repo_name stringlengths 9 75 | topic stringclasses 30
values | issue_number int64 1 203k | title stringlengths 1 976 | body stringlengths 0 254k | state stringclasses 2
values | created_at stringlengths 20 20 | updated_at stringlengths 20 20 | url stringlengths 38 105 | labels listlengths 0 9 | user_login stringlengths 1 39 | comments_count int64 0 452 |
|---|---|---|---|---|---|---|---|---|---|---|---|
axnsan12/drf-yasg | django | 720 | api/swagger/?format=openapi response 500 |

| open | 2021-06-04T09:25:46Z | 2025-03-07T12:13:02Z | https://github.com/axnsan12/drf-yasg/issues/720 | [
"triage"
] | dpreal | 2 |
liangliangyy/DjangoBlog | django | 77 | 新增文章时如果同时指定 tag,保存时会出错 | 出错原因应该是同时写入两张表blog_article和 blog_article_tags,写入顺序错了。估计是先写入blog_article_tags,因为外键不存在而报错。 | closed | 2018-01-12T08:28:27Z | 2018-01-14T02:43:27Z | https://github.com/liangliangyy/DjangoBlog/issues/77 | [] | xmyangz | 2 |
reloadware/reloadium | flask | 178 | Reloadium fails to start | ## Describe the bug*
'RW_IDE_NAME': 'PyCharm 2023.3.2',
'RW_IDE_PLUGINVERSION': '1.3.4',
```Traceback (most recent call last):
File "C:\Users\Administrator\.reloadium\package\3.9\reloadium\corium\l111ll1111ll1l11Il1l1.py", line 189, in ll1l1ll11l1lllllIl1l1
File "D:\a\reloadware\reloadware\reload\packa... | closed | 2024-01-10T10:04:22Z | 2024-02-20T14:16:08Z | https://github.com/reloadware/reloadium/issues/178 | [] | hyabean | 2 |
RomelTorres/alpha_vantage | pandas | 190 | Add github actions for builds to automatically run | So that tests run on every PR automatically, and those PRs that don't pass tests can't be approved.
Github actions is new (as of a few months ago) but looks really powerful. | closed | 2020-02-13T19:42:24Z | 2021-11-19T18:45:13Z | https://github.com/RomelTorres/alpha_vantage/issues/190 | [
"enhancement"
] | PatrickAlphaC | 1 |
lyhue1991/eat_tensorflow2_in_30_days | tensorflow | 53 | Suggest a virtual environment. | I suggest a virtual environment for this tutorial
For one thing, it decouples the change in new relase of tf and the development environment we use. it saves authors' effort to answer tf version related problem and delegate them back to tf developers.
And it also saves readers effort to figure out missing package.... | open | 2020-06-11T13:58:49Z | 2020-06-11T14:01:39Z | https://github.com/lyhue1991/eat_tensorflow2_in_30_days/issues/53 | [] | neilteng | 0 |
unionai-oss/pandera | pandas | 1,169 | Allow checks based on data types | **Is your feature request related to a problem? Please describe.**
Imagine you have this schema:
```
schema = pa.DataFrameSchema({
"a": pa.Column(int, checks=pa.Check.le(10)),
"b": pa.Column(float, checks=pa.Check.lt(-1.2)),
"c": pa.Column(str, checks=pa.Check.le(20)),
})
```
In above schema, c... | open | 2023-04-27T19:26:52Z | 2023-06-12T16:55:50Z | https://github.com/unionai-oss/pandera/issues/1169 | [
"enhancement"
] | NeerajMalhotra-QB | 1 |
graphdeco-inria/gaussian-splatting | computer-vision | 379 | Can't resume training from checkpoint | Hi there. I'm training on some photos of and alleyway. I've done 7k iterations in one session (loss 0.18). 30k iterations in the next session (loss 0.09). So obviously I'd like to keep training BUT this time I'd like to use the checkpoint saved at 30k iterations as my starting point to save training time.
Unfortunat... | open | 2023-10-24T00:37:32Z | 2024-01-17T23:16:45Z | https://github.com/graphdeco-inria/gaussian-splatting/issues/379 | [] | shokomon | 2 |
huggingface/datasets | pytorch | 7,399 | Synchronize parameters for various datasets | ### Describe the bug
[IterableDatasetDict](https://huggingface.co/docs/datasets/v3.2.0/en/package_reference/main_classes#datasets.IterableDatasetDict.map) map function is missing the `desc` parameter. You can see the equivalent map function for [Dataset here](https://huggingface.co/docs/datasets/v3.2.0/en/package_refe... | open | 2025-02-14T09:15:11Z | 2025-02-19T11:50:29Z | https://github.com/huggingface/datasets/issues/7399 | [] | grofte | 2 |
benbusby/whoogle-search | flask | 277 | [BUG] Captcha request on every page | **Describe the bug**
This page is now showing every time.
**Deployment Method**
- [X] Heroku (one-click deploy) --> Europe server
- [ ] Docker
- [ ] `run` executable
- [ ] pip/pipx
- [ ] Other: [des... | closed | 2021-04-10T08:10:12Z | 2021-04-11T03:53:18Z | https://github.com/benbusby/whoogle-search/issues/277 | [
"bug"
] | federicotorrielli | 5 |
yeongpin/cursor-free-vip | automation | 100 | i cannot use cursor for this reason,it seems like there are somthing wrong about the temporary mailbox ,,,,, |  | closed | 2025-02-25T08:19:48Z | 2025-03-06T04:23:34Z | https://github.com/yeongpin/cursor-free-vip/issues/100 | [
"bug"
] | kitaharam | 12 |
ScrapeGraphAI/Scrapegraph-ai | machine-learning | 795 | Error: cannot access local variable 'browser' where it is not associated with a value | **Describe the bug**
Whereas locally my code works on the bare machine and in a docker container, in Kubernetes, I get some very weird errors related to " cannot access local variable 'browser' where it is not associated with a value"
**To Reproduce**
Steps to reproduce the behavior:
- I guess just run a simple s... | open | 2024-11-11T16:05:21Z | 2025-01-08T03:36:25Z | https://github.com/ScrapeGraphAI/Scrapegraph-ai/issues/795 | [] | aleenprd | 6 |
dask/dask | pandas | 11,260 | Dask-expr - Extremely slow with using .compute | I installed the basic dask version using "pip install dask". When running, I receive a FutureWarning:
> Dask dataframe query planning is disabled because dask-expr is not installed. You can install it with 'pip install dask[dataframe]' or 'conda install dask'. This will raise in a future version.
I proceeded to in... | open | 2024-07-29T18:46:35Z | 2024-07-31T09:16:55Z | https://github.com/dask/dask/issues/11260 | [
"dataframe",
"dask-expr"
] | NCSUFeNiX | 3 |
scrapy/scrapy | python | 5,981 | Document the wrong reactor problem | We need a doc section about the "wrong reactor is installed" problem and how to debug and fix/work around it. | closed | 2023-07-20T09:19:18Z | 2023-08-04T11:26:52Z | https://github.com/scrapy/scrapy/issues/5981 | [
"enhancement",
"docs",
"asyncio"
] | wRAR | 0 |
taverntesting/tavern | pytest | 214 | Parametrize absence of parameter | Hello,
We want to test the absence of a parameter in combination with the rest of parameters, so if we have:
```
marks:
- parametrize:
key: p1
vals:
- "a"
- "b"
- parametrize:
key: optional_parameter
vals:
- "1"
-
```
The tests performed would ... | closed | 2018-11-28T16:36:27Z | 2018-12-09T13:54:55Z | https://github.com/taverntesting/tavern/issues/214 | [] | elurisoto | 1 |
huggingface/peft | pytorch | 2,063 | question about training time | ### System Info
Dear authors,
I have a question regarding the training time utilizing the peft package. I tried using LoRA with a swin transformer to reduce the parameter size.
