repo_name stringlengths 9 75 | topic stringclasses 30
values | issue_number int64 1 203k | title stringlengths 1 976 | body stringlengths 0 254k | state stringclasses 2
values | created_at stringlengths 20 20 | updated_at stringlengths 20 20 | url stringlengths 38 105 | labels listlengths 0 9 | user_login stringlengths 1 39 | comments_count int64 0 452 |
|---|---|---|---|---|---|---|---|---|---|---|---|
jupyter-book/jupyter-book | jupyter | 2,304 | Aligning figures in grid to the bottom. | I am trying to put 2 figures side-by-side.
I am currently trying the `grid` directive for this.
```
:::::{grid} 2
::::{grid-item}
:::{figure} assets/fish_swimming.svg
:height: 350px
Caption 1
:::
::::
::::{grid-item}
:::{figure} assets/stanford_bunny.svg
:height: 225px
Caption 2
:::
::::
:::::
```
This shows the figures as attached:
<img width="743" alt="Image" src="https://github.com/user-attachments/assets/8a6c1af9-6a81-43c4-8265-a78189f7c866" />
How do I bottom align the figures?
Is there a better way to produce subfigures so that I can have proper referencing of subfigures as Fig1a and Fig1b etc. I know the figure directive can be used for that but I want a two column figure style with both bottom aligned. | open | 2025-01-22T16:58:02Z | 2025-01-22T16:58:18Z | https://github.com/jupyter-book/jupyter-book/issues/2304 | [] | atharvaaalok | 0 |
wkentaro/labelme | computer-vision | 792 | Ubuntu 18 , not able to launch issue | File "/usr/lib/python3/dist-packages/pkg_resources/__init__.py", line 783, in resolve
raise VersionConflict(dist, req).with_context(dependent_req)
pkg_resources.ContextualVersionConflict: (Pillow 5.1.0 (/usr/lib/python3/dist-packages), Requirement.parse('pillow>=6.2.0'), {'matplotlib'})
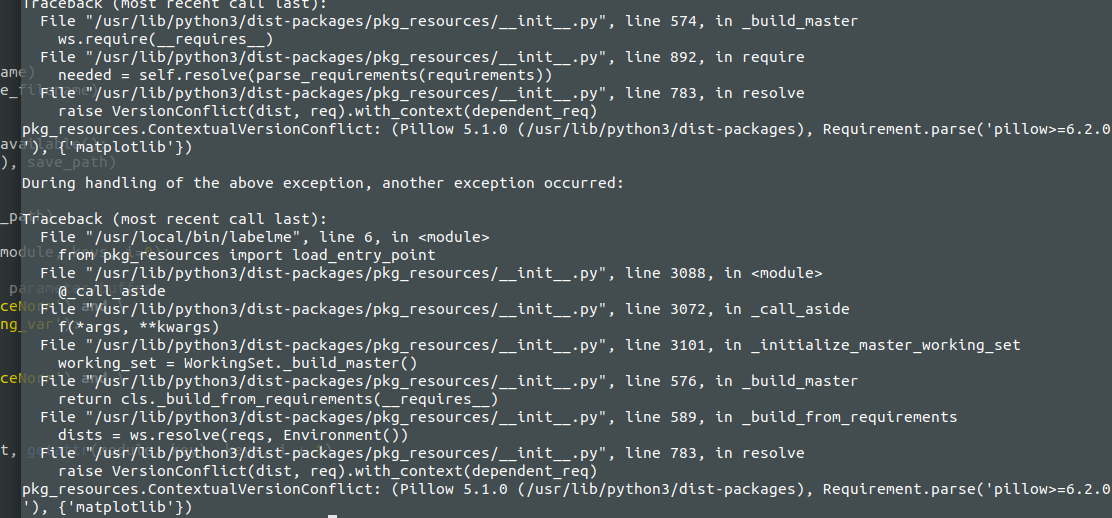
| closed | 2020-10-26T11:41:04Z | 2020-12-11T08:22:23Z | https://github.com/wkentaro/labelme/issues/792 | [] | sreshu | 1 |
Evil0ctal/Douyin_TikTok_Download_API | fastapi | 539 | fetch_user_post 接口只能读取第一页数据? | 获取用户帖子列表的接口只能获取第一页的数据吗?这个作者有687个视频
**`https://douyin.wtf/api/tiktok/web/fetch_user_post?secUid=MS4wLjABAAAAWtC4Km0mPiqpO8CM4JnOTG7sTMqs6ionh6AWF9sFb1dVtKiafyCwNz10DGf2UFk8&cursor=1&count=35&coverFormat=2`**
`{
code: 200,
router: "/api/tiktok/web/fetch_user_post",
data: {
cursor: "1",
extra: {
fatal_item_ids: [ ],
logid: "20250118172756C056BC4051BAEA68F65D",
now: 1737221277000
},
hasMore: false,
log_pb: {
impr_id: "20250118172756C056BC4051BAEA68F65D"
},
statusCode: 0,
status_code: 0,
status_msg: ""
}
}` | closed | 2025-01-17T05:58:49Z | 2025-01-21T04:29:39Z | https://github.com/Evil0ctal/Douyin_TikTok_Download_API/issues/539 | [
"BUG",
"enhancement"
] | Bruse-Lee | 1 |
piskvorky/gensim | machine-learning | 3,162 | Doc2Vec: when we have string tags, build_vocab with update removes previous index | <!--
**IMPORTANT**:
- Use the [Gensim mailing list](https://groups.google.com/forum/#!forum/gensim) to ask general or usage questions. Github issues are only for bug reports.
- Check [Recipes&FAQ](https://github.com/RaRe-Technologies/gensim/wiki/Recipes-&-FAQ) first for common answers.
Github bug reports that do not include relevant information and context will be closed without an answer. Thanks!
-->
#### Problem description
I'm trying to resume training my Doc2Vec model with string tags, but `model.build_vocab` removes all previous index from `model.dv`.
#### Steps/code/corpus to reproduce
A simple example to reproduce this:
```python
import string
from gensim.test.utils import common_texts
from gensim.models.doc2vec import Doc2Vec, TaggedDocument
documents = [TaggedDocument(doc, [tag]) for tag, doc in zip(string.ascii_lowercase, common_texts)]
documents1 = documents[:6]
documents2 = documents[6:]
model = Doc2Vec(vector_size=5, window=2, min_count=1)
model.build_vocab(documents1)
model.train(documents1, total_examples=len(documents1), epochs=5)
model.save('model')
model = Doc2Vec.load('model')
print('Vector count after train:', len(model.dv))
print('Keys:', model.dv.index_to_key)
model.build_vocab(documents2, update=True)
model.train(documents2, total_examples=model.corpus_count, epochs=model.epochs)
print('Vector count after update:', len(model.dv))
print('Keys:', model.dv.index_to_key)
model.save('model')
model = Doc2Vec.load('model')
print('Vector count after load:', len(model.dv))
print('Keys:', model.dv.index_to_key)
```
Output:
```
Vector count after train: 6
Keys: ['a', 'b', 'c', 'd', 'e', 'f']
Vector count after update: 3
Keys: ['g', 'h', 'i']
Vector count after load: 3
Keys: ['g', 'h', 'i']
```
And we have an interesting behavior:
```python
print('b' in model.dv)
# True
print(model.dv['b'])
# [ 0.00524729 -0.19762747 -0.10339681 -0.19433555 0.04022206]
```
The tag seems still exists in the model after updating, but `len` and `index_to_key` do not show this.
At the same time the code with int tags works correctly (it seems to me):
```python
documents = [TaggedDocument(doc, [tag]) for tag, doc in enumerate(common_texts)]
documents1 = documents[:6]
documents2 = documents[6:]
...
```
```
Vector count after train: 6
Keys: [0, 1, 2, 3, 4, 5]
Vector count after update: 9
Keys: [0, 1, 2, 3, 4, 5, 6, 7, 8]
Vector count after load: 9
Keys: [0, 1, 2, 3, 4, 5, 6, 7, 8]
```
#### Versions
```
Windows-10-10.0.19041-SP0
Python 3.9.0 (tags/v3.9.0:9cf6752, Oct 5 2020, 15:34:40) [MSC v.1927 64 bit (AMD64)]
Bits 64
NumPy 1.20.3
SciPy 1.6.1
gensim 4.0.1
FAST_VERSION 0
```
| closed | 2021-06-04T05:54:12Z | 2022-03-17T20:46:48Z | https://github.com/piskvorky/gensim/issues/3162 | [] | espdev | 13 |
python-visualization/folium | data-visualization | 1,813 | Make the Map pickable | **Is your feature request related to a problem? Please describe.**
As an extension of https://github.com/python-visualization/branca/pull/99, I want to be able to cache a map for an application, to switch quickly between them.
**Describe the solution you'd like**
I proposed a first solution in the following PR : https://github.com/python-visualization/folium/pull/1812
The solution is partial and would greatly appreciate any help, as I feel a pickable Map could be helpful in many ways.
**Additional context**
Map before correction :

Map after correction

**Implementation**
See https://github.com/python-visualization/folium/pull/1812
| closed | 2023-10-06T14:45:17Z | 2023-10-16T12:07:50Z | https://github.com/python-visualization/folium/issues/1813 | [
"enhancement"
] | BastienGauthier | 8 |
deepspeedai/DeepSpeed | pytorch | 6,639 | No module named 'op_builder' | deepspeed-0.15.2
AMD 5600x
Rtx 4060Ti
ERROR:
ModuleNotFoundError: No module named 'op_builder'
try install it ,but :
pip install op-builder
ERROR: Could not find a version that satisfies the requirement op-builder (from versions: none)
ERROR: No matching distribution found for op-builder
help!!! | closed | 2024-10-18T04:02:44Z | 2024-11-05T23:31:42Z | https://github.com/deepspeedai/DeepSpeed/issues/6639 | [
"windows"
] | hujiquan | 8 |
ranaroussi/yfinance | pandas | 1,976 | Currency data for statements | ### Describe bug
Currency data for financial statements
For some stocks, like BP.L, the share price data is GBp whereas the financial statements are in USD.
The currency of GBp is available in fast_info but the currency (USD) for the financial statements does not appear to be available.
Am I missing something?
### Simple code that reproduces your problem
import yfinance as yf
ticker = yf.Ticker("BP.L")
ticker.info['currency']
### Debug log
not a bug
### Bad data proof
ticker.info['currency']
'GBp'
### `yfinance` version
0.2.40
### Python version
_No response_
### Operating system
_No response_ | closed | 2024-07-04T19:29:24Z | 2024-07-05T20:11:48Z | https://github.com/ranaroussi/yfinance/issues/1976 | [] | mking007 | 3 |
JaidedAI/EasyOCR | deep-learning | 1,324 | Fine-tuned CRAFT model works much slower on CPU than default one. | I fine-tuned CRAFT model according to this guide: https://github.com/JaidedAI/EasyOCR/tree/master/trainer/craft
But this model works 5 times slower than default model 'craft_mlt_25k' on some server CPUs (on some CPUs speeds are same). What can it be? Is 'craft_mlt_25k' quantized in some way? | open | 2024-10-18T09:21:37Z | 2024-12-09T02:27:33Z | https://github.com/JaidedAI/EasyOCR/issues/1324 | [] | romanvelichkin | 1 |
matplotlib/matplotlib | data-science | 28,872 | [Bug]: Why is there an offset between grey bars and width of arrows in upper limits (reproducible data and code provided) | ### Bug summary
For the image shown below, which shows upper limits in red arrows between two variables X and Y and on the right side there is Z axis showing value of grey bars, why is their an offset between grey bars and width of arrows.

### Code for reproduction
```Python
def plot(Xmin, Xmax):
datafile = '' # Placeholder for the data file path
try:
data = np.loadtxt(datafile)
xmin = data[:, 0]
xmax = data[:, 1]
yvalue = data[:, 2]
yerror = data[:, 3]
zvalue = data[:, 4]
upperBound = data[:, 5]
# Compute the midpoint of the x-axis
x = np.sqrt(xmin * xmax)
xerr = np.array([x - xmin, xmax - x])
# Compute y-axis values
y = x**2 * yvalue / (xmax - xmin)
yerr = x**2 * yerror / (xmax - xmin)
y_ul = x**2 * upperBound / (xmax - xmin)
y_ulerr = np.array([0.5 * y_ul, [0] * len(y)])
# Create the plot
fig, ax = plt.subplots()
# Plot the data points where z > 9
ax.errorbar(x[zvalue > 9], y[zvalue > 9], xerr=xerr[:, zvalue > 9], yerr=yerr[zvalue > 9],
fmt='.k', color='blue', label='Detected', markersize=12)
# Plot upper limits where z < 9, in red
ax.errorbar(x[zvalue < 9], y_ul[zvalue < 9], xerr=xerr[:, zvalue < 9], yerr=y_ulerr[:, zvalue < 9],
fmt='.k', color='red', uplims=True, label='Upper Limit', markersize=12)
# Set axis limits and scales
ax.set_xlim(a,b)
ax.set_ylim(c, d)
ax.set_xscale('log')
ax.set_yscale('log')
# Set axis labels with bold font
ax.set_xlabel(r'X [units]', fontweight='bold')
ax.set_ylabel(r'Y [units]', fontweight='bold', fontsize=9)
# Plot z-values on a secondary axis
ax2 = ax.twinx()
ax2.bar(x, zvalue, width=(xmax - xmin), color='gray', edgecolor='gray', alpha=0.5)
ax2.set_ylim(bottom=0)
ax2.set_xscale('log')
ax2.set_ylabel('Z value', fontweight='bold')
# Return the file paths (placeholders)
return png_file, pdf_file
except Exception as e:
print(f"An error occurred: {e}")
```
### Actual outcome

When we use x = np.sqrt(xmin * xmax) which is the geometrical mean their is an offset between grey bars and width of arrows in upper limits. But when we use arithmetic mean x = 0.5 (xmin + xmax) the upper limit error bars are alligned with the grey bars. Why is it not alligning using geometric mean. The data to reproduce the above plot is at https://github.com/siddhantmannaiith/data/blob/main/data.dat where the first two columns are xmin and xmax. If you need any furthur details to reproduce the above plot please let me know.
### Expected outcome
In expected outcome the upper limits should align with the grey bars which happens when we use arithmetic mean but does not happen with geometric mean. I have provided the data at https://github.com/siddhantmannaiith/data/blob/main/data.dat to replicate the above plot. First two columns are xmin and xmax.
### Additional information
I have provided the data at https://github.com/siddhantmannaiith/data/blob/main/data.dat to replicate the above plot. First two columns are xmin and xmax.
### Operating system
Ubuntu
### Matplotlib Version
3.7.1
### Matplotlib Backend
_No response_
### Python version
_No response_
### Jupyter version
_No response_
### Installation
pip | closed | 2024-09-24T07:03:15Z | 2024-09-24T13:48:56Z | https://github.com/matplotlib/matplotlib/issues/28872 | [
"status: needs clarification",
"status: needs revision"
] | siddhantmannaiith | 1 |
ARM-DOE/pyart | data-visualization | 1,333 | NEXRAD Non-Reflectivity Values | Building on the discussion here - https://openradar.discourse.group/t/reflectivity-and-velocity-resolution-from-aws/97
The main question is whether we are parsing NEXRAD Level 2 files with the proper digital resolution.
Reflectivity should be at 8 bit resolution (every 0.5 dBZ), but other fields such as radial velocity and spectrum width should be higher? With differences in values on the order of 0.1 m/s, etc.
@dopplershift do you know where to find this information so we can verify we are treating this properly? Or do you know if the current implementation in MetPy/Py-ART handle this properly? | closed | 2022-11-22T15:49:15Z | 2022-11-22T19:28:04Z | https://github.com/ARM-DOE/pyart/issues/1333 | [] | mgrover1 | 2 |
aminalaee/sqladmin | sqlalchemy | 152 | Many to many field setup error | ### Checklist
- [X] The bug is reproducible against the latest release or `master`.
- [X] There are no similar issues or pull requests to fix it yet.
### Describe the bug
I am trying to update m2m field in form but i am getting error `"sqlalchemy.exc.InvalidRequestError: Can't attach instance another instance with key is already present in this session"`
### Steps to reproduce the bug
_No response_
### Expected behavior
_No response_
### Actual behavior
_No response_
### Debugging material
_No response_
### Environment
Macos , python 3.9
### Additional context
_No response_ | closed | 2022-05-14T16:16:25Z | 2024-06-15T13:33:42Z | https://github.com/aminalaee/sqladmin/issues/152 | [] | dasaderto | 10 |
pallets-eco/flask-sqlalchemy | flask | 512 | NoForeignKeysError with polymorphism and schemas | ```python
from flask import Flask
from flask_sqlalchemy import SQLAlchemy
app = Flask(__name__)
app.config['SQLALCHEMY_TRACK_MODIFICATIONS'] = False
app.config['SQLALCHEMY_DATABASE_URI'] = 'postgresql:///test'
db = SQLAlchemy(app)
class Item(db.Model):
__tablename__ = 'items'
__table_args__ = {'schema': 'foo'} # this causes the error
__mapper_args__ = {'polymorphic_on': 'type', 'polymorphic_identity': None}
id = db.Column(db.Integer, primary_key=True)
type = db.Column(db.Integer, nullable=False)
parent_id = db.Column(db.ForeignKey('foo.items.id'), index=True, nullable=True)
children = db.relationship('Item', backref=db.backref('parent', remote_side=[id]))
class SubItem(Item):
__mapper_args__ = {
'polymorphic_identity': 1
}
```
traceback:
```pythontraceback
Traceback (most recent call last):
File "flasksatest.py", line 21, in <module>
class SubItem(Item):
File "/home/adrian/dev/indico/env/lib/python2.7/site-packages/flask_sqlalchemy/__init__.py", line 602, in __init__
DeclarativeMeta.__init__(self, name, bases, d)
File "/home/adrian/dev/indico/env/lib/python2.7/site-packages/sqlalchemy/ext/declarative/api.py", line 64, in __init__
_as_declarative(cls, classname, cls.__dict__)
File "/home/adrian/dev/indico/env/lib/python2.7/site-packages/sqlalchemy/ext/declarative/base.py", line 88, in _as_declarative
_MapperConfig.setup_mapping(cls, classname, dict_)
File "/home/adrian/dev/indico/env/lib/python2.7/site-packages/sqlalchemy/ext/declarative/base.py", line 103, in setup_mapping
cfg_cls(cls_, classname, dict_)
File "/home/adrian/dev/indico/env/lib/python2.7/site-packages/sqlalchemy/ext/declarative/base.py", line 135, in __init__
self._early_mapping()
File "/home/adrian/dev/indico/env/lib/python2.7/site-packages/sqlalchemy/ext/declarative/base.py", line 138, in _early_mapping
self.map()
File "/home/adrian/dev/indico/env/lib/python2.7/site-packages/sqlalchemy/ext/declarative/base.py", line 534, in map
**self.mapper_args
File "<string>", line 2, in mapper
File "/home/adrian/dev/indico/env/lib/python2.7/site-packages/sqlalchemy/orm/mapper.py", line 671, in __init__
self._configure_inheritance()
File "/home/adrian/dev/indico/env/lib/python2.7/site-packages/sqlalchemy/orm/mapper.py", line 978, in _configure_inheritance
self.local_table)
File "<string>", line 2, in join_condition
File "/home/adrian/dev/indico/env/lib/python2.7/site-packages/sqlalchemy/sql/selectable.py", line 979, in _join_condition
(a.description, b.description, hint))
sqlalchemy.exc.NoForeignKeysError: Can't find any foreign key relationships between 'items' and 'items'.
```
It works fine if I do not use a custom schema. | closed | 2017-06-27T09:44:42Z | 2020-12-05T20:46:22Z | https://github.com/pallets-eco/flask-sqlalchemy/issues/512 | [
"tablename"
] | ThiefMaster | 3 |
amidaware/tacticalrmm | django | 954 | Server Side Checks | **Is your feature request related to a problem? Please describe.**
We sometimes need to monitor externally hosted services and cannot install an agent
**Describe the solution you'd like**
I would really like if the RMM server itself could carry out some very basic checks similar to the agents. i.e. ping checks, website availability checks (check for status 200) etc. This would be helpful as sometimes we monitor externally hosted services such as web servers, mail servers and VPN gateways where we cannot install an agent
Love this software! Keep up the good work! | closed | 2022-01-25T19:46:00Z | 2022-01-27T11:38:57Z | https://github.com/amidaware/tacticalrmm/issues/954 | [] | daveclev12 | 4 |
pydata/bottleneck | numpy | 9 | Writing beyond the range of an array | The low-level functions nanstd_3d_int32_axis1 and nanstd_3d_int64_axis1, called by bottleneck.nanstd() for 3d input, wrote beyond the memory owned by the output array if arr.shape[1] == 0 and arr.shape[0] > arr.shape[2], where arr is the input array.
Thanks to Christoph Gohlke for finding an example to demonstrate the bug.
| closed | 2011-03-08T20:24:41Z | 2011-03-08T20:51:45Z | https://github.com/pydata/bottleneck/issues/9 | [
"bug"
] | kwgoodman | 1 |
Textualize/rich | python | 2,825 | [REQUEST] Tweak default colors for RichHandler | Hello, first of all, thank you for rich! I use it in pretty much all my projects.
I have a _very minor_ suggestion regarding the _default_ colors for logging levels.
I know we can customize them using [themes](https://rich.readthedocs.io/en/latest/style.html#style-themes) (and I already do!).
I use mostly info/warning/error levels for logging, and depending on the terminal used, _warnings and errors_ render almost identically.
