeze k8s ...` and you attempt to open the exposed URL in the browser (it uses a random forwarded port, e.g. http://localhost:13582/ ) then you get the following errors:
Cross-Origin Request Blocked: The Same Origin Policy disallows reading the remote ... | closed | 2025-03-07T22:29:05Z | 2025-03-09T08:18:50Z | https://github.com/apache/airflow/issues/47521 | [
"kind:bug",
"area:dev-env",
"area:dev-tools",
"kind:meta",
"area:UI"
] | jscheffl | 2 |
chmp/ipytest | pytest | 123 | Command line / CI-CD / IDE execution | Firstly, thanks for an awesome tool. This is exactly what I was looking for.
The docstring to `_impl.run` states:
```
**NOTE:** In the default configuration `ipytest.run()` will not raise
exceptions, when tests fail. To raise exceptions on test errors, e.g.,
inside a CI/CD context, use `ipytest.autoconfig(... | closed | 2025-02-01T08:41:27Z | 2025-02-17T07:09:44Z | https://github.com/chmp/ipytest/issues/123 | [] | MusicalNinjaDad | 6 |
pallets/flask | python | 5,245 | NameError: name 'Response' is not defined | I have a view like this:
```
@pydantic.validate_arguments(config={"arbitrary_types_allowed": True})
def my_view(
)-> flask.typing.ResponseReturnValue:
...
```
This view is failing during tests:
```
@pd.validate_arguments(config={"arbitrary_types_allowed": True})
pydantic/decorator.py:36: in pydantic... | closed | 2023-08-31T07:11:47Z | 2023-09-15T00:05:39Z | https://github.com/pallets/flask/issues/5245 | [] | warvariuc | 4 |
sqlalchemy/alembic | sqlalchemy | 1,386 | Autogenerate renders TypeDecorator instance instead of underlying impl type | **Describe the bug**
This isn't a bug _per se_, but a small improvement for autogenerate when using TypeDecorator.
When a TypeDecorator is used in a column definition, e.g.:
```py
"""
File: app/models/foo.py
"""
from sqlalchemy.dialects.postgresql import JSONB
from sqlalchemy.types import TypeDecorator
.... | open | 2024-01-05T05:50:59Z | 2024-09-18T12:28:11Z | https://github.com/sqlalchemy/alembic/issues/1386 | [
"autogenerate - rendering",
"use case",
"cookbook requested"
] | saifelse | 7 |
scikit-multilearn/scikit-multilearn | scikit-learn | 12 | Implement a general ensemble of classifiers classifier | J. Read, B. Pfahringer, G. Holmes, E. Frank, Classifier chains for multi-label classification, in: Proceedings of the 20th European Conference on Machine Learning, 2009, pp. 254–269.
Ensembles of classifier chains (ECC ) [16] are an ensemble multi-label classification technique that uses classifier chains as a base ... | open | 2014-12-06T14:30:17Z | 2023-03-14T16:56:55Z | https://github.com/scikit-multilearn/scikit-multilearn/issues/12 | [
"enhancement",
"help wanted"
] | niedakh | 3 |
coqui-ai/TTS | deep-learning | 3,656 | [Bug] bug in tts_to_file | ### Describe the bug
```python
from TTS.api import TTS
tts_ins = TTS('tts_models/multilingual/multi-dataset/xtts_v2')
tts_ins.tts_to_file(
text='因为树脂这个材料它比较容易染色所以久了之后呢树脂贴片就没有刚开始那么漂亮了那再来就是树脂这个材料它比较软。',
file_path="output.wav",
speaker="Ana Florence", # 使用默认的人声
language="zh-cn",
split_sente... | closed | 2024-04-01T09:18:39Z | 2024-06-26T16:49:22Z | https://github.com/coqui-ai/TTS/issues/3656 | [
"bug",
"wontfix"
] | forthcoming | 1 |
FactoryBoy/factory_boy | django | 965 | KeyError: 'locale' | #### Description
KeyError: 'locale' generated when calling Faker.
factory/faker.py:46: KeyError
Seems that the `extra` dictionary is empty, but `locale` key is expected.