```
model = SwinModel.from_pretrained('./swin-large-patch4-window7-224-in22k').cuda()
config = LoraConfig(
r=16,
lora_alpha=... | closed | 2024-09-12T07:02:48Z | 2024-10-25T15:03:38Z | https://github.com/huggingface/peft/issues/2063 | [] | harborsarah | 5 |
google/seq2seq | tensorflow | 146 | Nr. of steps vs. nr. of epochs | I was wondering is there is a way to define the nr. of epochs you want a certain model to run. After reading the tutorial, I am only aware of a way to easily change the nr. of training steps (which isn't the same as training a nr. of epochs, unless I am mistaken).
Thanks in advance | closed | 2017-04-05T15:39:15Z | 2017-04-10T14:44:24Z | https://github.com/google/seq2seq/issues/146 | [] | ghost | 3 |
piskvorky/gensim | data-science | 3,039 | Documentation Notebooks | Hello, I was going through some documentation notebooks, and noticed that many of them ([Poincare Embeddings](https://github.com/RaRe-Technologies/gensim/blob/develop/docs/notebooks/Poincare%20Tutorial.ipynb), [WikiNews](https://github.com/RaRe-Technologies/gensim/blob/develop/docs/notebooks/wikinews-bigram-en.ipynb), ... | open | 2021-02-02T01:27:23Z | 2021-02-03T08:52:25Z | https://github.com/piskvorky/gensim/issues/3039 | [
"bug",
"documentation",
"difficulty easy",
"impact MEDIUM",
"reach LOW"
] | bhargavvader | 5 |
ranaroussi/yfinance | pandas | 1,986 | incorrect 52 week high and low (compare with Yahoo) | ### Describe bug
incorrect 52 week high and low values (compare with Yahoo). It is the same as regular market day high and low values.
### Simple code that reproduces your problem
```
import yfinance as yf
msft = yf.Ticker("PTT.BK")
hist = msft.history(period="1mo")
msft.history_metadata
```
### Debug log
N/A... | closed | 2024-07-16T09:51:23Z | 2024-07-19T03:43:54Z | https://github.com/ranaroussi/yfinance/issues/1986 | [] | leaderdevil | 4 |
google-research/bert | tensorflow | 960 | convert tf1 pretrained bert checkpoint to tf2 | I've trained a custom bert on my own data on tf1. now that i updated to tf2, i'm facing the issue on converting the checkpoint i have to something that's compatible with tf2. I couldn't find any converter that's working (or at least that generates a checkpoint not an h5 file).
I tried converting the checkpoint to py... | closed | 2019-12-11T15:22:40Z | 2024-10-16T18:49:03Z | https://github.com/google-research/bert/issues/960 | [] | fadybaly | 0 |
django-import-export/django-import-export | django | 1,760 | Failing test in different python environments | **Describe the bug**
In some environments one test fails:
```
(.venv) t14 ➜ django-import-export git:(feat/improve-docker-tests) python -V
Python 3.11.6
(.venv) t14 ➜ django-import-export git:(feat/improve-docker-tests) ./tests/manage.py test core --settings=settings -k test_import_data_error_saving_model
... | closed | 2024-02-28T07:13:24Z | 2024-03-13T10:15:41Z | https://github.com/django-import-export/django-import-export/issues/1760 | [
"bug"
] | bmihelac | 0 |
matplotlib/matplotlib | data-visualization | 29,778 | [Bug]: interpolation_stage="data" removes too many pixels in the vicinity of nans in upsampled, interpolated images | ### Bug summary
Currently, when upsampling images with interpolation_stage="data", upsampled pixels are set to nan if*any* of the underlying data points is nan. This leads to much wider "nan-propagation" than interpolation_stage="rgba".
### Code for reproduction
```Python
from pylab import *
a = tril(arange(1., 26... | open | 2025-03-19T00:06:01Z | 2025-03-21T02:34:02Z | https://github.com/matplotlib/matplotlib/issues/29778 | [
"topic: images"
] | anntzer | 1 |
ultralytics/yolov5 | deep-learning | 13,419 | How to generate the proper yolo style yaml? | ### Search before asking
- [X] I have searched the YOLOv5 [issues](https://github.com/ultralytics/yolov5/issues) and [discussions](https://github.com/ultralytics/yolov5/discussions) and found no similar questions.
### Question
Recently I'm working on something with yolo, and I have developed my own model and train ... | open | 2024-11-18T14:15:23Z | 2024-11-18T21:04:29Z | https://github.com/ultralytics/yolov5/issues/13419 | [
"question",
"detect"
] | tobymuller233 | 2 |
xzkostyan/clickhouse-sqlalchemy | sqlalchemy | 90 | Table engine does not reflected | I'm trying to reflect the existing table using SQL Expression Language:
```python
from sqlalchemy import create_engine, MetaData, Table
engine = create_engine('clickhouse://default:@localhost:8123/MyDatabase')
metadata = MetaData()
MyTable = Table('MyTable', metadata, autoload=True, autoload_with=engine)
```
B... | closed | 2020-03-12T10:42:43Z | 2022-05-28T11:45:27Z | https://github.com/xzkostyan/clickhouse-sqlalchemy/issues/90 | [] | IlyaBel | 2 |
s3rius/FastAPI-template | graphql | 25 | Fix kubernetes configs. | Currently we have several problems in kubernetes configs.
- [x] migrator job doesn't have limits on cpu and ram;
- [x] redis env value converted to boolean after formatting;
- [x] wrong indent in yamls;
- [x] invalid format for CMD in Dockerfile.
If you find more, please fell free to add comments. | closed | 2021-10-01T00:22:48Z | 2021-10-02T13:06:05Z | https://github.com/s3rius/FastAPI-template/issues/25 | [] | s3rius | 4 |
noirbizarre/flask-restplus | api | 160 | Proper OAuth 1/2 support | Flask-RESTPlus needs to provide a proper OAuth1/2 support:
- [ ] Oauth security definition support
- [ ] Swagger-UI OAuth configuration
- [ ] Automatic parameters extraction from oauthlib/flask-oauthlib (definition + scopes)
- [ ] Postman export
| open | 2016-04-21T11:22:02Z | 2018-09-27T11:40:33Z | https://github.com/noirbizarre/flask-restplus/issues/160 | [
"enhancement"
] | noirbizarre | 2 |
ExpDev07/coronavirus-tracker-api | rest-api | 167 | Added simple Telegram BOT | I'm using this tracker as a data source for a Telegram BOT: @CovidWORLDbot
Thanks. | closed | 2020-03-24T16:42:41Z | 2020-04-19T18:09:41Z | https://github.com/ExpDev07/coronavirus-tracker-api/issues/167 | [
"user-created"
] | tonjo | 1 |
Kav-K/GPTDiscord | asyncio | 446 | [BUG] Bot not responding to mentions in reply | **Describe the bug**
Bot does not detect/respond to @ mentions in replies to itself
**To Reproduce**
Steps to reproduce the behaviour:
1. Set BOT_TAGGABLE=True in .env
2. Send message @ bot
3. When bot replies, click reply to message in discord.
4. See error
**Expected behaviour**
When the bot is mentione... | closed | 2023-12-11T07:07:05Z | 2023-12-31T10:08:29Z | https://github.com/Kav-K/GPTDiscord/issues/446 | [
"bug"
] | jeffe | 3 |
pyppeteer/pyppeteer | automation | 459 | newPage() causes infinite await loop | I'm running the following code:
```
import asyncio
from pyppeteer import launch
# Get page
url = 'https://quotes.toscrape.com/'
async def main():
browser = await launch(headless=False)
page = await browser.newPage()
await page.goto(url)
await page.screenshot({"path": "example.png"})
... | open | 2024-01-04T10:47:55Z | 2024-02-10T13:43:12Z | https://github.com/pyppeteer/pyppeteer/issues/459 | [] | Ollenmire | 5 |
piskvorky/gensim | machine-learning | 2,942 | Segfault when training doc2vec | #### Problem description
When attempting to train doc2vec, gensim segfaults.