When googling "Python colored logs", the top solutions (in my case) use yellow-ish for warnings and red for errors.
Granted, many of the top results use `coloredlogs`, but in any case I see:
* [stack overflow top answer](https://stackoverflow.com/questions/384076/how-can-i-color-python-logging-output)
* [PyPI coloredlogs](https://pypi.org/project/coloredlogs/)
* [a blog post](https://alexandra-zaharia.github.io/posts/make-your-own-custom-color-formatter-with-python-logging/)
* [another blog post](https://betterstack.com/community/questions/how-to-color-python-logging-output/)
* [PyPI colorlog](https://pypi.org/project/colorlog/)
So I was wondering if you would be willing to tweak the default `'logging.level.warning'` to something closer to yellow, to be more in line with this, and give a bit more distinction to warnings and errors.
Anyway, I am perfectly happy with customization through themes!
| closed | 2023-02-22T14:39:27Z | 2024-07-01T10:43:43Z | https://github.com/Textualize/rich/issues/2825 | [
"accepted"
] | alexprengere | 3 |
modin-project/modin | data-science | 7,350 | Possible issue with `dropna(how="all")` not deleting data from partition on ray. | When processing a large dataframe with modin running on ray, if I had previously dropped invalid rows, it runs into an issue by accessing data from the new dataframe (after dropna).
It looks like the data is not released from ray, or maybe modin `dropna` operation is not removing it properly.
It works fine if I run an operation where modin defaults to pandas.
# EXAMPLE:
```
import modin.pandas as pd
data = [
{"record": 1, "data_set": [0,0,0,0], "index": 1},
{"record": 2, "data_set": [0,0,0,0], "index": 2},
{"record": 3, "data_set": [0,0,0,0], "index": 3},
{"record": 4, "data_set": [0,0,0,0], "index": 4},
{"record": 5, "data_set": [0,0,0,0], "index": 5},
{"record": 6, "data_set": [0,0,0,0], "index": 6},
{"record": 7, "data_set": [0,0,0,0], "index": 7},
{"record": 8, "data_set": [0,0,0,0], "index": 8},
{"record": 9, "data_set": [0,0,0,0], "index": 9},
{"record": 10, "data_set": [0,0,0,0], "index": 10},
] * 10000
modin_df = pd.DataFrame(data)
# process and remove unwanted rows
# imagine this as a more complex than just filtering by index
modin_df = modin_df.apply(lambda x: x if x["index"] < 5 else None, axis=1).dropna(how="all")
# try to access data_set column
# imagine this as a more complex processing job
modin_df.apply(lambda x: x["data_set"], axis=1)
```
# ERROR:
<details>
```python-traceback
{
"name": "RayTaskError(KeyError)",
"message": "ray::_apply_func() (pid=946, ip=10.169.23.29)
At least one of the input arguments for this task could not be computed:
ray.exceptions.RayTaskError: ray::_deploy_ray_func() (pid=942, ip=10.169.23.29)
File \"pandas/_libs/index.pyx\", line 138, in pandas._libs.index.IndexEngine.get_loc
File \"pandas/_libs/index.pyx\", line 165, in pandas._libs.index.IndexEngine.get_loc
File \"pandas/_libs/hashtable_class_helper.pxi\", line 5745, in pandas._libs.hashtable.PyObjectHashTable.get_item
File \"pandas/_libs/hashtable_class_helper.pxi\", line 5753, in pandas._libs.hashtable.PyObjectHashTable.get_item
KeyError: 'data_set'
The above exception was the direct cause of the following exception:
ray::_deploy_ray_func() (pid=942, ip=10.169.23.29)
File \"/Users/brunoj/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/core/execution/ray/implementations/pandas_on_ray/partitioning/virtual_partition.py\", line 313, in _deploy_ray_func
File \"/Users/brunoj/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/core/dataframe/pandas/partitioning/axis_partition.py\", line 419, in deploy_axis_func
File \"/Users/brunoj/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/core/dataframe/pandas/dataframe/dataframe.py\", line 1788, in _tree_reduce_func
File \"/Users/brunoj/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/core/storage_formats/pandas/query_compiler.py\", line 3084, in <lambda>
File \"/home/ray/anaconda3/lib/python3.9/site-packages/pandas/core/frame.py\", line 9568, in apply
return op.apply().__finalize__(self, method=\"apply\")
File \"/home/ray/anaconda3/lib/python3.9/site-packages/pandas/core/apply.py\", line 764, in apply
return self.apply_standard()
File \"/home/ray/anaconda3/lib/python3.9/site-packages/pandas/core/apply.py\", line 891, in apply_standard
results, res_index = self.apply_series_generator()
File \"/home/ray/anaconda3/lib/python3.9/site-packages/pandas/core/apply.py\", line 907, in apply_series_generator
results[i] = self.f(v)
File \"/Users/brunoj/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/utils.py\", line 611, in wrapper
File \"/var/folders/lz/4cs_fypj0ld8x6kyk9rbkl400000gn/T/ipykernel_24081/3890645143.py\", line 24, in <lambda>
File \"/home/ray/anaconda3/lib/python3.9/site-packages/pandas/core/series.py\", line 981, in __getitem__
return self._get_value(key)
File \"/home/ray/anaconda3/lib/python3.9/site-packages/pandas/core/series.py\", line 1089, in _get_value
loc = self.index.get_loc(label)
File \"/home/ray/anaconda3/lib/python3.9/site-packages/pandas/core/indexes/base.py\", line 3804, in get_loc
raise KeyError(key) from err
KeyError: 'data_set'",
"stack": "---------------------------------------------------------------------------
RayTaskError(KeyError) Traceback (most recent call last)
Cell In[79], line 24
20 modin_df = modin_df.apply(lambda x: x if x[\"index\"] < 5 else None, axis=1).dropna(how=\"all\")
22 # try to access data_set column
23 # imagine this as a more complex processing job
---> 24 modin_df.apply(lambda x: x[\"data_set\"], axis=1)
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/logging/logger_decorator.py:128, in enable_logging.<locals>.decorator.<locals>.run_and_log(*args, **kwargs)
113 \"\"\"
114 Compute function with logging if Modin logging is enabled.
115
(...)
125 Any
126 \"\"\"
127 if LogMode.get() == \"disable\":
--> 128 return obj(*args, **kwargs)
130 logger = get_logger()
131 logger_level = getattr(logger, log_level)
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/pandas/dataframe.py:419, in DataFrame.apply(self, func, axis, raw, result_type, args, **kwargs)
416 else:
417 output_type = DataFrame
--> 419 return output_type(query_compiler=query_compiler)
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/logging/logger_decorator.py:128, in enable_logging.<locals>.decorator.<locals>.run_and_log(*args, **kwargs)
113 \"\"\"
114 Compute function with logging if Modin logging is enabled.
115
(...)
125 Any
126 \"\"\"
127 if LogMode.get() == \"disable\":
--> 128 return obj(*args, **kwargs)
130 logger = get_logger()
131 logger_level = getattr(logger, log_level)
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/pandas/series.py:144, in Series.__init__(self, data, index, dtype, name, copy, fastpath, query_compiler)
130 name = data.name
132 query_compiler = from_pandas(
133 pandas.DataFrame(
134 pandas.Series(
(...)
142 )
143 )._query_compiler
--> 144 self._query_compiler = query_compiler.columnarize()
145 if name is not None:
146 self.name = name
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/logging/logger_decorator.py:128, in enable_logging.<locals>.decorator.<locals>.run_and_log(*args, **kwargs)
113 \"\"\"
114 Compute function with logging if Modin logging is enabled.
115
(...)
125 Any
126 \"\"\"
127 if LogMode.get() == \"disable\":
--> 128 return obj(*args, **kwargs)
130 logger = get_logger()
131 logger_level = getattr(logger, log_level)
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/core/storage_formats/base/query_compiler.py:1236, in BaseQueryCompiler.columnarize(self)
1232 if self._shape_hint == \"column\":
1233 return self
1235 if len(self.columns) != 1 or (
-> 1236 len(self.index) == 1 and self.index[0] == MODIN_UNNAMED_SERIES_LABEL
1237 ):
1238 return self.transpose()
1239 return self
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/core/storage_formats/pandas/query_compiler.py:87, in _get_axis.<locals>.<lambda>(self)
74 \"\"\"
75 Build index labels getter of the specified axis.
76
(...)
84 callable(PandasQueryCompiler) -> pandas.Index
85 \"\"\"
86 if axis == 0:
---> 87 return lambda self: self._modin_frame.index
88 else:
89 return lambda self: self._modin_frame.columns
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/core/dataframe/pandas/dataframe/dataframe.py:522, in PandasDataframe._get_index(self)
520 index, row_lengths = self._index_cache.get(return_lengths=True)
521 else:
--> 522 index, row_lengths = self._compute_axis_labels_and_lengths(0)
523 self.set_index_cache(index)
524 if self._row_lengths_cache is None:
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/logging/logger_decorator.py:128, in enable_logging.<locals>.decorator.<locals>.run_and_log(*args, **kwargs)
113 \"\"\"
114 Compute function with logging if Modin logging is enabled.
115
(...)
125 Any
126 \"\"\"
127 if LogMode.get() == \"disable\":
--> 128 return obj(*args, **kwargs)
130 logger = get_logger()
131 logger_level = getattr(logger, log_level)
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/core/dataframe/pandas/dataframe/dataframe.py:626, in PandasDataframe._compute_axis_labels_and_lengths(self, axis, partitions)
624 if partitions is None:
625 partitions = self._partitions
--> 626 new_index, internal_idx = self._partition_mgr_cls.get_indices(axis, partitions)
627 return new_index, list(map(len, internal_idx))
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/logging/logger_decorator.py:128, in enable_logging.<locals>.decorator.<locals>.run_and_log(*args, **kwargs)
113 \"\"\"
114 Compute function with logging if Modin logging is enabled.
115
(...)
125 Any
126 \"\"\"
127 if LogMode.get() == \"disable\":
--> 128 return obj(*args, **kwargs)
130 logger = get_logger()
131 logger_level = getattr(logger, log_level)
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/core/dataframe/pandas/partitioning/partition_manager.py:933, in PandasDataframePartitionManager.get_indices(cls, axis, partitions, index_func)
931 if len(target):
932 new_idx = [idx.apply(func) for idx in target[0]]
--> 933 new_idx = cls.get_objects_from_partitions(new_idx)
934 else:
935 new_idx = [pandas.Index([])]
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/logging/logger_decorator.py:128, in enable_logging.<locals>.decorator.<locals>.run_and_log(*args, **kwargs)
113 \"\"\"
114 Compute function with logging if Modin logging is enabled.
115
(...)
125 Any
126 \"\"\"
127 if LogMode.get() == \"disable\":
--> 128 return obj(*args, **kwargs)
130 logger = get_logger()
131 logger_level = getattr(logger, log_level)
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/core/dataframe/pandas/partitioning/partition_manager.py:874, in PandasDataframePartitionManager.get_objects_from_partitions(cls, partitions)
870 partitions[idx] = part.force_materialization()
871 assert all(
872 [len(partition.list_of_blocks) == 1 for partition in partitions]
873 ), \"Implementation assumes that each partition contains a single block.\"
--> 874 return cls._execution_wrapper.materialize(
875 [partition.list_of_blocks[0] for partition in partitions]
876 )
877 return [partition.get() for partition in partitions]
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/core/execution/ray/common/engine_wrapper.py:92, in RayWrapper.materialize(cls, obj_id)
77 @classmethod
78 def materialize(cls, obj_id):
79 \"\"\"
80 Get the value of object from the Plasma store.
81
(...)
90 Whatever was identified by `obj_id`.
91 \"\"\"
---> 92 return ray.get(obj_id)
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/ray/_private/auto_init_hook.py:21, in wrap_auto_init.<locals>.auto_init_wrapper(*args, **kwargs)
18 @wraps(fn)
19 def auto_init_wrapper(*args, **kwargs):
20 auto_init_ray()
---> 21 return fn(*args, **kwargs)
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/ray/_private/client_mode_hook.py:102, in client_mode_hook.<locals>.wrapper(*args, **kwargs)
98 if client_mode_should_convert():
99 # Legacy code
100 # we only convert init function if RAY_CLIENT_MODE=1
101 if func.__name__ != \"init\" or is_client_mode_enabled_by_default:
--> 102 return getattr(ray, func.__name__)(*args, **kwargs)
103 return func(*args, **kwargs)
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/ray/util/client/api.py:42, in _ClientAPI.get(self, vals, timeout)
35 def get(self, vals, *, timeout=None):
36 \"\"\"get is the hook stub passed on to replace `ray.get`
37
38 Args:
39 vals: [Client]ObjectRef or list of these refs to retrieve.
40 timeout: Optional timeout in milliseconds
41 \"\"\"
---> 42 return self.worker.get(vals, timeout=timeout)
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/ray/util/client/worker.py:433, in Worker.get(self, vals, timeout)
431 op_timeout = max_blocking_operation_time
432 try:
--> 433 res = self._get(to_get, op_timeout)
434 break
435 except GetTimeoutError:
File ~/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/ray/util/client/worker.py:461, in Worker._get(self, ref, timeout)
459 logger.exception(\"Failed to deserialize {}\".format(chunk.error))
460 raise
--> 461 raise err
462 if chunk.total_size > OBJECT_TRANSFER_WARNING_SIZE and log_once(
463 \"client_object_transfer_size_warning\"
464 ):
465 size_gb = chunk.total_size / 2**30
RayTaskError(KeyError): ray::_apply_func() (pid=946, ip=10.169.23.29)
At least one of the input arguments for this task could not be computed:
ray.exceptions.RayTaskError: ray::_deploy_ray_func() (pid=942, ip=10.169.23.29)
File \"pandas/_libs/index.pyx\", line 138, in pandas._libs.index.IndexEngine.get_loc
File \"pandas/_libs/index.pyx\", line 165, in pandas._libs.index.IndexEngine.get_loc
File \"pandas/_libs/hashtable_class_helper.pxi\", line 5745, in pandas._libs.hashtable.PyObjectHashTable.get_item
File \"pandas/_libs/hashtable_class_helper.pxi\", line 5753, in pandas._libs.hashtable.PyObjectHashTable.get_item
KeyError: 'data_set'
The above exception was the direct cause of the following exception:
ray::_deploy_ray_func() (pid=942, ip=10.169.23.29)
File \"/Users/brunoj/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/core/execution/ray/implementations/pandas_on_ray/partitioning/virtual_partition.py\", line 313, in _deploy_ray_func
File \"/Users/brunoj/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/core/dataframe/pandas/partitioning/axis_partition.py\", line 419, in deploy_axis_func
File \"/Users/brunoj/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/core/dataframe/pandas/dataframe/dataframe.py\", line 1788, in _tree_reduce_func
File \"/Users/brunoj/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/core/storage_formats/pandas/query_compiler.py\", line 3084, in <lambda>
File \"/home/ray/anaconda3/lib/python3.9/site-packages/pandas/core/frame.py\", line 9568, in apply
return op.apply().__finalize__(self, method=\"apply\")
File \"/home/ray/anaconda3/lib/python3.9/site-packages/pandas/core/apply.py\", line 764, in apply
return self.apply_standard()
File \"/home/ray/anaconda3/lib/python3.9/site-packages/pandas/core/apply.py\", line 891, in apply_standard
results, res_index = self.apply_series_generator()
File \"/home/ray/anaconda3/lib/python3.9/site-packages/pandas/core/apply.py\", line 907, in apply_series_generator
results[i] = self.f(v)
File \"/Users/brunoj/.pyenv/versions/3.9.18/envs/che/lib/python3.9/site-packages/modin/utils.py\", line 611, in wrapper
File \"/var/folders/lz/4cs_fypj0ld8x6kyk9rbkl400000gn/T/ipykernel_24081/3890645143.py\", line 24, in <lambda>
File \"/home/ray/anaconda3/lib/python3.9/site-packages/pandas/core/series.py\", line 981, in __getitem__
return self._get_value(key)
File \"/home/ray/anaconda3/lib/python3.9/site-packages/pandas/core/series.py\", line 1089, in _get_value
loc = self.index.get_loc(label)
File \"/home/ray/anaconda3/lib/python3.9/site-packages/pandas/core/indexes/base.py\", line 3804, in get_loc
raise KeyError(key) from err
KeyError: 'data_set'"
}
```
</details>
# INSTALLED VERSION
```
UserWarning: Setuptools is replacing distutils.
INSTALLED VERSIONS
------------------
commit : f5f9ae993ba5ed26461d3c9d26fbefecab88ee69
python : 3.9.18.final.0
python-bits : 64
OS : Darwin
OS-release : 23.5.0
Version : Darwin Kernel Version 23.5.0: Wed May 1 20:12:58 PDT 2024; root:xnu-10063.121.3~5/RELEASE_ARM64_T6000
machine : arm64
processor : arm
byteorder : little
LC_ALL : None
LANG : None
LOCALE : None.UTF-8
Modin dependencies
------------------
modin : 0.31.0+5.gf5f9ae99
ray : 2.23.0
dask : 2024.7.1
distributed : 2024.7.1
pandas dependencies
-------------------
pandas : 2.2.2
numpy : 1.26.4
pytz : 2024.1
dateutil : 2.9.0.post0
setuptools : 69.5.1
pip : 24.0
Cython : None
pytest : None
hypothesis : None
sphinx : None
blosc : None
feather : None
xlsxwriter : None
lxml.etree : None
html5lib : None
pymysql : None
psycopg2 : None
jinja2 : 3.1.4
IPython : 8.18.1
pandas_datareader : None
adbc-driver-postgresql: None
adbc-driver-sqlite : None
bs4 : 4.12.3
bottleneck : None
dataframe-api-compat : None
fastparquet : None
fsspec : 2024.6.1
gcsfs : 2024.6.1
matplotlib : 3.9.1
numba : None
numexpr : None
odfpy : None
openpyxl : None
pandas_gbq : 0.23.1
pyarrow : 14.0.2
pyreadstat : None
python-calamine : None
pyxlsb : None
s3fs : 2024.6.1
scipy : 1.13.1
sqlalchemy : None
tables : None
tabulate : 0.9.0
xarray : None
xlrd : None
zstandard : None
tzdata : 2024.1
qtpy : None
pyqt5 : None
``` | open | 2024-07-23T11:05:03Z | 2024-07-25T21:22:44Z | https://github.com/modin-project/modin/issues/7350 | [
"bug 🦗",
"P0"
] | brunojensen | 1 |
serengil/deepface | machine-learning | 882 | Why is the euclidean distance calculated that way and not using np.linalign.norm? | Just curios.
| closed | 2023-11-02T19:31:27Z | 2023-11-02T22:29:47Z | https://github.com/serengil/deepface/issues/882 | [
"question"
] | ghost | 1 |
vimalloc/flask-jwt-extended | flask | 57 | Version Logs? | @vimalloc Should we have a log of changes to clarify if newer releases might break previous versions? | closed | 2017-06-15T14:06:16Z | 2017-07-02T18:58:59Z | https://github.com/vimalloc/flask-jwt-extended/issues/57 | [] | rlam3 | 4 |
unionai-oss/pandera | pandas | 1,759 | Pass additional `Check` kwargs into `register_check_method` | **Is your feature request related to a problem? Please describe.**
Adding a custom error message to an inline custom check works now due to you're recent commits, because you can pass an error argument to the `Check` object. Thank you by the way. I could have missed it, but is there a way to extend that ability class based custom checks? I assumed maybe `register_check_method` or `Field` would take additional arguments for the init of the Check object, but they don't seem to.
**Describe the solution you'd like**
I was able to hack something together, but I'm not really qualified to muck around or contribute to a project like this. Even though I'd love to.
```python
def register_check_method( # pylint:disable=too-many-branches
check_fn=None,
*,
statistics: Optional[List[str]] = None,
supported_types: Optional[Union[type, Tuple, List]] = None,
check_type: Union[CheckType, str] = "vectorized",
strategy=None,
**kwargs # + ln 142
):
```
```python
if check_fn is None:
return partial(
register_check_method,
statistics=statistics,
supported_types=supported_types,
check_type=check_type,
strategy=strategy,
**kwargs # + ln 233
)
```
```python
if check_fn is None:
return partial(
register_check_method,
statistics=statistics,
supported_types=supported_types,
check_type=check_type,
strategy=strategy,
**kwargs # + ln 233
)
```
```python
def validate_check_kwargs(check_kwargs):
check_kwargs = check_kwargs | kwargs # + ln 259
msg = (
f"'{check_fn.__name__} has check_type={check_type}. "
"Providing the following arguments will have no effect: "
"{}. Remove these arguments to avoid this warning."