#### To Reproduce
running unit test with tox
python 3.10; factory-boy==3.2.1 faker==13.15.0
##### Model / Factory code
```python
@pytest.... | closed | 2022-07-17T06:20:52Z | 2024-06-18T07:42:50Z | https://github.com/FactoryBoy/factory_boy/issues/965 | [
"Bug"
] | radohristov | 16 |
snarfed/granary | rest-api | 208 | [reddit] support @self, @friends, and @all in get_activities() | right now it only supports search and activity_id, so bridgy only finds posts that link to your site. this would fill in the rest of bridgy's backfeed functionality for people who POSSE their posts to reddit without backlinks. cc @stedn | open | 2020-04-30T23:14:42Z | 2020-04-30T23:14:42Z | https://github.com/snarfed/granary/issues/208 | [] | snarfed | 0 |
2noise/ChatTTS | python | 246 | 为什么会自己加字? | 比如:`白 日 依 山 尽 , 黄 河 入 海 流。 欲 穷 千 里目 , 更 上 一 层 楼 。`
生成文字为:`白 日 依 山 尽 [uv_break] , 黄 河 入 海 流 [uv_break] 。 欲 穷 千 里 目 [uv_break] , 那 更 上 一 层 楼 [uv_break] 。`
为什么要加一个**那**字,有办法保持原文吗? | closed | 2024-06-04T04:45:58Z | 2024-06-19T03:50:12Z | https://github.com/2noise/ChatTTS/issues/246 | [] | zhouhao27 | 3 |
netbox-community/netbox | django | 18,577 | Translation Error in Device View: "No está atormentado" should be "No tiene rack asignado" | ### Language
Spanish
### ISO 639-1 code
es
### Volunteer
Yes
### Comments
The correct translation for this line should be:
"**No tiene rack asignado**" instead of "**No está atormentado**."
"Atormentado" translates to "tormented" and is not related to the rack device's status. The intended meaning refers to w... | closed | 2025-02-05T14:35:50Z | 2025-02-05T14:42:02Z | https://github.com/netbox-community/netbox/issues/18577 | [] | SorianoTech | 1 |
healthchecks/healthchecks | django | 847 | SMTP session exception | Hello together,
unfortunately, the update to version 2.9.2 works not for me. Some hours after this update, i got a error message with smtp session exception. With version 2.8.1 i don't get any error messages.
The error log says that the db isn't reachable, but other application that use the same db still work as ... | closed | 2023-06-20T10:01:02Z | 2023-07-02T09:37:52Z | https://github.com/healthchecks/healthchecks/issues/847 | [] | techsolo12 | 5 |
MaartenGr/BERTopic | nlp | 1,240 | Output size for supervised topic modeling (multi-class classification) | I am trying to change the output size of my model to allow the classification of multiple classes. Following a common PyTorch/CNN strategy, this would be the equivalent of returning something like `nn.Linear(K, 30)`, in which I could compare the output of my model [0.0123, 0.245, 0.113, ..., 0.136] of shape `[30]` with... | closed | 2023-05-08T15:04:15Z | 2023-09-27T08:56:42Z | https://github.com/MaartenGr/BERTopic/issues/1240 | [] | wlcosta | 2 |
iperov/DeepFaceLab | machine-learning | 667 | Remove lens distortion from src images before training? | Question: Would it make sense to remove lens distortion before you extract the faces and start training? When src images are collected from a huge variety of sources, lens distortion is quite a big variable.
I guess todays method is to train with uncorrected images, and then rebuild uncorrected.
It would be inte... | closed | 2020-03-21T17:41:39Z | 2020-03-22T18:32:06Z | https://github.com/iperov/DeepFaceLab/issues/667 | [] | Tesla32X | 2 |
graphistry/pygraphistry | jupyter | 304 | [FEA] List support in hypergraph entity extraction | **Is your feature request related to a problem? Please describe.**
When plotting some event data, some of the columns had entity lists ("topics", "attending_groups", ...) that it'd help to auto-link
**Describe the solution you'd like**
```python
df = pd.DataFrame({'event': ['e1', 'e2'], 'attendees': [ ['a', '... | open | 2022-02-03T05:42:05Z | 2022-02-03T05:42:05Z | https://github.com/graphistry/pygraphistry/issues/304 | [
"enhancement"
] | lmeyerov | 0 |
rthalley/dnspython | asyncio | 612 | Allow configuring dns.rdata._chunksize | [`dns.rdata._chunksize`](https://github.com/rthalley/dnspython/blob/65c4a968686dd0c3bca1ed1a5fd13fa0d2f1b441/dns/rdata.py#L35) currently is a private setting.