#### Steps/code/corpus to reproduce
I run the code:
```
import faulthandler
import gensim
faulthandler.enable()
model = gensim.models.doc2vec.Doc2Vec(corpus_file = "yelp_tripadvisor_linesentence.txt", vector_size=250, min_count=1... | closed | 2020-09-10T22:59:30Z | 2020-09-15T15:06:46Z | https://github.com/piskvorky/gensim/issues/2942 | [] | Paul-E | 5 |
coqui-ai/TTS | deep-learning | 4,017 | VITS model gives bad results (training an italian tts model) | ### Describe the bug
Hi everyone. I'm new to the world of ML, so I'm not used to training AI models...
I really want to create my own TTS model using coqui's VITS trainer, so I've done a lot of research about it. I configured some dataset parameters and configuration functions and then started training. For the tra... | closed | 2024-10-09T20:29:06Z | 2024-12-28T11:58:24Z | https://github.com/coqui-ai/TTS/issues/4017 | [
"bug",
"wontfix"
] | iDavide | 6 |
GibbsConsulting/django-plotly-dash | plotly | 105 | Using Bootstraps grid within a dash app | I'm attempting to place a formatted dash application into my django site. For the rest of my web application I'm using bootstrap 4.1.3 via CDN.
`<link rel="stylesheet" href="https://stackpath.bootstrapcdn.com/bootstrap/4.1.3/css/bootstrap.min.css" integrity="sha384-MCw98/SFnGE8fJT3GXwEOngsV7Zt27NXFoaoApmYm81iuXoPkFOJ... | closed | 2019-01-11T01:14:18Z | 2019-07-29T18:12:45Z | https://github.com/GibbsConsulting/django-plotly-dash/issues/105 | [] | vantaka2 | 7 |
d2l-ai/d2l-en | data-science | 1,738 | Typo in 5.2.2.1. Built-in Initialization | The TensorFlow tab for the final block in this section of 5.2.2.1 has a typo in the initialization.
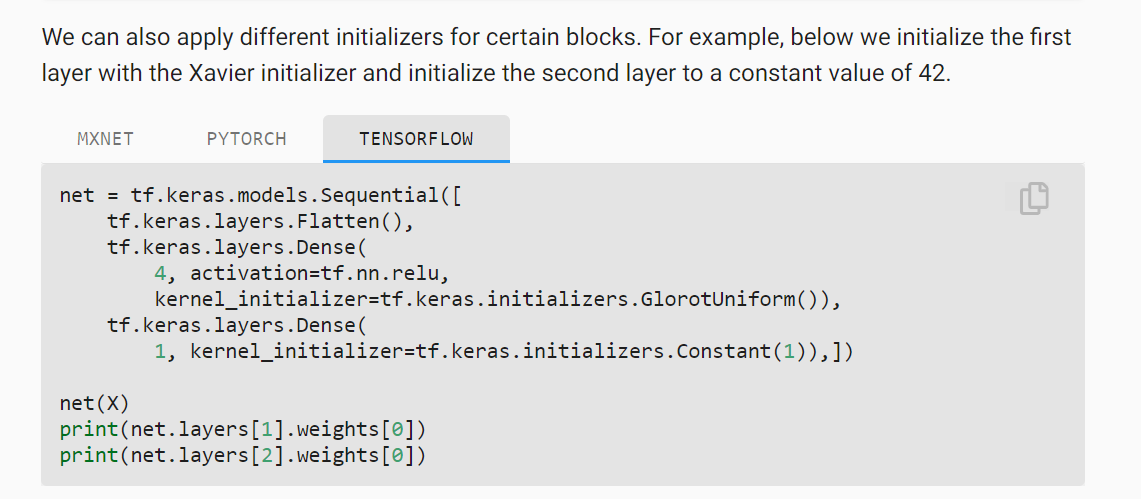

The text says the constant initialization is 42 while the code uses a constant of 1. | closed | 2021-04-27T00:10:46Z | 2021-04-27T01:46:49Z | https://github.com/d2l-ai/d2l-en/issues/1738 | [] | bgreenawald | 0 |
zalandoresearch/fashion-mnist | computer-vision | 50 | benchmark: update on GRU+SVM with Dropout | Hey @hanxiao , it's me again. I saw an update in the dataset, regarding duplicate samples. I did another training using my GRU+SVM (with Dropout) model (from #8 ) on the updated dataset. Here's the result:
```
Epoch : 0 completed out of 100, loss : 316.9036560058594, accuracy : 0.734375
Epoch : 1 completed out of ... | closed | 2017-09-01T14:57:58Z | 2017-09-01T15:24:35Z | https://github.com/zalandoresearch/fashion-mnist/issues/50 | [] | AFAgarap | 2 |
healthchecks/healthchecks | django | 772 | API providing project information | Hi,
At the moment there is no way, using the api, to get information about the project, if only the `API key` is known.
It would be great to have access to basic project information, like:
- Project Name
- Team Access
Best
m42e | closed | 2023-01-07T07:38:05Z | 2023-08-04T07:05:05Z | https://github.com/healthchecks/healthchecks/issues/772 | [
"feature"
] | m42e | 2 |
scikit-learn/scikit-learn | machine-learning | 30,397 | Unknown TypeError after updating to 1.5.2 | ### Describe the bug
I am not sure if this is an update bug or a compatibility issue for an older python version with scikit-learn.
### Steps/Code to Reproduce
```
from sklearn.model_selection import train_test_split
```
### Expected Results
no output
### Actual Results
```
-------------------------------... | closed | 2024-12-03T10:12:46Z | 2024-12-13T14:09:23Z | https://github.com/scikit-learn/scikit-learn/issues/30397 | [] | krishpy99 | 4 |
LibreTranslate/LibreTranslate | api | 746 | Chinese (zh) is not available as a target language from English (en) | Chinese (zh) is not available as a target language from English (en)
`curl -X 'POST' \
'http://localhost:5000/translate' \
-H 'accept: application/json' \
-H 'Content-Type: application/x-www-form-urlencoded' \
-d 'q=hello&source=en&target=zh&format=text&alternatives=3&api_key=xxxxxxxx-xxxx-xxxx-xxxx-xxxxxxxxxxx... | open | 2025-02-22T03:36:05Z | 2025-02-26T03:29:02Z | https://github.com/LibreTranslate/LibreTranslate/issues/746 | [
"possible bug"
] | joulong | 1 |
wkentaro/labelme | deep-learning | 1,459 | When i create ai polygon met the error | ### Provide environment information
python --version 3.8;labelme --vesion 5.5
2024-06-18 13:49:28,599 [INFO ] __init__:get_config:67- Loading config file from:
2024-06-18 13:49:36,430 [DEBUG ] canvas:initializeAiModel:139- Initializing AI model: 'EfficientSam (accuracy)'
/home/anaconda3/envs/labelme/lib/python3... | open | 2024-06-18T05:52:43Z | 2024-06-18T05:52:43Z | https://github.com/wkentaro/labelme/issues/1459 | [
"issue::bug"
] | Jll0716 | 0 |
sgl-project/sglang | pytorch | 4,417 | [Bug] MTP and cuda graph stuck at initialization on 2 h100 nodes | ### Checklist
- [x] 1. I have searched related issues but cannot get the expected help.
- [x] 2. The bug has not been fixed in the latest version.