)
```
...
| open | 2024-07-20T19:07:11Z | 2025-01-05T20:15:42Z | https://github.com/unionai-oss/pandera/issues/1759 | [
"enhancement"
] | typkrft | 1 |
dunossauro/fastapi-do-zero | sqlalchemy | 175 | Repositório do Paulo Cesar Peixoto (PC) | | Link do projeto | Seu @ no git | Comentário (opcional) |
| --- | --- | --- |
| [fast_zero_api](https://github.com/peixoto-pc/fast_api_zero) | [@peixoto-pc ](https://github.com/peixoto-pc)| Implementação do material do curso sem alterações | | closed | 2024-06-14T21:57:08Z | 2024-06-15T00:55:29Z | https://github.com/dunossauro/fastapi-do-zero/issues/175 | [] | peixoto-pc | 1 |
graphql-python/graphene-sqlalchemy | sqlalchemy | 211 | AssertionError: Found different types with the same name in the schema | I have two Classes Products and SalableProducts in my Models (SalableProducts inherits from Products so it has every field of it's database), in my Schema here is what i did
```python
class Product(SQLAlchemyObjectType):
class Meta:
model = ProductModel
interfaces = (relay.Node, )
class ProductConnections(relay.Connection):
class Meta:
node = Product
```
```python
class SalableProduct(SQLAlchemyObjectType):
class Meta:
model = SalableProductModel
interfaces = (relay.Node, )
class SalableProductConnections(relay.Connection):
class Meta:
node = SalableProduct
```
and here is my Query class :
```python
class Query(graphene.ObjectType):
node = relay.Node.Field()
all_products = SQLAlchemyConnectionField(ProductConnections)
all_salable_products = SQLAlchemyConnectionField(SalableProductConnections)
```
When i run my server i got this error :
AssertionError: Found different types with the same name in the schema: product_status, product_status. | open | 2019-04-29T11:04:01Z | 2022-07-18T21:24:07Z | https://github.com/graphql-python/graphene-sqlalchemy/issues/211 | [] | Rafik-Belkadi | 19 |
clovaai/donut | computer-vision | 221 | donut-base-finetuned-cord-v2 Demo not working properly in Gradio Space web demo | There is a run time error when the demo link is launched. I think there might be some dependency issue for models build on CORD dataset for document parsing. | open | 2023-07-02T03:01:20Z | 2023-11-25T07:48:35Z | https://github.com/clovaai/donut/issues/221 | [] | being-invincible | 1 |
Colin-b/pytest_httpx | pytest | 40 | Support for httpx > 0.17.x | `httpx` released a [new version](https://github.com/encode/httpx/releases/tag/0.18.0). Currently the `httpx` is limited to [`0.17.*`](https://github.com/Colin-b/pytest_httpx/blob/develop/setup.py#L41).
It would be nice if `pytest-httpx` is updated.
Thanks | closed | 2021-04-27T15:42:14Z | 2021-05-04T18:28:42Z | https://github.com/Colin-b/pytest_httpx/issues/40 | [
"enhancement"
] | fabaff | 3 |
streamlit/streamlit | deep-learning | 10,673 | st.data_editor: Pasting a copied range fails when the bottom-right cell is empty or None | ### Checklist
- [x] I have searched the [existing issues](https://github.com/streamlit/streamlit/issues) for similar issues.
- [x] I added a very descriptive title to this issue.
- [x] I have provided sufficient information below to help reproduce this issue.
### Summary
When copying a selected range of cells from st.data_editor and pasting it into another part of the table, the paste operation does not work if the bottom-right cell of the copied selection contains either an empty string ("") or None.
**Conditions**
Copying a range of cells works as expected.
However, when pasting the copied content into another part of the table, nothing happens if the bottom-right cell of the copied selection is empty ("") or None.
Sample table:

Copying a selected range of cells from st.data_editor:

Pasting it into another part of the table does not work:

### Reproducible Code Example
[](https://issues.streamlitapp.com/?issue=gh-10673)
```Python
import streamlit as st
import pandas as pd
samples = {
"col1": ["test11", "test12", "test13"],
"col2": ["test21", "", "test23"],
"col3": ["test31", "test32", None]
}
df = pd.DataFrame(samples)
editor = st.data_editor(df, num_rows="dynamic")
```
### Steps To Reproduce
1. Run the sample code.
2. Select a range of cells, e.g., from row 1, column 1 (test11) to row 2, column 2 ("").
3. Copy the selected range (Ctrl+C).
4. Try pasting it into row 2, column 1 (Ctrl+V).
5. Observe that the paste operation does not work.
### Expected Behavior
The selected range should be pasted successfully, regardless of whether the bottom-right cell is empty ("") or None.
### Current Behavior
Copying fails when the bottom-right cell of the selection is empty ("") or None.
### Is this a regression?
- [ ] Yes, this used to work in a previous version.
### Debug info
- Streamlit version: 1.43.0
- Python version: 3.11.0
- Operating System: Windows11
- Browser: Chrome
### Additional Information
_No response_ | open | 2025-03-07T05:57:07Z | 2025-03-14T02:55:26Z | https://github.com/streamlit/streamlit/issues/10673 | [
"type:bug",
"status:confirmed",
"priority:P3",
"feature:st.data_editor"
] | hirokika | 4 |
KaiyangZhou/deep-person-reid | computer-vision | 563 | torchreid | It's not 'torchreid.utils', it should be 'torchreid.reid.utils' | open | 2023-11-10T11:34:07Z | 2023-11-10T11:34:07Z | https://github.com/KaiyangZhou/deep-person-reid/issues/563 | [] | motherflunker | 0 |
strawberry-graphql/strawberry | graphql | 3,431 | Should we hide fields that starts with `_` by default? | We have strawberry.Private to hide field, but I was wondering if we should automatically hide fields that start with a underscore, since it is a common convention in Python to use it for "private" fields.
What do you all think? | open | 2024-04-02T11:55:00Z | 2025-03-20T15:56:39Z | https://github.com/strawberry-graphql/strawberry/issues/3431 | [] | patrick91 | 0 |
modin-project/modin | pandas | 7,385 | FEAT: Add type annotations to frontend methods | **Is your feature request related to a problem? Please describe.**
Many frontend methods are missing type annotations on parameters or return types, which are necessary for downstream extension libraries to generate annotations in documentation. | open | 2024-09-05T22:08:38Z | 2024-09-05T22:08:52Z | https://github.com/modin-project/modin/issues/7385 | [
"P3",
"Interfaces and abstractions"
] | noloerino | 0 |
deezer/spleeter | deep-learning | 161 | [Discussion] No GPU stress | Curious as to why activity monitor on Windows tells me that my GPU is barely used during `spleeter-gpu`.
I know for a fact that `spleeter-gpu` is running because my spleeter sessions complete A LOT faster now compared to when I run `spleeter-cpu`.
Usage: 3%
Dedicated GPU-Memory: 0,7 / 8,0 GB
GPU-Memory: 0,8 / 16,0 GB
Shared GPU-Memory: 0,1 / 8,0 GB
Why is this happening?
How can I take advantage of this?
Could on-board graphics be involved here? | closed | 2019-12-05T04:16:41Z | 2019-12-18T15:03:51Z | https://github.com/deezer/spleeter/issues/161 | [
"question"
] | aidv | 2 |
airtai/faststream | asyncio | 1,059 | Feature: subscribers should be resilient to segmentation faults | Thank you for FastStream, I really enjoy the use of pydantic here :smiley:
**Is your feature request related to a problem? Please describe.**
Segmentation faults can happen in the process of handling a message, when involving a library causing the segmentation fault (I don't think it is possible to cause a segmentation fault with native Python code).
When a segmentation fault occurs in the FastStream application which consumes the message, the application stops and is not restarted (that is for faststream[rabbit]==0.3.6; for 0.2.5, the application was hanging defunct). The message is not processed nor redirected to a dead letter queue, for example (in the case of a RabbitMQ cluster).
**Describe the solution you'd like**
I suggest that the message causing the segmentation fault does not stop the application, which would react the same way as if the message had raised an error/exception: the message is rejected and the subscriber keeps on consuming the next message.
**Feature code example**
I published this project to demonstrate how a segmentation fault stops the subscriber application: https://github.com/lucsorel/sigseg-faststream.
**Describe alternatives you've considered**
Being resilient to segmentation faults might involve handling each message in a sub-process for the main process to be resilient to segmentation faults. | closed | 2023-12-15T16:11:37Z | 2024-07-09T15:47:33Z | https://github.com/airtai/faststream/issues/1059 | [
"enhancement"
] | lucsorel | 10 |
PaddlePaddle/models | computer-vision | 5,368 | 下载不了pix2pix模型 | https://paddle-gan-models.bj.bcebos.com/pix2pix_G.tar.gz
https://www.paddlepaddle.org.cn/modelbasedetail/pix2pix | open | 2021-11-10T02:03:24Z | 2024-02-26T05:08:29Z | https://github.com/PaddlePaddle/models/issues/5368 | [] | zhenzi0322 | 0 |
ageitgey/face_recognition | python | 1,341 | Landmark detection is pretty slow :( | * face_recognition version: 1.3.0
* Python version: 3.9.5
* Operating System: Mac OS 10.14.6
I am detecting face landmarks. Mostly nose bridge in order to crop the images later. I found it pretty slow to do it. About 6 seconds per image.
Is there a way to speed up the process ? Can I look only for the nose bridge landmarks somehow ? Would that be faster ? Also file size might be a problem ?
Any help is appreciated !

`from PIL import Image, ImageDraw
import face_recognition
# Load the jpg file into a numpy array
image = face_recognition.load_image_file("test.jpg")
# Find all facial features in all the faces in the image
face_landmarks_list = face_recognition.face_landmarks(image)
print("I found {} face(s) in this photograph.".format(len(face_landmarks_list)))
# Create a PIL imagedraw object so we can draw on the picture
pil_image = Image.fromarray(image)
d = ImageDraw.Draw(pil_image)
for face_landmarks in face_landmarks_list:
# Print the location of each facial feature in this image
for facial_feature in face_landmarks.keys():
print("The {} in this face has the following points: {}".format(facial_feature, face_landmarks[facial_feature]))
# Let's trace out each facial feature in the image with a line!
for facial_feature in face_landmarks.keys():
d.line(face_landmarks[facial_feature], width=2)
# Show the picturew
pil_image.show()
cv2.imwrite('result.png', test)` | open | 2021-07-09T11:12:31Z | 2022-07-01T19:11:19Z | https://github.com/ageitgey/face_recognition/issues/1341 | [] | schwarzwals | 4 |
quokkaproject/quokka | flask | 663 | jinja2.exceptions.UndefinedError: 'theme' is undefined | when i create a block or page, and try to view i get:
```
2018-06-12 11:51:20,001 - werkzeug - ERROR - Error on request:
Traceback (most recent call last):
File "/Users/kyle/git/blog/venv/lib/python3.6/site-packages/werkzeug/serving.py", line 270, in run_wsgi
execute(self.server.app)
File "/Users/kyle/git/blog/venv/lib/python3.6/site-packages/werkzeug/serving.py", line 258, in execute
application_iter = app(environ, start_response)
File "/Users/kyle/git/blog/venv/lib/python3.6/site-packages/flask/app.py", line 2309, in __call__
return self.wsgi_app(environ, start_response)
File "/Users/kyle/git/blog/venv/lib/python3.6/site-packages/flask/app.py", line 2295, in wsgi_app
response = self.handle_exception(e)
File "/Users/kyle/git/blog/venv/lib/python3.6/site-packages/flask/app.py", line 1748, in handle_exception
return self.finalize_request(handler(e), from_error_handler=True)
File "/Users/kyle/git/blog/venv/lib/python3.6/site-packages/quokka/core/error_handlers.py", line 54, in server_error_page
return render_template("errors/server_error.html"), 500
File "/Users/kyle/git/blog/venv/lib/python3.6/site-packages/flask/templating.py", line 135, in render_template
context, ctx.app)
File "/Users/kyle/git/blog/venv/lib/python3.6/site-packages/flask/templating.py", line 117, in _render
rv = template.render(context)
File "/Users/kyle/git/blog/venv/lib/python3.6/site-packages/jinja2/asyncsupport.py", line 76, in render
return original_render(self, *args, **kwargs)
File "/Users/kyle/git/blog/venv/lib/python3.6/site-packages/jinja2/environment.py", line 1008, in render
return self.environment.handle_exception(exc_info, True)
File "/Users/kyle/git/blog/venv/lib/python3.6/site-packages/jinja2/environment.py", line 780, in handle_exception
reraise(exc_type, exc_value, tb)
File "/Users/kyle/git/blog/venv/lib/python3.6/site-packages/jinja2/_compat.py", line 37, in reraise
raise value.with_traceback(tb)
File "/Users/kyle/git/blog/venv/lib/python3.6/site-packages/quokka/templates/errors/server_error.html", line 2, in top-level template code
{% extends theme("base.html") %}
jinja2.exceptions.UndefinedError: 'theme' is undefined
``` | closed | 2018-06-12T18:53:57Z | 2018-07-23T20:16:48Z | https://github.com/quokkaproject/quokka/issues/663 | [] | jstacoder | 1 |
geopandas/geopandas | pandas | 2,478 | BUG: AttributeError about datetimelike values when reading file with Fiona engine | - [x] I have checked that this issue has not already been reported.
- [x] I have confirmed this bug exists on the latest version of geopandas (on the date of issue, last version is 0.11.0).
- [ ] (optional) I have confirmed this bug exists on the main branch of geopandas.
---
**Note**: Please read [this guide](https://matthewrocklin.com/blog/work/2018/02/28/minimal-bug-reports) detailing how to provide the necessary information for us to reproduce your bug.
#### Code Sample, a copy-pastable example
```python
import geopandas as gpd
path_to_file = <path to the file attached>
gdf = gpd.read_file(path_to_file)
```
#### Problem description
The attached file contains date-like columns. With the last version of geopandas I'm not able to read it anymore, as it raises an AttributeError `AttributeError: Can only use .dt accessor with datetimelike values`.
It was not the case with previous version of geopandas. The date-like columns were identified as object columns but at least no error were raised.
If I switch the engine from Fiona to Pyogrio to read the file, there is no error raised and all columns are well detected as datetime columns.
#### Expected Output
Have all columns parsed with no error raised. Here the output I obtained with engine Pyogrio instead of Fiona:
```
>>> gdf.dtypes
cleabs object
nature object
nature_detaillee object
toponyme object
statut_du_toponyme object
fictif bool
etat_de_l_objet object
date_creation datetime64[ns]
date_modification datetime64[ns]
date_d_apparition datetime64[ns]
date_de_confirmation datetime64[ns]
sources object
identifiants_sources object
precision_planimetrique float64
geometry geometry
```
#### Output of ``geopandas.show_versions()``
<details>
<pre>
SYSTEM INFO
-----------
python : 3.8.10 (default, Mar 15 2022, 12:22:08) [GCC 9.4.0]
executable : my_env/bin/python
machine : Linux-5.13.0-51-generic-x86_64-with-glibc2.29
GEOS, GDAL, PROJ INFO
---------------------
GEOS : None
GEOS lib : None
GDAL : 3.4.1
GDAL data dir: my_env/lib/python3.8/site-packages/fiona/gdal_data
PROJ : 8.2.0
PROJ data dir: my_env/lib/python3.8/site-packages/pyproj/proj_dir/share/proj
PYTHON DEPENDENCIES
-------------------
geopandas : 0.11.0
pandas : 1.4.3
fiona : 1.8.21
numpy : 1.23.0
shapely : 1.8.2
rtree : None
pyproj : 3.3.1
matplotlib : None
mapclassify: None
geopy : None
psycopg2 : None
geoalchemy2: None
pyarrow : None
pygeos : None
</pre>
</details>
Attached file: [grosfi.ch/GzMKuv5CHgu](https://www.grosfichiers.com/GzMKuv5CHgu) | closed | 2022-06-27T14:32:37Z | 2022-07-24T09:10:04Z | https://github.com/geopandas/geopandas/issues/2478 | [
"regression"
] | paumillet | 3 |
hankcs/HanLP | nlp | 594 | 在提取关键词的过程中,根据词性过滤得出的关键词 | 在提取关键词的过程中,我想根据词性过滤,只留下词性为名词的关键词。
我的想法是在分词的步骤中,就按照词性去过滤,只留下名词,但是在标准分词源码部分,并没有找到关于词性的代码,hankcs能否指点一下小弟呢?不胜感激
| closed | 2017-07-31T10:30:37Z | 2020-01-01T11:08:34Z | https://github.com/hankcs/HanLP/issues/594 | [
"ignored"
] | cpeixin | 4 |
yunjey/pytorch-tutorial | pytorch | 62 | Evaluation mode in Resnet | I have a question about the evaluation mode. I found that in the resnet tutorial, the network is not set to evaluation through resnet.eval(). Will this affect the testing accuracy? Thanks! | closed | 2017-09-20T08:59:55Z | 2017-10-12T04:59:22Z | https://github.com/yunjey/pytorch-tutorial/issues/62 | [] | zhangmozhe | 1 |
milesmcc/shynet | django | 73 | You were added to awesome-humane-tech | This is just a FYI issue to notify that you were added to the curated awesome-humane-tech in the 'Analytics' category, and - if you like that - are now entitled to wear our badge:
[](https://github.com/humanetech-community/awesome-humane-tech)
By adding this to the README:
```markdown
[](https://github.com/humanetech-community/awesome-humane-tech)
```
https://github.com/humanetech-community/awesome-humane-tech | closed | 2020-08-15T06:25:15Z | 2020-08-15T15:53:15Z | https://github.com/milesmcc/shynet/issues/73 | [
"meta"
] | aschrijver | 1 |
ScrapeGraphAI/Scrapegraph-ai | machine-learning | 544 | Scrapegraph returns relative path URLs instead of absolute path **Possible Bug?** | **Describe the bug**
When using gpt4o as the llm and scraping a webpage to return a list of links, sometimes the paths returned are :
- relative paths (OR)
- full path with an incorrect prefix/domain usually "http://example.com"
The behaviour was consistent until 3 days ago i.e. it always returned full paths on a large dataset as well. Since then, I had to uninstall Scrapegraph and reinstall the library and that's when this issue started popping up.
**Expected behavior**
For example : asking to scrape a website `www.some-actual-website.com` and return a list of webpages that contain information about the contact details of the company, used to consistently/always return a json like :
```
{"list_of_urls": "['www.some-actual-website.com/about','www.some-actual-website.com/contact-us']"}
```
However, now I get either :
```
{"list_of_urls": "['https://example.com/about', 'https://example.com/contact-us']"}
```
OR
```
{"list_of_urls": "['/about','/contact-us']"}
```
I'm curious , shouldn't the list of URLs being parsed/scraped be a straightforward output? Is the final output always produced by the LLM?
**Desktop (please complete the following information):**
- Ubuntu 22.04
- Chromium Browser with Playwright
| closed | 2024-08-13T10:28:49Z | 2024-09-12T14:14:34Z | https://github.com/ScrapeGraphAI/Scrapegraph-ai/issues/544 | [] | sandeepchittilla | 12 |
s3rius/FastAPI-template | asyncio | 219 | taskiq scheduler does not run ... | I followed the documentation for Taskiq [here](https://taskiq-python.github.io/available-components/schedule-sources.html#redisschedulesource) to set up scheduler in my tkq.py file, like following:
```
result_backend = RedisAsyncResultBackend(
redis_url=str(settings.redis_url.with_path("/1")),
)
broker = ListQueueBroker(
str(settings.redis_url.with_path("/1")),
).with_result_backend(result_backend)
scheduler = TaskiqScheduler(broker=broker, sources=[LabelScheduleSource(broker)])
```
And I created an example task:
```
@broker.task(schedule=[{"cron": "*/1 * * * *", "cron_offset": None, "time": None, "args": [10], "kwargs": {}, "labels": {}}])
async def heavy_task(a: int) -> int:
if broker.is_worker_process:
logger.info("heavy_task: {} is in worker process!!!", a)
else:
logger.info("heavy_task: {} NOT in worker process", a)
return 100 + a
```
In the docker-compose.yml file, I start the broker and scheduler like so:
```
taskiq-worker:
<<: *main_app
labels: []
command:
- taskiq
- worker
- market_insights.tkq:scheduler && market_insights.tkq:broker
```
However, the taskiq scheduler does not seem to do anything. I guess I must be missing something. Can some experts help? Thanks | closed | 2024-07-17T04:39:18Z | 2024-07-20T21:13:29Z | https://github.com/s3rius/FastAPI-template/issues/219 | [] | rcholic | 3 |
globaleaks/globaleaks-whistleblowing-software | sqlalchemy | 3,654 | Deleting the information provided by the whistleblower without deleting all the report | ### Proposal
Our proposal is to add a new feature that allows the recipient to deleat the information provided by the whistleblower without deleating all the report, so that the recipient can keep the communication with the whistleblower through the comments. This way, for exemple, whistleblowers can be informed that their denunciations have not been accepted, while we accomplish the rule of deletting the information after decidding whether to start or not an investigation (rule that we mention below).
### Motivation and context
* The current functionalities of the platform permit deleating the report. But, by doing that, it also deletes the possibility of keeping the communication through the comments with the whistleblower.