However, some other applications require certain chunk sizes (or no chunking at all, as is achieved when `_chunksize` is falsy). For example, the PowerDNS API... | closed | 2020-12-14T17:21:24Z | 2021-01-05T20:02:34Z | https://github.com/rthalley/dnspython/issues/612 | [] | peterthomassen | 3 |
matplotlib/cheatsheets | matplotlib | 38 | Legend placement error | Legend placement numbers in the cheatsheet are inconsistent with matplotlib documentation. (e.g. for lower-right, loc=3 in the cheatsheet instead of loc=4).
https://matplotlib.org/3.1.1/api/_as_gen/matplotlib.pyplot.legend.html | closed | 2020-07-20T09:33:56Z | 2020-07-28T08:06:33Z | https://github.com/matplotlib/cheatsheets/issues/38 | [] | bertrandcz | 1 |
huggingface/diffusers | pytorch | 11,144 | FlaxUNet2DConditionModel is not initialized with correct dtypes | ### Describe the bug
The FlaxUNet2DConditionModel allows specifying the dtype of the weights. Supplying a dtype different from float32 does not seem to be propagated to the actual model. This is imo different from https://github.com/huggingface/diffusers/issues/2068 since the afaik the code has correct dtype initializ... | open | 2025-03-23T17:47:33Z | 2025-03-24T15:59:04Z | https://github.com/huggingface/diffusers/issues/11144 | [
"bug"
] | wittenator | 3 |
jupyterlab/jupyter-ai | jupyter | 381 | azure-chat-openai | <!-- Welcome! Thank you for contributing. These HTML comments will not render in the issue.
I want connect Jupyter_ai magic commands using azure open ai models ..Any suggestion how to pass deployement id and Engine name?
###
Before creating a new issue:
* Search for relevant issues
* Follow the issue reportin... | open | 2023-09-07T11:08:21Z | 2024-02-09T04:29:50Z | https://github.com/jupyterlab/jupyter-ai/issues/381 | [
"bug"
] | bmshambu | 7 |
ExpDev07/coronavirus-tracker-api | fastapi | 210 | Reimplementation of country_code - why? | The reimplementation of the `country_code` function in the 73f02fb does at least 3 dictionary lookups for any input: 3 lookups in the best-case and 5 lookups in worst-case scenarios:
```python
# 1. # 2.
if not country in is_3166_1 and country in synonyms:
country... | closed | 2020-03-26T23:38:11Z | 2020-03-28T10:35:04Z | https://github.com/ExpDev07/coronavirus-tracker-api/issues/210 | [
"enhancement"
] | Bost | 11 |
chainer/chainer | numpy | 7,838 | `chainer.backend.copyto` cannot copy chainerx array to cupy | * Code to reproduce
```python
import chainer
import numpy
for dst_device in ['@numpy', '@cupy:0', '@intel64']:
for src_device in ['native', 'cuda:0']:
print((dst_device, src_device))
dst = chainer.get_device(dst_device).send(
numpy.array([1, 2], numpy.float32))
