code stringlengths 2.5k 150k | kind stringclasses 1 value |
|---|---|
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Exercise-1:-Compute-MSE" data-toc-modified-id="Exercise-1:-Compute-MSE-1"><span class="toc-item-num">1 </span>Exercise 1: Compute MSE</a></span></li><li><span><a href="#Interactive-Demo:-MSE-Explorer" data-toc-modified-id="Interactive-Demo:-MSE-Explorer-2"><span class="toc-item-num">2 </span>Interactive Demo: MSE Explorer</a></span><ul class="toc-item"><li><span><a href="#Exercise-2:-Solve-for-the-Optimal-Estimator" data-toc-modified-id="Exercise-2:-Solve-for-the-Optimal-Estimator-2.1"><span class="toc-item-num">2.1 </span>Exercise 2: Solve for the Optimal Estimator</a></span></li></ul></li><li><span><a href="#Least-Squares-Optimization-Derivation" data-toc-modified-id="Least-Squares-Optimization-Derivation-3"><span class="toc-item-num">3 </span>Least Squares Optimization Derivation</a></span></li></ul></div>
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W1D3_ModelFitting/W1D3_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Neuromatch Academy: Week 1, Day 3, Tutorial 1
# Model Fitting: Linear regression with MSE
**Content creators**: Pierre-Étienne Fiquet, Anqi Wu, Alex Hyafil with help from Byron Galbraith
**Content reviewers**: Lina Teichmann, Saeed Salehi, Patrick Mineault, Ella Batty, Michael Waskom
___
#Tutorial Objectives
This is Tutorial 1 of a series on fitting models to data. We start with simple linear regression, using least squares optimization (Tutorial 1) and Maximum Likelihood Estimation (Tutorial 2). We will use bootstrapping to build confidence intervals around the inferred linear model parameters (Tutorial 3). We'll finish our exploration of regression models by generalizing to multiple linear regression and polynomial regression (Tutorial 4). We end by learning how to choose between these various models. We discuss the bias-variance trade-off (Tutorial 5) and Cross Validation for model selection (Tutorial 6).
In this tutorial, we will learn how to fit simple linear models to data.
- Learn how to calculate the mean-squared error (MSE)
- Explore how model parameters (slope) influence the MSE
- Learn how to find the optimal model parameter using least-squares optimization
---
**acknowledgements:**
- we thank Eero Simoncelli, much of today's tutorials are inspired by exercises asigned in his mathtools class.
---
# Setup
```
import numpy as np
import matplotlib.pyplot as plt
#@title Figure Settings
import ipywidgets as widgets # interactive display
%config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
#@title Helper functions
def plot_observed_vs_predicted(x, y, y_hat, theta_hat):
""" Plot observed vs predicted data
Args:
x (ndarray): observed x values
y (ndarray): observed y values
y_hat (ndarray): predicted y values
"""
fig, ax = plt.subplots()
ax.scatter(x, y, label='Observed') # our data scatter plot
ax.plot(x, y_hat, color='r', label='Fit') # our estimated model
# plot residuals
ymin = np.minimum(y, y_hat)
ymax = np.maximum(y, y_hat)
ax.vlines(x, ymin, ymax, 'g', alpha=0.5, label='Residuals')
ax.set(
title=fr"$\hat{{\theta}}$ = {theta_hat:0.2f}, MSE = {mse(x, y, theta_hat):.2f}",
xlabel='x',
ylabel='y'
)
ax.legend()
```
---
# Section 1: Mean Squared Error (MSE)
```
#@title Video 1: Linear Regression & Mean Squared Error
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="HumajfjJ37E", width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
```
**Linear least squares regression** is an old but gold optimization procedure that we are going to use for data fitting. Least squares (LS) optimization problems are those in which the objective function is a quadratic function of the
parameter(s) being optimized.
Suppose you have a set of measurements, $y_{n}$ (the "dependent" variable) obtained for different input values, $x_{n}$ (the "independent" or "explanatory" variable). Suppose we believe the measurements are proportional to the input values, but are corrupted by some (random) measurement errors, $\epsilon_{n}$, that is:
$$y_{n}= \theta x_{n}+\epsilon_{n}$$
for some unknown slope parameter $\theta.$ The least squares regression problem uses **mean squared error (MSE)** as its objective function, it aims to find the value of the parameter $\theta$ by minimizing the average of squared errors:
\begin{align}
\min _{\theta} \frac{1}{N}\sum_{n=1}^{N}\left(y_{n}-\theta x_{n}\right)^{2}
\end{align}
We will now explore how MSE is used in fitting a linear regression model to data. For illustrative purposes, we will create a simple synthetic dataset where we know the true underlying model. This will allow us to see how our estimation efforts compare in uncovering the real model (though in practice we rarely have this luxury).
First we will generate some noisy samples $x$ from [0, 10) along the line $y = 1.2x$ as our dataset we wish to fit a model to.
```
# @title
# @markdown Execute this cell to generate some simulated data
# setting a fixed seed to our random number generator ensures we will always
# get the same psuedorandom number sequence
np.random.seed(121)
# Let's set some parameters
theta = 1.2
n_samples = 30
# Draw x and then calculate y
x = 10 * np.random.rand(n_samples) # sample from a uniform distribution over [0,10)
noise = np.random.randn(n_samples) # sample from a standard normal distribution
y = theta * x + noise
# Plot the results
fig, ax = plt.subplots()
ax.scatter(x, y) # produces a scatter plot
ax.set(xlabel='x', ylabel='y');
```
Now that we have our suitably noisy dataset, we can start trying to estimate the underlying model that produced it. We use MSE to evaluate how successful a particular slope estimate $\hat{\theta}$ is for explaining the data, with the closer to 0 the MSE is, the better our estimate fits the data.
## Exercise 1: Compute MSE
In this exercise you will implement a method to compute the mean squared error for a set of inputs $x$, measurements $y$, and slope estimate $\hat{\theta}$. We will then compute and print the mean squared error for 3 different choices of theta
```
def mse(x, y, theta_hat):
"""Compute the mean squared error
Args:
x (ndarray): An array of shape (samples,) that contains the input values.
y (ndarray): An array of shape (samples,) that contains the corresponding
measurement values to the inputs.
theta_hat (float): An estimate of the slope parameter
Returns:
float: The mean squared error of the data with the estimated parameter.
"""
####################################################
## TODO for students: compute the mean squared error
# Fill out function and remove
# raise NotImplementedError("Student exercise: compute the mean squared error")
####################################################
# Compute the estimated y
y_hat = x * theta_hat
# Compute mean squared error
mse = np.mean((y_hat - y)**2)
return mse
# Uncomment below to test your function
theta_hats = [0.75, 1.0, 1.5]
for theta_hat in theta_hats:
print(f"theta_hat of {theta_hat} has an MSE of {mse(x, y, theta_hat):.2f}")
# to_remove solution
def mse(x, y, theta_hat):
"""Compute the mean squared error
Args:
x (ndarray): An array of shape (samples,) that contains the input values.
y (ndarray): An array of shape (samples,) that contains the corresponding
measurement values to the inputs.
theta_hat (float): An estimate of the slope parameter
Returns:
float: The mean squared error of the data with the estimated parameter.
"""
# Compute the estimated y
y_hat = theta_hat * x
# Compute mean squared error
mse = np.mean((y - y_hat)**2)
return mse
theta_hats = [0.75, 1.0, 1.5]
for theta_hat in theta_hats:
print(f"theta_hat of {theta_hat} has an MSE of {mse(x, y, theta_hat):.2f}")
```
The result should be:
theta_hat of 0.75 has an MSE of 9.08\
theta_hat of 1.0 has an MSE of 3.0\
theta_hat of 1.5 has an MSE of 4.52
We see that $\hat{\theta} = 1.0$ is our best estimate from the three we tried. Looking just at the raw numbers, however, isn't always satisfying, so let's visualize what our estimated model looks like over the data.
```
#@title
#@markdown Execute this cell to visualize estimated models
fig, axes = plt.subplots(ncols=3, figsize=(18, 4))
for theta_hat, ax in zip(theta_hats, axes):
# True data
ax.scatter(x, y, label='Observed') # our data scatter plot
# Compute and plot predictions
y_hat = theta_hat * x
ax.plot(x, y_hat, color='r', label='Fit') # our estimated model
ax.set(
title= fr'$\hat{{\theta}}$= {theta_hat}, MSE = {mse(x, y, theta_hat):.2f}',
xlabel='x',
ylabel='y'
);
axes[0].legend()
```
## Interactive Demo: MSE Explorer
Using an interactive widget, we can easily see how changing our slope estimate changes our model fit. We display the **residuals**, the differences between observed and predicted data, as line segments between the data point (observed response) and the corresponding predicted response on the model fit line.
```
#@title
#@markdown Make sure you execute this cell to enable the widget!
@widgets.interact(theta_hat=widgets.FloatSlider(1.0, min=0.0, max=2.0))
def plot_data_estimate(theta_hat):
y_hat = theta_hat * x
plot_observed_vs_predicted(x, y, y_hat, theta_hat)
```
While visually exploring several estimates can be instructive, it's not the most efficient for finding the best estimate to fit our data. Another technique we can use is choose a reasonable range of parameter values and compute the MSE at several values in that interval. This allows us to plot the error against the parameter value (this is also called an **error landscape**, especially when we deal with more than one parameter). We can select the final $\hat{\theta}$ ($\hat{\theta}_{MSE}$) as the one which results in the lowest error.
```
# @title
# @markdown Execute this cell to loop over theta_hats, compute MSE, and plot results
# Loop over different thetas, compute MSE for each
theta_hat_grid = np.linspace(-2.0, 4.0)
errors = np.zeros(len(theta_hat_grid))
for i, theta_hat in enumerate(theta_hat_grid):
errors[i] = mse(x, y, theta_hat)
# Find theta that results in lowest error
best_error = np.min(errors)
theta_hat = theta_hat_grid[np.argmin(errors)]
# Plot results
fig, ax = plt.subplots()
ax.plot(theta_hat_grid, errors, '-o', label='MSE', c='C1')
ax.axvline(theta, color='g', ls='--', label=r"$\theta_{True}$")
ax.axvline(theta_hat, color='r', ls='-', label=r"$\hat{{\theta}}_{MSE}$")
ax.set(
title=fr"Best fit: $\hat{{\theta}}$ = {theta_hat:.2f}, MSE = {best_error:.2f}",
xlabel=r"$\hat{{\theta}}$",
ylabel='MSE')
ax.legend();
```
We can see that our best fit is $\hat{\theta}=1.18$ with an MSE of 1.45. This is quite close to the original true value $\theta=1.2$!
---
# Section 2: Least-squares optimization
While the approach detailed above (computing MSE at various values of $\hat\theta$) quickly got us to a good estimate, it still relied on evaluating the MSE value across a grid of hand-specified values. If we didn't pick a good range to begin with, or with enough granularity, we might miss the best possible estimator. Let's go one step further, and instead of finding the minimum MSE from a set of candidate estimates, let's solve for it analytically.
We can do this by minimizing the cost function. Mean squared error is a convex objective function, therefore we can compute its minimum using calculus. Please see video or appendix for this derivation! After computing the minimum, we find that:
\begin{align}
\hat\theta = \frac{\vec{x}^\top \vec{y}}{\vec{x}^\top \vec{x}}
\end{align}
This is known as solving the normal equations. For different ways of obtaining the solution, see the notes on [Least Squares Optimization](https://www.cns.nyu.edu/~eero/NOTES/leastSquares.pdf) by Eero Simoncelli.
### Exercise 2: Solve for the Optimal Estimator
In this exercise, you will write a function that finds the optimal $\hat{\theta}$ value using the least squares optimization approach (the equation above) to solve MSE minimization. It shoud take arguments $x$ and $y$ and return the solution $\hat{\theta}$.
We will then use your function to compute $\hat{\theta}$ and plot the resulting prediction on top of the data.
```
def solve_normal_eqn(x, y):
"""Solve the normal equations to produce the value of theta_hat that minimizes
MSE.
Args:
x (ndarray): An array of shape (samples,) that contains the input values.
y (ndarray): An array of shape (samples,) that contains the corresponding
measurement values to the inputs.
Returns:
float: the value for theta_hat arrived from minimizing MSE
"""
################################################################################
## TODO for students: solve for the best parameter using least squares
# Fill out function and remove
# raise NotImplementedError("Student exercise: solve for theta_hat using least squares")
################################################################################
# Compute theta_hat analytically
# theta_hat = np.sum(x * y)/np.sum(x * x)
theta_hat = (x.T @ y) / (x.T @ x)
return theta_hat
# Uncomment below to test your function
theta_hat = solve_normal_eqn(x, y)
y_hat = theta_hat * x
plot_observed_vs_predicted(x, y, y_hat, theta_hat)
# to_remove solution
def solve_normal_eqn(x, y):
"""Solve the normal equations to produce the value of theta_hat that minimizes
MSE.
Args:
x (ndarray): An array of shape (samples,) that contains the input values.
y (ndarray): An array of shape (samples,) that contains the corresponding
measurement values to the inputs.
Returns:
float: the value for theta_hat arrived from minimizing MSE
"""
theta_hat = (x.T @ y) / (x.T @ x)
return theta_hat
theta_hat = solve_normal_eqn(x, y)
y_hat = theta_hat * x
with plt.xkcd():
plot_observed_vs_predicted(x, y, y_hat, theta_hat)
```
We see that the analytic solution produces an even better result than our grid search from before, producing $\hat{\theta} = 1.21$ with MSE = 1.43!
---
# Summary
- Linear least squares regression is an optimization procedure that can be used for data fitting:
- Task: predict a value for $y$ given $x$
- Performance measure: $\textrm{MSE}$
- Procedure: minimize $\textrm{MSE}$ by solving the normal equations
- **Key point**: We fit the model by defining an *objective function* and minimizing it.
- **Note**: In this case, there is an *analytical* solution to the minimization problem and in practice, this solution can be computed using *linear algebra*. This is *extremely* powerful and forms the basis for much of numerical computation throughout the sciences.
---
# Appendix
## Least Squares Optimization Derivation
We will outline here the derivation of the least squares solution.
We first set the derivative of the error expression with respect to $\theta$ equal to zero,
\begin{align}
\frac{d}{d\theta}\frac{1}{N}\sum_{i=1}^N(y_i - \theta x_i)^2 = 0 \\
\frac{1}{N}\sum_{i=1}^N-2x_i(y_i - \theta x_i) = 0
\end{align}
where we used the chain rule. Now solving for $\theta$, we obtain an optimal value of:
\begin{align}
\hat\theta = \frac{\sum_{i=1}^N x_i y_i}{\sum_{i=1}^N x_i^2}
\end{align}
Which we can write in vector notation as:
\begin{align}
\hat\theta = \frac{\vec{x}^\top \vec{y}}{\vec{x}^\top \vec{x}}
\end{align}
This is known as solving the *normal equations*. For different ways of obtaining the solution, see the notes on [Least Squares Optimization](https://www.cns.nyu.edu/~eero/NOTES/leastSquares.pdf) by Eero Simoncelli.
| github_jupyter |
```
import pandas as pd
def map_income(income):
if income<=9036.8:
return 'Low'
else:
return 'High'
average_number = pd.read_csv('../../data/misc/languages.csv', sep=';', decimal=',')
average_number = average_number[['Language', 'Average']]
average_number
```
# Preply
```
preply = pd.read_csv('../../data/fair/preply.csv', index_col=0)
preply = pd.merge(preply, average_number, how='left', left_on='language', right_on='Language')
preply = preply.rename(columns={'Average': 'average_num_teachers'})
preply['income_level'] = preply['income_level'].apply(map_income)
preply.head()
preply.columns
preply = preply[preply['average_num_teachers'] >= 100]
from scipy.stats import ks_2samp
import numpy as np
def hypothesis_test(group1, group2, alpha=0.1):
st, p_value = ks_2samp(group1, group2)
if p_value<alpha:
return st, p_value
else:
return st, p_value
def compute_aggregated_feature_top_k(df, top_k, language_col, aggregation_col1, aggregation_col2, target_cols, group1_1, group1_2, group2_1, group2_2):
results = pd.DataFrame(columns=['language', 'top_k', 'target_col', 'measure', 'High|Men', 'High|Women', 'Low|Men', 'Low|Women', 'count_High|Men', 'count_High|Women', 'count_Low|Men', 'count_Low|Women'])
for lang in df[language_col].unique():
temp = df[df[language_col]==lang]
temp = temp.sort_values(by='position', ascending=True)
for target in target_cols:
temp = temp.dropna(subset=[target])
if top_k is not None:
temp = temp.head(top_k)
temp[target] = pd.to_numeric(temp[target], errors='coerce')
g1 = temp.loc[(temp[aggregation_col1]==group1_1) & (temp[aggregation_col2]==group2_1)][target].values
g2 = temp.loc[(temp[aggregation_col1]==group1_1) & (temp[aggregation_col2]==group2_2)][target].values
g3 = temp.loc[(temp[aggregation_col1]==group1_2) & (temp[aggregation_col2]==group2_1)][target].values
g4 = temp.loc[(temp[aggregation_col1]==group1_2) & (temp[aggregation_col2]==group2_2)][target].values
g1_count = len(g1)
g2_count = len(g2)
g3_count = len(g3)
g4_count = len(g4)
g1_mean = g1.mean() if g1_count else None
g2_mean = g2.mean() if g2_count else None
g3_mean = g3.mean() if g3_count else None
g4_mean = g4.mean() if g4_count else None
results = results.append({'language': lang, 'top_k': len(temp), 'target_col': target, 'measure': 'mean',
'High|Men': g1_mean, 'High|Women': g2_mean, 'Low|Men': g3_mean, 'Low|Women': g4_mean, 'count_High|Men': g1_count, 'count_High|Women': g2_count, 'count_Low|Men': g3_count, 'count_Low|Women': g4_count}, ignore_index=True)
g1_median = np.median(g1) if g1_count else None
g2_median = np.median(g2) if g2_count else None
g3_median = np.median(g3) if g3_count else None
g4_median = np.median(g4) if g4_count else None
results = results.append({'language': lang, 'top_k': len(temp), 'target_col': target, 'measure': 'median',
'High|Men': g1_median, 'High|Women': g2_median, 'Low|Men': g3_median, 'Low|Women': g4_median, 'count_High|Men': g1_count, 'count_High|Women': g2_count, 'count_Low|Men': g3_count, 'count_Low|Women': g4_count}, ignore_index=True)
return results
preply_results_100 = compute_aggregated_feature_top_k(preply, 100, 'language', 'income_level', 'gender_tuned', ['price'], 'High', 'Low', 'male', 'female')
preply_results_100
preply_results_100.to_csv('../data/results/features_analysis/combined/preply.csv')
```
# Italki
```
italki = pd.read_csv('../data/results/final_dataframes/italki.csv', index_col=0)
italki = italki.drop_duplicates(subset=['user_id', 'language'])
italki = pd.merge(italki, average_number, how='left', left_on='language', right_on='Language')
italki = italki.rename(columns={'Average': 'average_num_teachers'})
italki['income_level'] = italki['income_level'].apply(map_income)
italki.head()
italki = italki[italki['average_num_teachers'] >= 100]
italki.columns
italki_results_100 = compute_aggregated_feature_top_k(italki, 100, 'language', 'income_level', 'gender_tuned', ['price'], 'High', 'Low', 'male', 'female')
italki_results_100
italki_results_100.to_csv('../data/results/features_analysis/combined/italki.csv')
```
# Verbling
```
verbling = pd.read_csv('../data/results/final_dataframes/verbling.csv', index_col=0)
verbling = verbling.drop_duplicates(subset=['first_name', 'last_name', 'language'])
verbling = pd.merge(verbling, average_number, how='left', left_on='language', right_on='Language')
verbling = verbling.rename(columns={'Average': 'average_num_teachers'})
verbling['income_level'] = verbling['income_level'].apply(map_income)
verbling.head()
verbling = verbling[verbling['average_num_teachers'] >= 100]
verbling.columns
verbling_results_100 = compute_aggregated_feature_top_k(verbling, 100, 'language', 'income_level', 'gender_tuned', ['price'], 'High', 'Low', 'male', 'female')
verbling_results_100.to_csv('../data/results/features_analysis/combined/verbling.csv')
verbling_results_100
verbling_results_100.to_csv('../data/results/features_analysis/combined/verbling.csv')
```
| github_jupyter |
# The Quarterly Japanese Economic Model (Q-JEM)
This workbook implement the "The Quarterly Japanese Economic Model (Q-JEM): 2019 version".
At http://www.boj.or.jp/en/research/wps_rev/wps_2019/wp19e07.htm/ you will find the working paper describing
the model and a zipfile containing all the relevant information needed to use the model.
The model logic has been transformed from Eview equation to ModelFlow Business logic and the dataseries has been transformed to a Pandas Dataframe.
In this workbook the impulse responses from the working paper section 3.1.1, 3.1.2, 3.1.3, and 3.1.4 has been recreated.
# Import Python libraries
```
import pandas as pd
import sys
sys.path.append('modelflow/')
from modelsandbox import newmodel
import modelnet as mn
```
# Create model and dataframe
```
with open('QJEM/model/fqjem.frm','rt') as f:
fqjem =f.read()
baseline = pd.read_pickle('QJEM/data/dfqjem.pk')
mqjem = newmodel(fqjem)
mqjem.use_preorder = 1 # make a block decomposition of the model
turbo = 0 # 1 sppeds up by compiling with Numba, 0 use straight python
```
# Define some variable labels
```
legend = {
'GDP' : 'Real gross domestic product, S.A.',
'CP' : 'Real private consumption, S.A.',
'U' : 'Unemployment rate, S.A.',
'PGDP' : 'GDP deflator',
'USGDP' : 'Real gross domestic product of the United States, S.A.',
'NUSGDP': 'Output gap of the rest of the world',
'EX': 'Real exports of goods and services, S.A.',
'IM' : 'Real imports of goods and services, S.A.',
'INV' : 'Real private non-residential investment, S.A.',
'CORE_CPI' : 'Consumer price index (all items, less fresh food), S.A.'
}
```
# Run the baseline
```
res = mqjem(baseline,antal=50,first_test = 1,ljit=turbo,chunk=49,silent=1)
```
# Make experiment with Foreign GDP +1 percent point.
```
instruments = [ 'V_NUSGAP','V_USGAP']
target = baseline.loc['2005q1':,['USGDP','NUSGDP']].mfcalc('''\
USGDP = USGDP*1.01
NUSGDP = NUSGDP*1.01
''',silent=1)
resalt = mqjem.control(baseline,target,instruments,silent=1)
```
# Display the results
```
disp = mqjem['GDP CP INV EX IM CORE_CPI'].difpctlevel.mul100.rename(legend).plot(
colrow=2,sharey=0,title='Impact of Foreign GDP +1 percent',top=0.9)
```
# Lower Oil prices
```
instruments = [ 'V_POIL']
target = baseline.loc['2005q1':,['POIL']].mfcalc('''\
POIL = POIL*0.9
''',silent=1)
resalt = mqjem.control(baseline,target,instruments,silent=1)
disp = mqjem['GDP CP INV EX IM CORE_CPI'].difpctlevel.rename(legend).plot(
colrow=2,sharey=0,title='Impact of 10 percent permanent decrease in oil price',top=0.9)
```
# Combine the two previous experiments
```
instruments = [ 'V_POIL','V_NUSGAP','V_USGAP']
target = baseline.loc['2005q1':,['POIL','USGDP','NUSGDP']].mfcalc('''\
POIL = POIL*0.9
USGDP = USGDP*1.01
NUSGDP = NUSGDP*1.01
''',silent=1)
resalt = mqjem.control(baseline,target,instruments,silent=1)
disp = mqjem['GDP CP INV EX IM CORE_CPI'].difpctlevel.mul100.rename(legend).plot(
colrow=2,sharey=0,title='Impact of Foreign gdp GDP +1 and percent 10 percent permanent decrease in oil price',top=0.9)
```
# A permanent depreciation of exchange rates.
```
instruments = [ 'V_FXYEN']
target = baseline.loc['2005q1':,['FXYEN']].mfcalc('''\
FXYEN = FXYEN*1.1
''',silent=1)
resalt = mqjem.control(baseline,target,instruments,silent=1)
disp = mqjem['GDP CP INV EX IM CORE_CPI'].difpctlevel.mul100.rename(legend).plot(
colrow=2,sharey=0,title='Impact of Foreign gdp GDP +1 and percent 10 percent permanent decrease in oil price',top=0.9)
```
# Draw the causal structure
```
fig2 = mn.draw_adjacency_matrix(mqjem.endograph,mqjem.precoreepiorder,mqjem._superstrongblock,mqjem._superstrongtype,
size=(12,12))
```
# How is CPQ determined
```
mqjem.cpq.draw(up=2,down=2,HR=0,svg=1,transdic= {'ZPI*' : 'ZPI'}) # we condens all ZPI to one, to make the chart easy
```
## Also with values
The result can be inspected in the graph/subfolder in PDF format.
```
mqjem.cpq.draw(up=1,down=1,HR=0,svg=1,transdic= {'ZPI*' : 'ZPI'},last=1) # we condens all ZPI to one, to make the chart easy
```
# Another Example
```
mqjem.ex.draw(up=1,down=1)
mqjem.ex
```
| github_jupyter |
# Temperature graph with inset of relevant region
This example demonstrates the use of a single 3D data cube with time, latitude and longitude dimensions to plot a temperature series for a single latitude coordinate, with an inset plot of the data region.
```
import matplotlib.pyplot as plt
import numpy as np
import iris
import cartopy.crs as ccrs
import iris.quickplot as qplt
import iris.plot as iplt
import iris
cube = iris.load_cube(iris.sample_data_path('ostia_monthly.nc'))
cube
```
Slice into cube to retrieve data for the inset map showing the data region
```
region = cube[-1,...]
region
```
Average over latitude to reduce cube to 1 dimension
```
plot_line = region.collapsed('latitude', iris.analysis.MEAN)
plot_line
```
Add a single subplot (axes). Could also use `ax_main = plt.subplot()`.
```
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(12, 8))
ax_main = fig.add_subplot(1, 1, 1);
fig
```
Produce a quick plot of the 1D cube
```
import iris.quickplot as qplt
qplt.plot(plot_line, axes=ax_main)
fig
```
Adjust x limits to match the data.
Because we supplied our initial axes object, the limits are not automatically scaled, and are at their defaults of 0, 0
```
ax_main.set_xlim(0, plot_line.coord('longitude').points.max())
fig
```
Adjust the y limits so that the inset map won't clash with main plot
```
ax_main.set_ylim(294, 310)
ax_main.set_title('Meridional Mean Temperature')
fig
```
Add grid lines
```
ax_main.grid()
fig
```
Add a second set of axes specifying the fractional coordinates within the figure with bottom left corner at x=0.55, y=0.58 with width 0.3 and height 0.25.
We also specify the projection, the coast lines, and ask for the global area to be displayed. The latter prevents the figure form just showing the selection of interest, i.e., the small strip along the equator.
```
import cartopy.crs as ccrs
import iris.plot as iplt
ax_sub = fig.add_axes([0.55, 0.58, 0.3, 0.25],
projection=ccrs.Mollweide(central_longitude=180))
ax_sub.set_global()
ax_sub.coastlines()
fig
```
Use iris.plot (iplt) here so colour bar properties can be specified.
Also use a sequential colour scheme to reduce confusion for those with colour-blindness
```
import iris.plot as iplt
image = iplt.pcolormesh(region, cmap='Blues', axes=ax_sub)
fig
```
Manually set the orientation and tick marks on your colour bar. We specify the `image` variable as input to the colour bar: this is the color mesh we created before.
```
import numpy as np
ticklist = np.linspace(np.min(region.data), np.max(region.data), 4)
fig.colorbar(image, orientation='horizontal', ticks=ticklist)
ax_sub.set_title('Data Region')
fig
```
| github_jupyter |
# 9. Learning Rate를 수정해 보고 싶다.
* Learning Rate는 Optimizer에서 결정됨. 따라서 문자열 Parameter가 아닌 객체를 생성해서 적용해야 함.
* [Optimizer 의 종류와 특성 (Momentum, RMSProp, Adam)](https://onevision.tistory.com/entry/Optimizer-의-종류와-특성-Momentum-RMSProp-Adam)
---
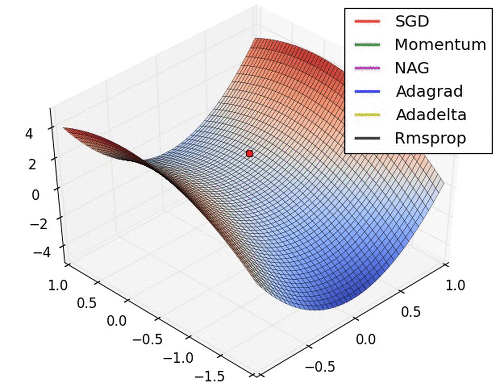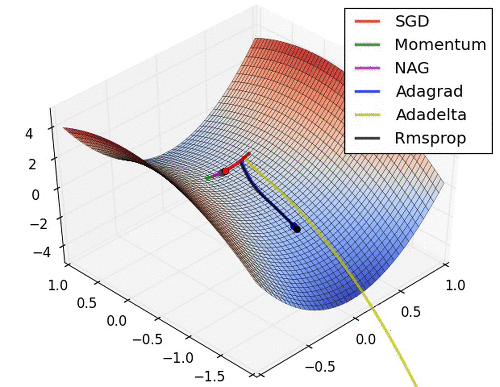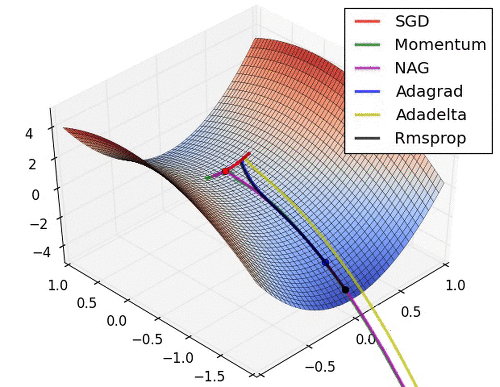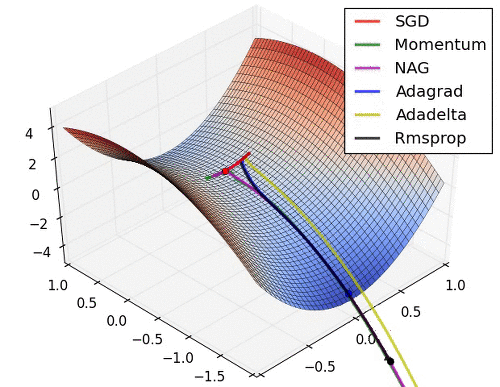
hyper parameter 설정
```
batch_size = 10
num_classes = 3
epochs = 200
```
라이브러리 및 함수 선언
```
import numpy as np
from tensorflow import keras
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
import matplotlib.pyplot as plt
def hist_view(hist):
print('## training loss and acc ##')
fig, loss_ax = plt.subplots()
acc_ax = loss_ax.twinx()
loss_ax.plot(hist.history['loss'], 'y', label='train loss')
loss_ax.plot(hist.history['val_loss'], 'r', label='val loss')
loss_ax.set_xlabel('epoch')
loss_ax.set_ylabel('loss')
loss_ax.legend(loc='center')
acc_ax.plot(hist.history['accuracy'], 'b', label='train acc')
acc_ax.plot(hist.history['val_accuracy'], 'g', label='val acc')
acc_ax.set_ylabel('accuracy')
acc_ax.legend(loc='center right')
plt.show()
```
데이터 로드
```
# from sklearn import datasets
# iris = datasets.load_iris()
# x=iris.data
import pandas as pd
![ ! -f iris0.csv ]&&wget http://j.finfra.com/_file/iris0.csv
iris=pd.read_csv("iris0.csv")
```
input data 와 target data 설정
```
x=iris.iloc[:,0:4].values
y_text=iris.iloc[:,4:5]
sets=y_text.drop_duplicates()["Species"].tolist()
encoder={k:v for v,k in enumerate(sets)}
y_num=[ encoder[i] for i in y_text["Species"].tolist() ]
```
훈련 데이터와 평가 데이터 나누기
```
# iris.target → y
y= keras.utils.to_categorical(y_num, num_classes)
from sklearn.model_selection import train_test_split
x_train,x_test,y_train,y_test=train_test_split(x,y,test_size=0.3,random_state=0)
x_train.shape,x_test.shape,y_train.shape,y_test.shape
```
모델 생성
```
model = Sequential()
model.add(Dense(6, activation='tanh', input_shape=(4,)))
model.add(Dense(4, activation=keras.activations.tanh))
model.add(Dense(num_classes, activation='softmax'))
model.summary()
```
## Optimizer 종류 확인
Keras 에서 사용 가능한 Optimizer 종류를 확인
```
from keras import optimizers
dir(optimizers)
```
## Learning Rate 수정
learning Rate 를 수정한 optimizer를 적용한다.
```
adam=optimizers.adam_v2.Adam(0.001)
model.compile(loss='categorical_crossentropy',
optimizer=adam,
metrics=['accuracy'])
```
Model 훈련
```
# batch_size,epochs
hist=model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=0,
validation_data=(x_test, y_test))
```
## 결과 확인
```
hist_view(hist)
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
decoder = {k:v for k,v in enumerate( sets )}
decoder
r=np.argmax(model.predict(x_test[:10,:]), axis=-1)
[decoder[i] for i in r]
```
# 이제 Match Nomalization과 Dropout도 적용해 보자
```
```
| github_jupyter |
# Lecture 4: Subduction
This notebook uses the data in the `data` folder and model outputs to make figures for the lecture.
## General instructions
This is a [Jupyter notebook](https://jupyter.org/) running in [Jupyter Lab](https://jupyterlab.readthedocs.io/en/stable/). The notebook is a programming environment that mixes code (the parts with `[1]: ` or similar next to them) and formatted text/images/equations with [Markdown](https://www.markdownguide.org/basic-syntax) (like this part right here).
Quick start guide:
* **Edit** any cell (blocks of code or text) by double clicking on it.
* **Execute** a code or Markdown cell by typing `Shift + Enter` after selecting it.
* The current active cell is the one with a **blue bar next to it**.
* You can run cells **in any order** as long as the code sequence makes sense (it's best to go top-to-bottom, though).
* To copy any file to the current directory, drag and drop it to the file browser on the left side.
* Notebook files have the extension `.ipynb`.
## Import things
As always, start by importing the packages that we need.
```
import gzip
import bz2
import matplotlib.pyplot as plt
import numpy as np
import scipy as sp
import xarray as xr
import pandas as pd
import pyproj
import cmocean
import cartopy.crs as ccrs
import cartopy.feature as cfeature
import verde as vd
import harmonica as hm
import boule as bl
```
## Load the datasets
We'll plot a profile from several datasets seen in class. First, we'll have to load all the data.
### Topography
```
etopo1 = xr.open_dataarray("../../data/earth_relief_10m.grd").rename("topography").rename(dict(lon="longitude", lat="latitude")) * 0.001
topography = vd.grid_to_table(etopo1)
topography
```
### Gravity
```
with bz2.open("../../data/EIGEN-6C4-gravity-30m.gdf.bz2", "rt") as decompressed:
gravity_grid = hm.load_icgem_gdf(decompressed)
gravity_grid["disturbance"] = gravity_grid.gravity_ell - bl.WGS84.normal_gravity(gravity_grid.latitude, gravity_grid.height_over_ell)
gravity = vd.grid_to_table(gravity_grid)
gravity
```
### Global seismicity catalog
```
# Use the gzip library to open the compressed file directly, like magic!
with gzip.open("../../data/jan76_dec17.ndk.gz", "rt") as quake_file:
# Start off with an empty list and we'll populate it with quakes one at a time
quakes = []
# Read and iterate over the lines of the file one by one.
# Use enumerate to get the line number as well.
for line_number, line in enumerate(quake_file):
# The data we want (location and depth) are only in every 5th line
if line_number % 5 == 0:
# The line looks like this:
# MLI 1976/01/01 01:29:39.6 -28.61 -177.64 59.0 6.2 0.0 KERMADEC ISLANDS REGION
# Split the line along spaces
parts = line.split()
# Unpack the location and depth into variables
latitude, longitude, depth = [float(i) for i in parts[3:6]]
# Add the 3 variables to the quakes list
if depth > 70:
quakes.append([latitude, longitude, depth])
quakes = np.array(quakes, dtype="float32")
print(quakes)
```
### Sediment thickness model
```
with gzip.open("../../data/sedmap.gmt.gz", "rt") as gridfile:
sediments = pd.read_csv(gridfile, delim_whitespace=True, names=["longitude", "latitude", "sediments"])
sediments
```
## Wadati-Benioff zone
```
plt.figure(figsize=(6, 8))
ax = plt.axes(projection=ccrs.LambertConformal(central_longitude=140, central_latitude=40))
ax.set_title("GlobalCMT hypocenters (1976-2017)\nfor earthquakes deeper than 70km")
ax.set_extent((125, 155, 20, 55))
ax.add_feature(
cfeature.NaturalEarthFeature(
'physical',
'land',
'50m',
edgecolor='#aaaaaa',
facecolor="#dddddd",
linewidth=0.5,
),
zorder=0,
)
# Scatter takes the point locations, size, and color
tmp = ax.scatter(
quakes[:, 1],
quakes[:, 0],
s=20,
c=quakes[:, 2],
cmap="inferno",
transform=ccrs.PlateCarree(),
)
# Add a color bar related to the scatter we just plotted
plt.colorbar(tmp, label="depth [km]", orientation="horizontal", aspect=50, pad=0.01, shrink=0.9)
plt.subplots_adjust(top=0.94, bottom=-0.07, right=1, left=0, hspace=0, wspace=0)
plt.savefig("japan-trench-globalcmt.png", dpi=200)
plt.show()
```
## Mariana Trench profile
```
def make_profiles(start, end, region, projection):
interpolator = vd.ScipyGridder(method="cubic")
profile = []
data_names = ["topography", "disturbance", "sediments"]
data_vars = [topography, gravity, sediments]
for name, table in zip(data_names, data_vars):
table = table[vd.inside((table.longitude, table.latitude), region)]
coords = projection(table.longitude.values, table.latitude.values)
interpolator.fit(coords, table[name])
profile.append(
interpolator.profile(
start,
end,
size=300,
projection=projection,
data_names=[name],
dims=["latitude", "longitude"],
)
)
profile = pd.concat(profile, axis=1)
# Remove duplicate columns
profile = profile.loc[:,~profile.columns.duplicated()]
# m to km
profile["distance"] *= 0.001
return profile
mariana = make_profiles(
end=(150, 13),
start=(144, 15),
region=(135, 155, 5, 20),
projection=pyproj.Proj(proj="merc"),
)
fig, ax = plt.subplots(1, 1, sharex=True, figsize=(6, 3))
bounds = [mariana.distance.min(), mariana.distance.max()]
max_depth = -10
ax.fill_between(bounds, [0, 0], max_depth, color='#bbbbbb')
#ax.fill_between(bounds, [0, 0], max_depth, color='#629fe3')
ax.fill_between(mariana.distance, mariana.topography, max_depth, color='#333333')
ax.set_ylabel('topography/bathymetry\n[km]')
ax.set_xlabel('Distance [km]')
ax.set_xlim(*bounds)
ax.set_ylim(max_depth, mariana.topography.max() + 4.5)
text_args = dict(horizontalalignment='center', verticalalignment='center')
ax.text(60, 1.5, "Mariana\nPlate", **text_args)
ax.text(160, 1.5, "Island\narc", **text_args)
ax.text(350, 1.5, "Mariana\nTrench", **text_args)
ax.text(500, -3, "forebulge", **text_args)
ax.text(650, 1.5, "Pacific\nPlate", **text_args)
plt.tight_layout(w_pad=0, h_pad=0)
plt.savefig("profiles-mariana.png", dpi=200)
plt.show()
```
## Flexure
```
ranges = mariana.distance > 340
reference_bathymetry = -6
flexure = mariana.topography[ranges] - reference_bathymetry
distance = mariana.distance[ranges].values
x0 = distance[np.where(flexure > 0)[0][0]]
x_forebulge = x0 + 55
flexure_forebulge = 0.5
plt.figure(figsize=(6, 4))
plt.title("Bathymetry profile at the Mariana trench")
plt.plot(mariana.distance[ranges], mariana.topography[ranges], "-k", linewidth=1.5)
plt.plot(
[mariana.distance[ranges].min(), mariana.distance[ranges].max()],
[reference_bathymetry, reference_bathymetry],
"--",
color="#666666",
linewidth=1,
)
plt.text(345, reference_bathymetry + 0.1, "reference")
plt.xlabel(r"distance [km]")
plt.ylabel(r"bathymetry [km]")
plt.xlim(mariana.distance[ranges].min(), mariana.distance[ranges].max())
plt.ylim(-9, -4.5)
plt.tight_layout()
plt.savefig("mariana-flexure-observed.png", dpi=200)
plt.show()
plt.figure(figsize=(6, 4))
plt.title("Flexure profile at the Mariana trench")
plt.plot(distance, flexure, "-k", linewidth=1.5)
plt.plot([distance.min(), distance.max()], [0, 0], "--", color="#666666", linewidth=1)
plt.plot([x0, x0], [-3, 2], "--", color="#666666", linewidth=1)
plt.plot([x_forebulge, x_forebulge], [-3, 2], "--", color="#666666", linewidth=1)
plt.text(x0 + 5, -2, "$x_0 = {:.0f}$\n km".format(x0))
plt.text(x_forebulge + 5, -2, "$x_b = {:.0f}$\n km".format(x_forebulge))
plt.plot([distance.min(), distance.max()], [flexure_forebulge, flexure_forebulge], "--", color="#666666", linewidth=1)
plt.text(distance.min() + 10, flexure_forebulge + 0.1, r"$w_b = {:.1f}$ km".format(flexure_forebulge))
plt.xlabel(r"distance $x$ [km]")
plt.ylabel(r"flexure $w$ [km]")
plt.xlim(distance.min(), distance.max())
plt.ylim(-3, 1.5)
plt.tight_layout()
plt.savefig("mariana-flexure-scaled.png", dpi=200)
plt.show()
def flexure_end_load(x, x0, xforebulge, flexure_forebulge):
xscaled = (x - x0) / (xforebulge - x0)
flexure = (
flexure_forebulge
* np.exp(np.pi / 4)
* np.sqrt(2)
* np.exp(- np.pi / 4 * xscaled)
* np.sin(np.pi / 4 * xscaled)
)
return flexure
predicted_flexure = flexure_end_load(distance, x0, x_forebulge, flexure_forebulge)
plt.figure(figsize=(6, 4))
plt.title("Flexure profile at the Mariana trench")
plt.plot([distance.min(), distance.max()], [0, 0], "--", color="#666666", linewidth=1)
plt.plot(distance, flexure, "-k", linewidth=1.5, label="observed")
plt.plot(distance, predicted_flexure, "--r", linewidth=1.5, label="predicted")
plt.legend(loc="lower right")
plt.plot([x0, x0], [-3, 2], "--", color="#666666", linewidth=1)
plt.plot([x_forebulge, x_forebulge], [-3, 2], "--", color="#666666", linewidth=1)
plt.text(x0 + 5, -2, "$x_0 = {:.0f}$\n km".format(x0))
plt.text(x_forebulge + 5, -2, "$x_b = {:.0f}$\n km".format(x_forebulge))
plt.plot([distance.min(), distance.max()], [flexure_forebulge, flexure_forebulge], "--", color="#666666", linewidth=1)
plt.text(distance.min() + 10, flexure_forebulge + 0.1, r"$w_b = {:.1f}$ km".format(flexure_forebulge))
plt.xlabel(r"distance $x$ [km]")
plt.ylabel(r"flexure $w$ [km]")
plt.xlim(distance.min(), distance.max())
plt.ylim(-3, 1.5)
plt.tight_layout()
plt.savefig("mariana-flexure-predicted.png", dpi=200)
plt.show()
def elastic_thickness(x0, xforebulge):
gravity = 9.8e-3 # km/s²
density_contrast = 3.3e12 - 1.0e12 # kg/km³
young_modulus = 70e12 # kg/(s²km)
poisson_modulus = 0.25 # dimensionless
flexural_rigidity = (
4**3 * density_contrast * gravity / np.pi**4
* (xforebulge - x0)**4
) # kg.km²/s²
thickness = np.power(
flexural_rigidity * 12 * (1 - poisson_modulus**2) / young_modulus,
1/3
)
return thickness
print("Effective elastic thickness {:.0f} km".format(elastic_thickness(x0, x_forebulge)))
```
## Gravity/bathymetry profiles
```
def plot_profiles(profile):
fig, axes = plt.subplots(2, 1, sharex=True, figsize=(7, 5))
bounds = [profile.distance.min(), profile.distance.max()]
ax = axes[0]
ax.plot(profile.distance, profile.disturbance, "-k")
ax.plot(bounds, [0, 0], "--", color="gray", linewidth=1)
ax.set_ylabel("gravity disturbance\n[mGal]")
max_depth = -10
ax = axes[1]
ax.fill_between(bounds, [0, 0], max_depth, color='#bbbbbb')
#ax.fill_between(bounds, [0, 0], max_depth, color='#629fe3')
ax.fill_between(profile.distance, profile.topography, max_depth, color='#333333')
ax.set_ylabel('topography/bathymetry\n[km]')
ax.set_xlabel('Distance [km]')
ax.set_xlim(*bounds)
ax.set_ylim(max_depth, profile.topography.max() + 5)
plt.tight_layout(w_pad=0, h_pad=0)
return fig, axes
japan = make_profiles(
end=(153, 30.5),
start=(130, 45),
region=(125, 155, 20, 55),
projection=pyproj.Proj(proj="merc"),
)
fig, axes = plot_profiles(japan)
text_args = dict(horizontalalignment='center', verticalalignment='center')
axes[-1].text(200, 3, "Asia", **text_args)
axes[-1].text(900, 3, "Sea of Japan\n(East Sea)", **text_args)
axes[-1].text(1500, 3, "Japan", **text_args)
axes[-1].text(1900, 3, "trench", **text_args)
axes[-1].text(2500, 3, "Pacific\nPlate", **text_args)
plt.savefig("profiles-japan.png", dpi=200)
plt.show()
plt.figure(figsize=(6, 8))
ax = plt.axes(projection=ccrs.LambertConformal(central_longitude=140, central_latitude=40))
ax.set_extent((125, 155, 20, 55))
etopo1.plot(
ax=ax,
cmap=cmocean.cm.topo,
transform=ccrs.PlateCarree(),
cbar_kwargs=dict(label="relief [km]", orientation="horizontal", aspect=50, pad=0.01, shrink=0.9),
zorder=0,
)
ax.plot(
japan.longitude,
japan.latitude,
"-w",
linewidth=2,
transform=ccrs.PlateCarree(),
)
plt.subplots_adjust(top=0.94, bottom=-0.07, right=1, left=0, hspace=0, wspace=0)
plt.savefig("japan-relief.png", dpi=200)
plt.show()
```
We can predict gravity assuming it's all caused by the flexure of the Pacific plate. Calculations will be done using a Bouguer plate approximation.
```
where = japan.distance > japan.distance[np.argmin(japan.topography) - 1]
distance = japan.distance[where].values
flexure = japan.topography[where] + 6
disturbance = japan.disturbance[where]
plt.figure(figsize=(6, 4))
plt.title("Bathymetry of the Pacific East of Japan")
plt.plot(distance, japan.topography[where], "-k")
plt.xlim(distance.min(), distance.max())
plt.xlabel("distance [km]")
plt.ylabel("bathymetry [km]")
plt.tight_layout()
plt.savefig("bathymetry-japan.png", dpi=200)
plt.show()
x0 = distance[np.where(flexure > 0)[0][0]]
x_forebulge = x0 + 95
flexure_forebulge = 0.3
predicted_flexure = flexure_end_load(distance, x0, x_forebulge, flexure_forebulge)
plt.figure(figsize=(6, 4))
plt.title("Flexure of the Pacific plate East of Japan")
plt.plot(distance, flexure, "-k", label="observed")
plt.plot(distance, predicted_flexure, "--b", label="predicted")
plt.plot([distance.min(), distance.max()], [0, 0], "--", color="#666666", linewidth=1)
plt.plot([x0, x0], [-3, 2], "--", color="#666666", linewidth=1)
plt.plot([x_forebulge, x_forebulge], [-3, 2], "--", color="#666666", linewidth=1)
plt.text(x0 + 5, -0.8, "$x_0 = {:.0f}$ km".format(x0))
plt.text(x_forebulge + 5, -1.2, "$x_b = {:.0f}$ km".format(x_forebulge))
plt.plot([distance.min(), distance.max()], [flexure_forebulge, flexure_forebulge], "--", color="#666666", linewidth=1)
plt.text(2500, flexure_forebulge + 0.1, r"$w_b = {:.1f}$ km".format(flexure_forebulge))
plt.legend(loc="lower right")
plt.xlim(distance.min(), distance.max())
plt.ylim(-1.5, flexure.max() + 0.1)
plt.xlabel("distance [km]")
plt.ylabel("flexure [km]")
plt.tight_layout()
plt.savefig("flexure-japan.png", dpi=200)
plt.show()
# Use 0 because bouguer_correction does density_water - density_crust
predicted = (
hm.bouguer_correction(predicted_flexure * 1000, density_crust=(2800 - 1000), density_water=0)
+ hm.bouguer_correction(predicted_flexure * 1000, density_crust=(3300 - 2800), density_water=0)
)
plt.figure(figsize=(6, 4))
plt.title("Gravity disturbance in the Pacific plate\nEast of Japan")
plt.plot(distance, disturbance, "-k", label="observed")
plt.plot(distance, predicted, "--r", label="predicted")
plt.legend(loc="lower right")
plt.xlim(distance.min(), distance.max())
plt.ylim(-140, 50)
plt.xlabel("distance [km]")
plt.ylabel("gravity disturbance [mGal]")
plt.tight_layout()
plt.savefig("disturbance-predicted-japan.png", dpi=200)
plt.grid()
```
## Foreland
```
andes = make_profiles(
start=(-75, -18),
end=(-55, -18),
region=(-80, -40, -30, -15),
projection=pyproj.Proj(proj="merc"),
)
fig, axes = plot_profiles(andes)
text_args = dict(horizontalalignment='center', verticalalignment='center')
axes[-1].text(200, 6, "Pacific", **text_args)
axes[-1].text(800, 7, "Andes", **text_args)
axes[-1].text(1400, 6, "foreland\nbasins", **text_args)
axes[-1].text(2000, 6, "South\nAmerica", **text_args)
plt.savefig("profiles-andes.png", dpi=200)
plt.show()
plt.figure(figsize=(6, 8))
ax = plt.axes(projection=ccrs.Mercator(central_longitude=-60))
ax.set_extent((-90, -30, -40, 20))
etopo1.plot(
ax=ax,
cmap=cmocean.cm.topo,
transform=ccrs.PlateCarree(),
cbar_kwargs=dict(label="relief [km]", orientation="horizontal", aspect=50, pad=0.01, shrink=0.9),
zorder=0,
)
ax.plot(
andes.longitude,
andes.latitude,
"-w",
linewidth=2,
transform=ccrs.PlateCarree(),
)
plt.subplots_adjust(top=0.94, bottom=-0.07, right=1, left=0, hspace=0, wspace=0)
plt.savefig("andes-relief.png", dpi=200)
plt.show()
where = andes.distance > 1380
distance = andes.distance[where].values
basement = -andes.sediments[where]
disturbance = andes.disturbance[where]
fig, axes = plt.subplots(2, 1, figsize=(6, 5), sharex=True)
ax = axes[0]
ax.set_title("Gravity disturbance and basement depth\nin the Andean foreland")
ax.plot(distance, disturbance, "-k")
ax.plot([distance.min(), distance.max()], [0, 0], "--", color="#666666", linewidth=1)
ax.set_ylabel("gravity disturbance [mGal]")
ax = axes[1]
ax.plot(distance, basement, "-k")
ax.set_ylabel("basement depth [km]")
ax.set_xlim(distance.min(), distance.max())
ax.set_xlabel("distance [km]")
plt.tight_layout()
plt.savefig("andes-observations", dpi=200)
plt.show()
flexure = basement + 0.7
x0 = distance[np.where((flexure > 0) & (distance > 1400))[0][1]]
x_forebulge = 1770
flexure_forebulge = 0.4
predicted_flexure = flexure_end_load(distance, x0, x_forebulge, flexure_forebulge)
print("Effective elastic thickness {:.0f} km".format(elastic_thickness(x0, x_forebulge)))
plt.figure(figsize=(6, 4))
plt.title("Flexure of the Andean foreland basins")
plt.plot(distance, flexure, "-k", label="observed")
plt.plot(distance, predicted_flexure, "--b", label="predicted")
plt.plot([distance.min(), distance.max()], [0, 0], "--", color="#666666", linewidth=1)
plt.plot([x0, x0], [-3, 2], "--", color="#666666", linewidth=1)
plt.plot([x_forebulge, x_forebulge], [-3, 2], "--", color="#666666", linewidth=1)
plt.text(x0 + 5, -1.5, "$x_0 = {:.0f}$ km".format(x0))
plt.text(x_forebulge + 5, -2, "$x_b = {:.0f}$ km".format(x_forebulge))
plt.plot([distance.min(), distance.max()], [flexure_forebulge, flexure_forebulge], "--", color="#666666", linewidth=1)
plt.text(1400, flexure_forebulge + 0.1, r"$w_b = {:.1f}$ km".format(flexure_forebulge))
plt.legend(loc="lower right")
plt.xlim(distance.min(), distance.max())
plt.ylim(-3, flexure.max() + 0.5)
plt.xlabel("distance [km]")
plt.ylabel("flexure [km]")
plt.tight_layout()
plt.savefig("flexure-andes.png", dpi=200)
plt.show()
# Use 0 because bouguer_correction does density_water - density_crust
predicted = (
hm.bouguer_correction(predicted_flexure * 1000, density_crust=(2800 - 2400), density_water=0)
+ hm.bouguer_correction(predicted_flexure * 1000, density_crust=(3300 - 2800), density_water=0)
)
plt.figure(figsize=(6, 4))
plt.title("Gravity disturbance in the Andean foreland")
plt.plot(distance, disturbance, "-k", label="observed")
plt.plot(distance, predicted, "--r", label="predicted")
plt.legend(loc="lower right")
plt.xlim(distance.min(), distance.max())
#plt.ylim(-250, 50)
plt.xlabel("distance [km]")
plt.ylabel("gravity disturbance [mGal]")
plt.tight_layout()
plt.savefig("disturbance-predicted-andes.png", dpi=200)
plt.grid()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/madhavjk/cricket_analytics/blob/main/Impact_of_toss.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# IMPACT OF TOSS ON OUTCOME OF A MATCH
<br>In this notebook, we will be analysing the impact of toss on the outcome of a match. and the steps we will be following are as follows-<br><br><br>
<b> 1) Defining the context: </b>
For any analysis, we will first need to develop an idea of the data that is required and also how are we going to breakdown the analysis into different categories. After finalising the structure of data to be used, the next task is to shortlist the parameters which we'll be using for performing the analysis.<br><br>
<b> 2) Getting the data: </b>
Now, the next task is to get the data in the required format discussed in previous step.<br><br>
<b> 3) Pre-Processing the data: </b>
After we get the data, the next step is to remove the extra information and clean the dataset to create a new one having the information which can be used for calculating parameters.<br><br>
<b> 4) Calculating the parameters: </b>
Now, we need to get the mathematical value of the parameters which we have shortlisted in the first step. So, in this step, we will use the preprocessed data to calculate the required parameters.<br><br>
<b> 5) Conclusion: </b>
Then using the calculated parameters, we try to find a trend from them and using this trend, we make a conclusion of our analysis.
Now, let's look at each of these 5 steps one by one for our analysis on impact of toss on outcome of a match.
### Defining the context
For sake of simplicity, we will only be considering the test matches, but the same analysis can be extrapolated to ODIs and T20Is as well. In this analysis, we will be loking at the impact of toss on the outcome of a match, so the data that we require should have the list of all the test matches that are played, their result, and the toss result. So, the necessary columns are team name, match result and toss result, for each test match happened. Also, for this analysis, we will be considering the recent generation's matches only, i.e. from 2016-present. This period is also selected because of the sudden increase in the of the use data from 2016. Now, since we are looking at test matches, there are 3 results possible for a match (win, lose, or draw), so the parameter that we should look for analysis should not be only focused on winning matches, but also on drawn test matches. So, for this analysis, we will consider lose percentage of a team, after they have one the toss and lost the toss. To summarize, <b>the data we will require is the scorecard data of each match, having result of the match and toss result, and the parameter we will be using for analysing the impact of toss will be the losing percentage of the team after they have won/lost the toss.</b>
### Getting the data
The data required for this analysis in the above mentioned format can be found on statsguru website (by ESPNCricinfo) by applying concerned filters. The procedure to go to the required page is:<br>
1) Go to https://stats.espncricinfo.com/ci/engine/stats/index.html and click on team tab. <br>
2) Select "Home Venue" in Home or Away row and "Match Results" in View Format row. <br>
3) Give the Starting date from 01 Jan 2016 and click Submit Query.<br>
4) Go to page number 2 and copy the url. We have copied the url of page number 2 because when the first page is loaded, there is no mention of page number in the URL, but in the URL of page 2, there is a part as "page=2" which will be useful for us in automating the scraping for all pages in one go.<br><br>
After we have the URL of the webpage, now we will create a web-scraper to scrape all the pages of the query that we gave. First we will create a scaper to scrape one page, and then we will repeat the same steps for all the pages to scrape all the pages.
```
import pandas as pd
from bs4 import BeautifulSoup
from urllib.request import urlopen
url = 'https://stats.espncricinfo.com/ci/engine/stats/index.html?class=1;home_or_away=1;orderby=start;orderbyad=reverse;page=1;spanmin1=01+Jan+2016;spanval1=span;template=results;type=team;view=results' # URL of page number 1
```
To open the webpage in the python notebook, we will use "urlopen" function. It will create a HTTP Response between python notebook and the webpage.
```
text = urlopen(url) # opening the webpage in python
print(text)
```
Using the link that is being created, we will get the HTML of the webpage using BeautifulSoup library
```
soup = BeautifulSoup(text, "lxml") # getting the HTML of the webpage using BeautifulSoup(_HTTP_Response_, "lxml")
print(soup)
```
Now, we will analyse the webpage and try to find tags and attributes which makes our desired table different from all other elements on webpage. So, after analysing the webpage, it is found that the table of our interest is different from other elements with tag as table and class as engineTable. Also amongst all the tables with class engineTable, it is the only table having caption tag in it. So now, in the HTML of the webpage, we need to find all the tables with class engineTable, and amongst all those tables, we need to find the table having a caption tag inside it.
```
table = soup.findAll('table', attrs = {'class' : 'engineTable'}) # finding all the tables with class as engineTable in the HTML of webpage
int_table = 0 # defining a variable which will store the HTML of the table of our interest
for temp_table in table: # running a loop over all the tables with class engineTable
caption_tag = temp_table.findAll('caption') # Looking for caption tag in each table's HTML
if(len(caption_tag) > 0): # checking if there is any captio tag in the table
int_table = temp_table # if a caption tag is found then save this table as the table of our interest
print(int_table)
```
After getting the HTML code of the required table, now we will look for all the rows inside the table and then scrape the data row-wise
```
tr_list = int_table.findAll('tr', attrs = {'class': 'data1'}) # getting all the rows in the table
print(len(tr_list))
```
As it can be seen that now we have the HTML of all the 50 rows stored in tr_list variable with each element in tr_list as HTML of one row. Now, we will get column data of each row separately and save that into a list. So, for each row, we will look for all td tags and then for each td tag, we will clear all HTML tags from the code to get only the cleantext or the data that is written in that particular cell. Then we will append this cleantext into the row data and then we will append the entire ro data to the master data, i.e. the data for all the matches.
```
master_data = [] # list to contain all the rows in our table
for tr in tr_list:
td_list = tr.findAll('td') # finding all the td tags in a row
row_data = [] # list to save the data of each row
for td in td_list:
td_str = str(td) # converting HTML of the td tag from BeautifulSoup HTML code to string datatype
cleantext = BeautifulSoup(td_str, "lxml").get_text() # removing all the unecssary tags from the HTML of td to get only the text written in that tag on webpage
row_data.append(cleantext) # appendng that data to row data
master_data.append(row_data) # appending the entire row data to our master data (contaiing all rows)
master_data
```
The above process is to get the data from 1 webpage, but in our query, we have total 5 webpages. Each of the webpage are same, only the data is different. So now we need to repeat the above process exactly for page 2, 3, 4, and 5. For doing so, we will use a for loop where our variable 'k' will vary from 1 to 5, and repeat the same lines of code for each page. The only difference in URL of these pages is "page=1" (for page 1), "page=2" (for page 2), "page=3" (for page 3) and so on till page=5. So, we will vary our k from 1 to 5 and edit the URL with page=k for each iteration, and run the above blocks of code for each page.
```
master_data = [] # list to contain all the rows in our tables of all the pages of the query
for k in range(1,6): # varying value of k from 1 to 5
url = 'https://stats.espncricinfo.com/ci/engine/stats/index.html?class=1;home_or_away=1;orderby=start;orderbyad=reverse;page=' + str(k) + ';spanmin1=01+Jan+2016;spanval1=span;template=results;type=team;view=results' # URL of page number 'k'
text = urlopen(url) # opening the webpage in python
soup = BeautifulSoup(text, "lxml") # getting the HTML of the webpage using BeautifulSoup(_HTTP_Response_, "lxml")
table = soup.findAll('table', attrs = {'class' : 'engineTable'}) # finding all the tables with class as engineTable in the HTML of webpage
int_table = 0 # defining a variable which will store the HTML of the table of our interest
for temp_table in table: # running a loop over all the tables with class engineTable
caption_tag = temp_table.findAll('caption') # Looking for caption tag in each table's HTML
if(len(caption_tag) > 0): # checking if there is any captio tag in the table
int_table = temp_table # if a caption tag is found then save this table as the table of our interest
tr_list = int_table.findAll('tr', attrs = {'class': 'data1'}) # getting all the rows in the table
for tr in tr_list:
td_list = tr.findAll('td') # finding all the td tags in a row
row_data = [] # list to save the data of each row
for td in td_list:
td_str = str(td) # converting HTML of the td tag from BeautifulSoup HTML code to string datatype
cleantext = BeautifulSoup(td_str, "lxml").get_text() # removing all the unecssary tags from the HTML of td to get only the text written in that tag on webpage
row_data.append(cleantext) # appendng that data to row data
master_data.append(row_data) # appending the entire row data to our master data (contaiing all rows)
print(len(master_data))
```
There were total 210 matches between 01 Jan 2016 and 25 Feb 2021, and we are also getting the same number of rows in our master_data. Hence our scraper has scraped the complete data from statsguru required for this analysis, now lets convert this master_data to a dataframe, so that we can use pandas then for working with the dataset.
```
master_data_df = pd.DataFrame(master_data)
master_data_df
# renaming the columns
master_data_df.columns = ["Home", "Result", "Margin", "Toss", "Bat", "None1", "Opposition" , "Ground", "Date", "None2"]
master_data_df
#saving the data scraped
master_data_df.to_csv('toss_data_2016_2021.csv')
```
## Pre-processing the data
Now, we have the data of all the matches from 2016-present, the next step is to pre process the data so as to make it easier for calculating parameter which in our case is losing percentage. To calculate losing percentage of each team, we need to have a column in our dataset which should show whether the team who won the toss lost the match (0:if the team winning the toss does not lose the match, 1: if the team winning the toss, lose the match). Now, to create this column, we will need to columns, one is the name of the team who won the toss, and second name of the team who lost the match (in case of draw, the column value will be draw). So let us now create these 2 columns first.<br><br><br>
Before we create these columns, let us first remove the unecessary columns from the table. Column "None1" and "None2" are empty columns, so we need to drop both of these columns. Also, in opposition column, before the name of oppositiion team, "v " is written, so we need to remove that "v " from the opposition column.
```
#dropping the columns
master_data_df = master_data_df.drop(['None1', 'None2'], axis = 1) # .drop(list_of_columns_to_be_dropped)
# axis=0, if we have to remove rows and axis=1, if we have to remove columns
master_data_df
# removing "v " from opposition column
master_data_df["Opposition"] = master_data_df["Opposition"].str.replace("v ", "")
# we have replaced "v " with empty string so that we now have only the name of the team in Opposition column
master_data_df
```
Now, let us create the two columns, the first column will be the name of the team who won the toss, because in our table, we only have won and lost in the toss column, it does not have the name of the team who won the toss. To create this column, we will run a for loop on the entire dataframe and then for each row we will see if the Toss is won, then the team who won the toss is Home team, else it is the opposition team.
```
toss_team = [] # a list to contain each row's value of new column
for index, row in master_data_df.iterrows(): # for loop on the dataframe
if(row["Toss"] == "won"): # if the toss value is won, then the team who won the toss is Home team
toss_team.append(row["Home"])
else: # if the toss value is lost, then the team who won the toss is opposition
toss_team.append(row["Opposition"])
toss_team
# Adding this column to dataframe
master_data_df["toss_team"] = toss_team
master_data_df
```
Now we will add the next column as the name of the team who lost the match (Draw in case of a draw). To add this column, we will again look at each row separately. For each row, if the Result is won, then the team lost the match is opposition, if the Result is lost, then the team lost the match is home, or if the Result is draw, then the match was drawn.
```
team_lost = [] # a list to contain the name of the team who lost the match for each row
for index, row in master_data_df.iterrows(): # for loop for the entire dataframe
if(row["Result"] == "won"): # if the Result is won, then the team lost the match is opposition
team_lost.append(row["Opposition"])
elif(row["Result"] == "lost"): # if the Result is lost, then the team lost the match is Home
team_lost.append(row["Home"])
else: # if the Result is draw, then the match is draw
team_lost.append("draw")
team_lost
# Adding this column to dataframe
master_data_df["team_lost"] = team_lost
master_data_df
```
Now we will add the final column i.e. whether the team who won the toss, has lost the match or not.<br>
0: if the team who won the toss, didn't lose the match<br>
1: if the team who won the toss, lost the match
```
won_toss_lost_match = [] # defing a list that will contain the 0/1 values for each row in the dataframe
for index, row in master_data_df.iterrows(): # for loop over the entire dataframe
if(row["toss_team"] == row["team_lost"]): # if the team who lost the toss, has also lost the match, then the value will be 1
won_toss_lost_match.append(1)
else: # if the team who lost the toss, has not lost the match, then the value will be 0
won_toss_lost_match.append(0)
won_toss_lost_match
# Adding the column to dataframe
master_data_df["toss_data"] = won_toss_lost_match
master_data_df
```
## Calculating the parameters
Now, we have the data in the desired format, we need to calculate the lose percentage for each team after they have won the toss. To do so, first of all we will need to find all the unique teams in our dataset and after that, for each team, we will see the total number of instances where the team has won the toss, and then we will look at the the total number of instances where the value of "toss_data" is 1 for that team. Using these 2 values we will get the losing percentage of the team ((total instances of "toss_data = 1")/(total instances where team has won the toss))
Let us now first find the unique teams in our dataset:
```
teams = master_data_df["toss_team"].unique().tolist() # first we use the unique() function to find all the unique values in
# the "toss_team" column and then converted that output array into a list using tolist() function because we are more
# comfortable working with lists.
teams
```
After finding the unique list of teams, now we will look at each team separately and calculate losing percentage of each team. For each team, first we will filter the dataset to contain only the rows where toss_team is the team we are considering, and then we will find the total number of occurances of 1 in toss_data column in this filtered dataframe. After that we will divide this number by the total number of rows for that team to calculate lose percentage.
```
loss_percentage = [] # loss percentage list for each team
for team in teams: # running a for loop on teams list, taking one team per iteration
filtered_df = master_data_df[master_data_df["toss_team"] == team] # filtering the data where toss_team is equal to the name of our concerened team
num_of_matches = len(filtered_df) # calculating total number of matches where the team has won the toss
lost_df = filtered_df[filtered_df["toss_data"] == 1] # filtering only the rows where team has lost the match after winning the toss
lost_count = len(lost_df) # calculating number of times the team has lost the match after winning the toss
loss_percent = lost_count/num_of_matches*100 # calculating loss percentage
temp = [] # creating an empty list to store losing percentage and team name
temp.append(team) # adding team name in temp list
temp.append(loss_percent) # adding loss_percent in temp list
loss_percentage.append(temp) # adding temp list to the main loss percentage list
loss_percentage
```
## Conclusion
We will not be considering Afghanistan and Ireland as the sample size is less for them. For India, Australia, New Zealand and South Africa, losing percentage is at max 20% after winning the toss, which indicate that these teams uses the toss for their benefit and toss plays an important role in the outcome of the match. Also, for India it is only 5% which indicates that team India knows how to use the toss to their benefit very well. Teams like England, Bangladesh, West Indies and Pakistan are yet to figure out how to use the toss for their benefit as their losing percentages are high even after winning the toss. So, to conclude, it can be said that in test matches, toss plays an important role for top teams as they have less losing percentages whereas other teams are yet to take advantage of toss in tests.
| github_jupyter |
```
import h2o
import time
from h2o.estimators.glm import H2OGeneralizedLinearEstimator
from h2o.estimators.gbm import H2OGradientBoostingEstimator
from h2o.estimators.random_forest import H2ORandomForestEstimator
from h2o.estimators.deeplearning import H2ODeepLearningEstimator
# Explore a typical Data Science workflow with H2O and Python
#
# Goal: assist the manager of CitiBike of NYC to load-balance the bicycles
# across the CitiBike network of stations, by predicting the number of bike
# trips taken from the station every day. Use 10 million rows of historical
# data, and eventually add weather data.
# Connect to a cluster
h2o.init()
from h2o.h2o import _locate # private function. used to find files within h2o git project directory.
# Set this to True if you want to fetch the data directly from S3.
# This is useful if your cluster is running in EC2.
data_source_is_s3 = False
def mylocate(s):
if data_source_is_s3:
return "s3n://h2o-public-test-data/" + s
else:
return _locate(s)
# Pick either the big or the small demo.
# Big data is 10M rows
small_test = [mylocate("bigdata/laptop/citibike-nyc/2013-10.csv")]
big_test = [mylocate("bigdata/laptop/citibike-nyc/2013-07.csv"),
mylocate("bigdata/laptop/citibike-nyc/2013-08.csv"),
mylocate("bigdata/laptop/citibike-nyc/2013-09.csv"),
mylocate("bigdata/laptop/citibike-nyc/2013-10.csv"),
mylocate("bigdata/laptop/citibike-nyc/2013-11.csv"),
mylocate("bigdata/laptop/citibike-nyc/2013-12.csv"),
mylocate("bigdata/laptop/citibike-nyc/2014-01.csv"),
mylocate("bigdata/laptop/citibike-nyc/2014-02.csv"),
mylocate("bigdata/laptop/citibike-nyc/2014-03.csv"),
mylocate("bigdata/laptop/citibike-nyc/2014-04.csv"),
mylocate("bigdata/laptop/citibike-nyc/2014-05.csv"),
mylocate("bigdata/laptop/citibike-nyc/2014-06.csv"),
mylocate("bigdata/laptop/citibike-nyc/2014-07.csv"),
mylocate("bigdata/laptop/citibike-nyc/2014-08.csv")]
# ----------
# 1- Load data - 1 row per bicycle trip. Has columns showing the start and end
# station, trip duration and trip start time and day. The larger dataset
# totals about 10 million rows
print "Import and Parse bike data"
data = h2o.import_file(path=big_test)
# ----------
# 2- light data munging: group the bike starts per-day, converting the 10M rows
# of trips to about 140,000 station&day combos - predicting the number of trip
# starts per-station-per-day.
# Convert start time to: Day since the Epoch
startime = data["starttime"]
secsPerDay=1000*60*60*24
data["Days"] = (startime/secsPerDay).floor()
data.describe()
# Now do a monster Group-By. Count bike starts per-station per-day. Ends up
# with about 340 stations times 400 days (140,000 rows). This is what we want
# to predict.
grouped = data.group_by(["Days","start station name"])
bpd = grouped.count().get_frame() # Compute bikes-per-day
bpd.set_name(2,"bikes")
bpd.show()
bpd.describe()
bpd.dim
# Quantiles: the data is fairly unbalanced; some station/day combos are wildly
# more popular than others.
print "Quantiles of bikes-per-day"
bpd["bikes"].quantile().show()
# A little feature engineering
# Add in month-of-year (seasonality; fewer bike rides in winter than summer)
secs = bpd["Days"]*secsPerDay
bpd["Month"] = secs.month().asfactor()
# Add in day-of-week (work-week; more bike rides on Sunday than Monday)
bpd["DayOfWeek"] = secs.dayOfWeek()
print "Bikes-Per-Day"
bpd.describe()
# ----------
# 3- Fit a model on train; using test as validation
# Function for doing class test/train/holdout split
def split_fit_predict(data):
global gbm0,drf0,glm0,dl0
# Classic Test/Train split
r = data['Days'].runif() # Random UNIForm numbers, one per row
train = data[ r < 0.6]
test = data[(0.6 <= r) & (r < 0.9)]
hold = data[ 0.9 <= r ]
print "Training data has",train.ncol,"columns and",train.nrow,"rows, test has",test.nrow,"rows, holdout has",hold.nrow
bike_names_x = data.names
bike_names_x.remove("bikes")
# Run GBM
s = time.time()
gbm0 = H2OGradientBoostingEstimator(ntrees=500, # 500 works well
max_depth=6,
learn_rate=0.1)
gbm0.train(x =bike_names_x,
y ="bikes",
training_frame =train,
validation_frame=test)
gbm_elapsed = time.time() - s
# Run DRF
s = time.time()
drf0 = H2ORandomForestEstimator(ntrees=250, max_depth=30)
drf0.train(x =bike_names_x,
y ="bikes",
training_frame =train,
validation_frame=test)
drf_elapsed = time.time() - s
# Run GLM
if "WC1" in bike_names_x: bike_names_x.remove("WC1")
s = time.time()
glm0 = H2OGeneralizedLinearEstimator(Lambda=[1e-5], family="poisson")
glm0.train(x =bike_names_x,
y ="bikes",
training_frame =train,
validation_frame=test)
glm_elapsed = time.time() - s
# Run DL
s = time.time()
dl0 = H2ODeepLearningEstimator(hidden=[50,50,50,50], epochs=50)
dl0.train(x =bike_names_x,
y ="bikes",
training_frame =train,
validation_frame=test)
dl_elapsed = time.time() - s
# ----------
# 4- Score on holdout set & report
train_r2_gbm = gbm0.model_performance(train).r2()
test_r2_gbm = gbm0.model_performance(test ).r2()
hold_r2_gbm = gbm0.model_performance(hold ).r2()
# print "GBM R2 TRAIN=",train_r2_gbm,", R2 TEST=",test_r2_gbm,", R2 HOLDOUT=",hold_r2_gbm
train_r2_drf = drf0.model_performance(train).r2()
test_r2_drf = drf0.model_performance(test ).r2()
hold_r2_drf = drf0.model_performance(hold ).r2()
# print "DRF R2 TRAIN=",train_r2_drf,", R2 TEST=",test_r2_drf,", R2 HOLDOUT=",hold_r2_drf
train_r2_glm = glm0.model_performance(train).r2()
test_r2_glm = glm0.model_performance(test ).r2()
hold_r2_glm = glm0.model_performance(hold ).r2()
# print "GLM R2 TRAIN=",train_r2_glm,", R2 TEST=",test_r2_glm,", R2 HOLDOUT=",hold_r2_glm
train_r2_dl = dl0.model_performance(train).r2()
test_r2_dl = dl0.model_performance(test ).r2()
hold_r2_dl = dl0.model_performance(hold ).r2()
# print " DL R2 TRAIN=",train_r2_dl,", R2 TEST=",test_r2_dl,", R2 HOLDOUT=",hold_r2_dl
# make a pretty HTML table printout of the results
header = ["Model", "R2 TRAIN", "R2 TEST", "R2 HOLDOUT", "Model Training Time (s)"]
table = [
["GBM", train_r2_gbm, test_r2_gbm, hold_r2_gbm, round(gbm_elapsed,3)],
["DRF", train_r2_drf, test_r2_drf, hold_r2_drf, round(drf_elapsed,3)],
["GLM", train_r2_glm, test_r2_glm, hold_r2_glm, round(glm_elapsed,3)],
["DL ", train_r2_dl, test_r2_dl, hold_r2_dl , round(dl_elapsed,3) ],
]
h2o.H2ODisplay(table,header)
# --------------
# Split the data (into test & train), fit some models and predict on the holdout data
split_fit_predict(bpd)
# Here we see an r^2 of 0.91 for GBM, and 0.71 for GLM. This means given just
# the station, the month, and the day-of-week we can predict 90% of the
# variance of the bike-trip-starts.
# ----------
# 5- Now lets add some weather
# Load weather data
wthr1 = h2o.import_file(path=[mylocate("bigdata/laptop/citibike-nyc/31081_New_York_City__Hourly_2013.csv"),
mylocate("bigdata/laptop/citibike-nyc/31081_New_York_City__Hourly_2014.csv")])
# Peek at the data
wthr1.describe()
# Lots of columns in there! Lets plan on converting to time-since-epoch to do
# a 'join' with the bike data, plus gather weather info that might affect
# cyclists - rain, snow, temperature. Alas, drop the "snow" column since it's
# all NA's. Also add in dew point and humidity just in case. Slice out just
# the columns of interest and drop the rest.
wthr2 = wthr1[["Year Local","Month Local","Day Local","Hour Local","Dew Point (C)","Humidity Fraction","Precipitation One Hour (mm)","Temperature (C)","Weather Code 1/ Description"]]
wthr2.set_name(wthr2.index("Precipitation One Hour (mm)"), "Rain (mm)")
wthr2.set_name(wthr2.index("Weather Code 1/ Description"), "WC1")
wthr2.describe()
# Much better!
# Filter down to the weather at Noon
wthr3 = wthr2[ wthr2["Hour Local"]==12 ]
# Lets now get Days since the epoch... we'll convert year/month/day into Epoch
# time, and then back to Epoch days. Need zero-based month and days, but have
# 1-based.
wthr3["msec"] = h2o.H2OFrame.mktime(year=wthr3["Year Local"], month=wthr3["Month Local"]-1, day=wthr3["Day Local"]-1, hour=wthr3["Hour Local"])
secsPerDay=1000*60*60*24
wthr3["Days"] = (wthr3["msec"]/secsPerDay).floor()
wthr3.describe()
# msec looks sane (numbers like 1.3e12 are in the correct range for msec since
# 1970). Epoch Days matches closely with the epoch day numbers from the
# CitiBike dataset.
# Lets drop off the extra time columns to make a easy-to-handle dataset.
wthr4 = wthr3.drop("Year Local").drop("Month Local").drop("Day Local").drop("Hour Local").drop("msec")
# Also, most rain numbers are missing - lets assume those are zero rain days
rain = wthr4["Rain (mm)"]
rain[ rain.isna() ] = 0
wthr4["Rain (mm)"] = rain
# ----------
# 6 - Join the weather data-per-day to the bike-starts-per-day
print "Merge Daily Weather with Bikes-Per-Day"
bpd_with_weather = bpd.merge(wthr4,allLeft=True,allRite=False)
bpd_with_weather.describe()
bpd_with_weather.show()
# 7 - Test/Train split again, model build again, this time with weather
split_fit_predict(bpd_with_weather)
```
| github_jupyter |
# Temporal Difference: On-policy n-Tuple Expected Sarsa, Stochastic
```
import numpy as np
```
## Create environment
```
def create_environment_states():
"""Creates environment states.
Returns:
num_states: int, number of states.
num_terminal_states: int, number of terminal states.
num_non_terminal_states: int, number of non terminal states.
"""
num_states = 16
num_terminal_states = 2
num_non_terminal_states = num_states - num_terminal_states
return num_states, num_terminal_states, num_non_terminal_states
def create_environment_actions(num_non_terminal_states):
"""Creates environment actions.
Args:
num_non_terminal_states: int, number of non terminal states.
Returns:
max_num_actions: int, max number of actions possible.
num_actions_per_non_terminal_state: array[int], number of actions per
non terminal state.
"""
max_num_actions = 4
num_actions_per_non_terminal_state = np.repeat(
a=max_num_actions, repeats=num_non_terminal_states)
return max_num_actions, num_actions_per_non_terminal_state
def create_environment_successor_counts(num_states, max_num_actions):
"""Creates environment successor counts.
Args:
num_states: int, number of states.
max_num_actions: int, max number of actions possible.
Returns:
num_state_action_successor_states: array[int], number of successor
states s' that can be reached from state s by taking action a.
"""
num_state_action_successor_states = np.repeat(
a=1, repeats=num_states * max_num_actions)
num_state_action_successor_states = np.reshape(
a=num_state_action_successor_states,
newshape=(num_states, max_num_actions))
return num_state_action_successor_states
def create_environment_successor_arrays(
num_non_terminal_states, max_num_actions):
"""Creates environment successor arrays.
Args:
num_non_terminal_states: int, number of non terminal states.
max_num_actions: int, max number of actions possible.
Returns:
sp_idx: array[int], state indices of new state s' of taking action a
from state s.
p: array[float], transition probability to go from state s to s' by
taking action a.
r: array[float], reward from new state s' from state s by taking
action a.
"""
sp_idx = np.array(
object=[1, 0, 14, 4,
2, 1, 0, 5,
2, 2, 1, 6,
4, 14, 3, 7,
5, 0, 3, 8,
6, 1, 4, 9,
6, 2, 5, 10,
8, 3, 7, 11,
9, 4, 7, 12,
10, 5, 8, 13,
10, 6, 9, 15,
12, 7, 11, 11,
13, 8, 11, 12,
15, 9, 12, 13],
dtype=np.int64)
p = np.repeat(
a=1.0, repeats=num_non_terminal_states * max_num_actions * 1)
r = np.repeat(
a=-1.0, repeats=num_non_terminal_states * max_num_actions * 1)
sp_idx = np.reshape(
a=sp_idx,
newshape=(num_non_terminal_states, max_num_actions, 1))
p = np.reshape(
a=p,
newshape=(num_non_terminal_states, max_num_actions, 1))
r = np.reshape(
a=r,
newshape=(num_non_terminal_states, max_num_actions, 1))
return sp_idx, p, r
def create_environment():
"""Creates environment.
Returns:
num_states: int, number of states.
num_terminal_states: int, number of terminal states.
num_non_terminal_states: int, number of non terminal states.
max_num_actions: int, max number of actions possible.
num_actions_per_non_terminal_state: array[int], number of actions per
non terminal state.
num_state_action_successor_states: array[int], number of successor
states s' that can be reached from state s by taking action a.
sp_idx: array[int], state indices of new state s' of taking action a
from state s.
p: array[float], transition probability to go from state s to s' by
taking action a.
r: array[float], reward from new state s' from state s by taking
action a.
"""
(num_states,
num_terminal_states,
num_non_terminal_states) = create_environment_states()
(max_num_actions,
num_actions_per_non_terminal_state) = create_environment_actions(
num_non_terminal_states)
num_state_action_successor_states = create_environment_successor_counts(
num_states, max_num_actions)
(sp_idx,
p,
r) = create_environment_successor_arrays(
num_non_terminal_states, max_num_actions)
return (num_states,
num_terminal_states,
num_non_terminal_states,
max_num_actions,
num_actions_per_non_terminal_state,
num_state_action_successor_states,
sp_idx,
p,
r)
```
## Set hyperparameters
```
def set_hyperparameters():
"""Sets hyperparameters.
Returns:
num_episodes: int, number of episodes to train over.
maximum_episode_length: int, max number of timesteps for an episode.
num_qs: int, number of state-action-value functions Q_i(s, a).
alpha: float, alpha > 0, learning rate.
epsilon: float, 0 <= epsilon <= 1, exploitation-exploration trade-off,
higher means more exploration.
gamma: float, 0 <= gamma <= 1, amount to discount future reward.
"""
num_episodes = 10000
maximum_episode_length = 200
num_qs = 3
alpha = 0.1
epsilon = 0.1
gamma = 1.0
return num_episodes, maximum_episode_length, num_qs, alpha, epsilon, gamma
```
## Create value function and policy arrays
```
def create_value_function_arrays(num_qs, num_states, max_num_actions):
"""Creates value function arrays.
Args:
num_qs: int, number of state-action-value functions Q_i(s, a).
num_states: int, number of states.
max_num_actions: int, max number of actions possible.
Returns:
q: array[float], keeps track of the estimated value of each
state-action pair Q_i(s, a).
"""
q = np.repeat(a=0.0, repeats=num_qs * num_states * max_num_actions)
q = np.reshape(a=q, newshape=(num_qs, num_states, max_num_actions))
return q
def create_policy_arrays(num_non_terminal_states, max_num_actions):
"""Creates policy arrays.
Args:
num_non_terminal_states: int, number of non terminal states.
max_num_actions: int, max number of actions possible.
Returns:
policy: array[float], learned stochastic policy of which
action a to take in state s.
"""
policy = np.repeat(
a=1.0 / max_num_actions,
repeats=num_non_terminal_states * max_num_actions)
policy = np.reshape(
a=policy,
newshape=(num_non_terminal_states, max_num_actions))
return policy
```
## Create algorithm
```
# Set random seed so that everything is reproducible
np.random.seed(seed=0)
def initialize_epsiode(num_non_terminal_states):
"""Initializes epsiode with initial state.
Args:
num_non_terminal_states: int, number of non terminal states.
Returns:
init_s_idx: int, initial state index from set of non terminal states.
"""
# Randomly choose an initial state from all non-terminal states
init_s_idx = np.random.randint(
low=0, high=num_non_terminal_states, dtype=np.int64)
return init_s_idx
def epsilon_greedy_policy_from_state_action_function(
max_num_actions, q, epsilon, s_idx, policy):
"""Create epsilon-greedy policy from state-action value function.
Args:
max_num_actions: int, max number of actions possible.
q: array[float], keeps track of the estimated value of each
state-action pair Q_i(s, a).
epsilon: float, 0 <= epsilon <= 1, exploitation-exploration trade-off,
higher means more exploration.
s_idx: int, current state index.
policy: array[float], learned stochastic policy of which action a to
take in state s.
Returns:
policy: array[float], learned stochastic policy of which action a to
take in state s.
"""
# Combine state-action value functions
q = np.sum(a=q[:, s_idx, :], axis=0)
# Save max state-action value and find the number of actions that have the
# same max state-action value
max_action_value = np.max(a=q)
max_action_count = np.count_nonzero(a=q == max_action_value)
# Apportion policy probability across ties equally for state-action pairs
# that have the same value and zero otherwise
if max_action_count == max_num_actions:
max_policy_prob_per_action = 1.0 / max_action_count
remain_prob_per_action = 0.0
else:
max_policy_prob_per_action = (1.0 - epsilon) / max_action_count
remain_prob_per_action = epsilon / (max_num_actions - max_action_count)
policy[s_idx, :] = np.where(
q == max_action_value,
max_policy_prob_per_action,
remain_prob_per_action)
return policy
def loop_through_episode(
num_non_terminal_states,
max_num_actions,
num_state_action_successor_states,
sp_idx,
p,
r,
num_qs,
q,
policy,
alpha,
epsilon,
gamma,
maximum_episode_length,
s_idx):
"""Loops through episode to iteratively update policy.
Args:
num_non_terminal_states: int, number of non terminal states.
max_num_actions: int, max number of actions possible.
num_state_action_successor_states: array[int], number of successor
states s' that can be reached from state s by taking action a.
sp_idx: array[int], state indices of new state s' of taking action a
from state s.
p: array[float], transition probability to go from state s to s' by
taking action a.
r: array[float], reward from new state s' from state s by taking
action a.
num_qs: int, number of state-action-value functions Q_i(s, a).
q: array[float], keeps track of the estimated value of each
state-action pair Q_i(s, a).
policy: array[float], learned stochastic policy of which
action a to take in state s.
alpha: float, alpha > 0, learning rate.
epsilon: float, 0 <= epsilon <= 1, exploitation-exploration trade-off,
higher means more exploration.
gamma: float, 0 <= gamma <= 1, amount to discount future reward.
maximum_episode_length: int, max number of timesteps for an episode.
s_idx: int, current state index.
Returns:
q: array[float], keeps track of the estimated value of each
state-action pair Q_i(s, a).
policy: array[float], learned stochastic policy of which
action a to take in state s.
"""
# Loop through episode steps until termination
for t in range(0, maximum_episode_length):
# Choose policy for chosen state by epsilon-greedy choosing from the
# state-action-value function
policy = epsilon_greedy_policy_from_state_action_function(
max_num_actions, q, epsilon, s_idx, policy)
# Get epsilon-greedy action
a_idx = np.random.choice(
a=max_num_actions, p=policy[s_idx, :])
# Get reward
successor_state_transition_idx = np.random.choice(
a=num_state_action_successor_states[s_idx, a_idx],
p=p[s_idx, a_idx, :])
reward = r[s_idx, a_idx, successor_state_transition_idx]
# Get next state
next_s_idx = sp_idx[s_idx, a_idx, successor_state_transition_idx]
# Update state action value equally randomly selecting from the
# state-action-value functions
updating_q_idx = np.random.randint(low=0, high=num_qs, dtype=np.int64)
q, policy, s_idx = update_q(
num_non_terminal_states,
max_num_actions,
policy,
alpha,
epsilon,
gamma,
s_idx,
a_idx,
reward,
next_s_idx,
updating_q_idx,
num_qs,
q)
if next_s_idx >= num_non_terminal_states:
break # episode terminated since we ended up in a terminal state
return q, policy
def update_q(
num_non_terminal_states,
max_num_actions,
policy,
alpha,
epsilon,
gamma,
s_idx,
a_idx,
reward,
next_s_idx,
updating_q_idx,
num_qs,
q):
"""Updates state-action-value function using multiple estimates.
Args:
num_non_terminal_states: int, number of non terminal states.
max_num_actions: int, max number of actions possible.
policy: array[float], learned stochastic policy of which
action a to take in state s.
alpha: float, alpha > 0, learning rate.
epsilon: float, 0 <= epsilon <= 1, exploitation-exploration trade-off,
higher means more exploration.
gamma: float, 0 <= gamma <= 1, amount to discount future reward.
s_idx: int, current state index.
a_idx: int, current action index.
reward: float, current reward from taking action a_idx in state s_idx.
next_s_idx: int, next state index.
updating_q_idx: int, index to which Q_i(s, a) we'll be updating.
num_qs: int, number of state-action-value functions Q_i(s, a).
q: array[float], keeps track of the estimated value of each
state-action pair Q_i(s, a).
Returns:
q: array[float], keeps track of the estimated value of each
state-action pair Q_i(s, a).
policy: array[float], learned stochastic policy of which
action a to take in state s.
s_idx: int, new current state index.
"""
# Check to see if we actioned into a terminal state
if next_s_idx >= num_non_terminal_states:
delta = reward - q[updating_q_idx, s_idx, a_idx]
q[updating_q_idx, s_idx, a_idx] += alpha * delta
else:
# Get next action, using expectation value
q_indices = np.arange(num_qs)
not_updating_q_idx = np.random.choice(
a=np.extract(condition=q_indices != updating_q_idx, arr=q_indices))
not_updating_v_expected_value_on_policy = np.sum(
a=policy[next_s_idx, :] * q[not_updating_q_idx, next_s_idx, :])
# Calculate state-action-function expectation
delta = gamma * not_updating_v_expected_value_on_policy
delta -= q[updating_q_idx, s_idx, a_idx]
q[updating_q_idx, s_idx, a_idx] += alpha * (reward + delta)
# Update state and action to next state and action
s_idx = next_s_idx
return q, policy, s_idx
def on_policy_temporal_difference_n_tuple_expected_sarsa(
num_non_terminal_states,
max_num_actions,
num_state_action_successor_states,
sp_idx,
p,
r,
num_qs,
q,
policy,
alpha,
epsilon,
gamma,
maximum_episode_length,
num_episodes):
"""Loops through episodes to iteratively update policy.
Args:
num_non_terminal_states: int, number of non terminal states.
max_num_actions: int, max number of actions possible.
num_state_action_successor_states: array[int], number of successor
states s' that can be reached from state s by taking action a.
sp_idx: array[int], state indices of new state s' of taking action a
from state s.
p: array[float], transition probability to go from state s to s' by
taking action a.
r: array[float], reward from new state s' from state s by taking
action a.
num_qs: int, number of state-action-value functions Q_i(s, a).
q: array[float], keeps track of the estimated value of each
state-action pair Q_i(s, a).
policy: array[float], learned stochastic policy of which
action a to take in state s.
alpha: float, alpha > 0, learning rate.
epsilon: float, 0 <= epsilon <= 1, exploitation-exploration trade-off,
higher means more exploration.
gamma: float, 0 <= gamma <= 1, amount to discount future reward.
maximum_episode_length: int, max number of timesteps for an episode.
num_episodes: int, number of episodes to train over.
Returns:
q: array[float], keeps track of the estimated value of each
state-action pair Q_i(s, a).
policy: array[float], learned stochastic policy of which
action a to take in state s.
"""
for episode in range(0, num_episodes):
# Initialize episode to get initial state
init_s_idx = initialize_epsiode(num_non_terminal_states)
# Loop through episode and update the policy
q, policy = loop_through_episode(
num_non_terminal_states,
max_num_actions,
num_state_action_successor_states,
sp_idx,
p,
r,
num_qs,
q,
policy,
alpha,
epsilon,
gamma,
maximum_episode_length,
init_s_idx)
return q, policy
```
## Run algorithm
```
def run_algorithm():
"""Runs the algorithm."""
(num_states,
num_terminal_states,
num_non_terminal_states,
max_num_actions,
num_actions_per_non_terminal_state,
num_state_action_successor_states,
sp_idx,
p,
r) = create_environment()
(num_episodes,
maximum_episode_length,
num_qs,
alpha,
epsilon,
gamma) = set_hyperparameters()
q = create_value_function_arrays(num_qs, num_states, max_num_actions)
policy = create_policy_arrays(num_non_terminal_states, max_num_actions)
# Print initial arrays
print("\nInitial state-action value function")
print(q)
print("\nInitial policy")
print(policy)
# Run on policy temporal difference n-tuple expected sarsa
q, policy = on_policy_temporal_difference_n_tuple_expected_sarsa(
num_non_terminal_states,
max_num_actions,
num_state_action_successor_states,
sp_idx,
p,
r,
num_qs,
q,
policy,
alpha,
epsilon,
gamma,
maximum_episode_length,
num_episodes)
# Print final results
print("\nFinal state-action value function")
print(q)
print("\nFinal policy")
print(policy)
run_algorithm()
```
| github_jupyter |
This is an example showing the prediction latency of various scikit-learn estimators.
The goal is to measure the latency one can expect when doing predictions either in bulk or atomic (i.e. one by one) mode.
The plots represent the distribution of the prediction latency as a boxplot.
#### New to Plotly?
Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
<br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
<br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
### Version
```
import sklearn
sklearn.__version__
```
### Imports
This tutorial imports [StandardScaler](http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html#sklearn.preprocessing.StandardScaler), [train_test_split](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html#sklearn.model_selection.train_test_split), [scoreatpercentile](http://docs.scipy.org/doc/scipy-0.11.0/reference/generated/scipy.stats.scoreatpercentile.html#scipy.stats.scoreatpercentile), [make_regression](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.make_regression.html#sklearn.datasets.make_regression), [RandomForestRegressor](http://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestRegressor.html#sklearn.ensemble.RandomForestRegressor), [Ridge](http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.Ridge.html#sklearn.linear_model.Ridge), [SGDRegressor](http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.SGDRegressor.html#sklearn.linear_model.SGDRegressor), [SVR](http://scikit-learn.org/stable/modules/generated/sklearn.svm.SVR.html#sklearn.svm.SVR) and [shuffle](http://scikit-learn.org/stable/modules/generated/sklearn.utils.shuffle.html#sklearn.utils.shuffle).
```
from __future__ import print_function
from collections import defaultdict
from plotly import tools
import plotly.plotly as py
import plotly.graph_objs as go
import time
import gc
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from scipy.stats import scoreatpercentile
from sklearn.datasets.samples_generator import make_regression
from sklearn.ensemble.forest import RandomForestRegressor
from sklearn.linear_model.ridge import Ridge
from sklearn.linear_model.stochastic_gradient import SGDRegressor
from sklearn.svm.classes import SVR
from sklearn.utils import shuffle
```
### Calculations
```
fig1 = tools.make_subplots(rows=4, cols=1,
subplot_titles=(
'Prediction Time per instance - Atomic, 100 feats',
'Prediction Time per instance - Bulk(100), 100 feats',
'Evolution of Prediction Time with #Features ',
'Prediction Throughput for different estimators (%d '
'features)' % configuration['n_features']))
def _not_in_sphinx():
# Hack to detect whether we are running by the sphinx builder
return '__file__' in globals()
def atomic_benchmark_estimator(estimator, X_test, verbose=False):
"""Measure runtime prediction of each instance."""
n_instances = X_test.shape[0]
runtimes = np.zeros(n_instances, dtype=np.float)
for i in range(n_instances):
instance = X_test[[i], :]
start = time.time()
estimator.predict(instance)
runtimes[i] = time.time() - start
if verbose:
print("atomic_benchmark runtimes:", min(runtimes), scoreatpercentile(
runtimes, 50), max(runtimes))
return runtimes
def bulk_benchmark_estimator(estimator, X_test, n_bulk_repeats, verbose):
"""Measure runtime prediction of the whole input."""
n_instances = X_test.shape[0]
runtimes = np.zeros(n_bulk_repeats, dtype=np.float)
for i in range(n_bulk_repeats):
start = time.time()
estimator.predict(X_test)
runtimes[i] = time.time() - start
runtimes = np.array(list(map(lambda x: x / float(n_instances), runtimes)))
if verbose:
print("bulk_benchmark runtimes:", min(runtimes), scoreatpercentile(
runtimes, 50), max(runtimes))
return runtimes
def benchmark_estimator(estimator, X_test, n_bulk_repeats=30, verbose=False):
"""
Measure runtimes of prediction in both atomic and bulk mode.
Parameters
----------
estimator : already trained estimator supporting `predict()`
X_test : test input
n_bulk_repeats : how many times to repeat when evaluating bulk mode
Returns
-------
atomic_runtimes, bulk_runtimes : a pair of `np.array` which contain the
runtimes in seconds.
"""
atomic_runtimes = atomic_benchmark_estimator(estimator, X_test, verbose)
bulk_runtimes = bulk_benchmark_estimator(estimator, X_test, n_bulk_repeats,
verbose)
return atomic_runtimes, bulk_runtimes
def generate_dataset(n_train, n_test, n_features, noise=0.1, verbose=False):
"""Generate a regression dataset with the given parameters."""
if verbose:
print("generating dataset...")
X, y, coef = make_regression(n_samples=n_train + n_test,
n_features=n_features, noise=noise, coef=True)
random_seed = 13
X_train, X_test, y_train, y_test = train_test_split(
X, y, train_size=n_train, random_state=random_seed)
X_train, y_train = shuffle(X_train, y_train, random_state=random_seed)
X_scaler = StandardScaler()
X_train = X_scaler.fit_transform(X_train)
X_test = X_scaler.transform(X_test)
y_scaler = StandardScaler()
y_train = y_scaler.fit_transform(y_train[:, None])[:, 0]
y_test = y_scaler.transform(y_test[:, None])[:, 0]
gc.collect()
if verbose:
print("ok")
return X_train, y_train, X_test, y_test
def benchmark(configuration):
"""Run the whole benchmark."""
X_train, y_train, X_test, y_test = generate_dataset(
configuration['n_train'], configuration['n_test'],
configuration['n_features'])
stats = {}
for estimator_conf in configuration['estimators']:
print("Benchmarking", estimator_conf['instance'])
estimator_conf['instance'].fit(X_train, y_train)
gc.collect()
a, b = benchmark_estimator(estimator_conf['instance'], X_test)
stats[estimator_conf['name']] = {'atomic': a, 'bulk': b}
cls_names = [estimator_conf['name'] for estimator_conf in configuration[
'estimators']]
runtimes = [1e6 * stats[clf_name]['atomic'] for clf_name in cls_names]
boxplot_runtimes(runtimes, 'atomic', configuration, 1)
runtimes = [1e6 * stats[clf_name]['bulk'] for clf_name in cls_names]
boxplot_runtimes(runtimes, 'bulk (%d)' % configuration['n_test'],
configuration, 2)
def n_feature_influence(estimators, n_train, n_test, n_features, percentile):
"""
Estimate influence of the number of features on prediction time.
Parameters
----------
estimators : dict of (name (str), estimator) to benchmark
n_train : nber of training instances (int)
n_test : nber of testing instances (int)
n_features : list of feature-space dimensionality to test (int)
percentile : percentile at which to measure the speed (int [0-100])
Returns:
--------
percentiles : dict(estimator_name,
dict(n_features, percentile_perf_in_us))
"""
percentiles = defaultdict(defaultdict)
for n in n_features:
print("benchmarking with %d features" % n)
X_train, y_train, X_test, y_test = generate_dataset(n_train, n_test, n)
for cls_name, estimator in estimators.items():
estimator.fit(X_train, y_train)
gc.collect()
runtimes = bulk_benchmark_estimator(estimator, X_test, 30, False)
percentiles[cls_name][n] = 1e6 * scoreatpercentile(runtimes,
percentile)
return percentiles
def benchmark_throughputs(configuration, duration_secs=0.1):
"""benchmark throughput for different estimators."""
X_train, y_train, X_test, y_test = generate_dataset(
configuration['n_train'], configuration['n_test'],
configuration['n_features'])
throughputs = dict()
for estimator_config in configuration['estimators']:
estimator_config['instance'].fit(X_train, y_train)
start_time = time.time()
n_predictions = 0
while (time.time() - start_time) < duration_secs:
estimator_config['instance'].predict(X_test[[0]])
n_predictions += 1
throughputs[estimator_config['name']] = n_predictions / duration_secs
return throughputs
```
### Plot Results
Boxplot Runtimes
```
def boxplot_runtimes(runtimes, pred_type, configuration, subplot):
"""
Plot a new `Figure` with boxplots of prediction runtimes.
Parameters
----------
runtimes : list of `np.array` of latencies in micro-seconds
cls_names : list of estimator class names that generated the runtimes
pred_type : 'bulk' or 'atomic'
"""
cls_infos = ['%s<br>(%d %s)' % (estimator_conf['name'],
estimator_conf['complexity_computer'](
estimator_conf['instance']),
estimator_conf['complexity_label']) for
estimator_conf in configuration['estimators']]
box_plot1 = go.Box(y=runtimes[0],showlegend=False,name=cls_infos[0],
fillcolor='rgba(0.4,225, 128, 128)',
line=dict(color="black", width=1))
box_plot2 = go.Box(y=runtimes[1],showlegend=False,name=cls_infos[1],
fillcolor='rgba(0.4,225, 128, 128)',
line=dict(color="black", width=1))
box_plot3 = go.Box(y=runtimes[2],showlegend=False,name=cls_infos[2],
fillcolor='rgba(0.4,225, 128, 128)',
line=dict(color="black", width=1))
fig1.append_trace(box_plot1, subplot, 1)
fig1.append_trace(box_plot2, subplot, 1)
fig1.append_trace(box_plot3, subplot, 1)
axis='yaxis'+str(subplot)
fig1['layout'][axis].update(title='Prediction Time (us)')
axis='xaxis'+str(subplot)
fig1['layout'][axis].update(ticks='Prediction Time (us)')
```
Plot n_features influence.
```
def plot_n_features_influence(percentiles, percentile):
for i, cls_name in enumerate(percentiles.keys()):
x = np.array(sorted([n for n in percentiles[cls_name].keys()]))
y = np.array([percentiles[cls_name][n] for n in x])
line_plot = go.Scatter(x=x, y=y,
showlegend=False,
mode='lines',
line=dict(color="red"))
fig1.append_trace(line_plot, 3, 1)
fig1['layout']['xaxis3'].update(title='#Features')
fig1['layout']['yaxis3'].update(title='Prediction Time at %d%%-ile (us)' % percentile)
def plot_benchmark_throughput(throughputs, configuration):
fig, ax = plt.subplots(figsize=(10, 6))
cls_infos = ['%s<br>(%d %s)' % (estimator_conf['name'],
estimator_conf['complexity_computer'](
estimator_conf['instance']),
estimator_conf['complexity_label']) for
estimator_conf in configuration['estimators']]
cls_values = [throughputs[estimator_conf['name']] for estimator_conf in
configuration['estimators']]
bar_plot = go.Bar(x=cls_infos, y= cls_values,
showlegend=False, marker=dict(
color=['red', 'green', 'blue']))
fig1.append_trace(bar_plot, 4, 1)
fig1['layout']['yaxis4'].update(title='Throughput (predictions/sec)')
```
Plot data
```
start_time = time.time()
# benchmark bulk/atomic prediction speed for various regressors
configuration = {
'n_train': int(1e3),
'n_test': int(1e2),
'n_features': int(1e2),
'estimators': [
{'name': 'Linear Model',
'instance': SGDRegressor(penalty='elasticnet', alpha=0.01,
l1_ratio=0.25, fit_intercept=True),
'complexity_label': 'non-zero coefficients',
'complexity_computer': lambda clf: np.count_nonzero(clf.coef_)},
{'name': 'RandomForest',
'instance': RandomForestRegressor(),
'complexity_label': 'estimators',
'complexity_computer': lambda clf: clf.n_estimators},
{'name': 'SVR',
'instance': SVR(kernel='rbf'),
'complexity_label': 'support vectors',
'complexity_computer': lambda clf: len(clf.support_vectors_)},
]
}
benchmark(configuration)
# benchmark n_features influence on prediction speed
percentile = 90
percentiles = n_feature_influence({'ridge': Ridge()},
configuration['n_train'],
configuration['n_test'],
[100, 250, 500], percentile)
plot_n_features_influence(percentiles, percentile)
# benchmark throughput
throughputs = benchmark_throughputs(configuration)
plot_benchmark_throughput(throughputs, configuration)
stop_time = time.time()
print("example run in %.2fs" % (stop_time - start_time))
fig1['layout'].update(height=2000)
py.iplot(fig1)
```
### License
Authors:
Eustache Diemert <eustache@diemert.fr>
License:
BSD 3 clause
```
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'Prediction-Latency.ipynb', 'scikit-learn/plot-prediction-latency/', 'Prediction Latency | plotly',
' ',
title = 'Prediction Latency | plotly',
name = 'Prediction Latency',
has_thumbnail='true', thumbnail='thumbnail/prediction-latency.jpg',
language='scikit-learn', page_type='example_index',
display_as='real_dataset', order=9,ipynb='~Diksha_Gabha/2674')
```
| github_jupyter |
# Library design for CTP-04, 25kb Chr21 tracing 7x21
by Pu Zheng and Jun-Han Su
This library design is for human chr21 tracing at 25 kb resolution
```
#minimum imports:
import time,os,sys,glob
import cPickle as pickle
import numpy as np
import khmer
sys.path.append(r'/n/home13/pzheng/Documents/python-functions/python-functions-library')
from LibraryConstruction import fastaread,fastawrite,fastacombine
import LibraryDesigner as ld
import LibraryConstruction as lc
```
## 3 Assign region into TADs
```
def Match_TADs(report_folder, TAD_ref, save_folder, save=True, verbose=True):
'''Function to match regions with a TAD reference
Inputs:
report_folder: directory for extracted probe files, string
TAD_ref: full filename for TAD reference, string
save_folder: directory for saving results id->TAD, string
save: whether save result, bool (default: True)
verbose: wether say something! bool (default: True)
Outputs:
'''
import os, glob, sys
import LibraryDesigner as ld
import numpy as np
import cPickle as pickle
def Read_TAD_ref(TAD_ref=TAD_ref):
_tad_dics = [];
with open(TAD_ref) as _ref_handle:
_lines = _ref_handle.readlines();
for _line in _lines:
_chrom = _line.split(':')[0]
_reg_str = _line.split(':')[1].split('\n')[0];
_start,_stop = _reg_str.split('-');
_tad_dic = {'chr':_chrom, 'start':int(_start), 'stop':int(_stop)}
_tad_dics.append(_tad_dic);
return sorted(_tad_dics, key=lambda d:d['start']);
def Region_to_TAD(tad_dics, report_filename):
_pb = ld.pb_reports_class()
_pb.load_pbr(report_filename)
# get its region status
_reg_id = int(_pb.pb_reports_keep.values()[0]['reg_name'].split('reg')[1].split('_')[1])
_chrom = _pb.pb_reports_keep.values()[0]['reg_name'].split(':')[0]
_start, _stop = _pb.pb_reports_keep.values()[0]['reg_name'].split(':')[1].split('_')[0].split('-')
_start = int(_start);
_stop = int(_stop);
if _start > _stop:
_start, _stop = _stop, _start
_reg_len = abs(_stop - _start)
# initialize tad identity of this region
_tad_id = -1;
for i in range(len(tad_dics)):
_dic = tad_dics[i];
if _chrom == _dic['chr']:
_overlap = min(_stop, _dic['stop']) - max(_start, _dic['start']);
if _overlap > _reg_len / 2:
_tad_id = i; # assign tad id
break
return _reg_id, _tad_id, len(_pb.pb_reports_keep)
def Extra_Region_Assigning(tad_id_dic):
'''Try to assign region to TADs as much as possible
'''
# calculate how many region has been assigned to each TAD
_v,_c = np.unique(tad_id_dic.values(),return_counts=True)
_reg_num_dic = dict(zip(_v,_c)) # dictionary for region number of each TAD
# maximum gap size to be filled
_gap_max = 4
# new_id_dic
_new_id_dic = tad_id_dic.copy();
# Starting filling gaps!
_gap = 0;
_prev_value = -1;
for _key, _value in sorted(_new_id_dic.items()):
# start a gap
if _gap == 0 and _value == -1:
_prev_tad = _prev_value
_gap = 1; # turn on gap
_key_ingap = [_key] # start recording keys in gap
# continue a gap
elif _gap == 1 and _value == -1:
_key_ingap.append(_key)
# stop a gap!
elif _gap == 1 and _value > -1:
_gap = 0; # stop counting gap
_next_tad = _value
# if the gap is not huge, try to make up
if len(_key_ingap) <= _gap_max:
if _prev_tad == -1: # don't fill any gap at beginning
continue
elif len(_key_ingap)/2*2 == len(_key_ingap): # gap size is even number
for i in range(len(_key_ingap)/2):
_new_id_dic[_key_ingap[i]] = _prev_tad
_new_id_dic[_key_ingap[i+len(_key_ingap)/2]] = _next_tad
else: # gap size is odd number
for i in range(len(_key_ingap)/2):
_new_id_dic[_key_ingap[i]] = _prev_tad
_new_id_dic[_key_ingap[i+len(_key_ingap)/2+1]] = _next_tad
if _reg_num_dic[_prev_tad] <= _reg_num_dic[_next_tad]:
_new_id_dic[_key_ingap[len(_key_ingap)/2]] = _prev_tad
else:
_new_id_dic[_key_ingap[len(_key_ingap)/2]] = _next_tad
_prev_value = _value # store previous tad info
return _new_id_dic
def Save_dics(save_folder, tad_dics, reg_len_dic, new_id_dic):
# save tad dics
tad_dic_file = open(save_folder+os.sep+'TAD_dic_list.pkl','w');
pickle.dump(tad_dics, tad_dic_file);
tad_dic_file.close()
# save region length dic
reg_len_dic_file = open(save_folder+os.sep+'region_length.pkl','w');
pickle.dump(reg_len_dic, reg_len_dic_file);
reg_len_dic_file.close()
# save region_to_tad dic
reg_to_tad_file = open(save_folder+os.sep+'region_to_TAD.pkl','w');
pickle.dump(new_id_dic, reg_to_tad_file);
reg_to_tad_file.close()
if verbose:
print '- Start reading TAD reference', TAD_ref
tad_dics = Read_TAD_ref() # load tad info
if verbose:
print "-- Number of Tads in reference:", len(tad_dics)
if verbose:
print '- Start reading probe reports'
files = glob.glob(report_folder + os.sep + '*.pbr') # extract all probe files
tad_id_dic = {} # store assigned tad id
reg_len_dic = {} # store number of probes in each region
for _file in sorted(files, key=lambda fl:int(fl.split('.pbr')[0].split('_')[-1])):
reg_id, tad_id, reg_len = Region_to_TAD(tad_dics, _file)
tad_id_dic[reg_id] = tad_id; # update tad id dic
reg_len_dic[reg_id] = reg_len; # update region length dic
if verbose:
print '--', os.path.basename(_file), 'tad_id:', tad_id, 'size:', reg_len
# update tads by putting gaps
new_id_dic = Extra_Region_Assigning(tad_id_dic)
if save:
Save_dics(save_folder=save_folder,
tad_dics=tad_dics,
reg_len_dic=reg_len_dic,
new_id_dic=new_id_dic);
return tad_dics, tad_id_dic, reg_len_dic, new_id_dic
# get previous outputs
max_pb_num = 400;
min_pb_num = 150;
# probes from 9x36
probe_folder=r'/n/boslfs/LABS/zhuang_lab/User/pzheng/Libraries/CTP-04/chr21_9by36/reports/centered_merged400';
# this library is 7x21
library_folder = r'/n/boslfs/LABS/zhuang_lab/User/pzheng/Libraries/CTP-04/chr21_7by21';
tad_dics, tad_id_dic, reg_len_dic, new_id_dic= Match_TADs(probe_folder,
TAD_ref=library_folder+os.sep+'chr21_TADs.bed',
save_folder=library_folder)
```
## 4 Generate encoding scheme
```
def Generate_Encoding(master_folder, reg_id_dic,
total_encoding={}, sub_encoding={},
n_color=3, n_reg=10, n_hyb=5, min_region_times=2,
sequential_for_nonencoding=True,
save=True, save_filename='other_encoding', verbose=True):
'''Design encoding scheme for all excluded regions in total_encoding from sub_encoding
Inputs:
master_folder, directory for this library, string
reg_id_dic: dictionary for region id -> tad id, dic
total_encoding: dictionary for total encoding generated by previous code, if not given then design a new encoding dic
sub_encoding: sub encoding in test experiment, if not given then design a new encoding dic
n_color: number of colors, int
n_reg: number of region per decoding unit, int
n_hyb: number of hybes per decoding unit, int
min_region_times: minimum region appearing times, int
save: whether save final result, bool
save_filename: filename of pickle file of this encoding scheme, string
verbose: whether say something!, bool
Outputs:
other_encoding: region_number -> color=i, cluster=j, region=k, barcodes->...
hyb_matrix: hybridization matrix, n_reg by n_hyb
assign_regs: matrix of assigning region to clusters, n_color by n_cluster by n_reg
assign_tads: matrix of assigning tad to clusters, n_color by n_cluster by n_reg
'''
def _Compare_Encodings(_total_encoding=total_encoding, _sub_encoding=sub_encoding, subset=True, _verbose=verbose):
'''Function to compare two encoding scheme'''
if _verbose:
print "-- comparing two encoding schemes with size", len(_total_encoding.keys()),'and',len(_sub_encoding.keys())
# check whether sub_encoding is really a subset of total_encoding
for _k,_v in sorted(_sub_encoding.items()):
if _k not in _total_encoding:
if subset:
raise ValueError('-- key '+str(k)+' not exists in total encoding scheme')
else:
raise Warning('-- key '+str(k)+' not exists in total encoding scheme')
# extract existing regions
_other_encoding = {}
for _k,_v in sorted(_total_encoding.items()):
if _k not in _sub_encoding:
_other_encoding[_k] = _v
return _other_encoding
def _TAD_to_Region(reg_id_dic, _reg_encodings, _verbose=verbose):
'''Function to inverse region->TAD dictionary'''
if _verbose:
print '-- Converting region->TAD dic into TAD->[regions]';
_tad_to_region = {}
for k, v in reg_id_dic.iteritems():
if k in _reg_encodings:
_tad_to_region[v] = _tad_to_region.get(v, [])
_tad_to_region[v].append(k)
_tad_to_region.pop(-1, None);
return _tad_to_region;
def _Generate_Hyb_Matrix(n_reg=n_reg, n_hyb=n_hyb, min_region_times=min_region_times, _verbose=verbose):
'''Function to generate hybridization matrix
Input: number of regions
number of hybridizations
the minimal time that each region appears. default:1
Output: A hybridization matrix'''
if _verbose:
print '-- Generating hybridization matrix for region='+str(n_reg)+', hyb='+str(n_hyb);
# generate all possible all_codess
all_codes =[] # list for all possible binary all_codess
for i in range(2**n_hyb):
hybe_0 = np.zeros(n_hyb,dtype=int)
binrep = [int(c) for c in str("{0:#b}".format(i))[2:]]
#print str("{0:#b}".format(i))[2:]
hybe_0[-len(binrep):]=binrep
all_codes.append(hybe_0)
all_codes = np.array(all_codes)
all_codes = all_codes[np.sum(all_codes,-1)>0]
# Choose candicate codes
_code_sums = np.sum(all_codes,axis=-1)
_code_sums[_code_sums < min_region_times]=np.max(_code_sums)+1 # remove codes that dont satisfy minimal region showup times
_max_region_time = np.sort(_code_sums)[n_reg] # maximum region appearance
if min_region_times == _max_region_time: # Case 1: all regions has the same code
_nchoose = n_reg
_cand_codes = all_codes[_code_sums == _max_region_time];
_sims = []
for _i in range(20000):
_sim = _cand_codes[np.random.choice(range(len(_cand_codes)), _nchoose, replace=False)]
_sims.append(_sim)
_sim_keep = _sims[np.argmin([np.var(np.sum(_sim,axis=0)) for _sim in _sims])]
_hyb_matrix = np.array(list(_sim_keep))
else: # Case 2: use lower-choose codes first, and then use higher codes
_used_codes = list(all_codes[_code_sums < _max_region_time]) # use up all shorter codes
_nchoose = n_reg-len(_used_codes) # other codes to be chosen
_cand_codes = all_codes[_code_sums == _max_region_time]
_sims = []
for _i in range(20000):
_sim = _cand_codes[np.random.choice(range(len(_cand_codes)), _nchoose, replace=False)]
_sims.append(_sim)
_sim_keep = _sims[np.argmin([np.var(np.sum(_sim,axis=0)) for _sim in _sims])]
_used_codes+=list(_sim_keep)
_hyb_matrix = np.array(_used_codes).astype(np.int)
return _hyb_matrix
def _Assign_Cluster(_reg_encodings, _tad_to_region, _n_reg=n_reg, _n_color=n_color,
_sequential_for_nonencoding=sequential_for_nonencoding, _verbose=verbose):
'''Assign regions into clusters'''
from math import ceil
from copy import copy
if _verbose:
print '-- Assigning clusters for all regions';
# region color
_reg_colors = [[] for _color in range(_n_color)]
_mode_counter = 0; # used for balancing mode_n results into n categories
# split regions in each tad into different colors
for _k,_v in sorted(_tad_to_region.items()):
for _color in range(_n_color):
_reg_list = _v[(_mode_counter+_color)%_n_color::_n_color];
_reg_colors[_color].append(_reg_list);
_mode_counter += 1
for _color in range(_n_color):
print "--- Number of regions in channel", str(_color)+":", sum([len(_l) for _l in _reg_colors[_color]])
# calculate number of clusters in each color
_n_cluster = int(ceil(len(_reg_encodings)/float(_n_color*n_reg)))
# initialize encoding matrix
_assign_regs = [-np.ones([_n_cluster, _n_reg],dtype=np.int) for _color in range(n_color)]
_left_regs = [[] for _color in range(_n_color)];
# loop through each color and each group to assign regions into encodings
for _color in range(_n_color):
_rlist = copy(sorted(_reg_colors[_color],key=lambda v:-len(v)));
_cluster = 0;
while len(_rlist) >= _n_reg:
for _reg in range(_n_reg):
_assign_regs[_color][_cluster, _reg] = _rlist[_reg].pop(0)
# store into reg_encodings
_reg_encodings[_assign_regs[_color][_cluster, _reg]]['color'] = _color;
_reg_encodings[_assign_regs[_color][_cluster, _reg]]['cluster'] = _cluster;
_reg_encodings[_assign_regs[_color][_cluster, _reg]]['region'] = _reg;
# clean all empty lists
while [] in _rlist:
_rlist.remove([]);
# sort again
_rlist = sorted(_rlist, key=lambda v:(-len(v),v[0]));
# next cluster
_cluster += 1
print len(_rlist)
if len(_rlist) < 34:
print [len(_lst) for _lst in _rlist]
# for the rest of unassigned regions
if _rlist:
# check if the rest of unassigned regions can fit into decoding scheme
if False in [len(_lst) == 1 for _lst in _rlist]:
for _lst in _rlist:
if not _sequential_for_nonencoding:
_left_regs[_color] += _lst
for _reg in _lst:
_reg_encodings[_reg]['color'] = _color
if _sequential_for_nonencoding:
_reg_encodings[_reg]['cluster'] = 'sequential'
_reg_encodings[_reg]['region'] = 'sequential'
_left_regs[_color] = np.array( _left_regs[_color])
else: # every region can fit into
_assign_regs[_color][ _cluster, :len(_rlist)] = np.array(_rlist).reshape(-1) # store the rest
for _left_reg in range(len(_rlist)):
_reg_encodings[_assign_regs[_color][_cluster, _left_reg]]['color'] = _color;
_reg_encodings[_assign_regs[_color][_cluster, _left_reg]]['cluster'] = _cluster;
_reg_encodings[_assign_regs[_color][_cluster, _left_reg]]['region'] = _left_reg;
# remove empty rows
for _row, _row_regs in enumerate(_assign_regs[_color]):
if (_row_regs < 0).all():
break
if _verbose:
print "--", _row, 'groups of encoding in color', _color
_assign_regs[_color] = _assign_regs[_color][slice(0,_row)]
return _reg_encodings, _assign_regs, _left_regs;
def _Assign_Decoding_Barcodes(_reg_encodings, assign_regs, hyb_matrix,
n_color=n_color, n_reg=n_reg, n_hyb=n_hyb,
_sequential_for_nonencoding=sequential_for_nonencoding,
_verbose=verbose):
'''Assign barcode (orders) used for decoding'''
if _verbose:
print '-- Assigning decoding barcodes.'
# Sanity check
if len(assign_regs) != n_color:
raise EOFError('wrong number of colors!');
for _a_regs in assign_regs:
if _a_regs.shape[1] != n_reg:
raise EOFError('wrong region assignment dimention for each group')
_barcode_set = 0; # barcode to be assigned
for _color in range(n_color):
# collect number of clusters per color
n_cluster = np.shape(assign_regs[_color])[0];
for _cluster in range(n_cluster):
for _reg in range(n_reg):
if assign_regs[_color][_cluster,_reg] >= 0:
_reg_encodings[assign_regs[_color][_cluster,_reg]]['bc_decoding'] = [n_hyb*_barcode_set+ i for i, j in enumerate(hyb_matrix[_reg]) if j == 1]
_barcode_set += 1; # next barcode set (size of n_hyb)
# assign sequential for encoding ones
if _sequential_for_nonencoding:
_next_barcode = _barcode_set * n_hyb
for _reg,_info in sorted(_reg_encodings.items()):
if _info['cluster'] == 'sequential' or _info['region'] == 'sequential' and not _info['bc_decoding']:
_reg_encodings[_reg]['bc_decoding'] = [_next_barcode for _i in range(np.max((hyb_matrix > 0).sum(1)))]
_next_barcode += 1
return _reg_encodings;
def _Check_Decoding_Barcodes(_reg_encodings, hyb_matrix,
_sequential_for_nonencoding=sequential_for_nonencoding, _verbose=verbose):
'''Function to check whether decoding barcode works fine'''
if _verbose:
print '--- Checking decoding barcodes.'
reg_bc_num=hyb_matrix.sum(1).max()
hyb_bc_num=hyb_matrix.sum(0).max()
bc_list = [];
for k,v in _reg_encodings.iteritems():
if not _sequential_for_nonencoding:
if not v['bc_decoding']:
continue
if len(v['bc_decoding']) > reg_bc_num or len(v['bc_decoding']) <=0:
print '--- wrong barcode size per region';
return False
bc_list += v['bc_decoding'];
# record unique barcodes
barcodes, barcode_counts = np.unique(bc_list, return_counts=True)
print barcodes
# check barcode usage per hybe
validate = False not in [n<=hyb_bc_num and n>0 for n in barcode_counts]
print '---', validate
return validate
def _Assign_Unique_Barcodes(_reg_encodings, _verbose=verbose):
'''Assign barcode (orders) used for unique sequential'''
if _verbose:
print '-- Assigning unique barcodes.'
# record all decoding barcodes and TAD barcodes
used_bcs = []
for k,v in _reg_encodings.iteritems():
if v['bc_decoding']:
used_bcs += v['bc_decoding']
# unique barcodes should start right after
unique_bc_start = max(used_bcs);
reg_new_id = 0;
for k,v in sorted(_reg_encodings.items()):
_reg_encodings[k]['bc_unique'] = reg_new_id + unique_bc_start;
_reg_encodings[k]['id'] = reg_new_id;
reg_new_id += 1;
return _reg_encodings
# Initialize
if not total_encoding:
if verbose:
print "- Initializing";
reg_encodings = {};
for key, value in reg_id_dic.items():
if value >= 0:
reg_encodings[key] = {'TAD':value, 'id':None, 'color':None,
'cluster':None, 'region': None,
'bc_decoding':None, 'bc_unique':None}
else: # total encoding is given
if verbose:
print "- Loading total encoding"
reg_encodings = total_encoding;
if sub_encoding: # sub encoding is given
if verbose:
print "- Excluding regions in sub encoding."
reg_encodings = _Compare_Encodings(reg_encodings, sub_encoding)
if verbose:
print "-- number of regions in current encoding scheme:", len(reg_encodings)
# prepare dictionary and hyb matrix
if verbose:
print "- Prepare dictionary and hyb matrix";
# creat tad to region dictionary
tad_to_region = _TAD_to_Region(reg_id_dic, reg_encodings);
# generate hybe matrix
hyb_matrix = _Generate_Hyb_Matrix()
if verbose:
print "- Calculate color, cluster assignment";
# assign cluster
reg_encodings, assign_regs, left_regs = _Assign_Cluster(reg_encodings, tad_to_region);
if verbose:
print "- Assign barcodes";
# assign decoding barcodes
reg_encodings = _Assign_Decoding_Barcodes(reg_encodings, assign_regs, hyb_matrix)
# check decoding barcodes
decoding_check = _Check_Decoding_Barcodes(reg_encodings, hyb_matrix)
# assign unique barcodes
reg_encodings = _Assign_Unique_Barcodes(reg_encodings)
if save:
import cPickle as pickle
import os
save_filename = master_folder + os.sep + str(save_filename)+'.pkl';
if verbose:
print "- Save to file:", save_filename
savefile = open(save_filename, 'w');
pickle.dump(reg_encodings, savefile)
return reg_encodings, hyb_matrix, assign_regs, left_regs
# You can continue here!
region_folder=r'/n/boslfs/LABS/zhuang_lab/User/pzheng/Libraries/CTP-04/chr21_7by21';
# dic for region -> tad
if not 'new_id_dic' in vars():
new_id_dic = pickle.load(open(region_folder+os.sep+'region_to_TAD.pkl','r'))
total_encoding, hyb_matrix, assign_regs, left_regs = Generate_Encoding(region_folder,
new_id_dic,
n_color=3, n_reg=21, n_hyb=7,
sequential_for_nonencoding=False,
save_filename='total_encoding')
```
## Extract subset
```
def Sub_Library_Encoding(total_encoding, hyb_matrix, assign_regs, reg_id_dic,
sub_library_size,
min_reg_in_tad=2,
random_select=True,
save=True, save_dir=r'/n/boslfs/LABS/zhuang_lab/User/pzheng/Libraries/CTP-04/chr21_7by21',
generate_other_encoding=True,
verbose=True):
'''Extract a sub library for total library and redesign encodings
Inputs:
_reg_encoding: region_number -> color=i, cluster=j, region=k, barcodes->...
hyb_matrix: hybridization matrix, n_reg by n_hyb
assign_regs: matrix of assigning region to clusters, n_color by n_cluster by n_reg
reg_id_dic: dictionary for region -> tad, dic
sub_library_size: number of regions in the sub library, int
min_reg_in_tad: criteria for selecting sub library, at least 2 regions in each new tad, int
save: whether save, bool
save_dir: directory for saving, str
generate_other_encoding: whether generate encoding for the rest of regions, bool (default: True)
verbose: whether say something!, bool
Outputs:
sub_encodings: encoding scheme for sub library
other_encodings: encoding scheme for the rest of library
'''
# imports
import numpy as np;
def _TAD_in_Cluster(_assign_regs, reg_id_dic=reg_id_dic, _verbose=verbose):
# input parameters
n_color = len(_assign_regs); # number of colors
# initialize
_assign_tads = [-np.ones(np.shape(_assign_regs[_color]), dtype=np.int) for _color in range(n_color)]
for _color in range(n_color):
n_cluster = _assign_regs[_color].shape[0]; # number of clusters per color
n_reg = _assign_regs[_color].shape[1]; # number of regions, defined by hyb matrix
for _cluster in range(n_cluster):
for _reg in range(n_reg):
if _assign_regs[_color][_cluster, _reg] >= 0:
_assign_tads[_color][_cluster, _reg] = reg_id_dic[_assign_regs[_color][_cluster, _reg]]
return _assign_tads;
def _Select_Sub_Encodings(total_encoding=total_encoding, assign_regs=assign_regs,
sub_library_size=sub_library_size, min_reg_in_tad=min_reg_in_tad,
_verbose=verbose):
if _verbose:
print "-- Starting sub library searching";
# convert assign_cluster into assign_tad
assign_tads = _TAD_in_Cluster(assign_regs);
# get all available tads
_all_tads = np.unique(np.concatenate(assign_tads,0))
_all_tads = _all_tads[_all_tads >= 0]
print '--- all available tads:',len(_all_tads)
# record parameters
n_color = len(assign_regs); # number of colors
n_reg = assign_regs[0].shape[1]; # number of regions, defined by hyb matrix
_select_clusters = int(sub_library_size / n_reg) # number total selected clusters (in all colors)
if _verbose:
print "--- color: "+str(n_color), "region: "+str(n_reg), "selected clusters: "+str(_select_clusters)
# Split select clusters in different colors equally
n_chooses = []
for _color in range(n_color):
_choose = (_select_clusters-sum(n_chooses)) / (n_color-_color)
n_chooses.append(_choose)
n_chooses = sorted(n_chooses)
n_chooses.reverse()
print n_chooses
if random_select: # Randomly generate region picking
j=0
min_reg = 0
else:
j=0
_cids = [range(n_ch) for n_ch in n_chooses]
_ctads = [assign_tads[_color][:n_ch,:] for _color, n_ch in zip(range(n_color),n_chooses)]
_tads, _cts = np.unique(np.concatenate(_ctads,1), return_counts=True);
if len(_tads) == len(_all_tads):
min_reg = np.min(_cts) # the minimum occurance of TADs
else:
min_reg = 0
while min_reg < min_reg_in_tad:
_cids = []; # chosen id list
_ctads = []; # chosen tad matrix parts
for _color in range(n_color):
n_cluster = assign_regs[_color].shape[0]
if (assign_regs[_color][-1,:] > 0).all():
# chosen ids
_cids.append([sorted(np.random.choice(n_cluster, n_chooses[_color], replace=False))])
else:
# chosen ids
_cids.append([sorted(np.random.choice(n_cluster-1, n_chooses[_color], replace=False))])
# chosen tads
_ctads.append(assign_tads[_color][_cids[_color], :]);
# get unique set
_tads, _cts = np.unique(np.concatenate(_ctads,1), return_counts=True);
# start updating once all TADs show up
if len(_tads) == len(_all_tads):
min_reg = np.min(_cts) # the minimum occurance of TADs
j+=1;
if _verbose:
print "--- Number of searches:", j;
print "-- Finishing library searching, constructing sub library";
# Storing information into reg matrix
_sub_regs = -np.ones([n_color, n_chooses[0], n_reg]);
_other_regs = -np.ones([n_color, n_cluster-n_chooses[-1], n_reg]);
for _color in range(n_color):
_sub_regs[_color,:n_chooses[_color],:] = assign_regs[_color][_cids[_color],:] # sub region
_oid = list(set(np.arange(n_cluster)) - set(sorted(np.random.choice(22,5,replace=False)))) #other region
_other_regs[_color,:len(_oid),:] = assign_regs[_color][_oid, :];
# Initialize encoding region list
_sub_encodings, _other_encodings = {},{};
for _r in np.unique(_sub_regs):
if _r >=0:
#_sub_encodings[int(_r)] = total_encoding[int(_r)];
_sub_encodings[int(_r)] = {'TAD':total_encoding[int(_r)]['TAD'],
'color':total_encoding[int(_r)]['color'],
'cluster':None,
'id':None,
'region':total_encoding[int(_r)]['region'],
'bc_decoding':None, 'bc_unique':None}
for _r in np.unique(_other_regs):
if _r >=0:
#_other_encodings[int(_r)] = total_encoding[int(_r)];
_other_encodings[int(_r)] = {'TAD':total_encoding[int(_r)]['TAD'],
'color':total_encoding[int(_r)]['color'],
'cluster':None,
'id':None,
'region':total_encoding[int(_r)]['region'],
'bc_decoding':None, 'bc_unique':None}
return _sub_encodings, _sub_regs, _other_encodings, _other_regs
def _Assign_All_Barcodes(_reg_encodings, _assign_regs, _hyb_matrix=hyb_matrix, _verbose=verbose):
'''Assembled function to update all barcodes'''
# record parameters
n_color = _assign_regs.shape[0]; # number of colors
n_cluster = _assign_regs.shape[1]; # number of groups per color
n_reg = _hyb_matrix.shape[0]; # number of regions per cluster, defined by hyb matrix
n_hyb = _hyb_matrix.shape[1]; # number of hybes per cluster
if _verbose:
print "--- color: "+str(n_color), "region: "+str(n_reg), "hybs: "+str(n_hyb);
def _Assign_Decoding_Barcodes(_reg_encodings, _assign_regs=_assign_regs, _hyb_matrix=_hyb_matrix,
n_color=n_color, n_cluster=n_cluster,
n_reg=n_reg, n_hyb=n_hyb, _verbose=verbose):
'''Assign barcode (orders) used for decoding'''
if _verbose:
print '-- Assigning decoding barcodes.'
# Sanity check
if np.shape(_assign_regs)[0] != n_color or np.shape(_assign_regs)[2] != n_reg:
raise EOFError('wrong input dimension!');
# collect number of clusters per color
_barcode_set = 0; # barcode to be assigned
for _color in range(n_color):
for _cluster in range(n_cluster):
for _reg in range(n_reg):
if _assign_regs[_color,_cluster,_reg] >= 0:
_reg_encodings[_assign_regs[_color,_cluster,_reg]]['cluster'] = _cluster
_reg_encodings[_assign_regs[_color,_cluster,_reg]]['bc_decoding'] = [n_hyb*_barcode_set+i for i, j in enumerate(_hyb_matrix[_reg]) if j == 1]
#print [n_hyb*_barcode_set+i for i, j in enumerate(_hyb_matrix[_reg]) if j == 1]
_barcode_set += 1; # next barcode set (size of n_hyb)
return _reg_encodings;
def _Assign_Unique_Barcodes(__reg_encodings, _verbose=verbose):
'''Assign barcode (orders) used for unique sequential'''
if _verbose:
print '-- Assigning unique barcodes.'