* The Spanish transposition law of the Directive (EU) 2019/1937 of the European Parliament and of the Council, prescribes that the data provided by the whistleblower can be kept in the information system only for the time necessary to decide on the appropriateness of starting an investigation (article 32.3 Ley 2/2023, de 20 de febrero). | open | 2023-09-25T06:57:18Z | 2023-09-27T11:45:34Z | https://github.com/globaleaks/globaleaks-whistleblowing-software/issues/3654 | [
"T: Feature"
] | jowis | 5 |
junyanz/pytorch-CycleGAN-and-pix2pix | pytorch | 1,113 | exporting Pix2Pix to onnx | Hello,
PyTorch complains that I used DataParallel to train the model and that it can't be exported because of it
so I have to remove that info somehow. But I can't figure out how to do it
I tried this [workaround](https://stackoverflow.com/questions/44230907/keyerror-unexpected-key-module-encoder-embedding-weight-in-state-dict) :
I'm using a modified test.py script in Google Colab
```
import os
from options.test_options import TestOptions
from data import create_dataset
from models import create_model
from util.visualizer import save_images
from util import html
import torch
if name == 'main':
opt = TestOptions().parse() # get test options
# hard-code some parameters for test
opt.num_threads = 0 # test code only supports num_threads =
opt.batch_size = 1 # test code only supports batch_size = 1
opt.serial_batches = True # disable data shuffling; comment this line if results on randomly chosen images are needed.
opt.no_flip = True # no flip; comment this line if results on flipped images are needed.
opt.display_id = -1 # no visdom display; the test code saves the results to a HTML file.
dataset = create_dataset(opt) # create a dataset given opt.dataset_mode and other options
model = create_model(opt) # create a model given opt.model and other options
model.setup(opt) # regular setup: load and print networks; create schedulers
# original saved file with DataParallel
state_dict = torch.load('/content/drive/My Drive/Training Data/checkpoints/human2cat_pix2pix/latest_net_G.pth')
# create new OrderedDict that does not contain module.
from collections import OrderedDict
new_state_dict = OrderedDict()
for k, v in state_dict.items():
name = k[7:] # remove module.
new_state_dict[name] = v
# load params
model.netG.load_state_dict(new_state_dict)
dummy = torch.randn(10, 3, 256, 256)
torch.onnx.export(model.netG, dummy, './out.onnx')
```
But I get `RuntimeError: Error(s) in loading state_dict for DataParallel:`
Do you have any suggestions for how to export to ONNB with a model trained with DataParallel?
thanks | open | 2020-08-03T10:01:22Z | 2022-09-16T08:21:58Z | https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/issues/1113 | [] | ReallyRad | 4 |
nerfstudio-project/nerfstudio | computer-vision | 3,184 | License of the gsplat | can you share the lic of the gsplat model ? | closed | 2024-05-31T03:49:02Z | 2024-05-31T05:47:27Z | https://github.com/nerfstudio-project/nerfstudio/issues/3184 | [] | sumanttyagi | 4 |
newpanjing/simpleui | django | 63 | 与django-import-export集成的时候遇到的兼容性问题 | **bug描述**
简单的描述下遇到的bug:
按照simpleui_demo的用了django_import_export,然后导入导出的图标跟过滤搜索的这些重叠了

建议是否可以增加配置把过滤搜索这些框下降一行吗?
**重现步骤**
1.
2.
3.
**环境**
Django==2.2.1
django-import-export==1.2.0
django-simpleui==2.1
**其他描述**
| closed | 2019-05-24T07:27:50Z | 2020-03-08T03:06:09Z | https://github.com/newpanjing/simpleui/issues/63 | [
"bug"
] | pandadriver | 4 |
awesto/django-shop | django | 112 | Bug in template samples | The current templates do not reflect the ManyToMany model between categories and products.
Is there an other way to submit patches, rather than Email?
diff --git a/shop/templates/shop/product_detail.html b/shop/templates/shop/product_detail.html
index 70d4ade..323b061 100644
--- a/shop/templates/shop/product_detail.html
+++ b/shop/templates/shop/product_detail.html
@@ -10,8 +10,8 @@
{{object.unit_price}}<br />
-{% if object.category %}
-{{object.category.name}}
+{% if object.categories %}
+{% for cat in object.categories.all %} {{ cat.name }} {% endfor %}
{% else %}
(Product is at root category)
{% endif %}
diff --git a/shop/templates/shop/product_list.html b/shop/templates/shop/product_list.html
index c7314c0..6439c39 100644
--- a/shop/templates/shop/product_list.html
+++ b/shop/templates/shop/product_list.html
@@ -13,8 +13,8 @@
{{object.unit_price}}<br />
-{% if object.category %}
-{{object.category.name}}<br />
+{% if object.categories %}
+{% for cat in object.categories.all %} {{ cat.name }}<br /> {% endfor %}
{% else %}
(Product is at root category)<br />
{% endif %}
| closed | 2011-11-01T10:00:21Z | 2016-02-02T14:09:11Z | https://github.com/awesto/django-shop/issues/112 | [] | jrief | 2 |
FactoryBoy/factory_boy | django | 465 | Model returned from .create() doesn't have an id | I might be missing something, but the docs say that `create()` returns a saved model, but if I simply do `UserFactory.create().id` I get back `None`, yet if I do `user = UserFactory.create(); user.save()` then the `user.id` is actually set. | closed | 2018-04-05T00:01:04Z | 2018-05-05T00:07:49Z | https://github.com/FactoryBoy/factory_boy/issues/465 | [
"Q&A"
] | darthdeus | 3 |
Sanster/IOPaint | pytorch | 348 | 1 Click Installer : AttributeError: 'LaMa' object has no attribute 'is_local_sd_model' | I was using the cleaner fine but when I tried to boot it up today it throws this error on all models. Ran config to see if there were any updates but no luck. Some help would be appreciated.
```
[2023-07-18 17:13:08,790] ERROR in app: Exception on /inpaint [POST]
Traceback (most recent call last):
File "S:\lama-cleaner\installer\lib\site-packages\flask\app.py", line 2528, in wsgi_app
response = self.full_dispatch_request()
File "S:\lama-cleaner\installer\lib\site-packages\flask\app.py", line 1825, in full_dispatch_request
rv = self.handle_user_exception(e)
File "S:\lama-cleaner\installer\lib\site-packages\flask_cors\extension.py", line 176, in wrapped_function
return cors_after_request(app.make_response(f(*args, **kwargs)))
File "S:\lama-cleaner\installer\lib\site-packages\flask\app.py", line 1823, in full_dispatch_request
rv = self.dispatch_request()
File "S:\lama-cleaner\installer\lib\site-packages\flask\app.py", line 1799, in dispatch_request
return self.ensure_sync(self.view_functions[rule.endpoint])(**view_args)
File "S:\lama-cleaner\installer\lib\site-packages\lama_cleaner\server.py", line 291, in process
res_np_img = model(image, mask, config)
File "S:\lama-cleaner\installer\lib\site-packages\lama_cleaner\model_manager.py", line 63, in __call__
self.switch_controlnet_method(control_method=config.controlnet_method)
File "S:\lama-cleaner\installer\lib\site-packages\lama_cleaner\model_manager.py", line 88, in switch_controlnet_method
if self.model.is_local_sd_model:
AttributeError: 'LaMa' object has no attribute 'is_local_sd_model'
127.0.0.1 - - [18/Jul/2023 17:13:08] "POST /inpaint HTTP/1.1" 500 -
``` | closed | 2023-07-18T07:15:20Z | 2023-07-18T13:45:53Z | https://github.com/Sanster/IOPaint/issues/348 | [] | Acephalia | 1 |
benlubas/molten-nvim | jupyter | 260 | [Feature Request] Text-Objects for jupytext "py:percent" format | [Jupytext](https://jupytext.readthedocs.io/en/latest/index.html) is a versatile tool for converting between Jupyter notebooks (`.ipynb`) and Python scripts (`.py`) and vice versa. It provides support for various formats, including the ["percent" format](https://jupytext.readthedocs.io/en/latest/formats-scripts.html#the-percent-format), which adds notebook cell metadata as comments in Python scripts.
It would be useful if `molten.nvim` included a custom text object to represent Jupyter notebook cells in Python scripts using this "percent" format.
#### Use Case
For example, consider the following Python script converted from a Jupyter notebook with three cells:
```python
# %% [markdown] # ┓
# This is a multiline. # ┠ first cell (markdown type)
# Markdown cell # ┃
# ┛
# %% [markdown] # ┓
# Another Markdown cell # ┠ second cell (markdown type)
# ┃
# ┛
# %% # ┓
# This is a code cell # ┠ third cell (code type)
class A(): # ┃
def one(): # ┃
return 1 # ┃
# ┃
def two(): # ┃
return 2 # ┛
```
The custom text object would identify and operate on these cell structures. This would allow users to:
1. Navigate between Jupyter cells easily (e.g., move to the next/previous cell).
2. Evaluate cells individually or sequentially using `MoltenEvaluateOperator`.
3. Bind familiar key mappings like `Shift-Enter` or `Ctrl-Enter` to run the current Jupyter cell and move to the next one, emulating the experience of running cells in a Jupyter notebook.
#### Proposed Solution
- Implement a custom text object to recognize Jupyter cells in Python scripts using the "percent" format.
- Add support for evaluating these cells through `MoltenEvaluateOperator`.
- Optionally, provide default key mappings for running and navigating cells (`Shift-Enter`/`Ctrl-Enter`).
#### Benefits
This feature would enhance the experience of working with Python scripts derived from Jupyter notebooks. | closed | 2024-12-04T19:05:56Z | 2024-12-04T22:16:47Z | https://github.com/benlubas/molten-nvim/issues/260 | [
"enhancement"
] | S1M0N38 | 1 |
jupyter/nbgrader | jupyter | 1,326 | assignment_dir issues | The `c.Exchange.assignment_dir` config setting is not behaving as expected when fetching assignments through the web interface (assignment list). We have a Jupyterhub setup where the notebooks of the users are placed in `~/Jupyter`:
in `jupyterhub_config.py`:
`c.Spawner.notebook_dir = '~/Jupyter'`
Additionally, we use following settings in `nbgrader_config.py` of the normal users:
```
c = get_config()
c.CourseDirectory.course_id = "somecourse"
c.Exchange.path_includes_course = True
```
### not setting `c.Exchange.assignment_dir`
When fetching assignments though the web interface, the files are placed in the home directory of users, instead of in `~/Jupyter`
e.g. `~/somecourse/ps1` instead of `~/Jupyter/somecourse/ps1`
### setting `c.Exchange.assignment_dir` to a the `Spawner.notebook_dir`
Files are placed in `~/Jupyter/Jupyter/somecourse/ps1` rather than `~/Jupyter/somecourse/ps1`
### setting `c.Exchange.assignment_dir` to a relative path
e.g. `c.Exchange.assignment_dir = 'foo'`
Files are placed in `~/foo/ps1` instead of `~/Jupyter/foo`
### setting `c.Exchange.assignment_dir` to an absolute path
We initially solved the issue by setting
```c.Exchange.assignment_dir = os.path.expanduser("~/Jupyter")```
This places files in `~/Jupyter/somecourse/ps1` as expected.
However, this introduces a bug with the `(view feedback)` links. They now link to:
`https://hostname/jupyter/user/someuser/tree/home/someuser/Jupyter/somecourse/ps1/feedback/2020-04-14%2018:01:49.392273%20UTC`
Note the absolute path following `tree`
Where we should have
`https://hostname/jupyter/user/someuser/tree/somecourse/ps1/feedback/2020-04-14%2018:01:49.392273%20UTC`
(removal of `home/someuser/Jupyter/`)
Note this does not happen with the links to the notebooks within a fetched course. The paths used to generate the links are relative in:
```
course_id: "somecourse"
assignment_id: "ps1"
status: "fetched"
path: "somecourse/ps1"
notebooks: [
{notebook_id: "problem1", path: "somecourse/ps1/problem1.ipynb"},
…]
```
The paths listed for feedback are absolute in `local_feedback_path: /home/someuser/Jupyter/somecourse/ps1/feedback/...`
### Recap
I think there are two issues: 1) inconsistencies in how `c.Exchange.assignment_dir` is handled, and 2) the path to feedback files should be handled the same way the path to notebooks are handled in the assignment list (see also https://github.com/jupyter/nbgrader/issues/1317 ).
### `nbgrader --version`
```
Python version 3.7.3 (default, Jun 25 2019, 16:36:57)
[GCC 5.5.0]
nbgrader version 0.6.1
```
### `jupyterhub --version` (if used with JupyterHub)
```
1.0.0
```
### `jupyter notebook --version`
```
5.7.8
```
| open | 2020-04-15T10:50:53Z | 2020-04-15T12:58:20Z | https://github.com/jupyter/nbgrader/issues/1326 | [] | bomma | 0 |
pandas-dev/pandas | python | 60,560 | BUG: inconsistent return types from __getitem__ vs iteration | ### Pandas version checks
- [X] I have checked that this issue has not already been reported.
- [X] I have confirmed this bug exists on the [latest version](https://pandas.pydata.org/docs/whatsnew/index.html) of pandas.
- [X] I have confirmed this bug exists on the [main branch](https://pandas.pydata.org/docs/dev/getting_started/install.html#installing-the-development-version-of-pandas) of pandas.
### Reproducible Example
```python
import numpy as np
import pandas as pd
print(np.__version__) # 2.0.2
print(pd.__version__) # 2.2.3
data = pd.Series([333, 555])
# accessing scalar via __getitem__ returns <class 'numpy.int64'>
print(type(data[0]))
# accessing scalar via iteration returns <class 'int'>
print(type(next(iter(data))))
```
### Issue Description
numpy 2.0 recently changed its [representation of scalars](https://numpy.org/devdocs/release/2.0.0-notes.html#representation-of-numpy-scalars-changed) to include type information. However, pandas produces inconsistent return types when one is accessing scalars with `__getitem__` vs iterating over items, as demonstrated in the example code snippet.
This inconsistency is showing up in downstream projects like NetworkX: https://github.com/networkx/networkx/issues/7763#issuecomment-2532716537
### Expected Behavior
pandas should produce consistent return types when one is accessing scalars with `__getitem__` vs iterating over items
### Installed Versions
<details>
INSTALLED VERSIONS
------------------
commit : 0691c5cf90477d3503834d983f69350f250a6ff7
python : 3.12.8
python-bits : 64
OS : Linux
OS-release : 6.8.0-50-generic
Version : #51-Ubuntu SMP PREEMPT_DYNAMIC Sat Nov 9 17:58:29 UTC 2024
machine : x86_64
processor : x86_64
byteorder : little
LC_ALL : None
LANG : en_US.UTF-8
LOCALE : en_US.UTF-8
pandas : 2.2.3
numpy : 2.0.2
pytz : 2024.1
dateutil : 2.9.0.post0
pip : 24.3.1
Cython : None
sphinx : 8.1.3
IPython : 8.30.0
adbc-driver-postgresql: None
adbc-driver-sqlite : None
bs4 : 4.12.3
blosc : None
bottleneck : 1.4.2
dataframe-api-compat : None
fastparquet : None
fsspec : None
html5lib : None
hypothesis : None
gcsfs : None
jinja2 : 3.1.4
lxml.etree : 5.3.0
matplotlib : 3.9.3
numba : 0.60.0
numexpr : 2.10.2
odfpy : None
openpyxl : None
pandas_gbq : None
psycopg2 : None
pymysql : None
pyarrow : None
pyreadstat : None
pytest : 8.3.4
python-calamine : None
pyxlsb : None
s3fs : None
scipy : 1.14.1
sqlalchemy : None
tables : None
tabulate : None
xarray : None
xlrd : None
xlsxwriter : None
zstandard : 0.23.0
tzdata : 2024.2
qtpy : None
pyqt5 : None
</details>
| open | 2024-12-13T16:17:51Z | 2025-02-11T23:54:01Z | https://github.com/pandas-dev/pandas/issues/60560 | [
"Bug",
"Needs Discussion",
"API - Consistency"
] | gboeing | 6 |
tortoise/tortoise-orm | asyncio | 1,242 | Correct way to override model constructor | Hello. Apologies if I overlooked something but I have combed the documentation trying to figure out a working technique for overriding the instance `__init__`/constructor method (i.e. to set up some non-db-backed instance vars etc, after the instance is initialized - whether when being pulled out of the db or from direct instantiation). I tried overriding `__init__` (as it is defined with the signature [here](https://github.com/tortoise/tortoise-orm/blob/db9c36cd5e4257f6cecd5488a1de8f915b329dd4/tortoise/models.py#L663)) and, calling `super.__init__` at the top of the overriding method body, but does not seem to ever get called. I may be missing something. If it helps any to clarify I'm either looking for how to do this override directly or if there is a callback etc (something like the equivalent of Rails/ActiveRecord's `after_initialize` callback). Thank you 🙏 | open | 2022-09-05T13:42:00Z | 2022-09-05T13:42:00Z | https://github.com/tortoise/tortoise-orm/issues/1242 | [] | AlgoDev1 | 0 |
encode/databases | sqlalchemy | 269 | Password containing digits and hashmark cannot be used (MySQL) | `databases==0.4.1`
If using a URL like:
`mysql://user_name:Xx7#4xxXX77xx@localhost/db_name`
I get the following error:
```
ERROR: Traceback (most recent call last):
File "/home/rkrell/work/lib/python3.7/site-packages/starlette/routing.py", line 526, in lifespan
async for item in self.lifespan_context(app):
File "/home/rkrell/work/lib/python3.7/site-packages/starlette/routing.py", line 467, in default_lifespan
await self.startup()
File "/home/rkrell/work/lib/python3.7/site-packages/starlette/routing.py", line 502, in startup
await handler()
File "./main.py", line 154, in startup
await database.connect()
File "/home/rkrell/work/lib/python3.7/site-packages/databases/core.py", line 84, in connect
await self._backend.connect()
File "/home/rkrell/work/lib/python3.7/site-packages/databases/backends/mysql.py", line 63, in connect
port=self._database_url.port or 3306,
File "/home/rkrell/work/lib/python3.7/site-packages/databases/core.py", line 448, in port
return self.components.port
File "/usr/lib/python3.7/urllib/parse.py", line 169, in port
port = int(port, 10)
ValueError: invalid literal for int() with base 10: 'Xx7'
``` | closed | 2020-11-27T11:38:08Z | 2020-11-30T20:29:48Z | https://github.com/encode/databases/issues/269 | [] | rkrell | 2 |
pyro-ppl/numpyro | numpy | 1,567 | Incorrect batch shape of low_rank normal | <img width="1156" alt="Screenshot 2023-03-28 at 15 55 18" src="https://user-images.githubusercontent.com/26022201/228352296-1eaa5d43-325b-438c-90af-e4635ed08fee.png">
The LowRankNormal has a batch shape of (5,), is this expected? I think it should have a batch shape of (,), just like the MultivariateNormal case.
```python
D = 5
K = 2
W_shape = (D, K)
W = jnp.ones(W_shape)
loc_x = jnp.zeros(D)
cov_diag = jnp.eye(D)
cov_mat = W @ W.T + cov_diag
dis = dist.LowRankMultivariateNormal(loc=loc_x, cov_factor=W, cov_diag=cov_diag)
print(dis.event_shape)
print(dis.batch_shape)
dis = dist.MultivariateNormal(covariance_matrix=cov_mat)
print(dis.event_shape)
print(dis.batch_shape)
``` | closed | 2023-03-28T19:58:33Z | 2023-03-28T20:48:34Z | https://github.com/pyro-ppl/numpyro/issues/1567 | [
"question"
] | xidulu | 2 |
junyanz/pytorch-CycleGAN-and-pix2pix | deep-learning | 1,082 | Template Dataset Doubt | Hello,
I am working with cycle gans ,on grayscale images.
I changed the channels to 1 and wrote the custom dataloader. My data is stored as tensors i.e as .pt files after being preprocessed and not into the training and testing folders in any directory.
I have split the training A and training B dataset in my dataloader and want to pass it to the model.
Can you tell me which lines I will have to change? | closed | 2020-06-26T20:36:04Z | 2020-06-28T20:45:23Z | https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/issues/1082 | [] | SurbhiKhushu | 7 |
Gerapy/Gerapy | django | 110 | 不能在 /tmp 目录下找到生成的egg包 | 两台环境一样的机器,其中一台不能正常的跑起来。在eggs/content_spider下已经生成了对应的egg包,但是运行的时候出现如下错误。
File "/tmp/content_spider-1561777690-PvDfIJ.egg/content_spider/pipelines.py", line 38, in __init__
IOError: [Errno 2] No such file or directory。 | open | 2019-06-29T03:31:54Z | 2019-06-29T03:31:54Z | https://github.com/Gerapy/Gerapy/issues/110 | [] | iamdaguduizhang | 0 |
ets-labs/python-dependency-injector | asyncio | 655 | Container dependencies issue, multiple files, classes | Traceback (most recent call last):
File "<frozen importlib._bootstrap>", line 1178, in _find_and_load
File "<frozen importlib._bootstrap>", line 1149, in _find_and_load_unlocked
File "<frozen importlib._bootstrap>", line 690, in _load_unlocked
File "<frozen importlib._bootstrap_external>", line 940, in exec_module
File "<frozen importlib._bootstrap>", line 241, in _call_with_frames_removed
File "D:\Projects\python\user-test\user\route\user_route.py", line 17, in <module>
@user_router.get(
^^^^^^^^^^^^^^^^
File "C:\Users\Eugene\AppData\Local\Programs\Python\Python311\Lib\site-packages\fastapi\routing.py", line 630, in decorator
self.add_api_route(
File "C:\Users\Eugene\AppData\Local\Programs\Python\Python311\Lib\site-packages\fastapi\routing.py", line 569, in add_api_route
route = route_class(
^^^^^^^^^^^^
File "C:\Users\Eugene\AppData\Local\Programs\Python\Python311\Lib\site-packages\fastapi\routing.py", line 442, in __init__
get_parameterless_sub_dependant(depends=depends, path=self.path_format),
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\Eugene\AppData\Local\Programs\Python\Python311\Lib\site-packages\fastapi\dependencies\utils.py", line 135, in get_parameterless_sub_dependant
return get_sub_dependant(depends=depends, dependency=depends.dependency, path=path)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\Eugene\AppData\Local\Programs\Python\Python311\Lib\site-packages\fastapi\dependencies\utils.py", line 158, in get_sub_dependant
sub_dependant = get_dependant(
^^^^^^^^^^^^^^
File "C:\Users\Eugene\AppData\Local\Programs\Python\Python311\Lib\site-packages\fastapi\dependencies\utils.py", line 281, in get_dependant
endpoint_signature = get_typed_signature(call)
^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\Eugene\AppData\Local\Programs\Python\Python311\Lib\site-packages\fastapi\dependencies\utils.py", line 249, in get_typed_signature
signature = inspect.signature(call)
^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\Eugene\AppData\Local\Programs\Python\Python311\Lib\inspect.py", line 3278, in signature
return Signature.from_callable(obj, follow_wrapped=follow_wrapped,
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\Eugene\AppData\Local\Programs\Python\Python311\Lib\inspect.py", line 3026, in from_callable
return _signature_from_callable(obj, sigcls=cls,
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\Eugene\AppData\Local\Programs\Python\Python311\Lib\inspect.py", line 2615, in _signature_from_callable
raise ValueError('callable {!r} is not supported by signature'.format(obj))
ValueError: callable <dependency_injector.providers.Factory(<class 'user.infrastructure.query.user_uow.UserUnitOfWork'>) at 0x27549c2cc40> is not supported by signature
python-BaseException | open | 2022-12-29T13:37:27Z | 2022-12-29T13:38:47Z | https://github.com/ets-labs/python-dependency-injector/issues/655 | [] | Spenchik | 1 |
mars-project/mars | numpy | 2,934 | [BUG] mars shuffle function not well-distributed | <!--
Thank you for your contribution!