src = c... | closed | 2019-07-31T00:48:09Z | 2019-08-07T04:29:16Z | https://github.com/chainer/chainer/issues/7838 | [
"cat:bug",
"prio:high",
"pr-ongoing"
] | toslunar | 0 |
huggingface/datasets | computer-vision | 6,897 | datasets template guide :: issue in documentation YAML | ### Describe the bug
There is a YAML error at the top of the page, and I don't think it's supposed to be there
### Steps to reproduce the bug
1. Browse to [this tutorial document](https://github.com/huggingface/datasets/blob/main/templates/README_guide.md)
2. Observe a big red error at the top
3. The rest of the ... | closed | 2024-05-13T17:33:59Z | 2024-05-16T14:28:17Z | https://github.com/huggingface/datasets/issues/6897 | [] | bghira | 2 |
PeterL1n/BackgroundMattingV2 | computer-vision | 102 | 请问可以分布式计算吗? | 您好,我现在手里有8块3090的GPU卡,我修改了以下代码
model = model.to(device).eval()
model.load_state_dict(torch.load(args.model_checkpoint, map_location=device), strict=False)
为
model = model.to(device).eval()
BM = model.load_state_dict(torch.load(args.model_checkpoint, map_location=device), strict=False)
BM = nn.DataParallel(BM)
... | closed | 2021-05-20T07:07:27Z | 2021-08-10T15:11:04Z | https://github.com/PeterL1n/BackgroundMattingV2/issues/102 | [] | zhanghonglishanzai | 8 |
reloadware/reloadium | pandas | 198 | [Feature Request] attach/detach `reloadium` at runtime | It would be nice if it was possible to attach `reloadium` at runtime:
- e.g. when you're debugging some script as usual and then you realize it would be helpful to have reloadium features now, then you would be able to load it, debug the issue and unload it and keep working
- in my case I work on Blender (3D soft... | open | 2024-07-14T17:55:41Z | 2024-07-15T15:04:09Z | https://github.com/reloadware/reloadium/issues/198 | [] | Andrej730 | 2 |
graphql-python/graphene-django | graphql | 1,065 | CSRF cookie being set regardless of whether csrf middleware is used | My project does not use CSRF middleware:
```python
MIDDLEWARE = [
# 'django.middleware.csrf.CsrfViewMiddleware'
}
```
And the view is set as exempt from CSRF:
```python
urlpatterns = [
path("graphql", csrf_exempt(GraphQLView.as_view())),
]
```
But despite this, a CSRF token is still *always*... | open | 2020-11-21T14:50:13Z | 2020-11-21T14:50:13Z | https://github.com/graphql-python/graphene-django/issues/1065 | [
"🐛bug"
] | samirelanduk | 0 |
ivy-llc/ivy | pytorch | 28,100 | Fix Frontend Failing Test: tensorflow - attribute.paddle.real | To-do List: https://github.com/unifyai/ivy/issues/27499 | closed | 2024-01-28T19:04:22Z | 2024-01-29T13:08:52Z | https://github.com/ivy-llc/ivy/issues/28100 | [
"Sub Task"
] | Sai-Suraj-27 | 0 |
521xueweihan/HelloGitHub | python | 2,756 | 【开源自荐】一款基于 bpf 的 dns 查询实时追踪工具 | ## 推荐项目