# record all decoding barcodes and TAD barcodes
used_bcs = []
for k,v in sorted(__reg_encodings.items()):
if v['bc_decoding']:
used_bcs += v['bc_decoding']
unique_bc_start = max(used_bcs)+1;
reg_new_id = 0;
for k,v in sorted(__reg_encodings.items()):
__reg_encodings[k]['bc_unique'] = reg_new_id + unique_bc_start;
__reg_encodings[k]['id'] = reg_new_id;
reg_new_id += 1;
return __reg_encodings
# assign decoding barcodes
_reg_encodings = _Assign_Decoding_Barcodes(_reg_encodings)
# assign unique barcodes
_reg_encodings = _Assign_Unique_Barcodes(_reg_encodings)
return _reg_encodings
# Select sub library
if verbose:
print "- Select sub library."
sub_encodings, sub_regs, other_encodings, other_regs= _Select_Sub_Encodings()
# Re_assign barcodes
if verbose:
print "- Reassign barcodes for sub library."
sub_encodings = _Assign_All_Barcodes(sub_encodings, sub_regs);
if generate_other_encoding:
if verbose:
print "- Reassign barcodes for the rest of library."
other_encodings = _Assign_All_Barcodes(other_encodings, other_regs);
if save:
import cPickle as pickle
import os
sub_filename = save_dir + os.sep + 'sub_encoding.pkl';
if verbose:
print "- Save to file:", sub_filename
# save
pickle.dump(sub_encodings, open(sub_filename,'w'))
if generate_other_encoding:
other_filename = save_dir + os.sep + 'other_encoding.pkl';
if verbose:
print "- Save to file:", other_filename
# save
pickle.dump(other_encodings, open(other_filename,'w'))
if generate_other_encoding:
return sub_encodings, other_encodings
else:
return sub_encodings
sub_encoding = Sub_Library_Encoding(total_encoding, hyb_matrix, assign_regs, reg_id_dic, 126,
min_reg_in_tad=2,
generate_other_encoding=False, save_dir=region_folder)
```
## 3.5 Decoding barcode check
```
def _Check_Decoding_Barcodes(reg_encodings, hyb_matrix, _verbose=True):
'''Function to check whether decoding barcode works fine'''
if _verbose:
print '--- Checking decoding barcodes.'
reg_bc_num=hyb_matrix.sum(1).max()
hyb_bc_num=hyb_matrix.sum(0).max()
bc_list = [];
for k,v in reg_encodings.iteritems():
if len(v['bc_decoding']) > reg_bc_num or len(v['bc_decoding']) <=0:
print '--- wrong barcode size per region';
return False
bc_list += v['bc_decoding'];
# record unique barcodes
barcodes, barcode_counts = np.unique(bc_list, return_counts=True)
print barcodes
print barcode_counts
# check barcode usage per hybe
validate = False not in [n<=hyb_bc_num and n>0 for n in barcode_counts]
print '---', validate
return validate
_Check_Decoding_Barcodes(sub_encoding, hyb_matrix)
```
# 4. Patch Barcode Sequence to Reads
```
# minimal imports for biopython
from Bio import SeqIO
from Bio.Seq import Seq
from Bio.Alphabet import IUPAC
from Bio.SeqRecord import SeqRecord
import os,glob,time
import numpy as np
```
## 4.2 Read all PCR primers
```
primer_dir = r'/n/boslfs/LABS/zhuang_lab/User/pzheng/Primers';
fwd_primer_filename = 'forward_primers_keep.fasta';
rev_primer_filename = 'reverse_primers_keep.fasta';
# read all forward primers
with open(primer_dir+os.sep+fwd_primer_filename, "rU") as handle:
fwd_primers = [];
for record in SeqIO.parse(handle, "fasta"):
fwd_primers.append(record);
# read all forward primers
with open(primer_dir+os.sep+rev_primer_filename, "rU") as handle:
rev_primers = [];
for record in SeqIO.parse(handle, "fasta"):
rev_primers.append(record);
print "Primers loaded: forward: "+str(len(fwd_primers))+", reverse: "+str(len(rev_primers));
```
## 4.3 read all probe reports and generate primary probes
```
import cPickle as pickle
library_folder =r'/n/boslfs/LABS/zhuang_lab/User/pzheng/Libraries/CTP-04/chr21_7by21';
fprimer = fwd_primers[1];
print '- forward primer:', fprimer
rprimer = rev_primers[4];
print '- reverse primer:', rprimer
def Patch_Barcodes_by_Color(reg_encodings, probe_folder, library_folder,
fwd_primer,rev_primer,
barcode_folder=r'/n/boslfs/LABS/zhuang_lab/User/pzheng/Barcodes',
filename_combo='combo_readouts_', filename_unique='unique_readouts_',
bc_combo='bc_decoding', bc_unique='bc_unique',
save_subfolder=r'final_probes',
add_rand_gap=0,
save=True, verbose=True):
'''Function to patch barcodes to designed probes
Inputs:
reg_encodings: encoding scheme for the barcode, dictionary(generated previously)
library_folder: master directory for this library, string
report_subfolder: sub-directory for probe reports, string
barcode_folder: directory for barcode files, string
filename_combo: readout filename for combo, string
filename_unique: readout filename for unique, string
bc_combo: key name for combo, string
bc_unique: key name for unique, string
save_subfolder: sub-directory for save files, string
add_rand_gap: whether adding (or length) of random gaps between barcodes, int
save: whether save, bool
verbose: whether say something, bool
Outputs:
total library SeqRecord
'''
# minimal imports
from Bio import SeqIO
from Bio.Seq import Seq
from Bio.Alphabet import IUPAC
from Bio.SeqRecord import SeqRecord
import numpy as np;
import glob, os, sys, time
import LibraryDesigner as ld
def _load_readouts_per_color(_filename_feature, _colors, _barcode_folder=barcode_folder, _verbose=verbose):
'''Function to load readouts from given filenames'''
# check readout source
_filenames = glob.glob(barcode_folder + os.sep + _filename_feature + '*.fasta')
if len(_filenames) < len(_colors):
raise ValueError('More colors are used than provided readout files for', _filename_feature)
# initialize
_readouts = [[] for _fl in sorted(_filenames)];
if _verbose:
print "-- loading readout seqs from colors:", len(_filenames)
for _i, _fl in enumerate(sorted(_filenames)):
with open(_fl, 'r') as _handle:
for _record in SeqIO.parse(_handle, "fasta"):
_readouts[_i].append(_record);
return _readouts
def _generate_reg_readout_dic(_encodings, _colors, bc_key, bc_list):
'''Function to build dic from region to readouts for a certain kind'''
print '-- generating readout dic'
_bc_ids = [{} for _c in _colors];
for _reg, _info in sorted(_encodings.items()):
if bc_key not in _info or not _info[bc_key]:
raise KeyError('No barcode '+bc_key+' in region', _reg)
_bc_ids[_info['color']][_reg] = _info[bc_key]
# change raw readout id to ids sorted by color
_orders = [{} for _id_list in _bc_ids];
# loop through all ids, get id order by color
for _color, _id_list in enumerate(_bc_ids):
_ids = np.unique(np.array(list(_id_list.values())))
for _reg, _id_per_reg in sorted(_id_list.items()):
if isinstance(_id_per_reg, list):
_orders[_color][_reg] = [int(np.where(_ids==_i)[0]) for _i in _id_per_reg]
else:
_orders[_color][_reg] = int(np.where(_ids==_id_per_reg)[0])
# generate region to readout dictionary
_reg_bc_dic = [{} for _id_list in _orders];
# loop through order list and replace order id to readout record
for _color, _id_list in enumerate(_orders):
for _reg, _id_per_reg in sorted(_id_list.items()):
if isinstance(_id_per_reg, list):
_reg_bc_dic[_color][_reg] = [ bc_list[_color][_i] for _i in _id_per_reg]
else:
_reg_bc_dic[_color][_reg] = bc_list[_color][_id_per_reg]
return _reg_bc_dic
# check inputs:
if verbose:
print "- Check inputs"
# all given colors in encoding
_colors = np.unique([v['color'] for v in reg_encodings.values()])
if verbose:
print "-- number of color used in encoding:", len(_colors)
if verbose:
print "- Load Readout sequences"
_unique_bc_list = _load_readouts_per_color(filename_unique, _colors)
_combo_bc_list = _load_readouts_per_color(filename_combo,_colors)
# check barcode id for each region is given
if verbose:
print "- Match region to readout sequences"
_unique_dic = _generate_reg_readout_dic(reg_encodings, _colors, bc_unique, _unique_bc_list)
_combo_dic = _generate_reg_readout_dic(reg_encodings, _colors, bc_combo, _combo_bc_list)
def _patch_barcode_per_file(_file, _reg_encodings,
_unique_dic, _combo_dic,
_bc_combo=bc_combo, _bc_unique=bc_unique,
_fwd_primer=fwd_primer, _rev_primer=rev_primer,
_add_rand_gap=add_rand_gap, _verbose=verbose):
from random import choice
import os
if _verbose:
print "-- patch barcodes for:", _file
# load probe report
_pb = ld.pb_reports_class()
_pb.load_pbr(_file)
# extract encoding info:
_reg = int(os.path.basename(_file).split('_')[1].split('.')[0])
_info = _reg_encodings[_reg]
##_encoding = _file_encodings[_file];
# initialize, save all infos here
_plist = [];
_precords = [];
for _probe_seq, _probe in sorted(_pb.pb_reports_keep.items(), key=lambda (k,v):v['name']):
# extract sequence
_probe['seq'] = _probe_seq
# extract all encoding info from reg_encodings
_probe['reg_index'] = _info['id']
_probe['color'] = _info['color']
if 'gene' in _info.keys():
_probe['gene'] = _info['gene']
# extract barcode info
_probe[_bc_combo] = _combo_dic[_probe['color']][_reg]
_probe[_bc_unique] = _unique_dic[_probe['color']][_reg]
# extract primer info:
_probe['fwd_primer'] = _fwd_primer;
_probe['rev_primer'] = _rev_primer;
## generate_whole sequence
# fwd_primer(20)
# barcode 1 [from list, 1], (reverse-complement of last 20)
# barcode 2, (reverse-complement of last 20)
# target sequence
# barcode 3, (reverse-complement of last 20)
# barcode 4 [from list, 1], (reverse-complement of last 20)
# rev_primer, (reverse-complement of last 20)
_seq_list = []; # start
_seq_list.append(_probe['fwd_primer'].seq[-20:]) # fwd primer
if isinstance(_probe[_bc_unique],list): # barcode unique
_seq_list += [_bc.seq[-20:].reverse_complement() for _bc in _probe[_bc_unique]];
else:
_seq_list.append(_probe[_bc_unique].seq[-20:].reverse_complement())
if isinstance(_probe[_bc_combo],list): # barcode combo
_seq_list += [_bc.seq[-20:].reverse_complement() for _bc in _probe[_bc_combo]];
else:
_seq_list.append(_probe[_bc_combo].seq[-20:].reverse_complement())
_seq_list.insert(-2, Seq(_probe['seq']) ) # target sequence in the middle
_seq_list.append(_probe['rev_primer'].seq[-20:].reverse_complement()) # reverse primer
# result
dna_alphabet = ['A','A','C','G','T','T']; # used for adding random gap, if needed
_total_seq = Seq('');
for j in range(len(_seq_list)):
_seq = _seq_list[j]
_total_seq += _seq;
if j > 0 and j < len(_seq_list)-2:
_total_seq += ''.join([choice(dna_alphabet) for i in range(_add_rand_gap)]);
_probe['total_seq'] = _total_seq;
## Generate total_name:
# chr21:10350001-10400001_reg_208_gene_chr21_pb_41577 (from base name)
# primer_[4,11]
# barcodes_75,109,[]
# base name
_total_name = _probe['name'].split('reg_')[0] + 'reg_'+str(_probe['reg_index']);
if 'gene' in _probe['name']:
_total_name += '_gene' + _probe['name'].split('gene')[1]
elif 'gene' in _probe.keys():
_total_name += '_gene_'+_probe['gene'];
# primer name
_primer_sets = [int(_probe['fwd_primer'].id.split('_')[-1]), int(_probe['rev_primer'].id.split('_')[-1])]
_total_name += '_primer_'+str(_primer_sets).replace(' ','')
# barcode name
_barcode_sets = [];
## unique
if isinstance(_probe[_bc_unique],list): # barcode combo
_barcode_sets.append([rec.id for rec in _probe[_bc_unique]]);
else:
_barcode_sets.append(_probe[_bc_unique].id);
## combo
if isinstance(_probe[_bc_combo],list): # barcode combo
_barcode_sets.append([rec.id for rec in _probe[_bc_combo]]);
else:
_barcode_sets.append(_probe[_bc_combo].id);
_total_name += '_barcodes_'+str(_barcode_sets).replace(' ','')
# color
_total_name += '_color_'+str(_probe['color'])
## save
_probe['total_name'] = _total_name;
## Append
_plist.append(_probe) # to plist
_precords.append(SeqRecord(_total_seq, id=_total_name, description='', name=_total_name)); # to seq record
return _plist, _precords
# load probe reports:
pb_files = [fl for fl in glob.glob(probe_folder+os.sep+r'*.pbr') if int(os.path.basename(fl).split('_')[1].split('.')[0]) in reg_encodings.keys()]
if verbose:
print "- Load probe reports, total_num:", len(pb_files);
# initialize
_pb_lists, _pb_records = [],[];
# loop through all files
for fl in sorted(pb_files, key=lambda f:int(f.split('.pbr')[0].split('reg_')[-1])):
_list, _records = _patch_barcode_per_file(fl, reg_encodings, _unique_dic, _combo_dic);
_pb_lists.append(_list);
_pb_records += _records
# save:
if save:
if not os.path.exists(library_folder + os.sep + save_subfolder):
os.makedirs(library_folder + os.sep + save_subfolder)
list_savefile = library_folder + os.sep + save_subfolder + os.sep + 'list.pkl';
pb_savefile = library_folder + os.sep + save_subfolder + os.sep + 'candidate_probes.fasta';
if verbose:
print "- Saving list to:", list_savefile
pickle.dump(_pb_lists, open(list_savefile,'w'));
if verbose:
print "- Saving probes to:", pb_savefile
with open(pb_savefile, 'w') as output_handle:
SeqIO.write(_pb_records, output_handle, 'fasta');
return _pb_lists, _pb_records
pb_lists, pb_records = Patch_Barcodes_by_Color(reg_encodings=sub_encoding, probe_folder=probe_folder,
library_folder=library_folder,
fwd_primer=fprimer, rev_primer=rprimer,
add_rand_gap=0, save=True)
```
# 5. Check probes
```
# minimal imports for biopython
import cPickle as pickle
from Bio import SeqIO
from Bio.Seq import Seq
from Bio.Alphabet import IUPAC
from Bio.SeqRecord import SeqRecord
import os,glob,time
import numpy as np
import khmer
library_folder =r'/n/boslfs/LABS/zhuang_lab/User/pzheng/Libraries/CTP-04/chr21_7by21';
probe_subfolder = 'final_probes'
if not 'sub_encoding' in vars():
print '- loading sub_encodings'
sub_encoding = pickle.load(open(library_folder+os.sep+'sub_encoding.pkl','r'))
if not 'pb_records' in vars():
print '- loading all probes'
with open(library_folder+os.sep+probe_subfolder+os.sep+'candidate_probes.fasta', "r") as handle:
pb_records = [];
for record in SeqIO.parse(handle, "fasta"):
pb_records.append(record);
def Check_Probes(pb_records, master_folder,
primer_set,
fwd_primer_filename = 'forward_primers_keep',
rev_primer_filename = 'reverse_primers_keep',
primer_folder=r'/n/boslfs/LABS/zhuang_lab/User/pzheng/Primers',
num_color=3,
barcode_folder=r'/n/boslfs/LABS/zhuang_lab/User/pzheng/Barcodes',
filename_combo='combo_readouts_', filename_unique='unique_readouts_',
bc_combo='bc_decoding', bc_unique='bc_unique',
report_folder=r'reports/centered_merged',save_subfolder=r'final_probes',
save_filename='filtered_probes.fasta',
add_rand_gap=0, total_bc=3, barcode_len=20, target_len=42,
word_size=17, max_internal_hits=50, max_genome_hits=200,
index_folder=r'/n/boslfs/LABS/zhuang_lab/User/pzheng/Indeces/human/hg38',
save=True, verbose=True):
# imports
import os,glob,sys
sys.path.append(r'/n/home13/pzheng/Documents/python-functions/python-functions-library')
from LibraryConstruction import fastaread,fastawrite,fastacombine
import LibraryDesigner as ld
import numpy as np
import matplotlib.pyplot as plt
def load_primers(_primer_set, _fwd_primer_filename,
_rev_primer_filename, _primer_folder,
_verbose=True):
'''Function to load primers'''
_fwd_fl = _primer_folder + os.sep + _fwd_primer_filename + '.fasta'
_rev_fl = _primer_folder + os.sep + _rev_primer_filename + '.fasta'
_fprimer, _rprimer = None, None
# read forward primers
with open(_fwd_fl, "r") as handle:
for record in SeqIO.parse(handle, "fasta"):
if int(record.id.split('_')[-1]) == _primer_set[0]:
_fprimer = record
break
# read forward primers
with open(_rev_fl, "r") as handle:
for record in SeqIO.parse(handle, "fasta"):
if int(record.id.split('_')[-1]) == _primer_set[-1]:
_rprimer = record
break
if _verbose:
print "Primers loaded: forward: "+_fprimer.id+", reverse: "+_rprimer.id;
return _fprimer, _rprimer
def _load_readouts_per_color(_filename_feature, _colors, _barcode_folder, _verbose=True):
'''Function to load readouts from given filenames'''
# check readout source
_filenames = glob.glob(barcode_folder + os.sep + _filename_feature + '*.fasta')
if len(_filenames) < len(_colors):
raise ValueError('More colors are used than provided readout files for', _filename_feature)
# initialize
_readouts = [[] for _fl in sorted(_filenames)];
if _verbose:
print "-- loading readout seqs from colors:", len(_filenames)
for _i, _fl in enumerate(sorted(_filenames)):
with open(_fl, 'r') as _handle:
for _record in SeqIO.parse(_handle, "fasta"):
_readouts[_i].append(_record);
return _readouts
def _check_primer_usage(pb_records, fwd_primer, rev_primer,
_verbose=True):
'''Check whether forward or reverse primer are used in all probes'''
if _verbose:
print "-- Checking primer usage, total probes:", len(pb_records)
fwd_len = len(fwd_primer.seq);
rev_len = len(rev_primer.seq[-20:].reverse_complement());
for record in pb_records:
if record.seq[:fwd_len] != fwd_primer.seq:
if _verbose:
print "--- Forward primer incorrect!"
return False
if record.seq[-rev_len:] != rev_primer.seq[-20:].reverse_complement():
if _verbose:
print "--- Forward primer incorrect!"
return False
return True # if no error applies
def _check_region_size(pb_records):
'''Generate a dirctionary '''
# get region size from probe names
_reg_size_dic = {}
for record in pb_records:
reg_id = int(record.id.split('_reg_')[1].split('_')[0]);
if reg_id not in _reg_size_dic.keys():
_reg_size_dic[reg_id] = 1; # if not in key, create
else:
_reg_size_dic[reg_id] += 1; # otherwise, add count
return _reg_size_dic, True;
def _check_region_to_barcode(pb_records, combo_bcs, unique_bcs,
total_bc=3):
'''Generate map from region id to barcodes used in this region'''
import re
_reg_to_barcode = {}
for record in pb_records:
# region id
reg_id = int(record.id.split('_reg_')[1].split('_')[0]);
if reg_id not in _reg_to_barcode.keys():
# barcode ids
stv_matches = re.findall('\'Stv_(.+?)\'', record.id, re.DOTALL)
ndb_matches = re.findall('\'NDB_(.+?)\'', record.id, re.DOTALL)
stv_names = ['Stv_'+str(stv_id) for stv_id in stv_matches]
ndb_names = ['NDB_'+str(ndb_id) for ndb_id in ndb_matches]
_reg_to_barcode[reg_id] = stv_names+ndb_names
## barcode check
_barcode_check = True;
# barcode names
bc_names = [];
for _bc_list in combo_bcs: # combo barcodes
bc_names += [_bc.id for _bc in _bc_list]
for _bc_list in unique_bcs: # unique barcodes
bc_names += [_bc.id for _bc in _bc_list]
# search through previous dictionary
for reg,bcs in sorted(_reg_to_barcode.items()):
for bc in bcs:
if len(bcs) != total_bc:
print "-- Error in barcode number for region:", reg
_barcode_check = False
break
if bc not in bc_names:
print "-- Wrong barcode name for barcode: "+str(bc)+", region: "+str(reg)
_barcode_check = False
break
return _reg_to_barcode, _barcode_check;
def _parsing_probe_sequence(record, add_rand_gap=add_rand_gap,
barcode_len=barcode_len, target_len=target_len):
'''parse a probe sequence to acquire all barcode binding sites'''
# take in a seq record, parse the sequence and return a list of all included barcodes (20mer,RC)
barcode_list = [];
_main_seq = record.seq[20:-20];
# trim last 2 barcodes
for i in range(2):
barcode_list.append(_main_seq[-barcode_len:]);
_main_seq = _main_seq[:-(barcode_len+add_rand_gap)];
# trim all barcodes from the beginning
while len(_main_seq) > target_len:
barcode_list.append(_main_seq[:barcode_len]);
_main_seq = _main_seq[(barcode_len+add_rand_gap):];
return barcode_list;
def _finding_barcode_name(barcode_list, _combo_bcs, _unique_bcs,
barcode_len=barcode_len, total_bc=total_bc):
'''Given barcode list generated by parsing probe, return a list of barcode names'''
_name_list = [];
_all_bcs = [];
for _bc_list in _combo_bcs: # combo barcodes
_all_bcs += _bc_list
for _bc_list in _unique_bcs: # unique barcodes
_all_bcs += _bc_list
for _site in barcode_list:
for bc in _all_bcs:
if bc.seq[-barcode_len:] == _site.reverse_complement():
_name_list.append(bc.id);
break;
if len(_name_list) < total_bc:
print "-- Failed in finding some barcodes."
return False
return _name_list;
def _check_barcode_to_region(reg_to_barcode,
pb_records, _combo_bcs, _unique_bcs):
'''Generate map from barcode id to region id'''
_barcode_to_reg = {}
_reg_id_exists = []
for record in pb_records:
reg_id = int(record.id.split('_reg_')[1].split('_')[0]);
if reg_id in _reg_id_exists:
continue;
else:
_barcode_list = _parsing_probe_sequence(record)
_name_list = _finding_barcode_name(_barcode_list, _combo_bcs, _unique_bcs)
for _n in _name_list:
if _n not in _barcode_to_reg.keys(): # create if not in dic
_barcode_to_reg[_n] = [reg_id]
else: # otherwise, append
_barcode_to_reg[_n].append(reg_id)
_reg_id_exists.append(reg_id)
## check region distribution
# invert dic from reg_to_barcode
_inv_dic = {}
for reg,bcs in sorted(reg_to_barcode.items()):
for bc in bcs:
if bc not in _inv_dic.keys():
_inv_dic[bc] = [reg];
else:
_inv_dic[bc].append(reg);
# compare
_region_check=True
for bc, regs in sorted(_inv_dic.items()):
if bc not in _barcode_to_reg.keys():
print "-- "+str(bc)+" not in barcode_to_region dic!"
_region_check = False
break
else:
if sorted(regs) != sorted(_barcode_to_reg[bc]):
print "-- "+str(bc)+" and region"+str(regs)+" not compatible with barcode_to_region dic!"
_region_check = False
break
return _barcode_to_reg, _region_check
def _check_barcode_to_color(pb_records, _combo_bcs, _unique_bcs,
_save=save, _master_folder=master_folder, _save_subfolder=save_subfolder):
'''If multi_color is applied, generate a barcode_to_color dic for adaptor design'''
# get barcodes
_barcode_names = []
for _bc_list in combo_bcs: # combo barcodes
_barcode_names += [_bc.id for _bc in _bc_list]
for _bc_list in unique_bcs: # unique barcodes
_barcode_names += [_bc.id for _bc in _bc_list]
# initialize color dic
_barcode_to_color = {};
_exist_regs = [];
# search through all probes
for record in pb_records:
_reg_id = int(record.id.split('_reg_')[1].split('_')[0]);
if _reg_id in _exist_regs:
continue
else:
_exist_regs.append(_reg_id);
_color = int(str(record.id).split('color_')[1])
_barcode_list = _parsing_probe_sequence(record)
_name_list = _finding_barcode_name(_barcode_list, _combo_bcs, _unique_bcs)
for _name in _name_list:
if _name in _barcode_names:
if _name not in _barcode_to_color.keys():
_barcode_to_color[_name] = [_color]
else:
_barcode_to_color[_name].append(_color);
# keep the unique colors
_barcode_to_unique_color = {}
for k,v in sorted(_barcode_to_color.items()):
_barcode_to_unique_color[k] = np.unique(v)
if _save:
import csv
if not os.path.exists(master_folder+os.sep+save_subfolder):
os.mkdir(master_folder+os.sep+save_subfolder)
with open(master_folder+os.sep+save_subfolder+os.sep+'color-usage.csv','w') as output_handle:
fieldnames = ['barcode', 'color']
writer = csv.DictWriter(output_handle, fieldnames=fieldnames)
writer.writeheader()
for _barcode, _color in sorted(_barcode_to_unique_color.items(), key=lambda (k,v):int(k.split('_')[1])):
writer.writerow({'barcode': _barcode, 'color': _color})
return _barcode_to_unique_color
def _construct_internal_map(_master_folder=master_folder, _save_subfolder=save_subfolder,
_pb_records=pb_records, _word_size=word_size):
'''Using functions in LibraryDesign, compute an internal khmer map'''
# save temp fasta
with open(_master_folder+os.sep+_save_subfolder+os.sep+'temp.fasta','w') as _output_handle:
SeqIO.write(_pb_records, _output_handle, 'fasta');
_int_map = khmer.Countgraph(_word_size, 1e9, 2)
_int_map.set_use_bigcount(True)
_nms,_seqs = fastaread(_master_folder+os.sep+_save_subfolder+os.sep+'temp.fasta')
for _seq in _seqs:
_int_map.consume(_seq.upper())
# remove temp file
os.remove(_master_folder+os.sep+_save_subfolder+os.sep+'temp.fasta')
return _int_map
def _check_barcode_in_probes(barcode_to_reg, reg_size_dic, int_map,
_combo_bcs, _unique_bcs,
barcode_len=barcode_len, max_internal_hits=max_internal_hits):
'''Check barcode appearance in probes, whether that match barcode_to_region scheme'''
# load all barcodes
_all_bcs = [];
for _bc_list in _combo_bcs: # combo barcodes
_all_bcs += _bc_list
for _bc_list in _unique_bcs: # unique barcodes
_all_bcs += _bc_list
_barcode_in_probes = {}
for bc_name, regs in sorted(barcode_to_reg.items()):
bc = None
for _bc in _all_bcs:
if bc_name == _bc.id:
bc = _bc
break
bc_hits = int_map.get_kmer_counts( str(bc.seq[-barcode_len:].reverse_complement()).upper());
if max(bc_hits) - min(bc_hits) > max_internal_hits:
print "-- Barcode: "+str(bc)+" has more off-target in different part of itself!"
return False
else:
regs,reg_cts = np.unique(regs, return_counts=True);
bc_in_probe = 0;
for reg,ct in zip(regs,reg_cts):
bc_in_probe += reg_size_dic[reg] * ct;
if max(bc_hits) - bc_in_probe > max_internal_hits:
print bc_hits, regs, bc_in_probe, max_internal_hits
print "-- Barcode: "+str(bc)+" has more off-target than threshold!"
return False
_barcode_in_probes[bc_name] = bc_in_probe;
return _barcode_in_probes, True
def _check_between_probes(int_map, pb_records, _max_internal_hits, target_len,
_make_plot=False, _verbose=True):
def __extract_targeting_sequence(record, target_len=target_len, barcode_len=20):
return record.seq[-barcode_len*2-20-target_len: -barcode_len*2-20]
_internal_hits = [];
_kept_pb_records = [];
_removed_count = 0;
for record in pb_records:
target_seq = str(__extract_targeting_sequence(record)).upper()
_rec_hits = sum(int_map.get_kmer_counts(target_seq));
_internal_hits.append(_rec_hits);
if _rec_hits <= _max_internal_hits:
_kept_pb_records.append(record)
else:
_removed_count += 1
print '--- Max_internal_hits is', _max_internal_hits, 'while this seq got hits:', _rec_hits
if _make_plot:
plt.figure();
plt.hist(_internal_hits);
plt.show();
if _verbose:
print "-- total probes removed by internal screening:", _removed_count
return _kept_pb_records, _removed_count
def _check_against_genome(pb_records, max_genome_hits, index_folder,
_make_plot=False, _verbose=True):
'''Use Khmer to compare probe against genome'''
hg38 = khmer.load_countgraph(index_folder+os.sep+'full_word17_.kmer')
_removed_count = 0;
_genome_hits = []
_keep_pb_records = [];
for record in pb_records:
_kmer_hits = hg38.get_kmer_counts(str(record.seq).upper());
_genome_hits.append(sum(_kmer_hits));
if sum(_kmer_hits) > max_genome_hits:
print '--- Max_genome_hits is: '+str(max_genome_hits)+", this seq got hits: "+ str(sum(_kmer_hits))
_removed_count += 1;
else:
_keep_pb_records.append(record);
if _make_plot:
plt.figure();
plt.hist(_genome_hits);
plt.show();
if _verbose:
print "-- total probes removed by genome screening:", _removed_count
return _keep_pb_records, _removed_count # if nothing goes wrong
def _plot_info():
pass
if verbose:
print "- 0. Loading primers and readouts"
## load primer
fprimer, rprimer = load_primers(primer_set, fwd_primer_filename, rev_primer_filename, primer_folder, verbose)
## load readouts
combo_bcs = _load_readouts_per_color(filename_combo, list(range(num_color)), barcode_folder, _verbose=verbose)
unique_bcs = _load_readouts_per_color(filename_unique, list(range(num_color)), barcode_folder, _verbose=verbose)
## check primers
primer_usage = _check_primer_usage(pb_records, fprimer, rprimer, _verbose=verbose)
if verbose:
print "\n- 1.Passing primer usage check? -", primer_usage
## check region size
reg_size_dic, size_pass = _check_region_size(pb_records)
if verbose:
print "\n- 2.Passing region size check? -", size_pass
for k,v in sorted(reg_size_dic.items()):
print k,':',v
## check region to barcode
reg_to_barcode, reg2bc = _check_region_to_barcode(pb_records, combo_bcs, unique_bcs)
if verbose:
print "\n- 3.Passing region to barcode mapping check? -", reg2bc
for k,v in sorted(reg_to_barcode.items(), key=lambda (k,v):k):
print k,':',v
## check barcode to region (this step must be run after step 3)
barcode_to_reg, bc2reg = _check_barcode_to_region(reg_to_barcode, pb_records, combo_bcs, unique_bcs)
if verbose:
print "\n- 4.Passing barcode to region mapping check? -", bc2reg
for k,v in sorted(barcode_to_reg.items(), key=lambda (k,v):[k[0],int(k.split('_')[1])]):
print k,':',v
## check barcode to region (this step must be run after step 3)
barcode_to_color = _check_barcode_to_color(pb_records, combo_bcs, unique_bcs)
if verbose:
print "\n- 5.Calculating barcode to color dictionary."
for k,v in sorted(barcode_to_color.items(), key=lambda (k,v):[k[0],int(k.split('_')[1])]):
print k,':',v
## Construct an internal map
int_map = _construct_internal_map();
if verbose:
print "\n- 6.Constructing internal khmer map";
## Check barcodes total counts in probes
barcode_in_probes, _bc_counting = _check_barcode_in_probes(barcode_to_reg, reg_size_dic, int_map, combo_bcs, unique_bcs)
if verbose:
print "\n- 7.Passing if counting barcode appearance times in probes:", _bc_counting;
## Check against each other
if verbose:
print "\n- 8.Checking probes within library to avoid excess hits";
kept_records, failed_internal_num = _check_between_probes(int_map, pb_records, max_internal_hits, target_len)
## Check against genome
if verbose:
print "\n- 9.Checking full probes against genome";
kept_records, failed_genome_num = _check_against_genome(kept_records, max_genome_hits, index_folder);
# check region size for kept probes
_reg_size_dic = {}
for record in pb_records:
reg_id = int(record.id.split('_reg_')[1].split('_')[0]);
if reg_id not in _reg_size_dic.keys():
_reg_size_dic[reg_id] = 1; # if not in key, create
else:
_reg_size_dic[reg_id] += 1; # otherwise, add count
if verbose:
print "-- re-check region size:"
for k,v in sorted(_reg_size_dic.items()):
print k,':',v
print "-- final probe number:", len(kept_records)
if save:
pb_savefile = master_folder + os.sep + save_subfolder + os.sep + save_filename;
if verbose:
print "\n- 10.Saving probes to:", pb_savefile
with open(pb_savefile, 'w') as output_handle:
SeqIO.write(kept_records, output_handle, 'fasta');
return kept_records, _reg_size_dic
kept_records, kept_size_dic = Check_Probes(pb_records, library_folder, [2,9],
save=True)
```
## 6. Blast
```
# minimal imports for biopython
import cPickle as pickle
from Bio import SeqIO
from Bio.Seq import Seq
from Bio.Alphabet import IUPAC
from Bio.SeqRecord import SeqRecord
import os,glob,time
import numpy as np
import khmer
# blast
from Bio.Blast.Applications import NcbiblastnCommandline
from Bio.Blast import NCBIXML
import time
library_folder = r'/n/boslfs/LABS/zhuang_lab/User/pzheng/Libraries/CTP-04/chr21_7by21'
probe_subfolder = r'final_probes'
kept_probe_filename = 'filtered_probes.fasta'
kept_probe_list_filename = 'list.pkl'
probes = [];
with open(library_folder+os.sep+probe_subfolder+os.sep+kept_probe_filename,'r') as handle:
for record in SeqIO.parse(handle, "fasta"):
probes.append(record)
```
### 6.1 Load Probes
```
probes = [];
with open(library_folder+os.sep+probe_subfolder+os.sep+kept_probe_filename,'r') as handle:
for record in SeqIO.parse(handle, "fasta"):
probes.append(record)
```
### 6.2 Functions
```
def acquire_blast_counts(blast_record, hard_thres=42, soft_thres=20, verbose=False):
'''filter for genome blast record
Input:
blast_record: xml format of blast result, generated by NCBIWWW.qblast or NCBIblastnCommandLine
hard_threshold: only one hsp_score match larger than this is allowed, int
soft_threshold: hsp_score larger than this should be counted, int
verbose: it says a lot of things!
Output:
_hard_count: hits for hard threshold
_soft_count: hits for soft threshold'''
_hard_count, _soft_count = 0, 0
if verbose:
print ('- '+blast_record.query_id, ' hits =',len(blast_record.alignments));
# extract information
hsp_scores = []
hsp_aligns = []
for alignment in blast_record.alignments:
for hsp in alignment.hsps:
hsp_scores.append(hsp.score)
hsp_aligns.append(hsp.align_length)
# hard threshold
keep_hard = [hsp_align >= hard_thres and hsp_score >= hard_thres for hsp_score,hsp_align in zip(hsp_scores,hsp_aligns)];
_hard_count = sum(keep_hard)
if verbose:
print ("-- hard count:", _hard_count);
# soft threshold
keep_soft = [hsp_align >= soft_thres and hsp_score >= soft_thres for hsp_score,hsp_align in zip(hsp_scores,hsp_aligns)];
_soft_count = sum(keep_soft)
if verbose:
print("-- soft count:", _soft_count);
return _hard_count, _soft_count
# do blast by region
def split_library_by_region(probes, info_key='_reg_', index_key='_pb_'):
'''Function to split a pb_record list into list of list,
each list belongs to one region'''
_pb_dic = {};
for _pb in probes:
if info_key in _pb.id:
_reg = int(_pb.id.split(info_key)[-1].split('_')[0])
if _reg not in _pb_dic:
_pb_dic[_reg] = [_pb]
else:
_pb_dic[_reg].append(_pb)
for _reg, _pbs in sorted(_pb_dic.items()):
_pbs = sorted(_pbs, key=lambda p:int(p.id.split(index_key)[-1].split('_')[0]))
_pb_dic[_reg] = _pbs
return _pb_dic
```
### 6.3 Blast!
```
pb_dic = split_library_by_region(probes)
print library_folder
```
### Blast by regions
```
# Folders
blast_subfolder = 'blast'
blast_folder = library_folder + os.sep + blast_subfolder
if not os.path.exists(blast_folder):
os.mkdir(blast_folder);
# parameters
force=False
verbose=True
verbose_parse=False
hard_counts, soft_counts = [],[]
for _reg, _pbs in sorted(pb_dic.items()):
if verbose:
print ("- region", _reg)
_start = time.time()
if force or not os.path.exists(blast_folder+os.sep+'blast_reg_'+str(_reg)+'.xml'):
if verbose:
print "-- writing file:", blast_folder+os.sep+'probe_reg_'+str(_reg)+'.fasta'
# save these number of probes into temp.fasta
with open(blast_folder+os.sep+'probe_reg_'+str(_reg)+'.fasta', "w") as output_handle:
SeqIO.write(_pbs, output_handle, "fasta")
if verbose:
print ("-- blasting region:", _reg)
# Run BLAST and parse the output as XML
output = NcbiblastnCommandline(query=blast_folder+os.sep+'probe_reg_'+str(_reg)+'.fasta',
num_threads=8,
db=r'/n/boslfs/LABS/zhuang_lab/User/pzheng/Genomes/human/hg38/hg38',
evalue=500,
word_size=10,
out=blast_folder+os.sep+'blast_reg_'+str(_reg)+'.xml',
outfmt=5)()[0]
if verbose:
_after_blast = time.time()
print("--- Total time for blast:", _after_blast-_start)
else:
_after_blast = time.time()
# parsing output:
if verbose:
print ("-- parsing blast result for region:", _reg)
blast_records = NCBIXML.parse(open(blast_folder+os.sep+'blast_reg_'+str(_reg)+'.xml'))
# save a vector to store keep or not
hcs,scs=[],[]
for blast_record in blast_records:
#print blast_record.query_id, len(blast_record.alignments)
hc, sc = acquire_blast_counts(blast_record, hard_thres=42, soft_thres=20, verbose=verbose_parse);
hcs.append(hc)
scs.append(sc)
if verbose:
_after_parse = time.time()
print("--- Total time for parsing:", _after_parse - _after_blast)
# save hard counts and soft counts stats
hard_counts.append(hcs)
soft_counts.append(scs)
import matplotlib.pyplot as plt
%matplotlib notebook
plt.figure()
plt.hist(np.concatenate(hard_counts,axis=0))
plt.figure()
plt.hist(np.concatenate(soft_counts,axis=0), bins=range(0,1000,100))
```
## 7. Post Processing after blast
```
def Screening_Probes_by_Blast(library_folder, probe_per_region, blast_subfolder='blast',
probe_subfolder='final_probes', probe_filename='filtered_probes.fasta',
soft_count_th=30,
smallest_region_ratio=0.75,
save=True, save_filename='blast_centered_probes.fasta',
verbose=True,):
'''Read blast results in blast folder and probe in filtered__probes, keep'''
# folders
_blast_folder = library_folder + os.sep + blast_subfolder
_probe_folder = library_folder + os.sep + probe_subfolder
# load probes
_probes = [];
with open(_probe_folder+os.sep+probe_filename,'r') as _handle:
for _record in SeqIO.parse(_handle, "fasta"):
_probes.append(_record)
if verbose:
print "- Number of probes loaded:", len(_probes)
# parse loaded probes by region
_pb_dic = split_library_by_region(_probes)
if verbose:
print "- Number of regions in this library:", len(_pb_dic);
# dictionary to store whether keep this probe
_keep_dic = {} # whether keep because of blast only
_kept_pb_dic = {}
_hard_count_list = []
_soft_count_list = []
# loop through all regions
for _reg, _pbs in sorted(_pb_dic.items()):
if verbose:
print "-- checking probes in region:", _reg
_keep_dic[_reg] = np.ones(len(_pbs), dtype=np.bool) # initialize with True
# parse blast result of this region
blast_records = NCBIXML.parse(open(_blast_folder+os.sep+'blast_reg_'+str(_reg)+'.xml', 'r'))
# loop through each probe in this region
_hard_cts, _soft_cts = [],[]
for _pbid, blast_record in enumerate(blast_records):
_hc, _sc = acquire_blast_counts(blast_record, hard_thres=42, soft_thres=20, verbose=verbose_parse);
_hard_cts.append(_hc)
_soft_cts.append(_sc)
if _hc > 2 or _hc < 1: # if this probe has no hit, or more than 2 hits, remove
_keep_dic[_reg][_pbid] = False
continue
if _sc > soft_count_th: # if this probe has too many soft counts (20mer hits)
_keep_dic[_reg][_pbid] = False
continue
# after looped through this region, check the hard counts
_hard_cts = np.array(_hard_cts);
_soft_cts = np.array(_soft_cts);
_hard_count_list.append(_hard_cts)
_soft_count_list.append(_soft_cts)
if verbose:
print "--- number of probes:", len(_pbs), ", kept by blast:", sum(_keep_dic[_reg]), ", if remove dups:", sum(_keep_dic[_reg] * (_hard_cts==1))
# check duplicated probes
if sum(_keep_dic[_reg] * (_hard_cts==1)) / float(sum(_keep_dic[_reg])) >= smallest_region_ratio and sum(_keep_dic[_reg]) >= smallest_region_ratio*probe_per_region:
print '--- remove duplicated probes'
_keep_dic[_reg] = _keep_dic[_reg] * (_hard_cts==1)
# generate list of kept probes
_kept_pbs = [_pb for _pb,_k in zip(_pbs, _keep_dic[_reg]) if _k]
# keep the center of this region
if sum(_keep_dic[_reg]) > probe_per_region:
if verbose:
print "--- keep centered probes"
_start, _end = _pbs[0].id.split(':')[1].split('_')[0].split('-')
_start, _end = int(_start), int(_end)
_reg_len = np.abs(_end - _start)
_kept_center_pbs = []
for _pb in sorted(_kept_pbs, key=lambda p: np.abs(int(p.id.split('pb_')[1].split('_')[0])-_reg_len/2) ):
_kept_center_pbs.append(_pb)
if len(_kept_center_pbs) >= probe_per_region:
break
_kept_pb_dic[_reg] = sorted(_kept_center_pbs ,key=lambda p:int(p.id.split('pb_')[1].split('_')[0]))
else:
_kept_pb_dic[_reg] = sorted(_kept_pbs ,key=lambda p:int(p.id.split('pb_')[1].split('_')[0]))
if verbose:
print '-- number of probes kept for this region:', len(_kept_pb_dic[_reg])
# SUMMARIZE
_kept_probe_list = []
if verbose:
print "- summarize";
for _reg, _pbs in sorted(_kept_pb_dic.items()):
if verbose:
print "-- region:", _reg, ", number of probes:",len(_pbs);
_kept_probe_list += _pbs
print "- Number of probes kept:", len(_kept_probe_list)
if save:
if verbose:
print "- Saving to file:", _probe_folder + os.sep + save_filename
with open(_probe_folder + os.sep + save_filename, 'w') as _output_handle:
SeqIO.write(_kept_probe_list, _output_handle, 'fasta');
return _kept_probe_list, _keep_dic, _hard_count_list, _soft_count_list
kept_pbs, blast_keep_dic, hard_count_list, soft_count_list = Screening_Probes_by_Blast(library_folder, 250)
library_folder = r'/n/boslfs/LABS/zhuang_lab/User/pzheng/Libraries/CTP-04/chr21_7by21'
probe_subfolder = 'final_probes'
blast_probe_filename = 'blast_centered_probes.fasta'
kept_pbs = [];
with open(library_folder+os.sep+probe_subfolder+os.sep+blast_probe_filename,'r') as handle:
for record in SeqIO.parse(handle, "fasta"):
kept_pbs.append(record)
print len(kept_pbs)
kept_blasted_pbs, kept_blasted_size_dic = Check_Probes(kept_pbs, library_folder, [2,9],
save=True, save_filename='filtered_blast_centered_probes.fasta')
```
| github_jupyter |
# First visit Monte Carlo Policy Evaluation
┌─┬─┬─┬─┐
│0 │1 │2 │3 │
├─┼─┼─┼─┤
│4 │X │5 │6 │
├─┼─┼─┼─┤
│7 │8 │9 │10│
└─┴─┴─┴─┘
3 : 1점, 6 : -1점
0 왼 1 오 2 위 3 아래
# V
```
import numpy as np
from copy import deepcopy
testmode = 0
# set parameters ###############################################################
epoch = 10000
# set parameters ###############################################################
# state
states = [0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10]
N_STATES = len(states)
# action
actions = [0, 1, 2, 3]
N_ACTIONS = len(actions)
# transition probabilities
P = np.empty((N_STATES, N_ACTIONS, N_STATES))
# 0 1 2 3 4 5 6 7 8 9 10
P[ 0, 0, :] = [ .9, 0, 0, 0, .1, 0, 0, 0, 0, 0, 0]
P[ 0, 1, :] = [ .1, .8, 0, 0, .1, 0, 0, 0, 0, 0, 0]
P[ 0, 2, :] = [ .9, .1, 0, 0, 0, 0, 0, 0, 0, 0, 0]
P[ 0, 3, :] = [ .1, .1, 0, 0, .8, 0, 0, 0, 0, 0, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 1, 0, :] = [ .8, .2, 0, 0, 0, 0, 0, 0, 0, 0, 0]
P[ 1, 1, :] = [ 0, .2, .8, 0, 0, 0, 0, 0, 0, 0, 0]
P[ 1, 2, :] = [ .1, .8, .1, 0, 0, 0, 0, 0, 0, 0, 0]
P[ 1, 3, :] = [ .1, .8, .1, 0, 0, 0, 0, 0, 0, 0, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 2, 0, :] = [ 0, .8, .1, 0, 0, .1, 0, 0, 0, 0, 0]
P[ 2, 1, :] = [ 0, 0, .1, .8, 0, .1, 0, 0, 0, 0, 0]
P[ 2, 2, :] = [ 0, .1, .8, .1, 0, 0, 0, 0, 0, 0, 0]
P[ 2, 3, :] = [ 0, .1, 0, .1, 0, .8, 0, 0, 0, 0, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 3, 0, :] = [ 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0]
P[ 3, 1, :] = [ 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0]
P[ 3, 2, :] = [ 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0]
P[ 3, 3, :] = [ 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 4, 0, :] = [ .1, 0, 0, 0, .8, 0, 0, .1, 0, 0, 0]
P[ 4, 1, :] = [ .1, 0, 0, 0, .8, 0, 0, .1, 0, 0, 0]
P[ 4, 2, :] = [ .8, 0, 0, 0, .2, 0, 0, 0, 0, 0, 0]
P[ 4, 3, :] = [ 0, 0, 0, 0, .2, 0, 0, .8, 0, 0, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 5, 0, :] = [ 0, 0, .1, 0, 0, .8, 0, 0, 0, .1, 0]
P[ 5, 1, :] = [ 0, 0, .1, 0, 0, 0, .8, 0, 0, .1, 0]
P[ 5, 2, :] = [ 0, 0, .8, 0, 0, .1, .1, 0, 0, 0, 0]
P[ 5, 3, :] = [ 0, 0, 0, 0, 0, .1, .1, 0, 0, .8, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 6, 0, :] = [ 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0]
P[ 6, 1, :] = [ 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0]
P[ 6, 2, :] = [ 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0]
P[ 6, 3, :] = [ 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 7, 0, :] = [ 0, 0, 0, 0, .1, 0, 0, .9, 0, 0, 0]
P[ 7, 1, :] = [ 0, 0, 0, 0, .1, 0, 0, .1, .8, 0, 0]
P[ 7, 2, :] = [ 0, 0, 0, 0, .8, 0, 0, .1, .1, 0, 0]
P[ 7, 3, :] = [ 0, 0, 0, 0, 0, 0, 0, .9, .1, 0, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 8, 0, :] = [ 0, 0, 0, 0, 0, 0, 0, .8, .2, 0, 0]
P[ 8, 1, :] = [ 0, 0, 0, 0, 0, 0, 0, 0, .2, .8, 0]
P[ 8, 2, :] = [ 0, 0, 0, 0, 0, 0, 0, .1, .8, .1, 0]
P[ 8, 3, :] = [ 0, 0, 0, 0, 0, 0, 0, .1, .8, .1, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 9, 0, :] = [ 0, 0, 0, 0, 0, .1, 0, 0, .8, .1, 0]
P[ 9, 1, :] = [ 0, 0, 0, 0, 0, .1, 0, 0, 0, .1, .8]
P[ 9, 2, :] = [ 0, 0, 0, 0, 0, .8, 0, 0, .1, 0, .1]
P[ 9, 3, :] = [ 0, 0, 0, 0, 0, 0, 0, 0, .1, .8, .1]
# 0 1 2 3 4 5 6 7 8 9 10
P[10, 0, :] = [ 0, 0, 0, 0, 0, 0, .1, 0, 0, .8, .1]
P[10, 1, :] = [ 0, 0, 0, 0, 0, 0, .1, 0, 0, 0, .9]
P[10, 2, :] = [ 0, 0, 0, 0, 0, 0, .8, 0, 0, .1, .1]
P[10, 3, :] = [ 0, 0, 0, 0, 0, 0, 0, 0, 0, .1, .9]
# rewards
R = -0.02 * np.ones((N_STATES, N_ACTIONS))
R[3,:] = 1.
R[6,:] = -1.
# discount factor
gamma = 0.99
# policy
if 0:
# bad policy
policy = np.empty((N_STATES, N_ACTIONS))
policy[0,:] = [0,1,0,0]
policy[1,:] = [0,1,0,0]
policy[2,:] = [0,1,0,0]
policy[3,:] = [0,1,0,0]
policy[4,:] = [0,0,0,1]
policy[5,:] = [0,1,0,0]
policy[6,:] = [0,1,0,0]
policy[7,:] = [0,1,0,0]
policy[8,:] = [0,1,0,0]
policy[9,:] = [0,0,1,0]
policy[10,:] = [0,0,1,0]
elif 0:
# random policy
policy = 0.25*np.ones((N_STATES, N_ACTIONS))
elif 0:
# optimal policy
policy = np.empty((N_STATES, N_ACTIONS))
policy[0,:] = [0,1,0,0]
policy[1,:] = [0,1,0,0]
policy[2,:] = [0,1,0,0]
policy[3,:] = [0,1,0,0]
policy[4,:] = [0,0,1,0]
policy[5,:] = [0,0,1,0]
policy[6,:] = [0,0,1,0]
policy[7,:] = [0,0,1,0]
policy[8,:] = [1,0,0,0]
policy[9,:] = [1,0,0,0]
policy[10,:] = [1,0,0,0]
elif 1:
# optimal policy + noise
# we use optimal policy with probability 1/(1+ep)
# we use random policy with probability ep/(1+ep)
ep = 0.1
policy = np.empty((N_STATES, N_ACTIONS))
policy[0,:] = [0,1,0,0]
policy[1,:] = [0,1,0,0]
policy[2,:] = [0,1,0,0]
policy[3,:] = [0,1,0,0]
policy[4,:] = [0,0,1,0]
policy[5,:] = [0,0,1,0]
policy[6,:] = [0,0,1,0]
policy[7,:] = [0,0,1,0]
policy[8,:] = [1,0,0,0]
policy[9,:] = [1,0,0,0]
policy[10,:] = [1,0,0,0]
policy = policy + (ep/4)*np.ones((N_STATES, N_ACTIONS))
policy = policy / np.sum(policy, axis=1).reshape((N_STATES,1))
# Every-Visit Monte-Carlo Policy Evaluation
# n_visits records number of visits for each state
# cum_gains records cumulative gains, i.e., sum of gains for each state
# where
# gain = reward + gamma * next_reward + gamma^2 * ...
n_visits = np.zeros(N_STATES)
cum_gains = np.zeros(N_STATES)
for _ in range(epoch):
if testmode == 1 :
print ("안녕")
print (_,"번 째")
# simulation_history records visited states including the terminal states 3 and 6
# reward_history records occured rewards including final rewards 1. and -1.
simulation_history = []
reward_history = []
# indicate game is not over yet
done = False
# choose initial state randomly, not from 3 or 6
s = np.random.choice([0, 1, 2, 4, 5, 7, 8, 9, 10])
NUM = 0
while not done:
if testmode == 1 :
print (NUM)
# choose action using current policy
a = np.random.choice(actions, p=policy[s, :])
simulation_history.append(s)
reward_history.append(R[s,a])
if testmode == 1 :
print ("si_hi : ", simulation_history)
print ("re_hi : ", reward_history)
# choose next state using transition probabilities
s1 = np.random.choice(states, p=P[s, a, :])
if s1 == 3:
# if game is over,
# ready to break while loop by letting done = True
# append end result to simulation_history
done = True
simulation_history.append(s1)
reward_history.append(R[s1,0])
elif s1 == 6:
# if game is over,
# ready to break while loop by letting done = True
# append end result to simulation_history
done = True
simulation_history.append(s1)
reward_history.append(R[s1,0])
else:
# if game is not over, continue playing game
s = s1
NUM += 1
# reward_history records occured rewards including final rewards 1 and -1
simulation_history = np.array(simulation_history)
reward_history = np.array(reward_history)
n = len(reward_history)
# gain_history records occured gains
gain_history = deepcopy(reward_history)
if testmode == 1 :
print ("gain_pre : ", gain_history)
for i, reward in enumerate(reward_history[:-1][::-1]):
gain_history[n-i-2] = reward + gamma * gain_history[n-i-2+1]
if testmode == 1 :
print ("i:",i,"reward",reward)
print (gain_history)
if testmode == 1 :
print ("gain_post :", gain_history)
check = [-1 for i in range(N_STATES)]
for i in range(N_STATES):
bbbbbb=-1
for j in range(n) :
if simulation_history[j] == i and bbbbbb == -1 :
bbbbbb = j
check [i] = bbbbbb
if testmode == 1 :
print ("check :", check)
# update n_visits and cum_gains
for i in range(N_STATES):
if check[i] != -1:
n_visits[i] += 1
cum_gains[i] += gain_history[check[i]]
if testmode == 1 :
print ("n_vi : ", n_visits)
print ("cum_ga : ", cum_gains)
V = cum_gains / (n_visits + 1.0e-8)
print(V)
```
# Q
```
# test
testmode = 0
import numpy as np
from copy import deepcopy
# set parameters ###############################################################
epoch = 50000
# set parameters ###############################################################
# state
states = [0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10]
N_STATES = len(states)
# action
actions = [0, 1, 2, 3]
N_ACTIONS = len(actions)
# transition probabilities
P = np.empty((N_STATES, N_ACTIONS, N_STATES))
# 0 1 2 3 4 5 6 7 8 9 10
P[ 0, 0, :] = [ .9, 0, 0, 0, .1, 0, 0, 0, 0, 0, 0]
P[ 0, 1, :] = [ .1, .8, 0, 0, .1, 0, 0, 0, 0, 0, 0]
P[ 0, 2, :] = [ .9, .1, 0, 0, 0, 0, 0, 0, 0, 0, 0]
P[ 0, 3, :] = [ .1, .1, 0, 0, .8, 0, 0, 0, 0, 0, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 1, 0, :] = [ .8, .2, 0, 0, 0, 0, 0, 0, 0, 0, 0]
P[ 1, 1, :] = [ 0, .2, .8, 0, 0, 0, 0, 0, 0, 0, 0]
P[ 1, 2, :] = [ .1, .8, .1, 0, 0, 0, 0, 0, 0, 0, 0]
P[ 1, 3, :] = [ .1, .8, .1, 0, 0, 0, 0, 0, 0, 0, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 2, 0, :] = [ 0, .8, .1, 0, 0, .1, 0, 0, 0, 0, 0]
P[ 2, 1, :] = [ 0, 0, .1, .8, 0, .1, 0, 0, 0, 0, 0]
P[ 2, 2, :] = [ 0, .1, .8, .1, 0, 0, 0, 0, 0, 0, 0]
P[ 2, 3, :] = [ 0, .1, 0, .1, 0, .8, 0, 0, 0, 0, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 3, 0, :] = [ 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0]
P[ 3, 1, :] = [ 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0]
P[ 3, 2, :] = [ 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0]
P[ 3, 3, :] = [ 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 4, 0, :] = [ .1, 0, 0, 0, .8, 0, 0, .1, 0, 0, 0]
P[ 4, 1, :] = [ .1, 0, 0, 0, .8, 0, 0, .1, 0, 0, 0]
P[ 4, 2, :] = [ .8, 0, 0, 0, .2, 0, 0, 0, 0, 0, 0]
P[ 4, 3, :] = [ 0, 0, 0, 0, .2, 0, 0, .8, 0, 0, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 5, 0, :] = [ 0, 0, .1, 0, 0, .8, 0, 0, 0, .1, 0]
P[ 5, 1, :] = [ 0, 0, .1, 0, 0, 0, .8, 0, 0, .1, 0]
P[ 5, 2, :] = [ 0, 0, .8, 0, 0, .1, .1, 0, 0, 0, 0]
P[ 5, 3, :] = [ 0, 0, 0, 0, 0, .1, .1, 0, 0, .8, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 6, 0, :] = [ 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0]
P[ 6, 1, :] = [ 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0]
P[ 6, 2, :] = [ 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0]
P[ 6, 3, :] = [ 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 7, 0, :] = [ 0, 0, 0, 0, .1, 0, 0, .9, 0, 0, 0]
P[ 7, 1, :] = [ 0, 0, 0, 0, .1, 0, 0, .1, .8, 0, 0]
P[ 7, 2, :] = [ 0, 0, 0, 0, .8, 0, 0, .1, .1, 0, 0]
P[ 7, 3, :] = [ 0, 0, 0, 0, 0, 0, 0, .9, .1, 0, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 8, 0, :] = [ 0, 0, 0, 0, 0, 0, 0, .8, .2, 0, 0]
P[ 8, 1, :] = [ 0, 0, 0, 0, 0, 0, 0, 0, .2, .8, 0]
P[ 8, 2, :] = [ 0, 0, 0, 0, 0, 0, 0, .1, .8, .1, 0]
P[ 8, 3, :] = [ 0, 0, 0, 0, 0, 0, 0, .1, .8, .1, 0]
# 0 1 2 3 4 5 6 7 8 9 10
P[ 9, 0, :] = [ 0, 0, 0, 0, 0, .1, 0, 0, .8, .1, 0]
P[ 9, 1, :] = [ 0, 0, 0, 0, 0, .1, 0, 0, 0, .1, .8]
P[ 9, 2, :] = [ 0, 0, 0, 0, 0, .8, 0, 0, .1, 0, .1]
P[ 9, 3, :] = [ 0, 0, 0, 0, 0, 0, 0, 0, .1, .8, .1]
# 0 1 2 3 4 5 6 7 8 9 10
P[10, 0, :] = [ 0, 0, 0, 0, 0, 0, .1, 0, 0, .8, .1]
P[10, 1, :] = [ 0, 0, 0, 0, 0, 0, .1, 0, 0, 0, .9]
P[10, 2, :] = [ 0, 0, 0, 0, 0, 0, .8, 0, 0, .1, .1]
P[10, 3, :] = [ 0, 0, 0, 0, 0, 0, 0, 0, 0, .1, .9]
# rewards
R = -0.02 * np.ones((N_STATES, N_ACTIONS))
R[3,:] = 1.
R[6,:] = -1.
# discount factor
gamma = 0.99
# policy
if 0:
# bad policy
policy = np.empty((N_STATES, N_ACTIONS))
policy[0,:] = [0,1,0,0]
policy[1,:] = [0,1,0,0]
policy[2,:] = [0,1,0,0]
policy[3,:] = [0,1,0,0]
policy[4,:] = [0,0,0,1]
policy[5,:] = [0,1,0,0]
policy[6,:] = [0,1,0,0]
policy[7,:] = [0,1,0,0]
policy[8,:] = [0,1,0,0]
policy[9,:] = [0,0,1,0]
policy[10,:] = [0,0,1,0]
elif 0:
# random policy
policy = 0.25*np.ones((N_STATES, N_ACTIONS))
elif 0:
# optimal policy
policy = np.empty((N_STATES, N_ACTIONS))
policy[0,:] = [0,1,0,0]
policy[1,:] = [0,1,0,0]
policy[2,:] = [0,1,0,0]
policy[3,:] = [0,1,0,0]
policy[4,:] = [0,0,1,0]
policy[5,:] = [0,0,1,0]
policy[6,:] = [0,0,1,0]
policy[7,:] = [0,0,1,0]
policy[8,:] = [1,0,0,0]
policy[9,:] = [1,0,0,0]
policy[10,:] = [1,0,0,0]
elif 1:
# optimal policy + noise
# we use optimal policy with probability 1/(1+ep)
# we use random policy with probability ep/(1+ep)
ep = 0.1
policy = np.empty((N_STATES, N_ACTIONS))
policy[0,:] = [0,1,0,0]
policy[1,:] = [0,1,0,0]
policy[2,:] = [0,1,0,0]
policy[3,:] = [0,1,0,0]
policy[4,:] = [0,0,1,0]
policy[5,:] = [0,0,1,0]
policy[6,:] = [0,0,1,0]
policy[7,:] = [0,0,1,0]
policy[8,:] = [1,0,0,0]
policy[9,:] = [1,0,0,0]
policy[10,:] = [1,0,0,0]
policy = policy + (ep/4)*np.ones((N_STATES, N_ACTIONS))
policy = policy / np.sum(policy, axis=1).reshape((N_STATES,1))
# Every-Visit Monte-Carlo Policy Evaluation
# n_visits records number of visits for each state and action
# cum_gains records cumulative gains, i.e., sum of gains for each state and action
# where
# gain = reward + gamma * next_reward + gamma^2 * ...
# Previously for V
# n_visits = np.zeros(N_STATES)
# cum_gains = np.zeros(N_STATES)
print ("policy")
print (policy)
n_visits = np.zeros((N_STATES, N_ACTIONS))
cum_gains = np.zeros((N_STATES, N_ACTIONS))
for _ in range(epoch):
if _ % 5000 == 0 :
print (_,"/",epoch)
if testmode == 1 :
print ("안녕")
print (_,"번 째")
# simulation_history records visited states and actions including the terminal states 3 and 6
# reward_history records occured rewards including final rewards 1 and -1
simulation_history = []
reward_history = []
# indicate game is not over yet
done = False
# choose initial state randomly, not from 3 or 6
s = np.random.choice([0, 1, 2, 4, 5, 7, 8, 9, 10])
NUM = 0
while not done:
# choose action using current policy
a = np.random.choice(actions, p=policy[s, :])
# Previously for V
# simulation_history.append(s)
simulation_history.append((s,a))
reward_history.append(R[s,a])
if testmode == 1 :
print ("si_hi : ", simulation_history)
print ("re_hi : ", reward_history)
# choose next state using transition probabilities
s1 = np.random.choice(states, p=P[s, a, :])
if s1 == 3:
# if game is over,
# ready to break while loop by letting done = True
# append end result to simulation_history
done = True
# Previously for V
# simulation_history.append(s1)
simulation_history.append((s1,0))
reward_history.append(R[s1,0])
elif s1 == 6:
# if game is over,
# ready to break while loop by letting done = True
# append end result to simulation_history
done = True
# Previously for V
# simulation_history.append(s1)
simulation_history.append((s1,0))
reward_history.append(R[s1,0])
else:
# if game is not over, continue playing game
s = s1
NUM += 1
# reward_history records occured rewards including final rewards 1 and -1
reward_history = np.array(reward_history)
n = len(reward_history)
# gain_history records occured gains
gain_history = deepcopy(reward_history)
for i, reward in enumerate(reward_history[:-1][::-1]):
gain_history[n-i-2] = reward + gamma * gain_history[n-i-2+1]
# update n_visits and cum_gains
# Previously for V
# for i in range(N_STATES):
# n_visits[i] += np.sum(simulation_history==i)
# cum_gains[i] += np.sum(gain_history[simulation_history==i])
if testmode == 1 :
print ("simu_hi : ", simulation_history)
print ("gain_hi : ", gain_history)
check = [-1 for i in range(N_STATES)]
for i in range(N_STATES):
bbbbbb=-1
for j in range(n) :
if simulation_history[j][0] == i and bbbbbb ==-1 :
bbbbbb = j
check [i] = bbbbbb
if testmode == 1 :
print ("check :", check)
for i, (s, a) in enumerate(simulation_history):
if check[s]==i :
n_visits[s, a] += 1.
cum_gains[s, a] += gain_history[i]
if testmode == 1 :
print ("n_vi : ", n_visits)
print ("cum_ga : ", cum_gains)
Q = cum_gains / (n_visits + 1.0e-8)
print("Q", Q)
```
| github_jupyter |
# Better Long-Term Stock Forecasts
by [Magnus Erik Hvass Pedersen](http://www.hvass-labs.org/)
/ [GitHub](https://github.com/Hvass-Labs/FinanceOps) / [Videos on YouTube](https://www.youtube.com/playlist?list=PL9Hr9sNUjfsmlHaWuVxIA0pKL1yjryR0Z)
## Introduction
The [previous paper](https://github.com/Hvass-Labs/FinanceOps/blob/master/01_Forecasting_Long-Term_Stock_Returns.ipynb) showed a strong predictive relationship between the P/Sales ratio and long-term returns of some individual stocks and the S&P 500 stock-market index.
However, there was a considerable amount of noise in those scatter-plots, because we considered fixed investment periods of exactly 10 years, for example. So even though the P/Sales ratio was a strong predictor for the mispricing at the buy-time, it was impossible to predict the mispricing at the sell-time, because the stock-market could be in a bubble or in a crash 10 years into the future, which would distort the estimated returns.
This paper presents a simple solution, which is to consider the average returns for all investment periods between 7 and 15 years, and then make a scatter-plot of the mean returns versus the P/Sales ratio. This produces incredibly smooth curves for estimating the future long-term returns of the S&P 500 and some individual stocks.
Along with the [previous paper](https://github.com/Hvass-Labs/FinanceOps/blob/master/01_Forecasting_Long-Term_Stock_Returns.ipynb), this is a very important discovery and it has implications for many areas of both theoretical and applied finance. It means that the U.S. stock-market as a whole is not "efficient" and does not follow a purely "random walk" in the long-term. It is possible to estimate the future long-term return of the stock-market and some individual stocks from just a single indicator variable.
## Python Imports
This Jupyter Notebook is implemented in Python v. 3.6 and requires various packages for numerical computations and plotting. See the installation instructions in the README-file.
```
%matplotlib inline
# Imports from Python packages.
import matplotlib.pyplot as plt
from matplotlib.ticker import FuncFormatter
import pandas as pd
import numpy as np
import os
# Imports from FinanceOps.
from curve_fit import CurveFitReciprocal
from data_keys import *
from data import load_index_data, load_stock_data
from returns import prepare_mean_ann_returns
```
## Load Data
We now load all the financial data we will be using.
```
# Define the ticker-names for the stocks we consider.
ticker_SP500 = "S&P 500"
ticker_JNJ = "JNJ"
ticker_K = "K"
ticker_PG = "PG"
ticker_WMT = "WMT"
# Load the financial data for the stocks.
df_SP500 = load_index_data(ticker=ticker_SP500)
df_JNJ = load_stock_data(ticker=ticker_JNJ)
df_K = load_stock_data(ticker=ticker_K)
df_PG = load_stock_data(ticker=ticker_PG)
df_WMT = load_stock_data(ticker=ticker_WMT)
```
## Plotting Functions
These are helper-functions used for making plots.
```
def plot_psales(df, ticker, start_date=None):
"""
Plot the P/Sales ratio.
:param df: Pandas DataFrame with PSALES.
:param ticker: Ticker-name for the stock or index.
:param start_date: Start-date for the plot.
:return: Nothing.
"""
psales = df[PSALES][start_date:].dropna()
psales.plot(title=ticker + " - P/Sales", grid=True)
def plot_ann_returns(ticker, df, key=PSALES,
min_years=7, max_years=15,
use_colors=True):
"""
Create a single scatter-plot with P/Sales or P/Book
vs. Mean Annualized Returns for e.g. 7-15 years.
:param ticker: Ticker-name for the stock or index.
:param df: Pandas DataFrame containing key and TOTAL_RETURN.
:param key: Name of data-column to use e.g. PSALES or PBOOK.
:param min_years: Min number of years for return periods.
:param max_years: Max number of years for return periods.
:param use_colors: Boolean whether to use colors in plot.
:return: Nothing.
"""
# Prepare the data.
# x is the P/Sales or P/Book and y is the Mean Ann. Returns.
x, y = prepare_mean_ann_returns(df=df, key=key,
min_years=min_years,
max_years=max_years)
# Create a single plot.
fig = plt.figure(figsize=(10, 10))
ax = fig.add_subplot(211)
# Scatter-plot.
if use_colors:
# Give each dot in the scatter-plot a shade of blue
# according to the date of the data-point.
ax.scatter(x, y,
c=list(range(len(x))), cmap='Blues',
alpha=1.0, marker='o')
else:
# Use the same color for all dots.
ax.scatter(x, y, marker='o')
# First part of the title.
title1 = "[{0}] {1} vs. {2}-{3} Years Mean Ann. Return"
title1 = title1.format(ticker, key, min_years, max_years)
# X-values for plotting fitted curves.
x_min = np.min(x)
x_max = np.max(x)
x_range = np.arange(x_min, x_max, (x_max/x_min)/1000)
# Plot reciprocal curve-fit.
curve_fit_reciprocal = CurveFitReciprocal(x=x, y=y)
y_pred = curve_fit_reciprocal.predict(x=x_range)
ax.plot(x_range, y_pred, color='red')
# Title with these curve-fit parameters.
title2 = "Mean Ann. Return = {0:.1%} / " + key + " + {1:.1%}"
title2 = title2.format(*curve_fit_reciprocal.params)
# Combine and set the plot-title.
title = "\n".join([title1, title2])
ax.set_title(title)
# Set axis labels.
ax.set_xlabel(key)
ax.set_ylabel("Mean Ann. Return")
# Convert y-ticks to percentages.
# We use a custom FuncFormatter because PercentFormatter
# is inconsistent with string-formatters used elsewhere.
formatter = FuncFormatter(lambda y, _: '{:.0%}'.format(y))
ax.yaxis.set_major_formatter(formatter)
# Show grid.
ax.grid()
# Show the plot.
plt.show()
```
## Case Study: S&P 500
The S&P 500 is a stock-market index consisting of the stocks of 500 of the largest companies in USA. The S&P 500 covers about 80% of the whole U.S. stock-market in terms of size so it is useful as a gauge for the entire U.S. stock-market.
We consider the Total Return of the S&P 500 which is what you would get from investing in the S&P 500 and re-investing all dividends back into the S&P 500. We ignore all taxes here.
The following scatter-plot shows the P/Sales ratio versus the Mean Annualized Returns of the S&P 500 for periods between 7 and 15 years.
For each day we calculate the Total Return of the S&P 500 over the next 7-15 years, then we calculate the Mean Annualized Return from those, and then we put a blue dot in the scatter-plot for that date's P/Sales ratio and the Mean Annualized Return we just calculated. This process is continued for all days in the time-series, until we have calculated and plotted the P/Sales vs. Mean Annualized Return for all days.
As can be seen from this scatter-plot, the P/Sales ratio is a very strong predictor for long investment periods between 7-15 years. We call the fitted red curve for the "return curve".
```
plot_ann_returns(ticker=ticker_SP500, df=df_SP500, key=PSALES,
min_years=7, max_years=15, use_colors=True)
```
We can forecast the future long-term returns using the fitted "return curve" from the scatter-plot above. Towards the end of 2017, the P/Sales ratio was almost 2.2 for the S&P 500, which was about the previous high point of the "Dot-Com" bubble around year 2000.
```
df_SP500[PSALES].dropna().tail(1)
plot_psales(df=df_SP500, ticker=ticker_SP500)
```
So if you purchased the S&P 500 in December 2017 at this P/Sales ratio and will keep the investment for more than 7 years, while reinvesting all dividends during those years (all taxes are ignored), then the formula forecasts an annualized return of about 1.35%:
$$
Annualized\ Return = 14.4\% / (P/Sales) - 5.2\% = 14.4\% / 2.2 - 5.2\% \simeq 1.35\%
$$
The formula cannot predict exactly what will happen in the future, because there might be a stock-market bubble or a crash in any given year. The formula merely predicts an average annualized return for long-term investments of about 7-15 years in the S&P 500.
## Case Study: Johnson & Johnson (JNJ)
Now let us consider individual companies instead of a whole stock-market index. The first company we consider is Johnson & Johnson with the ticker symbol JNJ. This is a very large company with over 130.000 employees worldwide that manufacture a wide range of health-care related products.
When we plot the P/Sales ratio versus the mean annualized return for 7-15 year periods, we see that the "return curve" fits quite well although there appears to be a few separate "return curves" for P/Sales ratios roughly between 2 and 3.
The blue shades in the scatter-plot indicate the time of the data-points and suggest that the separate curves belong to different periods of time. More research would be needed to establish why these periods have different "return curves". Perhaps the periods had significantly different profit-margins or sales-growth.
```
plot_ann_returns(ticker=ticker_JNJ, df=df_JNJ, key=PSALES,
min_years=7, max_years=15, use_colors=True)
```
Towards the end of 2017 the P/Sales ratio was about 4.9 which is close to the all-time historical highs experienced during the stock-market bubble around year 2000.
```
df_JNJ[PSALES].dropna().tail(1)
plot_psales(df=df_JNJ, ticker=ticker_JNJ)
```
Using the formula for the fitted "return curve" from the scatter-plot above, we get this forecasted long-term return:
$$
Annualized\ Return \simeq 77.9\% / (P/Sales) - 8.9\%
\simeq 77.9\% / 4.9 - 8.9\% \simeq 7.0\%
$$
So according to this formula, the annualized return of the JNJ stock will be around 7.0% if you own the stock for at least 7 years, when dividends are reinvested and ignoring taxes.
Again there is the caveat that it is impossible to predict whether there will be a stock-market bubble or crash several years into the future, so the forecasted return is an average for 7-15 year investment periods.
## Case Study: Procter & Gamble (PG)
Another very large company is Procter & Gamble with the ticker symbol PG, which sells a wide range of consumer products and has almost 100.000 employees.
If we plot the P/Sales ratio versus the mean annualized return we get an incredibly regular curve of data-points. The red line shows a reciprocal curve-fit, which is apparently not the correct formula for this data, as it doesn't fit so well at the ends. You are encouraged to try and find a better curve-fit and a theoretical explanation why your formula is better.
```
plot_ann_returns(ticker=ticker_PG, df=df_PG, key=PSALES,
min_years=7, max_years=15)
```
When we plot the historical P/Sales ratio, we see that at the end of 2017 it was around 3.5 which was near its all-time high experienced during the bubble around year 2000.
```
plot_psales(df=df_PG, ticker=ticker_PG)
```
Using the fitted reciprocal curve from the scatter-plot above, we get a forecasted return of about 6.1% per year, when dividends are reinvested without taxes:
$$
Annualized\ Return \simeq 24.4\% / (P/Sales) - 0.9\% \simeq
24.4\% / 3.5 - 0.9\% \simeq 6.1\%
$$
But it should again be noted that this formula doesn't fit so well towards the ends of the data, and looking at the scatter-plot suggests a slightly lower return of maybe 5.5%.
## Case Study: Kellogg's (K)
The next company is Kellogg's which trades under the ticker symbol K. The company has about 33.000 employees and is especially known for making breakfast cereals.
When we plot the P/Sales ratio versus the mean annualized return it shows a strong trend that higher P/Sales ratios gives lower long-term returns, although the curve-fit is not as good as for the other companies we studied above, especially for lower P/Sales ratios.
The blue shades show the time of the data-points. It can be hard to see in this plot, but for P/Sales ratios between 1.50 and 1.75, there is a "blob" of light-blue data-points well above the fitted red curve. This clearly indicates that the outlying data-points belong to a specific period in time. But we would have to do more research into the financial data for that period, to uncover the reason why the returns are so different.
```
plot_ann_returns(ticker=ticker_K, df=df_K, key=PSALES,
min_years=7, max_years=15, use_colors=True)
```
Towards the end of 2017 the P/Sales ratio was about 1.8 which was actually very close to the historical average.
```
df_K[PSALES].dropna().mean()
plot_psales(df=df_K, ticker=ticker_K)
```
Using the fitted "return curve" from the scatter-plot above with the P/Sales ratio of 1.8 we get the forecasted return:
$$
Annualized\ Return \simeq 27.5\% / (P/Sales) - 6.2\% \simeq
27.5\% / 1.8 - 6.2\% \simeq 9.1\%
$$
So a forecasted return of about 9.1% per year over the next 7-15 years when dividends are reinvested without taxes. That is about 2% (percentage points) higher than the return forecasted for JNJ and 3% higher than forecasted for PG above.
## Case Study: Wal-Mart (WMT)
Now let us consider the company Wal-Mart which trades under the ticker symbol WMT. It is an extremely large retail-company with about 2.3 million employees.
If we plot the P/Sales ratio versus the mean annualized return, we see that the red curve fits very poorly. There seems to be several separate trends in the data, and the blue shades indicate that the trends belong to different periods in time. But more research into the company's financial history would be needed to uncover the reason for this, perhaps it is because of significantly different sales-growth, profit margins, etc.
```
plot_ann_returns(ticker=ticker_WMT, df=df_WMT, key=PSALES,
min_years=7, max_years=15, use_colors=True)
```
## Conclusion
We have shown that the P/Sales ratio is a very strong predictor for the long-term returns of the S&P 500 index and some individual stocks.
In the [previous paper](https://github.com/Hvass-Labs/FinanceOps/blob/master/01_Forecasting_Long-Term_Stock_Returns.ipynb) we considered fixed investment periods of e.g. 10 years, which meant that the investment return depended on the P/Sales ratio both at the time of buying and selling. This distorted the data because sometimes the stock-market would be in a bubble or crash 10 years later.
In this paper we presented a simple solution by considering all investment periods between 7 and 15 years, and then using the average return instead. This averages out the distorting effects of future bubbles and crashes, so we get much more smooth data that only depends on the P/Sales ratio at the buy-time.
We then fitted a reciprocal "return curve" to the scatter-plots, and although it generally had a very tight fit, it was not so accurate towards the end-points, thus suggesting that the reciprocal formula is not entirely correct for this data. It would be of great interest to not only find a mathematical model that fits better, but also a theoretical explanation why that model makes sense. Perhaps such a model would also allow us to use smaller amounts of data and take into account the changing economics of a business. Perhaps we could use such a model to forecast returns of more companies where the basic method does not work so well, such as Wal-Mart as demonstrated above.
It should be stressed that the forecasted returns will also depend on a *qualitative* assessment of the company. If the company's future will be significantly different from its historical sales, profit-margins and growth, then the forecasted returns will be inaccurate. That is why this forecasting method is perhaps best used on broad stock-market indices such as the S&P 500, or companies whose products and markets are expected to be highly predictable long into the future.
## Research Ideas
You are strongly encouraged to do more research on this topic. If you make any new discoveries then please let me know your results.
To my knowledge, there are no academic studies of predicting the long-term returns of stocks and stock-markets as we have done here. This work has presented the basic idea and methodology, but a lot more research can be done on this subject and it may impact many areas of both theoretical and applied finance.
Here are a few more research ideas to get you started, in addition to the ideas from the [previous paper](https://github.com/Hvass-Labs/FinanceOps/blob/master/01_Forecasting_Long-Term_Stock_Returns.ipynb):
- Try other investment periods, for example 5 to 10 years. How does it change the scatter-plots and the fitted "return curves"?
- Try using P/Book as the predictor signal. How does that affect the plots? Why?
- Although the data in some of these scatter-plots is incredibly smooth, the reciprocal curve does not fit the data exactly, which suggests that it is the wrong formula for this data. Can you find a better formula and perhaps give a theoretical explanation why that is better?
- What is the reason that some companies such as Wal-Mart have several different trend-lines in the scatter-plot? You will probably need to investigate the historical financial data to uncover the reason. Can you modify the forecasting method to somehow take this into account?
## License (MIT)
Copyright (c) 2015-18 by [Magnus Erik Hvass Pedersen](http://www.hvass-labs.org/)
Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
| github_jupyter |
# Memory usage for FlexDriver and CPU driver
This notebook estimates the memory usage for an mlx5 driver for RDMA and compares it to the memory usage for FlexDriver.
```
%config InlineBackend.figure_format = 'retina'
from copy import copy
import numpy as np
import pandas as pd
import plotnine as p9
from common import *
def log2(value):
return np.ceil(np.log2(value)).astype('int')
def round_pow2(value):
return np.left_shift(1, log2(value))
def cuckoo_hash(tag_size, value_size):
'''Compute the size of the cuckoo hash table in bytes.'''
num_banks = 4
num_entries = 2 * round_pow2(value_size) / num_banks
tag_width = log2(tag_size)
value_width = log2(value_size)
return num_banks * num_entries * (tag_width + value_width) / 8
def bdp(df):
'''Compute the bandwidth-delay product.'''
linerate = df['linerate_gbps']
df[f'packet_rate_mpps'] = linerate / (df['min_packet_size_bytes'] + 20) * 125
for dir in ('tx', 'rx'):
latency = df[f'latency_{dir}_usec']
df[f'{dir}_bdp_bytes'] = np.ceil(linerate * latency * 125)
df[f'{dir}_bdp_descs'] = np.ceil(df['packet_rate_mpps'] * latency)
def outstanding_buffer_fragmentation(df):
'''Compute necessary buffer size for outstanding receive/transmit packets,
taking fragmentation into account.'''
# Use double the computed size for fragmentation (assumes minimal packet size >= block size)
assert (df['fld_tx_data_block_bytes'] <= df['min_packet_size_bytes']).all()
rx_stride = 64 # byte
assert (rx_stride <= df['min_packet_size_bytes']).all()
# Standard SRQ uses same computation as unshared tx buffer
for dir in ('tx', 'rx'):
df[f'{dir}_descs'] = df[f'{dir}_bdp_descs']
# FLD
outstanding_fragmentation_bytes = df[f'{dir}_bdp_bytes'].rename(f'{dir}_outstanding_fragmentation_bytes') * 2
# not FLD
outstanding_fragmentation_bytes = outstanding_fragmentation_bytes.where(df['fld'],
df[f'{dir}_descs'] * df['max_packet_size_bytes'])
df[f'{dir}_outstanding_bytes'] = np.maximum(df[f'{dir}_bdp_bytes'], outstanding_fragmentation_bytes)
def srq(df):
'''Compute the size of the receive queue with FLD and with a standard driver.'''
df['rwqe_bytes'] = np.where(df['fld'], 32, 16)
df['cqe_bytes'] = np.where(df['fld'], 120 / 8, 64)
rx_descs_pow2 = round_pow2(df['rx_descs'])
df['srq_wq_bytes'] = (rx_descs_pow2 * df['rwqe_bytes']).where(~df['fld'], 0)
df['rcq_bytes'] = rx_descs_pow2 * df['cqe_bytes']
df['srq_doorbell_record_bytes'] = 4
wqebb_bytes = 64
fld_wqe_bytes = 8
def txq(df):
'''Compute the size of the transmit queues.'''
tx_descs_pow2 = round_pow2(df['tx_descs'])
df['twqe_bytes'] = np.where(df['fld'], fld_wqe_bytes, wqebb_bytes)
txq_wq_bytes_fld = tx_descs_pow2 * df['twqe_bytes']
txq_wq_bytes_no_fld = tx_descs_pow2 * df['num_tx_queues'] * df['twqe_bytes']
df['txq_wq_trans_table_tag_size'] = tx_descs_pow2 * df['num_tx_queues']
txq_wq_translation_table_bytes_fld = cuckoo_hash(tag_size=df['txq_wq_trans_table_tag_size'],
value_size=df['tx_descs'])
df['txq_data_trans_table_tag_size'] = (df['tx_outstanding_bytes'] * df['num_tx_queues'] / df['fld_tx_data_block_bytes'])
txq_data_translation_table_bytes_fld = cuckoo_hash(tag_size=df['txq_data_trans_table_tag_size'],
value_size=df['tx_outstanding_bytes'] / df['fld_tx_data_block_bytes'])
df['txq_wq_bytes'] = txq_wq_bytes_fld.where(df['fld'], txq_wq_bytes_no_fld)
df['txq_wq_translation_table_bytes'] = txq_wq_translation_table_bytes_fld.where(df['fld'], 0)
df['txq_data_translation_table_bytes'] = txq_data_translation_table_bytes_fld.where(df['fld'], 0)
df['tcq_bytes'] = tx_descs_pow2 * df['cqe_bytes']
df['txq_doorbell_record_bytes'] = 4 * df['num_tx_queues']
metadata_fields = ['srq_wq_bytes', 'rcq_bytes', 'txq_wq_bytes', 'tcq_bytes',
'srq_doorbell_record_bytes', 'txq_doorbell_record_bytes',
'txq_wq_translation_table_bytes',
'txq_data_translation_table_bytes']
data_fields = [f'{dir}_outstanding_bytes' for dir in ('tx', 'rx')]
def mem_usage(df):
'''Compute usage of the different driver components with FlexDriver and for
a CPU driver.'''
keys = list(df.columns)
bdp(df)
outstanding_buffer_fragmentation(df)
srq(df)
txq(df)
df = df.set_index(keys)
df['total_pi_bytes'] = df['txq_doorbell_record_bytes'] + df['srq_doorbell_record_bytes']
df['total_cq_bytes'] = df['rcq_bytes'] + df['tcq_bytes']
df['total_metadata_bytes'] = df.groupby(keys)[metadata_fields].sum().sum(axis=1)
df['total_data_bytes'] = df.groupby(keys)[data_fields].sum().sum(axis=1)
df['total_bytes'] = df['total_metadata_bytes'] + df['total_data_bytes']
return df.reset_index()
# Scenarios: FLD / CPU x Linerate x No. TX queues
df = (pd.DataFrame({'fld': (False, True)})
.merge(pd.DataFrame({'linerate_gbps': [50 << i for i in range(4)]}), how='cross')
.merge(pd.DataFrame({'num_tx_queues': [1 << i for i in range(0, 13)]}), how='cross')
).assign(latency_tx_usec=25, latency_rx_usec=5, min_packet_size_bytes=256, max_packet_size_bytes=1 << 14, fld_tx_data_block_bytes=256)
df = mem_usage(df)
# add KiB/MiB units results
for col in df.columns:
if col.endswith('bytes'):
df[col.split('_bytes')[0] + '_kib'] = df[col] / (1 << 10)
df[col.split('_bytes')[0] + '_mib'] = df[col] / (1 << 20)
interesting_rows = (df.linerate_gbps.isin((100,))) & (df.num_tx_queues == 512)
keys = ['linerate_gbps', 'fld', 'num_tx_queues']
table_2a_cols = ['min_packet_size_bytes', 'max_packet_size_kib', 'latency_rx_usec', 'latency_tx_usec', 'packet_rate_mpps', 'tx_bdp_descs', 'rx_bdp_descs', 'tx_bdp_kib', 'rx_bdp_kib']
df[interesting_rows][keys + table_2a_cols]
table_2b_cols = ['twqe_bytes', 'rwqe_bytes', 'cqe_bytes']
df[interesting_rows][keys + table_2b_cols]
table_3_cols = [
'txq_wq_mib', 'txq_wq_kib',
'txq_wq_translation_table_kib',
'tx_outstanding_mib', 'tx_outstanding_kib',
'txq_data_translation_table_kib',
'rx_outstanding_mib', 'rx_outstanding_kib',
'total_cq_kib',
'srq_wq_mib', 'srq_wq_kib',
'total_pi_bytes',
'total_mib',
'total_kib']
df[interesting_rows][keys + table_3_cols]
# Color palette
software_color, fld_color, fpga_color = pal(3)
def linerate_labeller(linerate):
return f'{linerate} Gbps'
def log2_labeller(breaks):
return [f'$2^{{{i}}}$' for i in np.log2(breaks).astype('int')]
def minor_breaks(limits):
major = [10 ** i for i in range(0, 5)]
res = [m * 1.5 for m in major
if m * 1.5 < limits[1]]
return res
limit_xcku15p_df = pd.DataFrame({'type': ['BRAM', 'URAM', 'LUTRAM'],
'Mibit': [34.6, 36, 9.8]})
limit_xcku15p_mib = limit_xcku15p_df['Mibit'].sum() / 8
grid_color = p9.options.current_theme.themeables['panel_grid_major'].theme_element.properties['color']
ticks_color = "#000000"
df['total_mib'] = df['total_bytes'] / (2 ** 20)
df['fld_name'] = np.where(df['fld'], 'FLD', 'software')
plt = (p9.ggplot(df, p9.aes(x='num_tx_queues', y='total_mib', linetype='fld_name', color='fld_name')) +
p9.geom_line() +
p9.geom_hline(mapping=p9.aes(linetype=['XCKU15P limit'], color=['XCKU15P limit'], yintercept=limit_xcku15p_mib)) +
p9.scale_x_continuous(trans='log2', name='# transmit queues', labels=log2_labeller) +
p9.scale_y_continuous(trans='log10', name='shared memory [MiB]',
minor_breaks=minor_breaks) +
p9.facet_grid('~ linerate_gbps', labeller=linerate_labeller) +
p9.theme(
figure_size=(fig_width, 1),
subplots_adjust={'top': 0.77},
axis_ticks_minor_y=p9.element_line(ticks_color),
panel_grid_minor_y=p9.element_line(color=grid_color, size=0.5),
) +
top_legend + \
p9.scale_color_manual(values=[fld_color, software_color, fpga_color])
)
plt
plt.save('mem_usage.pdf')
reduction = copy(df)
reduction['tx_meta_total_bytes'] = reduction['txq_wq_bytes'] + reduction['txq_wq_translation_table_bytes']
reduction['tx_data_total_bytes'] = reduction['tx_outstanding_bytes'] + reduction['txq_data_translation_table_bytes']
reduction_cols = ['tx_meta_total_bytes', 'tx_data_total_bytes', 'total_cq_bytes', 'srq_wq_bytes', 'rx_outstanding_bytes', 'total_bytes']
reduction = reduction.pivot_table(values=reduction_cols, columns=['fld'], index=['linerate_gbps', 'num_tx_queues'])
def slice_fld(df, fld):
new_df = df.loc[:, [(col, fld) for col in reduction_cols]]
new_df.columns = new_df.columns.droplevel(1)
return new_df
no_fld = slice_fld(reduction, False)
fld = slice_fld(reduction, True)
ratio = no_fld / fld
ratio.columns = pd.MultiIndex.from_tuples((c, 'ratio') for c in ratio.columns)
reduction = pd.concat([reduction, ratio], axis=1)#.sort_values(by=1, axis=1)
reduction.columns.names = ['type', 'col']
reduction = reduction.sort_values(by=['type', 'col'], axis=1)
reduction[[(col, 'ratio') for col in reduction_cols]].loc[(100, 512)]
```
| github_jupyter |
# Kaggle Seattle Airbnb
## Project Info
- Author: [Zacks Shen](https://www.linkedin.com/in/zacks-shen/)
- Contributor: [Kevin Chu](https://www.linkedin.com/in/yen-duo-chu/)
> [GitHub: Code with Full Output](https://github.com/ZacksAmber/Kaggle-Seattle-Airbnb)<br>
> [Kaggle: Code](https://www.kaggle.com/zacksshen/kaggle-seattle-airbnb)<br>
> [Medium: Article](https://medium.com/@zacks.shen/how-airbnb-evaluates-the-listings-b35c5a7890cb)
---
### Reference
> [Kaggle](https://www.kaggle.com/airbnb/seattle)<br>
> [Data Source](http://insideairbnb.com/seattle)<br>
> [Map](http://insideairbnb.com/get-the-data.html)
---
### Why Price Prediction?
Price prediction is beneficial for **Perfect Competition**, a triple-win market.
- Hosts have an intuitive opportunity to compare their services and amenities with competitors. As competition intensifies, the overall service quality and market - size of the rental housing market will be improved.
- Price prediction models are reliable references for data-driven decision-making. With price suggestions from Airbnb, hosts can make different business strategies.
- Airbnb users have more choices at lower price or higher quality.
- Airbnb can grow faster including attract more users and hosts, leading to opertional and data center cost reduction and profit growth.
---
### Cons
The model accuracy will be lower than expected due to missing some essential features.
- The transaction data: The datasets did not contain transaction data.
- We don't know if the listings were booked or just unavailable.
- We will explore how to find the confirmed orders under the Exploratory Data Analysis part.
- The search engine data: The datasets do not contain searching histories.
---
## Conclusion
Hosts could adjust the following features for improving their competitiveness.
- Decrease
- `cleaning_fee`
- `security_deposit`
- Change
- `cancellation_policy`
- `bed_type`
- `beds`
- `guests_included`
- `extra_people`
- `accommodates`
- Amenities:
- `Indoor Fireplace`
- `Cable TV`
- `TV`
- `Doorman`
- `Dryer`
- `Air Conditioning`
- `Gym`
- `Family/Kid Friendly`
- `Kitchen`
- `Washer`
- Hard to change:
- `neighbourhood_group_cleansed`
- `room_type`
- `bathrooms`
- `bedrooms`
Although we don't have the transaction data, and the number of reviews did not play an essential role in price prediction, I'm sure such features will influence the rank on the search engine.
Airbnb uses machine learning models to intensify the competition. They can provide Price Tips, Smart Pricing, and improvement suggestions to the host. As a host, you can combine the models and good strategies to maximize your competitiveness and profit. Airbnb users are also benefited from the perfect competition.
---
# Airbnb Exploration
## Who needs the prediction? The host.
<details>
<summary><b>Click to see Price Tips example</b></summary>
<img src='https://raw.githubusercontent.com/ZacksAmber/PicGo/master/img/20210919210229.png' alt='The price is suggested by Airbnb'>
<img src='https://raw.githubusercontent.com/ZacksAmber/PicGo/master/img/20210919210423.png' >
</details>
---
## How Airbnb determines the price of a listing?
Currently, Airbnb has at least two Machine Learning models for price prediction. They are both for hosts:
### Price Tips
> **Price Tips**: Price tips are automated nightly price recommendations that you can choose to save. They are based on the type and location of your listing, the season, demand, and other factors. Even if you use price tips, you always control your price and can override the tips at any time. — Refered from Airbnb.<br>
<details>
<summary><b>Click to see Price Tips</b></summary>
<picture>
<img src='https://raw.githubusercontent.com/ZacksAmber/PicGo/master/img/20211009230743.png' alt='Price Tips'>
<img src='https://raw.githubusercontent.com/ZacksAmber/PicGo/master/img/20211009230813.png' alt='Smart Pricing with Price Tips'>
</picture>
</details>
---
### Smart Pricing
> **Smart Pricing**: Smart Pricing lets you set your prices to automatically go up or down based on changes in demand for listings like yours. You are always responsible for your price, so Smart Pricing is controlled by other pricing settings you choose, and you can adjust nightly prices any time. Smart Pricing is based on the type and location of your listing, the season, demand, and other factors. — Refered from Airbnb.
The following features are defined by Airbnb:
- How many people are searching for listings like yours
- The dates they’re looking at
- Whether other listings are getting booked
- Your listing’s best qualities
- Your neiborhood: To calculate pricing based on location, Smart Pricing looks at whether your listing is in a city neighborhood, a suburb, or a more spread-out area.
- Review rate: The number and quality of your reviews is another key factor in Smart Pricing.
- Completed trips: If you honor most confirmed reservations, your prices can go higher within the minimum and maximum range you set.
- Your listing' amenities: Wi-fi, washer/dryer, and air conditioning are especially important, but Smart Pricing looks at all your amenities.
<details>
<summary><b>Click to see Smart Pricing</b></summary>
<picture>
<img src='https://raw.githubusercontent.com/ZacksAmber/PicGo/master/img/20210919213149.png'>
<img src='https://raw.githubusercontent.com/ZacksAmber/PicGo/master/img/20210919213824.png'>
<img src='https://raw.githubusercontent.com/ZacksAmber/PicGo/master/img/20210919214931.png'>
<img src='https://raw.githubusercontent.com/ZacksAmber/PicGo/master/img/20210919214948.png'>
<img src='https://raw.githubusercontent.com/ZacksAmber/PicGo/master/img/20210919215026.png'>
<img src='https://raw.githubusercontent.com/ZacksAmber/PicGo/master/img/20210919215039.png'>
<img src='https://raw.githubusercontent.com/ZacksAmber/PicGo/master/img/20210919215201.png'>
<img src='https://raw.githubusercontent.com/ZacksAmber/PicGo/master/img/20211003122330.png'>
</picture>
</details>
---
## What is Calendar?
The calendar contains three types of price:
- Base price set by the host.
- One time special price for event set by the host.
- Price based on the demand, which is automatically set by **Smart Pricing**.
<details>
<summary><b>Click to see Calendar</b></summary>
See how I turned on Smart Pricing, set a one time price at a specific day, and set a day unavailable.
<picture>
<img src='https://raw.githubusercontent.com/ZacksAmber/PicGo/master/img/20211003123104.png'>
<img src='https://raw.githubusercontent.com/ZacksAmber/PicGo/master/img/20211003123145.png'>
<img src='https://raw.githubusercontent.com/ZacksAmber/PicGo/master/img/20211003123708.png'>
<img src='https://raw.githubusercontent.com/ZacksAmber/PicGo/master/img/20211003123609.png'>
</picture>
</details>
---
# Dependencies
```
# Statistics
import pandas as pd
import numpy as np
import math as mt
# Data Visualization
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
import plotly.express as px
import plotly.graph_objects as go
import plotly.io as pio
px.defaults.width = 1200
px.defaults.height = 800
# plotly.io Settings for both plotly.graph_objects and plotly.express
pio.templates.default = "plotly_white" # "plotly", "plotly_white", "plotly_dark", "ggplot2", "seaborn", "simple_white", "none"
"""
pio.kaleido.scope.default_format = 'svg'
pio.kaleido.scope.default_scale = 1
"""
# Data Preprocessing - Standardization, Encoding, Imputation
from sklearn.preprocessing import StandardScaler # Standardization
from sklearn.preprocessing import Normalizer # Normalization
from sklearn.preprocessing import OneHotEncoder # One-hot Encoding
from sklearn.preprocessing import OrdinalEncoder # Ordinal Encoding
from category_encoders import MEstimateEncoder # Target Encoding
from sklearn.preprocessing import PolynomialFeatures # Create Polynomial Features
from sklearn.impute import SimpleImputer # Imputation
# Exploratory Data Analysis - Feature Engineering
from sklearn.preprocessing import PolynomialFeatures
from sklearn.feature_selection import mutual_info_regression
from sklearn.decomposition import PCA
# Modeling - ML Pipelines
from sklearn.pipeline import Pipeline
from sklearn.compose import ColumnTransformer
from sklearn.model_selection import train_test_split
from sklearn.model_selection import KFold
# Modeling - Algorithms
from xgboost import XGBRegressor
from lightgbm import LGBMRegressor
#from catboost import CatBoostRegressor
from sklearn.linear_model import LinearRegression
from sklearn.ensemble import RandomForestRegressor, GradientBoostingRegressor
# ML - Evaluation
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import cross_val_score
# ML - Tuning
import optuna
#from sklearn.model_selection import GridSearchCV
# Settings
# Settings for Seaborn
sns.set_theme(context='notebook', style='ticks', palette="bwr_r", font_scale=0.7, rc={"figure.dpi":240, 'savefig.dpi':240})
```
---
## Load Datasets
- listings: The listings' information including nightly price (base price).
- reviews: All of the past reviews for each listing.
- calendar: The availability for each listing with the base price, special price, or smart price.
```
import os
kaggle_project = 'seattle'
# Import dataset from local directory './data' or from Kaggle
data_dir = ('./data/201601' if os.path.exists('data') else f'/kaggle/input/{kaggle_project}')
# print all files in data_dir
for dirname, _, filenames in os.walk(data_dir):
for filename in filenames:
print(os.path.join(dirname, filename))
# Import three datasets
reviews = pd.read_csv(f'{data_dir}/reviews.csv')
calendar = pd.read_csv(f'{data_dir}/calendar.csv')
listings = pd.read_csv(f'{data_dir}/listings_kfold.csv') if os.path.exists(f'{data_dir}/listings_kfold.csv') else pd.read_csv(f'{data_dir}/listings.csv')
```
---
## Cross-Validation KFold
```
def generate_listings_kfold():
# Mark the train dataset with kfold = 5
listings = pd.read_csv(f'{data_dir}/listings.csv')
if os.path.exists(f'{data_dir}/listings_kfold.csv'):
os.remove(f'{data_dir}/listings_kfold.csv')
kf = KFold(n_splits=5, shuffle=True, random_state=42)
for fold, (train_idx, valid_idx) in enumerate(kf.split(X=listings)):
listings.loc[valid_idx, "kfold"] = fold
listings.to_csv(f'listings_kfold.csv', index=False)
generate_listings_kfold()
listings = pd.read_csv(f'listings_kfold.csv')
# After assigning kfold
# If error, run the above function then re-load listings_kfold.csv
listings.loc[:, ['id', 'kfold']].head()
```
---
## Load Metadata
The [metadata](https://docs.google.com/spreadsheets/d/1M_qah-ym6O8vDcSmoKAP-lbZRPHUey83R_DJaW3LXfs/edit?usp=sharing) was analyzed and made by [Zacks Shen](https://www.linkedin.com/in/zacks-shen/) and [Kevin Chu](https://www.linkedin.com/in/yen-duo-chu/).
The medata data includes includes `Label`, `Data Type`, `Description`, `ML`, `Reason`:
- `Label`: the column name
- `Data Type`: the data type of the column
- `Description`: the label usage based on our observation from Airbnb
- `ML`: is it useful for Machine Learning
- `0`: cannot be used for ML since it is meaningless, or it's hard to measuring (e.g. listing photos)
- `1`: must be used for ML due to the official description by Airbnb
- `2`: possible be used for ML due to our assessment
- `Reason`: why the label can or cannot be used for ML (may empty)
```
# Define sheet id and base url
sheet_id = "1M_qah-ym6O8vDcSmoKAP-lbZRPHUey83R_DJaW3LXfs"
base_url = f"https://docs.google.com/spreadsheets/d/{sheet_id}/gviz/tq?tqx=out:csv&sheet="
# Load metadata for three datasets
listings_metadata = pd.read_csv(base_url+"listings")
calendar_metadata = pd.read_csv(base_url+"calendar")
reviews_metadata = pd.read_csv(base_url+"reviews")
```
---
# ETL Pipeline
The ETL pipeline provides data transformation and formatting. Thus, we can calculate the data and perform machine learning with the correct data format.
```
class ETL_pipeline:
def __init__(self, data_frame):
self.df = data_frame
# Data type transformation
def _transformation(self, data_frame):
df = data_frame
# Convert dollar columns from object to float
# Remove '$' and ','
dollar_cols = ['price', 'weekly_price', 'monthly_price', 'extra_people', 'security_deposit', 'cleaning_fee']
for dollar_col in dollar_cols:
df[dollar_col] = df[dollar_col].replace('[\$,]', '', regex=True).astype(float)
# Convert dollar columns from object to float
# Remove '%'
percent_cols = ['host_response_rate', 'host_acceptance_rate']
for percent_col in percent_cols:
df[percent_col] = df[percent_col].replace('%', '', regex=True).astype(float)
# Replace the following values in property_type to Unique space due to small sample size
unique_space = ["Barn",
"Boat",
"Bus",
"Camper/RV",
"Treehouse",
"Campsite",
"Castle",
"Cave",
"Dome House",
"Earth house",
"Farm stay",
"Holiday park",
"Houseboat",
"Hut",
"Igloo",
"Island",
"Lighthouse",
"Plane",
"Ranch",
"Religious building",
"Shepherd’s hut",
"Shipping container",
"Tent",
"Tiny house",
"Tipi",
"Tower",
"Train",
"Windmill",
"Yurt",
"Riad",
"Pension",
"Dorm",
"Chalet"]
df.property_type = df.property_type.replace(unique_space, "Unique space", regex=True)
# Convert 't', 'f' to 1, 0
tf_cols = ['host_is_superhost', 'instant_bookable', 'require_guest_profile_picture', 'require_guest_phone_verification']
for tf_col in tf_cols:
df[tf_col] = df[tf_col].replace('f', 0, regex=True)
df[tf_col] = df[tf_col].replace('t', 1, regex=True)
return df
# Parse listings
def parse_listings(self):
"""Parse listings.
"""
df = self.df
df = self._transformation(df)
return df
def parse_reviews(self):
"""Parse reviews.
"""
df = self.df
df.date = pd.to_datetime(df.date)
return df
# Parse calendar
def parse_calender(self):
"""Paser calendar.
"""
df = self.df
# Convert date from object to datetime
df.date = pd.to_datetime(df.date)
# Convert price from object to float
# Convert '$' and ',' to ''
df.price = df.price.replace('[\$,]', '', regex=True).astype(float)
# Convert 't', 'f' to 1, 0
df['available'] = df['available'].replace('f', 0, regex=True)
df['available'] = df['available'].replace('t', 1, regex=True)
return df
# e.g. Before parsing
listings.loc[:4, ['id', 'price']]
listings = ETL_pipeline(listings).parse_listings()
reviews = ETL_pipeline(reviews).parse_reviews()
calendar = ETL_pipeline(calendar).parse_calender()
# e.g. After parsing
listings.loc[:4, ['id', 'price']]
```
---
# ML Pipeline
`EDA_demand` calculates the demand for each listing from csv reviews.<br>
`ML Pipeline` imputes and transforms data for Machine Learning.
```
class EDA_demand:
def __init__(self):
pass
def reviews_rate_vs_unavailability(self, period=30):
"""Calculate the booked listing from file calendar.
Args:
period (int): Positive integer. Default is 30.
Returns:
Pandas DataFrame.
"""
assert (0 < period <= 365) & isinstance(period, int), "period must be an integer and greater than 0"
self.period = period
#
# Calculate review rate & unavailability
#
# reviews Rate: review / days
"""
SELECT
listing_id,
COUNT(listing_id) / DATEDIFF(20160104+1, MIN(date)) AS reviews_per_day
FROM reviews
GROUP BY listing_id
"""
# Extract the first reviews date for each listing
func = lambda df: pd.Series({'first_day': df.date.min()})
df_reviews_per_day = pd.DataFrame(reviews.groupby('listing_id').apply(func))
# Define last scraped date
last_scraped = listings.last_scraped.unique()[0]
last_scraped = pd.Timestamp(last_scraped)
df_reviews_per_day['last_day'] = last_scraped + pd.DateOffset(days=1)
# Calculate the datediff
df_reviews_per_day['datediff'] = df_reviews_per_day.last_day - df_reviews_per_day.first_day
df_reviews_per_day['datediff'] = df_reviews_per_day['datediff'].dt.days
# Calculate the reviews Rate
df_reviews_per_day['reviews_per_day'] = reviews.groupby('listing_id').size() / df_reviews_per_day['datediff']
"""
SELECT listing_id, SUM(IF(available = 0, 1, 0))
FROM calendar
WHERE DATEDIFF(date, 20160104) <= period
GROUP BY listing_id
"""
last_day = last_scraped + pd.DateOffset(days=period-1)
filter = calendar.date <= (last_day)
func = lambda df: pd.Series({f'unavailability_{period}_unscaled': sum(df.available == 0)}) # Scaling available to day scale
df_unavailability = pd.DataFrame(calendar[filter].groupby('listing_id').apply(func))
df_unavailability[f'unavailability_{period}'] = df_unavailability[f'unavailability_{period}_unscaled'] / period
#df_unavailability['first_day'] = last_scraped
#df_unavailability['last_day'] = last_day
self.df_unavailability = df_unavailability
# Join two tables
df_unavailability_reviews = df_unavailability.join(df_reviews_per_day, how='left')
df_unavailability_reviews.reviews_per_day.fillna(value=0, inplace=True)
#df_unavailability_reviews.loc[:, [f'unavailability_{period}_unscaled', f'unavailability_{period}', 'reviews_per_day']]
# Find outliers (unavailable rather than booked)
# Extrat quantiles
reviews_rate_25 = df_unavailability_reviews.reviews_per_day.quantile(q=0.25, interpolation='higher')
unavailability_75 = df_unavailability_reviews[f'unavailability_{period}'].quantile(q=0.75, interpolation='higher')
# Low reviews rate: 0.010376
filter1 = df_unavailability_reviews.reviews_per_day < reviews_rate_25
# High unavailability: 0.660274
filter2 = df_unavailability_reviews[f'unavailability_{period}'] > unavailability_75
outliers = df_unavailability_reviews[filter1 & filter2]
df_unavailability_reviews['demand'] = df_unavailability_reviews[f'unavailability_{period}_unscaled']
df_unavailability_reviews.loc[outliers.index, 'demand'] = period - df_unavailability_reviews.loc[outliers.index, 'demand']
self.outliers = outliers
self.df_unavailability_reviews = df_unavailability_reviews
return self.df_unavailability_reviews
def plot(self, outliers=True):
"""Display plot or describe the relationship between reviews per day and unavailabilities to filter the outliers of demand.
Args:
outlier (bool): Display outliers or not. Default is True
Returns:
Plotly instance
"""
period = self.period
if outliers is True:
idx = self.outliers.index
df = self.df_unavailability_reviews.loc[idx, :]
else:
idx = self.df_unavailability_reviews.index.drop(self.outliers.index)
df = self.df_unavailability_reviews.loc[idx, :]
assert df.shape[0] > 0, "No records"
fig = px.line(df,
x=df.index,
y=[f'unavailability_{period}', 'reviews_per_day'],
color_discrete_sequence=['rgb(71, 92, 118, 0.9)', 'rgb(250, 211, 102, 0.9)']
)
fig.update_layout(title=f'Unavailability per day vs. reviews per day<br>Outliers', xaxis_title='index', yaxis_title='Rate')
return fig
class ML_pipeline:
"""ML Pipeline for listings.
"""
def __init__(self, data_frame, features, target, days=365):
"""
Args:
data_frame (Pandas DataFrame): listings.
features (list): The Machine Learning features.
target (str): price
days (int): The days after 2016-01-04 for calculating demand.
"""
import warnings
warnings.filterwarnings("ignore") # ignore target encoding warnings
# Get demand
demand = EDA_demand().reviews_rate_vs_unavailability(days)
# The index will change to id
data_frame = data_frame.set_index('id').join(demand['demand'], how='inner')
features.append(target)
data_frame = data_frame[features]
# Encode amenities
data_frame = self._encode_amentities(data_frame)
data_frame.pop('amenities')
self.data_frame = data_frame
# encode amentities
def _encode_amentities(self, data_frame):
# Replace amenities from {}" to ''
data_frame.amenities.replace('[{}"]', '', regex=True, inplace=True)
# Split amenities with ,
amenities = data_frame.amenities.str.split(',', expand=True)
"""All amenities
'24-Hour Check-in',
'Air Conditioning',
'Breakfast',
'Buzzer/Wireless Intercom',
'Cable TV',
'Carbon Monoxide Detector',
'Cat(s)',
'Dog(s)',
'Doorman',
'Dryer',
'Elevator in Building',
'Essentials',
'Family/Kid Friendly',
'Fire Extinguisher',
'First Aid Kit',
'Free Parking on Premises',
'Gym',
'Hair Dryer',
'Hangers',
'Heating',
'Hot Tub',
'Indoor Fireplace',
'Internet',
'Iron',
'Kitchen',
'Laptop Friendly Workspace',
'Lock on Bedroom Door',
'Other pet(s)',
'Pets Allowed',
'Pets live on this property',
'Pool',
'Safety Card',
'Shampoo',
'Smoke Detector',
'Smoking Allowed',
'Suitable for Events',
'TV',
'Washer',
'Washer / Dryer',
'Wheelchair Accessible',
'Wireless Internet'
"""
# For each col, extract the unique amenities
amenities_uniques = []
for col in amenities.columns:
amenities_uniques += list(amenities[col].unique())
# Remove the duplicate values
amenities_uniques = set(amenities_uniques)
amenities_uniques.remove('')
amenities_uniques.remove(None)
# Only two rows have Washer / Dryer, and they both have washer and dryer
amenities_uniques.remove('Washer / Dryer')
# When 'Pets live on this property' is True, one or more from 'Cat(s)', 'Dog(s)', 'Other pet(s)' will appear
# Encoding amenities
amenities_enc = pd.DataFrame()
for amenity in amenities_uniques:
amenities_enc[amenity] = data_frame.amenities.str.contains(amenity, regex=False)
# Rename the columns with prefix amenity_
amenities_enc.columns = [f"amenity_{col}" for col in amenities_enc.columns]
# Concat encoded amenities and data_frame
data_frame = pd.concat([data_frame, amenities_enc], axis=1)
return data_frame
def _imputation(self, X_train, X_valid, y_train, y_valid):
X_train, X_valid, y_train, y_valid = X_train.copy(), X_valid.copy(), y_train.copy(), y_valid.copy()
# Zero imputation
# Reason:
zero_imp = SimpleImputer(missing_values=np.nan, strategy='constant', fill_value=0)
zero_features = ['reviews_per_month', 'host_response_rate', 'host_is_superhost', 'security_deposit', 'cleaning_fee']
X_train_zero_imp = pd.DataFrame(zero_imp.fit_transform(X_train[zero_features]))
X_valid_zero_imp = pd.DataFrame(zero_imp.transform(X_valid[zero_features]))
X_train_zero_imp.columns = zero_features
X_valid_zero_imp.columns = zero_features
X_train_zero_imp.index = X_train.index
X_valid_zero_imp.index = X_valid.index
X_train_zero_imp = X_train_zero_imp.astype(float)
X_valid_zero_imp = X_valid_zero_imp.astype(float)
# Mean imputation
# Reason:
mean_imp = SimpleImputer(missing_values=np.nan, strategy='mean')
mean_features = ['host_acceptance_rate', 'review_scores_accuracy', 'review_scores_checkin',
'review_scores_value', 'review_scores_location', 'review_scores_cleanliness',
'review_scores_communication', 'review_scores_rating']
X_train_mean_imp = pd.DataFrame(mean_imp.fit_transform(X_train[mean_features]))
X_valid_mean_imp = pd.DataFrame(mean_imp.transform(X_valid[mean_features]))
X_train_mean_imp.columns = mean_features
X_valid_mean_imp.columns = mean_features
X_train_mean_imp.index = X_train.index
X_valid_mean_imp.index = X_valid.index
X_train_mean_imp = X_train_mean_imp.astype(float)
X_valid_mean_imp = X_valid_mean_imp.astype(float)
# Mode imputation
# Reason:
mode_imp = SimpleImputer(missing_values=np.nan, strategy='most_frequent')
mode_features = ['bathrooms', 'bedrooms', 'beds', 'property_type']
X_train_mode_imp = pd.DataFrame(mode_imp.fit_transform(X_train[mode_features]))
X_valid_mode_imp = pd.DataFrame(mode_imp.transform(X_valid[mode_features]))
X_train_mode_imp.columns = mode_features
X_valid_mode_imp.columns = mode_features
X_train_mode_imp.index = X_train.index
X_valid_mode_imp.index = X_valid.index
X_train_mode_imp[['bathrooms', 'bedrooms', 'beds']] = X_train_mode_imp[['bathrooms', 'bedrooms', 'beds']].astype(int)
X_valid_mode_imp[['bathrooms', 'bedrooms', 'beds']] = X_valid_mode_imp[['bathrooms', 'bedrooms', 'beds']].astype(int)
# Replace the unimputated columns
for feature in zero_features:
X_train[feature] = X_train_zero_imp[feature]
X_valid[feature] = X_valid_zero_imp[feature]
for feature in mean_features:
X_train[feature] = X_train_mean_imp[feature]
X_valid[feature] = X_valid_mean_imp[feature]
for feature in mode_features:
X_train[feature] = X_train_mode_imp[feature]
X_valid[feature] = X_valid_mode_imp[feature]
return X_train, X_valid, y_train, y_valid
def _one_hot_encoding(self, X_train, X_valid, y_train, y_valid):
X_train, X_valid, y_train, y_valid = X_train.copy(), X_valid.copy(), y_train.copy(), y_valid.copy()
oe_enc_features = ['cancellation_policy', 'require_guest_profile_picture', 'require_guest_phone_verification',
'neighbourhood_group_cleansed', 'property_type', 'instant_bookable', 'room_type', 'bed_type']
oe = OrdinalEncoder()
X_train[oe_enc_features] = oe.fit_transform(X_train[oe_enc_features])
X_valid[oe_enc_features] = oe.transform(X_valid[oe_enc_features])
return X_train, X_valid, y_train, y_valid
def _target_encoding(self, X_train, X_valid, y_train, y_valid):
X_train, X_valid, y_train, y_valid = X_train.copy(), X_valid.copy(), y_train.copy(), y_valid.copy()
target_enc_features = ['cancellation_policy', 'require_guest_profile_picture', 'require_guest_phone_verification',
'neighbourhood_group_cleansed', 'property_type', 'instant_bookable', 'room_type', 'bed_type']
# Create the encoder instance. Choose m to control noise.
target_enc = MEstimateEncoder(cols=target_enc_features, m=5.0)
X_train = target_enc.fit_transform(X_train, y_train)
X_valid = target_enc.transform(X_valid)
return X_train, X_valid, y_train, y_valid
def getData(self, kfold, target_encoding=True):
data_frame = self.data_frame.copy()
# Split train and valid
X_train = data_frame[data_frame.kfold != kfold]
X_valid = data_frame[data_frame.kfold == kfold]
y_train = X_train.pop('price')
y_valid = X_valid.pop('price')
# Imputation
X_train, X_valid, y_train, y_valid = self._imputation(X_train, X_valid, y_train, y_valid)
# Target Encoding
if target_encoding:
X_train, X_valid, y_train, y_valid = self._target_encoding(X_train, X_valid, y_train, y_valid)
else:
X_train, X_valid, y_train, y_valid = self._one_hot_encoding(X_train, X_valid, y_train, y_valid)
return X_train, X_valid, y_train, y_valid
# e.g. Before ML pipeline
listings.loc[:2, ['id', 'neighbourhood_group_cleansed', 'property_type', 'amenities', 'price']]
# e.g. After ML pipeline
features = ['host_acceptance_rate', 'neighbourhood_group_cleansed', 'property_type', 'room_type',
'bathrooms', 'bedrooms', 'beds', 'bed_type', 'number_of_reviews', 'review_scores_rating',
'review_scores_accuracy', 'review_scores_cleanliness', 'review_scores_checkin', 'review_scores_communication',
'review_scores_location', 'review_scores_value', 'reviews_per_month', 'host_response_rate', 'host_is_superhost',
'accommodates', 'security_deposit', 'cleaning_fee', 'guests_included', 'extra_people', 'minimum_nights',
'maximum_nights', 'instant_bookable', 'cancellation_policy', 'require_guest_profile_picture',
'require_guest_phone_verification', 'amenities', 'demand', 'kfold']
ml_pipeline = ML_pipeline(data_frame=listings, features=features, target='price')
X_train, X_valid, y_train, y_valid = ml_pipeline.getData(kfold=0, target_encoding=True) # perform target encoding
X = pd.concat([X_train, X_valid], axis=0)
y = pd.concat([y_train, y_valid])
X['price'] = y
X.loc[[241032, 953595, 3308979], ['neighbourhood_group_cleansed', 'property_type', 'price']]
# e.g. After ML pipeline
X.loc[[241032, 953595, 3308979], 'amenity_Elevator in Building':]
```
---
# EDA and Feature Engineering
## Heatmap 1.0
Pandas `.corr()` method calculates the Pearson correlation coefficient between every two features.<br>
And heatmap shows the relationship more clearly.<br>
Here, `accommodates`, `bathrooms`, `bedrooms`, `beds`, `security_deposit`, `cleaning_fee`, `guests_included`, `extra_people` have relatively higher correlation coefficient than other features with `price`.
```
features = listings_metadata[(listings_metadata.ML == 1) | (listings_metadata.ML == 2)].Label.to_list() # Official & Possible ML features
features.append('price') # Add target
plt.figure(dpi=800)
sns.heatmap(listings[features].corr(), cmap="rocket", annot=True, annot_kws={"fontsize": 3});
```
---
## Heatmap 2.0
Since Pandas `.corr()` only calculates the numeric data. I performed target encoding then drew the heatmap again.
`room_type`, `neighbourhood_group_cleansed`, `bed_type`, `cancellation_policy` were categorical data that cannot be calculated the correlation coefficient. But after target encoding, they do have a very high impact on price.
For the amenities, `TV`, `Hot Tub`, `Kitchen`, `Indoor Fireplace`, `Dryer`, `Family/Kid Friendly`, `Doorman`, `Gym`, `Cable TV`, `Washer`, `Air Conditioning` are the more critical than other amenities.
All available amenities:
- 24-Hour Check-in
- Air Conditioning
- Breakfast
- Buzzer/Wireless Intercom
- Cable TV
- Carbon Monoxide Detector
- Cat(s)
- Dog(s)
- Doorman
- Dryer
- Elevator in Building
- Essentials
- Family/Kid Friendly
- Fire Extinguisher
- First Aid Kit
- Free Parking on Premises
- Gym
- Hair Dryer
- Hangers
- Heating
- Hot Tub
- Indoor Fireplace
- Internet
- Iron
- Kitchen
- Laptop Friendly Workspace
- Lock on Bedroom Door
- Other pet(s)
- Pets Allowed
- Pets live on this property
- Pool
- Safety Card
- Shampoo
- Smoke Detector
- Smoking Allowed
- Suitable for Events
- TV
- Washer
- Washer / Dryer
- Wheelchair Accessible
- Wireless Internet
```
def clean_corr(df, target, threshold):
"""Return df.corr() that greater or equal than threshold.
Args:
df (dataframe): Pandas dataframe.
target (str): The name of target.
threshold (float): The miniumu required correlation coefficient.
Returns:
df.corr()
"""
df = df.corr().copy()
for col in df.columns:
if abs(df.loc[col, target]) < threshold:
df.drop(col, axis=0, inplace=True)
df.drop(col, axis=1, inplace=True)
return df
X_corr = clean_corr(X, 'price', 0.1)
plt.figure(dpi=800)
sns.heatmap(X_corr, cmap="rocket", annot=True, annot_kws={"fontsize": 3});
```
---
## Map
For more ideas, visualizations of all Seattle datasets can be found [here](http://insideairbnb.com/seattle/).
> Reference: [Scatter Plots on Mapbox in Python](https://plotly.com/python/scattermapbox/#multiple-markers)
```
#px.set_mapbox_access_token(open(".mapbox_galaxy").read())
px.set_mapbox_access_token('pk.eyJ1IjoiemFja3NhbWJlciIsImEiOiJjazc3MXI1NjQwMXIzM25vMnBtMWtpNWFjIn0.FHxYZnEoStWmap8EQe2l-g')
fig = px.scatter_mapbox(listings,
lat='latitude',
lon='longitude',
color='neighbourhood_group_cleansed',
size='price',
color_continuous_scale=px.colors.cyclical.IceFire,
hover_name='id',
hover_data=['listing_url', 'property_type', 'room_type'],
size_max=15,
zoom=10,
title='Map of price group by neighbourhood_group_cleansed')
fig.show()
```
---
## Histogram
Take a glance at the following high correlation coefficient features.
`accommodates`, `bathrooms`, `bedrooms`, `beds`, `security_deposit`, `cleaning_fee`, `guests_included`, `extra_people`, `room_type`, `neighbourhood_group_cleansed`, `bed_type`, `cancellation_policy`
```
fig = px.histogram(listings.dropna(subset=['property_type'], axis=0),
x='price',
histnorm='percent',
color='property_type',
title='Histogram of price vs. propery_type')
fig.show()
fig = px.histogram(listings.dropna(subset=['room_type'], axis=0),
x='price',
histnorm='percent',
color='room_type',
title='Histogram of price vs. room_type')
fig.show()
fig = px.histogram(listings.dropna(subset=['beds'], axis=0),
x='price',
histnorm='percent',
color='beds',
title='Histogram of price vs. (number of) beds')
fig.show()
```
---
## Parallel Coordinates Plot
Reference: [Parallel Coordinates Plot in Python](https://plotly.com/python/parallel-coordinates-plot/)
```
features = ['accommodates', 'bathrooms', 'bedrooms', 'beds', 'security_deposit', 'cleaning_fee',
'guests_included', 'extra_people', 'room_type', 'neighbourhood_group_cleansed',
'bed_type', 'cancellation_policy']
fig = px.parallel_coordinates(listings[features + ['price']].dropna(), color = "price",
color_continuous_scale = px.colors.diverging.Tealrose, color_continuous_midpoint = 2)
fig.show()
```
---
## Mutual Information
```
def make_mi_scores(X, y):
X = X.copy()
# Mutual Information required all data be integers
for colname in X.select_dtypes(["object", "category"]):
X[colname], _ = X[colname].factorize() # factorize() returns code and uniques
# All discrete features should now have integer dtypes
discrete_features = [pd.api.types.is_integer_dtype(t) for t in X.dtypes]
mi_scores = mutual_info_regression(X, y, discrete_features=discrete_features, random_state=0)
mi_scores = pd.Series(mi_scores, name="MI Scores", index=X.columns)
mi_scores = mi_scores.sort_values(ascending=False)
return mi_scores
%%time
features = ['host_acceptance_rate', 'neighbourhood_group_cleansed', 'property_type', 'room_type',
'bathrooms', 'bedrooms', 'beds', 'bed_type', 'number_of_reviews', 'review_scores_rating',
'review_scores_accuracy', 'review_scores_cleanliness', 'review_scores_checkin', 'review_scores_communication',
'review_scores_location', 'review_scores_value', 'reviews_per_month', 'host_response_rate', 'host_is_superhost',
'accommodates', 'security_deposit', 'cleaning_fee', 'guests_included', 'extra_people', 'minimum_nights',
'maximum_nights', 'instant_bookable', 'cancellation_policy', 'require_guest_profile_picture',
'require_guest_phone_verification', 'amenities', 'demand', 'kfold']
X.drop(columns=['kfold', 'price'], inplace=True)
# Review the MI score from all data
all_mi_scores = pd.DataFrame(make_mi_scores(X, y))
all_mi_scores.style.bar(align='mid', color=['#d65f5f', '#5fba7d'])
```
---
## Outliers and Real Demand
> Reference: [Supply and demand](https://en.wikipedia.org/wiki/Supply_and_demand)
<img src="https://upload.wikimedia.org/wikipedia/commons/thumb/7/7a/Supply-and-demand.svg/1920px-Supply-and-demand.svg.png" width=50%>
In [microeconomics](https://en.wikipedia.org/wiki/Microeconomics), **supply and demand** is an economic model of price determination in a market. The price _P_ of a product is determined by a balance between production at each price (supply _S_) and the desires of those with purchasing power at each price (demand _D_). The diagram shows a positive shift in demand from $D_1$ to $D_2$, resulting in an increase in price (_P_) and quantity sold (_Q_) of the product.
However, the booked listings and unavailable listings are both `available = f` in the dataset **calendar**. As I mentioned above, we don't have the transaction data. Therefore, I designed a simple but effective model to filter the booked listings.
For example:
[listing 3402376](https://www.airbnb.com/rooms/3402376) has 5 reviews in total (1st review was in Sep 2014) but `available = 'f'` for 365 days. As you can see, the listing was not booked by someone for a whole year. Instead, the host set the listing as unavailable for 365 days.
```
calendar
# The calendar recorded the availability of each listing in the next 365 days
calendar.groupby('listing_id').size()
# When the available is False, the price is NaN (Not a Number)
print(calendar.isna().sum())
print(calendar[calendar.available == 1].price.isna().sum())
print(calendar[calendar.available == 0].price.isna().sum())
```
As I mentioned above, we CANNOT know the real demand since we don't have transaction data and search engine data.<br>
However, we can use a statistical method to filter the outliers.
- I extracted the unavailability for each listing in the next 365 days then divided it by 365. So the maximum `unavailability_365` is 1, which means the listing was unavailable for a year or was booked by someone for a year; the minimum `unavailability_365` is 0, which means the listing was ready for booking every day.
- I extracted the number of reviews for each listing started from the first view day and ended with 2016-01-04 (the scraped date). Then I calculated the number of reviews per day for each listing. Therefore, the maximum `reviews_per_day` is 1, which means the listing got reviews every day; the minimum `reviews_per_day` is 0, which means the listing never got any reviews.
You can determine the threshold. But here I used `unavailability_365 > 75% quantile` and `0.010376 < 25% quantile` as the filter to split the dataset into two parts.<br>
As you can see, the relative normal data in the calendar shows the intersections on the plot.<br>
However, the possible outliers have a wide border between `reviews_per_day` and `unavailability_365`.
```
# The possible outliers
eda_demand = EDA_demand()
demand = eda_demand.reviews_rate_vs_unavailability(365)
demand.describe()
eda_demand.plot(outliers=False)
eda_demand.plot(outliers=True)
```
---
# Machine Learning
The models in this project are [XGBoost](https://xgboost.readthedocs.io/en/latest/) and [LightGBM](https://lightgbm.readthedocs.io/en/latest/).<br>
## ML Baseline
The baseline can provide an insight into the performance of different data preprocessing strategies, such as encoding methods.<br>
Here, I chose `target encoding`. First, it had a better performance than ordinal encoding. Second, we already knew the categorical data have potential levels for a different price.<br>
e.g. different `roomt_type` has different `price` histogram.<br>
Features marked as `ML1` were defined by Airbnb for Machine Learning models.<br>
Then, as you can see, `ML1 + ML2` is better than `ML1`, which means we found more features that were useful for Machine Learning.<br>
```
"""
One Hot Encoding
ML1, kfold: 0. RMSE: 56.481995126446456
ML1, kfold: 1. RMSE: 66.83960978199953
ML1, kfold: 2. RMSE: 61.957734603524976
ML1, kfold: 3. RMSE: 62.69133725976135
ML1, kfold: 4. RMSE: 55.715497896362415
ML1. Average RMSE: 60.73723493361895
ML1 + ML2, kfold: 0. RMSE: 52.568454955844246
ML1 + ML2, kfold: 1. RMSE: 63.234791588163155
ML1 + ML2, kfold: 2. RMSE: 58.68112865265134
ML1 + ML2, kfold: 3. RMSE: 60.09474908722824
ML1 + ML2, kfold: 4. RMSE: 47.693034296085685
ML1 + ML2. Average RMSE: 56.45443171599453
Target Encoding
ML1, kfold: 0. RMSE: 56.64589093002433
ML1, kfold: 1. RMSE: 62.44468185143068
ML1, kfold: 2. RMSE: 60.40781093438012
ML1, kfold: 3. RMSE: 63.666798642194124
ML1, kfold: 4. RMSE: 52.226979216000906
ML1. Average RMSE: 59.07843231480604
ML1 + ML2, kfold: 0. RMSE: 52.92355341945994
ML1 + ML2, kfold: 1. RMSE: 65.04777557551235
ML1 + ML2, kfold: 2. RMSE: 58.69704656344895
ML1 + ML2, kfold: 3. RMSE: 55.149794448218394
ML1 + ML2, kfold: 4. RMSE: 49.509631025616585
ML1 + ML2. Average RMSE: 56.26556020645124
"""
def baseline(target_encoding=True):
#import warnings
#warnings.filterwarnings("ignore")
features = ['host_acceptance_rate', 'neighbourhood_group_cleansed', 'property_type', 'room_type',
'bathrooms', 'bedrooms', 'beds', 'bed_type', 'number_of_reviews', 'review_scores_rating',
'review_scores_accuracy', 'review_scores_cleanliness', 'review_scores_checkin', 'review_scores_communication',
'review_scores_location', 'review_scores_value', 'reviews_per_month', 'host_response_rate', 'host_is_superhost',
'accommodates', 'security_deposit', 'cleaning_fee', 'guests_included', 'extra_people', 'minimum_nights',
'maximum_nights', 'instant_bookable', 'cancellation_policy', 'require_guest_profile_picture',
'require_guest_phone_verification', 'amenities', 'demand', 'kfold']
ml_pipeline = ML_pipeline(data_frame=listings, features=features, target='price')
amenities = ['amenity_Washer', 'amenity_Air Conditioning', 'amenity_TV',
'amenity_Kitchen', 'amenity_Wheelchair Accessible',
'amenity_Free Parking on Premises', 'amenity_Doorman',
'amenity_Cable TV', 'amenity_Smoke Detector',
'amenity_Pets live on this property', 'amenity_Internet',
'amenity_Hangers', 'amenity_Family/Kid Friendly',
'amenity_First Aid Kit', 'amenity_Indoor Fireplace', 'amenity_Gym',
'amenity_Suitable for Events', 'amenity_Breakfast', 'amenity_Cat(s)',
'amenity_Lock on Bedroom Door', 'amenity_Smoking Allowed',
'amenity_Dog(s)', 'amenity_Shampoo', 'amenity_Hair Dryer',
'amenity_Carbon Monoxide Detector', 'amenity_Wireless Internet',
'amenity_Hot Tub', 'amenity_Safety Card',
'amenity_Buzzer/Wireless Intercom', 'amenity_Pool',
'amenity_Elevator in Building', 'amenity_Pets Allowed',
'amenity_Fire Extinguisher', 'amenity_Other pet(s)',
'amenity_Laptop Friendly Workspace', 'amenity_Essentials',
'amenity_Iron', 'amenity_Dryer', 'amenity_24-Hour Check-in',
'amenity_Heating']
# Define sheet id and base url
sheet_id = "1M_qah-ym6O8vDcSmoKAP-lbZRPHUey83R_DJaW3LXfs"
base_url = f"https://docs.google.com/spreadsheets/d/{sheet_id}/gviz/tq?tqx=out:csv&sheet="
# Load metadata for three datasets
listings_metadata = pd.read_csv(base_url+"listings")
calendar_metadata = pd.read_csv(base_url+"calendar")
reviews_metadata = pd.read_csv(base_url+"reviews")
# ML1
ml1 = listings_metadata[listings_metadata.ML == 1].Label.to_list()
useless_features = ['availability_30', 'availability_60', 'availability_90', 'availability_365', 'first_review', 'last_review', 'amenities']
for useless_feature in useless_features:
ml1.remove(useless_feature)
ml1.append('demand')
ml1 += amenities
AVG_RMSE = []
for kfold in range(5):
X_train, X_test, y_train, y_test = ml_pipeline.getData(kfold=kfold, target_encoding=target_encoding)
model = XGBRegressor(random_state=kfold, n_jobs=-1)
model.fit(X_train[ml1], y_train)
test_preds = model.predict(X_test[ml1])
RMSE = mean_squared_error(y_test, test_preds, squared=False)
print(f"ML1, kfold: {kfold}. RMSE: {RMSE}")
AVG_RMSE.append(RMSE)
print(f"ML1. Average RMSE: {np.mean(AVG_RMSE)}\n")
# ML1 + ML2
ml1 = listings_metadata[listings_metadata.ML == 1].Label.to_list()
useless_features = ['availability_30', 'availability_60', 'availability_90', 'availability_365', 'first_review', 'last_review', 'amenities']
for useless_feature in useless_features:
ml1.remove(useless_feature)
ml2 = listings_metadata[listings_metadata.ML == 2].Label.to_list()
ml2.append('demand')
ml2 = ml1 + ml2 + amenities
AVG_RMSE = []
for kfold in range(5):
X_train, X_test, y_train, y_test = ml_pipeline.getData(kfold=kfold, target_encoding=target_encoding)
model = XGBRegressor(random_state=kfold, n_jobs=-1)
model.fit(X_train[ml2], y_train)
test_preds = model.predict(X_test[ml2])
RMSE = mean_squared_error(y_test, test_preds, squared=False)
print(f"ML1 + ML2, kfold: {kfold}. RMSE: {RMSE}")
AVG_RMSE.append(RMSE)
print(f"ML1 + ML2. Average RMSE: {np.mean(AVG_RMSE)}\n")
baseline(target_encoding=True)
```
---
## Model Tuning
The Hyperparameter tuning platform I used is [Optuna](https://optuna.org/).<br>
I implemented a logger to write the tuning results in the local log file.<br>
After all tunings are finished, the program will sent an email to my mailbox with the best hyperparameters.
- **To enable this feature**, go to [configure your gmail first](#gmail-configuration).
P.S: If your computer does not support GPU accleration, uncomment code `For CPU` and comment code `For GPU`.
**If you want to train your model, DO NOT RUN the following code in this notebook.**<br>
Instead, make another notebook for model tuning. Please following this [link](https://github.com/ZacksAmber/Kaggle-Seattle-Airbnb/blob/main/ML_Tuning.ipynb).
---
### Define Logger
```
import logging
# Define logger
logger = logging.getLogger('ML')
# Set level for logger
logger.setLevel(logging.DEBUG)
# Define the handler and formatter for file logging
log_file = 'ML'
fileHandler = logging.FileHandler(f'{log_file}.log') # Define FileHandler
fileHandler.setLevel(logging.INFO) # Set level
fileFormatter = logging.Formatter('%(asctime)s - %(name)s - %(levelname)s - %(message)s') # Define formatter
fileHandler.setFormatter(fileFormatter) # Set formatter
logger.addHandler(fileHandler) # Add handler to logger
```
---
### Define Features for ML
```
# Define sheet id and base url
sheet_id = "1M_qah-ym6O8vDcSmoKAP-lbZRPHUey83R_DJaW3LXfs"
base_url = f"https://docs.google.com/spreadsheets/d/{sheet_id}/gviz/tq?tqx=out:csv&sheet="
# Load metadata for three datasets
listings_metadata = pd.read_csv(base_url+"listings")
calendar_metadata = pd.read_csv(base_url+"calendar")
reviews_metadata = pd.read_csv(base_url+"reviews")
amenities = ['amenity_Washer', 'amenity_Air Conditioning', 'amenity_TV',
'amenity_Kitchen', 'amenity_Wheelchair Accessible',
'amenity_Free Parking on Premises', 'amenity_Doorman',
'amenity_Cable TV', 'amenity_Smoke Detector',
'amenity_Pets live on this property', 'amenity_Internet',
'amenity_Hangers', 'amenity_Family/Kid Friendly',
'amenity_First Aid Kit', 'amenity_Indoor Fireplace', 'amenity_Gym',
'amenity_Suitable for Events', 'amenity_Breakfast', 'amenity_Cat(s)',
'amenity_Lock on Bedroom Door', 'amenity_Smoking Allowed',
'amenity_Dog(s)', 'amenity_Shampoo', 'amenity_Hair Dryer',
'amenity_Carbon Monoxide Detector', 'amenity_Wireless Internet',
'amenity_Hot Tub', 'amenity_Safety Card',
'amenity_Buzzer/Wireless Intercom', 'amenity_Pool',
'amenity_Elevator in Building', 'amenity_Pets Allowed',
'amenity_Fire Extinguisher', 'amenity_Other pet(s)',
'amenity_Laptop Friendly Workspace', 'amenity_Essentials',
'amenity_Iron', 'amenity_Dryer', 'amenity_24-Hour Check-in',
'amenity_Heating']
# ML1 + ML2
ml1 = listings_metadata[listings_metadata.ML == 1].Label.to_list()
useless_features = ['availability_30', 'availability_60', 'availability_90', 'availability_365', 'first_review', 'last_review', 'amenities']
for useless_feature in useless_features:
ml1.remove(useless_feature)
ml2 = listings_metadata[listings_metadata.ML == 2].Label.to_list()
ml2.append('demand')
ml2 = ml1 + ml2 + amenities
```
---
### Tuning Configurations
```
# Silence Optuna
optuna.logging.set_verbosity(optuna.logging.WARNING)
# Define number of trails
n_trials = 200
```
---
### Model Tuning: XGBoost
```
def objective(trial):
"""Modeling tuning with Target encoding.
"""
features = ['host_acceptance_rate', 'neighbourhood_group_cleansed', 'property_type', 'room_type',
'bathrooms', 'bedrooms', 'beds', 'bed_type', 'number_of_reviews', 'review_scores_rating',
'review_scores_accuracy', 'review_scores_cleanliness', 'review_scores_checkin', 'review_scores_communication',
'review_scores_location', 'review_scores_value', 'reviews_per_month', 'host_response_rate', 'host_is_superhost',
'accommodates', 'security_deposit', 'cleaning_fee', 'guests_included', 'extra_people', 'minimum_nights',
'maximum_nights', 'instant_bookable', 'cancellation_policy', 'require_guest_profile_picture',
'require_guest_phone_verification', 'amenities', 'demand', 'kfold']
ml_pipeline = ML_pipeline(data_frame=listings, features=features, target='price')
RMSE_AVG = []
for kfold in range(5):
X_train, X_valid, y_train, y_valid = ml_pipeline.getData(kfold=kfold, target_encoding=True)
X_train, X_valid = X_train[ml2], X_valid[ml2]
# Hyperparameters for XGBoost
xgb_params = {
'lambda': trial.suggest_loguniform('lambda', 1e-3, 10.0),
'alpha': trial.suggest_loguniform('alpha', 1e-3, 10.0),
'reg_lambda': trial.suggest_loguniform("reg_lambda", 1e-8, 100.0),
'reg_alpha': trial.suggest_loguniform("reg_alpha", 1e-8, 100.0),
'colsample_bytree': trial.suggest_float("colsample_bytree", 0.1, 1.0),
'subsample': trial.suggest_float("subsample", 0.1, 1.0),
'learning_rate': trial.suggest_float("learning_rate", 1e-2, 0.3, log=True),
'n_estimators': trial.suggest_int('n_estimators', 100, 10000),
'max_depth': trial.suggest_int("max_depth", 1, 7),
'random_state': trial.suggest_categorical('random_state', [0, 42, 2021]),
'min_child_weight': trial.suggest_int('min_child_weight', 1, 300)
}
# For GPU
model = XGBRegressor(
tree_method='gpu_hist',
gpu_id=0,
predictor='gpu_predictor',
**xgb_params)
'''
# For CPU
model = XGBRegressor(**xgb_params)
'''
model.fit(
X_train, y_train,
early_stopping_rounds=300,
eval_set=[(X_valid, y_valid)],
verbose=5000
)
valid_preds = model.predict(X_valid)
RMSE = mean_squared_error(y_valid, valid_preds, squared=False)
RMSE_AVG.append(RMSE)
return np.mean(RMSE_AVG)
'''
%%time
study = optuna.create_study(direction='minimize', study_name=f'XGBoost {n_trials} trails')
study.optimize(objective, n_trials=n_trials, show_progress_bar=False) # set n_triasl
logger.info(f"Study name: {study.study_name}")
logger.info(f"Best value: {study.best_value}")
logger.info(f"Best paras: {study.best_params}")
logger.info("Mission Complete! --------------")
'''
```
---
### Model Tuning: LightGBM
```
def objective(trial):
"""Modeling tuning with Target encoding.
"""
features = ['host_acceptance_rate', 'neighbourhood_group_cleansed', 'property_type', 'room_type',
'bathrooms', 'bedrooms', 'beds', 'bed_type', 'number_of_reviews', 'review_scores_rating',
'review_scores_accuracy', 'review_scores_cleanliness', 'review_scores_checkin', 'review_scores_communication',
'review_scores_location', 'review_scores_value', 'reviews_per_month', 'host_response_rate', 'host_is_superhost',
'accommodates', 'security_deposit', 'cleaning_fee', 'guests_included', 'extra_people', 'minimum_nights',
'maximum_nights', 'instant_bookable', 'cancellation_policy', 'require_guest_profile_picture',
'require_guest_phone_verification', 'amenities', 'demand', 'kfold']
ml_pipeline = ML_pipeline(data_frame=listings, features=features, target='price')
RMSE_AVG = []
for kfold in range(5):
X_train, X_valid, y_train, y_valid = ml_pipeline.getData(kfold=kfold, target_encoding=True)
X_train, X_valid = X_train[ml2], X_valid[ml2]
# Hyperparameters for LightGBM
lgb_params = {
'random_state': trial.suggest_categorical('random_state', [0, 42, 2021]),
'num_iterations': trial.suggest_int('num_iterations', 100, 10000),
'learning_rate': trial.suggest_float("learning_rate", 1e-2, 0.3, log=True),
'max_depth': trial.suggest_int('max_depth', 1, 7),
'num_leaves': trial.suggest_int('num_leaves', 2, 100),
'min_data_in_leaf': trial.suggest_int('min_data_in_leaf', 100, 2000),
'lambda_l1': trial.suggest_loguniform('lambda_l1', 1e-8, 10.0),
'lambda_l2': trial.suggest_loguniform('lambda_l2', 1e-8, 10.0),
'feature_fraction': trial.suggest_uniform('feature_fraction', 0.01, 0.99),
'bagging_fraction': trial.suggest_uniform('bagging_fraction', 0.01, 0.99),
'bagging_freq': trial.suggest_int('bagging_freq', 1, 7),
'min_child_samples': trial.suggest_int('min_child_samples', 5, 100),
}
# For GPU
model = LGBMRegressor(
device='gpu',
gpu_platform_id=0,
gpu_device_id=0,
n_jobs=-1,
metric='rmse',
**lgb_params
)
'''
# For CPU
model = LGBMRegressor(**lgb_params)
'''
model.fit(
X_train, y_train,
early_stopping_rounds=300,
eval_set=[(X_valid, y_valid)],
verbose=5000
)
valid_preds = model.predict(X_valid)
RMSE = mean_squared_error(y_valid, valid_preds, squared=False)
RMSE_AVG.append(RMSE)
return np.mean(RMSE_AVG)
'''
%%time
study = optuna.create_study(direction='minimize', study_name=f'LGBoost {n_trials} trails')
study.optimize(objective, n_trials=n_trials, show_progress_bar=False) # set n_triasl
logger.info(f"Study name: {study.study_name}")
logger.info(f"Best value: {study.best_value}")
logger.info(f"Best paras: {study.best_params}")
logger.info("Mission Complete! --------------")
'''
```
---
### Gmail Configuration<a id='gmail-configuration'></a>
> [How to Send Emails with Gmail using Python](https://stackabuse.com/how-to-send-emails-with-gmail-using-python/)
```
def gmail(YOUR_GMAIL, YOUR_APP_PASSWORD, SEND_TO):
"""Send the ML tuning result to one or more email addresses.
Args:
YOUR_GMAIL (str): Your gmail address.
YOUR_APP_PASSWORD (str): Your APP Password for gmail.
SEND_TO (str or list): The target emails.
"""
gmail_user = YOUR_GMAIL
gmail_password = YOUR_APP_PASSWORD # Google App Password
import smtplib
from email.message import EmailMessage
msg = EmailMessage()
msg["From"] = YOUR_GMAIL
msg["Subject"] = "Seattle Airbnb ML Tuning"
msg["To"] = SEND_TO
msg.set_content(f"""\
{n_trials} Trials are done.
Mission Complete!""")
with open('ML.log', 'rb') as f:
content = f.read()
msg.add_attachment(content, maintype='application', subtype='log', filename='ML.log')
server = smtplib.SMTP_SSL('smtp.gmail.com', 465)
server.login(gmail_user, gmail_password)
server.send_message(msg)
server.close()
#gmail(YOUR_GMAIL, YOUR_APP_PASSWORD, SEND_TO)
```
---
## Model Blending
> Reference: [Ensemble Learning: Stacking, Blending & Voting](https://towardsdatascience.com/ensemble-learning-stacking-blending-voting-b37737c4f483)
After hyperparameter tuning, we have a set of beter hyperparameters for XGBoost and LightGBM. Then I performed a model blending for a better ML performance.
My 200 Trails ($200 \times 5$ in total) hyperparameters:
- XGBoost
{'lambda': 0.029949323233957558, 'alpha': 0.47821306780284645, 'reg_lambda': 0.03007272817610808, 'reg_alpha': 5.7650942972599255e-05, 'colsample_bytree': 0.32733907049678806, 'subsample': 0.9397958925107069, 'learning_rate': 0.016087339011505105, 'n_estimators': 4117, 'max_depth': 6, 'random_state': 42, 'min_child_weight': 5}
- LightGBM
{'random_state': 42, 'num_iterations': 5549, 'learning_rate': 0.07313607774375752, 'max_depth': 5, 'num_leaves': 75, 'min_data_in_leaf': 100, 'lambda_l1': 1.3379869858112054e-06, 'lambda_l2': 0.00025091437242776726, 'feature_fraction': 0.5910800704597817, 'bagging_fraction': 0.9553891294481797, 'bagging_freq': 6, 'min_child_samples': 23}
```
class Model_Blending:
def __init__(self, data_frame, features_etl, features_ml):
data_frame = data_frame.copy()
self.ml_pipeline = ML_pipeline(data_frame=data_frame, features=features_etl, target='price')
self.features_ml = features_ml
def _xgboost_reg(self, xgb_params):
"""
# For GPU
model = XGBRegressor(
tree_method='gpu_hist',
gpu_id=0,
predictor='gpu_predictor',
n_jobs=-1,
**xgb_params
)
"""
# For CPU
model = XGBRegressor(**xgb_params)
return model
def _lightgbm_reg(self, lgb_params):
"""
# For GPU
model = LGBMRegressor(
device='gpu',
gpu_platform_id=0,
gpu_device_id=0,
n_jobs=-1,
metric='rmse',
**lgb_params
)
"""
# For CPUT
model = LGBMRegressor(**lgb_params)
return model
def blending(self, model: str, params: dict):
'''Model blending. Generate 5 predictions according to 5 folds.
Args:
model: One of xgboost or lightgbm.
params: Hyperparameters for XGBoost or LightGBM.
Returns:
None
'''
assert model in ['xgboost', 'lightgbm'], "ValueError: model must be one of ['xgboost', 'lightgbm']!"
final_valid_predictions = {}
scores = []
for fold in range(5):
X_train, X_valid, y_train, y_valid = self.ml_pipeline.getData(kfold=fold, target_encoding=True)
X_train, X_valid = X_train[self.features_ml], X_valid[self.features_ml] # Add many amenities
# Get X_valid_ids
X_valid_ids = list(X_valid.index)
print(f"Training ...")
# Define model
if model == 'xgboost':
reg = self._xgboost_reg(params)
elif model == 'lightgbm':
reg = self._lightgbm_reg(params)
# Modeling - Training
reg.fit(
X_train, y_train,
early_stopping_rounds=300,
eval_set=[(X_valid, y_valid)],
verbose=False
)
# Modeling - Inference
valid_preds = reg.predict(X_valid)
final_valid_predictions.update(dict(zip(X_valid_ids, valid_preds))) # loop 5 times with different valid id
rmse = mean_squared_error(y_valid, valid_preds, squared=False)
scores.append(rmse)
print(f'Fold: {fold}, RMSE: {rmse}')
# Export results
if not os.path.exists('output'):
os.mkdir('output')
final_valid_predictions = pd.DataFrame.from_dict(final_valid_predictions, orient="index").reset_index()
final_valid_predictions.columns = ["id", f"{model}_pred"]
final_valid_predictions.to_csv(f"output/{model}_valid_pred.csv", index=False)
print('-----------------------------------------------------------------')
print(f'Average RMSE: {np.mean(scores)}, STD of RMSE: {np.std(scores)}')
def predict(self, models: list):
df_valids = pd.read_csv(f'output/{models[0]}_valid_pred.csv')
models.remove(models[0])
for model in models:
df = pd.read_csv(f'output/{model}_valid_pred.csv')
df_valids = df_valids.set_index('id').join(df.set_index('id'), how='inner')
# Calculate the average predictions
df_valids['mean_valids'] = df_valids.mean(axis=1)
# Join listings price to df_valids
df_valids['price'] = listings.set_index('id')['price']
# Use the average predictions to validate the target
return mean_squared_error(df_valids.price, df_valids['mean_valids'], squared=False)
features_etl = ['host_acceptance_rate', 'neighbourhood_group_cleansed', 'property_type', 'room_type',
'bathrooms', 'bedrooms', 'beds', 'bed_type', 'number_of_reviews', 'review_scores_rating',
'review_scores_accuracy', 'review_scores_cleanliness', 'review_scores_checkin', 'review_scores_communication',
'review_scores_location', 'review_scores_value', 'reviews_per_month', 'host_response_rate', 'host_is_superhost',
'accommodates', 'security_deposit', 'cleaning_fee', 'guests_included', 'extra_people', 'minimum_nights',
'maximum_nights', 'instant_bookable', 'cancellation_policy', 'require_guest_profile_picture',
'require_guest_phone_verification', 'amenities', 'demand', 'kfold']
features_ml = ml2
xgb_params = {'lambda': 0.029949323233957558, 'alpha': 0.47821306780284645, 'reg_lambda': 0.03007272817610808,
'reg_alpha': 5.7650942972599255e-05, 'colsample_bytree': 0.32733907049678806, 'subsample': 0.9397958925107069,
'learning_rate': 0.016087339011505105, 'n_estimators': 4117, 'max_depth': 6, 'random_state': 42, 'min_child_weight': 5}
lgb_params = {'random_state': 42, 'num_iterations': 5549, 'learning_rate': 0.07313607774375752, 'max_depth': 5, 'num_leaves': 75,
'min_data_in_leaf': 100, 'lambda_l1': 1.3379869858112054e-06, 'lambda_l2': 0.00025091437242776726,
'feature_fraction': 0.5910800704597817, 'bagging_fraction': 0.9553891294481797, 'bagging_freq': 6, 'min_child_samples': 23}
model_blending = Model_Blending(listings, features_etl, features_ml)
model_blending.blending(model='xgboost', params=xgb_params)
model_blending.blending(model='lightgbm', params=lgb_params)
model_blending.predict(models=['xgboost', 'lightgbm'])
```
The model blending result is not much better than any single model since I only used two models with target encoding.<br>
You can combine different models with different encoding strategies even different features to improve the overall performance.<br>
For instance, you can combine XGBoost, LightGBM, and CatBoost with one-hot encoding, target encoding, ordinal encoding, and polynomial encoding. Then you have $3 \times 4$ models for model blending.
---
## Model Stacking
> Reference: [Ensemble Learning: Stacking, Blending & Voting](https://towardsdatascience.com/ensemble-learning-stacking-blending-voting-b37737c4f483)<br>
> Reference: [How To Use “Model Stacking” To Improve Machine Learning Predictions](https://medium.com/geekculture/how-to-use-model-stacking-to-improve-machine-learning-predictions-d113278612d4)
Model Stacking is a way to improve model predictions by combining the outputs of multiple models and running them through another machine learning model called a meta-learner.
After hyperparameter tuning, we have a set of beter hyperparameters for XGBoost and LightGBM. Then I performed a model blending for a better ML performance.
My 200 Trails ($200 \times 5$ in total) hyperparameters:
- XGBoost
{'lambda': 0.029949323233957558, 'alpha': 0.47821306780284645, 'reg_lambda': 0.03007272817610808, 'reg_alpha': 5.7650942972599255e-05, 'colsample_bytree': 0.32733907049678806, 'subsample': 0.9397958925107069, 'learning_rate': 0.016087339011505105, 'n_estimators': 4117, 'max_depth': 6, 'random_state': 42, 'min_child_weight': 5}
- LightGBM
{'random_state': 42, 'num_iterations': 5549, 'learning_rate': 0.07313607774375752, 'max_depth': 5, 'num_leaves': 75, 'min_data_in_leaf': 100, 'lambda_l1': 1.3379869858112054e-06, 'lambda_l2': 0.00025091437242776726, 'feature_fraction': 0.5910800704597817, 'bagging_fraction': 0.9553891294481797, 'bagging_freq': 6, 'min_child_samples': 23}
```
class Model_Stacking:
def __init__(self, data_frame, features_etl, features_ml):
data_frame = data_frame.copy()
self.ml_pipeline = ML_pipeline(data_frame=data_frame, features=features_etl, target='price')
self.features_ml = features_ml
def _xgboost_reg(self, xgb_params):
"""
# For GPU
model = XGBRegressor(
tree_method='gpu_hist',
gpu_id=0,
predictor='gpu_predictor',
n_jobs=-1,
**xgb_params
)
"""
# For CPU
model = XGBRegressor(**xgb_params)
return model
def _lightgbm_reg(self, lgb_params):
"""
# For GPU
model = LGBMRegressor(
device='gpu',
gpu_platform_id=0,
gpu_device_id=0,
n_jobs=-1,
metric='rmse',
**lgb_params
)
"""
# For CPUT
model = LGBMRegressor(**lgb_params)
return model
def stacking(self, model: str, params: dict):
'''Model blending. Generate 5 predictions according to 5 folds.
Args:
model: One of xgboost or lightgbm.
params: Hyperparameters for XGBoost or LightGBM.