- [x] 3. Please note that if the bug-related issue you submitted lacks corresponding environment info and a minimal reproducible demo, it will be challenging for us to repr... | closed | 2025-03-14T08:01:56Z | 2025-03-14T08:31:31Z | https://github.com/sgl-project/sglang/issues/4417 | [] | victorserbu2709 | 1 |
jonaswinkler/paperless-ng | django | 895 | [BUG] Contents are not recognized | Hello together!
Currently I'm testing paperless-ng and I've come across a strange thing, but I'm not sure if it's a bug or not. I have set up a mailbox that I retrieve via IMap. Here I have forwarded a mail with 2 attachments. They were also extracted, so far so good. But when I looked at the documents in Paperless ... | closed | 2021-04-11T14:24:59Z | 2021-04-17T20:17:58Z | https://github.com/jonaswinkler/paperless-ng/issues/895 | [] | prodigy7 | 3 |
BeanieODM/beanie | pydantic | 121 | Read and Write concerns | Any plans to implement https://pymongo.readthedocs.io/en/3.12.0/api/pymongo/write_concern.html
Seems fairly striaght forward to do though unsure how you would want to handle the API for it. Happy to mock something up. | closed | 2021-09-30T09:54:36Z | 2023-04-04T02:21:37Z | https://github.com/BeanieODM/beanie/issues/121 | [
"Stale"
] | zrothberg | 3 |
browser-use/browser-use | python | 963 | Handling pop-up windows from websites | ### Problem Description
I was testing brwoser-use and was impressed performance.
However, I noticed that browser-use may potentially get stuck on websites that has pop up windows after the website is loaded.
(For example, the cookie acknowledgement pop up window for this web page: https://www.mdpi.com/2673-947X/5/1/... | open | 2025-03-06T20:21:58Z | 2025-03-11T08:42:22Z | https://github.com/browser-use/browser-use/issues/963 | [
"enhancement"
] | HengyueL | 1 |
donnemartin/system-design-primer | python | 542 | Calculation of attorney's fees | open | 2021-05-29T13:52:01Z | 2022-04-23T13:17:40Z | https://github.com/donnemartin/system-design-primer/issues/542 | [
"needs-review"
] | yezhuoying | 1 | |
jmcnamara/XlsxWriter | pandas | 465 | worksheet.repeat_rows(last) not functioning as worksheet.repeat_rows(0, last) | Code in worksheet.py line 3142, sucked down from PIP as of a few minutes ago:
if last_row is None:
last_row = first_row
# Convert rows to 1 based.
I believe, you need to then set first_row=0 in that if-clause. Not sure how this happened; it used to work fine in previous versions. | closed | 2017-09-07T17:44:39Z | 2017-09-07T18:37:54Z | https://github.com/jmcnamara/XlsxWriter/issues/465 | [] | kjosib | 4 |
benbusby/whoogle-search | flask | 593 | [BUG] xml.etree.ElementTree.ParseError: not well-formed (invalid token) | **Describe the bug**
```
ERROR:app:Exception on /autocomplete [POST]
Traceback (most recent call last):
File "/usr/local/lib/python3.8/site-packages/flask/app.py", line 2446, in wsgi_app
response = self.full_dispatch_request()
File "/usr/local/lib/python3.8/site-packages/flask/app.py", line 1951, in full_... | closed | 2021-12-27T11:00:23Z | 2021-12-28T18:38:35Z | https://github.com/benbusby/whoogle-search/issues/593 | [
"bug"
] | bruvv | 1 |
deepset-ai/haystack | machine-learning | 8,310 | AutoMerging-Retriever: support more document stores | - Pinecone doesn't support it because apparently, one cannot filter by `id`:
- https://docs.pinecone.io/guides/data/query-data#querying-by-record-id
- https://community.pinecone.io/t/does-pinecone-support-filtering-by-vector-id/3039/2
- Weaviate needs to be properly tested
- All the others document stores work, w... | closed | 2024-08-29T13:32:10Z | 2024-09-19T07:44:02Z | https://github.com/deepset-ai/haystack/issues/8310 | [
"P2"
] | davidsbatista | 2 |
stanfordnlp/stanza | nlp | 523 | Introducing external Chinese tokenizer into the pipeline makes 'tokenize_no_ssplit=True' not working | Hi, after careful comparison with jieba tokenizer with your Chinese tokenizer, I am using jieba tokenizer for my downstream NER task, as it shows better performance on Chinese names and locations identification.
I do NER in batchces in order to speed up, so I add "\n\n" between all the segmented sentences as mentio... | closed | 2020-11-18T08:54:00Z | 2021-02-02T00:33:24Z | https://github.com/stanfordnlp/stanza/issues/523 | [
"bug",
"fixed on dev"
] | twang18 | 6 |
mirumee/ariadne | graphql | 1,234 | convert_names_case on default resolvers doesn't work well with directives | For example in the current test https://github.com/mirumee/ariadne/blob/e03d333244d1521aad4ba9a5a811d9ade117d410/tests/test_directives.py#L60
For simplicity if I change the test case to have both `convert_names_case=True` and `@upper` it will fail
```python
def test_field_definition_directive_replaces_field_resolver_w... | open | 2025-02-19T23:45:51Z | 2025-02-20T00:02:44Z | https://github.com/mirumee/ariadne/issues/1234 | [] | zwangBLP | 0 |
dpgaspar/Flask-AppBuilder | flask | 1,643 | Issue get user data in class inherits modelview body | ### Issue get user data in class inherits modelview body
- i want to access user data in class Customers(ModelView) to set show_fieldsets columns or base_permissions based on user data
- when i call get_user_type like this it return function name
- when i call get_user_type() i get error: **RuntimeError: work... | closed | 2021-05-19T11:07:28Z | 2022-04-17T16:24:31Z | https://github.com/dpgaspar/Flask-AppBuilder/issues/1643 | [
"question",
"stale"
] | Basma18 | 4 |
satwikkansal/wtfpython | python | 206 | Prehistory | Hi Satwik! If you are interested in antediluvian times, check out https://github.com/MarcinCiura/6-gotchas :)
Cheers! | closed | 2020-05-22T09:35:34Z | 2024-10-11T11:46:16Z | https://github.com/satwikkansal/wtfpython/issues/206 | [] | MarcinCiura | 2 |
flairNLP/flair | nlp | 3,033 | TypeError: RobertaModel.__init__() got an unexpected keyword argument 'force_max_length' when loading model with SequenceTagger | When loading a fine-tuned model, I am getting the above error locally and in a Docker container. I am using flair==0.11.3, torch==1.11 and transformers==4.21.3.
When loading the model within a notebook hosted on a cloud platform, the loading runs just fine. But locally (I am using an M1 MacBook) I am getting that er... | closed | 2022-12-16T12:36:54Z | 2022-12-16T14:22:58Z | https://github.com/flairNLP/flair/issues/3033 | [
"question"
] | agademic | 1 |
pytest-dev/pytest-django | pytest | 755 | pytest.ini is ignored when using manage.py test with pytest-django | When I run pytest with coverage from the command line I get right results.
But when I try to run it with manage.py test coverage results are wrong.
Model fields are ignored.
What I figured out, is that manage.py test ignores pytest.ini file. If I delete it tests won't fail.
It says "Django settings: seabattle.setti... | closed | 2019-08-16T19:23:59Z | 2020-10-16T19:36:27Z | https://github.com/pytest-dev/pytest-django/issues/755 | [] | nvishnya | 2 |
plotly/dash | dash | 2,878 | [BUG] `id` passed through `dcc.Loading` not visible in DOM | **Describe your context**
Hello guys 👋
I am currently trying to pass an `id` to the dcc.Loading component or its parent container and I would like the `id` to be visible in the DOM such that I can target the CSS of the components inside the `dcc.Loading` via ID.