Please review https://github.com/mars-project/mars/blob/master/CONTRIBUTING.rst before opening an issue.
-->
**Describe the bug**
Groupby shuffle keys for different groups are not well-distributed. In a online case which has 10000_0000 lines and chunk size is 20_0000, some gorups has about 24000 keys, but most groups has less than 5000 keys. The overall pecess is dominated by large keys group, and the execution is 5 times slower than expected.
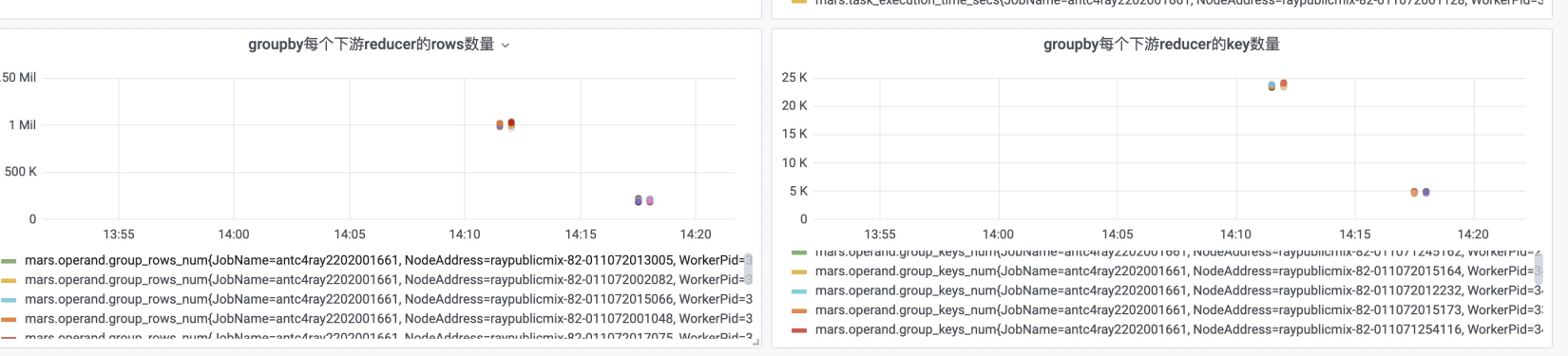
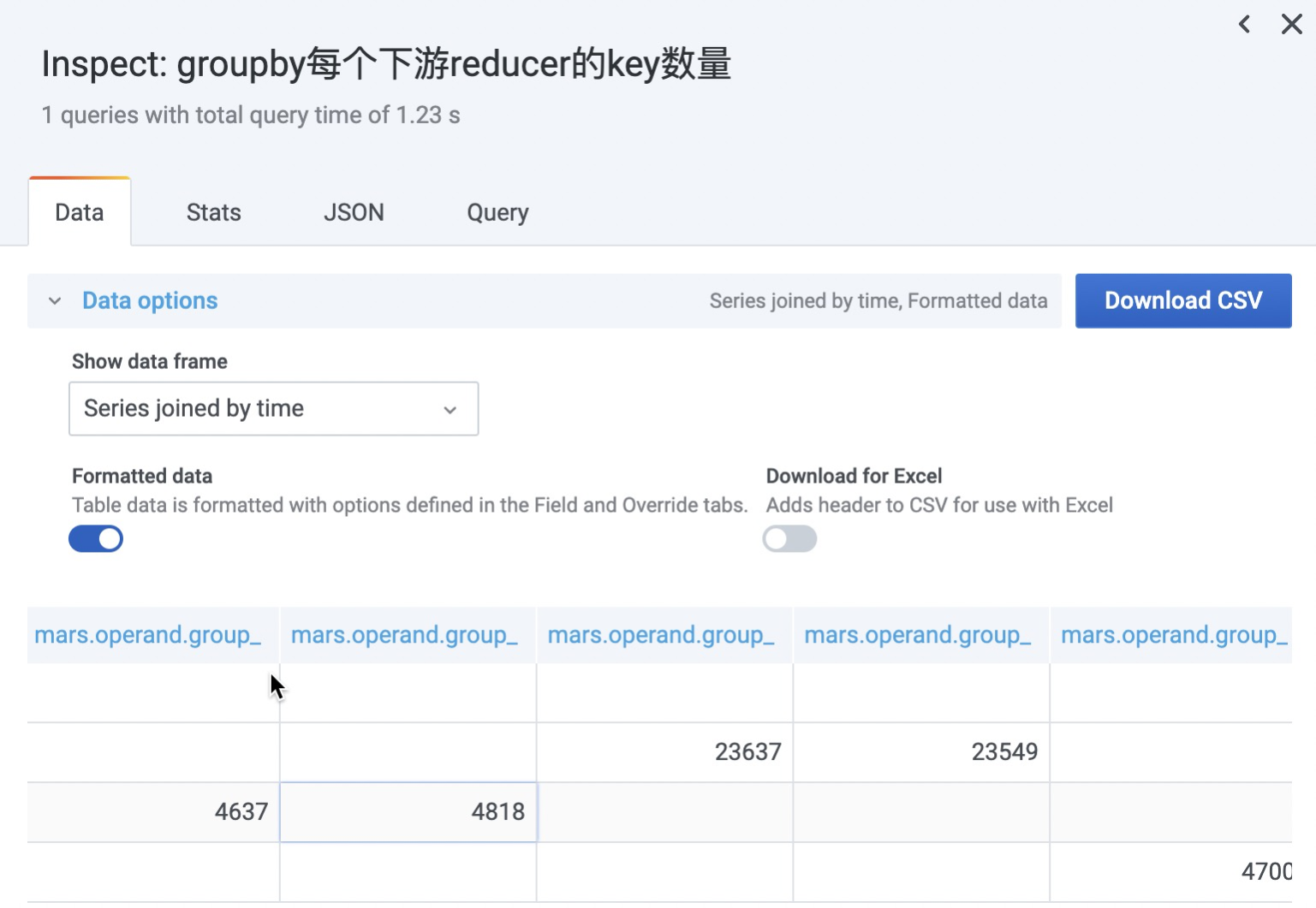
**To Reproduce**
To help us reproducing this bug, please provide information below:
1. Your Python version: 3.7
2. The version of Mars you use: master
3. Versions of crucial packages, such as numpy, scipy and pandas
4. Full stack of the error.
5. Minimized code to reproduce the error.
**Expected behavior**
The keys should be well-distributed. This is not a data skew. For data skew, some key groups will have much more data thant other group, but the issue is that some chunks has much more keys than other chunks.
| open | 2022-04-19T07:47:47Z | 2022-04-19T07:47:47Z | https://github.com/mars-project/mars/issues/2934 | [] | chaokunyang | 0 |
AirtestProject/Airtest | automation | 422 | check_app检测不到app时报错,而不是返回False | **描述问题bug**
check_app文档对返回值的描述是:
`True or False whether the package exists on the device or not`
实际上,找不到app时会抛出异常,而不会返回False。
```
File "G:\Workspace\PyCharm\AirtestSign\venv\lib\site-packages\airtest\core\android\android.py", line 105, in check_app
return self.adb.check_app(package)
File "G:\Workspace\PyCharm\AirtestSign\venv\lib\site-packages\airtest\core\android\adb.py", line 1169, in check_app
raise AirtestError('package "{}" not found'.format(package))
airtest.core.error.AirtestError: 'package "com.xxx.yyy" not found'
```
**期待结果**
android.py中的实现应该捕捉adb.py里的异常,然后返回False,而不是直接return调用
**复现步骤**
调用check_app检测一个设备上没安装的应用。
**python 版本:** `python3.5`
**airtest 版本:** `1.0.26`
| closed | 2019-06-11T15:55:09Z | 2019-06-13T02:45:29Z | https://github.com/AirtestProject/Airtest/issues/422 | [] | WalkerMe | 3 |
yuka-friends/Windrecorder | streamlit | 92 | bug: 当磁盘上不存在对应视频文件时,“一日之时”无法正确处理异常 | ```
File "C:\Users\Anton\AppData\Local\pypoetry\Cache\virtualenvs\windrecorder-QcaNLmW7-py3.10\lib\site-packages\streamlit\runtime\scriptrunner\script_runner.py", line 534, in _run_script
exec(code, module.__dict__)
File "D:\git\Windrecorder\webui.py", line 88, in <module>
windrecorder.ui.oneday.render()
File "D:\git\Windrecorder\windrecorder\ui\oneday.py", line 398, in render
show_and_locate_video_timestamp_by_filename_and_time(day_video_file_name, shown_timestamp)
File "D:\git\Windrecorder\windrecorder\ui\oneday.py", line 500, in show_and_locate_video_timestamp_by_filename_and_time
video_file = open(videofile_path, "rb")
```

| closed | 2024-01-04T17:15:22Z | 2024-01-05T14:25:17Z | https://github.com/yuka-friends/Windrecorder/issues/92 | [
"bug",
"P2"
] | Antonoko | 2 |
mlflow/mlflow | machine-learning | 14,546 | [BUG] LangGraph MemorySaver checkpointer usage with MLflow | ### Issues Policy acknowledgement
- [x] I have read and agree to submit bug reports in accordance with the [issues policy](https://www.github.com/mlflow/mlflow/blob/master/ISSUE_POLICY.md)
### Where did you encounter this bug?
Databricks
### MLflow version
- MLflow version: 2.18
### System information
- **OS Platform and Distribution (e.g., Linux Ubuntu 16.04)**: Databricks DBR 16.0 ML
- **Python version**: 3.12.3
### Describe the problem
Hi everyone.
I am working on a graph that utilizes the MemorySaver class to incorporate short-term memory. This will enable me to maintain a multi-turn conversation with the user by storing the chat history.
I am using the MLflow "models from code" feature but I'm getting an error because when the model is invoked it requires the config parameter with a thread_id:
```
ValueError("Checkpointer requires one or more of the following 'configurable' keys: ['thread_id', 'checkpoint_ns', 'checkpoint_id']")Traceback (most recent call last)
```
The graph compilation is:
```
# Compile
memory = MemorySaver()
graph = graph_builder.compile(checkpointer=memory)
```
How to register a LangGraph graph in MLflow that uses the MemorySaver to store the chat history in the short-term memory?
Thanks!
### Tracking information
<!-- PLEASE KEEP BACKTICKS AND CHECK PREVIEW -->
```shell
REPLACE_ME
```
### Code to reproduce issue
<!-- PLEASE KEEP BACKTICKS AND CHECK PREVIEW -->
```
REPLACE_ME
```
### Stack trace
<!-- PLEASE KEEP BACKTICKS AND CHECK PREVIEW -->
```
REPLACE_ME
```
### Other info / logs
<!-- PLEASE KEEP BACKTICKS AND CHECK PREVIEW -->
```
REPLACE_ME
```
### What component(s) does this bug affect?
- [ ] `area/artifacts`: Artifact stores and artifact logging
- [ ] `area/build`: Build and test infrastructure for MLflow
- [x] `area/deployments`: MLflow Deployments client APIs, server, and third-party Deployments integrations
- [ ] `area/docs`: MLflow documentation pages
- [ ] `area/examples`: Example code
- [x] `area/model-registry`: Model Registry service, APIs, and the fluent client calls for Model Registry
- [x] `area/models`: MLmodel format, model serialization/deserialization, flavors
- [ ] `area/recipes`: Recipes, Recipe APIs, Recipe configs, Recipe Templates
- [ ] `area/projects`: MLproject format, project running backends
- [ ] `area/scoring`: MLflow Model server, model deployment tools, Spark UDFs
- [ ] `area/server-infra`: MLflow Tracking server backend
- [ ] `area/tracking`: Tracking Service, tracking client APIs, autologging
### What interface(s) does this bug affect?
- [ ] `area/uiux`: Front-end, user experience, plotting, JavaScript, JavaScript dev server
- [ ] `area/docker`: Docker use across MLflow's components, such as MLflow Projects and MLflow Models
- [ ] `area/sqlalchemy`: Use of SQLAlchemy in the Tracking Service or Model Registry
- [ ] `area/windows`: Windows support
### What language(s) does this bug affect?
- [ ] `language/r`: R APIs and clients
- [ ] `language/java`: Java APIs and clients
- [ ] `language/new`: Proposals for new client languages
### What integration(s) does this bug affect?
- [ ] `integrations/azure`: Azure and Azure ML integrations
- [ ] `integrations/sagemaker`: SageMaker integrations
- [x] `integrations/databricks`: Databricks integrations | closed | 2025-02-11T21:58:34Z | 2025-02-17T00:03:23Z | https://github.com/mlflow/mlflow/issues/14546 | [
"bug",
"area/model-registry",
"area/models",
"integrations/databricks",
"area/deployments"
] | scardonal | 7 |
ultralytics/yolov5 | pytorch | 13,390 | training "Memory Error" on Window | ### Search before asking
- [X] I have searched the YOLOv5 [issues](https://github.com/ultralytics/yolov5/issues) and found no similar bug report.
### YOLOv5 Component
_No response_
### Bug
I tried to run the training app of yours on my Window machine. it has just loaded some stuffs, moving stuffs around, a few caches... then it crashed.
`(yolo) C:\Users\baoth\OneDrive\Desktop\yolo\yolov5>python train.py --epochs 10 --img 640 --batch 16 --data ../data.yaml --weights yolov5s.pt
train: weights=yolov5s.pt, cfg=, data=../data.yaml, hyp=data\hyps\hyp.scratch-low.yaml, epochs=10, batch_size=16, imgsz=640, rect=False, resume=False, nosave=Fal
se, noval=False, noautoanchor=False, noplots=False, evolve=None, evolve_population=data\hyps, resume_evolve=None, bucket=, cache=None, image_weights=False, devic
e=, multi_scale=False, single_cls=False, optimizer=SGD, sync_bn=False, workers=8, project=runs\train, name=exp, exist_ok=False, quad=False, cos_lr=False, label_s
moothing=0.0, patience=100, freeze=[0], save_period=-1, seed=0, local_rank=-1, entity=None, upload_dataset=False, bbox_interval=-1, artifact_alias=latest, ndjson_console=False, ndjson_file=False
github: up to date with https://github.com/ultralytics/yolov5
YOLOv5 v7.0-378-g2f74455a Python-3.12.4 torch-2.5.0+cpu CPU
hyperparameters: lr0=0.01, lrf=0.01, momentum=0.937, weight_decay=0.0005, warmup_epochs=3.0, warmup_momentum=0.8, warmup_bias_lr=0.1, box=0.05, cls=0.5, cls_pw=1
.0, obj=1.0, obj_pw=1.0, iou_t=0.2, anchor_t=4.0, fl_gamma=0.0, hsv_h=0.015, hsv_s=0.7, hsv_v=0.4, degrees=0.0, translate=0.1, scale=0.5, shear=0.0, perspective=0.0, flipud=0.0, fliplr=0.5, mosaic=1.0, mixup=0.0, copy_paste=0.0
Comet: run 'pip install comet_ml' to automatically track and visualize YOLOv5 runs in Comet
TensorBoard: Start with 'tensorboard --logdir runs\train', view at http://localhost:6006/
Overriding model.yaml nc=80 with nc=3
from n params module arguments
0 -1 1 3520 models.common.Conv [3, 32, 6, 2, 2]
1 -1 1 18560 models.common.Conv [32, 64, 3, 2]
2 -1 1 18816 models.common.C3 [64, 64, 1]
3 -1 1 73984 models.common.Conv [64, 128, 3, 2]
4 -1 2 115712 models.common.C3 [128, 128, 2]
5 -1 1 295424 models.common.Conv [128, 256, 3, 2]
6 -1 3 625152 models.common.C3 [256, 256, 3]
7 -1 1 1180672 models.common.Conv [256, 512, 3, 2]
8 -1 1 1182720 models.common.C3 [512, 512, 1]
9 -1 1 656896 models.common.SPPF [512, 512, 5]
10 -1 1 131584 models.common.Conv [512, 256, 1, 1]
11 -1 1 0 torch.nn.modules.upsampling.Upsample [None, 2, 'nearest']
12 [-1, 6] 1 0 models.common.Concat [1]
13 -1 1 361984 models.common.C3 [512, 256, 1, False]
14 -1 1 33024 models.common.Conv [256, 128, 1, 1]
15 -1 1 0 torch.nn.modules.upsampling.Upsample [None, 2, 'nearest']
16 [-1, 4] 1 0 models.common.Concat [1]
17 -1 1 90880 models.common.C3 [256, 128, 1, False]
18 -1 1 147712 models.common.Conv [128, 128, 3, 2]
19 [-1, 14] 1 0 models.common.Concat [1]
20 -1 1 296448 models.common.C3 [256, 256, 1, False]
21 -1 1 590336 models.common.Conv [256, 256, 3, 2]
22 [-1, 10] 1 0 models.common.Concat [1]
23 -1 1 1182720 models.common.C3 [512, 512, 1, False]
24 [17, 20, 23] 1 21576 models.yolo.Detect [3, [[10, 13, 16, 30, 33, 23], [30, 61, 62, 45, 59, 119], [116, 90, 156, 198, 373, 326]], [128, 256, 512]]
Model summary: 214 layers, 7027720 parameters, 7027720 gradients, 16.0 GFLOPs
Transferred 343/349 items from yolov5s.pt
optimizer: SGD(lr=0.01) with parameter groups 57 weight(decay=0.0), 60 weight(decay=0.0005), 60 bias
train: Scanning C:\Users\baoth\OneDrive\Desktop\yolo\train\labels.cache... 996 images, 0 backgrounds, 0 corrupt: 100%|██████████| 996/996 [00:00<?, ?it/s]
val: Scanning C:\Users\baoth\OneDrive\Desktop\yolo\valid\labels.cache... 61 images, 0 backgrounds, 0 corrupt: 100%|██████████| 61/61 [00:00<?, ?it/s]
Traceback (most recent call last):
File "<string>", line 1, in <module>
File "C:\Users\baoth\miniconda3\Lib\multiprocessing\spawn.py", line 122, in spawn_main
exitcode = _main(fd, parent_sentinel)
^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\baoth\miniconda3\Lib\multiprocessing\spawn.py", line 131, in _main
prepare(preparation_data)
File "C:\Users\baoth\miniconda3\Lib\multiprocessing\spawn.py", line 246, in prepare
_fixup_main_from_path(data['init_main_from_path'])
File "C:\Users\baoth\miniconda3\Lib\multiprocessing\spawn.py", line 297, in _fixup_main_from_path
main_content = runpy.run_path(main_path,
^^^^^^^^^^^^^^^^^^^^^^^^^
File "<frozen runpy>", line 286, in run_path
File "<frozen runpy>", line 98, in _run_module_code
File "<frozen runpy>", line 88, in _run_code
File "C:\Users\baoth\OneDrive\Desktop\yolo\yolov5\train.py", line 47, in <module>
import val as validate # for end-of-epoch mAP
^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\baoth\OneDrive\Desktop\yolo\yolov5\val.py", line 60, in <module>
from utils.plots import output_to_target, plot_images, plot_val_study
File "C:\Users\baoth\OneDrive\Desktop\yolo\yolov5\utils\plots.py", line 15, in <module>
import seaborn as sn
File "C:\Users\baoth\miniconda3\Lib\site-packages\seaborn\__init__.py", line 7, in <module>
from .categorical import * # noqa: F401,F403
^^^^^^^^^^^^^^^^^^^^^^^^^^
File "C:\Users\baoth\miniconda3\Lib\site-packages\seaborn\categorical.py", line 19, in <module>
from seaborn._stats.density import KDE
File "C:\Users\baoth\miniconda3\Lib\site-packages\seaborn\_stats\density.py", line 10, in <module>
from scipy.stats import gaussian_kde
File "C:\Users\baoth\miniconda3\Lib\site-packages\scipy\stats\__init__.py", line 610, in <module>
from ._stats_py import *
File "<frozen importlib._bootstrap>", line 1360, in _find_and_load
File "<frozen importlib._bootstrap>", line 1331, in _find_and_load_unlocked
File "<frozen importlib._bootstrap>", line 935, in _load_unlocked
File "<frozen importlib._bootstrap_external>", line 991, in exec_module
File "<frozen importlib._bootstrap_external>", line 1087, in get_code
File "<frozen importlib._bootstrap_external>", line 1187, in get_data
MemoryError
`
### Environment
yolov5s, Window, no cuda
### Minimal Reproducible Example
_No response_
### Additional
_No response_
### Are you willing to submit a PR?
- [ ] Yes I'd like to help by submitting a PR! | open | 2024-10-28T17:41:57Z | 2024-11-09T13:10:21Z | https://github.com/ultralytics/yolov5/issues/13390 | [
"bug"
] | suws0501 | 2 |
jupyter-widgets-contrib/ipycanvas | jupyter | 188 | Dynamic MultiCanvas | Thank you for this library!