Returns:
None
'''
assert model in ['xgboost', 'lightgbm'], "ValueError: model must be one of ['xgboost', 'lightgbm']!"
final_valid_predictions = {}
scores = []
for fold in range(5):
X_train, X_valid, y_train, y_valid = self.ml_pipeline.getData(kfold=fold, target_encoding=True)
X_train, X_valid = X_train[self.features_ml], X_valid[self.features_ml] # Add many amenities
# Get X_valid_ids
X_valid_ids = list(X_valid.index)
print(f"Training ...")
# Define model
if model == 'xgboost':
reg = self._xgboost_reg(params)
elif model == 'lightgbm':
reg = self._lightgbm_reg(params)
# Modeling - Training
reg.fit(
X_train, y_train,
early_stopping_rounds=300,
eval_set=[(X_valid, y_valid)],
verbose=False
)
# Modeling - Inference
valid_preds = reg.predict(X_valid)
final_valid_predictions.update(dict(zip(X_valid_ids, valid_preds))) # loop 5 times with different valid id
rmse = mean_squared_error(y_valid, valid_preds, squared=False)
scores.append(rmse)
print(f'Fold: {fold}, RMSE: {rmse}')
# Export results
if not os.path.exists('output'):
os.mkdir('output')
final_valid_predictions = pd.DataFrame.from_dict(final_valid_predictions, orient="index").reset_index()
final_valid_predictions.columns = ["id", f"{model}_pred"]
final_valid_predictions.to_csv(f"output/{model}_valid_pred.csv", index=False)
print('-----------------------------------------------------------------')
print(f'Average RMSE: {np.mean(scores)}, STD of RMSE: {np.std(scores)}')
def predict(self, models: list):
df_valids = pd.read_csv(f'output/{models[0]}_valid_pred.csv')
models.remove(models[0])
for model in models:
df = pd.read_csv(f'output/{model}_valid_pred.csv')
df_valids = df_valids.set_index('id').join(df.set_index('id'), how='inner')
# Join listings price to df_valids
df_valids['price'] = listings.set_index('id')['price']
# Implement a simple regressor such as linear regression
linear_reg = LinearRegression()
# Define X, y
X, y = df_valids.iloc[:, :len(models)], df_valids.price
# Use the models validations as training set for predictions
scores = cross_val_score(linear_reg, X, y, cv=5, scoring='neg_root_mean_squared_error')
scores = -scores
return np.mean(scores)
features_etl = ['host_acceptance_rate', 'neighbourhood_group_cleansed', 'property_type', 'room_type',
'bathrooms', 'bedrooms', 'beds', 'bed_type', 'number_of_reviews', 'review_scores_rating',
'review_scores_accuracy', 'review_scores_cleanliness', 'review_scores_checkin', 'review_scores_communication',
'review_scores_location', 'review_scores_value', 'reviews_per_month', 'host_response_rate', 'host_is_superhost',
'accommodates', 'security_deposit', 'cleaning_fee', 'guests_included', 'extra_people', 'minimum_nights',
'maximum_nights', 'instant_bookable', 'cancellation_policy', 'require_guest_profile_picture',
'require_guest_phone_verification', 'amenities', 'demand', 'kfold']
features_ml = ml2
xgb_params = {'lambda': 0.029949323233957558, 'alpha': 0.47821306780284645, 'reg_lambda': 0.03007272817610808,
'reg_alpha': 5.7650942972599255e-05, 'colsample_bytree': 0.32733907049678806, 'subsample': 0.9397958925107069,
'learning_rate': 0.016087339011505105, 'n_estimators': 4117, 'max_depth': 6, 'random_state': 42, 'min_child_weight': 5}
lgb_params = {'random_state': 42, 'num_iterations': 5549, 'learning_rate': 0.07313607774375752, 'max_depth': 5, 'num_leaves': 75,
'min_data_in_leaf': 100, 'lambda_l1': 1.3379869858112054e-06, 'lambda_l2': 0.00025091437242776726,
'feature_fraction': 0.5910800704597817, 'bagging_fraction': 0.9553891294481797, 'bagging_freq': 6, 'min_child_samples': 23}
model_stacking = Model_Stacking(listings, features_etl, features_ml)
model_stacking.stacking('xgboost', xgb_params)
model_stacking.stacking('lightgbm', lgb_params)
model_stacking.predict(models=['xgboost', 'lightgbm'])
```
The model stacking result is not much better than any single model since I only used two models with target encoding.<br>
You can combine different models with different encoding strategies even different features to improve the overall performance.<br>
For instance, you can combine XGBoost, LightGBM, and CatBoost with one-hot encoding, target encoding, ordinal encoding, and polynomial encoding. Then you have $3 \times 4$ models for model stacking.
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
%aimport utils_1_1
import pandas as pd
import numpy as np
import altair as alt
from altair_saver import save
import datetime
import dateutil.parser
from os.path import join
from constants_1_1 import SITE_FILE_TYPES
from utils_1_1 import (
read_loinc_df,
get_site_file_paths,
get_site_file_info,
get_site_ids,
read_full_daily_counts_df,
get_visualization_subtitle,
get_country_color_map,
apply_theme,
merge_single_site_country_adult_name,
)
from web import for_website
alt.data_transformers.disable_max_rows(); # Allow using rows more than 5000
```
## The 4CE Health Systems Participating spreadsheet must be downloaded from Google Sheets and moved to
```
data/Health_Systems_Participating.csv
```
because we need the number of hospitals, beds, and inpatient discharges per year in order to compute the country-level rates of change.
```
isHospitalizationData = False # Use hospitalization data?
min_date = datetime.datetime(2020, 1, 27) + datetime.timedelta(hours=1)
max_date = datetime.datetime(2020, 9, 29) + datetime.timedelta(hours=1)
DATA_RELEASE = "2020-09-28"
COHORT = "Adult"
MERGE_SINGLE_SITE_COUNTRIES = False
CATEGORY = "category"
CATEGORY_OF_INTEREST = "new_positive_cases"
COUNTRY_POPULATION = {
# From https://data.worldbank.org/indicator/SP.POP.TOTL
"France": 67059887,
"USA": 328239523,
"Germany": 83132799,
"Italy": 60297396,
"Singapore": 5703569,
"Spain": 47076781,
"UK": 66834405,
}
COUNTRY_HOSP_DISCHARGE = {
# From https://data.oecd.org/healthcare/hospital-discharge-rates.htm
"France": 18553.0,
"USA": 10906.2, # https://hcup-us.ahrq.gov/faststats/NationalTrendsServlet
"Germany": 25478.4,
"Italy": 11414.6,
"Singapore": 12700.4, # https://www.moh.gov.sg/resources-statistics/healthcare-institution-statistics/hospital-admission-rates-by-age-and-sex/hospital-admission-rates-by-age-and-sex-2017
"Spain": 10470.5,
"UK": 12869.4,
}
# Un-comment if you want to merge Spain and Singapore
#COUNTRY_POPULATION["Spain + Singapore"] = COUNTRY_POPULATION["Spain"] + COUNTRY_POPULATION["Singapore"]
#COUNTRY_HOSP_DISCHARGE["Spain + Singapore"] = COUNTRY_HOSP_DISCHARGE["Spain"] + COUNTRY_HOSP_DISCHARGE["Singapore"]
df = read_full_daily_counts_df()
df.head()
```
## Remove pediatric sites
```
df = df.loc[df["pediatric"] == False]
df = df.drop(columns=["pediatric"])
df.head()
# Remove RP401 non-pediatric data since RP401 is only listed as a pediatric site
df = df.loc[~df["siteid"].isin(["RP401"])]
df = df.replace(-99, np.nan)
df = df.replace(-999, np.nan)
df["num_sites"] = 1
# We only need the JHU data for the countries that exist in the 4CE data.
COUNTRIES = df["country"].unique().tolist()
COUNTRIES
df["date"] = df["calendar_date"]
df = df.drop(columns=["calendar_date"])
```
## Load participating sites metadata
```
sites_df = pd.read_csv(join("..", "data", "Health_Systems_Participating.tsv"), sep='\t', skiprows=2, header=None, thousands=',')
sites_column_map = {
0: "site_name",
1: "siteid",
2: "city",
3: "country",
4: "patient_type",
6: "adult_num_hosp",
7: "adult_num_beds",
8: "adult_num_yearly_discharge",
10: "ped_num_hosp",
11: "ped_num_beds",
12: "ped_num_yearly_discharge",
}
sites_df = sites_df.rename(columns=sites_column_map)
sites_df = sites_df[list(sites_column_map.values())]
sites_df["pediatric"] = sites_df["patient_type"].apply(lambda t: t == "Pediatric")
sites_df = sites_df.dropna(subset=["site_name"])
sites_df["siteid"] = sites_df["siteid"].apply(lambda x: x.upper())
sites_df.tail()
# Drop the pediatric hospitals
sites_df = sites_df.loc[sites_df["pediatric"] == False]
```
## Take intersection of sites that have provided valid num_yearly_discharge counts and sites that have provided daily counts data
```
sites_df["adult_num_hosp"] = sites_df["adult_num_hosp"].apply(lambda x: str(x).replace(",", "")).astype(float)
sites_df["adult_num_beds"] = sites_df["adult_num_beds"].apply(lambda x: str(x).replace(",", "")).astype(float)
sites_df["adult_num_yearly_discharge"] = sites_df["adult_num_yearly_discharge"].apply(lambda x: str(x).replace(",", "")).astype(float)
sites_df = sites_df.set_index("siteid")
def get_num_hosp(sid):
try:
return sites_df.at[sid, "adult_num_hosp"] if pd.notna(sites_df.at[sid, "adult_num_hosp"]) else 1
except KeyError:
return 1
df["num_hosps"] = df["siteid"].apply(get_num_hosp)
sites_df.head()
sites_df = sites_df.reset_index()
sites_df = sites_df.dropna(subset=["adult_num_yearly_discharge"])
sites_in_sites_df = sites_df["siteid"].unique().tolist()
sites_in_df = df["siteid"].unique().tolist()
intersecting_sites = set(sites_in_sites_df).intersection(set(sites_in_df))
sites_df = sites_df.loc[sites_df["siteid"].isin(intersecting_sites)]
df = df.loc[df["siteid"].isin(intersecting_sites)]
intersecting_sites
# Get number of sites after restricting to pediatrics
# and after taking the intersection
NUM_SITES = len(df["siteid"].unique().tolist())
sites_in_sites_df
```
## If site is missing data for a particular date, use the most recent previous data point for that date
```
def convert_date(date_str):
try:
return dateutil.parser.parse(date_str)
except:
return np.nan
max_date_str = str(max_date).split(" ")[0]
all_date_country_df = pd.DataFrame()
for siteid, cd_df in df.groupby(["siteid"]):
min_date = cd_df["date"].min()
min_date_str = str(min_date).split(" ")[0]
num_days = (dateutil.parser.parse(max_date_str) - dateutil.parser.parse(min_date_str)).days
cd_df = cd_df.copy()
cd_df["date"] = cd_df["date"].astype(str)
prev_date_row = None
for day_offset in range(num_days):
curr_date = dateutil.parser.parse(min_date_str) + datetime.timedelta(days=day_offset)
curr_date_str = str(curr_date).split(" ")[0]
try:
curr_date_row = cd_df.loc[cd_df["date"] == curr_date_str].to_dict('records')[0]
prev_date_row = curr_date_row
except:
prev_date_row['date'] = curr_date_str
prev_date_row['num_sites'] = 0
prev_date_row['num_hosps'] = 0
cd_df = cd_df.append(prev_date_row, ignore_index=True)
all_date_country_df = all_date_country_df.append(cd_df, ignore_index=True)
all_date_country_df["date"] = all_date_country_df["date"].apply(convert_date)
df = all_date_country_df
```
## Subtract severe patients from all patients to get the "never severe"-like count
```
country_color_map = get_country_color_map(merge_single_site_countries=MERGE_SINGLE_SITE_COUNTRIES)
if MERGE_SINGLE_SITE_COUNTRIES:
df["country"] = df["country"].apply(merge_single_site_country_adult_name)
country_sum_df = df.groupby(["country", "date"]).sum().reset_index()
country_sum_df.head()
COUNTRIES = country_sum_df["country"].unique().tolist()
country_sum_df.tail()
country_sum_df["num_patients_in_hospital_on_this_date_minus_severe"] = df["num_patients_in_hospital_on_this_date"] - df["num_patients_in_hospital_and_severe_on_this_date"]
country_sum_df["cumulative_patients_all_minus_severe"] = df["cumulative_patients_all"] - df["cumulative_patients_severe"]
country_sum_temp_df = pd.DataFrame(index=[], data=[], columns=[])
for country, country_df in country_sum_df.groupby("country"):
country_df = country_df.copy()
country_df["cum_diff_all"] = np.concatenate((np.array([np.nan]), np.diff(country_df["cumulative_patients_all"].values)))
country_df["cum_diff_severe"] = np.concatenate((np.array([np.nan]), np.diff(country_df["cumulative_patients_severe"].values)))
country_df["cum_diff_dead"] = np.concatenate((np.array([np.nan]), np.diff(country_df["cumulative_patients_dead"].values)))
country_df["cum_diff_all_minus_severe"] = country_df["cum_diff_all"] - country_df["cum_diff_severe"]
country_df["cum_diff_all"] = country_df["cum_diff_all"].clip(lower=0)
country_df["cum_diff_severe"] = country_df["cum_diff_severe"].clip(lower=0)
country_df["cum_diff_dead"] = country_df["cum_diff_dead"].clip(lower=0)
country_df["cum_diff_all_minus_severe"] = country_df["cum_diff_all_minus_severe"].clip(lower=0)
"""
country_df["count"] = np.concatenate((np.array([np.nan]), np.diff(country_df["cumulative_count"].values)))
country_df["cumulative_count"] = country_df["cumulative_count"].replace(0, np.nan)
country_df["N0"] = country_df["cumulative_count"].shift(1) # N0 is the total case up to the day before
country_df["n1"] = country_df["count"] # n1 is the case number this day
country_df["n2"] = country_df["n1"].shift(1) # n2 is the case number yesterday
country_df["percent_increase"] = (country_df["n1"] / country_df["N0"]) * 100
country_df['R'] = country_df["percent_increase"] # TODO: is this correct?
# TODO: update CI formula
country_df['C'] = country_df['R'] - 1
country_df['standard_error'] = country_df.apply(lambda obs: (obs['R']+np.power(obs['R'], 2))/obs['n2'], axis='columns')
country_df['95_CI_below'] = country_df.apply(lambda obs: obs['C'] - 1.96*np.sqrt(obs['standard_error']), axis='columns')
country_df['95_CI_above'] = country_df.apply(lambda obs: obs['C'] + 1.96*np.sqrt(obs['standard_error']), axis='columns')
country_df = country_df.replace([np.inf, -np.inf], np.nan)
"""
country_sum_temp_df = country_sum_temp_df.append(country_df, ignore_index=True)
country_sum_df = country_sum_temp_df
country_sum_df.tail()
```
## Obtain country-level daily counts from JHU CSSE
```
jhu_url = "https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_confirmed_global.csv"
jhu_df = pd.read_csv(jhu_url)
jhu_df = jhu_df.rename(columns={"Country/Region": "country", "Province/State": "state"})
jhu_df = jhu_df.drop(columns=["Lat", "Long"])
# Countries have different ids in the JHU data than in the 4CE data
country_map = {
"US": "USA",
"United Kingdom": "UK"
}
jhu_df["country"] = jhu_df["country"].apply(lambda c: country_map[c] if c in country_map else c)
jhu_df = jhu_df.loc[~pd.notna(jhu_df["state"])]
jhu_df = jhu_df.drop(columns=["state"])
if MERGE_SINGLE_SITE_COUNTRIES:
jhu_df["country"] = jhu_df["country"].apply(merge_single_site_country_adult_name)
jhu_df = jhu_df.loc[jhu_df["country"].isin(COUNTRIES)]
jhu_df = jhu_df.melt(id_vars=["country"], var_name="date", value_name="cumulative_count")
jhu_df["date"] = jhu_df["date"].astype(str)
jhu_df = jhu_df.groupby(["country", "date"]).sum().reset_index()
########################################################
# If you want to use public hospitalization data, add here!
########################################################
if(isHospitalizationData):
### France
fr = pd.read_csv("../data/hospitalization_france.csv")
fr.date = fr.date.astype(str)
fr.new_hospitalization = fr.apply(lambda x: x.new_hospitalization_per_100000 / 100000 * COUNTRY_POPULATION[x.country] / 7, axis='columns')
fr = fr.rename(columns={'new_hospitalization': 'cumulative_count'})
fr.cumulative_count = fr.cumulative_count.cumsum() # already sorted by date, so no need to sort
fr.to_csv("../data/hospitalization_france_calculated.csv")
jhu_df = jhu_df[jhu_df.country != 'France']
jhu_df = jhu_df.append(fr)
### Germany
de = pd.read_csv("../data/hospitalization_germany.csv")
de.date = de.date.astype(str)
de.new_hospitalization = de.apply(lambda x: x.new_hospitalization_per_100000 / 100000 * COUNTRY_POPULATION[x.country] / 7, axis='columns')
de = de.rename(columns={'new_hospitalization': 'cumulative_count'})
de.cumulative_count = de.cumulative_count.cumsum() # already sorted by date, so no need to sort
de.to_csv("../data/hospitalization_germany_calculated.csv")
jhu_df = jhu_df[jhu_df.country != 'Germany']
jhu_df = jhu_df.append(de)
### USA
us = pd.read_csv("../data/hospitalization_usa.csv")
us.date = us.date.astype(str)
#... # we already have raw new hospitalization admission data
us = us.rename(columns={'new_hospitalization': 'cumulative_count'})
us.cumulative_count = us.cumulative_count.cumsum() # already sorted by date, so no need to sort
us.to_csv("../data/hospitalization_usa_calculated.csv")
jhu_df = jhu_df[jhu_df.country != 'USA']
jhu_df = jhu_df.append(us)
### Spain
es = pd.read_csv("../data/hospitalization_spain.csv")
es.date = es.date.astype(str)
es.new_hospitalization = es.apply(lambda x: x.new_hospitalization_per_100000 / 100000 * COUNTRY_POPULATION[x.country] / 7, axis='columns')
es = es.rename(columns={'new_hospitalization': 'cumulative_count'})
es.cumulative_count = es.cumulative_count.cumsum() # already sorted by date, so no need to sort
es.to_csv("../data/hospitalization_spain_calculated.csv")
es.country = 'Spain + Singapore'
jhu_df = jhu_df[jhu_df.country != 'Spain + Singapore']
jhu_df = jhu_df.append(es)
########################################################
# End of data replacement
########################################################
jhu_df["date"] = jhu_df["date"].apply(convert_date)
jhu_df = jhu_df.sort_values(by="date", ascending=True)
jhu_df = jhu_df.loc[(jhu_df["date"] >= min_date) & (jhu_df["date"] <= max_date)]
jhu_df["date_str"] = jhu_df["date"].astype(str)
jhu_df_freeze = jhu_df.copy()
jhu_roc_df = pd.DataFrame(index=[], data=[], columns=["country", "date", "cumulative_count"])
for country, country_df in jhu_df.groupby("country"):
country_df = country_df.copy()
country_df["count"] = np.concatenate((np.array([np.nan]), np.diff(country_df["cumulative_count"].values)))
country_df["cumulative_count"] = country_df["cumulative_count"].replace(0, np.nan)
country_df["N0"] = country_df["cumulative_count"].shift(1) # N0 is the total case up to the day before
country_df["n1"] = country_df["count"] # n1 is the case number this day
country_df["n2"] = country_df["n1"].shift(1) # n2 is the case number yesterday
country_df["percent_increase"] = (country_df["n1"] / country_df["N0"]) * 100
country_df['R'] = country_df["percent_increase"] # TODO: is this correct?
# TODO: update CI formula
country_df['C'] = country_df['R'] - 1
country_df['standard_error'] = country_df.apply(lambda obs: (obs['R']+np.power(obs['R'], 2))/obs['n2'], axis='columns')
country_df['95_CI_below'] = country_df.apply(lambda obs: obs['C'] - 1.96*np.sqrt(obs['standard_error']), axis='columns')
country_df['95_CI_above'] = country_df.apply(lambda obs: obs['C'] + 1.96*np.sqrt(obs['standard_error']), axis='columns')
country_df = country_df.replace([np.inf, -np.inf], np.nan)
jhu_roc_df = jhu_roc_df.append(country_df, ignore_index=True)
jhu_roc_df
def get_jhu_cumulative_count(date_str, country):
try:
return jhu_roc_df.loc[(jhu_roc_df["date_str"] == date_str) & (jhu_roc_df["country"] == country)].reset_index().iloc[0]["cumulative_count"]
except:
return 0
# Start plotting after country has 100 cases
count_threshold = 100
jhu_roc_df["jhu_past_100"] = jhu_roc_df["cumulative_count"] >= count_threshold
# Transform the 4CE data to obtain normalized change values.
def compute_change_4ce(df_dc, cumulative_count_colname, daily_count_colname, category):
df_dc["cumulative_count"] = df_dc[cumulative_count_colname]
df_dc["count"] = df_dc[daily_count_colname]
# Sort dates
df_dc["date"] = df_dc["date"].astype(str)
df_dc["date"] = df_dc["date"].apply(convert_date)
df_dc = df_dc.sort_values(by="date", ascending=True)
df_dc = df_dc.loc[(df_dc["date"] >= min_date) & (df_dc["date"] <= max_date)]
df_dc["date_str"] = df_dc["date"].astype(str)
df_dc_freeze = df_dc.copy()
dc_roc_df = pd.DataFrame(index=[], data=[], columns=["country", "date"])
for country, country_df in df_dc.groupby("country"):
country_df = country_df.copy()
country_df["N0"] = country_df["cumulative_count"].shift(1) # N0 is the total case up to the day before
country_df["n1"] = country_df["count"] # n1 is the case number this day
country_df["n2"] = country_df["n1"].shift(1) # n2 is the case number yesterday
country_df["percent_increase"] = (country_df["n1"] / country_df["N0"]) * 100
country_df['R'] = country_df["percent_increase"] # TODO: is this correct?
# TODO: update CI formula
country_df['C'] = country_df['R'] - 1
country_df['standard_error'] = country_df.apply(lambda obs: (obs['R']+np.power(obs['R'], 2))/obs['n2'], axis='columns')
country_df['95_CI_below'] = country_df.apply(lambda obs: obs['C'] - 1.96*np.sqrt(obs['standard_error']), axis='columns')
country_df['95_CI_above'] = country_df.apply(lambda obs: obs['C'] + 1.96*np.sqrt(obs['standard_error']), axis='columns')
country_df = country_df.replace([np.inf, -np.inf], np.nan)
dc_roc_df = dc_roc_df.append(country_df, ignore_index=True)
dc_roc_df.head()
dc_roc_df = dc_roc_df[["country", "date", "num_sites", "num_hosps", "cumulative_count", "count", "date_str", "N0", "n1", "n2", "percent_increase", "R", "C", "standard_error", "95_CI_below", "95_CI_above"]]
dc_roc_df["category"] = category
dc_roc_df["jhu_past_100"] = dc_roc_df.apply(lambda row: get_jhu_cumulative_count(row["date_str"], row["country"]) >= count_threshold, axis='columns')
return dc_roc_df
dc_roc_df_all = compute_change_4ce(country_sum_df, "cumulative_patients_all", "cum_diff_all", "All")
dc_roc_df_severe = compute_change_4ce(country_sum_df, "cumulative_patients_severe", "cum_diff_severe", "Severe")
dc_roc_df_all_minus_severe = compute_change_4ce(country_sum_df, "cumulative_patients_all_minus_severe", "cum_diff_all_minus_severe", "All minus Severe")
dc_roc_df = dc_roc_df_all.append(dc_roc_df_severe, ignore_index=True).append(dc_roc_df_all_minus_severe, ignore_index=True)
dc_roc_df.to_csv("../data/dc_roc_df.csv")
```
### Transform data for plots faceted by country
```
jhu_roc_df = jhu_roc_df.copy()
dc_roc_df = dc_roc_df.copy()
jhu_roc_df["source"] = "JHU CSSE"
dc_roc_df["source"] = "4CE"
join_df = jhu_roc_df.append(dc_roc_df, ignore_index=True)
join_df["country_source"] = join_df.apply(lambda row: row["country"] + "_" + row["source"], axis='columns')
join_df.head()
```
## Normalized New Daily Cases
First, obtain total hospital discharges for each country.
$\texttt{country_total} = \text{country total in-patient-discharge}$
$\texttt{country_4CE_total} = \text{total in-patient-discharge in our sites within that country}$
$F0 = \frac{\texttt{country_total}}{\texttt{country_4CE_total}}$
$F0$ is used to normalize.
- For new figure that shows daily case # per 100K, we will instead show
- $\texttt{RATE} = \texttt{N_case} * F1$
- where $F1 = F0 * \frac{100K}{\texttt{country population}}$
- then the standard error for $\texttt{RATE}$ will be $\sqrt(\texttt{RATE}*F1)$ and the confidence interval will be $\texttt{RATE} \pm 1.96*\sqrt(\texttt{RATE}*F1)$
```
# Get daily new cases from cumulative_count
norm_jhu = jhu_roc_df.copy()
# Ensure not to calculate diff between country
# norm_jhu["count"] = norm_jhu["cumulative_count"].diff()
u_countries = norm_jhu.country.unique()
for c in u_countries:
norm_jhu.loc[norm_jhu.country == c, "count"] = norm_jhu.loc[norm_jhu.country == c, "cumulative_count"].diff()
norm_jhu.loc[norm_jhu["date"] == "2020-01-28", "count"] = np.nan # Make sure the start count is NaN
norm_jhu[norm_jhu.country == "Germany"]
norm_4ce = dc_roc_df.copy()
```
## Compute F0 value for each country using COUNTRY_POPULATION numbers
```
country_sites_df = sites_df.groupby("country").sum().reset_index()
country_sites_df = country_sites_df[["country", "adult_num_hosp", "adult_num_beds", "adult_num_yearly_discharge"]]
country_sites_df = country_sites_df.rename(columns={
"adult_num_hosp": "4ce_num_hosp",
"adult_num_beds": "4ce_num_beds",
"adult_num_yearly_discharge": "4ce_num_yearly_discharge",
})
country_sites_df["country_num_yearly_discharge_per_100000"] = country_sites_df["country"].apply(lambda c: COUNTRY_HOSP_DISCHARGE[c])
country_sites_df["country_population"] = country_sites_df["country"].apply(lambda c: COUNTRY_POPULATION[c])
if MERGE_SINGLE_SITE_COUNTRIES:
country_sites_df["country"] = country_sites_df["country"].apply(merge_single_site_country_adult_name)
country_sites_df = country_sites_df.groupby(["country"]).sum().reset_index()
country_sites_df["F1"] = country_sites_df["country_num_yearly_discharge_per_100000"] / country_sites_df["4ce_num_yearly_discharge"]
country_sites_df["F0"] = country_sites_df["F1"] * (country_sites_df["country_population"] / 100000)
country_sites_df
country_sites_df.to_csv("../data/CASE_RATE_CONSTANTS.csv")
F0 = dict(zip(country_sites_df["country"].values.tolist(), country_sites_df["F0"].values.tolist()))
F0
# Append the F0 values to each df
norm_jhu["population"] = norm_jhu["country"].apply(lambda x: COUNTRY_POPULATION[x])
norm_jhu["F0"] = norm_jhu["country"].apply(lambda x: F0[x])
norm_4ce["population"] = norm_4ce["country"].apply(lambda x: COUNTRY_POPULATION[x])
norm_4ce["F0"] = norm_4ce["country"].apply(lambda x: F0[x])
# Compute adjusted counts
norm_jhu["adjusted_count"] = norm_jhu["count"]
norm_jhu["F1"] = 100000 / norm_jhu["population"]
norm_jhu["RATE"] = norm_jhu["count"] * norm_jhu["F1"]
norm_jhu["RATE_7_day_avg"] = norm_jhu["RATE"].rolling(7).mean().shift(-3)
norm_jhu["std_error"] = norm_jhu["F1"] * norm_jhu["RATE"]
norm_jhu["std_error"] = norm_jhu["std_error"].apply(lambda x: np.sqrt(x))
norm_jhu["ci_above"] = norm_jhu["RATE_7_day_avg"] + 1.96 * norm_jhu["std_error"]
norm_jhu["ci_below"] = norm_jhu["RATE_7_day_avg"] - 1.96 * norm_jhu["std_error"]
norm_4ce["adjusted_count"] = norm_4ce["F0"] * norm_4ce["count"]
norm_4ce["F1"] = norm_4ce["F0"] * 100000 / norm_4ce["population"]
norm_4ce["RATE"] = norm_4ce["count"] * norm_4ce["F1"]
for c in COUNTRIES:
c_filter = norm_4ce["country"] == c
norm_4ce.loc[c_filter, "RATE_7_day_avg"] = norm_4ce.loc[c_filter, "RATE"].rolling(7).mean().shift(-3)
# norm_4ce["RATE_7_day_avg"] = norm_4ce["RATE"].rolling(7).mean().shift(-3)
norm_4ce["std_error"] = norm_4ce["F1"] * norm_4ce["RATE"]
norm_4ce["std_error"] = norm_4ce["std_error"].apply(lambda x: np.sqrt(x))
norm_4ce["ci_above"] = norm_4ce["RATE_7_day_avg"] + 1.96 * norm_4ce["std_error"]
norm_4ce["ci_below"] = norm_4ce["RATE_7_day_avg"] - 1.96 * norm_4ce["std_error"]
norm_jhu_min_col = norm_jhu[['country', 'date', 'count', 'adjusted_count', 'RATE_7_day_avg', 'source', 'population', 'F0', 'F1', 'RATE', 'std_error', 'ci_above', 'ci_below', 'jhu_past_100']]
norm_4ce_min_col = norm_4ce[['country', 'date', 'count', 'adjusted_count', 'RATE_7_day_avg', 'source', 'population', 'F0', 'F1', 'RATE', 'std_error', 'ci_above', 'ci_below', 'jhu_past_100', 'num_sites', 'num_hosps', 'category']]
norm_jhu_min_col.head(10)
norm_4ce_min_col.head(10)
```
## Temporary: select the All category of 4CE only (ignoring severity)
```
norm_4ce_min_col = norm_4ce_min_col.loc[norm_4ce_min_col["category"] != "All minus Severe"]
norm_jhu_min_col = norm_jhu_min_col.copy()
norm_4ce_min_col = norm_4ce_min_col.copy()
print(norm_4ce_min_col["category"].unique())
if isHospitalizationData:
norm_jhu_min_col["category"] = "Hospital admission rate"
else:
norm_jhu_min_col["category"] = "JHU CSSE: Positive Cases"
norm_4ce_min_col["category"] = norm_4ce_min_col["category"].apply(lambda x: "4CE: " + x)
norm_df = norm_jhu_min_col.append(norm_4ce_min_col, ignore_index=True)
norm_df["country_source"] = norm_df.apply(lambda row: row["country"] + "_" + row["source"], axis='columns')
norm_df.to_csv("../data/norm_df.csv")
norm_fce_df = norm_df.loc[norm_df['source'] == '4CE'].copy()
norm_df
norm_df["category"].unique()
```
## Temporary: drop Spain since so few discharges per year, it causes the rates to be way off
```
#norm_df = norm_df.loc[norm_df["country"] != "Spain"]
#norm_fce_df = norm_fce_df.loc[norm_fce_df["country"] != "Spain"]
min_date_3 = min_date + datetime.timedelta(days=3)
max_date_3 = max_date - datetime.timedelta(days=3)
norm_df = norm_df.loc[(norm_df["jhu_past_100"]) & (norm_df["date"] >= min_date_3) & (norm_df["date"] <= max_date_3)]
title = "Country-Level Positive Case Rate, Comparison to JHU CSSE Data"
# Selection
source_selection = alt.selection_multi(fields=["source"], bind="legend")
min_date = norm_df["date"].min()
max_date = norm_df["date"].max()
norm_fce_df = norm_fce_df.loc[(norm_fce_df["date"] >= min_date) & (norm_fce_df["date"] <= max_date)]
# Domains
date_domain = [alt.DateTime(year=min_date.year, month=min_date.month, date=min_date.day), alt.DateTime(year=max_date.year, month=max_date.month, date=max_date.day)]
sites_domain = [0, norm_fce_df["num_hosps"].max() + 1]
patients_domain = [0, norm_fce_df["count"].max() + 1]
rate_domain = [0, norm_fce_df["RATE_7_day_avg"].max() + 1]
country_names = COUNTRIES
COUNTRY_COLORS = [country_color_map[c] for c in country_names]
country_source_names = [c + "_" + "4CE" for c in country_names] + [c + "_" + "JHU CSSE" for c in country_names]
color_scale = alt.Scale(domain=country_names, range=COUNTRY_COLORS)
join_color_scale = alt.Scale(domain=country_source_names, range=COUNTRY_COLORS + ["#707070"] * len(country_names))
country_width = 170
nearest = alt.selection_single(encodings=['x', 'y'], on="mouseover", nearest=True, empty="none", clear="mouseout")
y_selection = alt.selection_interval(encodings=["y"], bind="scales", init={"y": rate_domain})
date_brush = alt.selection(type='interval', encodings=['x'])
# Additional Visual Elements
tooltip = [
alt.Tooltip("source", title="Data source"),
alt.Tooltip("country", title="Country"),
alt.Tooltip("count", title="Daily Cases"),
alt.Tooltip("adjusted_count", title="Adjusted Daily Cases"),
alt.Tooltip("RATE_7_day_avg", title="Daily Case Rate, 7-day Average", format=".2f"),
alt.Tooltip("date", title="Date"),
alt.Tooltip("ci_below", title="95% CI upper bound", format=".2f"),
alt.Tooltip("ci_above", title="95% CI lower bound", format=".2f")
]
rule = alt.Chart().mark_rule(color="red", size=0.5).encode(
x="date:T"
).transform_filter(
nearest
)
line = alt.Chart(norm_df).transform_filter(source_selection).mark_line(opacity=0.7).encode(
x=alt.X("date:T", title=None, axis=alt.Axis(labelBound=True), scale=alt.Scale(padding=5)),
y=alt.Y("RATE_7_day_avg:Q", axis=alt.Axis(title="Adjusted daily case rate, 7 day average"), scale=alt.Scale(zero=False, nice=False, domain=rate_domain, padding=5)),
strokeDash=alt.StrokeDash("source:N", scale=alt.Scale(domain=["4CE", "JHU CSSE"], range=[[0,0], [3,3]]),
legend=alt.Legend(title="Data Source")),
color=alt.Color("country_source:N", scale=join_color_scale, legend=None),
tooltip=tooltip
).properties(width=country_width, height=200)
errorband = line.transform_filter(alt.datum["source"] == "4CE").mark_errorband().encode(
x=alt.X(f"date:T", title=None, axis=alt.Axis(labelBound=True)),
y=alt.Y(f"sum(ci_below):Q", title=""),
y2=alt.Y2(f"sum(ci_above):Q", title=""),
color=alt.Color(f"country:N", scale=color_scale, legend=alt.Legend(title=None)),
tooltip=tooltip
)
circle = (
line.mark_circle()
.encode(
size=alt.condition(~nearest, alt.value(5), alt.value(30))
)
.add_selection(nearest)
)
num_sites_bar_bg = (
alt.Chart(norm_fce_df)
.mark_bar(size=2)
.encode(
x=alt.X("date:T", scale=alt.Scale(domain=date_domain, padding=5), title=None, axis=alt.Axis(labelBound=True)),
y=alt.Y("num_sites:Q", axis=alt.Axis(title="# of sites"), scale=alt.Scale(domain=sites_domain)),
color=alt.value("gray"),
tooltip=tooltip
)
.properties(width=country_width, height=60)
)
num_sites_bar = (
num_sites_bar_bg
.encode(
color=alt.Color("country:N", scale=color_scale, legend=None),
)
.transform_filter(date_brush)
)
num_patients_bar_bg = (
alt.Chart(norm_fce_df)
.mark_bar(size=2)
.encode(
x=alt.X("date:T", scale=alt.Scale(domain=date_domain, padding=5), title=None, axis=alt.Axis(labelBound=True)),
y=alt.Y("count:Q", axis=alt.Axis(title="# of new cases"), scale=alt.Scale(domain=patients_domain)),
color=alt.value("gray"),
tooltip=tooltip
)
.properties(width=country_width, height=60)
)
num_patients_bar = (
num_patients_bar_bg
.encode(
color=alt.Color("country:N", scale=color_scale, legend=None),
)
.transform_filter(date_brush)
)
top = (
alt.layer(line, errorband, circle, rule, data=norm_df)
.facet(
column=alt.Column("country:N"), bounds="flush" #header=alt.Header(labels=False)
)
.add_selection(y_selection)
.transform_filter(date_brush)
)
num_sites_bottom = (
alt.layer(num_sites_bar_bg, num_sites_bar, rule, data=norm_fce_df)
.facet(
column=alt.Column("country:N", header=alt.Header(labels=False)), bounds="flush"
)
.add_selection(nearest)
.add_selection(date_brush)
)
num_patients_bottom = (
alt.layer(num_patients_bar_bg, num_patients_bar, rule, data=norm_fce_df)
.facet(
column=alt.Column("country:N", header=alt.Header(labels=False)), bounds="flush"
)
.add_selection(nearest)
.add_selection(date_brush)
)
plot = (
alt.vconcat(top, num_patients_bottom, num_sites_bottom, spacing=5)
.resolve_scale(color="shared", x="independent")
.properties(title={
"text": title,
"subtitle": get_visualization_subtitle(data_release=DATA_RELEASE, cohort=COHORT, num_sites=NUM_SITES),
"subtitleColor": "gray",
"dx": 60
})
.add_selection(source_selection)
)
plot = apply_theme(
plot,
axis_label_font_size=10,
axis_title_font_size=12,
axis_title_padding=8,
legend_orient="bottom",
legend_symbol_type="stroke",
legend_title_orient="left",
legend_title_font_size=14,
label_font_size=12
).configure_header(title=None, labelPadding=3, labelFontSize=13)
for_website(plot, "Daily Count", "country-level rate of positive cases")
plot
norm_df = norm_df.dropna(subset=["RATE_7_day_avg"])
norm_df = norm_df.loc[~((norm_df["source"] == "JHU CSSE") & (norm_df["RATE_7_day_avg"] < 0))]
# Revision 2021-06-28:
# - remove Singapore, Italy and Spain
# - show data time points until Sep. 29th, 2020
norm_df = norm_df.loc[~norm_df["country"].isin(["Singapore", "Italy"])].copy()
norm_df = norm_df.loc[norm_df["category"] != "4CE: Severe"].copy()
norm_fce_df = norm_fce_df.loc[~norm_fce_df["country"].isin(["Singapore", "Italy"])].copy()
## Change category since we are now using differen data sources
if isHospitalizationData:
pub_data_category_str = 'Hospital admission rate'
norm_df.loc[norm_df.source == "JHU CSSE", 'source'] = pub_data_category_str
norm_fce_df.loc[norm_fce_df.source == "JHU CSSE", 'source'] = pub_data_category_str
else:
pub_data_category_str = "JHU CSSE"
###
min_date = norm_df["date"].min()
max_date = datetime.date(year=2020, month=9, day=29)
date_domain = [alt.DateTime(year=min_date.year, month=min_date.month, date=min_date.day), alt.DateTime(year=max_date.year, month=max_date.month, date=max_date.day)]
date_brush = alt.selection(type='interval', encodings=['x'])
y_selection = alt.selection(type='interval', encodings=["y"], bind="scales", init={"y": rate_domain})
tooltip = [
alt.Tooltip("source", title="Data source"),
alt.Tooltip("country", title="Country"),
alt.Tooltip("count", title="Daily Cases"),
alt.Tooltip("adjusted_count", title="Adjusted Daily Cases"),
alt.Tooltip("RATE_7_day_avg", title="Daily Case Rate, 7-day Average", format=".2f"),
alt.Tooltip("date", title="Date"),
alt.Tooltip("num_hosps", title="Number of 4CE hospitals"),
alt.Tooltip("ci_below", title="95% CI upper bound", format=".2f"),
alt.Tooltip("ci_above", title="95% CI lower bound", format=".2f")
]
chart = alt.Chart(norm_df)
if isHospitalizationData:
yTitle = "Hospital admission rate, 7-day average"
else:
yTitle = "Adjusted daily case rate, 7-day average"
line = alt.Chart(norm_df).mark_line(opacity=0.7, size=3).encode(
x=alt.X("date:T", title=None, axis=alt.Axis(labelBound=True), scale=alt.Scale(domain=date_domain, padding=5)),
y=alt.Y("RATE_7_day_avg:Q", axis=alt.Axis(title=yTitle), scale=alt.Scale(zero=False, nice=False, domain=rate_domain, padding=5)),
strokeDash=alt.StrokeDash("source:N", scale=alt.Scale(domain=[pub_data_category_str, "4CE"], range=[[3,3], [0,0]]), legend=None), #, legend=alt.Legend(title="Data Source", symbolStrokeWidth=4)),
color=alt.Color("source:N", legend=alt.Legend(title=None, symbolStrokeWidth=6)), # scale=alt.Scale(range=['#E79F00', '#57B4E9', '#D45E00']))
tooltip=tooltip
).properties(width=country_width, height=200)
errorband = line.transform_filter(alt.datum["source"] == "4CE").mark_errorband().encode(
x=alt.X(f"date:T", title=None, axis=alt.Axis(labelBound=True)),
y=alt.Y(f"sum(ci_below):Q", title=""),
y2=alt.Y2(f"sum(ci_above):Q", title=""),
tooltip=tooltip
)
top = (
alt.layer(line, errorband)
.facet(
column=alt.Column("country:N"), bounds="flush" #header=alt.Header(labels=False)
)
.transform_filter(date_brush)
.add_selection(y_selection)
)
# BOTTOM
if isHospitalizationData:
midYTitle = '4CE admissions'
else:
midYTitle = '4CE cases'
num_patients_bar = alt.Chart(norm_fce_df).mark_bar(size=2).encode(
x=alt.X("date:T", scale=alt.Scale(domain=date_domain, padding=5), title=None, axis=alt.Axis(labelBound=True, labels=False, ticks=False)),
y=alt.Y("count:Q", axis=alt.Axis(title=midYTitle), scale=alt.Scale(domain=patients_domain)),
color=alt.value("gray"),
tooltip=tooltip
).properties(width=country_width, height=70).facet(
column=alt.Column("country:N", header=alt.Header(labels=False)), bounds="flush"
).transform_filter(date_brush).add_selection(date_brush)
num_sites_bar = alt.Chart(norm_fce_df).mark_bar(size=2).encode(
x=alt.X("date:T", scale=alt.Scale(domain=date_domain, padding=5), title=None, axis=alt.Axis(labelBound=True)),
y=alt.Y("num_hosps:Q", axis=alt.Axis(title="4CE hospitals"), scale=alt.Scale(domain=sites_domain)),
color=alt.value("gray"),
tooltip=tooltip
).properties(width=country_width, height=50).facet(
column=alt.Column("country:N", header=alt.Header(labels=False)), bounds="flush"
).transform_filter(date_brush).add_selection(date_brush)
if isHospitalizationData:
title = "Hospital Admission Rate by Country"
else:
title = "Country-Level Positive Case Rate, Comparison to JHU CSSE Data"
plot = (
alt.vconcat(top, num_patients_bar, num_sites_bar, spacing=5)
.properties(title={
"text": title,
"subtitle": "",
"subtitleColor": "gray",
"dx": 14
})
)
plot = apply_theme(
plot,
axis_label_font_size=10,
axis_title_font_size=10,
axis_title_padding=8,
legend_orient="bottom",
legend_symbol_type="stroke",
legend_title_orient="left",
legend_title_font_size=14,
label_font_size=12
).configure_header(title=None, labelPadding=3, labelFontSize=13)
for_website(plot, f"Case Rate Comparison {COHORT}", f"4CE vs {pub_data_category_str}", df=norm_df)
plot
```
| github_jupyter |
```
# MASTER ONLY
%load_ext prog_edu_assistant_tools.magics
from prog_edu_assistant_tools.magics import report, autotest
```
```
# ASSIGNMENT METADATA
assignment_id: "q13"
```
## 練習
1. 数値 `x` の絶対値を求める関数 `absolute(x)` を定義してください。
(Pythonには `abs` という組み込み関数が用意されていますが。)
定義ができたら、その次のセルを実行して、`True` のみが表示されることを確認してください。
```
# EXERCISE METADATA
exercise_id: "Absolute"
```
```
%%solution
def absolute(x):
# BEGIN SOLUTION
if x<0:
return -x
else:
return x
# END SOLUTION
print(absolute(5) == 5)
print(absolute(-5) == 5)
print(absolute(0) == 0)
%%inlinetest AutograderTest_Absolute
assert 'absolute' in globals(), "Did you define a function named 'absolute' in the solution cell?"
assert str(absolute.__class__) == "<class 'function'>", f"Did you define 'absolute' as a function? There was a {absolute.__class__} instead"
def test_absolute(x, expected, hint):
actual = absolute(x)
assert actual is not None, f"Did you return a value from the function? Got {actual}"
try:
float(actual)
except ValueError:
assert False, f"Did you return a number from the function? Got {actual}"
assert actual == expected, f"{hint} Expected {expected}, but got {actual}"
test_absolute(x=1, expected=1, hint="Did you handle positive numbers correctly?")
test_absolute(x=-1, expected=1, hint="Did you handle negative numbers correctly?")
test_absolute(x=0, expected=0, hint="Did you handle 0 correctly?")
test_absolute(x=3.1415, expected=3.1415, hint="Did you handle non-integer numbers correctly?")
test_absolute(x=-3.1415, expected=3.1415, hint="Did you handle non-integer numbers correctly?")
```
## 練習
1. `x` が正ならば 1、負ならば -1、ゼロならば 0 を返す `sign(x)` という関数を定義してください。
定義ができたら、その次のセルを実行して、`True` のみが表示されることを確認してください。
```
# EXERCISE METADATA
exercise_id: "Sign"
```
```
%%solution
def sign(x):
if x<0:
return -1
if x>0:
return 1
return 0
print(sign(5) == 1)
print(sign(-5) == -1)
print(sign(0) == 0)
```
## 練習の解答
```
%%inlinetest AutograderTest_Sign
assert 'sign' in globals(), "Did you define a function named 'sign' in the solution cell?"
assert str(sign.__class__) == "<class 'function'>", f"Did you define 'absolute' as a function? There was a {sign.__class__} instead"
def test_sign(x, expected, hint):
actual = sign(x)
assert actual is not None, f"Did you return a value from the function? Got {actual}"
try:
float(actual)
except ValueError:
assert False, f"Did you return a number from the function? Got {actual}"
assert actual == expected, f"{hint} Expected {expected}, but got {actual}"
test_sign(x=1, expected=1, hint="Does the function return 1 for positive numbers?")
test_sign(x=-1, expected=-1, hint="Does the function return -1 for negative numbers?")
test_sign(x=0, expected=0, hint="Does the function return 0 for 0?")
test_sign(x=0.5, expected=1, hint="Does the function return 1 for positive numbers less than 1?")
test_sign(x=21345, expected=1, hint="Does the function return 1 for positive numbers greater than 1?")
test_sign(x=-0.5, expected=-1, hint="Does the function return 1 for negative numbers greater than -1?")
test_sign(x=-21345, expected=-1, hint="Does the function return 1 for negative numbers less than -1?")
```
| github_jupyter |
## TODO
* `gb_accession` and `gisaid_accession` are not found for new sequences, how do we concat to `metadata.csv` without them?
* metadata format for NCBI
* support tools for manual sanity checks
```
from bjorn import *
from bjorn_support import *
from onion_trees import *
import gffutils
import math
from mutations import *
input_fasta = "/home/al/analysis/mutations/S501Y/msa_reference.fa"
meta_fp = "/home/al/analysis/mutations/S501Y/metadata_2020-12-20_12-24.tsv"
out_dir = "/home/al/analysis/mutations/S501Y/"
ref_fp = "/home/al/data/test_inputs/NC045512.fasta"
patient_zero = 'NC_045512.2'
## keep only seqs contained in meta_file and save to fasta file
## concat with internal SD file
## generate MSA
meta = pd.read_csv(meta_fp, sep='\t')
meta.columns
# consensus_data = SeqIO.to_dict(SeqIO.parse(seqs_fp, "fasta"))
strains = meta['strain'].unique().tolist()
len(strains)
print(f"Loading Alignment file at: {input_fasta}")
cns = AlignIO.read(input_fasta, 'fasta')
print(f"Initial cleaning...")
seqs, ref_seq = process_cns_seqs(cns, patient_zero,
start_pos=0, end_pos=30000)
print(f"Creating a dataframe...")
seqsdf = (pd.DataFrame(index=seqs.keys(),
data=seqs.values(),
columns=['sequence'])
.reset_index()
.rename(columns={'index': 'idx'}))
def find_replacements(x, ref):
return [f'{i}:{n}' for i, n in enumerate(x)
if n!=ref[i] and n!='-' and n!='n']
print(f"Identifying mutations...")
# for each sample, identify list of substitutions (position:alt)
seqsdf['replacements'] = seqsdf['sequence'].apply(find_replacements, args=(ref_seq,))
# wide-to-long data manipulation
seqsdf = seqsdf.explode('replacements')
# seqsdf
seqsdf['pos'] = -1
# populate position column
seqsdf.loc[~seqsdf['replacements'].isna(), 'pos'] = (seqsdf.loc[~seqsdf['replacements'].isna(), 'replacements']
.apply(lambda x: int(x.split(':')[0])))
# filter out non-substitutions
seqsdf = seqsdf.loc[seqsdf['pos']!=-1]
print(f"Mapping Genes to mutations...")
# identify gene of each substitution
seqsdf['gene'] = seqsdf['pos'].apply(map_gene_to_pos)
seqsdf = seqsdf.loc[~seqsdf['gene'].isna()]
# seqsdf
# filter our substitutions in non-gene positions
seqsdf = seqsdf.loc[seqsdf['gene']!='nan']
print(f"Compute codon numbers...")
# compute codon number of each substitution
seqsdf['codon_num'] = seqsdf.apply(compute_codon_num, args=(GENE2POS,), axis=1)
print(f"Fetch reference codon...")
# fetch the reference codon for each substitution
seqsdf['ref_codon'] = seqsdf.apply(get_ref_codon, args=(ref_seq, GENE2POS), axis=1)
print(f"Fetch alternative codon...")
# fetch the alternative codon for each substitution
seqsdf['alt_codon'] = seqsdf.apply(get_alt_codon, args=(GENE2POS,), axis=1)
print(f"Map amino acids...")
# fetch the reference and alternative amino acids
seqsdf['ref_aa'] = seqsdf['ref_codon'].apply(get_aa)
seqsdf['alt_aa'] = seqsdf['alt_codon'].apply(get_aa)
# filter out substitutions with non-amino acid alternates (bad consensus calls)
seqsdf = seqsdf.loc[seqsdf['alt_aa']!='nan']
print(f"Fuse with metadata...")
# load and join metadata
meta = pd.read_csv(meta_fp, sep='\t')
seqsdf = pd.merge(seqsdf, meta, left_on='idx', right_on='strain')
seqsdf['date'] = pd.to_datetime(seqsdf['date_submitted'])
seqsdf['month'] = seqsdf['date'].dt.month
seqsdf.columns
seqsdf.loc[seqsdf['location'].isna(), 'location'] = 'unk'
out_dir = Path('/home/al/analysis/mutations/gisaid')
seqsdf.drop(columns=['sequence']).to_csv(out_dir/'gisaid_replacements_19-12-2020.csv', index=False)
seqsdf[['idx', 'sequence']].to_csv(out_dir/'gisaid_sequences_19-12-2020.csv', index=False)
seqsdf = pd.read_csv('/home/al/analysis/mutations/gisaid/gisaid_replacements_19-12-2020.csv')
seqsdf = seqsdf[seqsdf['host']=='Human']
print(f"Aggregate final results...")
# aggregate on each substitutions, compute number of samples and other attributes
subs = (seqsdf.groupby(['gene', 'pos', 'ref_aa', 'codon_num', 'alt_aa'])
.agg(
num_samples=('idx', 'nunique'),
first_detected=('date', 'min'),
last_detected=('date', 'max'),
num_locations=('location', 'nunique'),
location_counts=('location', lambda x: np.unique(x, return_counts=True)),
num_divisions=('division', 'nunique'),
division_counts=('division', lambda x: np.unique(x, return_counts=True)),
num_countries=('country', 'nunique'),
country_counts=('country', lambda x: np.unique(x, return_counts=True))
)
.reset_index())
# 1-based nucleotide position coordinate system
subs['pos'] = subs['pos'] + 1
# subs.sort_values('num_samples', ascending=False).iloc[0]['country_counts']
subs['locations'] = subs['location_counts'].apply(lambda x: list(x[0]))
subs['location_counts'] = subs['location_counts'].apply(lambda x: list(x[1]))
subs['divisions'] = subs['division_counts'].apply(lambda x: list(x[0]))
subs['division_counts'] = subs['division_counts'].apply(lambda x: list(x[1]))
subs['countries'] = subs['country_counts'].apply(lambda x: list(x[0]))
subs['country_counts'] = subs['country_counts'].apply(lambda x: list(x[1]))
print(f"Aggregate final results...")
# aggregate on each substitutions, compute number of samples and other attributes
subs_mnth = (seqsdf.groupby(['month', 'gene', 'pos', 'ref_aa', 'codon_num', 'alt_aa'])
.agg(
num_samples=('idx', 'nunique'),
first_detected_mnth=('date', 'min'),
last_detected_mnth=('date', 'max'),
num_locations=('location', 'nunique'),
# locations=('location', lambda x: list(np.unique(x))),
location_counts=('location', lambda x: np.unique(x, return_counts=True)),
num_divisions=('division', 'nunique'),
division_counts=('division', lambda x: np.unique(x, return_counts=True)),
num_countries=('country', 'nunique'),
# countries=('country', lambda x: list(np.unique(x))),
country_counts=('country', lambda x: np.unique(x, return_counts=True)),
)
.reset_index())
# 1-based nucleotide position coordinate system
subs_mnth['pos'] = subs_mnth['pos'] + 1
subs_mnth = pd.merge(subs_mnth, subs[['gene', 'pos', 'alt_aa', 'first_detected', 'last_detected']], on=['gene', 'pos', 'alt_aa'])
out_dir = Path('/home/al/analysis/mutations/gisaid')
subs.to_csv(out_dir/'gisaid_substitutions_aggregated_19-12-2020.csv', index=False)
top_s_mnthly = (subs_mnth[subs_mnth['gene']=='S'].sort_values('num_samples', ascending=False)
.drop_duplicates(subset=['gene', 'codon_num', 'alt_aa'])
.iloc[:50]
.reset_index(drop=True))
muts_of_interest = []
for i, mutation in top_s_mnthly.iterrows():
locs = mutation['location_counts'][0]
for l in locs:
if 'san diego' in l.lower():
muts_of_interest.append(i)
muts_of_interest
def is_in(x, loc):
for i in x[0]:
if loc in i.lower():
return True
return False
top_s_mnthly['isin_SD'] = top_s_mnthly['location_counts'].apply(is_in, args=('san diego',))
top_s_mnthly['isin_CA'] = top_s_mnthly['division_counts'].apply(is_in, args=('california',))
top_s_mnthly['isin_US'] = top_s_mnthly['country_counts'].apply(is_in, args=('usa',))
top_s_mnthly.to_csv("/home/al/analysis/mutations/gisaid/top_S_mutations_monthly.csv", index=False)
```
### Integrate GISAID information with ALab variants table
```
out_dir = Path('/home/al/analysis/mutations/gisaid')
subs = pd.read_csv(out_dir/'gisaid_substitutions_aggregated_19-12-2020.csv')
gisaid_subs = (subs.rename(columns={'num_samples': 'gisaid_num_samples', 'first_detected': 'gisaid_1st_detected', 'last_detected': 'gisaid_last_detected',
'num_locations': 'gisaid_num_locations', 'locations': 'gisaid_locations', 'location_counts': 'gisaid_location_counts',
'num_divisions': 'gisaid_num_states','divisions': 'gisaid_states', 'division_counts': 'gisaid_state_counts',
'num_countries': 'gisaid_num_countries', 'countries': 'gisaid_countries', 'country_counts': 'gisaid_country_counts'})
.drop(columns=['ref_aa', 'pos']))
gisaid_subs.columns
# gisaid_subs.sort_values('gisaid_num_samples', ascending=False).iloc[0]['gisaid_country_counts']
our_subs = pd.read_csv("/home/al/analysis/mutations/alab_git/substitutions_22-12-2020_orig.csv")
our_subs.shape
all_subs = pd.merge(our_subs, gisaid_subs, on=['gene', 'codon_num', 'alt_aa'], how='left').drop_duplicates(subset=['gene', 'codon_num', 'alt_aa'])
all_subs.columns
all_subs.sort_values('num_samples', ascending=False)
subs.loc[(subs['gene']=='S')&(subs['alt_aa']=='L')&(subs['codon_num']==957)]
cols = ['month', 'ref_aa', 'codon_num', 'alt_aa', 'first_detected',
'last_detected', 'num_samples', 'num_countries',
'countries', 'country_counts', 'num_locations', 'locations', 'location_counts' ,
'first_detected_mnth', 'last_detected_mnth']
# (subs_mnth[(subs_mnth['gene']=='S') & (subs_mnth['month']==12)]
# .sort_values('num_samples', ascending=False)
# .drop_duplicates(subset=['codon_num', 'alt_aa'], keep='first')
# .iloc[:50]
# .reset_index(drop=True))[cols]
# keys_df = seqsdf[['idx', 'sequence']]
# keys_df.to_csv('gisaid_replacements.csv', index=False)
sd = []
for d in seqsdf['location'].dropna().unique():
if 'san diego' in d.lower():
sd.append(d)
ca = []
for d in seqsdf['division'].unique():
if 'cali' in d.lower():
ca.append(d)
# cols = ['idx', 'location', 'division', 'pos']
# seqsdf.loc[(seqsdf['codon_num']==681) & (seqsdf['gene']=='S')][cols]
```
## Deletions
```
input_fasta = "/home/al/analysis/mutations/S501Y/msa_reference.fa"
meta_fp = "/home/al/analysis/mutations/S501Y/metadata_2020-12-20_12-24.tsv"
out_dir = "/home/al/analysis/mutations/S501Y/"
ref_fp = "/home/al/data/test_inputs/NC045512.fasta"
patient_zero = 'NC_045512.2'
min_del_len = 1
start_pos = 265
end_pos = 29674
# read MSA file
consensus_data = AlignIO.read(input_fasta, 'fasta')
# prcess MSA to remove insertions and fix position coordinate systems
seqs, ref_seq = process_cns_seqs(consensus_data, patient_zero, start_pos=start_pos, end_pos=end_pos)
# load into dataframe
seqsdf = (pd.DataFrame(index=seqs.keys(), data=seqs.values(),
columns=['sequence'])
.reset_index().rename(columns={'index': 'idx'}))
# load and join metadata
meta = pd.read_csv(meta_fp, sep='\t')
print(seqsdf.shape)
seqsdf = pd.merge(seqsdf, meta, left_on='idx', right_on='strain')
print(seqsdf.shape)
# # clean and process sample collection dates
# seqsdf = seqsdf.loc[(seqsdf['collection_date']!='Unknown')
# & (seqsdf['collection_date']!='1900-01-00')]
# seqsdf.loc[seqsdf['collection_date'].str.contains('/'), 'collection_date'] = seqsdf['collection_date'].apply(lambda x: x.split('/')[0])
seqsdf['date'] = pd.to_datetime(seqsdf['date_submitted'])
# compute length of each sequence
seqsdf['seq_len'] = seqsdf['sequence'].str.len()
# identify deletion positions
seqsdf['del_positions'] = seqsdf['sequence'].apply(find_deletions)
seqsdf.columns
seqsdf = seqsdf[seqsdf['host']=='Human']
# sequences with one or more deletions
del_seqs = seqsdf.loc[seqsdf['del_positions'].str.len() > 0]
del_seqs = del_seqs.explode('del_positions')
# compute length of each deletion
del_seqs['del_len'] = del_seqs['del_positions'].apply(len)
# only consider deletions longer than 2nts
del_seqs = del_seqs[del_seqs['del_len'] >= min_del_len]
# fetch coordinates of each deletion
del_seqs['relative_coords'] = del_seqs['del_positions'].apply(get_indel_coords)
del_seqs.loc[del_seqs['location'].isna(), 'location'] = 'unk'
# group sample by the deletion they share
del_seqs = (del_seqs.groupby(['relative_coords', 'del_len'])
.agg(
samples=('idx', 'unique'),
num_samples=('idx', 'nunique'),
first_detected=('date', 'min'),
last_detected=('date', 'max'),
# locations=('location', lambda x: list(np.unique(x))),
location_counts=('location', lambda x: np.unique(x, return_counts=True)),
# divisions=('division', lambda x: list(np.unique(x))),
division_counts=('division', lambda x: np.unique(x, return_counts=True)),
# countries=('country', lambda x: list(np.unique(x))),
country_counts=('country', lambda x: np.unique(x, return_counts=True)),
)
.reset_index()
.sort_values('num_samples'))
del_seqs['type'] = 'deletion'
# adjust coordinates to account for the nts trimmed from beginning e.g. 265nts
del_seqs['absolute_coords'] = del_seqs['relative_coords'].apply(adjust_coords, args=(start_pos+1,))
del_seqs['pos'] = del_seqs['absolute_coords'].apply(lambda x: int(x.split(':')[0]))
# approximate the gene where each deletion was identified
del_seqs['gene'] = del_seqs['pos'].apply(map_gene_to_pos)
del_seqs = del_seqs.loc[~del_seqs['gene'].isna()]
# filter our substitutions in non-gene positions
del_seqs = del_seqs.loc[del_seqs['gene']!='nan']
# compute codon number of each substitution
del_seqs['codon_num'] = del_seqs.apply(compute_codon_num, args=(GENE2POS,), axis=1)
# fetch the reference codon for each substitution
del_seqs['ref_codon'] = del_seqs.apply(get_ref_codon, args=(ref_seq, GENE2POS), axis=1)
# fetch the reference and alternative amino acids
del_seqs['ref_aa'] = del_seqs['ref_codon'].apply(get_aa)
# record the 5 nts before each deletion (based on reference seq)
del_seqs['prev_5nts'] = del_seqs['absolute_coords'].apply(lambda x: ref_seq[int(x.split(':')[0])-5:int(x.split(':')[0])])
# record the 5 nts after each deletion (based on reference seq)
del_seqs['next_5nts'] = del_seqs['absolute_coords'].apply(lambda x: ref_seq[int(x.split(':')[1])+1:int(x.split(':')[1])+6])
del_seqs['locations'] = del_seqs['location_counts'].apply(lambda x: list(x[0]))
del_seqs['location_counts'] = del_seqs['location_counts'].apply(lambda x: list(x[1]))
del_seqs['divisions'] = del_seqs['division_counts'].apply(lambda x: list(x[0]))
del_seqs['division_counts'] = del_seqs['division_counts'].apply(lambda x: list(x[1]))
del_seqs['countries'] = del_seqs['country_counts'].apply(lambda x: list(x[0]))
del_seqs['country_counts'] = del_seqs['country_counts'].apply(lambda x: list(x[1]))
del_seqs.sort_values('num_samples', ascending=False)
del_seqs.to_csv('/home/al/analysis/mutations/gisaid/gisaid_deletions_aggregated_19-12-2020.csv', index=False)
del_seqs.columns
gisaid_dels = (del_seqs.rename(columns={'num_samples': 'gisaid_num_samples', 'first_detected': 'gisaid_1st_detected', 'last_detected': 'gisaid_last_detected',
'locations': 'gisaid_locations', 'location_counts': 'gisaid_location_counts',
'divisions': 'gisaid_states', 'division_counts': 'gisaid_state_counts',
'countries': 'gisaid_countries', 'country_counts': 'gisaid_country_counts'})
.drop(columns=['ref_aa', 'pos', 'type', 'samples', 'ref_codon', 'prev_5nts', 'next_5nts', 'relative_coords', 'del_len']))
our_dels = pd.read_csv("/home/al/analysis/mutations/alab_git/deletions_22-12-2020_orig.csv")
# our_dels
cols = ['type', 'gene', 'absolute_coords', 'del_len', 'pos',
'ref_aa', 'codon_num', 'num_samples',
'first_detected', 'last_detected', 'locations',
'location_counts', 'gisaid_num_samples',
'gisaid_1st_detected', 'gisaid_last_detected', 'gisaid_countries', 'gisaid_country_counts',
'gisaid_states', 'gisaid_state_counts', 'gisaid_locations', 'gisaid_location_counts', 'samples',
'ref_codon', 'prev_5nts', 'next_5nts'
]
our_dels = pd.merge(our_dels, gisaid_dels, on=['gene', 'codon_num', 'absolute_coords'], how='left')
our_dels[cols]
our_dels[cols].sort_values('num_samples', ascending=False).to_csv("/home/al/analysis/mutations/alab_git/deletions_22-12-2020.csv", index=False)
align_fasta_reference(seqs_fp, num_cpus=25, ref_fp=ref_fp)
```
## CNS Mutations Report
```
analysis_folder = Path('/home/al/code/HCoV-19-Genomics/consensus_sequences/')
meta_fp = Path('/home/al/code/HCoV-19-Genomics/metadata.csv')
ref_path = Path('/home/gk/code/hCoV19/db/NC045512.fasta')
patient_zero = 'NC_045512.2'
in_fp = '/home/al/analysis/mutations/S501Y/msa_aligned.fa'
subs = identify_replacements(in_fp, meta_fp)
subs.head()
dels = identify_deletions(in_fp, meta_fp, patient_zero)
dels
dels[dels['gene']=='S'].sort_values('num_samples', ascending=False)#.to_csv('S_deletions_consensus.csv', index=False)
identify_insertions(in_fp, patient_zero).to_csv('test.csv', index=False)
```
## dev
```
GENE2POS = {
'5UTR': {'start': 0, 'end': 265},
'ORF1ab': {'start': 265, 'end': 21555},
'S': {'start': 21562, 'end': 25384},
'ORF3a': {'start': 25392, 'end': 26220},
'E': {'start': 26244, 'end': 26472},
'M': {'start': 26522, 'end': 27191},
'ORF6': {'start': 27201, 'end': 27387},
'ORF7a': {'start': 27393, 'end': 27759},
'ORF7b': {'start': 27755, 'end': 27887},
'ORF8': {'start': 27893, 'end': 28259},
'N': {'start': 28273, 'end': 29533},
'ORF10': {'start': 29557, 'end': 29674},
'3UTR': {'start': 29674, 'end': 29902}
}
in_dir = '/home/al/analysis/mutations/fa/'
out_dir = '/home/al/analysis/mutations/msa/'
!rm -r /home/al/analysis/mutations
!mkdir /home/al/analysis/mutations
!mkdir /home/al/analysis/mutations/fa
for filename in analysis_folder.listdir():
if (filename.endswith('fa') or filename.endswith('fasta')):
copy(filename, '/home/al/analysis/mutations/fa/')
# print(filename)
copy(ref_path, in_dir)
in_dir = '/home/al/analysis/mutations/fa/'
out_dir = '/home/al/analysis/mutations/msa'
concat_fasta(in_dir, out_dir)
align_fasta_reference('/home/al/analysis/mutations/msa.fa', num_cpus=12, ref_fp=ref_path)
cns = AlignIO.read('/home/al/analysis/mutations/msa_aligned.fa', 'fasta')
ref_seq = get_seq(cns, patient_zero)
len(ref_seq)
seqs = get_seqs(cns, 0, 30000)
seqsdf = (pd.DataFrame(index=seqs.keys(), data=seqs.values(), columns=['sequence'])
.reset_index().rename(columns={'index': 'idx'}))
# seqsdf
def find_replacements(x, ref):
return [f'{i}:{n}' for i, n in enumerate(x)
if n!=ref[i] and n!='-' and n!='n']
seqsdf['replacements'] = seqsdf['sequence'].apply(find_replacements, args=(ref_seq,))
seqsdf = seqsdf.explode('replacements')
seqsdf['pos'] = -1
seqsdf.loc[~seqsdf['replacements'].isna(), 'pos'] = seqsdf.loc[~seqsdf['replacements'].isna(), 'replacements'].apply(lambda x: int(x.split(':')[0]))
seqsdf = seqsdf.loc[seqsdf['pos']!=-1]
def compute_codon_num(x, gene2pos: dict):
pos = x['pos']
ref_pos = gene2pos[x['gene']]['start']
return math.ceil((pos - ref_pos + 1) / 3)
seqsdf['gene'] = seqsdf['pos'].apply(map_gene_to_pos)
seqsdf = seqsdf.loc[~seqsdf['gene'].isna()]
seqsdf = seqsdf.loc[seqsdf['gene']!='nan']
seqsdf['codon_num'] = seqsdf.apply(compute_codon_num, args=(GENE2POS,), axis=1)
def get_ref_codon(x, ref_seq, gene2pos: dict):
ref_pos = gene2pos[x['gene']]['start']
codon_start = ref_pos + ((x['codon_num'] - 1) * 3)
return ref_seq[codon_start: codon_start+3].upper()
seqsdf['ref_codon'] = seqsdf.apply(get_ref_codon, args=(ref_seq, GENE2POS), axis=1)
def get_alt_codon(x, gene2pos: dict):
ref_pos = gene2pos[x['gene']]['start']
codon_start = ref_pos + ((x['codon_num'] - 1) * 3)
return x['sequence'][codon_start: codon_start+3].upper()
seqsdf['alt_codon'] = seqsdf.apply(get_alt_codon, args=(GENE2POS,), axis=1)
def get_aa(codon: str):
CODON2AA = {
'ATA':'I', 'ATC':'I', 'ATT':'I', 'ATG':'M',
'ACA':'T', 'ACC':'T', 'ACG':'T', 'ACT':'T',
'AAC':'N', 'AAT':'N', 'AAA':'K', 'AAG':'K',
'AGC':'S', 'AGT':'S', 'AGA':'R', 'AGG':'R',
'CTA':'L', 'CTC':'L', 'CTG':'L', 'CTT':'L',
'CCA':'P', 'CCC':'P', 'CCG':'P', 'CCT':'P',
'CAC':'H', 'CAT':'H', 'CAA':'Q', 'CAG':'Q',
'CGA':'R', 'CGC':'R', 'CGG':'R', 'CGT':'R',
'GTA':'V', 'GTC':'V', 'GTG':'V', 'GTT':'V',
'GCA':'A', 'GCC':'A', 'GCG':'A', 'GCT':'A',
'GAC':'D', 'GAT':'D', 'GAA':'E', 'GAG':'E',
'GGA':'G', 'GGC':'G', 'GGG':'G', 'GGT':'G',
'TCA':'S', 'TCC':'S', 'TCG':'S', 'TCT':'S',
'TTC':'F', 'TTT':'F', 'TTA':'L', 'TTG':'L',
'TAC':'Y', 'TAT':'Y', 'TAA':'_', 'TAG':'_',
'TGC':'C', 'TGT':'C', 'TGA':'_', 'TGG':'W',
}
return CODON2AA.get(codon, 'nan')
seqsdf['ref_aa'] = seqsdf['ref_codon'].apply(get_aa)
seqsdf['alt_aa'] = seqsdf['alt_codon'].apply(get_aa)
seqsdf = seqsdf.loc[seqsdf['alt_aa']!='nan']
seqsdf.columns
meta = pd.read_csv(meta_fp)
print(seqsdf['idx'].unique().shape)
seqsdf = pd.merge(seqsdf, meta, left_on='idx', right_on='fasta_hdr')
print(seqsdf['idx'].unique().shape)
seqsdf = seqsdf.loc[(seqsdf['collection_date']!='Unknown')
& (seqsdf['collection_date']!='1900-01-00')]
seqsdf.loc[seqsdf['collection_date'].str.contains('/'), 'collection_date'] = seqsdf['collection_date'].apply(lambda x: x.split('/')[0])
seqsdf['date'] = pd.to_datetime(seqsdf['collection_date'])
seqsdf['date'].min()
# (seqsdf.groupby(['gene', 'ref_aa', 'codon_num', 'alt_aa'])
# .agg(
# num_samples=('ID', 'nunique')))
def uniq_locs(x):
return np.unique(x)
def loc_counts(x):
_, counts = np.unique(x, return_counts=True)
return counts
subs = (seqsdf.groupby(['gene', 'pos', 'ref_aa', 'codon_num', 'alt_aa'])
.agg(
num_samples=('ID', 'nunique'),
first_detected=('date', 'min'),
last_detected=('date', 'max'),
locations=('location', uniq_locs),
location_counts=('location', loc_counts),
samples=('ID', 'unique')
)
.reset_index())
subs['pos'] = subs['pos'] + 1
(subs[subs['gene']=='S'].sort_values('num_samples', ascending=False)
.to_csv('S_mutations_consensus.csv', index=False))
```
## Consolidate metadata ID and fasta headers
```
def fix_header(x):
if 'Consensus' in x:
return x.split('_')[1]
else:
return x.split('/')[2]
seqsdf['n_ID'] = seqsdf['idx'].apply(fix_header)
seqsdf['n_ID'] = seqsdf['n_ID'].str.replace('ALSR', 'SEARCH')
meta = pd.read_csv(meta_fp)
meta['n_ID'] = meta['ID'].apply(lambda x: '-'.join(x.split('-')[:2]))
seqsdf['n_ID'] = seqsdf['n_ID'].apply(lambda x: '-'.join(x.split('-')[:2]))
tmp = pd.merge(seqsdf, meta, on='n_ID')
# tmp[tmp['ID'].str.contains('2112')]
# seqsdf
set(meta['n_ID'].unique()) - set(tmp['n_ID'].unique())
seqsdf['idx'].unique().shape
meta['ID'].unique().shape
s = seqsdf[['n_ID', 'idx']].drop_duplicates()
new_meta = pd.merge(meta, s, on='n_ID', how='left')
(new_meta.drop(columns=['n_ID'])
.rename(columns={'idx': 'fasta_hdr'})
.to_csv('metadata.csv', index=False))
new_meta.shape
new_meta
len(ref_seq)
```
| github_jupyter |
# Unit 12 - Tales from the Crypto
---
## 1. Sentiment Analysis
Use the [newsapi](https://newsapi.org/) to pull the latest news articles for Bitcoin and Ethereum and create a DataFrame of sentiment scores for each coin.
Use descriptive statistics to answer the following questions:
1. Which coin had the highest mean positive score?
2. Which coin had the highest negative score?
3. Which coin had the highest positive score?
```
# Initial imports
import os
from pathlib import Path
import pandas as pd
from dotenv import load_dotenv
import nltk as nltk
from newsapi import NewsApiClient
nltk.download('vader_lexicon')
from nltk.sentiment.vader import SentimentIntensityAnalyzer
analyzer = SentimentIntensityAnalyzer()
%matplotlib inline
# Read your api key environment variable
# YOUR CODE HERE!
api_key = os.getenv("news_api")
# Create a newsapi client
# YOUR CODE HERE!
newsapi = NewsApiClient(api_key=api_key)
# Fetch the Bitcoin news articles
# YOUR CODE HERE!
btc_headlines = newsapi.get_everything(
q="bitcoin",
language="en",
page_size=100,
sort_by="relevancy"
)
# Print total articles
print(f"Total articles about Bitcoin: {btc_headlines['totalResults']}")
# Show sample article
btc_headlines["articles"][0]
# Fetch the Ethereum news articles
# YOUR CODE HERE!
eth_headlines = newsapi.get_everything(
q="ethereum",
language="en",
page_size=100,
sort_by="relevancy"
)
# Print total articles
print(f"Total articles about Ethereum: {eth_headlines['totalResults']}")
# Show sample article
eth_headlines["articles"][0]
# Create the Bitcoin sentiment scores DataFrame
# YOUR CODE HERE!
btc_sentiments = []
for article in btc_headlines["articles"]:
try:
text = article["content"]
date = article["publishedAt"][:10]
sentiment = analyzer.polarity_scores(text)
compound = sentiment["compound"]
pos = sentiment["pos"]
neu = sentiment["neu"]
neg = sentiment["neg"]
btc_sentiments.append({
"text": text,
"date": date,
"compound": compound,
"positive": pos,
"negative": neg,
"neutral": neu
})
except AttributeError:
pass
# Create DataFrame
btc_df = pd.DataFrame(btc_sentiments)
# Reorder DataFrame columns
cols = ["date", "text", "compound", "positive", "negative", "neutral"]
btc_df = btc_df[cols]
btc_df.head()
# Create the Ethereum sentiment scores DataFrame
# YOUR CODE HERE!
eth_sentiments = []
for article in eth_headlines["articles"]:
try:
text = article["content"]
date = article["publishedAt"][:10]
sentiment = analyzer.polarity_scores(text)
compound = sentiment["compound"]
pos = sentiment["pos"]
neu = sentiment["neu"]
neg = sentiment["neg"]
eth_sentiments.append({
"text": text,
"date": date,
"compound": compound,
"positive": pos,
"negative": neg,
"neutral": neu
})
except AttributeError:
pass
# Create DataFrame
eth_df = pd.DataFrame(eth_sentiments)
# Reorder DataFrame columns
cols = ["date", "text", "compound", "positive", "negative", "neutral"]
eth_df = eth_df[cols]
eth_df.head()
# Describe the Bitcoin Sentiment
# YOUR CODE HERE!
btc_df.describe()
# Describe the Ethereum Sentiment
# YOUR CODE HERE!
eth_df.describe()
```
### Questions:
Q: Which coin had the highest mean positive score?
A:
Q: Which coin had the highest compound score?
A:
Q. Which coin had the highest positive score?
A:
```
# 1. BTC had the higher mean positive score
# 2. BTC had the highest compaound score
# 3. BTC had the highest positive score
```
---
## 2. Natural Language Processing
---
### Tokenizer
In this section, you will use NLTK and Python to tokenize the text for each coin. Be sure to:
1. Lowercase each word.
2. Remove Punctuation.
3. Remove Stopwords.
```
from nltk.tokenize import word_tokenize, sent_tokenize
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer, PorterStemmer
from string import punctuation
import re
# Instantiate the lemmatizer
# YOUR CODE HERE!
wnl = WordNetLemmatizer()
# Create a list of stopwords
# YOUR CODE HERE!
stop = stopwords.words('english')
# Expand the default stopwords list if necessary
# YOUR CODE HERE!
stop.append("u")
stop.append("it'")
stop.append("'s'")
stop.append("n't")
stop.append('...')
stop.append('``')
stop.append('char')
stop.append("''")
stop = set(stop)
# Complete the tokenizer function
def tokenizer(text):
"""Tokenizes text."""
# Remove the punctuation from text
words = word_tokenize(text)
# Create a tokenized list of the words
words = list(filter(lambda w: w.lower(), words))
# Lemmatize words into root words
words = list (filter(lambda t: t not in punctuation, words))
# Convert the words to lowercase
words = list(filter(lambda t: t.lower() not in stop, words))
# Remove the stop words
tokens = [wnl.lemmatize(word) for word in words]
return tokens
# Create a new tokens column for Bitcoin
# YOUR CODE HERE!
btc_df['tokens'] = btc_df.text.apply(tokenizer)
btc_df.head()
# Create a new tokens column for Ethereum
# YOUR CODE HERE!
eth_df['tokens'] = eth_df.text.apply(tokenizer)
eth_df.head()
```
---
### NGrams and Frequency Analysis
In this section you will look at the ngrams and word frequency for each coin.
1. Use NLTK to produce the n-grams for N = 2.
2. List the top 10 words for each coin.
```
from collections import Counter
from nltk import ngrams
# Generate the Bitcoin N-grams where N=2
# YOUR CODE HERE!
N = 2
grams = ngrams(tokenizer(btc_df.text.str.cat()), N)
Counter(grams).most_common(20)
# Generate the Ethereum N-grams where N=2
# YOUR CODE HERE!
N = 2
grams = ngrams(tokenizer(eth_df.text.str.cat()), N)
Counter(grams).most_common(20)
# Function token_count generates the top 10 words for a given coin
def token_count(tokens, N=3):
"""Returns the top N tokens from the frequency count"""
return Counter(tokens).most_common(N)
# Use token_count to get the top 10 words for Bitcoin
# YOUR CODE HERE!
all_tokens = tokenizer(btc_df.text.str.cat())
token_count(all_tokens, 10)
# Use token_count to get the top 10 words for Ethereum
# YOUR CODE HERE!
all_tokens = tokenizer(eth_df.text.str.cat())
token_count(all_tokens, 10)
```
---
### Word Clouds
In this section, you will generate word clouds for each coin to summarize the news for each coin
```
from wordcloud import WordCloud
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
import matplotlib as mpl
mpl.rcParams['figure.figsize'] = [20.0, 10.0]
# Generate the Bitcoin word cloud
# YOUR CODE HERE!
def wordcloud(text, title=""):
df_cloud = WordCloud(width=500, colormap='GnBu_r').generate(text)
plt.imshow(df_cloud)
plt.axis("off")
fontdict = {"fontsize":50, "fontweight" : "bold"}
plt.title(title, fontdict=fontdict)
plt.show()
wordcloud(btc_df.text.str.cat(), title="Bitcoin Word Cloud")
# Generate the Ethereum word cloud
# YOUR CODE HERE!
wordcloud(eth_df.text.str.cat(), title="Ethereum Word Cloud")
```
---
## 3. Named Entity Recognition
In this section, you will build a named entity recognition model for both Bitcoin and Ethereum, then visualize the tags using SpaCy.
```
import spacy
from spacy import displacy
# Download the language model for SpaCy
# !python -m spacy download en_core_web_sm
# Load the spaCy model
nlp = spacy.load('en_core_web_sm')
```
---
### Bitcoin NER
```
# Concatenate all of the Bitcoin text together
# YOUR CODE HERE!
all_btc_text = btc_df.text.str.cat()
all_btc_text
# Run the NER processor on all of the text
# YOUR CODE HERE!
doc = nlp(all_btc_text)
# Add a title to the document
# YOUR CODE HERE!
doc.user_data["title"] = "Bitcoin NER"
# Render the visualization
# YOUR CODE HERE!
displacy.render(doc, style='ent', jupyter=True)
# List all Entities
# YOUR CODE HERE!
for ent in doc.ents:
print(ent.text, ent.label_)
```
---
### Ethereum NER
```
# Concatenate all of the Ethereum text together
# YOUR CODE HERE!
all_eth_text = eth_df.text.str.cat()
all_eth_text
# Run the NER processor on all of the text
# YOUR CODE HERE!
doc = nlp(all_eth_text)
# Add a title to the document
# YOUR CODE HERE!
doc.user_data["title"] = "Ethereum NER"
# Render the visualization
# YOUR CODE HERE!
displacy.render(doc, style='ent', jupyter=True)
# List all Entities
# YOUR CODE HERE!
for ent in doc.ents:
print(ent.text, ent.label_)
```
---
| github_jupyter |
```
#Load the modules for data manipulation.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
#Load the module for visualizing NYC.
import geopandas as gpd
#Load the module for suppressing unwanted error messages.
import warnings
warnings.filterwarnings('ignore')
#Adjust visualization settings.
#sns.set_style()
plt.style.use('ggplot')
%matplotlib inline
#Load the Shapefile for NYC into a dataframe. The Shapefile is used to visualize NYC, and was retrieved from NYC Open Data.
street_map = gpd.read_file('geo_export_629b3182-5516-4755-987b-f4106c7d8b99.shp')
#Load the dataset that will be used for this project. Omit unnecessary columns.
shops = pd.read_csv('shops_pre_EDA.csv')
shops = shops.loc[:, 'name':]
#Dunkin' Donuts appears under two different names, which will be fixed now.
shops.loc[shops['name'] == "Dunkin Donuts", 'name'] = "Dunkin' Donuts"
#Function which restricts shops by the number of reviews
def reviews_range(df, minimum, maximum):
"""
Returns the sub-dataframe of df such that minimum <= reviews <= maximum.
"""
return df[df['reviews'].between(minimum, maximum)]
#Visualizing the coffee shop network before filtering out bad points.
plt.style.use('ggplot')
fig, ax = plt.subplots(figsize=(8,8))
ax.set(xlabel='Longitude', ylabel='Latitude', title='Coffee shops in NYC and surrounding region')
street_map.plot(ax=ax, alpha=1, color='grey')
plt.scatter(x='longitude', y='latitude', s=5, data=shops)
ax.patch.set_facecolor('white');
```
Given that we are only interested in NYC coffee shops, the visualization above reveals many points that do not belong in the dataset. Any point which satisfies at least one of the following three bullets needs to be removed:
•The longitude is outside of NYC's longitude range (which can be removed used a longitude filter). <br />
•The latitude is outside of NYC's latitude range (which can be removed used a latitude filter). <br />
•The coordinates are within NYC's longitude/latitude ranges but the point is not in NYC (which can be removed using appropriately-chosen rectangles). <br />
```
#Estimates for the longitude and latitude ranges of NYC, taken directly from Google Maps.
min_longitude_NYC = -74.255610
max_longitude_NYC = -73.700049
min_latitude_NYC = 40.496165
max_latitude_NYC = 40.915577
#Repeated application of the following functions will remove the points described in the third bullet above.
def remove_north_west_rectangle(df, corner_lat, corner_lon):
"""
df is a dataframe with 'longitude' and 'latitude' columns.
All points that are north-west of (corner_lat, corner_lon) will be filtered out of df.
"""
return df[~((df['longitude']<corner_lon) & (df['latitude']>corner_lat))]
def remove_north_east_rectangle(df, corner_lat, corner_lon):
"""
df is a dataframe with 'longitude' and 'latitude' columns.
All points that are north-east of (corner_lat, corner_lon) will be filtered out of df.
"""
return df[~((df['longitude']>corner_lon) & (df['latitude']>corner_lat))]
def remove_south_east_rectangle(df, corner_lat, corner_lon):
"""
df is a dataframe with 'longitude' and 'latitude' columns.
All points that are south-east of (corner_lat, corner_lon) will be filtered out of df.
"""
return df[~((df['longitude']>corner_lon) & (df['latitude']<corner_lat))]
#Lists of coordinates that will be used as inputs for remove_north_west_rectangle(), remove_north_east_rectangle() and remove_south_east_rectangle().
#The coordinates were selected directly from Google Maps in such a way that no points in NYC are removed.
north_west_rectangle_corner = [(40.652578, -74.027496), (40.647619, -74.114938), (40.685464, -74.028357),
(40.707065, -74.020988), (40.739028, -74.013958), (40.757455, -74.010169),
(40.774145, -73.996326), (40.799351, -73.980336), (40.817706, -73.967623),
(40.838019, -73.951854), (40.857452, -73.942018), (40.882348, -73.929711),
(40.908047, -73.916567), (40.910700, -73.890578), (40.800000, -73.970000),
(40.560000, -74.200000)]
north_east_rectangle_corner = [(40.905783, -73.882172), (40.891769, -73.834559), (40.879946, -73.789872),
(40.857882, -73.779249), (40.781774, -73.759754), (40.764539, -73.723468)]
south_east_rectangle_corner = [(40.720591, -73.725117), (40.644738, -73.739240), (40.737579, -73.700744),
(40.731155, -73.704537), (40.731099, -73.704888), (40.729896, -73.705880),
(40.728241, -73.706477)]
#Remove points where the longitude falls outside of the longitude range of NYC or the latitude falls outside of the latitude range of NYC.
shops = shops[shops['longitude'].between(min_longitude_NYC, max_longitude_NYC) &\
shops['latitude'].between(min_latitude_NYC, max_latitude_NYC)]
#Remove rectangles of points within the longitude/latitude ranges of NYC which are not in NYC.
for (lat, lon) in north_west_rectangle_corner:
shops = remove_north_west_rectangle(shops, lat, lon)
for (lat, lon) in north_east_rectangle_corner:
shops = remove_north_east_rectangle(shops, lat, lon)
for (lat, lon) in south_east_rectangle_corner:
shops = remove_south_east_rectangle(shops, lat, lon)
#Reset the row indices of shops to make indexing easier: 0,1,2,...
shops.index = list(range(shops.shape[0]))
#Visualizing the coffee shop network after filtering out bad points.
plt.style.use('ggplot')
fig, ax = plt.subplots(figsize=(8,8))
ax.set(xlabel='Longitude', ylabel='Latitude', title='Coffee shops in NYC')
street_map.plot(ax=ax, alpha=1, color='black')
plt.scatter(x='longitude', y='latitude', s=5, data=shops)
ax.patch.set_facecolor('white');
#Visualizing review counts in increasing order.
reviews_sorted = shops['reviews'].sort_values()
number_shops = len(reviews_sorted)
#Define the indices for the points (0,1,2...), to be used for the x-axis values in the scatterplot.
indices = list(range(number_shops))
fig, ax = plt.subplots(figsize = (6,4))
plt.scatter(indices, reviews_sorted, s=4, color='blue')
plt.xlabel('Shop Index')
plt.ylabel('Number of Reviews')
plt.title("Pareto Principle: Review count follows an exponential distribution")
ax.patch.set_facecolor('white');
#Pareto's Law: the highest 20% of shops by reviews contribute 69% of the total review count across all shops.
shops_lowest_80 = reviews_sorted[:int(0.8*number_shops)]
shops_highest_20 = reviews_sorted[int(0.8*number_shops):]
sum(shops_highest_20) / (sum(shops_highest_20) + sum(shops_lowest_80))
#Summary statistics for the distribution of reviews
shops['reviews'].describe().astype(int)
#Univariate analysis of reviews restricted to reviews<=1000; the continuous analog of a histogram.
#The dataframe for the KDE plot
shops_reviews_1_1000 = reviews_range(shops, 1, 1000)
#Creating the KDE plot.
fig, ax = plt.subplots(1, figsize=(6,4))
sns.kdeplot(shops_reviews_1_1000['reviews'], ax=ax, legend=False, color='blue')
ax.set(xlabel='Number of Reviews', title='KDE Plot (reviews <= 1000): Number of reviews is skewed right')
ax.patch.set_facecolor('white');
#Univariate analysis of price.
#Creating the dataframe for the bar chart.
price_counts = shops['price'].value_counts()
price_counts_df = pd.DataFrame({'price':price_counts.index, 'count':price_counts.values})
#Creating the bar chart.
fig, ax = plt.subplots(1, figsize=(6,4))
ax = sns.barplot(x='price', y='count', data=price_counts_df, order=['x','xx','xxx','xxxx'], color='blue')
ax.set(xlabel='Price Level', ylabel='Number of Shops', title="Most shops are in the two lowest price levels")
ax.patch.set_facecolor('white');
#Univariate analysis of ratings.
#Creating the dataframe for the bar chart.
rating_counts = shops['rating'].value_counts()
rating_counts_df = pd.DataFrame({'rating':rating_counts.index, 'count':rating_counts.values})
#Creating the bar chart.
fig, ax = plt.subplots(1, figsize=(6,4))
ax = sns.barplot(x='rating', y='count', data=rating_counts_df, color='blue')
ax.set(xlabel='Rating', ylabel='Number of Shops', title="Ratings are skewed left")
ax.patch.set_facecolor('white');
#Bivariate analysis of ratings vs. price group.
#Define the relevant dataframes to be used in sns.barplot(). This could also be done using groupby.
#There is one dataframe for each price level (x, xx, xxx, xxxx) and each gives a count for the different values of 'rating'.
rating_counts_x = shops[shops['price']=='x']['rating'].value_counts()
rating_counts_x_df = pd.DataFrame({'rating':rating_counts_x.index, 'count':rating_counts_x.values})
rating_counts_xx = shops[shops['price']=='xx']['rating'].value_counts()
rating_counts_xx_df = pd.DataFrame({'rating':rating_counts_xx.index, 'count':rating_counts_xx.values})
rating_counts_xxx = shops[shops['price']=='xxx']['rating'].value_counts()
rating_counts_xxx_df = pd.DataFrame({'rating':rating_counts_xxx.index, 'count':rating_counts_xxx.values})
rating_counts_xxxx = shops[shops['price']=='xxxx']['rating'].value_counts()
rating_counts_xxxx_df = pd.DataFrame({'rating':rating_counts_xxxx.index, 'count':rating_counts_xxxx.values})
#Create the bar charts.
fig, ax = plt.subplots(1,4,figsize=(24,6))
sns.barplot(x='rating', y='count', data=rating_counts_x_df, ax=ax[0], color='purple')
sns.barplot(x='rating', y='count', data=rating_counts_xx_df, ax=ax[1], color='purple')
sns.barplot(x='rating', y='count', data=rating_counts_xxx_df, ax=ax[2], color='purple')
sns.barplot(x='rating', y='count', data=rating_counts_xxxx_df, ax=ax[3], color='purple')
ax[0].set(xlabel='Rating', ylabel="Number of Shops", title='Price Level: x')
ax[1].set(xlabel='Rating', ylabel="Number of Shops", title='Price Level: xx')
ax[2].set(xlabel='Rating', ylabel="Number of Shops", title='Price Level: xxx')
ax[3].set(xlabel='Rating', ylabel="Number of Shops", title='Price Level: xxxx')
ax[0].patch.set_facecolor('white')
ax[1].patch.set_facecolor('white')
ax[2].patch.set_facecolor('white')
ax[3].patch.set_facecolor('white');
#Bivariate analysis of reviews vs. price.
fig, ax = plt.subplots(1, figsize=(6,4))
sns.boxplot(x='price', y='reviews', data=reviews_range(shops,1,1000), ax=ax, order=['x','xx','xxx','xxxx'], color='purple')
ax.set(xlabel='Price Level', ylabel="Number of Reviews", title='Reviews by price (reviews <= 1000)')
ax.patch.set_facecolor('white');
#Bivariate analysis of reviews vs. rating.
fig, ax = plt.subplots(1, figsize=(9,6))
sns.scatterplot(x='rating', y='reviews', data=shops, color='purple')
ax.set(xlabel='Rating', ylabel="Number of Reviews", title='Higher-rated shops receive more reviews')
ax.patch.set_facecolor('white');
#Multivariate analysis of average reviews grouped by rating and price.
#Build an 11x4 dataframe where each cell corresponds to a unique (rating,price) pair, and the corresponding number of reviews.
reviews_heatmap_df = pd.DataFrame({'rating':[0,1,2,3,4,5,6,7,8,9,10]*4,
'price': ['x']*11 + ['xx']*11 + ['xxx']*11 + ['xxxx']*11,
'mean_reviews': [0]*44,
'median_reviews': [0]*44})
#Fill in the values for mean_reviews and median_reviews in reviews_heatmap_df.
for row in range(44):
price = reviews_heatmap_df.loc[row, 'price']
rating = reviews_heatmap_df.loc[row, 'rating']
shops_filtered = shops.loc[(shops['rating']==rating) & (shops['price']==price)]
mean_reviews = np.mean(shops_filtered['reviews'])
median_reviews = np.median(shops_filtered['reviews'])
reviews_heatmap_df.loc[row, 'mean_reviews'] = mean_reviews
reviews_heatmap_df.loc[row, 'median_reviews'] = median_reviews
#Creating the first heatmap.
fig, ax = plt.subplots(figsize=(12,6))
sns.heatmap(reviews_heatmap_df.pivot("rating", "price", "mean_reviews"), cmap='OrRd',cbar_kws={'label': 'Mean Number of Reviews'})
ax.set(xlabel='Price', ylabel='Rating', title='Expensive, high rated shops have more reviews (by mean)')
ax.invert_yaxis();
#Creating the second heatmap.
fig, ax = plt.subplots(figsize=(12,6))
sns.heatmap(reviews_heatmap_df.pivot("rating", "price", "median_reviews"), cmap='OrRd',cbar_kws={'label': 'Median Number of Reviews'})
ax.set(xlabel='Price Level', ylabel='Rating', title='Expensive, high rated shops have more reviews (by median)')
ax.invert_yaxis();
#The file that is to be loaded for the first version of the simulator.
shops.to_csv('shops_post_EDA.csv', index=False)
```
| github_jupyter |
# PASTIS matrix from E-fields
This notebook calculates PASTIS matrices for the low, mid, and high order modes from single-mode E-fields in the focal plane.
It also calculates matrices on the low order wavefront sensor (LOWFS) and out of band wavefront sensor (OBWFS).
```
import os
import time
from shutil import copy
from astropy.io import fits
import astropy.units as u
import hcipy
import numpy as np
import pastis.util as util
from pastis.config import CONFIG_PASTIS
from pastis.e2e_simulators.luvoir_imaging import LuvoirA_APLC
```
## Initial setup and parameters
Set up data paths for input and output
```
root_dir = CONFIG_PASTIS.get('local', 'local_data_path')
coronagraph_design = 'small' # user provides
overall_dir = util.create_data_path(root_dir, telescope='luvoir_'+coronagraph_design)
resDir = os.path.join(overall_dir, 'matrix_numerical')
print(resDir)
```
Read from configfile how many modes each DM should be able to do
```
max_LO = CONFIG_PASTIS.getint('dm_objects', 'number_of_low_order_modes')
max_MID = CONFIG_PASTIS.getint('dm_objects', 'number_of_mid_order_modes')
max_HI = CONFIG_PASTIS.getint('dm_objects', 'number_of_high_order_modes')
num_DM_act = CONFIG_PASTIS.getint('dm_objects', 'number_of_continuous_dm_actuators')
print(f'max_LO: {max_LO}')
print(f'max_MID: {max_MID}')
print(f'max_HI: {max_HI}')
print(f'num_DM_act: {num_DM_act}')
```
Read some more required parameters from the configfile
```
nb_seg = CONFIG_PASTIS.getint('LUVOIR', 'nb_subapertures')
wvln = CONFIG_PASTIS.getfloat('LUVOIR', 'lambda') * 1e-9 # m
diam = CONFIG_PASTIS.getfloat('LUVOIR', 'diameter') # m
nm_aber = CONFIG_PASTIS.getfloat('LUVOIR', 'calibration_aberration') * 1e-9 # m
sampling = CONFIG_PASTIS.getfloat('LUVOIR', 'sampling')
print('LUVOIR apodizer design: {}'.format(coronagraph_design))
print()
print('Wavelength: {} m'.format(wvln))
print('Telescope diameter: {} m'.format(diam))
print('Number of segments: {}'.format(nb_seg))
print()
print('Sampling: {} px per lambda/D'.format(sampling))
# Create necessary directories if they don't exist yet
os.makedirs(resDir, exist_ok=True)
os.makedirs(os.path.join(resDir, 'OTE_images'), exist_ok=True)
os.makedirs(os.path.join(resDir, 'psfs'), exist_ok=True)
# Copy configfile to resulting matrix directory
util.copy_config(resDir)
# Create LUVOIR simulator instance
optics_input = os.path.join(util.find_repo_location(), CONFIG_PASTIS.get('LUVOIR', 'optics_path_in_repo'))
luvoir = LuvoirA_APLC(optics_input, coronagraph_design, sampling)
```
Generate the deformable mirrors that are required.
**This will take quite some time**
```
luvoir.create_global_zernike_mirror(max_LO)
luvoir.create_segmented_mirror(max_MID)
luvoir.create_ripple_mirror(max_HI)
luvoir.create_continuous_deformable_mirror(num_DM_act)
# Figur out the total number of "actuators" (= effective modes) for each DM
n_LO = luvoir.zernike_mirror.num_actuators
n_MID = luvoir.sm.num_actuators
n_HI = luvoir.ripple_mirror.num_actuators
n_DM = luvoir.dm.num_actuators
print(f'n_LO: {n_LO}')
print(f'n_MID: {n_MID}')
print(f'n_HI: {n_HI}')
print(f'n_DM: {n_DM}')
```
Define some parameters that are needed for the subsampled LOWFS output.
```
z_pup_downsample = CONFIG_PASTIS.getfloat('numerical', 'z_pup_downsample')
N_pup_z = int(luvoir.pupil_grid.shape[0] / z_pup_downsample)
grid_zernike = hcipy.field.make_pupil_grid(N_pup_z, diameter=luvoir.diam)
```
## Flatten all DMs and create unaberrated reference PSF
```
LO_modes = np.zeros(n_LO)
MID_modes = np.zeros(n_MID)
HI_modes = np.zeros(n_HI)
DM_modes = np.zeros(n_DM)
luvoir.zernike_mirror.actuators = LO_modes
luvoir.sm.actuators = MID_modes
luvoir.ripple_mirror.actuators = HI_modes
luvoir.dm.actuators = DM_modes
# Calculate the unaberrated coro and direct PSFs in INTENSITY
unaberrated_coro_psf, ref = luvoir.calc_psf(ref=True, display_intermediate=False)
# Define the normalization factor for contrast units
norm = np.max(ref)
# Calculate the unaberrated coro and direct PSFs in E-FIELDS
nonaberrated_coro_psf, ref, efield = luvoir.calc_psf(ref=True, display_intermediate=False, return_intermediate='efield')
Efield_ref = nonaberrated_coro_psf.electric_field
```
## Generate LOW-order PASTIS matrix from E-fields
```
print('Generating the E-fields for LOW modes in science plane')
print(f'number of LO modes: {n_LO}')
# Create lists that will hold measured fields
print(f'Calibration aberration used: {nm_aber} m')
start_time = time.time()
focus_fieldS = []
focus_fieldS_Re = []
focus_fieldS_Im = []
for pp in range(0, n_LO):
print(f'Working on mode {pp}/{n_LO}')
# Apply calibration aberration to used mode
LO_modes = np.zeros(n_LO)
LO_modes[pp] = nm_aber / 2
luvoir.zernike_mirror.actuators = LO_modes
# Calculate coronagraphic E-field and add to lists
aberrated_coro_psf, inter = luvoir.calc_psf(display_intermediate=False, return_intermediate='efield')
focus_field1 = aberrated_coro_psf
focus_fieldS.append(focus_field1)
focus_fieldS_Re.append(focus_field1.real)
focus_fieldS_Im.append(focus_field1.imag)
# Construct the PASTIS matrix from the E-fields
mat_fast = np.zeros([n_LO, n_LO]) # create empty matrix
for i in range(0, n_LO):
for j in range(0, n_LO):
test = np.real((focus_fieldS[i].electric_field - Efield_ref) * np.conj(focus_fieldS[j].electric_field - Efield_ref))
dh_test = (test / norm) * luvoir.dh_mask
contrast = np.mean(dh_test[np.where(luvoir.dh_mask != 0)])
mat_fast[i, j] = contrast
# Normalize by the calibration aberration
matrix_pastis = np.copy(mat_fast)
matrix_pastis /= np.square(nm_aber * 1e9)
```
Save results out to disk
```
filename_matrix = 'PASTISmatrix_num_LO_' + str(max_LO)
hcipy.write_fits(matrix_pastis, os.path.join(resDir, filename_matrix + '.fits'))
print('Matrix saved to:', os.path.join(resDir, filename_matrix + '.fits'))
filename_matrix = 'EFIELD_Re_matrix_num_LO_' + str(max_LO)
hcipy.write_fits(focus_fieldS_Re, os.path.join(resDir, filename_matrix + '.fits'))
print('Efield Real saved to:', os.path.join(resDir, filename_matrix + '.fits'))
filename_matrix = 'EFIELD_Im_matrix_num_LO_' + str(max_LO)
hcipy.write_fits(focus_fieldS_Im, os.path.join(resDir, filename_matrix + '.fits'))
print('Efield Imag saved to:', os.path.join(resDir, filename_matrix + '.fits'))
end_time = time.time()
print('Runtime for LO modes:', end_time - start_time, 'sec =', (end_time - start_time) / 60, 'min')
print('Data saved to {}'.format(resDir))
```
## Generate MID-order PASTIS matrix from E-fields
```
print('Generating the Efield for MID modes in science plane')
print(f'number of MID modes: {n_MID}')
LO_modes = np.zeros(n_LO)
MID_modes = np.zeros(n_MID)
HI_modes = np.zeros(n_HI)
# Create lists that will hold measured fields
print(f'Calibration aberration used: {nm_aber} m')
start_time = time.time()
focus_fieldS = []
focus_fieldS_Re = []
focus_fieldS_Im = []
for pp in range(0, n_MID):
print(f'Working on mode {pp}/{n_MID}')
# Apply calibration aberration to used mode
MID_modes = np.zeros(n_MID)
MID_modes[pp] = nm_aber / 2
luvoir.sm.actuators = MID_modes
# Calculate coronagraphic E-field and add to lists
aberrated_coro_psf, inter = luvoir.calc_psf(display_intermediate=False, return_intermediate='efield')
focus_field1 = aberrated_coro_psf
focus_fieldS.append(focus_field1)
focus_fieldS_Re.append(focus_field1.real)
focus_fieldS_Im.append(focus_field1.imag)
# Construct the PASTIS matrix from the E-fields
mat_fast = np.zeros([n_MID, n_MID])
for i in range(0, n_MID):
for j in range(0, n_MID):
test = np.real((focus_fieldS[i].electric_field - Efield_ref) * np.conj(focus_fieldS[j].electric_field - Efield_ref))
dh_test = (test / norm) * luvoir.dh_mask
contrast = np.mean(dh_test[np.where(luvoir.dh_mask != 0)])
mat_fast[i, j] = contrast
# Normalize by the calibration aberration
matrix_pastis = np.copy(mat_fast)
matrix_pastis /= np.square(nm_aber * 1e9)
filename_matrix = 'PASTISmatrix_num_MID_' + str(max_MID)
hcipy.write_fits(matrix_pastis, os.path.join(resDir, filename_matrix + '.fits'))
print('Matrix saved to:', os.path.join(resDir, filename_matrix + '.fits'))
filename_matrix = 'EFIELD_Re_matrix_num_MID_' + str(max_MID)
hcipy.write_fits(focus_fieldS_Re, os.path.join(resDir, filename_matrix + '.fits'))
print('Efield Real saved to:', os.path.join(resDir, filename_matrix + '.fits'))
filename_matrix = 'EFIELD_Im_matrix_num_MID_' + str(max_MID)
hcipy.write_fits(focus_fieldS_Im, os.path.join(resDir, filename_matrix + '.fits'))
print('Efield Imag saved to:', os.path.join(resDir, filename_matrix + '.fits'))
end_time = time.time()
print('Runtime for MID modes:', end_time - start_time, 'sec =', (end_time - start_time) / 60, 'min')
print('Data saved to {}'.format(resDir))
```
## Generate MID-order PASTIS matrix from E-fields
```
print('Generating the Efield for HI modes in science plane')
print(f'number of HI modes: {n_HI}')
LO_modes = np.zeros(n_LO)
MID_modes = np.zeros(n_MID)
HI_modes = np.zeros(n_HI)
# Create lists that will hold measured fields
print(f'Calibration aberration used: {nm_aber} m')
start_time = time.time()
focus_fieldS = []
focus_fieldS_Re = []
focus_fieldS_Im = []
for pp in range(0, n_HI):
print(f'Working on mode {pp}/{n_HI}')
# Apply calibration aberration to used mode
HI_modes = np.zeros(n_HI)
HI_modes[pp] = nm_aber / 2
luvoir.ripple_mirror.actuators = HI_modes
# Calculate coronagraphic E-field and add to lists
aberrated_coro_psf, inter = luvoir.calc_psf(display_intermediate=False, return_intermediate='efield')
focus_field1 = aberrated_coro_psf
focus_fieldS.append(focus_field1)
focus_fieldS_Re.append(focus_field1.real)
focus_fieldS_Im.append(focus_field1.imag)
# Construct the PASTIS matrix from the E-fieldsmat_fast = np.zeros([n_HI, n_HI])
for i in range(0, n_HI):
for j in range(0, n_HI):
test = np.real((focus_fieldS[i].electric_field - Efield_ref) * np.conj(focus_fieldS[j].electric_field - Efield_ref))
dh_test = (test / norm) * luvoir.dh_mask
contrast = np.mean(dh_test[np.where(luvoir.dh_mask != 0)])
mat_fast[i, j] = contrast
# Normalize by the calibration aberration
matrix_pastis = np.copy(mat_fast)
matrix_pastis /= np.square(nm_aber * 1e9)
filename_matrix = 'PASTISmatrix_num_HI_' + str(max_HI)
hcipy.write_fits(matrix_pastis, os.path.join(resDir, filename_matrix + '.fits'))
print('Matrix saved to:', os.path.join(resDir, filename_matrix + '.fits'))
filename_matrix = 'EFIELD_Re_matrix_num_HI_' + str(max_HI)
hcipy.write_fits(focus_fieldS_Re, os.path.join(resDir, filename_matrix + '.fits'))
print('Efield Real saved to:', os.path.join(resDir, filename_matrix + '.fits'))
filename_matrix = 'EFIELD_Im_matrix_num_HI_' + str(max_HI)
hcipy.write_fits(focus_fieldS_Im, os.path.join(resDir, filename_matrix + '.fits'))
print('Efield Imag saved to:', os.path.join(resDir, filename_matrix + '.fits'))
end_time = time.time()
print('Runtime for HI modes:', end_time - start_time, 'sec =', (end_time - start_time) / 60, 'min')
print('Data saved to {}'.format(resDir))
```
## Generate LOW-order matrix on LOWFS
```
print('Generating the Efield for LOW modes through LOWFS')
print('number of LO modes'.format(n_LO))
```
Flatten DMs
```
LO_modes = np.zeros(n_LO)
MID_modes = np.zeros(n_MID)
HI_modes = np.zeros(n_HI)
luvoir.zernike_mirror.actuators = LO_modes
luvoir.sm.actuators = MID_modes
luvoir.ripple_mirror.actuators = HI_modes
zernike_ref = luvoir.calc_low_order_wfs()
```
Calculate unaberrated reference E-field on Zernike WFS on a subsampled grid.
```
zernike_ref_sub_real = hcipy.field.subsample_field(zernike_ref.real, z_pup_downsample, grid_zernike, statistic='mean')
zernike_ref_sub_imag = hcipy.field.subsample_field(zernike_ref.imag, z_pup_downsample, grid_zernike, statistic='mean')
Efield_ref = zernike_ref_sub_real + 1j*zernike_ref_sub_imag
# Create lists that will hold measured fields
print(f'Calibration aberration used: {nm_aber} m')
start_time = time.time()
focus_fieldS = []
focus_fieldS_Re = []
focus_fieldS_Im = []
for pp in range(0, n_LO):
print(f'Working on mode {pp}/{n_LO}')
# Apply calibration aberration to used mode
LO_modes = np.zeros(n_LO)
LO_modes[pp] = nm_aber / 2
luvoir.zernike_mirror.actuators = LO_modes
# Calculate E-field on Zernike WFS and add to lists
zernike_meas = luvoir.calc_low_order_wfs()
zernike_meas_sub_real = hcipy.field.subsample_field(zernike_meas.real, z_pup_downsample, grid_zernike,statistic='mean')
zernike_meas_sub_imag = hcipy.field.subsample_field(zernike_meas.imag, z_pup_downsample, grid_zernike,statistic='mean')
focus_field1 = zernike_meas_sub_real + 1j * zernike_meas_sub_imag
focus_fieldS.append(focus_field1)
focus_fieldS_Re.append(focus_field1.real)
focus_fieldS_Im.append(focus_field1.imag)
filename_matrix = 'EFIELD_LOWFS_Re_matrix_num_LO_' + str(max_LO)
hcipy.write_fits(focus_fieldS_Re, os.path.join(resDir, filename_matrix + '.fits'))
print('Efield Real saved to:', os.path.join(resDir, filename_matrix + '.fits'))
filename_matrix = 'EFIELD_LOWFS_Im_matrix_num_LO_' + str(max_LO)
hcipy.write_fits(focus_fieldS_Im, os.path.join(resDir, filename_matrix + '.fits'))
print('Efield Imag saved to:', os.path.join(resDir, filename_matrix + '.fits'))
end_time = time.time()
print('Runtime for LO modes and LOWFS:', end_time - start_time, 'sec =', (end_time - start_time) / 60, 'min')
print('Data saved to {}'.format(resDir))
```
## Generate MID-order matrix on OBWFS
```
print('Generating the Efield for MID modes to OBWFS')
print('number of MID modes'.format(n_MID))
# Flatten DMs
LO_modes = np.zeros(n_LO)
MID_modes = np.zeros(n_MID)
HI_modes = np.zeros(n_HI)
luvoir.zernike_mirror.actuators = LO_modes
luvoir.sm.actuators = MID_modes
luvoir.ripple_mirror.actuators = HI_modes
# Calculate unaberrated reference E-field on Zernike WFS on a subsampled grid.
zernike_ref = luvoir.calc_out_of_band_wfs()
zernike_ref_sub_real = hcipy.field.subsample_field(zernike_ref.real, z_pup_downsample, grid_zernike, statistic='mean')
zernike_ref_sub_imag = hcipy.field.subsample_field(zernike_ref.imag, z_pup_downsample, grid_zernike, statistic='mean')
Efield_ref = zernike_ref_sub_real + 1j*zernike_ref_sub_imag
# Create lists that will hold measured fields
print(f'Calibration aberration used: {nm_aber} m')
start_time = time.time()
focus_fieldS = []
focus_fieldS_Re = []
focus_fieldS_Im = []
for pp in range(0, n_MID):
print(f'Working on mode {pp}/{n_MID}')
# Apply calibration aberration to used mode
MID_modes = np.zeros(n_MID)
MID_modes[pp] = nm_aber / 2
luvoir.sm.actuators = MID_modes
# Calculate E-field on OBWFS and add to lists
zernike_meas = luvoir.calc_out_of_band_wfs()
zernike_meas_sub_real = hcipy.field.subsample_field(zernike_meas.real, z_pup_downsample, grid_zernike, statistic='mean')
zernike_meas_sub_imag = hcipy.field.subsample_field(zernike_meas.imag, z_pup_downsample, grid_zernike, statistic='mean')
focus_field1 = zernike_meas_sub_real + 1j * zernike_meas_sub_imag
focus_fieldS.append(focus_field1)
focus_fieldS_Re.append(focus_field1.real)
focus_fieldS_Im.append(focus_field1.imag)
filename_matrix = 'EFIELD_OBWFS_Re_matrix_num_MID_' + str(max_MID)
hcipy.write_fits(focus_fieldS_Re, os.path.join(resDir, filename_matrix + '.fits'))
print('Efield Real saved to:', os.path.join(resDir, filename_matrix + '.fits'))
filename_matrix = 'EFIELD_OBWFS_Im_matrix_num_MID_' + str(max_MID)
hcipy.write_fits(focus_fieldS_Im, os.path.join(resDir, filename_matrix + '.fits'))
print('Efield Imag saved to:', os.path.join(resDir, filename_matrix + '.fits'))
end_time = time.time()
print('Runtime for MID modes and OBWFS:', end_time - start_time, 'sec =', (end_time - start_time) / 60, 'min')
print('Data saved to {}'.format(resDir))
```
| github_jupyter |
# Image analysis with fMRI 3D images imported with LORIS API
This is a tutorial to show how to use Loris' API to download MRI images. It also contains a few examples of how the data can be used to run basic data analysis.
This tutorial is also available as a Google colab notebook so you can run it directly from your browser. To access it, click on the button below: <a href="https://colab.research.google.com/github/spell00/Loris/blob/2020-08-06-JupyterCreateImageDataset/docs/notebooks/LORIS-API_Part3-Create_image_dataset.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
# Uncomment and run to install the packages required to run the notebook
# !pip3 install tqdm
# !pip3 install numpy
# !pip3 install nibabel
# !pip3 install sklearn
# !pip3 install matplotlib
# !pip3 install nilearn
```
## Setup
```
import getpass # For input prompt not to show what is entered
import json # Provide convenient functions to handle json objects
import re # For regular expression
import requests # To handle http requests
import nibabel as nib
import numpy as np
import warnings
from tqdm import tqdm_notebook as tqdm # To make a nice progress bar
import os
import itertools
os.chdir('..')
warnings.simplefilter('ignore') # Because I am using unverified ssl certificates
def prettyPrint(string):
print(json.dumps(string, indent=2, sort_keys=True))
import argparse
import torch
import torch.nn as nn
import numpy as np
import json
from torch.utils.data import DataLoader
from tensorboardX import SummaryWriter
from fmri.utils.activations import Swish, Mish
from fmri.utils.CycleAnnealScheduler import CycleScheduler
from fmri.utils.dataset import load_checkpoint, save_checkpoint, MRIDataset
from fmri.utils.transform_3d import Normalize, RandomRotation3D, ColorJitter3D, Flip90, Flip180, Flip270, XFlip, YFlip, \
ZFlip
from fmri.models.supervised.MLP import MLP
from fmri.utils.plot_performance import plot_performance
import torchvision
from torchvision import transforms
from ax.service.managed_loop import optimize
import random
import nibabel as nib
from fmri.utils.utils import validation_spliter
import nilearn.plotting as nlplt
```
## Getting the data
The data on https://demo.loris.ca are only for development purposes. Nevertheless, with this in mind, we will use it for demonstration purposes only. In this tutorial, we will download all the T1 and T2 raw images from every project.
```
images_path = 'D:\\workbench\\projects\\AutoTKV_MouseMRI-master\\AllTrainingImages\\images\\'
targets_path = 'D:\\workbench\\projects\\AutoTKV_MouseMRI-master\\AllTrainingImages\\targets\\'
all_set = MRIDataset(images_path, targets_path, transform=None, resize=False)
spliter = validation_spliter(all_set, cv=5)
valid_set, train_set = spliter.__next__()
train_loader = DataLoader(train_set,
num_workers=0,
shuffle=True,
batch_size=1,
pin_memory=False,
drop_last=True)
valid_loader = DataLoader(valid_set,
num_workers=0,
shuffle=True,
batch_size=1,
pin_memory=False,
drop_last=True)
sample_x, sample_target = next(iter(valid_set))
sample_x = sample_x.numpy().squeeze()
sample_target = sample_target.numpy().squeeze()
np.round(sample.shape) / 2
def np_to_nifti(sample):
coords = np.round(sample.shape) / 2
t1_fullimage = nib.Nifti1Image(sample_x, np.eye(4))
return nlplt.plot_anat(t1_fullimage, (128, 128, 10))
np_to_nifti(sample_x)
np_to_nifti(sample_target)
def _resize_data(data, new_size=(160, 160, 160)):
initial_size_x = data.shape[0]
initial_size_y = data.shape[1]
initial_size_z = data.shape[2]
new_size_x = new_size[0]
new_size_y = new_size[1]
new_size_z = new_size[2]
delta_x = initial_size_x / new_size_x
delta_y = initial_size_y / new_size_y
delta_z = initial_size_z / new_size_z
new_data = np.zeros((new_size_x, new_size_y, new_size_z))
for x, y, z in itertools.product(range(new_size_x),
range(new_size_y),
range(new_size_z)):
new_data[x][y][z] = data[int(x * delta_x)][int(y * delta_y)][int(z * delta_z)]
return new_data
sample_x_14x14x14 = _resize_data(sample_x, (14, 14, 14))
t1_fullimage = nib.Nifti1Image(sample_x_14x14x14, np.eye(4))
nlplt.plot_anat(t1_fullimage, (7, 7, 7))
nlplt.show()
training_images_dir = 'D:\workbench\projects\AutoTKV_MouseMRI-master\AllTrainingImages\images'
training_targets_dir = 'D:\workbench\projects\AutoTKV_MouseMRI-master\AllTrainingImages\targets'
```
#### Then, we get the information necessary to retrieve all images from all the projects and store them in a dictionnary.
```
# The dictionary to store the images
images_dict = {
"raw": {
't1': [],
't2': []
},
"32x32x32": {
't1': [],
't2': []
},
"128x128x128": {
't1': [],
't2': []
}
}
# Progress bar for downloads
pbar = tqdm(total=sum([len([meta for meta in imagesMeta[p]['Images'] if meta['ScanType'] in ['t1', 't2']]) for p in projectnames]))
for project in projectnames:
for i, meta in enumerate(imagesMeta[project]['Images']):
if(meta['ScanType'] not in ['t1', 't2']):
continue
r = requests.get(baseurl + meta['Link'],
headers = {'Authorization': 'Bearer %s' % token})
page = r.content
filename = meta['Link'].split('/')[-1]
t = meta['ScanType']
# The images need to be saved first.
# Only t1 and t2 images are kept.
if (t in ['t1', 't2']):
file_ = open(filename, 'wb')
else:
continue
file_.write(page)
file_.close()
img = nib.load(filename)
# The images are not necessary for the rest of this tutorial.
os.remove(filename)
img = img.get_fdata()
# The images are save in the dictionary
if(meta['ScanType'] == 't1'):
images_dict["raw"]["t1"] += [img]
if(meta['ScanType'] == 't2'):
images_dict["raw"]["t2"] += [img]
pbar.update(1)
```
## Preprocessing
In this section, we'll explore a few preprocessing methods that might make the models learned perform better.
### Resize images
In this tutorial, T1 and T2 images are compared. They are of similar sizes (160x256x224 and 160x256x256 for T1 and T2, respectively), but they need to be exactly the same size for any subsequent analysis.
In machine learning, it is common practice to reduce large images before training a model. Large images have the advantage of containing more information, but it comes with a tradeoff known as the Curse of dimensionality. Having a high dimensionality can make it much easier to have good performances on the training set, but the models trained overfit more easily to the training data and perform poorly on the validation and test data.
Of course, reducing images too much will also harm the performance of the model trained. There is no rule of thumb or algorithm to get the optimal size of images to be used in a specific task, so it might be a good idea to try a few different reductions.
This tutorial will explore 2 dimensions. Both will cubes (all sides have the same length): 128x128x128 and 32x32x32. The later dimensions might be a huge reduction, but the 3D images still have 32,768 dimensions (each voxel being a dimension), which is still huge, but much more manageable than the larger reduction, which has 2,097,152 dimensions. In order to decide which reduction to use, we will observe the data using a Principal Component Analysis (PCA). It will give an idea of whether the data has lost too much information to use it in a classification task.
Ultimately, it might be necessary to use both strategies to test if one is better than the other. In case both strategies appear to be equal, Ockham's razor principle suggest the images with fewer voxels should be used. In this case, the notion of equality is somewhat arbitrary and might depend on the task to be accomplished.
```
def resize_image(image, new_size=(160, 160, 160)):
"""
Function to resize an image.
Args:
image (Numpy array of shape (Length, Width, Depth)): image to transform
new_size (3-Tuple) : The new image length, width and Depth
"""
initial_size_x = image.shape[0]
initial_size_y = image.shape[1]
initial_size_z = image.shape[2]
new_size_x = new_size[0]
new_size_y = new_size[1]
new_size_z = new_size[2]
delta_x = initial_size_x / new_size_x
delta_y = initial_size_y / new_size_y
delta_z = initial_size_z / new_size_z
new_image = np.zeros((new_size_x, new_size_y, new_size_z))
for x, y, z in itertools.product(range(new_size_x),
range(new_size_y),
range(new_size_z)):
new_image[x][y][z] = image[int(x * delta_x)][int(y * delta_y)][int(z * delta_z)]
return new_image
```
We need to create new directeories to save the resized T1 and T2 images.
#### Resize and normalize all T1 images
```
from sklearn.preprocessing import Normalizer
pbar = tqdm(total=len(images_dict['raw']['t1']))
for t1 in images_dict['raw']["t1"]:
t1_32 = resize_image(t1, (32, 32, 32))
t1_32 = Normalizer().fit_transform(t1_32.reshape([1, -1]))
t1_32 = t1_32.reshape([-1, 32, 32, 32])
images_dict['32x32x32']['t1'] += [t1_32]
t1_128 = resize_image(t1, (128, 128, 128))
t1_128 = Normalizer().fit_transform(t1_128.reshape([1, -1]))
t1_128 = t1_128.reshape([-1, 128, 128, 128])
images_dict['128x128x128']['t1'] += [t1_128]
pbar.update(1)
"""
We don't need to save the images for this tutorial, but the package nibabel
can be used to save the images to disk like this:
img = nib.Nifti1Image(image_to_save, np.eye(4))
img.to_filename("/path/to/new_file_name.nii")
"""
# Make numpy arrays from the lists of numpy arrays
images_dict['32x32x32']['t1'] = np.stack(images_dict['32x32x32']['t1'])
images_dict['128x128x128']['t1'] = np.stack(images_dict['128x128x128']['t1'])
```
#### Resize and normalize T2 images
```
pbar = tqdm(total=len(images_dict['raw']['t2']))
for t2 in images_dict['raw']["t2"]:
t2_32 = resize_image(t2, (32, 32, 32))
t2_32 = Normalizer().fit_transform(t2_32.reshape([1, -1]))
t2_32 = t2_32.reshape([-1, 32, 32, 32])
images_dict['32x32x32']['t2'] += [t2_32]
t2_128 = resize_image(t2, (128, 128, 128))
t2_128 = Normalizer().fit_transform(t2_128.reshape([1, -1]))
t2_128 = t2_128.reshape([-1, 128, 128, 128])
images_dict['128x128x128']['t2'] += [t2_128]
pbar.update(1)
# Make numpy arrays from the lists of numpy arrays
images_dict['32x32x32']['t2'] = np.stack(images_dict['32x32x32']['t2'])
images_dict['128x128x128']['t2'] = np.stack(images_dict['128x128x128']['t2'])
```
### Visualisation with nilearn
Visualisation of the raw images and the 2 reductions for T1 and T2 images.
#### T1 images
```
# This package is used to plot a section of the 3D images
import nilearn.plotting as nlplt
print("Original (160x256x224)")
t1_fullimage = nib.Nifti1Image(images_dict['raw']['t1'][0], np.eye(4))
nlplt.plot_anat(t1_fullimage, (80, 128, 112))
nlplt.show()
print("128x128x128")
img_t1_128 = nib.Nifti1Image(resize_image(images_dict['raw']['t1'][0], (128, 128, 128)), np.eye(4))
nlplt.plot_anat(img_t1_128, (64, 64, 64))
nlplt.show()
print("32x32x32")
img_t1_32 = nib.Nifti1Image(resize_image(images_dict['raw']['t1'][0], (32, 32, 32)), np.eye(4))
nlplt.plot_anat(img_t1_32, (16, 16, 16))
nlplt.show()
```
#### T2 images
```
print("Original (160x256x256)")
t2_fullimage = nib.Nifti1Image(images_dict['raw']['t2'][0], np.eye(4))
nlplt.plot_anat(t2_fullimage, (80, 128, 112))
nlplt.show()
print("128x128x128")
img_t2_128 = nib.Nifti1Image(resize_image(images_dict['raw']['t2'][0], (128, 128, 128)), np.eye(4))
nlplt.plot_anat(img_t2_128, (64, 64, 64))
nlplt.show()
print("32x32x32")
img_t2_32 = nib.Nifti1Image(resize_image(images_dict['raw']['t2'][0], (32, 32, 32)), np.eye(4))
nlplt.plot_anat(img_t2_32, (16, 16, 16))
nlplt.show()
```
## Unsupervised learning: Principal Component Analysis
Principal Component Analysis (PCA) is a popular method used for dimensioanlity reduction, which is a good first step to vizualise the data to analyse and can give insight for the subsequent steps of the analysis. Dimensionality reduction can also be used to transform the data before using it to train a ML model.
```
# sklearn needs the data to be flattened
images_dict['32x32x32']['t1'] = images_dict['32x32x32']['t1'].reshape(
[images_dict['32x32x32']['t1'].shape[0], -1]
)
images_dict['128x128x128']['t1'] = images_dict['128x128x128']['t1'].reshape(
[images_dict['128x128x128']['t1'].shape[0], -1]
)
images_dict['32x32x32']['t2'] = images_dict['32x32x32']['t2'].reshape(
[images_dict['32x32x32']['t2'].shape[0], -1]
)
images_dict['128x128x128']['t2'] = images_dict['128x128x128']['t2'].reshape(
[images_dict['128x128x128']['t2'].shape[0], -1]
)
#@title The orginal T1 images have a total of 9175040 voxels.
from IPython.display import Markdown as md
md("The sizes for the 32x32x32 and 128x128x128 images are \
{} and {}, respectively. They represent {}% and \
{}% of the original size.".format(images_dict['32x32x32']['t1'].shape[1],
images_dict['128x128x128']['t1'].shape[1],
np.round(images_dict['32x32x32']['t1'].shape[1] / 9175040 * 100, 2),
np.round(images_dict['128x128x128']['t1'].shape[1] / 9175040 * 100, 2),
)
)
from sklearn.decomposition import PCA
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
pca32 = PCA(n_components=2)
pca32.fit(
np.concatenate([
images_dict['32x32x32']['t1'][:30],
images_dict['32x32x32']['t2'][:30]
], 0)
)
# Some samples (usually ~10-20%) are used as validation data that will not
# be used to train the model.
t1_transform_train = pca32.transform(images_dict['32x32x32']['t1'][:30])
t2_transform_train = pca32.transform(images_dict['32x32x32']['t2'][:30])
t1_transform_valid = pca32.transform(images_dict['32x32x32']['t1'][30:])
t2_transform_valid = pca32.transform(images_dict['32x32x32']['t2'][30:])
plt.figure(figsize=(12,6))
blues = ['b' for _ in range(len(images_dict['32x32x32']['t1'][:30]))]
greens = ['g' for _ in range(len(images_dict['32x32x32']['t2'][:30]))]
reds = ['r' for _ in range(len(images_dict['32x32x32']['t1'][30:]))]
cyans = ['c' for _ in range(len(images_dict['32x32x32']['t2'][30:]))]
blue_patch = mpatches.Patch(color='b', label='T1 (train)')
green_patch = mpatches.Patch(color='g', label='T2 (train)')
red_patch = mpatches.Patch(color='r', label='T1 (valid)')
cyan_patch = mpatches.Patch(color='c', label='T2 (valid)')
plt.scatter(t1_transform_train[:, 0], t1_transform_train[:, 1], c=blues)
plt.scatter(t2_transform_train[:, 0], t2_transform_train[:, 1], c=greens)
plt.scatter(t1_transform_valid[:, 0], t1_transform_valid[:, 1], c=reds)
plt.scatter(t2_transform_valid[:, 0], t2_transform_valid[:, 1], c=cyans)
plt.title('PCA of images resized to 32x32x32')
plt.legend()
plt.xlabel('Component 1')
plt.ylabel('Component 2')
plt.legend(handles=[blue_patch, green_patch, red_patch, cyan_patch])
plt.show()
plt.close()
pca128 = PCA(n_components=2)
pca128.fit(
np.concatenate([
images_dict['128x128x128']['t1'][:30],
images_dict['128x128x128']['t2'][:30]
], 0)
)
t1_transform_train = pca128.transform(images_dict['128x128x128']['t1'][:30])
t2_transform_train = pca128.transform(images_dict['128x128x128']['t2'][:30])
t1_transform_valid = pca128.transform(images_dict['128x128x128']['t1'][30:])
t2_transform_valid = pca128.transform(images_dict['128x128x128']['t2'][30:])
plt.figure(figsize=(12,6))
plt.scatter(t1_transform_train[:, 0], t1_transform_train[:, 1], c=blues)
plt.scatter(t2_transform_train[:, 0], t2_transform_train[:, 1], c=greens)
plt.scatter(t1_transform_valid[:, 0], t1_transform_valid[:, 1], c=reds)
plt.scatter(t2_transform_valid[:, 0], t2_transform_valid[:, 1], c=cyans)
plt.title('PCA of images resized to 128x128x128')
plt.xlabel('Component 1')
plt.ylabel('Component 2')
plt.legend(handles=[blue_patch, green_patch, red_patch, cyan_patch])
plt.show()
plt.close()
#@title The orginal T1 images have a total of 9175040 voxels.
from IPython.display import Markdown as md
md("For the 128x128x128 voxel images, the first component of the PCA "
"explains ~{}% of the variance of the images and the second ~{}%. "
"For the 32x32x32 images, the first component explains {}% of the "
"variance and the second {}%".format(
np.round(pca128.explained_variance_ratio_[0] * 100, 2),
np.round(pca128.explained_variance_ratio_[1] * 100, 2),
np.round(pca32.explained_variance_ratio_[0] * 100, 2),
np.round(pca32.explained_variance_ratio_[1] * 100, 2),
))
```
## Basic machine learning classification model
The classification in this tutorial is trivial, so a simple linear model like a logistic regression classifier should be able to learn hot to perfectly classify the images for both image sizes.
```
from sklearn.linear_model import LogisticRegression
print('32x32x32')
lr32 = LogisticRegression()
labels = [0 for x in range(len(images_dict['32x32x32']['t1'][:30]))] + \
[1 for x in range(len(images_dict['32x32x32']['t2'][:30]))]
labels_valid = [0 for x in range(len(images_dict['32x32x32']['t1'][30:]))] + \
[1 for x in range(len(images_dict['32x32x32']['t2'][30:]))]
lr32.fit(
np.concatenate([
images_dict['32x32x32']['t1'][:30],
images_dict['32x32x32']['t2'][:30]
], 0),
labels
)
# Labels T1 are 0s and T2 are 1
labels_t1_train = [0 for _ in preds_t1]
labels_t1_valid = [0 for _ in preds_t1_valid]
labels_t2_train = [1 for _ in preds_t2]
labels_t2_valid = [1 for _ in preds_t2_valid]
preds_t1 = lr32.predict(images_dict['32x32x32']['t1'][:30])
preds_t2 = lr32.predict(images_dict['32x32x32']['t2'][:30])
preds_t1_valid = lr32.predict(images_dict['32x32x32']['t1'][30:])
preds_t2_valid = lr32.predict(images_dict['32x32x32']['t2'][30:])
accuracy = sum([1 if pred == target else 0 for (pred, target) in zip(
np.concatenate((preds_t1_train, preds_t2_train)),
np.concatenate((labels_t1_train, labels_t2_train)))]
) / len(labels)
accuracy_valid = sum([1 if pred == target else 0 for (pred, target) in zip(
np.concatenate((preds_t1_valid, preds_t2_valid)),
np.concatenate((labels_t1_valid, labels_t2_valid)))]
) / len(labels_valid)
print('Train Accuracy: ', accuracy)
print('Valid Accuracy: ', accuracy_valid)
print('128x128x128')
lr128 = LogisticRegression()
labels = [0 for x in range(len(images_dict['128x128x128']['t1'][:30]))] + \
[1 for x in range(len(images_dict['128x128x128']['t2'][:30]))]
labels_valid = [0 for x in range(len(images_dict['128x128x128']['t1'][30:]))] + \
[1 for x in range(len(images_dict['32x32x32']['t2'][30:]))]
lr128.fit(
np.concatenate([
images_dict['128x128x128']['t1'][:30],
images_dict['128x128x128']['t2'][:30]
], 0),
labels
)
preds_t1_train = lr128.predict(images_dict['128x128x128']['t1'][:30])
preds_t2_train = lr128.predict(images_dict['128x128x128']['t2'][:30])
preds_t1_valid = lr128.predict(images_dict['128x128x128']['t1'][30:])
preds_t2_valid = lr128.predict(images_dict['128x128x128']['t2'][30:])
accuracy = sum([1 if pred == target else 0 for (pred, target) in zip(
np.concatenate((preds_t1_train, preds_t2_train)),
np.concatenate((labels_t1_train, labels_t2_train)))]
) / len(labels)
accuracy_valid = sum([1 if pred == target else 0 for (pred, target) in zip(
np.concatenate((preds_t1_valid, preds_t2_valid)),
np.concatenate((labels_t1_valid, labels_t2_valid)))]
) / len(labels_valid)
print('Train Accuracy: ', accuracy)
print('Valid Accuracy: ', accuracy_valid)
```
| github_jupyter |
```
%%html
<style>
/*#These style overrides are for printing in b/w. The Jupyter style comes from
CodeMirror. The CodeMirror overrides all begin with .cm below.
*/
div.input_area {
border: 4px solid #cfcfcf;
border-radius: 4px;
background: white;
line-height: 1.21429em;
}
div.cell{color: black;}
/*
These are all style overrides from the CodeMirror theme
Good source of names:
https://github.com/draperjames/one-dark-notebook/blob/master/custom.css
*/
/*
.cm-s-ipython span.cm-string {
color: black;
}
.cm-s-ipython span.cm-link {
color: black;
}
.cm-s-ipython span.cm-keyword {
color: black;
}
.cm-s-ipython .CodeMirror-linenumber {
color: black;
font-size: 10px;
}
.cm-s-ipython span.cm-comment {
color: black;
}
.cm-s-ipython span.cm-builtin {
color: black;
}
.cm-s-ipython span.cm-variable {
color: black;
}
.cm-s-ipython span.cm-variable-2 {
color: black;
}
.cm-s-ipython span.cm-variable-3 {
color: black;
}
.cm-s-default span.cm-bracket {
color: black;
}
.cm-s-ipython span.cm-operator {
color: black;
}
.cm-s-ipython span.cm-number {
color: black;
}
*/
</style>
import os
from jupyter_core.paths import jupyter_config_dir
jupyter_dir = jupyter_config_dir()
print(jupyter_dir)
print(os.path.exists(jupyter_dir))
```
# <center> <u> A Deep Dive Into Comprehensions </u></center>
### <center> <i>Patrick Barton barton.pj@gmail.com</center> </i>
Comprehensions provide an elegant and efficient means of producing collections in Python. They also allow on-the-fly application of filters.
Even if you don't use them yourself, it's worthwhile learning how to read them because you'll find plenty of them "in the wild", embedded in Python libraries you'll want to understand. And they can be might handy once you get the hang of them.
## Use Cases
Here are a couple situations where you may want to use comprehensions:
<u>Case 1</u>: You inherited a messy data table from your predecessor. It contains a bunch of values "Val_1", "Val_2", etc. each calculated for a different year. The column headings show up something like "Val_1_2025". You want to sort them and screen only for the year 2025, retaining the 'catch' in a new object.
With a comprehension, you could pull that off with a single, short line of code:
```
columns = ['Val_3_2020', 'Val_1_2025', 'Val_3_2023', 'Val_3_2024', 'Val_3_2024', 'Val_3_2023', 'Val_3_2025',
'Val_2_2023', 'Val_1_2023', 'Val_2_2025', 'Val_1_2022', 'Val_3_2023', 'Val_2_2024', 'Val_3_2021',
'Val_1_2023', 'Val_3_2024', 'Val_3_2021', 'Val_1_2020', 'Val_3_2024', 'Val_1_2022']
[ col for col in sorted(columns) if '2025' in col]
```
<u>Case 2</u>: You just need some sensible index names and placeholder values for a pandas <b>Series</b> object and want to do it efficiently. You could go:
```
import pandas as pd
pd.Series(data = [val for val in range(5)], index = ["Year_{}".format(yr) for yr in range(1905, 1910)])
```
A notable downside to comprehensions is that they can be difficult to decypher for those not familiar with the syntax. Don't believe me? Here's an example:
```
[y for level in three_d_array if level[2][2] %2 for x in level if x[1] > 3 for y in x if not y % 3 ]
```
A bit daunting, I'll admit. But fear not. When you've completed this unit, you'll be able to figure out what it produces with your pocket protector tied behind your back. Besides, there's nothing you can do with comprehensions that you can't do another way, so you don't really <u>need</u> to nail them. When you're done reading this unit, you'll be able to convert this into a (likely more easily-understood), verbose format and never have to look at again.
## Basics
But let's begin at the beginning. Here's a simple example of a list comprehension:
```
iterable = "hey"
[item for item in iterable]
```
As you can see, we've done is created a new list object. It's identical to the following, more verbose, code (with a couple subtle differences):
```
as_list = []
for char in iterable:
as_list.append(char)
as_list
```
What are the differences? In the verbose mode we had to add the name 'as_list' to the namespace and create an empty <b>list</b> instance. We also had to add the iterating variable 'item' to the project namespace.
Using the <b>list</b> comprehension, we had to do neither. The comprehension has its own "mini-namespace" which is created on the fly, and which goes out of context as soon as the comprehension has completed.
Now, lets build this up a bit at a time - it'll be easier to remember the steps this way.
The iterable object can be a <b>list</b>, <b>set</b>, <b>tuple</b>, generator expression, or just about anything with a __next__() method defined will do. Here's an example with a range object.
```
[i for i in range(10, 20, 2)]
```
Now, since we've produced the iterating variable 'i', we can make any use of it to perform a "preprocessing operation" to produce the item added to the <b>list</b> at each iteration.
Let's say we wanted to find which printable character is mapped to a <b>range</b> of ordinal code points. Here's an easy way to manage it:
```
print([chr(code_point) for code_point in range(50, 65) ])
```
Good so far? Now, we can go upmarket a bit and add a filter to "screen in" a subset of the values produced by the iterating expression. The filter bit goes on the right.
Here's an example where we're looking only for the numbers evenly divisible by 3:
```
print([i for i in range(10, 20, 2) if not i % 3 ] )
```
The filter can be any valid expression that can Python can evaluate as Boolean (<b>True</b>/<b>False</b>).
As an aside, Python is super-flexible in this regard. Any non-empty object and any non-zero number will evaluated as <b>True</b> when subjected to a Boolean test. The "%" is Python's modulo operator - it returns the remainder of a "floor" (rounded down) division e.g., 13 % 3 --> 1
## 2-D Objects (adding an extra internal variable)
Let's circle back to the notion of a comprehension's namespace. As mentioned earlier, the comprehension whistles up its own "mini-namespace" - the iterating variable 'item' is defined anew as the comprehension is calculated. To demonstrate:
```
item = "Snoopy the beagle" #setting 'item' in the main namespace
[ item for item in {'x', 'y', 'z'} ] #re-using 'item' as an iterating variable
print(item)
```
Introduction of a name to the internal namespace is done via the defining the iterating expression.
We've created an internal definition of 'item' and an internal definition of the <b>set</b> {'x', 'y', 'z'}. That's a seperate object from the object of the same name in the mainline code, and the value did not propogate.
If you need to create additional iterating variables, you need to create additional <b>for</b> statements.
Let's say you had an array-like structure, represented here as a <b>list</b> of <b>lists</b>, and you wanted to create a flattened version of it. You would want two iterating variables - one for the rows and the other for the columns. So you can go:
```
nested = [ [1,2,3],
[4,5,6],
[7,8, 9] ]
flat = [y for x in nested for y in x]
flat
```
```
[y
for x in nested
for y in x]
```
## 3-D and Beyond
You can extend a nested <b>list</b> comprehension to an arbitrarily-large number of levels. Here we'll discuss the example presented at the beginning of this unit along with a concrete example.
```
#This is the array we'll flatten
three_d_array = [[[11, 12, 13], [14, 15, 16], [17, 18, 19]], #top 'level'
[[21, 22, 23], [24, 25, 26], [27, 28, 29]], #middle 'level'
[[31, 32, 33], [34, 35, 36], [37, 38, 39]]] #bottom 'level'
flat_three_d = [y for level in three_d_array if level[2][2] %2 for x in level if x[1] > 3 for y in x if not y % 3 ]
flat_three_d
```
Here's an annotated version of the comprehension in verbose form. You'll note that we have three iterating variables, 'level', 'x', and 'y' and a filter associated with each:
```
flat_three_d = []
for level in three_d_array:
if level[2][2] %2: #is the last element of the last list in the 'level' odd?
for x in level:
if x[1] > 3: #is the second element of of each list larger than 3?
for y in x:
if not y %3: #is the individual element divisible by 3?
flat_three_d.append(y) #... if so, the number 'screens in' to the finished list
flat_three_d
```
Following the pattern in the 2-d example, we'll apply indentation to make it a bit more readable.
```
flat_three_d = [y #we'll add 'y' to the list here
for level in three_d_array #loop through the 'levels'
if level[2][2] %2 #...apply the 'level filter'
for x in level #loop thru the 'x' dimension
if x[1] > 3 #...apply the 'x' filter
for y in x #loop thru the 'y' dimension
if not y % 3 ] #...apply the 'y' filter
flat_three_d
```
## Going the Other Way - Building Data Structures
Just as you can flatten high-dimensional data, you can also build data structures starting from a flat data source. Just for fun, let's try to make one of these:
```
target = [[1, 2, 3], [4, 5, 6], [7, 8, 9], [10, 11, 12]]
```
So we're shooting for an array-like structure comprised of a <b>list</b> of <b>list</b> objects. Each of the internal <b>list</b> objects contains consecutive integers. Each new <b>list</b> picks off where the old one left off.
So, we're going to need a computational way to figure out the starting point of each internal <b>list</b>. Then populate the rest if it. One way to code this is:
```
#[ [start1 = 1, start1+1, start1+2 ], [start2 = 4, start2+1, start2_2], ... ]
outer = []
for y in range(0, 10, 3): #number just before start 0, 3, 6, 9
inner = []
for x in range(1, 4): # numbers to add: 1, 2, 3
inner.append(x + y)
outer.append(inner)
print(outer)
print("Nailed it? {}.".format(outer==target))
```
Alternatively, we could use nested comprehensions where we use a outer 'y' loop to find the initial values; and the inner 'x' loop to populate individual <b>lists</b>. The comprehension should be structured thusly:
```
[ [x + y
for x in range (1, 4)]
for y in range(0, 10, 3)
]
# more simply as a 'one-liner'
#inner loop makes inner lists ...with every cycle of the outer loop
[[x + y for x in range(1,4) ] for y in range(0, 10, 3)]
```
Creating a 3-d structure is just a little more complex. Here's a quick-and-dirty way to create a stack of array-like <b>'list-of-lists'</b> structure. This is handy code fragment to keep around for testing - you'll note that you can determine the x, y, and z coordinates based on the data values alone.
```
[ #new stack
[ #new row
[z*100 + y*10 + x for x in range(1,4)] #a list within a row ('inner loop')
for y in range(4) # for statement populates the list elements
] #end of new row
for z in range(1,4) #for statement to add a new layer layer within the stack
] #end of new stack
# ...as a 'one-liner'
#<-----------------------------containg list ('the stack')-------------------------------------->
# <------------------a row------------------------------------------->
# <-------------an inner list----------->
[ [ [z*100 + y*10 + x for x in range(1,4) ] for y in range(4)] for z in range(1,4) ]
```
## Other 'Flavors' of Comprehensions
Besides the <b>list</b> comprehension Python supports lots of others including comprehensions for <b>set</b> and <b>dict</b> objects, as well as one that creates a generator object. The good news is that they have about the same syntax. Here are some examples:
```
#Set comprehension - just use opposing {curly braces} instead of [square brackets]. Note that the set object deduplicates.