Please provide us your environment, so we can ea... | closed | 2024-06-07T10:41:21Z | 2024-06-18T13:22:13Z | https://github.com/plotly/dash/issues/2878 | [
"good first issue"
] | huong-li-nguyen | 4 |
tortoise/tortoise-orm | asyncio | 1,742 | Timezone-aware datetime returns incorrect timezone for SQLite | **Describe the bug**
When I create a new object in a SQLite database, the timezone being put into the database, and the timezone being retrieved from the database, are different from the value I originally inputted.
**To Reproduce**
```py
import datetime
from tortoise import Tortoise, Model, fields, run_async
... | closed | 2024-10-17T11:07:36Z | 2024-10-19T23:44:54Z | https://github.com/tortoise/tortoise-orm/issues/1742 | [] | seriaati | 3 |
gee-community/geemap | jupyter | 1,358 | GEEMAP installation not possible on Mac M1 Max using Anaconda | Hi Qiusheng. I'm trying to install geemap, but I am unable to install the most recent version of geemap because Anaconda is not supporting python versions less than 3.8 on Apple M1 chips. From what I understand, geemap was built on python 3.7, which is why I believe I am having issues installing the latest version of... | closed | 2022-12-02T23:29:02Z | 2022-12-05T19:46:09Z | https://github.com/gee-community/geemap/issues/1358 | [] | melrohde | 7 |
microsoft/unilm | nlp | 1,622 | How can I have dit document layout analysis checkpoints? | **Describe**
Model I am using is Dit object detection, when I want to run inference, I found the checkpoint is unavailable.
It is possible for us to have it? Is there any suggestion for me to the the object detection model fine tune?
Thank you in advance.
| open | 2024-09-10T20:36:25Z | 2024-09-10T20:36:25Z | https://github.com/microsoft/unilm/issues/1622 | [] | WYY220062 | 0 |
pbugnion/gmaps | jupyter | 80 | How do you just set the origin/center longitude and latitude to a particular area? | The documentation doesn't show a basic example for this?
| closed | 2016-08-27T14:42:08Z | 2016-09-04T06:35:37Z | https://github.com/pbugnion/gmaps/issues/80 | [] | CMCDragonkai | 4 |
voila-dashboards/voila | jupyter | 1,441 | Extension registration requires a kernel? | <!--To help us understand and resolve your issue, please fill out the form to the best of your ability.-->
<!--You can feel free to delete the sections that do not apply.-->
### Problem
We're migrating from voila 0.4 to 0.5. We have an extension we've built for jupyter that we had been using in voila 0.4. Howe... | closed | 2024-02-05T15:30:30Z | 2024-02-05T21:55:38Z | https://github.com/voila-dashboards/voila/issues/1441 | [
"documentation"
] | ClaytonAstrom | 9 |
abhiTronix/vidgear | dash | 303 | [Bug]:NetGear is not OPENCV 4.5.5 Compatible | closed | 2022-05-06T18:04:16Z | 2022-05-07T09:18:03Z | https://github.com/abhiTronix/vidgear/issues/303 | [
"INVALID :stop_sign:"
] | rubar-tech | 5 | |
ydataai/ydata-profiling | jupyter | 1,303 | Feature Request - Add support for Pandas 2 | ### Missing functionality
Add support for Pandas 2
### Proposed feature
I'd just like to be able to install ydata-profiling and Pandas 2 into the same environment
### Alternatives considered
_No response_
### Additional context
_No response_ | open | 2023-04-05T01:03:14Z | 2023-06-12T17:29:44Z | https://github.com/ydataai/ydata-profiling/issues/1303 | [
"feature request 💬"
] | owenlamont | 5 |
horovod/horovod | deep-learning | 3,921 | the repuirements installing Horovod in Conda in CUDA12 | **Environment:**
1. Framework: (TensorFlow, Keras, PyTorch, MXNet)
2. Framework version:PyTorch
3. Horovod version:
4. MPI version:4.1.5
5. CUDA version:12
6. NCCL version:
7. Python version:
8. Spark / PySpark version:
9. Ray version:
10. OS and version:
11. GCC version:9+
12. CMake version:
**Checklist... | open | 2023-05-11T06:13:11Z | 2023-05-11T06:13:11Z | https://github.com/horovod/horovod/issues/3921 | [
"bug"
] | CoconutSweet999 | 0 |
ShishirPatil/gorilla | api | 774 | [BFCL] Error when consecutively generate from multiple oss_models with vllm | I have plenty of models and I want to test them all. So I run a bash script like this:
```
bfcl generate --model "model_1" --test-category simple
bfcl generate --model "model_2" --test-category simple
...
```
The first generation was good. But the second run and thereafter encountered the following error:
![imag... | closed | 2024-11-21T09:28:01Z | 2024-11-27T08:05:49Z | https://github.com/ShishirPatil/gorilla/issues/774 | [
"BFCL-General"
] | YifanHao | 2 |
ultralytics/ultralytics | machine-learning | 19,111 | YOLO with dinov2 as backbone | ### Search before asking
- [x] I have searched the Ultralytics YOLO [issues](https://github.com/ultralytics/ultralytics/issues) and [discussions](https://github.com/orgs/ultralytics/discussions) and found no similar questions.
### Question
Hello @Y-T-G ,
I saw your code to support different backbones from torchvis... | open | 2025-02-06T23:57:42Z | 2025-02-14T13:34:43Z | https://github.com/ultralytics/ultralytics/issues/19111 | [
"enhancement",
"question"
] | SebastianJanampa | 6 |
amdegroot/ssd.pytorch | computer-vision | 202 | Add pretrained weights for COCO dataset [feature request] | It would be great if there are weights for a the model trained on the COCO dataset. | open | 2018-07-11T09:37:56Z | 2018-07-13T09:45:58Z | https://github.com/amdegroot/ssd.pytorch/issues/202 | [] | sotte | 1 |
keras-team/keras | deep-learning | 20,106 | Unrecognized keyword arguments passed to LSTM: {'batch_input_shape' | model = Sequential()
model.add(LSTM(4, batch_input_shape=(1, X_train.shape[1], X_train.shape[2]), stateful=True))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
model.fit(X_train, y_train, epochs=100, batch_size=1, verbose=1, shuffle=False)
ValueError: Unrecognized keyword argumen... | closed | 2024-08-09T19:19:55Z | 2025-03-22T12:16:24Z | https://github.com/keras-team/keras/issues/20106 | [
"type:support",
"stat:awaiting response from contributor"
] | Ineedsomehelpah | 7 |
microsoft/unilm | nlp | 1,342 | [Kosmos-G] Docker failed to run demo | I try to run docker demo for test kosmos-g, but it's still has failed, error with torchscale,
Can anyone give docker fix?
tks | open | 2023-10-26T01:55:55Z | 2023-10-26T01:55:55Z | https://github.com/microsoft/unilm/issues/1342 | [] | trangtv57 | 0 |
deepspeedai/DeepSpeed | deep-learning | 6,737 | [BUG] CUDA out of memory error when using a customized model at deepspeed.initialize(). | **Describe the bug**
In my own implementation, I combine a large language model and a speculator model. And my goal is to train the speculator model to make it better at predicting the n+2, n+3... tokens. I have read the doc of deepspeed, and I think it supports any customized model on top of nn.Module. But I have enco... | closed | 2024-11-11T14:50:56Z | 2024-11-15T01:36:13Z | https://github.com/deepspeedai/DeepSpeed/issues/6737 | [
"bug",
"training"
] | 962086838 | 4 |
python-gino/gino | asyncio | 559 | Inserting millions of rows. Much much slower than SQLite. Am I doing it wrong? How can I improve throughput? | * GINO version: 0.8.3
* Python version: 3.7.3
* asyncpg version: 0.18.3
* aiocontextvars version: 0.2.2
* PostgreSQL version: 9.6
### Description
I am building something similar/based on: https://github.com/p3pperp0tts/leaks_parser.