I have a couple of questions regarding `MultiCanvas` objects.
1. Is it safe to dynamically add canvases to a `MultiCanvas` object?
2. Is this bad for performance? More specifically, is there a point beyond which performance regresses - say 10 canvases vs. 100 canvases?
```python
from ipycanvas import MultiCanvas, Canvas
# (1)
canvas = MultiCanvas(n_canvases=1, width=100, height=100)
canvas._canvases.append(Canvas(width=100, height=100))
``` | open | 2021-04-11T13:47:36Z | 2022-04-07T13:07:31Z | https://github.com/jupyter-widgets-contrib/ipycanvas/issues/188 | [
"enhancement"
] | rsomani95 | 4 |
tiangolo/uwsgi-nginx-flask-docker | flask | 86 | Container runs Python 2.7 instead of Python 3.6 | i have an app structured as follow:
```
app
app
main.py
Dockerfile
uwsgi.ini
docker-compose.yml
```
Dockerfile:
```
FROM tiangolo/uwsgi-nginx-flask:python3.6
RUN pip3 install gensim pymongo pandas numpy
```
docker-compose.yml:
```
version: '2'
services:
s_a:
build: ./app
links:
- s_db
volumes:
- ./app/app:/app
ports:
- "8080:80"
s_db:
image: mongo
ports:
- "27370:27017"
volumes:
- ./app/mongodb:/data/db
```
uwsgi.ini:
```
[uwsgi]
module = app.main
callable = app
master = true
processes = 10
```
when up `docker-compose.yml` i get the error:

I don't know why the runs on Python 2.7 instead of Python 3.6.
Where is the problem ? | closed | 2018-09-25T12:15:43Z | 2018-10-14T20:36:48Z | https://github.com/tiangolo/uwsgi-nginx-flask-docker/issues/86 | [] | pymooner | 5 |
graphql-python/graphene | graphql | 1,298 | Question on using argument in Graphene | i am trying to pass size argument to graphene along with object mapping and it gives error
```
try:
import graphene
import json
except Exception as e:
print("Error : {} ".format(e))
global DATA
DATA = [
{
"name":"Soumil",
"age":23,
"language" : ["Python", "c++"]
},
{
"name":"Seymur",
"age":27,
"language" : ["Python", "c++"]
},
{
"name":"Test",
"age":23,
"language" : ["Python", "c++"]
}
]
class Person(graphene.ObjectType):
name = graphene.String()
age = graphene.Int()
language = graphene.List(graphene.String)
class Users(graphene.ObjectType):
users = graphene.List(Person, size=graphene.Int(default_value=2))
def resolve_users(root, info):
return DATA
schema = graphene.Schema(query=Users)
#print(schema)
# ====================== Query 1 ==================================
query_string1 = '''
query {
users (size : 2) {
name,
age
}
}
'''
result = schema.execute(query_string1)
print(json.dumps(result.data, indent=3))
# =================================================================
``` | closed | 2021-01-06T00:40:27Z | 2021-01-06T11:16:49Z | https://github.com/graphql-python/graphene/issues/1298 | [] | soumilshah1995 | 2 |
JaidedAI/EasyOCR | deep-learning | 771 | Dont want detection. Only want Recogntion | I do not want to perform detection first, then recognition. I want to perform recognition straight on the image i pass.
Code:
```
self.ocr = easyocr.Reader(
["en"],
gpu=False,
detector=False,
quantize=True,
recognizer=True,
)
return self.ocr.readtext(path_to_img)
```
But I get this error
```
File "/home/ahmad/Desktop/FYP/venv/lib/python3.7/site-packages/easyocr/easyocr.py", line 397, in readtext
add_margin, False)
File "/home/ahmad/Desktop/FYP/venv/lib/python3.7/site-packages/easyocr/easyocr.py", line 279, in detect
text_box_list = get_textbox(self.detector, img, canvas_size, mag_ratio,
AttributeError: 'Reader' object has no attribute 'detector'
```
``` | open | 2022-07-03T17:34:16Z | 2024-05-06T02:56:18Z | https://github.com/JaidedAI/EasyOCR/issues/771 | [] | ahmadmustafaanis | 5 |
jupyter-incubator/sparkmagic | jupyter | 885 | Support notebook >= 7 | #825 highlighted a problem where `notebook >= 7.0.0` causes `sparkmagic` installation to fail due to removed `jupyter-nbextension` command.
A [workaround](https://github.com/jupyter-incubator/sparkmagic/blob/6eab8aadfa3c61a6247868836b2a8df086e1b649/Dockerfile.jupyter#L33-L36) has been added to the Docker image that downgrades `notebook` version to `6.x.x`.
This ticket to provide a long-term solution and remove the workaround. | closed | 2024-02-21T03:31:34Z | 2024-12-15T01:58:47Z | https://github.com/jupyter-incubator/sparkmagic/issues/885 | [
"kind:bug"
] | sergiimk | 2 |
zappa/Zappa | django | 900 | [Migrated] Remote function invocation does not need quotes around function | Originally from: https://github.com/Miserlou/Zappa/issues/2162 by [LaundroMat](https://github.com/LaundroMat)
To invoke your function remotely, the docs say:
zappa invoke production 'my_app.my_function'
But for me (Windows 10, python3.8, Zappa0.51.0), this returns an error
[ERROR] ModuleNotFoundError: No module named "'my_app"
Note the single quote... To invoke the function remotely, do not use the single quotes and do this instead:
zappa invoke production my_app.my_function
| closed | 2021-02-20T13:03:31Z | 2022-08-05T10:36:44Z | https://github.com/zappa/Zappa/issues/900 | [] | jneves | 1 |
iperov/DeepFaceLab | deep-learning | 803 | Step 4: take the picture out of the picture and make a mistake | [wf] Face type ( f/wf/head ?:help ) :
wf
[0] Max number of faces from image ( ?:help ) :
0
[512] Image size ( 256-2048 ?:help ) :
512
[90] Jpeg quality ( 1-100 ?:help ) :
90
[n] Write debug images to aligned_debug? ( y/n ) : n
Extracting faces...
Traceback (most recent call last):
File "C:\DeepFaceLab_NVIDIA\_internal\DeepFaceLab\main.py", line 324, in <module>
arguments.func(arguments)
File "C:\DeepFaceLab_NVIDIA\_internal\DeepFaceLab\main.py", line 45, in process_extract
force_gpu_idxs = [ int(x) for x in arguments.force_gpu_idxs.split(',') ] if arguments.force_gpu_idxs is not None else None,
File "C:\DeepFaceLab_NVIDIA\_internal\DeepFaceLab\mainscripts\Extractor.py", line 840, in main
device_config=device_config).run()
File "C:\DeepFaceLab_NVIDIA\_internal\DeepFaceLab\core\joblib\SubprocessorBase.py", line 199, in run
raise Exception ("Unable to start Subprocessor '%s' " % (self.name))
Exception: Unable to start Subprocessor 'Extractor'

| open | 2020-07-01T14:55:39Z | 2023-06-08T23:20:15Z | https://github.com/iperov/DeepFaceLab/issues/803 | [] | shiranII | 5 |
iMerica/dj-rest-auth | rest-api | 162 | Tests failing in master with no new *source code changes. | Looks like it might be related to Django-All-Auth | closed | 2020-11-03T01:48:03Z | 2020-11-11T17:00:33Z | https://github.com/iMerica/dj-rest-auth/issues/162 | [] | iMerica | 0 |
WZMIAOMIAO/deep-learning-for-image-processing | pytorch | 703 | hi | 计算损失时
for name, x in inputs.items:
报错:
AttributeError: 'Tensor' object has no attribute 'items'
请问这个是什么原因导致的呢!
| closed | 2022-12-01T05:43:33Z | 2022-12-03T05:20:49Z | https://github.com/WZMIAOMIAO/deep-learning-for-image-processing/issues/703 | [] | 7788fine | 1 |
scikit-learn/scikit-learn | machine-learning | 30,222 | Changelog check on towncrier false positive case | Observed on this PR: https://github.com/scikit-learn/scikit-learn/pull/30209
This run: https://github.com/scikit-learn/scikit-learn/actions/runs/11681055082/job/32525320042?pr=30209
The PR needs to add PR number to existing changelog, and changes another affected changelog, therefore there are 3 changelog files affected in the PR. However, the changelog checker complains with:
```
Not all changelog file number(s) match this pull request number (30209):
doc/whats_new/upcoming_changes/sklearn.calibration/30171.api.rst
doc/whats_new/upcoming_changes/sklearn.frozen/29705.major-feature.rst
doc/whats_new/upcoming_changes/sklearn.frozen/30209.major-feature.rst
```
Which I'd say is a false positive.
cc @lesteve | open | 2024-11-05T09:28:21Z | 2024-11-18T10:14:55Z | https://github.com/scikit-learn/scikit-learn/issues/30222 | [
"Bug",
"Build / CI"
] | adrinjalali | 1 |
tflearn/tflearn | tensorflow | 1,171 | OSS License compatibility question | There’s some possible confusion on the license of your repository when you combine other open-source code.
The module `tflearn/vendor/arg_scope.py` claims its license as **Apache-2.0**. However, the license of your whole project is shown as **the MIT license** in LICENSE, i.e., less strict than Apache-2.0 on license terms, which has impacted the whole license compatibility in your repository and may bring legal and financial risks.
You can select another proper license for your repository, or write a custom license with license exceptions if some license terms couldn’t be summed up consistently
| open | 2023-01-14T05:33:02Z | 2023-01-14T05:33:02Z | https://github.com/tflearn/tflearn/issues/1171 | [] | Ashley123456789 | 0 |
gradio-app/gradio | data-visualization | 10,747 | Initial states saved in ClearButton can be corrupted | ### Describe the bug
ClearButton saves the initial value of State component and then can use it to clear the current value of State.
It works perfectly, but there are some situations when this saved initial value can be corrupted and ClearButton stops working
Below is a simple example of such a situation:
1) This demo just shows one number which can be increased by 1 or can be reset to 0
2) What's wrong with this demo? After you press the ClearButton once, it will stop reseting number to 0
3) Why does it happen? When ClearButton initializes, it deep-copies the init value of State and there are 2 different value objects (1 belongs to State and 1 belongs to ClearButton). But when ClearButton resets the value of State, it returns its own saved value without deepcopying. So there is only 1 value object (common to ClearButton and to State), and that's the problem.
4) increase_button changes the value of State by reference. Therefore, when there is only 1 value object, increase_button gets the opportunity to change the init value saved in ClearButton :(
### Have you searched existing issues? 🔎
- [x] I have searched and found no existing issues
### Reproduction
```python
import gradio as gr
def increase_number(x):
x[0] += 1
with gr.Blocks() as demo:
state = gr.State([0])
text = gr.Textbox(lambda x: x[0], inputs=state)
increase_button = gr.Button('Increase number by 1').click(increase_number, inputs=state).then(lambda x: x[0], inputs=state, outputs=text)
gr.ClearButton(state).click(lambda x: print(id(x)), inputs=state)
if __name__ == "__main__":
demo.launch()
```
### Screenshot
_No response_
### Logs
```shell
```
### System Info
```shell
Gradio Environment Information:
------------------------------
Operating System: Darwin
gradio version: 5.20.0
gradio_client version: 1.7.2
------------------------------------------------
gradio dependencies in your environment:
aiofiles: 23.2.1
anyio: 4.8.0
audioop-lts is not installed.
fastapi: 0.115.8
ffmpy: 0.5.0
gradio-client==1.7.2 is not installed.
groovy: 0.1.2
httpx: 0.28.1
huggingface-hub: 0.28.1
jinja2: 3.1.5
markupsafe: 2.1.5
numpy: 1.26.4
orjson: 3.10.15
packaging: 24.2
pandas: 2.2.3
pillow: 11.1.0
pydantic: 2.10.6
pydub: 0.25.1
python-multipart: 0.0.20
pyyaml: 6.0.2
ruff: 0.9.6
safehttpx: 0.1.6
semantic-version: 2.10.0
starlette: 0.45.3
tomlkit: 0.13.2
typer: 0.15.1
typing-extensions: 4.12.2
urllib3: 2.3.0
uvicorn: 0.34.0
authlib; extra == 'oauth' is not installed.
itsdangerous; extra == 'oauth' is not installed.
gradio_client dependencies in your environment:
fsspec: 2025.2.0
httpx: 0.28.1
huggingface-hub: 0.28.1
packaging: 24.2
typing-extensions: 4.12.2
websockets: 14.2
```
### Severity
I can work around it | closed | 2025-03-06T14:24:14Z | 2025-03-06T21:52:46Z | https://github.com/gradio-app/gradio/issues/10747 | [
"bug"
] | phos-phophy | 0 |
kevlened/pytest-parallel | pytest | 114 | Fatal Python error: _enter_buffered_busy: could not acquire lock for <_io.BufferedWriter name=5> at interpreter shutdown, possibly due to daemon threads | Hello, while trying this library on [Gradio ](https://github.com/gradio-app/gradio)project, I encountered this error, will share the reproduction below.
commit 98242fe3632c20511300ac63b774290e4fdf8313
```
➜ gradio git:(queue-refactor-backend) ✗ pytest --tests-per-worker 5 test/test_event_queue.py
================================================================================================================== test session starts ===================================================================================================================
platform win32 -- Python 3.9.10, pytest-7.0.0, pluggy-1.0.0
rootdir: F:\SecondaryDownloads\git_repos\gradio
plugins: anyio-3.5.0, asyncio-0.18.3, cov-3.0.0, parallel-0.1.1
asyncio: mode=legacy
collected 1 item
pytest-parallel: 1 worker (process), 1 test per worker (thread)
INTERNALERROR> Traceback (most recent call last):
INTERNALERROR> File "f:\secondarydownloads\git_repos\gradio\venv\lib\site-packages\_pytest\main.py", line 268, in wrap_session
INTERNALERROR> session.exitstatus = doit(config, session) or 0
INTERNALERROR> File "f:\secondarydownloads\git_repos\gradio\venv\lib\site-packages\_pytest\main.py", line 322, in _main
INTERNALERROR> config.hook.pytest_runtestloop(session=session)
INTERNALERROR> File "f:\secondarydownloads\git_repos\gradio\venv\lib\site-packages\pluggy\_hooks.py", line 265, in __call__
INTERNALERROR> return self._hookexec(self.name, self.get_hookimpls(), kwargs, firstresult)
INTERNALERROR> File "f:\secondarydownloads\git_repos\gradio\venv\lib\site-packages\pluggy\_manager.py", line 80, in _hookexec
INTERNALERROR> return self._inner_hookexec(hook_name, methods, kwargs, firstresult)
INTERNALERROR> File "f:\secondarydownloads\git_repos\gradio\venv\lib\site-packages\pluggy\_callers.py", line 60, in _multicall
INTERNALERROR> return outcome.get_result()
INTERNALERROR> File "f:\secondarydownloads\git_repos\gradio\venv\lib\site-packages\pluggy\_result.py", line 60, in get_result
INTERNALERROR> raise ex[1].with_traceback(ex[2])
INTERNALERROR> File "f:\secondarydownloads\git_repos\gradio\venv\lib\site-packages\pluggy\_callers.py", line 39, in _multicall
INTERNALERROR> res = hook_impl.function(*args)
INTERNALERROR> File "f:\secondarydownloads\git_repos\gradio\venv\lib\site-packages\pytest_parallel\__init__.py", line 313, in pytest_runtestloop
INTERNALERROR> process.start()
INTERNALERROR> File "C:\Program Files\WindowsApps\PythonSoftwareFoundation.Python.3.9_3.9.2800.0_x64__qbz5n2kfra8p0\lib\multiprocessing\process.py", line 121, in start
INTERNALERROR> self._popen = self._Popen(self)
INTERNALERROR> File "C:\Program Files\WindowsApps\PythonSoftwareFoundation.Python.3.9_3.9.2800.0_x64__qbz5n2kfra8p0\lib\multiprocessing\context.py", line 224, in _Popen
INTERNALERROR> return _default_context.get_context().Process._Popen(process_obj)
INTERNALERROR> File "C:\Program Files\WindowsApps\PythonSoftwareFoundation.Python.3.9_3.9.2800.0_x64__qbz5n2kfra8p0\lib\multiprocessing\context.py", line 327, in _Popen
INTERNALERROR> return Popen(process_obj)
INTERNALERROR> File "C:\Program Files\WindowsApps\PythonSoftwareFoundation.Python.3.9_3.9.2800.0_x64__qbz5n2kfra8p0\lib\multiprocessing\popen_spawn_win32.py", line 93, in __init__
INTERNALERROR> reduction.dump(process_obj, to_child)
INTERNALERROR> File "C:\Program Files\WindowsApps\PythonSoftwareFoundation.Python.3.9_3.9.2800.0_x64__qbz5n2kfra8p0\lib\multiprocessing\reduction.py", line 60, in dump
INTERNALERROR> ForkingPickler(file, protocol).dump(obj)
INTERNALERROR> AttributeError: Can't pickle local object 'ArgumentParser.__init__.<locals>.identity'
================================================================================================================== 2 warnings in 2.62s ===================================================================================================================
Exception in thread Thread-1:
Traceback (most recent call last):
File "C:\Program Files\WindowsApps\PythonSoftwareFoundation.Python.3.9_3.9.2800.0_x64__qbz5n2kfra8p0\lib\multiprocessing\connection.py", line 317, in _recv_bytes
nread, err = ov.GetOverlappedResult(True)
BrokenPipeError: [WinError 109] The pipe has been ended
During handling of the above exception, another exception occurred:
Traceback (most recent call last):
File "C:\Program Files\WindowsApps\PythonSoftwareFoundation.Python.3.9_3.9.2800.0_x64__qbz5n2kfra8p0\lib\threading.py", line 973, in _bootstrap_inner
self.run()
File "C:\Program Files\WindowsApps\PythonSoftwareFoundation.Python.3.9_3.9.2800.0_x64__qbz5n2kfra8p0\lib\threading.py", line 910, in run
self._target(*self._args, **self._kwargs)
File "f:\secondarydownloads\git_repos\gradio\venv\lib\site-packages\pytest_parallel\__init__.py", line 359, in process_responses
event_name, kwargs = queue.get()
Traceback (most recent call last):
File "<string>", line 1, in <module>
File "C:\Program Files\WindowsApps\PythonSoftwareFoundation.Python.3.9_3.9.2800.0_x64__qbz5n2kfra8p0\lib\multiprocessing\spawn.py", line 107, in spawn_main
new_handle = reduction.duplicate(pipe_handle,
File "C:\Program Files\WindowsApps\PythonSoftwareFoundation.Python.3.9_3.9.2800.0_x64__qbz5n2kfra8p0\lib\multiprocessing\reduction.py", line 79, in duplicate
return _winapi.DuplicateHandle(
OSError: [WinError 6] The handle is invalid
Fatal Python error: _enter_buffered_busy: could not acquire lock for <_io.BufferedWriter name=5> at interpreter shutdown, possibly due to daemon threads
Python runtime state: finalizing (tstate=0000022F10A55240)
Current thread 0x000049a4 (most recent call first):
<no Python frame>
``` | open | 2022-07-05T09:26:45Z | 2022-07-05T09:26:45Z | https://github.com/kevlened/pytest-parallel/issues/114 | [] | omerXfaruq | 0 |
matplotlib/matplotlib | matplotlib | 29,275 | [Bug]: clip_on=False dosen't work | ### Bug summary
i create ax as the log area, when i use divider to create a axLin as linear area , the point in axlin range can display the whole scatter, but the point in ax range, clip_on=False seems dosen't work!

### Code for reproduction
```Python
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1 import make_axes_locatable
fig, ax = plt.subplots(figsize=(11.69, 8.27))
ax.set_xscale('log')
ax.set_yscale('log')
ax.set_xlim(1, 10000) # x 轴范围
ax.set_ylim(10, 1000) # y 轴范围
ax.plot([2000], [999], color='black', linestyle='-'
, marker='o', markersize=2, linewidth=0.5, clip_on=False)
divider = make_axes_locatable(ax)
axLin = divider.append_axes("top", size='44.8%', pad=0, sharex=ax)
axLin.set_yscale('linear')
axLin.set_ylim(1000,1007)
axLin.plot([1000], [1000.001], color='black', linestyle='-'
, marker='o', markersize=2, linewidth=0.5, clip_on=False)
plt.savefig('plot1.svg', format='svg')
plt.show()
```
### Actual outcome

### Expected outcome

### Additional information
_No response_
### Operating system
windows10
### Matplotlib Version
3.9.2
### Matplotlib Backend
_No response_
### Python version
_No response_
### Jupyter version
_No response_
### Installation
None | closed | 2024-12-10T10:31:19Z | 2024-12-19T15:18:20Z | https://github.com/matplotlib/matplotlib/issues/29275 | [
"Community support"
] | thomaslilu | 5 |
axnsan12/drf-yasg | rest-api | 61 | ReDoc failed to render this spec | if i add `'USE_SESSION_AUTH': False` or `'SHOW_REQUEST_HEADERS': True` to `SWAGGER_SETTINGS` i get page that said
> Oops... ReDoc failed to render this spec can't assign to property "_displayType" on false: not an object
just want to ask, is this something that should not appear or its normal? | closed | 2018-02-18T16:33:53Z | 2018-02-18T21:03:38Z | https://github.com/axnsan12/drf-yasg/issues/61 | [] | DimasInchidi | 4 |
PaddlePaddle/ERNIE | nlp | 195 | 我想用你们训练好的模型参数来训练我们的文章数据拿到词向量做检索 召回相似的文章 现在怎么拿到经过ERNIE 编码后Embedding对应的句子呢 | 已经通过 ernie_encoder.py 抽取出到输入句子的 Embedding 表示并做了向量检索,现在拿不到向量对应的句子表示 | closed | 2019-07-09T10:53:21Z | 2019-07-10T08:00:57Z | https://github.com/PaddlePaddle/ERNIE/issues/195 | [] | qq1074123922 | 7 |
charlesq34/pointnet | tensorflow | 303 | How to visualize the semantic segmentation results through ROS | Hi, thank you again for sharing your work. I have successfully trained and tested, however i would like to know how i could possible visualize the semantic segmentation on ROS like the teaser you have shown.