{ value**2 for value in range(-5, 5)}
#Dict comprehension - same as a set comprehension, but with a key:value pairing on the left.
{char:ord(char) for char in ['A', '*', 'x', '|']}
#Generator comprehension - same as a list comprehension, but with (parens) instead of [square brackets].
gen = (color for color in ['red', 'blue', 'green'])
print (gen)
print(next(gen))
print(next(gen))
print(next(gen))
```
## Further Reading
If you're interested in learning more about comprehensions in general, or in exploring their history, application in other languages, etc. Wikipedia has an excellent article here:
https://en.wikipedia.org/wiki/List_comprehension#History
| github_jupyter |
# Checking stimuli for balance
This notebook helps to ensure that the generated stimuli are roughly balanced between positive and negative trials.
```
import os
import numpy as np
from PIL import Image
import pandas as pd
import json
import pymongo as pm
from glob import glob
from IPython.display import clear_output
import ast
import itertools
import random
import h5py
from tqdm import tqdm
import matplotlib.pyplot as plt
#display all columns
pd.set_option('display.max_columns', None)
def list_files(paths, ext='mp4'):
"""Pass list of folders if there are stimuli in multiple folders.
Make sure that the containing folder is informative, as the rest of the path is ignored in naming.
Also returns filenames as uploaded to S3"""
if type(paths) is not list:
paths = [paths]
results = []
names = []
for path in paths:
results += [y for x in os.walk(path) for y in glob(os.path.join(x[0], '*.%s' % ext))]
names += [os.path.basename(os.path.dirname(y))+'_'+os.path.split(y)[1].split('.')[0] for x in os.walk(path) for y in glob(os.path.join(x[0], '*.%s' % ext))]
# hdf5s = [r.split("_img.")[0]+".hdf5" for r in results]
hdf5s = [r.split("_img.")[0] for r in results]
return results,names,hdf5s
local_stem = 'XXX' #CHANGE THIS ⚡️
dirnames = [d.split('/')[-1] for d in glob(local_stem+'/*')]
data_dirs = [local_stem + d for d in dirnames]
stimulus_extension = "hdf5" #what's the file extension for the stims? Provide without dot
## get a list of paths to each one
full_stim_paths,filenames, full_hdf5_paths = list_files(data_dirs,stimulus_extension)
full_map_paths, mapnames, _ = list_files(data_dirs, ext = 'png') #generate filenames and stimpaths for target/zone map
print('We have {} stimuli to evaluate.'.format(len(full_stim_paths)))
stim_IDs = [name.split('.')[0] for name in filenames]
set_names= ['_'.join(s.split('_')[:-2]) for s in stim_IDs]
## convert to pandas dataframe
M = pd.DataFrame([stim_IDs,set_names]).transpose()
M.columns = ['stim_ID','set_name']
# if needed, add code to add additional columns
# Add trial labels to the metadata using the stimulus metadata.json
target_hit_zone_labels = dict()
for _dir in data_dirs:
with open(_dir + '/metadata.json', 'rb') as f:
trial_metas = json.load(f)
for i,meta in enumerate(trial_metas):
stim_name = meta['stimulus_name']
if stim_name == 'None': #recreate stimname from order in metadata
stim_name = str(i).zfill(4)
stim_name = _dir.split('/')[-1] + '_' + stim_name
# if stim_name[-4:] != "_img": stim_name+='_img' #stimnames need to end in "_img"
label = meta['does_target_contact_zone']
target_hit_zone_labels[stim_name] = label
print("num positive labels: %d" % sum(list(target_hit_zone_labels.values())))
print("num negative labels: %d" % (len(target_hit_zone_labels) - sum(list(target_hit_zone_labels.values()))))
print("ratio",sum(list(target_hit_zone_labels.values())) / (len(target_hit_zone_labels) - sum(list(target_hit_zone_labels.values()))))
# make new df with all metadata
GT = pd.DataFrame([list(target_hit_zone_labels.keys()), list(target_hit_zone_labels.values())]).transpose()
GT.columns = ['stim_ID', 'target_hit_zone_label']
# merge with M
M = M.merge(GT, on='stim_ID')
print("added labels %s" % list(GT.columns[1:]))
metadata = {} #holds all the metadata for all stimuli
for name,hdf5_path in tqdm(list(zip([f.split('.')[0] for f in filenames],full_hdf5_paths))):
#load hdf5
# print("loading",hdf5_path)
try:
hdf5 = h5py.File(hdf5_path,'r') #get the static part of the HDF5
stim_name = str(np.array(hdf5['static']['stimulus_name']))
metadatum = {} #metadata for the current stimulus
for key in hdf5['static'].keys():
datum = np.array(hdf5['static'][key])
if datum.shape == (): datum = datum.item() #unwrap non-arrays
metadatum[key] = datum
#close file
hdf5.close()
metadata[name] = metadatum
except Exception as e:
print("Error with",hdf5_path,":",e)
continue
```
Insert those metadatas into M:
```
for index in M.index:
stim_name = M.at[index,'stim_ID']
for key,value in metadata[stim_name].items():
M.at[index,key] = str(value) #insert every item as string
M
M['label'] = M['target_hit_zone_label'].astype(int)
def get_set_base(name):
"""Group together stims independent of distractors/room.
Assumes a naming scheme with `setname_{tdw,box}_occluderinfo`"""
return name.split("_tdw")[0].split("_box")[0]
M['set_base'] = M['set_name'].apply(get_set_base)
```
## Analysis
How many stimuli?
```
len(M)
```
How many of trials are positive (1) rather than negative (0)?
```
np.mean(M['label'])
```
How many of trials *per set name* are positive (1) rather than negative (0)?
```
M.groupby('set_name').agg({'stim_ID':['count'],'label':['mean']})
```
How many of trials *per set base* (independent of room or occluders—assumes a naming scheme with `setname_{tdw,box}_occluderinfo`) are positive (1) rather than negative (0)?
```
M.groupby('set_base').agg({'stim_ID':['count'],'label':['mean']})
```
| github_jupyter |
<a href="https://colab.research.google.com/github/zahraDehghanian97/Poetry_Generator/blob/master/Word_Poem_generator.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import tensorflow as tf
from tensorflow import keras
import numpy as np
import pickle
from nltk.metrics import accuracy ,ConfusionMatrix
from nltk.translate.bleu_score import sentence_bleu
seqLength = 20
BATCH_SIZE = 64
BUFFER_SIZE = 100
embedding_dim = 256
rnn_units = 1024
```
# make data ready
```
filepath = "/content/drive/MyDrive/Colab Notebooks/my_shahname_represntation.txt"
with open(filepath, "rb") as f:
corpus , test = pickle.load(f)
corpus = corpus.replace("\t"," \t ").replace("\n", " \n ")
corpusList = [w for w in corpus.split(' ')]
corpus_words = [i for i in corpusList if i]
map(str.strip, corpus_words)
vocab = sorted(set(corpus_words))
print(len(corpus_words))
vocab_size = len(vocab)
word2idx = {u: i for i, u in enumerate(vocab)}
idx2words = np.array(vocab)
word_as_int = np.array([word2idx[c] for c in corpus_words])
def split_input_target(chunk):
input_text = chunk[:-1]
target_text = chunk[1:]
return input_text, target_text
# examples_per_epoch = len(corpus_words)//(seqLength + 1)
wordDataset = tf.data.Dataset.from_tensor_slices(word_as_int)
sequencesOfWords = wordDataset.batch(seqLength + 1, drop_remainder=True)
dataset = sequencesOfWords.map(split_input_target)
dataset = dataset.shuffle(BUFFER_SIZE).batch(BATCH_SIZE, drop_remainder=True)
```
# LSTM Model
```
def create_model_lstm(vocab_size, embedding_dim, rnn_units, batch_size):
model = tf.keras.Sequential()
model.add(tf.keras.layers.Embedding(vocab_size, embedding_dim,batch_input_shape=[batch_size, None]))
model.add(tf.keras.layers.LSTM(rnn_units,return_sequences=True,stateful=True,recurrent_initializer='glorot_uniform'))
model.add(tf.keras.layers.LSTM(rnn_units,return_sequences=True,stateful=True,recurrent_initializer='glorot_uniform'))
model.add(tf.keras.layers.Dense(vocab_size))
return model
lstm_model = create_model_lstm(vocab_size = len(vocab), embedding_dim=embedding_dim, rnn_units=rnn_units, batch_size=BATCH_SIZE)
lstm_model.compile(optimizer='adam', loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True), metrics=['accuracy'])
history = lstm_model.fit(dataset, epochs=50)
main_lstm_model = create_model_lstm(vocab_size = len(vocab), embedding_dim=embedding_dim, rnn_units=rnn_units, batch_size=1)
main_lstm_model.set_weights(lstm_model.get_weights())
# main_lstm_model = tf.keras.models.load_model('/content/drive/MyDrive/Colab Notebooks/word_lstm.h5')
main_lstm_model.summary()
def generate_text(model, start_string):
num_generate = 200
start_string_list =[]
for w in start_string.split(' '):
if w in word2idx :
start_string_list.append(w)
input_eval = [word2idx[s] for s in start_string_list]
input_eval = tf.expand_dims(input_eval, 0)
text_generated = []
model.reset_states()
for i in range(num_generate):
predictions = model(input_eval)
predictions = tf.squeeze(predictions, 0)
predicted_id = tf.random.categorical(predictions, num_samples=1)[-1,0].numpy()
input_eval = tf.expand_dims([predicted_id], 0)
text_generated.append(idx2words[predicted_id])
return (start_string + ' '.join(text_generated))
print(generate_text(main_lstm_model, start_string=u"چنین گفت رستم به اسفندیار"))
main_lstm_model.save("/content/drive/MyDrive/Colab Notebooks/word_lstm.h5")
```
# Test
```
BLEU_scores = []
accuracy_scores = []
poem = test[0]
start = poem[:25]
generated_poem = generate_text(main_lstm_model, start_string=start)
BLEU_scores.append(sentence_bleu(poem, generated_poem))
len_min = min(len(poem),len(generated_poem))
accuracy_scores.append(accuracy(poem[:len_min], generated_poem[:len_min]))
print("-----------------------")
print("start sentence : ",start)
print(generated_poem)
print("BLEU score = ",BLEU_scores[-1])
print("Accuracy score = ",accuracy_scores[-1])
print("Confusion matrix =")
print(ConfusionMatrix(poem[:len_min], generated_poem[:len_min]))
counter = 0
for poem in test :
counter+=1
start = poem[:25]
generated_poem = generate_text(main_lstm_model, start_string=start)
BLEU_scores.append(sentence_bleu(poem, generated_poem))
len_min = min(len(poem),len(generated_poem))
accuracy_scores.append(accuracy(poem[:len_min], generated_poem[:len_min]))
print("-----------------------")
print("sentence number : ",counter)
print("BLEU score = ",BLEU_scores[-1])
print("Accuracy score = ",accuracy_scores[-1])
print("<<------------final report----------->>")
print("number of test set = ",len(test))
print("mean BLEU score = ",np.mean(BLEU_scores))
print("mean Accuracy score = ",np.mean(accuracy_scores))
```
| github_jupyter |
```
from mmdet.apis import init_detector, inference_detector, show_result_pyplot, show_result
import mmcv
import time
import json
import numpy as np
import glob
import os
config_file = 'configs/faster_rcnn_x101_64x4d_fpn_TSD_zalo.py'
# download the checkpoint from model zoo and put it in `checkpoints/`
checkpoint_file = 'checkpoints/faster_rcnn_x101_64x4d_fpn_TSD/epoch_6.pth'
public_test = "data/za_traffic_2020/traffic_public_test/images/*.png"
save_folder = "./6/"
score_thr = 0.3
class_names = ["1. No entry", "2. No parking / waiting", \
"3. No turning", "4. Max Speed", \
"5. Other prohibition signs", "6. Warning", "7. Mandatory"]
# build the model from a config file and a checkpoint file
model = init_detector(config_file, checkpoint_file, device='cuda:0')
submit_list = []
for test_file_path in glob.glob(public_test):
img_file = os.path.basename(test_file_path)
img_name, _ = img_file.split('.')
print (img_name)
img_im_temp = mmcv.imread(test_file_path)
result = inference_detector(model, test_file_path)
img_out, out_bboxes, out_labels = show_result(test_file_path, result, class_names, show=False)
out_scores = out_bboxes[:, -1]
inds = out_scores > score_thr
out_bboxes = out_bboxes[inds, :]
out_labels = out_labels[inds]
mmcv.imwrite(img_out, os.path.join(save_folder, img_file))
for idx, (bbox, label) in enumerate(zip(out_bboxes, out_labels)):
temp_ = dict()
temp_["image_id"] = int(img_name)
bbox_elem0, bbox_elem1, bbox_elem2, bbox_elem3, score = bbox
bbox_width = bbox_elem2 - bbox_elem0
bbox_height = bbox_elem3 - bbox_elem1
temp_["category_id"] = label
temp_["bbox"] = [round(bbox_elem0, 2), round(bbox_elem1, 2), round(bbox_width, 2), round(bbox_height, 2)]
temp_["score"] = score
print (bbox_elem0, bbox_elem1, bbox_elem2, bbox_elem3)
print (round(bbox_elem0, 2), round(bbox_elem1, 2), round(bbox_width, 2), round(bbox_height, 2))
print ('{:.2f} {:.2f} {:.2f} {:.2f}'.format(bbox_elem0, bbox_elem1, bbox_width, bbox_height))
print (label)
# print (img_im_temp.shape)
# img_im_patch = img_im_temp[int(bbox_elem1):int(bbox_elem3),
# int(bbox_elem0):int(bbox_elem2), :]
# print (img_im_patch.shape)
# mmcv.imwrite(img_im_patch, "{}_{}_{}.png".format(img_name, idx, label))
submit_list.append(temp_)
submit_list
with open("./submit_{}.json".format(time.time()), "w") as file:
file.writelines(str(submit_list))
out_file = open("./submit_{}.json".format(time.time()), "w")
json.dump(submit_list, out_file, indent = 6)
out_file.close()
```
| github_jupyter |
# Assignment 4: Word Embeddings
Welcome to the fourth (and last) programming assignment of Course 2!
In this assignment, you will practice how to compute word embeddings and use them for sentiment analysis.
- To implement sentiment analysis, you can go beyond counting the number of positive words and negative words.
- You can find a way to represent each word numerically, by a vector.
- The vector could then represent syntactic (i.e. parts of speech) and semantic (i.e. meaning) structures.
In this assignment, you will explore a classic way of generating word embeddings or representations.
- You will implement a famous model called the continuous bag of words (CBOW) model.
By completing this assignment you will:
- Train word vectors from scratch.
- Learn how to create batches of data.
- Understand how backpropagation works.
- Plot and visualize your learned word vectors.
Knowing how to train these models will give you a better understanding of word vectors, which are building blocks to many applications in natural language processing.
## Outline
- [1 The Continuous bag of words model](#1)
- [2 Training the Model](#2)
- [2.0 Initialize the model](#2)
- [Exercise 01](#ex-01)
- [2.1 Softmax Function](#2.1)
- [Exercise 02](#ex-02)
- [2.2 Forward Propagation](#2.2)
- [Exercise 03](#ex-03)
- [2.3 Cost Function](#2.3)
- [2.4 Backproagation](#2.4)
- [Exercise 04](#ex-04)
- [2.5 Gradient Descent](#2.5)
- [Exercise 05](#ex-05)
- [3 Visualizing the word vectors](#3)
<a name='1'></a>
# 1. The Continuous bag of words model
Let's take a look at the following sentence:
>**'I am happy because I am learning'**.
- In continuous bag of words (CBOW) modeling, we try to predict the center word given a few context words (the words around the center word).
- For example, if you were to choose a context half-size of say $C = 2$, then you would try to predict the word **happy** given the context that includes 2 words before and 2 words after the center word:
> $C$ words before: [I, am]
> $C$ words after: [because, I]
- In other words:
$$context = [I,am, because, I]$$
$$target = happy$$
The structure of your model will look like this:
<div style="width:image width px; font-size:100%; text-align:center;"><img src='word2.png' alt="alternate text" width="width" height="height" style="width:600px;height:250px;" /> Figure 1 </div>
Where $\bar x$ is the average of all the one hot vectors of the context words.
<div style="width:image width px; font-size:100%; text-align:center;"><img src='mean_vec2.png' alt="alternate text" width="width" height="height" style="width:600px;height:250px;" /> Figure 2 </div>
Once you have encoded all the context words, you can use $\bar x$ as the input to your model.
The architecture you will be implementing is as follows:
\begin{align}
h &= W_1 \ X + b_1 \tag{1} \\
a &= ReLU(h) \tag{2} \\
z &= W_2 \ a + b_2 \tag{3} \\
\hat y &= softmax(z) \tag{4} \\
\end{align}
```
# Import Python libraries and helper functions (in utils2)
import nltk
from nltk.tokenize import word_tokenize
import numpy as np
from collections import Counter
from utils2 import sigmoid, get_batches, compute_pca, get_dict
# Download sentence tokenizer
nltk.data.path.append('.')
# Load, tokenize and process the data
import re # Load the Regex-modul
with open('shakespeare.txt') as f:
data = f.read() # Read in the data
data = re.sub(r'[,!?;-]', '.',data) # Punktuations are replaced by .
data = nltk.word_tokenize(data) # Tokenize string to words
data = [ ch.lower() for ch in data if ch.isalpha() or ch == '.'] # Lower case and drop non-alphabetical tokens
print("Number of tokens:", len(data),'\n', data[:15]) # print data sample
# Compute the frequency distribution of the words in the dataset (vocabulary)
fdist = nltk.FreqDist(word for word in data)
print("Size of vocabulary: ",len(fdist) )
print("Most frequent tokens: ",fdist.most_common(20) ) # print the 20 most frequent words and their freq.
```
#### Mapping words to indices and indices to words
We provide a helper function to create a dictionary that maps words to indices and indices to words.
```
# get_dict creates two dictionaries, converting words to indices and viceversa.
word2Ind, Ind2word = get_dict(data)
V = len(word2Ind)
print("Size of vocabulary: ", V)
# example of word to index mapping
print("Index of the word 'king' : ",word2Ind['king'] )
print("Word which has index 2743: ",Ind2word[2743] )
```
<a name='2'></a>
# 2 Training the Model
### Initializing the model
You will now initialize two matrices and two vectors.
- The first matrix ($W_1$) is of dimension $N \times V$, where $V$ is the number of words in your vocabulary and $N$ is the dimension of your word vector.
- The second matrix ($W_2$) is of dimension $V \times N$.
- Vector $b_1$ has dimensions $N\times 1$
- Vector $b_2$ has dimensions $V\times 1$.
- $b_1$ and $b_2$ are the bias vectors of the linear layers from matrices $W_1$ and $W_2$.
The overall structure of the model will look as in Figure 1, but at this stage we are just initializing the parameters.
<a name='ex-01'></a>
### Exercise 01
Please use [numpy.random.rand](https://numpy.org/doc/stable/reference/random/generated/numpy.random.rand.html) to generate matrices that are initialized with random values from a uniform distribution, ranging between 0 and 1.
**Note:** In the next cell you will encounter a random seed. Please **DO NOT** modify this seed so your solution can be tested correctly.
```
# UNQ_C1 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: initialize_model
def initialize_model(N,V, random_seed=1):
'''
Inputs:
N: dimension of hidden vector
V: dimension of vocabulary
random_seed: random seed for consistent results in the unit tests
Outputs:
W1, W2, b1, b2: initialized weights and biases
'''
np.random.seed(random_seed)
### START CODE HERE (Replace instances of 'None' with your code) ###
# W1 has shape (N,V)
W1 = np.random.rand(N,V)
# W2 has shape (V,N)
W2 = np.random.rand(V,N)
# b1 has shape (N,1)
b1 = np.random.rand(N,1)
# b2 has shape (V,1)
b2 = np.random.rand(V,1)
### END CODE HERE ###
return W1, W2, b1, b2
# Test your function example.
tmp_N = 4
tmp_V = 10
tmp_W1, tmp_W2, tmp_b1, tmp_b2 = initialize_model(tmp_N,tmp_V)
assert tmp_W1.shape == ((tmp_N,tmp_V))
assert tmp_W2.shape == ((tmp_V,tmp_N))
print(f"tmp_W1.shape: {tmp_W1.shape}")
print(f"tmp_W2.shape: {tmp_W2.shape}")
print(f"tmp_b1.shape: {tmp_b1.shape}")
print(f"tmp_b2.shape: {tmp_b2.shape}")
```
##### Expected Output
```CPP
tmp_W1.shape: (4, 10)
tmp_W2.shape: (10, 4)
tmp_b1.shape: (4, 1)
tmp_b2.shape: (10, 1)
```
<a name='2.1'></a>
### 2.1 Softmax
Before we can start training the model, we need to implement the softmax function as defined in equation 5:
<br>
$$ \text{softmax}(z_i) = \frac{e^{z_i} }{\sum_{i=0}^{V-1} e^{z_i} } \tag{5} $$
- Array indexing in code starts at 0.
- $V$ is the number of words in the vocabulary (which is also the number of rows of $z$).
- $i$ goes from 0 to |V| - 1.
<a name='ex-02'></a>
### Exercise 02
**Instructions**: Implement the softmax function below.
- Assume that the input $z$ to `softmax` is a 2D array
- Each training example is represented by a column of shape (V, 1) in this 2D array.
- There may be more than one column, in the 2D array, because you can put in a batch of examples to increase efficiency. Let's call the batch size lowercase $m$, so the $z$ array has shape (V, m)
- When taking the sum from $i=1 \cdots V-1$, take the sum for each column (each example) separately.
Please use
- numpy.exp
- numpy.sum (set the axis so that you take the sum of each column in z)
```
# UNQ_C2 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: softmax
def softmax(z):
'''
Inputs:
z: output scores from the hidden layer
Outputs:
yhat: prediction (estimate of y)
'''
### START CODE HERE (Replace instances of 'None' with your own code) ###
# Calculate yhat (softmax)
yhat = np.exp(z) / np.exp(z).sum(axis=0)
### END CODE HERE ###
return yhat
# Test the function
tmp = np.array([[1,2,3],
[1,1,1]
])
tmp_sm = softmax(tmp)
display(tmp_sm)
```
##### Expected Ouput
```CPP
array([[0.5 , 0.73105858, 0.88079708],
[0.5 , 0.26894142, 0.11920292]])
```
<a name='2.2'></a>
### 2.2 Forward propagation
<a name='ex-03'></a>
### Exercise 03
Implement the forward propagation $z$ according to equations (1) to (3). <br>
\begin{align}
h &= W_1 \ X + b_1 \tag{1} \\
a &= ReLU(h) \tag{2} \\
z &= W_2 \ a + b_2 \tag{3} \\
\end{align}
For that, you will use as activation the Rectified Linear Unit (ReLU) given by:
$$f(h)=\max (0,h) \tag{6}$$
<details>
<summary>
<font size="3" color="darkgreen"><b>Hints</b></font>
</summary>
<p>
<ul>
<li>You can use numpy.maximum(x1,x2) to get the maximum of two values</li>
<li>Use numpy.dot(A,B) to matrix multiply A and B</li>
</ul>
</p>
```
# UNQ_C3 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: forward_prop
def forward_prop(x, W1, W2, b1, b2):
'''
Inputs:
x: average one hot vector for the context
W1, W2, b1, b2: matrices and biases to be learned
Outputs:
z: output score vector
'''
### START CODE HERE (Replace instances of 'None' with your own code) ###
# Calculate h
h = np.dot(W1,x)+b1
# Apply the relu on h (store result in h)
h = np.maximum(0,h)
# Calculate z
z = np.dot(W2,h)+b2
### END CODE HERE ###
return z, h
# Test the function
# Create some inputs
tmp_N = 2
tmp_V = 3
tmp_x = np.array([[0,1,0]]).T
tmp_W1, tmp_W2, tmp_b1, tmp_b2 = initialize_model(N=tmp_N,V=tmp_V, random_seed=1)
print(f"x has shape {tmp_x.shape}")
print(f"N is {tmp_N} and vocabulary size V is {tmp_V}")
# call function
tmp_z, tmp_h = forward_prop(tmp_x, tmp_W1, tmp_W2, tmp_b1, tmp_b2)
print("call forward_prop")
print()
# Look at output
print(f"z has shape {tmp_z.shape}")
print("z has values:")
print(tmp_z)
print()
print(f"h has shape {tmp_h.shape}")
print("h has values:")
print(tmp_h)
```
##### Expected output
```CPP
x has shape (3, 1)
N is 2 and vocabulary size V is 3
call forward_prop
z has shape (3, 1)
z has values:
[[0.55379268]
[1.58960774]
[1.50722933]]
h has shape (2, 1)
h has values:
[[0.92477674]
[1.02487333]]
```
<a name='2.3'></a>
## 2.3 Cost function
- We have implemented the *cross-entropy* cost function for you.
```
# compute_cost: cross-entropy cost functioN
def compute_cost(y, yhat, batch_size):
# cost function
logprobs = np.multiply(np.log(yhat),y) + np.multiply(np.log(1 - yhat), 1 - y)
cost = - 1/batch_size * np.sum(logprobs)
cost = np.squeeze(cost)
return cost
# Test the function
tmp_C = 2
tmp_N = 50
tmp_batch_size = 4
tmp_word2Ind, tmp_Ind2word = get_dict(data)
tmp_V = len(word2Ind)
tmp_x, tmp_y = next(get_batches(data, tmp_word2Ind, tmp_V,tmp_C, tmp_batch_size))
print(f"tmp_x.shape {tmp_x.shape}")
print(f"tmp_y.shape {tmp_y.shape}")
tmp_W1, tmp_W2, tmp_b1, tmp_b2 = initialize_model(tmp_N,tmp_V)
print(f"tmp_W1.shape {tmp_W1.shape}")
print(f"tmp_W2.shape {tmp_W2.shape}")
print(f"tmp_b1.shape {tmp_b1.shape}")
print(f"tmp_b2.shape {tmp_b2.shape}")
tmp_z, tmp_h = forward_prop(tmp_x, tmp_W1, tmp_W2, tmp_b1, tmp_b2)
print(f"tmp_z.shape: {tmp_z.shape}")
print(f"tmp_h.shape: {tmp_h.shape}")
tmp_yhat = softmax(tmp_z)
print(f"tmp_yhat.shape: {tmp_yhat.shape}")
tmp_cost = compute_cost(tmp_y, tmp_yhat, tmp_batch_size)
print("call compute_cost")
print(f"tmp_cost {tmp_cost:.4f}")
```
##### Expected output
```CPP
tmp_x.shape (5778, 4)
tmp_y.shape (5778, 4)
tmp_W1.shape (50, 5778)
tmp_W2.shape (5778, 50)
tmp_b1.shape (50, 1)
tmp_b2.shape (5778, 1)
tmp_z.shape: (5778, 4)
tmp_h.shape: (50, 4)
tmp_yhat.shape: (5778, 4)
call compute_cost
tmp_cost 9.9560
```
<a name='2.4'></a>
## 2.4 Training the Model - Backpropagation
<a name='ex-04'></a>
### Exercise 04
Now that you have understood how the CBOW model works, you will train it. <br>
You created a function for the forward propagation. Now you will implement a function that computes the gradients to backpropagate the errors.
```
# UNQ_C4 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: back_prop
def back_prop(x, yhat, y, h, W1, W2, b1, b2, batch_size):
'''
Inputs:
x: average one hot vector for the context
yhat: prediction (estimate of y)
y: target vector
h: hidden vector (see eq. 1)
W1, W2, b1, b2: matrices and biases
batch_size: batch size
Outputs:
grad_W1, grad_W2, grad_b1, grad_b2: gradients of matrices and biases
'''
### START CODE HERE (Replace instanes of 'None' with your code) ###
# Compute l1 as W2^T (Yhat - Y)
# Re-use it whenever you see W2^T (Yhat - Y) used to compute a gradient
l1 = np.dot(W2.T,(yhat-y))
# Apply relu to l1
l1 = np.maximum(0, l1)
# Compute the gradient of W1
grad_W1 = (1/batch_size)*np.dot(l1,x.T)
# Compute the gradient of W2
grad_W2 = (1/batch_size)*np.dot(yhat-y,h.T)
# Compute the gradient of b1
grad_b1 = np.sum((1/batch_size)*np.dot(l1,x.T),axis=1,keepdims=True)
# Compute the gradient of b2
grad_b2 = np.sum((1/batch_size)*np.dot(yhat-y,h.T),axis=1,keepdims=True)
### END CODE HERE ###
return grad_W1, grad_W2, grad_b1, grad_b2
# Test the function
tmp_C = 2
tmp_N = 50
tmp_batch_size = 4
tmp_word2Ind, tmp_Ind2word = get_dict(data)
tmp_V = len(word2Ind)
# get a batch of data
tmp_x, tmp_y = next(get_batches(data, tmp_word2Ind, tmp_V,tmp_C, tmp_batch_size))
print("get a batch of data")
print(f"tmp_x.shape {tmp_x.shape}")
print(f"tmp_y.shape {tmp_y.shape}")
print()
print("Initialize weights and biases")
tmp_W1, tmp_W2, tmp_b1, tmp_b2 = initialize_model(tmp_N,tmp_V)
print(f"tmp_W1.shape {tmp_W1.shape}")
print(f"tmp_W2.shape {tmp_W2.shape}")
print(f"tmp_b1.shape {tmp_b1.shape}")
print(f"tmp_b2.shape {tmp_b2.shape}")
print()
print("Forwad prop to get z and h")
tmp_z, tmp_h = forward_prop(tmp_x, tmp_W1, tmp_W2, tmp_b1, tmp_b2)
print(f"tmp_z.shape: {tmp_z.shape}")
print(f"tmp_h.shape: {tmp_h.shape}")
print()
print("Get yhat by calling softmax")
tmp_yhat = softmax(tmp_z)
print(f"tmp_yhat.shape: {tmp_yhat.shape}")
tmp_m = (2*tmp_C)
tmp_grad_W1, tmp_grad_W2, tmp_grad_b1, tmp_grad_b2 = back_prop(tmp_x, tmp_yhat, tmp_y, tmp_h, tmp_W1, tmp_W2, tmp_b1, tmp_b2, tmp_batch_size)
print()
print("call back_prop")
print(f"tmp_grad_W1.shape {tmp_grad_W1.shape}")
print(f"tmp_grad_W2.shape {tmp_grad_W2.shape}")
print(f"tmp_grad_b1.shape {tmp_grad_b1.shape}")
print(f"tmp_grad_b2.shape {tmp_grad_b2.shape}")
```
##### Expected output
```CPP
get a batch of data
tmp_x.shape (5778, 4)
tmp_y.shape (5778, 4)
Initialize weights and biases
tmp_W1.shape (50, 5778)
tmp_W2.shape (5778, 50)
tmp_b1.shape (50, 1)
tmp_b2.shape (5778, 1)
Forwad prop to get z and h
tmp_z.shape: (5778, 4)
tmp_h.shape: (50, 4)
Get yhat by calling softmax
tmp_yhat.shape: (5778, 4)
call back_prop
tmp_grad_W1.shape (50, 5778)
tmp_grad_W2.shape (5778, 50)
tmp_grad_b1.shape (50, 1)
tmp_grad_b2.shape (5778, 1)
```
<a name='2.5'></a>
## Gradient Descent
<a name='ex-05'></a>
### Exercise 05
Now that you have implemented a function to compute the gradients, you will implement batch gradient descent over your training set.
**Hint:** For that, you will use `initialize_model` and the `back_prop` functions which you just created (and the `compute_cost` function). You can also use the provided `get_batches` helper function:
```for x, y in get_batches(data, word2Ind, V, C, batch_size):```
```...```
Also: print the cost after each batch is processed (use batch size = 128)
```
# UNQ_C5 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: gradient_descent
def gradient_descent(data, word2Ind, N, V, num_iters, alpha=0.03):
'''
This is the gradient_descent function
Inputs:
data: text
word2Ind: words to Indices
N: dimension of hidden vector
V: dimension of vocabulary
num_iters: number of iterations
Outputs:
W1, W2, b1, b2: updated matrices and biases
'''
W1, W2, b1, b2 = initialize_model(N,V, random_seed=282)
batch_size = 128
iters = 0
C = 2
for x, y in get_batches(data, word2Ind, V, C, batch_size):
### START CODE HERE (Replace instances of 'None' with your own code) ###
# Get z and h
z, h = forward_prop(x, W1, W2, b1, b2)
# Get yhat
yhat = softmax(z)
# Get cost
cost = compute_cost(y, yhat, batch_size)
if ( (iters+1) % 10 == 0):
print(f"iters: {iters + 1} cost: {cost:.6f}")
# Get gradients
grad_W1, grad_W2, grad_b1, grad_b2 = back_prop(x, yhat, y, h, W1, W2, b1, b2, batch_size)
# Update weights and biases
W1 = W1 - alpha * grad_W1
W2 = W2 - alpha * grad_W2
b1 = b1 - alpha * grad_b1
b2 = b2 - alpha * grad_b2
### END CODE HERE ###
iters += 1
if iters == num_iters:
break
if iters % 100 == 0:
alpha *= 0.66
return W1, W2, b1, b2
# test your function
C = 2
N = 50
word2Ind, Ind2word = get_dict(data)
V = len(word2Ind)
num_iters = 150
print("Call gradient_descent")
W1, W2, b1, b2 = gradient_descent(data, word2Ind, N, V, num_iters)
```
##### Expected Output
```CPP
iters: 10 cost: 0.789141
iters: 20 cost: 0.105543
iters: 30 cost: 0.056008
iters: 40 cost: 0.038101
iters: 50 cost: 0.028868
iters: 60 cost: 0.023237
iters: 70 cost: 0.019444
iters: 80 cost: 0.016716
iters: 90 cost: 0.014660
iters: 100 cost: 0.013054
iters: 110 cost: 0.012133
iters: 120 cost: 0.011370
iters: 130 cost: 0.010698
iters: 140 cost: 0.010100
iters: 150 cost: 0.009566
```
Your numbers may differ a bit depending on which version of Python you're using.
<a name='3'></a>
## 3.0 Visualizing the word vectors
In this part you will visualize the word vectors trained using the function you just coded above.
```
# visualizing the word vectors here
from matplotlib import pyplot
%config InlineBackend.figure_format = 'svg'
words = ['king', 'queen','lord','man', 'woman','dog','wolf',
'rich','happy','sad']
embs = (W1.T + W2)/2.0
# given a list of words and the embeddings, it returns a matrix with all the embeddings
idx = [word2Ind[word] for word in words]
X = embs[idx, :]
print(X.shape, idx) # X.shape: Number of words of dimension N each
result= compute_pca(X, 2)
pyplot.scatter(result[:, 0], result[:, 1])
for i, word in enumerate(words):
pyplot.annotate(word, xy=(result[i, 0], result[i, 1]))
pyplot.show()
```
You can see that man and king are next to each other. However, we have to be careful with the interpretation of this projected word vectors, since the PCA depends on the projection -- as shown in the following illustration.
```
result= compute_pca(X, 4)
pyplot.scatter(result[:, 3], result[:, 1])
for i, word in enumerate(words):
pyplot.annotate(word, xy=(result[i, 3], result[i, 1]))
pyplot.show()
```
| github_jupyter |
## Face and Facial Keypoint detection
After you've trained a neural network to detect facial keypoints, you can then apply this network to *any* image that includes faces. The neural network expects a Tensor of a certain size as input and, so, to detect any face, you'll first have to do some pre-processing.
1. Detect all the faces in an image using a face detector (we'll be using a Haar Cascade detector in this notebook).
2. Pre-process those face images so that they are grayscale, and transformed to a Tensor of the input size that your net expects. This step will be similar to the `data_transform` you created and applied in Notebook 2, whose job was tp rescale, normalize, and turn any iimage into a Tensor to be accepted as input to your CNN.
3. Use your trained model to detect facial keypoints on the image.
---
In the next python cell we load in required libraries for this section of the project.
```
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
%matplotlib inline
```
#### Select an image
Select an image to perform facial keypoint detection on; you can select any image of faces in the `images/` directory.
```
import cv2
# load in color image for face detection
image = cv2.imread('images/obamas.jpg')
# switch red and blue color channels
# --> by default OpenCV assumes BLUE comes first, not RED as in many images
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
# plot the image
fig = plt.figure(figsize=(9,9))
plt.imshow(image)
```
## Detect all faces in an image
Next, you'll use one of OpenCV's pre-trained Haar Cascade classifiers, all of which can be found in the `detector_architectures/` directory, to find any faces in your selected image.
In the code below, we loop over each face in the original image and draw a red square on each face (in a copy of the original image, so as not to modify the original). You can even [add eye detections](https://docs.opencv.org/3.4.1/d7/d8b/tutorial_py_face_detection.html) as an *optional* exercise in using Haar detectors.
An example of face detection on a variety of images is shown below.
<img src='images/haar_cascade_ex.png' width=80% height=80%/>
```
# load in a haar cascade classifier for detecting frontal faces
face_cascade = cv2.CascadeClassifier('detector_architectures/haarcascade_frontalface_default.xml')
# run the detector
# the output here is an array of detections; the corners of each detection box
# if necessary, modify these parameters until you successfully identify every face in a given image
faces = face_cascade.detectMultiScale(image, 1.2, 2)
# make a copy of the original image to plot detections on
image_with_detections = image.copy()
# loop over the detected faces, mark the image where each face is found
for (x,y,w,h) in faces:
# draw a rectangle around each detected face
# you may also need to change the width of the rectangle drawn depending on image resolution
cv2.rectangle(image_with_detections,(x,y),(x+w,y+h),(255,0,0),3)
fig = plt.figure(figsize=(9,9))
plt.imshow(image_with_detections)
```
## Loading in a trained model
Once you have an image to work with (and, again, you can select any image of faces in the `images/` directory), the next step is to pre-process that image and feed it into your CNN facial keypoint detector.
First, load your best model by its filename.
```
import torch
from models import Net
net = Net()
## TODO: load the best saved model parameters (by your path name)
## You'll need to un-comment the line below and add the correct name for *your* saved model
net.load_state_dict(torch.load('saved_models/keypoints_model.pt'))
## print out your net and prepare it for testing (uncomment the line below)
net.eval()
```
## Keypoint detection
Now, we'll loop over each detected face in an image (again!) only this time, you'll transform those faces in Tensors that your CNN can accept as input images.
### TODO: Transform each detected face into an input Tensor
You'll need to perform the following steps for each detected face:
1. Convert the face from RGB to grayscale
2. Normalize the grayscale image so that its color range falls in [0,1] instead of [0,255]
3. Rescale the detected face to be the expected square size for your CNN (224x224, suggested)
4. Reshape the numpy image into a torch image.
**Hint**: The sizes of faces detected by a Haar detector and the faces your network has been trained on are of different sizes. If you find that your model is generating keypoints that are too small for a given face, try adding some padding to the detected `roi` before giving it as input to your model.
You may find it useful to consult to transformation code in `data_load.py` to help you perform these processing steps.
### TODO: Detect and display the predicted keypoints
After each face has been appropriately converted into an input Tensor for your network to see as input, you can apply your `net` to each face. The ouput should be the predicted the facial keypoints. These keypoints will need to be "un-normalized" for display, and you may find it helpful to write a helper function like `show_keypoints`. You should end up with an image like the following with facial keypoints that closely match the facial features on each individual face:
<img src='images/michelle_detected.png' width=30% height=30%/>
```
image_copy = np.copy(image)
i = 0
# loop over the detected faces from your haar cascade
for (x,y,w,h) in faces:
i =+1
plt.figure(figsize=(20,10))
ax = plt.subplot(1, len(faces), i)
# Select the region of interest that is the face in the image
a = 50
roi = image_copy[y-a:y+h+a, x-a:x+w+a]
## TODO: Convert the face region from RGB to grayscale
gray = cv2.cvtColor(roi, cv2.COLOR_RGB2GRAY)
## TODO: Normalize the grayscale image so that its color range falls in [0,1] instead of [0,255]
gray = gray/255
## TODO: Rescale the detected face to be the expected square size for your CNN (224x224, suggested)
gray = cv2.resize(gray,(224,224))
## TODO: Reshape the numpy image shape (H x W x C) into a torch image shape (C x H x W)
gray = np.expand_dims(gray,0)
tensor = torch.FloatTensor(np.array([gray]))
net.eval()
## TODO: Make facial keypoint predictions using your loaded, trained network
predicted_key_pts = net(tensor)
predicted_key_pts = predicted_key_pts.view(predicted_key_pts.size()[0], 68, -1)
predicted_key_pts = predicted_key_pts.data
predicted_key_pts = predicted_key_pts.numpy()[0]
## TODO: Display each detected face and the corresponding keypoints
# undo normalization of keypoints
predicted_key_pts = predicted_key_pts*50.0+100
roi = cv2.resize(roi,(224,224))
plt.imshow(roi)
plt.scatter(predicted_key_pts[:, 0], predicted_key_pts[:, 1], s=20, marker='.', c='m')
plt.axis("off")
plt.show()
```
| github_jupyter |
# Logistic Regression with a Neural Network mindset
Welcome to your first (required) programming assignment! You will build a logistic regression classifier to recognize cats. This assignment will step you through how to do this with a Neural Network mindset, and so will also hone your intuitions about deep learning.
**Instructions:**
- Do not use loops (for/while) in your code, unless the instructions explicitly ask you to do so.
**You will learn to:**
- Build the general architecture of a learning algorithm, including:
- Initializing parameters
- Calculating the cost function and its gradient
- Using an optimization algorithm (gradient descent)
- Gather all three functions above into a main model function, in the right order.
## 1 - Packages ##
First, let's run the cell below to import all the packages that you will need during this assignment.
- [numpy](www.numpy.org) is the fundamental package for scientific computing with Python.
- [h5py](http://www.h5py.org) is a common package to interact with a dataset that is stored on an H5 file.
- [matplotlib](http://matplotlib.org) is a famous library to plot graphs in Python.
- [PIL](http://www.pythonware.com/products/pil/) and [scipy](https://www.scipy.org/) are used here to test your model with your own picture at the end.
```
import numpy as np
import matplotlib.pyplot as plt
import h5py
import scipy
from PIL import Image
from scipy import ndimage
from lr_utils import load_dataset
%matplotlib inline
```
## 2 - Overview of the Problem set ##
**Problem Statement**: You are given a dataset ("data.h5") containing:
- a training set of m_train images labeled as cat (y=1) or non-cat (y=0)
- a test set of m_test images labeled as cat or non-cat
- each image is of shape (num_px, num_px, 3) where 3 is for the 3 channels (RGB). Thus, each image is square (height = num_px) and (width = num_px).
You will build a simple image-recognition algorithm that can correctly classify pictures as cat or non-cat.
Let's get more familiar with the dataset. Load the data by running the following code.
```
# Loading the data (cat/non-cat)
train_set_x_orig, train_set_y, test_set_x_orig, test_set_y, classes = load_dataset()
```
We added "_orig" at the end of image datasets (train and test) because we are going to preprocess them. After preprocessing, we will end up with train_set_x and test_set_x (the labels train_set_y and test_set_y don't need any preprocessing).
Each line of your train_set_x_orig and test_set_x_orig is an array representing an image. You can visualize an example by running the following code. Feel free also to change the `index` value and re-run to see other images.
```
# Example of a picture
index = 25
plt.imshow(train_set_x_orig[index])
print ("y = " + str(train_set_y[:, index]) + ", it's a '" + classes[np.squeeze(train_set_y[:, index])].decode("utf-8") + "' picture.")
```
Many software bugs in deep learning come from having matrix/vector dimensions that don't fit. If you can keep your matrix/vector dimensions straight you will go a long way toward eliminating many bugs.
**Exercise:** Find the values for:
- m_train (number of training examples)
- m_test (number of test examples)
- num_px (= height = width of a training image)
Remember that `train_set_x_orig` is a numpy-array of shape (m_train, num_px, num_px, 3). For instance, you can access `m_train` by writing `train_set_x_orig.shape[0]`.
```
### START CODE HERE ### (≈ 3 lines of code)
m_train = None
m_test = None
num_px = None
m_train = train_set_x_orig.shape[0]
m_test = test_set_x_orig.shape[0]
num_px = train_set_x_orig.shape[2]
### END CODE HERE ###
print ("Number of training examples: m_train = " + str(m_train))
print ("Number of testing examples: m_test = " + str(m_test))
print ("Height/Width of each image: num_px = " + str(num_px))
print ("Each image is of size: (" + str(num_px) + ", " + str(num_px) + ", 3)")
print ("train_set_x shape: " + str(train_set_x_orig.shape))
print ("train_set_y shape: " + str(train_set_y.shape))
print ("test_set_x shape: " + str(test_set_x_orig.shape))
print ("test_set_y shape: " + str(test_set_y.shape))
```
**Expected Output for m_train, m_test and num_px**:
<table style="width:15%">
<tr>
<td>**m_train**</td>
<td> 209 </td>
</tr>
<tr>
<td>**m_test**</td>
<td> 50 </td>
</tr>
<tr>
<td>**num_px**</td>
<td> 64 </td>
</tr>
</table>
For convenience, you should now reshape images of shape (num_px, num_px, 3) in a numpy-array of shape (num_px $*$ num_px $*$ 3, 1). After this, our training (and test) dataset is a numpy-array where each column represents a flattened image. There should be m_train (respectively m_test) columns.
**Exercise:** Reshape the training and test data sets so that images of size (num_px, num_px, 3) are flattened into single vectors of shape (num\_px $*$ num\_px $*$ 3, 1).
A trick when you want to flatten a matrix X of shape (a,b,c,d) to a matrix X_flatten of shape (b$*$c$*$d, a) is to use:
```python
X_flatten = X.reshape(X.shape[0], -1).T # X.T is the transpose of X
```
```
# Reshape the training and test examples
### START CODE HERE ### (≈ 2 lines of code)
train_set_x_flatten = None
test_set_x_flatten = None
train_set_x_flatten = train_set_x_orig.reshape(train_set_x_orig.shape[0], -1).T
test_set_x_flatten = test_set_x_orig.reshape(test_set_x_orig.shape[0], -1).T
### END CODE HERE ###
print ("train_set_x_flatten shape: " + str(train_set_x_flatten.shape))
print ("train_set_y shape: " + str(train_set_y.shape))
print ("test_set_x_flatten shape: " + str(test_set_x_flatten.shape))
print ("test_set_y shape: " + str(test_set_y.shape))
print ("sanity check after reshaping: " + str(train_set_x_flatten[0:5,0]))
```
**Expected Output**:
<table style="width:35%">
<tr>
<td>**train_set_x_flatten shape**</td>
<td> (12288, 209)</td>
</tr>
<tr>
<td>**train_set_y shape**</td>
<td>(1, 209)</td>
</tr>
<tr>
<td>**test_set_x_flatten shape**</td>
<td>(12288, 50)</td>
</tr>
<tr>
<td>**test_set_y shape**</td>
<td>(1, 50)</td>
</tr>
<tr>
<td>**sanity check after reshaping**</td>
<td>[17 31 56 22 33]</td>
</tr>
</table>
To represent color images, the red, green and blue channels (RGB) must be specified for each pixel, and so the pixel value is actually a vector of three numbers ranging from 0 to 255.
One common preprocessing step in machine learning is to center and standardize your dataset, meaning that you substract the mean of the whole numpy array from each example, and then divide each example by the standard deviation of the whole numpy array. But for picture datasets, it is simpler and more convenient and works almost as well to just divide every row of the dataset by 255 (the maximum value of a pixel channel).
<!-- During the training of your model, you're going to multiply weights and add biases to some initial inputs in order to observe neuron activations. Then you backpropogate with the gradients to train the model. But, it is extremely important for each feature to have a similar range such that our gradients don't explode. You will see that more in detail later in the lectures. !-->
Let's standardize our dataset.
```
train_set_x = train_set_x_flatten/255.
test_set_x = test_set_x_flatten/255.
```
<font color='blue'>
**What you need to remember:**
Common steps for pre-processing a new dataset are:
- Figure out the dimensions and shapes of the problem (m_train, m_test, num_px, ...)
- Reshape the datasets such that each example is now a vector of size (num_px \* num_px \* 3, 1)
- "Standardize" the data
## 3 - General Architecture of the learning algorithm ##
It's time to design a simple algorithm to distinguish cat images from non-cat images.
You will build a Logistic Regression, using a Neural Network mindset. The following Figure explains why **Logistic Regression is actually a very simple Neural Network!**
<img src="images/LogReg_kiank.png" style="width:650px;height:400px;">
**Mathematical expression of the algorithm**:
For one example $x^{(i)}$:
$$z^{(i)} = w^T x^{(i)} + b \tag{1}$$
$$\hat{y}^{(i)} = a^{(i)} = sigmoid(z^{(i)})\tag{2}$$
$$ \mathcal{L}(a^{(i)}, y^{(i)}) = - y^{(i)} \log(a^{(i)}) - (1-y^{(i)} ) \log(1-a^{(i)})\tag{3}$$
The cost is then computed by summing over all training examples:
$$ J = \frac{1}{m} \sum_{i=1}^m \mathcal{L}(a^{(i)}, y^{(i)})\tag{6}$$
**Key steps**:
In this exercise, you will carry out the following steps:
- Initialize the parameters of the model
- Learn the parameters for the model by minimizing the cost
- Use the learned parameters to make predictions (on the test set)
- Analyse the results and conclude
## 4 - Building the parts of our algorithm ##
The main steps for building a Neural Network are:
1. Define the model structure (such as number of input features)
2. Initialize the model's parameters
3. Loop:
- Calculate current loss (forward propagation)
- Calculate current gradient (backward propagation)
- Update parameters (gradient descent)
You often build 1-3 separately and integrate them into one function we call `model()`.
### 4.1 - Helper functions
**Exercise**: Using your code from "Python Basics", implement `sigmoid()`. As you've seen in the figure above, you need to compute $sigmoid( w^T x + b) = \frac{1}{1 + e^{-(w^T x + b)}}$ to make predictions. Use np.exp().
```
# GRADED FUNCTION: sigmoid
import numpy as np
def sigmoid(z):
"""
Compute the sigmoid of z
Arguments:
z -- A scalar or numpy array of any size.
Return:
s -- sigmoid(z)
"""
### START CODE HERE ### (≈ 1 line of code)
s = None
### END CODE HERE ###
s = 1/(1+np.exp(-z))
return s
print ("sigmoid([0, 2]) = " + str(sigmoid(np.array([0,2]))))
```
**Expected Output**:
<table>
<tr>
<td>**sigmoid([0, 2])**</td>
<td> [ 0.5 0.88079708]</td>
</tr>
</table>
### 4.2 - Initializing parameters
**Exercise:** Implement parameter initialization in the cell below. You have to initialize w as a vector of zeros. If you don't know what numpy function to use, look up np.zeros() in the Numpy library's documentation.
```
# GRADED FUNCTION: initialize_with_zeros
def initialize_with_zeros(dim):
"""
This function creates a vector of zeros of shape (dim, 1) for w and initializes b to 0.
Argument:
dim -- size of the w vector we want (or number of parameters in this case)
Returns:
w -- initialized vector of shape (dim, 1)
b -- initialized scalar (corresponds to the bias)
"""
### START CODE HERE ### (≈ 1 line of code)
w = None
b = None
w = np.zeros((dim,1))
b = 0
### END CODE HERE ###
assert(w.shape == (dim, 1))
assert(isinstance(b, float) or isinstance(b, int))
return w, b
dim = 2
w, b = initialize_with_zeros(dim)
print ("w = " + str(w))
print ("b = " + str(b))
```
**Expected Output**:
<table style="width:15%">
<tr>
<td> ** w ** </td>
<td> [[ 0.]
[ 0.]] </td>
</tr>
<tr>
<td> ** b ** </td>
<td> 0 </td>
</tr>
</table>
For image inputs, w will be of shape (num_px $\times$ num_px $\times$ 3, 1).
### 4.3 - Forward and Backward propagation
Now that your parameters are initialized, you can do the "forward" and "backward" propagation steps for learning the parameters.
**Exercise:** Implement a function `propagate()` that computes the cost function and its gradient.
**Hints**:
Forward Propagation:
- You get X
- You compute $A = \sigma(w^T X + b) = (a^{(1)}, a^{(2)}, ..., a^{(m-1)}, a^{(m)})$
- You calculate the cost function: $J = -\frac{1}{m}\sum_{i=1}^{m}y^{(i)}\log(a^{(i)})+(1-y^{(i)})\log(1-a^{(i)})$
Here are the two formulas you will be using:
$$ \frac{\partial J}{\partial w} = \frac{1}{m}X(A-Y)^T\tag{7}$$
$$ \frac{\partial J}{\partial b} = \frac{1}{m} \sum_{i=1}^m (a^{(i)}-y^{(i)})\tag{8}$$
```
# GRADED FUNCTION: propagate
def propagate(w, b, X, Y):
"""
Implement the cost function and its gradient for the propagation explained above
Arguments:
w -- weights, a numpy array of size (num_px * num_px * 3, 1)
b -- bias, a scalar
X -- data of size (num_px * num_px * 3, number of examples)
Y -- true "label" vector (containing 0 if non-cat, 1 if cat) of size (1, number of examples)
Return:
cost -- negative log-likelihood cost for logistic regression
dw -- gradient of the loss with respect to w, thus same shape as w
db -- gradient of the loss with respect to b, thus same shape as b
Tips:
- Write your code step by step for the propagation. np.log(), np.dot()
"""
m = X.shape[1]
# FORWARD PROPAGATION (FROM X TO COST)
### START CODE HERE ### (≈ 2 lines of code)
A = None # compute activation
cost = None # compute cost
A = sigmoid(np.dot(w.T,X)+b)
cost = (-1/m) * np.sum(Y * np.log(A) + (1-Y) * np.log(1-A))
### END CODE HERE ###
# BACKWARD PROPAGATION (TO FIND GRAD)
### START CODE HERE ### (≈ 2 lines of code)
dw = None
db = None
dw = (1/m)*np.dot(X,(A-Y).T)
db = (np.sum(A-Y))/m
### END CODE HERE ###
assert(dw.shape == w.shape)
assert(db.dtype == float)
cost = np.squeeze(cost)
assert(cost.shape == ())
grads = {"dw": dw,
"db": db}
return grads, cost
w, b, X, Y = np.array([[1.],[2.]]), 2., np.array([[1.,2.,-1.],[3.,4.,-3.2]]), np.array([[1,0,1]])
grads, cost = propagate(w, b, X, Y)
print ("dw = " + str(grads["dw"]))
print ("db = " + str(grads["db"]))
print ("cost = " + str(cost))
```
**Expected Output**:
<table style="width:50%">
<tr>
<td> ** dw ** </td>
<td> [[ 0.99845601]
[ 2.39507239]]</td>
</tr>
<tr>
<td> ** db ** </td>
<td> 0.00145557813678 </td>
</tr>
<tr>
<td> ** cost ** </td>
<td> 5.801545319394553 </td>
</tr>
</table>
### 4.4 - Optimization
- You have initialized your parameters.
- You are also able to compute a cost function and its gradient.
- Now, you want to update the parameters using gradient descent.
**Exercise:** Write down the optimization function. The goal is to learn $w$ and $b$ by minimizing the cost function $J$. For a parameter $\theta$, the update rule is $ \theta = \theta - \alpha \text{ } d\theta$, where $\alpha$ is the learning rate.
```
# GRADED FUNCTION: optimize
def optimize(w, b, X, Y, num_iterations, learning_rate, print_cost = False):
"""
This function optimizes w and b by running a gradient descent algorithm
Arguments:
w -- weights, a numpy array of size (num_px * num_px * 3, 1)
b -- bias, a scalar
X -- data of shape (num_px * num_px * 3, number of examples)
Y -- true "label" vector (containing 0 if non-cat, 1 if cat), of shape (1, number of examples)
num_iterations -- number of iterations of the optimization loop
learning_rate -- learning rate of the gradient descent update rule
print_cost -- True to print the loss every 100 steps
Returns:
params -- dictionary containing the weights w and bias b
grads -- dictionary containing the gradients of the weights and bias with respect to the cost function
costs -- list of all the costs computed during the optimization, this will be used to plot the learning curve.
Tips:
You basically need to write down two steps and iterate through them:
1) Calculate the cost and the gradient for the current parameters. Use propagate().
2) Update the parameters using gradient descent rule for w and b.
"""
costs = []
for i in range(num_iterations):
# Cost and gradient calculation (≈ 1-4 lines of code)
### START CODE HERE ###
grads, cost = propagate(w,b, X, Y)
### END CODE HERE ###
# Retrieve derivatives from grads
dw = grads["dw"]
db = grads["db"]
# update rule (≈ 2 lines of code)
### START CODE HERE ###
w = w - learning_rate*dw
b = b - learning_rate*db
### END CODE HERE ###
# Record the costs
if i % 100 == 0:
costs.append(cost)
# Print the cost every 100 training iterations
if print_cost and i % 100 == 0:
print ("Cost after iteration %i: %f" %(i, cost))
params = {"w": w,
"b": b}
grads = {"dw": dw,
"db": db}
return params, grads, costs
params, grads, costs = optimize(w, b, X, Y, num_iterations= 100, learning_rate = 0.009, print_cost = False)
print ("w = " + str(params["w"]))
print ("b = " + str(params["b"]))
print ("dw = " + str(grads["dw"]))
print ("db = " + str(grads["db"]))
```
**Expected Output**:
<table style="width:40%">
<tr>
<td> **w** </td>
<td>[[ 0.19033591]
[ 0.12259159]] </td>
</tr>
<tr>
<td> **b** </td>
<td> 1.92535983008 </td>
</tr>
<tr>
<td> **dw** </td>
<td> [[ 0.67752042]
[ 1.41625495]] </td>
</tr>
<tr>
<td> **db** </td>
<td> 0.219194504541 </td>
</tr>
</table>
**Exercise:** The previous function will output the learned w and b. We are able to use w and b to predict the labels for a dataset X. Implement the `predict()` function. There are two steps to computing predictions:
1. Calculate $\hat{Y} = A = \sigma(w^T X + b)$
2. Convert the entries of a into 0 (if activation <= 0.5) or 1 (if activation > 0.5), stores the predictions in a vector `Y_prediction`. If you wish, you can use an `if`/`else` statement in a `for` loop (though there is also a way to vectorize this).
```
# GRADED FUNCTION: predict
def predict(w, b, X):
'''
Predict whether the label is 0 or 1 using learned logistic regression parameters (w, b)
Arguments:
w -- weights, a numpy array of size (num_px * num_px * 3, 1)
b -- bias, a scalar
X -- data of size (num_px * num_px * 3, number of examples)
Returns:
Y_prediction -- a numpy array (vector) containing all predictions (0/1) for the examples in X
'''
m = X.shape[1]
Y_prediction = np.zeros((1,m))
w = w.reshape(X.shape[0], 1)
# Compute vector "A" predicting the probabilities of a cat being present in the picture
### START CODE HERE ### (≈ 1 line of code)
A = sigmoid(np.dot(w.T,X)+b)
### END CODE HERE ###
for i in range(A.shape[1]):
# Convert probabilities A[0,i] to actual predictions p[0,i]
### START CODE HERE ### (≈ 4 lines of code)
Y_prediction[0,i] = A[0,i]>0.5
### END CODE HERE ###
assert(Y_prediction.shape == (1, m))
return Y_prediction
w = np.array([[0.1124579],[0.23106775]])
b = -0.3
X = np.array([[1.,-1.1,-3.2],[1.2,2.,0.1]])
print ("predictions = " + str(predict(w, b, X)))
```
**Expected Output**:
<table style="width:30%">
<tr>
<td>
**predictions**
</td>
<td>
[[ 1. 1. 0.]]
</td>
</tr>
</table>
<font color='blue'>
**What to remember:**
You've implemented several functions that:
- Initialize (w,b)
- Optimize the loss iteratively to learn parameters (w,b):
- computing the cost and its gradient
- updating the parameters using gradient descent
- Use the learned (w,b) to predict the labels for a given set of examples
## 5 - Merge all functions into a model ##
You will now see how the overall model is structured by putting together all the building blocks (functions implemented in the previous parts) together, in the right order.
**Exercise:** Implement the model function. Use the following notation:
- Y_prediction_test for your predictions on the test set
- Y_prediction_train for your predictions on the train set
- w, costs, grads for the outputs of optimize()
```
# GRADED FUNCTION: model
def model(X_train, Y_train, X_test, Y_test, num_iterations = 2000, learning_rate = 0.5, print_cost = False):
"""
Builds the logistic regression model by calling the function you've implemented previously
Arguments:
X_train -- training set represented by a numpy array of shape (num_px * num_px * 3, m_train)
Y_train -- training labels represented by a numpy array (vector) of shape (1, m_train)
X_test -- test set represented by a numpy array of shape (num_px * num_px * 3, m_test)
Y_test -- test labels represented by a numpy array (vector) of shape (1, m_test)
num_iterations -- hyperparameter representing the number of iterations to optimize the parameters
learning_rate -- hyperparameter representing the learning rate used in the update rule of optimize()
print_cost -- Set to true to print the cost every 100 iterations
Returns:
d -- dictionary containing information about the model.
"""
### START CODE HERE ###
# initialize parameters with zeros (≈ 1 line of code)
w, b = initialize_with_zeros(X_train.shape[0])
# Gradient descent (≈ 1 line of code)
parameters, grads, costs = optimize(w, b, X_train, Y_train, num_iterations, learning_rate, print_cost)
# Retrieve parameters w and b from dictionary "parameters"
w = parameters["w"]
b = parameters["b"]
# Predict test/train set examples (≈ 2 lines of code)
Y_prediction_test = predict(w, b, X_test)
Y_prediction_train = predict(w,b ,X_train)
### END CODE HERE ###
# Print train/test Errors
print("train accuracy: {} %".format(100 - np.mean(np.abs(Y_prediction_train - Y_train)) * 100))
print("test accuracy: {} %".format(100 - np.mean(np.abs(Y_prediction_test - Y_test)) * 100))
d = {"costs": costs,
"Y_prediction_test": Y_prediction_test,
"Y_prediction_train" : Y_prediction_train,
"w" : w,
"b" : b,
"learning_rate" : learning_rate,
"num_iterations": num_iterations}
return d
```
Run the following cell to train your model.
```
d = model(train_set_x, train_set_y, test_set_x, test_set_y, num_iterations = 2000, learning_rate = 0.005, print_cost = True)
```
**Expected Output**:
<table style="width:40%">
<tr>
<td> **Cost after iteration 0 ** </td>
<td> 0.693147 </td>
</tr>
<tr>
<td> <center> $\vdots$ </center> </td>
<td> <center> $\vdots$ </center> </td>
</tr>
<tr>
<td> **Train Accuracy** </td>
<td> 99.04306220095694 % </td>
</tr>
<tr>
<td>**Test Accuracy** </td>
<td> 70.0 % </td>
</tr>
</table>
**Comment**: Training accuracy is close to 100%. This is a good sanity check: your model is working and has high enough capacity to fit the training data. Test error is 68%. It is actually not bad for this simple model, given the small dataset we used and that logistic regression is a linear classifier. But no worries, you'll build an even better classifier next week!
Also, you see that the model is clearly overfitting the training data. Later in this specialization you will learn how to reduce overfitting, for example by using regularization. Using the code below (and changing the `index` variable) you can look at predictions on pictures of the test set.
```
# Example of a picture that was wrongly classified.
index = 1
plt.imshow(test_set_x[:,index].reshape((num_px, num_px, 3)))
print ("y = " + str(test_set_y[0,index]) + ", you predicted that it is a \"" + classes[d["Y_prediction_test"][0,index]].decode("utf-8") + "\" picture.")
```
Let's also plot the cost function and the gradients.
```
# Plot learning curve (with costs)
costs = np.squeeze(d['costs'])
plt.plot(costs)
plt.ylabel('cost')
plt.xlabel('iterations (per hundreds)')
plt.title("Learning rate =" + str(d["learning_rate"]))
plt.show()
```
**Interpretation**:
You can see the cost decreasing. It shows that the parameters are being learned. However, you see that you could train the model even more on the training set. Try to increase the number of iterations in the cell above and rerun the cells. You might see that the training set accuracy goes up, but the test set accuracy goes down. This is called overfitting.
## 6 - Further analysis (optional/ungraded exercise) ##
Congratulations on building your first image classification model. Let's analyze it further, and examine possible choices for the learning rate $\alpha$.
#### Choice of learning rate ####
**Reminder**:
In order for Gradient Descent to work you must choose the learning rate wisely. The learning rate $\alpha$ determines how rapidly we update the parameters. If the learning rate is too large we may "overshoot" the optimal value. Similarly, if it is too small we will need too many iterations to converge to the best values. That's why it is crucial to use a well-tuned learning rate.
Let's compare the learning curve of our model with several choices of learning rates. Run the cell below. This should take about 1 minute. Feel free also to try different values than the three we have initialized the `learning_rates` variable to contain, and see what happens.
```
learning_rates = [0.01, 0.001, 0.0001]
models = {}
for i in learning_rates:
print ("learning rate is: " + str(i))
models[str(i)] = model(train_set_x, train_set_y, test_set_x, test_set_y, num_iterations = 1500, learning_rate = i, print_cost = False)
print ('\n' + "-------------------------------------------------------" + '\n')
for i in learning_rates:
plt.plot(np.squeeze(models[str(i)]["costs"]), label= str(models[str(i)]["learning_rate"]))
plt.ylabel('cost')
plt.xlabel('iterations (hundreds)')
legend = plt.legend(loc='upper center', shadow=True)
frame = legend.get_frame()
frame.set_facecolor('0.90')
plt.show()
```
**Interpretation**:
- Different learning rates give different costs and thus different predictions results.
- If the learning rate is too large (0.01), the cost may oscillate up and down. It may even diverge (though in this example, using 0.01 still eventually ends up at a good value for the cost).
- A lower cost doesn't mean a better model. You have to check if there is possibly overfitting. It happens when the training accuracy is a lot higher than the test accuracy.
- In deep learning, we usually recommend that you:
- Choose the learning rate that better minimizes the cost function.
- If your model overfits, use other techniques to reduce overfitting. (We'll talk about this in later videos.)
## 7 - Test with your own image (optional/ungraded exercise) ##
Congratulations on finishing this assignment. You can use your own image and see the output of your model. To do that:
1. Click on "File" in the upper bar of this notebook, then click "Open" to go on your Coursera Hub.
2. Add your image to this Jupyter Notebook's directory, in the "images" folder
3. Change your image's name in the following code
4. Run the code and check if the algorithm is right (1 = cat, 0 = non-cat)!
```
## START CODE HERE ## (PUT YOUR IMAGE NAME)
my_image = "my_image.jpg" # change this to the name of your image file
## END CODE HERE ##
# We preprocess the image to fit your algorithm.
fname = "images/" + my_image
image = np.array(ndimage.imread(fname, flatten=False))
my_image = scipy.misc.imresize(image, size=(num_px,num_px)).reshape((1, num_px*num_px*3)).T
my_predicted_image = predict(d["w"], d["b"], my_image)
plt.imshow(image)
print("y = " + str(np.squeeze(my_predicted_image)) + ", your algorithm predicts a \"" + classes[int(np.squeeze(my_predicted_image)),].decode("utf-8") + "\" picture.")
```
<font color='blue'>
**What to remember from this assignment:**
1. Preprocessing the dataset is important.
2. You implemented each function separately: initialize(), propagate(), optimize(). Then you built a model().
3. Tuning the learning rate (which is an example of a "hyperparameter") can make a big difference to the algorithm. You will see more examples of this later in this course!
Finally, if you'd like, we invite you to try different things on this Notebook. Make sure you submit before trying anything. Once you submit, things you can play with include:
- Play with the learning rate and the number of iterations
- Try different initialization methods and compare the results
- Test other preprocessings (center the data, or divide each row by its standard deviation)
Bibliography:
- http://www.wildml.com/2015/09/implementing-a-neural-network-from-scratch/
- https://stats.stackexchange.com/questions/211436/why-do-we-normalize-images-by-subtracting-the-datasets-image-mean-and-not-the-c
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# 畳み込みニューラルネットワーク (Convolutional Neural Networks)
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/images/cnn">
<img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/ja/tutorials/images/cnn.ipynb">
<img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/ja/tutorials/images/cnn.ipynb">
<img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
View source on GitHub</a>
</td>
</table>
このチュートリアルでは、MNIST の数の分類をするための、シンプルな[畳み込みニューラルネットワーク](https://developers.google.com/machine-learning/glossary/#convolutional_neural_network) (CNN: Convolutional Neural Network) の学習について説明します。このシンプルなネットワークは MNIST テストセットにおいて、99%以上の精度を達成します。このチュートリアルでは、[Keras Sequential API](https://www.tensorflow.org/guide/keras)を使用するため、ほんの数行のコードでモデルの作成と学習を行うことができます。
Note: GPU を使うことで CNN をより早く学習させることができます。もし、このノートブックを Colab で実行しているならば、*編集 -> ノートブックの設定 -> ハードウェアアクセラレータ -> GPU* から無料のGPUを有効にすることができます。
### TensorFlowのインポート
```
from __future__ import absolute_import, division, print_function, unicode_literals
!pip install tensorflow-gpu==2.0.0-beta1
import tensorflow as tf
from tensorflow.keras import datasets, layers, models
```
### MNISTデータセットのダウンロードと準備
```
(train_images, train_labels), (test_images, test_labels) = datasets.mnist.load_data()
train_images = train_images.reshape((60000, 28, 28, 1))
test_images = test_images.reshape((10000, 28, 28, 1))
# ピクセルの値を 0~1 の間に正規化
train_images, test_images = train_images / 255.0, test_images / 255.0
```
### 畳み込みの基礎部分の作成
下記の6行のコードは、一般的なパターンで畳み込みの基礎部分を定義しています: [Conv2D](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Conv2D) と [MaxPooling2D](https://www.tensorflow.org/api_docs/python/tf/keras/layers/MaxPool2D) レイヤーのスタック。
入力として、CNN はバッチサイズを無視して、shape (image_height, image_width, color_channels) のテンソルをとります。color channels について、MNIST は1つ (画像がグレースケールのため) の color channels がありますが、カラー画像には3つ (R, G, B) があります。この例では、MNIST 画像のフォーマットである shape (28, 28, 1) の入力を処理するように CNN を構成します。これを行うには、引数 `input_shape` を最初のレイヤーに渡します。
```
model = models.Sequential()
model.add(layers.Conv2D(32, (3, 3), activation='relu', input_shape=(28, 28, 1)))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
```
ここまでのモデルのアーキテクチャを表示してみましょう。
```
model.summary()
```
上記より、すべての Conv2D と MaxPooling2D レイヤーの出力は shape (height, width, channels) の 3D テンソルであることがわかります。width と height の寸法は、ネットワークが深くなるにつれて縮小する傾向があります。各 Conv2D レイヤーの出力チャネルの数は、第一引数 (例: 32 または 64) によって制御されます。通常、width とheight が縮小すると、各 Conv2D レイヤーにさらに出力チャネルを追加する余裕が (計算上) できます。
### 上に Dense レイヤーを追加
モデルを完成するために、(shape (3, 3, 64) の) 畳み込みの基礎部分からの最後の出力テンソルを、1つ以上の Dense レイヤーに入れて分類を実行します。現在の出力は 3D テンソルですが、Dense レイヤーは入力としてベクトル (1D) を取ります。まず、3D 出力を 1D に平滑化 (または展開) してから、最上部に1つ以上の Dense レイヤーを追加します。MNIST は 10 個の出力クラスを持ちます。そのため、我々は最後の Dense レイヤーの出力を 10 にし、softmax関数を使用します。
```
model.add(layers.Flatten())
model.add(layers.Dense(64, activation='relu'))
model.add(layers.Dense(10, activation='softmax'))
```
これが私たちのモデルの完全なアーキテクチャです。
```
model.summary()
```
ご覧のとおり、2 つの Dense レイヤーを通過する前に、(3, 3, 64) の出力は shape (576) のベクターに平滑化されました。
### モデルのコンパイルと学習
```
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(train_images, train_labels, epochs=5)
```
### モデルの評価
```
test_loss, test_acc = model.evaluate(test_images, test_labels)
print(test_acc)
```
ご覧のとおり、我々のシンプルな CNN は 99% 以上のテスト精度を達成しています。数行のコードにしては悪くありません!違うスタイルでの CNN の書き方 (Keras Subclassing API や GradientTape を使ったもの) については[ここ](https://github.com/tensorflow/docs/blob/master/site/en/tutorials/quickstart/advanced.ipynb)を参照してください。
| github_jupyter |
# 函数
- 函数可以用来定义可重复代码,组织和简化
- 一般来说一个函数在实际开发中为一个小功能
- 一个类为一个大功能
- 同样函数的长度不要超过一屏
Python中的所有函数实际上都是有返回值(return None),
如果你没有设置return,那么Python将不显示None.
如果你设置return,那么将返回出return这个值.
```
def HJN():
print('Hello')
return 1000
b=HJN()
print(b)
HJN
def panduan(number):
if number % 2 == 0:
print('O')
else:
print('J')
panduan(number=1)
panduan(2)
```
## 定义一个函数
def function_name(list of parameters):
do something

- 以前使用的random 或者range 或者print.. 其实都是函数或者类
函数的参数如果有默认值的情况,当你调用该函数的时候:
可以不给予参数值,那么就会走该参数的默认值
否则的话,就走你给予的参数值.
```
import random
def hahah():
n = random.randint(0,5)
while 1:
N = eval(input('>>'))
if n == N:
print('smart')
break
elif n < N:
print('太小了')
elif n > N:
print('太大了')
```
## 调用一个函数
- functionName()
- "()" 就代表调用
```
def H():
print('hahaha')
def B():
H()
B()
def A(f):
f()
A(B)
```

## 带返回值和不带返回值的函数
- return 返回的内容
- return 返回多个值
- 一般情况下,在多个函数协同完成一个功能的时候,那么将会有返回值

- 当然也可以自定义返回None
## EP:

```
def main():
print(min(min(5,6),(51,6)))
def min(n1,n2):
a = n1
if n2 < a:
a = n2
main()
```
## 类型和关键字参数
- 普通参数
- 多个参数
- 默认值参数
- 不定长参数
## 普通参数
## 多个参数
## 默认值参数
## 强制命名
```
def U(str_):
xiaoxie = 0
for i in str_:
ASCII = ord(i)
if 97<=ASCII<=122:
xiaoxie +=1
elif xxxx:
daxie += 1
elif xxxx:
shuzi += 1
return xiaoxie,daxie,shuzi
U('HJi12')
```
## 不定长参数
- \*args
> - 不定长,来多少装多少,不装也是可以的
- 返回的数据类型是元组
- args 名字是可以修改的,只是我们约定俗成的是args
- \**kwargs
> - 返回的字典
- 输入的一定要是表达式(键值对)
- name,\*args,name2,\**kwargs 使用参数名
```
def TT(a,b)
def TT(*args,**kwargs):
print(kwargs)
print(args)
TT(1,2,3,4,6,a=100,b=1000)
{'key':'value'}
TT(1,2,4,5,7,8,9,)
def B(name1,nam3):
pass
B(name1=100,2)
def sum_(*args,A='sum'):
res = 0
count = 0
for i in args:
res +=i
count += 1
if A == "sum":
return res
elif A == "mean":
mean = res / count
return res,mean
else:
print(A,'还未开放')
sum_(-1,0,1,4,A='var')
'aHbK134'.__iter__
b = 'asdkjfh'
for i in b :
print(i)
2,5
2 + 22 + 222 + 2222 + 22222
```
## 变量的作用域
- 局部变量 local
- 全局变量 global
- globals 函数返回一个全局变量的字典,包括所有导入的变量
- locals() 函数会以字典类型返回当前位置的全部局部变量。
```
a = 1000
b = 10
def Y():
global a,b
a += 100
print(a)
Y()
def YY(a1):
a1 += 100
print(a1)
YY(a)
print(a)
```
## 注意:
- global :在进行赋值操作的时候需要声明
- 官方解释:This is because when you make an assignment to a variable in a scope, that variable becomes local to that scope and shadows any similarly named variable in the outer scope.
- 
# Homework
- 1

```
def getPentagonalNumber():
n=1
hang=0
for n in range(1,101):
s=n*(3*n-1)/2
print('%d'%s,end=' ')
hang +=1
if hang %10 ==0:
print( )
getPentagonalNumber()
```
- 2

```
def sumDigits(n):
while n > 10 and n< 1000:
b=n%10
c=n//10
d=c%10
f=c//10
print(b+d+f)
break
sumDigits(234)
```
- 3

```
def displaySortedNumbers():
num1,num2,num3=map(float,input('输入三个数:').split(','))
if num1>num2>num3:
print(num3,num2,num1)
if num1>num3>num2:
print(num2,num3,num1)
if num2>num1>num3:
print(num3,num1,num2)
if num2>num3>num1:
print(num1,num3,num2)
if num3>num1>num2:
print(num2,num1.num3)
if num3>num2>num1:
print(num1,num2,num3)
displaySortedNumbers()
```
- 4

```
def futureInvestmentValue(investmentAmount,monthlyInterestRate,years):
```
- 5

```
def printChars(ch1,ch2,numberPerLine):
count=0
while ch1!=chr(ord(ch2)+1):
print(ch1,end=' ')
i=ord(ch1)
i+=1
ch1=chr(i)
count+=1
if count%10==0:
print()
printChars('I','Z',10)
```
- 6

```
def numberOfDaysInAYear(year):
for year in range(2010,2021):
if year%4==0 and year%100!=0 or year%400==0:
print('%d年有366天'%year)
else:
print('%d年有365天'%year)
numberOfDaysInAYear(2013)
```
- 7

```
def distance(x1,y1,x2,y2):
x=(x1-x2)**2
y=(y1-y2)**2
jl=(x+y)**0.5
print(jl)
distance(2,5,6,3)
```
- 8

```
a=[]
for j in range(2,100):
for i in range(2,j):
if (j % i) == 0:
break
else:
a.append(j)
for z in a:
x=2**z-1
while x<=31:
print(z,x)
break
```
- 9


```
import time
import time
localtime = time.asctime(time.localtime(time.time()))
print("本地时间为 :", localtime)
2019 - 1970
```
- 10

```
import random
i=random.randrange(1,7)
j=random.randrange(1,7)
if i+j==2 or i+j==3 or i+j==12 :
print('you rolled %d+%d=%d'%(i,j,i+j))
print('you lose')
elif i+j==7 or i+j==11:
print('you rolled %d+%d=%d'%(i,j,i+j))
print('you win')
else:
print('you rolled %d+%d=%d'%(i,j,i+j))
print('point is %d'%(i+j))
n=i+j
i=random.randrange(1,7)
j=random.randrange(1,7)
for z in range(1,100):
if i+j==7:
print('you rolled %d+%d=%d'%(i,j,i+j))
print('you lose')
break
elif i+j==n :
print('you rolled %d+%d=%d'%(i,j,i+j))
print('you win')
break
else:
i=random.randrange(1,7)
j=random.randrange(1,7)
```
- 11
### 去网上寻找如何用Python代码发送邮件
```
import smtplib
from email.mime.text import MIMEText
from email.mime.multipart import MIMEMultipart
from email.header import Header
from email.utils import parseaddr, formataddr
# 格式化邮件地址
def formatAddr(s):
name, addr = parseaddr(s)
return formataddr((Header(name, 'utf-8').encode(), addr))
def sendMail(body):
smtp_server = 'smtp.163.com'
from_mail = 'baojingtongzhi@163.com'
mail_pass = 'xxx'
to_mail = ['962510244@qq.com', 'zhenliang369@163.com']
# 构造一个MIMEMultipart对象代表邮件本身
msg = MIMEMultipart()
# Header对中文进行转码
msg['From'] = formatAddr('管理员 <%s>' % from_mail).encode()
msg['To'] = ','.join(to_mail)
msg['Subject'] = Header('监控', 'utf-8').encode()
msg.attach(MIMEText(body, 'html', 'utf-8'))
try:
s = smtplib.SMTP()
s.connect(smtp_server, "25")
s.login(from_mail, mail_pass)
s.sendmail(from_mail, to_mail, msg.as_string()) # as_string()把MIMEText对象变成str
s.quit()
except smtplib.SMTPException as e:
print "Error: %s" % e
if __name__ == "__main__":
body = """
<h1>测试邮件</h1>
<h2 style="color:red">This is a test</h1>
"""
sendMail(body)
```
| github_jupyter |
# 第0章 そもそも量子コンピュータとは?
近年、マスコミでも「量子コンピュータ」というワードを耳にすることが多い。「名前だけは聞いたことあるけど、どんなものかはよくわからない…」そんな方のために、この章では、量子コンピュータの概要を説明する。
(なお、「量子」コンピュータの業界では、現在のコンピュータを「古典」コンピュータと呼んでおり、Qunatum Native Dojoでもそれに従う。)
## 量子コンピュータというアイディア
量子コンピュータのアイディア自体は古く、エッセイ「ご冗談でしょう、ファインマンさん」でも有名な物理学者リチャード・ファインマンが、1982年に「自然をシミュレーションしたければ、量子力学の原理でコンピュータを作らなくてはならない」と述べたことに端を発する[1]。そして、1985年にオックスフォード大学の物理学者デイビット・ドイチュによって量子コンピュータが理論的に定式化された[2]。
量子コンピュータと従来の古典コンピュータの最も異なる点は、その情報の表し方である。古典コンピュータの内部では、情報は0か1、どちらか1つの状態を取ることのできる「古典ビット」で表現される。これに対し、量子コンピュータの内部では、情報は0と1の両方の状態を**同時に**取ることのできる「量子ビット」で表現される(詳細はこれからQuantum Native Dojoで学んでいくのでご安心を)。量子コンピュータは、量子ビットを用いて多数の計算を同時に行い、その結果をうまく処理することで、古典コンピュータと比べて飛躍的に高速に計算を行える**場合がある**。
## どのように役に立つのか
さきほど**場合がある**と強調したのは、現実にある全ての問題(タスク)を、量子コンピュータが高速に計算できるわけではないからだ。むしろ、量子コンピュータの方が古典コンピュータより高速なのかどうか未だに分かっていない問題の方が圧倒的に多い。それでも、一部の問題については量子コンピュータの方が**現状の**古典コンピュータのアルゴリズムよりも高速であることが証明されている。その代表例が素因数分解である。
素因数分解は、$30 = 2\times3\times5$ などの簡単な場合なら暗算でも計算できるが、分解すべき整数の桁数が大きくなってくると、最高レベルの速度を持つスーパーコンピュータでさえ、その計算には年単位・あるいは宇宙の寿命ほどの時間が必要になる。この「解くのに時間がかかる」ことを利用したのが、現在の暗号通信・情報セキュリティに広く用いられているRSA暗号である。しかし、1995年に米国の数学者ピーター・ショアが、古典よりも**圧倒的に速く**素因数分解を行う量子アルゴリズムを見つけた[3]ことで、一気に量子コンピュータに注目が集まることになった。
素因数分解の他にも、量子コンピュータの方が現状の古典コンピュータよりも高速であると証明されている問題がいくつかある。例えば、整理されていないデータから目的のデータを探し出す探索問題(8-2節`グローバーのアルゴリズム`)や、連立一次方程式の解を求める問題(7-4節`Harrow-Hassidim-Lloydアルゴリズム`)などである。これらの問題は、流体解析・電磁気解析・機械学習など、現代社会を支える様々な科学技術計算に活用できる。さらに、万物は元をたどれば量子力学に従っているため、ファインマンやドイチュが考えたように、究極の自然現象シミュレーターとして量子コンピュータを活用し、物質設計や素材開発を行うことも考案されている。
このように、量子コンピュータによって世の中の全ての計算が高速化される訳でないものの、現代社会に与えるインパクトは計り知れないものがある。
## 量子誤り訂正
ここまで述べたのは、量子コンピュータの理論的な話である。理論的に速く計算できると証明できても、応用するには実際に計算を行うハードウェアが必要になる。量子コンピュータのハードウェアを製作する研究は世界中で広く行われているが、課題はまだまだ多いのが現状である。もっとも大きな課題の一つが、ノイズである。量子ビットは古典ビットと比べて磁場や温度揺らぎなどの外部ノイズを非常に受けやすく、保持していた情報をすぐに失ってしまう。2019年現在でも、数個〜数十個程度の量子ビットを連結し、より安定に長く動作させる方法を探っているような段階である。
そのようなノイズの問題を克服するために研究されているのが、量子誤り訂正の技術である。量子誤り訂正は、計算途中に生じた量子ビットの誤り(エラー)を検知し、本来の状態に訂正する技術で、理論的には様々な手法が提案されている(なお、我々が普段使っている古典コンピュータにも古典ビットの誤り訂正機能が搭載されている。この機能のおかけで、我々はPC内のデータが突然無くなることを気にせずに暮らせるのである)。しかし、量子誤り訂正は単なる量子ビットの作製・動作実現よりもはるかに技術的難易度が高く、誤り訂正機能を持った量子ビットを作るには少なくともあと10年は必要であると言われている。
そして、前項であげた量子コンピュータの様々な応用を実用的な規模で実行するには、誤り訂正機能を持った量子ビットが1000個単位で必要となることから、**真の量子コンピュータの現実的応用は数十年先になる**と考えられている。
## NISQ (Noisy Intermidiate-Scale Quantum) デバイスの時代
では我々が量子コンピュータの恩恵を受けるには、あと何十年も待たないといけないのだろうか。「そんなことで手をこまねいているわけにはいかない!」ということで、科学者たちは様々な方法で量子コンピュータの有用性を示そうと模索している。その中で現在最も注目されており、世界中で研究が進められているのが**Noisy Intermediate-Scale Quantum (NISQ)** デバイスという量子コンピュータである。
NISQデバイスの定義は「ノイズを含む(誤り訂正機能を持たない)、中規模(〜数百qubit)な量子デバイス」であり、これであれば数年以内の実用化が可能であると考えられている。NISQデバイスは、誤り訂正機能がないので限られた量子アルゴリズムしか実行できないものの、量子化学計算や機械学習といった領域で、現在の古典コンピュータを凌駕する性能を発揮すると予想されている(第4章・第5章・第6章参照)。
NISQデバイスまで含めれば、量子コンピュータが応用され始めるのはそう遠くない未来なのである。このQuantum Native Dojoを通して、そうした量子コンピュータ応用の最先端に触れるための知識を皆様に身につけていただければ幸いである。
## 参考文献
1. “Simulating physics with computers”, R. P. Feybmann, International Journal of Theoretical Physics **21**, 467 [(pdfリンク)](https://people.eecs.berkeley.edu/~christos/classics/Feynman.pdf)
2. “Quantum theory, the Church-Turing principle and the universal quantum computer” Proceedings of the Royal Society of London A **400**, 97 (1985) [(pdfリンク)](https://people.eecs.berkeley.edu/~christos/classics/Deutsch_quantum_theory.pdf)
3. “Polynomial-Time Algorithms for Prime Factorization and Discrete Logarithms on a Quantum Computer”, IAM J.Sci.Statist.Comput. **26** (1997) 1484 [(pdfリンク)](https://arxiv.org/pdf/quant-ph/9508027.pdf)
## 量子コンピュータの基礎から応用までに関するスライド
量子コンピュータの動作原理や応用アプリケーション、政府・企業の動きに関したさらに詳しい解説は、
[Quantum Summit 2019の講演をまとめたスライド](https://speakerdeck.com/qunasys/quantum-summit-2019)をご覧ください。
[](https://speakerdeck.com/qunasys/quantum-summit-2019)
---
## コラム:量子ビット・量子ゲート操作を物理的にどう実現するか
実際の量子コンピュータを構成する量子ビットはいったいどのように作られ、量子ゲート操作はどのように実行されているのだろうか。
量子ビットを実現する方法は1995年頃から複数の有望な方式(物理系)が提案されており、超伝導回路方式・イオントラップ方式・光方式などがある。各方式によって、現在実現できている量子ビット数や量子ビットの寿命(コヒーレンス時間)、エラー率等に違いがあり、世界各国で研究が盛んに進められている。
数々の量子ビット実現方式の中で、最も広く知られている方式は超伝導回路を用いた超伝導量子ビットである。これは1999年に当時NECに所属していた中村泰信(現東京大学教授)・蔡兆申(現東京理科大学教授)らによって世界で初めて製作された量子ビットで、超伝導物質を用いたジョセフソン接合と呼ばれる微細な構造を作ることで量子ビットを実現している。量子ゲート操作は、マイクロ波(電磁波の一種)のパルスをターゲットの量子ビットに送ることで実現される。また、量子ビットの測定は測定用の超伝導回路を量子ビットにつけることで行われる。
超伝導回路方式は、GoogleやRigetti conmputingが数十量子ビットの素子の開発を発表するなど、2019年3月現在で最も有望な量子コンピュータ実現方式であると言える。
量子ビットの実現方法について、より深く知りたい場合には以下を参考にされたい:
- Qmedia 量子コンピュータを実現するハードウェア
- (前編) https://www.qmedia.jp/making-quantum-hardware-1/
- (後編) https://www.qmedia.jp/making-quantum-hardware-2/
- レビュー論文:“Quantum Computing”, T. D. Ladd _et al._, Nature, **464**, 45 (2010). https://arxiv.org/abs/1009.2267
- Nielsen-Chuang 第7章 `Quantum computers: physical realization`
| github_jupyter |
# 1. Read the dataset
```
import pandas as pd
import seaborn as sns
data = pd.read_csv("pubg - Dr. Darshan Ingle.csv")
data.head()
```
# 2. Check the datatype of all the columns.
```
#checking datatype of all columns
data.dtypes
```
# 3. Find the summary of all the numerical columns and write your findings about it.
```
#Summary of all numerical columns
data.describe()
```
As seen above, describe() function gives the stats of only the numerical columns. Each column consists a "count" of 10000 values. The "mean" parameter indicates the mean value of all the columns. The "std" parameter represents standard deviation, which is the deviation from the mean. The "min" and "max" parameters represent the maximum and minimum values respectively of each column. For example, consider the column "killPlace". Now, 25% indicates that 25% of the players have the kill place of 24 or less. Same goes for 50% and 75%.
```
data['kills'].describe()
```
# 4. The average person kills how many players?
# 5. 99% of people have how many kills?
# 6. The most kills ever recorded are how much?
```
#Observations from "kills" column
average_kills = data['kills'].mean()
print("The average person kills: ", average_kills)
Q99 = data["kills"].quantile(0.99)
print("99% of people have", Q99, "kills")
max_kills = data['kills'].max()
print("The maximum kills ever recorded are: ",max_kills,"kills")
```
# 7. Print all the columns of the dataframe.
```
#printing all the columns of the dataframe
data.columns
```
# 8. Comment on distribution of the match's duration. Use seaborn.
```
#distribution of match duration
import matplotlib.pyplot as plt
plt.figure(figsize=(15,5))
import seaborn as sns
sns.distplot(data["matchDuration"]);
data["matchDuration"].describe()
```
As seen from the Seaborn's distplot above, the longest duration match played is between 1350 and 1400 seconds. The second longest duration is between 1850 and 1890 seconds.
# 9. Comment on distribution of the walk distance. Use seaborn.
```
#distribution of walk distance
%matplotlib inline
import matplotlib.pyplot as plt
plt.figure(figsize=(15,5))
import seaborn as sns
sns.distplot(data["walkDistance"],bins=10);
```
The maximum walk distance seen from the above Seaborn's distplot is approx. 1000 mts
# 10. Plot distribution of the match's duration vs walk distance one below the other.
```
# matchDuration and walkDistance plots one below the other
import matplotlib.pyplot as plt
import seaborn as sns
plt.figure(figsize=(15,8))
plt.subplot(2,1,1)
sns.distplot(data["matchDuration"])
plt.subplot(2,1,2)
sns.distplot(data["walkDistance"]);
```
# 11. Plot distribution of the match's duration vs walk distance side by side.
```
# matchDuration and walkDistance plots side by side
import matplotlib.pyplot as plt
import seaborn as sns
plt.figure(figsize=(15,5))
plt.subplot(1,2,1)
sns.distplot(data["matchDuration"])
plt.subplot(1,2,2)
sns.distplot(data["walkDistance"]);
```
# 12. Pairplot the dataframe. Comment on kills vs damage dealt, Comment on maxPlace vs numGroups
```
columns = data[["kills","damageDealt","maxPlace","numGroups"]]
columns.head()
import seaborn as sns
sns.pairplot(columns);
```
Pairplot observations:
1. kills vs damage dealt
More the kills, damage dealt is more. There's a linear relationship as seen in the plot above.
2. maxPlace vs numGroups
There's a positive linear relationship between maxPlace and numGroups
```
import seaborn as sns
sns.distplot(data["maxPlace"]);
solo = data[data["numGroups"]>50]
duos = data[(data['numGroups']>25) & (data['numGroups']<=50)]
squad = data[data["numGroups"]<=25]
solo.head()
#for solo
solo_col = solo[["kills","damageDealt","maxPlace","numGroups"]]
sns.pairplot(solo_col)
#for duos
duo_col = duos[["kills","damageDealt","maxPlace","numGroups"]]
sns.pairplot(duo_col)
#for squad
squad_col = squad[["kills","damageDealt","maxPlace","numGroups"]]
sns.pairplot(squad_col)
```
# 13. How many unique values are there in 'matchType' and what are their counts?
```
# Unique values and their count in "matchType" column
data["matchType"].value_counts()
```
# 14. Plot a barplot of ‘matchType’ vs 'killPoints'. Write your inferences.
```
plt.figure(figsize=(15,8))
sns.barplot(x="matchType", y="killPoints", data = data)
plt.xticks(rotation=90);
```
As seen from the bar plot above, maximum kill points are achieved for the match types normal-squad-fpp and normal-duo-fpp
The kill points are the least for solo and solo-fpp match types
# 15. Plot a barplot of ‘matchType’ vs ‘weaponsAcquired’. Write your inferences
```
plt.figure(figsize=(15,8))
sns.barplot(x="matchType", y="weaponsAcquired", data = data)
plt.xticks(rotation=90);
```
As seen from the bar plot above, normal-solo-fpp and normal-squad-fpp have acquired the maximum number of weapons
Whereas, crashtpp and crashtpp have acquired the least number of weapons
# 16. Find the Categorical columns
```
#listing all the categorical columns
categorical = data.select_dtypes(include = ['object'])
categorical.columns
```
# 17. Plot a boxplot of ‘matchType’ vs ‘winPlacePerc’. Write your inferences
```
plt.figure(figsize=(10,8))
sns.boxplot(x='matchType',y='winPlacePerc',data=data)
plt.xticks(rotation=90);
```
The match type "normal-duo-fpp" has the highest spread of winPlacePerc. Therefore, its probability of winning is more.
# 18. Plot a boxplot of ‘matchType’ vs ‘matchDuration’. Write your inferences
```
plt.figure(figsize=(10,8))
sns.boxplot(x='matchType',y='matchDuration',data=data)
plt.xticks(rotation=90);
```
The match type "normal-squad-fpp" has the highest spread of matchDuration
# 19. Change the orientation of the above plot to horizontal
```
#changing the orientation of above plot to horizontal
plt.figure(figsize=(15,10))
sns.boxplot(x='matchDuration',y='matchType',data=data,orient="h");
```
# 20. Add a new column called ‘KILL’ which contains the sum of following columns viz. headshotKills, teamKills, roadKills.
```
#Adding a new column "KILL" to the dataset
data["KILL"] = data["headshotKills"]+data["teamKills"]+data["roadKills"]
data.head()
data["KILL"].describe()
```
# 21. Round off column ‘winPlacePerc’ to 2 decimals.
```
#rounding off column "winPlacePerc" to 2 decimal places
data['winPlacePerc'] = round(data['winPlacePerc'] , 2)
data['winPlacePerc'].head()
```
# 22. Take a sample of size 50 from the column damageDealt for 100 times and calculate its mean. Plot it on a histogram and comment on its distribution.
```
#taking sample of size 50 from the column "damageDealt" and calculating its mean for 100 times
l = []
x=0
while x<100:
sample = data["damageDealt"].sample(n=50).mean()
sample = round(sample,2)
l.append(sample)
x = x + 1
print(l)
#plotting histogram for damageDealt
%matplotlib inline
plt.figure(figsize=(10,8))
plt.hist(l,histtype="step");
plt.title("Damage Dealt by the player");
```
As seen from the Histogram above, the highest frequency (~27) for Damage dealt occurs in the interval of 118-125
| github_jupyter |
# Basic Plotting with matplotlib
You can show matplotlib figures directly in the notebook by using the `%matplotlib notebook` and `%matplotlib inline` magic commands.
`%matplotlib notebook` provides an interactive environment.
```
%matplotlib notebook
import matplotlib as mpl
mpl.get_backend()
import matplotlib.pyplot as plt
plt.plot?
# because the default is the line style '-',
# nothing will be shown if we only pass in one point (3,2)
plt.plot(3, 2)
# we can pass in '.' to plt.plot to indicate that we want
# the point (3,2) to be indicated with a marker '.'
plt.plot(3, 2, '.')
```
Let's see how to make a plot without using the scripting layer.
```
# First let's set the backend without using mpl.use() from the scripting layer
from matplotlib.backends.backend_agg import FigureCanvasAgg
from matplotlib.figure import Figure
# create a new figure
fig = Figure()
# associate fig with the backend
canvas = FigureCanvasAgg(fig)
# add a subplot to the fig
ax = fig.add_subplot(111)
# plot the point (3,2)
ax.plot(3, 2, '.')
# save the figure to test.png
# you can see this figure in your Jupyter workspace afterwards by going to
# https://hub.coursera-notebooks.org/
canvas.print_png('test.png')
```
We can use html cell magic to display the image.
```
%%html
<img src='test.png' />
# create a new figure
plt.figure()
# plot the point (3,2) using the circle marker
plt.plot(3, 2, 'o')
# get the current axes
ax = plt.gca()
# Set axis properties [xmin, xmax, ymin, ymax]
ax.axis([0,6,0,10])
# create a new figure
plt.figure()
# plot the point (1.5, 1.5) using the circle marker
plt.plot(1.5, 1.5, 'o')
# plot the point (2, 2) using the circle marker
plt.plot(2, 2, 'o')
# plot the point (2.5, 2.5) using the circle marker
plt.plot(2.5, 2.5, 'o')
# get current axes
ax = plt.gca()
# get all the child objects the axes contains
ax.get_children()
```
# Scatterplots
```
import numpy as np
x = np.array([1,2,3,4,5,6,7,8])
y = x
plt.figure()
plt.scatter(x, y) # similar to plt.plot(x, y, '.'), but the underlying child objects in the axes are not Line2D
import numpy as np
x = np.array([1,2,3,4,5,6,7,8])
y = x
# create a list of colors for each point to have
# ['green', 'green', 'green', 'green', 'green', 'green', 'green', 'red']
colors = ['green']*(len(x)-1)
colors.append('red')
plt.figure()
# plot the point with size 100 and chosen colors
plt.scatter(x, y, s=100, c=colors)
# convert the two lists into a list of pairwise tuples
zip_generator = zip([1,2,3,4,5], [6,7,8,9,10])
print(list(zip_generator))
# the above prints:
# [(1, 6), (2, 7), (3, 8), (4, 9), (5, 10)]
zip_generator = zip([1,2,3,4,5], [6,7,8,9,10])
# The single star * unpacks a collection into positional arguments
print(*zip_generator)
# the above prints:
# (1, 6) (2, 7) (3, 8) (4, 9) (5, 10)
# use zip to convert 5 tuples with 2 elements each to 2 tuples with 5 elements each
print(list(zip((1, 6), (2, 7), (3, 8), (4, 9), (5, 10))))
# the above prints:
# [(1, 2, 3, 4, 5), (6, 7, 8, 9, 10)]
zip_generator = zip([1,2,3,4,5], [6,7,8,9,10])
# let's turn the data back into 2 lists
x, y = zip(*zip_generator) # This is like calling zip((1, 6), (2, 7), (3, 8), (4, 9), (5, 10))
print(x)
print(y)
# the above prints:
# (1, 2, 3, 4, 5)
# (6, 7, 8, 9, 10)
plt.figure()
# plot a data series 'Tall students' in red using the first two elements of x and y
plt.scatter(x[:2], y[:2], s=100, c='red', label='Tall students')
# plot a second data series 'Short students' in blue using the last three elements of x and y
plt.scatter(x[2:], y[2:], s=100, c='blue', label='Short students')
# add a label to the x axis
plt.xlabel('The number of times the child kicked a ball')
# add a label to the y axis
plt.ylabel('The grade of the student')
# add a title
plt.title('Relationship between ball kicking and grades')
# add a legend (uses the labels from plt.scatter)
plt.legend()
# add the legend to loc=4 (the lower right hand corner), also gets rid of the frame and adds a title
plt.legend(loc=4, frameon=False, title='Legend')
# get children from current axes (the legend is the second to last item in this list)
plt.gca().get_children()
# get the legend from the current axes
legend = plt.gca().get_children()[-2]
# you can use get_children to navigate through the child artists
legend.get_children()[0].get_children()[1].get_children()[0].get_children()
# import the artist class from matplotlib
from matplotlib.artist import Artist
def rec_gc(art, depth=0):
if isinstance(art, Artist):
# increase the depth for pretty printing
print(" " * depth + str(art))
for child in art.get_children():
rec_gc(child, depth+2)
# Call this function on the legend artist to see what the legend is made up of
rec_gc(plt.legend())
```
# Line Plots
```
import numpy as np
linear_data = np.array([1,2,3,4,5,6,7,8])
exponential_data = linear_data**2
plt.figure()
# plot the linear data and the exponential data
plt.plot(linear_data, '-o', exponential_data, '-o')
# plot another series with a dashed red line
plt.plot([22,44,55], '--r')
plt.xlabel('Some data')
plt.ylabel('Some other data')
plt.title('A title')
# add a legend with legend entries (because we didn't have labels when we plotted the data series)
plt.legend(['Baseline', 'Competition', 'Us'])
# fill the area between the linear data and exponential data
plt.gca().fill_between(range(len(linear_data)),
linear_data, exponential_data,
facecolor='blue',
alpha=0.25)
range(len(linear_data))
```
Let's try working with dates!
```
plt.figure()
observation_dates = np.arange('2017-01-01', '2017-01-09', dtype='datetime64[D]')
plt.plot(observation_dates, linear_data, '-o', observation_dates, exponential_data, '-o')
```
Let's try using pandas
```
import pandas as pd
plt.figure()
observation_dates = np.arange('2017-01-01', '2017-01-09', dtype='datetime64[D]')
observation_dates = map(pd.to_datetime, observation_dates) # trying to plot a map will result in an error
plt.plot(observation_dates, linear_data, '-o', observation_dates, exponential_data, '-o')
plt.figure()
observation_dates = np.arange('2017-01-01', '2017-01-09', dtype='datetime64[D]')
observation_dates = list(map(pd.to_datetime, observation_dates)) # convert the map to a list to get rid of the error
plt.plot(observation_dates, linear_data, '-o', observation_dates, exponential_data, '-o')
x = plt.gca().xaxis
# rotate the tick labels for the x axis
for item in x.get_ticklabels():
item.set_rotation(45)
# adjust the subplot so the text doesn't run off the image
plt.subplots_adjust(bottom=0.25)
ax = plt.gca()
ax.set_xlabel('Date')
ax.set_ylabel('Units')
ax.set_title('Exponential vs. Linear performance')
# you can add mathematical expressions in any text element
ax.set_title("Exponential ($x^2$) vs. Linear ($x$) performance")
```
# Bar Charts
```
plt.figure()
xvals = range(len(linear_data))
plt.bar(xvals, linear_data, width = 0.3)
new_xvals = []
# plot another set of bars, adjusting the new xvals to make up for the first set of bars plotted
for item in xvals:
new_xvals.append(item+0.3)
plt.bar(new_xvals, exponential_data, width = 0.3 ,color='red')
from random import randint
linear_err = [randint(0,15) for x in range(len(linear_data))]
# This will plot a new set of bars with errorbars using the list of random error values
plt.bar(xvals, linear_data, width = 0.3, yerr=linear_err)
# stacked bar charts are also possible
plt.figure()
xvals = range(len(linear_data))
plt.bar(xvals, linear_data, width = 0.3, color='b')
plt.bar(xvals, exponential_data, width = 0.3, bottom=linear_data, color='r')
# or use barh for horizontal bar charts
plt.figure()
xvals = range(len(linear_data))
plt.barh(xvals, linear_data, height = 0.3, color='b')
plt.barh(xvals, exponential_data, height = 0.3, left=linear_data, color='r')
```
| github_jupyter |
# Introduction to Taxi ETL Job
This is the Taxi ETL job to generate the input datasets for the Taxi XGBoost job.
## Prerequirement
### 1. Download data
All data could be found at https://www1.nyc.gov/site/tlc/about/tlc-trip-record-data.page
### 2. Download needed jars
* [cudf-21.10.0-cuda11.jar](https://repo1.maven.org/maven2/ai/rapids/cudf/21.10.0/)
* [rapids-4-spark_2.12-21.10.0.jar](https://repo1.maven.org/maven2/com/nvidia/rapids-4-spark_2.12/21.10.0/rapids-4-spark_2.12-21.10.0.jar)
### 3. Start Spark Standalone
Before running the script, please setup Spark standalone mode
### 4. Add ENV
```
$ export SPARK_JARS=cudf-21.10.0-cuda11.jar,rapids-4-spark_2.12-21.10.0.jar
$ export PYSPARK_DRIVER_PYTHON=jupyter
$ export PYSPARK_DRIVER_PYTHON_OPTS=notebook
```
### 5. Start Jupyter Notebook with plugin config
```
$ pyspark --master ${SPARK_MASTER} \
--jars ${SPARK_JARS} \
--conf spark.plugins=com.nvidia.spark.SQLPlugin \
--conf spark.rapids.sql.incompatibleDateFormats.enabled=true \
--conf spark.rapids.sql.csv.read.integer.enabled=true \
--conf spark.rapids.sql.csv.read.long.enabled=true \
--conf spark.rapids.sql.csv.read.double.enabled=true \
--py-files ${SPARK_PY_FILES}
```
## Import Libs
```
import time
import os
import math
from pyspark.sql import SparkSession
from pyspark.sql.functions import *
from pyspark.sql.types import *
```
## Script Settings
### File Path Settings
* Define input/output file path
```
# You need to update them to your real paths! You can download the dataset
# from https://www1.nyc.gov/site/tlc/about/tlc-trip-record-data.page
# or you can just unzip datasets/taxi-small.tar.gz and use the provided
# sample dataset datasets/taxi/taxi-etl-input-small.csv
dataRoot = os.getenv('DATA_ROOT', '/root')
rawPath = dataRoot + '/spark-rapids-examples/datasets/taxi/taxi-etl-input-small.csv'
outPath = dataRoot + '/spark-rapids-examples/datasets/taxi/output'
```
## Function and Object Define
### Define the constants
* Define input file schema
```
raw_schema = StructType([
StructField('vendor_id', StringType()),
StructField('pickup_datetime', StringType()),
StructField('dropoff_datetime', StringType()),
StructField('passenger_count', IntegerType()),
StructField('trip_distance', DoubleType()),
StructField('pickup_longitude', DoubleType()),
StructField('pickup_latitude', DoubleType()),
StructField('rate_code', StringType()),
StructField('store_and_fwd_flag', StringType()),
StructField('dropoff_longitude', DoubleType()),
StructField('dropoff_latitude', DoubleType()),
StructField('payment_type', StringType()),
StructField('fare_amount', DoubleType()),
StructField('surcharge', DoubleType()),
StructField('mta_tax', DoubleType()),
StructField('tip_amount', DoubleType()),
StructField('tolls_amount', DoubleType()),
StructField('total_amount', DoubleType()),
])
```
* Define some ETL functions
```
def drop_useless(data_frame):
return data_frame.drop(
'dropoff_datetime',
'payment_type',
'surcharge',
'mta_tax',
'tip_amount',
'tolls_amount',
'total_amount')
def encode_categories(data_frame):
categories = [ 'vendor_id', 'rate_code', 'store_and_fwd_flag' ]
for category in categories:
data_frame = data_frame.withColumn(category, hash(col(category)))
return data_frame.withColumnRenamed("store_and_fwd_flag", "store_and_fwd")
def fill_na(data_frame):
return data_frame.fillna(-1)
def remove_invalid(data_frame):
conditions = [
( 'fare_amount', 0, 500 ),
( 'passenger_count', 0, 6 ),
( 'pickup_longitude', -75, -73 ),
( 'dropoff_longitude', -75, -73 ),
( 'pickup_latitude', 40, 42 ),
( 'dropoff_latitude', 40, 42 ),
]
for column, min, max in conditions:
data_frame = data_frame.filter('{} > {} and {} < {}'.format(column, min, column, max))
return data_frame
def convert_datetime(data_frame):
datetime = col('pickup_datetime')
return (data_frame
.withColumn('pickup_datetime', to_timestamp(datetime))
.withColumn('year', year(datetime))
.withColumn('month', month(datetime))
.withColumn('day', dayofmonth(datetime))
.withColumn('day_of_week', dayofweek(datetime))
.withColumn(
'is_weekend',
col('day_of_week').isin(1, 7).cast(IntegerType())) # 1: Sunday, 7: Saturday
.withColumn('hour', hour(datetime))
.drop('pickup_datetime'))
def add_h_distance(data_frame):
p = math.pi / 180
lat1 = col('pickup_latitude')
lon1 = col('pickup_longitude')
lat2 = col('dropoff_latitude')
lon2 = col('dropoff_longitude')
internal_value = (0.5
- cos((lat2 - lat1) * p) / 2
+ cos(lat1 * p) * cos(lat2 * p) * (1 - cos((lon2 - lon1) * p)) / 2)
h_distance = 12734 * asin(sqrt(internal_value))
return data_frame.withColumn('h_distance', h_distance)
```
* Define main ETL function
```
def pre_process(data_frame):
processes = [
drop_useless,
encode_categories,
fill_na,
remove_invalid,
convert_datetime,
add_h_distance,
]
for process in processes:
data_frame = process(data_frame)
return data_frame
```
## Run ETL Process and Save the Result
* Create Spark Session and create dataframe
```
spark = (SparkSession
.builder
.appName("Taxi-ETL")
.getOrCreate())
reader = (spark
.read
.format('csv'))
reader.schema(raw_schema).option('header', 'True')
raw_data = reader.load(rawPath)
```
* Run ETL Process and Save the Result
```
start = time.time()
etled_train, etled_eval, etled_trans = pre_process(raw_data).randomSplit(list(map(float, (80,20,0))))
etled_train.write.mode("overwrite").parquet(outPath+'/train')
etled_eval.write.mode("overwrite").parquet(outPath+'/eval')
etled_trans.write.mode("overwrite").parquet(outPath+'/trans')
end = time.time()
print(end - start)
spark.stop()
```
| github_jupyter |
```
# #colabを使う方はこちらを使用ください。
# !pip install torch==0.4.1
# !pip install torchvision==0.2.1
# !pip install numpy==1.14.6
# !pip install matplotlib==2.1.2
# !pip install pillow==5.0.0
# !pip install opencv-python==3.4.3.18
# !pip install torchtext==0.3.1
import torch
import torch.nn as nn
import torch.nn.init as init
import torch.optim as optim
import torch.nn.functional as F
#torchtextを使用
from torchtext import data
from torchtext import vocab
from torchtext import datasets
%matplotlib inline
import numpy as np
from matplotlib import pyplot as plt
# データとモデルに.to(device)を指定してgpuの計算資源を使用する。
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
device
```
#文章生成
## データの読み込み
```
tokenize = lambda x: x.split()
# 前処理用の機能のFieldをセットアップ
#Field
TEXT = data.Field(sequential=True, tokenize=tokenize, lower=True, batch_first=True)
# データを取得
# The Penn Treebankデータセット。
train_dataset, val_dataset, test_dataset = datasets.PennTreebank.splits(TEXT)
TEXT.build_vocab(train_dataset, vectors=vocab.GloVe(name='6B', dim=300))
#全単語数
vocab_size = len(TEXT.vocab)
print(vocab_size)
# 単語の件数のtop10
print(TEXT.vocab.freqs.most_common(10))
# 単語
print(TEXT.vocab.itos[:10])
#埋め込みベクトルを取得
word_embeddings = TEXT.vocab.vectors
# ハイパーパラメータ
embedding_length = 300
hidden_size = 256
batch_size = 32
# BPTTIteratorは言語モデル用のイテレータ作成を行います。
# textとtarget属性を持ちます。
train_iter, val_iter, test_iter = data.BPTTIterator.splits((train_dataset, val_dataset, test_dataset)
, batch_size=32, bptt_len=30, repeat=False)
print(len(train_iter))
print(len(val_iter))
print(len(test_iter))
for i, batch in enumerate(train_iter):
print("データの形状確認")
print(batch.text.size())
print(batch.target.size())
#BPTTIteratorがBatch firstになってない件は2018/11/24時点では#462がPull requestsがされています。
print("permuteでバッチを先にする")
print(batch.text.permute(1, 0).size())
print(batch.target.permute(1, 0).size())
print("データ目の形状とデータを確認")
text = batch.text.permute(1, 0)
target = batch.target.permute(1, 0)
print(text[1,:].size())
print(target[1,:].size())
print(text[1,:].tolist())
print(target[1,:].tolist())
print("データの単語列を表示")
print([TEXT.vocab.itos[data] for data in text[1,:].tolist()])
print([TEXT.vocab.itos[data] for data in target[1,:].tolist()])
break
```
## ネットワークを定義
```
class LstmLangModel(nn.Module):
def __init__(self, batch_size, hidden_size, vocab_size, embedding_length, weights):
super(LstmLangModel, self).__init__()
self.batch_size = batch_size
self.hidden_size = hidden_size
self.vocab_size = vocab_size
self.embed = nn.Embedding(vocab_size, embedding_length)
self.embed.weight.data.copy_(weights)
self.lstm = nn.LSTM(embedding_length, hidden_size, batch_first=True)
self.fc = nn.Linear(hidden_size, vocab_size)
def forward(self, x, h):
x = self.embed(x)
output_seq, (h, c) = self.lstm(x, h)
# 出力を変形する (batch_size*sequence_length, 隠れ層のユニット数hidden_size)
out = output_seq.reshape(output_seq.size(0)*output_seq.size(1), output_seq.size(2))
out = self.fc(out)
return out, (h, c)
net = LstmLangModel(batch_size, hidden_size, vocab_size, embedding_length, word_embeddings)
net = net.to(device)
# 損失関数、最適化関数を定義
criterion = nn.CrossEntropyLoss()
optim = optim.Adam(filter(lambda p: p.requires_grad, net.parameters()))
```
## 学習
```
num_epochs = 200
train_loss_list = []
# Truncated backpropagation
# 逆伝播を途中で打ち切る
def detach(states):
return [state.detach() for state in states]
for epoch in range(num_epochs):
train_loss = 0
# 初期隠れ状態とセル状態を設定する
states = (torch.zeros(1, batch_size, hidden_size).to(device),
torch.zeros(1, batch_size, hidden_size).to(device))
#train
net.train()
for i, batch in enumerate(train_iter):
text = batch.text.to(device)
labels = batch.target.to(device)
#LSTMの形状に合わせて入力もバッチを先にする。
text = text.permute(1, 0)
labels = labels.permute(1, 0)
optim.zero_grad()
states = detach(states)
outputs, states = net(text, states)
loss = criterion(outputs, labels.reshape(-1))
train_loss += loss.item()
loss.backward()
optim.step()
avg_train_loss = train_loss / len(train_iter)
print ('Epoch [{}/{}], Loss: {loss:.4f}, Perplexity: {perp:5.2f}'
.format(epoch+1, num_epochs, i+1, loss=avg_train_loss, perp=np.exp(avg_train_loss)))
train_loss_list.append(avg_train_loss)
```
## 生成
```
num_samples = 1000 # サンプリングされる単語の数
# モデルをテストする
net.eval()
with torch.no_grad():
text = ""
# 初期隠れ状態とセル状態を設定する
states = (torch.zeros(1, 1, hidden_size).to(device),
torch.zeros(1, 1, hidden_size).to(device))
# ランダムに1単語のIDを選択
input = torch.multinomial(torch.ones(vocab_size), num_samples=1).unsqueeze(1).to(device)
# print("input word", TEXT.vocab.itos[input])
for i in range(num_samples):
# print("input word", TEXT.vocab.itos[input])
output, states = net(input, states)
word_id = output.max(1)[1].item()
# 次のタイムステップのために単語IDを入力
input.fill_(word_id)
# 単語IDから文字を取得
word = TEXT.vocab.itos[word_id]
# textに書き込む
word = '\n' if word == '<eos>' else word + ' '
text += word
# textを表示
print(text)
```
| github_jupyter |
# Top coding, bottom coding and zero coding
## Outliers
An outlier is a data point which is significantly different from the remaining data. “An outlier is an observation which deviates so much from the other observations as to arouse suspicions that it was generated by a different mechanism.” [D. Hawkins. Identification of Outliers, Chapman and Hall , 1980].
Statistics such as the mean and variance are very susceptible to outliers. In addition, **some Machine Learning models are indeed sensitive to outliers and their performance might be impaired by them**. Thus, it is common practice to engineer the features to minimise the impact of outliers on the performance of these algorithms.
### Nature of outliers
- Genuine extremely high or extremely low values
- Introduced due to mechanical error (wrong measurement)
- Introduced by replacing missing values (NA) by a value out of the distribution (as described in previous lectures)
In some cases, the presence of outliers is informative, and therefore they deserve further study. In this course I will tackle the engineering of those values that do not add any particular extra information, and could as well be eliminated.
## How can we pre-process outliers?
- Mean/median imputation or random sampling
- Discrestisation
- Discard the outliers: process also called Trimming
- Top-coding, bottom-coding and zero-coding: also known as windsorization
### Mean/median imputation or random sampling
If we have reasons to believe that the outliers are due to mechanical error or problems during measurement. This means, if the outliers are in nature similar to missing data, then any of the methods discussed for missing data can be applied to replace outliers. Because the number of outliers is in nature small (otherwise they would not be outliers), it is reasonable to use the mean/median imputation to replace them.
### Discretisation
Discretisation is the transformation of continuous variables into discrete variables. It involves assigning the variable values into defined groups. For example, for the variable age, we could group the observations (people) into buckets / groups like: 0-20, 21-40, 41-60, > 61. This grouping of the variables in ranges is called discretisation. As you can see, any outlier (extremely high) value of age would be included in the > 61 group, therefore minimising its impact. I will discuss more on the different discretisation methods in the "Discretisation" section of this course.
### Trimming
Trimming refers to the removal of the extreme values of a sample. In this procedure, the outliers are identified and those observations removed from the sample. On the down side, these values, may contain useful information for other variables included in the dataset. Thus, likely, we may choose not to remove these observations and handle outliers by top / bottom coding as described below.
## Top-coding, bottom-coding and zero-coding.
**Top-conding**, widely used in econometrics and statistics, means capping the maximum of a distribution at an arbitrarily set value. A top-coded variable is one for which data points whose values are above an upper bound are censored. This means in practical terms that all values above the upper band will be arbitrarily set to the upper band.
Top-coding is common practice in survey data, before it is released to the public. It is used to preserve the anonymity of respondents. For example, high earners may be easily identifiable by their earnings. Thus, by implementing top-coding, that outlier is capped at a certain maximum value and therefore looks like many other observations, it is not uniquely identifiable any more. Top-coding can be also applied to prevent possibly-erroneous outliers from being published.
Bottom-coding is analogous, but on the left side of the distribution. This is, all values below a certain threshold, are capped at that threshold. If the threshold is zero, then it is known as **zero-coding**, e.g. if amounts below zero are reported as zero. Good examples would be the variable "age", or the variable "earnings". It is not possible to have negative age or a negative salary, thus, it is reasonable to cap the lowest values at zero. Any observation with a value under zero must have been introduced by mistake.
Top-coding and bottom-coding are indeed used in practice to remove outliers of variables and therefore prevent model over-fitting. For an example in a financial institution, look at my talk in [pydata](https://www.google.co.uk/url?sa=t&rct=j&q=&esrc=s&source=web&cd=2&cad=rja&uact=8&ved=0ahUKEwiEtaG7p6fXAhVI2hoKHWqQBsMQtwIILTAB&url=https%3A%2F%2Fwww.youtube.com%2Fwatch%3Fv%3DKHGGlozsRtA&usg=AOvVaw13tQ7UEv3w1k_RLsEbB3aB).
#### Note
Top-coding may affect estimates of the standard errors of the variable, or change the variable distribution, by censoring those values at the far end of the tails.
### Identifying outliers
#### Extreme Value Analysis
The most basic form of outlier detection is Extreme Value Analysis of 1-dimensional data. The key for this method is to determine the statistical tails of the underlying distribution of the variable, and then finding the values that sit at the very end of the tails.
In the typical scenario, the distribution of the variable is Gaussian and thus outliers will lie outside the mean plus or minus 3 times the standard deviation of the variable.
If the variable is not normally distributed, a general approach is to calculate the quantiles, and then the interquantile range (IQR), as follows:
IQR = 75th quantile - 25th quantile
An outlier will sit outside the following upper and lower boundaries:
Upper boundary = 75th quantile + (IQR * 1.5)
Lower boundary = 25th quantile - (IQR * 1.5)
or for extreme cases:
Upper boundary = 75th quantile + (IQR * 3)
Lower boundary = 25th quantile - (IQR * 3)
=======================================================================
Below I will demonstrate top-coding in real-life datasets. We have seen an intuition of how this improves machine learning algorithms in the lecture "Outliers" in the section "Type of problems within variables".
=============================================================================
## Real Life example:
### Predicting Survival on the Titanic: understanding society behaviour and beliefs
Perhaps one of the most infamous shipwrecks in history, the Titanic sank after colliding with an iceberg, killing 1502 out of 2224 people on board. Interestingly, by analysing the probability of survival based on few attributes like gender, age, and social status, we can make very accurate predictions on which passengers would survive. Some groups of people were more likely to survive than others, such as women, children, and the upper-class. Therefore, we can learn about the society priorities and privileges at the time.
### Lending Club
**Lending Club** is a peer-to-peer Lending company based in the US. They match people looking to invest money with people looking to borrow money. When investors invest their money through Lending Club, this money is passed onto borrowers, and when borrowers pay their loans back, the capital plus the interest passes on back to the investors. It is a win for everybody as they can get typically lower loan rates and higher investor returns.
If you want to learn more about Lending Club follow this link:
https://www.lendingclub.com/
The Lending Club dataset contains complete loan data for all loans issued through the 2007-2015, including the current loan status (Current, Late, Fully Paid, etc.) and latest payment information. Features (aka variables) include credit scores, number of finance inquiries, address including zip codes and state, and collections among others. Collections indicates whether the customer has missed one or more payments and the team is trying to recover their money.
The file is a matrix of about 890 thousand observations and 75 variables. More detail on this dataset can be found in Kaggle's website: https://www.kaggle.com/wendykan/lending-club-loan-data
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
from sklearn.model_selection import train_test_split
pd.set_option('display.max_columns', None) # to display the total number columns present in the dataset
```
## Titanic dataset
```
# let's load the titanic dataset
data = pd.read_csv('titanic.csv')
data.head()
```
### Top-coding important
Top-coding and bottom-coding, as any other feature pre-processing step, should be determined over the training set, and then transferred onto the test set. This means that we should find the upper and lower bounds in the training set only, and use those bands to cap the values in the test set.
```
# divide dataset into train and test set
X_train, X_test, y_train, y_test = train_test_split(data, data.Survived,
test_size=0.3,
random_state=0)
X_train.shape, X_test.shape
```
There are 2 numerical variables in this dataset, Fare and Age. So let's go ahead and find out whether there are values that we could consider outliers
### Fare
```
# First let's plot a histogram to get an idea of the distribution
fig = X_train.Fare.hist(bins=50)
fig.set_title('Fare Paid Distribution')
fig.set_xlabel('Fare')
fig.set_ylabel('Number of Passengers')
sns.kdeplot(X_train.Fare)
```
Because the distribution of Fare is skewed, we should estimate outliers using the quantile method instead of the Gaussian distribution.
```
# visualising outliers using boxplots and whiskers, which provides the quantiles
# and inter-quantile range, with the outliers sitting outside the error bars.
# All the dots in the plot below are outliers according to the 1.5 IQR rule
fig = sns.boxplot(y='Fare', data=X_train)
fig.set_xlabel('Fare')
fig.set_ylabel('Number of Passengers')
```
The outliers, according to the above plot, lie all at the right side of the distribution. This is, some people paid extremely high prices for their tickets.
Therefore, in this variable, only extremely high values will affect the performance of our machine learning models, and we need to do therefore top-coding. Bottom coding in this case it is not necessary. At least not to improve the performance of the machine learning algorithms.
```
# let's look at the values of the quantiles so we can calculate the upper and lower boundaries for the outliers
X_train.Fare.describe()
# top coding: upper boundary for outliers according to interquantile proximity rule
IQR = data.Fare.quantile(0.75) - data.Fare.quantile(0.25)
Upper_fence = X_train.Fare.quantile(0.75) + (IQR * 3)
Upper_fence
```
The upper boundary, above which every value is considered an outlier is a cost of 100 dollars for the Fare.
```
# lets look at the actual number of passengers that paid more than USS 100
print('total passengers: {}'.format(X_train.shape[0]))
print('passengers that paid more than 100: {}'.format(X_train[X_train.Fare>100].shape[0]))
print('percentage of outliers: {}'.format(X_train[X_train.Fare>100].shape[0]/np.float(X_train.shape[0])))
# top-coding: capping the variable Fare at 100
X_train.loc[X_train.Fare>100, 'Fare'] = 100
X_test.loc[X_test.Fare>100, 'Fare'] = 100
X_train.Fare.max(), X_test.Fare.max()
```
This is all we need to remove outliers from a machine learning perspective.
However, note that in the dataset, there are also a few passengers that paid zero for their tickets
```
X_train[X_train.Fare==0].shape
X_train[X_train.Fare==0]
```
The majority of them do not have a Cabin assigned, and could therefore have jumped on the boat illegally. Alternatively, there could also be that that information could not be retrieved, so we do not know how much they paid. But we do know that the cheapest ticket was 5 dollars, see below:
```
X_train[X_train.Fare!=0]['Fare'].min()
```
In situations like this, it is best to discuss with the data owner (in business, someone who knows the data well) the nature of the data, and the importance of the variable.
If the 0 values in this case mean that the data could not be retrieved properly, and therefore is in nature an NaN, one could choose to replace them by a random sample or mean/median imputation, or to do bottom-coding.
If the case of zero corresponds otherwise to people jumping on the boat illegally, one may choose to leave them as zero.
### Age
```
# First let's plot the histogram to get an idea of the distribution
fig = X_train.Age.hist(bins=50)
fig.set_title('Age Distribution')
fig.set_xlabel('Age')
fig.set_ylabel('Number of Passengers')
sns.kdeplot(X_train.Age)
```
Although it does not look strictly normal, we could assume normality and use the Gaussian approach to find outliers. See below.
```
# now let's plot the boxplots and whiskers
fig = sns.boxplot(y='Age', data=X_train)
fig.set_xlabel('Age')
fig.set_ylabel('Number of Passengers')
```
Again, for this variable the outliers lie only on the right of the distribution. Therefore we only need to introduce top-coding.
```
# and let's get the numbers to calculate the upper boundary
X_train.Age.describe()
# Assuming normality
Upper_boundary = X_train.Age.mean() + 3* X_train.Age.std()
Upper_boundary
# let's find out whether there are outliers according to the above boundaries
# remember that Age has ~ 20% missing values
total_passengers = np.float(X_train.shape[0])
print('total passengers: {}'.format(X_train.Age.dropna().shape[0]/total_passengers))
print('passengers older than 73 (Gaussian app): {}'.format(X_train[X_train.Age>73].shape[0]/total_passengers))
X_train.loc[X_train.Age>73, 'Age'] = 73
X_test.loc[X_test.Age>73, 'Age'] = 73
X_train.Age.max(), X_test.Age.max()
```
In the test set, there were no outliers, as the maximum Age value is 70, below the value we used to cap outliers.
## Loan book from Lending Club
```
# we will examine only the income variable, as this is one that typically shows outliers.
# a few people are high earners, and the remaining of the borrowers fall within a normal-ish distribution
data = pd.read_csv('loan.csv', usecols=['annual_inc'], nrows=30000)
data.head()
fig = data.annual_inc.hist(bins=500)
fig.set_xlim(0,500000)
sns.boxplot(y='annual_inc', data=data)
```
As expected, outliers sit on the right of the distribution. Therefore, we will perform top-coding.
```
data.annual_inc.describe()
# because the distribution is not completely normal, I choose to examine outliers with the interquantal
# distance
IQR = data.annual_inc.quantile(0.75) - data.annual_inc.quantile(0.25)
Upper_fence = data.annual_inc.quantile(0.75) + (IQR * 1.5)
Upper_fence_ext = data.annual_inc.quantile(0.75) + (IQR * 3)
Upper_fence, Upper_fence_ext
# let's look at the percentage of high earners within each extreme bucket
total_borrowers = np.float(data.shape[0])
print('total borrowers: {}'.format(data.annual_inc.shape[0]/total_borrowers))
print('borrowers than earn > 146k: {}'.format(data[data.annual_inc>146000].shape[0]/total_borrowers))
print('borrowers than earn > 210k: {}'.format(data[data.annual_inc>210000].shape[0]/total_borrowers))
# top-coding
data['annual_capped'] = np.where(data.annual_inc>210000, 210000, data.annual_inc)
data.describe()
```
We see the effect of capping on the overall distribution of the variable. The standard deviation is smaller, and so is the maximum value.
```
fig = plt.figure(figsize=(12,8))
ax = fig.add_subplot(111)
data.annual_inc.plot(kind='kde', ax=ax)
data.annual_capped.plot(kind='kde', ax=ax, color = 'red')
lines, labels = ax.get_legend_handles_labels()
labels = ['Income original', 'Income capped']
ax.legend(lines, labels, loc='best')
ax.set_xlim(0,500000)
```
We can observe the effect of top codding on the variable distribution. The maximum value corresponds now to the value we set as a cap. And we observe a peak in that value, that indicates that people that earn more than the cap, are now grouped together under a capped maximum salary.
**That is all for this demonstration. I hope you enjoyed the notebook, and see you in the next one.**
| github_jupyter |
```
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
titanic = pd.read_csv('./titanic.csv')
titanic.head(3)
```
## Summary statistics
### Summarizing numerical data
- .mean()
- .median()
- .min()
- .maxx()
- .var()
- .std()
- .sum()
- .quantile()
```
titanic['Age'].mean()
titanic['Age'].mode()
titanic.Age.min()
titanic.Age.max()
titanic['Age'].var() #<--Return unbiased variance over requested axis.
titanic['Age'].quantile() #<--Return values at the given quantile over requested axis.
titanic['Age'].std()
titanic['Age'].sum()
```
### summarizing dates
### .agg() method
##### on Single column
```
def pct30(column): return column.quantile(0.3)
titanic['Age'].agg(pct30)#<-- applying agg() on a column using simple function
titanic['Age'].agg(lambda x: x.quantile(.3)) #<-- using lambda function
```
##### on multiple column
```
titanic[['Age', 'Fare']].agg(lambda x: x.quantile(0.3))
```
##### multiple summaries
```
def pct30(column): return column.quantile(0.3)
def pct40(column): return column.quantile(0.4)
titanic['Age'].agg([pct30,pct40])
```
### cumulative statistics
- .cumsum()
- .cummax()
- .cummin()
- .cumprod()
```
pd.DataFrame(titanic['Age'].cumsum()).head(4)
```
## Counting
#### Dropping duplicate names
```
titanic.drop_duplicates(subset = "Pclass")
titanic.drop_duplicates(subset = ["Pclass", 'SibSp'])
```
#### .values_count()
```
pd.DataFrame(titanic['Age'].value_counts())
pd.DataFrame(titanic['Age'].value_counts(sort=True))
pd.DataFrame(titanic['Age'].value_counts(normalize=True))
```
## Group summary satistics
```
titanic[titanic['Sex'] == 'male']['Age'].mean()
titanic[titanic['Sex'] == 'female']['Age'].mean()
titanic.groupby('Sex')['Age'].mean()
titanic.groupby(['Survived', 'Sex'])['Age'].count() # < -- multiple group
titanic.groupby('Sex')['Age'].agg(['count', 'min', 'max'])# <-- multiple stats
titanic.groupby(['Survived', 'Sex'])[['Age', 'SibSp']].mean()
titanic.groupby(['Survived', 'Sex'])[['Age', 'SibSp']].agg(['count', 'min', 'max'])
```
## Pivot tables
**Signature**:
titanic.pivot_table(
values=None,
index=None,
columns=None,
aggfunc='mean',
fill_value=None,
margins=False,
dropna=True,
margins_name='All',
observed=False,
)
```
titanic.groupby('Sex')['Age'].mean()
#pivot and implicitly define agffunc=np.mean
titanic.pivot_table(values = 'Age', index='Sex')
#explicitly define statistics i:e np.median
titanic.pivot_table(values= 'Age', index='Sex', aggfunc=np.median)
#multiple statistics
titanic.pivot_table(values='Age', index='Sex', aggfunc=[np.std, np.median])
```
#### pivot on two varibales
```
#in groupby
# titanic.groupby(['Survived','Sex'])['Age'].mean().unstack()
#pivot on two varibales
titanic.pivot_table(values='Age', index='Sex', columns='Survived')
```
#### filling missing values in pivot table
```
titanic.pivot_table(values='Age', index='Sex', columns='Survived', fill_value=0)
```
#### summing with pivot table
```
titanic.pivot_table(values='Age',
index='Sex',
columns='Survived',
fill_value=0,
margins=True)
titanic.pivot_table(values='Age',
index='Sex',
columns='Survived',
fill_value=0,
margins=True,
margins_name='mean')
```
| github_jupyter |
```
pip install spacy
!pip3 install nltk
pip install pyLDAvis
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from sklearn.decomposition import NMF, LatentDirichletAllocation
from time import time
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import timeit
from time import time
import gensim
import gensim.corpora as corpora
from gensim.utils import simple_preprocess
from gensim.models import CoherenceModel
import matplotlib.pyplot as plt
%matplotlib inline
import pyLDAvis
import pyLDAvis.gensim
import nltk
from nltk.corpus import stopwords
from nltk.tokenize import TweetTokenizer
from collections import Counter
from wordcloud import WordCloud
import os
from google.colab import drive
drive.mount('/content/drive')
root_folder = '/content/drive/My Drive/IITR Project'
data_folder = '/content/drive/My Drive/IITR Project/data'
df_news=pd.read_csv(os.path.join(data_folder,'RedditNews-1.csv'))
df_news.head()
```
#Topic Modeling of news from df_news
Data Preprocessing - Removing hyphen
```
df_news = df_news.replace(np.nan, ' ', regex=True)
df_news = df_news.replace('b\"|b\'|\\\\|\\\"', '', regex=True)
```
TOKENISATION
Using Spacy Lemmatizer
```
testdata = df_news['News']
#testdataNews = df_news['News']
#testdata = testdataNews[:100]
import spacy
nlp = spacy.load('en', disable=['parser', 'tagger', 'ner'])
from spacy.lang.en import English
parser = English()
stop_words=spacy.lang.en.stop_words.STOP_WORDS
print(stop_words)
len(stop_words)
def spacy_lemma_text(text):
token = []
doc = nlp(text)
tokens = [tok.lemma_.lower().strip() for tok in doc if tok.lemma_ != '-PRON-']
tokens = [tok for tok in tokens if tok not in stop_words]
tokens = [tok for tok in tokens if len(tok) >= 3]
#tokens = ' '.join(tokens)
return tokens
filtered_words_text = testdata.apply(spacy_lemma_text)
def join_tokens_text(text):
token = []
doc = nlp(text)
tokens = [tok.lemma_.lower().strip() for tok in doc if tok.lemma_ != '-PRON-']
tokens = [tok for tok in tokens if tok not in stop_words]
tokens = [tok for tok in tokens if len(tok) >= 3]
tokens = ' '.join(tokens)
return tokens
filtered_join_text = testdata.apply(join_tokens_text)
filtered_words = filtered_words_text.to_list()
id2word = corpora.Dictionary(filtered_words_text)
# Create Corpus
texts = filtered_words_text
# Term Document Frequency
corpus = [id2word.doc2bow(text) for text in texts]
# View
print(corpus[:1])
[[(id2word[id], freq) for id, freq in cp] for cp in corpus[:1]]
def compute_coherence_values(dictionary, corpus, texts, limit, start=2, step=3):
coherence_values = []
model_list = []
for num_topics in range(start, limit, step):
model = gensim.models.ldamodel.LdaModel(corpus=corpus, num_topics=num_topics, id2word=dictionary, update_every=1, chunksize=100, passes=25)
model_list.append(model)
coherencemodel = CoherenceModel(model=model, texts=texts, dictionary=dictionary, coherence='c_v')
coherence_values.append(coherencemodel.get_coherence())
return model_list, coherence_values
model_list_Lda,coherence_values_Lda=compute_coherence_values(dictionary=id2word,corpus=corpus,texts=filtered_words,start=8,limit=12,step=1)
if not os.path.exists(os.path.join(data_folder, 'ldamodels')):
os.makedirs(os.path.join(data_folder, 'ldamodels'))
for i in range(len(model_list_Lda)):
model_list_Lda[i].save(os.path.join(data_folder, 'ldamodels', 'ldamodels-{}.model'.format(i+2)))
!ls '/content/drive/My Drive/IITR Project/data/ldamodels'
import seaborn as sns
sns.set()
limit=12; start=8; step=1;
x = range(start, limit, step)
plt.plot(x, coherence_values_Lda)
plt.xlabel("Num Topics")
plt.ylabel("Coherence score")
plt.legend(("coherence_values"), loc='best')
plt.show()
ldamodel = gensim.models.ldamodel.LdaModel(corpus=corpus, num_topics=11, id2word=id2word, update_every=1, chunksize=100, passes=10)
coherencemodel = CoherenceModel(model=ldamodel, texts=filtered_words, dictionary=id2word, coherence='c_v')
coherence_values = coherencemodel.get_coherence()
print(coherence_values)
def format_topics_sentences(ldamodel=ldamodel, corpus=corpus, texts=testdata):
# Init output
sent_topics_df = pd.DataFrame()
# Get main topic in each document
for i, row in enumerate(ldamodel[corpus]):
row = sorted(row, key=lambda x: (x[1]), reverse=True)
# Get the Dominant topic, Perc Contribution and Keywords for each document
for j, (topic_num, prop_topic) in enumerate(row):
if j == 0: # => dominant topic
wp = ldamodel.show_topic(topic_num)
topic_keywords = ", ".join([word for word, prop in wp])
sent_topics_df = sent_topics_df.append(pd.Series([int(topic_num), round(prop_topic,4), topic_keywords]), ignore_index=True)
else:
break
sent_topics_df.columns = ['Dominant_Topic', 'Perc_Contribution', 'Topic_Keywords']
# Add original text to the end of the output
contents = pd.Series(texts)
sent_topics_df = pd.concat([sent_topics_df, contents], axis=1)
return(sent_topics_df)
df_topic_sents_keywords = format_topics_sentences(ldamodel=ldamodel, corpus=corpus, texts=testdata)
# Format
df_dominant_topic = df_topic_sents_keywords.reset_index()
df_dominant_topic.columns = ['Document_No', 'Dominant_Topic', 'Topic_Perc_Contrib', 'Keywords', 'Text']
# Show
df_dominant_topic.head(5)
df_dominant_topic
for idx,row in df_dominant_topic.iterrows():
print( row['Document_No'], row['Keywords'].split(',')[:5]
)
# Group top 5 sentences under each topic
sent_topics_sorted = pd.DataFrame()
sent_topics_outdf_grpd = df_topic_sents_keywords.groupby('Dominant_Topic')
for i, grp in sent_topics_outdf_grpd:
sent_topics_sorted = pd.concat([sent_topics_sorted,
grp.sort_values(['Perc_Contribution'], ascending=[0]).head(1)],
axis=0)
# Reset Index
sent_topics_sorted.reset_index(drop=True, inplace=True)
# Format
sent_topics_sorted.columns = ['Topic_Num', "Topic_Perc_Contrib", "Keywords", "Text"]
# Show
sent_topics_sorted
for idx, row in sent_topics_sorted.iterrows():
print('Topic number {}'.format(int(row['Topic_Num'])))
print('Keywords: {}'.format(row['Keywords']))
print()
print(row['Text'])
print()
# Number of Documents for Each Topic
topic_counts = df_topic_sents_keywords['Dominant_Topic'].value_counts()
# Percentage of Documents for Each Topic
topic_contribution = round(topic_counts/topic_counts.sum(), 4)
# Topic Number and Keywords
topic_num_keywords = sent_topics_sorted[['Topic_Num', 'Keywords']]
# Concatenate Column wise
df_dominant_topics = pd.concat([topic_num_keywords, topic_counts, topic_contribution], axis=1)
# Change Column names
df_dominant_topics.columns = ['Dominant_Topic', 'Topic_Keywords', 'Num_Documents', 'Percent_Documents']
# Show
df_dominant_topics
with pd.option_context('display.max_rows', None, 'display.max_columns', None):
pd.set_option('display.max_colwidth', -1)
display(df_dominant_topics)
df_dominant_topics.to_csv('df_dominant_topics.csv', index=False)
for top in ldamodel.print_topics():
print(top)
# Visualize the topics
pyLDAvis.enable_notebook()
vis = pyLDAvis.gensim.prepare(ldamodel, corpus, id2word)
vis
#Function to generate the word cloud
def create_word_cloud(string):
cloud = WordCloud(background_color = "white", max_words = 200)
cloud.generate(string)
plt.imshow(cloud, interpolation='bilinear')
plt.axis("off")
plt.show()
def convertListToString(s):
str1 = " "
for ele in s:
str1 += ele + ' '
return str1
def Extract(lst):
return [item[0] for item in lst]
create_word_cloud(convertListToString(Extract(ldamodel.show_topic(0))))
create_word_cloud(convertListToString(Extract(ldamodel.show_topic(1))))
create_word_cloud(convertListToString(Extract(ldamodel.show_topic(2))))
create_word_cloud(convertListToString(Extract(ldamodel.show_topic(3))))
create_word_cloud(convertListToString(Extract(ldamodel.show_topic(4))))
create_word_cloud(convertListToString(Extract(ldamodel.show_topic(5))))
create_word_cloud(convertListToString(Extract(ldamodel.show_topic(6))))
create_word_cloud(convertListToString(Extract(ldamodel.show_topic(7))))
create_word_cloud(convertListToString(Extract(ldamodel.show_topic(8))))
create_word_cloud(convertListToString(Extract(ldamodel.show_topic(9))))
create_word_cloud(convertListToString(Extract(ldamodel.show_topic(10))))
```
| github_jupyter |
```
import torch
import torch.nn as nn
from torchvision import models
from torchvision import transforms
import argparse
import datetime
import os
from tensorboardX import SummaryWriter
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn import metrics
class MRNet(nn.Module):
def __init__(self):
super().__init__()
self.pretrained_model = models.vgg16(pretrained=True)
self.pooling_layer = nn.AdaptiveAvgPool2d(1)
self.classifer = nn.Linear(512, 1)
def forward(self, x):
x = torch.squeeze(x, dim=0)
features = self.pretrained_model.features(x)
pooled_features = self.pooling_layer(features)
pooled_features = pooled_features.view(pooled_features.size(0), -1)
flattened_features = torch.max(pooled_features, 0, keepdim=True)[0]
output = self.classifer(flattened_features)
return output
cnn_name="VGG16"
injury_type="acl"
mri_view="axial"
injury_name=injury_type.capitalize()+"_Tear"
class MRDataset(torch.utils.data.Dataset):
def __init__(self, root_dir, task, plane, train=True, transform=None, weights=None):
super().__init__()
self.task = task
self.plane = plane
self.root_dir = root_dir
self.train = train
if self.train:
self.folder_path = self.root_dir + 'train/{0}/'.format(plane)
self.records = pd.read_csv(
self.root_dir + 'train-{0}.csv'.format(task), header=None, names=['id', 'label'])
else:
transform = None
self.folder_path = self.root_dir + 'valid/{0}/'.format(plane)
self.records = pd.read_csv(
self.root_dir + 'valid-{0}.csv'.format(task), header=None, names=['id', 'label'])
self.records['id'] = self.records['id'].map(
lambda i: '0' * (4 - len(str(i))) + str(i))
self.paths = [self.folder_path + filename +
'.npy' for filename in self.records['id'].tolist()]
self.labels = self.records['label'].tolist()
self.transform = transform
if weights is None:
pos = np.sum(self.labels)
neg = len(self.labels) - pos
self.weights = [1, neg / pos]
else:
self.weights = weights
def __len__(self):
return len(self.paths)
def __getitem__(self, index):
array = np.load(self.paths[index])
label = self.labels[index]
label = torch.FloatTensor([label])
if self.transform:
array = self.transform(array)
else:
array = np.stack((array,)*3, axis=1)
array = torch.FloatTensor(array)
if label.item() == 1:
weight = np.array([self.weights[1]])
weight = torch.FloatTensor(weight)
else:
weight = np.array([self.weights[0]])
weight = torch.FloatTensor(weight)
return array, label, weight
def th_iterproduct(*args):
return torch.from_numpy(np.indices(args).reshape((len(args),-1)).T)
def th_affine2d(x, matrix, mode='bilinear', center=True):
"""
2D Affine image transform on torch.Tensor
Arguments
---------
x : torch.Tensor of size (C, H, W)
image tensor to be transformed
matrix : torch.Tensor of size (3, 3) or (2, 3)
transformation matrix
mode : string in {'nearest', 'bilinear'}
interpolation scheme to use
center : boolean
whether to alter the bias of the transform
so the transform is applied about the center
of the image rather than the origin
Example
-------
>>> import torch
>>> from torchsample.utils import *
>>> x = torch.zeros(2,1000,1000)
>>> x[:,100:1500,100:500] = 10
>>> matrix = torch.FloatTensor([[1.,0,-50],
... [0,1.,-50]])
>>> xn = th_affine2d(x, matrix, mode='nearest')
>>> xb = th_affine2d(x, matrix, mode='bilinear')
"""
if matrix.dim() == 2:
matrix = matrix[:2,:]
matrix = matrix.unsqueeze(0)
elif matrix.dim() == 3:
if matrix.size()[1:] == (3,3):
matrix = matrix[:,:2,:]
A_batch = matrix[:,:,:2]
if A_batch.size(0) != x.size(0):
A_batch = A_batch.repeat(x.size(0),1,1)
b_batch = matrix[:,:,2].unsqueeze(1)
# make a meshgrid of normal coordinates
_coords = th_iterproduct(x.size(1),x.size(2))
coords = _coords.unsqueeze(0).repeat(x.size(0),1,1).float()
if center:
# shift the coordinates so center is the origin
coords[:,:,0] = coords[:,:,0] - (x.size(1) / 2. - 0.5)
coords[:,:,1] = coords[:,:,1] - (x.size(2) / 2. - 0.5)
# apply the coordinate transformation
new_coords = coords.bmm(A_batch.transpose(1,2)) + b_batch.expand_as(coords)
if center:
# shift the coordinates back so origin is origin
new_coords[:,:,0] = new_coords[:,:,0] + (x.size(1) / 2. - 0.5)
new_coords[:,:,1] = new_coords[:,:,1] + (x.size(2) / 2. - 0.5)
# map new coordinates using bilinear interpolation
if mode == 'nearest':
x_transformed = th_nearest_interp2d(x.contiguous(), new_coords)
elif mode == 'bilinear':
x_transformed = th_bilinear_interp2d(x.contiguous(), new_coords)
return x_transformed
def th_random_choice(a, n_samples=1, replace=True, p=None):
"""
Parameters
-----------
a : 1-D array-like
If a torch.Tensor, a random sample is generated from its elements.