Parsing GB of text files, and inputting to database. The difference between... | closed | 2019-10-07T00:29:48Z | 2019-10-12T11:47:59Z | https://github.com/python-gino/gino/issues/559 | [
"question"
] | brizzbane | 8 |
identixone/fastapi_contrib | pydantic | 170 | Why Serializer do not output "_id" field? | My model need output _id field to frontend of my project.
```python
def dict(self, *args, **kwargs) -> dict:
"""
Removes excluded fields based on `Meta` and `kwargs`
:return: dict of serializer data fields
"""
exclude = kwargs.get("exclude")
if not exclude:
... | open | 2021-05-10T06:43:33Z | 2021-06-03T14:06:33Z | https://github.com/identixone/fastapi_contrib/issues/170 | [] | ChandlerBent | 1 |
ludwig-ai/ludwig | computer-vision | 3,960 | Dependency issue | **Describe the bug**
When importing ludwig.backend and initializing the ray cluster I am getting the following error:
/Users/robertheise/Documents/SD/accelator/venv/lib/python3.11/site-packages/bitsandbytes/cextension.py:34: UserWarning: The installed version of bitsandbytes was compiled without GPU support. 8-bit o... | closed | 2024-03-08T21:22:40Z | 2024-03-11T18:31:29Z | https://github.com/ludwig-ai/ludwig/issues/3960 | [] | robhheise | 8 |
BeanieODM/beanie | pydantic | 574 | Search operators without fields | ### Discussed in https://github.com/roman-right/beanie/discussions/570
<div type='discussions-op-text'>
<sup>Originally posted by **akriese** May 23, 2023</sup>
Hi, I want to be able to use query operators like Eq, GT etc. without fields. One simple use case is, that I want to use ElemMatch on a list of numbers.... | open | 2023-05-25T19:24:05Z | 2024-12-08T21:53:54Z | https://github.com/BeanieODM/beanie/issues/574 | [
"feature request"
] | roman-right | 0 |
PablocFonseca/streamlit-aggrid | streamlit | 259 | Row selection events not reported | Row selection events are not reported. I took the examples that one finds in the web. They do not report what they are supposed to do. | closed | 2024-03-30T18:29:46Z | 2024-04-23T01:17:21Z | https://github.com/PablocFonseca/streamlit-aggrid/issues/259 | [] | mmf431 | 14 |
ets-labs/python-dependency-injector | flask | 240 | Add Python 3.8 support | Python 3.8.0 is available since Oct of 2019 and it's needed to start supporting it.
Links:
- https://www.python.org/downloads/release/python-380/ | closed | 2020-01-24T02:09:13Z | 2020-01-29T18:33:53Z | https://github.com/ets-labs/python-dependency-injector/issues/240 | [
"enhancement"
] | rmk135 | 0 |
keras-team/keras | data-science | 21,076 | calculate score calculation within callback | ```python
import keras
def get_model():
model = keras.Sequential()
model.add(keras.layers.Dense(1))
model.compile(
optimizer=keras.optimizers.RMSprop(learning_rate=0.1),
loss="mean_squared_error",
metrics=["mean_absolute_error"],
)
return model
(x_train, y_train), (x_test, ... | open | 2025-03-20T20:18:57Z | 2025-03-21T11:10:29Z | https://github.com/keras-team/keras/issues/21076 | [
"keras-team-review-pending"
] | pure-rgb | 2 |
hankcs/HanLP | nlp | 1,009 | pyhanlp 在python3环境下data-for-1.6.8.zip 解压缩乱码 | <!--
注意事项和版本号必填,否则不回复。若希望尽快得到回复,请按模板认真填写,谢谢合作。
-->
## 注意事项
请确认下列注意事项:
* 我已仔细阅读下列文档,都没有找到答案:
- [首页文档](https://github.com/hankcs/HanLP)
- [wiki](https://github.com/hankcs/HanLP/wiki)
- [常见问题](https://github.com/hankcs/HanLP/wiki/FAQ)
* 我已经通过[Google](https://www.google.com/#newwindow=1&q=HanLP)和[issue区检... | closed | 2018-11-02T13:31:40Z | 2020-01-01T10:56:11Z | https://github.com/hankcs/HanLP/issues/1009 | [
"ignored"
] | passerbythesun | 2 |
arogozhnikov/einops | tensorflow | 252 | Support JAX's distributed arrays | **Describe the bug**
Using `...` in `einops.rearrange` introduces extraneous reshape operations, where multiple dimensions are flattened into 1D and then reshaped back.
This is typically fine, but can be problematic in (at least) two contexts:
1. When using JAX's [distributed arrays](https://jax.readthedocs.io... | closed | 2023-04-21T01:21:52Z | 2023-10-02T03:50:52Z | https://github.com/arogozhnikov/einops/issues/252 | [
"enhancement"
] | shoyer | 4 |
collerek/ormar | pydantic | 632 | poetry add fails | When I run `poetry add ormar` I see this error:
```
$ poetry add ormar
Using version ^0.11.0 for ormar
Updating dependencies
Resolving dependencies... (0.0s)
SolverProblemError
Because no versions of ormar match >0.11.0,<0.12.0
and ormar (0.11.0) depends on SQLAlchemy (>=1.3.18,<=1.4.31), ormar (... | closed | 2022-04-30T00:04:42Z | 2022-05-04T07:59:20Z | https://github.com/collerek/ormar/issues/632 | [
"bug"
] | mturilin | 1 |
brightmart/text_classification | tensorflow | 113 | tensorflow.python.framework.errors_impl.NotFoundError: Key is_training not found in checkpoint | Hi Mr.Brightmart
@brightmart when I try to run fast text, I got an error like this.
I try to run a01_FastText,
steps:
1. python p6_fastTextB_train_multilabel.py
everything works fine, then I got fast_text_checkpoint_multi folder with file tree like following: fast_text_checkpoint_multi
├── checkpoint
... | open | 2019-03-21T04:37:53Z | 2019-03-21T04:37:53Z | https://github.com/brightmart/text_classification/issues/113 | [] | luhawk803 | 0 |
python-gino/gino | asyncio | 238 | Can't load plugin: sqlalchemy.dialects:postgresql.asyncpg | * GINO version: 0.7.3
* Python version: Python 3.6.3
* asyncpg version: 0.15.0
* aiocontextvars version: 0.1.2
* PostgreSQL version: postgresql-9.2.23-3.el7_4.x86_64
I have an app using gino that works well on two Fedora machines running Python 3.6.5
However, on a CentOS 7 machine running Python 3.6.3, gino ref... | closed | 2018-06-02T17:52:20Z | 2025-03-12T15:09:02Z | https://github.com/python-gino/gino/issues/238 | [
"invalid"
] | elad661 | 3 |
benbusby/whoogle-search | flask | 186 | [FEATURE] Config file for preset Configuration | **Describe the feature you'd like to see added**
Well, it would be wonderful if there's a config file in the repo where Whoogle default Configuration is defined before and can be set before deploying.
**Additional context**
Like, variables, Country, City, IsDarkModeOn etc. I know there is a Load function for loadi... | closed | 2021-01-29T20:37:08Z | 2021-03-28T18:37:23Z | https://github.com/benbusby/whoogle-search/issues/186 | [
"enhancement"
] | mizzunet | 4 |
plotly/dash-table | dash | 579 | [FEATURE] Sorting/Filtering of selected rows | This is basically asking for https://github.com/plotly/dash-table-experiments/issues/60
I face a similar issue - a large data table where the selected rows are not necessarily visible in the current page. This means the users need to jump through significant hoops to locate the selected rows. | open | 2019-09-10T07:47:00Z | 2021-03-02T14:09:50Z | https://github.com/plotly/dash-table/issues/579 | [
"dash-type-enhancement"
] | orenbenkiki | 3 |
NVIDIA/pix2pixHD | computer-vision | 110 | Cuda run out of memory. Pytorch 1.0 - Cuda 10.0 | I am testing pix2pixHD. It works on my local machine, but it raise an error in a cloud server machine. The strange think is that the server machine is more powerful.