Eager to hear from you soon.
Best,
Rohith. | closed | 2022-09-21T14:13:17Z | 2022-10-20T14:25:32Z | https://github.com/charlesq34/pointnet/issues/303 | [] | rohithsaro | 0 |
flasgger/flasgger | rest-api | 13 | Fix UI style | Change the UI style to not break the header

| closed | 2016-01-11T12:55:09Z | 2017-03-24T20:06:46Z | https://github.com/flasgger/flasgger/issues/13 | [] | rochacbruno | 0 |
NVIDIA/pix2pixHD | computer-vision | 102 | Training with VGG feature loss is quite slower than without using it? or I miss something? | Training with VGG feature loss is quite slower than without using it? or I miss something? Thank you in advance! | open | 2019-02-21T20:07:38Z | 2019-04-28T07:08:24Z | https://github.com/NVIDIA/pix2pixHD/issues/102 | [] | happsky | 1 |
junyanz/pytorch-CycleGAN-and-pix2pix | pytorch | 1,172 | Can I execute test.py in other computer without a GPU? | I am wondering If I got well trained model to a computer that didn't have GPU, is there a way I can do ?
I want use the model in other computer that don't have a GPU, the model is trained by my other computer. | closed | 2020-10-28T12:58:40Z | 2022-08-19T06:30:32Z | https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/issues/1172 | [] | darrenleeleelee1 | 1 |
piskvorky/gensim | machine-learning | 3,340 | ldaseqmodel convergence | <!--
**IMPORTANT**:
- Use the [Gensim mailing list](https://groups.google.com/forum/#!forum/gensim) to ask general or usage questions. Github issues are only for bug reports.
- Check [Recipes&FAQ](https://github.com/RaRe-Technologies/gensim/wiki/Recipes-&-FAQ) first for common answers.
Github bug reports that do not include relevant information and context will be closed without an answer. Thanks!
-->
#### Problem description
https://github.com/RaRe-Technologies/gensim/blob/742fb188dc6de03a42411510bf5b45e26574b328/gensim/models/ldaseqmodel.py#L303
This line in `ldaseqmodel.py` seems preventing the early termination of the algorithm. Set the `convergence` to 1 whenever the convergence criterion is met makes it must exhaust the `em_max_iter` hence cannot terminate earlier.
#### Versions
Please provide the output of:
```python
import platform; print(platform.platform())
import sys; print("Python", sys.version)
import struct; print("Bits", 8 * struct.calcsize("P"))
import numpy; print("NumPy", numpy.__version__)
import scipy; print("SciPy", scipy.__version__)
import gensim; print("gensim", gensim.__version__)
from gensim.models import word2vec;print("FAST_VERSION", word2vec.FAST_VERSION)
```
gensim version 4.1.2 | open | 2022-04-29T01:12:11Z | 2022-04-29T01:12:11Z | https://github.com/piskvorky/gensim/issues/3340 | [] | trkwyk | 0 |
tensorflow/tensor2tensor | machine-learning | 1,016 | Unable to download translate_ende_wmt32k using t2t-datagen | ### Description
Downloading dataset "translate_ende_wmt32k" with t2t-datagen results in following error.
tensorflow.python.framework.errors_impl.NotFoundError: /tmp/t2t_datagen/training/news-commentary-v13.de-en.en; No such file or directory
I did not have this issue while downloading translate_ende_wmt_bpe32k.
PROBLEM=translate_ende_wmt32k
MODEL=transformer
HPARAMS=transformer_base_single_gpu
t2t-datagen --data_dir=$DATA_DIR --tmp_dir=$TMP_DIR --problem=$PROBLEM
100% completed
Traceback (most recent call last):
File "/home/prashant/.local/bin/t2t-datagen", line 27, in <module>
tf.app.run()
File "/home/prashant/.local/lib/python2.7/site-packages/tensorflow/python/platform/app.py", line 125, in run
_sys.exit(main(argv))
File "/home/prashant/.local/bin/t2t-datagen", line 23, in main
t2t_datagen.main(argv)
File "/home/prashant/.local/lib/python2.7/site-packages/tensor2tensor/bin/t2t_datagen.py", line 190, in main
generate_data_for_registered_problem(problem)
File "/home/prashant/.local/lib/python2.7/site-packages/tensor2tensor/bin/t2t_datagen.py", line 240, in generate_data_for_registered_problem
problem.generate_data(data_dir, tmp_dir, task_id)
File "/home/prashant/.local/lib/python2.7/site-packages/tensor2tensor/data_generators/text_problems.py", line 294, in generate_data
self.generate_encoded_samples(data_dir, tmp_dir, split)), paths)
File "/home/prashant/.local/lib/python2.7/site-packages/tensor2tensor/data_generators/text_problems.py", line 254, in generate_encoded_samples
generator = self.generate_samples(data_dir, tmp_dir, dataset_split)
File "/home/prashant/.local/lib/python2.7/site-packages/tensor2tensor/data_generators/translate.py", line 55, in generate_samples
tag))
File "/home/prashant/.local/lib/python2.7/site-packages/tensor2tensor/data_generators/translate.py", line 148, in compile_data
lang1_filepath, lang2_filepath):
File "/home/prashant/.local/lib/python2.7/site-packages/tensor2tensor/data_generators/text_problems.py", line 552, in text2text_txt_iterator
txt_line_iterator(source_txt_path), txt_line_iterator(target_txt_path)):
File "/home/prashant/.local/lib/python2.7/site-packages/tensor2tensor/data_generators/text_problems.py", line 545, in txt_line_iterator
for line in f:
File "/home/prashant/.local/lib/python2.7/site-packages/tensorflow/python/lib/io/file_io.py", line 214, in next
retval = self.readline()
File "/home/prashant/.local/lib/python2.7/site-packages/tensorflow/python/lib/io/file_io.py", line 183, in readline
self._preread_check()
File "/home/prashant/.local/lib/python2.7/site-packages/tensorflow/python/lib/io/file_io.py", line 85, in _preread_check
compat.as_bytes(self.__name), 1024 * 512, status)
File "/home/prashant/.local/lib/python2.7/site-packages/tensorflow/python/framework/errors_impl.py", line 519, in __exit__
c_api.TF_GetCode(self.status.status))
tensorflow.python.framework.errors_impl.NotFoundError: /tmp/t2t_datagen/training/news-commentary-v13.de-en.en; No such file or directory
$TMP_DIR has downloaded files, but the path is different than what script is looking for.
ls -l /tmp/t2t_datagen
total 110512
drwxrwxr-x 2 prashant prashant 4096 Feb 21 2018 training-parallel-nc-v13
-rw-rw-r-- 1 prashant prashant 113157482 Aug 23 14:17 training-parallel-nc-v13.tgz
ls -la /tmp/t2t_datagen/training-parallel-nc-v13
total 313256
drwxrwxr-x 2 prashant prashant 4096 Feb 21 2018 .
drwxrwxr-x 3 prashant prashant 4096 Aug 23 14:17 ..
-rw-r--r-- 1 prashant prashant 32894113 Feb 21 2018 news-commentary-v13.cs-en.cs
-rw-r--r-- 1 prashant prashant 29823721 Feb 21 2018 news-commentary-v13.cs-en.en
-rw-r--r-- 1 prashant prashant 48226262 Feb 21 2018 news-commentary-v13.de-en.de
-rw-r--r-- 1 prashant prashant 39610338 Feb 21 2018 news-commentary-v13.de-en.en
-rw-r--r-- 1 prashant prashant 34376953 Feb 21 2018 news-commentary-v13.ru-en.en
-rw-r--r-- 1 prashant prashant 69178183 Feb 21 2018 news-commentary-v13.ru-en.ru
-rw-r--r-- 1 prashant prashant 35525461 Feb 21 2018 news-commentary-v13.zh-en.en
-rw-r--r-- 1 prashant prashant 31113639 Feb 21 2018 news-commentary-v13.zh-en.zh
### Environment information
```
OS: Ubuntu 16.04.4
$ pip freeze | grep tensor
tensor2tensor==1.8.0
tensorboard==1.9.0
tensorflow==1.5.0
tensorflow-gpu==1.9.0
tensorflow-tensorboard==1.5.1
$ python -V
Python 2.7.12
### For bugs: reproduction and error logs
```
# Steps to reproduce:
t2t-datagen --data_dir=$DATA_DIR --tmp_dir=$TMP_DIR --problem=$PROBLEM
# Error logs:
...
```
| open | 2018-08-23T19:31:25Z | 2018-09-12T03:08:43Z | https://github.com/tensorflow/tensor2tensor/issues/1016 | [] | pksubbarao | 5 |
gradio-app/gradio | data-science | 10,481 | gr.ImageEditor does not support source="webcam" for direct image capture | ### Describe the bug
gr.ImageEditor does not support `sources="webcam"`, preventing direct image capture from a webcam. Although the webcam icon appears in the UI, clicking it does not activate the webcam or allow image capture.
Gradio Version: `5.13.2`
Browser: Google Chrome
If `gr.Image` is used with `sources="webcam"`, the webcam functions correctly, capturing images as expected. However, `gr.ImageEditor` does not seem to support this feature, requiring an additional step to transfer the image from `gr.Image` to `gr.ImageEditor`.
This limitation makes it less convenient for users who want to edit images directly after capturing them from a webcam. It would be beneficial if `gr.ImageEditor` could support source="webcam" natively.
### Have you searched existing issues? 🔎
- [x] I have searched and found no existing issues
### Reproduction
```python
import gradio as gr
image_input = gr.ImageEditor(sources=['webcam'])
```
### Screenshot

### Logs
```shell
None
```
### System Info
```shell
Gradio Version: `5.13.2`
```
### Severity
Blocking usage of gradio | open | 2025-02-01T12:25:18Z | 2025-03-07T23:58:29Z | https://github.com/gradio-app/gradio/issues/10481 | [
"bug",
"🖼️ ImageEditor"
] | kuri54 | 6 |
cobrateam/splinter | automation | 583 | How do I load page with disabling images? | I am doing this:
browser = Browser('firefox', profile_preferences=proxy_settings)
path = #Given the path
browser.visit(path)
soup = BeautifulSoup(browser.html, 'html.parser')
For faster performance, I want to load pages by disabling the images. I tried editing the "permissions.default.image" parameter to 2 in "about:config" of firefox, but it resets everytime firefox is under remote control.
TL;DR: How do I pass "permissions.default.image" in splinter.browser.visit() request?
Thanks | closed | 2018-01-30T19:02:21Z | 2018-09-26T09:01:47Z | https://github.com/cobrateam/splinter/issues/583 | [
"question"
] | bondeanikets | 2 |
mwaskom/seaborn | matplotlib | 2,811 | kdeplot log normalization | Is it possible to apply a log normalization to a bivariate density plot? I only see the `log_scale` parameter which can apply a log scale to either one or both of the variables, but I want to scale the density values. With Matplotlib.pyplot.hist2d there is a norm parameter to which I can pass `norm=matplotlib.colors.LogNorm()` to apply a log normalization. Is this functionality available for kdeplot? | closed | 2022-05-16T15:03:54Z | 2022-05-17T16:19:05Z | https://github.com/mwaskom/seaborn/issues/2811 | [] | witherscp | 2 |
indico/indico | sqlalchemy | 5,951 | [A11Y] "Skip access checks" checkbox not associated with label | **Describe the bug**
The checkbox has a visible label but it is not semantically linked to it.
**Screenshots**
<img width="267" alt="image" src="https://github.com/indico/indico/assets/65413/2e57bf05-0899-4dc1-a353-daf533d47f7f">
**Additional context**
- https://www.w3.org/WAI/WCAG21/Understanding/labels-or-instructions
| open | 2023-09-26T10:45:01Z | 2023-09-26T10:45:01Z | https://github.com/indico/indico/issues/5951 | [
"bug"
] | foxbunny | 0 |
vaexio/vaex | data-science | 1,978 | unable to open files | I tried to open files in hdf5 and pkl format but none of them worked. I alwas get the error ""OSError: Cannot open raw_data/24h/20190101_dl_raw.pkl nobody knows how to read it.".
What could be the reson for this? | closed | 2022-03-18T08:29:42Z | 2022-03-18T13:47:12Z | https://github.com/vaexio/vaex/issues/1978 | [] | janwyler | 4 |
globaleaks/globaleaks-whistleblowing-software | sqlalchemy | 3,075 | "Do not expose users names" not working after upgrading 4.4.3 to 4.4.4 | After upgrading a system it's showing up users given names even thou the tick box is marked "do not expose users names". Tried to switch the feature on and off, rebooted VM. | closed | 2021-10-25T14:01:59Z | 2021-10-26T09:14:34Z | https://github.com/globaleaks/globaleaks-whistleblowing-software/issues/3075 | [] | simohks | 1 |
matplotlib/mplfinance | matplotlib | 388 | Problem with Bollinger Bands | Hi, I'm having a problem with the upper Bollinger band being misaligned, and I suspect that it is because the upper band is being treated as a lower band when I graph it. When I print the dataframe, the values seem correct, but the graph is definitely incorrect.
Here are both bands together:
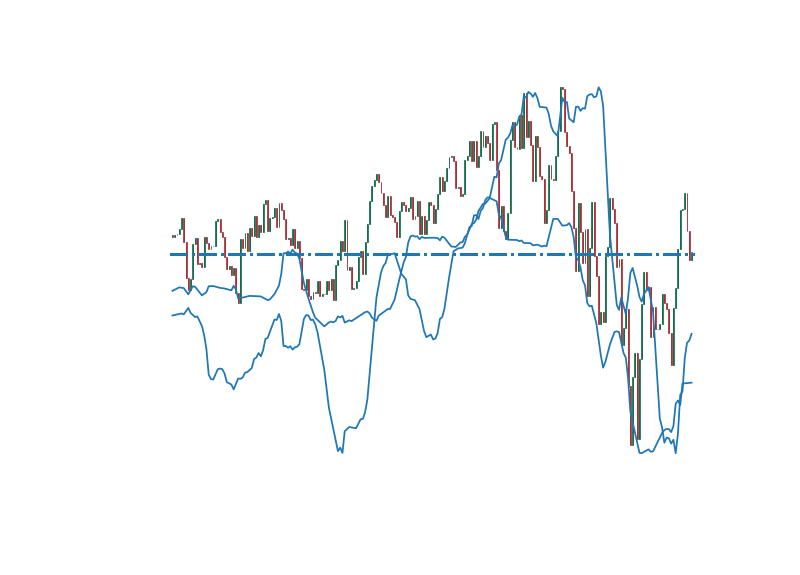
Here is the Lower Band (which seems correct):
Here is the Upper Band (which seems incorrect):
Here is the line of code that creates the graph:
```python
mpf.plot(df2, type='candle', style='charles', hlines=dict(hlines=[0],linestyle='-.'), axisoff=True,
addplot=mpf.make_addplot(df2[['UpperB', 'LowerB']]), savefig=newpath)
```
I'm not sure if this is a bug, or if I am just doing something wrong, but any help would be greatly appreciated. | closed | 2021-05-03T06:37:20Z | 2021-05-03T22:16:23Z | https://github.com/matplotlib/mplfinance/issues/388 | [
"question"
] | hedge0 | 2 |
graphql-python/graphene-django | graphql | 1,020 | GRAPHQL_SCHEMA is not included in the docs' testing example | **Note: for support questions, please use stackoverflow**. This repository's issues are reserved for feature requests and bug reports.
* **What is the current behavior?**
I was following [the docs' article on testing](https://docs.graphene-python.org/projects/django/en/latest/testing/) and noticed that the example code raises the following error:
`AttributeError: Variable GRAPHQL_SCHEMA not defined in GraphQLTestCase.`
* **If the current behavior is a bug, please provide the steps to reproduce and if possible a minimal demo of the problem** via
a github repo, https://repl.it or similar (you can use this template as a starting point: https://repl.it/@jkimbo/Graphene-Django-Example).
[Here is the code from docs](https://gist.github.com/karmek-k/e361a5896ee03c1c87f5dd062af69644).
Put it somewhere in a Django project with graphene-django installed and run `python3 manage.py test`.
* **What is the expected behavior?**
The tests should pass.
* **What is the motivation / use case for changing the behavior?**
It took me a while to investigate the problem. Hopefully this change may save others' time.
* **Please tell us about your environment:**
- Version: Python 3.8.3
- Platform: Manjaro Linux with KDE Plasma
* **Other information** (e.g. detailed explanation, stacktraces, related issues, suggestions how to fix, links for us to have context, eg. stackoverflow)
The article should mention the GRAPHQL_SCHEMA property and how to set it properly. | closed | 2020-08-07T20:07:27Z | 2023-06-11T19:11:41Z | https://github.com/graphql-python/graphene-django/issues/1020 | [
"🐛bug"
] | karmek-k | 2 |
Lightning-AI/LitServe | fastapi | 438 | Add `input_audio` Support to OpenAISpec Request | ## 🚀 Feature : Add `input_audio` Support to OpenAISpec Request
### Summary
Add support for `input_audio` in OpenAISpec to align with multimodal that accept audio inputs. This will extend the spec to handle audio data, making it more compatible with OpenIAI API.
Reference: [OpenAI API Reference](https://platform.openai.com/docs/api-reference/chat/create?lang=python)
<img width="623" alt="Image" src="https://github.com/user-attachments/assets/022615cd-0db5-4a3b-aa9f-3fe4abac9f82" />
### Motivation
As models like [Phi-4-multimodal-instruct](https://huggingface.co/microsoft/Phi-4-multimodal-instruct) support audio, adding this to OpenAISpec will keep it up-to-date and versatile, allowing seamless audio input handling.
| closed | 2025-02-27T07:28:41Z | 2025-02-27T11:26:41Z | https://github.com/Lightning-AI/LitServe/issues/438 | [
"enhancement"
] | bhimrazy | 0 |
jupyterlab/jupyter-ai | jupyter | 1,173 | Deleting messages breaks future replies | Testing the Jupyter AI 3.0.0.a0 prerelease and the behavior of deleting messages.
## Description
There are a couple of issues in how chat works when you delete messages.
First, create a new chat with Jupyter AI configured. It will reply as expected:

Now click the trashcan button in your message to delete it. This deletes both the human message and the AI message following it.

Now reload the page, and you will see that the human message remains deleted, but the AI message has returned:

Now, with a human message in the deleted state, try another chat message to the AI. Jupyter AI throws an exceptions in the chat windows:

The interesting pieces is the last line of the traceback:
```
botocore.exceptions.EventStreamError: An error occurred (validationException) when calling the InvokeModelWithResponseStream operation: messages.0: all messages must have non-empty content except for the optional final assistant message
```
## Expected behavior
* As a user, I want to delete AI messages separate from those that I type.
* As a user, if I delete an AI message, I want it to always be deleted in the future (across opening/closing the document or reloading the page).
* As a user, I want chat with AI to work, even when humans have deleted messages.
## Context
Using the following conda environment:
```
name: jupyter-ai-testing
dependencies:
- python
- pip:
- jupyterlab==4.2.5
- jupyter-ai==3.0.0a0
- langchain-aws
``` | open | 2024-12-30T17:50:49Z | 2024-12-30T21:18:08Z | https://github.com/jupyterlab/jupyter-ai/issues/1173 | [
"bug"
] | ellisonbg | 0 |
autogluon/autogluon | scikit-learn | 4,151 | I have already downloaded the CUDA version of torch, why does it automatically uninstall my CUDA when I pip install autogluon and turn it into the CPU version? | **Bug Report Checklist**
<!-- Please ensure at least one of the following to help the developers troubleshoot the problem: -->
- [ ] I provided code that demonstrates a minimal reproducible example. <!-- Ideal, especially via source install -->
- [ ] I confirmed bug exists on the latest mainline of AutoGluon via source install. <!-- Preferred -->
- [ ] I confirmed bug exists on the latest stable version of AutoGluon. <!-- Unnecessary if prior items are checked -->
**Describe the bug**
<!-- A clear and concise description of what the bug is. -->

**Expected behavior**
<!-- A clear and concise description of what you expected to happen. -->
**To Reproduce**
<!-- A minimal script to reproduce the issue. Links to Colab notebooks or similar tools are encouraged.
If the code is too long, feel free to put it in a public gist and link it in the issue: https://gist.github.com.