If an int, the random sample is generated as if a was torch.range(n)
n_samples : int, optional
Number of samples to draw. Default is None, in which case a
single value is returned.
replace : boolean, optional
Whether the sample is with or without replacement
p : 1-D array-like, optional
The probabilities associated with each entry in a.
If not given the sample assumes a uniform distribution over all
entries in a.
Returns
--------
samples : 1-D ndarray, shape (size,)
The generated random samples
"""
if isinstance(a, int):
a = torch.arange(0, a)
if p is None:
if replace:
idx = torch.floor(torch.rand(n_samples)*a.size(0)).long()
else:
idx = torch.randperm(len(a))[:n_samples]
else:
if abs(1.0-sum(p)) > 1e-3:
raise ValueError('p must sum to 1.0')
if not replace:
raise ValueError('replace must equal true if probabilities given')
idx_vec = torch.cat([torch.zeros(round(p[i]*1000))+i for i in range(len(p))])
idx = (torch.floor(torch.rand(n_samples)*999)).long()
idx = idx_vec[idx].long()
selection = a[idx]
if n_samples == 1:
selection = selection[0]
return selection
def th_bilinear_interp2d(input, coords):
"""
bilinear interpolation in 2d
"""
x = torch.clamp(coords[:,:,0], 0, input.size(1)-2)
x0 = x.floor()
x1 = x0 + 1
y = torch.clamp(coords[:,:,1], 0, input.size(2)-2)
y0 = y.floor()
y1 = y0 + 1
stride = torch.LongTensor(input.stride())
x0_ix = x0.mul(stride[1]).long()
x1_ix = x1.mul(stride[1]).long()
y0_ix = y0.mul(stride[2]).long()
y1_ix = y1.mul(stride[2]).long()
input_flat = input.view(input.size(0),-1)
vals_00 = input_flat.gather(1, x0_ix.add(y0_ix))
vals_10 = input_flat.gather(1, x1_ix.add(y0_ix))
vals_01 = input_flat.gather(1, x0_ix.add(y1_ix))
vals_11 = input_flat.gather(1, x1_ix.add(y1_ix))
xd = x - x0
yd = y - y0
xm = 1 - xd
ym = 1 - yd
x_mapped = (vals_00.mul(xm).mul(ym) +
vals_10.mul(xd).mul(ym) +
vals_01.mul(xm).mul(yd) +
vals_11.mul(xd).mul(yd))
return x_mapped.view_as(input)
class Rotate(object):
def __init__(self,
value,
interp='bilinear',
lazy=False):
"""
Randomly rotate an image between (-degrees, degrees). If the image
has multiple channels, the same rotation will be applied to each channel.
Arguments
---------
rotation_range : integer or float
image will be rotated between (-degrees, degrees) degrees
interp : string in {'bilinear', 'nearest'} or list of strings
type of interpolation to use. You can provide a different
type of interpolation for each input, e.g. if you have two
inputs then you can say `interp=['bilinear','nearest']
lazy : boolean
if true, only create the affine transform matrix and return that
if false, perform the transform on the tensor and return the tensor
"""
self.value = value
self.interp = interp
self.lazy = lazy
def __call__(self, *inputs):
if not isinstance(self.interp, (tuple,list)):
interp = [self.interp]*len(inputs)
else:
interp = self.interp
theta = math.pi / 180 * self.value
rotation_matrix = torch.FloatTensor([[math.cos(theta), -math.sin(theta), 0],
[math.sin(theta), math.cos(theta), 0],
[0, 0, 1]])
if self.lazy:
return rotation_matrix
else:
outputs = []
for idx, _input in enumerate(inputs):
input_tf = th_affine2d(_input,
rotation_matrix,
mode=interp[idx],
center=True)
outputs.append(input_tf)
return outputs if idx > 1 else outputs[0]
class Translate(object):
def __init__(self,
value,
interp='bilinear',
lazy=False):
"""
Arguments
---------
value : float or 2-tuple of float
if single value, both horizontal and vertical translation
will be this value * total height/width. Thus, value should
be a fraction of total height/width with range (-1, 1)
interp : string in {'bilinear', 'nearest'} or list of strings
type of interpolation to use. You can provide a different
type of interpolation for each input, e.g. if you have two
inputs then you can say `interp=['bilinear','nearest']
"""
if not isinstance(value, (tuple,list)):
value = (value, value)
if value[0] > 1 or value[0] < -1:
raise ValueError('Translation must be between -1 and 1')
if value[1] > 1 or value[1] < -1:
raise ValueError('Translation must be between -1 and 1')
self.height_range = value[0]
self.width_range = value[1]
self.interp = interp
self.lazy = lazy
def __call__(self, *inputs):
if not isinstance(self.interp, (tuple,list)):
interp = [self.interp]*len(inputs)
else:
interp = self.interp
tx = self.height_range * inputs[0].size(1)
ty = self.width_range * inputs[0].size(2)
translation_matrix = torch.FloatTensor([[1, 0, tx],
[0, 1, ty],
[0, 0, 1]])
if self.lazy:
return translation_matrix
else:
outputs = []
for idx, _input in enumerate(inputs):
input_tf = th_affine2d(_input,
translation_matrix,
mode=interp[idx],
center=True)
outputs.append(input_tf)
return outputs if idx > 1 else outputs[0]
class RandomRotate(object):
def __init__(self,
rotation_range,
interp='bilinear',
lazy=False):
"""
Randomly rotate an image between (-degrees, degrees). If the image
has multiple channels, the same rotation will be applied to each channel.
Arguments
---------
rotation_range : integer or float
image will be rotated between (-degrees, degrees) degrees
interp : string in {'bilinear', 'nearest'} or list of strings
type of interpolation to use. You can provide a different
type of interpolation for each input, e.g. if you have two
inputs then you can say `interp=['bilinear','nearest']
lazy : boolean
if true, only create the affine transform matrix and return that
if false, perform the transform on the tensor and return the tensor
"""
self.rotation_range = rotation_range
self.interp = interp
self.lazy = lazy
def __call__(self, *inputs):
degree = random.uniform(-self.rotation_range, self.rotation_range)
if self.lazy:
return Rotate(degree, lazy=True)(inputs[0])
else:
outputs = Rotate(degree,
interp=self.interp)(*inputs)
return outputs
class RandomTranslate(object):
def __init__(self,
translation_range,
interp='bilinear',
lazy=False):
"""
Randomly translate an image some fraction of total height and/or
some fraction of total width. If the image has multiple channels,
the same translation will be applied to each channel.
Arguments
---------
translation_range : two floats between [0, 1)
first value:
fractional bounds of total height to shift image
image will be horizontally shifted between
(-height_range * height_dimension, height_range * height_dimension)
second value:
fractional bounds of total width to shift image
Image will be vertically shifted between
(-width_range * width_dimension, width_range * width_dimension)
interp : string in {'bilinear', 'nearest'} or list of strings
type of interpolation to use. You can provide a different
type of interpolation for each input, e.g. if you have two
inputs then you can say `interp=['bilinear','nearest']
lazy : boolean
if true, only create the affine transform matrix and return that
if false, perform the transform on the tensor and return the tensor
"""
if isinstance(translation_range, float):
translation_range = (translation_range, translation_range)
self.height_range = translation_range[0]
self.width_range = translation_range[1]
self.interp = interp
self.lazy = lazy
def __call__(self, *inputs):
# height shift
random_height = random.uniform(-self.height_range, self.height_range)
# width shift
random_width = random.uniform(-self.width_range, self.width_range)
if self.lazy:
return Translate([random_height, random_width],
lazy=True)(inputs[0])
else:
outputs = Translate([random_height, random_width],
interp=self.interp)(*inputs)
return outputs
class RandomFlip(object):
def __init__(self, h=True, v=False, p=0.5):
"""
Randomly flip an image horizontally and/or vertically with
some probability.
Arguments
---------
h : boolean
whether to horizontally flip w/ probability p
v : boolean
whether to vertically flip w/ probability p
p : float between [0,1]
probability with which to apply allowed flipping operations
"""
self.horizontal = h
self.vertical = v
self.p = p
def __call__(self, x, y=None):
x = x.numpy()
if y is not None:
y = y.numpy()
# horizontal flip with p = self.p
if self.horizontal:
if random.random() < self.p:
x = x.swapaxes(2, 0)
x = x[::-1, ...]
x = x.swapaxes(0, 2)
if y is not None:
y = y.swapaxes(2, 0)
y = y[::-1, ...]
y = y.swapaxes(0, 2)
# vertical flip with p = self.p
if self.vertical:
if random.random() < self.p:
x = x.swapaxes(1, 0)
x = x[::-1, ...]
x = x.swapaxes(0, 1)
if y is not None:
y = y.swapaxes(1, 0)
y = y[::-1, ...]
y = y.swapaxes(0, 1)
if y is None:
# must copy because torch doesnt current support neg strides
return torch.from_numpy(x.copy())
else:
return torch.from_numpy(x.copy()),torch.from_numpy(y.copy())
from torchvision import transforms
import random
import math
augmentor=transforms.Compose([
transforms.Lambda(lambda x: torch.Tensor(x)),
RandomRotate(25),
RandomTranslate([0.11, 0.11]),
RandomFlip(),
transforms.Lambda(lambda x: x.repeat(3, 1, 1, 1).permute(1, 0, 2, 3))
])
train_dataset = MRDataset('../input/kneescans/MRNet-v1.0/', injury_type,mri_view , train=True,transform=augmentor)
validation_dataset = MRDataset('../input/kneescans/MRNet-v1.0/', injury_type, mri_view, train=False)
# train_dataset = MRDataset('D:/MRNet-v1.0/MRNet-v1.0/', injury_type,mri_view , train=True,transform=augmentor)
# validation_dataset = MRDataset('D:/MRNet-v1.0/MRNet-v1.0/', injury_type, mri_view, train=False)
train_loader = torch.utils.data.DataLoader(
train_dataset, batch_size=1, shuffle=True, num_workers=0, drop_last=False)
validation_loader = torch.utils.data.DataLoader(
validation_dataset, batch_size=1, shuffle=True, num_workers=0, drop_last=False)
if torch.cuda.is_available():
device=torch.device("cuda:0")
print("running on GPU")
else:
device=torch.device("cpu")
print("running on cpu")
mrnet = MRNet().to(device)
optimizer = torch.optim.Adam(mrnet.parameters(), lr=1e-5, weight_decay=0.1)
scheduler = torch.optim.lr_scheduler.StepLR(
optimizer, step_size=3, gamma=0.5)
best_val_loss = float('inf')
best_val_auc = float(0)
num_epochs = 15
def train_model(model, train_loader, epoch, num_epochs, optimizer):
print("train_model")
print("Epoch:",epoch+1)
_ = model.train()
if torch.cuda.is_available():
device=torch.device("cuda:0")
print("running on GPU")
else:
device=torch.device("cpu")
print("running on cpu")
model.to(device)
y_preds = []
y_trues = []
losses = []
for i, (image, label, weight) in enumerate(train_loader):
print(i)
optimizer.zero_grad()
if torch.cuda.is_available():
image = image.to(device)
label = label.to(device)
weight = weight.to(device)
prediction = model.forward(image.float())
loss = torch.binary_cross_entropy_with_logits(
prediction[0], label[0], weight=weight[0])
loss.backward()
optimizer.step()
y_pred = torch.sigmoid(prediction).item()
y_true = int(label.item())
y_preds.append(y_pred)
y_trues.append(y_true)
try:
auc = metrics.roc_auc_score(y_trues, y_preds)
except:
auc = 0.5
loss_value = loss.item()
losses.append(loss_value)
train_loss_epoch = np.round(np.mean(losses), 4)
train_auc_epoch = np.round(auc, 4)
return train_loss_epoch, train_auc_epoch
def evaluate_model(model, val_loader, epoch, num_epochs):
print("Eval_model")
_ = model.eval()
if torch.cuda.is_available():
device=torch.device("cuda:0")
print("running on GPU")
else:
device=torch.device("cpu")
print("running on cpu")
model.to(device)
y_trues = []
y_preds = []
losses = []
for i, (image, label, weight) in enumerate(val_loader):
print(i)
if torch.cuda.is_available():
image = image.to(device)
label = label.to(device)
weight = weight.to(device)
prediction = model.forward(image.float())
loss = torch.binary_cross_entropy_with_logits(
prediction[0], label[0], weight=weight[0])
loss_value = loss.item()
losses.append(loss_value)
probas = torch.sigmoid(prediction)
y_trues.append(int(label[0]))
y_preds.append(probas[0].item())
try:
auc = metrics.roc_auc_score(y_trues, y_preds)
except:
auc = 0.5
val_loss_epoch = np.round(np.mean(losses), 4)
val_auc_epoch = np.round(auc, 4)
return val_loss_epoch, val_auc_epoch
train_loss_dict={}
train_auc_dict={}
val_loss_dict={}
val_auc_dict={}
for epoch in range(num_epochs):
train_loss, train_auc = train_model(
mrnet, train_loader, epoch, num_epochs, optimizer)
val_loss, val_auc = evaluate_model(
mrnet, validation_loader, epoch, num_epochs)
train_auc_dict[epoch+1]=train_auc
train_loss_dict[epoch+1]=train_loss
val_loss_dict[epoch+1]=val_loss
val_auc_dict[epoch+1]=val_auc
print("train loss : {0} | train auc {1} | val loss {2} | val auc {3}".format(
train_loss, train_auc, val_loss, val_auc))
scheduler.step()
print('-' * 30)
import matplotlib.pylab as plt
train_auc_lists = sorted(train_auc_dict.items()) # sorted by key, return a list of tuples
plt.style.use("ggplot")
plt.figure()
x, y = zip(*train_auc_lists) # unpack a list of pairs into two tuples
plt.xlabel("Number_of_Epochs")
plt.ylabel("Training_Accuracy")
plt.title("{}(Data Augmentation) - \nTrain/Accuracy ({}/{})".format(cnn_name,injury_name,mri_view.capitalize()+"_View"))
plt.plot(x, y)
#plt.savefig("VGG19(Data Augmentation) - Train/Accuracy (ACL_Tear/Sagittal_View).png")
plt.show()
import matplotlib.pylab as plt
train_loss_lists = sorted(train_loss_dict.items()) # sorted by key, return a list of tuples
plt.style.use("ggplot")
plt.figure()
x, y = zip(*train_loss_lists) # unpack a list of pairs into two tuples
plt.xlabel("Number_of_Epochs")
plt.ylabel("Training_Loss")
plt.title("{}(Data Augmentation) - \nTrain/Loss ({}/{})".format(cnn_name,injury_name,mri_view.capitalize()+"_View"))
plt.plot(x, y)
#plt.savefig("VGG19(Data Augmentation) - Train/Loss (ACL_Tear/Sagittal_View).png")
plt.show()
import matplotlib.pylab as plt
val_auc_lists = sorted(val_auc_dict.items()) # sorted by key, return a list of tuples
plt.style.use("ggplot")
plt.figure()
x, y = zip(*val_auc_lists) # unpack a list of pairs into two tuples
plt.xlabel("Number_of_Epochs")
plt.ylabel("Validation_Accuracy")
plt.title("{}(Data Augmentation) - \nValidation/Accuracy ({}/{})".format(cnn_name,injury_name,mri_view.capitalize()+"_View"))
plt.plot(x, y)
#plt.savefig("VGG19(Data Augmentation) - Validation/Accuracy (ACL_Tear/Sagittal_View).png")
plt.show()
import matplotlib.pylab as plt
val_loss_lists = sorted(val_loss_dict.items()) # sorted by key, return a list of tuples
plt.style.use("ggplot")
plt.figure()
x, y = zip(*val_loss_lists) # unpack a list of pairs into two tuples
plt.xlabel("Number_of_Epochs")
plt.ylabel("Validation_Loss")
plt.title("{}(Data Augmentation) - \nValidation/Loss ({}/{})".format(cnn_name,injury_name,mri_view.capitalize()+"_View"))
plt.plot(x, y)
#plt.savefig("VGG19(Data Augmentation) - Validation/Loss (ACL_Tear/Sagittal_View).png")
plt.show()
import matplotlib.pylab as plt
plt.style.use("ggplot")
plt.figure()
x, y = zip(*train_auc_lists)
x1, y1 = zip(*val_auc_lists)
plt.plot(x, y,label="train_auc")
plt.plot(x1, y1,label="val_auc")
# plt.title("VGG19(Data Augmentation) - \nTrain_Acc/Val_Acc (ACL_Tear/Sagittal_View)")
plt.title("{}(Data Augmentation) - \nTrain_Acc/Val_Acc ({}/{})".format(cnn_name,injury_name,mri_view.capitalize()+"_View"))
plt.xlabel("Epoch #")
plt.ylabel("Accuracy")
plt.legend(loc="lower left")
plt.show()
import matplotlib.pylab as plt
plt.style.use("ggplot")
plt.figure()
x, y = zip(*train_loss_lists)
x1, y1 = zip(*val_loss_lists)
plt.plot(x, y,label="train_loss")
plt.plot(x1, y1,label="val_loss")
# plt.title("VGG19(Data Augmentation) - \nTrain_Loss/Val_Loss (ACL_Tear/Sagittal_View)")
plt.title("{}(Data Augmentation) - \nTrain_Loss/Val_Loss ({}/{})".format(cnn_name,injury_name,mri_view.capitalize()+"_View"))
plt.xlabel("Epoch #")
plt.ylabel("Loss")
plt.legend(loc="lower left")
plt.show()
import matplotlib.pylab as plt
plt.style.use("ggplot")
plt.figure()
x, y = zip(*train_loss_lists)
x1, y1 = zip(*val_loss_lists)
x2, y2 = zip(*val_auc_lists)
x3, y3= zip(*train_auc_lists)
plt.plot(x, y,label="train_loss")
plt.plot(x1, y1,label="val_loss")
plt.plot(x2, y2,label="val_acc")
plt.plot(x3, y3,label="train_acc")
# plt.title("VGG19(Data Augmentation) - \nTrain_Acc/Val_Acc & Train_Loss/Val_Loss \n(ACL_Tear/Sagittal_View)")
plt.title("{}(Data Augmentation) - \nTrain_Acc/Val_Acc & Train_Loss/Val_Loss \n({}/{})".format(cnn_name,injury_name,mri_view.capitalize()+"_View"))
plt.xlabel("Epoch #")
plt.ylabel("Loss/Accuracy")
plt.legend(loc="lower left")
#plt.savefig("VGG19(Data Augmentation) - Validation/Loss (ACL_Tear/Sagittal_View).png")
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/u-masao/YutaroOgawa_pytorch_advanced/blob/master/1_image_classification/1-3_transfer_learning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# はじめに
『つくりながら学ぶ! PyTorchによる発展ディープラーニング』
のサンプルコードを Google Colaboratory で動作にしました。
オリジナルリポジトリ
> https://github.com/YutaroOgawa/pytorch_advanced
```
! git clone https://github.com/YutaroOgawa/pytorch_advanced.git
! ln -s pytorch_advanced/1_image_classification/data data
import os
import urllib.request
import zipfile
data_dir="./data"
# ImageNetのclass_indexをダウンロードする
# Kerasで用意されているものです
# https://github.com/fchollet/deep-learning-models/blob/master/imagenet_utils.py
url = "https://s3.amazonaws.com/deep-learning-models/image-models/imagenet_class_index.json"
save_path = os.path.join(data_dir, "imagenet_class_index.json")
if not os.path.exists(save_path):
urllib.request.urlretrieve(url, save_path)
# 1.3節で使用するアリとハチの画像データをダウンロードし解凍します
# PyTorchのチュートリアルで用意されているものです
# https://pytorch.org/tutorials/beginner/transfer_learning_tutorial.html
url = "https://download.pytorch.org/tutorial/hymenoptera_data.zip"
save_path = os.path.join(data_dir, "hymenoptera_data.zip")
if not os.path.exists(save_path):
urllib.request.urlretrieve(url, save_path)
# ZIPファイルを読み込み
zip = zipfile.ZipFile(save_path)
zip.extractall(data_dir) # ZIPを解凍
zip.close() # ZIPファイルをクローズ
# ZIPファイルを消去
os.remove(save_path)
```
# 1.3「転移学習」で少量データの分類を実現する方法
- 本ファイルでは、学習済みのVGGモデルを使用し、転移学習でアリとハチの画像を分類するモデルを学習します
# 学習目標
1. 画像データからDatasetを作成できるようになる
2. DataSetからDataLoaderを作成できるようになる
3. 学習済みモデルの出力層を任意の形に変更できるようになる
4. 出力層の結合パラメータのみを学習させ、転移学習が実装できるようになる
```
# パッケージのimport
import glob
import os.path as osp
import random
import numpy as np
import json
from PIL import Image
from tqdm import tqdm
import matplotlib.pyplot as plt
%matplotlib inline
import torch
import torch.nn as nn
import torch.optim as optim
import torch.utils.data as data
import torchvision
from torchvision import models, transforms
# 乱数のシードを設定
torch.manual_seed(1234)
np.random.seed(1234)
random.seed(1234)
```
# DataSetを作成
```
# 入力画像の前処理をするクラス
# 訓練時と推論時で処理が異なる
class ImageTransform():
"""
画像の前処理クラス。訓練時、検証時で異なる動作をする。
画像のサイズをリサイズし、色を標準化する。
訓練時はRandomResizedCropとRandomHorizontalFlipでデータオーギュメンテーションする。
Attributes
----------
resize : int
リサイズ先の画像の大きさ。
mean : (R, G, B)
各色チャネルの平均値。
std : (R, G, B)
各色チャネルの標準偏差。
"""
def __init__(self, resize, mean, std):
self.data_transform = {
'train': transforms.Compose([
transforms.RandomResizedCrop(
resize, scale=(0.5, 1.0)), # データオーギュメンテーション
transforms.RandomHorizontalFlip(), # データオーギュメンテーション
transforms.ToTensor(), # テンソルに変換
transforms.Normalize(mean, std) # 標準化
]),
'val': transforms.Compose([
transforms.Resize(resize), # リサイズ
transforms.CenterCrop(resize), # 画像中央をresize×resizeで切り取り
transforms.ToTensor(), # テンソルに変換
transforms.Normalize(mean, std) # 標準化
])
}
def __call__(self, img, phase='train'):
"""
Parameters
----------
phase : 'train' or 'val'
前処理のモードを指定。
"""
return self.data_transform[phase](img)
# 訓練時の画像前処理の動作を確認
# 実行するたびに処理結果の画像が変わる
# 1. 画像読み込み
image_file_path = './data/goldenretriever-3724972_640.jpg'
img = Image.open(image_file_path) # [高さ][幅][色RGB]
# 2. 元の画像の表示
plt.imshow(img)
plt.show()
# 3. 画像の前処理と処理済み画像の表示
size = 224
mean = (0.485, 0.456, 0.406)
std = (0.229, 0.224, 0.225)
transform = ImageTransform(size, mean, std)
img_transformed = transform(img, phase="train") # torch.Size([3, 224, 224])
# (色、高さ、幅)を (高さ、幅、色)に変換し、0-1に値を制限して表示
img_transformed = img_transformed.numpy().transpose((1, 2, 0))
img_transformed = np.clip(img_transformed, 0, 1)
plt.imshow(img_transformed)
plt.show()
# アリとハチの画像へのファイルパスのリストを作成する
def make_datapath_list(phase="train"):
"""
データのパスを格納したリストを作成する。
Parameters
----------
phase : 'train' or 'val'
訓練データか検証データかを指定する
Returns
-------
path_list : list
データへのパスを格納したリスト
"""
rootpath = "./data/hymenoptera_data/"
target_path = osp.join(rootpath+phase+'/**/*.jpg')
print(target_path)
path_list = [] # ここに格納する
# globを利用してサブディレクトリまでファイルパスを取得する
for path in glob.glob(target_path):
path_list.append(path)
return path_list
# 実行
train_list = make_datapath_list(phase="train")
val_list = make_datapath_list(phase="val")
train_list
# アリとハチの画像のDatasetを作成する
class HymenopteraDataset(data.Dataset):
"""
アリとハチの画像のDatasetクラス。PyTorchのDatasetクラスを継承。
Attributes
----------
file_list : リスト
画像のパスを格納したリスト
transform : object
前処理クラスのインスタンス
phase : 'train' or 'test'
学習か訓練かを設定する。
"""
def __init__(self, file_list, transform=None, phase='train'):
self.file_list = file_list # ファイルパスのリスト
self.transform = transform # 前処理クラスのインスタンス
self.phase = phase # train or valの指定
def __len__(self):
'''画像の枚数を返す'''
return len(self.file_list)
def __getitem__(self, index):
'''
前処理をした画像のTensor形式のデータとラベルを取得
'''
# index番目の画像をロード
img_path = self.file_list[index]
img = Image.open(img_path) # [高さ][幅][色RGB]
# 画像の前処理を実施
img_transformed = self.transform(
img, self.phase) # torch.Size([3, 224, 224])
# 画像のラベルをファイル名から抜き出す
if self.phase == "train":
label = img_path[30:34]
elif self.phase == "val":
label = img_path[28:32]
# ラベルを数値に変更する
if label == "ants":
label = 0
elif label == "bees":
label = 1
return img_transformed, label
# 実行
train_dataset = HymenopteraDataset(
file_list=train_list, transform=ImageTransform(size, mean, std), phase='train')
val_dataset = HymenopteraDataset(
file_list=val_list, transform=ImageTransform(size, mean, std), phase='val')
# 動作確認
index = 0
print(train_dataset.__getitem__(index)[0].size())
print(train_dataset.__getitem__(index)[1])
```
# DataLoaderを作成
```
# ミニバッチのサイズを指定
batch_size = 32
# DataLoaderを作成
train_dataloader = torch.utils.data.DataLoader(
train_dataset, batch_size=batch_size, shuffle=True)
val_dataloader = torch.utils.data.DataLoader(
val_dataset, batch_size=batch_size, shuffle=False)
# 辞書型変数にまとめる
dataloaders_dict = {"train": train_dataloader, "val": val_dataloader}
# 動作確認
batch_iterator = iter(dataloaders_dict["train"]) # イテレータに変換
inputs, labels = next(
batch_iterator) # 1番目の要素を取り出す
print(inputs.size())
print(labels)
```
# ネットワークモデルの作成する
```
# 学習済みのVGG-16モデルをロード
# VGG-16モデルのインスタンスを生成
use_pretrained = True # 学習済みのパラメータを使用
net = models.vgg16(pretrained=use_pretrained)
print(net)
# VGG16の最後の出力層の出力ユニットをアリとハチの2つに付け替える
net.classifier[6] = nn.Linear(in_features=4096, out_features=2)
print(net)
# 訓練モードに設定
net.train()
print('ネットワーク設定完了:学習済みの重みをロードし、訓練モードに設定しました')
```
# 損失関数を定義
```
# 損失関数の設定
criterion = nn.CrossEntropyLoss()
```
# 最適化手法を設定
```
# 転移学習で学習させるパラメータを、変数params_to_updateに格納する
params_to_update = []
# 学習させるパラメータ名
update_param_names = ["classifier.6.weight", "classifier.6.bias"]
# 学習させるパラメータ以外は勾配計算をなくし、変化しないように設定
for name, param in net.named_parameters():
if name in update_param_names:
param.requires_grad = True
params_to_update.append(param)
print(name)
else:
param.requires_grad = False
# params_to_updateの中身を確認
print("-----------")
print(params_to_update)
# 最適化手法の設定
optimizer = optim.SGD(params=params_to_update, lr=0.001, momentum=0.9)
```
# 学習・検証を実施
```
# モデルを学習させる関数を作成
def train_model(net, dataloaders_dict, criterion, optimizer, num_epochs):
# epochのループ
for epoch in range(num_epochs):
print('Epoch {}/{}'.format(epoch+1, num_epochs))
print('-------------')
# epochごとの学習と検証のループ
for phase in ['train', 'val']:
if phase == 'train':
net.train() # モデルを訓練モードに
else:
net.eval() # モデルを検証モードに
epoch_loss = 0.0 # epochの損失和
epoch_corrects = 0 # epochの正解数
# 未学習時の検証性能を確かめるため、epoch=0の訓練は省略
if (epoch == 0) and (phase == 'train'):
continue
# データローダーからミニバッチを取り出すループ
for inputs, labels in tqdm(dataloaders_dict[phase]):
# optimizerを初期化
optimizer.zero_grad()
# 順伝搬(forward)計算
with torch.set_grad_enabled(phase == 'train'):
outputs = net(inputs)
loss = criterion(outputs, labels) # 損失を計算
_, preds = torch.max(outputs, 1) # ラベルを予測
# 訓練時はバックプロパゲーション
if phase == 'train':
loss.backward()
optimizer.step()
# イタレーション結果の計算
# lossの合計を更新
epoch_loss += loss.item() * inputs.size(0)
# 正解数の合計を更新
epoch_corrects += torch.sum(preds == labels.data)
# epochごとのlossと正解率を表示
epoch_loss = epoch_loss / len(dataloaders_dict[phase].dataset)
epoch_acc = epoch_corrects.double(
) / len(dataloaders_dict[phase].dataset)
print('{} Loss: {:.4f} Acc: {:.4f}'.format(
phase, epoch_loss, epoch_acc))
# 学習・検証を実行する
num_epochs=2
train_model(net, dataloaders_dict, criterion, optimizer, num_epochs=num_epochs)
```
以上
| github_jupyter |
# EnKF Assumption Experiments
### Keiran Suchak
Assumptions to test:
* Normality of prior
* Normality of likelihood
* Subsequent normality of posterior
This notebook will make use of the `multivariate_normality()` function from the `pingouin` package to perform multidimensional normality tests.
## Imports
```
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pickle
import pingouin as pg
import seaborn as sns
import sys
%matplotlib inline
sys.path.append('../../../stationsim/')
from ensemble_kalman_filter import EnsembleKalmanFilter, EnsembleKalmanFilterType
from stationsim_gcs_model import Model
np.random.seed(28)
```
## Functions
```
def tidy_dataframe(df, independent_col: str, dependent_cols: list):
output = list()
for i, row in df.iterrows():
for col in dependent_cols:
d = {independent_col: row[independent_col],
'variable': col,
'value': row[col]}
output.append(d)
output = pd.DataFrame(output)
return output
```
## Experiment 0: Testing `pg.multivariate_normality()`
Create a sample of 5000 $x$-$y$ coordinates from a 2-dimensional normal distribution.
```
mean = [0, 0]
cov = [[1, 0], [0, 100]]
x, y = np.random.multivariate_normal(mean, cov, 5000).T
```
Plot samples in $x$-$y$ space.
```
plt.figure()
plt.plot(x, y, 'x')
plt.xlabel('x')
plt.ylabel('y')
plt.show()
```
Test for normality.
```
X = pd.DataFrame({'x': x, 'y': y})
pg.multivariate_normality(X, alpha=0.05)
```
The test did not find sufficient evidence to reject the null hypothesis, i.e. the data are normally distributed.
Let us now consider data drawn from a distribution that is not gaussian.
In this case, we draw the $x$-$y$ coordinates from two uniform distributions, \[0.0, 1.0\).
```
x, y = np.random.random_sample((2, 5000))
```
Plot samples in $x$-$y$ space.
```
plt.figure()
plt.plot(x, y, 'x')
plt.xlabel('x')
plt.ylabel('y')
plt.show()
```
Test for normality.
```
X = pd.DataFrame({'x': x, 'y': y})
pg.multivariate_normality(X, alpha=0.05)
```
The test correctly finds sufficient evidence to reject the null hypothesis that the data are normally distributed.
We can make a couple of functions to generate normally- and uniformly-distributed samples of arbitrary size and check that the test works for different sample sizes.
```
def normal_sample_2d(N):
mean = [0, 0]
cov = [[1, 0], [0, 100]]
x, y = np.random.multivariate_normal(mean, cov, N).T
X = pd.DataFrame({'x': x, 'y': y})
return X
def uniform_sample_2d(N):
x, y = np.random.random_sample((2, N))
X = pd.DataFrame({'x': x, 'y': y})
return X
def test_multidim_normality(X):
t = pg.multivariate_normality(X)
return t.normal
```
Now we can run through a collection of different sample sizes, each time generating that number of random samples from both normal and uniform distributions and testing whether `pg.multivariate_normality()` found the samples to be normally distributed, or whether sufficient evidence was found to reject the null hypothesis.
The sample sizes to be used are `[10, 20, 50, 100, 200, 500, 1000, 2000]`.
This selection has been chosen to observe how the test performs on different scales of sample size.
In each case, the testing process shall be run $20$ times to account for the randomness of the samples and the fact that the test may incorrectly consider normally distributed data to be non-normal (or vice-versa).
```
results = list()
sample_sizes = [10, 20, 50, 100, 200, 500, 1000, 2000]
n_runs = 20
for ss in sample_sizes:
for _ in range(n_runs):
d = {'sample_size': ss}
normal_sample = normal_sample_2d(ss)
uniform_sample = uniform_sample_2d(ss)
d['gaussian'] = test_multidim_normality(normal_sample)
d['non-gaussian'] = test_multidim_normality(uniform_sample)
results.append(d)
```
Let's convert these results into a dataframe.
```
results = pd.DataFrame(results)
results.head()
```
We can now find the proprtion of cases in each scenario for which the test correctly accepted/rejected the null hypothesis.
```
proportions = list()
for ss in sample_sizes:
tdf = results.loc[results['sample_size']==ss, ]
d = {'sample_size': ss}
d['gaussian'] = tdf['gaussian'].sum() / len(tdf['gaussian'])
d['non-gaussian'] = tdf['non-gaussian'].sum() / len(tdf['non-gaussian'])
proportions.append(d)
```
Again, converting this to a dataframe.
```
proportions = pd.DataFrame(proportions)
proportions.head()
plt.figure()
plt.semilogx(proportions['sample_size'], proportions['gaussian'],
label='np.random.multivariate_normal')
plt.semilogx(proportions['sample_size'], proportions['non-gaussian'],
label='np.random.random_sample')
plt.xlabel('Sample size')
plt.ylabel('Proportion accepted as gaussian')
plt.legend()
plt.show()
```
From the above figure, we can see that the test correctly identifies data from `np.random.multivariate_normal()` as gaussian the majority of the time for all sample sizes.
We can also see that, for very small sample sizes (i.e. $N<50$), the test typically does not find sufficient evidence to reject the null hypothesis of normality for non-gaussian data.
We should, therefore, ensure that our sample sizes are sufficiently large when using the test with data from the Ensemble Kalman Filter.
It is also worth considering how this scales with the dimensions of the data - when working with state vectors from the Ensemble Kalman Filter, we consider our sample size to be the filter's ensemble size and twice the model population size to be our number of dimensions.
In order to test this, we will need updated version of the functions used to generate samples - the new versions of these functions should generalise such that we can generate $m$-dimensional data samples.
```
def __convert_to_df(Y, m):
d = {'var_{0}'.format(i): Y[i] for i in range(m)}
X = pd.DataFrame(d)
return X
def normal_sample_d(N, m):
mean = np.zeros(m)
cov = np.identity(m)
Y = np.random.multivariate_normal(mean, cov, N).T
X = __convert_to_df(Y, m)
return X
def uniform_sample_d(N, m):
Y = np.random.random_sample((m, N))
X = __convert_to_df(Y, m)
return X
```
Now that we have constructed the functions, let us test them out with $200$ samples from $10$-dimensional distribtutions.
We can start by sampling from the multivariate normal distribution:
```
z = normal_sample_d(200, 10)
z
z.shape
```
And just to check, we can plot the first $2$ dimensions.
```
plt.figure()
plt.plot(z['var_0'], z['var_1'], 'x')
plt.xlabel('x')
plt.ylabel('y')
plt.show()
```
Similarly we can sample from the uniform distribution.
```
z = uniform_sample_d(200, 10)
z
```
And we can, again, plot the first $2$ dimensions.
```
plt.figure()
plt.plot(z['var_0'], z['var_1'], 'x')
plt.xlabel('x')
plt.ylabel('y')
plt.show()
```
Well it looks like these functions work!
Now we can make use of them in conjunction with `pg.multivariate_normality()` to see how the normality test responds to different population sizes and ensemble sizes (otherwise referred to as different dimensionalities and sample sizes).
```
results = list()
sample_sizes = [50, 100, 200, 500, 1000]
dimensionalities = list(range(5, 105, 5))
n_runs = 20
for ss in sample_sizes:
for dimensionality in dimensionalities:
print(f'Running sample size={ss}, dimensionality={dimensionality}')
for _ in range(n_runs):
# Make dictionary for gaussian data
normal_sample = normal_sample_d(ss, dimensionality)
test_result = test_multidim_normality(normal_sample)
d = {'sample_size': ss,
'dimensionality': dimensionality,
'kind': 'gaussian',
'result': test_result}
results.append(d)
# Make dictionary for non-gaussian data
uniform_sample = uniform_sample_d(ss, dimensionality)
test_result = test_multidim_normality(uniform_sample)
d = {'sample_size': ss,
'dimensionality': dimensionality,
'kind': 'non-gaussian',
'result': test_result}
results.append(d)
```
Convert results to a dataframe.
```
results = pd.DataFrame(results)
results.head()
results['dimensionality'].unique()
```
We now wish to visualise how the number of
To do this, we first create a filtered dataset where we filter out the rows for which the tests returned false.
```
results_f = results.loc[results['result']==True, ['sample_size', 'dimensionality', 'kind']]
results_f.head()
results_f['dimensionality'].unique()
```
We can now create our kde-plot, segregating the data for gaussian and non-gaussian samples.
```
g = sns.FacetGrid(results_f, col='dimensionality', col_wrap=4)
g.map_dataframe(sns.histplot, x='sample_size', hue='kind', kde=True, log_scale=True)
g.set_axis_labels('Sample size', 'Gaussian sample sets')
g.add_legend()
```
## Experiment 1: Pure forecasting
In this experiment, we will not be assimilating any data, i.e. each step of the model ensemble will only consist of the forecast process.
Following each forecast step, the ensemble will be tested for normality using the `multivariate_normality()` function from the `pingouin` package.
In order to keep the process simple at this stage, a small population of agents will be used, allowing us to use a large ensemble size.
This will act as a preliminary experiment towards demonstrating the normality of the ensemble prior distribution.
```
results = list()
# Set up filter parameters
ensemble_size = 50
pop_size = 2
state_vec_length = 2 * pop_size
# Initialise filter with StationSim and params
filter_params = {'vanilla_ensemble_size': ensemble_size,
'state_vector_length': state_vec_length,
'mode': EnsembleKalmanFilterType.STATE}
model_params = {'pop_total': pop_size,
'station': 'Grand_Central',
'do_print': False}
enkf = EnsembleKalmanFilter(Model, filter_params, model_params,
filtering=False, benchmarking=True)
while enkf.active:
enkf.baseline_step()
results.append(enkf.vanilla_state_ensemble.copy())
len(results)
all_xs = list()
all_ys = list()
for i in range(10, len(results), len(results)//10):
state = results[i]
xs = state[::2]
ys = state[1::2]
all_xs.append(xs)
all_ys.append(ys)
plt.figure()
plt.scatter(xs, ys, s=5, marker='.')
plt.xlim((0, 750))
plt.ylim((0, 750))
plt.show()
norm_results = list()
for state in results:
stateT = state.T
normality = pg.multivariate_normality(stateT, alpha=1)
norm_results.append(normality.normal)
sum(norm_results)
print(all_xs[0])
print(all_ys[0])
for xs in all_xs:
plt.figure()
for x in xs:
plt.hist(x, alpha=0.5)
plt.show()
for ys in all_ys:
plt.figure()
for y in ys:
plt.hist(y, alpha=0.5)
plt.show()
```
| github_jupyter |
This is the first of a series about bitcoin futures on the exchange [Deribit.com](https://www.deribit.com). We will download the data we need to do our future analysis and get some background on how futures work.
Over the entire series I plan to (1) explore historical prices on bitcoin futures, (2) discover potential arbitrage opportunities and (3) do a full investment analysis of the potential profits from arbitrage strategies.
- [Part 1 - Getting the data]({% post_url 2019-05-11-bitcoin-futures-arbitrage-part-1 %})
- [Part 2 - Were there arbitrage profits in the past?]({% post_url 2019-05-12-bitcoin-futures-arbitrage-part-2 %})
- [Part 3 - Perpetual futures 101]({% post_url 2019-05-20-bitcoin-futures-arbitrage-part-3 %})
- [Part 4 - Arbitrage Profit Analysis]({% post_url 2019-05-24-bitcoin-futures-arbitrage-part-4 %})
## Background Information
Users on Deribit can trade in cryptocurrency derivates including options and futures. However, users are not able to buy or sell cryptocurrency on the platform. It doesn't accept traditional fiat currency; all deposits and withdrawals are done in crypto. Both the futures and options are cash-settled, simply transferring the profits and losses between the bitcoin accounts of the platform users.
However, derivative prices are always based on an underlying asset price. The bitcoin futures and options at Deribit have payoffs based on the price of bitcoin in USD.
Thus, Deribit needs a way to track the fiat price of Bitcoin to determine the payoffs on options and futures. This brings us to our first important concept.
### The Deribit Bitcoin Index
Deribit's Bitcoin index is **the** price of bitcoin by which all their derivatives profits are based on. It is an average of the current mid-price on 6 large cryptocurrency exchanges. Further information can be found [here](https://www.deribit.com/main#/indexes). Whenever I mention the *index* I mean this.
### Regular Futures vs Perpetual Futures
Deribit sells two types of futures, regular and perpetual.
Any listings with an expiry date are just typical futures contracts. The specification can be found [here](https://www.deribit.com/pages/docs/futures). Wiki and Investopedia can provide good explanations of how they work. When I refer to a *future* in this document, I mean a typical future.
Perpetual futures are very different and have unusual characteristics. We will discuss them in length in part 2. When I refer to a *perpetual*, I mean this special type of perpetual future.
## Historical Future Price Data
Let's download the data! Below is some python code that you can use to get all trade data for any instrument. I used this to get the sales price of every transaction for all Deribit futures in the last half of 2018 and 2019, which I will start exploring in part 2.
```
import requests
from time import sleep
def download(instrument_names):
"""Downloads all past data for the provided instument names
Parameters
----------
instrument_names: iterable
The list of instrument names to download.
"""
for name in instrument_names:
with open(f'downloads/{name}.txt', 'w') as txt:
txt.write('timestamp,instrument_name,price,index_price\n')
has_more = True
seq = 1
count = 1000
while has_more:
url = f'https://www.deribit.com/api/v2/public/get_last_trades_by_instrument?instrument_name={name}&start_seq={seq}&count={count}&include_old=true'
r = None
for _ in range(5):
while r is None:
try:
r = requests.get(url, timeout=5)
except Timeout:
sleep(2)
pass
r = r.json()
for trade in r['result']['trades']:
timestamp = trade['timestamp']
instrument_name = trade['instrument_name']
price = trade['price']
index_price = trade['index_price']
txt.write(f'{timestamp},{instrument_name},{price},{index_price}\n')
seq += count
has_more = r['result']['has_more']
def get_instrument_names(currency='BTC', kind='future', expired='true'):
"""Get instrument names
Parameters
----------
currency: string
The currency of instrument names to download
kind: string
'future' or 'option'
expired: bool
past instruments too or only current ones
"""
url = f'https://www.deribit.com/api/v2/public/get_instruments?currency={currency}&kind={kind}&expired={expired}'
r = None
for x in range(5):
while r is None:
try:
r = requests.get(url, timeout=5)
except Timeout:
sleep(2)
pass
r = r.json()
for instrument in r['result']:
yield instrument['instrument_name']
```
| github_jupyter |
# BHPToolkit Spring 2020 Workshop: EMRISur1dq1e4 Proejct Tutorial
Some portions of this notebook are also found in the notebook
[EMRISur1dq1e4.ipynb](https://github.com/BlackHolePerturbationToolkit/EMRISurrogate/blob/master/EMRISur1dq1e4.ipynb). The waveform model is described in [arXiv:1910.10473](https://arxiv.org/abs/1910.10473).
EMRISur1dq1e4 is a surrogate gravitational-waveform model for non-spinning black hole binary systems with mass-ratios varying from 3 to $10^4$. This surrogate model is trained on waveform data generated by point-particle black hole perturbation theory (ppBHPT), with the total mass rescaling paramter tuned to NR simulations according to the paper's Eq. 4. Available modes are $\{(2,2), (2,1), (3,3), (3,2), (3,1), (4,4), (4,3), (4,2), (5,5), (5,4), (5,3)\}$. The $m<0$ modes can be deduced from the m>0 modes due to symmetry of the system about the orbital plane.
**NOTE**: This notebook rquires the file emri_sur_load.py and EMRISur1dq1e4.h5 datafile in the same directory for waveform generation
# Setup
1. You should now have the programs git and Jupyter as well as the Python packages scipy, h5py, and hashlib
2. clone the EMRISurrogate code and this tutorial
```shell
>>> git clone https://github.com/BlackHolePerturbationToolkit/EMRISurrogate.git
>>> cd EMRISurrogate # move into the new directory
>>> jupyter notebook BHPToolkit_Spring2020_Tutorial.ipynb # launch this tutorial
```
(no-git fallback plan: download the zip file from https://github.com/BlackHolePerturbationToolkit/EMRISurrogate)
```
# If your setup is correct, then this block of code should execute...
import matplotlib.pyplot as plt
import numpy as np
import hashlib
import h5py
```
# Lesson 1: The data
Surrogate models are numerical models. They require code *and* data
The EMRI surrogate model's data is [hosted on zenodo](https://zenodo.org/record/3612600#.XsoAP3VKg5k).
Our first task is to download the data:
```shell
>>> wget https://zenodo.org/record/3612600/files/EMRISur1dq1e4.h5
```
or click the "download" button and move the file to the same folder as this notebook.
## Make sure your data is at the latest version
The data file has a unique hash, which from zenodo is d145958484738e0c7292e084a66a96fa.
If the surrogate model is updated then EMRISur1dq1e4.h5 will be replaced with a newer file.
The [EMRI surrogate code](https://github.com/BlackHolePerturbationToolkit/EMRISurrogate/blob/master/emri_sur_load.py#L48) checks that your local h5 file's hash matches the most recent version.
<br>
<img src="hash_check.png" width="940" />
```
# data integrity: lets check that your file's hash is correct
# Current hash: d145958484738e0c7292e084a66a96fa
def md5(fname):
hash_md5 = hashlib.md5()
with open(fname, "rb") as f:
for chunk in iter(lambda: f.read(4096), b""):
hash_md5.update(chunk)
return hash_md5.hexdigest()
print( md5('EMRISur1dq1e4.h5') )
# now import the some functions to help with evaluating the model
# This step requires that you've cloaned the git
# project EMRISurrogate and have the data file listed above
import emri_sur_load as emriSur
```
## Data file's structure
Recall that the full surrogate model is
\begin{align}
h_{\tt S}(t,\theta,\phi;q) & = \sum_{\ell,m} h_{\tt S}^{\ell,m}(t;q) {}_{-2}Y_{\ell m} (\theta,\phi) \\
h_{\tt S}^{\ell,m}(t;q) & = A_{\tt S}^{\ell,m}(t;q) \exp(- \mathrm{i} \phi_{\tt S}^{\ell,m}(t;q)) \,,
\end{align}
and we build models for $A_{\tt S}^{\ell,m}$, $\phi_{\tt S}^{\ell,m}$. Each $A_{\tt S}^{\ell,m}$ and $\phi_{\tt S}^{\ell,m}$ is represented as an empirical interpolant (EI) with parametric fits that the time nodes.
Lets check that our data file matches this structure.
```
# Each mode is a separate surrogate model
# open the file
fp = h5py.File("EMRISur1dq1e4.h5","r")
# inspect the data groups
print("EMRI surrogate data groups: ",fp.keys() )
print("\n\n" )
# grab the 22 mode data group
sur_22 = fp["l2_m2"]
# inspect (2,2)-mode's data groups
print("22 mode surrogate data groups: ",sur_22.keys() )
```
## The surrogate model for $A^{22}(t;q)$
Do to an unfortunate choice made a long time ago, the quantitites without any extra postfix denote ampitude date: B_phase is the basis for the phase, while B is the basis for the amplitude. So we have the following data for the 22 mode's amplitude:
### Information about the amplitude's parametric dependence
* 'fit_type_amp': model for the amplitude's dependence with mass ratio
* 'parameterization': how we parameterize the amplitude data
* 'fit_max' / 'fit_min' : smallest / largest values of parametric domain
* 'n_spline_knots': number of spline knots
* 'spline_knots': location of spline knots
* 'fitparams_amp': spline parameters
* 'degree': degree of the spline used
### Information about the amplitude's temporal dependence
* 'B': basis
* 'eim_indicies': location of empirical interpolation nodes
* 'times': temporal domain on which the amplitude is modeled
The model is given by
\begin{align}
A_{\tt S}^{22}(t;q) = \sum_{i=0}^9 A(T_i^\mathrm{EIM};q) B_i(t) \,,
\end{align}
where $B_i(t)$ is the $i^{th}$ basis function and $T_i^\mathrm{EIM}$ is the $i^{th}$ empirical interpolation time node. We see that, $A(T_i^\mathrm{EIM};q)$ are the coefficients defining the amplitude's expansion in terms of the basis.
Data found in "Information about the amplitude's parametric dependence" is used to model each coefficient, $A(T_i^\mathrm{EIM};q)$, over the paramter space.
```
# Lets inspect the basis for the 22 mode's amplitude
# Because each basis is a linear combination of amplitudes,
# the basis will look kinda like an waveform amplitude.
# The basis will not look like any standard function (polynomials, sine/cosine, etc...)
### plot the ith cardinal basis function ###
i = 0
times = sur_22['times']
B = sur_22['B']
plt.figure(1)
plt.plot(times,B[:,i])
plt.title('Cardinal basis %i for amplitude'%i)
plt.show()
# Lets inspect the empirical interpolation time nodes for the
# 22 mode's amplitude.
# These nodes are used to define the interpolation problem: the
# surrogate interpolates the waveform training data at these times.
# The time nodes will automatically be placed in regions of higher
# activitiy. Unlike uniform, Chebyshev, or other common time nodes
# the EI nodes are adaptive to the problem
### plot the location of all emprical interpolation points ###
plt.figure(2)
eim_indicies = sur_22['eim_indicies'][:]
plt.plot(times,B[:,0]) # 0^th basis function
plt.plot(times[np.sort(eim_indicies)],np.zeros_like(eim_indicies),'r*')
plt.title('EI time node locations')
plt.figure(3)
eim_indicies = sur_22['eim_indicies'][:]
plt.plot(times,B[:,0]) # 0^th basis function
plt.plot(times[np.sort(eim_indicies)],np.zeros_like(eim_indicies),'r*')
plt.xlim([-500,100])
plt.title('EI time node locations')
```
# Lesson 2: Evaluating the model
Input: mass ratio q, and (optionally) modes to generate = [(2,1),(2,2),(3,1),(3,2),(3,3),(4,2),(4,3),(4,4),(5,3),(5,4),(5,5)]
Output: dictionary of modes, $h_{\tt S}^{\ell,m}(t;q)$
\begin{align}
h_{\tt S}(t,\theta,\phi;q) & = \sum_{\ell,m} h_{\tt S}^{\ell,m}(t;q) {}_{-2}Y_{\ell m} (\theta,\phi)
\end{align}
Both h and t are in geometric units
```
q = 10.0 # mass ratio
time, h = emriSur.generate_surrogate(q) # h is a dictionary with modes as its key
# h is a dictionary of modes
h_22=h[(2,2)]
h_21=h[(2,1)]
plt.figure(figsize=(14,4))
plt.plot(time,np.real(h_22),label='{2,2} mode')
plt.plot(time,np.real(h_21),label='{2,1} mode')
plt.xlabel('t',fontsize=12)
plt.ylabel('h(t)',fontsize=12)
plt.legend(fontsize=12)
#plt.savefig('emri_sur_q_%f.png'%q)
plt.show()
# waveform in mks units (roughly what the detector would observe)
G=6.674*1e-11
MSUN_SI = 1.9885469549614615e+30
PC_SI = 3.085677581491367e+16
C_SI = 299792458.0
# values of M and dL
M=80.0*MSUN_SI
dL=100.0* PC_SI
# scaling of time and h(t)
time=time*(G*M/C_SI**3)
ht22=np.array(h[(2,2)])*(G*M/C_SI**3)/dL
plt.figure(figsize=(14,4))
plt.title('M=80, dL=100.0')
plt.plot(time,np.real(ht22),label='{2,2} mode')
plt.xlabel('t',fontsize=12)
plt.ylabel('h(t)',fontsize=12)
plt.legend(fontsize=12)
#plt.savefig('emri_sur_q_%f_physical.png'%q)
plt.show()
```
# Lesson 3: EMRI surrogate vs output from the Teukolsky solver
Our underlying model (denoted $h^{\ell,m}_{\tt S}$ below) is for the output of the Teukolsky solver for point-particle perturbation theory.
However, the EMRI surrogate model (denoted $h^{\ell,m}_{\tt S, \alpha}$ below) is calibrated to numerical relativity waveforms at comparable mass binaries with a single parameter $\alpha$:
\begin{align*}
h^{\ell,m}_{\tt S, \alpha}(t ; q)= {\alpha} h^{\ell,m}_{\tt S}\left( t \alpha;q \right) \,,
\end{align*}
Which is enacted in emri_sur_load.py file as shown below:
<img src="Alpha_scaling_in_code.png" width="840" />
### To generate output from the Teukolsky solver (no NR calibration) simply set $\alpha = 1$ in lines 110 and 118
(Issue tracker: feature request to allow for this without needing to modify the code)
<img src="Alpha_scaling_in_code.png" width="840" />
# Summary
1. To understand how to use the EMRI surrogate model, please consult the Jupyter noteobook found with the [EMRISur1dq1e4 proejct](https://github.com/BlackHolePerturbationToolkit/EMRISurrogate/blob/master/EMRISur1dq1e4.ipynb) in addition to this notebook.
2. Future EMRI surrogate models or improvements will be added here.
3. If you find any issues or have suggestions please open up an issue (or, better yet, a pull request!)
<br>
<img src="Issue1.png" width="840" />
| github_jupyter |
<h1>Содержание<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Список-(list)" data-toc-modified-id="Список-(list)-1">Список (list)</a></span></li><li><span><a href="#Дек-(collections.deque)" data-toc-modified-id="Дек-(collections.deque)-2">Дек (collections.deque)</a></span></li><li><span><a href="#Множество-(set)" data-toc-modified-id="Множество-(set)-3">Множество (set)</a></span></li><li><span><a href="#Словарь-(dict)" data-toc-modified-id="Словарь-(dict)-4">Словарь (dict)</a></span></li></ul></div>
# Время работы операций в контейнерах
## Список (list)
| Операция | Результат | В среднем | В худшем случае |
| --- | --- | --- | --- |
| `x in s` | True, если в s есть элемент, равный x, иначе False | O(n) ||
| `x not in s` | False, если в s есть элемент, равный x, иначе True| O(n) ||
| `s + t` | Конкатенация s и t | O(len(s) + len(t)) | O(len(s) + len(t)) |
| `s * k` или `k * s` | Добавление s к самому себе k раз | O(len(s) * k) | O(len(s) * k) |
| `s[i]` | Взять i-ый элемент | O(1) | O(1) |
| `s[i] = x` | Установить i-ый элемент | O(1) | O(1) |
| `del s[i]` | Удалить i-ый элемент | O(n) | O(n) |
| `s[i:j]` | Взять срез s от i до j | O(j - i) | O(j - i) |
| `s[i:j:k]` | Взять срез s от i до j с шагом k | O((j - i) // k) | O((j - i) // k) |
| `s[i:j] = t` | Изменение среза | O(len(s) + len(t)) | O(len(s) + len(t)) |
| `del s[i:j]` | Удаление среза | O(n) | O(n) |
| `for x in s` | Итерация | O(n) | O(n) |
| `len(s)` | Длина s | O(1) | O(1) |
| `min(s)` | Минимальный элемент s | O(n) | |
| `max(s)` | Максимальный элемент s | O(n) | |
| `s.index(x[, i[, j]])` | Индекс первого вхождения x в s (в диапазоне от i до j) | O(n) | O(n) |
| `s.count(x)` | Количество вхождений x в s | O(n) | O(n) |
| `s.append(x)` | Добавление элемента x в конец s| O(1) | O(1) |
| `s.sort()` | Сортировка | O(n log n) | O(n log n) |
| `s.extend(t)` | Добавление всех элементов t в в конец s| O(n + m) | O(n + m) |
| `s.pop()` | Вытащить последний элемент | O(1) | O(1) |
| `s.pop(i)` | Вытащить i-ый элемент | O(n - i) | O(n - i) |
| `s.insert(i, x)` | Вставить на i-ое место элемент x | O(n) | O(n) |
## Дек (collections.deque)
| Операция | Результат | В среднем | В худшем случае |
| --- | --- | --- | --- |
| `q.append(x)` | Добавить элемент x в конец дека | O(1) | O(1) |
| `q.appendleft(x)` | Добавить элемент x в начало дека| O(1) | O(1) |
| `q.pop()` | Вытащить последний элемент | O(1) | O(1) |
| `q.popleft()` | Вытащить первый элемент | O(1) | O(1) |
| `q.extend(t)` | Добавить все элементы t в конец дека | O(len(t)) | O(len(t)) |
| `q.extendleft(t)` | Добавить все элементы t в начало дека | O(len(t)) | O(len(t)) |
| `q.rotate(x)` | Сделать циклически сдвиг на x элементов | O(x) | O(x) |
| `q.remove(x)` | Удалить элемент x | O(n) | O(n) |
## Множество (set)
<table>
<thead><tr>
<th>Операция</th>
<th>Результат</th>
<th>В среднем</th>
<th>В худшем случае</th>
</tr>
</thead>
<tbody>
<tr>
<td><code>x in s</code></td>
<td>Вхождение элемента x в s</td>
<td>O(1)</td>
<td>O(n)</td>
</tr>
<tr>
<td><code>s | t</code></td>
<td>Объединение множеств</td>
<td>O(len(s) + len(t))</td>
<td></td>
</tr>
<tr>
<td><code>s & t</code></td>
<td>Пересечение множеств</td>
<td>O(min(len(s), len(t)))</td>
<td>O(len(s) * len(t))</td>
</tr>
<tr>
<td><code>s1 & s2 & ... & sn</code></td>
<td>Пересечение нескольких множеств</td>
<td></td>
<td>(n - 1) * O(l), l = max(len(s1), len(s2), ..., len(sn))</td>
</tr>
<tr>
<td><code>s - t</code></td>
<td>Разность множеств</td>
<td>O(len(s))</td>
<td></td>
</tr>
<tr>
<td><code>s -= t</code></td>
<td>Разность множеств, вычисляется в s</td>
<td>O(len(t))</td>
<td></td>
</tr>
<tr>
<td><code>s ^ t</code></td>
<td>Симметрическая разность</td>
<td>O(len(s))</td>
<td>O(len(s) * len(t))</td>
</tr>
<tr>
<td><code>s ^= t</code></td>
<td>Симметрическая разность, вычисляется в s</td>
<td>O(len(t))</td>
<td>O(len(t) * len(s))</td>
</tr>
<tr>
<td><code>s.add(x)</code></td>
<td>Добавить элемент x в s</td>
<td>O(1)</td>
<td>O(n)</td>
</tr>
<tr>
<td><code>s.remove(x)</code></td>
<td>Удалить элемент x из s, кидает ошибку, если элемента нет</td>
<td>O(1)</td>
<td>O(n)</td>
</tr>
<tr>
<td><code>s.discard(x)</code></td>
<td>Удалить элемент x из s, если он есть</td>
<td>O(1)</td>
<td>O(n)</td>
</tr>
<tr>
<td><code>s.pop()</code></td>
<td>Вытащить какой-то элемент из s</td>
<td>O(1)</td>
<td>O(n)</td>
</tr>
<tr>
<td><code>s.clear()</code></td>
<td>Удалить все элементы из s</td>
<td>O(n)</td>
<td>O(n)</td>
</tr>
</tbody>
</table>
## Словарь (dict)
| Операция | Результат | В среднем | В худшем случае |
| --- | --- | --- | --- |
| `d[key]` | Взять элемент по ключу key | O(1) | O(n) |
| `d.get(key, default)` | Взять элемент по ключу key. Если его нет, взять default | O(1) | O(n) |
| `d[key] = value` | Присвоить элемент по ключу key | O(1) | O(n) |
| `del d[key]` | Удалить элемент по ключу key | O(1) | O(n) |
| `for key, value in d.items()` | Итерация | O(n) | O(n) |
| `key in d` | Проверка существования ключа key | O(n) | O(n) |
| github_jupyter |
```
!pip install tensorflow-datasets
import numpy as np
import tensorflow as tf
import tensorflow_datasets as tfds
from tensorflow import keras
tfds.list_builders()
# get information on data
builder = tfds.builder('cats_vs_dogs')
builder.info
!kaggle datasets list
!kaggle datasets download -d alessiocorrado99/animals10
import sys
from matplotlib import pyplot
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.models import Sequential, Model, load_model
from tensorflow.keras.layers import Dense, Flatten, Dropout, Activation, Conv2D, MaxPooling2D, BatchNormalization
from tensorflow.keras.optimizers import SGD
DATADIR = "large_data/"
CATEGORIES = ['furry', 'notfurry']
SIZE = 200
RATIO = 0.2
BATCH_SIZE = 32
EPOCHS = 20
print('Constants initialized.')
import os
from matplotlib import pyplot as plt
def display_examples(folder):
"""
Display 25 images from the images array with its corresponding labels
"""
path = os.path.join('large_dataset/dataset', folder)
fig = plt.figure(figsize=(10,10))
fig.suptitle("Some examples of images of the dataset", fontsize=16)
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
images = os.listdir(path)
idx = np.random.randint(0, len(images))
arr = cv2.imread(os.path.join(path, images[idx]))
arr = cv2.cvtColor(arr, cv2.COLOR_BGR2RGB)
plt.imshow(arr, cmap=plt.cm.binary)
plt.xlabel(folder)
plt.show()
display_examples('furry')
from zipfile import ZipFile
# Create a ZipFile Object and load dataset.zip
with ZipFile('large_dataset.zip', 'r') as zipObj:
# Extract all the contents of zip file in dataset directory
zipObj.extractall('large_dataset')
print('Dataset extracted.')
from os import makedirs, listdir, path # create directories
from shutil import copyfile # move/swap files
from random import seed, random # generate random dispersion
def create_dir(dataset_path):
# organize dataset into a useful structure
# create directories
subdirs = ['train', 'test']
for subdir in subdirs:
# create label subdirectories
for label in CATEGORIES:
new = path.join(dataset_path, subdir, label)
makedirs(new, exist_ok=True)
# seed random number generator
seed(1)
# copy training dataset images into subdirectories
for dir in CATEGORIES:
head = path.join(dataset_path, dir)
for file in listdir(head):
src = path.join(head, file)
dst = subdirs[0]
if random() < RATIO:
dst = subdirs[1]
check = file.rsplit("_", 1)[1]
if check.startswith(dir):
dst = path.join(dataset_path, dst, dir, file)
copyfile(src, dst)
print('Directories are successfully created! Ready for modelling.')
create_dir('large_dataset/dataset')
# !pip install tensorflow
import os
from tensorflow.keras.preprocessing.image import ImageDataGenerator
path = 'large_dataset/dataset'
# create data generators
train_datagen = ImageDataGenerator(rotation_range=15,
rescale=1./255,
shear_range=0.1,
zoom_range=0.2,
horizontal_flip=True,
width_shift_range=0.1,
height_shift_range=0.1)
# keep test images unmodified
test_datagen = ImageDataGenerator(rescale=1.0/255.0)
# prepare iterators
train_it = train_datagen.flow_from_directory(os.path.join(path, 'train'),
class_mode='binary', batch_size=BATCH_SIZE, target_size=(SIZE, SIZE))
test_it = test_datagen.flow_from_directory(os.path.join(path, 'test'),
class_mode='binary', batch_size=BATCH_SIZE, target_size=(SIZE, SIZE))
print('Initiated data generators.')
model = Sequential()
model.add(Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same', input_shape=(200, 200, 3)))
model.add(BatchNormalization())
model.add(MaxPooling2D((2, 2)))
model.add(Dropout(0.25))
model.add(Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
model.add(BatchNormalization())
model.add(MaxPooling2D((2, 2)))
model.add(Dropout(0.25))
# model.add(Conv2D(128, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
# model.add(BatchNormalization())
# model.add(MaxPooling2D((2, 2)))
# model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(256, activation='relu', kernel_initializer='he_uniform'))
model.add(Dropout(0.5))
model.add(Dense(1, activation='sigmoid'))
# compile model
# opt = SGD(lr=0.001, momentum=0.9)
model.compile(optimizer='rmsprop', loss='binary_crossentropy', metrics=['accuracy'])
model.summary()
from keras.callbacks import EarlyStopping, ReduceLROnPlateau, ModelCheckpoint
earlystop = EarlyStopping(patience = 5)
learning_rate_reduction = ReduceLROnPlateau(monitor = 'val_accuracy', patience = 2, verbose = 1, factor = 0.5, min_lr = 0.00001)
mc = ModelCheckpoint('furspect_best.h5', monitor='val_accuracy', mode='max', verbose=1)
callbacks = [earlystop,learning_rate_reduction, mc]
print('Callbacks are setup.')
# fit model
history = model.fit(train_it, steps_per_epoch=len(train_it),
validation_data=test_it, validation_steps=len(test_it), epochs=EPOCHS, verbose=1, callbacks=[callbacks], shuffle=True)
# evaluate model
_, acc = model.evaluate(test_it, steps=len(test_it), verbose=1)
print('> %.3f' % (acc * 100.0))
def summarize_diagnostics(history):
# plot loss
pyplot.subplot(211)
pyplot.title('Cross Entropy Loss')
pyplot.plot(history.history['loss'], color='blue', label='train')
pyplot.plot(history.history['val_loss'], color='orange', label='test')
# plot accuracy
pyplot.subplot(212)
pyplot.title('Classification Accuracy')
pyplot.plot(history.history['accuracy'], color='blue', label='train')
pyplot.plot(history.history['val_accuracy'], color='orange', label='test')
# save plot to file
filename = sys.argv[0].split('/')[-1]
pyplot.show()
pyplot.savefig(filename + '_plot.png')
pyplot.close()
summarize_diagnostics(history)
# !pip install matplotlib
!pip install scipy
!apt-get install python-scipy
import sys
from matplotlib import pyplot
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.models import Sequential, Model, load_model
from tensorflow.keras.layers import Dense, Flatten, Dropout, Activation, Conv2D, MaxPooling2D, BatchNormalization
from tensorflow.keras.optimizers import SGD
def create_model():
# define cnn model (threeblock, dropout 25/50, batchnormalization, dense=512, rmsprop)
model = Sequential()
model.add(Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same', input_shape=(200, 200, 3)))
model.add(BatchNormalization())
model.add(MaxPooling2D((2, 2)))
model.add(Dropout(0.25))
model.add(Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
model.add(BatchNormalization())
model.add(MaxPooling2D((2, 2)))
model.add(Dropout(0.25))
# model.add(Conv2D(128, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
# model.add(BatchNormalization())
# model.add(MaxPooling2D((2, 2)))
# model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(256, activation='relu', kernel_initializer='he_uniform'))
model.add(Dropout(0.5))
model.add(Dense(1, activation='sigmoid'))
# compile model
# opt = SGD(lr=0.001, momentum=0.9)
model.compile(optimizer='rmsprop', loss='binary_crossentropy', metrics=['accuracy'])
return model
def load_path(path):
model = load_model(path)
print(model.summary())
return model
def summarize_diagnostics(history):
# plot loss
pyplot.subplot(211)
pyplot.title('Cross Entropy Loss')
pyplot.plot(history.history['loss'], color='blue', label='train')
pyplot.plot(history.history['val_loss'], color='orange', label='test')
# plot accuracy
pyplot.subplot(212)
pyplot.title('Classification Accuracy')
pyplot.plot(history.history['accuracy'], color='blue', label='train')
pyplot.plot(history.history['val_accuracy'], color='orange', label='test')
# save plot to file
filename = sys.argv[0].split('/')[-1]
pyplot.show()
pyplot.savefig(filename + '_plot.png')
pyplot.close()
def callbacks():
from keras.callbacks import EarlyStopping, ReduceLROnPlateau
earlystop = EarlyStopping(patience = 5)
learning_rate_reduction = ReduceLROnPlateau(monitor = 'val_accuracy',patience = 2,verbose = 1,factor = 0.5, min_lr = 0.00001)
mc = ModelCheckpoint('furspect_best.h5', monitor='val_accuracy', mode='max', verbose=1)
callbacks = [earlystop,learning_rate_reduction, mc]
print('Callbacks are setup.')
return callbacks
def run_harness():
# model = load_path('furspect_model2_1b.h5')
model = create_model()
cb = callbacks()
print('Loaded model.')
# fit model
history = model.fit(train_it, steps_per_epoch=len(train_it),
validation_data=test_it, validation_steps=len(test_it), epochs=EPOCHS, verbose=1, callbacks=[cb], shuffle=True)
# evaluate model
_, acc = model.evaluate(test_it, steps=len(test_it), verbose=1)
print('> %.3f' % (acc * 100.0))
summarize_diagnostics(history)
run_harness()
```
```
from tensorflow.keras.models import load_model
model = load_model('furspect_best_2.h5')
# For a more creative and expressive way – you can draw a diagram of the architecture (hint – take a look at the keras.utils.vis_utils function).
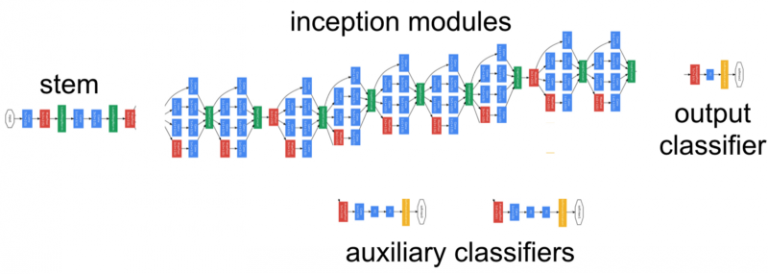
from matplotlib import pyplot as plt
top_layer = model.layers[0]
plt.imshow(top_layer.get_weights()[0][:, :, :, 0].squeeze(), cmap='gray')
# attempting to visualize filters (separated by R, G, B)
from matplotlib import pyplot as plt
# retrieve weights from first convolutional hidden layer
filters, biases = model.layers[0].get_weights()
# normalize filter values to 0-1
f_min, f_max = filters.min(), filters.max()
filters = (filters - f_min) / (f_max - f_min)
#plot filters
n_filters, ix = 6, 1
for i in range(n_filters):
f = filters[:, :, :, i]
for j in range(3):
ax = plt.subplot(n_filters, 3, ix)
ax.set_xticks([])
ax.set_yticks([])
plt.imshow(f[:, :, j], cmap='gray')
ix += 1
plt.show()
model.layers[0].get_weights()
# attempting to visualize feature maps
# idea: understand what features of an input are detected/preserved
# in these feature maps.
# generally, feature maps close to input -> small/fine-grained detail
# generally, feature maps close to output -> large general features
# output featuremap after first convolutional layer (index 1)
from tensorflow.keras.models import Model
model = Model(inputs=model.inputs, outputs=model.layers[0].output)
from PIL import Image
import numpy as np
from skimage import transform
from tensorflow.keras.models import load_model
from matplotlib import pyplot as plt
import cv2
import os
def load_image():
path = 'large_dataset/dataset/'
folder = ['furry/', 'notfurry/']
chosen = folder[np.random.randint(0, 2)]
# chosen = folder[0]
dir = os.path.join(path, chosen)
images = os.listdir(dir)
# load all images into a list
index = np.random.randint(0, len(images))
file = images[index]
np_image = Image.open(os.path.join(dir, file))
np_image = np.array(np_image).astype('float32')/255
np_image = transform.resize(np_image, (200, 200, 3))
np_image = np.expand_dims(np_image, axis=0)
# final_images.append(np_image)
return np_image
image = load_image()
feature_maps = model.predict(image)
ix = 1
for _ in range(8):
for _ in range(4):
# specify subplot and turn of axis
ax = plt.subplot(8, 4, ix)
ax.set_xticks([])
ax.set_yticks([])
# plot filter channel in grayscale
plt.imshow(feature_maps[0, :, :, ix-1], cmap='gray')
ix += 1
# show the figure
plt.show()
# plot all 32 maps
# redefine model to output right after the first hidden layer
from tensorflow.keras.models import load_model
model = load_model('furspect_best_2.h5')
ixs = [0, 4]
outputs = [model.layers[i].output for i in ixs]
model = Model(inputs=model.inputs, outputs=outputs)
from PIL import Image
import numpy as np
from skimage import transform
from tensorflow.keras.models import load_model
from matplotlib import pyplot as plt
import cv2
import os
def load_image():
path = 'large_dataset/dataset/'
folder = ['furry/', 'notfurry/']
chosen = folder[np.random.randint(0, 2)]
# chosen = folder[0]
dir = os.path.join(path, chosen)
images = os.listdir(dir)
# load all images into a list
index = np.random.randint(0, len(images))
file = images[index]
np_image = Image.open(os.path.join(dir, file))
np_image = np.array(np_image).astype('float32')/255
np_image = transform.resize(np_image, (200, 200, 3))
np_image = np.expand_dims(np_image, axis=0)
# final_images.append(np_image)
return np_image
image = load_image()
feature_maps = model.predict(image)
square = 8
for fmap in feature_maps:
# plot all 64 maps in an 8x8 squares
num = (fmap[0, :, :, :].shape)[2]
if num == 32:
col = 8
row = 4
elif num == 64:
col = 8
row = 8
print(num)
ix = 1
for _ in range(col):
for _ in range(row):
# specify subplot and turn of axis
ax = plt.subplot(col, row, ix)
ax.set_xticks([])
ax.set_yticks([])
# plot filter channel in grayscale
plt.imshow(fmap[0, :, :, ix-1], cmap='gray')
ix += 1
# show the figure
plt.show()
'''
We can see that the feature maps closer
to the input of the model capture a lot of
fine detail in the image and that as we progress
deeper into the model, the feature maps show less and less detail.
This pattern was to be expected,
as the model abstracts the features from
the image into more general concepts that can be used to make a
classification. Although it is not clear from the final image
that the model saw a bird, we generally lose the ability to
interpret these deeper feature maps.
'''
# Importance of visualizing a CNN Model
'''
It turned out that in the researchers’ dataset,
photos of camouflaged tanks had been taken on cloudy days,
while photos of plain forest had been taken on sunny days.
The neural network had learned to distinguish cloudy days from
sunny days, instead of distinguishing camouflaged tanks from an
empty forest.
'''
# redefine model to output right after the first hidden layer
from tensorflow.keras.models import load_model
model = load_model('furspect_best_2.h5')
model.summary()
!pip install pydot
!pip install graphviz
from tensorflow.keras.utils import plot_model
plot_model(model, to_file='/tmp/furspect_model_2_plot.png', show_shapes=True)
# analyzing activation maps
# understand what sort of input patterns activate particulra filter
!pip install keras-vis
!pip install tensorflow
from vis.visualization import visualize_activation
from vis.utils import utils
from keras import activations
from matplotlib import pyplot as plt
%matplotlib inline
plt.rcParams['figure.figsize'] = (18, 6)
# Utility to search for layer index by name.
# Alternatively we can specify this as -1 since it corresponds to the last layer.
layer_idx = utils.find_layer_idx(model, 'preds')
# Swap softmax with linear
model.layers[layer_idx].activation = activations.linear
model = utils.apply_modifications(model)
# This is the output node we want to maximize.
filter_idx = 0
img = visualize_activation(model, layer_idx, filter_indices=filter_idx)
plt.imshow(img[..., 0])
# summarize filter shapes
for layer in model.layers:
# check for convolutional layer
if 'conv' not in layer.name:
continue
# get filter weights
filters, biases = layer.get_weights()
print(layer.name, filters.shape)
from matplotlib import pyplot
# retrieve weights from the second hidden layer
filters, biases = model.layers[1].get_weights()
# normalize filter values to 0-1 so we can visualize them
f_min, f_max = filters.min(), filters.max()
filters = (filters - f_min) / (f_max - f_min)
# plot first few filters
n_filters, ix = 6, 1
for i in range(n_filters):
# get the filter
f = filters[:, :, :, i]
# plot each channel separately
for j in range(3):
# specify subplot and turn of axis
ax = pyplot.subplot(n_filters, 3, ix)
ax.set_xticks([])
ax.set_yticks([])
# plot filter channel in grayscale
pyplot.imshow(f[:, :, j], cmap='gray')
ix += 1
# show the figure
pyplot.show()
# summarize feature map shapes
for i in range(len(model.layers)):
layer = model.layers[i]
# check for convolutional layer
if 'conv' not in layer.name:
continue
# summarize output shape
print(i, layer.name, layer.output.shape)
# plot feature map of first conv layer for given image
from keras.preprocessing.image import load_img
from keras.preprocessing.image import img_to_array
from keras.models import Model
from matplotlib import pyplot
from numpy import expand_dims
# redefine model to output right after the first hidden layer (warning: will edit model)
model = Model(inputs=model.inputs, outputs=model.layers[1].output)
model.summary()
# load the image with the required shape
img = load_img('bird.jpg', target_size=(224, 224))
# convert the image to an array
img = img_to_array(img)
# expand dimensions so that it represents a single 'sample'
img = expand_dims(img, axis=0)
# prepare the image (e.g. scale pixel values for the vgg)
#img = preprocess_input(img) revise for model
# get feature map for first hidden layer
feature_maps = model.predict(img)
# plot all 64 maps in an 8x8 squares
square = 8
ix = 1
for _ in range(square):
for _ in range(square):
# specify subplot and turn of axis
ax = pyplot.subplot(square, square, ix)
ax.set_xticks([])
ax.set_yticks([])
# plot filter channel in grayscale
pyplot.imshow(feature_maps[0, :, :, ix-1], cmap='gray')
ix += 1
# show the figure
pyplot.show()
# visualize feature maps output from each block in the vgg model
from keras.preprocessing.image import load_img
from keras.preprocessing.image import img_to_array
from keras.models import Model
from matplotlib import pyplot
from numpy import expand_dims
ixs = [2, 5, 9, 13, 17]
outputs = [model.layers[i].output for i in ixs]
model = Model(inputs=model.inputs, outputs=outputs)
# load the image with the required shape
img = load_img('bird.jpg', target_size=(224, 224))
# convert the image to an array
img = img_to_array(img)
# expand dimensions so that it represents a single 'sample'
img = expand_dims(img, axis=0)
# prepare the image (e.g. scale pixel values for the vgg)
# img = preprocess_input(img) revise for model
# get feature map for first hidden layer
feature_maps = model.predict(img)
# plot the output from each block
square = 8
for fmap in feature_maps:
# plot all 64 maps in an 8x8 squares
ix = 1
for _ in range(square):
for _ in range(square):
# specify subplot and turn of axis
ax = pyplot.subplot(square, square, ix)
ax.set_xticks([])
ax.set_yticks([])
# plot filter channel in grayscale
pyplot.imshow(fmap[0, :, :, ix-1], cmap='gray')
ix += 1
# show the figure
pyplot.show()
for output_idx in np.arange(10):
# Lets turn off verbose output this time to avoid clutter and just see the output.
img = visualize_activation(model, layer_idx, filter_indices=output_idx, input_range=(0., 1.))
plt.figure()
plt.title('Networks perception of {}'.format(output_idx))
plt.imshow(img[..., 0])
def iter_occlusion(image, size=8):
# taken from https://www.kaggle.com/blargl/simple-occlusion-and-saliency-maps
occlusion = np.full((size * 5, size * 5, 1), [0.5], np.float32)
occlusion_center = np.full((size, size, 1), [0.5], np.float32)
occlusion_padding = size * 2
# print('padding...')
image_padded = np.pad(image, ( \
(occlusion_padding, occlusion_padding), (occlusion_padding, occlusion_padding), (0, 0) \
), 'constant', constant_values = 0.0)
for y in range(occlusion_padding, image.shape[0] + occlusion_padding, size):
for x in range(occlusion_padding, image.shape[1] + occlusion_padding, size):
tmp = image_padded.copy()
tmp[y - occlusion_padding:y + occlusion_center.shape[0] + occlusion_padding, \
x - occlusion_padding:x + occlusion_center.shape[1] + occlusion_padding] \
= occlusion
tmp[y:y + occlusion_center.shape[0], x:x + occlusion_center.shape[1]] = occlusion_center
yield x - occlusion_padding, y - occlusion_padding, \
tmp[occlusion_padding:tmp.shape[0] - occlusion_padding, occlusion_padding:tmp.shape[1] - occlusion_padding]
i = 23 # for example
data = val_x[i]
correct_class = np.argmax(val_y[i])
# input tensor for model.predict
inp = data.reshape(1, 28, 28, 1)
# image data for matplotlib's imshow
img = data.reshape(28, 28)
# occlusion
img_size = img.shape[0]
occlusion_size = 4
print('occluding...')
heatmap = np.zeros((img_size, img_size), np.float32)
class_pixels = np.zeros((img_size, img_size), np.int16)
from collections import defaultdict
counters = defaultdict(int)
for n, (x, y, img_float) in enumerate(iter_occlusion(data, size=occlusion_size)):
X = img_float.reshape(1, 28, 28, 1)
out = model.predict(X)
#print('#{}: {} @ {} (correct class: {})'.format(n, np.argmax(out), np.amax(out), out[0][correct_class]))
#print('x {} - {} | y {} - {}'.format(x, x + occlusion_size, y, y + occlusion_size))
heatmap[y:y + occlusion_size, x:x + occlusion_size] = out[0][correct_class]
class_pixels[y:y + occlusion_size, x:x + occlusion_size] = np.argmax(out)
counters[np.argmax(out)] += 1
class_idx = 0
indices = np.where(val_y[:, class_idx] == 1.)[0]
# pick some random input from here.
idx = indices[0]
# Lets sanity check the picked image.
from matplotlib import pyplot as plt
%matplotlib inline
plt.rcParams['figure.figsize'] = (18, 6)
plt.imshow(val_x[idx][..., 0])
from vis.visualization import visualize_saliency
from vis.utils import utils
from keras import activations
# Utility to search for layer index by name.
# Alternatively we can specify this as -1 since it corresponds to the last layer.
layer_idx = utils.find_layer_idx(model, 'preds')
# Swap softmax with linear
model.layers[layer_idx].activation = activations.linear
model = utils.apply_modifications(model)
grads = visualize_saliency(model, layer_idx, filter_indices=class_idx, seed_input=val_x[idx])
# Plot with 'jet' colormap to visualize as a heatmap.
plt.imshow(grads, cmap='jet')
# This corresponds to the Dense linear layer.
for class_idx in np.arange(10):
indices = np.where(val_y[:, class_idx] == 1.)[0]
idx = indices[0]
f, ax = plt.subplots(1, 4)
ax[0].imshow(val_x[idx][..., 0])
for i, modifier in enumerate([None, 'guided', 'relu']):
grads = visualize_saliency(model, layer_idx, filter_indices=class_idx,
seed_input=val_x[idx], backprop_modifier=modifier)
if modifier is None:
modifier = 'vanilla'
ax[i+1].set_title(modifier)
ax[i+1].imshow(grads, cmap='jet')
from vis.visualization import visualize_cam
# This corresponds to the Dense linear layer.
for class_idx in np.arange(10):
indices = np.where(val_y[:, class_idx] == 1.)[0]
idx = indices[0]
f, ax = plt.subplots(1, 4)
ax[0].imshow(val_x[idx][..., 0])
for i, modifier in enumerate([None, 'guided', 'relu']):
grads = visualize_cam(model, layer_idx, filter_indices=class_idx,
seed_input=val_x[idx], backprop_modifier=modifier)
if modifier is None:
modifier = 'vanilla'
ax[i+1].set_title(modifier)
ax[i+1].imshow(grads, cmap='jet')
def print_mislabeled_images(class_names, test_images, test_labels, pred_labels):
"""
Print 25 examples of mislabeled images by the classifier, e.g when test_labels != pred_labels
"""
BOO = (test_labels == pred_labels)
mislabeled_indices = np.where(BOO == 0)
mislabeled_images = test_images[mislabeled_indices]
mislabeled_labels = pred_labels[mislabeled_indices]
title = "Some examples of mislabeled images by the classifier:"
display_examples(class_names, mislabeled_images, mislabeled_labels)
print_mislabeled_images(class_names, test_images, test_labels, pred_labels)
predictions = model.predict(test_images) # Vector of probabilities
pred_labels = np.argmax(predictions, axis = 1) # We take the highest probability
display_random_image(class_names, test_images, pred_labels)
def plot_accuracy_loss(history):
"""
Plot the accuracy and the loss during the training of the nn.
"""
fig = plt.figure(figsize=(10,5))
# Plot accuracy
plt.subplot(221)
plt.plot(history.history['acc'],'bo--', label = "acc")
plt.plot(history.history['val_acc'], 'ro--', label = "val_acc")
plt.title("train_acc vs val_acc")
plt.ylabel("accuracy")
plt.xlabel("epochs")
plt.legend()
# Plot loss function
plt.subplot(222)
plt.plot(history.history['loss'],'bo--', label = "loss")
plt.plot(history.history['val_loss'], 'ro--', label = "val_loss")
plt.title("train_loss vs val_loss")
plt.ylabel("loss")
plt.xlabel("epochs")
plt.legend()
plt.show()
plot_accuracy_loss(history)
CM = confusion_matrix(test_labels, pred_labels)
ax = plt.axes()
sn.heatmap(CM, annot=True,
annot_kws={"size": 10},
xticklabels=class_names,
yticklabels=class_names, ax = ax)
ax.set_title('Confusion matrix')
plt.show()
### Figures
plt.subplots(figsize=(10,10))
for i, class_name in enumerate(class_names):
plt.scatter(C1[train_labels == i][:1000], C2[train_labels == i][:1000], label = class_name, alpha=0.4)
plt.legend()
plt.title("PCA Projection")
plt.show()
# !pip install scikit-image
# !pip install opencv-python
from PIL import Image
import numpy as np
from skimage import transform
from tensorflow.keras.models import load_model
from matplotlib import pyplot as plt
import cv2
def load_image():
path = 'large_dataset/dataset/'
folder = ['furry/', 'notfurry/']
chosen = folder[np.random.randint(0, 2)]
# chosen = folder[0]
dir = os.path.join(path, chosen)
images = os.listdir(dir)
# load all images into a list
index = np.random.randint(0, len(images))
file = images[index]
plt.figure()
arr = cv2.imread(os.path.join(dir, file))
arr = cv2.cvtColor(arr, cv2.COLOR_BGR2RGB)
plt.imshow(arr, cmap=plt.cm.binary)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.title('Image #{} : '.format(index) + file.rsplit("_", 1)[1])
plt.show()
np_image = Image.open(os.path.join(dir, file))
np_image = np.array(np_image).astype('float32')/255
np_image = transform.resize(np_image, (200, 200, 3))
np_image = np.expand_dims(np_image, axis=0)
# final_images.append(np_image)
return np_image
def predict_image(model):
image = load_image()
model = load_model(model)
res = model.predict(image, batch_size=10)
final = 0
consensus = "It's a furry!."
if res > 0.5:
final = 1
if final == 1:
consensus = "It's not a furry."
print('Result: {0}'.format(res))
print('Consensus: {0}'.format(consensus))
print(res)
return res
predict_image('furspect_best_2.h5')
from PIL import Image
import numpy as np
from skimage import transform
def load(count):
path = 'large_dataset/dataset'
folder = ['furry/', 'notfurry/']
# chosen = folder[np.random.randint(0, 2)]
chosen = folder[0]
dir = os.path.join(path, chosen)
images = os.listdir(dir)
# load all images into a list
final_images = []
c = 0
for image in images:
if c >= count:
break
file = image
# ignore webm
if 'webm' in file:
continue
np_image = Image.open(os.path.join(dir, file))
np_image = np.array(np_image).astype('float32')/255
np_image = transform.resize(np_image, (200, 200, 3))
np_image = np.expand_dims(np_image, axis=0)
final_images.append(np_image)
c += 1
return np.vstack(final_images)
def predict(model, count):
images = load(count)
model = load_model(model)
res = model.predict(images, batch_size=10)
return res
pred_array = predict('furspect_best_2.h5', 100)
sum = 0
for i in pred_array:
if i[0] < 0.50:
sum += 1
print(sum / len(pred_array)) # prediction accuracy
```
| github_jupyter |
# Convolutional Neural Networks: Application
Welcome to Course 4's second assignment! In this notebook, you will:
- Implement helper functions that you will use when implementing a TensorFlow model
- Implement a fully functioning ConvNet using TensorFlow
**After this assignment you will be able to:**
- Build and train a ConvNet in TensorFlow for a classification problem
We assume here that you are already familiar with TensorFlow. If you are not, please refer the *TensorFlow Tutorial* of the third week of Course 2 ("*Improving deep neural networks*").
## 1.0 - TensorFlow model
In the previous assignment, you built helper functions using numpy to understand the mechanics behind convolutional neural networks. Most practical applications of deep learning today are built using programming frameworks, which have many built-in functions you can simply call.
As usual, we will start by loading in the packages.
```
import math
import numpy as np
import h5py
import matplotlib.pyplot as plt
import scipy
from PIL import Image
from scipy import ndimage
import tensorflow as tf
from tensorflow.python.framework import ops
from cnn_utils import *
%matplotlib inline
np.random.seed(1)
```
Run the next cell to load the "SIGNS" dataset you are going to use.
```
# Loading the data (signs)
X_train_orig, Y_train_orig, X_test_orig, Y_test_orig, classes = load_dataset()
```
As a reminder, the SIGNS dataset is a collection of 6 signs representing numbers from 0 to 5.
<img src="images/SIGNS.png" style="width:800px;height:300px;">
The next cell will show you an example of a labelled image in the dataset. Feel free to change the value of `index` below and re-run to see different examples.
```
# Example of a picture
index = 6
plt.imshow(X_train_orig[index])
print ("y = " + str(np.squeeze(Y_train_orig[:, index])))
```
In Course 2, you had built a fully-connected network for this dataset. But since this is an image dataset, it is more natural to apply a ConvNet to it.
To get started, let's examine the shapes of your data.
```
X_train = X_train_orig/255.
X_test = X_test_orig/255.
Y_train = convert_to_one_hot(Y_train_orig, 6).T
Y_test = convert_to_one_hot(Y_test_orig, 6).T
print ("number of training examples = " + str(X_train.shape[0]))
print ("number of test examples = " + str(X_test.shape[0]))
print ("X_train shape: " + str(X_train.shape))
print ("Y_train shape: " + str(Y_train.shape))
print ("X_test shape: " + str(X_test.shape))
print ("Y_test shape: " + str(Y_test.shape))
conv_layers = {}
```
### 1.1 - Create placeholders
TensorFlow requires that you create placeholders for the input data that will be fed into the model when running the session.
**Exercise**: Implement the function below to create placeholders for the input image X and the output Y. You should not define the number of training examples for the moment. To do so, you could use "None" as the batch size, it will give you the flexibility to choose it later. Hence X should be of dimension **[None, n_H0, n_W0, n_C0]** and Y should be of dimension **[None, n_y]**. [Hint](https://www.tensorflow.org/api_docs/python/tf/placeholder).
```
# GRADED FUNCTION: create_placeholders
def create_placeholders(n_H0, n_W0, n_C0, n_y):
"""
Creates the placeholders for the tensorflow session.
Arguments:
n_H0 -- scalar, height of an input image
n_W0 -- scalar, width of an input image
n_C0 -- scalar, number of channels of the input
n_y -- scalar, number of classes
Returns:
X -- placeholder for the data input, of shape [None, n_H0, n_W0, n_C0] and dtype "float"
Y -- placeholder for the input labels, of shape [None, n_y] and dtype "float"
"""
### START CODE HERE ### (≈2 lines)
X = tf.placeholder(tf.float32, (None, n_H0, n_W0, n_C0))
Y = tf.placeholder(tf.float32, (None, n_y))
### END CODE HERE ###
return X, Y
X, Y = create_placeholders(64, 64, 3, 6)
print ("X = " + str(X))
print ("Y = " + str(Y))
```
**Expected Output**
<table>
<tr>
<td>
X = Tensor("Placeholder:0", shape=(?, 64, 64, 3), dtype=float32)
</td>
</tr>
<tr>
<td>
Y = Tensor("Placeholder_1:0", shape=(?, 6), dtype=float32)
</td>
</tr>
</table>
### 1.2 - Initialize parameters
You will initialize weights/filters $W1$ and $W2$ using `tf.contrib.layers.xavier_initializer(seed = 0)`. You don't need to worry about bias variables as you will soon see that TensorFlow functions take care of the bias. Note also that you will only initialize the weights/filters for the conv2d functions. TensorFlow initializes the layers for the fully connected part automatically. We will talk more about that later in this assignment.
**Exercise:** Implement initialize_parameters(). The dimensions for each group of filters are provided below. Reminder - to initialize a parameter $W$ of shape [1,2,3,4] in Tensorflow, use:
```python
W = tf.get_variable("W", [1,2,3,4], initializer = ...)
```
[More Info](https://www.tensorflow.org/api_docs/python/tf/get_variable).
```
# GRADED FUNCTION: initialize_parameters
def initialize_parameters():
"""
Initializes weight parameters to build a neural network with tensorflow. The shapes are:
W1 : [4, 4, 3, 8]
W2 : [2, 2, 8, 16]
Returns:
parameters -- a dictionary of tensors containing W1, W2
"""
tf.set_random_seed(1) # so that your "random" numbers match ours
initi = tf.contrib.layers.xavier_initializer(seed = 0)
### START CODE HERE ### (approx. 2 lines of code)
W1 = tf.get_variable('W1', [4, 4, 3, 8], initializer=initi)
W2 = tf.get_variable('W2', [2, 2, 8, 16], initializer=initi)
#W1 = tf.contrib.layers.xavier_initializer(seed = 0)
#W2 = tf.contrib.layers.xavier_initializer(seed = 0)
### END CODE HERE ###
parameters = {"W1": W1,
"W2": W2}
return parameters
tf.reset_default_graph()
with tf.Session() as sess_test:
parameters = initialize_parameters()
init = tf.global_variables_initializer()
sess_test.run(init)
print("W1 = " + str(parameters["W1"].eval()[1,1,1]))
print("W2 = " + str(parameters["W2"].eval()[1,1,1]))
```
** Expected Output:**
<table>
<tr>
<td>
W1 =
</td>
<td>
[ 0.00131723 0.14176141 -0.04434952 0.09197326 0.14984085 -0.03514394 <br>
-0.06847463 0.05245192]
</td>
</tr>
<tr>
<td>
W2 =
</td>
<td>
[-0.08566415 0.17750949 0.11974221 0.16773748 -0.0830943 -0.08058 <br>
-0.00577033 -0.14643836 0.24162132 -0.05857408 -0.19055021 0.1345228 <br>
-0.22779644 -0.1601823 -0.16117483 -0.10286498]
</td>
</tr>
</table>
### 1.2 - Forward propagation
In TensorFlow, there are built-in functions that carry out the convolution steps for you.
- **tf.nn.conv2d(X,W1, strides = [1,s,s,1], padding = 'SAME'):** given an input $X$ and a group of filters $W1$, this function convolves $W1$'s filters on X. The third input ([1,f,f,1]) represents the strides for each dimension of the input (m, n_H_prev, n_W_prev, n_C_prev). You can read the full documentation [here](https://www.tensorflow.org/api_docs/python/tf/nn/conv2d)
- **tf.nn.max_pool(A, ksize = [1,f,f,1], strides = [1,s,s,1], padding = 'SAME'):** given an input A, this function uses a window of size (f, f) and strides of size (s, s) to carry out max pooling over each window. You can read the full documentation [here](https://www.tensorflow.org/api_docs/python/tf/nn/max_pool)
- **tf.nn.relu(Z1):** computes the elementwise ReLU of Z1 (which can be any shape). You can read the full documentation [here.](https://www.tensorflow.org/api_docs/python/tf/nn/relu)
- **tf.contrib.layers.flatten(P)**: given an input P, this function flattens each example into a 1D vector it while maintaining the batch-size. It returns a flattened tensor with shape [batch_size, k]. You can read the full documentation [here.](https://www.tensorflow.org/api_docs/python/tf/contrib/layers/flatten)
- **tf.contrib.layers.fully_connected(F, num_outputs):** given a the flattened input F, it returns the output computed using a fully connected layer. You can read the full documentation [here.](https://www.tensorflow.org/api_docs/python/tf/contrib/layers/fully_connected)
In the last function above (`tf.contrib.layers.fully_connected`), the fully connected layer automatically initializes weights in the graph and keeps on training them as you train the model. Hence, you did not need to initialize those weights when initializing the parameters.
**Exercise**:
Implement the `forward_propagation` function below to build the following model: `CONV2D -> RELU -> MAXPOOL -> CONV2D -> RELU -> MAXPOOL -> FLATTEN -> FULLYCONNECTED`. You should use the functions above.
In detail, we will use the following parameters for all the steps:
- Conv2D: stride 1, padding is "SAME"
- ReLU
- Max pool: Use an 8 by 8 filter size and an 8 by 8 stride, padding is "SAME"
- Conv2D: stride 1, padding is "SAME"
- ReLU
- Max pool: Use a 4 by 4 filter size and a 4 by 4 stride, padding is "SAME"
- Flatten the previous output.
- FULLYCONNECTED (FC) layer: Apply a fully connected layer without an non-linear activation function. Do not call the softmax here. This will result in 6 neurons in the output layer, which then get passed later to a softmax. In TensorFlow, the softmax and cost function are lumped together into a single function, which you'll call in a different function when computing the cost.
```
# GRADED FUNCTION: forward_propagation
def forward_propagation(X, parameters):
"""
Implements the forward propagation for the model:
CONV2D -> RELU -> MAXPOOL -> CONV2D -> RELU -> MAXPOOL -> FLATTEN -> FULLYCONNECTED
Arguments:
X -- input dataset placeholder, of shape (input size, number of examples)
parameters -- python dictionary containing your parameters "W1", "W2"
the shapes are given in initialize_parameters
Returns:
Z3 -- the output of the last LINEAR unit
"""
# Retrieve the parameters from the dictionary "parameters"
W1 = parameters['W1']
W2 = parameters['W2']
### START CODE HERE ###
# CONV2D: stride of 1, padding 'SAME'
Z1 = tf.nn.conv2d(X, W1, [1, 1, 1, 1], padding='SAME')
# RELU
A1 = tf.nn.relu(Z1)
# MAXPOOL: window 8x8, sride 8, padding 'SAME'
P1 = tf.nn.max_pool(A1, [1,8,8,1], [1,8,8,1], padding='SAME')
# CONV2D: filters W2, stride 1, padding 'SAME'
Z2 = tf.nn.conv2d(P1, W2, [1, 1, 1, 1], padding = 'SAME')
# RELU
A2 = tf.nn.relu(Z2)
# MAXPOOL: window 4x4, stride 4, padding 'SAME'
P2 = tf.nn.max_pool(A2, [1,4,4,1], [1,4,4,1], padding='SAME')
# FLATTEN
P2 = tf.contrib.layers.flatten(P2)
# FULLY-CONNECTED without non-linear activation function (not not call softmax).
# 6 neurons in output layer. Hint: one of the arguments should be "activation_fn=None"
Z3 = tf.contrib.layers.fully_connected(P2, 6, activation_fn=None)
### END CODE HERE ###
return Z3
tf.reset_default_graph()
with tf.Session() as sess:
np.random.seed(1)
X, Y = create_placeholders(64, 64, 3, 6)
parameters = initialize_parameters()
Z3 = forward_propagation(X, parameters)
init = tf.global_variables_initializer()
sess.run(init)
a = sess.run(Z3, {X: np.random.randn(2,64,64,3), Y: np.random.randn(2,6)})
print("Z3 = " + str(a))
```
**Expected Output**:
<table>
<td>
Z3 =
</td>
<td>
[[-0.44670227 -1.57208765 -1.53049231 -2.31013036 -1.29104376 0.46852064] <br>
[-0.17601591 -1.57972014 -1.4737016 -2.61672091 -1.00810647 0.5747785 ]]
</td>
</table>
### 1.3 - Compute cost
Implement the compute cost function below. You might find these two functions helpful:
- **tf.nn.softmax_cross_entropy_with_logits(logits = Z3, labels = Y):** computes the softmax entropy loss. This function both computes the softmax activation function as well as the resulting loss. You can check the full documentation [here.](https://www.tensorflow.org/api_docs/python/tf/nn/softmax_cross_entropy_with_logits)
- **tf.reduce_mean:** computes the mean of elements across dimensions of a tensor. Use this to sum the losses over all the examples to get the overall cost. You can check the full documentation [here.](https://www.tensorflow.org/api_docs/python/tf/reduce_mean)
** Exercise**: Compute the cost below using the function above.
# GRADED FUNCTION: compute_cost
def compute_cost(Z3, Y):
"""
Computes the cost
Arguments:
Z3 -- output of forward propagation (output of the last LINEAR unit), of shape (6, number of examples)
Y -- "true" labels vector placeholder, same shape as Z3
Returns:
cost - Tensor of the cost function
"""
### START CODE HERE ### (1 line of code)
cost = tf.nn.softmax_cross_entropy_with_logits(logits=Z3, labels=Y)
cost = tf.reduce_mean(cost)
### END CODE HERE ###
return cost
```
tf.reset_default_graph()
with tf.Session() as sess:
np.random.seed(1)
X, Y = create_placeholders(64, 64, 3, 6)
parameters = initialize_parameters()
Z3 = forward_propagation(X, parameters)
cost = compute_cost(Z3, Y)
init = tf.global_variables_initializer()
sess.run(init)
a = sess.run(cost, {X: np.random.randn(4,64,64,3), Y: np.random.randn(4,6)})
print("cost = " + str(a))
```
**Expected Output**:
<table>
<td>
cost =
</td>
<td>
2.91034
</td>
</table>
## 1.4 Model
Finally you will merge the helper functions you implemented above to build a model. You will train it on the SIGNS dataset.
You have implemented `random_mini_batches()` in the Optimization programming assignment of course 2. Remember that this function returns a list of mini-batches.
**Exercise**: Complete the function below.
The model below should:
- create placeholders
- initialize parameters
- forward propagate
- compute the cost
- create an optimizer
Finally you will create a session and run a for loop for num_epochs, get the mini-batches, and then for each mini-batch you will optimize the function. [Hint for initializing the variables](https://www.tensorflow.org/api_docs/python/tf/global_variables_initializer)
```
# GRADED FUNCTION: model
def model(X_train, Y_train, X_test, Y_test, learning_rate = 0.009,
num_epochs = 100, minibatch_size = 64, print_cost = True):
"""
Implements a three-layer ConvNet in Tensorflow:
CONV2D -> RELU -> MAXPOOL -> CONV2D -> RELU -> MAXPOOL -> FLATTEN -> FULLYCONNECTED
Arguments:
X_train -- training set, of shape (None, 64, 64, 3)
Y_train -- test set, of shape (None, n_y = 6)
X_test -- training set, of shape (None, 64, 64, 3)
Y_test -- test set, of shape (None, n_y = 6)
learning_rate -- learning rate of the optimization
num_epochs -- number of epochs of the optimization loop
minibatch_size -- size of a minibatch
print_cost -- True to print the cost every 100 epochs
Returns:
train_accuracy -- real number, accuracy on the train set (X_train)
test_accuracy -- real number, testing accuracy on the test set (X_test)
parameters -- parameters learnt by the model. They can then be used to predict.
"""
ops.reset_default_graph() # to be able to rerun the model without overwriting tf variables
tf.set_random_seed(1) # to keep results consistent (tensorflow seed)
seed = 3 # to keep results consistent (numpy seed)
(m, n_H0, n_W0, n_C0) = X_train.shape
n_y = Y_train.shape[1]
costs = [] # To keep track of the cost
# Create Placeholders of the correct shape
### START CODE HERE ### (1 line)
X = tf.placeholder(tf.float32, shape=[None, n_H0, n_W0, n_C0], name='X')
Y = tf.placeholder(tf.float32, shape = [None, n_y], name = 'Y')
### END CODE HERE ###
# Initialize parameters
### START CODE HERE ### (1 line)
parameters = initialize_parameters()
### END CODE HERE ###
# Forward propagation: Build the forward propagation in the tensorflow ,graph
### START CODE HERE ### (1 line)
Z3 = forward_propagation(X, parameters)
### END CODE HERE ###
# Cost function: Add cost function to tensorflow graph
### START CODE HERE ### (1 line)
cost = compute_cost(Z3, Y)
### END CODE HERE ###
# Backpropagation: Define the tensorflow optimizer. Use an AdamOptimizer that minimizes the cost.
### START CODE HERE ### (1 line)
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(loss=cost)
### END CODE HERE ###
# Initialize all the variables globally
init = tf.global_variables_initializer()
# Start the session to compute the tensorflow graph
with tf.Session() as sess:
# Run the initialization
sess.run(init)
# Do the training loop
for epoch in range(num_epochs):
minibatch_cost = 0.
num_minibatches = int(m / minibatch_size) # number of minibatches of size minibatch_size in the train set
seed = seed + 1
minibatches = random_mini_batches(X_train, Y_train, minibatch_size, seed)
for minibatch in minibatches:
# Select a minibatch
(minibatch_X, minibatch_Y) = minibatch
# IMPORTANT: The line that runs the graph on a minibatch.
# Run the session to execute the optimizer and the cost, the feedict should contain a minibatch for (X,Y).
### START CODE HERE ### (1 line)
_ , temp_cost = sess.run([optimizer, cost], feed_dict={X: minibatch_X, Y: minibatch_Y})
### END CODE HERE ###
minibatch_cost += temp_cost / num_minibatches
# Print the cost every epoch
if print_cost == True and epoch % 5 == 0:
print ("Cost after epoch %i: %f" % (epoch, minibatch_cost))
if print_cost == True and epoch % 1 == 0:
costs.append(minibatch_cost)
# plot the cost
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per tens)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
# Calculate the correct predictions
predict_op = tf.argmax(Z3, 1)
correct_prediction = tf.equal(predict_op, tf.argmax(Y, 1))
# Calculate accuracy on the test set
accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
print(accuracy)
train_accuracy = accuracy.eval({X: X_train, Y: Y_train})
test_accuracy = accuracy.eval({X: X_test, Y: Y_test})
print("Train Accuracy:", train_accuracy)
print("Test Accuracy:", test_accuracy)
return train_accuracy, test_accuracy, parameters
```
Run the following cell to train your model for 100 epochs. Check if your cost after epoch 0 and 5 matches our output. If not, stop the cell and go back to your code!
```
_, _, parameters = model(X_train, Y_train, X_test, Y_test)
```
**Expected output**: although it may not match perfectly, your expected output should be close to ours and your cost value should decrease.
<table>
<tr>
<td>
**Cost after epoch 0 =**
</td>
<td>
1.917929
</td>
</tr>
<tr>
<td>
**Cost after epoch 5 =**
</td>
<td>
1.506757
</td>
</tr>
<tr>
<td>
**Train Accuracy =**
</td>
<td>
0.940741
</td>
</tr>
<tr>
<td>
**Test Accuracy =**
</td>
<td>
0.783333
</td>
</tr>
</table>
Congratulations! You have finised the assignment and built a model that recognizes SIGN language with almost 80% accuracy on the test set. If you wish, feel free to play around with this dataset further. You can actually improve its accuracy by spending more time tuning the hyperparameters, or using regularization (as this model clearly has a high variance).
Once again, here's a thumbs up for your work!
```
fname = "images/thumbs_up.jpg"
image = np.array(ndimage.imread(fname, flatten=False))
my_image = scipy.misc.imresize(image, size=(64,64))
plt.imshow(my_image)
```
| github_jupyter |
<table>
<tr><td><img style="height: 150px;" src="images/geo_hydro1.jpg"></td>
<td bgcolor="#FFFFFF">
<p style="font-size: xx-large; font-weight: 900; line-height: 100%">AG Dynamics of the Earth</p>
<p style="font-size: large; color: rgba(0,0,0,0.5);">Juypter notebooks</p>
<p style="font-size: large; color: rgba(0,0,0,0.5);">Georg Kaufmann</p>
</td>
</tr>
</table>
# Angewandte Geophysik II: Kap 6: Magnetik
# Magnetfeldmodellierung
----
*Georg Kaufmann,
Geophysics Section,
Institute of Geological Sciences,
Freie Universität Berlin,
Germany*
```
import numpy as np
import matplotlib.pyplot as plt
import ipywidgets as widgets
# define profile
xmin = -500.
xmax = +500.
xstep = 101
x = np.linspace(xmin,xmax,xstep)
```
For the **magnetic induction** $\vec{B}$ [T], we define
$$
\vec{B} = \mu_0 \vec{H}
$$
with $\mu_0=4 \pi \times 10^{-7}$ Vs/A/m the **permeability of vacuum**,
and $\vec{H}$ [A/m] the **magnetic field strength**.
For the **magnetisation** $\vec{M}$ [A/m] we define
$$
\vec{M} = \chi \vec{H}
$$
with $\chi$ [-] the **susceptibility**.
## Sphere
<img src=figures/sketch_monopole.jpg style=width:10cm>
$$
\begin{array}{rcl}
B_z & = & \frac{\mu_0}{4\pi} \frac{M V}{z^3}
\frac{1}{r_1}
\left( 2 + \frac{3 x}{z \tan{\alpha}} - \frac{x^2}{z^2} \right) \\
B_x & = & \frac{\mu_0}{4\pi} \frac{M V}{z^3}
\frac{1}{r_1}
\left( \frac{1}{\tan{\alpha}} \frac{2 x^2}{z^2} + \frac{3x}{z} \right)
\end{array}
$$
```
def B_sphere(x,D=100.,R=20.,alpha=90.,M=0.04):
mu0 = 4.e-7*np.pi
r1 = np.sqrt((1. + x**2 / D**2)**5)
factor = mu0 / 4. / np.pi * M * 4. / 3. * np.pi * R**3 / D**3
# magnetic induction of sphere
Bx = factor * (((2.*x**2/D**2-1.)/np.tan(alpha*np.pi/180.) + 3.*x/D) / r1)
Bz = factor * ((2. + 3.*x/D/np.tan(alpha*np.pi/180.) - x**2 / D**2) / r1)
return Bx,Bz
def plot_sphere(f1=False,f2=False,f3=False,f4=False,f5=False):
D = [100,100,100,100,100]
R = [20,30,40,30,30]
alpha = [90,90,90,90,90]
M = [0.04,0.04,0.04,0.02,0.08]
fig,axs = plt.subplots(2,1,figsize=(12,8))
axs[0].set_xlim([-500,500])
axs[0].set_xticks([x for x in np.linspace(-400,400,9)])
axs[0].set_xlabel('Profile [m]')
axs[0].set_ylim([-1.5,2.5])
axs[0].set_yticks([y for y in np.linspace(-1.0,2.0,5)])
axs[0].set_ylabel('Bx,Bz [nT]')
axs[0].plot(x,1.e9*B_sphere(x)[0],linewidth=1.0,linestyle='-',color='black',label='B$_x$ - sphere')
axs[0].plot(x,1.e9*B_sphere(x)[1],linewidth=1.0,linestyle=':',color='black',label='B$_z$ - sphere')
if (f1):
axs[0].plot(x,1.e9*B_sphere(x,D=D[0],R=R[0],M=M[0],alpha=alpha[0])[0],linewidth=2.0,linestyle='-',color='black',
label='D='+str(D[0])+',R='+str(R[0])+',M='+str(M[0])+',alpha='+str(alpha[0]))
axs[0].plot(x,1.e9*B_sphere(x,D=D[0],R=R[0],M=M[0],alpha=alpha[0])[1],linewidth=2.0,linestyle=':',color='black')
if (f2):
axs[0].plot(x,1.e9*B_sphere(x,D=D[1],R=R[1],M=M[1],alpha=alpha[1])[0],linewidth=2.0,linestyle='-',color='red',
label='D='+str(D[1])+',R='+str(R[1])+',M='+str(M[1])+',alpha='+str(alpha[1]))
axs[0].plot(x,1.e9*B_sphere(x,D=D[1],R=R[1],M=M[1],alpha=alpha[1])[1],linewidth=2.0,linestyle=':',color='red')
if (f3):
axs[0].plot(x,1.e9*B_sphere(x,D=D[2],R=R[2],M=M[2],alpha=alpha[2])[0],linewidth=2.0,linestyle='-',color='orange',
label='D='+str(D[2])+',R='+str(R[2])+',M='+str(M[2])+',alpha='+str(alpha[2]))
axs[0].plot(x,1.e9*B_sphere(x,D=D[2],R=R[2],M=M[2],alpha=alpha[2])[1],linewidth=2.0,linestyle=':',color='orange')
if (f4):
axs[0].plot(x,1.e9*B_sphere(x,D=D[3],R=R[3],M=M[3],alpha=alpha[3])[0],linewidth=2.0,linestyle='-',color='green',
label='D='+str(D[3])+',R='+str(R[3])+',M='+str(M[3])+',alpha='+str(alpha[3]))
axs[0].plot(x,1.e9*B_sphere(x,D=D[3],R=R[3],M=M[3],alpha=alpha[3])[1],linewidth=2.0,linestyle=':',color='green')
if (f5):
axs[0].plot(x,1.e9*B_sphere(x,D=D[4],R=R[4],M=M[4],alpha=alpha[4])[0],linewidth=2.0,linestyle='-',color='blue',
label='D='+str(D[4])+',R='+str(R[4])+',M='+str(M[4])+',alpha='+str(alpha[4]))
axs[0].plot(x,1.e9*B_sphere(x,D=D[4],R=R[4],M=M[4],alpha=alpha[4])[1],linewidth=2.0,linestyle=':',color='blue')
axs[0].legend()
axs[1].set_xlim([-500,500])
axs[1].set_xticks([x for x in np.linspace(-400,400,9)])
#axs[1].set_xlabel('Profile [m]')
axs[1].set_ylim([250,0])
axs[1].set_yticks([y for y in np.linspace(0.,200.,5)])
axs[1].set_ylabel('Depth [m]')
angle = [theta for theta in np.linspace(0,2*np.pi,41)]
if (f1):
axs[1].plot(R[0]*np.cos(angle),D[0]+R[0]*np.sin(angle),linewidth=2.0,linestyle='-',color='black')
if (f2):
axs[1].plot(R[1]*np.cos(angle),D[1]+R[1]*np.sin(angle),linewidth=2.0,linestyle='-',color='red')
if (f3):
axs[1].plot(R[2]*np.cos(angle),D[2]+R[2]*np.sin(angle),linewidth=2.0,linestyle='-',color='orange')
if (f4):
axs[1].plot(R[3]*np.cos(angle),D[3]+R[3]*np.sin(angle),linewidth=2.0,linestyle='-',color='green')
if (f5):
axs[1].plot(R[4]*np.cos(angle),D[4]+R[4]*np.sin(angle),linewidth=2.0,linestyle='-',color='blue')
plot_sphere(f2=True)
# call interactive module
w = dict(
f1=widgets.Checkbox(value=True,description='eins',continuous_update=False,disabled=False),
#a1=widgets.FloatSlider(min=0.,max=2.,step=0.1,value=1.0),
f2=widgets.Checkbox(value=False,description='zwei',continuous_update=False,disabled=False),
f3=widgets.Checkbox(value=False,description='drei',continuous_update=False,disabled=False),
f4=widgets.Checkbox(value=False,description='vier',continuous_update=False,disabled=False),
f5=widgets.Checkbox(value=False,description='fuenf',continuous_update=False,disabled=False))
output = widgets.interactive_output(plot_sphere, w)
box = widgets.HBox([widgets.VBox([*w.values()]), output])
display(box)
```
... done
| github_jupyter |
# Self-Driving Car Engineer Nanodegree
## Project: **Finding Lane Lines on the Road**
***
In this project, you will use the tools you learned about in the lesson to identify lane lines on the road. You can develop your pipeline on a series of individual images, and later apply the result to a video stream (really just a series of images). Check out the video clip "raw-lines-example.mp4" (also contained in this repository) to see what the output should look like after using the helper functions below.
Once you have a result that looks roughly like "raw-lines-example.mp4", you'll need to get creative and try to average and/or extrapolate the line segments you've detected to map out the full extent of the lane lines. You can see an example of the result you're going for in the video "P1_example.mp4". Ultimately, you would like to draw just one line for the left side of the lane, and one for the right.
In addition to implementing code, there is a brief writeup to complete. The writeup should be completed in a separate file, which can be either a markdown file or a pdf document. There is a [write up template](https://github.com/udacity/CarND-LaneLines-P1/blob/master/writeup_template.md) that can be used to guide the writing process. Completing both the code in the Ipython notebook and the writeup template will cover all of the [rubric points](https://review.udacity.com/#!/rubrics/322/view) for this project.
---
Let's have a look at our first image called 'test_images/solidWhiteRight.jpg'. Run the 2 cells below (hit Shift-Enter or the "play" button above) to display the image.
**Note: If, at any point, you encounter frozen display windows or other confounding issues, you can always start again with a clean slate by going to the "Kernel" menu above and selecting "Restart & Clear Output".**
---
**The tools you have are color selection, region of interest selection, grayscaling, Gaussian smoothing, Canny Edge Detection and Hough Tranform line detection. You are also free to explore and try other techniques that were not presented in the lesson. Your goal is piece together a pipeline to detect the line segments in the image, then average/extrapolate them and draw them onto the image for display (as below). Once you have a working pipeline, try it out on the video stream below.**
---
<figure>
<img src="examples/line-segments-example.jpg" width="380" alt="Combined Image" />
<figcaption>
<p></p>
<p style="text-align: center;"> Your output should look something like this (above) after detecting line segments using the helper functions below </p>
</figcaption>
</figure>
<p></p>
<figure>
<img src="examples/laneLines_thirdPass.jpg" width="380" alt="Combined Image" />
<figcaption>
<p></p>
<p style="text-align: center;"> Your goal is to connect/average/extrapolate line segments to get output like this</p>
</figcaption>
</figure>
**Run the cell below to import some packages. If you get an `import error` for a package you've already installed, try changing your kernel (select the Kernel menu above --> Change Kernel). Still have problems? Try relaunching Jupyter Notebook from the terminal prompt. Also, consult the forums for more troubleshooting tips.**
## Import Packages
```
#importing some useful packages
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import numpy as np
import cv2
%matplotlib inline
```
## Read in an Image
```
#reading in an image
image = mpimg.imread('test_images/solidWhiteRight.jpg')
#printing out some stats and plotting
print('This image is:', type(image), 'with dimensions:', image.shape)
plt.imshow(image) # if you wanted to show a single color channel image called 'gray', for example, call as plt.imshow(gray, cmap='gray')
```
## Ideas for Lane Detection Pipeline
**Some OpenCV functions (beyond those introduced in the lesson) that might be useful for this project are:**
`cv2.inRange()` for color selection
`cv2.fillPoly()` for regions selection
`cv2.line()` to draw lines on an image given endpoints
`cv2.addWeighted()` to coadd / overlay two images
`cv2.cvtColor()` to grayscale or change color
`cv2.imwrite()` to output images to file
`cv2.bitwise_and()` to apply a mask to an image
**Check out the OpenCV documentation to learn about these and discover even more awesome functionality!**
## Helper Functions
Below are some helper functions to help get you started. They should look familiar from the lesson!
```
import math
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import PolynomialFeatures
def grayscale(img):
"""Applies the Grayscale transform
This will return an image with only one color channel
but NOTE: to see the returned image as grayscale
(assuming your grayscaled image is called 'gray')
you should call plt.imshow(gray, cmap='gray')"""
return cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# Or use BGR2GRAY if you read an image with cv2.imread()
# return cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
def canny(img, low_threshold, high_threshold):
"""Applies the Canny transform"""
return cv2.Canny(img, low_threshold, high_threshold)
def gaussian_blur(img, kernel_size):
"""Applies a Gaussian Noise kernel"""
return cv2.GaussianBlur(img, (kernel_size, kernel_size), 0)
def region_of_interest(img, vertices):
"""
Applies an image mask.
Only keeps the region of the image defined by the polygon
formed from `vertices`. The rest of the image is set to black.
`vertices` should be a numpy array of integer points.
"""
#defining a blank mask to start with
mask = np.zeros_like(img)
#defining a 3 channel or 1 channel color to fill the mask with depending on the input image
if len(img.shape) > 2:
channel_count = img.shape[2] # i.e. 3 or 4 depending on your image
ignore_mask_color = (255,) * channel_count
else:
ignore_mask_color = 255
#filling pixels inside the polygon defined by "vertices" with the fill color
cv2.fillPoly(mask, vertices, ignore_mask_color)
#returning the image only where mask pixels are nonzero
masked_image = cv2.bitwise_and(img, mask)
return masked_image
def linear_regression_find_points(lines,imp):
"""
Apply Linear Regression function to find the left line and right line model.
Then use the model to preditct top and bottom points of the line.
"""
points = lines[:,[0,1]]
points_y = points[:,1].reshape(-1,1)
#polynomial_features= PolynomialFeatures(degree=2)
#points_y = polynomial_features.fit_transform(points_y)
reg = LinearRegression().fit(points_y, points[:,0])
#imp_b = polynomial_features.fit_transform(np.array([[imp]]))
imp_b = np.array([[imp]])
bottom = (int(reg.predict(imp_b)[0]),imp)
imp_t = np.array([[320]])
#imp_t = polynomial_features.fit_transform(np.array([[320]]))
top = (int(reg.predict(imp_t)[0]),320)
return [bottom,top]
def draw_lines(img, lines, color=[255, 0, 0], thickness=10):
"""
NOTE: this is the function you might want to use as a starting point once you want to
average/extrapolate the line segments you detect to map out the full
extent of the lane (going from the result shown in raw-lines-example.mp4
to that shown in P1_example.mp4).
Think about things like separating line segments by their
slope ((y2-y1)/(x2-x1)) to decide which segments are part of the left
line vs. the right line. Then, you can average the position of each of
the lines and extrapolate to the top and bottom of the lane.
This function draws `lines` with `color` and `thickness`.
Lines are drawn on the image inplace (mutates the image).
If you want to make the lines semi-transparent, think about combining
this function with the weighted_img() function below
"""
imp = img.shape[0]
left = []
right = []
left_slope = []
right_slope = []
for line in lines:
for x1,y1,x2,y2 in line:
slope = (y2-y1)/(x2-x1)
if slope <= 0:
left.append([int((x1+x2)/2),int((y1+y2)/2),slope])
else:
right.append([int((x1+x2)/2),int((y1+y2)/2),slope])
left = [x for x in left if x[2]>-50]
right =[x for x in right if x[2]<50]
left = np.array(left)
right = np.array(right)
left_slope= right[:,2]
max_slope = np.max(left_slope)
min_slope = np.min(left_slope)
print(max_slope,min_slope)
right_bottom,right_top = linear_regression_find_points(right,imp)
left_bottom, left_top =linear_regression_find_points(left,imp)
cv2.line(img, left_bottom, left_top, color, thickness)
cv2.line(img, right_bottom, right_top, color, thickness)
"""
left_sorted_lines = left_lines[left_lines[:,1].argsort()]
left_bottom = tuple(left_sorted_lines[-1])
left_top = tuple(left_sorted_lines[0])
right_sorted_lines = right_lines[right_lines[:,1].argsort()]
right_bottom = tuple(right_sorted_lines[0])
right_top = tuple(right_sorted_lines[-1])
kl = (left_bottom[1]-left_top[1])/(left_bottom[0]-left_top[0])
#kl = np.median(np.sort((np.array(left_slope))))
bl = left_bottom[1]-kl*left_bottom[0]
kr = (right_bottom[1]-right_top[1])/(right_bottom[0]-right_top[0])
br = right_bottom[1]-kr*right_bottom[0]
vir_l_bottom = (int((imp-bl)/kl),imp)
vir_r_bottom = (int((imp-br)/kr),imp)
vir_l_top = (int((320-bl)/kl),320)
vir_r_top = (int((320-br)/kr),320)
cv2.line(img, vir_l_bottom, vir_l_top, color, thickness)
cv2.line(img, vir_r_bottom, vir_r_top, color, thickness)
"""
def hough_lines(img, rho, theta, threshold, min_line_len, max_line_gap):
"""
`img` should be the output of a Canny transform.
Returns an image with hough lines drawn.
"""
lines = cv2.HoughLinesP(img, rho, theta, threshold, np.array([]), minLineLength=min_line_len, maxLineGap=max_line_gap)
line_img = np.zeros((img.shape[0], img.shape[1], 3), dtype=np.uint8)
draw_lines(line_img, lines)
return line_img
# Python 3 has support for cool math symbols.
def weighted_img(img, initial_img, α=0.8, β=1., γ=0.):
"""
`img` is the output of the hough_lines(), An image with lines drawn on it.
Should be a blank image (all black) with lines drawn on it.
`initial_img` should be the image before any processing.
The result image is computed as follows:
initial_img * α + img * β + γ
NOTE: initial_img and img must be the same shape!
"""
return cv2.addWeighted(initial_img, α, img, β, γ)
```
## Test Images
Build your pipeline to work on the images in the directory "test_images"
**You should make sure your pipeline works well on these images before you try the videos.**
```
import os
os.listdir("test_images/")
```
## Build a Lane Finding Pipeline
Build the pipeline and run your solution on all test_images. Make copies into the `test_images_output` directory, and you can use the images in your writeup report.
Try tuning the various parameters, especially the low and high Canny thresholds as well as the Hough lines parameters.
```
# TODO: Build your pipeline that will draw lane lines on the test_images
# then save them to the test_images_output directory.
images_names = os.listdir("test_images/")
# readin images from images folder
for i in range(len(images_names)):
image = mpimg.imread('test_images/'+images_names[i])
# grayscale the image
gray = grayscale(image)
# Define a kernel size and apply Gaussian smoothing
kernel_size = 5
blur_gray = gaussian_blur(gray, kernel_size)
# Define our parameters for Canny and apply
low_threshold = 50
high_threshold = 150
edges = canny(blur_gray, low_threshold, high_threshold)
# This time we are defining a four sided polygon to mask
imshape = image.shape
vertices = np.array([[(0,imshape[0]),(450, 325), (490, 325), (imshape[1],imshape[0])]], dtype=np.int32)
# Only keep the interest region of the image
masked_edges = region_of_interest(edges, vertices)
# Define the Hough transform parameters
# Make a blank the same size as our image to draw on
rho = 1 # distance resolution in pixels of the Hough grid
theta = np.pi/180 # angular resolution in radians of the Hough grid
threshold = 1 # minimum number of votes (intersections in Hough grid cell)
min_line_length = 5 #minimum number of pixels making up a line
max_line_gap = 1 # maximum gap in pixels between connectable line segments
line_image = np.copy(image)*0 # creating a blank to draw lines on
# Run Hough on edge detected image
# Output "lines" is an array containing endpoints of detected line segments
lines_image = hough_lines(masked_edges, rho, theta, threshold, min_line_length, max_line_gap)
# Create a "color" binary image to combine with line image
color_edges = np.dstack((edges, edges, edges))
lines_edges = weighted_img(lines_image, image, 0.8, 1, 0)
plt.imsave("test_images_output/"+images_names[i], lines_edges)
os.listdir("test_images/")
```
## Test on Videos
You know what's cooler than drawing lanes over images? Drawing lanes over video!
We can test our solution on two provided videos:
`solidWhiteRight.mp4`
`solidYellowLeft.mp4`
**Note: if you get an import error when you run the next cell, try changing your kernel (select the Kernel menu above --> Change Kernel). Still have problems? Try relaunching Jupyter Notebook from the terminal prompt. Also, consult the forums for more troubleshooting tips.**
**If you get an error that looks like this:**
```
NeedDownloadError: Need ffmpeg exe.
You can download it by calling:
imageio.plugins.ffmpeg.download()
```
**Follow the instructions in the error message and check out [this forum post](https://discussions.udacity.com/t/project-error-of-test-on-videos/274082) for more troubleshooting tips across operating systems.**
```
# Import everything needed to edit/save/watch video clips
from moviepy.editor import VideoFileClip
from IPython.display import HTML
def process_image(image):
# NOTE: The output you return should be a color image (3 channel) for processing video below
# TODO: put your pipeline here,
# you should return the final output (image where lines are drawn on lanes)
#mage = mpimg.imread('test_images/'+images_names[i])
# grayscale the image
gray = grayscale(image)
# Define a kernel size and apply Gaussian smoothing
kernel_size = 5
blur_gray = gaussian_blur(gray, kernel_size)
# Define our parameters for Canny and apply
low_threshold = 50
high_threshold = 150
edges = canny(blur_gray, low_threshold, high_threshold)
# This time we are defining a four sided polygon to mask
imshape = image.shape
vertices = np.array([[(0,imshape[0]),(450, 320),(490, 320), (imshape[1],imshape[0])]], dtype=np.int32)
# Only keep the interest region of the image
masked_edges = region_of_interest(edges, vertices)
# Define the Hough transform parameters
# Make a blank the same size as our image to draw on
rho = 1 # distance resolution in pixels of the Hough grid
theta = np.pi/180 # angular resolution in radians of the Hough grid
threshold = 1 # minimum number of votes (intersections in Hough grid cell)
min_line_length = 5 #minimum number of pixels making up a line
max_line_gap = 1 # maximum gap in pixels between connectable line segments
line_image = np.copy(image)*0 # creating a blank to draw lines on
# Run Hough on edge detected image
# Output "lines" is an array containing endpoints of detected line segments
lines_image = hough_lines(masked_edges, rho, theta, threshold, min_line_length, max_line_gap)
# Create a "color" binary image to combine with line image
color_edges = np.dstack((edges, edges, edges))
lines_edges = weighted_img(lines_image, image, 0.8, 1, 0)
return lines_edges
```
Let's try the one with the solid white lane on the right first ...
```
white_output = 'test_videos_output/solidWhiteRight.mp4'
## To speed up the testing process you may want to try your pipeline on a shorter subclip of the video
## To do so add .subclip(start_second,end_second) to the end of the line below
## Where start_second and end_second are integer values representing the start and end of the subclip
## You may also uncomment the following line for a subclip of the first 5 seconds
##clip1 = VideoFileClip("test_videos/solidWhiteRight.mp4").subclip(0,5)
clip1 = VideoFileClip("test_videos/solidWhiteRight.mp4")
white_clip = clip1.fl_image(process_image) #NOTE: this function expects color images!!
%time white_clip.write_videofile(white_output, audio=False)
```
Play the video inline, or if you prefer find the video in your filesystem (should be in the same directory) and play it in your video player of choice.
```
HTML("""
<video width="960" height="540" controls>
<source src="{0}">
</video>
""".format(white_output))
```
## Improve the draw_lines() function
**At this point, if you were successful with making the pipeline and tuning parameters, you probably have the Hough line segments drawn onto the road, but what about identifying the full extent of the lane and marking it clearly as in the example video (P1_example.mp4)? Think about defining a line to run the full length of the visible lane based on the line segments you identified with the Hough Transform. As mentioned previously, try to average and/or extrapolate the line segments you've detected to map out the full extent of the lane lines. You can see an example of the result you're going for in the video "P1_example.mp4".**
**Go back and modify your draw_lines function accordingly and try re-running your pipeline. The new output should draw a single, solid line over the left lane line and a single, solid line over the right lane line. The lines should start from the bottom of the image and extend out to the top of the region of interest.**
Now for the one with the solid yellow lane on the left. This one's more tricky!
```
yellow_output = 'test_videos_output/solidYellowLeft.mp4'
## To speed up the testing process you may want to try your pipeline on a shorter subclip of the video
## To do so add .subclip(start_second,end_second) to the end of the line below
## Where start_second and end_second are integer values representing the start and end of the subclip
## You may also uncomment the following line for a subclip of the first 5 seconds
##clip2 = VideoFileClip('test_videos/solidYellowLeft.mp4').subclip(0,5)
clip2 = VideoFileClip('test_videos/solidYellowLeft.mp4')
yellow_clip = clip2.fl_image(process_image)
%time yellow_clip.write_videofile(yellow_output, audio=False)
HTML("""
<video width="960" height="540" controls>
<source src="{0}">
</video>
""".format(yellow_output))
```
## Writeup and Submission
If you're satisfied with your video outputs, it's time to make the report writeup in a pdf or markdown file. Once you have this Ipython notebook ready along with the writeup, it's time to submit for review! Here is a [link](https://github.com/udacity/CarND-LaneLines-P1/blob/master/writeup_template.md) to the writeup template file.
## Optional Challenge
Try your lane finding pipeline on the video below. Does it still work? Can you figure out a way to make it more robust? If you're up for the challenge, modify your pipeline so it works with this video and submit it along with the rest of your project!
```
challenge_output = 'test_videos_output/challenge.mp4'
## To speed up the testing process you may want to try your pipeline on a shorter subclip of the video
## To do so add .subclip(start_second,end_second) to the end of the line below
## Where start_second and end_second are integer values representing the start and end of the subclip
## You may also uncomment the following line for a subclip of the first 5 seconds
##clip3 = VideoFileClip('test_videos/challenge.mp4').subclip(0,5)
clip3 = VideoFileClip('test_videos/challenge.mp4')
challenge_clip = clip3.fl_image(process_image)
%time challenge_clip.write_videofile(challenge_output, audio=False)
HTML("""
<video width="960" height="540" controls>
<source src="{0}">
</video>
""".format(challenge_output))
```
| github_jupyter |
# Face Generation
In this project, you'll define and train a DCGAN on a dataset of faces. Your goal is to get a generator network to generate *new* images of faces that look as realistic as possible!
The project will be broken down into a series of tasks from **loading in data to defining and training adversarial networks**. At the end of the notebook, you'll be able to visualize the results of your trained Generator to see how it performs; your generated samples should look like fairly realistic faces with small amounts of noise.
### Get the Data
You'll be using the [CelebFaces Attributes Dataset (CelebA)](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html) to train your adversarial networks.
This dataset is more complex than the number datasets (like MNIST or SVHN) you've been working with, and so, you should prepare to define deeper networks and train them for a longer time to get good results. It is suggested that you utilize a GPU for training.
### Pre-processed Data
Since the project's main focus is on building the GANs, we've done *some* of the pre-processing for you. Each of the CelebA images has been cropped to remove parts of the image that don't include a face, then resized down to 64x64x3 NumPy images. Some sample data is show below.
<img src='assets/processed_face_data.png' width=60% />
> If you are working locally, you can download this data [by clicking here](https://s3.amazonaws.com/video.udacity-data.com/topher/2018/November/5be7eb6f_processed-celeba-small/processed-celeba-small.zip)
This is a zip file that you'll need to extract in the home directory of this notebook for further loading and processing. After extracting the data, you should be left with a directory of data `processed_celeba_small/`
```
# can comment out after executing
#!unzip processed_celeba_small.zip
data_dir = 'processed_celeba_small/'
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import pickle as pkl
import matplotlib.pyplot as plt
import numpy as np
import problem_unittests as tests
#import helper
%matplotlib inline
```
## Visualize the CelebA Data
The [CelebA](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html) dataset contains over 200,000 celebrity images with annotations. Since you're going to be generating faces, you won't need the annotations, you'll only need the images. Note that these are color images with [3 color channels (RGB)](https://en.wikipedia.org/wiki/Channel_(digital_image)#RGB_Images) each.
### Pre-process and Load the Data
Since the project's main focus is on building the GANs, we've done *some* of the pre-processing for you. Each of the CelebA images has been cropped to remove parts of the image that don't include a face, then resized down to 64x64x3 NumPy images. This *pre-processed* dataset is a smaller subset of the very large CelebA data.
> There are a few other steps that you'll need to **transform** this data and create a **DataLoader**.
#### Exercise: Complete the following `get_dataloader` function, such that it satisfies these requirements:
* Your images should be square, Tensor images of size `image_size x image_size` in the x and y dimension.
* Your function should return a DataLoader that shuffles and batches these Tensor images.
#### ImageFolder
To create a dataset given a directory of images, it's recommended that you use PyTorch's [ImageFolder](https://pytorch.org/docs/stable/torchvision/datasets.html#imagefolder) wrapper, with a root directory `processed_celeba_small/` and data transformation passed in.
```
# necessary imports
import torch
from torchvision import datasets
from torchvision import transforms
def get_dataloader(batch_size, image_size, data_dir='processed_celeba_small/'):
"""
Batch the neural network data using DataLoader
:param batch_size: The size of each batch; the number of images in a batch
:param img_size: The square size of the image data (x, y)
:param data_dir: Directory where image data is located
:return: DataLoader with batched data
"""
# TODO: Implement function and return a dataloader
transform = transforms.Compose([transforms.Resize(image_size),
transforms.ToTensor()])
dataset = datasets.ImageFolder(data_dir,transform=transform)
dataloader = torch.utils.data.dataloader.DataLoader(dataset, batch_size=batch_size,shuffle=True)
return dataloader
```
## Create a DataLoader
#### Exercise: Create a DataLoader `celeba_train_loader` with appropriate hyperparameters.
Call the above function and create a dataloader to view images.
* You can decide on any reasonable `batch_size` parameter
* Your `image_size` **must be** `32`. Resizing the data to a smaller size will make for faster training, while still creating convincing images of faces!
```
# Define function hyperparameters
batch_size = 32
img_size = 32
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# Call your function and get a dataloader
celeba_train_loader = get_dataloader(batch_size, img_size)
```
Next, you can view some images! You should seen square images of somewhat-centered faces.
Note: You'll need to convert the Tensor images into a NumPy type and transpose the dimensions to correctly display an image, suggested `imshow` code is below, but it may not be perfect.
```
# helper display function
def imshow(img):
npimg = img.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# obtain one batch of training images
dataiter = iter(celeba_train_loader)
images, _ = dataiter.next() # _ for no labels
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(20, 4))
plot_size=20
for idx in np.arange(plot_size):
ax = fig.add_subplot(2, plot_size/2, idx+1, xticks=[], yticks=[])
imshow(images[idx])
```
#### Exercise: Pre-process your image data and scale it to a pixel range of -1 to 1
You need to do a bit of pre-processing; you know that the output of a `tanh` activated generator will contain pixel values in a range from -1 to 1, and so, we need to rescale our training images to a range of -1 to 1. (Right now, they are in a range from 0-1.)
```
# TODO: Complete the scale function
def scale(x, feature_range=(-1, 1)):
''' Scale takes in an image x and returns that image, scaled
with a feature_range of pixel values from -1 to 1.
This function assumes that the input x is already scaled from 0-1.'''
# assume x is scaled to (0, 1)
# scale to feature_range and return scaled x
min, max = feature_range
x = x * (max - min) + min
return x
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# check scaled range
# should be close to -1 to 1
img = images[0]
scaled_img = scale(img)
print('Min: ', scaled_img.min())
print('Max: ', scaled_img.max())
```
---
# Define the Model
A GAN is comprised of two adversarial networks, a discriminator and a generator.
## Discriminator
Your first task will be to define the discriminator. This is a convolutional classifier like you've built before, only without any maxpooling layers. To deal with this complex data, it's suggested you use a deep network with **normalization**. You are also allowed to create any helper functions that may be useful.
#### Exercise: Complete the Discriminator class
* The inputs to the discriminator are 32x32x3 tensor images
* The output should be a single value that will indicate whether a given image is real or fake
```
def conv(in_channels, out_channels, kernal_size=3, stride =2, padding=0, batch_norm = False):
layers =[]
layers.append(nn.Conv2d(in_channels, out_channels, kernel_size =kernal_size, stride =stride, padding=padding, bias =False))
if batch_norm:
layers.append(nn.BatchNorm2d(out_channels))
return nn.Sequential(*layers)
import torch.nn as nn
import torch.nn.functional as F
class Discriminator(nn.Module):
def __init__(self, conv_dim):
"""
Initialize the Discriminator Module
:param conv_dim: The depth of the first convolutional layer
"""
super(Discriminator, self).__init__()
# complete init function
self.conv_dim = conv_dim
# out - 16
self.conv1 = conv(3, conv_dim, kernal_size=4, padding=1, batch_norm=False)
# out - 8
self.conv2 = conv(conv_dim, conv_dim*2, kernal_size=4, padding=1, batch_norm=True)
# out - 4
self.conv3 = conv(conv_dim*2, conv_dim*4, kernal_size=4, padding=1, batch_norm=True)
# out - 2
self.conv4 = conv(conv_dim*4, conv_dim*8, kernal_size=4, padding=1, batch_norm=True)
# out - 1
self.conv5 = conv(conv_dim*8, conv_dim*16, kernal_size=4, padding=1, batch_norm=False)
self.fc = nn.Linear(conv_dim*16, 1)
def forward(self, x):
"""
Forward propagation of the neural network
:param x: The input to the neural network
:return: Discriminator logits; the output of the neural network
"""
# define feedforward behavior
out = F.leaky_relu(self.conv1(x),0.2)
out = F.leaky_relu(self.conv2(out),0.2)
out = F.leaky_relu(self.conv3(out),0.2)
out = F.leaky_relu(self.conv4(out),0.2)
out = F.leaky_relu(self.conv5(out),0.2)
#print(out.shape)
out = out.view(-1, self.conv_dim*16)
#print(out.shape)
out = self.fc(out)
#print(out.shape)
return out
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_discriminator(Discriminator)
```
## Generator
The generator should upsample an input and generate a *new* image of the same size as our training data `32x32x3`. This should be mostly transpose convolutional layers with normalization applied to the outputs.
#### Exercise: Complete the Generator class
* The inputs to the generator are vectors of some length `z_size`
* The output should be a image of shape `32x32x3`
```
def dconv(in_channels, out_channels, kernal_size=3, stride =2, padding=0, batch_norm = False):
layers =[]
layers.append(nn.ConvTranspose2d(in_channels, out_channels, kernel_size =kernal_size, stride =stride, padding=padding))
if batch_norm:
layers.append(nn.BatchNorm2d(out_channels))
return nn.Sequential(*layers)
class Generator(nn.Module):
def __init__(self, z_size, conv_dim):
"""
Initialize the Generator Module
:param z_size: The length of the input latent vector, z
:param conv_dim: The depth of the inputs to the *last* transpose convolutional layer
"""
super(Generator, self).__init__()
# complete init function
self.z_size = z_size
self.conv_dim = conv_dim
self.fc = nn.Linear(z_size, conv_dim*8*2*2)
# out - 4
self.dconv1 = dconv(conv_dim*8, conv_dim*4, kernal_size=4, padding=1, batch_norm=True)
# out - 8
self.dconv2 = dconv(conv_dim*4, conv_dim*2, kernal_size=4, padding=1, batch_norm=True)
# out - 16
self.dconv3 = dconv(conv_dim*2, conv_dim, kernal_size=4, padding=1, batch_norm=True)
# out - 32
self.dconv4 = dconv(conv_dim, 3, kernal_size=4, padding=1, batch_norm=False)
# # out - 2
# self.dconv4 = dconv(conv_dim*4, conv_dim*8, kernal_size=4, padding=1, batch_norm=True)
# # out - 1
# self.dconv5 = dconv(conv_dim*16, conv_dim*32, kernal_size=4, padding=1, batch_norm=False)
def forward(self, x):
"""
Forward propagation of the neural network
:param x: The input to the neural network
:return: A 32x32x3 Tensor image as output
"""
# define feedforward behavior
out = self.fc(x)
out = out.view(-1, self.conv_dim*8,2,2)
out = F.relu(self.dconv1(out))
out = F.relu(self.dconv2(out))
out = F.relu(self.dconv3(out))
out = F.tanh(self.dconv4(out))
return out
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_generator(Generator)
```
## Initialize the weights of your networks
To help your models converge, you should initialize the weights of the convolutional and linear layers in your model. From reading the [original DCGAN paper](https://arxiv.org/pdf/1511.06434.pdf), they say:
> All weights were initialized from a zero-centered Normal distribution with standard deviation 0.02.
So, your next task will be to define a weight initialization function that does just this!
You can refer back to the lesson on weight initialization or even consult existing model code, such as that from [the `networks.py` file in CycleGAN Github repository](https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/blob/master/models/networks.py) to help you complete this function.
#### Exercise: Complete the weight initialization function
* This should initialize only **convolutional** and **linear** layers
* Initialize the weights to a normal distribution, centered around 0, with a standard deviation of 0.02.
* The bias terms, if they exist, may be left alone or set to 0.
```
def weights_init_normal(m):
"""
Applies initial weights to certain layers in a model .
The weights are taken from a normal distribution
with mean = 0, std dev = 0.02.