Here the details:
**LOCAL MACHINE**
Ubuntu 16.04
GPU: Geforce GTX 1050 - 4GB GPU Memory
Pytorch version: 0.4.0
Cuda 9.0
**SERVER MACHINE**
U... | open | 2019-03-22T14:52:35Z | 2019-05-06T15:43:46Z | https://github.com/NVIDIA/pix2pixHD/issues/110 | [] | ghost | 3 |
oegedijk/explainerdashboard | dash | 162 | TypeError: can only concatenate str (not "int") to str | from sklearn.linear_model import LogisticRegression
model = LogisticRegression()
model.fit(X_train_norm, y_train)
explainer = ClassifierExplainer(model, X_test_norm, y_test,
shap='linear',
X_background=X_train,
... | closed | 2021-12-06T12:34:17Z | 2021-12-23T19:15:57Z | https://github.com/oegedijk/explainerdashboard/issues/162 | [] | andrecasotti | 1 |
onnx/onnx | scikit-learn | 6,267 | [1.16.2/1.17?] ONNX build Windows | # Bug Report
### Is the issue related to model conversion?
No. I can't even perform imports.
### Describe the bug
My projects are permissive with respect to which `onnx` PyPI package version is installed. `onnx 1.16.2` came out this morning and broke my projects.
For example, in a turnkeyml environment that... | open | 2024-08-01T14:26:01Z | 2025-03-14T07:51:06Z | https://github.com/onnx/onnx/issues/6267 | [
"bug",
"announcement"
] | jeremyfowers | 45 |
joouha/euporie | jupyter | 31 | [Feature request] Toggle top menu | Thanks for the great project!
Is there an option to toggle whether the top menu (`File`, `Edit`, etc.) is displayed, and can this be activated by a keyboard shortcut? Having the ability to remove the top menu at times would aid a minimalist setup greatly. | closed | 2022-08-31T03:59:46Z | 2022-08-31T17:03:26Z | https://github.com/joouha/euporie/issues/31 | [] | jjeffrey | 2 |
plotly/dash | jupyter | 2,423 | Add loading attribute to html.Img component | **Is your feature request related to a problem? Please describe.**
I'm trying to lazy load images using the in built browser functionality, but I can't because that's not exposed in the html.Img component.
**Describe the solution you'd like**
I'd like the loading attribute to be added to the html.Img built in comp... | open | 2023-02-13T12:15:58Z | 2024-08-13T19:26:45Z | https://github.com/plotly/dash/issues/2423 | [
"feature",
"P3"
] | LiamLombard | 1 |
chatanywhere/GPT_API_free | api | 73 | 为什么IDEA中搜索不到ChatGPT呢 | closed | 2023-08-02T22:48:44Z | 2023-08-11T02:24:39Z | https://github.com/chatanywhere/GPT_API_free/issues/73 | [] | ghost | 1 | |
microsoft/nni | tensorflow | 5,656 | InputChoice raises issues with TPE search strategy in NAS | **Describe the issue**:
In version 3.0rc1 TPE does not seem to be compatible with the InputChoice primitive. In addition, I found TPE tuner is default to minimize (I'm assuming because hpo tuners all minimize), but this isn't coherent with https://github.com/microsoft/nni/issues/5626#issuecomment-1615350440 , perhaps ... | open | 2023-08-03T17:17:37Z | 2023-08-17T04:52:55Z | https://github.com/microsoft/nni/issues/5656 | [] | sw33zy | 1 |
piskvorky/gensim | data-science | 3,377 | Install gensim fails because C code is not ANSI-compliant | #### Problem description
Can't use `pip install gensim` to install the latest `gensim`
#### Steps/code/corpus to reproduce
```
building 'gensim.similarities.fastss' extension
creating build/temp.linux-x86_64-3.6/gensim/similarities
gcc -pthread -Wno-unused-result -Wsign-compare -DNDEBUG -O2 -g -pipe... | closed | 2022-08-10T09:49:06Z | 2022-08-22T12:51:26Z | https://github.com/piskvorky/gensim/issues/3377 | [] | hstk30 | 1 |
assafelovic/gpt-researcher | automation | 609 | No module named 'gpt_researcher.retrievers.custom' | I am trying to run multi-agent researcher but I am getting following error:
> Traceback (most recent call last):
File "C:\Users\Tomas\Documents\Python Workspace\gpt-researcher\multi_agents\main.py", line 32, in <module>
asyncio.run(main())
File "C:\Users\Tomas\anaconda3\envs\gpt\Lib\asyncio\runners.py", l... | closed | 2024-06-18T17:39:09Z | 2024-06-19T06:13:34Z | https://github.com/assafelovic/gpt-researcher/issues/609 | [] | JustUser1410 | 3 |
jupyterlab/jupyter-ai | jupyter | 1,138 | Dev install on CI times out | ## Description
See the 42-minute workflow run in #1129: https://github.com/jupyterlab/jupyter-ai/actions/runs/12167591800/job/33939176577?pr=1129
Relevant logs below:
```
@jupyter-ai/core: INFO: This is taking longer than usual. You might need to provide the dependency resolver with stricter constraints to re... | closed | 2024-12-04T21:59:16Z | 2024-12-05T14:59:45Z | https://github.com/jupyterlab/jupyter-ai/issues/1138 | [
"bug"
] | dlqqq | 7 |
WZMIAOMIAO/deep-learning-for-image-processing | deep-learning | 737 | MAP和AP50 | 如果我的数据集只有一个类别,这时候输出的指标里MAP和AP50应该差不多吧?为什么MAP才0.3,AP50倒是有0.7。怎么修改相应指标呢,如果我想输出其他的指标,例如准确率,召回率或者自定义的一些指标 | open | 2023-05-20T06:12:37Z | 2023-05-20T06:12:37Z | https://github.com/WZMIAOMIAO/deep-learning-for-image-processing/issues/737 | [] | thestars-maker | 0 |
cvat-ai/cvat | tensorflow | 8,723 | Cant communicate cvat Api from my dockerized application | ### Actions before raising this issue
- [X] I searched the existing issues and did not find anything similar.
- [X] I read/searched [the docs](https://docs.cvat.ai/docs/)
### Steps to Reproduce
1) Docker compose up - CVAT application
2) Docker compose up - my application
3) Call CVAT REST API from my application
... | closed | 2024-11-20T07:46:52Z | 2024-11-20T07:55:33Z | https://github.com/cvat-ai/cvat/issues/8723 | [
"bug",
"invalid"
] | Nishanth-KR | 1 |
huggingface/datasets | deep-learning | 6,695 | Support JSON file with an array of strings | Support loading a dataset from a JSON file with an array of strings.
See: https://huggingface.co/datasets/CausalLM/Refined-Anime-Text/discussions/1 | closed | 2024-02-26T12:35:11Z | 2024-03-08T14:16:25Z | https://github.com/huggingface/datasets/issues/6695 | [
"enhancement"
] | albertvillanova | 1 |
pydantic/pydantic-settings | pydantic | 300 | Validation error for 3 levels of nested dicts in v2.3.0 | Hello,
My model does not work anymore with the latest version of pydantic-settings.