In short, we are going to copy-paste your code to run it and we expect to get the same result as you. -->
**Screenshots / Logs**
<!-- If applicable, add screenshots or logs to help explain your problem. -->
**Installed Versions**
<!-- Please run the following code snippet: -->
<details>
```python
# Replace this code with the output of the following:
from autogluon.core.utils import show_versions
show_versions()
```
</details>
| closed | 2024-04-30T07:38:41Z | 2024-05-02T07:45:39Z | https://github.com/autogluon/autogluon/issues/4151 | [
"bug: unconfirmed",
"Needs Triage"
] | psv666 | 1 |
kizniche/Mycodo | automation | 1,074 | Controllers and Inputs not functioning after reboot | STOP right now, and please first look to see if the issue you're about to submit is already an open or recently closed issue at https://github.com/kizniche/Mycodo/issues
Please DO NOT OPEN AN ISSUE:
- If your Mycodo version is not the latest release version, please update your device before submitting your issue (unless your issue is related to not being able to upgrade). Your problem might already be solved.
- If your issue has been addressed before. If you have any new information that may aid in solving the issue, post it in the issue that already exists.
If you are going to post a new issue, next read How to Write a Good Bug Report at https://forum.kylegabriel.com/t/how-to-write-a-good-bug-report/71
Please complete as many of the sections below, if applicable, to provide the most information that may help with investigating your issue. Replace the text in brackets with your own text describing the issue. The details requested potentially affect which options to pursue. The small amount of time you spend completing the template will also help those providing assistance by reducing the time required to help you.
### Describe the problem/bug
Inputs and functions show active, but not working after a reboot. Inputs and functions have to be manually disabled/enabled before they work.
### Versions:
- Mycodo Version: 8.11.0
- Raspberry Pi Version: 4B
- Raspbian OS Version: Raspberry Pi OS Lite kernel 5.10.17
### Reproducibility
Please list specific setup details that are involved and the steps to reproduce the behavior:
1. Reboot/halt system and restart
2. Setup --> Input shows all inputs active
3. Setup --> Function shows all previously enabled functions active
4. Live data shows no sensor data
5. Disabling the input or function produces an error indicating that the input or function ID was not found (see screenshot)
6. Enabling the input or function is successful without error
7. Data now appears in the Live Data screen for the enabled input and the function is now running
### Expected behavior
All previously enabled inputs and functions should start and provide data after reboot/halt/power failure.
### Screenshots
### Additional context
I want to avoid having to manually deactivate/active inputs and functions after a reboot and avoid trouble in the greenhouse after a power failure.
| closed | 2021-08-24T15:10:07Z | 2021-08-25T02:03:29Z | https://github.com/kizniche/Mycodo/issues/1074 | [] | kharberts | 12 |
huggingface/datasets | nlp | 7,400 | 504 Gateway Timeout when uploading large dataset to Hugging Face Hub | ### Description
I encountered consistent 504 Gateway Timeout errors while attempting to upload a large dataset (approximately 500GB) to the Hugging Face Hub. The upload fails during the process with a Gateway Timeout error.
I will continue trying to upload. While it might succeed in future attempts, I wanted to report this issue in the meantime.
### Reproduction
- I attempted the upload 3 times
- Each attempt resulted in the same 504 error during the upload process (not at the start, but in the middle of the upload)
- Using `dataset.push_to_hub()` method
### Environment Information
```
- huggingface_hub version: 0.28.0
- Platform: Linux-6.8.0-52-generic-x86_64-with-glibc2.39
- Python version: 3.11.10
- Running in iPython ?: No
- Running in notebook ?: No
- Running in Google Colab ?: No
- Running in Google Colab Enterprise ?: No
- Token path ?: /home/hotchpotch/.cache/huggingface/token
- Has saved token ?: True
- Who am I ?: hotchpotch
- Configured git credential helpers: store
- FastAI: N/A
- Tensorflow: N/A
- Torch: 2.5.1
- Jinja2: 3.1.5
- Graphviz: N/A
- keras: N/A
- Pydot: N/A
- Pillow: 10.4.0
- hf_transfer: N/A
- gradio: N/A
- tensorboard: N/A
- numpy: 1.26.4
- pydantic: 2.10.6
- aiohttp: 3.11.11
- ENDPOINT: https://huggingface.co
- HF_HUB_CACHE: /home/hotchpotch/.cache/huggingface/hub
- HF_ASSETS_CACHE: /home/hotchpotch/.cache/huggingface/assets
- HF_TOKEN_PATH: /home/hotchpotch/.cache/huggingface/token
- HF_STORED_TOKENS_PATH: /home/hotchpotch/.cache/huggingface/stored_tokens
- HF_HUB_OFFLINE: False
- HF_HUB_DISABLE_TELEMETRY: False
- HF_HUB_DISABLE_PROGRESS_BARS: None
- HF_HUB_DISABLE_SYMLINKS_WARNING: False
- HF_HUB_DISABLE_EXPERIMENTAL_WARNING: False
- HF_HUB_DISABLE_IMPLICIT_TOKEN: False
- HF_HUB_ENABLE_HF_TRANSFER: False
- HF_HUB_ETAG_TIMEOUT: 10
- HF_HUB_DOWNLOAD_TIMEOUT: 10
```
### Full Error Traceback
```python
Traceback (most recent call last):
File "/home/hotchpotch/src/github.com/hotchpotch/fineweb-2-edu-classifier-japanese/.venv/lib/python3.11/site-packages/huggingface_hub/utils/_http.py", line 406, in hf_raise_for_status
response.raise_for_status()
File "/home/hotchpotch/src/github.com/hotchpotch/fineweb-2-edu-classifier-japanese/.venv/lib/python3.11/site-packages/requests/models.py", line 1024, in raise_for_status
raise HTTPError(http_error_msg, response=self)
requests.exceptions.HTTPError: 504 Server Error: Gateway Time-out for url: https://huggingface.co/datasets/hotchpotch/fineweb-2-edu-japanese.git/info/lfs/objects/batch
The above exception was the direct cause of the following exception:
Traceback (most recent call last):
File "/home/hotchpotch/src/github.com/hotchpotch/fineweb-2-edu-classifier-japanese/create_edu_japanese_ds/upload_edu_japanese_ds.py", line 12, in <module>
ds.push_to_hub("hotchpotch/fineweb-2-edu-japanese", private=True)
File "/home/hotchpotch/src/github.com/hotchpotch/fineweb-2-edu-classifier-japanese/.venv/lib/python3.11/site-packages/datasets/dataset_dict.py", line 1665, in push_to_hub
split_additions, uploaded_size, dataset_nbytes = self[split]._push_parquet_shards_to_hub(
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/home/hotchpotch/src/github.com/hotchpotch/fineweb-2-edu-classifier-japanese/.venv/lib/python3.11/site-packages/datasets/arrow_dataset.py", line 5301, in _push_parquet_shards_to_hub
api.preupload_lfs_files(
File "/home/hotchpotch/src/github.com/hotchpotch/fineweb-2-edu-classifier-japanese/.venv/lib/python3.11/site-packages/huggingface_hub/hf_api.py", line 4215, in preupload_lfs_files
_upload_lfs_files(
File "/home/hotchpotch/src/github.com/hotchpotch/fineweb-2-edu-classifier-japanese/.venv/lib/python3.11/site-packages/huggingface_hub/utils/_validators.py", line 114, in _inner_fn
return fn(*args, **kwargs)
^^^^^^^^^^^^^^^^^^^
File "/home/hotchpotch/src/github.com/hotchpotch/fineweb-2-edu-classifier-japanese/.venv/lib/python3.11/site-packages/huggingface_hub/_commit_api.py", line 395, in _upload_lfs_files
batch_actions_chunk, batch_errors_chunk = post_lfs_batch_info(
^^^^^^^^^^^^^^^^^^^^
File "/home/hotchpotch/src/github.com/hotchpotch/fineweb-2-edu-classifier-japanese/.venv/lib/python3.11/site-packages/huggingface_hub/utils/_validators.py", line 114, in _inner_fn
return fn(*args, **kwargs)
^^^^^^^^^^^^^^^^^^^
File "/home/hotchpotch/src/github.com/hotchpotch/fineweb-2-edu-classifier-japanese/.venv/lib/python3.11/site-packages/huggingface_hub/lfs.py", line 168, in post_lfs_batch_info
hf_raise_for_status(resp)
File "/home/hotchpotch/src/github.com/hotchpotch/fineweb-2-edu-classifier-japanese/.venv/lib/python3.11/site-packages/huggingface_hub/utils/_http.py", line 477, in hf_raise_for_status
raise _format(HfHubHTTPError, str(e), response) from e
huggingface_hub.errors.HfHubHTTPError: 504 Server Error: Gateway Time-out for url: https://huggingface.co/datasets/hotchpotch/fineweb-2-edu-japanese.git/info/lfs/objects/batch
```
| open | 2025-02-14T02:18:35Z | 2025-02-14T23:48:36Z | https://github.com/huggingface/datasets/issues/7400 | [] | hotchpotch | 4 |
coqui-ai/TTS | python | 2,456 | [Bug] yourTTS Python API French not working any more (KeyError: 'fr') | ### Describe the bug
Python API for yourTTS is no longer working for French (working well for English). I used the exactly same example as the given one on readme.md. This was working on Feb 18, 2023.
It gives me a "KeyError: 'fr'"
Here is the colab link: https://colab.research.google.com/drive/1YuiWxDCbLsw5dvEd9YQpHwDBMo7Ngtvx#scrollTo=Xqy_I3zNbbEG
### To Reproduce
from TTS.api import TTS
tts = TTS(model_name="tts_models/multilingual/multi-dataset/your_tts")
tts.tts_to_file("C'est le clonage de la voix.", speaker_wav="clone.wav", language="fr", file_path="output_fr.wav")
Here is the colab link: https://colab.research.google.com/drive/1YuiWxDCbLsw5dvEd9YQpHwDBMo7Ngtvx#scrollTo=Xqy_I3zNbbEG
### Expected behavior
"output_fr.wav" created
### Logs
```shell
> Text splitted to sentences.
["C'est le clonage de la voix."]
---------------------------------------------------------------------------
KeyError Traceback (most recent call last)
<ipython-input-7-bb0038b4fd2c> in <module>
----> 1 tts.tts_to_file("C'est le clonage de la voix.", speaker_wav="clone.wav", language="fr", file_path="output_fr.wav")
2 frames
/usr/local/lib/python3.9/dist-packages/TTS/api.py in tts_to_file(self, text, speaker, language, speaker_wav, file_path)
218 Output file path. Defaults to "output.wav".
219 """
--> 220 wav = self.tts(text=text, speaker=speaker, language=language, speaker_wav=speaker_wav)
221 self.synthesizer.save_wav(wav=wav, path=file_path)
/usr/local/lib/python3.9/dist-packages/TTS/api.py in tts(self, text, speaker, language, speaker_wav)
181 self._check_arguments(speaker=speaker, language=language, speaker_wav=speaker_wav)
182
--> 183 wav = self.synthesizer.tts(
184 text=text,
185 speaker_name=speaker,
/usr/local/lib/python3.9/dist-packages/TTS/utils/synthesizer.py in tts(self, text, speaker_name, language_name, speaker_wav, style_wav, style_text, reference_wav, reference_speaker_name)
251
252 elif language_name and isinstance(language_name, str):
--> 253 language_id = self.tts_model.language_manager.name_to_id[language_name]
254
255 elif not language_name:
KeyError: 'fr'
```
### Environment
```shell
{
"CUDA": {
"GPU": [],
"available": false,
"version": "11.7"
},
"Packages": {
"PyTorch_debug": false,
"PyTorch_version": "2.0.0+cu117",
"TTS": "0.12.0",
"numpy": "1.21.6"
},
"System": {
"OS": "Linux",
"architecture": [
"64bit",
""
],
"processor": "x86_64",
"python": "3.8.0",
"version": "#36~22.04.1-Ubuntu SMP PREEMPT_DYNAMIC Fri Feb 17 15:17:25 UTC 2"
}
}
```
### Additional context
_No response_ | closed | 2023-03-24T16:30:21Z | 2023-03-27T06:30:00Z | https://github.com/coqui-ai/TTS/issues/2456 | [
"bug"
] | SiaH319 | 4 |
microsoft/unilm | nlp | 991 | Incorrect Window Size - BEATs | **Describe the bug**
Model I am using (UniLM, MiniLM, LayoutLM ...): BEATs
The problem arises when using:
* [x] the official example scripts: (give details below)
* [ ] my own modified scripts: (give details below)
A clear and concise description of what the bug is.
When I try to use a finetune trained model of BEATs with my own audio file, I get the following error. The code is identical to the load Fine-tuned Models section of the README in BEATs except that I load own audio file with `torchaudio` and then use its size to create the padding mask. I've ensured that the sampling rate of the audio file is 16khz.
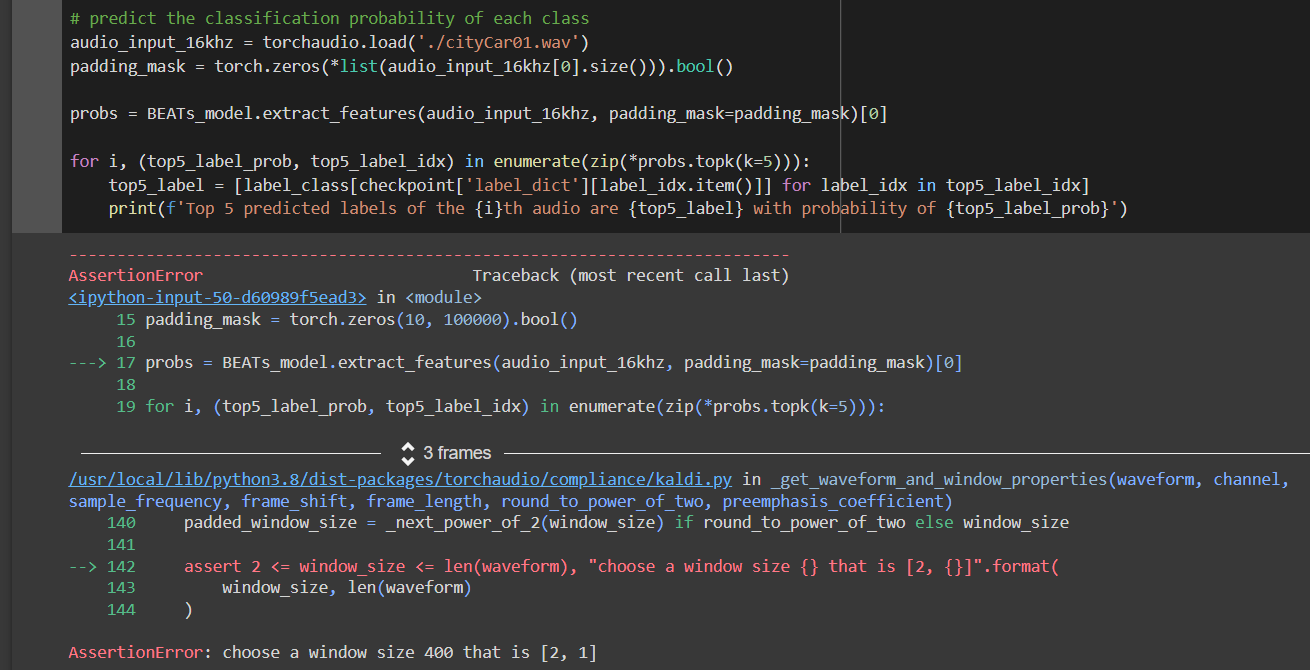
**To Reproduce**
Steps to reproduce the behavior:
1. Replace `audio_input_16khz` by using `torchaudio.load()` with your own ~10s audio file.
2. Replace `padding_mask` with the size of the loaded audio file.
**Expected behavior**
A clear and concise description of what you expected to happen.
Labels of the audio file should be printed.
- Platform: Google Colaboratory
- Python version: 3.8
- PyTorch version (GPU?): 1.13.1
| closed | 2023-02-02T21:37:39Z | 2023-02-03T04:51:07Z | https://github.com/microsoft/unilm/issues/991 | [] | jeremyng353 | 1 |
explosion/spaCy | data-science | 13,147 | The en_core_web_trf model results in zero output | ### Discussed in https://github.com/explosion/spaCy/discussions/13145
<div type='discussions-op-text'>
<sup>Originally posted by **HarounAbdelsamad** November 22, 2023</sup>
I tried training the en_core_web_trf model based on datasets i have but after training and evaluation the fscore, recall and precision are all zero. I tried using the small model works fine. I changed the code so that the transformer component is added to the pipe and also use another config file for this. Here is my code for reference:
Could anybody help me or direct me towards the issue?
[code.txt](https://github.com/explosion/spaCy/files/13442430/code.txt)
</div> | closed | 2023-11-23T08:04:50Z | 2023-12-24T00:02:25Z | https://github.com/explosion/spaCy/issues/13147 | [
"training",
"feat / transformer"
] | HarounAbdelsamad | 2 |
encode/databases | asyncio | 113 | Native decimal support | Hi,
first thank you for this great library. I was missing native decimal support (decimals are rounded) and sketched a solution in https://github.com/encode/databases/pull/112 - seems like it can be safely enabled for Postgres and MySQL. Hope this would be useful for you.
Cheers
Jakub
| closed | 2019-06-25T13:33:01Z | 2019-06-26T06:25:11Z | https://github.com/encode/databases/issues/113 | [] | coobas | 2 |
huggingface/transformers | nlp | 36,040 | `Llama-3.2-11B-Vision-Instruct` (`mllama`) FSDP fails if grad checkpointing is enabled | ### System Info
1 node with 4 A100 40GB GPUs launched by SkyPilot (`A100:4`) on GCP
### Who can help?
### What happened?
FSDP SFT fine-tuning of `meta-llama/Llama-3.2-90B-Vision-Instruct` on 1 node with 4 `A100-40GB` GPU-s with TRL trainer (`trl.SFTTrainer`) started to fail for us after upgrade to `transformers>=4.46`, including `transformers==4.48.2`:
Sample error for `sdpa` attention:
```
[rank2]: return self._call_impl(*args, **kwargs)
[rank2]: File "/home/gcpuser/miniconda3/lib/python3.10/site-packages/torch/nn/modules/module.py", line 1747, in _call_impl
[rank2]: return forward_call(*args, **kwargs)
[rank2]: File "/home/gcpuser/miniconda3/lib/python3.10/site-packages/transformers/models/mllama/modeling_mllama.py", line 798, in forward
[rank2]: attn_output = torch.nn.functional.scaled_dot_product_attention(
[rank2]: RuntimeError: The expanded size of the tensor (46) must match the existing size (23) at non-singleton dimension 3. Target sizes: [2, 32, 23, 46]. Tensor sizes: [2, 1, 23, 23]
```
It fails with similar error messages for `eager` attention as well.
This affects both full-finetuning and LoRA tuning.
Disabling grad checkpointing (w/ smaller batch size) resolves the error.
Note that if we install `transformers>=4.45.2,<4.46` then training works w/o the error under the same settings w/ gradient checkpointing on or off. It's likely the regression is related to this attention refactor: https://github.com/huggingface/transformers/pull/35235
### Steps to reproduce the bug
1. Install `transformers>=4.48.2,<4.49`, `trl>=0.13.0,<0.14`
2. FSDP tune `meta-llama/Llama-3.2-90B-Vision-Instruct` using `torchrun`
Accelerate environment variables for FSDP:
` {'ACCELERATE_DYNAMO_BACKEND': 'NO', 'ACCELERATE_DYNAMO_MODE': 'default', 'ACCELERATE_DYNAMO_USE_FULLGRAPH': 'False', 'ACCELERATE_DYNAMO_USE_DYNAMIC': 'False', 'FSDP_CPU_RAM_EFFICIENT_LOADING': 'true', 'FSDP_USE_ORIG_PARAMS': 'true', 'ACCELERATE_USE_FSDP': 'true', 'FSDP_SHARDING_STRATEGY': 'HYBRID_SHARD', 'FSDP_OFFLOAD_PARAMS': 'false', 'FSDP_BACKWARD_PREFETCH': 'BACKWARD_PRE', 'FSDP_FORWARD_PREFETCH': 'false', 'FSDP_STATE_DICT_TYPE': 'FULL_STATE_DICT', 'FSDP_AUTO_WRAP_POLICY': 'TRANSFORMER_BASED_WRAP', 'FSDP_MIN_NUM_PARAMS': '100000', 'FSDP_TRANSFORMER_CLS_TO_WRAP': 'MllamaSelfAttentionDecoderLayer,MllamaCrossAttentionDecoderLayer,MllamaVisionEncoderLayer', 'FSDP_SYNC_MODULE_STATES': 'true', 'FSDP_ACTIVATION_CHECKPOINTING': 'true'}
`
### Information
- [ ] The official example scripts
- [ ] My own modified scripts
### Tasks
- [ ] An officially supported task in the `examples` folder (such as GLUE/SQuAD, ...)
- [ ] My own task or dataset (give details below)
### Reproduction
I don't yet have a standalone repro script for this issue (it was reproduced as part of a different system). If it's a requirement, and you can't easily reproduce the issue using your own scripts based on the description above, please let me know .
### Expected behavior
No error | open | 2025-02-05T01:23:16Z | 2025-03-08T17:55:39Z | https://github.com/huggingface/transformers/issues/36040 | [
"bug"
] | nikg4 | 3 |
errbotio/errbot | automation | 920 | Add setting in the flows so they don't prompt for the next step | ### I am...
* [x] Suggesting a new feature
### I am running...
* Errbot version: 4.3.4
* OS version: MacOS Sierra
* Python version: 3.5
* Using a virtual environment: yes
### Issue description
While in flow, there is always a prompt to the next step. I would like to add an flag to make this optional | closed | 2016-12-02T17:41:52Z | 2016-12-06T19:53:01Z | https://github.com/errbotio/errbot/issues/920 | [
"newcomer-friendly",
"feature: plugins",
"#usability"
] | avivl | 0 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.