:param m: A module or layer in a network
"""
# classname will be something like:
# `Conv`, `BatchNorm2d`, `Linear`, etc.
classname = m.__class__.__name__
# TODO: Apply initial weights to convolutional and linear layers
if classname =='Conv2d':
w = torch.empty(m.weight.data.shape)
m.weight.data = nn.init.normal_(w,mean=0, std=0.02)
if classname =='Linear':
w = torch.empty(m.weight.data.shape)
m.weight.data = nn.init.normal_(w,mean=0, std=0.02)
```
## Build complete network
Define your models' hyperparameters and instantiate the discriminator and generator from the classes defined above. Make sure you've passed in the correct input arguments.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
def build_network(d_conv_dim, g_conv_dim, z_size):
# define discriminator and generator
D = Discriminator(d_conv_dim)
G = Generator(z_size=z_size, conv_dim=g_conv_dim)
# initialize model weights
D.apply(weights_init_normal)
G.apply(weights_init_normal)
print(D)
print()
print(G)
return D, G
```
#### Exercise: Define model hyperparameters
```
# Define model hyperparams
d_conv_dim = 32
g_conv_dim = 32
z_size = 100
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
D, G = build_network(d_conv_dim, g_conv_dim, z_size)
```
### Training on GPU
Check if you can train on GPU. Here, we'll set this as a boolean variable `train_on_gpu`. Later, you'll be responsible for making sure that
>* Models,
* Model inputs, and
* Loss function arguments
Are moved to GPU, where appropriate.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import torch
# Check for a GPU
train_on_gpu = torch.cuda.is_available()
if not train_on_gpu:
print('No GPU found. Please use a GPU to train your neural network.')
else:
print('Training on GPU!')
```
---
## Discriminator and Generator Losses
Now we need to calculate the losses for both types of adversarial networks.
### Discriminator Losses
> * For the discriminator, the total loss is the sum of the losses for real and fake images, `d_loss = d_real_loss + d_fake_loss`.
* Remember that we want the discriminator to output 1 for real images and 0 for fake images, so we need to set up the losses to reflect that.
### Generator Loss
The generator loss will look similar only with flipped labels. The generator's goal is to get the discriminator to *think* its generated images are *real*.
#### Exercise: Complete real and fake loss functions
**You may choose to use either cross entropy or a least squares error loss to complete the following `real_loss` and `fake_loss` functions.**
```
def real_loss(D_out, smooth=False):
'''Calculates how close discriminator outputs are to being real.
param, D_out: discriminator logits
return: real loss'''
batch_size = D_out.size(0)
if smooth:
labels = torch.ones(batch_size)*0.9
else:
labels = torch.ones(batch_size)
if train_on_gpu:
labels = labels.cuda()
criterion = nn.BCEWithLogitsLoss()
loss = criterion(D_out.squeeze(), labels)
return loss
def fake_loss(D_out):
'''Calculates how close discriminator outputs are to being fake.
param, D_out: discriminator logits
return: fake loss'''
batch_size = D_out.size(0)
labels = torch.zeros(batch_size)
if train_on_gpu:
labels = labels.cuda()
criterion = nn.BCEWithLogitsLoss()
loss = criterion(D_out.squeeze(), labels)
return loss
```
## Optimizers
#### Exercise: Define optimizers for your Discriminator (D) and Generator (G)
Define optimizers for your models with appropriate hyperparameters.
```
import torch.optim as optim
lr = 0.0002
beta1 = 0.5
beta2 = 0.999
# Create optimizers for the discriminator D and generator G
d_optimizer = optim.Adam(D.parameters(),lr = lr, betas=[beta1,beta2])
g_optimizer = optim.Adam(G.parameters(),lr = lr, betas=[beta1,beta2])
```
---
## Training
Training will involve alternating between training the discriminator and the generator. You'll use your functions `real_loss` and `fake_loss` to help you calculate the discriminator losses.
* You should train the discriminator by alternating on real and fake images
* Then the generator, which tries to trick the discriminator and should have an opposing loss function
#### Saving Samples
You've been given some code to print out some loss statistics and save some generated "fake" samples.
#### Exercise: Complete the training function
Keep in mind that, if you've moved your models to GPU, you'll also have to move any model inputs to GPU.
```
def train(D, G, n_epochs, print_every=50):
'''Trains adversarial networks for some number of epochs
param, D: the discriminator network
param, G: the generator network
param, n_epochs: number of epochs to train for
param, print_every: when to print and record the models' losses
return: D and G losses'''
# move models to GPU
if train_on_gpu:
D.cuda()
G.cuda()
# keep track of loss and generated, "fake" samples
samples = []
losses = []
# Get some fixed data for sampling. These are images that are held
# constant throughout training, and allow us to inspect the model's performance
sample_size=16
fixed_z = np.random.uniform(-1, 1, size=(sample_size, z_size))
fixed_z = torch.from_numpy(fixed_z).float()
# move z to GPU if available
if train_on_gpu:
fixed_z = fixed_z.cuda()
# epoch training loop
for epoch in range(n_epochs):
# batch training loop
for batch_i, (real_images, _) in enumerate(celeba_train_loader):
batch_size = real_images.size(0)
real_images = scale(real_images)
if train_on_gpu:
real_images = real_images.cuda()
# ===============================================
# YOUR CODE HERE: TRAIN THE NETWORKS
# ===============================================
# 1. Train the discriminator on real and fake images
d_optimizer.zero_grad()
d_real_out = D(real_images)
#print(d_real_out.shape)
d_real_loss = real_loss(d_real_out)
z = np.random.uniform(-1, 1, size=(batch_size, z_size))
z = torch.from_numpy(z).float()
if train_on_gpu:
z = z.cuda()
d_fake_out = G(z)
d_fake_out_bloss = D(d_fake_out)
#print(d_fake_out_bloss.shape)
d_fake_out_loss = fake_loss(d_fake_out_bloss)
#print(d_fake_out_bloss.shape)
d_loss = d_real_loss + d_fake_out_loss
#print(d_total_loss)
d_loss.backward()
d_optimizer.step()
g_optimizer.zero_grad()
# 2. Train the generator with an adversarial loss
z = np.random.uniform(-1, 1, size=(batch_size, z_size))
z = torch.from_numpy(z).float()
if train_on_gpu:
z = z.cuda()
fake_out = G(z)
D_fake = D(fake_out)
g_loss = real_loss(D_fake)
g_loss.backward()
g_optimizer.step()
# ===============================================
# END OF YOUR CODE
# ===============================================
# Print some loss stats
if batch_i % print_every == 0:
# append discriminator loss and generator loss
losses.append((d_loss.item(), g_loss.item()))
# print discriminator and generator loss
print('Epoch [{:5d}/{:5d}] | d_loss: {:6.4f} | g_loss: {:6.4f}'.format(
epoch+1, n_epochs, d_loss.item(), g_loss.item()))
## AFTER EACH EPOCH##
# this code assumes your generator is named G, feel free to change the name
# generate and save sample, fake images
G.eval() # for generating samples
samples_z = G(fixed_z)
samples.append(samples_z)
G.train() # back to training mode
# Save training generator samples
with open('train_samples.pkl', 'wb') as f:
pkl.dump(samples, f)
# finally return losses
return losses
```
Set your number of training epochs and train your GAN!
```
# set number of epochs
n_epochs = 100
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# call training function
from workspace_utils import active_session
with active_session():
losses = train(D, G, n_epochs=n_epochs)
```
## Training loss
Plot the training losses for the generator and discriminator, recorded after each epoch.
```
fig, ax = plt.subplots()
losses = np.array(losses)
plt.plot(losses.T[0], label='Discriminator', alpha=0.5)
plt.plot(losses.T[1], label='Generator', alpha=0.5)
plt.title("Training Losses")
plt.legend()
```
## Generator samples from training
View samples of images from the generator, and answer a question about the strengths and weaknesses of your trained models.
```
# helper function for viewing a list of passed in sample images
def view_samples(epoch, samples):
fig, axes = plt.subplots(figsize=(16,4), nrows=2, ncols=8, sharey=True, sharex=True)
for ax, img in zip(axes.flatten(), samples[epoch]):
img = img.detach().cpu().numpy()
img = np.transpose(img, (1, 2, 0))
img = ((img + 1)*255 / (2)).astype(np.uint8)
ax.xaxis.set_visible(False)
ax.yaxis.set_visible(False)
im = ax.imshow(img.reshape((32,32,3)))
# Load samples from generator, taken while training
with open('train_samples.pkl', 'rb') as f:
samples = pkl.load(f)
_ = view_samples(-1, samples)
```
### Question: What do you notice about your generated samples and how might you improve this model?
When you answer this question, consider the following factors:
* The dataset is biased; it is made of "celebrity" faces that are mostly white
* Model size; larger models have the opportunity to learn more features in a data feature space
* Optimization strategy; optimizers and number of epochs affect your final result
**Answer:**
- As mentioned in the above text, celebrity photos are mostly white. Need to collect different samples of other color to make a general imporved model to generate all sorts of color.
- Go with more deeper model on generator and discriminator with more convolution layers.
- After many iterations I have settled down with the used hyperparameters. Model can be trained somemore epochs for a better model.
- Create more transformations, so better pictures will be generated which looks like real one.
- Tune around beta values for the optimizer to improve the model.
- Input higher quality input images and add more convolutions on generator and discriminator.
### Submitting This Project
When submitting this project, make sure to run all the cells before saving the notebook. Save the notebook file as "dlnd_face_generation.ipynb" and save it as a HTML file under "File" -> "Download as". Include the "problem_unittests.py" files in your submission.
| github_jupyter |
```
try:
import openmdao.api as om
except ImportError:
!python -m pip install openmdao[notebooks]
import openmdao.api as om
```
# Writing Plugins
OpenMDAO was designed to allow you to code up your own components,
groups, etc., and to use them within the framework, but what if you want others to be able to
discover and use your creations? The OpenMDAO plugin system was created to make that easier.
Before laying out the steps to follow in order to create your plugin, a brief discussion of
entry points is in order. An entry point is simply a string passed into the `setup()` function
in the `setup.py` file for your python package. The string has the form:
```
'my_ep_name=my_plugin_module_path:my_module_attribute'
```
where typically, `my_module_attribute` is a class or a function.
The plugin system uses entry points in order to provide local discovery, and in some cases to
support adding new functionality to openmdao, e.g., adding new openmdao command line tools.
Every entry point is associated with an entry point group, and
the entry point groups that openmdao recognizes are shown in the table below:
| Entry Point Group | Type | Entry Point Refers To |
|-------------------------- |------------------- |-------------------------------------------------------------|
| openmdao_component | Component | class or factory funct |
| openmdao_group | Group | class or factory funct |
| openmdao_driver | Driver | class or factory funct |
| openmdao_lin_solver | LinearSolver | class or factory funct |
| openmdao_nl_solver | NonlinearSolver | class or factory funct |
| openmdao_surrogate_model | SurrogateModel | class or factory funct |
| openmdao_case_recorder | CaseRecorder | class or factory funct |
| openmdao_case_reader | BaseCaseReader | funct returning (file_ext, class or factor funct) |
| openmdao_command | command line tool | funct returning (setup_parser_func, exec_func, help_string) |
## 'Typical' Plugins
Most OpenMDAO plugins are created simply by registering an entry point that refers
to the class definition of the plugin or to some factory function that returns an instance of
the plugin. The following entry point types are all handled in this way:
- component
- group
- driver
- nl_solver
- lin_solver
- surrogate_model
- case_recorder
For these types of plugins, the entry point does nothing other than allow them to be listed using
the [openmdao list_installed](list-installed) command.
Here's an example of how to specify the *entry_points* arg to the *setup* call in `setup.py`
for a component plugin class called `MyComponent` in a package called `my_plugins_package`
in a module called `my_comp_plugin.py`:
```
entry_points={
'openmdao_component': [
'mycompplugin=my_plugins_package.my_comp_plugin:MyComponent'
]
}
```
Note that the actual entry point name, `mycompplugin` in the example above, isn't used for
anything in the case of a 'typical' plugin.
## CaseReader Plugins
The entry point for a case reader should point to a function that returns a tuple of the form
(file_extension, class), where *file_extension* contains the leading dot, for example '.sql',
and *class* could either be the class definition of the plugin or a factory function returning
an instance of the plugin. The file extension is used to provide an automatic mapping to the
correct case reader based on the file extension of the file being read.
## Command Line Tool Plugins
An entry point for an OpenMDAO command line tool plugin should point to a function that returns
a tuple of the form (setup_parser_func, exec_func, help_string). For example:
```
def _hello_setup():
"""
This command prints a hello message after final setup.
"""
return (_hello_setup_parser, _hello_exec, 'Print hello message after final setup.')
```
The *setup_parser_func* is a function taking a single *parser* argument that adds any arguments
expected by the plugin to the *parser* object. The *parser* is an *argparse.ArgumentParser* object.
For example, the following code sets up a subparser for a `openmdao hello` command that adds a file
argument and a `--repeat` option:
```
def _hello_setup_parser(parser):
"""
Set up the openmdao subparser (using argparse) for the 'openmdao hello' command.
Parameters
----------
parser : argparse subparser
The parser we're adding options to.
"""
parser.add_argument('-r', '--repeat', action='store', dest='repeats',
default=1, type=int, help='Number of times to say hello.')
parser.add_argument('file', metavar='file', nargs=1,
help='Script to execute.')
```
The *exec_func* is a function that performs whatever action is necessary for the command line
tool plugin to operate. Typically this will involve registering another function that is to
execute at some point during the execution of a script file. For example, the following
function registers a function that prints a `hello` message, specifying that it should execute
after the `Problem._final_setup` method.
```
def _hello_exec(options, user_args):
"""
This registers the hook function and executes the user script.
Parameters
----------
options : argparse Namespace
Command line options.
user_args : list of str
Args to be passed to the user script.
"""
script = options.file[0]
def _hello_after_final_setup(prob):
for i in range(options.repeats):
print('*** hello ***')
exit() # If you want to exit after your command, you must explicitly do that here
# register the hook to execute after Problem.final_setup
_register_hook('final_setup', class_name='Problem', post=_hello_after_final_setup)
# load and execute the given script as __main__
_load_and_exec(script, user_args)
```
The final entry in the tuple returned by the function referred to by the entry point
(in this case *_hello_setup*)
is a string containing a high level description of the command. This description will be displayed
along with the name of the command when a user runs `openmdao -h`.
Here's an example of how to specify the *entry_points* arg to the *setup* call in `setup.py`
for our command line tool described above if it were inside of a package called `my_plugins_package`
in a file called `hello_cmd.py`:
```
entry_points={
'openmdao_command': [
'hello=my_plugins_package.hello_cmd:_hello_setup'
]
}
```
In this case, the name of our entry point, `hello`, will be the name of the openmdao command line
tool, so the user will activate the tool by typing `openmdao hello`.
## Local Discovery
After a python package containing OpenMDAO plugins has been installed in a user's python
environment, they will be able to print a list of installed plugins using the
[openmdao list_installed](list-installed) command.
For example, if a package called `foobar` is installed, we could list all of the plugins
found in that package using the following command:
```
openmdao list_installed -i foobar
```
The `list_installed` command simply goes through all of the entry points it finds in any of the
openmdao entry point groups described above and displays them.
## Global Discovery Using github
Entry point groups are also used for global discovery of plugins. They can be used (in slightly
modified form, with underscores replaced with dashes) as *topic* strings in a github repository
in order to allow a user to perform a global search over all of github to find any openmdao related
plugin packages.
## Plugin Creation from Scratch
To create an OpenMDAO plugin from scratch, it may be helpful to use the [openmdao scaffold](ref-scaffold) tool. It will automatically generate the directory structure for a python package and will define the entry point of a type that you specify. For example, to create a scaffold for a python package called mypackage that contains a component plugin that's an ExplicitComponent called MyComp, do the following:
```
openmdao scaffold --base=ExplicitComponent --class=MyComp --package=mypackage
```
To instead create a package containing an openmdao command line tool called `hello` in
a package called `myhello`, do the following:
```
openmdao scaffold --cmd=hello --package=myhello
```
## Converting Existing Classes to Plugins
If you already have a package containing components, groups, etc. that work in the OpenMDAO
framework, all you need to do to register them as plugins is to define an entry point in
your `setup.py` file for each one.
You can use the [openmdao compute_entry_points](compute-entry-points) command
line tool to help you do this. Running the tool with your installed package name will print
out a list of all of the openmdao entry points required to register any openmdao compatible
classes it finds in your package. For example, if your package is called `mypackage`, you
can list its entry points using
```
openmdao compute_entry_points mypackage
```
The entry points will be printed out in a form that can be pasted as a *setup* argument into
your `setup.py` file.
## Plugin Checklist
To recap, to **fully** integrate your plugin into the OpenMDAO plugin infrastructure, you must do all
of the following:
1. The plugin will be part of a pip-installable python package.
2. An entry point will be added to the appropriate entry point group (see above) of the
*entry_points* argument passed to the *setup* call in the *setup.py* file for the python package containing the plugin.
3. If the package resides in a public **github** repository, the `openmdao` topic will be added
to the repository, along with topics for each openmdao entry point group (with underscores
converted to dashes, e.g., `openmdao_component` becomes `openmdao-component`) that
contains an openmdao entry point found in the package.
4. If the package resides on the Python Package Index (PyPI), the string `openmdao` should be
mentioned in the package summary.
5. To support the future ability to query PyPI package keywords, any openmdao entry point
groups used by the package should be added to the `keywords` argument to the *setup*
call in the *setup.py* file for the package.
| github_jupyter |
## Breakout session
We want to reproduce the Global Land-Ocean Temperature index plot below (available at this <A HREF="http://data.giss.nasa.gov/gistemp/graphs_v3/"> GISS website</A>). Download the data <A HREF="http://data.giss.nasa.gov/gistemp/graphs_v3/Fig.A2.txt">this link</A> and recreate the plot **as closely as you can** (don't worry about the two green points or matching the font). We've broken up this problem into steps to get you started. Feel free to follow them as you wish, or be a lone wolf and solve this on your own.
<img src="http://data.giss.nasa.gov/gistemp/graphs_v3/Fig.A2.gif">
1. Save the data as a text file, and then read the data into a numpy array. You'll have to deal with the extra lines at the top and bottom of the file, as well as the missing values (the asterixes) that appear. There are multiple ways to solve these problems; do what works for you. If you get stuck, remember that we talked about slicing arrays at the beginning of this lecture.
2. Plot "Annual Mean" as a set of black squares connected by a thick black line. Try Googling something about "matplotlib square markers". Don't forget to give it a label so it can go in the legend later.
3. Plot "5-Year Running Mean" as a thick red line. Don't forget the label.
4. Set the x and y ranges of your plot so that they match the ones in the plot we are trying to recreate.
5. Add a grid. The default grid that shows up is fine; don't worry about making it match *exactly*.
6. Label the y axis. Don't worry about the "degree" symbol for now.
7. Add the title.
8. Add the legend (default settings are fine).
BONUS TASKS:
1. Notice how, in the the plot above, there are labels at every 20 years along the x axis but grid marks at every ten (and similar for the y axis)? Figure out how to recreate this.
2. On that note, match the formatting they have for the y axis ticks.
3. Make all the font sizes match (yours are probably smaller than you'd like).
4. Add in the "degree" symbol for the y axis label. Hint: LaTeX or unicode.
5. Move the legend to the right place. Remove the box (the "frame") that surrounds the legend. Notice how your legend has a line with *two* markers for "Annual mean" and their legend has a line with only one marker? Match that.
6. Move the title up a little bit so the spacing better matches the plot above.
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
# I've chosen to save the "Annual means" and "5-year means" data sets into two separate sets of x- and y-values:
annual_means = np.genfromtxt('../Fig.A2.txt', dtype=float, skip_header=4, skip_footer=2, usecols=(0,1))
fiveyear_means = np.genfromtxt('../Fig.A2.txt', dtype=float, skip_header=6, skip_footer=4, usecols=(0,2))
plt.figure(figsize=(12,8))
plt.plot(annual_means[:,0], annual_means[:,1], 's-', color='k', lw=2, label="Annual Mean")
plt.plot(fiveyear_means[:,0], fiveyear_means[:,1], color='r', lw=2, label="5-year Running Mean")
plt.xlim(1880, 2015)
plt.ylim(-0.5, 0.7)
## Note: The way I have below is a brute-force method. If you do a little Googling you can
## find a more elegant solution using minor ticks. That having been said, this is probaly the
## easiest way to match the way their y axis labels EXACTLY.
myXTicks = np.arange(1880, 2020, 10)
myXTickLabels = ('1880', '', '1900', '', '1920', '', '1940', '', '1960', '', '1980', '', '2000')
plt.xticks(myXTicks, myXTickLabels, size=18)
myYTicks = np.arange(-0.5, 0.7, 0.1)
myYTickLabels = ('', '-.4', '', '-.2', '', '0.', '', '.2', '', '.4', '', '.6')
plt.yticks(myYTicks, myYTickLabels, size=18)
plt.grid(True)
plt.ylabel(r'Temperature Anomaly ($^\circ$C)', size=20)
plt.title('Global Land-Ocean Temperature Index', size=24, y=1.03)
plt.legend(loc=(0.1, 0.7), fontsize=20, numpoints=1, frameon=0)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/HomelyAi/stylegan2-ada/blob/main/style_mixing.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
!nvidia-smi -L
%tensorflow_version 1.x
from IPython.display import display,Image as Image2
!git clone https://github.com/vishnukool/stylegan2-ada.git
%cd /content/stylegan2-ada
!pip install awscli
network_pkl = 'network-snapshot-009800.pkl'
!aws s3 cp s3://homely-pretrained-models/$network_pkl ./
from PIL import Image
def showImages(images):
widths, heights = zip(*(i.size for i in images))
total_width = sum(widths)
max_height = max(heights)
new_im = Image.new('RGB', (total_width, max_height))
x_offset = 0
for im in images:
new_im.paste(im, (x_offset,0))
x_offset += im.size[0]
display(new_im)
# -- Gray Traditional --
# https://www.ikea.com/gb/en/images/products/ektorp-2-seat-sofa__0818544_pe774475_s5.jpg
# projectPath1 = 'data/a5.jpg'
# outPath1 = 'out/a5'
# -- Recessed Arm Pink --
# imageUrl = 'https://secure.img1-fg.wfcdn.com/im/41020583/resize-h900-w900%5Ecompr-r85/1069/106985690/Jonas+66.75%27%27+Recessed+Arm+Loveseat+with+Reversible+Cushions.jpg'
# projectPath1 = 'data/a6.jpg'
# outPath1 = 'out/a6'
# -- Pillow top Arm --
# imageUrl = 'https://secure.img1-fg.wfcdn.com/im/98861474/resize-h310-w310%5Ecompr-r85/1456/145637496/52.75%27%27+Faux+Leather+Pillow+Top+Arm+Reclining.jpg'
# projectPath1 = 'data/a7.jpg'
# outPath1 = 'out/a7'
# -- Gray Modern --
# imageUrl = 'https://www.boconcept.com/on/demandware.static/-/Sites-master-catalog/default/dw7429ad62/images/710000/719958.jpg'
# projectPath1 = 'data/a8.jpg'
# outPath1 = 'out/a8'
# -- White Cushy --
# imageUrl = 'https://i.imgur.com/dweqIyf.jpg'
# projectPath1 = 'data/a9.jpg'
# outPath1 = 'out/a9'
# -- Orange Leather Apt Therapy --
# imageUrl = 'https://i.imgur.com/C9hZKgN.jpg'
# projectPath1 = 'data/a10.jpg'
# outPath1 = 'out/a10'
# -- Black Leathery --
# imageUrl = 'https://encrypted-tbn0.gstatic.com/images?q=tbn:ANd9GcSFKW8GkZjfjqVYto6-C8m-6DiwTNeYF2C-8RAgrPq8Au1EUlQkl0Yzs1ycgYILOVXBb6KWNTo&usqp=CAc'
# projectPath1 = 'data/a11.jpg'
# outPath1 = 'out/a11'
# ----
# -- Recessed Arm --
# imageUrl = 'https://secure.img1-fg.wfcdn.com/im/23871536/resize-h800-w800%5Ecompr-r85/8461/84611427/Cheeky+49.5%27%27+Recessed+Arm+Loveseat.jpg'
# projectPath1 = 'data/a12.jpg'
# outPath1 = 'out/a12'
# -- Recessed Arm 2 --
# imageUrl = 'https://secure.img1-fg.wfcdn.com/im/53536085/resize-h310-w310%5Ecompr-r85/5520/55205111/default_name.jpg'
# projectPath1 = 'data/a13.jpg'
# outPath1 = 'out/a13'
# -- Recessed Arm 3 --
# imageUrl = 'https://secure.img1-fg.wfcdn.com/im/65284946/resize-h310-w310%5Ecompr-r85/9910/99107984/Abbott+78%27%27+Recessed+Arm+Sofa.jpg'
# projectPath1 = 'data/a14.jpg'
# outPath1 = 'out/a14'
# -- Wayfair Dark Blue --
# imageUrl = 'https://secure.img1-fg.wfcdn.com/im/67481968/resize-h800-w800%5Ecompr-r85/1175/117510521/56.69%27%27+Recessed+Arm+Loveseat.jpg'
# projectPath1 = 'data/a15.jpg'
# outPath1 = 'out/a15'
# -- Single Seat Gray --
# imageUrl = 'https://loveseat.mx/media/catalog/product/l/o/love_seat_skara_quantum_ash_1.png'
# projectPath1 = 'data/a16.jpg'
# outPath1 = 'out/a16'
# -- Yellow --
# imageUrl = 'https://encrypted-tbn0.gstatic.com/images?q=tbn:ANd9GcTvY7t9efXRvQ0Dv5btk8-6g9EtL-N8uTeGng&usqp=CAU'
# projectPath1 = 'data/a17.jpg'
# outPath1 = 'out/a17'
# -- Blue Tufted, trouble tufting --
# imageUrl = 'https://cdn.shopify.com/s/files/1/2660/5202/products/shopify-image_a6fa3eb1-c388-4ce9-a914-c9ea3e4384cf_1400x.jpg?v=1614976781'
# projectPath1 = 'data/a18.jpg'
# outPath1 = 'out/a18'
# -- Gray Love Seat with Pillows --
# imageUrl = 'https://images.furnituredealer.net/img/products%2Fbenchcraft%2Fcolor%2Fmercado_8460435-b1.jpg'
# projectPath1 = 'data/a19.jpg'
# outPath1 = 'out/a19'
# -- Wayfair Traditional bench seat --
# imageUrl = 'https://secure.img1-fg.wfcdn.com/im/34805391/resize-h800-w800%5Ecompr-r85/5279/52791200/Cali+49.5%27%27+Loveseat.jpg'
# projectPath1 = 'data/a20.jpg'
# outPath1 = 'out/a20'
# -- Wayfair Flared Arm --
# imageUrl = 'https://secure.img1-fg.wfcdn.com/im/83196554/resize-h900-w900%5Ecompr-r85/4293/42930166/Erinn+67.25%27%27+Flared+Arm+Loveseat.jpg'
# projectPath1 = 'data/a21.jpg'
# outPath1 = 'out/a21'
# -- Armless --
# imageUrl = 'https://secure.img1-fg.wfcdn.com/im/89601253/resize-h800-w800%5Ecompr-r85/4277/42778262/Valeria+55.1%27%27+Armless+Loveseat.jpg'
# projectPath1 = 'data/a22.jpg'
# outPath1 = 'out/a22'
# -- Single Seat but shows 2 seats --
# imageUrl = 'https://i.imgur.com/wBkyw6l.jpg'
# projectPath1 = 'data/a23.jpg'
# outPath1 = 'out/a23'
# -- Double Cushion Arm problem --
# imageUrl = 'https://moblum.com/media/catalog/product/cache/dc1ae9f78f90a12f86cb0d1e3aec3fb2/s/a/sal54325s1-ruan-gris-obscuro-love_seat-01_1.jpg'
# projectPath1 = 'data/a24.jpg'
# outPath1 = 'out/a24'
# -- Asymmetric pillow top arm --
# imageUrl = 'https://mueblesmexico.com/wp-content/uploads/2020/06/Love-Seat-Acord-Warest-Silver.jpg'
# projectPath1 = 'data/a25.jpg'
# outPath1 = 'out/a25'
# -- Sloping Arm, single seat problem --
# imageUrl = 'https://mueblesmexico.com/wp-content/uploads/2020/06/Love-Seat-Alicia-Curri-Demin.jpg'
# projectPath1 = 'data/a27.jpg'
# outPath1 = 'out/a27'
# -- Pillow Arm problem
# imageUrl = 'https://cdn1.coppel.com/images/catalog/pm/4237953-3.jpg'
# projectPath1 = 'data/a28.jpg'
# outPath1 = 'out/a28'
# -- Heavy tuft and arm
# imageUrl = 'https://mueblesmexico.com/wp-content/uploads/2020/06/Love-Seat-Bronx-Velvet-Mustard.jpg'
# projectPath1 = 'data/a29.jpg'
# outPath1 = 'out/a29'
# -- No Idea why --
# imageUrl = 'https://cdn1.coppel.com/images/catalog/pm/4767593-3.jpg'
# projectPath1 = 'data/a30.jpg'
# outPath1 = 'out/a30'
# -- Single seat, sloping arm --
# imageUrl = 'https://mueblesmexico.com/wp-content/uploads/2020/06/Love-Seat-Gandhi-Curri-Demin.jpg'
# projectPath1 = 'data/a31.jpg'
# outPath1 = 'out/a31'
# -- Multi pillow problem --
# imageUrl = 'https://d3uq4j19mzp2q8.cloudfront.net/media/catalog/product/cache/16/image/1200x590/9df78eab33525d08d6e5fb8d27136e95/s/a/sal69587k1-c-f-1-w_1.jpg'
# projectPath1 = 'data/a32.jpg'
# outPath1 = 'out/a32'
# ------------------ ROTATOIN IMAGES --------------------------------
# https://images-na.ssl-images-amazon.com/images/I/61gWcr%2BtWYL._SP1017,567,0%7C518vFSohJkL.jpg,61qEo-+QkUL.jpg,61ysawsCBXL.jpg,51MboY31LXL.jpg,61rRd195GGL.jpg_.jpg
# projectPath1 = 'data/b1.jpg'
# outPath1 = 'out/b1'
# https://www.amazon.com/Versatile-Modern-Storage-Loveseat-Charcoal/dp/B072LT9YC3/ref=sxin_12_ac_d_rm?ac_md=1-1-YXJtbGVzcyBsb3Zlc2VhdA%3D%3D-ac_d_rm_rm_rm&cv_ct_cx=armless+sofa&dchild=1&keywords=armless+sofa&pd_rd_i=B072LT9YC3&pd_rd_r=d8e2d496-4a0f-4955-994d-03b1349e4f0c&pd_rd_w=YdWrj&pd_rd_wg=e1RPV&pf_rd_p=bdf723b2-f1c3-4d1c-967e-60197e162550&pf_rd_r=T2A173RHSM9XN3WPVFCS&psc=1&qid=1627955380&sr=1-2-12d4272d-8adb-4121-8624-135149aa9081
# projectPath1 = 'data/b2.jpg'
# outPath1 = 'out/b2'
# The above link stretched to make it longer
# projectPath1 = 'data/b5.jpg'
# outPath1 = 'out/b5'
# https://www.wayfair.com/furniture/pdp/hashtag-home-valeria-551-armless-loveseat-w004143937.html
# projectPath1 = 'data/b9.jpg'
# outPath1 = 'out/b9'
# https://www.amazon.com/POLY-BARK-Napa-Variation-Cognac/dp/B07QPQCQKQ/ref=sxin_13_ac_d_rm?ac_md=3-1-bGVhdGhlciBzb2Zh-ac_d_rm_rm_rm&cv_ct_cx=sofa&dchild=1&keywords=sofa&pd_rd_i=B07QPQCQKQ&pd_rd_r=df1a7480-5af0-4909-ab06-cdf21b4eb431&pd_rd_w=PhGXb&pd_rd_wg=Y1fba&pf_rd_p=bdf723b2-f1c3-4d1c-967e-60197e162550&pf_rd_r=24ZDVCWSW4VA3N8XVHJN&psc=1&qid=1628008261&sr=1-2-12d4272d-8adb-4121-8624-135149aa9081
# projectPath1 = 'data/c1.jpg'
# outPath1 = 'out/c1'
# https://www.amazon.com/Zinus-Classic-Upholstered-Living-Couch/dp/B079PP84CC/ref=sr_1_5?dchild=1&keywords=sofa&qid=1628008831&s=home-garden&sr=1-5
# Side View
# projectPath1 = 'data/c2.jpg'
# outPath1 = 'out/c2'
projectPath1 = 'data/c4.jpg'
outPath1 = 'out/c4'
!python projector.py --outdir=$outPath1 --target=$projectPath1 --save-video=False --network=./network-snapshot-009800.pkl
showImages([Image.open('./' + outPath1+'/target.png'), Image.open('./' + outPath1+'/proj.png')])
import dnnlib
import dnnlib.tflib as tflib
import pickle
import numpy as np
from PIL import Image
Gs_syn_kwargs = {
'output_transform': dict(func=tflib.convert_images_to_uint8, nchw_to_nhwc=True),
'randomize_noise': False,
'minibatch_size': 4
}
tflib.init_tf()
with dnnlib.util.open_url(network_pkl) as fp:
_G, _D, Gs = pickle.load(fp)
angleLatentPath = 'out/a27'
styleLatentPath = 'out/c3'
# display(Image2(filename='./' + angleLatentPath+'/proj.png'))
# display(Image2(filename='./' + styleLatentPath+'/proj.png'))
with np.load(angleLatentPath + '/dlatents.npz') as latent1:
with np.load(styleLatentPath + '/dlatents.npz') as latent2:
lat1 = latent1['dlatents']
lat2 = latent2['dlatents']
col_styles = [3,4,5,6,7,8,9,10,11,12,13]
lat1[0][col_styles] = lat2[0][col_styles]
image = Gs.components.synthesis.run(lat1, **Gs_syn_kwargs)[0]
mixImg = Image.fromarray(image, 'RGB')
showImages([Image.open('./' + angleLatentPath+'/proj.png'),Image.open('./' + styleLatentPath+'/proj.png') ,mixImg])
```
| github_jupyter |
```
import matplotlib.pyplot as plt
import numpy as np
from pylj import mc, comp, util, sample, pairwise
```
# Atomistic simulation
The use of computers in chemistry is becoming more common as computers are increasing in power and the algorithms are becoming more accurate and efficient. Computer simulation of atomic and molecular species takes two "flavours":
- **Quantum calculations**: where approximations are made to the Schrödinger equation to allow for its calculation for a multi-electron system (more information about this can be found in Atkins & Friedman's *Molecular Quantum Mechanics*),
- **Classical simulations**: where model interactions between atoms are developed to allow chemical systems to be simulated,
This exercise will focus on the latter.
## Classical simulation
One of the most popular models used to simulate interparticle interactions is known as the **Lennard-Jones** function. This models the London dispersion interactions between atoms. Hopefully the London dispersion interactions are familiar as consisting of:
- **van der Waals attraction**: the attraction between atoms that occurs as a result of the formation of instantenous dipole formation,
- **Pauli's exclusion principle**: the repulsion which stops atoms overlapping as no two electrons can have the same quantum state.
This means that the Lennard-Jones function is attractive when particles are close enough to induce dipole formation but **very** repulsive when the particles are too close. The Lennard-Jones function has the following form,
$$ E(r) = \frac{A}{r^{12}} - \frac{B}{r^6}, $$
where $E(r)$ is the potential energy at a given distance $r$, while $A$ and $B$ are parameters specific to the nature of the given interaction. In the cell below, write a function to calculate the energy of an interaction between two argon atoms for given range of distances using the Lennard-Jones function as the model.
```
def lennard_jones(A, B, r):
return ◽◽◽
```
Then use this function to plot the potential energy from $r=3$ Å to $r=8$ Å, and discuss the shape and sign of this function in terms of the attractive and repulsive regimes, when the values of ε and σ are as follows.
| A/$\times10^{-134}$Jm$^{12}$ | B/$\times10^{-78}$Jm$^6$ |
|:------:|:------:|
| 1.36 | 9.27 |
```
r = np.linspace( ◽◽◽, ◽◽◽, ◽◽◽ )
E = ◽◽◽
plt.plot( ◽◽◽, ◽◽◽ )
plt.xlabel( ◽◽◽ )
plt.ylabel( ◽◽◽ )
plt.show()
```
We are going to use the software pylj. This is a Python library, which means it is a collection of functions to enable the simulation by Monte Carlo. An example of a function in pylj is the `pairwise.lennard_jones_energy`, which can be used similar to the `lennard_jones` function that you have defined above.
```
r = np.linspace( 3e-10, 8e-10, 100 )
E = pairwise.lennard_jones_energy(1.36e-134, 9.27e-78, r)
plt.plot( r, E )
plt.xlabel( 'r/m' )
plt.ylabel( 'E/J' )
plt.show()
```
In addition to the functions associated directly with the Monte Carlo simulations, pylj is also useful for the visualisation of simulations (an example of a visualiation environment in pylj is shown below).
<img src="fig1.png" width="500px">
*Figure 1. The Energy sampling class in pylj.*
In this exercise, we will use pylj to help to build a Monte Carlo simulation where the total energy of the system is found using a Lennard-Jones potential.
## Monte Carlo
Monte Carlo or MC is a broad class of simulation method for determining equilibrium properties about a system. Monte Carlo is in many ways similar to [molecular dynamics](../molecular_dynamics/intro_to_molecular_dynamics.ipynb) however it makes use of statistical mechanics in lieu of molecular dynamics. Therefore, instead of dynamical information, the information generated is according to Boltzmann probabilities. The algorithm that is used in Monte Carlo simulations are as follows:
1. initialise the system,
2. calculate the energy of the system,
3. suggest some change to the system,
4. calculate the energy with this change,
5. if some condition is true, accept the change,
6. go to step 3.
This process continues for as long as the scientist is interested in.
### Initialisation
Lets try and use pylj to initialise our system, to do this we use the function `mc.initialise`. This function takes 4 inputs:
- number of particles,
- temperature of the simulation,
- size of the simulation cell,
- how the particles should be distributed.
The first line is to set up the visualisation environment within the Jupyter notebook, and the third line then sets and plots the particular enivornment that you want (in this case the `JustCell` environment).
```
%matplotlib notebook
simulation = mc.initialise(16, 300, 20, 'square')
sample_system = sample.JustCell(simulation)
```
In the above example, a `square` distribution of particles is assigned, however, it is also possible to distribute the particles randomly through the cell (with the keyword `random`), try this out in the cell above.
### Calculate energy
The next step is to find the energy of the system. This is achieved by summing the value for each interaction in the system. The example above has 16 atoms, the number of interactions in a *pairwise* method such as Monte Carlo is found with the following equation,
$$ \frac{(N - 1)N}{2}, $$
define a function to determine the number of interactions below.
```
def number_of_interactions( ◽◽◽ ):
◽◽◽
```
pylj has a function for the determination of the energy of each interaction, `pairwise.compute_energy`. This can be run as follows.
```
simulation.compute_energy()
old_energies = simulation.energies
print(old_energies)
```
If the particles are more than 15 Å (or half the box length if the box length is less than 30 Å), the energy is automatically assigned as 0, since the particles are considered to be far enough apart that there interaction is negligible.
### Make a change
Then a change is made to the system, in Monte Carlo generally this can be a wide variety of types of change, such as the number of particles, their position, and the volume of the system. We will focus on modifying the positions of the particles, which is achieved through the following pair of functions.
```
random_particle = simulation.select_random_particle()
simulation.new_random_position(random_particle)
```
If we then print the energies of the interactions again hopefully it is possible to see the difference.
```
simulation.compute_energy()
new_energies = simulation.energies
print(new_energies)
```
### Find the new energy
The total energy of the system in Monte Carlo is generally in the units of joules per atom (or molecule). Define below a function to calculate the total energy from the energy of the individual interactions.
```
def total_energy( ◽◽◽, ◽◽◽ ):
◽◽◽
```
### Comparing the energies
It is then possible to compare the energy of the system before and after the move.
```
if total_energy( new_energy, ◽◽◽ ) < total_energy( old_energy, ◽◽◽ ):
print('The energy has decreased!')
else:
print('The energy increased :(')
```
However, we are not just interested in if the energy has decreased as we want Boltzmann probabilities of the possible states. This requires the Metropolis condition, where the change is accepted if,
$$ n < \exp{\bigg(\frac{E_{\text{new}} - E_{\text{old}}}{k_BT}\bigg)}, $$
where $n\sim U(0, 1)$, $k_B$ is the Boltzmann constant, and $T$ is the temperature. This is implemented in pylj using the `mc.metropolis` function.
```
mc.metropolis(300, total_energy(old_energies, ◽◽◽ ), total_energy(new_energies, ◽◽◽ ))
```
This returns `True` if the move should be accepted and `False` if it should be rejected. Plot the new configuration of your system using the following cell and comment in whether the move was accepted or rejected (note that this updates the original plotting of the cell so you need to scroll up to observe the change).
```
sample_system.update(simulation)
```
### Accept and reject
pylj also has functions built in to accept and reject the moves that are proposed. These take the following forms,
```python
accept()
reject()
```
Using this knowledge, attempt to build you own Monte Carlo simulation loop, using the `update` to present the changes to the simulation visualisation (note that this is the slowest part of the computation so it is recommended that you don't perform this every iteration).
```
# Cell for building MC simulation.
```
| github_jupyter |
# What’s New In Python 3.10
> **See also:**
>
> * [What’s New In Python 3.10](https://docs.python.org/3.10/whatsnew/3.10.html)
```
import sys
assert sys.version_info[:2] >= (3, 10)
```
## Better error messages
### Syntax Errors
* When parsing code that contains unclosed parentheses or brackets the interpreter now includes the location of the unclosed bracket of parentheses instead of displaying `SyntaxError: unexpected EOF`.
* `SyntaxError` exceptions raised by the interpreter will now highlight the full error range of the expression that consistutes the syntax error itself, instead of just where the problem is detected.
* Specialised messages for `SyntaxError` exceptions have been added e.g. for
* missing `:` before blocks
* unparenthesised tuples in comprehensions targets
* missing commas in collection literals and between expressions
* missing `:` and values in dictionary literals
* usage of `=` instead of `==` in comparisons
* usage of `*` in f-strings
### Indentation Errors
* Many `IndentationError` exceptions now have more context.
### Attribute Errors
* `AttributeError` will offer suggestions of similar attribute names in the object that the exception was raised from.
### Name Errors
* `NameError` will offer suggestions of similar variable names in the function that the exception was raised from.
## Structural Pattern Matching
Many functional languages have a `match` expression, for example [Scala](https://www.scala-lang.org/files/archive/spec/2.11/08-pattern-matching.html), [Rust](https://doc.rust-lang.org/reference/expressions/match-expr.html), [F#](https://docs.microsoft.com/en-us/dotnet/fsharp/language-reference/pattern-matching).
A `match` statement takes an expression and compares it to successive patterns given as one or more case blocks. This is superficially similar to a switch statement in C, Java or JavaScript, but much more powerful.
### `match`
The simplest form compares a subject value against one or more literals:
```
def http_error(status):
match status:
case 400:
return "Bad request"
case 401:
return "Unauthorized"
case 403:
return "Forbidden"
case 404:
return "Not found"
case 418:
return "I'm a teapot"
case _:
return "Something else"
```
> **Note:**
>
> Only in this case `_` acts as a wildcard that never fails and **not** as a variable name.
The cases not only check for equality, but rebind variables that match the specified pattern. For example:
```
NOT_FOUND = 404
retcode = 200
match retcode:
case NOT_FOUND:
print('not found')
print(f"Current value of {NOT_FOUND=}")
```
> «If this poorly-designed feature is really added to Python, we lose a principle I’ve always taught students: ‹if you see an undocumented constant, you can always name it without changing the code’s meaning.› The Substitution Principle, learned in algebra? It’ll no longer apply.» – [Brandon Rhodes](https://twitter.com/brandon_rhodes/status/1360226108399099909)
> «… the semantics of this can be quite different from switch. The cases don't simply check equality, they rebind variables that match the specified pattern.» – [Jake VanderPlas](https://twitter.com/jakevdp/status/1359870794877132810)
### Symbolic constants
Patterns may use named constants. These must be dotted names to prevent them from being interpreted as capture variable:
```
from enum import Enum
class Color(Enum):
RED = 0
GREEN = 1
BLUE = 2
color = Color(2)
match color:
case color.RED:
print("I see red!")
case color.GREEN:
print("Grass is green")
case color.BLUE:
print("I'm feeling the blues :(")
```
> «… "case CONSTANT" actually matching everything and assigning to a variable named CONSTANT» – [Armin Ronacher](https://twitter.com/mitsuhiko/status/1359263136994516999)
> **See also:**
>
> * [Structural pattern matching for Python](https://lwn.net/Articles/827179/)
> * [PEP 622 – Structural Pattern Matching](https://www.python.org/dev/peps/pep-0622) superseded by
> * [PEP 634: Specification](https://www.python.org/dev/peps/pep-0634)
> * [PEP 635: Motivation and Rationale](https://www.python.org/dev/peps/pep-0635)
> * [PEP 636: Tutorial](https://www.python.org/dev/peps/pep-0636)
> * [github.com/gvanrossum/patma/](https://github.com/gvanrossum/patma/)
> * [playground-622.ipynb on binder](https://mybinder.org/v2/gh/gvanrossum/patma/master?urlpath=lab/tree/playground-622.ipynb)
> * [Tobias Kohn: On the Syntax of Pattern Matching in Python](https://tobiaskohn.ch/index.php/2018/09/18/pattern-matching-syntax-in-python/)
| github_jupyter |
# Import the Dataset
```
import pandas as pd
messages = pd.read_csv('SMSSpamCollection', sep='\t',
names=["label", "message"])
```
# Importing Dependencies
```
%matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
import re
import nltk
import numpy as np
nltk.download('stopwords')
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.metrics import accuracy_score
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
ps = PorterStemmer()
```
# Data preprocessing
```
messages.head()
messages.isnull().sum()
len(messages)
```
# Histogram visualisation for ham and spam
```
plt.figure(figsize=(8,5))
sns.countplot(x='label', data=messages)
plt.xlabel('message')
plt.ylabel('Count')
plt.show()
```
# Data Cleaning
```
corpus = []
for i in range(0, len(messages)):
review = re.sub('[^a-zA-Z]', ' ', messages['message'][i])
review = review.lower()
review = review.split()
review = [ps.stem(word) for word in review if not word in stopwords.words('english')]
review = ' '.join(review)
corpus.append(review)
```
# Creating the Bag of Words model
```
cv = CountVectorizer(max_features=2500)
X = cv.fit_transform(corpus).toarray()
```
# Extracting dependent variable from the dataset
```
y = pd.get_dummies(messages['label'])
y = y.iloc[:, 1].values
y
```
# Train Test Split
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20, random_state=0)
```
# Accuracy
```
best_accuracy = 0.0
alpha_val = 0.0
for i in np.arange(0.0,1.1,0.1):
temp_classifier = MultinomialNB(alpha=i)
temp_classifier.fit(X_train, y_train)
temp_y_pred = temp_classifier.predict(X_test)
score = accuracy_score(y_test, temp_y_pred)
print("Accuracy score for alpha={} is: {}%".format(round(i,1), round(score*100,2)))
if score>best_accuracy:
best_accuracy = score
alpha_val = i
print('***************************************************')
print('The best accuracy is {}% with alpha value as {}'.format(round(best_accuracy*100, 2), round(alpha_val,1)))
acc_s = accuracy_score(y_test, y_pred)*100
print("Accuracy Score {} %".format(round(acc_s,2)))
```
# Naive Bayes
```
classifier = MultinomialNB(alpha=0.1)
classifier.fit(X_train, y_train)
y_pred = classifier.predict(X_test)
y_pred
```
# Prediction
```
def predict_spam(sample_message):
sample_message = re.sub(pattern='[^a-zA-Z]',repl=' ', string = sample_message)
sample_message = sample_message.lower()
sample_message_words = sample_message.split()
sample_message_words = [word for word in sample_message_words if not word in set(stopwords.words('english'))]
ps = PorterStemmer()
final_message = [ps.stem(word) for word in sample_message_words]
final_message = ' '.join(final_message)
temp = cv.transform([final_message]).toarray()
return classifier.predict(temp)
result = ['SPAM!','HAM']
msg = "Hi! kindly submit your assignment"
if predict_spam(msg):
print(result[0])
else:
print(result[1])
msg = 'Click to invest now:'
if predict_spam(msg):
print(result[0])
else:
print(result[1])
```
| github_jupyter |
<center>
<img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/IDSNlogo.png" width="300" alt="cognitiveclass.ai logo" />
</center>
# 2D Numpy in Python
Estimated time needed: **20** minutes
## Objectives
After completing this lab you will be able to:
* Operate comfortably with `numpy`
* Perform complex operations with `numpy`
<h2>Table of Contents</h2>
<div class="alert alert-block alert-info" style="margin-top: 20px">
<ul>
<li><a href="create">Create a 2D Numpy Array</a></li>
<li><a href="access">Accessing different elements of a Numpy Array</a></li>
<li><a href="op">Basic Operations</a></li>
</ul>
</div>
<hr>
<h2 id="create">Create a 2D Numpy Array</h2>
```
# Import the libraries
import numpy as np
import matplotlib.pyplot as plt
```
Consider the list <code>a</code>, which contains three nested lists **each of equal size**.
```
# Create a list
a = [[11, 12, 13], [21, 22, 23], [31, 32, 33]]
a
```
We can cast the list to a Numpy Array as follows:
```
# Convert list to Numpy Array
# Every element is the same type
A = np.array(a)
A
```
We can use the attribute <code>ndim</code> to obtain the number of axes or dimensions, referred to as the rank.
```
# Show the numpy array dimensions
A.ndim
```
Attribute <code>shape</code> returns a tuple corresponding to the size or number of each dimension.
```
# Show the numpy array shape
A.shape
```
The total number of elements in the array is given by the attribute <code>size</code>.
```
# Show the numpy array size
A.size
```
<hr>
<h2 id="access">Accessing different elements of a Numpy Array</h2>
We can use rectangular brackets to access the different elements of the array. The correspondence between the rectangular brackets and the list and the rectangular representation is shown in the following figure for a 3x3 array:
<img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%205/images/NumTwoEg.png" width="500" />
We can access the 2nd-row, 3rd column as shown in the following figure:
<img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%205/images/NumTwoFT.png" width="400" />
We simply use the square brackets and the indices corresponding to the element we would like:
```
# Access the element on the second row and third column
A[1, 2]
```
We can also use the following notation to obtain the elements:
```
# Access the element on the second row and third column
A[1][2]
```
Consider the elements shown in the following figure
<img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%205/images/NumTwoFF.png" width="400" />
We can access the element as follows:
```
# Access the element on the first row and first column
A[0][0]
```
We can also use slicing in numpy arrays. Consider the following figure. We would like to obtain the first two columns in the first row
<img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%205/images/NumTwoFSF.png" width="400" />
This can be done with the following syntax:
```
# Access the element on the first row and first and second columns
A[0][0:2]
```
Similarly, we can obtain the first two rows of the 3rd column as follows:
```
# Access the element on the first and second rows and third column
A[0:2, 2]
```
Corresponding to the following figure:
<img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%205/images/2D_numpy.png" width="550"><br />
<h2 id="op">Basic Operations</h2>
We can also add arrays. The process is identical to matrix addition. Matrix addition of <code>X</code> and <code>Y</code> is shown in the following figure:
<img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%205/images/NumTwoAdd.png" width="500" />
The numpy array is given by <code>X</code> and <code>Y</code>
```
# Create a numpy array X
X = np.array([[1, 0], [0, 1]])
X
# Create a numpy array Y
Y = np.array([[2, 1], [1, 2]])
Y
```
We can add the numpy arrays as follows.
```
# Add X and Y
Z = X + Y
Z
```
Multiplying a numpy array by a scaler is identical to multiplying a matrix by a scaler. If we multiply the matrix <code>Y</code> by the scaler 2, we simply multiply every element in the matrix by 2, as shown in the figure.
<img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%205/images/NumTwoDb.png" width="500" />
We can perform the same operation in numpy as follows
```
# Create a numpy array Y
Y = np.array([[2, 1], [1, 2]])
Y
# Multiply Y with 2
Z = 2 * Y
Z
```
Multiplication of two arrays corresponds to an element-wise product or <em>Hadamard product</em>. Consider matrix <code>X</code> and <code>Y</code>. The Hadamard product corresponds to multiplying each of the elements in the same position, i.e. multiplying elements contained in the same color boxes together. The result is a new matrix that is the same size as matrix <code>Y</code> or <code>X</code>, as shown in the following figure.
<img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-PY0101EN-SkillsNetwork/labs/Module%205/images/NumTwoMul.png" width="500" />
We can perform element-wise product of the array <code>X</code> and <code>Y</code> as follows:
```
# Create a numpy array Y
Y = np.array([[2, 1], [1, 2]])
Y
# Create a numpy array X
X = np.array([[1, 0], [0, 1]])
X
# Multiply X with Y
Z = X * Y
Z
```
We can also perform matrix multiplication with the numpy arrays <code>A</code> and <code>B</code> as follows:
First, we define matrix <code>A</code> and <code>B</code>:
```
# Create a matrix A
A = np.array([[0, 1, 1], [1, 0, 1]])
A
# Create a matrix B
B = np.array([[1, 1], [1, 1], [-1, 1]])
B
```
We use the numpy function <code>dot</code> to multiply the arrays together.
```
# Calculate the dot product
Z = np.dot(A,B)
Z
# Calculate the sine of Z
np.sin(Z)
```
We use the numpy attribute <code>T</code> to calculate the transposed matrix
```
# Create a matrix C
C = np.array([[1,1],[2,2],[3,3]])
C
# Get the transposed of C
C.T
```
<h2>Quiz on 2D Numpy Array</h2>
Consider the following list <code>a</code>, convert it to Numpy Array.
```
# Write your code below and press Shift+Enter to execute
a = [[1, 2, 3, 4], [5, 6, 7, 8], [9, 10, 11, 12]]
A = np.array(a)
A
```
<details><summary>Click here for the solution</summary>
```python
A = np.array(a)
A
```
</details>
Calculate the numpy array size.
```
# Write your code below and press Shift+Enter to execute
A.size
```
<details><summary>Click here for the solution</summary>
```python
A.size
```
</details>
Access the element on the first row and first and second columns.
```
# Write your code below and press Shift+Enter to execute
A[0][1]
```
<details><summary>Click here for the solution</summary>
```python
A[0][0:2]
```
</details>
Perform matrix multiplication with the numpy arrays <code>A</code> and <code>B</code>.
```
# Write your code below and press Shift+Enter to execute
B = np.array([[0, 1], [1, 0], [1, 1], [-1, 0]])
C = np.dot(A,B)
C
```
<details><summary>Click here for the solution</summary>
```python
X = np.dot(A,B)
X
```
</details>
<hr>
<h2>The last exercise!</h2>
<p>Congratulations, you have completed your first lesson and hands-on lab in Python. However, there is one more thing you need to do. The Data Science community encourages sharing work. The best way to share and showcase your work is to share it on GitHub. By sharing your notebook on GitHub you are not only building your reputation with fellow data scientists, but you can also show it off when applying for a job. Even though this was your first piece of work, it is never too early to start building good habits. So, please read and follow <a href="https://cognitiveclass.ai/blog/data-scientists-stand-out-by-sharing-your-notebooks/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01" target="_blank">this article</a> to learn how to share your work.
<hr>
## Author
<a href="https://www.linkedin.com/in/joseph-s-50398b136/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkPY0101ENSkillsNetwork19487395-2021-01-01" target="_blank">Joseph Santarcangelo</a>
## Other contributors
<a href="www.linkedin.com/in/jiahui-mavis-zhou-a4537814a">Mavis Zhou</a>
## Change Log
| Date (YYYY-MM-DD) | Version | Changed By | Change Description |
|---|---|---|---|
| 2021-01-05 | 2.2 | Malika | Updated the solution for dot multiplication |
| 2020-09-09 | 2.1 | Malika | Updated the screenshot for first two rows of the 3rd column |
| 2020-08-26 | 2.0 | Lavanya | Moved lab to course repo in GitLab |
| | | | |
| | | | |
<hr/>
## <h3 align="center"> © IBM Corporation 2020. All rights reserved. <h3/>
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Eager Execution
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/guide/eager"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/eager.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/eager.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
Note: これらのドキュメントは私たちTensorFlowコミュニティが翻訳したものです。コミュニティによる 翻訳は**ベストエフォート**であるため、この翻訳が正確であることや[英語の公式ドキュメント](https://www.tensorflow.org/?hl=en)の 最新の状態を反映したものであることを保証することはできません。 この翻訳の品質を向上させるためのご意見をお持ちの方は、GitHubリポジトリ[tensorflow/docs](https://github.com/tensorflow/docs)にプルリクエストをお送りください。 コミュニティによる翻訳やレビューに参加していただける方は、 [docs-ja@tensorflow.org メーリングリスト](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ja)にご連絡ください。
TensorflowのEager Executionは、計算グラフの作成と評価を同時におこなう命令的なプログラミングを行うための環境です:
オペレーションはあとで実行するための計算グラフでなく、具体的な計算結果の値を返します。
この方法を用いることにより、初心者にとってTensorFlowを始めやすくなり、またモデルのデバッグも行いやすくなります。
さらにコードの記述量も削減されます。
このガイドの内容を実行するためには、対話的インタープリタ`python`を起動し、以下のコードサンプルを実行してください。
Eager Executionは研究や実験のための柔軟な機械学習環境として、以下を提供します。
* *直感的なインタフェース*—Pythonのデータ構造を使用して、コードをナチュラルに記述することができます。スモールなモデルとデータに対してすばやく実験を繰り返すことができます。
* *より簡単なデバッグ*—opsを直接呼び出すことで、実行中のモデルを調査したり、変更をテストすることができます。Python標準のデバッグツールを用いて即座にエラーのレポーティングができます。
* *自然な制御フロー*—TensorFlowのグラフ制御フローの代わりにPythonの制御フローを利用するため、動的なモデルのパラメータ変更をシンプルに行うことができます。
Eager ExecutionはTensorflowのほとんどのオペレーションとGPUアクセラレーションをサポートします。
Eager Executionの実行例については、以下を参照してください。
[tensorflow/contrib/eager/python/examples](https://github.com/tensorflow/tensorflow/tree/master/tensorflow/contrib/eager/python/examples).
Note: いくつかのモデルはEager Executionを有効化することでオーバヘッドが増える可能性があります。
パフォーマンス改善を行っていますが、もしも問題を発見したら、バグ報告してベンチマークを共有してください。
## セットアップと基本的な使い方
Eager Executionをはじめるためには、プログラムやコンソールセッションの最初に、`tf.enable_eager_execution()`を追加してください。
プログラムが呼び出すほかのモジュールにこのオペレーションを追加しないでください。
```
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow as tf
tf.enable_eager_execution()
```
これでTensorFlowのオペレーションを実行してみましょう。結果はすぐに返されます。
```
tf.executing_eagerly()
x = [[2.]]
m = tf.matmul(x, x)
print("hello, {}".format(m))
```
Eager Executionを有効化することで、TensorFlowの挙動は変わります—TensorFlowは即座に式を評価して結果をPythonに返すようになります。
`tf.Tensor` オブジェクトは計算グラフのノードへのシンボリックハンドルの代わりに具体的な値を参照します。
セッションの中で構築して実行する計算グラフが存在しないため、`print()`やデバッガを使って容易に結果を調べることができます。
勾配計算を終了することなくテンソル値を評価、出力、およびチェックすることができます。
Eager Executionは、[NumPy](http://www.numpy.org/)と一緒に使うことができます。
NumPyのオペレーションは、`tf.Tensor`を引数として受け取ることができます。
TensorFlow [math operations](https://www.tensorflow.org/api_guides/python/math_ops) はPythonオブジェクトとNumpy arrayを`tf.Tensor`にコンバートします。
`tf.Tensor.numpy`メソッドはオブジェクトの値をNumPyの`ndarray`形式で返します。
```
a = tf.constant([[1, 2],
[3, 4]])
print(a)
# ブロードキャストのサポート
b = tf.add(a, 1)
print(b)
# オペレータのオーバーロードがサポートされている
print(a * b)
# NumPy valueの使用
import numpy as np
c = np.multiply(a, b)
print(c)
# Tensorからnumpyの値を得る
print(a.numpy())
# => [[1 2]
# [3 4]]
```
`tf.contrib.eager` モジュールは、Eager ExecutionとGraph Executionの両方の環境で利用可能なシンボルが含まれており、[Graph Execution](#work_with_graphs)方式での記述に便利です:
```
tfe = tf.contrib.eager
```
## 動的な制御フロー
Eager Executionの主要なメリットは、モデルを実行する際にホスト言語のすべての機能性が利用できることです。
たとえば、[fizzbuzz](https://en.wikipedia.org/wiki/Fizz_buzz)が簡単に書けます:
```
def fizzbuzz(max_num):
counter = tf.constant(0)
max_num = tf.convert_to_tensor(max_num)
for num in range(1, max_num.numpy()+1):
num = tf.constant(num)
if int(num % 3) == 0 and int(num % 5) == 0:
print('FizzBuzz')
elif int(num % 3) == 0:
print('Fizz')
elif int(num % 5) == 0:
print('Buzz')
else:
print(num.numpy())
counter += 1
fizzbuzz(15)
```
この関数はテンソル値に依存する条件式を持ち、実行時にこれらの値を表示します。
## モデルの構築
多くの機械学習モデルはレイヤーを積み重ねによって成り立っています。Eager ExecutionでTensorFlowを使うときは、自分でレイヤーの内容を記述してもいいし、もしくは `tf.keras.layers`パッケージで提供されるレイヤーを使うこともできます。
レイヤーを表現するためには任意のPythonオブジェクトを使用できますが、
TensorFlowには便利な基本クラスとして `tf.keras.layers.Layer`があります。
このクラスを継承した独自のレイヤーを実装してみます:
```
class MySimpleLayer(tf.keras.layers.Layer):
def __init__(self, output_units):
super(MySimpleLayer, self).__init__()
self.output_units = output_units
def build(self, input_shape):
# buildメソッドは、レイヤーが初めて使われたときに呼ばれます
# build()で変数を作成すると、それらのshapeを入力のshapeに依存させることができ、
# ユーザがshapeを完全に指定する必要はありません。
# 既に完全なshapeが決まっている場合は、__init__()の中で変数を作成することもできます。
self.kernel = self.add_variable(
"kernel", [input_shape[-1], self.output_units])
def call(self, input):
# __call__の代わりにcall()を上書きします。
return tf.matmul(input, self.kernel)
```
`MySimpleLayer`の代わりに、その機能のスーパーセットを持っている`tf.keras.layers.Dense`レイヤーを使用してください
(このレイヤーはバイアスを加えることもできるもできます)。
レイヤーをモデルに組み立てるとき、レイヤーの線形スタックである
モデルを表すために `tf.keras.Sequential`を使うことができます。この書き方は基本的なモデルを扱いやすいです。
```
model = tf.keras.Sequential([
tf.keras.layers.Dense(10, input_shape=(784,)), # 入力のshapeを指定する必要がある
tf.keras.layers.Dense(10)
])
```
もしくは、 `tf.keras.Model`を継承してモデルをクラスにまとめます。
これはレイヤー自身であるレイヤーのコンテナで、 `tf.keras.Model`オブジェクトが他の` tf.keras.Model`オブジェクトを含むことを可能にします。
Alternatively, organize models in classes by inheriting from `tf.keras.Model`.
This is a container for layers that is a layer itself, allowing `tf.keras.Model`
objects to contain other `tf.keras.Model` objects.
```
class MNISTModel(tf.keras.Model):
def __init__(self):
super(MNISTModel, self).__init__()
self.dense1 = tf.keras.layers.Dense(units=10)
self.dense2 = tf.keras.layers.Dense(units=10)
def call(self, input):
"""Run the model."""
result = self.dense1(input)
result = self.dense2(result)
result = self.dense2(result) # dense2レイヤーを再利用します reuse variables from dense2 layer
return result
model = MNISTModel()
```
入力のshapeは最初のレイヤーに初めて入力データを渡すときにセットされるため、
モデル構築時に`tf.keras.Model`クラスに設定する必要はありません。
`tf.keras.layers`クラスは独自のモデル変数を作成し、包含します。このモデル変数は、それを含むレイヤーオブジェクトのライフタイムにひもづきます。レイヤー変数を共有するには、それらのオブジェクトを共有します。
## Eager Executionにおける学習
### 勾配の計算
[自動微分](https://en.wikipedia.org/wiki/Automatic_differentiation)はニューラルネットワークの学習で利用される[バックプロパゲーション](https://en.wikipedia.org/wiki/Backpropagation)などの機械学習アルゴリズムの実装を行う上で便利です。
Eager Executionでは、勾配計算をあとで行うためのオペレーションをトレースするために`tf.GradientTape` を利用します。
`tf.GradientTape` はトレースしない場合に最大のパフォーマンスを提供するオプトイン機能です。各呼び出し中に異なるオペレーションが発生する可能性があるため、すべてのforward-passオペレーションは一つの「テープ」に記録されます。勾配を計算するには、テープを逆方向に再生してから破棄します。特定の `tf.GradientTape`は一つのグラデーションしか計算できません。後続の呼び出しは実行時エラーをスローします。
```
w = tf.Variable([[1.0]])
with tf.GradientTape() as tape:
loss = w * w
grad = tape.gradient(loss, w)
print(grad) # => tf.Tensor([[ 2.]], shape=(1, 1), dtype=float32)
```
### モデル学習
以下のexampleはMNISTという手書き数字分類を行うマルチレイヤーモデルを作成します。
Eager Execution環境における学習可能なグラフを構築するためのオプティマイザーとレイヤーAPIを提示します。
```
# mnistデータのを取得し、フォーマットする
(mnist_images, mnist_labels), _ = tf.keras.datasets.mnist.load_data()
dataset = tf.data.Dataset.from_tensor_slices(
(tf.cast(mnist_images[...,tf.newaxis]/255, tf.float32),
tf.cast(mnist_labels,tf.int64)))
dataset = dataset.shuffle(1000).batch(32)
# モデルを構築する
mnist_model = tf.keras.Sequential([
tf.keras.layers.Conv2D(16,[3,3], activation='relu'),
tf.keras.layers.Conv2D(16,[3,3], activation='relu'),
tf.keras.layers.GlobalAveragePooling2D(),
tf.keras.layers.Dense(10)
])
```
学習を行わずとも、モデルを呼び出して、Eager Executionにより、出力を検査することができます:
```
for images,labels in dataset.take(1):
print("Logits: ", mnist_model(images[0:1]).numpy())
```
kerasモデルは組み込みで学習のループを回すメソッド`fit`がありますが、よりカスタマイズが必要な場合もあるでしょう。 Eager Executionを用いて実装された学習ループのサンプルを以下に示します:
```
optimizer = tf.train.AdamOptimizer()
loss_history = []
for (batch, (images, labels)) in enumerate(dataset.take(400)):
if batch % 10 == 0:
print('.', end='')
with tf.GradientTape() as tape:
logits = mnist_model(images, training=True)
loss_value = tf.losses.sparse_softmax_cross_entropy(labels, logits)
loss_history.append(loss_value.numpy())
grads = tape.gradient(loss_value, mnist_model.trainable_variables)
optimizer.apply_gradients(zip(grads, mnist_model.trainable_variables),
global_step=tf.train.get_or_create_global_step())
import matplotlib.pyplot as plt
plt.plot(loss_history)
plt.xlabel('Batch #')
plt.ylabel('Loss [entropy]')
```
### 値とオプティマイザ
`tf.Variable` オブジェクトは、学習中にアクセスされるミュータブルな`tf.Tensor`値を格納し、自動微分を容易にします。
モデルのパラメータは、変数としてクラスにカプセル化できます。
`tf.GradientTape`と共に` tf.Variable`を使うことでモデルパラメータはよりカプセル化されます。たとえば、上の
の自動微分の例は以下のように書き換えることができます:
```
class Model(tf.keras.Model):
def __init__(self):
super(Model, self).__init__()
self.W = tf.Variable(5., name='weight')
self.B = tf.Variable(10., name='bias')
def call(self, inputs):
return inputs * self.W + self.B
# 3 * 2 + 2を近似するトイデータセット
NUM_EXAMPLES = 2000
training_inputs = tf.random_normal([NUM_EXAMPLES])
noise = tf.random_normal([NUM_EXAMPLES])
training_outputs = training_inputs * 3 + 2 + noise
# オプティマイズ対象のloss関数
def loss(model, inputs, targets):
error = model(inputs) - targets
return tf.reduce_mean(tf.square(error))
def grad(model, inputs, targets):
with tf.GradientTape() as tape:
loss_value = loss(model, inputs, targets)
return tape.gradient(loss_value, [model.W, model.B])
# 定義:
# 1. モデル
# 2. モデルパラメータに関する損失関数の導関数
# 3. 導関数に基づいて変数を更新するストラテジ。
model = Model()
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.01)
print("Initial loss: {:.3f}".format(loss(model, training_inputs, training_outputs)))
# 学習ループ
for i in range(300):
grads = grad(model, training_inputs, training_outputs)
optimizer.apply_gradients(zip(grads, [model.W, model.B]),
global_step=tf.train.get_or_create_global_step())
if i % 20 == 0:
print("Loss at step {:03d}: {:.3f}".format(i, loss(model, training_inputs, training_outputs)))
print("Final loss: {:.3f}".format(loss(model, training_inputs, training_outputs)))
print("W = {}, B = {}".format(model.W.numpy(), model.B.numpy()))
```
## Eager Executionの途中でオブジェクトのステータスを使用する
Graph Executionでは、プログラムの状態(変数など)はglobal collectionに格納され、それらの存続期間は `tf.Session`オブジェクトによって管理されます。
対照的に、Eager Executionの間、状態オブジェクトの存続期間は、対応するPythonオブジェクトの存続期間によって決定されます。
### 変数とオブジェクト
Eager Executionの間、変数はオブジェクトへの最後の参照が削除され、その後削除されるまで存続します。
```
if tf.test.is_gpu_available():
with tf.device("gpu:0"):
v = tf.Variable(tf.random_normal([1000, 1000]))
v = None # vは既にGPUメモリ上を使用しないようにする
```
### オブジェクトベースの保存
`tf.train.Checkpoint`はチェックポイントを用いて`tf.Variable`を保存および復元することができます:
```
x = tf.Variable(10.)
checkpoint = tf.train.Checkpoint(x=x)
x.assign(2.) # 変数に新しい値を割り当てて保存する
checkpoint_path = './ckpt/'
checkpoint.save('./ckpt/')
x.assign(11.) # 保存後に変数の値を変更する
# チェックポイントから値を復元する
checkpoint.restore(tf.train.latest_checkpoint(checkpoint_path))
print(x) # => 2.0
```
モデルを保存して読み込むために、 `tf.train.Checkpoint`は隠れ変数なしにオブジェクトの内部状態を保存します。 `モデル`、 `オプティマイザ`、そしてグローバルステップの状態を記録するには、それらを `tf.train.Checkpoint`に渡します。
```
import os
import tempfile
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(16,[3,3], activation='relu'),
tf.keras.layers.GlobalAveragePooling2D(),
tf.keras.layers.Dense(10)
])
optimizer = tf.train.AdamOptimizer(learning_rate=0.001)
checkpoint_dir = tempfile.mkdtemp()
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
root = tf.train.Checkpoint(optimizer=optimizer,
model=model,
optimizer_step=tf.train.get_or_create_global_step())
root.save(checkpoint_prefix)
root.restore(tf.train.latest_checkpoint(checkpoint_dir))
```
### オブジェクト指向メトリクス
`tfe.metrics`はオブジェクトとして保存されます。新しいデータを呼び出し可能オブジェクトに渡してメトリクスを更新し、 `tfe.metrics.result`メソッドを使って結果を取得します。次に例を示します:
```
m = tfe.metrics.Mean("loss")
m(0)
m(5)
m.result() # => 2.5
m([8, 9])
m.result() # => 5.5
```
#### サマリとTensorBoard
[TensorBoard](../guide/summaries_and_tensorboard.md) はモデルの学習プロセスを理解、デバッグ、最適化するための可視化ツールです。プログラムの実行中に書き込まれるサマリイベントを使用します。
`tf.contrib.summary`はEager ExecutionとGraph Executionの両方の環境と互換性があります。
`tf.contrib.summary.scalar`のようなサマリオペレーションはモデル構築の間に挿入されます。
たとえば、100のグローバルステップごとにサマリを記録するには、次のようにします。
```
global_step = tf.train.get_or_create_global_step()
logdir = "./tb/"
writer = tf.contrib.summary.create_file_writer(logdir)
writer.set_as_default()
for _ in range(10):
global_step.assign_add(1)
# record_summariesメソッドをincludeする必要がある
with tf.contrib.summary.record_summaries_every_n_global_steps(100):
# ここにモデルのコードを記述する
tf.contrib.summary.scalar('global_step', global_step)
!ls tb/
```
## 高度な自動分類トピック
### 動的なモデル
`tf.GradientTape`は動的モデルでも使うことができます。
以下の[バックトラックライン検索](https://wikipedia.org/wiki/Backtracking_line_search)
アルゴリズムの例は、複雑な制御フローにも関わらず
勾配があり、微分可能であることを除いて、通常のNumPyコードのように見えます:
```
def line_search_step(fn, init_x, rate=1.0):
with tf.GradientTape() as tape:
# 変数は自動的に記録されるが、手動でTensorを監視する
tape.watch(init_x)
value = fn(init_x)
grad = tape.gradient(value, init_x)
grad_norm = tf.reduce_sum(grad * grad)
init_value = value
while value > init_value - rate * grad_norm:
x = init_x - rate * grad
value = fn(x)
rate /= 2.0
return x, value
```
### 勾配計算のための追加機能
`tf.GradientTape`は強力な勾配計算インタフェースですが、
自動微分に利用できる別の[Autograd](https://github.com/HIPS/autograd)スタイルのAPIもあります。
これらの関数はテンソルと勾配関数のみを使って、`tf.variables`を使わずに数式コードを書く場合に便利です:
* `tfe.gradients_function`—引数をとり、入力関数パラメータの導関数を計算する関数を返します。
入力パラメータはスカラ値を返さなければなりません。返された関数が
されると、 `tf.Tensor`オブジェクトのリストを返します:入力関数のそれぞれの
引数に対して一つの要素。重要なものすべてを関数パラメータとして渡さなければならないので、
多くのtrainableパラメータに依存している場合、これは扱いにくくなります。
* `tfe.value_and_gradients_function`—` tfe.gradients_function`に似ていますが、
返された関数が呼び出されると、その引数に関する入力関数の導関数のリストに加えて、入力関数からの値を返します。
次の例では、 `tfe.gradients_function`は引数として` square` 関数を取り、その入力に関して `square`の偏微分
導関数を計算する関数を返します。 `3`における` square`の微分を計算するために、 `grad(3.0)`は `6`を返します。
```
def square(x):
return tf.multiply(x, x)
grad = tfe.gradients_function(square)
square(3.).numpy()
grad(3.)[0].numpy()
# 平方の二次導関数:
gradgrad = tfe.gradients_function(lambda x: grad(x)[0])
gradgrad(3.)[0].numpy()
# 3次導関数はNoneになる:
gradgradgrad = tfe.gradients_function(lambda x: gradgrad(x)[0])
gradgradgrad(3.)
# フロー制御:
def abs(x):
return x if x > 0. else -x
grad = tfe.gradients_function(abs)
grad(3.)[0].numpy()
grad(-3.)[0].numpy()
```
### カスタム勾配
カスタム勾配は、Eager ExecutionとGraph Executionの両方の環境で、勾配を上書きする簡単な方法です。 フォワード関数では、
入力、出力、または中間結果に関する勾配を定義します。たとえば、逆方向パスにおいて勾配のノルムを切り取る簡単な方法は次のとおりです:
```
@tf.custom_gradient
def clip_gradient_by_norm(x, norm):
y = tf.identity(x)
def grad_fn(dresult):
return [tf.clip_by_norm(dresult, norm), None]
return y, grad_fn
```
カスタム勾配は、一連の演算に対して数値的に安定した勾配を提供するために共通的に使用されます。
```
def log1pexp(x):
return tf.log(1 + tf.exp(x))
grad_log1pexp = tfe.gradients_function(log1pexp)
# 勾配計算はx = 0のときにはうまくいきます。
grad_log1pexp(0.)[0].numpy()
# しかし、x = 100のときは数値的不安定により失敗します。
grad_log1pexp(100.)[0].numpy()
```
ここで、 `log1pexp`関数はカスタム勾配を用いて解析的に単純化することができます。
以下の実装は、フォワードパスの間に計算された `tf.exp(x)`の値を
再利用します—冗長な計算を排除することでより効率的になります:
```
@tf.custom_gradient
def log1pexp(x):
e = tf.exp(x)
def grad(dy):
return dy * (1 - 1 / (1 + e))
return tf.log(1 + e), grad
grad_log1pexp = tfe.gradients_function(log1pexp)
# 上と同様に、勾配計算はx = 0のときにはうまくいきます。
grad_log1pexp(0.)[0].numpy()
# また、勾配計算はx = 100でも機能します。
grad_log1pexp(100.)[0].numpy()
```
## パフォーマンス
Eager Executionの間、計算は自動的にGPUにオフロードされます。計算を実行するデバイスを指定したい場合は、
`tf.device( '/ gpu:0')`ブロック(もしくはCPUを指定するブロック)で囲むことで指定できます:
```
import time
def measure(x, steps):
# TensorFlowはGPUを初めて使用するときに初期化するため、時間計測対象からは除外する。
tf.matmul(x, x)
start = time.time()
for i in range(steps):
x = tf.matmul(x, x)
# tf.matmulは、行列乗算が完了する前に戻ることができます。
# (たとえば、CUDAストリームにオペレーションをエンキューした後に戻すことができます)。
# 以下のx.numpy()呼び出しは、すべてのキューに入れられたオペレーションが完了したことを確認します。
# (そして結果をホストメモリにコピーするため、計算時間は単純なmatmulオペレーションよりも多くのことを含む時間になります。)
_ = x.numpy()
end = time.time()
return end - start
shape = (1000, 1000)
steps = 200
print("Time to multiply a {} matrix by itself {} times:".format(shape, steps))
# CPU上で実行するとき:
with tf.device("/cpu:0"):
print("CPU: {} secs".format(measure(tf.random_normal(shape), steps)))
# GPU上で実行するとき(GPUが利用できれば):
if tfe.num_gpus() > 0:
with tf.device("/gpu:0"):
print("GPU: {} secs".format(measure(tf.random_normal(shape), steps)))
else:
print("GPU: not found")
```
`tf.Tensor`オブジェクトはそのオブジェクトに対するオペレーションを実行するために別のデバイスにコピーすることができます:
```
if tf.test.is_gpu_available():
x = tf.random_normal([10, 10])
x_gpu0 = x.gpu()
x_cpu = x.cpu()
_ = tf.matmul(x_cpu, x_cpu) # CPU上で実行するとき
_ = tf.matmul(x_gpu0, x_gpu0) # GPU:0上で実行するとき
if tfe.num_gpus() > 1:
x_gpu1 = x.gpu(1)
_ = tf.matmul(x_gpu1, x_gpu1) # GPU:1で実行するとき
```
### ベンチマーク
GPUでの
[ResNet50](https://github.com/tensorflow/tensorflow/tree/master/tensorflow/contrib/eager/python/examples/resnet50)
の学習のような、計算量の多いモデルの場合は、Eager ExecutionのパフォーマンスはGraph Executionのパフォーマンスに匹敵します。
しかし、この2つの環境下のパフォーマンスの違いは計算量の少ないモデルではより大きくなり、小さなたくさんのオペレーションからなるモデルでホットコードパスを最適化するためにやるべきことがあります。
## Graph Executionの実行
Eager Executionは開発とデバッグをより対話的にしますが、
TensorFlowのGraph Executionは分散学習、パフォーマンスの最適化、そしてプロダクション環境へのデプロイの観点で利点があります。
しかし、Graph Executionのコードの記述方法、標準的なのPythonコードの書き方と異なり、デバッグがより難しく感じるかもしれません。
Graph Execution形式のモデルの構築と学習のために、Pythonプログラムは最初に計算グラフを構築し、
それからC++ベースのランタイムで実行するために`Session.run`を呼び出し、グラフを渡します。この機能の特徴は以下のとおりです:
* 静的なautodiffによる自動微分
* プラットフォームに依存しないサーバーへの簡単なデプロイ
* グラフベースの最適化(共通的な部分式の削除、定数の畳み込みなど)
* コンパイルとカーネルフュージョン
* 自動分散とレプリケーション(分散システムへのノード配置)
Eager Executionのコードは、Graph Executionのコードよりもデプロイが難しいです:モデルから
計算グラフを生成するか、またはサーバ上で直接Pythonランタイムからコードを実行する必要があります。
### 互換性のあるコードの記述
Eager Execution環境で記述されたコードは、Eager Executionが有効になっていない新しいPythonセッションでおなじコードを実行するだけで
おなじコードのままGraph Executionで実行することができます。
ほとんどのTensorFlowオペレーションはEager Executionで動作しますが、注意すべき点がいくつかあります:
* 入力処理にはキューの代わりに `tf.data`を使います。この方法はより高速で簡単です。
* `tf.keras.layers`や`tf.keras.Model`のような、オブジェクト指向のレイヤーAPIを使用します—これらのAPIは変数のための明示的なストレージを持っているためです。
* ほとんどのモデルコードは、Eager ExecutionとGraph Executionにおなじように機能しますが、例外があります。
(たとえば、Pythonによる制御フローで入力に基づいて演算を変更する動的モデルなど)
* 一度`tf.enable_eager_execution`によってEager Executionが有効化されると、それを無効化することはできません。
Graph Executionに戻すには、新しいPythonセッションを開始する必要があります。
以上が、Eager Execution *と* Graph Executionの両方のためのコードを書くためのベストプラクティスです。これによって、
Eager Executionによる対話的な実験とデバッガビリティを享受することができ、かつGraph Executionによる分散パフォーマンスの恩恵を受けることができます。
Eager Executionを用いてコードを記述、デバッグ、実験を繰り返したのちにプロダクションへのデプロイのためにモデルパスをimportします。
モデル変数を保存および復元するには `tf.train.Checkpoint`を使います。これはEager ExecutionとGraph Executionの両環境の互換性を担保します。
以下にEager Executionのサンプル集があります:
[tensorflow/contrib/eager/python/examples](https://github.com/tensorflow/tensorflow/tree/master/tensorflow/contrib/eager/python/examples)
### Graph Execution環境でEager Executionを使う
`tfe.py_func`を使ってTensorFlowGraph Execution環境でEager Executionを選択的に可能にすることができます。
この機能は、 `tf.enable_eager_execution()`が呼ばれていないときに使うことができます。
```
def my_py_func(x):
x = tf.matmul(x, x) # tfオペレーションを使用することができる
print(x) # しかしEager Executionで実行される!
return x
with tf.Session() as sess:
x = tf.placeholder(dtype=tf.float32)
# Graph Execution環境でEager Executionを呼び出す
pf = tfe.py_func(my_py_func, [x], tf.float32)
sess.run(pf, feed_dict={x: [[2.0]]}) # [[4.0]]
```
| github_jupyter |
<a href="https://colab.research.google.com/github/krmiddlebrook/intro_to_deep_learning/blob/master/machine_learning/lesson%202%20-%20logistic%20regression/challenges/logistic-regression-pokemon.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Logistic regression with Pokemon data
The goal in this challenge is to build a logistic regression classifier to distinguish the Type 1 Grass Pokemon from other Pokemon using features about each one. Use Tensorflow to build, train, and evaluate the model.
Challenges:
1. Load and prepare the Pokemon dataset.
2. Build the model.
3. Train the model.
4. Evaluate the model.
5. Draw conclusions.
```
# import the libraries we need
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# TensorFlow and tf.keras
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
```
# 1. Load and prepare the data
Load the dataset into a pandas dataframe, and prepare it for the model.
Hints:
- Define the features ($\mathbf{x}$) and labels ($y$). You will probably want to use the Pandas `map` function to convert Pokemon with the Type 1 feature to the proper numerical representation, think "Grass" or not.
- split the dataset into training and test sets
- separate the features and labels in training set and test set
```
data_url = 'https://raw.githubusercontent.com/krmiddlebrook/intro_to_deep_learning/master/datasets/pokemon.csv'
# your code here
```
# 2. Build your model
Build a model to model the relationship between the features $x$ (multiple features) and labels $y$.
Hints:
- use the `Sequential` class to define a container for the layers of your model.
- use the `layers.Dense` class with the sigmoid activation to define your logistic model layer
- define your loss function with "binary_crossentropy"
- configure the optimization algorithm with stochastic gradient descent
- track the accuracy metric
- glue the model, loss function, optimizer, and metrics together
```
# your code here
model = keras.Sequential([
])
```
# 3. Train your model
Now that you have a model, it's time to train it. Train your model for 100 epochs (i.e., iterations), and record the training and validation metrics in the history object.
```
# your code here
```
Visualize the mean squared error metric over the training process. Hint: create a line chart with the epoch (x) and the accuracy (y) variables.
```
# your code here
```
# 4. Evaluate the model
Now that the model is trained, it's time to evaluate it using the test dataset, which you did not use when training the model. This gives you a sense of how well the model predicts unseen data, which is the case when you use it in the real world. Make sure to evaluate the model and visualize it's predictions against the true values.
Hints:
- use the `evaluate` method to test the model
- use the `predict` method to make predictions given test features.
- visualize the predictions against the real labels using matplotlib's pyplot API methods like `scatter` and `plot`.
```
# your code here
```
# 5. Draw conclusions
Write up your conclusions about the model. Report the goal, the model design, and the results. Make sure to contextualize the model results as best you can.
| github_jupyter |
```
import logging
import logging.handlers
logger = logging.getLogger(__name__)
logging.basicConfig(level=logging.DEBUG, format='[%(asctime)s]: %(levelname)s: %(message)s')
from sklearn.metrics import precision_recall_fscore_support as prf
from sklearn.metrics import roc_auc_score, accuracy_score
from sklearn import preprocessing
from sklearn.metrics import auc, roc_curve
import pandas as pd
import numpy as np
import csv
import pickle
import warnings
warnings.filterwarnings('ignore')
dxpr_path_dir = '../tests/feature_selection/'
```
## Load Variable Names
```
def load_names(pickle_file):
with open(pickle_file, 'rb') as f:
names = pickle.load(f)
names.remove('NDX') # since number of diagnoses/procedures are often assigned weeks and even months after discharge, these numbers might not be available in time for early prediction model to use them
names.remove('NPR')
names.remove('NCHRONIC')
return names
names = load_names('../saved_models/input_variables.pkl')
```
## Load Data
```
input_path = "../../../data/hcup/nis/all_year_combination/cleaned_for_repcv_imputed.pickle"
def load_data_full(datafile, names, rescale = True):
fn_pt_trn = pd.read_pickle(datafile)
logger.info(list(fn_pt_trn.columns))
fn_pt_trn.dropna(subset = ['DIED'], inplace = True)
input_df_trn = fn_pt_trn[names]
input_x_trn = input_df_trn.convert_objects(convert_numeric=True)
input_y_died_trn = [int(x) for x in list(fn_pt_trn.DIED)]
if rescale:
scaler = preprocessing.StandardScaler().fit(input_x_trn)
input_x_trn = scaler.transform(input_x_trn)
X = input_x_trn
y = input_y_died_trn
y = np.asarray(y)
return X, y
x, y = load_data_full(input_path, names)
```
## Train/Test (70%/30%) Split
```
from sklearn.model_selection import train_test_split
train_x, test_x, train_y, test_y = train_test_split(x, y, test_size=0.3, random_state=None, stratify = y, shuffle = True)
train_x = np.array(train_x)
test_x = np.array(test_x)
print(train_x.shape)
print(train_y.shape)
print(test_x.shape)
print(test_y.shape)
```
## Machine Learning Models Development
```
from sklearn.metrics import precision_recall_fscore_support as prf
from sklearn.metrics import accuracy_score as acc
from sklearn.metrics import roc_curve, auc, log_loss, confusion_matrix
from sklearn.model_selection import GridSearchCV
from sklearn.calibration import CalibratedClassifierCV
# find the best cutoff point based on Youden's Index
def find_best_threshold(fpr, tpr, thresholds):
optimal_idx = np.argmax(tpr - fpr)
optimal_threshold = thresholds[optimal_idx]
return optimal_threshold
# Model evaluation with fitted model and data used for evaluation. It returns all commonly used evaluation scores.
def evaluation_scores(fitted_model, x_test, y_test, mimic = False, mode = 'macro'):
y_true = y_test
# whether use mimic learning or not.
if mimic == False:
y_pred_prob = fitted_model.predict_proba(x_test)
else:
y_pred_prob = sigmoid(fitted_model.predict(x_test))
y_pred_prob = np.stack((1-y_pred_prob, y_pred_prob), axis = -1)
loss = log_loss(y_true, y_pred_prob)
fpr, tpr, thresholds = roc_curve(y_true, y_pred_prob[:,1])
# get performance based on the best cutoff defined by Youden's J Index
optimal_threshold = find_best_threshold(fpr, tpr, thresholds)
y_pred = y_pred_prob[:,1] >= optimal_threshold
y_pred_prf = y_pred_prob[:,1] >= 0.5
y_pred = y_pred.astype(int)
tn, fp, fn, tp = confusion_matrix(y_true, y_pred).ravel()
precision, recall, f_score, _ = prf(y_true = y_true, y_pred = y_pred_prf, average = mode)
sensitivity = tp/(tp + fn)
specificity = tn/(tn + fp)
roc_auc = auc(fpr, tpr)
ppv = tp/(tp + fp)
accuracy = acc(y_true = y_true, y_pred = y_pred)
return fpr, tpr, tn, fp, fn, tp, precision, recall, f_score, sensitivity, specificity, roc_auc, ppv, accuracy, loss, optimal_threshold
```
### Ridge Logistic Regression
```
from sklearn.linear_model import LogisticRegression
tuned_parameters_lr = {"C": [0.001, 0.01, 0.1, 1, 100, 1000]}
lr_clf = LogisticRegression(penalty = 'l2')
gs_lr = GridSearchCV(lr_clf, tuned_parameters_lr, cv = 3, scoring = 'roc_auc')
gs_lr.fit(train_x, train_y)
gs_lr.best_params_
fpr_lr, tpr_lr, tn_lr, fp_lr, fn_lr, tp_lr, precision_lr, recall_lr, f_score_lr, sensitivity_lr, specificity_lr, roc_auc_lr, ppv_lr, accuracy_lr, loss_lr, optimal_threshold_lr = evaluation_scores(gs_lr, test_x, test_y)
logger.info('precision: %.4f; recall: %4f; f_score: %4f; sensitivity: %4f; specificity: %4f; AUROC: %4f; optimal cutoff: %.4f'
%(precision_lr, recall_lr, f_score_lr, sensitivity_lr, specificity_lr, roc_auc_lr, optimal_threshold_lr))
```
### Gradient Boosting Tree
```
from sklearn.ensemble import GradientBoostingClassifier
gbdt_clf = GradientBoostingClassifier()
tuned_parameters_gbdt = {"n_estimators": [100, 200, 300, 400, 500]}
gs_gbdt = GridSearchCV(gbdt_clf, tuned_parameters_gbdt, cv = 3, scoring = 'roc_auc')
gs_gbdt.fit(train_x, train_y)
gs_gbdt.best_params_
fpr_gbdt, tpr_gbdt, tn_gbdt, fp_gbdt, fn_gbdt, tp_gbdt, precision_gbdt, recall_gbdt, f_score_gbdt, sensitivity_gbdt, specificity_gbdt, roc_auc_gbdt, ppv_gbdt, accuracy_gbdt, loss_gbdt, optimal_threshold_gbdt = evaluation_scores(gs_gbdt, test_x, test_y)
logger.info('precision: %.4f; recall: %.4f; f_score: %.4f; sensitivity: %.4f; specificity: %.4f; AUROC: %.4f; optimal cutoff: %.4f'
%(precision_gbdt, recall_gbdt, f_score_gbdt, sensitivity_gbdt, specificity_gbdt, roc_auc_gbdt, optimal_threshold_gbdt))
```
### Neural Network
```
import tensorflow as tf
import random
# Get validation set for parameter tuning
learn_x, val_x, learn_y, val_y = train_test_split(train_x, train_y, test_size=0.3, random_state=None, stratify = train_y, shuffle = True)
def batch_generator(batch_size, batch_index, input_x, input_y):
input_x = np.array(input_x)
input_y = np.array(input_y)
if (batch_index+1)*batch_size < len(input_x):
return input_x[batch_index*batch_size: (batch_index+1)*batch_size], input_y[batch_index*batch_size: (batch_index+1)*batch_size]
else:
select = random.sample(range(batch_index * batch_size), (batch_index+1)*batch_size - len(input_x))
return np.concatenate([input_x[batch_index*batch_size: len(input_x)], input_x[select]]), np.concatenate([input_y[batch_index*batch_size: len(input_y)], input_y[select]])
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
class Model():
pass
def declare_placeholders(self):
self.input_batch = tf.placeholder(dtype=tf.float32, shape=[None, 82], name='input_batch')
self.ground_truth_labels = tf.placeholder(dtype=tf.int32, shape=[None, None], name='ground_truth_labels')
# self.lengths = tf.placeholder(dtype=tf.int32, shape=[None], name='lengths')
self.dropout_ph = tf.placeholder_with_default(tf.cast(0.5, tf.float32), shape=[], name = 'dropout_ph')
self.learning_rate_ph = tf.placeholder(dtype=tf.float32, shape=[], name='learning_rate_ph')
self.temperature = tf.placeholder(dtype=tf.float32, shape=[], name='softmax_temperature')
Model.__declare_placeholders = classmethod(declare_placeholders)
def build_layers(self, n_hidden):
self.layer_1 = tf.layers.dense(inputs = self.input_batch, units = n_hidden, activation = tf.nn.relu)
self.bn_1 = tf.layers.batch_normalization(inputs = self.layer_1)
self.layer_2 = tf.layers.dense(inputs = self.bn_1, units = n_hidden, activation = tf.nn.relu)
self.bn_2 = tf.layers.batch_normalization(inputs = self.layer_2)
self.layer_3 = tf.layers.dense(inputs = self.bn_2, units = n_hidden, activation = tf.nn.relu)
self.bn_3 = tf.layers.batch_normalization(inputs = self.layer_3)
self.layer_4 = tf.layers.dense(inputs = self.bn_3, units = n_hidden, activation = tf.nn.relu)
self.bn_4 = tf.layers.batch_normalization(inputs = self.layer_4)
self.layer_5 = tf.layers.dense(inputs = self.bn_4, units = n_hidden, activation = tf.nn.relu)
self.bn_5 = tf.layers.batch_normalization(inputs = self.layer_5)
self.layer_6 = tf.layers.dense(inputs = self.bn_5, units = n_hidden, activation = tf.nn.relu)
self.dropout_6 = tf.layers.dropout(self.layer_6, rate = self.dropout_ph)
self.logits = tf.layers.dense(self.dropout_6, 2, activation=None)
Model.__build_layers = classmethod(build_layers)
def compute_predictions(self):
self.softmax_output = tf.nn.softmax(logits = self.logits/self.temperature)
self.predictions = tf.argmax(input = self.softmax_output, axis = -1)
Model.__compute_predictions = classmethod(compute_predictions)
def compute_loss(self):
ground_truth_labels_one_hot = tf.one_hot(self.ground_truth_labels, 2)
# self.loss_tensor = focal_loss(y_true = ground_truth_labels_one_hot, y_pred = self.logits)
# self.loss_tensor = focal_loss(onehot_labels = ground_truth_labels_one_hot, cls_preds = self.logits)
self.loss_tensor = tf.nn.softmax_cross_entropy_with_logits(labels = ground_truth_labels_one_hot, logits = self.logits)
# mask = tf.cast(tf.not_equal(self.input_batch, PAD_index), tf.float32)
self.loss = tf.reduce_mean(self.loss_tensor)
Model.__compute_loss = classmethod(compute_loss)
def perform_optimization(self):
self.optimizer = tf.train.AdamOptimizer(learning_rate = self.learning_rate_ph)
self.grads_and_vars = self.optimizer.compute_gradients(self.loss)
clip_norm = tf.cast(1.0, tf.float32)
self.grads_and_vars_2 = [(tf.clip_by_norm(grad, clip_norm), var) for grad, var in self.grads_and_vars]
self.train_op = self.optimizer.apply_gradients(self.grads_and_vars_2)
Model.__perform_optimization = classmethod(perform_optimization)
def init_model(self, n_hidden):
self.__declare_placeholders()
self.__build_layers(n_hidden)
self.__compute_predictions()
self.__compute_loss()
self.__perform_optimization()
Model.__init__ = classmethod(init_model)
def train_on_batch(self, session, x_batch, y_batch, learning_rate, dropout_keep_probability, temperature):
feed_dict = {self.input_batch: x_batch,
self.ground_truth_labels: y_batch,
self.learning_rate_ph: learning_rate,
self.dropout_ph: dropout_keep_probability,
self.temperature: temperature}
session.run(self.train_op, feed_dict=feed_dict)
Model.train_on_batch = classmethod(train_on_batch)
def predict_for_batch(self, session, x_batch, temperature):#, lengths):
feed_dict = {self.input_batch: x_batch, self.temperature: temperature}#), self.lengths: lengths}
predictions_proba = session.run(self.softmax_output, feed_dict = feed_dict)
predictions = session.run(self.predictions, feed_dict = feed_dict)
return predictions_proba, predictions
Model.predict_for_batch = classmethod(predict_for_batch)
tf.reset_default_graph()
model = Model(n_hidden = 512)
```
#### Parameter Tuning
```
import math
batch_size = 256
n_epochs = 32
learning_rate = 1e-5
#learning_rate_decay = 1.2
dropout_keep_probability = 0.5
temperature = 4
n_batches = math.ceil(len(learn_x)//batch_size)
def evaluation(model, session, data, label, temperature):
from sklearn.metrics import precision_recall_fscore_support as prf
from sklearn.metrics import auc, roc_curve
y_pred_prob, y_pred = model.predict_for_batch(sess, data, temperature)
# print(y_pred_prob[0:2])
# print(y_pred[0:2])
y_true = label
fpr, tpr, thresholds = roc_curve(y_true, y_pred_prob[:,1])
# get performance based on the best cutoff defined by Youden's J Index
optimal_threshold = find_best_threshold(fpr, tpr, thresholds)
y_pred = y_pred_prob[:,1] >= optimal_threshold
y_pred_prf = y_pred_prob[:,1] >= 0.5
y_pred = y_pred.astype(int)
tn, fp, fn, tp = confusion_matrix(y_true, y_pred).ravel()
precision, recall, f_score, _ = prf(y_true = y_true, y_pred = y_pred_prf, average = 'macro')
sensitivity = tp/(tp + fn)
specificity = tn/(tn + fp)
roc_auc = auc(fpr, tpr)
ppv = tp/(tp + fp)
accuracy = acc(y_true = y_true, y_pred = y_pred)
logger.info('precision: %.4f; recall: %.4f; f_score: %.4f; sensitivity: %.4f; specificity: %.4f; AUROC: %.4f; optimal cutoff: %.4f'
%(precision, recall, f_score, sensitivity, specificity, roc_auc, optimal_threshold))
return y_pred_prob, y_pred, roc_auc, fpr, tpr
sess = tf.Session()
sess.run(tf.global_variables_initializer())
# early stop with a patience of 5
baseline = 0
patience = 3
print('Start training... \n')
for epoch in range(n_epochs):
# For each epoch evaluate the model on train and validation data
print('-' * 20 + ' Epoch {} '.format(epoch+1) + 'of {} '.format(n_epochs) + '-' * 20)
print('Train data evaluation:')
evaluation(model, sess, learn_x, learn_y, temperature)
print('Validation data evaluation:')
_, _, score, _, _ = evaluation(model, sess, val_x, val_y, temperature)
if score > baseline:
baseline = score
else:
patience -= 1
if patience <= 0:
break
# Train the model
for batch_index in range(n_batches):
print(batch_index/n_batches,end="\r")
dt = batch_generator(batch_size, batch_index, learn_x, learn_y)
feed_dict = {model.input_batch: dt[0],
model.ground_truth_labels: dt[1].reshape(batch_size,1),
model.learning_rate_ph: learning_rate,
model.dropout_ph: dropout_keep_probability}
model.train_on_batch(sess, dt[0], dt[1].reshape(batch_size, 1), learning_rate, dropout_keep_probability, temperature)
# Decaying the learning rate
# learning_rate = learning_rate / learning_rate_decay
# break
print('...training finished.')
```
#### Training and Testing
```
n_epochs = 12
n_batches = math.ceil(len(train_x)//batch_size)
sess = tf.Session()
sess.run(tf.global_variables_initializer())
print('Start training... \n')
for epoch in range(n_epochs):
# For each epoch evaluate the model on train and validation data
print('-' * 20 + ' Epoch {} '.format(epoch+1) + 'of {} '.format(n_epochs) + '-' * 20)
print('Train data evaluation:')
evaluation(model, sess, train_x, train_y, temperature)
print('Validation data evaluation:')
_, _, roc_auc_nn, fpr_nn, tpr_nn= evaluation(model, sess, test_x, test_y, temperature)
# Train the model
for batch_index in range(n_batches):
print(batch_index/n_batches,end="\r")
dt = batch_generator(batch_size, batch_index, train_x, train_y)
feed_dict = {model.input_batch: dt[0],
model.ground_truth_labels: dt[1].reshape(batch_size,1),
model.learning_rate_ph: learning_rate,
model.dropout_ph: dropout_keep_probability}
model.train_on_batch(sess, dt[0], dt[1].reshape(batch_size, 1), learning_rate, dropout_keep_probability, temperature)
# Decaying the learning rate
# learning_rate = learning_rate / learning_rate_decay
# break
print('...training finished.')
```
### Support Vector Machine
```
from sklearn import svm
tuned_parameters = {"C": [2e-2, 2e-1, 2, 2e1, 2e2]}
gs_svm = GridSearchCV(svm.LinearSVC(), tuned_parameters, cv=3,
scoring='roc_auc', n_jobs = 5)
gs_svm.fit(train_x, train_y)
gs_svm.best_params_
svm_clf = CalibratedClassifierCV(svm.LinearSVC(C = 0.2, class_weight = 'balanced'))
svm_clf.fit(train_x, train_y)
fpr_svm, tpr_svm, tn_svm, fp_svm, fn_svm, tp_svm, precision_svm, recall_svm, f_score_svm, sensitivity_svm, specificity_svm, roc_auc_svm, ppv_svm, accuracy_svm, loss_svm, optimal_threshold_svm = evaluation_scores(svm_clf, test_x, test_y)
logger.info('precision: %.4f; recall: %.4f; f_score: %.4f; sensitivity: %.4f; specificity: %.4f; AUROC: %.4f; optimal cutoff: %.4f'
%(precision_svm, recall_svm, f_score_svm, sensitivity_svm, specificity_svm, roc_auc_svm, optimal_threshold_svm))
```
## ROC Plots
```
import matplotlib
%matplotlib inline
import numpy as np
import pickle
from scipy import interp
matplotlib.use('agg')
import matplotlib.pyplot as plt
matplotlib.rcParams['figure.figsize'] = [10, 10]
fprs = np.linspace(0, 1, 100)
tprs_lr = interp(fprs, fpr_lr, tpr_lr)
tprs_gbdt = interp(fprs, fpr_gbdt, tpr_gbdt)
tprs_nn = interp(fprs, fpr_nn, tpr_nn)
tprs_svm = interp(fprs, fpr_svm, tpr_svm)
# baseline
plt.plot([0, 1], [0, 1], linestyle='--', lw=2, color='#000000',
label='Random guess', alpha=.8)
# Ridge Logistic Regression
plt.plot(fprs, tprs_lr, color='g',
label=r'Ridge Logistic Regression (AUC = %0.2f)' % roc_auc_lr,
lw=2, alpha=.8)
# Gradient Boosting Tree
plt.plot(fprs, tprs_gbdt, color='r',
label=r'Gradient Boosting Tree (AUC = %0.2f)' % roc_auc_gbdt,
lw=2, alpha=.8)
# Artificial Neural Network
plt.plot(fprs, tprs_nn, color='#6e2c00',
label=r'Artificial Neural Network (AUC = %0.2f)' % roc_auc_nn,
lw=2, alpha=.8)
# Support Vector Machine
plt.plot(fprs, tprs_svm, color='y',
label=r'Support Vector Machine (AUC = %0.2f)' % roc_auc_svm,
lw=2, alpha=.8)
plt.xlim([-0.05, 1.05])
plt.xticks(np.arange(0, 1, step=0.1))
plt.ylim([-0.05, 1.05])
plt.yticks(np.arange(0, 1, step=0.1))
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver Operating Characteristic for Model Comparison')
plt.legend(loc="lower right", fontsize="xx-large")
plt.grid()
plt.show()
```
## Variable Importance Analysis
### Gradient Boosting Tree
```
pd.DataFrame(list(zip(names, gs_gbdt.best_estimator_.feature_importances_)), columns = ['var_name', 'importance_score']).sort_values(by = ["importance_score"], ascending = False).head()
```
### Ridge Logistic Regression
```
pd.DataFrame(list(zip(names,gs_lr.best_estimator_.coef_[0])), columns = ['var_name', 'coef']).sort_values(by = ["coef"], ascending = False).head()
```
| github_jupyter |
<img src="../Images/Level1Beginner.png" alt="Beginner" width="128" height="128" align="right">
## Ejercicios - Funciones
### Cuenta espacios
Escribe una función que reciba una cadena y devuelva la cantidad de espacios en blancos que tiene la cadena.
<span><img src="../Images/selected.png" alt="Selected" width="32" height="32" align="right"></span>
```
# definición de la función
def white_space_count(text) :
u"""Returns white space characteres count."""
# descripción - "Qué hace"
# implentación - "Cómo lo hace"
pass
# código que prueba la función
print(white_space_count("Hola que tal"))
```
### Tiene dígitos
Escribe una función que reciba una cadena y devuelva verdadero si tiene algún dígito entre los caracteres o falso si no los tiene.
```
# definición de la función
def has_digits(text) :
u""""Returns True if the text string has digits"""
# descripción - "Qué hace"
# implentación - "Cómo lo hace"
pass
# código que prueba la función
```
### Cuenta palabras
Escribe una función que reciba una cadena y devuelva la cantidad de palabras que tiene la cadena.
```
# definición de la función
def word_count(text) :
u"""Returns words count. """
# descripción - "Qué hace"
# implentación - "Cómo lo hace"
pass
# código que prueba la función
```
### Cuenta dígitos
Escribe una función que reciba una cadena y devuelva la cantidad de dígitos que tiene la cadena.
```
# definición de la función
def digits_count(text) :
u"""Returns digits count. """
# descripción - "Qué hace"
# implentación - "Cómo lo hace"
pass
# código que prueba la función
```
### Palabras largas
Escribe una función que reciba una cadena y un número y devuelva una lista con las palabras de la cadena cuya longitud es mayor al número recibido.
```
# definición de la función
def long_words(text, _length) :
u"""Return a list with long words. """
# cógido que implementa el "Qué hace"
pass
# código que prueba la función
```
### Horas extras
Escribe una función que calcule y devuelva el salario de un empleado de manera que cobre 1.5 veces el valor de cada hora que exceda las 40 mensuales.
```
# definición de la función
```
### Calificaciones
Escriba una función que determine y devuelva la denominación de una calificación de acuerdo al siguiente detalle:
\>= 0.9 - Sobresaliente
\>= 0.8 - Notable
\>= 0.7 - Bien
\>= 0.6 - Suficiente
\< 0.6 - Insuficiente
Los valores fuera del rango 0.0 y 1.0 no deben aceptarse.
```
# definición de la función
```
### Validar una entrada
Escribe una función que solicite el ingreso de un número entre dos valores inclusive, si se ingresa un número fuera del rango el código solicita nuevamente que se ingrese el número tantas vecese como sea necesario hasta cumplir con la validación.
```
# definición de la función
```
### Máximo Común Divisor
Escribe la función que permita obtener el máximo común divisor de dos números.
```
# definición de la función
```
### Palabra alfabética
Se dice que una palabra es alfabética si todas sus letras están ordenadas alfabéticamente, por ejemplo "chino".
Escriba una función que reciba una palabra y devuelva verdadero si la palabra es alfabética y falso si no lo es
```
# definición de la función
```
### Criptografía básica
Dado un alfabeto (secuencia de símbolos o caracteres de un lenguaje) una técnica de encriptar es tomar un texto y un número y reemplazar cada caracter del texto por el caracter que se encuentra tantas posiciones más a la derecha como el valor del número; cuando se alcanza el final del alfabeto se vuelve a comenzar por el primer símbolo del alfabeto.
Escriba una función que reciba una cadena y un número y devuelva la cadena encriptada según la técnica indicada anteriormente de acuerdo a un alfabeto que contenga los símbolos para números, mayúsculas y minúsculas.
```
# definición de la función
```
### Máximo Común Divisor de varios números
Escribe la función que permita obtener el máximo común divisor de al menos tres números.
```
# definición de la función
```
### Números primos
Escribe la función que determina si un número es primo o no.
```
# definición de la función
```
### Cuenta números
Escribe una función que reciba una cadena y devuelva la cantidad de números (no dígitos) que tiene la cadena.
```
# definición de la función
```
| github_jupyter |
```
#necessary imports
import numpy as np
import scipy
from scipy.special import gamma, factorial
import scipy.special as sc
import mpmath as mp
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D #for 3D surface plots
import math
from cmath import phase
from scipy.ndimage.filters import gaussian_filter1d
plt.rc('xtick',labelsize=20)
plt.rc('ytick',labelsize=20)
"""
Reproducing plots from the following paper on point mass and SIS lens models
R. Takahashi and T. Nakamura, “Wave effects in the gravitational lensing of gravitational waves from chirping binaries,” The Astrophysical Journal, vol. 595, pp. 1039–1051, 2003.
"""
#Point mass gravitational lens model
#magnitude of amplification factor plot
#legends on source position
y=[0.1,0.25,0.5,1.0]
plt.figure(figsize=(10,8))
#computing magnification factor magnitude for varying values of dimensionless frequencies
for j in range(0,4):
n=10000
w=np.linspace(0.01,27,n)
func1=np.exp((np.pi*w)/4)
z=1-((1j/2)*w)
func2=abs(scipy.special.gamma(z))
a=(1j/2)*w
b=np.ones(n)
c=(1j/2)*w*y[j]*y[j]
func3=np.zeros(n)
for i in range(0,n):
func3[i]=abs(mp.hyp1f1(a[i],b[i],c[i]))
F=func1*func2*func3
plt.loglog(w, F) #plot
plt.grid(True, which="both", ls="-")
plt.xlabel('w=8πMf (dimensionless)', fontsize=20)
plt.xlim(0.01,27)
plt.ylim(0.1,10)
plt.ylabel('|F| (dimensionless)', fontsize=20)
plt.legend(['y = 0.1', 'y = 0.25', 'y = 0.5', 'y = 1.0'], loc='upper left', fontsize=20)
plt.show()
#legends on w
w=[20,5,1]
plt.figure(figsize=(10,8))
#computing magnification factor magnitude for varying values of y
for j in range(0,3):
n=10000
y=np.linspace(0.01,1.5,n)
func1=np.exp((np.pi*w[j])/4)
z=1-((1j/2)*w[j])
func2=abs(scipy.special.gamma(z))
for i in range(0,n):
a[i]=(1j/2)*w[j]
b=np.ones(n)
c=(1j/2)*w[j]*y*y
func3=np.zeros(n)
for i in range(0,n):
func3[i]=abs(mp.hyp1f1(a[i],b[i],c[i]))
F=func1*func2*func3
plt.loglog(y, F) #plot
plt.grid(True, which="both", ls="-")
plt.xlabel('y (dimensionless)', fontsize=20)
plt.xlim(np.amin(y),np.amax(y))
plt.ylim(0.1,20)
plt.ylabel('|F| (dimensionless)', fontsize=20)
plt.legend(['w = 20', 'w = 5', 'w = 1'], loc='upper right', fontsize=20)
plt.show()
#phase of amplification factor plot
y=[0.1,0.25,0.5,1.0]
plt.figure(figsize=(10,8))
xm=np.zeros(4)
for j in range(0,4):
n=10000
w=np.linspace(0.01,27,n)
func1=np.exp((np.pi*w)/4)
z=1-((1j/2)*w)
func2=(scipy.special.gamma(z))
a=(1j/2)*w
b=np.ones(n)
c=(1j/2)*w*y[j]*y[j]
func3=np.zeros(n,dtype='complex64')
for i in range(0,n):
func3[i]=(mp.hyp1f1(a[i],b[i],c[i]))
xm=0.5*(y[j]+np.sqrt(y[j]*y[j]+4.0))
phim=0.5*((xm-y[j])**2)-np.log(xm)
func4=np.zeros(n,dtype='complex64')
for i in range(0,n):
func4[i]=mp.exp(0.5j*w[i]*(np.log(0.5*w[i])-2.0*phim))
F=abs(func1*func2*func3)
Phi=-1j*np.log((func1*func2*func3*func4)/abs((func1*func2*func3*func4)))
plt.grid(True, which="both", ls="-")
plt.plot(w, Phi)
plt.xlabel('w=8πMf (dimensionless)', fontsize=20)
plt.xscale('log')
plt.xlim(np.amin(w),np.amax(w))
plt.ylabel('\u03B8(F) (in radians)', fontsize=20)
plt.legend(['y = 0.1', 'y = 0.25', 'y = 0.5', 'y = 1.0'], loc='upper left', fontsize=20)
plt.show()
w=[20,5,1]
plt.figure(figsize=(10,8))
for j in range(0,3):
n=10000
y=np.linspace(0.01,1.5,n)
func1=np.exp((np.pi*w[j])/4)
z=1-((1j/2)*w[j])
func2=(scipy.special.gamma(z))
for i in range(0,n):
a[i]=(1j/2)*w[j]
b=np.ones(n)
c=(1j/2)*w[j]*y*y
func3=np.zeros(n,dtype='complex64')
for i in range(0,n):
func3[i]=(mp.hyp1f1(a[i],b[i],c[i]))
xm=0.5*(y+np.sqrt(y*y+4.0))
phim=0.5*((xm-y)**2)-np.log(xm)
func4=np.zeros(n,dtype='complex64')
for i in range(0,n):
func4[i]=mp.exp(0.5j*w[j]*(np.log(0.5*w[j])-2.0*phim[i]))
F=abs(func1*func2*func3)
Phi=-1j*np.log((func1*func2*func3*func4)/abs((func1*func2*func3*func4)))
plt.grid(True, which="both", ls="-")
plt.plot(y, Phi)
plt.xlabel('y (dimensionless)',fontsize=20)
plt.xscale('log')
plt.ylabel('\u03B8(F) (in radians)',fontsize=20)
plt.xlim(np.amin(y),np.amax(y))
plt.legend(['w = 20', 'w = 5', 'w = 1'], loc='upper left',fontsize=20)
plt.show()
import numpy as np
import matplotlib.pyplot as plt
from mpmath import gamma, hyp1f1, factorial, exp, pi,log
from cmath import phase
plt.rc('xtick',labelsize=20)
plt.rc('ytick',labelsize=20)
#singular isothermal sphere (SIS) gravitational lens model
#defining the summation in F function
def integrand(n,w,y):
return gamma(1+n/2.0)*hyp1f1(1+n/2.0,1.0,-0.5j*w*y*y)*((2*w*exp(1j*3*pi/2))**(n/2))/factorial(n)
#computing phase and magnitude of amplification factor for 4 values of y and varying values of w
N = 1000
wvec = np.linspace(0.01,27,N)
Fvec = np.zeros(N)
Ph1 = np.zeros(N, dtype='complex64')
Ph2 = np.zeros(N, dtype='complex64')
Ph3 = np.zeros(N, dtype='complex64')
Ph4 = np.zeros(N, dtype='complex64')
y=0.1
for i,w in enumerate(wvec):
delta = 1.0
F = 0.0
n = 0
while delta>1e-6:
dF = integrand(n,w,y)
F += dF
delta = np.abs(dF)
n += 1
Fvec[i] = abs(exp(0.5j*w*y*y)*F)
Ph1[i]=-1j*log(exp(0.5j*w*((y*y)+(2*y)+1))*F/abs((exp(0.5j*w*((y*y)+(2*y)+1))*F)))
plt.figure(figsize=(10,8))
plt.loglog(wvec,Fvec)
y=0.25
for i,w in enumerate(wvec):
delta = 1.0
F = 0.0
n = 0
while delta>1e-6:
dF = integrand(n,w,y)
F += dF
delta = np.abs(dF)
n += 1
Fvec[i] = abs(exp(0.5j*w*y*y)*F)
Ph2[i]=-1j*log((exp(0.5j*w*((y*y)+(2*y)+1))*F)/abs((exp(0.5j*w*((y*y)+(2*y)+1))*F)))
plt.loglog(wvec,Fvec)
y=0.5
for i,w in enumerate(wvec):
delta = 1.0
F = 0.0
n = 0
while delta>1e-6:
dF = integrand(n,w,y)
F += dF
delta = np.abs(dF)
n += 1
Fvec[i] = abs(exp(0.5j*w*y*y)*F)
Ph3[i]=-1j*log(exp(0.5j*w*((y*y)+(2*y)+1))*F/abs((exp(0.5j*w*((y*y)+(2*y)+1))*F)))
plt.loglog(wvec,Fvec)
y=1.0
for i,w in enumerate(wvec):
delta = 1.0
F = 0.0
n = 0
while delta>1e-6:
dF = integrand(n,w,y)
F += dF
delta = np.abs(dF)
n += 1
Fvec[i] = abs(exp(0.5j*w*y*y)*F)
Ph4[i]=-1j*log(exp(0.5j*w*((y*y)+(2*y)+1))*F/abs((exp(0.5j*w*((y*y)+(2*y)+1))*F)))
#magnitude plots
plt.loglog(wvec,Fvec)
plt.grid(True, which="both", ls="-")
plt.xlim(0.01,27)
plt.ylim(0.1,10)
plt.xlabel('w=8πMf (dimensionless)', fontsize=20)
plt.ylabel('|F| (dimensionless)', fontsize=20)
plt.legend(['y = 0.1', 'y = 0.25', 'y = 0.5', 'y = 1.0'], loc='upper left', fontsize=20)
plt.show()
#phase plots
plt.figure(figsize=(10,8))
plt.plot(wvec,Ph1)
plt.plot(wvec,Ph2)
plt.plot(wvec,Ph3)
plt.plot(wvec,Ph4)
plt.xscale('log')
plt.xlim(np.amin(wvec),np.amax(wvec))
plt.xlabel('w=8πMf (dimensionless)', fontsize=20)
plt.ylabel('\u03B8(F) (in radians)', fontsize=20)
plt.grid(True, which="both", ls="-")
plt.legend(['y = 0.1', 'y = 0.25', 'y = 0.5', 'y = 1.0'], loc='upper left', fontsize=20)
plt.show()
#phase and magnitude values computation for 4 values of w and varying values of y
a=[20,5,1]
yvec=np.linspace(0.01,1.5,N)
plt.figure(figsize=(10,8))
for j in range(0,3):
w=a[j]
for i,y in enumerate(yvec):
delta = 1.0
F = 0.0
n = 0
while delta>1e-3:
dF = integrand(n,w,y)
F += dF
delta = np.abs(dF)
n += 1
Fvec[i] = abs(exp(0.5j*w*y*y)*F)
if(j==0):
Ph1[i]=-1j*log(exp(0.5j*w*((y*y)+(2*y)+1))*F/abs((exp(0.5j*w*((y*y)+(2*y)+1))*F)))
elif(j==1):
Ph2[i]=-1j*log(exp(0.5j*w*((y*y)+(2*y)+1))*F/abs((exp(0.5j*w*((y*y)+(2*y)+1))*F)))
elif(j==2):
Ph3[i]=-1j*log(exp(0.5j*w*((y*y)+(2*y)+1))*F/abs((exp(0.5j*w*((y*y)+(2*y)+1))*F)))
plt.loglog(yvec,Fvec)
#magnitude plot
plt.grid(True, which="both", ls="-")
plt.xlabel('y (dimensionless)', fontsize=20)
plt.xlim(np.amin(yvec),np.amax(yvec))
plt.ylim(0.1,20)
plt.ylabel('|F| (dimensionless)', fontsize=20)
plt.legend(['w = 20', 'w = 5', 'w = 1'], loc='upper right', fontsize=20)
plt.show()
#phase plot
plt.figure(figsize=(10,8))
plt.plot(yvec,Ph1)
plt.plot(yvec,Ph2)
plt.plot(yvec,Ph3)
plt.xscale('log')
plt.xlim(np.amin(yvec),np.amax(yvec))
plt.xlabel('y (dimensionless)', fontsize=20)
plt.ylabel('\u03B8(F) (in radians)', fontsize=20)
plt.grid(True, which="both", ls="-")
plt.legend(['w = 20', 'w = 5', 'w = 1'], loc='upper left', fontsize=20)
plt.show()
```
| github_jupyter |
## IBM Quantum Challenge Fall 2021
# Challenge 2: OLED 분자들의 밴드갭 계산
<div id='problem'></div>
<div class="alert alert-block alert-info">
최고의 경험을 위해 오른쪽 상단의 계정 메뉴에서 **light** 워크스페이스 테마로 전환하는 것을 추천합니다.
## 소개
유기 발광 다이오드(Organic Light Emitting Diode) 또는 OLED는 전류를 인가하면 빛을 내는, 얇고 유연한 TV 및 휴대폰 디스플레이 제조의 기초 소자로 최근 몇 년 동안 점점 인기를 얻고 있습니다.
최근 연구([**Gao et al., 2021**](https://www.nature.com/articles/s41524-021-00540-6))에서는 페닐설포닐-카바졸(PSPCz) 분자에서 고에너지 상태의 전자 전이를 관측했으며, 이는 OLED 기술에 유용한 열 활성화 지연 형광(TADF) 이미터에 활용 될 수있습니다. OLED를 만들기 위해 현재 사용되는 양자 효율이 25%로 제한된 기존의 형광 포어와 비교하여 TADF 방출기는 100% 내부 양자 효율(IQP)-흡수된 광자를 방출하는 회로 또는 시스템에서 전하 캐리어의 비율-로 작동하는 OLED를 생산할 가능성을 지니고 있습니다. 효율성의 큰 증가는 제조업체들이 휴대폰 처럼 낮은 소비 전력을 요구하는 기기에 사용할 OLED를 생산할 수 있다는 것을 의미하며, 이는 결과적으로 가정, 사무실, 박물관 그리고 그보다 더 많은 넓은 면적의 공간들이 값싸고 에너지 효율이 높은 조명원을 사용하게 되는 미래의 개발로 이어질 수 있음을 의미합니다!
<center><img src="resources/JSR_img6_1920w.jpg" width="600"></center>
### Why quantum?
양자 컴퓨터는 고전적인 컴퓨터보다 양자 장치에서 양자 역학 시스템을 모델링하는 것이 지닌 자연스러움으로 인해 복잡한 분자 및 재료의 전자 구조 및 동적 특성을 연구하는 데 귀중한 도구가 될 것으로 기대되고 있습니다. IBM Quantum과 파트너의 최근 공동 연구 프로젝트는 효율적인 OLED를 위한 TADF의 들뜬 상태 계산의 정확도를 향상시키는 방법을 성공적으로 개발하여 상용 재료의 들뜬 상태 계산에 양자 컴퓨터를 적용한 세계 최초의 연구 사례가 되었습니다(위에 링크된 논문 참조).
이러한 배경 정보와 함께, 효율적인 유기발광다이오드(OLED) 소자의 제작에 잠재적으로 사용될 수 있는, 산업적 화학 화합물의 "들뜬 상태" 또는 높은 에너지 상태에 대한 양자 계산을 설명하고자 합니다.
## 도전 과제
<div class="alert alert-block alert-success">
<b>목표</b>
이 도전의 목표는 양자 알고리즘을 사용하여 TADF 물질의 들뜬 상태 에너지를 신뢰성 있게 계산하는 것입니다. 그러기 위해서 이 노트북을 통해 고전적 근사 단계와 보다 정확한 양자 계산 단계 간의 작업 부하 분할을 허용하는 최첨단 하이브리드 고전-양자 방식의 화학 모델링을 도입합니다.
1. **도전 2a & 2b**: 원자 궤도(AO), 분자 궤도 (MO), 능동 공간 변환(Active Space Transformation)을 사용하여 궤도의 수를 줄이는 방법을 이해합니다.
2. **도전 2c & 2d**: NumPy 및 VQE(Variational Quantum Eigensolver)를 사용하여 PSPCz 분자의 바닥 상태 에너지를 계산합니다.
3. **도전 2e**: 양자 운동 방정식(QEOM) 알고리즘을 사용하여 PSPCz 분자의 들뜬 상태 에너지를 계산합니다.
4. **도전 2f**: Qiskit Runtime을 사용하여 클라우드(시뮬레이터 또는 실제 양자 시스템)에서 VQE를 실행합니다.
</div>
<div class="alert alert-block alert-info">
시작하기에 앞서,[**Qiskit Nature Demo Session with Max Rossmannek**](https://youtu.be/UtMVoGXlz04?t=38)을 시청하고 데모에 사용된 [**demo notebook**](https://github.com/qiskit-community/qiskit-application-modules-demo-sessions/tree/main/qiskit-nature)을 통해 전자 구조 계산을 미리 배워보길 추천합니다.
</div>
### 1. 드라이버(Driver)
Qiskit과 고전적인 화학 코드 사이의 인터페이스를 드라이버라고 합니다. `PSI4Driver`, `PyQuanteDriver`, `PySCFDriver` 등이 있습니다.
아래의 셀에서 드라이버(주어진 기저 집합과 분자의 구조 정보에 대한 하트리-폭 계산)를 실행하여, 양자 알고리즘을 적용하기 위해 알아야 하는 대상 분자에 대한 모든 필요한 정보를 얻습니다.
```
from qiskit_nature.drivers import Molecule
from qiskit_nature.drivers.second_quantization import ElectronicStructureDriverType, ElectronicStructureMoleculeDriver
# PSPCz molecule
geometry = [['C', [ -0.2316640, 1.1348450, 0.6956120]],
['C', [ -0.8886300, 0.3253780, -0.2344140]],
['C', [ -0.1842470, -0.1935670, -1.3239330]],
['C', [ 1.1662930, 0.0801450, -1.4737160]],
['C', [ 1.8089230, 0.8832220, -0.5383540]],
['C', [ 1.1155860, 1.4218050, 0.5392780]],
['S', [ 3.5450920, 1.2449890, -0.7349240]],
['O', [ 3.8606900, 1.0881590, -2.1541690]],
['C', [ 4.3889120, -0.0620730, 0.1436780]],
['O', [ 3.8088290, 2.4916780, -0.0174650]],
['C', [ 4.6830900, 0.1064460, 1.4918230]],
['C', [ 5.3364470, -0.9144080, 2.1705280]],
['C', [ 5.6895490, -2.0818670, 1.5007820]],
['C', [ 5.4000540, -2.2323130, 0.1481350]],
['C', [ 4.7467230, -1.2180160, -0.5404770]],
['N', [ -2.2589180, 0.0399120, -0.0793330]],
['C', [ -2.8394600, -1.2343990, -0.1494160]],
['C', [ -4.2635450, -1.0769890, 0.0660760]],
['C', [ -4.5212550, 0.2638010, 0.2662190]],
['C', [ -3.2669630, 0.9823890, 0.1722720]],
['C', [ -2.2678900, -2.4598950, -0.3287380]],
['C', [ -3.1299420, -3.6058560, -0.3236210]],
['C', [ -4.5179520, -3.4797390, -0.1395160]],
['C', [ -5.1056310, -2.2512990, 0.0536940]],
['C', [ -5.7352450, 1.0074800, 0.5140960]],
['C', [ -5.6563790, 2.3761270, 0.6274610]],
['C', [ -4.4287740, 3.0501460, 0.5083650]],
['C', [ -3.2040560, 2.3409470, 0.2746950]],
['H', [ -0.7813570, 1.5286610, 1.5426490]],
['H', [ -0.7079140, -0.7911480, -2.0611600]],
['H', [ 1.7161320, -0.2933710, -2.3302930]],
['H', [ 1.6308220, 2.0660550, 1.2427990]],
['H', [ 4.4214900, 1.0345500, 1.9875450]],
['H', [ 5.5773000, -0.7951290, 3.2218590]],
['H', [ 6.2017810, -2.8762260, 2.0345740]],
['H', [ 5.6906680, -3.1381740, -0.3739110]],
['H', [ 4.5337010, -1.3031330, -1.6001680]],
['H', [ -1.1998460, -2.5827750, -0.4596910]],
['H', [ -2.6937370, -4.5881470, -0.4657540]],
['H', [ -5.1332290, -4.3740010, -0.1501080]],
['H', [ -6.1752900, -2.1516170, 0.1987120]],
['H', [ -6.6812260, 0.4853900, 0.6017680]],
['H', [ -6.5574610, 2.9529350, 0.8109620]],
['H', [ -4.3980410, 4.1305040, 0.5929440]],
['H', [ -2.2726630, 2.8838620, 0.1712760]]]
molecule = Molecule(geometry=geometry, charge=0, multiplicity=1)
driver = ElectronicStructureMoleculeDriver(molecule=molecule,
basis='631g*',
driver_type=ElectronicStructureDriverType.PYSCF)
```
<div class="alert alert-block alert-success">
<b> 도전 2a</b>
질문: PSPCz 분자의 다음의 값들을 찾아봅시다.
1. C, H, N, O, S 원자들은 각각 몇개입니까?
1. 원자들의 총 몇 개입니까?
1. 원자 궤도는 전부 몇개입니까 (AO) ?
1. 분자 궤도는 전부 몇개입니까 (MO) ?
</div>
<div class="alert alert-block alert-danger">
**원자의 궤도를 어떻게 셀 수 있나요?**
궤도의 숫자는 베이시스와 관련이 있습니다. 아래의 숫자는 이 도전에 사용될 `631g*` 베이시스의 숫자들입니다.
- C: 1s, 2s2p, 3s3p3d = 1+4+9 = 14
- H: 1s, 2s = 1+1 = 2
- N: 1s, 2s2p, 3s3p3d = 1+4+9 = 14
- O: 1s, 2s2p, 3s3p3d = 1+4+9 = 14
- S: 1s, 2s2p, 3s3p3d, 4s4p = 1+4+9+4 = 18
```
num_ao = {
'C': 14,
'H': 2,
'N': 14,
'O': 14,
'S': 18,
}
##############################
# Provide your code here
num_C_atom =
num_H_atom =
num_N_atom =
num_O_atom =
num_S_atom =
num_atoms_total =
num_AO_total =
num_MO_total =
##############################
answer_ex2a ={
'C': num_C_atom,
'H': num_H_atom,
'N': num_N_atom,
'O': num_O_atom,
'S': num_S_atom,
'atoms': num_atoms_total,
'AOs': num_AO_total,
'MOs': num_MO_total
}
print(answer_ex2a)
# Check your answer and submit using the following code
from qc_grader import grade_ex2a
grade_ex2a(answer_ex2a)
```
위의 연습에서 발견했듯이 PSPCz는 많은 원자와 많은 원자 궤도로 구성된 거대한 분자입니다. 거대한 분자는 현재의 양자 시스템으로 직접 계산하기 어렵습니다. 그러나 이 도전에서는 밴드갭에만 관심이 있으므로 HOMO(Highest Occuppied Molecular Orbital) 및 LUMO(Lowest Unoccuppied Molecular Orbital)의 에너지를 계산하는 것만으로도 충분합니다. 이 부분에서는 분자 궤도의 수를 2개(HOMO 및 LUMO)로 줄이기 위해 능동 공간 변환(Active Space Transformation) 기술을 사용해 보도록 하겠습니다:
$$E_g = E_{LUMO} - E_{HOMO}$$
<center><img src="resources/Molecule_HOMO-LUMO_diagram.svg" width="600"></center>
이 그림의 동그라미들은 궤도에 있는 전자입니다; HOMO에 위치한 전자가 충분히 높은 주파수의 에너지나 빛을 흡수하면 전자는 LUMO 상태로 점프하여 올라갑니다.
이번 PSPCz 분자의 문제에서 들뜬 상태는 첫 번째 단일항(singlet) 및 삼중항(triplet) 상태로 제한합니다. 단일항 상태에서 시스템의 모든 전자는 스핀 쌍을 이루며 공간에서 가능한 한 방향 선택할 수 있습니다. 단일 또는 삼중항 상태는 두 전자 중 하나를 더 높은 에너지 수준으로 들뜨게하여 생성됩니다. 들뜬 전자는 단일 들뜬 상태에서 동일한 스핀 방향을 유지하는 반면, 삼중 들뜬 상태에서 들뜬 전자는 바닥 상태 전자와 동일한 스핀 방향을 가집니다.
<center><img src="resources/spin.jpg" width="300"><figcaption>Spin in the ground and excited states</figcaption></center>
삼중항 상태에서 전자 스핀의 한 세트는 짝을 이루지 않습니다. 즉, 공간의 세 방향 축을 스핀이 선택 할 수 있습니다. PSPCz (a, d)와 그 변형 2F-PSPCz (b, e), 4F-PSPCz (c, f)의 삼중항 상태에서 최적화된 구조의 LUMO (a-c)와 HOMO (e-f)는 다음 그림과 같을 것입니다.
<center><img src="resources/oled_paper_fig2.jpg" width="600"></center>
<center><img src="resources/oled_paper_fig1.jpg" width="600"></center>
능동 공간 변환 방법을 사용하여, 가능한 가장 작은 활성 공간인 단일항와 삼중항으로 계산을 제한함과 동시에 비핵심 전자 상태를 배제하면서 시스템에 대한 높은 수준의 표현력을 유지하면서 적은 수의 큐비트로 이 에너지를 계산할 수 있습니다.
```
from qiskit_nature.drivers.second_quantization import HDF5Driver
driver_reduced = HDF5Driver("resources/PSPCz_reduced.hdf5")
properties = driver_reduced.run()
from qiskit_nature.properties.second_quantization.electronic import ElectronicEnergy
electronic_energy = properties.get_property(ElectronicEnergy)
print(electronic_energy)
```
`(AO) 1-Body Terms`은 대상 분자가 가진 전체 430개의 분자 궤도에 대한 430개의 원자 궤도를 표현하는 (430 x 430) 행렬을 지니고 있습니다. `ActiveSpaceTransformation`(미리 계산되어 있음) 이후에, 분자 궤도 `(MO) 1-Body Terms`의 수는 (2x2) 행렬로 감소합니다.
<div class="alert alert-block alert-success">
<b> 도전 2b</b>
질문: 속성 프레임워크(Property framework)를 사용하여 아래 질문에 대한 답하시오.
1. 능동 공간 변환 후 시스템의 전자 수는 얼마입니까?
1. 분자 궤도는 몇 개입니까? What is the number of molecular orbitals?
1. Spin-orbital은 몇 개입니까? What is the number of spin-orbitals?
1. Jordan-Wigner 매핑으로 이 분자를 시뮬레이션하려면 몇 개의 큐비트가 필요합니까?
</div>
```
from qiskit_nature.properties.second_quantization.electronic import ParticleNumber
##############################
# Provide your code here
particle_number =
num_electron =
num_MO =
num_SO =
num_qubits =
##############################
answer_ex2b = {
'electrons': num_electron,
'MOs': num_MO,
'SOs': num_SO,
'qubits': num_qubits
}
print(answer_ex2b)
# Check your answer and submit using the following code
from qc_grader import grade_ex2b
grade_ex2b(answer_ex2b)
```
### 2. 전자 구조 문제
다음 단계로, 큐비트에 매핑하기 전, ElectronicStructureProblem을 사용하여 페르미온 연산자 목록을 작성합니다. 여러분의 분자 시스템의 바닥 상태를 계산하기 위한 첫번째 단계입니다. 바닥 상태를 계산하는 자세한 내용은 [**this tutorial**](https://qiskit.org/documentation/nature/tutorials/03_ground_state_solvers.html)에서 확인할 수 있습니다.
<center><img src="resources/H2_gs.png" width="300"></center>
```
from qiskit_nature.problems.second_quantization import ElectronicStructureProblem
##############################
# Provide your code here
es_problem =
##############################
second_q_op = es_problem.second_q_ops()
print(second_q_op[0])
```
### 3. 큐비트 변환기(QubitConverter)
시뮬레이션에 사용할 매핑의 종류를 설정할 수 있게 해줍니다.
```
from qiskit_nature.converters.second_quantization import QubitConverter
from qiskit_nature.mappers.second_quantization import JordanWignerMapper, ParityMapper, BravyiKitaevMapper
##############################
# Provide your code here
qubit_converter =
##############################
qubit_op = qubit_converter.convert(second_q_op[0])
print(qubit_op)
```
### 4. 초기 상태
이론 부분에서 설명한 것과 같이 화학 계산에서 사용하기 좋은 초기 상태는 HF 상태입니다. 다음과 같이 초기화할 수 있습니다:
```
from qiskit_nature.circuit.library import HartreeFock
##############################
# Provide your code here
init_state =
##############################
init_state.draw()
```
### 5. 안사츠(Ansatz)
바닥 상태를 계산하기 위해서는 좋은 양자 회로를 선택하는 것이 몹시 중요합니다.
Qiskit circuit library를 사용해 여러분만의 회로를 작성하는 예제들을 아래에서 확인해 보세요.
```
from qiskit.circuit.library import EfficientSU2, TwoLocal, NLocal, PauliTwoDesign
from qiskit_nature.circuit.library import UCCSD, PUCCD, SUCCD
##############################
# Provide your code here
ansatz =
##############################
ansatz.decompose().draw()
```
## 바닥 상태 에너지 계산
### Numpy를 사용해 계산하기
학습 목적으로, 행렬의 대각화를 통해 이 문제를 정확하게 푼 값을 VQE가 얻어야 하는 목표값으로 삼을 수 있습니다. 이 행렬의 차원은 분자 궤도의 수가 늘어남에 따라 기하급수적으로 확장되기 때문에 문제의 대상인 큰 분자의 경우 계산에 다소 시간이 소요 될수 있습니다. 아주 커다란 분자 시스템을 계산할 경우 파동 함수를 저장할 메모리 공간이 부족할 수 있습니다.
<center><img src="resources/vqe.png" width="600"></center>
```
from qiskit.algorithms import NumPyMinimumEigensolver
from qiskit_nature.algorithms import GroundStateEigensolver
##############################
# Provide your code here
numpy_solver =
numpy_ground_state_solver =
numpy_results =
##############################
exact_energy = numpy_results.computed_energies[0]
print(f"Exact electronic energy: {exact_energy:.6f} Hartree\n")
print(numpy_results)
# Check your answer and submit using the following code
from qc_grader import grade_ex2c
grade_ex2c(numpy_results)
```
### VQE로 계산하기
다음 단계는 VQE를 사용하여 바닥 상태 에너지를 계산하는 것입니다. 여러분의 전자 문제의 해결까지 이제 절반가량 왔습니다!
```
from qiskit.providers.aer import StatevectorSimulator, QasmSimulator
from qiskit.algorithms.optimizers import COBYLA, L_BFGS_B, SPSA, SLSQP
##############################
# Provide your code here
backend =
optimizer =
##############################
from qiskit.algorithms import VQE
from qiskit_nature.algorithms import VQEUCCFactory, GroundStateEigensolver
from jupyterplot import ProgressPlot
import numpy as np
error_threshold = 10 # mHartree
np.random.seed(5) # fix seed for reproducibility
initial_point = np.random.random(ansatz.num_parameters)
# for live plotting
pp = ProgressPlot(plot_names=['Energy'],
line_names=['Runtime VQE', f'Target + {error_threshold}mH', 'Target'])
intermediate_info = {
'nfev': [],
'parameters': [],
'energy': [],
'stddev': []
}
def callback(nfev, parameters, energy, stddev):
intermediate_info['nfev'].append(nfev)
intermediate_info['parameters'].append(parameters)
intermediate_info['energy'].append(energy)
intermediate_info['stddev'].append(stddev)
pp.update([[energy, exact_energy+error_threshold/1000, exact_energy]])
##############################
# Provide your code here
vqe =
vqe_ground_state_solver =
vqe_results =
##############################
print(vqe_results)
error = (vqe_results.computed_energies[0] - exact_energy) * 1000 # mHartree
print(f'Error is: {error:.3f} mHartree')
# Check your answer and submit using the following code
from qc_grader import grade_ex2d
grade_ex2d(vqe_results)
```
## 들뜬 상태 계산
### QEOM을 사용한 계산
이번에는 여러분의 분자 해밀토니안의 들뜬 상태를 계산해 보도록 하겠습니다. 시스템은 위에서 이미 정의했으므로, 이번에는 양자 운동 방정식(qEOM) 알고리즘을 사용하여 들뜬 상태의 에너지를 아래의 유사 고유값 문제로 풀어보도록 하겠습니다.
<center><img src="resources/math-1.svg" width="400"></center>
에서
<center><img src="resources/math-2.svg" width="300"></center>
이며, 각 매트릭스 요소는 각각에 해당하는 기저 상태와 함께 양자 컴퓨터에서 측정되어야 합니다.
더 깊은 이해를 위해, 들뜬 상태 계산에 대한 [**this tutorial**](https://qiskit.org/documentation/nature/tutorials/04_excited_states_solvers.html)을 읽어보시길 권장하며, qEOM 이론 자체는 [**corresponding paper by Ollitrault et al., 2019**](https://arxiv.org/abs/1910.12890)을 읽어보시길 권장합니다.
```
from qiskit_nature.algorithms import QEOM
##############################
# Provide your code here
qeom_excited_state_solver =
qeom_results =
##############################
print(qeom_results)
# Check your answer and submit using the following code
from qc_grader import grade_ex2e
grade_ex2e(qeom_results)
```
최종적으로, 위에서 얻은 두 세트의 들뜬 상태의 에너지와 바닥 상태의 에너지 차이를 계산하여 밴드 갭 또는 에너지 갭(전자가 바닥 상태에서 들뜬 상태로 벗어나는 데 필요한 최소 에너지량)을 얻게 됩니다.
```
bandgap = qeom_results.computed_energies[1] - qeom_results.computed_energies[0]
bandgap # in Hartree
```
## Qiskit Runtime을 사용하여 클라우드에서 VQE를 실행하기
Qiskit Runtime은 IBM Quantum이 제공하는 새로운 아키텍처로, 많은 반복이 필요한 계산을 간소화합니다. Qiskit runtime은 개선된 하이브리드 양자/고전 프로세스 내에서 실험의 실행 속도를 눈에 띄게 개선합니다.
인증된 사용자는 Qiskit Runtime을 사용해 자신과 다른 사용자가 사용할 수 있도록 Qiskit으로 작성된 양자 프로그램을 업로드할 수 있습니다. Qiskit 양자 프로그램은 Qiskit Runtime 프로그램이라고도 불리며 특정 입력을 받아 양자 및 고전적인 계산을 수행하고 원하는 경우 반응형으로 중간 결과를 제공하고 처리 결과를 반환하는 파이썬으로 작성된 코드를 의미합니다. 사용자 본인 혹은 다른 인증된 사용자는 프로그램 동작에 필요한 입력 매개 변수를 전송하여 업로드된 양자 프로그램을 호출해 사용할 수 있습니다.
<center><img src="resources/qiskit-runtime1.gif" width="600"></center>
<center><img src="resources/runtime_arch.png" width="600"></center>
Qiskit Runtime을 사용하여 VQE를 실행하기 위해서는, 로컬 VQE 실행 단계에서의 변화는 거의 없으므로, VQE 클래스를 VQEProgram 클래스로 교체해야 합니다. 두 가지 모두 동일한 MinimumEigensolver 인터페이스를 따르기때문에 compute_minimum_eigenvalue 메서드를 공유하여 알고리즘이 실행되고 같은 유형의 결과 개체가 반환합니다. 단지 이니셜라이저의 서명이 약간 다를 뿐입니다.
먼저 Qiskit Runtime 서비스에 액세스할 수 있는 공급자와 회로를 실행할 백엔드를 선택합니다.
Qiskit Runtime에 대한 추가정보는, [**VQEProgram**](https://qiskit.org/documentation/partners/qiskit_runtime/tutorials/vqe.html#Runtime-VQE:-VQEProgram)과 [**Leveraging Qiskit Runtime**](https://qiskit.org/documentation/nature/tutorials/07_leveraging_qiskit_runtime.html) 튜토리얼을 참고 하십시오.
```
from qc_grader.util import get_challenge_provider
provider = get_challenge_provider()
backend = provider.get_backend('ibmq_qasm_simulator')
from qiskit_nature.runtime import VQEProgram
error_threshold = 10 # mHartree
# for live plotting
pp = ProgressPlot(plot_names=['Energy'],
line_names=['Runtime VQE', f'Target + {error_threshold}mH', 'Target'])
intermediate_info = {
'nfev': [],
'parameters': [],
'energy': [],
'stddev': []
}
def callback(nfev, parameters, energy, stddev):
intermediate_info['nfev'].append(nfev)
intermediate_info['parameters'].append(parameters)
intermediate_info['energy'].append(energy)
intermediate_info['stddev'].append(stddev)
pp.update([[energy,exact_energy+error_threshold/1000, exact_energy]])
##############################
# Provide your code here
optimizer = {
'name': 'QN-SPSA', # leverage the Quantum Natural SPSA
# 'name': 'SPSA', # set to ordinary SPSA
'maxiter': 100,
}
runtime_vqe =
##############################
```
<div class="alert alert-block alert-success">
**도전 2f 채점 설명**
이 도전 과제의 채점은 이전의 연습들과 약간 다릅니다.