Here is a test that reproduces my issue (`env` is the fixture from the pydantic-settings tests):
```python
def test_nested_dicts(env):
class Settings(BaseSettings):
nested: Dict[str, Dict[str, Dict[str, str]]]
... | closed | 2024-06-05T09:48:33Z | 2024-06-05T15:16:50Z | https://github.com/pydantic/pydantic-settings/issues/300 | [
"bug"
] | bpicardat | 7 |
alteryx/featuretools | scikit-learn | 2,284 | Add primitive for 2 digit Postal Code Prefix (US-only) | - As a user of Featuretools, I would like to do feature engineering for Postal Codes in USA.
- I would like to extract the 2 digit prefix:
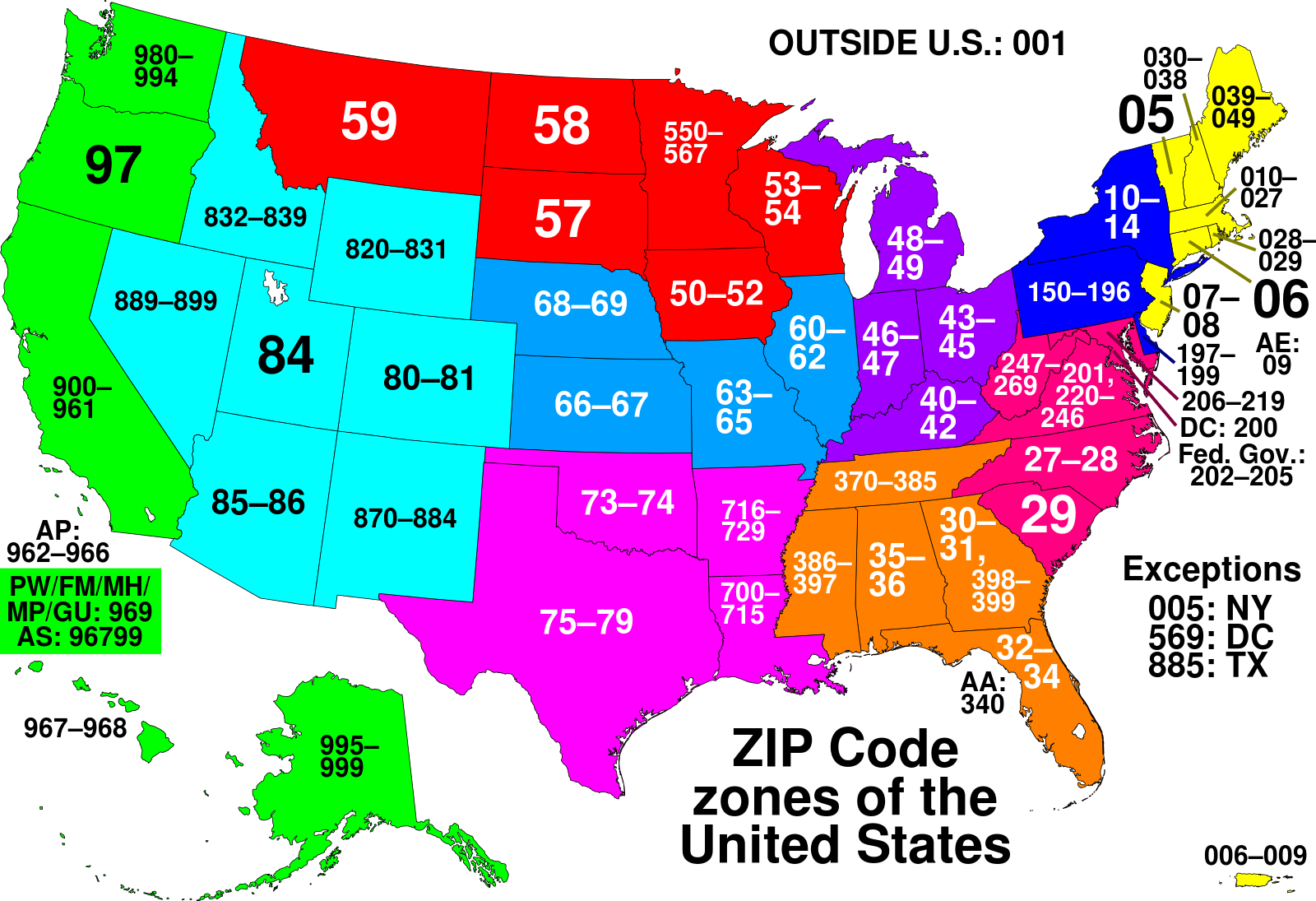
| closed | 2022-09-12T14:38:50Z | 2022-11-29T20:08:15Z | https://github.com/alteryx/featuretools/issues/2284 | [] | gsheni | 0 |
pytest-dev/pytest-mock | pytest | 405 | Failing tests for python 3.12 | When running contribution tests for python 3.12 env:
```bash
tox -e py312
```
I get the following output:
```
py312: install_package> python -I -m pip install --force-reinstall --no-deps /home/brandon/remotes/pytest-mock/.tox/.tmp/package/3/pytest-mock-3.12.1.dev10+g3d48ff9.tar.gz
py312: commands[0]> ... | closed | 2024-01-25T23:04:41Z | 2024-01-25T23:29:35Z | https://github.com/pytest-dev/pytest-mock/issues/405 | [] | blotero | 1 |
fastapi/sqlmodel | sqlalchemy | 150 | How to create computed columns ? | ### First Check
- [X] I added a very descriptive title to this issue.
- [X] I used the GitHub search to find a similar issue and didn't find it.
- [X] I searched the SQLModel documentation, with the integrated search.
- [X] I already searched in Google "How to X in SQLModel" and didn't find any information.
- [X] I al... | open | 2021-10-30T11:35:51Z | 2024-02-14T13:28:18Z | https://github.com/fastapi/sqlmodel/issues/150 | [
"question"
] | sorasful | 12 |
AUTOMATIC1111/stable-diffusion-webui | deep-learning | 15,903 | [Fix Guide] ImportError: cannot import name 'packaging' from 'pkg_resources' | Recent update to the `setuptools=70.0.0` prevents some new installs of Web UI from launching
Error:
```py
ImportError: cannot import name 'packaging' from 'pkg_resources' (venv\lib\site-packages\pkg_resources\__init__.py)
```
initial issue post
- https://github.com/AUTOMATIC1111/stable-diffusion-webui/issues... | closed | 2024-05-28T16:57:51Z | 2024-05-28T16:58:54Z | https://github.com/AUTOMATIC1111/stable-diffusion-webui/issues/15903 | [
"announcement"
] | w-e-w | 0 |
influxdata/influxdb-client-python | jupyter | 403 | create_bucket trigger IndexError: list index out of range | <!--
Thank you for reporting a bug.
* Please add a :+1: or comment on a similar existing bug report instead of opening a new one.
* https://github.com/influxdata/influxdb-client-python/issues?utf8=%E2%9C%93&q=is%3Aissue+is%3Aopen+is%3Aclosed+sort%3Aupdated-desc+label%3Abug+
* Please check whether the bug c... | closed | 2022-02-13T15:51:22Z | 2022-03-18T07:27:23Z | https://github.com/influxdata/influxdb-client-python/issues/403 | [
"state: confirmed"
] | Bugazelle | 3 |
sinaptik-ai/pandas-ai | data-visualization | 1,360 | Unable to analyze the DataFrame when it contains data in list format. | ### System Info
OS version: MacOS Sonoma
Python version: 3.12.5
The current version of `pandasai` being used: 2.2.14
### 🐛 Describe the bug
I tried using pandasai `Agent` to analyze my data in DataFrame format, but I found that if the DataFrame contains data in list format, the analysis fails, and there are no er... | closed | 2024-09-08T06:30:09Z | 2024-12-15T16:08:08Z | https://github.com/sinaptik-ai/pandas-ai/issues/1360 | [
"bug"
] | ReeveWu | 1 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.