<!-- 这里是 HelloGitHub 月刊推荐项目的入口,欢迎自荐和推荐开源项目,唯一要求:请按照下面的提示介绍项目。-->
<!-- 点击上方 “Preview” 立刻查看提交的内容 -->
<!--仅收录 GitHub 上的开源项目,请填写 GitHub 的项目地址-->
- 项目地址:https://github.com/chenjiandongx/dnstrack
<!--请从中选择(C、C#、C++、CSS、Go、Java、JS、Kotlin、Objective-C、PHP、Python、Ruby、Rust、Swift、其它、书籍、机器学习)-->
- 类别:Go
... | open | 2024-05-29T05:02:15Z | 2024-05-29T06:05:47Z | https://github.com/521xueweihan/HelloGitHub/issues/2756 | [] | chenjiandongx | 0 |
localstack/localstack | python | 11,983 | bug: Batch of ECS FargateContainer doesn't pass secrets to environment variable | ### Is there an existing issue for this?
- [X] I have searched the existing issues
- #8492
- But, the different thing is this Issue is for Aws Batch.
### Current Behavior
Job Definition is setting with EcsFargateContainerDefinition.
The option of EcsFargateContainerDefinitionProp environment is passed to en... | open | 2024-12-04T03:10:42Z | 2024-12-05T13:40:04Z | https://github.com/localstack/localstack/issues/11983 | [
"type: bug",
"aws:batch",
"aws:ecs",
"aws:fargate",
"status: backlog"
] | BrianKim-git | 0 |
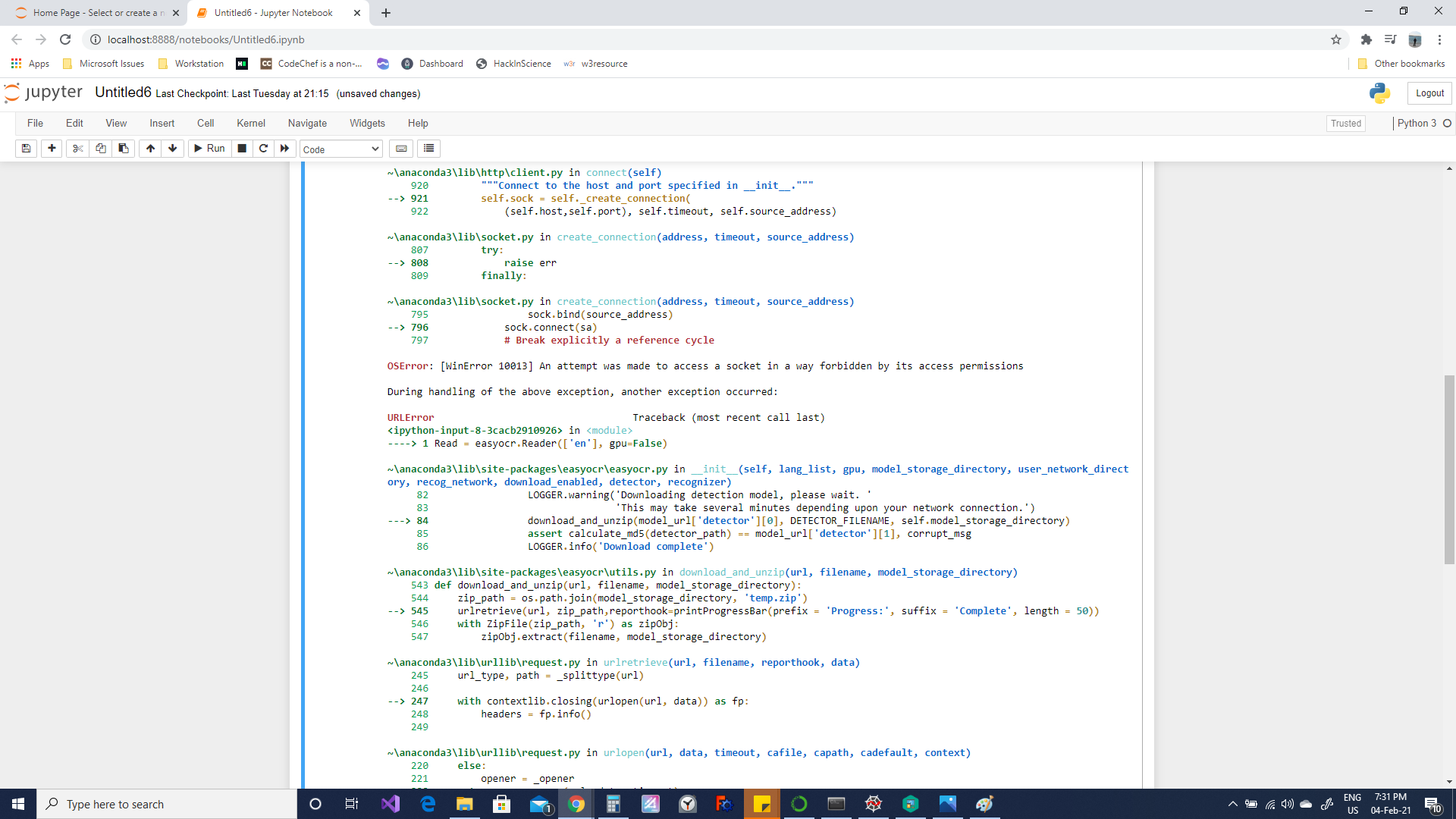
JaidedAI/EasyOCR | deep-learning | 366 | Error in downloading Languages in EasyOCR- OS Error and URL Error- Help | I face this error, kindly help, I am new to this OCR topic...

 or [Embedders](https://docs.haystack.deepset.ai/docs/choosing-the-right-embedder), on how to choose the right ... | open | 2025-03-19T16:02:42Z | 2025-03-20T15:44:51Z | https://github.com/deepset-ai/haystack/issues/9071 | [
"type:documentation",
"P2"
] | dfokina | 0 |
aiogram/aiogram | asyncio | 927 | Task was destroyed but it is pending | ## Context
I'm using aiogram to design my bot into telegrams, but as the load increased, I started getting a weird bug.
* Operating System: Linux (Heroku-22)
* Python Version: 3.9.13
* aiogram version: 2.20
* aiohttp version: 3.8.1
* uvloop version (if installed): 0.16.0
## Expected Behavior
I expect th... | closed | 2022-06-18T22:41:29Z | 2023-08-04T18:23:53Z | https://github.com/aiogram/aiogram/issues/927 | [
"needs triage",
"2.x"
] | koval01 | 3 |
AUTOMATIC1111/stable-diffusion-webui | pytorch | 16,136 | [Bug]: /sdapi/v1/txt2img returns 404 with --nowebui argument | ### Checklist
- [X] The issue exists after disabling all extensions
- [X] The issue exists on a clean installation of webui
- [ ] The issue is caused by an extension, but I believe it is caused by a bug in the webui
- [X] The issue exists in the current version of the webui
- [X] The issue has not been reported before... | closed | 2024-07-03T10:31:29Z | 2024-07-03T13:47:57Z | https://github.com/AUTOMATIC1111/stable-diffusion-webui/issues/16136 | [
"bug-report"
] | ibrahimsn98 | 1 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.