1. 우선 `prepare_ex2f`를 사용해 `runtime_vqe (VQEProgram)`, `qubit_converter (QubitConverter)`, `es_problem (ElectronicStructureProblem)`이 포함된 runtime job을 IBM Quantum으로 시뮬레이션 계산을 위해 전송합니다. 대기 상태에 따라 실행에 몇 분이 소요될 수 있습니다. 내부적으로 `prepare_ex2f`은 다음의 작업을 합니다:
```python
runtime_vqe_groundstate_solver = GroundStateEigensolver(qubit_converter, runtime_vqe)
runtime_vqe_result = runtime_vqe_groundstate_solver.solve(es_problem)
```
2. 작업이 완료된 후, `grade_ex2f`를 사용해 정답을 확인한 후 제출합니다.
```
# Submit a runtime job using the following code
from qc_grader import prepare_ex2f
runtime_job = prepare_ex2f(runtime_vqe, qubit_converter, es_problem)
# Check your answer and submit using the following code
from qc_grader import grade_ex2f
grade_ex2f(runtime_job)
print(runtime_job.result().get("eigenvalue"))
```
축하합니다! 첫 번째 Qiskit Runtime 프로그램을 제출하고 도전 과제를 성공했습니다.
하지만 즐거움은 끝나지 않았습니다! 여러분을 위해 양자 챌린지를 위한 전용 양자 시스템을 예약해 두었습니다. 채점되지 않는 보너스 연습으로 실제 양자 시스템에 VQE 런타임 작업을 실행해 보십시오!
<div class="alert alert-block alert-success">
** VQE를 실제 양자 시스템에서 실행시키기(부수적인)**
이 과제를 위해 전용 양자 시스템 [`ibm_perth`](https://quantum-computing.ibm.com/services?services=systems&system=ibm_perth)을 예약해 두었습니다. 실제 양자 시스템에 런타임 작업을 제출하려면 아래 단계를 따르십시오.
1. 백엔드 선택을 `ibm_perth`로 업데이트하고 `runtime_vqe`에 다시 전달하십시오.
```
backend = provider.get_backend('ibm_perth')
runtime_vqe = VQEProgram(...
backend=backend,
...)
```
2. `prepare_ex2f`의 `real_device`플래그를 `True`로 설정합니다.
3. `prepare_ex2f`를 실행하여 runtime 작업을 `ibm_perth`로 전달합니다.
</div>
<div class="alert alert-block alert-danger">
노트: Qiskit 런타임은 VQE 속도를 최대 5배까지 높입니다. 하지만 각 런타임 작업은 여전히 30~60분의 양자 프로세서 시간이 소요될 수 있습니다. 따라서 **작업** 을 완료하기 위한 대기열 시간은 작업을 제출하는 참가자 수에 따라 몇 시간 또는 며칠이 될 수도 있습니다.
모든 참가자에게 즐거운 경험을 제공하기 위해 먼저 시뮬레이터를 사용하여 다음 설정을 시도한 이후에만 실제 양자 시스템에 작업을 제출해 주십시오
1. `PartiyMapper`를 사용하고 `two_qubit_reduction=True`를 설정하여 필요한 큐비트 수를 2로 줄여서 VQE 프로그램이 바닥 상태 에너지에 더 빠르게 수렴하도록 합니다(더 적은 반복 횟수로).
1. 최적화 옵션 중 'maxiter=100 혹은 그 이하로 제한합니다. 시뮬레이터를 사용하여 최적의 낮은 반복 횟수를 찾습니다.
1. 시뮬레이터를 백엔드로 사용하여 `grade_ex2f`에 VQE를 전달하여 런타임 프로그램이 올바르게 작성 되었는지 먼저 확인합니다.
1. 더 많은 참가자가 실제 양자 시스템에서 런타임을 시도할 수 있도록 작업을 참가자당 1개로 제한합니다.
작업 실행이 너무 오래 걸리거나 챌린지가 종료되기 전에 실행할 수 없는 경우에도 걱정하지 마십시오. 이것은 부수적인 도전입니다. 실제 양자 시스템에서 작업을 실행하지 않고도 모든 도전 과제를 통과하고 디지털 배지를 취득 할 수 있습니다.
</div>
```
# Please change `real_device` flag to True if you want to send a job to the real quantum system
runtime_job_real_device = prepare_ex2f(runtime_vqe, qubit_converter, es_problem, real_device=False)
print(runtime_job_real_device.result().get("eigenvalue"))
```
## 추가 정보
**제작자:** Junye Huang, Samanvay Sharma
**한글번역:** 신소영, 김정원
**Version:** 1.0.0
| github_jupyter |
```
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import DataLoader
from torch.utils.data import sampler
import skorch
import torchvision.datasets as dset
import torchvision.transforms as T
import torchvision.models as models
import collections
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import cross_validate
import numpy as np
import matplotlib
# matplotlib.use('agg')
import matplotlib.pyplot as plt
from Conv3D_Dataset import ILDDataset
USE_GPU = True
dtype = torch.float32 # we will be using float throughout this tutorial
if USE_GPU and torch.cuda.is_available():
device = torch.device('cuda')
else:
device = torch.device('cpu')
# Constant to control how frequently we print train loss
print_every = 100
print('using device:', device)
NUM_TOTAL = 1979
NUM_TRAIN = 1600
#add path as absolute path for root dir
im_size = 128
lung_dataset_train = ILDDataset(csv_file=r'C:/Users/Akrofi/Desktop/CS 231/Project/train_labels.csv',
root_dir=r'C:/Users/Akrofi/Desktop/CS 231/Project/train',mask=True, train=True, resize=im_size)
#add path as absolute path for root dir
lung_dataset_test = ILDDataset(csv_file=r'C:/Users/Akrofi/Desktop/CS 231/Project/test_labels.csv',
root_dir=r'C:/Users/Akrofi/Desktop/CS 231/Project/test', mask=True, train=False, resize=im_size)
dataset_x = []
dataset_y = []
for i in range(20):
x = i
print(x)
a = lung_dataset_train[i][0].tolist()
dataset_x.append(a)
b = lung_dataset_train[i][1].tolist()
dataset_y.append(b)
a = np.asarray(dataset_x)
b = np.asarray(dataset_y)
print(len(a))
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(np.asarray(dataset_x),
np.asarray(dataset_y),
test_size=0.20,
random_state=42,
stratify = dataset_y)
print (y_test)
class_sample_count = np.array([len(np.where(y_train==t)[0]) for t in np.unique(y_train)])
weight = 1. / class_sample_count
samples_weight = np.array([weight[t] for t in y_train])
samples_weight = torch.from_numpy(samples_weight)
sampler = WeightedRandomSampler(samples_weight.type('torch.DoubleTensor'), len(samples_weight))
trainDataset = torch.utils.data.TensorDataset(torch.FloatTensor(X_train), torch.LongTensor(y_train.astype(int)))
validDataset = torch.utils.data.TensorDataset(torch.FloatTensor(X_test), torch.LongTensor(y_test.astype(int)))
trainLoader = torch.utils.data.DataLoader(dataset = trainDataset, batch_size=bs, num_workers=1, sampler = sampler)
testLoader = torch.utils.data.DataLoader(dataset = validDataset, batch_size=bs, shuffle=False, num_workers=1)
for i, (data, target) in enumerate(trainLoader):
print ("batch index {}, 0/1: {}/{}".format(
i, len(np.where(target.numpy()==0)[0]), len(np.where(target.numpy()==1)[0])))
loader_train = DataLoader(lung_dataset_train, batch_size= 16,
sampler=sampler.SubsetRandomSampler(range(NUM_TRAIN)))
loader_val = DataLoader(lung_dataset_train, batch_size=16,
sampler=sampler.SubsetRandomSampler(range(NUM_TRAIN, NUM_TOTAL)))
loader_test = DataLoader(lung_dataset_test, batch_size=16)
#show datasample
sample = lung_dataset_train[810]
plt.imshow(sample[0][3], cmap='gray')
plt.show()
print("label: " + str(sample[1]))
def flatten(x):
N = x.shape[0] # read in N, C, H, W
return x.view(N, -1) # "flatten" the C * H * W values into a single vector per image
class Flatten(nn.Module):
def forward(self, x):
return flatten(x)
def get_model():
"""
Used to fetch model for classification
"""
in_channel = 1
channel_1 = 32
channel_2 = 64
channel_3 = 32
num_classes = 3
model = nn.Sequential(
nn.Conv3d(in_channel,channel_1, kernel_size=5, padding=2, stride= 1 ),
nn.ReLU(),
nn.Conv3d(channel_1, channel_2, kernel_size=3, padding=1, stride=1),
nn.ReLU(),
nn.BatchNorm3d(num_features= channel_2),
nn.MaxPool3d(kernel_size=1,stride=1),
nn.Conv3d(channel_2, channel_3, kernel_size=3, padding=1, stride=1),
nn.ReLU(),
Flatten(),
nn.Linear(2097152, num_classes)
)
return model
def check_accuracy(loader, model, val =False, train=False):
if train == True:
print('Checking accuracy on training set')
elif val == True:
print('Checking accuracy on validation set')
else:
print('Checking accuracy on test set')
num_correct = 0
num_samples = 0
model.eval() # set model to evaluation mode
with torch.no_grad():
for x, y in loader:
x = x.to(device=device, dtype=dtype) # move to device, e.g. GPU
y = y.to(device=device, dtype=torch.long)
[N,C,H,W] = [*x.size()]
# print(y)
# a = np.split(y,4)
# y = np.zeros(len(a))
# for i in range(len(a)):
# b = collections.Counter(a[i]).most_common()[0][0]
# y[i] = (b)
# y = torch.LongTensor(y)
# y = y.to(device=device, dtype=torch.long)
scores = model(x.view(N, 1 , C, H , W))
_, preds = scores.max(1)
num_correct += (preds == y).sum()
num_samples += preds.size(0)
acc = float(num_correct) / num_samples
print('Got %d / %d correct (%.2f)' % (num_correct, num_samples, 100 * acc))
return acc
def train(model, optimizer, epochs=1, overfit=False):
"""
Inputs:
- model: A PyTorch Module giving the model to train.
- optimizer: An Optimizer object we will use to train the model
- epochs: (Optional) A Python integer giving the number of epochs to train for
Returns: Nothing, but prints model accuracies during training.
"""
model = model.to(device=device) # move the model parameters to CPU/GPU
best_acc = 0;
for e in range(epochs):
for t, (x, y) in enumerate(loader_train):
model.train() # put model to training mode
x = x.to(device=device, dtype=dtype) # move to device, e.g. GPU
y = y.to(device=device, dtype=torch.long)
[N,H,C,W] = [*x.size()]
# print(y)
# a = np.split(y,4)
# y = np.zeros(len(a))
# for i in range(len(a)):
# b = collections.Counter(a[i]).most_common()[0][0]
# y[i] = b
# y = torch.LongTensor(y)
# y = y.to(device=device, dtype=torch.long)
weights = torch.cuda.FloatTensor([1,1,1])
scores = model(x.view(N, 1, C, H , W))
Loss = nn.CrossEntropyLoss(weight = weights)
loss = Loss(scores, y)
reg = torch.tensor(2.5e-2, device= device, dtype=dtype)
l2_reg = torch.tensor(0. , device= device, dtype=dtype)
for param in model.parameters():
l2_reg += torch.norm(param)
loss += reg * l2_reg
# Zero out all of the gradients for the variables which the optimizer
# will update.
optimizer.zero_grad()
# This is the backwards pass: compute the gradient of the loss with
# respect to each parameter of the model.
loss.backward()
# Actually update the parameters of the model using the gradients
# computed by the backwards pass.
optimizer.step()
# if t % print_every == 0:
print('Iteration %d, loss = %.4f' % (t, loss.item()))
if(overfit):
check_accuracy(loader_train, model, _, train=True)
acc = check_accuracy(loader_validation, model, True, _)
if acc > best_acc:
best_acc = acc
best_model = model
print()
return best_model
model1 = get_model()
optimizer = optim.Adam(model1.parameters(), lr = 0.0001, weight_decay= 1e-4)
best_model = train(model1, optimizer, epochs= 10, overfit=True)
#Check Test set
check_accuracy(loader_test, best_model)
# Define model
```
| github_jupyter |
<a href="https://colab.research.google.com/github/sudeepds/mtech-datascience/blob/main/dvi_a2_ps6_wb.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
<img src = "https://www.bits-pilani.ac.in/Uploads/Campus/BITS_university_logo.gif">
<h1><center>Work Integrated Learning Programmes Division<br>
M.Tech (Data Science and Engineering)<br> Data Visualization And Interpretation (DSECL ZG555))<br>
First Semester, 2021-22
</center></h1>
<h2><center>Assignment 2 – PS6 [Weightage 13%]</center></h2>
## Instructions
<ol>
<li>Do not change the name of the data file that is shared with the problem statement.</li>
<li>If intermediate data files are created, retain in the present working directory and attach them during submission.</li>
<li>Retain the data file in the same directory as that of this workbook.</li>
<li>Retain the Visualizations that is produced in the file. Don't clear them away.</li>
<li>Submit only the .IPYNB file. Intermediate files to be attached as mentioned in (2).</li>
<li><b><div class="alert-warning">All the visuals should adhere to the visualization principles learnt in the Course and must be presentation ready.Most effective visuals would fetch maximum credits</div></b></li>
<li><b><div class="alert-warning">Submissions done via means other than CANVAS will strictly be NOT graded.</div></b></li>
</ol>
<style>
table {
font-family: arial, sans-serif;
border-collapse: collapse;
width: 100%;
}
td, th {
border: 1px solid #dddddd;
text-align: left;
padding: 8px;
}
tr:nth-child(even) {
background-color: #dddddd;
}
</style>
<h2>Group No: 006</h2>
<table>
<tr>
<th>Full Name</th>
<th>BITS ID</th>
</tr>
<tr>
<td>Lalu George</td>
<td>2020sc04489</td>
</tr>
<tr>
<td>Abhilash Pillai</td>
<td>2020sc04615</td>
</tr>
<tr>
<td>Sudeep Ghosh</td>
<td>2020sc04781</td>
</tr>
</table>
<h1>Objective</h1>
<h4>Analyse the scenario of automobile production in India.</h4>
## Download and Prep the Data: 1 Mark
<h4>Import the libraries needed</h4>
```
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
```
<h4> Load data and store in dataframe </h4>
```
df = pd.read_csv('dvi-a2-ps6-data.csv')
```
<h4>Find out what type of variables you are dealing with. This will help you find the right visualization method for that variable.</p>
```
df.info()
df.describe()
```
<h4>Perform data cleaning and mention the steps</h4>
```
# Checking number of null or NaN values
df.isnull().sum()
# Creating a new dataframe for data attributes
new_df= df.drop(['Category', 'Segment'], axis=1)
# Doing row wise forward fill
new_df= new_df.ffill(axis = 1)
# Filling 0 for the remaining NaN values.
new_df.fillna(value=0, inplace=True)
new_df
```
Steps performed for data cleaning:
> * Row wise forward fill for NaN values.
> * Performed 0 fill for NaN values till 2005-06 for Electric Two Wheelers.
# Visualisation Questions - 2 X 5 = 10 Marks
### Question 1
<h4>Compare the production of passenger vehicles and commercial vehicles in India from 2001-02 to 2012-13, in a single visual.<br><br>
Write the python code in the below cell to create appropriate visual to perform the above task.
<h4>Answer in markdown cells below the visual <br><br>
1.Summarise your findings from the visual.<br>
2.The reason for selecting the chart type you did. <br>
3.Mention the pre-attentive attributes used.(atleast 2)<br>
4.Mention the gestalt principles used.(atleast 2)<br>
5.Fill in the blanks:<br><br>
<i>1.In percentage terms, the total growth recorded for passenger vehicles is ______ during the period from 2001-02 to 2012-13.</i><br><br>
<i>2.In percentage terms, the total growth recorded for commercial vehicles is ______ during the period from 2001-02 to 2012-13.
```
new_df= df.drop(['Category', 'Segment'], axis=1)
x1 = new_df.columns
y1 = new_df.iloc[df.index[df['Segment'] == 'Total Commercial Vehicles (CVs)']].values.flatten().tolist()
y2 = new_df.iloc[df.index[df['Segment'] == 'Total Passenger Vehicles (PVs)']].values.flatten().tolist()
barWidth = 0.25
fig = plt.subplots(figsize =(12, 8))
br1 = np.arange(len(y1))
br2 = [x + barWidth for x in br1]
# Make the plot
plt.bar(br1, y1, color ='orange', width = barWidth,
edgecolor ='grey', label ='CVs')
plt.bar(br2, y2, color ='royalblue', width = barWidth,
edgecolor ='grey', label ='PVs')
# Adding Xticks
plt.xlabel('Year wise production', fontweight ='bold', fontsize = 15)
plt.ylabel('Numbers Produced(in lakhs)', fontweight ='bold', fontsize = 15)
plt.xticks([r + barWidth for r in range(len(y1))],
new_df.columns)
plt.legend()
plt.show()
# Calculating percentage
## Passenger Vehicles 2001-02 to 2012-13.
pv_df = new_df.iloc[df.index[df['Segment'] == 'Total Passenger Vehicles (PVs)']]
percentage = ((pv_df['2012-13'] - pv_df['2001-02'])/pv_df['2012-13']) * 100
print('In percentage terms, the total growth recorded for passenger vehicles is ', percentage.values[0],' during the period from 2001-02 to 2012-13')
## Commercial Vehicles 2001-02 to 2012-13.
cv_df = new_df.iloc[df.index[df['Segment'] == 'Total Commercial Vehicles (CVs)']]
percentage = ((cv_df['2012-13'] - cv_df['2001-02'])/cv_df['2012-13']) * 100
print('In percentage terms, the total growth recorded for commercial vehicles is ', percentage.values[0],' during the period from 2001-02 to 2012-13')
```
1.Summarise your findings from the visual:
* Passenger vehicles have seen a steady growth in production except between years 2008-09 and 2010-11 when it grew at a faster rate.
* Unlike Commercial Vehicles, Passenger vehicles have not seen downward trend in any year.
* Commercial vehicles have seen both upward and downward trends. A steady growth between year 2001-07 and then a fall in 2008-09. After that it grew again till 20011-12 before falling again.
* The ratio of vehicles produced between Passenger and commercial is around 3:1 in almost every year.
2.The reason for selecting the chart type you did: Side by side bar chart gives a better year on year comparison between both the categories.
3.Mention the pre-attentive attributes used.(atleast 2): Preattentive attributes used are
* Color
* Length: Length of each bar gives an impression of the volume produced.
4.Mention the gestalt principles used.(atleast 2):
* Proximity
* Figure and Ground
### Question 2
<h4>Analyse the passenger vehicle production(segment wise) during 2001-02 to 2012-13<br><br>
Write the python code in the below cell to create the appropriate visual(single visual) to perform the above task .
<h4>Answer in markdown cells below the visual <br><br>
1.Summarise your findings from the visual.<br>
2.The reason for selecting the chart type you did. <br>
3.Mention the pre-attentive attributes used.(atleast 2)<br>
4.Mention the gestalt principles used.(atleast 2)<br>
5.Fill in the blanks.<br>
<i>1.In percentage terms, the growth of production of passenger cars in India was ________ during the period from 2001-02 to 2012-13</i><br><br>
<i>2.In percentage terms, the growth the production of Multi-Utility Vehicles in India was ________ during the period from 2001-02 to 2012-13.
```
df.head(2)
x1 = new_df.columns
y1 = new_df.iloc[df.index[df['Segment'] == 'Passenger Cars']].values.flatten().tolist()
y2 = new_df.iloc[df.index[df['Segment'] == 'Multi-Utility Vehicles']].values.flatten().tolist()
barWidth = 0.25
fig = plt.subplots(figsize =(12, 8))
br1 = np.arange(len(y1))
br2 = [x + barWidth for x in br1]
# Make the plot
plt.bar(br1, y1, color ='royalblue', width = barWidth,
edgecolor ='grey', label ='Passenger Cars')
plt.bar(br2, y2, color ='orange', width = barWidth,
edgecolor ='grey', label ='Multi-Utility Vehicles')
# Adding Xticks
plt.xlabel('Year wise production', fontweight ='bold', fontsize = 15)
plt.ylabel('Numbers Produced(in lakhs)', fontweight ='bold', fontsize = 15)
plt.xticks([r + barWidth for r in range(len(y1))],
new_df.columns)
plt.legend()
plt.show()
# Creation of Data
x1 = new_df.columns
y1 = new_df.iloc[df.index[df['Segment'] == 'Passenger Cars']].values.flatten().tolist()
y2 = new_df.iloc[df.index[df['Segment'] == 'Multi-Utility Vehicles']].values.flatten().tolist()
# Plotting the Data
plt.plot(x1, y1, label='Passenger Cars')
plt.plot(x1, y2, label='Multi-Utility Vehicles')
plt.xlabel('Yearwise Production')
plt.ylabel('Numbers Produced(in lakhs)')
plt.title("Passenger Vehicle Yearwise Trend")
plt.rcParams["figure.figsize"] = (20,8)
plt.plot(y1, color = 'royalblue', linewidth = 3)
plt.plot(y2, color = 'orange')
plt.legend()
# Calculating percentage
## Passenger Vehicles 2001-02 to 2012-13.
pv_df = new_df.iloc[df.index[df['Segment'] == 'Passenger Cars']]
percentage = ((pv_df['2012-13'] - pv_df['2001-02'])/pv_df['2012-13']) * 100
print('In percentage terms, the growth of production of passenger cars in India was ', percentage.values[0],' during the period from 2001-02 to 2012-13')
## Commercial Vehicles 2001-02 to 2012-13.
cv_df = new_df.iloc[df.index[df['Segment'] == 'Multi-Utility Vehicles']]
percentage = ((cv_df['2012-13'] - cv_df['2001-02'])/cv_df['2012-13']) * 100
print('In percentage terms, the growth the production of Multi-Utility Vehicles in India was ', percentage.values[0],' during the period from 2001-02 to 2012-13')
```
1. Summarise your findings from the visual:
> * Passenger Cars segment grew faster during the period 2001-13 than Multi Utility Vehicles.
> * Both segments showed an opposite trend during the period 2010-13. When Passenger car grew, multi-utility fell where as when Multi-Utility grew Passenger car fell.
> * Passenger cars saw the fastest growth during 2008-12 where as Multi-Utility vehicles saw fastest growth during 2008-11.
2. The reason for selecting the chart type you did: This chart shows a year wise trend comparison between both the segments.
3. Mention the pre-attentive attributes used.(atleast 2):
> * Color
> * Size
4. Mention the gestalt principles used.(atleast 2):
> * Figure and Ground
> * Continuity
### Question 3
<h4>Compare the production of 2 wheeler and three wheelers in India from 2001-02 to 2012-13, in a single visual.<br><br>
Write the python code in the below cell to create appropriate visual to perform the above task.
<h4>Answer in markdown cells below the visual <br><br>
1.Summarise your findings from the visual.<br>
2.The reason for selecting the chart type you did. <br>
3.Mention the pre-attentive attributes used.(atleast 2)<br>
4.Mention the gestalt principles used.(atleast 2)<br>
```
x1 = new_df.columns
y1 = new_df.iloc[df.index[df['Segment'] == 'Total Two wheelers']].values.flatten().tolist()
y2 = new_df.iloc[df.index[df['Segment'] == 'Total Three Wheelers']].values.flatten().tolist()
barWidth = 0.25
fig = plt.subplots(figsize =(12, 8))
br1 = np.arange(len(y1))
br2 = [x + barWidth for x in br1]
# Make the plot
plt.bar(br1, y1, color ='royalblue', width = barWidth,
edgecolor ='grey', label ='Total Two Wheelers')
plt.bar(br2, y2, color ='orange', width = barWidth,
edgecolor ='grey', label ='Total Three Wheelers')
# Adding Xticks
plt.xlabel('Year wise production', fontweight ='bold', fontsize = 15)
plt.ylabel('Numbers Produced(in millions)', fontweight ='bold', fontsize = 15)
plt.xticks([r + barWidth for r in range(len(y1))],
new_df.columns)
plt.legend()
plt.show()
```
1.Summarise your findings from the visual.
> * Number of two wheeler saw an steady growth except a dip in 2007-08.
> * Two Wheelers saw it fastest growth in the period 2008-12.
> * Three Wheelers saw a dip in the period 2007-09 and again in 2012-13.
> * The ratio between numbers of Two Wheelers and Three wheelers has been close to 12:1.
2.The reason for selecting the chart type you did: It provides a side by side year wise comparison of both the categories.
3.Mention the pre-attentive attributes used.(atleast 2):
> * Color
> * Length: Length of each bar gives an impression of the volume produced.
4.Mention the gestalt principles used.(atleast 2):
> * Proximity
> * Figure and Ground
### Question 4
<h4>Analyse the two wheeler production in India(segment wise) during 2001-02 to 2012-13.<br><br>
Write the python code in the below cell to create the appropriate visual(single visual) to perform the above task .
<h4>Answer in markdown cells below the visual <br><br>
1.Summarise your findings from the visual.<br>
2.The reason for selecting the chart type you did. <br>
3.Mention the pre-attentive attributes used.(atleast 2)<br>
4.Mention the gestalt principles used.(atleast 2)<br>
```
# Creation of Data
x1 = new_df.columns
y1 = new_df.iloc[df.index[df['Segment'] == 'Scooter/Scooterettee']].values.flatten().tolist()
y2 = new_df.iloc[df.index[df['Segment'] == 'Motorcycles/Step-Throughs']].values.flatten().tolist()
y3 = new_df.iloc[df.index[df['Segment'] == 'Mopeds']].values.flatten().tolist()
y4 = new_df.iloc[df.index[df['Segment'] == 'Electric Two Wheelers']].values.flatten().tolist()
plt.xlabel('Yearwise Production')
plt.ylabel('Numbers Produced(in 10 millions)')
plt.title("Two Wheelers Yearwise Trend")
plt.rcParams["figure.figsize"] = (20,8)
# Plotting the Data
plt.plot(x1, y1, label='Scooter/Scooterettee', color='c')
plt.plot(x1, y2, label='Motorcycles/Step-Throughs', color='royalblue', linewidth=3)
plt.plot(x1, y3, label='Mopeds', color='orange')
plt.plot(x1, y4, label='Electric Two Wheelers', color='green')
plt.legend()
```
1.Summarise your findings from the visual.
> * Motorcycles/Step-Throughs segment registered a substantially faster growth than any other segment.
> * Mopeds production remained approximately same throughtout the period of 2001-13.
> * Scooter/Scooterettee segment showed significant growth only in the period 2009-13.
> * Electric Two Wheelers production remained almost consistent throughout the period 2006-10.
> * Numbers of Motorcycles/Step-Throughs is substantially higher than any other sector.
2.The reason for selecting the chart type you did: This chart gives a better visual of the yearwise trend of each segment while providing a inter segment comparison as well.
3.Mention the pre-attentive attributes used.(atleast 2):
> * Color
> * Size
4.Mention the gestalt principles used.(atleast 2
> * Figure and Ground
> * Continuity
### Question 5
<h4>Analyse the three wheeler production in India(segment wise) during 2001-02 to 2012-13.<br><br>
Write the python code in the below cell to create the appropriate visual(single visual) to perform the above task .
<h4>Answer in markdown cells below the visual <br><br>
1.Summarise your findings from the visual.<br>
2.The reason for selecting the chart type you did. <br>
3.Mention the pre-attentive attributes used.(atleast 2)<br>
4.Mention the gestalt principles used.(atleast 2)<br>
```
# Creation of Data
x1 = new_df.columns
y1 = new_df.iloc[df.index[(df['Segment'] == 'Passenger Carriers') & (df['Category'] == 'Three Wheelers')]].values.flatten().tolist()
y2 = new_df.iloc[df.index[(df['Segment'] == 'Goods Carriers') & (df['Category'] == 'Three Wheelers')]].values.flatten().tolist()
# Plotting the Data
plt.plot(x1, y1, label='Passenger Carriers', linewidth=3)
plt.plot(x1, y2, label='Goods Carriers')
plt.xlabel('Yearwise Production')
plt.ylabel('Numbers Produced')
plt.title("Three Wheelers Yearwise Trend")
plt.rcParams["figure.figsize"] = (20,8)
plt.legend()
```
1.Summarise your findings from the visual:
> * Both the segment showed an opposite trend during 2007-10.
> * Passenger Carrier has overall faster growth than Goods Carriers.
2.The reason for selecting the chart type you did: This chart gives an overall trend comparison between both the segments.
3.Mention the pre-attentive attributes used.(atleast 2):
> * Color
> * Size
4.Mention the gestalt principles used.(atleast 2)
> * Continuity
> * Figure and Ground
# Group's choice-2 Marks
#### Frame 1 (more) question which will help in the EDA(Exploratory Data Analysis) of the given data set and answer the same using the best visual.
1. Write the question in a markdown cell
2. Below the question, in a coding cell, write the python code to create the visual to answer the question
<h4> Answer in markdown cells below the visual <br><br>
1.Summarise your findings from the visual.<br>
2.The reason for selecting the chart type you did. <br>
3.Mention the pre-attentive attributes used.(atleast 2)<br>
4.Mention the gestalt principles used.(atleast 2)<br>
Question: Analyse the yearwise trend of M & HCVs and LCVs.
```
# Creation of Data
x1 = new_df.columns
y1 = new_df.iloc[df.index[df['Segment'] == 'Total M & HCVs']].values.flatten().tolist()
y2 = new_df.iloc[df.index[df['Segment'] == 'Total LCVs']].values.flatten().tolist()
# Plotting the Data
plt.plot(x1, y1, label='M & HCVs', color='lightgrey')
plt.plot(x1, y2, label='LCVs', color='royalblue', linewidth=3)
plt.xlabel('Yearwise Production')
plt.ylabel('Numbers Produced')
plt.title("Heavy, Mid and Light CVs Yearwise Trend")
plt.rcParams["figure.figsize"] = (20,8)
plt.legend()
```
1.Summarise your findings from the visual:
> * Both the segments showed a similar growth rate between year 2001-05.
> * LCVs surpasses M & HCVs between 2008-09. Post that, LCVs showed a higher growth rate.
2.The reason for selecting the chart type you did: As the intention was to analyse yearwise trend, trend lines suited best for it.
3.Mention the pre-attentive attributes used.(atleast 2):
> * Color
> * Size
4.Mention the gestalt principles used.(atleast 2)
> * Continuity
> * Figure and Ground
<h1><center> ************ END OF ASSIGNMENT ****************</center></h1>
| github_jupyter |
# 机器学习练习 1 - 线性回归
这个是另一位大牛写的,作业内容在根目录: [作业文件](ex1.pdf)
代码修改并注释:黄海广,haiguang2000@qq.com
## 单变量线性回归
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
path = 'ex1data1.txt'
data = pd.read_csv(path, header=None, names=['Population', 'Profit'])
data.head()
data.describe()
```
看下数据长什么样子
```
data.plot(kind='scatter', x='Population', y='Profit', figsize=(12,8))
plt.show()
```
现在让我们使用梯度下降来实现线性回归,以最小化成本函数。 以下代码示例中实现的方程在“练习”文件夹中的“ex1.pdf”中有详细说明。
首先,我们将创建一个以参数θ为特征函数的代价函数
$$J\left( \theta \right)=\frac{1}{2m}\sum\limits_{i=1}^{m}{{{\left( {{h}_{\theta }}\left( {{x}^{(i)}} \right)-{{y}^{(i)}} \right)}^{2}}}$$
其中:\\[{{h}_{\theta }}\left( x \right)={{\theta }^{T}}X={{\theta }_{0}}{{x}_{0}}+{{\theta }_{1}}{{x}_{1}}+{{\theta }_{2}}{{x}_{2}}+...+{{\theta }_{n}}{{x}_{n}}\\]
```
def computeCost(X, y, theta):
inner = np.power(((X * theta.T) - y), 2)
return np.sum(inner) / (2 * len(X))
```
让我们在训练集中添加一列,以便我们可以使用向量化的解决方案来计算代价和梯度。
```
data.insert(0, 'Ones', 1)
```
现在我们来做一些变量初始化。
```
# set X (training data) and y (target variable)
cols = data.shape[1]
X = data.iloc[:,0:cols-1]#X是所有行,去掉最后一列
y = data.iloc[:,cols-1:cols]#X是所有行,最后一列
```
观察下 X (训练集) and y (目标变量)是否正确.
```
X.head()#head()是观察前5行
y.head()
```
代价函数是应该是numpy矩阵,所以我们需要转换X和Y,然后才能使用它们。 我们还需要初始化theta。
```
X = np.matrix(X.values)
y = np.matrix(y.values)
theta = np.matrix(np.array([0,0]))
```
theta 是一个(1,2)矩阵
```
theta
```
看下维度
```
X.shape, theta.shape, y.shape
```
计算代价函数 (theta初始值为0).
```
computeCost(X, y, theta)
```
# batch gradient decent(批量梯度下降)
$${{\theta }_{j}}:={{\theta }_{j}}-\alpha \frac{\partial }{\partial {{\theta }_{j}}}J\left( \theta \right)$$
```
def gradientDescent(X, y, theta, alpha, iters):
temp = np.matrix(np.zeros(theta.shape))
parameters = int(theta.ravel().shape[1])
cost = np.zeros(iters)
for i in range(iters):
error = (X * theta.T) - y
for j in range(parameters):
term = np.multiply(error, X[:,j])
temp[0,j] = theta[0,j] - ((alpha / len(X)) * np.sum(term))
theta = temp
cost[i] = computeCost(X, y, theta)
return theta, cost
```
初始化一些附加变量 - 学习速率α和要执行的迭代次数。
```
alpha = 0.01
iters = 1000
```
现在让我们运行梯度下降算法来将我们的参数θ适合于训练集。
```
g, cost = gradientDescent(X, y, theta, alpha, iters)
g
```
最后,我们可以使用我们拟合的参数计算训练模型的代价函数(误差)。
```
computeCost(X, y, g)
```
现在我们来绘制线性模型以及数据,直观地看出它的拟合。
```
x = np.linspace(data.Population.min(), data.Population.max(), 100)
f = g[0, 0] + (g[0, 1] * x)
fig, ax = plt.subplots(figsize=(12,8))
ax.plot(x, f, 'r', label='Prediction')
ax.scatter(data.Population, data.Profit, label='Traning Data')
ax.legend(loc=2)
ax.set_xlabel('Population')
ax.set_ylabel('Profit')
ax.set_title('Predicted Profit vs. Population Size')
plt.show()
```
由于梯度方程式函数也在每个训练迭代中输出一个代价的向量,所以我们也可以绘制。 请注意,代价总是降低 - 这是凸优化问题的一个例子。
```
fig, ax = plt.subplots(figsize=(12,8))
ax.plot(np.arange(iters), cost, 'r')
ax.set_xlabel('Iterations')
ax.set_ylabel('Cost')
ax.set_title('Error vs. Training Epoch')
plt.show()
```
## 多变量线性回归
练习1还包括一个房屋价格数据集,其中有2个变量(房子的大小,卧室的数量)和目标(房子的价格)。 我们使用我们已经应用的技术来分析数据集。
```
path = 'ex1data2.txt'
data2 = pd.read_csv(path, header=None, names=['Size', 'Bedrooms', 'Price'])
data2.head()
```
对于此任务,我们添加了另一个预处理步骤 - 特征归一化。 这个对于pandas来说很简单
```
data2 = (data2 - data2.mean()) / data2.std()
data2.head()
```
现在我们重复第1部分的预处理步骤,并对新数据集运行线性回归程序。
```
# add ones column
data2.insert(0, 'Ones', 1)
# set X (training data) and y (target variable)
cols = data2.shape[1]
X2 = data2.iloc[:,0:cols-1]
y2 = data2.iloc[:,cols-1:cols]
# convert to matrices and initialize theta
X2 = np.matrix(X2.values)
y2 = np.matrix(y2.values)
theta2 = np.matrix(np.array([0,0,0]))
# perform linear regression on the data set
g2, cost2 = gradientDescent(X2, y2, theta2, alpha, iters)
# get the cost (error) of the model
computeCost(X2, y2, g2)
```
我们也可以快速查看这一个的训练进程。
```
fig, ax = plt.subplots(figsize=(12,8))
ax.plot(np.arange(iters), cost2, 'r')
ax.set_xlabel('Iterations')
ax.set_ylabel('Cost')
ax.set_title('Error vs. Training Epoch')
plt.show()
```
我们也可以使用scikit-learn的线性回归函数,而不是从头开始实现这些算法。 我们将scikit-learn的线性回归算法应用于第1部分的数据,并看看它的表现。
```
from sklearn import linear_model
model = linear_model.LinearRegression()
model.fit(X, y)
```
scikit-learn model的预测表现
```
x = np.array(X[:, 1].A1)
f = model.predict(X).flatten()
fig, ax = plt.subplots(figsize=(12,8))
ax.plot(x, f, 'r', label='Prediction')
ax.scatter(data.Population, data.Profit, label='Traning Data')
ax.legend(loc=2)
ax.set_xlabel('Population')
ax.set_ylabel('Profit')
ax.set_title('Predicted Profit vs. Population Size')
plt.show()
```
# 4. normal equation(正规方程)
正规方程是通过求解下面的方程来找出使得代价函数最小的参数的:$\frac{\partial }{\partial {{\theta }_{j}}}J\left( {{\theta }_{j}} \right)=0$ 。
假设我们的训练集特征矩阵为 X(包含了${{x}_{0}}=1$)并且我们的训练集结果为向量 y,则利用正规方程解出向量 $\theta ={{\left( {{X}^{T}}X \right)}^{-1}}{{X}^{T}}y$ 。
上标T代表矩阵转置,上标-1 代表矩阵的逆。设矩阵$A={{X}^{T}}X$,则:${{\left( {{X}^{T}}X \right)}^{-1}}={{A}^{-1}}$
梯度下降与正规方程的比较:
梯度下降:需要选择学习率α,需要多次迭代,当特征数量n大时也能较好适用,适用于各种类型的模型
正规方程:不需要选择学习率α,一次计算得出,需要计算${{\left( {{X}^{T}}X \right)}^{-1}}$,如果特征数量n较大则运算代价大,因为矩阵逆的计算时间复杂度为$O(n3)$,通常来说当$n$小于10000 时还是可以接受的,只适用于线性模型,不适合逻辑回归模型等其他模型
```
# 正规方程
def normalEqn(X, y):
theta = np.linalg.inv(X.T@X)@X.T@y#X.T@X等价于X.T.dot(X)
return theta
final_theta2=normalEqn(X, y)#感觉和批量梯度下降的theta的值有点差距
final_theta2
#梯度下降得到的结果是matrix([[-3.24140214, 1.1272942 ]])
```
在练习2中,我们将看看分类问题的逻辑回归。
| github_jupyter |
# PyPi Releases
PyPi Releases notebook has the basic structure to release a package at PyPi using a token.
http://pytzen.com
## PYPI
> The Python Package Index (PyPI) is a repository of software for the Python programming language. PyPI helps you find and install software developed and shared by the Python community. Package authors use PyPI to distribute their software. https://pypi.org/
## Update Version
```
# Get the PyPi token and package path from the config file
%run ../config.ipynb
# If it is the first upload to PyPi
first_run = False
old_version = '0.0.3'
new_version = '0.0.4'
```
### Package structure
```
# Change the working directory to the package repo path
%cd {PATH_RESEARCH}
!tree -L 1
```
### Configuration File
This the simple PYTZEN configuration file that will have the version updated:
```
with open('setup.cfg', 'r') as file:
print(file.read())
```
### `__init__.py` file update
This is the `__init__.py` file that will have the version updated:
```
with open('src/pytzen/__init__.py', 'r') as file:
print(file.read())
```
### Files Update
```
def update_version(path):
with open(path, 'r') as file_read:
content = file_read.read().replace(old_version, new_version)
with open(path, 'w') as file_write:
file_write.write(content)
update_version('setup.cfg')
update_version('src/pytzen/__init__.py')
```
### Update Dependences and Built Version
```
# Update the used libraries
!pip install -U pip -U build -U twine
if not first_run:
# Remove the distribution folder
!rm -rf dist
# Remove the egg folder
!rm -rf src/pytzen/pytzen.egg-info
# Build new versions of the deleted material
!python3 -m build
```
## Upload the new built version
### PYPI API Token
>API tokens provide an alternative way (instead of username and password) to authenticate when uploading packages to PyPI. You can create a token for an entire PyPI account, in which case, the token will work for all projects associated with that account. Alternatively, you can limit a token's scope to a specific project. https://pypi.org/help/#apitoken
### Twine
> Twine is a utility for publishing Python packages to PyPI and other repositories. It provides build system independent uploads of source and binary distribution artifacts for both new and existing projects. https://twine.readthedocs.io/en/stable/
```
-u USERNAME, --username USERNAME
The username to authenticate to the repository
(package index) as. (Can also be set via
TWINE_USERNAME environment variable.)
-p PASSWORD, --password PASSWORD
The password to authenticate to the repository
(package index) with. (Can also be set via
TWINE_PASSWORD environment variable.)
```
### Upload to PyPi
```
!twine upload dist/* -u __token__ -p {PYPI_TOKEN}
```
## References
https://semver.org/
https://git-scm.com/book/en/v2/Git-Basics-Tagging
https://stackoverflow.com/questions/27951206/bash-cell-magic-in-ipython-script
https://pypi.org/
https://packaging.python.org/en/latest/tutorials/packaging-projects/
https://stackoverflow.com/questions/52700692/a-guide-for-updating-packages-on-pypi
https://pypi.org/help/#apitoken
https://twine.readthedocs.io/en/stable/
| github_jupyter |
# Tutorial: Computing with shapes of landmarks in Kendall shape spaces
Lead author: Nina Miolane.
In this tutorial, we show how to use geomstats to perform a shape data analysis. Specifically, we aim to study the difference between two groups of data:
- optical nerve heads that correspond to normal eyes,
- optical nerve heads that correspond to glaucoma eyes.
We wish to investigate if there is a difference in these two groups, and if this difference is a difference in sizes of the optical nerve heads, or a difference in shapes (where the size has been quotiented out).
<img src="figures/optic_nerves.png" />
## Set up
```
import os
import sys
import warnings
sys.path.append(os.path.dirname(os.getcwd()))
warnings.filterwarnings('ignore')
%matplotlib inline
import matplotlib.colors as colors
import matplotlib.patches as mpatches
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from mpl_toolkits.mplot3d.art3d import Poly3DCollection
import geomstats.backend as gs
import geomstats.datasets.utils as data_utils
from geomstats.geometry.pre_shape import PreShapeSpace, KendallShapeMetric
```
We import the dataset of the optical nerve heads from 22 images of Rhesus monkeys’ eyes (11 monkeys), available in [[PE2015]](#References).
For each monkey, an experimental glaucoma was introduced in one eye, while the second
eye was kept as control. One seeks to observe differences between the glaucoma and the
control eyes. On each image, 5 anatomical landmarks were recorded:
- 1st landmark: superior aspect of the retina,
- 2nd landmark: side of the retina closest to the temporal bone of the skull,
- 3rd landmark: nose side of the retina,
- 4th landmark: inferior point,
- 5th landmark: optical nerve head deepest point.
Label 0 refers to a normal eye, and Label 1 to an eye with glaucoma.
```
nerves, labels, monkeys = data_utils.load_optical_nerves()
print(nerves.shape)
print(labels)
print(monkeys)
```
We extract the landmarks' sets corresponding to the two eyes' nerves of the first monkey, with their corresponding labels.
```
two_nerves = nerves[monkeys==0]
print(two_nerves.shape)
two_labels = labels[monkeys==0]
print(two_labels)
label_to_str = {0: 'Normal nerve', 1: 'Glaucoma nerve'}
label_to_color = {0: (102/255, 178/255, 255/255, 1.), 1: (255/255, 178/255, 102/255, 1.)}
fig = plt.figure()
ax = Axes3D(fig)
ax.set_xlim((2000, 4000))
ax.set_ylim((1000, 5000))
ax.set_zlim((-600, 200))
for nerve, label in zip(two_nerves, two_labels):
x = nerve[:, 0]
y = nerve[:, 1]
z = nerve[:, 2]
verts = [list(zip(x,y,z))]
poly = Poly3DCollection(verts, alpha=0.5)
color = label_to_color[int(label)]
poly.set_color(colors.rgb2hex(color))
poly.set_edgecolor('k')
ax.add_collection3d(poly)
patch_0 = mpatches.Patch(color=label_to_color[0], label=label_to_str[0], alpha=0.5)
patch_1 = mpatches.Patch(color=label_to_color[1], label=label_to_str[1], alpha=0.5)
plt.legend(handles=[patch_0, patch_1], prop={'size': 14})
plt.show()
```
We first try to detect if there are two groups of optical nerve heads, based on the 3D coordinates of the landmarks sets.
```
from geomstats.geometry.euclidean import EuclideanMetric
nerves_vec = nerves.reshape(22, -1)
eucl_metric = EuclideanMetric(nerves_vec.shape[-1])
eucl_dist = eucl_metric.dist_pairwise(nerves_vec)
plt.figure()
plt.imshow(eucl_dist);
```
We do not see any two clear clusters.
We want to investigate if there is a difference between these two groups of shapes - normal nerve versus glaucoma nerve - or if the main difference is merely relative to the global size of the landmarks' sets.
```
m_ambient = 3
k_landmarks = 5
preshape = PreShapeSpace(m_ambient=m_ambient, k_landmarks=k_landmarks)
matrices_metric = preshape.embedding_metric
sizes = matrices_metric.norm(preshape.center(nerves))
plt.figure(figsize=(6, 4))
for label, col in label_to_color.items():
label_sizes = sizes[labels==label]
plt.hist(label_sizes, color=col, label=label_to_str[label], alpha=0.5, bins=10)
plt.axvline(gs.mean(label_sizes), color=col)
plt.legend(fontsize=14)
plt.title('Sizes of optical nerves', fontsize=14);
```
The vertical lines represent the sample mean of each group (normal/glaucoma).
```
plt.figure(figsize=(6, 4))
plt.hist(sizes[labels==1] - sizes[labels==0], alpha=0.5)
plt.axvline(0, color='black')
plt.title('Difference in size of optical nerve between glaucoma and normal eyes', fontsize=14);
```
We perform a hypothesis test, testing if the two samples of sizes have the same average. We use the t-test for related samples, since the sample elements are paired: two eyes for each monkey.
```
from scipy import stats
signif_level = 0.05
tstat, pvalue = stats.ttest_rel(sizes[labels==0], sizes[labels==1])
print(pvalue < signif_level)
```
There is a significative difference, in optical nerve eyes' sizes, between the glaucoma and normal eye.
We want to investigate if there is a difference in shapes, where the size component has been quotiented out.
We project the data to the Kendall pre-shape space, which:
- centers the nerve landmark sets so that they share the same barycenter,
- normalizes the sizes of the landmarks' sets to 1.
```
nerves_preshape = preshape.projection(nerves)
print(nerves_preshape.shape)
print(preshape.belongs(nerves_preshape))
print(gs.isclose(matrices_metric.norm(nerves_preshape), 1.))
```
In order to quotient out the 3D orientation component, we align the landmark sets in the preshape space.
```
base_point = nerves_preshape[0]
nerves_shape = preshape.align(point=nerves_preshape, base_point=base_point)
```
The Kendall metric is a Riemannian metric that takes this alignment into account. It corresponds to the metric of the Kendall shape space, which is the manifold defined as the preshape space quotient by the action of the rotation in m_ambient dimensions, here in 3 dimensions.
```
kendall_metric = KendallShapeMetric(m_ambient=m_ambient, k_landmarks=k_landmarks)
```
We can use it to perform a tangent PCA in the Kendall shape space, and determine if we see a difference in the shapes of the optical nerves.
```
from geomstats.learning.pca import TangentPCA
tpca = TangentPCA(kendall_metric)
tpca.fit(nerves_shape)
plt.plot(
tpca.explained_variance_ratio_)
plt.xlabel("Number of principal tangent components", size=14)
plt.ylabel("Fraction of explained variance", size=14);
```
Two principal components already describe around 60% of the variance. We plot the data projected in the tangent space defined by these two principal components.
```
X = tpca.transform(nerves_shape)
plt.figure(figsize=(12, 12))
for label, col in label_to_color.items():
mask = labels == label
plt.scatter(X[mask, 0], X[mask, 1], color=col, s=100, label=label_to_str[label]);
plt.legend(fontsize=14);
for label, x, y in zip(monkeys, X[:, 0], X[:, 1]):
plt.annotate(
label,
xy=(x, y), xytext=(-20, 20),
textcoords='offset points', ha='right', va='bottom',
bbox=dict(boxstyle='round,pad=0.5', fc='white', alpha=0.5),
arrowprops=dict(arrowstyle = '->', connectionstyle='arc3,rad=0'))
plt.show()
```
The indices represent the monkeys' indices.
In contrast to the above study focusing on the optical nerves' sizes, visual inspection does not reveal any clusters between the glaucoma and normal optical nerves' shapes. We also do not see any obvious pattern between the two optical nerves of the same monkey.
This shows that the difference between the optical nerve heads mainly resides in the over sizes of the optical nerves.
```
dist_pairwise = kendall_metric.dist_pairwise(nerves_shape)
print(dist_pairwise .shape)
plt.figure()
plt.imshow(dist_pairwise);
```
We try a agglomerative hierarchical clustering to investigate if we can cluster in the Kendall shape space.
```
from geomstats.learning.agglomerative_hierarchical_clustering import AgglomerativeHierarchicalClustering
clustering = AgglomerativeHierarchicalClustering(distance='precomputed', n_clusters=2)
clustering.fit(dist_pairwise)
predicted_labels = clustering.labels_
print('True labels:', labels)
print('Predicted labels:', predicted_labels)
accuracy = gs.sum(labels==predicted_labels) / len(labels)
print(f'Accuracy: {accuracy:.2f}')
```
The accuracy is barely above the accuracy of a random classifier, that would assign 0 or 1 with probably 0.5 to each of the shapes. This confirms that the difference that exists between the two groups is mostly due to the landmarks' set size and not their shapes.
## References
.. [PE2015] Patrangenaru and L. Ellingson. Nonparametric Statistics on Manifolds and Their Applications to Object Data, 2015. https://doi.org/10.1201/b18969
| github_jupyter |
# Fibermagic Tutorial
Fibermagic is an open-source software package for the analysis of photometry data. It is written in Python and operates on pandas and numpy dataframes.
Fiber Photometry is a novel technique to capture neural activity in-vivo with a subsecond temporal precision. Genetically encoded fluorescent sensors are expressed by either transfection with an (AAV)-virus or transgenic expression. Strenght of fluorescence is dependent on the concentration of the neurotransmitter the sensor detects. Using an optic glas fiber, it is possible to image neural dynamics and transmitter release.

graphik source: https://en.wikipedia.org/wiki/Fiber_photometry
## Fiber Photometry using Neurophotometrics devices
There are a bunch of different photometry systems out there like TDT systems, Neurophotometrics or open-hardware approaches. In this tutorial, we will use data from Neurophotometrics as an example.

## Multiple Colors
With Neurophotometrics (NPM), it is possible to capture data from two fluorescent sensors simultaneously - if they emit light in different wave lengths. NPM can measure light of two different color spectra: Red and Green. Using this technology, it is possible, to e.g. express a green calcium sensor (e.g. GCaMP6f) and a red dopamine sensor (e.g. Rdlight1).

grafik source: https://www.researchgate.net/publication/264796291_Bright_and_fast_multi-colored_voltage_reporters_via_electrochromic_FRET/figures?lo=1
## Multiple Mice
NPM can record data from multiple mice at once - delivered through a single path chord that splits into several small cables that can be attached individually to a single mouse.
The end of the patch chord projects to a photosensor which captures the light. Here's how it looks like:

Before starting the recording, the researcher defines a region of interest for each cable and color. Then, NPM captures the light intensity for each region of interest seperately. All data streams are written into one single csv file per recording.
## Investigating a single raw data file
Let's have a look into an arbitrary recording file and try to understand the data.
```
!pip install fibermagic
import pandas as pd
from fibermagic.utils.download_dataset import download
download('tutorial')
df = pd.read_csv('rawdata_demo.csv')
df.head(10)
```
Each Region of Interest is represented as one column.
A simple frame counter and a timestamp indicate when a measurement happened.
The Flags column encodes different things: Which LED was on at the time of measurement, which input and/or output channel was on or off and whether the laser for optogenetic stimulation was on. NPM encodes all this information into a single number using a dual encoding.
Because the emission spectra of fluorescent proteins are partly overlapping, it is not possible to measure both, the red and green sensor, at the same time. Instead, only one LED is on at a time. That way, we get presice measurements.

We can use fibermagic to decode which LEDs were on at what measurement from the Flags column.
```
from fibermagic.IO.NeurophotometricsIO import extract_leds
if 'Flags' in df.columns: # legacy fix: Flags were renamed to LedState
df = df.rename(columns={'Flags': 'LedState'})
df = extract_leds(df).dropna()
df.head(10)
```
Now, we have all we need to plot the full trace of a single sensor of one mouse and inspect the raw data values. In this recording, Rdlight1 and GCaMP6f were expressed. Rdlight1 is excited by the 560 nm LED and emits light in the red color spectrum. GCaMP6f is excited by the 470 nm LED and emits light in the green spectrum. Now, let's spike into the raw data of Rdlight1 of a single mouse.
We use "plotly" for plotting here. Plotly is a plotting library for Python and other programming languages. It is interactive, which means we can zoom in and out and scroll through the data, which is exactly what we want to do at the moment.
```
import plotly.express as px
region0R_red = df[df.wave_len == 560][['FrameCounter', 'Region0R']]
px.line(region0R_red, x='FrameCounter', y='Region0R')
```
We can clearly see huge transients from dopamine activity in the raw data. Congratulations!
However, there are several issues with the data. Fiber Photometry is not measureing the difference between day and night. Changes in fluorescence are usually extremely small and very hard to measure. We basically measure light intensity with a super high precision. However, this means that the photometry system will also pick up every disturbance, even if very small.
Examples of unwanted noise are:
* Photobleaching of the Patch Chord
* Photobleaching caused by LED heating
* Photobleaching caused by destroyment of sensors
* Motion artifacts
* Loose or poorly attached cables

## Demodulation of Fiber Photometry Data
As we saw, the raw data is useless without further processing. There are a variety of different of different methods to remove photobleaching and motion artifacts, here are a few examples:
### Correction of Photobleaching:
* High-Pass Filtering: Photobleaching usually occurs at a very slow timescale. By applying a simple high-pass filter, e.g. a butterworth, it is possible to remove the gross artifacts of photobleaching, but it removes also slow changes in neurotransmitter concentration
* Biexponential decay: Photobleaching can be estimated by a decreasing exponential function. However, as it happens on two different timescales (e.g. fast LED-heating-based photobleaching and slow patch chord photobleaching), we need a biexponential decay that can be regressed to the data and then subtracted.
* airPLS: Adaptive iteratively reweighted penalized least squares. A more advanced method to remove artifacts. For more info, please see the paper: DOI: 10.1039/b922045c
### Correction of Motion Artifacts:
* Genetically encoded Neurotransmitter indicators have usually a useful attribute: If stimulated at around 410 nm (instead of e.g. 470 or 560), the excitation will be the same independent of the neurotransmitter concentration. Stimulating a this wave length is called recording an "isobestic". The isobestic is usefull to correct motion artifacts.
* If a transient is caused by neural activity, it should be detectable in the signal channel, but if it is caused by motion, it should be detectable in both, the isobestic and signal.
* We can remove motion artifacts if we fit the isobestic to the signal and subtract.
## Demodulate using airPLS
We can use fibermagic to calculate z-scores of dF/F using the airPLS algorithm. However, before we do that, we need to bring the dataframe into long-format.
```
NPM_RED = 560
NPM_GREEN = 470
NPM_ISO = 410
# dirty hack to come around dropped frames until we find better solution -
# it makes about 0.16 s difference
df.FrameCounter = df.index // len(df.wave_len.unique())
df = df.set_index('FrameCounter')
regions = [column for column in df.columns if 'Region' in column]
dfs = list()
for region in regions:
channel = NPM_GREEN if 'G' in region else NPM_RED
sdf = pd.DataFrame(data={
'Region': region,
'Channel': channel,
'Signal': df[region][df.wave_len == channel],
'Reference': df[region][df.wave_len == NPM_ISO]
}
)
dfs.append(sdf)
dfs = pd.concat(dfs).reset_index().set_index(['Region', 'Channel', 'FrameCounter'])
dfs
from fibermagic.core.demodulate import add_zdFF
dfs = add_zdFF(dfs, method='airPLS', remove=250)
dfs
fig = px.line(dfs.reset_index(), x='FrameCounter', y='zdFF (airPLS)', facet_row='Region', height=1000)
fig.for_each_annotation(lambda a: a.update(text=a.text.split("=")[-1]))
fig
```
## Analyzing and Synchronizing Behavioral Data
In almost all experiments, we want to collect behavioral data along with the neural data. We then want to correlate the mice's behavior with our neural recording. For example, the mouse might perform a lever pressing task in an operand box. The operand box collects data about the time when the lever is pressed and when a food reward is delivered.
Let's have a look into the log file produced by the operant box
```
logs = pd.read_csv('operand_box.log')
logs
```
As we see, the column "SI@0.0" records the type of event (e.g. left lever pressed; food delivered, etc...) and the time.
There are a variety of possibilities how to synchronize the logs with the FP data. In this case, an external generator generates a TTL pulse every 100 ms. The TTL pulse is captured by the operand box and logged as SI. The TTL pulse is also captured by the "input1" channel of NPM and saved in "input1.csv".
Luckily, fibermagic offers functionality to synchronize both.
```
from fibermagic.IO.NeurophotometricsIO import sync_from_TTL_gen
from pathlib import Path
logs = sync_from_TTL_gen(logs, Path('.'))
logs
```
We see that the column 'FrameCounter' was added to the dataframe. Now we know exactly where in the photometry data an event happened.
## Plot Perievents
With the synchronization done, we can finally extract a few seconds before and after each event and investigate if there is a common pattern. This is called calculating perievents.
```
logs['Region'] = 'Region6R'
logs = logs.set_index('Region', append=True).swaplevel()
logs
from fibermagic.core.perievents import perievents
peri = perievents(dfs.set_index('FrameCounter', append=True), logs[logs.Event=='FD'], window=5, frequency=25)
peri
fig = px.scatter(peri.reset_index(), x='Timestamp', y='Trial', color='zdFF (airPLS)', range_color=(-5,5),
color_continuous_scale=['blue', 'grey', 'red'], height=250).update_yaxes(autorange="reversed")
for scatter in fig.data:
scatter.marker.symbol = 'square'
fig.show()
```
# How to analyze big projects
So far, we analyzed one single mouse, one single recording, one single channel. However, in practice, we usually record from 10-20 mice per experiment and have multiple long recordings. This adds up easily to several hundret files together with all the logs produced. It would be very time consuming and error-prone if a researcher would have to analyze every single file on its own.
Fortunately, fibermagic offers functionality to process a full project at once and fully automatically. In addition, fibermagic is very fast and can process full projects within seconds. Ultimately, you can use the same functions, no matter if you load a full project or a single file.
The only thing you have to do is to structure your project into a tree of directories with one category per level. For example, you may want to structure your project into experimental group (condition/control), experimental procedure (e.g. PR2, PR5, PR8, PR11) and recording (e.g. R1, R2, R3). You have to organize your recordings into subdirectories.
```
== condition
=PR2
=R1
data.csv
input1.csv
regions_to_mouse.csv
time.csv
logs.csv
=R2
=R3
=PR5
=PR8
=PR11
= control
```
```
from fibermagic.IO.NeurophotometricsIO import read_project_logs, read_project_rawdata
download('fdrd2xadora_PR_Pilot')
help(read_project_rawdata)
help(read_project_logs)
project_path = Path(r'fdrd2xadora_PR_Pilot')
logs = read_project_logs(project_path,
['Paradigm'], ignore_dirs=['meta', 'processed'])
df = read_project_rawdata(project_path,
['Paradigm'], 'FED3.csv', ignore_dirs=['meta', 'processed'])
df = add_zdFF(df, smooth_win=10, remove=200).set_index('FrameCounter', append=True)
peri = perievents(df, logs[logs.Event == 'FD'], 5, 25)
output_path = project_path / 'processed'
df.to_csv(output_path / 'datastream.csv')
logs.to_csv(output_path / 'logs.csv')
peri.to_csv(output_path / 'perievents.csv')
peri = peri.reset_index()
fig = px.scatter(peri[peri.Channel==560], x='Timestamp', y='Trial', color='zdFF (airPLS)', facet_row='Paradigm', facet_col='Mouse',
range_color=(-5,5), facet_row_spacing=0, facet_col_spacing=0, width=140*len(peri.Mouse.unique()),
color_continuous_scale=['blue', 'grey', 'red']).update_yaxes(autorange="reversed")
for scatter in fig.data:
scatter.marker.symbol = 'square'
fig.for_each_annotation(lambda a: a.update(text=a.text.split("=")[-1]))
fig.update_xaxes(range = [peri.Timestamp.min(),peri.Timestamp.max()])
fig.update_xaxes(matches='x')
fig.show()
peri
```
| github_jupyter |
```
#Importing libraries
import numpy as np
import pandas as pd
import random
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
%matplotlib inline
#Defining independent variable as angles from 60deg to 300deg converted to radians
x = np.array([i*np.pi/180 for i in range(10,360,3)])
#Setting seed for reproducability
np.random.seed(10)
#Defining the target/dependent variable as sine of the independent variable
y = np.sin(x) + np.random.normal(0,0.15,len(x))
#Creating the dataframe using independent and dependent variable
data = pd.DataFrame(np.column_stack([x,y]),columns=['x','y'])
#Printing first 5 rows of the data
data.head()
#Plotting the dependent and independent variables
plt.figure(figsize=(12,8))
plt.plot(data['x'],data['y'],'.')
# polynomial regression with powers of x from 1 to 15
for i in range(2,16): #power of 1 is already there, hence starting with 2
colname = 'x_%d'%i #new var will be x_power
data[colname] = data['x']**i
data.head()
```
Creating test and train
```
data['randNumCol'] = np.random.randint(1, 6, data.shape[0])
train=data[data['randNumCol']<=3]
test=data[data['randNumCol']>3]
train = train.drop('randNumCol', axis=1)
test = test.drop('randNumCol', axis=1)
```
## Linear Regression
```
#Import Linear Regression model from scikit-learn.
from sklearn.linear_model import LinearRegression
#Separating the independent and dependent variables
X_train = train.drop('y', axis=1).values
y_train = train['y'].values
X_test = test.drop('y', axis=1).values
y_test = test['y'].values
#Linear regression with one features
independent_variable_train = X_train[:,0:1]
linreg = LinearRegression(normalize=True)
linreg.fit(independent_variable_train,y_train)
y_train_pred = linreg.predict(independent_variable_train)
rss_train = sum((y_train_pred-y_train)**2) / X_train.shape[0]
independent_variable_test = X_test[:,0:1]
y_test_pred = linreg.predict(independent_variable_test)
rss_test = sum((y_test_pred-y_test)**2)/ X_test.shape[0]
print("Training Error", rss_train)
print("Testing Error",rss_test)
plt.plot(X_train[:,0:1],y_train_pred)
plt.plot(X_train[:,0:1],y_train,'.')
#Linear regression with three features
independent_variable_train = X_train[:,0:3]
linreg = LinearRegression(normalize=True)
linreg.fit(independent_variable_train,y_train)
y_train_pred = linreg.predict(independent_variable_train)
rss_train = sum((y_train_pred-y_train)**2) / X_train.shape[0]
independent_variable_test = X_test[:,0:3]
y_test_pred = linreg.predict(independent_variable_test)
rss_test = sum((y_test_pred-y_test)**2)/ X_test.shape[0]
print("Training Error", rss_train)
print("Testing Error",rss_test)
plt.plot(X_train[:,0:1],y_train_pred)
plt.plot(X_train[:,0:1],y_train,'.')
#Linear regression with Seven features
independent_variable_train = X_train[:,0:9]
linreg = LinearRegression(normalize=True)
linreg.fit(independent_variable_train,y_train)
y_train_pred = linreg.predict(independent_variable_train)
rss_train = sum((y_train_pred-y_train)**2) / X_train.shape[0]
independent_variable_test = X_test[:,0:9]
y_test_pred = linreg.predict(independent_variable_test)
rss_test = sum((y_test_pred-y_test)**2)/ X_test.shape[0]
print("Training Error", rss_train)
print("Testing Error",rss_test)
plt.plot(X_train[:,0:1],y_train_pred)
plt.plot(X_train[:,0:1],y_train,'.')
# defining a function which will fit linear regression model, plot the results, and return the coefficients
def linear_regression(train_x, train_y, test_x, test_y, features, models_to_plot):
#Fit the model
linreg = LinearRegression(normalize=True)
linreg.fit(train_x,train_y)
train_y_pred = linreg.predict(train_x)
test_y_pred = linreg.predict(test_x)
#Check if a plot is to be made for the entered features
if features in models_to_plot:
plt.subplot(models_to_plot[features])
plt.tight_layout()
plt.plot(train_x[:,0:1],train_y_pred)
plt.plot(train_x[:,0:1],train_y,'.')
plt.title('Number of Predictors: %d'%features)
#Return the result in pre-defined format
rss_train = sum((train_y_pred-train_y)**2)/train_x.shape[0]
ret = [rss_train]
rss_test = sum((test_y_pred-test_y)**2)/test_x.shape[0]
ret.extend([rss_test])
ret.extend([linreg.intercept_])
ret.extend(linreg.coef_)
return ret
#Initialize a dataframe to store the results:
col = ['mrss_train','mrss_test','intercept'] + ['coef_Var_%d'%i for i in range(1,16)]
ind = ['Number_of_variable_%d'%i for i in range(1,16)]
coef_matrix_simple = pd.DataFrame(index=ind, columns=col)
#Define the number of features for which a plot is required:
models_to_plot = {1:231,3:232,6:233,9:234,12:235,15:236}
#Iterate through all powers and store the results in a matrix form
plt.figure(figsize=(12,8))
for i in range(1,16):
train_x = X_train[:,0:i]
train_y = y_train
test_x = X_test[:,0:i]
test_y = y_test
coef_matrix_simple.iloc[i-1,0:i+3] = linear_regression(train_x,train_y, test_x, test_y, features=i, models_to_plot=models_to_plot)
#Set the display format to be scientific for ease of analysis
pd.options.display.float_format = '{:,.2g}'.format
coef_matrix_simple
coef_matrix_simple[['mrss_train','mrss_test']].plot()
plt.xlabel('Features')
plt.ylabel('MRSS')
plt.legend(['train', 'test'])
```
## Ridge
<img src="ridge.png">
```
# Importing ridge from sklearn's linear_model module
from sklearn.linear_model import Ridge
#Set the different values of alpha to be tested
alpha_ridge = [0, 1e-8, 1e-4, 1e-3,1e-2, 1, 5, 10, 20, 25]
# defining a function which will fit ridge regression model, plot the results, and return the coefficients
def ridge_regression(train_x, train_y, test_x, test_y, alpha, models_to_plot={}):
#Fit the model
ridgereg = Ridge(alpha=alpha,normalize=True)
ridgereg.fit(train_x,train_y)
train_y_pred = ridgereg.predict(train_x)
test_y_pred = ridgereg.predict(test_x)
#Check if a plot is to be made for the entered alpha
if alpha in models_to_plot:
plt.subplot(models_to_plot[alpha])
plt.tight_layout()
plt.plot(train_x[:,0:1],train_y_pred)
plt.plot(train_x[:,0:1],train_y,'.')
plt.title('Plot for alpha: %.3g'%alpha)
#Return the result in pre-defined format
mrss_train = sum((train_y_pred-train_y)**2)/train_x.shape[0]
ret = [mrss_train]
mrss_test = sum((test_y_pred-test_y)**2)/test_x.shape[0]
ret.extend([mrss_test])
ret.extend([ridgereg.intercept_])
ret.extend(ridgereg.coef_)
return ret
#Initialize the dataframe for storing coefficients.
col = ['mrss_train','mrss_test','intercept'] + ['coef_Var_%d'%i for i in range(1,16)]
ind = ['alpha_%.2g'%alpha_ridge[i] for i in range(0,10)]
coef_matrix_ridge = pd.DataFrame(index=ind, columns=col)
#Define the alpha value for which a plot is required:
models_to_plot = {0:231, 1e-4:232, 1e-3:233, 1e-2:234, 1:235, 5:236}
#Iterate over the 10 alpha values:
plt.figure(figsize=(12,8))
for i in range(10):
coef_matrix_ridge.iloc[i,] = ridge_regression(train_x, train_y, test_x, test_y, alpha_ridge[i], models_to_plot)
#Set the display format to be scientific for ease of analysis
pd.options.display.float_format = '{:,.2g}'.format
coef_matrix_ridge
coef_matrix_ridge['mrss_train']
coef_matrix_ridge['mrss_test'].shape
coef_matrix_ridge[['mrss_train','mrss_test']].plot()
plt.xlabel('Alpha Values')
plt.ylabel('MRSS')
plt.legend(['train', 'test'])
alpha_ridge
#Printing number of zeros in each row of the coefficients dataset
coef_matrix_ridge.apply(lambda x: sum(x.values==0),axis=1)
```
## Lasso
<img src="Lasso.png">
```
#Importing Lasso model from sklearn's linear_model module
from sklearn.linear_model import Lasso
#Define the alpha values to test
alpha_lasso = [0, 1e-10, 1e-8, 1e-5,1e-4, 1e-3,1e-2, 1, 5, 10]
# defining a function which will fit lasso regression model, plot the results, and return the coefficients
def lasso_regression(train_x, train_y, test_x, test_y, alpha, models_to_plot={}):
#Fit the model
if alpha == 0:
lassoreg = LinearRegression(normalize=True)
lassoreg.fit(train_x, train_y)
train_y_pred = lassoreg.predict(train_x)
test_y_pred = lassoreg.predict(test_x)
else:
lassoreg = Lasso(alpha=alpha,normalize=True)
lassoreg.fit(train_x,train_y)
train_y_pred = lassoreg.predict(train_x)
test_y_pred = lassoreg.predict(test_x)
#Check if a plot is to be made for the entered alpha
if alpha in models_to_plot:
plt.subplot(models_to_plot[alpha])
plt.tight_layout()
plt.plot(train_x[:,0:1],train_y_pred)
plt.plot(train_x[:,0:1],train_y,'.')
plt.title('Plot for alpha: %.3g'%alpha)
#Return the result in pre-defined format
mrss_train = sum((train_y_pred-train_y)**2)/train_x.shape[0]
ret = [mrss_train]
mrss_test = sum((test_y_pred-test_y)**2)/test_x.shape[0]
ret.extend([mrss_test])
ret.extend([lassoreg.intercept_])
ret.extend(lassoreg.coef_)
return ret
#Initialize the dataframe to store coefficients
col = ['mrss_train','mrss_test','intercept'] + ['coef_Var_%d'%i for i in range(1,16)]
ind = ['alpha_%.2g'%alpha_lasso[i] for i in range(0,10)]
coef_matrix_lasso = pd.DataFrame(index=ind, columns=col)
#Define the models to plot
models_to_plot = {0:231, 1e-5:232,1e-4:233, 1e-3:234, 1e-2:235, 1:236}
#Iterate over the 10 alpha values:
plt.figure(figsize=(12,8))
for i in range(10):
coef_matrix_lasso.iloc[i,] = lasso_regression(train_x, train_y, test_x, test_y, alpha_lasso[i], models_to_plot)
#Set the display format to be scientific for ease of analysis
pd.options.display.float_format = '{:,.2g}'.format
coef_matrix_lasso
coef_matrix_lasso[['mrss_train','mrss_test']].plot()
plt.xlabel('Alpha Values')
plt.ylabel('MRSS')
plt.legend(['train', 'test'])
coef_matrix_lasso.apply(lambda x: sum(x.values==0),axis=1)
```
| github_jupyter |
```
%matplotlib inline
%load_ext autoreload
%autoreload 2
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import networkx as nx
import utils # local file
plt.rcParams['font.family'] = 'serif'
plt.rcParams['font.size'] = 17.0
plt.rc('text', usetex=False)
```
## Temporal study of network parameters
```
def parse_time(team, matchid):
df = pd.read_csv('./2020_Problem_D_DATA/passingevents.csv')
df = df[df['MatchID'] == matchid]
df = df[df['TeamID'] == team]
df_full = pd.read_csv('./2020_Problem_D_DATA/fullevents.csv')
df_full = df_full[df_full['MatchID'] == matchid]
df_full = df_full[df_full['TeamID'] == team]
first_half_time = df_full[df_full['MatchPeriod'] == '1H'].iloc[-1]['EventTime']
df.loc[df['MatchPeriod'] == '2H', 'EventTime'] += first_half_time
return df
def temporal_network_params(team, matchid, length=50):
from network import build_network, plot_network, calc_network_params
df = parse_time(team, matchid)
df = df[df['TeamID'] == team]
df = df[df['MatchID'] == matchid]
time = df['EventTime'].values
net_param_dict = {}
for i in range(len(df)):
if length + i >= len(df):
break
if i % 20 == 0:
print(i)
G, pos, centrality_dict, geometrical_dist, unidirection_pass, weight_dict = build_network(
df[i:length + i], team, matchid)
network_params = calc_network_params(G)
local_time = time[length + i] - time[i]
network_params['delta_time'] = local_time
network_params['time'] = time[length + i]
net_param_dict[i] = network_params
df_net_param = pd.DataFrame(net_param_dict).T
return df_net_param
df_net_huskies = temporal_network_params('Huskies', 18)
df_net_opponent = temporal_network_params('Opponent18', 18)
plt.rc('text', usetex=True)
fig, axes = plt.subplots(4, 1, figsize=(11, 10), sharex=True)
keywords = [
'clustering_coeff', 'shortest_path', 'largest_eigenvalue', 'algebraic_conn'
]
ylabel_set = ['Clustering', 'Shortest Path',
r'$\lambda_1$', r'$\widetilde{\lambda_2}$']
tag_set = ['A', 'B', 'C', 'D', 'E', 'F']
for ind, key in enumerate(keywords):
ax = axes[ind]
ax.plot(df_net_huskies['time'].values / 60, df_net_huskies[key].values,
color='steelblue', marker='.', label='Huskies')
ax.plot(df_net_opponent['time'].values / 60, df_net_opponent[key].values,
color='orange', marker='.', label='Opponent')
ax.set_ylabel(ylabel_set[ind])
ax.set_xlabel('Time (min)')
ax.tick_params(direction='in', left=True, right=True,
bottom=True, top=True, labelleft=True, labelbottom=False)
if ind == 0:
ax.legend(loc='upper right')
ylim = ax.get_ylim()
ax.text(10, ylim[1] - (ylim[1] - ylim[0]) * 0.15,
r'$\textbf{' + tag_set[ind] + '}$',
fontsize=17,
fontweight='bold',
horizontalalignment='center',
verticalalignment='center')
axes[3].tick_params(direction='in', left=True, right=True,
bottom=True, top=True, labelleft=True, labelbottom=True)
plt.subplots_adjust(hspace=0)
#plt.savefig('./Draft/temporal-net-params-m18.pdf', dpi=200, bbox_inches='tight')
#plt.savefig('./Draft/temporal-net-params-m18.png', dpi=200, bbox_inches='tight')
```
## Temporal Classical Metrics
```
def cal_mean_position(df_pass, direction):
huskies_coor_mean = df_pass['EventOrigin_{}'.format(direction)].mean()
return huskies_coor_mean
def cal_centroid_disp(df_pass):
x_cen = cal_mean_position(df_pass, 'x')
y_cen = cal_mean_position(df_pass, 'y')
dist = np.sqrt(np.square(df_pass['EventOrigin_x'] - x_cen) + \
np.square(df_pass['EventOrigin_y'] - y_cen))
dispersion = np.std(dist, ddof=1)
return dispersion
def cal_advance(df_pass):
delta_x = np.abs(df_pass['EventDestination_x'] - df_pass['EventOrigin_x'])
delta_y = np.abs(df_pass['EventDestination_y'] - df_pass['EventOrigin_y'])
return delta_y.sum() / delta_x.sum()
def temporal_classical_metrics(team, matchid, length=50):
from network import build_network, plot_network, calc_network_params
df = parse_time(team, matchid)
df = df[df['TeamID'] == team]
df = df[df['MatchID'] == matchid]
time = df['EventTime'].values
metrics_dict = {}
for i in range(len(df)):
if length + i >= len(df):
break
metric_params = {}
metric_params['x_cen'] = cal_mean_position(df[i:length + i], 'x')
metric_params['y_cen'] = cal_mean_position(df[i:length + i], 'y')
metric_params['cen_disp'] = cal_centroid_disp(df[i:length + i])
metric_params['advance'] = cal_advance(df[i:length + i])
local_time = time[length + i] - time[i]
metric_params['delta_time'] = local_time
metric_params['time'] = time[length + i]
metrics_dict[i] = metric_params
df_metrics = pd.DataFrame(metrics_dict).T
return df_metrics
df_metrics_huskies = temporal_classical_metrics('Huskies', 18)
df_metrics_opponent = temporal_classical_metrics('Opponent18', 18)
plt.rc('text', usetex=True)
fig, axes = plt.subplots(4, 1, figsize=(11, 10), sharex=True)
keywords = [
'x_cen', 'y_cen', 'cen_disp', 'advance'
]
ylabel_set = [r'$<X>$', r'$<Y>$',
'Dispersion', r'$<\Delta Y>/<\Delta X>$']
tag_set = ['A', 'B', 'C', 'D', 'E', 'F']
for ind, key in enumerate(keywords):
ax = axes[ind]
ax.plot(df_metrics_huskies['time'].values / 60, df_metrics_huskies[key].values,
color='steelblue', marker='.', label='Huskies')
ax.plot(df_metrics_opponent['time'].values / 60, df_metrics_opponent[key].values,
color='orange', marker='.', label='Opponent')
ax.set_ylabel(ylabel_set[ind])
ax.set_xlabel('Time (min)')
ax.tick_params(direction='in', left=True, right=True,
bottom=True, top=True, labelleft=True, labelbottom=False)
if ind == 0:
ax.legend()
ylim = ax.get_ylim()
ax.text(10, ylim[1] - (ylim[1] - ylim[0]) * 0.15,
r'$\textbf{' + tag_set[ind] + '}$',
fontsize=17,
fontweight='bold',
horizontalalignment='center',
verticalalignment='center')
axes[3].tick_params(direction='in', left=True, right=True,
bottom=True, top=True, labelleft=True, labelbottom=True)
plt.subplots_adjust(hspace=0)
plt.savefig('./Draft/temporal-spatial-m18.pdf', dpi=200, bbox_inches='tight')
plt.savefig('./Draft/temporal-spatial-m18.png', dpi=200, bbox_inches='tight')
```
| github_jupyter |
# Peform statistical analyses of GNSS station locations and tropospheric zenith delays
**Author**: Simran Sangha, David Bekaert - Jet Propulsion Laboratory
This notebook provides an overview of the functionality included in the **`raiderStats.py`** program. Specifically, we outline examples on how to perform basic statistical analyses of GNSS station location and tropospheric zenith delay information over a user defined area of interest, span of time, and seasonal interval. In this notebook, we query GNSS stations spanning northern California between 2018 and 2019.
We will outline the following statistical analysis and filtering options:
- Restrict analyses to range of years
- Restrict analyses to range of months (i.e. seasonal interval)
- Illustrate station distribution and tropospheric zenith delay mean/standard deviation
- Illustrate gridded distribution and tropospheric zenith delay mean/standard deviation
- Generate variogram plots across specified time periods
<div class="alert alert-info">
<b>Terminology:</b>
- *GNSS*: Stands for Global Navigation Satellite System. Describes any satellite constellation providing global or regional positioning, navigation, and timing services.
- *tropospheric zenith delay*: The precise atmospheric delay satellite signals experience when propagating through the troposphere.
- *variogram*: Characterization of the difference between field values at two locations.
- *empirical variogram*: Provides a description of how the data are correlated with distance.
- *experimental variogram*: A discrete function calculated using a measure of variability between pairs of points at various distances
- *sill*: Limit of the variogram, tending to infinity lag distances.
- *range*: The distance in which the difference of the variogram from the sill becomes negligible, such that the data arre no longer autocorrelated.
</div>
## Prep: Initial setup of the notebook
Below we set up the directory structure for this notebook exercise. In addition, we load the required modules into our python environment using the **`import`** command.
```
import os
import numpy as np
import matplotlib.pyplot as plt
## Defining the home and data directories
tutorial_home_dir = os.path.abspath(os.getcwd())
work_dir = os.path.abspath(os.getcwd())
print("Tutorial directory: ", tutorial_home_dir)
print("Work directory: ", work_dir)
# Verifying if RAiDER is installed correctly
try:
from RAiDER import statsPlot
except:
raise Exception('RAiDER is missing from your PYTHONPATH')
os.chdir(work_dir)
```
## Overview of the raiderStats.py program
<a id='overview'></a>
The **`raiderStats.py`** program provides a suite of convinient statistical analyses of GNSS station locations and tropospheric zenith delays.
Running **`raiderStats.py`** with the **`-h`** option will show the parameter options and outline several basic, practical examples.
Let us explore these options:
```
!raiderStats.py -h
```
### 1. Basic user input options
#### Input CSV file (**`--file FNAME`**)
**REQUIRED** argument. Provide a valid CSV file as input through **`--file`** which lists the GNSS station IDs (ID), lat/lon coordinates (Lat,Lon), dates in YYYY-MM-DD format (Date), and the desired data field in units of meters.
Note that the complementary **`raiderDownloadGNSS.py`** format generates such a primary CSV file named **`CombinedGPS_ztd.csv`** that contains all such fields and is already formatted as expected by **`raiderStats.py`**. Please refer to the accompanying **`raiderDownloadGNSS/raiderDownloadGNSS_tutorial.ipynb `** for more details and practical examples.
#### Data column name (**`--column_name COL_NAME`**)
Specify name of data column in input CSV file through **`--column_name `** that you wish to perform statistical analyses on. Input assumed to be in units of meters.
Default input column name set to **`ZTD`**, the name assigned to tropospheric zenith delays populated under the **`CombinedGPS_ztd.csv`** file generated through **`raiderDownloadGNSS.py`**
#### Data column unit (**`--unit UNIT`**)
Specify unit of input data column through **`--unit `**. Again, input assumed to be in units of meters so it will be converted into meters if not already in meters.
### 2. Run parameters
#### Output directory (**`--workdir WORKDIR`**)
Specify directory to deposit all outputs into with **`--workdir`**. Absolute and relative paths are both supported.
By default, outputs will be deposited into the current working directory where the program is launched.
#### Number of CPUs to be used (**`--cpus NUMCPUS`**)
Specify number of cpus to be used for multiprocessing with **`--cpus`**. For most cases, multiprocessing is essential in order to access data and perform statistical analyses within a reasonable amount of time.
May specify **`--cpus all`** at your own discretion in order to leverage all available CPUs on your system.
By default 8 CPUs will be used.
#### Verbose mode (**`--verbose`**)
Specify **`--verbose`** to print all statements through entire routine.
Additionally, verbose mode will generate variogram plots per gridded station AND time-slice.
### 3. Optional controls for spatiotemporal subsetting
#### Geographic bounding box (**`--bbox BOUNDING_BOX`**)
An area of interest may be specified as `SNWE` coordinates using the **`--bbox`** option. Coordinates should be specified as a space delimited string surrounded by quotes. The common intersection between the user-specified spatial bounds and the spatial bounds computed from the station locations in the input file is then passed. This example below would restrict the analysis to stations over northern California:
**`--bbox '36 40 -124 -119'`**
If no area of interest is specified, by default the spatial bounds computed from the station locations in the input file as passed.
#### Gridcell spacing (**`--spacing SPACING`**)
Specify degree spacing of grid-cells for statistical analyses through **`--spacing`**
By default grid-cell spacing is set to 1°. If the specified grid-cell spacing is not a multiple of the spatial bounds of the dataset, the grid-cell spacing again defaults back to 1°.
#### Subset in time (**`-ti TIMEINTERVAL`**)
Define temporal bounds with **`-ti TIMEINTERVAL`** by specifying earliest YYYY-MM-DD date followed by latest date YYYY-MM-DD. For example: **`-ti 2018-01-01,2019-01-01`**
By default, bounds set to earliest and latest time found in input file.
#### Seasonal interval (**`-si SEASONALINTERVAL`**)
Define subset in time by a specific interval for each year (i.e. seasonal interval) with **`-si SEASONALINTERVAL`** by specifying earliest MM-DD time followed by latest MM-DD time. For example: **`-si '03-21 06-21'`**
### 4. Supported types of individual station scatter-plots
#### Plot station distribution (**`--station_distribution`**)
Illustrate each individual station with black markers.
#### Plot mean tropospheric zenith delay by station (**`--station_delay_mean`**)
Illustrate the tropospheric zenith delay mean for each station with a **`hot`** colorbar.
#### Plot standard deviation of tropospheric zenith delay by station (**`--station_delay_stdev`**)
Illustrate the tropospheric zenith delay standard deviation for each station with a **`hot`** colorbar.
### 5. Supported types of gridded station plots
#### Plot gridded station heatmap (**`--grid_heatmap`**)
Illustrate heatmap of gridded station array with a **`hot`** colorbar.
#### Plot gridded mean tropospheric zenith delay (**`--grid_delay_mean`**)
Illustrate gridded tropospheric zenith delay mean with a **`hot`** colorbar.
#### Plot gridded standard deviation of tropospheric zenith delay (**`--grid_delay_stdev`**)
Illustrate gridded tropospheric zenith delay standard deviation with a **`hot`** colorbar.
### 6. Supported types of variogram plots
#### Plot variogram (**`--variogramplot`**)
Passing **`--variogramplot`** toggles plotting of gridded station variogram, where gridded sill and range values for the experimental variogram fits are illustrated.
#### Apply experimental fit to binned variogram (**`--binnedvariogram`**)
Pass **`--binnedvariogram`** to apply experimental variogram fit to total binned empirical variograms for each time slice.
Default is to pass total unbinned empiricial variogram.
### 7. Optional controls for plotting
#### Plot format (**`--plot_format PLOT_FMT`**)
File format for saving plots. Default is PNG.
#### Colorbar bounds (**`--color_bounds CBOUNDS`**)
Set lower and upper-bounds for plot colorbars. For example: **`--color_bounds '0 100'`**
By default set to the dynamic range of the data.
#### Colorbar percentile limits (**`--colorpercentile COLORPERCENTILE COLORPERCENTILE`**)
Set lower and upper percentile for plot colorbars. For example: **`--colorpercentile '30 100'`**
By default set to 25% and 95%, respectively.
#### Variogram density threshold (**`--densitythreshold DENSITYTHRESHOLD`**)
For variogram plots, a given grid-cell is only valid if it contains this specified threshold of stations.
By default set to 10 stations.
#### Superimpose individual stations over gridded array (**`--stationsongrids`**)
In gridded plots, superimpose your gridded array with a scatterplot of station locations.
#### Draw gridlines (**`--drawgridlines`**)
In gridded plots, draw gridlines.
#### Generate all supported plots (**`--plotall`**)
Generate all supported plots, as outlined under sections #4, #5, and #6 above.
## Download prerequisite GNSS station location and tropospheric zenith delay information with the **`raiderDownloadGNSS.py`** program
<a id='downloads'></a>
Virtually access GNSS station location and zenith delay information for the years '2018,2019', at a UTC time of day 'HH:MM:SS' of '00:00:00', and across a geographic bounding box '36 40 -124 -119' spanning over Northern California.
The footprint of the specified geographic bounding box is again depicted in **Fig. 1**.
In addition to querying for multiple years, we will also experiment with using the maximum number of allowed CPUs to save some time! Recall again that the default number of CPUs used for parallelization is 8.
Note these features and similar examples are outlined in more detail in the companion notebook **`raiderDownloadGNSS/raiderDownloadGNSS_tutorial.ipynb`**
```
!raiderDownloadGNSS.py --out products --years '2018,2019' --returntime '00:00:00' --BBOX '36 40 -124 -119' --cpus all
```
All of the extracted tropospheric zenith delay information stored under **`GPS_delays`** is concatenated with the GNSS station location information stored under **`gnssStationList_overbbox.csv`** into a primary comprehensive file **`CombinedGPS_ztd.csv`**
**`CombinedGPS_ztd.csv`** may in turn be directly used to perform basic statistical analyses using **`raiderStats.py`**.
<img src="support_docs/bbox_footprint.png" alt="footprint" width="700">
<center><b>Fig. 1</b> Footprint of geopraphic bounding box used in examples 1 and 2. </center>
## Examples of the **`raiderStats.py`** program
<a id='examples'></a>
### Example 1. Generate all individual station scatter-plots, as listed under section #4 <a id='example_1'></a>
Using the file **`CombinedGPS_ztd.csv`** generated by **`raiderDownloadGNSS.py`** as input, produce plots illustrating station distribution, mean tropospheric zenith delay by station, and standard deviation of tropospheric zenith delay by station.
Restrict the temporal span of the analyses to all data acquired between 2018-01-01 and 2019-12-31, and restrict the spatial extent to a geographic bounding box '36 40 -124 -119' spanning over Northern California.
The footprint of the specified geographic bounding box is depicted in **Fig. 1**.
These basic spatiotemporal constraints will be inherited by all successive examples.
```
!raiderStats.py --file products/CombinedGPS_ztd.csv -w maps -b '36 40 -124 -119' -ti '2018-01-01 2019-12-31' --station_distribution --station_delay_mean --station_delay_stdev
```
Now we can take a look at the generated products:
```
!ls maps/figures
```
Here we visualize the spatial distribution of stations (*ZTD_station_distribution.png*) as black markers.
<img src="maps/figures/ZTD_station_distribution.png" alt="ZTD_station_distribution" width="700">
To generate this figure alone, run:
```
!raiderStats.py --file products/CombinedGPS_ztd.csv -w maps -b '36 40 -124 -119' -ti '2018-01-01 2019-12-31' --station_distribution
```
Here we visualize the mean tropospheric zenith delay by station (*ZTD_station_delay_mean.png*) with a **`hot`** colorbar.
<img src="maps/figures/ZTD_station_delay_mean.png" alt="ZTD_station_delay_mean" width="700">
To generate this figure alone, run:
```
!raiderStats.py --file products/CombinedGPS_ztd.csv -w maps -b '36 40 -124 -119' -ti '2018-01-01 2019-12-31' --station_delay_mean
```
Here we visualize the standard deviation of tropospheric zenith delay by station (*ZTD_station_delay_stdev.png*) with a **`hot`** colorbar.
<img src="maps/figures/ZTD_station_delay_stdev.png" alt="ZTD_station_delay_stdev" width="700">
To generate this figure alone, run:
```
!raiderStats.py --file products/CombinedGPS_ztd.csv -w maps -b '36 40 -124 -119' -ti '2018-01-01 2019-12-31' --station_delay_stdev
```
### Example 2. Generate all gridded station plots, as listed under section #5 <a id='example_2'></a>
Produce plots illustrating gridded station distribution, gridded mean tropospheric zenith delay, and gridded standard deviation of tropospheric zenith delay.
```
!rm -rf maps
!raiderStats.py --file products/CombinedGPS_ztd.csv -w maps -b '36 40 -124 -119' -ti '2018-01-01 2019-12-31' --grid_heatmap --grid_delay_mean --grid_delay_stdev
```
Now we can take a look at the generated products:
```
!ls maps/figures
```
Here we visualize the heatmap of gridded station array (*ZTD_grid_heatmap.png*) with a **`hot`** colorbar.
Note that the colorbar bounds are saturated, which demonstrates the utility of plotting options outlined under section #7 such as **`--color_bounds`** and **`--colorpercentile`**
<img src="maps/figures/ZTD_grid_heatmap.png" alt="ZTD_grid_heatmap" width="700">
To generate this figure alone, run:
```
!raiderStats.py --file products/CombinedGPS_ztd.csv -w maps -b '36 40 -124 -119' -ti '2018-01-01 2019-12-31' --grid_heatmap
```
Here we visualize the gridded mean tropospheric zenith delay (*ZTD_grid_delay_mean.png*) with a **`hot`** colorbar.
<img src="maps/figures/ZTD_grid_delay_mean.png" alt="ZTD_grid_delay_mean" width="700">
To generate this figure alone, run:
```
!raiderStats.py --file products/CombinedGPS_ztd.csv -w maps -b '36 40 -124 -119' -ti '2018-01-01 2019-12-31' --grid_delay_mean
```
Here we visualize the gridded standard deviation of tropospheric zenith delay (*ZTD_grid_delay_stdev.png*) with a **`hot`** colorbar.
<img src="maps/figures/ZTD_grid_delay_stdev.png" alt="ZTD_grid_delay_stdev" width="700">
To generate this figure alone, run:
```
!raiderStats.py --file products/CombinedGPS_ztd.csv -w maps -b '36 40 -124 -119' -ti '2018-01-01 2019-12-31' --grid_delay_stdev
```
### Example 3. Generate all gridded station plots, as listed under section #5 <a id='example_3'></a>
Produce plot illustrating gridded mean tropospheric zenith delay, superimposed with individual station locations.
Additionally, subset data in time for spring. I.e. **`'03-21 06-21'`**
```
!rm -rf maps
!raiderStats.py --file products/CombinedGPS_ztd.csv -w maps -b '36 40 -124 -119' -ti '2018-01-01 2019-12-31' --seasonalinterval '03-21 06-21' --grid_delay_mean --stationsongrids
```
Now we can take a look at the generated product:
```
!ls maps/figures
```
Here we visualize the gridded mean tropospheric zenith delay (*ZTD_grid_delay_mean.png*) with a **`hot`** colorbar, with superimposed station locations denoted by blue markers.
<img src="maps/figures/ZTD_grid_delay_mean.png" alt="ZTD_grid_delay_mean" width="700">
### Example 4. Generate variogram plots, as listed under section #5 <a id='example_4'></a>
Produce plot illustrating gridded mean tropospheric zenith delay, superimposed with individual station locations.
Additionally, subset data in time for spring. I.e. **`'03-21 06-21'`**
Finally, use the maximum number of allowed CPUs to save some time.
```
!rm -rf maps
!raiderStats.py --file products/CombinedGPS_ztd.csv -w maps -b '36 40 -124 -119' -ti '2018-01-01 2019-12-31' --seasonalinterval '03-21 06-21' -variogramplot -verbose --cpus all
```
Now we can take a look at the generated variograms:
```
!ls maps/variograms
```
There are several subdirectories corresponding to each grid-cell that each contain empirical and experimental variograms generated for each time-slice (e.g. **`grid6_timeslice20180321_justEMPvariogram.eps `** and **`grid6_timeslice20180321_justEXPvariogram.eps `**, respectively) and across the entire sampled time period (**`grid6_timeslice20180321–20190621_justEMPvariogram.eps `** and **`grid6_timeslice20180321–20190621_justEXPvariogram.epss `**, respectively)
Here we visualize the total empirical variogram corresponding to the entire sampled time period for grid-cell 6 in the array (*grid6_timeslice20180321–20190621_justEMPvariogram.eps*).
<img src="maps/variograms/grid6_timeslice20180321–20190621_justEMPvariogram.eps" alt="justEMPvariogram" width="700">
Here we visualize the total experimental variogram corresponding to the entire sampled time period for grid-cell 6 in the array (*grid6_timeslice20180321–20190621_justEXPvariogram.eps*).
<img src="maps/variograms/grid6_timeslice20180321–20190621_justEXPvariogram.eps" alt="justEXPvariogram" width="700">
The central coordinates for all grid-nodes that satisfy the specified station density threshold (**`--densitythreshold`**, by default 10 stations per grid-cell) for variogram plots are stored in a lookup table:
```
!head maps/variograms/gridlocation_lookup.txt
```
Now we can take a look at the other generated figures:
```
!ls maps/figures
```
Here we visualize the gridded experimental variogram range (*ZTD_range_heatmap.png*) with a **`hot`** colorbar.
<img src="maps/figures/ZTD_range_heatmap.png" alt="ZTD_range_heatmap" width="700">
Here we visualize the gridded experimental variogram sill (*ZTD_sill_heatmap.png*) with a **`hot`** colorbar.
<img src="maps/figures/ZTD_sill_heatmap.png" alt="ZTD_sill_heatmap" width="700">
| github_jupyter |
# IASI Data Experiment
```
import sys
sys.path.append('/home/emmanuel/projects/2019_egp/src')
from data.iasi import IASIOrbits, create_dataarray
from experiments.experiment_iasi import GPModels
from models.gp_models import SparseGP
import GPy
import numpy as np
import pandas as pd
import xarray as xr
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
from sklearn.externals import joblib
%matplotlib inline
%load_ext autoreload
%autoreload 2
```
## Experimental Parameters
```
n_components = 50
train_size = 5000
n_train_orbits = 'all'
n_test_orbits = 'all'
random_state = 123
chunksize = 4000
rng = np.random.RandomState(random_state)
orbits = [
'20131001120859',
'20131001102955',
'20131001015955',
'20131001202954',
'20131001185058',
'20131001084755',
'20131001152954',
'20131001170858',
'20131001221154',
'20131001135058',
'20131001034155',
'20131001070555',
'20131001052355']
print(len(orbits))
day = orbits[0][:8]
iasi_data = IASIOrbits(
n_components = n_components,
train_size=train_size,
input_noise_level=None,
noise_coefficient=1.0,
n_train_orbits=n_train_orbits,
n_test_orbits=n_test_orbits,
)
# Get Training Data
training_data = iasi_data.get_training_data()
```
### Training Region
```
train_xarray = create_dataarray(
training_data['y'],
training_data['lat'],
training_data['lon'],
day)
train_xarray.name = 'Temperature'
train_xarray.min(), train_xarray.max()
# Labeled Data
fig = plt.figure(figsize=(15, 5))
ax1 = fig.add_subplot(121, projection=ccrs.PlateCarree())
ax2 = fig.add_subplot(122)
plot_data = train_xarray.isel(time=0) - 273.15
plot_data.plot.imshow(ax=ax1, transform=ccrs.PlateCarree(),
cmap='RdBu_r', alpha=1.0, vmin=-10, vmax=30,
cbar_kwargs={'orientation': 'horizontal',
'label': 'Temperature'})
ax1.coastlines()
plot_data.plot.hist(ax=ax2, bins=100)
ax2.set_xlabel('Data Histogram')
# fig.savefig(fig_save_loc + 't1_bias.png')
plt.show()
```
### Save Training Data
```
save_path = '/home/emmanuel/projects/2019_egp/data/processed/'
# DATA
train_xarray.to_netcdf(f"{save_path}training_egp_v1.nc")
```
### Model Parameters and Training
```
# Model Parameters
init_params = True
n_inducing = 1000
x_variance = iasi_data.x_cov
n_restarts = 1
# Initialize Modeler
clf = GPModels(
init_params = init_params,
x_variance=x_variance,
n_inducing=n_inducing,
n_restarts=n_restarts,
)
# Fit Models
clf.train_models(
training_data['X'],
training_data['y']
);
```
#### Save Models
```
save_path = '/home/emmanuel/projects/2019_egp/data/processed/'
# MODELS
joblib.dump(clf, f"{save_path}models_v1.pckl");
clf.models['error'].kernel_
```
### Testing
```
iasi_data.n_test_orbits = 13
datasets = xr.Dataset()
n_test = list()
for xtest, labels, lon, lat, orbit in iasi_data.get_testing_data():
n_test.append(xtest.shape)
temp_dataset = xr.Dataset()
# Save Labels Dataarray
temp_dataset['Temperature'] = create_dataarray(labels, lat, lon, orbit)
for imodel, ypred, ystd in clf.test_models(xtest, noise_less=True):
# Create Predictions array
temp_dataset[f"{imodel}_pred"] = create_dataarray(ypred, lat, lon, orbit)
# Create Standard Deviations array
temp_dataset[f"{imodel}_std"] = create_dataarray(ystd, lat, lon, orbit)
datasets = datasets.merge(temp_dataset)
del temp_dataset
n_test = np.array(n_test)
```
## Save Results
```
save_path = '/home/emmanuel/projects/2019_egp/data/processed/'
# DATA
datasets.to_netcdf(f"{save_path}results_egp_v1.nc")
```
| github_jupyter |
# Testing differences between groups
```
# Import numerical, data and plotting libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
# Only show 4 decimals when printing
np.set_printoptions(precision=4)
# Show the plots in the notebook
%matplotlib inline
```
Imagine we have some some measures of psychopathy in 12 students. 4 students are from Berkeley, and 4 students are from MIT.
```
psychos = pd.read_csv('psycho_students.csv')
psychos
```
We find that the mean score for the Berkeley students is different from the mean score for the MIT students:
```
berkeley_students = psychos[psychos['university'] == 'Berkeley']
berkeley_students
mit_students = psychos[psychos['university'] == 'MIT']
mit_students
berkeley_scores = berkeley_students['psychopathy']
mit_scores = mit_students['psychopathy']
berkeley_scores.mean(), mit_scores.mean()
```
Here is the difference between the means:
```
mean_diff = berkeley_scores.mean() - mit_scores.mean()
mean_diff
```
That's the difference we see. But - if we take any 8 students from a single university, and take the mean of the first four, and the mean of the second four, there will almost certainly be a difference in the means, just because there's some difference across individuals in the psychopathy score. Is this difference we see unusual compared to the differences we would see if we took eight students from the same university, and compared the means of the first four and the second four?
For a moment, let us pretend that all our Berkeley and MIT students come from the same university. Then I can pool the Berkely and MIT students together.
```
all_pooled = list(berkeley_scores) + list(mit_scores)
all_pooled
```
If there is no difference between Berkeley and MIT, then it should be OK to just shuffle the students to a random order, like this:
```
from random import shuffle
shuffle(all_pooled)
all_pooled
```
Now I can just pretend that the first four students are from one university, and the last four are from another university. Then I can compare the means.
```
fake_berkeley = all_pooled[:4]
fake_mit = all_pooled[4:]
np.mean(fake_berkeley) - np.mean(fake_mit)
```
```
fake_differences = []
for i in range(10000):
np.random.shuffle(all_pooled)
diff = np.mean(all_pooled[:4]) - np.mean(all_pooled[4:])
fake_differences.append(diff)
```
The 10000 values we calculated form the *sampling distribution*. Let's have a look:
```
plt.hist(fake_differences)
plt.title("Sampling distribution of mean difference");
```
Where does the value we actually see, sit in this histogram? More specifically, how many of the values in this histogram are less then or equal to the value we actually see?
```
# We will count the number of fake_differences <= our observed
count = 0
# Go through each of the 10000 values one by one
for diff in fake_differences:
if diff <= mean_diff:
count = count + 1
proportion = count / 10000
proportion
```
That's the p value.
| github_jupyter |
# Deep Neural Network for Image Classification: Application
When you finish this, you will have finished the last programming assignment of Week 4, and also the last programming assignment of this course!
You will use use the functions you'd implemented in the previous assignment to build a deep network, and apply it to cat vs non-cat classification. Hopefully, you will see an improvement in accuracy relative to your previous logistic regression implementation.
**After this assignment you will be able to:**
- Build and apply a deep neural network to supervised learning.
Let's get started!
## 1 - Packages
Let's first import all the packages that you will need during this assignment.
- [numpy](www.numpy.org) is the fundamental package for scientific computing with Python.
- [matplotlib](http://matplotlib.org) is a library to plot graphs in Python.
- [h5py](http://www.h5py.org) is a common package to interact with a dataset that is stored on an H5 file.
- [PIL](http://www.pythonware.com/products/pil/) and [scipy](https://www.scipy.org/) are used here to test your model with your own picture at the end.
- dnn_app_utils provides the functions implemented in the "Building your Deep Neural Network: Step by Step" assignment to this notebook.
- np.random.seed(1) is used to keep all the random function calls consistent. It will help us grade your work.
```
import time
import numpy as np
import h5py
import matplotlib.pyplot as plt
import scipy
from PIL import Image
from scipy import ndimage
from dnn_app_utils_v3 import *
%matplotlib inline
plt.rcParams['figure.figsize'] = (5.0, 4.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
%load_ext autoreload
%autoreload 2
np.random.seed(1)
```
## 2 - Dataset
You will use the same "Cat vs non-Cat" dataset as in "Logistic Regression as a Neural Network" (Assignment 2). The model you had built had 70% test accuracy on classifying cats vs non-cats images. Hopefully, your new model will perform a better!
**Problem Statement**: You are given a dataset ("data.h5") containing:
- a training set of m_train images labelled as cat (1) or non-cat (0)
- a test set of m_test images labelled as cat and non-cat
- each image is of shape (num_px, num_px, 3) where 3 is for the 3 channels (RGB).
Let's get more familiar with the dataset. Load the data by running the cell below.
```
train_x_orig, train_y, test_x_orig, test_y, classes = load_data()
```
The following code will show you an image in the dataset. Feel free to change the index and re-run the cell multiple times to see other images.
```
# Example of a picture
index = 10
plt.imshow(train_x_orig[index])
print ("y = " + str(train_y[0,index]) + ". It's a " + classes[train_y[0,index]].decode("utf-8") + " picture.")
# Explore your dataset
m_train = train_x_orig.shape[0]
num_px = train_x_orig.shape[1]
m_test = test_x_orig.shape[0]
print ("Number of training examples: " + str(m_train))
print ("Number of testing examples: " + str(m_test))
print ("Each image is of size: (" + str(num_px) + ", " + str(num_px) + ", 3)")
print ("train_x_orig shape: " + str(train_x_orig.shape))
print ("train_y shape: " + str(train_y.shape))
print ("test_x_orig shape: " + str(test_x_orig.shape))
print ("test_y shape: " + str(test_y.shape))
```
As usual, you reshape and standardize the images before feeding them to the network. The code is given in the cell below.
<img src="images/imvectorkiank.png" style="width:450px;height:300px;">
<caption><center> <u>Figure 1</u>: Image to vector conversion. <br> </center></caption>
```
# Reshape the training and test examples
train_x_flatten = train_x_orig.reshape(train_x_orig.shape[0], -1).T # The "-1" makes reshape flatten the remaining dimensions
test_x_flatten = test_x_orig.reshape(test_x_orig.shape[0], -1).T
# Standardize data to have feature values between 0 and 1.
train_x = train_x_flatten/255.
test_x = test_x_flatten/255.
print ("train_x's shape: " + str(train_x.shape))
print ("test_x's shape: " + str(test_x.shape))
```
$12,288$ equals $64 \times 64 \times 3$ which is the size of one reshaped image vector.
## 3 - Architecture of your model
Now that you are familiar with the dataset, it is time to build a deep neural network to distinguish cat images from non-cat images.
You will build two different models:
- A 2-layer neural network
- An L-layer deep neural network
You will then compare the performance of these models, and also try out different values for $L$.
Let's look at the two architectures.
### 3.1 - 2-layer neural network
<img src="images/2layerNN_kiank.png" style="width:650px;height:400px;">
<caption><center> <u>Figure 2</u>: 2-layer neural network. <br> The model can be summarized as: ***INPUT -> LINEAR -> RELU -> LINEAR -> SIGMOID -> OUTPUT***. </center></caption>
<u>Detailed Architecture of figure 2</u>:
- The input is a (64,64,3) image which is flattened to a vector of size $(12288,1)$.
- The corresponding vector: $[x_0,x_1,...,x_{12287}]^T$ is then multiplied by the weight matrix $W^{[1]}$ of size $(n^{[1]}, 12288)$.
- You then add a bias term and take its relu to get the following vector: $[a_0^{[1]}, a_1^{[1]},..., a_{n^{[1]}-1}^{[1]}]^T$.
- You then repeat the same process.
- You multiply the resulting vector by $W^{[2]}$ and add your intercept (bias).
- Finally, you take the sigmoid of the result. If it is greater than 0.5, you classify it to be a cat.
### 3.2 - L-layer deep neural network
It is hard to represent an L-layer deep neural network with the above representation. However, here is a simplified network representation:
<img src="images/LlayerNN_kiank.png" style="width:650px;height:400px;">
<caption><center> <u>Figure 3</u>: L-layer neural network. <br> The model can be summarized as: ***[LINEAR -> RELU] $\times$ (L-1) -> LINEAR -> SIGMOID***</center></caption>
<u>Detailed Architecture of figure 3</u>:
- The input is a (64,64,3) image which is flattened to a vector of size (12288,1).
- The corresponding vector: $[x_0,x_1,...,x_{12287}]^T$ is then multiplied by the weight matrix $W^{[1]}$ and then you add the intercept $b^{[1]}$. The result is called the linear unit.
- Next, you take the relu of the linear unit. This process could be repeated several times for each $(W^{[l]}, b^{[l]})$ depending on the model architecture.
- Finally, you take the sigmoid of the final linear unit. If it is greater than 0.5, you classify it to be a cat.
### 3.3 - General methodology
As usual you will follow the Deep Learning methodology to build the model:
1. Initialize parameters / Define hyperparameters
2. Loop for num_iterations:
a. Forward propagation
b. Compute cost function
c. Backward propagation
d. Update parameters (using parameters, and grads from backprop)
4. Use trained parameters to predict labels
Let's now implement those two models!
## 4 - Two-layer neural network
**Question**: Use the helper functions you have implemented in the previous assignment to build a 2-layer neural network with the following structure: *LINEAR -> RELU -> LINEAR -> SIGMOID*. The functions you may need and their inputs are:
```python
def initialize_parameters(n_x, n_h, n_y):
...
return parameters
def linear_activation_forward(A_prev, W, b, activation):
...
return A, cache
def compute_cost(AL, Y):
...
return cost
def linear_activation_backward(dA, cache, activation):
...
return dA_prev, dW, db
def update_parameters(parameters, grads, learning_rate):
...
return parameters
```
```
### CONSTANTS DEFINING THE MODEL ####
n_x = 12288 # num_px * num_px * 3
n_h = 7
n_y = 1
layers_dims = (n_x, n_h, n_y)
# GRADED FUNCTION: two_layer_model
def two_layer_model(X, Y, layers_dims, learning_rate = 0.0075, num_iterations = 3000, print_cost=False):
"""
Implements a two-layer neural network: LINEAR->RELU->LINEAR->SIGMOID.
Arguments:
X -- input data, of shape (n_x, number of examples)
Y -- true "label" vector (containing 0 if cat, 1 if non-cat), of shape (1, number of examples)
layers_dims -- dimensions of the layers (n_x, n_h, n_y)
num_iterations -- number of iterations of the optimization loop
learning_rate -- learning rate of the gradient descent update rule
print_cost -- If set to True, this will print the cost every 100 iterations
Returns:
parameters -- a dictionary containing W1, W2, b1, and b2
"""
np.random.seed(1)
grads = {}
costs = [] # to keep track of the cost
m = X.shape[1] # number of examples
(n_x, n_h, n_y) = layers_dims
# Initialize parameters dictionary, by calling one of the functions you'd previously implemented
### START CODE HERE ### (≈ 1 line of code)
parameters = initialize_parameters(n_x, n_h, n_y)
### END CODE HERE ###
# Get W1, b1, W2 and b2 from the dictionary parameters.
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
# Loop (gradient descent)
for i in range(0, num_iterations):
# Forward propagation: LINEAR -> RELU -> LINEAR -> SIGMOID. Inputs: "X, W1, b1, W2, b2". Output: "A1, cache1, A2, cache2".
### START CODE HERE ### (≈ 2 lines of code)
A1, cache1 = linear_activation_forward(X, W1, b1, "relu")
A2, cache2 = linear_activation_forward(A1, W2, b2, "sigmoid")
### END CODE HERE ###
# Compute cost
### START CODE HERE ### (≈ 1 line of code)
cost = compute_cost(A2, Y)
### END CODE HERE ###
# Initializing backward propagation
dA2 = - (np.divide(Y, A2) - np.divide(1 - Y, 1 - A2))
# Backward propagation. Inputs: "dA2, cache2, cache1". Outputs: "dA1, dW2, db2; also dA0 (not used), dW1, db1".
### START CODE HERE ### (≈ 2 lines of code)
dA1, dW2, db2 = linear_activation_backward(dA2, cache2, "sigmoid")
dA0, dW1, db1 = linear_activation_backward(dA1, cache1, "relu")
### END CODE HERE ###
# Set grads['dWl'] to dW1, grads['db1'] to db1, grads['dW2'] to dW2, grads['db2'] to db2
grads['dW1'] = dW1
grads['db1'] = db1
grads['dW2'] = dW2
grads['db2'] = db2
# Update parameters.
### START CODE HERE ### (approx. 1 line of code)
parameters = update_parameters(parameters, grads, learning_rate)
### END CODE HERE ###
# Retrieve W1, b1, W2, b2 from parameters
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
# Print the cost every 100 training example
if print_cost and i % 100 == 0:
print("Cost after iteration {}: {}".format(i, np.squeeze(cost)))
if print_cost and i % 100 == 0:
costs.append(cost)
# plot the cost
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per tens)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
return parameters
```
Run the cell below to train your parameters. See if your model runs. The cost should be decreasing. It may take up to 5 minutes to run 2500 iterations. Check if the "Cost after iteration 0" matches the expected output below, if not click on the square (⬛) on the upper bar of the notebook to stop the cell and try to find your error.
```
parameters = two_layer_model(train_x, train_y, layers_dims = (n_x, n_h, n_y), num_iterations = 2500, print_cost=True)
```
**Expected Output**:
<table>
<tr>
<td> **Cost after iteration 0**</td>
<td> 0.6930497356599888 </td>
</tr>
<tr>
<td> **Cost after iteration 100**</td>
<td> 0.6464320953428849 </td>
</tr>
<tr>
<td> **...**</td>
<td> ... </td>
</tr>
<tr>
<td> **Cost after iteration 2400**</td>
<td> 0.048554785628770206 </td>
</tr>
</table>
Good thing you built a vectorized implementation! Otherwise it might have taken 10 times longer to train this.
Now, you can use the trained parameters to classify images from the dataset. To see your predictions on the training and test sets, run the cell below.
```
predictions_train = predict(train_x, train_y, parameters)
```
**Expected Output**:
<table>
<tr>
<td> **Accuracy**</td>
<td> 1.0 </td>
</tr>
</table>
```
predictions_test = predict(test_x, test_y, parameters)
```
**Expected Output**:
<table>
<tr>
<td> **Accuracy**</td>
<td> 0.72 </td>
</tr>
</table>
**Note**: You may notice that running the model on fewer iterations (say 1500) gives better accuracy on the test set. This is called "early stopping" and we will talk about it in the next course. Early stopping is a way to prevent overfitting.
Congratulations! It seems that your 2-layer neural network has better performance (72%) than the logistic regression implementation (70%, assignment week 2). Let's see if you can do even better with an $L$-layer model.
## 5 - L-layer Neural Network
**Question**: Use the helper functions you have implemented previously to build an $L$-layer neural network with the following structure: *[LINEAR -> RELU]$\times$(L-1) -> LINEAR -> SIGMOID*. The functions you may need and their inputs are:
```python
def initialize_parameters_deep(layers_dims):
...
return parameters
def L_model_forward(X, parameters):
...
return AL, caches
def compute_cost(AL, Y):
...
return cost
def L_model_backward(AL, Y, caches):
...
return grads
def update_parameters(parameters, grads, learning_rate):
...
return parameters
```
```
### CONSTANTS ###
layers_dims = [12288, 20, 7, 5, 1] # 4-layer model
# GRADED FUNCTION: L_layer_model
def L_layer_model(X, Y, layers_dims, learning_rate = 0.0075, num_iterations = 3000, print_cost=False):#lr was 0.009
"""
Implements a L-layer neural network: [LINEAR->RELU]*(L-1)->LINEAR->SIGMOID.
Arguments:
X -- data, numpy array of shape (number of examples, num_px * num_px * 3)
Y -- true "label" vector (containing 0 if cat, 1 if non-cat), of shape (1, number of examples)
layers_dims -- list containing the input size and each layer size, of length (number of layers + 1).
learning_rate -- learning rate of the gradient descent update rule
num_iterations -- number of iterations of the optimization loop
print_cost -- if True, it prints the cost every 100 steps
Returns:
parameters -- parameters learnt by the model. They can then be used to predict.
"""
np.random.seed(1)
costs = [] # keep track of cost
# Parameters initialization. (≈ 1 line of code)
### START CODE HERE ###
parameters = initialize_parameters_deep(layers_dims)
### END CODE HERE ###
# Loop (gradient descent)
for i in range(0, num_iterations):
# Forward propagation: [LINEAR -> RELU]*(L-1) -> LINEAR -> SIGMOID.
### START CODE HERE ### (≈ 1 line of code)
AL, caches = L_model_forward(X, parameters)
### END CODE HERE ###
# Compute cost.
### START CODE HERE ### (≈ 1 line of code)
cost = compute_cost(AL, Y)
### END CODE HERE ###
# Backward propagation.
### START CODE HERE ### (≈ 1 line of code)
grads = L_model_backward(AL, Y, caches)
### END CODE HERE ###
# Update parameters.
### START CODE HERE ### (≈ 1 line of code)
parameters = update_parameters(parameters, grads, learning_rate)
### END CODE HERE ###
# Print the cost every 100 training example
if print_cost and i % 100 == 0:
print ("Cost after iteration %i: %f" %(i, cost))
if print_cost and i % 100 == 0:
costs.append(cost)
# plot the cost
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per tens)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
return parameters
```
You will now train the model as a 4-layer neural network.
Run the cell below to train your model. The cost should decrease on every iteration. It may take up to 5 minutes to run 2500 iterations. Check if the "Cost after iteration 0" matches the expected output below, if not click on the square (⬛) on the upper bar of the notebook to stop the cell and try to find your error.
```
parameters = L_layer_model(train_x, train_y, layers_dims, num_iterations = 2500, print_cost = True)
```
**Expected Output**:
<table>
<tr>
<td> **Cost after iteration 0**</td>
<td> 0.771749 </td>
</tr>
<tr>
<td> **Cost after iteration 100**</td>
<td> 0.672053 </td>
</tr>
<tr>
<td> **...**</td>
<td> ... </td>
</tr>
<tr>
<td> **Cost after iteration 2400**</td>
<td> 0.092878 </td>
</tr>
</table>
```
pred_train = predict(train_x, train_y, parameters)
```
<table>
<tr>
<td>
**Train Accuracy**
</td>
<td>
0.985645933014
</td>
</tr>
</table>
```
pred_test = predict(test_x, test_y, parameters)
```
**Expected Output**:
<table>
<tr>
<td> **Test Accuracy**</td>
<td> 0.8 </td>
</tr>
</table>
Congrats! It seems that your 4-layer neural network has better performance (80%) than your 2-layer neural network (72%) on the same test set.
This is good performance for this task. Nice job!
Though in the next course on "Improving deep neural networks" you will learn how to obtain even higher accuracy by systematically searching for better hyperparameters (learning_rate, layers_dims, num_iterations, and others you'll also learn in the next course).
## 6) Results Analysis
First, let's take a look at some images the L-layer model labeled incorrectly. This will show a few mislabeled images.
```
print_mislabeled_images(classes, test_x, test_y, pred_test)
```
**A few types of images the model tends to do poorly on include:**
- Cat body in an unusual position
- Cat appears against a background of a similar color
- Unusual cat color and species
- Camera Angle
- Brightness of the picture
- Scale variation (cat is very large or small in image)
## 7) Test with your own image (optional/ungraded exercise) ##
Congratulations on finishing this assignment. You can use your own image and see the output of your model. To do that:
1. Click on "File" in the upper bar of this notebook, then click "Open" to go on your Coursera Hub.
2. Add your image to this Jupyter Notebook's directory, in the "images" folder
3. Change your image's name in the following code
4. Run the code and check if the algorithm is right (1 = cat, 0 = non-cat)!
```
## START CODE HERE ##
my_image = "my_image.jpg" # change this to the name of your image file
my_label_y = [1] # the true class of your image (1 -> cat, 0 -> non-cat)
## END CODE HERE ##
fname = "images/" + my_image
image = np.array(ndimage.imread(fname, flatten=False))
my_image = scipy.misc.imresize(image, size=(num_px,num_px)).reshape((num_px*num_px*3,1))
my_image = my_image/255.
my_predicted_image = predict(my_image, my_label_y, parameters)
plt.imshow(image)
print ("y = " + str(np.squeeze(my_predicted_image)) + ", your L-layer model predicts a \"" + classes[int(np.squeeze(my_predicted_image)),].decode("utf-8") + "\" picture.")
```
**References**:
- for auto-reloading external module: http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
| github_jupyter |
# Portfolio Optimization
- 포트폴리오 최적화 이론에 대한 정리 자료
- **포트폴리오**란, 다양한 자산에 분산하여 투자하는 것을 말함
- **분산투자**를 통해 변동성과 위험을 낮출 수 있음 (계란을 한 바구니에 담지 말라)
- **자산 배분이란?** 위험 대비 수익을 최대화하는 포트폴리오를 구성하는 것
---
#### Tactical Asset Allocation(TAA)
- 위험 대비 수익을 "단기적으로" 최대화
- Smart Beta...
#### Strategic Asset Allocation(SAA)
- 위험 대비 수익을 "장기적으로" 최대화
- 일반적으로 위험과 수익률은 비례 관계
- SAA 모델은 위험(변동성)이 낮으면서 수익률이 높은 포트폴리오를 만드는 것이 목표
- Markowitz, Black-Litterman Model...
---
#### 평균 수익률 (Expectation)
- 월 평균 수익률과 같이 기간이 포함된 경우라면 기하평균을 사용
- $E(R) = \sqrt[N]{R_1 \times R_2... \times R_i}$
- $E(R) = \frac{1}{N}\sum_{i=1}^{N}R_i$
#### 변동성 (Varience)
- 변동성(=위험)은 기댓값으로 부터 얼마나 떨어져있는지를 나타내는 분산과 동일
- $\sigma^2 = Var(R) = \frac{1}{N-1}\sum_{i=1}^{N}(R_i-\bar{R})$
#### 공분산 (Covarience)
- 확률변수가 2개 이상일 때 각 확률변수들이 얼마나 퍼져있는지를 나타내는 값
- $Cov(R^1, R^2) = E[(R^1-\bar{R^1})(R^2-\bar{R^2})] = \frac{1}{N-1}\sum_{i=1}^{N}(R^1-\bar{R^1})(R^2-\bar{R^2})$
#### 상관관계 (Correlation Coefficient)
- 확률변수의 절대적 크기에 영향을 받지 않도록 0과 1사이로 단위화시킨 값
- $\rho = \cfrac{Cov(X,Y)}{Std(X)Std(Y)}, (-1\leq\rho\leq1)$
---
## 포트폴리오 기대수익과 위험 측정
#### 포트폴리오 정의
- 주어진 예산 내에서 자산 별 투자 비중
- 투자할 자산 군을 결정하고 결정한 자산 별 수익률, 변동성 및 상관관계를 계산
- 변동성 대비 수익률이 가장 높은 포트폴리오를 구성하는 것이 목표
- $w = portfolio = [w_1, w_2, ... , w_N]^T, where \sum_{i=1}^{N}w_i = 1$
#### 포트폴리오의 기대 수익 (Weighted Average)
- 개별 자산의 기대수익률과 포트폴리오의 비중을 곱해서 합산
- $w = portfolio = [w_1, w_2, ... , w_N]^T, where \sum_{i=1}^{N}w_i = 1$
- $\mu_p = portfolio \times expectation = [w_1, w_2, ... , w_N][R_1, R_2, ... , R_N]^T$
#### 포트폴리오의 변동성 (=위험)
- $\sigma_p^2 = [w_1, w_2, ... , w_N]
\begin{bmatrix}
\sigma_{11} & \sigma_{12} & \cdots & \sigma_{1n} \\
\vdots & \vdots & \ddots & \vdots \\
\sigma_{n1} & \sigma_{n2} & \cdots & \sigma_{n^2}
\end{bmatrix}
\begin{bmatrix}
w_{1} \\
w_{2} \\
\vdots \\
w_{N} \\
\end{bmatrix}
$
```
import math
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
sns.set_style('darkgrid')
%matplotlib inline
def get_dataset(code, start, end):
df = pd.read_pickle("../dataset/{}.p".format(code))[::-1]
df = df[df['date'].between(start, end, inclusive=True)]
df = df.drop(['diff'], axis=1).set_index('date').sort_index()
return df
# KAKAO 2017-01-01 ~ 2018-03-30
# NAVER 2017-01-01 ~ 2018-03-30
kakao = get_dataset('035720', '2017-01-01', '2018-03-30')
naver = get_dataset('035420', '2017-01-01', '2018-03-30')
```
## NAVER vs KAKAO
- 2017-01-01 ~ 2018-03-30 기간의 종가 그래프
- 아래 그래프를 통해 두 종목의 변동성을 비교할 수 있을까?
```
plt.figure(figsize=(12,8))
plt.xlabel("Time Period")
plt.ylabel("Stock Price")
kakao_price = kakao['close']
naver_price = naver['close']
plt.plot(naver_price)
plt.plot(kakao_price)
plt.legend(['NAVER', 'KAKAO'], loc='upper left')
plt.figure(figsize=(12,8))
plt.xlabel("Time Period")
plt.ylabel("Stock Price")
kakao_price = kakao['close']
mean = kakao_price.replace(kakao_price, kakao_price.mean())
plt.plot(kakao_price)
plt.plot(mean)
plt.legend(['KAKAO', 'MEAN'], loc='upper left')
```
## Daily per change
```
plt.figure(figsize=(12,8))
real_returns = kakao_price.pct_change()
plt.bar(real_returns.index,real_returns)
```
## Mean-Varience on Single Stock
```
def income(start, end):
return round((end - start) / start * 100, 2)
def geometric_mean(iterable):
iterable = [i for i in iterable if int(i) is not 0]
a = np.log(iterable)
return np.exp(a.sum()/len(a))
point = kakao_price[0]
result = kakao_price.apply(lambda d: income(point, d))
result.head()
print("Mean of daily income: {}".format(np.mean(result)))
print("Geometric Mean of daily income: {}".format(geometric_mean(result)))
print("Varience of daily income: {}".format(np.var(result)))
print("Standard Deviation of daily income: {}".format(np.std(result)))
```
## Correlation
- r이 -1에 가까울 수록 음의 상관관계, +1에 가까울 수록 양의 상관관계
- r이 -0.1과 +0.1 사이이면, 거의 무시될 수 있는 상관관계
```
naver_price.corr(kakao_price, method='pearson')
```
## Mean-Varience on Portfolio
- 네이버(30%), 카카오(30%), 셀트리온(20%), SK이노베이션(20%)
```
def init_portfolio(stock, ratio, start, end):
dfs = []
for each in stock:
df = get_dataset(each, start, end)['close']
point = df[0]
result = df.apply(lambda d: income(point, d))
dfs.append(result)
return pd.concat(dfs, axis=1, keys=stock)
def port_mean_var(avg_ret_, var_covar_, w_):
port_ret = np.dot(w_, avg_ret_)
port_std = np.dot(np.dot(w_, var_covar_), w_.T)
return port_ret, port_std
stock = ['035420', '035720', '068270', '096770']
ratio = [0.3, 0.3, 0.2, 0.2]
df = init_portfolio(stock, ratio, '2017-01-01', '2018-03-30')
df.head()
avg_ret = df.mean()
var_covar = df.cov()
w = np.array(ratio).T
mean, var = port_mean_var(avg_ret, var_covar, w)
print("Mean of portfolio: {}".format(mean))
print("Varience of portfolio: {}".format(var))
```
| github_jupyter |
# Think Bayes
This notebook presents example code and exercise solutions for Think Bayes.
Copyright 2016 Allen B. Downey
MIT License: https://opensource.org/licenses/MIT
```
# Configure Jupyter so figures appear in the notebook
%matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
%config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import classes from thinkbayes2
from thinkbayes2 import Pmf, Suite
import thinkbayes2
import pandas as pd
import thinkplot
df = pd.read_csv('../data/flea_beetles.csv', delimiter='\t')
species = df.Species.tolist();
names = ['Con','Hep','Hei'];
angles = df.Angle.tolist();
widths = df.Width.tolist();
minW=min(widths)
maxW=max(widths)
nW=maxW-minW+1
minA=min(angles)
maxA=max(angles)
nA=maxA-minA+1
conW = thinkbayes2.EstimatedPdf([w for s,w in zip(species,widths) if s=='Con']).MakePmf(low=minW,high=maxW,n=nW,label='Con');
conA = thinkbayes2.EstimatedPdf([w for s,w in zip(species,angles) if s=='Con']).MakePmf(low=minA,high=maxA,n=nA,label='Con');
hepW = thinkbayes2.EstimatedPdf([w for s,w in zip(species,widths) if s=='Hep']).MakePmf(low=minW,high=maxW,n=nW,label='Hep');
hepA = thinkbayes2.EstimatedPdf([w for s,w in zip(species,angles) if s=='Hep']).MakePmf(low=minA,high=maxA,n=nA,label='Hep');
heiW = thinkbayes2.EstimatedPdf([w for s,w in zip(species,widths) if s=='Hei']).MakePmf(low=minW,high=maxW,n=nW,label='Hei');
heiA = thinkbayes2.EstimatedPdf([w for s,w in zip(species,angles) if s=='Hei']).MakePmf(low=minA,high=maxA,n=nA,label='Hei');
class beetle(Suite):
def Likelihood(self, data, hypo):
w,a = data;
if hypo=='Con':
return conW.Prob(w)*conA.Prob(a)
elif hypo=='Hep':
return hepW.Prob(w)*hepA.Prob(a)
elif hypo=='Hei':
return heiW.Prob(w)*heiA.Prob(a)
print('here')
thinkplot.Pdf(conW)
thinkplot.Pdf(hepW)
thinkplot.Pdf(heiW)
thinkplot.figure()
thinkplot.Pdf(conA)
thinkplot.Pdf(hepA)
thinkplot.Pdf(heiA)
thinkplot.figure()
beetles = beetle(names);
beetles.Update((140,15))
thinkplot.hist(beetles)
die = Pmf([1,0]);
roll = sum([die]*4);
class ur(Suite):
def Likelihood(self, data, hypo):
nTurns = hypo
dist = sum([roll]*nTurns)
return dist.Prob(data)
turns = ur(range(1,20));
for(i in range(100)):
thinkplot.hist(turns)
class Subclass(Suite):
def Likelihood(self, data, hypo):
"""Computes the likelihood of the data under the hypothesis.
data:
hypo:
"""
like = 1
return like
%psource thinkbayes2.EstimatedPdf
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
housing=pd.read_csv("housing.csv")
housing.head()
housing.info()
housing['ocean_proximity'].value_counts()
housing.describe()
%matplotlib inline
housing.hist(bins=50, figsize=(20,15))
plt.show()
```
## Has flaw, will break the next time you run it, even with random.seed, once data updated the randomness will change, hence at one point your model will gegt to see the whole data
(Avoid using this)
```
def train_test_split(data, test_size_ratio):
shuffled_indices=np.random.permutation(len(data))
test_size=int(len(data)*test_size_ratio)
test_indices=shuffled_indices[:test_size]
train_indices=shuffled_indices[test_size:]
return data.iloc[train_indices], data.iloc[test_indices]
train_set, test_set=train_test_split(housing, 0.2)
print(len(train_set))
print(len(test_set))
```
## Best approach to split the dataset
```
from sklearn.model_selection import train_test_split
train_set, test_set=train_test_split(housing, random_state=42, test_size=0.2)
```
# Output variable feature scaling
```
housing["income_cat"]=pd.cut(housing["median_income"], bins=[0., 1.5, 3, 4.5, 6., np.inf], labels=[1,2,3,4,5])
housing["income_cat"].hist()
from sklearn.model_selection import StratifiedShuffleSplit
split = StratifiedShuffleSplit(n_splits=1, test_size=0.2, random_state=42)
for train_index, test_index in split.split(housing, housing["income_cat"]):
strat_train_set = housing.loc[train_index]
strat_test_set = housing.loc[test_index]
strat_test_set["income_cat"].value_counts() / len(strat_test_set)
## Drop "income_cat"
for set_ in (strat_train_set, strat_test_set):
set_.drop("income_cat", axis=1, inplace=True)
housing=strat_train_set.copy()
housing.plot(kind="scatter", x="latitude", y="longitude", alpha=0.1)
housing.plot(kind="scatter", x="longitude", y="latitude", alpha=0.4,
s=housing["population"]/100, label="population", figsize=(10,7),
c="median_house_value", cmap=plt.get_cmap("jet"), colorbar=True,)
plt.legend()
corr_matrix=housing.corr()
corr_matrix
corr_matrix["median_house_value"].sort_values(ascending=False)
from pandas.plotting import scatter_matrix
attributes = ["median_house_value", "median_income", "total_rooms",
"housing_median_age"]
scatter_matrix(housing[attributes], figsize=(12, 8))
plt.show()
housing.plot(kind="scatter", x="median_income", y="median_house_value",
alpha=0.1)
housing["rooms_per_household"] = housing["total_rooms"]/housing["households"]
housing["bedrooms_per_room"] = housing["total_bedrooms"]/housing["total_rooms"]
housing["population_per_household"]=housing["population"]/housing["households"]
corr_matrix = housing.corr()
corr_matrix["median_house_value"].sort_values(ascending=False)
housing = strat_train_set.drop("median_house_value", axis=1)
housing_labels = strat_train_set["median_house_value"].copy()
median = housing["total_bedrooms"].median()
housing["total_bedrooms"].fillna(median, inplace=True)
from sklearn.impute import SimpleImputer
imputer = SimpleImputer(strategy="median")
housing_num = housing.drop("ocean_proximity", axis=1)
imputer.fit(housing_num)
X = imputer.transform(housing_num)
X
housing_tr = pd.DataFrame(X, columns=housing_num.columns,
index=housing_num.index)
housing_cat = housing[["ocean_proximity"]]
housing_cat.head(10)
from sklearn.preprocessing import OneHotEncoder
cat_encoder = OneHotEncoder()
housing_cat_1hot = cat_encoder.fit_transform(housing_cat)
housing_cat_1hot
from sklearn.base import BaseEstimator, TransformerMixin
rooms_ix, bedrooms_ix, population_ix, households_ix = 3, 4, 5, 6
class CombinedAttributesAdder(BaseEstimator, TransformerMixin):
def __init__(self, add_bedrooms_per_room = True): # no *args or **kargs
self.add_bedrooms_per_room = add_bedrooms_per_room
def fit(self, X, y=None):
return self # nothing else to do
def transform(self, X, y=None):
rooms_per_household = X[:, rooms_ix] / X[:, households_ix]
population_per_household = X[:, population_ix] / X[:, households_ix]
if self.add_bedrooms_per_room:
bedrooms_per_room = X[:, bedrooms_ix] / X[:, rooms_ix]
return np.c_[X, rooms_per_household, population_per_household,bedrooms_per_room]
else:
return np.c_[X, rooms_per_household, population_per_household]
attr_adder = CombinedAttributesAdder(add_bedrooms_per_room=False)
housing_extra_attribs = attr_adder.transform(housing.values)
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
num_pipeline = Pipeline([
('imputer', SimpleImputer(strategy="median")),
('attribs_adder', CombinedAttributesAdder()),
('std_scaler', StandardScaler()),
])
housing_num_tr = num_pipeline.fit_transform(housing_num)
from sklearn.compose import ColumnTransformer
num_attribs = list(housing_num)
cat_attribs = ["ocean_proximity"]
full_pipeline = ColumnTransformer([
("num", num_pipeline, num_attribs),
("cat", OneHotEncoder(), cat_attribs),
])
housing_prepared = full_pipeline.fit_transform(housing)
from sklearn.linear_model import LinearRegression
lin_reg = LinearRegression()
lin_reg.fit(housing_prepared, housing_labels)
some_data = housing.iloc[:5]
some_labels = housing_labels.iloc[:5]
some_data_prepared = full_pipeline.transform(some_data)
print("Predictions:", lin_reg.predict(some_data_prepared))
print("Labels:", list(some_labels))
from sklearn.metrics import mean_squared_error
housing_predictions = lin_reg.predict(housing_prepared)
lin_mse = mean_squared_error(housing_labels, housing_predictions)
lin_rmse = np.sqrt(lin_mse)
lin_rmse
from sklearn.tree import DecisionTreeRegressor
tree_reg = DecisionTreeRegressor()
tree_reg.fit(housing_prepared, housing_labels)
housing_predictions = tree_reg.predict(housing_prepared)
tree_mse = mean_squared_error(housing_labels, housing_predictions)
tree_rmse = np.sqrt(tree_mse)
tree_rmse
from sklearn.model_selection import cross_val_score
scores = cross_val_score(tree_reg, housing_prepared, housing_labels,
scoring="neg_mean_squared_error", cv=10)
tree_rmse_scores = np.sqrt(-scores)
def display_scores(scores):
print("Scores:", scores)
print("Mean:", scores.mean())
print("Standard deviation:", scores.std())
display_scores(tree_rmse_scores)
lin_scores = cross_val_score(lin_reg, housing_prepared, housing_labels, scoring="neg_mean_squared_error", cv=10)
lin_rmse_scores = np.sqrt(-lin_scores)
display_scores(lin_rmse_scores)
from sklearn.ensemble import RandomForestRegressor
forest_reg = RandomForestRegressor()
forest_reg.fit(housing_prepared, housing_labels)
rand_scores = cross_val_score(forest_reg, housing_prepared, housing_labels, scoring="neg_mean_squared_error", cv=10)
rand_rmse_scores = np.sqrt(-rand_scores)
display_scores(rand_rmse_scores)
from sklearn.model_selection import GridSearchCV
param_grid = [
{'n_estimators': [3, 10, 30], 'max_features': [2, 4, 6, 8]},
{'bootstrap': [False], 'n_estimators': [3, 10], 'max_features': [2, 3,
4]},
]
forest_reg = RandomForestRegressor()
grid_search = GridSearchCV(forest_reg, param_grid, cv=5,
scoring='neg_mean_squared_error',
return_train_score=True)
grid_search.fit(housing_prepared, housing_labels)
grid_search.best_params_
grid_search.best_estimator_
cvres = grid_search.cv_results_
for mean_score, params in zip(cvres["mean_test_score"], cvres["params"]):
print(np.sqrt(-mean_score), params)
feature_importances = grid_search.best_estimator_.feature_importances_
feature_importances
extra_attribs = ["rooms_per_hhold", "pop_per_hhold", "bedrooms_per_room"]
cat_encoder = full_pipeline.named_transformers_["cat"]
cat_one_hot_attribs = list(cat_encoder.categories_[0])
attributes = num_attribs + extra_attribs + cat_one_hot_attribs
sorted(zip(feature_importances, attributes), reverse=True)
final_model = grid_search.best_estimator_
X_test = strat_test_set.drop("median_house_value", axis=1)
y_test = strat_test_set["median_house_value"].copy()
X_test_prepared = full_pipeline.transform(X_test)
final_predictions = final_model.predict(X_test_prepared)
final_mse = mean_squared_error(y_test, final_predictions)
final_rmse = np.sqrt(final_mse)
final_rmse
from scipy import stats
confidence = 0.95
squared_errors = (final_predictions - y_test) ** 2
np.sqrt(stats.t.interval(confidence, len(squared_errors) - 1,loc=squared_errors.mean(),scale=stats.sem(squared_errors)))
```
| github_jupyter |
# Improved feature engineering
*Anders Poirel - 11-02-2020*
Ideas I'll be building on
- seperating by city (data has different structure between the cities, avoids needing to build a more complex model that captures feature interactions)
- using the lifecycle of the mosquito: new mostiquos become adults 1-3 weeks after eggs are laid in water. Therefore, we could expect a lot of cases if the previous 2-3week/~month was humid
**Note**: I realized I used *median* absolute error instead of mean absolute error in the previous notebook, which explain why my CV scores were so far from the test set scores!
```
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import sklearn
from os.path import join
DATA_PATH = '../data/raw/'
```
## Acquiring the data
```
X_test_o = pd.read_csv(join(DATA_PATH, 'dengue_features_test.csv'))
X_train_o = pd.read_csv(join(DATA_PATH, 'dengue_features_train.csv'))
y_train_o = pd.read_csv(join(DATA_PATH, 'dengue_labels_train.csv'))
```
### Preprocessing
```
X_train = pd.get_dummies(X_train_o, columns = ['city'], drop_first = True)
X_test = pd.get_dummies(X_test_o, columns = ['city'], drop_first = True)
X_train = X_train.drop('week_start_date', axis = 1)
X_test = X_test.drop('week_start_date', axis = 1)
```
Drop features that have correlation 1 with other features
```
X_train.drop(
['reanalysis_sat_precip_amt_mm', 'reanalysis_dew_point_temp_k',
'reanalysis_tdtr_k'],
axis = 1,
inplace = True
)
X_test.drop(
['reanalysis_sat_precip_amt_mm', 'reanalysis_dew_point_temp_k',
'reanalysis_tdtr_k'],
axis = 1,
inplace = True
)
y_train
```
### Precipitation at several time lags
First, we split the data by city:
```
X_train_sj = X_train[X_train['city_sj'] == 1]
X_train_iq = X_train[X_train['city_sj'] == 0]
X_test_sj = X_test[X_test['city_sj'] == 1]
X_test_iq = X_test[X_test['city_sj'] == 0]
y_train_sj = y_train[y_train['city'] == 'sj']['total_cases']
y_train_iq = y_train[y_train['city'] == 'iq']['total_cases']
def precip_n_weeks(k, n, precips):
if k - n < 0:
re turn .0
else:
return precips[k - n]
train_precip_sj = X_train_sj['precipitation_amt_mm']
train_precip_iq = X_train_iq['precipitation_amt_mm']
test_precip_sj = X_test_sj['precipitation_amt_mm']
test_precip_iq = X_test_iq['precipitation_amt_mm']
```
We re-index the series for Iquitos so that they start from 0 and our code can run properly
```
iq_train_index = list(range(len(train_precip_iq)))
iq_test_index = list(range(len(test_precip_iq)))
train_precip_iq.index = iq_train_index
test_precip_iq.index = iq_test_index
X_train_sj['precip_2'] = [precip_n_weeks(k, 2, train_precip_sj)
for k in range(len(train_precip_sj))]
X_train_sj['precip_3'] = [precip_n_weeks(k, 3, train_precip_sj)
for k in range(len(train_precip_sj))]
X_train_sj['precip_4'] = [precip_n_weeks(k, 4, train_precip_sj)
for k in range(len(train_precip_sj))]
X_test_sj['precip_2'] = [precip_n_weeks(k, 2, test_precip_sj)
for k in range(len(test_precip_sj))]
X_test_sj['precip_3'] = [precip_n_weeks(k, 3, test_precip_sj)
for k in range(len(test_precip_sj))]
X_test_sj['precip_4'] = [precip_n_weeks(k, 4, test_precip_sj)
for k in range(len(test_precip_sj))]
X_train_iq['precip_2'] = [precip_n_weeks(k, 2, train_precip_iq)
for k in range(len(train_precip_iq))]
X_train_iq['precip_3'] = [precip_n_weeks(k, 3, train_precip_iq)
for k in range(len(train_precip_iq))]
X_train_iq['precip_4'] = [precip_n_weeks(k, 4, train_precip_iq)
for k in range(len(train_precip_iq))]
X_test_iq['precip_2'] = [precip_n_weeks(k, 2, test_precip_iq)
for k in range(len(test_precip_iq))]
X_test_iq['precip_3'] = [precip_n_weeks(k, 3, test_precip_iq)
for k in range(len(test_precip_iq))]
X_test_iq['precip_4'] = [precip_n_weeks(k, 4, test_precip_iq)
for k in range(len(test_precip_iq))]
```
Let's check that this worked as intended:
```
X_test_sj.head(30)
```
## Building the models
```
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler, PolynomialFeatures
from sklearn.linear_model import LinearRegression, ElasticNet
from sklearn.model_selection import (cross_validate, TimeSeriesSplit,
RandomizedSearchCV)
```
#### ElasticNet with penalty
San Jose:
```
en_sj = Pipeline([
('scale', StandardScaler()),
('impute_m', SimpleImputer()),
('en', LinearRegression())
])
cv_res_sj = cross_validate(
estimator = en_sj,
X = X_train_sj,
y = y_train_sj,
cv = TimeSeriesSplit(n_splits = 10),
scoring = 'neg_mean_absolute_error',
n_jobs = -1
)
en_sj_score = np.mean(cv_res_sj['test_score'])
en_sj_score
en_sj.fit(X_train_sj, y_train_sj)
y_pred_sj = en_sj.predict(X_train_sj)
```
Iquitos:
```
en_iq = Pipeline([
('scale', StandardScaler()),
('impute_m', SimpleImputer()),
('en', ElasticNet(alpha = 10))
])
cv_res_iq = cross_validate(
estimator = en_iq,
X = X_train_iq,
y = y_train_iq,
cv = TimeSeriesSplit(n_splits = 10),
scoring = 'neg_mean_absolute_error',
n_jobs = -1
)
en_iq_score = np.mean(cv_res_iq['test_score'])
en_iq_score
y_train_iq.mean()
```
Something is really strange here... both models have large MAEs (close to the means values of the targets for each)
```
plt.style.use('default')
```
We get the date data for each city:
```
sj_dates = X_train_o[X_train_o['city'] == 'sj']['week_start_date']
iq_dates = X_train_o[X_train_o['city'] == 'iq']['week_start_date']
ax = plt.axes()
ax.plot(sj_dates, y_pred_sj)
ax.plot(sj_dates, y_train_sj)
```
It appears that the model is predicting very close to the mean
### Building a submission
```
submission = pd.read_csv(join(DATA_PATH, 'submission_format.csv'))
y_pred = poly_model_3.predict(X_test)
submission['total_cases'] = np.round(y_pred).astype(int)
submission
submission.to_csv('../models/baseline.csv', index = False)
```
| github_jupyter |
# Advanced Google Search for Machine Learning
> "Advanced Google Search for Machine Learning"
- toc: false
- branch: master
- badges: true
- comments: true
- categories: [machine-learning, google-search]
- image: images/cover_google_search_ml.png
- hide: false
- search_exclude: true
- metadata_key1: metadata_value1
- metadata_key2: metadata_value2

Searching on Google is one of the most important tasks in any development workflow.
It's the central skill for anyone looking practice their coding skills in any development job, be it machine learning or otherwise.
***In this article I want to share 9 tips for how to perform advanced google searches for machine learning***
## 1. Searching machine learning papers after a certain date
`site: arxiv.org machine learning after:2020`
## 2. Searching papers that mention a specific word or sentence within a specific interval date
`site: arxiv.org "artificial general intelligence 2019..2022"`
## 3. Searching repositories that use a specific framework
`site: github.com "pytorch" *& "RNN" after: 2020`
## 4. Searching tools of a certain type
`image annotation tool & "open source" after:2018`
## 5. Searching Images of a Certain Size and Type
`image:panda imagesize:256x256`
## 6. Search social media & hashtags
`@twitter transformers & #artificialintelligence`
## 7. Search for a price of GPUs or Hardware in ML
`GPU $600`
## 8. Exclude topics from a search with "-" operator
Put `-` in front of a word you want to leave out. For example,
``"cloud provider" & "machine learning" -Azure -"Google Cloud" -AWS`
(also search for without the exclude terms to show the difference)
## 9. Search for related sites
Put "`related:`" in front of a web address you already know. For example,
`related:kaggle.com`
## Other important operators:
- `filetype:` to search specific file types
- `intitle:` to search websites whose title contains a certain keyword
- `inurl:` to search websites containing a certain search term in the url
- `intext:` to search for websites containing a certain search term in the text
- `info:` to find information about a specific page
# Just Google It!
Any experienced developer in the machine learning field or anywhere else will
tell you that for pretty much any task, question or concern you have: "Just google
it!". Developing this skill is a must to boost your ability to self-learn
complicated topics that require the combination of a lot of information.
Just google it!
---
If you liked this post, [join Medium](https://lucas-soares.medium.com/membership), [follow](https://lucas-soares.medium.com/), [subscribe to my newsletter](https://lucas-soares.medium.com/subscribe). Also, subscribe to my [youtube channel](https://www.youtube.com/channel/UCu8WF59Scx9f3H1N_FgZUwQ) connect with me on [Tiktok](https://www.tiktok.com/@enkrateialucca?lang=en), [Twitter](https://twitter.com/LucasEnkrateia), [LinkedIn](https://www.linkedin.com/in/lucas-soares-969044167/), [Instagram](https://www.instagram.com/theaugmentedself/)! Thanks and see you next time! :)
---
# References
- https://www.youtube.com/watch?v=cEBkvm0-rg0&list=WL&index=15&t=2s
- https://ahrefs.com/blog/google-advanced-search-operators/
| github_jupyter |
<img src="https://upload.wikimedia.org/wikipedia/commons/4/47/Logo_UTFSM.png" width="200" alt="utfsm-logo" align="left"/>
# MAT281
### Aplicaciones de la Matemática en la Ingeniería
## Proyecto 01: Clasificación de dígitos
### Instrucciones
* Completa tus datos personales (nombre y rol USM) en siguiente celda.
* Debes _pushear_ tus cambios a tu repositorio personal del curso.
* Como respaldo, debes enviar un archivo .zip con el siguiente formato `mXX_projectYY_apellido_nombre.zip` a alonso.ogueda@gmail.com, debe contener todo lo necesario para que se ejecute correctamente cada celda, ya sea datos, imágenes, scripts, etc.
* Se evaluará:
- Soluciones
- Código
- Que Binder esté bien configurado.
- Al presionar `Kernel -> Restart Kernel and Run All Cells` deben ejecutarse todas las celdas sin error.
__Nombre__: Claudia Alvarez Latuz
__Rol__: 201610006-7
## Clasificación de dígitos
En este laboratorio realizaremos el trabajo de reconocer un dígito a partir de una imagen.
## Contenidos
* [K Nearest Neighbours](#k_nearest_neighbours)
* [Exploración de Datos](#data_exploration)
* [Entrenamiento y Predicción](#train_and_prediction)
* [Selección de Modelo](#model_selection)
<a id='k_neirest_neighbours'></a>
## K Nearest Neighbours
El algoritmo **k Nearest Neighbors** es un método no paramétrico: una vez que el parámetro $k$ se ha fijado, no se busca obtener ningún parámetro adicional.
Sean los puntos $x^{(i)} = (x^{(i)}_1, ..., x^{(i)}_n)$ de etiqueta $y^{(i)}$ conocida, para $i=1, ..., m$.
El problema de clasificación consiste en encontrar la etiqueta de un nuevo punto $x=(x_1, ..., x_m)$ para el cual no conocemos la etiqueta.
La etiqueta de un punto se obtiene de la siguiente forma:
* Para $k=1$, **1NN** asigna a $x$ la etiqueta de su vecino más cercano.
* Para $k$ genérico, **kNN** asigna a $x$ la etiqueta más popular de los k vecinos más cercanos.
El modelo subyacente a kNN es el conjunto de entrenamiento completo. A diferencia de otros métodos que efectivamente generalizan y resumen la información (como regresión logística, por ejemplo), cuando se necesita realizar una predicción, el algoritmo kNN mira **todos** los datos y selecciona los k datos más cercanos, para regresar la etiqueta más popular/más común. Los datos no se resumen en parámetros, sino que siempre deben mantenerse en memoria. Es un método por tanto que no escala bien con un gran número de datos.
En caso de empate, existen diversas maneras de desempatar:
* Elegir la etiqueta del vecino más cercano (problema: no garantiza solución).
* Elegir la etiqueta de menor valor (problema: arbitrario).
* Elegir la etiqueta que se obtendría con $k+1$ o $k-1$ (problema: no garantiza solución, aumenta tiempo de cálculo).
La cercanía o similaridad entre los datos se mide de diversas maneras, pero en general depende del tipo de datos y del contexto.
* Para datos reales, puede utilizarse cualquier distancia, siendo la **distancia euclidiana** la más utilizada. También es posible ponderar unas componentes más que otras. Resulta conveniente normalizar para poder utilizar la noción de distancia más naturalmente.
* Para **datos categóricos o binarios**, suele utilizarse la distancia de Hamming.
A continuación, una implementación de "bare bones" en numpy:
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
def knn_search(X, k, x):
""" find K nearest neighbours of data among D """
# Distancia euclidiana
d = np.linalg.norm(X - x, axis=1)
# Ordenar por cercania
idx = np.argsort(d)
# Regresar los k mas cercanos
id_closest = idx[:k]
return id_closest, d[id_closest].max()
def knn(X,Y,k,x):
# Obtener los k mas cercanos
k_closest, dmax = knn_search(X, k, x)
# Obtener las etiquetas
Y_closest = Y[k_closest]
# Obtener la mas popular
counts = np.bincount(Y_closest.flatten())
# Regresar la mas popular (cualquiera, si hay empate)
return np.argmax(counts), k_closest, dmax
def plot_knn(X, Y, k, x):
y_pred, neig_idx, dmax = knn(X, Y, k, x)
# plotting the data and the input point
fig = plt.figure(figsize=(8, 8))
plt.plot(x[0, 0], x[0, 1], 'ok', ms=16)
m_ob = Y[:, 0] == 0
plt.plot(X[m_ob, 0], X[m_ob, 1], 'ob', ms=8)
m_sr = Y[:,0] == 1
plt.plot(X[m_sr, 0], X[m_sr, 1], 'sr', ms=8)
# highlighting the neighbours
plt.plot(X[neig_idx, 0], X[neig_idx, 1], 'o', markerfacecolor='None', markersize=24, markeredgewidth=1)
# Plot a circle
x_circle = dmax * np.cos(np.linspace(0, 2*np.pi, 360)) + x[0, 0]
y_circle = dmax * np.sin(np.linspace(0, 2*np.pi, 360)) + x[0, 1]
plt.plot(x_circle, y_circle, 'k', alpha=0.25)
plt.show();
# Print result
if y_pred==0:
print("Prediccion realizada para etiqueta del punto = {} (circulo azul)".format(y_pred))
else:
print("Prediccion realizada para etiqueta del punto = {} (cuadrado rojo)".format(y_pred))
```
Puedes ejecutar varias veces el código anterior, variando el número de vecinos `k` para ver cómo afecta el algoritmo.
```
k = 3 # hyper-parameter
N = 100
X = np.random.rand(N, 2) # random dataset
Y = np.array(np.random.rand(N) < 0.4, dtype=int).reshape(N, 1) # random dataset
x = np.random.rand(1, 2) # query point
# performing the search
plot_knn(X, Y, k, x)
```
<a id='data_exploration'></a>
## Exploración de los datos
A continuación se carga el conjunto de datos a utilizar, a través del sub-módulo `datasets` de `sklearn`.
```
import pandas as pd
from sklearn import datasets
digits_dict = datasets.load_digits()
print(digits_dict["DESCR"])
digits_dict.keys()
digits_dict["target"]
```
A continuación se crea dataframe declarado como `digits` con los datos de `digits_dict` tal que tenga 65 columnas, las 64
primeras a la representación de la imagen en escala de grises (0-blanco, 255-negro) y la última correspondiente al dígito (`target`) con el nombre _target_.
```
digits = (
pd.DataFrame(
digits_dict["data"],
)
.rename(columns=lambda x: f"c{x:02d}")
.assign(target=digits_dict["target"])
.astype(int)
)
digits.head()
```
### Ejercicio 1
**_(10 puntos)_**
**Análisis exploratorio:** Realiza tu análisis exploratorio, no debes olvidar nada! Recuerda, cada análisis debe responder una pregunta.
Algunas sugerencias:
* ¿Cómo se distribuyen los datos?
* ¿Cuánta memoria estoy utilizando?
* ¿Qué tipo de datos son?
* ¿Cuántos registros por clase hay?
* ¿Hay registros que no se correspondan con tu conocimiento previo de los datos?
```
digits.describe().T[['min','max']]
digits.info(verbose = False, memory_usage='deep')
digits.assign(count=np.ones(len(digits["target"]))).groupby('target').sum()['count']
```
**Respuestas:**
_Los datos estan en un dataframe de (1797,65) descrito en el enunciado. Los valores de cXX van desde 0 hasta 16. La memoria utilizada es de 456.4 KB, los datos son del tipo int32 y la cantidad de registros por clase es la calculada en la celda anterior. Me sorprende que los datos vayan de 0 a 16 ya que al ser datos de una imagen en RGB deberían ir de 0 a 250._
### Ejercicio 2
**_(10 puntos)_**
**Visualización:** Para visualizar los datos utilizaremos el método `imshow` de `matplotlib`. Resulta necesario convertir el arreglo desde las dimensiones (1,64) a (8,8) para que la imagen sea cuadrada y pueda distinguirse el dígito. Superpondremos además el label correspondiente al dígito, mediante el método `text`. Esto nos permitirá comparar la imagen generada con la etiqueta asociada a los valores. Realizaremos lo anterior para los primeros 25 datos del archivo.
```
digits_dict["images"][0]
```
Visualiza imágenes de los dígitos utilizando la llave `images` de `digits_dict`.
Sugerencia: Utiliza `plt.subplots` y el método `imshow`. Puedes hacer una grilla de varias imágenes al mismo tiempo!
```
nx, ny = 5, 5
fig, axs = plt.subplots(nx, ny, figsize=(12, 12))
for i in range(nx):
for j in range(ny):
axs[i][j].imshow(digits_dict["images"][ny*i+j],cmap='gray_r')
```
<a id='train_and_prediction'></a>
## Entrenamiento y Predicción
Se utilizará la implementación de `scikit-learn` llamada `KNeighborsClassifier` (el cual es un _estimator_) que se encuentra en `neighbors`.
Utiliza la métrica por defecto.
```
from sklearn.neighbors import KNeighborsClassifier
X = digits.drop(columns="target").values
y = digits["target"].values
```
### Ejercicio 3
**_(10 puntos)_**
Entrenar utilizando todos los datos. Además, recuerda que `k` es un hiper-parámetro, por lo tanto prueba con distintos tipos `k` y obten el `score` desde el modelo.
```
k_array = np.arange(1, 101)
for k in k_array:
knn = KNeighborsClassifier(k)
knn.fit(X, y)
print('Utilizando todos los datos y k= {:.0f}'
.format(k),' se obtiene score= {:.4f}'
.format(knn.score(X, y)))
```
**Preguntas**
* ¿Cuál fue la métrica utilizada?
* ¿Por qué entrega estos resultados? En especial para k=1.
* ¿Por qué no se normalizó o estandarizó la matriz de diseño?
_La metrica utilizada es la que se da por defecto: **minkowski**. Para k=1 el score es perfecto ya que los datos que se ocupan para hacer fit son los mismos que se ocuparon para hacer training. No se normalizó o estandarizó la matriz de diseño pues aumentaría la complejidad._
### Ejercicio 4
**_(10 puntos)_**
Divide los datos en _train_ y _test_ utilizando la función preferida del curso. Para reproducibilidad utiliza `random_state=42`. A continuación, vuelve a ajustar con los datos de _train_ y con los distintos valores de _k_, pero en esta ocasión calcula el _score_ con los datos de _test_.
¿Qué modelo escoges?
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
for k in k_array:
knn = KNeighborsClassifier(k)
knn.fit(X_train, y_train)
print('Utilizando todos los datos y k= {:.0f}'
.format(k),' se obtiene score= {:.4f}'
.format(knn.score(X_test, y_test)))
```
<a id='model_selection'></a>
_Eligiría k=6 ya que es el que tiene mejor score._
## Selección de Modelo
### Ejercicio 5
**_(15 puntos)_**
**Curva de Validación**: Replica el ejemplo del siguiente [link](https://scikit-learn.org/stable/auto_examples/model_selection/plot_validation_curve.html#sphx-glr-auto-examples-model-selection-plot-validation-curve-py) pero con el modelo, parámetros y métrica adecuada.
¿Qué podrías decir de la elección de `k`?
```
from sklearn.model_selection import validation_curve
param_range = np.arange(1, 101)
train_scores, test_scores = validation_curve(KNeighborsClassifier(),X, y, param_name="n_neighbors",cv=5, param_range=param_range, n_jobs=1)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
plt.figure(figsize=(12, 8))
plt.title("Validation Curve with KNN")
plt.xlabel(r"$\gamma$")
plt.ylabel("Score")
plt.ylim(0.0, 1.1)
lw = 2
plt.semilogx(param_range, train_scores_mean, label="Training score",
color="darkorange", lw=lw)
plt.fill_between(param_range, train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std, alpha=0.2,
color="darkorange", lw=lw)
plt.semilogx(param_range, test_scores_mean, label="Cross-validation score",
color="navy", lw=lw)
plt.fill_between(param_range, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.2,
color="navy", lw=lw)
plt.legend(loc="best")
plt.show();
plt.figure(figsize=(12, 8))
plt.title("Validation Curve with KNN")
plt.xlabel(r"$\gamma$")
plt.ylabel("Score")
lw = 2
plt.semilogx(param_range, train_scores_mean, label="Training score",
color="darkorange", lw=lw)
plt.fill_between(param_range, train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std, alpha=0.2,
color="darkorange", lw=lw)
plt.semilogx(param_range, test_scores_mean, label="Cross-validation score",
color="navy", lw=lw)
plt.fill_between(param_range, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.2,
color="navy", lw=lw)
plt.legend(loc="best")
plt.show();
```
**Pregunta**
* ¿Qué refleja este gráfico?
* ¿Qué conclusiones puedes sacar a partir de él?
* ¿Qué patrón se observa en los datos, en relación a los números pares e impares? ¿Porqué sucede esto?
_El grafico muestra como varía el score máximo con k. Es claro que Training siempre esta por sobre Cros-validation. Se puede concluir que un número k menor es una buena opción ya que las curvan parecen ir decreciendo, en especifico para Training k=1 es el máximo score por lo visto en el ejercicio 3 y para Cros-validation parece ser k=2. Un patrón que se observa es que los valores van creciendo y decreciendo cuando cambian de par a impar, pero es no siempre. Esto se debe a que el algoritmo debe elegir entre empates cuando se tiene k par._
### Ejercicio 6
**_(15 puntos)_**
**Búsqueda de hiper-parámetros con validación cruzada:** Utiliza `sklearn.model_selection.GridSearchCV` para obtener la mejor estimación del parámetro _k_. Prueba con valores de _k_ desde 2 a 100.
```
from sklearn.model_selection import GridSearchCV
parameters = {'n_neighbors':np.arange(2, 100)}
digits_gscv = GridSearchCV(KNeighborsClassifier(), parameters, verbose=1, cv=5)
digits_results = digits_gscv.fit(X, y)
# Best params
digits_results.best_params_['n_neighbors']
```
**Pregunta**
* ¿Cuál es el mejor valor de _k_?
* ¿Es consistente con lo obtenido en el ejercicio anterior?
_El mejor valor fue k=2, lo cual es consistentes con el ejercicio anterior._
### Ejercicio 7
**_(10 puntos)_**
__Visualizando datos:__ A continuación se provee código para comparar las etiquetas predichas vs las etiquetas reales del conjunto de _test_.
* Define la variable `best_knn` que corresponde al mejor estimador `KNeighborsClassifier` obtenido.
* Ajusta el estimador anterior con los datos de entrenamiento.
* Crea el arreglo `y_pred` prediciendo con los datos de test.
_Hint:_ `digits_gscv.best_estimator_` te entrega una instancia `estimator` del mejor estimador encontrado por `GridSearchCV`.
```
best_knn = digits_gscv.best_estimator_
best_knn.fit(X_train,y_train)
y_pred = best_knn.predict(X_test)
# Mostrar los datos correctos
mask = (y_pred == y_test)
X_aux = X_test[mask]
y_aux_true = y_test[mask]
y_aux_pred = y_pred[mask]
# We'll plot the first 100 examples, randomly choosen
nx, ny = 5, 5
fig, ax = plt.subplots(nx, ny, figsize=(12,12))
for i in range(nx):
for j in range(ny):
index = j + ny * i
data = X_aux[index, :].reshape(8,8)
label_pred = str(int(y_aux_pred[index]))
label_true = str(int(y_aux_true[index]))
ax[i][j].imshow(data, interpolation='nearest', cmap='gray_r')
ax[i][j].text(0, 0, label_pred, horizontalalignment='center', verticalalignment='center', fontsize=10, color='green')
ax[i][j].text(7, 0, label_true, horizontalalignment='center', verticalalignment='center', fontsize=10, color='blue')
ax[i][j].get_xaxis().set_visible(False)
ax[i][j].get_yaxis().set_visible(False)
plt.show()
```
Modifique el código anteriormente provisto para que muestre los dígitos incorrectamente etiquetados, cambiando apropiadamente la máscara. Cambie también el color de la etiqueta desde verde a rojo, para indicar una mala etiquetación.
```
# Mostrar los datos incorrectos
mask = (y_pred != y_test)
X_aux = X_test[mask]
y_aux_true = y_test[mask]
y_aux_pred = y_pred[mask]
ni = X_aux.shape[0]
fig, ax = plt.subplots(1, ni, figsize=(12,12))
for i in range(ni):
data = X_aux[i, :].reshape(8,8)
label_pred = str(int(y_aux_pred[i]))
label_true = str(int(y_aux_true[i]))
ax[i].imshow(data, interpolation='nearest', cmap='gray_r')
ax[i].text(0, 0, label_pred, horizontalalignment='center', verticalalignment='center', fontsize=10, color='red')
ax[i].text(7, 0, label_true, horizontalalignment='center', verticalalignment='center', fontsize=10, color='blue')
ax[i].get_xaxis().set_visible(False)
ax[i].get_yaxis().set_visible(False)
plt.show()
```
**Pregunta**
* Solo utilizando la inspección visual, ¿Por qué crees que falla en esos valores?
_Visualmente la predicción falla pues los valores están mal dibujados y además son números similares (3 y 9, 1y 8, 0 y 6)_
### Ejercicio 8
**_(10 puntos)_**
**Matriz de confusión:** Grafica la matriz de confusión.
**Importante!** Al principio del curso se entregó una versión antigua de `scikit-learn`, por lo cual es importante que actualicen esta librearía a la última versión para hacer uso de `plot_confusion_matrix`. Hacerlo es tan fácil como ejecutar `conda update -n mat281 -c conda-forge scikit-learn` en la terminal de conda.
```
from sklearn.metrics import plot_confusion_matrix
fig, ax = plt.subplots(figsize=(12, 12))
plot_confusion_matrix(best_knn, X_test, y_test, ax=ax)
```
**Pregunta**
* ¿Cuáles son las etiquetas con mejores y peores predicciones?
* Con tu conocimiento previo del problema, ¿Por qué crees que esas etiquetas son las que tienen mejores y peores predicciones?
_Las etiquetas con mejores predicciones son 1,2,3,5 y 6 ya que no tienen ninguna incorrecta. Las peores son 4,7,8 y 9, donde 9 es la peor con 4 errores. Según como se tomaron los datos creo que las mejores son números que no son parecidos a ningún otro. Este no es el caso de los peores, ya que por ej. el número 9 fácilmente se confunde con otros._
### Ejercicio 9
**_(10 puntos)_**
**Curva de aprendizaje:** Replica el ejemplo del siguiente [link](https://scikit-learn.org/stable/auto_examples/model_selection/plot_learning_curve.html#sphx-glr-auto-examples-model-selection-plot-learning-curve-py) pero solo utilizando un modelo de KNN con el hiperparámetro _k_ seleccionado anteriormente.
```
def plot_learning_curve(estimator, title, X, y, axes=None, ylim=None, cv=None,
n_jobs=None, train_sizes=np.linspace(.1, 1.0, 5)):
"""
Generate 3 plots: the test and training learning curve, the training
samples vs fit times curve, the fit times vs score curve.
Parameters
----------
estimator : object type that implements the "fit" and "predict" methods
An object of that type which is cloned for each validation.
title : string
Title for the chart.
X : array-like, shape (n_samples, n_features)
Training vector, where n_samples is the number of samples and
n_features is the number of features.
y : array-like, shape (n_samples) or (n_samples, n_features), optional
Target relative to X for classification or regression;
None for unsupervised learning.
axes : array of 3 axes, optional (default=None)
Axes to use for plotting the curves.
ylim : tuple, shape (ymin, ymax), optional
Defines minimum and maximum yvalues plotted.
cv : int, cross-validation generator or an iterable, optional
Determines the cross-validation splitting strategy.
Possible inputs for cv are:
- None, to use the default 5-fold cross-validation,
- integer, to specify the number of folds.
- :term:`CV splitter`,
- An iterable yielding (train, test) splits as arrays of indices.
For integer/None inputs, if ``y`` is binary or multiclass,
:class:`StratifiedKFold` used. If the estimator is not a classifier
or if ``y`` is neither binary nor multiclass, :class:`KFold` is used.
Refer :ref:`User Guide <cross_validation>` for the various
cross-validators that can be used here.
n_jobs : int or None, optional (default=None)
Number of jobs to run in parallel.
``None`` means 1 unless in a :obj:`joblib.parallel_backend` context.
``-1`` means using all processors. See :term:`Glossary <n_jobs>`
for more details.
train_sizes : array-like, shape (n_ticks,), dtype float or int
Relative or absolute numbers of training examples that will be used to
generate the learning curve. If the dtype is float, it is regarded as a
fraction of the maximum size of the training set (that is determined
by the selected validation method), i.e. it has to be within (0, 1].
Otherwise it is interpreted as absolute sizes of the training sets.
Note that for classification the number of samples usually have to
be big enough to contain at least one sample from each class.
(default: np.linspace(0.1, 1.0, 5))
"""
if axes is None:
_, axes = plt.subplots(1, 3, figsize=(20, 5))
axes[0].set_title(title)
if ylim is not None:
axes[0].set_ylim(*ylim)
axes[0].set_xlabel("Training examples")
axes[0].set_ylabel("Score")
train_sizes, train_scores, test_scores, fit_times, _ = \
learning_curve(estimator, X, y, cv=cv, n_jobs=n_jobs,
train_sizes=train_sizes,
return_times=True)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
fit_times_mean = np.mean(fit_times, axis=1)
fit_times_std = np.std(fit_times, axis=1)
# Plot learning curve
axes[0].grid()
axes[0].fill_between(train_sizes, train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std, alpha=0.1,
color="r")
axes[0].fill_between(train_sizes, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.1,
color="g")
axes[0].plot(train_sizes, train_scores_mean, 'o-', color="r",
label="Training score")
axes[0].plot(train_sizes, test_scores_mean, 'o-', color="g",
label="Cross-validation score")
axes[0].legend(loc="best")
# Plot n_samples vs fit_times
axes[1].grid()
axes[1].plot(train_sizes, fit_times_mean, 'o-')
axes[1].fill_between(train_sizes, fit_times_mean - fit_times_std,
fit_times_mean + fit_times_std, alpha=0.1)
axes[1].set_xlabel("Training examples")
axes[1].set_ylabel("fit_times")
axes[1].set_title("Scalability of the model")
# Plot fit_time vs score
axes[2].grid()
axes[2].plot(fit_times_mean, test_scores_mean, 'o-')
axes[2].fill_between(fit_times_mean, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.1)
axes[2].set_xlabel("fit_times")
axes[2].set_ylabel("Score")
axes[2].set_title("Performance of the model")
return plt
from sklearn.model_selection import learning_curve
from sklearn.model_selection import ShuffleSplit
fig, axes = plt.subplots(3, 1, figsize=(10, 15))
title = "Learning Curves (KNeighborsClassifier)"
cv = ShuffleSplit(n_splits=100, test_size=0.2, random_state=0)
estimator = best_knn
plot_learning_curve(estimator, title, X, y, axes=axes, ylim=(0.7, 1.01),cv=cv, n_jobs=4)
plt.show()
```
**Pregunta**
* ¿Qué refleja este gráfico?
* ¿Qué conclusiones puedes sacar a partir de él?
* ¿En qué crees que hay que poner más atención a la hora de trabajar con un problema de clasificación?
_El primer gráfico muestra los Training score y Cross-validation score para diferentes training examples. De él se puede apreciar que la curva de score para Training y Cross-validation tienen el mismo comportamiento (parten bajos y luego crecen hasta 'estabilizarce'), pero Training esta por sobre Cross-validation._
_El segundo gráfico muestra el tiempo de ejecución para distintos training examples. Se puede concluir que el tiempo de ejecución aumente de manera considerable (quizás exponencial) en relación con el tamaño de la muestra._
_El tercer gráfico muestra la relación de los tiempos del segundo gráfico con los scores del primero. Nos dice que a partir del tiempo 0.04, el score no cambia considerablemente, por lo que uniendo esta información con la del gráfico anterior no serían necesarios más de 800 training examples._
_A mi parecer hay que poner más atención con el score en relación a los training examples ya que es algo fácil de manejar._
| github_jupyter |
# T1499.002 - Service Exhaustion Flood
Adversaries may target the different network services provided by systems to conduct a DoS. Adversaries often target DNS and web services, however others have been targeted as well.(Citation: Arbor AnnualDoSreport Jan 2018) Web server software can be attacked through a variety of means, some of which apply generally while others are specific to the software being used to provide the service.
One example of this type of attack is known as a simple HTTP flood, where an adversary sends a large number of HTTP requests to a web server to overwhelm it and/or an application that runs on top of it. This flood relies on raw volume to accomplish the objective, exhausting any of the various resources required by the victim software to provide the service.(Citation: Cloudflare HTTPflood)
Another variation, known as a SSL renegotiation attack, takes advantage of a protocol feature in SSL/TLS. The SSL/TLS protocol suite includes mechanisms for the client and server to agree on an encryption algorithm to use for subsequent secure connections. If SSL renegotiation is enabled, a request can be made for renegotiation of the crypto algorithm. In a renegotiation attack, the adversary establishes a SSL/TLS connection and then proceeds to make a series of renegotiation requests. Because the cryptographic renegotiation has a meaningful cost in computation cycles, this can cause an impact to the availability of the service when done in volume.(Citation: Arbor SSLDoS April 2012)
## Atomic Tests:
Currently, no tests are available for this technique.
## Detection
Detection of Endpoint DoS can sometimes be achieved before the effect is sufficient to cause significant impact to the availability of the service, but such response time typically requires very aggressive monitoring and responsiveness. Typical network throughput monitoring tools such as netflow, SNMP, and custom scripts can be used to detect sudden increases in circuit utilization.(Citation: Cisco DoSdetectNetflow) Real-time, automated, and qualitative study of the network traffic can identify a sudden surge in one type of protocol can be used to detect an attack as it starts.
In addition to network level detections, endpoint logging and instrumentation can be useful for detection. Attacks targeting web applications may generate logs in the web server, application server, and/or database server that can be used to identify the type of attack, possibly before the impact is felt.
Externally monitor the availability of services that may be targeted by an Endpoint DoS.
| github_jupyter |
```
''' IMPORTS, RELOAD THIS BLOCK WHEN NEEDED '''
from IPython.display import display
from GoogleImageSpider import *
from FaceClassifier import *
from ImageClassifier import *
import certifi, urllib3
import time
import json
import os
path = "trump_images"
start_time = time.time()
def print_time():
# TIMING CONTROL
elapsed_time = time.time() - start_time
mins = int(elapsed_time / 60)
secs = elapsed_time - (mins * 60)
print("Accumulative time: %02d:%02d" % (mins, int(secs % 60)))
''' FIRST STEP GET A "SMALL" SAMPLE OF TRUMP IMAGES, MAYBE 300 or 400 IMAGES '''
if not os.path.exists(path):
os.makedirs(path)
start_time = time.time()
# SECOND STEP DOWNLOAD IMAGES FROM EACH POKEMON NAME:
gis = GoogleImageSpider()
gis.get_images("trump", 300) #"trump pictures"
print ("Download and save..")
gis.save_images("trump", path)
gis.clear()
print ("Trump images downloaded")
print_time()
''' SECOND STEP: DETECT FACES AND EXPORT TO A NEW FOLDER '''
from PIL import Image
start_time = time.time()
destin_path = "faces_detected"
display_faces = False
face_size = 256
fc = FaceClassifier(face_resize=face_size)
#fc.train(show_train_images=True)
for image_filename in os.listdir(path):
image_path = path + "/" + image_filename
print ("Processing image: " + image_path)
#print (fc.predict(image_path))
data = fc.detect_face(image_path, grayscale_output = False)
for face in data:
if not face[0] is None:
img = Image.fromarray(face[0])
img.save(destin_path + "/" + image_filename)
if display_faces:
display(img)
print_time()
print ("Done!")
''' FOURTH STEP: IS TO TRAIN A NETWORK WITH THIS IMAGES (I MADE ONE SIMPLE
CONVOLUTIONAL CLASSIFIER, FEEL FREE TO UPDATE IT AND SEND A PUSH IT TO THE REPO)'''
from ImageClassifier import *
good_images_folder = "trump_classifier/good"
bad_images_folder = "trump_classifier/bad"
start_time = time.time()
ic = ImageClassifier(good_images_folder, bad_images_folder, 256, batch_size=50, training_epochs=2000, \
test_batch_percentage = 0) # Once proved that works, we need to use all possible inputs
#ic.test()
ic.train()
''' LAST STEP: FINALLY WE MIX ALL THAT TOGETHER, DOWNLOAD ALL THE DATASET, NORMALIZE,
AND LET OUR NEURAL NETWORK DISCRIMINATE THEM TO GET ONLY THE GOOD ONES.
THIS IS PRETTY MUCH COPY OF THE FIRST STEP CODE IMPROVED.
WE ARRIVE HERE IN LIKE 15 MINUTES, BUT NOW 100 IMAGES PER 949 POKEMONS WILL TAKE A WHILE..
TAKE A COFEE AND COME BACK IN A FEW HOURS ;)
GOOD NEWS IS THAT IS COMPLETELY FREE OF HUMAN INTERACTION '''
originals_temp_path = "trump_original_images"
source_path = "trump_destin_images"
good_images_folder = source_path+"/good"
bad_images_folder = source_path+"/bad"
images_to_download = 5000
start_time = time.time()
if not os.path.exists(source_path):
os.makedirs(source_path)
# DOWNLOAD IMAGES:
print ("Downloading images..")
gis = GoogleImageSpider()
gis.get_images("trump", images_to_download)
gis.save_images("trump", originals_temp_path)
print_time()
# DETECT AND CROP FACES:
print ("Detect and crop faces..")
face_size = 256
fc = FaceClassifier(face_resize=face_size)
for image_filename in os.listdir(originals_temp_path):
image_path = originals_temp_path + "/" + image_filename
print ("Processing image: " + image_path)
data = fc.detect_face(image_path, grayscale_output = False)
for face in data:
if not face[0] is None:
img = Image.fromarray(face[0])
img.save(source_path + "/" + image_filename)
print_time()
# RUN THE CNN TO CLASSSIFY GOOD AND BAD:
print ("Loading CNN..")
ic = ImageClassifier(None, None, 256)
ic.load()
print_time()
print ("Detect and crop faces..")
ic.run(source_path, good_images_folder, bad_images_folder, good_percent_treshold=85, delete_images=True)
print_time()
# END:
sys.stdout.flush() # For python command line
print ("Dataset finally done!")
```

| github_jupyter |
# T1018 - Remote System Discovery
Adversaries may attempt to get a listing of other systems by IP address, hostname, or other logical identifier on a network that may be used for Lateral Movement from the current system. Functionality could exist within remote access tools to enable this, but utilities available on the operating system could also be used such as [Ping](https://attack.mitre.org/software/S0097) or <code>net view</code> using [Net](https://attack.mitre.org/software/S0039). Adversaries may also use local host files (ex: <code>C:\Windows\System32\Drivers\etc\hosts</code> or <code>/etc/hosts</code>) in order to discover the hostname to IP address mappings of remote systems.
Specific to macOS, the <code>bonjour</code> protocol exists to discover additional Mac-based systems within the same broadcast domain.
Within IaaS (Infrastructure as a Service) environments, remote systems include instances and virtual machines in various states, including the running or stopped state. Cloud providers have created methods to serve information about remote systems, such as APIs and CLIs. For example, AWS provides a <code>DescribeInstances</code> API within the Amazon EC2 API and a <code>describe-instances</code> command within the AWS CLI that can return information about all instances within an account.(Citation: Amazon Describe Instances API)(Citation: Amazon Describe Instances CLI) Similarly, GCP's Cloud SDK CLI provides the <code>gcloud compute instances list</code> command to list all Google Compute Engine instances in a project, and Azure's CLI <code>az vm list</code> lists details of virtual machines.(Citation: Google Compute Instances)(Citation: Azure VM List)
## Atomic Tests
```
#Import the Module before running the tests.
# Checkout Jupyter Notebook at https://github.com/cyb3rbuff/TheAtomicPlaybook to run PS scripts.
Import-Module /Users/0x6c/AtomicRedTeam/atomics/invoke-atomicredteam/Invoke-AtomicRedTeam.psd1 - Force
```
### Atomic Test #1 - Remote System Discovery - net
Identify remote systems with net.exe.
Upon successful execution, cmd.exe will execute `net.exe view` and display results of local systems on the network that have file and print sharing enabled.
**Supported Platforms:** windows
#### Attack Commands: Run with `command_prompt`
```command_prompt
net view /domain
net view
```
```
Invoke-AtomicTest T1018 -TestNumbers 1
```
### Atomic Test #2 - Remote System Discovery - net group Domain Computers
Identify remote systems with net.exe querying the Active Directory Domain Computers group.
Upon successful execution, cmd.exe will execute cmd.exe against Active Directory to list the "Domain Computers" group. Output will be via stdout.
**Supported Platforms:** windows
#### Attack Commands: Run with `command_prompt`
```command_prompt
net group "Domain Computers" /domain
```
```
Invoke-AtomicTest T1018 -TestNumbers 2
```
### Atomic Test #3 - Remote System Discovery - nltest
Identify domain controllers for specified domain.
Upon successful execution, cmd.exe will execute nltest.exe against a target domain to retrieve a list of domain controllers. Output will be via stdout.
**Supported Platforms:** windows
#### Attack Commands: Run with `command_prompt`
```command_prompt
nltest.exe /dclist:domain.local
```
```
Invoke-AtomicTest T1018 -TestNumbers 3
```
### Atomic Test #4 - Remote System Discovery - ping sweep
Identify remote systems via ping sweep.
Upon successful execution, cmd.exe will perform a for loop against the 192.168.1.1/24 network. Output will be via stdout.
**Supported Platforms:** windows
#### Attack Commands: Run with `command_prompt`
```command_prompt
for /l %i in (1,1,254) do ping -n 1 -w 100 192.168.1.%i
```
```
Invoke-AtomicTest T1018 -TestNumbers 4
```
### Atomic Test #5 - Remote System Discovery - arp
Identify remote systems via arp.
Upon successful execution, cmd.exe will execute arp to list out the arp cache. Output will be via stdout.
**Supported Platforms:** windows
#### Attack Commands: Run with `command_prompt`
```command_prompt
arp -a
```
```
Invoke-AtomicTest T1018 -TestNumbers 5
```
### Atomic Test #6 - Remote System Discovery - arp nix
Identify remote systems via arp.
Upon successful execution, sh will execute arp to list out the arp cache. Output will be via stdout.
**Supported Platforms:** linux, macos
#### Dependencies: Run with `sh`!
##### Description: Check if arp command exists on the machine
##### Check Prereq Commands:
```sh
if [ -x "$(command -v arp)" ]; then exit 0; else exit 1; fi;
```
##### Get Prereq Commands:
```sh
echo "Install arp on the machine."; exit 1;
```
```
Invoke-AtomicTest T1018 -TestNumbers 6 -GetPreReqs
```
#### Attack Commands: Run with `sh`
```sh
arp -a | grep -v '^?'
```
```
Invoke-AtomicTest T1018 -TestNumbers 6
```
### Atomic Test #7 - Remote System Discovery - sweep
Identify remote systems via ping sweep.
Upon successful execution, sh will perform a ping sweep on the 192.168.1.1/24 and echo via stdout if an IP is active.
**Supported Platforms:** linux, macos
#### Attack Commands: Run with `sh`
```sh
for ip in $(seq 1 254); do ping -c 1 192.168.1.$ip; [ $? -eq 0 ] && echo "192.168.1.$ip UP" || : ; done
```
```
Invoke-AtomicTest T1018 -TestNumbers 7
```
### Atomic Test #8 - Remote System Discovery - nslookup
Powershell script that runs nslookup on cmd.exe against the local /24 network of the first network adaptor listed in ipconfig.
Upon successful execution, powershell will identify the ip range (via ipconfig) and perform a for loop and execute nslookup against that IP range. Output will be via stdout.
**Supported Platforms:** windows
Elevation Required (e.g. root or admin)
#### Attack Commands: Run with `powershell`
```powershell
$localip = ((ipconfig | findstr [0-9].\.)[0]).Split()[-1]
$pieces = $localip.split(".")
$firstOctet = $pieces[0]
$secondOctet = $pieces[1]
$thirdOctet = $pieces[2]
foreach ($ip in 1..255 | % { "$firstOctet.$secondOctet.$thirdOctet.$_" } ) {cmd.exe /c nslookup $ip}
```
```
Invoke-AtomicTest T1018 -TestNumbers 8
```
### Atomic Test #9 - Remote System Discovery - adidnsdump
This tool enables enumeration and exporting of all DNS records in the zone for recon purposes of internal networks
Python 3 and adidnsdump must be installed, use the get_prereq_command's to meet the prerequisites for this test.
Successful execution of this test will list dns zones in the terminal.
**Supported Platforms:** windows
Elevation Required (e.g. root or admin)
#### Dependencies: Run with `powershell`!
##### Description: Computer must have python 3 installed
##### Check Prereq Commands:
```powershell
if (python --version) {exit 0} else {exit 1}
```
##### Get Prereq Commands:
```powershell
echo "Python 3 must be installed manually"
```
##### Description: Computer must have pip installed
##### Check Prereq Commands:
```powershell
if (pip3 -V) {exit 0} else {exit 1}
```
##### Get Prereq Commands:
```powershell
echo "PIP must be installed manually"
```
##### Description: adidnsdump must be installed and part of PATH
##### Check Prereq Commands:
```powershell
if (cmd /c adidnsdump -h) {exit 0} else {exit 1}
```
##### Get Prereq Commands:
```powershell
pip3 install adidnsdump
```
```
Invoke-AtomicTest T1018 -TestNumbers 9 -GetPreReqs
```
#### Attack Commands: Run with `command_prompt`
```command_prompt
adidnsdump -u domain\user -p password --print-zones 192.168.1.1
```
```
Invoke-AtomicTest T1018 -TestNumbers 9
```
## Detection
System and network discovery techniques normally occur throughout an operation as an adversary learns the environment. Data and events should not be viewed in isolation, but as part of a chain of behavior that could lead to other activities, such as Lateral Movement, based on the information obtained.
Normal, benign system and network events related to legitimate remote system discovery may be uncommon, depending on the environment and how they are used. Monitor processes and command-line arguments for actions that could be taken to gather system and network information. Remote access tools with built-in features may interact directly with the Windows API to gather information. Information may also be acquired through Windows system management tools such as [Windows Management Instrumentation](https://attack.mitre.org/techniques/T1047) and [PowerShell](https://attack.mitre.org/techniques/T1059/001).
In cloud environments, the usage of particular commands or APIs to request information about remote systems may be common. Where possible, anomalous usage of these commands and APIs or the usage of these commands and APIs in conjunction with additional unexpected commands may be a sign of malicious use. Logging methods provided by cloud providers that capture history of CLI commands executed or API usage may be utilized for detection.
## Shield Active Defense
### Software Manipulation
Make changes to a system's software properties and functions to achieve a desired effect.
Software Manipulation allows a defender to alter or replace elements of the operating system, file system, or any other software installed and executed on a system.
#### Opportunity
There is an opportunity for the defender to observe the adversary and control what they can see, what effects they can have, and/or what data they can access.
#### Use Case
A defender can change the output of a recon commands to hide simulation elements you don’t want attacked and present simulation elements you want the adversary to engage with.
#### Procedures
Hook the Win32 Sleep() function so that it always performs a Sleep(1) instead of the intended duration. This can increase the speed at which dynamic analysis can be performed when a normal malicious file sleeps for long periods before attempting additional capabilities.
Hook the Win32 NetUserChangePassword() and modify it such that the new password is different from the one provided. The data passed into the function is encrypted along with the modified new password, then logged so a defender can get alerted about the change as well as decrypt the new password for use.
Alter the output of an adversary's profiling commands to make newly-built systems look like the operating system was installed months earlier.
Alter the output of adversary recon commands to not show important assets, such as a file server containing sensitive data.
| github_jupyter |
# BERT finetuning on Yelp NYC data
BERT, or Bidirectional Encoder Representations from Transformers, is a new method of fine tuning a pre-trained language model for specific NLP tasks. It has been shown to beat the SOTA methods in almost every domain, and is highly adaptable to a wide range of tasks. In this notebook, we will use a Google Colab cloud TPU to fine-tune the Large Uncased BERT model on our Yelp NYC hotel reviews. This notebook was created on Google Colab and then ported to our Jupyter notebook collection.
----
```
!pip install bert-tensorflow
import datetime
import json
import os
import pprint
import random
import string
import sys
import tensorflow as tf
import bert
from bert import run_classifier
from bert import optimization
from bert import tokenization
from sklearn.model_selection import train_test_split
import pandas as pd
import tensorflow as tf
import tensorflow_hub as hub
from datetime import datetime
```
This section is Colab-specific and is used for setting up the GCP bucket to save the model checkpoints and weights in and authenticating the TPU.
```
# Set the output directory for saving model file
# Optionally, set a GCP bucket location
OUTPUT_DIR = 'yelp_finetuning'#@param {type:"string"}
#@markdown Whether or not to clear/delete the directory and create a new one
DO_DELETE = False #@param {type:"boolean"}
#@markdown Set USE_BUCKET and BUCKET if you want to (optionally) store model output on GCP bucket.
USE_BUCKET = True #@param {type:"boolean"}
BUCKET = 'lucas0' #@param {type:"string"}
if USE_BUCKET:
OUTPUT_DIR = 'gs://{}/{}'.format(BUCKET, OUTPUT_DIR)
from google.colab import auth
auth.authenticate_user()
if DO_DELETE:
try:
tf.gfile.DeleteRecursively(OUTPUT_DIR)
except:
# Doesn't matter if the directory didn't exist
pass
tf.gfile.MakeDirs(OUTPUT_DIR)
print('***** Model output directory: {} *****'.format(OUTPUT_DIR))
```
Because a TPU is a Google Cloud hosted runtime, we need to manually set up the data. I ran the protobuffer processing script on the YelpNYC dataset offline and then uploaded the text-ified version to our Google Cloud Storage bucket to be pulled.
```
from tensorflow import keras
import os
import re
# Download and process the dataset files.
def download_and_load_dataset(force_download=False):
dataset = tf.keras.utils.get_file(
fname="normalizedNYCYelp.txt",
origin="https://storage.googleapis.com/lucas0/imdb_classification/normalizedNYCYelp.txt",
extract=False)
dfile = open(dataset).read()
reviews = dfile.split('\n, ')
return reviews
```
Because we don't have the scripts directory (for this notebook, we will find a way to use our scripts in the future), I had to process the data manually. I took 5000 training and testing samples from the data, following the notebook from Google Research fine-tuning BERT for IMDB review sentiment analysis.
```
import numpy as np
reviews = download_and_load_dataset()
select = reviews[1:10001]
data = {}
data['review'] = []
data['deceptive'] = []
for x in select:
data['review'].append(x.split('\n')[0].split(': ')[1].replace('"', '').strip())
data['deceptive'].append(0 if 'label' in x else 1)
dataDict = pd.DataFrame.from_dict(data)
train = dataDict[:5000]
test = dataDict[5001:10001]
DATA_COLUMN = 'review'
LABEL_COLUMN = 'deceptive'
# label_list is the list of labels, i.e. True, False or 0, 1 or 'dog', 'cat'
label_list = [0, 1]
```
BERT uses a special type of tokenizer that breaks down words into their sub-grams. BERT stores multiple embeddings for one word, so that the word 'bank' in 'river bank' is seen as different to 'bank holiday'. This is why it outperforms other models thus far. In this code, BERT creates training and testing input examples by running some data through the classifier, so that the tokenizer can be built in the correct format.
```
# Use the InputExample class from BERT's run_classifier code to create examples from the data
train_InputExamples = train.apply(lambda x: bert.run_classifier.InputExample(guid=None, # Globally unique ID for bookkeeping, unused in this example
text_a = x[DATA_COLUMN],
text_b = None,
label = x[LABEL_COLUMN]), axis = 1)
test_InputExamples = test.apply(lambda x: bert.run_classifier.InputExample(guid=None,
text_a = x[DATA_COLUMN],
text_b = None,
label = x[LABEL_COLUMN]), axis = 1)
BERT_MODEL_HUB = "https://tfhub.dev/google/bert_uncased_L-12_H-768_A-12/1"
def create_tokenizer_from_hub_module():
"""Get the vocab file and casing info from the Hub module."""
with tf.Graph().as_default():
bert_module = hub.Module(BERT_MODEL_HUB)
tokenization_info = bert_module(signature="tokenization_info", as_dict=True)
with tf.Session() as sess:
vocab_file, do_lower_case = sess.run([tokenization_info["vocab_file"],
tokenization_info["do_lower_case"]])
return bert.tokenization.FullTokenizer(
vocab_file=vocab_file, do_lower_case=do_lower_case)
tokenizer = create_tokenizer_from_hub_module()
```
Now we actually process our data. We convert the examples we generated from our data above into actual features. We will limit the sequences to be 128 characters long.
```
# We'll set sequences to be at most 128 tokens long.
MAX_SEQ_LENGTH = 128
# Convert our train and test features to InputFeatures that BERT understands.
train_features = bert.run_classifier.convert_examples_to_features(train_InputExamples, label_list, MAX_SEQ_LENGTH, tokenizer)
test_features = bert.run_classifier.convert_examples_to_features(test_InputExamples, label_list, MAX_SEQ_LENGTH, tokenizer)
```
Now we define our BERT model creation function! This simply creates and returns the network, and stores the loss in a publicly accessible Tensorflow variable.
```
def create_model(is_predicting, input_ids, input_mask, segment_ids, labels,
num_labels):
"""Creates a classification model."""
bert_module = hub.Module(
BERT_MODEL_HUB,
trainable=True)
bert_inputs = dict(
input_ids=input_ids,
input_mask=input_mask,
segment_ids=segment_ids)
bert_outputs = bert_module(
inputs=bert_inputs,
signature="tokens",
as_dict=True)
# Use "pooled_output" for classification tasks on an entire sentence.
# Use "sequence_outputs" for token-level output.
output_layer = bert_outputs["pooled_output"]
hidden_size = output_layer.shape[-1].value
output_weights = tf.get_variable(
"output_weights", [num_labels, hidden_size],
initializer=tf.truncated_normal_initializer(stddev=0.02))
output_bias = tf.get_variable(
"output_bias", [num_labels], initializer=tf.zeros_initializer())
with tf.variable_scope("loss"):
# Dropout helps prevent overfitting
output_layer = tf.nn.dropout(output_layer, keep_prob=0.9)
logits = tf.matmul(output_layer, output_weights, transpose_b=True)
logits = tf.nn.bias_add(logits, output_bias)
log_probs = tf.nn.log_softmax(logits, axis=-1)
# Convert labels into one-hot encoding
one_hot_labels = tf.one_hot(labels, depth=num_labels, dtype=tf.float32)
predicted_labels = tf.squeeze(tf.argmax(log_probs, axis=-1, output_type=tf.int32))
# If we're predicting, we want predicted labels and the probabiltiies.
if is_predicting:
return (predicted_labels, log_probs)
# If we're train/eval, compute loss between predicted and actual label
per_example_loss = -tf.reduce_sum(one_hot_labels * log_probs, axis=-1)
loss = tf.reduce_mean(per_example_loss)
return (loss, predicted_labels, log_probs)
```
This function passses the parameters and evaluates our model using the powerful tf.Estimator. It creates an optimizer based on the learning rate and steps, and creates the model with our labels, feature shapes, etc, and returns a function to be used by tf.Estimator with everything it needs inside.
```
# model_fn_builder actually creates our model function
# using the passed parameters for num_labels, learning_rate, etc.
def model_fn_builder(num_labels, learning_rate, num_train_steps,
num_warmup_steps):
"""Returns `model_fn` closure for TPUEstimator."""
def model_fn(features, labels, mode, params):
"""The `model_fn` for TPUEstimator."""
input_ids = features["input_ids"]
input_mask = features["input_mask"]
segment_ids = features["segment_ids"]
label_ids = features["label_ids"]
is_predicting = (mode == tf.estimator.ModeKeys.PREDICT)
# TRAIN and EVAL
if not is_predicting:
(loss, predicted_labels, log_probs) = create_model(
is_predicting, input_ids, input_mask, segment_ids, label_ids, num_labels)
train_op = bert.optimization.create_optimizer(
loss, learning_rate, num_train_steps, num_warmup_steps, use_tpu=False)
# Calculate evaluation metrics.
def metric_fn(label_ids, predicted_labels):
accuracy = tf.metrics.accuracy(label_ids, predicted_labels)
f1_score = tf.contrib.metrics.f1_score(
label_ids,
predicted_labels)
auc = tf.metrics.auc(
label_ids,
predicted_labels)
recall = tf.metrics.recall(
label_ids,
predicted_labels)
precision = tf.metrics.precision(
label_ids,
predicted_labels)
true_pos = tf.metrics.true_positives(
label_ids,
predicted_labels)
true_neg = tf.metrics.true_negatives(
label_ids,
predicted_labels)
false_pos = tf.metrics.false_positives(
label_ids,
predicted_labels)
false_neg = tf.metrics.false_negatives(
label_ids,
predicted_labels)
return {
"eval_accuracy": accuracy,
"f1_score": f1_score,
"auc": auc,
"precision": precision,
"recall": recall,
"true_positives": true_pos,
"true_negatives": true_neg,
"false_positives": false_pos,
"false_negatives": false_neg
}
eval_metrics = metric_fn(label_ids, predicted_labels)
if mode == tf.estimator.ModeKeys.TRAIN:
return tf.estimator.EstimatorSpec(mode=mode,
loss=loss,
train_op=train_op)
else:
return tf.estimator.EstimatorSpec(mode=mode,
loss=loss,
eval_metric_ops=eval_metrics)
else:
(predicted_labels, log_probs) = create_model(
is_predicting, input_ids, input_mask, segment_ids, label_ids, num_labels)
predictions = {
'probabilities': log_probs,
'labels': predicted_labels
}
return tf.estimator.EstimatorSpec(mode, predictions=predictions)
# Return the actual model function in the closure
return model_fn
```
Here we programatically calculate how many training and warmup steps to do based on what our batch size and number of training epochs are. We also set up how often we take model checkpoints, in case our instance goes down and we lose all our progress.
```
BATCH_SIZE = 32
LEARNING_RATE = 2e-5
NUM_TRAIN_EPOCHS = 3.0
# Warmup is a period of time where the learning rate
# is small and gradually increases--usually helps training.
WARMUP_PROPORTION = 0.1
# Model configs
SAVE_CHECKPOINTS_STEPS = 500
SAVE_SUMMARY_STEPS = 100
# Compute # train and warmup steps from batch size
num_train_steps = int(len(train_features) / BATCH_SIZE * NUM_TRAIN_EPOCHS)
num_warmup_steps = int(num_train_steps * WARMUP_PROPORTION)
print(num_train_steps, num_warmup_steps)
# Specify output directory and number of checkpoint steps to save
run_config = tf.estimator.RunConfig(
model_dir=OUTPUT_DIR,
save_summary_steps=SAVE_SUMMARY_STEPS,
save_checkpoints_steps=SAVE_CHECKPOINTS_STEPS)
```
Here we simply pass the steps calculated above to our model builder and then pass our model creator function to our estimator. Our estimator works as a sort of runtime manager, like keras.fit, where we specify batch size. It also sets up output directories for model checkpoints. We can see all the configs it sets up below in the output.
```
model_fn = model_fn_builder(
num_labels=len(label_list),
learning_rate=LEARNING_RATE,
num_train_steps=num_train_steps,
num_warmup_steps=num_warmup_steps)
estimator = tf.estimator.Estimator(
model_fn=model_fn,
config=run_config,
params={"batch_size": BATCH_SIZE})
```
This function simply returns a function that provides our input to the model.
```
# Create an input function for training. drop_remainder = True for using TPUs.
train_input_fn = bert.run_classifier.input_fn_builder(
features=train_features,
seq_length=MAX_SEQ_LENGTH,
is_training=True,
drop_remainder=True)
```
Now we begin training! Due to this being the large BERT model, it will take some time. However, as we're utilizing a Google Cloud TPU, this isn't a problem for us anymore.
```
print('Beginning Training!')
current_time = datetime.now()
estimator.train(input_fn=train_input_fn, max_steps=num_train_steps)
print("Training took time ", datetime.now() - current_time)
```
And there we have it. After training for a little over 6 hours, we managed to achieve a loss of 0.5. Now, in order to evaluate our model, we need to create a test input function in the same fashion and evaluate it in non-training mode (prediction mode).
```
test_input_fn = bert.run_classifier.input_fn_builder(
features=test_features,
seq_length=MAX_SEQ_LENGTH,
is_training=False,
drop_remainder=False)
estimator.evaluate(input_fn=test_input_fn, steps=None)
```
Our accuracy on this run seems to be about .65, which is surprisingly good to me considering there was very little tweaking of the model done to be more fitted to our task rather than IMDB sentiment classification. It did get more true negatives than any other class, which is great for us, as a true negative corresponds to a correct deceptive classification.
```
def getPrediction(in_sentences):
labels = ["Deceptive", "Genuine"]
input_examples = [run_classifier.InputExample(guid="", text_a = x, text_b = None, label = 0) for x in in_sentences] # here, "" is just a dummy label
input_features = run_classifier.convert_examples_to_features(input_examples, label_list, MAX_SEQ_LENGTH, tokenizer)
predict_input_fn = run_classifier.input_fn_builder(features=input_features, seq_length=MAX_SEQ_LENGTH, is_training=False, drop_remainder=False)
predictions = estimator.predict(predict_input_fn)
return [(sentence, prediction['probabilities'], labels[prediction['labels']]) for sentence, prediction in zip(in_sentences, predictions)]
```
Let's actually see its predictive power as a classifier. I wrote four 'deceptive' reviews, 2 negative and two positive.
```
pred_sentences = [
"This hotel was truly terrible. Will never be returning.",
"Rude staff, filthy room, crappy service. Won't be coming here again!",
"I loved everything about my stay. The Hilton was a fantastic hotel and I will be returning.",
"Absolutely fantastic! Great food, service and amenities. Looked after for the full duration."
]
predictions = getPrediction(pred_sentences)
predictions
```
This seems to align pretty much with our results, getting it 75% correct! Overall I think trained BERT representations used in conjunction with a finely tuned RNN architecture (with perhaps some convolution to enable cross-domain accuracy) could be the key to our success.
| github_jupyter |
## Facial Filters
Using your trained facial keypoint detector, you can now do things like add filters to a person's face, automatically. In this optional notebook, you can play around with adding sunglasses to detected face's in an image by using the keypoints detected around a person's eyes. Checkout the `images/` directory to see what pther .png's have been provided for you to try, too!
<img src="images/face_filter_ex.png" width=60% height=60%/>
Let's start this process by looking at a sunglasses .png that we'll be working with!
```
# import necessary resources
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import os
import cv2
# load in sunglasses image with cv2 and IMREAD_UNCHANGED
sunglasses = cv2.imread('images/sunglasses.png', cv2.IMREAD_UNCHANGED)
# plot our image
plt.imshow(sunglasses)
# print out its dimensions
print('Image shape: ', sunglasses.shape)
```
## The 4th dimension
You'll note that this image actually has *4 color channels*, not just 3 as your avg RGB image does. This is due to the flag we set `cv2.IMREAD_UNCHANGED`, which tells this to read in another color channel.
#### Alpha channel
It has the usual red, blue, and green channels any color image has, and the 4th channel respresents the **transparency level of each pixel** in the image; this is often called the **alpha** channel. Here's how the transparency channel works: the lower the value, the more transparent, or see-through, the pixel will become. The lower bound (completely transparent) is zero here, so any pixels set to 0 will not be seen; these look like white background pixels in the image above, but they are actually totally transparent.
This transparent channel allows us to place this rectangular image of sunglasses on an image of a face and still see the face area that is techically covered by the transparentbackground of the sunglasses image!
Let's check out the alpha channel of our sunglasses image in the next Python cell. Because many of the pixels in the background of the image have an alpha value of 0, we'll need to explicitly print out non-zero values if we want to see them.
```
# print out the sunglasses transparency (alpha) channel
alpha_channel = sunglasses[:,:,3]
print ('The alpha channel looks like this (black pixels = transparent): ')
plt.imshow(alpha_channel, cmap='gray')
# just to double check that there are indeed non-zero values
# let's find and print out every value greater than zero
values = np.where(alpha_channel != 0)
print ('The non-zero values of the alpha channel are: ')
print (values)
```
#### Overlaying images
This means that when we place this sunglasses image on top of another image, we can use the transparency channel as a filter:
* If the pixels are non-transparent (alpha_channel > 0), overlay them on the new image
#### Keypoint locations
In doing this, it's helpful to understand which keypoint belongs to the eyes, mouth, etc., so in the image below we also print the index of each facial keypoint directly on the image so you can tell which keypoints are for the eyes, eyebrows, etc.,
<img src="images/landmarks_numbered.jpg" width=50% height=50%/>
It may be useful to use keypoints that correspond to the edges of the face to define the width of the sunglasses, and the locations of the eyes to define the placement.
Next, we'll load in an example image. Below, you've been given an image and set of keypoints from the provided training set of data, but you can use your own CNN model to generate keypoints for *any* image of a face (as in Notebook 3) and go through the same overlay process!
```
# load in training data
key_pts_frame = pd.read_csv('data/training_frames_keypoints.csv')
# print out some stats about the data
print('Number of images: ', key_pts_frame.shape[0])
# helper function to display keypoints
def show_keypoints(image, key_pts):
"""Show image with keypoints"""
plt.imshow(image)
plt.scatter(key_pts[:, 0], key_pts[:, 1], s=20, marker='.', c='m')
# a selected image
n = 120
image_name = key_pts_frame.iloc[n, 0]
image = mpimg.imread(os.path.join('data/training/', image_name))
key_pts = key_pts_frame.iloc[n, 1:].as_matrix()
key_pts = key_pts.astype('float').reshape(-1, 2)
print('Image name: ', image_name)
plt.figure(figsize=(5, 5))
show_keypoints(image, key_pts)
plt.show()
```
Next, you'll see an example of placing sunglasses on the person in the loaded image.
Note that the keypoints are numbered off-by-one in the numbered image above, and so `key_pts[0,:]` corresponds to the first point (1) in the labelled image.
```
# Display sunglasses on top of the image in the appropriate place
# copy of the face image for overlay
image_copy = np.copy(image)
# top-left location for sunglasses to go
# 17 = edge of left eyebrow
x = int(key_pts[17, 0])
y = int(key_pts[17, 1])
# height and width of sunglasses
# h = length of nose
h = int(abs(key_pts[27,1] - key_pts[34,1]))
# w = left to right eyebrow edges
w = int(abs(key_pts[17,0] - key_pts[26,0]))
# read in sunglasses
sunglasses = cv2.imread('images/sunglasses.png', cv2.IMREAD_UNCHANGED)
# resize sunglasses
new_sunglasses = cv2.resize(sunglasses, (w, h), interpolation = cv2.INTER_CUBIC)
# get region of interest on the face to change
roi_color = image_copy[y:y+h,x:x+w]
# find all non-transparent pts
ind = np.argwhere(new_sunglasses[:,:,3] > 0)
# for each non-transparent point, replace the original image pixel with that of the new_sunglasses
for i in range(3):
roi_color[ind[:,0],ind[:,1],i] = new_sunglasses[ind[:,0],ind[:,1],i]
# set the area of the image to the changed region with sunglasses
image_copy[y:y+h,x:x+w] = roi_color
# display the result!
plt.imshow(image_copy)
```
#### Further steps
Look in the `images/` directory to see other available .png's for overlay! Also, you may notice that the overlay of the sunglasses is not entirely perfect; you're encouraged to play around with the scale of the width and height of the glasses and investigate how to perform [image rotation](https://docs.opencv.org/3.0-beta/doc/py_tutorials/py_imgproc/py_geometric_transformations/py_geometric_transformations.html) in OpenCV so as to match an overlay with any facial pose.
| github_jupyter |
## 1. Tweet classification: Trump vs. Trudeau
<p>So you think you can classify text? How about tweets? In this notebook, we'll take a dive into the world of social media text classification by investigating how to properly classify tweets from two prominent North American politicians: Donald Trump and Justin Trudeau.</p>
<p><img src="https://upload.wikimedia.org/wikipedia/commons/thumb/4/47/President_Donald_Trump_and_Prime_Minister_Justin_Trudeau_Joint_Press_Conference%2C_February_13%2C_2017.jpg/800px-President_Donald_Trump_and_Prime_Minister_Justin_Trudeau_Joint_Press_Conference%2C_February_13%2C_2017.jpg" alt="Donald Trump and Justin Trudeau shaking hands." height="50%" width="50%"></p>
<p><a href="https://commons.wikimedia.org/wiki/File:President_Donald_Trump_and_Prime_Minister_Justin_Trudeau_Joint_Press_Conference,_February_13,_2017.jpg">Photo Credit: Executive Office of the President of the United States</a></p>
<p>Tweets pose specific problems to NLP, including the fact they are shorter texts. There are also plenty of platform-specific conventions to give you hassles: mentions, #hashtags, emoji, links and short-hand phrases (ikr?). Can we overcome those challenges and build a useful classifier for these two tweeters? Yes! Let's get started.</p>
<p>To begin, we will import all the tools we need from scikit-learn. We will need to properly vectorize our data (<code>CountVectorizer</code> and <code>TfidfVectorizer</code>). And we will also want to import some models, including <code>MultinomialNB</code> from the <code>naive_bayes</code> module, <code>LinearSVC</code> from the <code>svm</code> module and <code>PassiveAggressiveClassifier</code> from the <code>linear_model</code> module. Finally, we'll need <code>sklearn.metrics</code> and <code>train_test_split</code> and <code>GridSearchCV</code> from the <code>model_selection</code> module to evaluate and optimize our model.</p>
```
# Set seed for reproducibility
import random; random.seed(53)
# Import all we need from sklearn
# ... YOUR CODE FOR TASK 1 ...
# ... YOUR CODE FOR TASK 1 ...
# ... YOUR CODE FOR TASK 1 ...
# ... YOUR CODE FOR TASK 1 ...
# ... YOUR CODE FOR TASK 1 ...
```
## 2. Transforming our collected data
<p>To begin, let's start with a corpus of tweets which were collected in November 2017. They are available in CSV format. We'll use a Pandas DataFrame to help import the data and pass it to scikit-learn for further processing.</p>
<p>Since the data has been collected via the Twitter API and not split into test and training sets, we'll need to do this. Let's use <code>train_test_split()</code> with <code>random_state=53</code> and a test size of 0.33, just as we did in the DataCamp course. This will ensure we have enough test data and we'll get the same results no matter where or when we run this code.</p>
```
import pandas as pd
# Load data
tweet_df = ...
# Create target
y = ...
# Split training and testing data
X_train, X_test, y_train, y_test = ...
```
## 3. Vectorize the tweets
<p>We have the training and testing data all set up, but we need to create vectorized representations of the tweets in order to apply machine learning.</p>
<p>To do so, we will utilize the <code>CountVectorizer</code> and <code>TfidfVectorizer</code> classes which we will first need to fit to the data.</p>
<p>Once this is complete, we can start modeling with the new vectorized tweets!</p>
```
# Initialize count vectorizer
count_vectorizer = ...
# Create count train and test variables
count_train = ...
count_test = ...
# Initialize tfidf vectorizer
tfidf_vectorizer = ...
# Create tfidf train and test variables
tfidf_train = ...
tfidf_test = ...
```
## 4. Training a multinomial naive Bayes model
<p>Now that we have the data in vectorized form, we can train the first model. Investigate using the Multinomial Naive Bayes model with both the <code>CountVectorizer</code> and <code>TfidfVectorizer</code> data. Which do will perform better? How come?</p>
<p>To assess the accuracies, we will print the test sets accuracy scores for both models.</p>
```
# Create a MulitnomialNB model
tfidf_nb = ...
# ... Train your model here ...
# Run predict on your TF-IDF test data to get your predictions
tfidf_nb_pred = ...
# Calculate the accuracy of your predictions
tfidf_nb_score = ...
# Create a MulitnomialNB model
count_nb = ...
# ... Train your model here ...
# Run predict on your count test data to get your predictions
count_nb_pred = ...
# Calculate the accuracy of your predictions
count_nb_score = ...
print('NaiveBayes Tfidf Score: ', tfidf_nb_score)
print('NaiveBayes Count Score: ', count_nb_score)
```
## 5. Evaluating our model using a confusion matrix
<p>We see that the TF-IDF model performs better than the count-based approach. Based on what we know from the NLP fundamentals course, why might that be? We know that TF-IDF allows unique tokens to have a greater weight - perhaps tweeters are using specific important words that identify them! Let's continue the investigation.</p>
<p>For classification tasks, an accuracy score doesn't tell the whole picture. A better evaluation can be made if we look at the confusion matrix, which shows the number correct and incorrect classifications based on each class. We can use the metrics, True Positives, False Positives, False Negatives, and True Negatives, to determine how well the model performed on a given class. How many times was Trump misclassified as Trudeau?</p>
```
%matplotlib inline
from datasets.helper_functions import plot_confusion_matrix
# Calculate the confusion matrices for the tfidf_nb model and count_nb models
tfidf_nb_cm = ...
count_nb_cm = ...
# Plot the tfidf_nb_cm confusion matrix
plot_confusion_matrix(tfidf_nb_cm, classes=..., title="TF-IDF NB Confusion Matrix")
# Plot the count_nb_cm confusion matrix without overwriting the first plot
plot_confusion_matrix(..., classes=..., title=..., figure=1)
```
## 6. Trying out another classifier: Linear SVC
<p>So the Bayesian model only has one prediction difference between the TF-IDF and count vectorizers -- fairly impressive! Interestingly, there is some confusion when the predicted label is Trump but the actual tweeter is Trudeau. If we were going to use this model, we would want to investigate what tokens are causing the confusion in order to improve the model. </p>
<p>Now that we've seen what the Bayesian model can do, how about trying a different approach? <a href="https://scikit-learn.org/stable/modules/generated/sklearn.svm.LinearSVC.html">LinearSVC</a> is another popular choice for text classification. Let's see if using it with the TF-IDF vectors improves the accuracy of the classifier!</p>
```
# Create a LinearSVM model
tfidf_svc = ...
# ... Train your model here ...
# Run predict on your tfidf test data to get your predictions
tfidf_svc_pred = ...
# Calculate your accuracy using the metrics module
tfidf_svc_score = ...
print("LinearSVC Score: %0.3f" % tfidf_svc_score)
# Calculate the confusion matrices for the tfidf_svc model
svc_cm = ...
# Plot the confusion matrix using the plot_confusion_matrix function
plot_confusion_matrix(svc_cm, classes=..., title="TF-IDF LinearSVC Confusion Matrix")
```
## 7. Introspecting our top model
<p>Wow, the LinearSVC model is even better than the Multinomial Bayesian one. Nice work! Via the confusion matrix we can see that, although there is still some confusion where Trudeau's tweets are classified as Trump's, the False Positive rate is better than the previous model. So, we have a performant model, right? </p>
<p>We might be able to continue tweaking and improving all of the previous models by learning more about parameter optimization or applying some better preprocessing of the tweets. </p>
<p>Now let's see what the model has learned. Using the LinearSVC Classifier with two classes (Trump and Trudeau) we can sort the features (tokens), by their weight and see the most important tokens for both Trump and Trudeau. What are the most Trump-like or Trudeau-like words? Did the model learn something useful to distinguish between these two men? </p>
```
from datasets.helper_functions import plot_and_return_top_features
# Import pprint from pprint
from pprint ...
# Get the top features using the plot_and_return_top_features function and your top model and tfidf vectorizer
top_features = ...
# pprint the top features
pprint(...)
```
## 8. Bonus: can you write a Trump or Trudeau tweet?
<p>So, what did our model learn? It seems like it learned that Trudeau tweets in French!</p>
<p>I challenge you to write your own tweet using the knowledge gained to trick the model! Use the printed list or plot above to make some inferences about what words will classify your text as Trump or Trudeau. Can you fool the model into thinking you are Trump or Trudeau?</p>
<p>If you can write French, feel free to make your Trudeau-impersonation tweet in French! As you may have noticed, these French words are common words, or, "stop words". You could remove both English and French stop words from the tweets as a preprocessing step, but that might decrease the accuracy of the model because Trudeau is the only French-speaker in the group. If you had a dataset with more than one French speaker, this would be a useful preprocessing step.</p>
<p>Future work on this dataset could involve:</p>
<ul>
<li>Add extra preprocessing (such as removing URLs or French stop words) and see the effects</li>
<li>Use GridSearchCV to improve both your Bayesian and LinearSVC models by finding the optimal parameters</li>
<li>Introspect your Bayesian model to determine what words are more Trump- or Trudeau- like</li>
<li>Add more recent tweets to your dataset using tweepy and retrain</li>
</ul>
<p>Good luck writing your impersonation tweets -- feel free to share them on Twitter!</p>
```
# Write two tweets as strings, one which you want to classify as Trump and one as Trudeau
trump_tweet = ...
trudeau_tweet = ...
# Vectorize each tweet using the TF-IDF vectorizer's transform method
# Note: `transform` needs the string in a list object (i.e. [trump_tweet])
trump_tweet_vectorized = ...
trudeau_tweet_vectorized = ...
# Call the predict method on your vectorized tweets
trump_tweet_pred = ...
trudeau_tweet_pred = ...
print("Predicted Trump tweet", trump_tweet_pred)
print("Predicted Trudeau tweet", trudeau_tweet_pred)
```
| github_jupyter |
```
# Required modules
import os
import sys
sys.path.append('../OceanVIM/notebooks/')
import numpy as np
import matplotlib.pyplot as plt
import xarray as xr
import pygmt
import VIM_tools as vt
import pyshtools
from scipy.interpolate import RectSphereBivariateSpline
%matplotlib inline
%load_ext autoreload
%autoreload 2
HM = np.loadtxt('grids/W19_l90.xyz')
lons = HM[:,0].reshape(720,1440)
lats = HM[:,1].reshape(720,1440)
sus = HM[:,2].reshape(720,1440)
sus = np.hstack((sus[:,720:],sus[:,:720]))
lons = np.hstack((lons[:,720:],lons[:,:720]+360))
#sus_test = np.zeros_like(sus)
#sus_test[350:480,100:140] = 5.
#sus_test = np.random.random(sus.shape)*10.
sus_test = np.copy(sus)*0.0001
plt.pcolormesh(sus_test)
plt.colorbar()
(HM_IVIM_Mr,
HM_IVIM_Mtheta,
HM_IVIM_Mphi) = vt.vis2magnetisation(np.flipud(sus_test))
vt.write_vh0_input(lons[0,:].flatten(),90-lats[:,0].flatten(),HM_IVIM_Mr,HM_IVIM_Mtheta,HM_IVIM_Mphi,filename='total_py.in')
os.system('vh0 < total_py.in')
# The skip is to remove lines for harmonic degrees 14 and lower
coeffs, lmaxout = pyshtools.shio.shread('./glm.out')#,skip=104)
r0 = 3389500.
alt = 150000.
result = pyshtools.SHMagCoeffs.from_array(coeffs, r0=r0)
resultg = result.expand(a=r0+alt, sampling=2, extend=True)
plt.figure(figsize=(20,10))
plt.pcolormesh(-resultg.rad.to_array(),vmin=-100,vmax=100,cmap=plt.cm.seismic)
# Load coefficients for Mars magnetic model of Langlais++ 2018
clm_lang, lmax = pyshtools.shio.shread('./shc/Langlais++_Mars.cof')
clmm = pyshtools.SHMagCoeffs.from_array(clm_lang, r0=3389500.)
clmm.plot_spectrum()
result.plot_spectrum()
HellasDistance = xr.open_dataarray('DistanceToHellasCentre.nc')
plt.contourf(HellasDistance.data)
plt.colorbar()
plt.show()
print(HellasDistance.data.max())
print(3390*np.pi)
inner_limit = 500.
outer_limit = 3000.
ramping = HellasDistance.data.copy()
ramping[ramping<inner_limit] = inner_limit
ramping[ramping>outer_limit] = outer_limit
ramping = (ramping-inner_limit)/(outer_limit-inner_limit)
plt.contourf(ramping)
plt.colorbar()
plt.show()
plt.figure(figsize=(16,4))
plt.subplot(121)
plt.contourf(sus_test)
plt.subplot(122)
plt.contourf(sus_test*ramping)
print(np.min(ramping))
print(np.max(ramping))
(HM_IVIM_Mr,
HM_IVIM_Mtheta,
HM_IVIM_Mphi) = vt.vis2magnetisation(np.flipud(sus_test*ramping))
vt.write_vh0_input(lons[0,:].flatten(),90-lats[:,0].flatten(),HM_IVIM_Mr,HM_IVIM_Mtheta,HM_IVIM_Mphi,filename='total_py.in')
os.system('vh0 < total_py.in')
# The skip is to remove lines for harmonic degrees 14 and lower
coeffs, lmaxout = pyshtools.shio.shread('./glm.out')#,skip=104)
#coeffs, lmaxout = pyshtools.shio.shread('./glm.out',skip=104)
r0 = 3389500.
alt = 10000.
result = pyshtools.SHMagCoeffs.from_array(coeffs, r0=r0)
resultg = result.expand(a=r0+alt, sampling=2, extend=True)
plt.figure(figsize=(20,10))
plt.pcolormesh(-resultg.rad.to_array(),vmin=-200,vmax=200,cmap=plt.cm.seismic)
```
| github_jupyter |
```
BRANCH = 'main'
"""
You can run either this notebook locally (if you have all the dependencies and a GPU) or on Google Colab.
Instructions for setting up Colab are as follows:
1. Open a new Python 3 notebook.
2. Import this notebook from GitHub (File -> Upload Notebook -> "GITHUB" tab -> copy/paste GitHub URL)
3. Connect to an instance with a GPU (Runtime -> Change runtime type -> select "GPU" for hardware accelerator)
4. Run this cell to set up dependencies.
"""
# If you're using Google Colab and not running locally, run this cell
# install NeMo
!python -m pip install git+https://github.com/NVIDIA/NeMo.git@$BRANCH#egg=nemo_toolkit[nlp]
# If you're not using Colab, you might need to upgrade jupyter notebook to avoid the following error:
# 'ImportError: IProgress not found. Please update jupyter and ipywidgets.'
! pip install ipywidgets
! jupyter nbextension enable --py widgetsnbextension
# Please restart the kernel after running this cell
from nemo.collections import nlp as nemo_nlp
from nemo.utils.exp_manager import exp_manager
import os
import wget
import torch
import pytorch_lightning as pl
from omegaconf import OmegaConf
```
In this tutorial, we are going to describe how to finetune BioMegatron - a [BERT](https://arxiv.org/abs/1810.04805)-like [Megatron-LM](https://arxiv.org/pdf/1909.08053.pdf) model pre-trained on large biomedical text corpus ([PubMed](https://pubmed.ncbi.nlm.nih.gov/) abstracts and full-text commercial use collection) - on the [NCBI Disease Dataset](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3951655/) for Named Entity Recognition.
The model size of Megatron-LM can be larger than BERT, up to multi-billion parameters, compared to 345 million parameters of BERT-large.
There are some alternatives of BioMegatron, most notably [BioBERT](https://arxiv.org/abs/1901.08746). Compared to BioBERT BioMegatron is larger by model size and pre-trained on larger text corpus.
A more general tutorial of using BERT-based models, including Megatron-LM, for downstream natural language processing tasks can be found [here](https://github.com/NVIDIA/NeMo/blob/main/tutorials/nlp/01_Pretrained_Language_Models_for_Downstream_Tasks.ipynb).
# Task Description
**Named entity recognition (NER)**, also referred to as entity chunking, identification or extraction, is the task of detecting and classifying key information (entities) in text.
For instance, **given sentences from medical abstracts, what diseases are mentioned?**<br>
In this case, our data input is sentences from the abstracts, and our labels are the precise locations of the named disease entities. Take a look at the information provided for the dataset.
For more details and general examples on Named Entity Recognition, please refer to the [Token Classification and Named Entity Recognition tutorial notebook](https://github.com/NVIDIA/NeMo/blob/main/tutorials/nlp/Token_Classification_Named_Entity_Recognition.ipynb).
# Dataset
The [NCBI-disease corpus](https://www.ncbi.nlm.nih.gov/CBBresearch/Dogan/DISEASE/) is a set of 793 PubMed abstracts, annotated by 14 annotators. The annotations take the form of HTML-style tags inserted into the abstract text using the clearly defined rules. The annotations identify named diseases, and can be used to fine-tune a language model to identify disease mentions in future abstracts, *whether those diseases were part of the original training set or not*.
Here's an example of what an annotated abstract from the corpus looks like:
```html
10021369 Identification of APC2, a homologue of the <category="Modifier">adenomatous polyposis coli tumour</category> suppressor . The <category="Modifier">adenomatous polyposis coli ( APC ) tumour</category>-suppressor protein controls the Wnt signalling pathway by forming a complex with glycogen synthase kinase 3beta ( GSK-3beta ) , axin / conductin and betacatenin . Complex formation induces the rapid degradation of betacatenin . In <category="Modifier">colon carcinoma</category> cells , loss of APC leads to the accumulation of betacatenin in the nucleus , where it binds to and activates the Tcf-4 transcription factor ( reviewed in [ 1 ] [ 2 ] ) . Here , we report the identification and genomic structure of APC homologues . Mammalian APC2 , which closely resembles APC in overall domain structure , was functionally analyzed and shown to contain two SAMP domains , both of which are required for binding to conductin . Like APC , APC2 regulates the formation of active betacatenin-Tcf complexes , as demonstrated using transient transcriptional activation assays in APC - / - <category="Modifier">colon carcinoma</category> cells . Human APC2 maps to chromosome 19p13 . 3 . APC and APC2 may therefore have comparable functions in development and <category="SpecificDisease">cancer</category> .
```
In this example, we see the following tags within the abstract:
```html
<category="Modifier">adenomatous polyposis coli tumour</category>
<category="Modifier">adenomatous polyposis coli ( APC ) tumour</category>
<category="Modifier">colon carcinoma</category>
<category="Modifier">colon carcinoma</category>
<category="SpecificDisease">cancer</category>
```
For our purposes, we will consider any identified category (such as "Modifier", "Specific Disease", and a few others) to generally be a "disease".
Let's download the dataset.
```
DATA_DIR = "DATA_DIR"
os.makedirs(DATA_DIR, exist_ok=True)
os.makedirs(os.path.join(DATA_DIR, 'NER'), exist_ok=True)
print('Downloading NCBI data...')
wget.download('https://www.ncbi.nlm.nih.gov/CBBresearch/Dogan/DISEASE/NCBI_corpus.zip', DATA_DIR)
! unzip -o {DATA_DIR}/NCBI_corpus.zip -d {DATA_DIR}
```
If you want to see more examples, you can explore the text of the corpus using the file browser to the left, or open files directly, for example typing a command like the following in a code-cell:
<pre><code>
! head -1 $DATA_DIR/NCBI_corpus_testing.txt
</code></pre>
We have two datasets derived from this corpus: a text classification dataset and a named entity recognition (NER) dataset. The text classification dataset labels the abstracts among three broad disease groupings. We'll use this simple split to demonstrate the NLP text classification task. The NER dataset labels individual words as diseases. This dataset will be used for the NLP NER task.
## Pre-process dataset
A pre-processed NCBI-disease dataset for NER can be found [here](https://github.com/spyysalo/ncbi-disease/tree/master/conll) or [here](https://github.com/dmis-lab/biobert#datasets).<br>
We download the files under {DATA_DIR/NER} directory.
```
wget.download('https://raw.githubusercontent.com/spyysalo/ncbi-disease/master/conll/train.tsv', os.path.join(DATA_DIR, 'NER'))
wget.download('https://raw.githubusercontent.com/spyysalo/ncbi-disease/master/conll/devel.tsv', os.path.join(DATA_DIR, 'NER'))
wget.download('https://raw.githubusercontent.com/spyysalo/ncbi-disease/master/conll/test.tsv', os.path.join(DATA_DIR, 'NER'))
NER_DATA_DIR = 'DATA_DIR/NER'
!ls -lh $NER_DATA_DIR
```
Convert these to a format that is compatible with [NeMo Token Classification module](https://github.com/NVIDIA/NeMo/blob/main/examples/nlp/token_classification/token_classification_train.py), using the [conversion script](https://github.com/NVIDIA/NeMo/blob/main/examples/nlp/token_classification/data/import_from_iob_format.py).
```
! mv DATA_DIR/NER/devel.tsv DATA_DIR/NER/dev.tsv
wget.download(f'https://raw.githubusercontent.com/NVIDIA/NeMo/{BRANCH}/examples/nlp/token_classification/data/import_from_iob_format.py')
! python import_from_iob_format.py --data_file=DATA_DIR/NER/train.tsv
! python import_from_iob_format.py --data_file=DATA_DIR/NER/dev.tsv
! python import_from_iob_format.py --data_file=DATA_DIR/NER/test.tsv
```
The NER task requires two files: the text sentences, and the labels. Run the next two cells to see a sample of the two files.
```
!head $NER_DATA_DIR/text_train.txt
!head $NER_DATA_DIR/labels_train.txt
```
### IOB Tagging
We can see that the abstract has been broken into sentences. Each sentence is then further parsed into words with labels that correspond to the original HTML-style tags in the corpus.
The sentences and labels in the NER dataset map to each other with _inside, outside, beginning (IOB)_ tagging. Anything separated by white space is a word, including punctuation. For the first sentence we have the following mapping:
```text
Identification of APC2 , a homologue of the adenomatous polyposis coli tumour suppressor .
O O O O O O O O B I I I O O
```
Recall the original corpus tags:
```html
Identification of APC2, a homologue of the <category="Modifier">adenomatous polyposis coli tumour</category> suppressor .
```
The beginning word of the tagged text, "adenomatous", is now IOB-tagged with a <span style="font-family:verdana;font-size:110%;">B</span> (beginning) tag, the other parts of the disease, "polyposis coli tumour" tagged with <span style="font-family:verdana;font-size:110%;">I</span> (inside) tags, and everything else tagged as <span style="font-family:verdana;font-size:110%;">O</span> (outside).
# Model configuration
Our Named Entity Recognition model is comprised of the pretrained [BERT](https://arxiv.org/pdf/1810.04805.pdf) model followed by a Token Classification layer.
The model is defined in a config file which declares multiple important sections. They are:
- **model**: All arguments that are related to the Model - language model, token classifier, optimizer and schedulers, datasets and any other related information
- **trainer**: Any argument to be passed to PyTorch Lightning
```
WORK_DIR = "WORK_DIR"
os.makedirs(WORK_DIR, exist_ok=True)
MODEL_CONFIG = "token_classification_config.yaml"
# download the model's configuration file
config_dir = WORK_DIR + '/configs/'
os.makedirs(config_dir, exist_ok=True)
if not os.path.exists(config_dir + MODEL_CONFIG):
print('Downloading config file...')
wget.download(f'https://raw.githubusercontent.com/NVIDIA/NeMo/{BRANCH}/examples/nlp/token_classification/conf/' + MODEL_CONFIG, config_dir)
else:
print ('config file is already exists')
# this line will print the entire config of the model
config_path = f'{WORK_DIR}/configs/{MODEL_CONFIG}'
print(config_path)
config = OmegaConf.load(config_path)
# Note: these are small batch-sizes - increase as appropriate to available GPU capacity
config.model.train_ds.batch_size=8
config.model.validation_ds.batch_size=8
print(OmegaConf.to_yaml(config))
```
# Model Training
## Setting up Data within the config
Among other things, the config file contains dictionaries called dataset, train_ds and validation_ds. These are configurations used to setup the Dataset and DataLoaders of the corresponding config.
We assume that both training and evaluation files are located in the same directory, and use the default names mentioned during the data download step.
So, to start model training, we simply need to specify `model.dataset.data_dir`, like we are going to do below.
Also notice that some config lines, including `model.dataset.data_dir`, have `???` in place of paths, this means that values for these fields are required to be specified by the user.
Let's now add the data directory path to the config.
```
# in this tutorial train and dev datasets are located in the same folder, so it is enought to add the path of the data directory to the config
config.model.dataset.data_dir = os.path.join(DATA_DIR, 'NER')
# if you want to decrease the size of your datasets, uncomment the lines below:
# NUM_SAMPLES = 1000
# config.model.train_ds.num_samples = NUM_SAMPLES
# config.model.validation_ds.num_samples = NUM_SAMPLES
```
## Building the PyTorch Lightning Trainer
NeMo models are primarily PyTorch Lightning modules - and therefore are entirely compatible with the PyTorch Lightning ecosystem.
Let's first instantiate a Trainer object
```
print("Trainer config - \n")
print(OmegaConf.to_yaml(config.trainer))
# lets modify some trainer configs
# checks if we have GPU available and uses it
cuda = 1 if torch.cuda.is_available() else 0
config.trainer.gpus = cuda
# for PyTorch Native AMP set precision=16
config.trainer.precision = 16 if torch.cuda.is_available() else 32
# remove distributed training flags
config.trainer.accelerator = 'DDP'
trainer = pl.Trainer(**config.trainer)
```
## Setting up a NeMo Experiment¶
NeMo has an experiment manager that handles logging and checkpointing for us, so let's use it:
```
exp_dir = exp_manager(trainer, config.get("exp_manager", None))
# the exp_dir provides a path to the current experiment for easy access
exp_dir = str(exp_dir)
exp_dir
# complete list of supported BERT-like models
print(nemo_nlp.modules.get_pretrained_lm_models_list())
# specify BERT-like model, you want to use
PRETRAINED_BERT_MODEL = "biomegatron-bert-345m-cased"
# add the specified above model parameters to the config
config.model.language_model.pretrained_model_name = PRETRAINED_BERT_MODEL
```
Now, we are ready to initialize our model. During the model initialization call, the dataset and data loaders we'll be prepared for training and evaluation.
Also, the pretrained BERT model will be downloaded, note it can take up to a few minutes depending on the size of the chosen BERT model.
```
model_ner = nemo_nlp.models.TokenClassificationModel(cfg=config.model, trainer=trainer)
```
## Monitoring training progress
Optionally, you can create a Tensorboard visualization to monitor training progress.
If you're not using Colab, refer to [https://www.tensorflow.org/tensorboard/tensorboard_in_notebooks](https://www.tensorflow.org/tensorboard/tensorboard_in_notebooks) if you're facing issues with running the cell below.
```
try:
from google import colab
COLAB_ENV = True
except (ImportError, ModuleNotFoundError):
COLAB_ENV = False
# Load the TensorBoard notebook extension
if COLAB_ENV:
%load_ext tensorboard
%tensorboard --logdir {exp_dir}
else:
print("To use tensorboard, please use this notebook in a Google Colab environment.")
# start model training
trainer.fit(model_ner)
```
# Inference
To see how the model performs, we can run generate prediction similar to the way we did it earlier
```
# let's first create a subset of our dev data
! head -n 100 {DATA_DIR}/NER/text_dev.txt > {DATA_DIR}/NER/sample_text_dev.txt
! head -n 100 {DATA_DIR}/NER/labels_dev.txt > {DATA_DIR}/NER/sample_labels_dev.txt
```
Now, let's generate predictions for the provided text file.
If labels file is also specified, the model will evaluate the predictions and plot confusion matrix.
```
model_ner.evaluate_from_file(
text_file=os.path.join(DATA_DIR, 'NER', 'sample_text_dev.txt'),
labels_file=os.path.join(DATA_DIR, 'NER', 'sample_labels_dev.txt'),
output_dir=exp_dir,
add_confusion_matrix=True,
normalize_confusion_matrix=True,
batch_size=1
)
# Please check matplotlib version if encountering any error plotting confusion matrix:
# https://stackoverflow.com/questions/63212347/importerror-cannot-import-name-png-from-matplotlib
```
## Training Script
If you have NeMo installed locally, you can also train the model with `nlp/token_classification/token_classification_train.py.`
To run training script, use:
`python token_classification_train.py model.dataset.data_dir=PATH_TO_DATA_DIR PRETRAINED_BERT_MODEL=biomegatron-bert-345m-cased`
The training could take several minutes and the result should look something like
```
[NeMo I 2020-05-22 17:13:48 token_classification_callback:82] Accuracy: 0.9882348032875798
[NeMo I 2020-05-22 17:13:48 token_classification_callback:86] F1 weighted: 98.82
[NeMo I 2020-05-22 17:13:48 token_classification_callback:86] F1 macro: 93.74
[NeMo I 2020-05-22 17:13:48 token_classification_callback:86] F1 micro: 98.82
[NeMo I 2020-05-22 17:13:49 token_classification_callback:89] precision recall f1-score support
O (label id: 0) 0.9938 0.9957 0.9947 22092
B (label id: 1) 0.8843 0.9034 0.8938 787
I (label id: 2) 0.9505 0.8982 0.9236 1090
accuracy 0.9882 23969
macro avg 0.9429 0.9324 0.9374 23969
weighted avg 0.9882 0.9882 0.9882 23969
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import numpy as np
import itertools
import h5py
import os
from scipy.stats import norm
import time
import matplotlib.pyplot as plt
import matplotlib.cm as cm
%matplotlib inline
import seaborn as sns
sns.set(style="ticks", color_codes=True, font_scale=1.5)
sns.set_style({"xtick.direction": "in", "ytick.direction": "in"})
```
We start by loading a few libraries, either external or from the smFS package. Above this point, you should be checking elsewhere in case anything failed. Below this point, we are to blame. First we modify the path, so that we can actually install the package and load the modules both for running Brownian dynamics and for running kinetics.
```
import sys
sys.path.append("..")
from smfsmodels import cossio
import kinetics
```
Now we define a couple of functions, that let us produce 2D-histograms and calculate kinetics.
```
def pmf2d(xq, qk):
fig, ax = plt.subplots(figsize=(6,5))
counts, ybins, xbins, image = ax.hist2d(xk, qk, \
bins=[np.linspace(-12,12,100), np.linspace(-12,12,100)])
pmf = -np.log(counts.transpose())
pmf -= np.min(pmf)
cs = ax.contourf(pmf, \
extent=[xbins.min(), xbins.max(), ybins.min(), ybins.max()], \
cmap=cm.rainbow, levels=np.arange(0,10 ,1))
cbar = plt.colorbar(cs)
ax.set_xlim(-12,12)
ax.set_ylim(-12,12)
ax.set_xlabel('$x$', fontsize=20)
ax.set_ylabel('$q$', fontsize=20)
plt.tight_layout()
def calc_rates(y):
lifeA, lifeB = kinetics.calc_life([y])
meanA = 1./np.exp(np.mean(np.log([x for x in lifeA if x>0])))
meanB = 1./np.exp(np.mean(np.log([x for x in lifeB if x>0])))
errorA = meanA/np.sqrt(len(lifeA))
errorB = meanA/np.sqrt(len(lifeB))
return np.mean([meanA, meanB]), np.mean([errorA, errorB])
def smooth(y, box_pts):
box = np.ones(box_pts)/box_pts
y_smooth = np.convolve(y, box, mode='same')
return y_smooth
```
#### Molecular potential of mean force
First we show the molecular potential of mean force on the extension coordinate, $x$, represented by a bistable potential.
```
x = np.linspace(-10, 10, 1000)
fig, ax = plt.subplots(2,1, figsize=(6,5), sharex=True)
Gx = [cossio.Gx(y, barrier=3., F12=0) for y in x]
dGqxdx = [cossio.dGqxdx(0, y, barrier=3., F12=0) for y in x]
ax[0].plot(x, Gx, lw=3)
ax[1].plot(x, dGqxdx, lw=3)
Gx = [cossio.Gx(y, barrier=3., F12=-1) for y in x]
dGqxdx = [cossio.dGqxdx(0, y, barrier=3., F12=-1) for y in x]
ax[0].plot(x, Gx, lw=3)
ax[1].plot(x, dGqxdx, lw=3)
Gx = [cossio.Gx(y, barrier=3., F12=1) for y in x]
dGqxdx = [cossio.dGqxdx(0, y, barrier=3., F12=1) for y in x]
ax[0].plot(x, Gx, lw=3)
ax[1].plot(x, dGqxdx, lw=3)
ax[0].set_ylabel('$G(x)$', fontsize=20)
ax[0].set_ylim(-9,5)
ax[1].set_xlabel('$x$', fontsize=20)
ax[1].set_ylabel('$\partial G(x)/\partial x$', fontsize=20)
ax[1].hlines(0, -10, 10, linestyle='dashed', linewidth=0.5)
ax[1].set_xlim(-10,10)
#ax[2].set_ylabel('$\partial^2 G(x)/\partial x^2$', fontsize=20)
plt.tight_layout(h_pad=0.2)
kl = 0.25
x = np.linspace(-15,15,100)
q = np.linspace(-15,15,50)
fig, ax = plt.subplots(1,3, figsize=(9,4), sharex=True, sharey=True)
for ii, f in enumerate([-1,0,1]):
G2d = np.ones((50, 100), float)*[cossio.Gx(y, barrier=5., F12=f) for y in x]
for i, j in itertools.product(range(50), range(100)):
G2d[i,j] += cossio.V(q[i], x[j], kl)
G2d = np.array(G2d)
G2d -= np.min(G2d)
cs = ax[ii].contourf(x, q, G2d, cmap=cm.rainbow, \
levels=np.arange(0,12,0.5), alpha=0.9)
#cbar = plt.colorbar(cs)
ax[ii].set_xlim(-15,15)
ax[ii].set_ylim(-15,15)
ax[1].set_xlabel('$x$', fontsize=20)
ax[0].set_ylabel('$q$', fontsize=20)
plt.tight_layout()
# Globals
dt = 5e-4
Dx = 1. # Diffusion coefficient for molecular coordinate
```
Next we invoke the `cossio` module to run the dynamics, whose function `run_brownian` will do the job. We are passing a number of input parameters, including the size of the free energy barrier and some variables for IO.
```
start = time.time()
x, q = [5., 5.]
t, xk, qk = cossio.run_brownian(x0=x, dt=dt, barrier=3., \
Dx=Dx, Dq=Dx, F12=-1, numsteps=int(1e8), \
fwrite=int(0.01/dt))
end = time.time()
print (end - start)
data = np.column_stack((t, xk, qk))
h5file = "data/cossio_kl%g_Dx%g_Dq%g_Fdt%g.h5"%(0, Dx, 0, dt)
try:
os.makedirs("data")
except OSError:
pass
with h5py.File(h5file, "w") as hf:
hf.create_dataset("data", data=data)
fig, ax = plt.subplots(figsize=(14,3))
ax.plot(t, xk, '.', ms=0.1)
ax.plot(t, smooth(xk, 1000), linewidth=0.5)
ax.set_ylim(-10,10)
ax.set_xlim(0,50000)
ax.set_yticks([-5.,0,5.])
ax.set_xlabel('time', fontsize=20)
ax.set_ylabel('x', fontsize=20)
start = time.time()
x, q = [-5., -5.]
t, xk, qk = cossio.run_brownian(x0=x, dt=dt, barrier=3., \
Dx=Dx, Dq=Dx, F12=1, numsteps=int(1e7), \
fwrite=int(0.01/dt))
end = time.time()
print (end - start)
data = np.column_stack((t, xk, qk))
h5file = "data/cossio_kl%g_Dx%g_Dq%g_Fdt%g.h5"%(0, Dx, 0, dt)
try:
os.makedirs("data")
except OSError:
pass
with h5py.File(h5file, "w") as hf:
hf.create_dataset("data", data=data)
fig, ax = plt.subplots(figsize=(14,3))
ax.plot(t, xk, '.', ms=0.1)
ax.plot(t, smooth(xk, 1000), linewidth=0.5)
ax.set_ylim(-10,10)
ax.set_xlim(0,50000)
ax.set_yticks([-5.,0,5.])
ax.set_xlabel('time', fontsize=20)
ax.set_ylabel('x', fontsize=20)
```
### Diffusion in two dimensions
```
x = np.linspace(-10,10,100)
G2d = np.ones((50, 100), float)*[cossio.Gx(y, barrier=5.) for y in x]
q = np.linspace(-12,12,50)
for i, j in itertools.product(range(50), range(100)):
G2d[i,j] += cossio.V(q[i], x[j], kl)
G2d = np.array(G2d)
G2d -= np.min(G2d)
import matplotlib.cm as cm
fig, ax = plt.subplots(figsize=(6,5))
cs = ax.contourf(x, q, G2d, cmap=cm.rainbow, levels=np.arange(0,10,1), alpha=0.9)
cbar = plt.colorbar(cs)
ax.set_xlim(-12,12)
ax.set_ylim(-12,12)
ax.set_xlabel('$x$', fontsize=20)
ax.set_ylabel('$q$', fontsize=20)
expGM = np.trapz(np.exp(-G2d), q, axis=0)
GM = -np.log(expGM)
expGA = np.trapz(np.exp(-G2d), x, axis=1)
GA = -np.log(expGA)
fig, ax = plt.subplots()
ax.plot(x, GM - np.min(GM), label='$G_M$', lw=4)
ax.plot(q, GA - np.min(GA), label='$G_A$', lw=4)
#ax.plot(bin_centers, [cossio.Gx(y) for y in bin_centers], '--', c='red', lw=3)
ax.set_xlim(-10,10)
ax.set_ylim(-1,7)
ax.set_xlabel('Extension', fontsize=20)
ax.set_ylabel('Free Energy', fontsize=20)
ax.legend(loc=1)
fig.tight_layout()
x, q = [5., 5.]
t, xk, qk = cossio.run_brownian(x0=x, dt=dt, barrier=5., kl=kl, \
Dx=Dx, Dq=Dx, numsteps=int(1e8), fwrite=int(0.1/dt))
fig, ax = plt.subplots(2,1, figsize=(10,5), sharex=True, sharey=True)
ax[0].plot(xk, linewidth=0.2)
ax[1].plot(qk, 'r', linewidth=0.2)
ax[1].set_xlabel('time', fontsize=20)
ax[1].set_ylabel('$q$', fontsize=20)
ax[0].set_ylabel('$x$', fontsize=20)
ax[0].set_ylim(-10, 10)
#ax[0].set_xlim(0, numsteps)
n
scaling = [5, 2, 1, 0.5, 0.2, 0.1, 0.05, 0.02, 0.01]
nsteps = [1e7, 2e7, 2e7, 2e7, 2e7, 2e7, 2e7, 5e7, 5e7, 5e7]
trajs = {}
for i,sc in enumerate(scaling):
trajs[sc] = {}
x, q = [5., 5.]
numsteps = int(nsteps[i])
xk, qk = cossio.run_brownian(x0=x, dt=dt, barrier=5., kl=kl, \
Dx=Dx, Dq=Dx*sc, numsteps=numsteps, fwrite=100)
trajs[sc]['x'] = xk
trajs[sc]['q'] = qk
fig, ax = plt.subplots(2,1, figsize=(10,5), sharex=True, sharey=True)
ax[0].plot(xk, linewidth=0.2, label='Dx/Dq=%g'%(1./sc))
ax[1].plot(qk, 'r', linewidth=0.2)
ax[1].set_xlabel('Time', fontsize=20)
ax[1].set_ylabel('$q$', fontsize=20)
ax[0].set_ylabel('$x$', fontsize=20)
ax[0].set_ylim(-10, 10)
ax[0].legend(loc=1)
kfq = []
kfx = []
for i, sc in enumerate(scaling):
kfx.append(calc_rates(trajs[sc]['x']))
kfq.append(calc_rates(trajs[sc]['q']))
fig, ax = plt.subplots()
rates = np.array(kfq)
ax.errorbar(1./np.array(scaling), rates[:,0]/rate0[0], \
rates[:,1]/rate0[0], marker='o')
ax.set_xscale('log')
ax.set_xlabel('$D_x/D_q$', fontsize=24)
ax.set_ylabel('$k_{MA}/k_M$', fontsize=24)
ax.set_xlim(2e-2,5e2)
plt.tight_layout()
scaling = [10]
nsteps = [1e8]
for i,sc in enumerate(scaling):
trajs[sc] = {}
x, q = [5., 5.]
numsteps = int(nsteps[i])
xk, qk = cossio.run_brownian(x0=x, dt=dt/5., barrier=5., kl=kl, \
Dx=Dx, Dq=Dx*sc, numsteps=numsteps, fwrite=1000)
trajs[sc]['x'] = xk
trajs[sc]['q'] = qk
fig, ax = plt.subplots(2,1, figsize=(10,5), sharex=True, sharey=True)
ax[0].plot(xk, linewidth=0.2, label='Dx/Dq=%g'%(1./sc))
ax[1].plot(qk, 'r', linewidth=0.2)
ax[1].set_xlabel('Time', fontsize=20)
ax[1].set_ylabel('$q$', fontsize=20)
ax[0].set_ylabel('$x$', fontsize=20)
ax[0].set_ylim(-10, 10)
ax[0].legend(loc=1)
scaling = [10, 5, 2, 1, 0.5, 0.2, 0.1, 0.05, 0.02, 0.01]
kfq = []
kfx = []
for i, sc in enumerate(scaling):
#kfx.append(calc_rates(trajs[sc]['x']))
kfq.append(calc_rates(trajs[sc]['q']))
fig, ax = plt.subplots()
rates = np.array(kfq)
ax.errorbar(1./np.array(scaling), rates[:,0]/rate0[0], \
rates[:,1]/rate0[0], marker='o')
ax.set_xscale('log')
ax.set_xlabel('$D_x/D_q$', fontsize=24)
ax.set_ylabel('$k_{MA}/k_M$', fontsize=24)
ax.set_xlim(2e-2,5e2)
plt.tight_layout()
```
| github_jupyter |
# Usage
This document is intended as a fast way to get an idea of what LDA can produce. Actual research should be done using a full experimental process including the use of the "LDA Job manager" notebook.
To make and inspect a quick topic model:
1. Make sure that you are using a fully functional notebook viewer, such as VS Code (best) or Jupyter Notebooks. Use options like the ability to collapse sections or input cells. Other options, like Jupyter Lab or custom web views, can be configured to work, but that's on you.
1. Prepare a dataset with at least columns for a unique document ID number and text you want to process, with a single textual response per row. LDA does not require preprocessed text to function, but it is easier to interpret results if you use the preprocessing notebook first.
1. Edit the data import section ([click here](#data)) with the path, columns names etc for your dataset.
1. Run the notebook
1. Look at the results in the model inspection section ([click here](#model-inspection))
1. If you want to try looking at particular subsets of your data look at the examples section ([click here](#examples-of-how-to-look-at-subsets-of-your-set-of-documents))
1. Keep in mind that LDA works best on a large textual dataset (many comments), where each comment is long. We didn't find the need to remove short comments, but you need long comments.
# Imports
## Libraries
```
import pandas as pd
import numpy as np
from scipy.stats.mstats import gmean
from ipywidgets import interact, Combobox
from IPython.display import display, display_html
from gensim.corpora import Dictionary
from gensim.models.ldamodel import LdaModel
from gensim.models import CoherenceModel
```
## Data
### Data Details
```
index_col = "unique_comment_ID" # Unique number/code for each document
text_col = "Preprocessed answer" # Text to be fed to LDA
nice_text_col = "answer" # Unprocessed text for viewing. Can be same as text_col
```
### Import Data
```
data_path = "/home/azureuser/cloudfiles/code/Data/pp-20210830_SES_and_SET.csv"
raw_df = pd.read_csv(data_path) # Import data
raw_df.set_index(index_col, inplace=True) # Set the document index as index
raw_df.dropna(subset=[text_col],inplace=True) # Remove all rows with empty or missing text
raw_df[text_col] = raw_df[text_col].astype('string') # make sure the text columns is all strings
```
### If your dataset is large, you may want to reduce the size of raw_df by selecting rows to reduce computation time intially. For instance, we normally choose to look at comments only from our newer SES survey, even though that makes up 250k of 1.5 million textual responses.
```
display(f"Number of comments: {len(raw_df)}")
raw_df.head(3)
```
## Gensim Components from Data
This section is helpful if you want to understand the various steps to feeding textual data into a computational framework like Gensim
### Tokenize Documents
```
texts = raw_df[[text_col]].applymap(str.split)
texts.head(2)
```
### Generate Dictionary
```
dictionary = Dictionary(texts[text_col])
display(f"Number of Words: {len(dictionary)}")
words = [*dictionary.token2id]
```
### Create Corpus
```
corpus = texts.applymap(dictionary.doc2bow)
corpus.head(2)
```
# Other Defintions
```
def display_side_by_side(*args):
html_str=''
for df in args:
html_str+=df.to_html() + ("\xa0" * 5) # Spaces
display_html(html_str.replace('<table','<table style="display:inline"'),raw=True)
```
# Topic Model Setup
You should not have to edit anything in this section.
## Helper Functions
```
def convert_row_to_term_score(row):
'''Converts a word-topic matrix to a term score matrix.
Input should be a series of probabilities (intent is that the term is the index)'''
normalizer = gmean(row) # Compute geometric mean of the word probabilities
term_score_row = row.apply(lambda b: b*(np.log(b/normalizer))) #applying the transformation
return term_score_row
```
## LDA Class Definition
```
class QuickLDA(object):
def __init__(self,doc_ids, num_topics = 7):
'''Takes a list of doc ids and creates all the LDA components'''
self.doc_ids = list(corpus.loc[doc_ids].index) # Making sure this is ordered correctly. Probably not necessary
self.num_topics = num_topics
self.sub_corpus = corpus.loc[doc_ids][text_col] # This is not a dataframe, just an iterable
self.num_docs = len(self.sub_corpus)
self.fit_lda()
self.score_lda()
self.make_term_matrices()
self.make_doc_topic_matrix()
def fit_lda(self):
lda = LdaModel(
id2word = dictionary,
passes = int(np.ceil(50000/self.num_docs)), # Extra fitting for small corpi
num_topics = self.num_topics,
alpha = "auto"
)
lda.update(self.sub_corpus)
self.lda = lda
def score_lda(self):
self.perplexity = 2**(-self.lda.log_perplexity(self.sub_corpus))
c_model = CoherenceModel(
model = self.lda,
texts = texts.loc[self.doc_ids][text_col], #Again can't have dataframe
dictionary = dictionary,
coherence = "c_v"
)
self.cv_score = c_model.get_coherence()
def make_term_matrices(self):
self.term_topic_matrix = pd.DataFrame(self.lda.get_topics()).transpose()
self.term_topic_matrix.rename(
index = dictionary.id2token,
inplace=True
)
self.term_score_matrix = self.term_topic_matrix.apply(convert_row_to_term_score,axis=1)
def make_doc_topic_matrix(self):
document_topic_matrix = pd.DataFrame(
[{doc_tuple[0]:doc_tuple[1] for doc_tuple in doc_tuple_list} for doc_tuple_list in self.lda[self.sub_corpus]])
# Fill Missing Values
document_topic_matrix.fillna(0,inplace = True)
# Sort columns by topic number
document_topic_matrix = document_topic_matrix.reindex(sorted(document_topic_matrix.columns), axis=1)
document_topic_matrix.index = self.sub_corpus.index
self.document_topic_matrix = document_topic_matrix
self.topic_means = document_topic_matrix.mean().apply(lambda x: round(x, 3))
```
## LDA Visuals Definitions
```
def plot_term(lda, word = "class"):
try:
display_html(f"<h4> Probability(term|topic) for \"{word}\"",raw=True)
display_html(lda.term_topic_matrix.loc[[word]].transpose().plot.bar(ylabel = "Conditional term probability",xlabel = "Topic"))
except KeyError as e: print("Waiting for valid input")
def get_top_responses(topic_name,number_responses,lda, doc_metadata = None, max_words = 1000):
doc_ids = lda.document_topic_matrix.sort_values(by=topic_name,ascending = False)
doc_ids = doc_ids.index.tolist()
doc_ids = list(filter(
lambda doc_id: len(texts.loc[doc_id][text_col]) < max_words,
doc_ids))
doc_ids = doc_ids[:number_responses]
# Print results
for doc_id in doc_ids:
if doc_metadata is not None: # Check if we want to display metadata with each comment
display(doc_metadata.loc[[doc_id]].style.hide_index())
display_html(" • " + raw_df.loc[doc_id][nice_text_col] + "<br><br><br>", raw = True)
```
# Examples of how to look at subsets of your set of documents
Below is a set of examples showing how to look at particular subsets and a fitting LDA for those subsets. If you have a dataframe you like, an easy way to get the list of document IDs is to use .index.tolist(). I give separate examples here, but you can combine, or bring in your own list of document IDs based on something else like sentiment analysis.
## Getting all doc_ids for a particular question
In this example I wanted to get all of the answers to "what specific change in clarity would help learning". I use the .isin method to ask if a particular column has a value in a list that I give. So in this case you could write a bunch of question IDs out.
```
# clarity_ids = raw_df[raw_df["question_ID"].isin(
# ["X840307","Your Document Code Here"]
# )].index.tolist()
# display_html("<h4>Sample Selected Texts:", raw=True)
# for row in raw_df.loc[clarity_ids][nice_text_col].head(3):
# display(row)
```
## Getting all Document IDs for a certain list of words
This example looks at all responses containing particular words and does the full LDA exploration for that set of documents.
```
# @interact(word = Combobox(options = words,continuous_update = False))
# def show_words(word):
# display_html("Type in here if you want to see what the kernel thinks are words", raw=True)
```
#### Each document will need to contain at least one word from this list
```
# req_words = ["canvas"]
```
The following code gets all responses for which the preprocessed answer contains a word from the req_words list. It generates a list of True/False for each word pairing that might agree between the two lists, then "any" collapses that into a single True if there was any agreement. The result of apply, which is a dataframe with True/False as it's main column, it used to select a subset of the larger data as usual, then the index is extracted as a list.
```
# word_doc_ids = texts[texts[text_col].apply(
# lambda tokenized_text: any(word in tokenized_text for word in req_words)
# )].index.tolist()
# display_html(f"<b>Number of doc ids: {len(word_doc_ids)}",raw=True)
# display_html("<h4>Sample Selected Texts:",raw= True)
# for row in raw_df.loc[word_doc_ids][nice_text_col].head(2):
# display(row)
# word_lda = QuickLDA(doc_ids=word_doc_ids,num_topics=8)
```
# Model Inspection
After an initial run of the notebook, you only need to rerun these cells and below to change your model and output.
```
doc_ids = raw_df[raw_df["survey"] == "SES"].index.tolist()
basic_lda = QuickLDA(doc_ids = doc_ids,num_topics= 7) # Fit a topic model on all of the supplied textual data
lda = basic_lda # Set the topic model to be inspected.
```
Check the topic means to make sure that it actually worked. If the topic means seem too focused on one topic, then you need to change the number of topics or select more documents.
```
display_html(f"<b> Coherence Score (c_v): </b> {lda.cv_score}",raw = True)
display_html(f"<b> Perplexity: </b> {lda.perplexity}",raw = True)
display(lda.topic_means)
```
### Explore the distribution of a particular term
```
@interact(word = Combobox(options = list(lda.term_score_matrix.index)), continuous_update = False)
def f(word):
plot_term(lda,word)
```
### Raw display of top words for all topics
```
@interact(show = False,num_top_words = (5,30,100))
def relevant_words(show,num_top_words = 14):
# Display top words per topic
if show:
for c in lda.term_score_matrix.columns:
print(f'\n Topic {c} -- {lda.topic_means[c]} \n',
lda.term_score_matrix[c]
.sort_values(ascending=False) #Sort most relevant words by their term score in column 'c'
.head(num_top_words) #Take top ten most relevant words
.index #The index is the word itself
.tolist() #Feel free to replace with some nicer display function
)
```
### Top Words per Topic
```
@interact(topic = lda.document_topic_matrix.columns, num = (5,100), cols = (1,10),include_term_score = True)
def top_words(topic,num = 30, cols = 4, include_term_score = True):
sorted_term_score = lda.term_score_matrix.sort_values(by = topic, ascending = False)[[topic]] # Prepare terms sorted by score
sorted_term_score.columns = ["Term Score"]
display_html(f"<h4><u> Most Relevant words for Topic {topic} ({lda.topic_means[topic]}):", raw = True) # Heading
if include_term_score:
per_col = int(np.ceil(num/cols)) # Figure out how many words to put per column
display_side_by_side(*[sorted_term_score.iloc[x: x + per_col] for x in range(0,num,per_col)]) # Display the columns. *[] used to partition the dataframe
else:
print(sorted_term_score.head(num).index.tolist()) # Print them out plainly if we want that for some reason.
```
### Top Comments by Topic
```
@interact(
topic = lda.document_topic_matrix.columns, # Choose a topic from the doc-topic matrix
number_responses = [1,5,10,20,100,1000], # Choose a number of responses
max_words = [5,10,20,50,1000], # Max number of words in the responses
include_topic_distributions = False # Choose whether you want to show the entry from the doc-topic matrix for each response
)
def top_resp(topic, number_responses = 5, include_topic_distributions = False, max_words = 1000):
if include_topic_distributions:
metadata = lda.document_topic_matrix # Set the metadata to display and populate it
else: metadata = None
display_html(f"<h2><u> Top Responses for Topic {topic} ({lda.topic_means[topic]}):", raw = True)
return get_top_responses(topic_name = topic, number_responses = number_responses, doc_metadata = metadata, lda = lda, max_words = max_words)
```
| github_jupyter |
# Acme: Quickstart
## Guide to installing Acme and training your first D4PG agent.
# <a href="https://colab.research.google.com/github/deepmind/acme/blob/master/examples/quickstart.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Select your environment library
Note: `dm_control` requires a valid Mujoco license.
```
environment_library = 'gym' # @param ['dm_control', 'gym']
```
## Add your Mujoco license here
Note: only required for `dm_control`.
```
mjkey = """
""".strip()
if not mjkey and environment_library == 'dm_control':
raise ValueError(
'A Mujoco license is required for `dm_control`, if you do not have on '
'consider selecting `gym` from the dropdown menu in the cell above.')
```
## Installation
### Install Acme
```
!pip install dm-acme
!pip install dm-acme[reverb]
!pip install dm-acme[tf]
```
### Install the environment library
Without a valid license you won't be able to use the `dm_control` environments but can still follow this colab using the `gym` environments.
If you have a personal Mujoco license (_not_ an institutional one), you may
need to follow the instructions at https://research.google.com/colaboratory/local-runtimes.html to run a Jupyter kernel on your local machine.
This will allow you to install `dm_control` by following instructions in
https://github.com/deepmind/dm_control and using a personal MuJoCo license.
```
#@test {"skip": true}
if environment_library == 'dm_control':
mujoco_dir = "$HOME/.mujoco"
# Install OpenGL dependencies
!apt-get update && apt-get install -y --no-install-recommends \
libgl1-mesa-glx libosmesa6 libglew2.0
# Get MuJoCo binaries
!wget -q https://www.roboti.us/download/mujoco200_linux.zip -O mujoco.zip
!unzip -o -q mujoco.zip -d "$mujoco_dir"
# Copy over MuJoCo license
!echo "$mjkey" > "$mujoco_dir/mjkey.txt"
# Install dm_control
!pip install dm_control
# Configure dm_control to use the OSMesa rendering backend
%env MUJOCO_GL=osmesa
# Check that the installation succeeded
try:
from dm_control import suite
env = suite.load('cartpole', 'swingup')
pixels = env.physics.render()
except Exception as e:
raise RuntimeError(
'Something went wrong during installation. Check the shell output above '
'for more information. If you do not have a valid Mujoco license, '
'consider selecting `gym` in the dropdown menu at the top of this '
'Colab.') from e
else:
del suite, env, pixels
elif environment_library == 'gym':
!pip install gym
```
### Install visualization packages
```
!sudo apt-get install -y xvfb ffmpeg
!pip install imageio
!pip install PILLOW
!pip install pyvirtualdisplay
```
## Import Modules
```
import IPython
from acme import environment_loop
from acme import specs
from acme import wrappers
from acme.agents.tf import d4pg
from acme.tf import networks
from acme.tf import utils as tf2_utils
from acme.utils import loggers
import numpy as np
import sonnet as snt
# Import the selected environment lib
if environment_library == 'dm_control':
from dm_control import suite
elif environment_library == 'gym':
import gym
# Imports required for visualization
import pyvirtualdisplay
import imageio
import base64
# Set up a virtual display for rendering.
display = pyvirtualdisplay.Display(visible=0, size=(1400, 900)).start()
```
## Load an environment
We can now load an environment. In what follows we'll create an environment and grab the environment's specifications.
```
if environment_library == 'dm_control':
environment = suite.load('cartpole', 'balance')
elif environment_library == 'gym':
environment = gym.make('MountainCarContinuous-v0')
environment = wrappers.GymWrapper(environment) # To dm_env interface.
else:
raise ValueError(
"Unknown environment library: {};".format(environment_library) +
"choose among ['dm_control', 'gym'].")
# Make sure the environment outputs single-precision floats.
environment = wrappers.SinglePrecisionWrapper(environment)
# Grab the spec of the environment.
environment_spec = specs.make_environment_spec(environment)
```
## Create a D4PG agent
```
#@title Build agent networks
# Get total number of action dimensions from action spec.
num_dimensions = np.prod(environment_spec.actions.shape, dtype=int)
# Create the shared observation network; here simply a state-less operation.
observation_network = tf2_utils.batch_concat
# Create the deterministic policy network.
policy_network = snt.Sequential([
networks.LayerNormMLP((256, 256, 256), activate_final=True),
networks.NearZeroInitializedLinear(num_dimensions),
networks.TanhToSpec(environment_spec.actions),
])
# Create the distributional critic network.
critic_network = snt.Sequential([
# The multiplexer concatenates the observations/actions.
networks.CriticMultiplexer(),
networks.LayerNormMLP((512, 512, 256), activate_final=True),
networks.DiscreteValuedHead(vmin=-150., vmax=150., num_atoms=51),
])
# Create a logger for the agent and environment loop.
agent_logger = loggers.TerminalLogger(label='agent', time_delta=10.)
env_loop_logger = loggers.TerminalLogger(label='env_loop', time_delta=10.)
# Create the D4PG agent.
agent = d4pg.D4PG(
environment_spec=environment_spec,
policy_network=policy_network,
critic_network=critic_network,
observation_network=observation_network,
sigma=1.0,
logger=agent_logger,
checkpoint=False
)
# Create an loop connecting this agent to the environment created above.
env_loop = environment_loop.EnvironmentLoop(
environment, agent, logger=env_loop_logger)
```
## Run a training loop
```
# Run a `num_episodes` training episodes.
# Rerun this cell until the agent has learned the given task.
env_loop.run(num_episodes=100)
```
## Visualize an evaluation loop
### Helper functions for rendering and vizualization
```
# Create a simple helper function to render a frame from the current state of
# the environment.
if environment_library == 'dm_control':
def render(env):
return env.physics.render(camera_id=0)
elif environment_library == 'gym':
def render(env):
return env.environment.render(mode='rgb_array')
else:
raise ValueError(
"Unknown environment library: {};".format(environment_library) +
"choose among ['dm_control', 'gym'].")
def display_video(frames, filename='temp.mp4'):
"""Save and display video."""
# Write video
with imageio.get_writer(filename, fps=60) as video:
for frame in frames:
video.append_data(frame)
# Read video and display the video
video = open(filename, 'rb').read()
b64_video = base64.b64encode(video)
video_tag = ('<video width="320" height="240" controls alt="test" '
'src="data:video/mp4;base64,{0}">').format(b64_video.decode())
return IPython.display.HTML(video_tag)
```
### Run and visualize the agent in the environment for an episode
```
timestep = environment.reset()
frames = [render(environment)]
while not timestep.last():
# Simple environment loop.
action = agent.select_action(timestep.observation)
timestep = environment.step(action)
# Render the scene and add it to the frame stack.
frames.append(render(environment))
# Save and display a video of the behaviour.
display_video(np.array(frames))
```
| github_jupyter |
```
!pip install jsonlines
import os
import sys
import csv, jsonlines
import numpy as np
import copy
import random
# Importing tensorflow
import tensorflow as tf
# Importing some more libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error as MSE
from sklearn import preprocessing
%matplotlib inline
import matplotlib.pyplot as plt
# Reading ratings file
ratings = pd.read_csv('ratings.csv', sep='\t', encoding='latin-1',
usecols=['user_id', 'movie_id', 'user_emb_id', 'movie_emb_id', 'rating'])
max_userid = ratings['user_id'].drop_duplicates().max()
max_movieid = ratings['movie_id'].drop_duplicates().max()
# Reading ratings file
users = pd.read_csv('users.csv', sep='\t', encoding='latin-1',
usecols=['user_id', 'gender', 'zipcode', 'age_desc', 'occ_desc'])
# Reading ratings file
movies = pd.read_csv('movies.csv', sep='\t', encoding='latin-1',
usecols=['movie_id', 'title', 'genres'])
# You need an interaction matrix of ratings where column and row align to user_id and movie_id
ratings = ratings.drop('user_emb_id', axis=1).drop('movie_emb_id', axis=1)
num_movies = ratings.movie_id.nunique()
num_users = ratings.user_id.nunique()
print("USERS: {} MOVIES: {}".format(num_users, num_movies))
import collections
import csv
import os
Rating = collections.namedtuple('Rating', ['user_id', 'item_id', 'rating'])
class Dataset(collections.namedtuple('Dataset', ['users', 'items', 'ratings'])):
#users: set[str]
#items: set[str]
#ratings: list[Rating]
__slots__ = ()
def __str__(self):
out = 'Users: {:,d}\n'.format(self.n_users)
out += 'Items: {:,d}\n'.format(self.n_items)
out += 'Ratings: {:,d}\n'.format(self.n_ratings)
return out
@property
def n_users(self):
return len(self.users)
@property
def n_items(self):
return len(self.items)
@property
def n_ratings(self):
return len(self.ratings)
def user_ratings(self, user_id):
return list(r for r in self.ratings if r.user_id == user_id)
def item_ratings(self, item_id):
return list(r for r in self.ratings if r.item_id == item_id)
def filter_ratings(self, users, items):
return list(((r.user_id, r.item_id), r.rating)
for r in self.ratings
if r.user_id in users
and r.item_id in items)
def new_dataset(ratings):
users = set(r.user_id for r in ratings)
items = set(r.item_id for r in ratings)
return Dataset(users, items, ratings)
small_dataset = new_dataset([Rating(x['user_id'], x['movie_id'], x['rating']) for i, x in ratings.iterrows()])
print('Dataset\n\n{}'.format(small_dataset))
from random import shuffle
def split_randomly(dataset, train_ratio=0.80):
ratings = dataset.ratings
shuffle(ratings)
size = int(len(ratings) * train_ratio)
train_ratings = ratings[:size]
test_ratings = ratings[size:]
return new_dataset(train_ratings), \
new_dataset(test_ratings)
train_valid_data, test_data = split_randomly(small_dataset)
train_data, valid_data = split_randomly(train_valid_data)
train_eval = list(((r.user_id, r.item_id), r.rating) for r in train_data.ratings)
print('Evaluation ratings for train: {:,d}'.format(len(train_eval)))
# only items in train will be available for validation
valid_items = train_data.items & valid_data.items
print('Items in train and validation: {:,d}'.format(len(valid_items)))
# users from validation that has any item from train
valid_users = set(r.user_id for r in valid_data.ratings if r.item_id in train_data.items)
print('Users in validation with train items: {:,d}'.format(len(valid_users)))
# only users in train are available for validation
valid_users &= train_data.users
print('Users in train and validation: {:,d}'.format(len(valid_users)))
valid_eval = valid_data.filter_ratings(valid_users, valid_items)
print('Evaluation ratings for validation: {:,d}'.format(len(valid_eval)))
# Map User <-> index
# Map Item <-> index
IndexMapping = collections.namedtuple('IndexMapping', ['users_to_idx',
'users_from_idx',
'items_to_idx',
'items_from_idx'])
def map_index(values):
values_from_idx = dict(enumerate(values))
values_to_idx = dict((value, idx) for idx, value in values_from_idx.items())
return values_to_idx, values_from_idx
def new_mapping(dataset):
users_to_idx, users_from_idx = map_index(dataset.users)
items_to_idx, items_from_idx = map_index(dataset.items)
return IndexMapping(users_to_idx, users_from_idx, items_to_idx, items_from_idx)
import tensorflow as tf
import numpy as np
from tensorflow.contrib.factorization import WALSModel
class ALSRecommenderModel:
def __init__(self, user_factors, item_factors, mapping):
self.user_factors = user_factors
self.item_factors = item_factors
self.mapping = mapping
def transform(self, x):
for user_id, item_id in x:
if user_id not in self.mapping.users_to_idx \
or item_id not in self.mapping.items_to_idx:
yield (user_id, item_id), 0.0
continue
i = self.mapping.users_to_idx[user_id]
j = self.mapping.items_to_idx[item_id]
u = self.user_factors[i]
v = self.item_factors[j]
r = np.dot(u, v)
yield (user_id, item_id), r
def recommend(self, user_id, num_items=10, items_exclude=set()):
i = self.mapping.users_to_idx[user_id]
u = self.user_factors[i]
V = self.item_factors
P = np.dot(V, u)
rank = sorted(enumerate(P), key=lambda p: p[1], reverse=True)
top = list()
k = 0
while k < len(rank) and len(top) < num_items:
j, r = rank[k]
k += 1
item_id = self.mapping.items_from_idx[j]
if item_id in items_exclude:
continue
top.append((item_id, r))
return top
class ALSRecommender:
def __init__(self, num_factors=10, num_iters=10, reg=1e-1):
self.num_factors = num_factors
self.num_iters = num_iters
self.regularization = reg
def fit(self, dataset, verbose=False):
with tf.Graph().as_default(), tf.Session() as sess:
input_matrix, mapping = self.sparse_input(dataset)
model = self.als_model(dataset)
self.train(model, input_matrix, verbose)
row_factor = model.row_factors[0].eval()
col_factor = model.col_factors[0].eval()
return ALSRecommenderModel(row_factor, col_factor, mapping)
def sparse_input(self, dataset):
mapping = new_mapping(dataset)
indices = [(mapping.users_to_idx[r.user_id],
mapping.items_to_idx[r.item_id])
for r in dataset.ratings]
values = [r.rating for r in dataset.ratings]
shape = (dataset.n_users, dataset.n_items)
return tf.SparseTensor(indices, values, shape), mapping
def als_model(self, dataset):
return WALSModel(
dataset.n_users,
dataset.n_items,
self.num_factors,
regularization=self.regularization,
unobserved_weight=0)
def train(self, model, input_matrix, verbose=False):
rmse_op = self.rmse_op(model, input_matrix) if verbose else None
row_update_op = model.update_row_factors(sp_input=input_matrix)[1]
col_update_op = model.update_col_factors(sp_input=input_matrix)[1]
model.initialize_op.run()
model.worker_init.run()
for _ in range(self.num_iters):
# Update Users
model.row_update_prep_gramian_op.run()
model.initialize_row_update_op.run()
row_update_op.run()
# Update Items
model.col_update_prep_gramian_op.run()
model.initialize_col_update_op.run()
col_update_op.run()
if verbose:
print('RMSE: {:,.3f}'.format(rmse_op.eval()))
def approx_sparse(self, model, indices, shape):
row_factors = tf.nn.embedding_lookup(
model.row_factors,
tf.range(model._input_rows),
partition_strategy="div")
col_factors = tf.nn.embedding_lookup(
model.col_factors,
tf.range(model._input_cols),
partition_strategy="div")
row_indices, col_indices = tf.split(indices,
axis=1,
num_or_size_splits=2)
gathered_row_factors = tf.gather(row_factors, row_indices)
gathered_col_factors = tf.gather(col_factors, col_indices)
approx_vals = tf.squeeze(tf.matmul(gathered_row_factors,
gathered_col_factors,
adjoint_b=True))
return tf.SparseTensor(indices=indices,
values=approx_vals,
dense_shape=shape)
def rmse_op(self, model, input_matrix):
approx_matrix = self.approx_sparse(model, input_matrix.indices, input_matrix.dense_shape)
err = tf.sparse_add(input_matrix, approx_matrix * (-1))
err2 = tf.square(err)
n = input_matrix.values.shape[0].value
return tf.sqrt(tf.sparse_reduce_sum(err2) / n)
als = ALSRecommender()
als_model = als.fit(train_data, verbose=True)
for k in range(10):
x, y = valid_eval[k]
_, y_hat = list(als_model.transform([x]))[0]
print(*x, y, y_hat)
def _rmse(model, data):
x, y = zip(*data)
y_hat = list(r_hat for _, r_hat in model.transform(x))
return np.sqrt(np.mean(np.square(np.subtract(y, y_hat))))
def eval_rmse(model):
rmse = _rmse(model, train_eval)
print('RMSE (train): {:,.3f}'.format(rmse))
rmse = _rmse(model, valid_eval)
print('RMSE (validation): {:,.3f}'.format(rmse))
eval_rmse(als_model)
als = ALSRecommender(num_factors=10, num_iters=10, reg=0.1)
print('Training...\n')
als_model = als.fit(train_data, verbose=True)
print('\nEvaluation...\n')
eval_rmse(als_model)
```
| github_jupyter |
```
# For PostGREs and SQLAlchemy
!pip install psycopg2 sqlalchemy
# Dependencies and Setup
# SQL Alchemy, Pandas, Matplotlib, NumpY
from sqlalchemy import create_engine
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# Create Engine and Setup to Connect to Database
engine = create_engine('postgres://postgres:paswordxxx@localhost:5432/employee_db')
conn = engine.connect()
# Query in the Salaries Table
salaries = pd.read_sql("SELECT * FROM salaries", conn)
salaries
```
# Create a histogram to visualize the most common salary ranges for employees.
```
Hist = salaries['salary']
plt.hist(Hist)
plt.title('Most common salary range for employees')
plt.xlabel('Salaries')
plt.ylabel('Counts')
plt.show()
```
# Create a bar chart of average salary by title.
```
# Query in the Titles Table
titles = pd.read_sql("SELECT * FROM titles", conn)
titles
# Query in the Employee Table
employee = pd.read_sql("SELECT * FROM employees", conn)
employee
# Merge salaries and employee dataframe on "emp_no"
merged = pd.merge(salaries, employee, on="emp_no", how="inner")
merged.head()
#Rename column "title_id" for "emp_title_id"
rename = titles.rename(columns={"title_id":"emp_title_id"})
rename
#Merging merged and rename dataframe on "emp_title_id" so we can see the title of each employee
merge2 = pd.merge(merged, rename, on="emp_title_id", how="inner")
merge2
# Groupby Title
grouped_df = merge2.groupby("title").mean()
grouped_df
#Drop the "emp_no" columns
title_salary_df = grouped_df.drop(columns = "emp_no")
title_salary_df
#Reset index
title_salary_df = title_salary_df.reset_index()
title_salary_df
# Set x_axis, y_axis & Tick Locations
x_axis = title_salary_df["title"]
ticks = np.arange(len(x_axis))
y_axis = title_salary_df["salary"]
# Create Bar Chart Based on Above Data
plt.bar(x_axis, y_axis, align="center", alpha=0.5, color=["pink", "b", "r", "orange", "y", "b", "g"])
# Create Ticks for Bar Chart's x_axis
plt.xticks(ticks, x_axis, rotation="vertical")
# Set Labels & Title
plt.ylabel("Salaries ($)")
plt.xlabel("Employee Titles")
plt.title("Average Employee Salary by Title")
# Save Figure
plt.savefig("./Images/average_salary_by_title.png")
# Show plot
plt.show()
```
# Epilogue
Evidence in hand, you march into your boss's office and present the visualization. With a sly grin, your boss thanks you for your work. On your way out of the office, you hear the words, "Search your ID number." You look down at your badge to see that your employee ID number is 499942.
```
Personal_Pay=pd.read_sql('Select * From Salaries Where emp_no=499942', conn)
Personal=pd.read_sql('Select * From Employees Where emp_no=499942', conn)
print(Personal_Pay)
Personal
#['salary'][0]
```
| github_jupyter |
# String Types
```
new_string = "This is a String" # storing a string
print('ID:', id(new_string)) # shows the object identifier (address)
print('Type:', type(new_string)) # shows the object type
print('Value:', new_string) # shows the object value
```
### Simple String
```
simple_string = 'Hello!' + " I'm a simple string"
print(simple_string)
```
### Multi-line String
```
# Note the \n (newline) escape character automatically created
multi_line_string = """Hello I'm
a multi-line
string!"""
multi_line_string
print(multi_line_string)
```
### Escape sequences
```
# Normal string with escape sequences leading to a wrong file path!
escaped_string = "C:\the_folder\new_dir\file.txt"
print(escaped_string) # will cause errors if we try to open a file here
# raw string keeping the backslashes in its normal form
raw_string = r'C:\the_folder\new_dir\file.txt'
print(raw_string)
```
### Unicode literals
```
# unicode string literals
string_with_unicode = 'H\u00e8llo!'
print(string_with_unicode)
more_unicode = 'I love Pizza 🍕! Shall we book a cab 🚕 to get pizza?'
print(more_unicode)
```
## Your Turn: How can we reverse the above string?
```
more_unicode[::-1] # reverses the string
```
# String Operations
### String Concatenation
```
'Hello 😊' + ' and welcome ' + 'to Python 🐍!'
'Hello 😊' ' and welcome ' 'to Python 🐍!'
s3 = ('This '
'is another way '
'to concatenate '
'several strings!')
s3
```
### Substring check
```
'way' in s3
'python' not in s3
```
### String Length
```
len(s3)
```
# String Indexing and Slicing
```
# creating a string
s = 'PYTHON'
s, type(s)
```
## String Indexing
```
# depicting string indexes
for index, character in enumerate(s):
print('Character ->', character, 'has index->', index)
s[0], s[1], s[2], s[3], s[4], s[5]
s[-1], s[-2], s[-3], s[-4], s[-5], s[-6]
```
## String Slicing
```
s[:]
s[1:4]
s[:3], s[3:]
```
## String slicing with offsets
```
s[::1] # no offset
s[::2] # print every 2nd character in string
```
# String Immutability
```
# strings are immutable hence assignment throws error
s[0] = 'X'
print('Original String id:', id(s))
# creates a new string
s = 'X' + s[1:]
print(s)
print('New String id:', id(s))
```
# Useful String methods
## Case Conversions
```
s = 'python is great'
s.capitalize()
s.upper()
s.title()
```
## String Replace
```
s.replace('python', 'NLP')
```
## Numeric Checks
```
'12345'.isdecimal()
'apollo11'.isdecimal()
```
## Alphabet Checks
```
'python'.isalpha()
'number1'.isalpha()
```
## Alphanumeric Checks
```
'total'.isalnum()
'abc123'.isalnum()
'1+1'.isalnum()
```
## String splitting and joining
```
s = 'I,am,a,comma,separated,string'
s
s.split(',')
' '.join(s.split(','))
# stripping whitespace characters
s = ' I am surrounded by spaces '
s
s.strip()
sentences = 'Python is great. NLP is also good.'
sentences.split('.')
```
# String formatting
## Formatting expressions with different data types - old style
```
'We have %d %s containing %.2f gallons of %s' %(2, 'bottles', 2.5, 'milk')
'We have %d %s containing %.2f gallons of %s' %(5.21, 'jugs', 10.86763, 'juice')
```
## Formatting strings using the format method - new style
```
'Hello {} {}, it is a great {} to meet you at {}'.format('Mr.', 'Jones', 'pleasure', 5)
'Hello {} {}, it is a great {} to meet you at {} o\' clock'.format('Sir', 'Arthur', 'honor', 9)
```
## Alternative ways of using string format
```
'I have a {food_item} and a {drink_item} with me'.format(drink_item='soda', food_item='sandwich')
'The {animal} has the following attributes: {attributes}'.format(animal='dog', attributes=['lazy', 'loyal'])
```
# Regular Expressions
```
s1 = 'Python is an excellent language'
s2 = 'I love the Python language. I also use Python to build applications at work!'
import re
pattern = 'python'
# match only returns a match if regex match is found at the beginning of the string
re.match(pattern, s1)
# pattern is in lower case hence ignore case flag helps
# in matching same pattern with different cases
re.match(pattern, s1, flags=re.IGNORECASE)
# printing matched string and its indices in the original string
m = re.match(pattern, s1, flags=re.IGNORECASE)
print('Found match {} ranging from index {} - {} in the string "{}"'.format(m.group(0),
m.start(),
m.end(), s1))
# match does not work when pattern is not there in the beginning of string s2
re.match(pattern, s2, re.IGNORECASE)
# illustrating find and search methods using the re module
re.search(pattern, s2, re.IGNORECASE)
re.findall(pattern, s2, re.IGNORECASE)
match_objs = re.finditer(pattern, s2, re.IGNORECASE)
match_objs
print("String:", s2)
for m in match_objs:
print('Found match "{}" ranging from index {} - {}'.format(m.group(0),
m.start(), m.end()))
# illustrating pattern substitution using sub and subn methods
re.sub(pattern, 'Java', s2, flags=re.IGNORECASE)
re.subn(pattern, 'Java', s2, flags=re.IGNORECASE)
# dealing with unicode matching using regexes
s = u'H\u00e8llo! this is Python 🐍'
s
re.findall(r'\w+', s)
re.findall(r"[A-Z]\w+", s)
emoji_pattern = r"['\U0001F300-\U0001F5FF'|'\U0001F600-\U0001F64F'|'\U0001F680-\U0001F6FF'|'\u2600-\u26FF\u2700-\u27BF']"
re.findall(emoji_pattern, s, re.UNICODE)
```
| github_jupyter |
# Passing ancillary information to evaluate()
This notebook explores passing anciallary information for evaluating individuals.
For example, each `leap_ec.distributed.individual.DistributedIndividual` supports adding a UUID for each newly created individual. This UUID string can be used to create a file or subdirectory name that can be later associated with that individual. For example, a deep-learner model can use that as a file name, or a file containing frames for a driving animation can be stored in a subdirectory that uses that UUID as a name. However, `Problem.evaluate()` has no direct support for passing that UUID to support either of those scenarios, or any other similar ancillary information.
This notebook poses two solutions: one where `*args, **kwargs` is added to `Problem.evluate()` and the other where a special `Decoder` that adds that ancillary information is given.
```
import random
from math import nan
from pprint import pformat
import leap_ec
from leap_ec.decoder import IdentityDecoder, Decoder
from leap_ec.problem import Problem, ScalarProblem
from leap_ec.distributed.individual import DistributedIndividual
```
## Tailoring an `Individual` and `Problem` to accept ancillary evaluation information
First we'll demonstrate extending `Individual` and tailoring a corresponding `Problem` subclass to allow for passing ancially information to `Problem.evaluate()`.
```
# Define a problem class that uses UUID during evaluation
class UUIDProblem(Problem):
def __init__(self):
super().__init__()
def evaluate(self, phenome, *args, **kwargs):
print(f'UUIDProblem.evaluate(), phenome: {str(phenome)}, uuid: {str(kwargs["uuid"])}')
# Just return a random number because we only care about ensuring the UUID made it to evaluate
return random.random()
def worse_than(self, first_fitness, second_fitness):
""" have to define because abstract method """
pass # for this example we don't need to implement anything here
def equivalent(self, first_fitness, second_fitness):
""" have to define because abstract method """
pass # for this example we don't need to implement anything here
# We need to subclass DistributedIndividual to ensure that it passes the UUID for evaluations;
# DistributedIndividual implicitly tags each newly created individual with a UUID, which is
# why we're using it for this example instead of Individual.
class MyDistributedIndividual(DistributedIndividual):
def __init__(self,genome, decoder=None, problem=None):
super().__init__(genome, decoder, problem) # superclass will also set self.uuid
def evaluate_imp(self):
""" We override Individual.evaluate_imp() to pass in the UUID
"""
return self.problem.evaluate(self.decode(), uuid=self.uuid)
# Now test this scheme out by creating an example individual
ind = MyDistributedIndividual([], decoder=IdentityDecoder(), problem=UUIDProblem())
fitness = ind.evaluate()
print(f'ind uuid: {ind.uuid}, ind.fitness: {ind.fitness}')
```
Since we had full control over `UUIDProblem.evaluate()` we could have explicitly added a `uuid` keyword parameter, especially since our tailored `MyDistributedIndividaul.evaluate()` was going to be the only `evaluate()` to call it. However, this did have a disadvantage in that we had to faithfully copy over the original `Individual.evaluate()` and hack to fit to ensure the exception handling semantics made it over.
## Extending a `Decoder` to contain ancillary data to pass to a tailored `Problem`
The next approach entails extending the `phenome`, itself, to contain the desired ancillary data. That is, `Problem.evalute()` accepts a `phenome` parameter, not a `genome` as happens with other, extand EA toolkits, and we can piggyback that ancillary data in the phenome. During the evaluation an individual's genome is decoded into a phenome meaningful to a given `Problem` instance. Of course for representations that are already phenotypic, such as real-valued vectors, the decoder can be `IdentifyDecoder` that just faithfully passes along the `genome` as the `phenome` without change.
Though there is generally a one-to-one mapping between the genome and phenome, it needn't be that way. That is, the phenotypic space can contain information not found in the genome. For example, alleles will dictate our hair color, but we can still dye hair a different color. So, extending the phenome to include additional information is a more "evolutionary correct" approach to using ancillary information.
In our test implementation we modify the `Decoder` abstract base class to have `decode` accept arguments for what individual attributes, such as the UUID, we want to pass to `Problem.evaluate()`. We once again use the familiar `*args, **kwargs` pattern.
(This is what changed in the base implementation.)
```
class Decoder(abc.ABC):
@abc.abstractmethod
def decode(self, genome, *args, **kwargs): # added optional args and kwargs
pass
```
So, now to use this modified version of the decoder.
```
# Define a problem class that uses UUID during evaluation
class DifferentUUIDProblem(ScalarProblem):
def __init__(self):
super().__init__(maximize=True)
def evaluate(self, phenome):
print(f'DifferentUUIDProblem.evaluate(), phenome: {pformat(repr(phenome))}')
# Just return a random number because we only care about ensuring the UUID made it to evaluate
return random.random()
class ExtraIdentityDecoder(Decoder):
def __init__(self):
super().__init__()
def decode(self, genome, *args, **kwargs):
print(f'decode(): genome: {genome}, args: {args}, kwargs: {kwargs}')
return {'phenome' : genome, 'args' : args, 'kwargs' : kwargs}
def __repr__(self):
return type(self).__name__ + "()"
class MyOtherDistributedIndividual(DistributedIndividual):
def __init__(self,genome, decoder=None, problem=None):
super().__init__(genome, decoder, problem) # superclass will also set self.uuid
def decode(self, *args, **kwargs): # Have to extend this, too
return self.decoder.decode(self.genome, args, kwargs)
def evaluate_imp(self):
""" Once again, we have to override this to tailor the call to the decoder.
"""
phenome = self.decode(self.genome, uuid=self.uuid)
return self.problem.evaluate(phenome)
# Now to test it out.
ind = MyOtherDistributedIndividual([], decoder=ExtraIdentityDecoder(), problem=DifferentUUIDProblem())
fitness = ind.evaluate()
print(f'ind uuid: {ind.uuid}, ind.fitness: {ind.fitness}')
```
| github_jupyter |
## Read CSVs
```
!ls
# to download the data go to https://drive.google.com/file/d/111xy2JSJ2sE-zQ2dLhOvntX1mt46EepY/view
import pandas as pd
t_file = pd.read_csv("csv_files/t_file.csv")
t_people = pd.read_csv("csv_files/t_people.csv")
t_change = pd.read_csv("csv_files/t_change.csv")
t_history = pd.read_csv("csv_files/t_history.csv")
t_revision = pd.read_csv("csv_files/t_revision.csv")
print("shape of t_file : {}".format(t_file.shape))
print("shape of t_people : {}".format(t_people.shape))
print("shape of t_change : {}".format(t_change.shape))
print("shape of t_history : {}".format(t_history.shape))
print("shape of t_revision : {}".format(t_revision.shape))
```
## Dtypes setup
```
# use pandas to_datetime function to convert the string datatype to datetime
t_change.ch_createdTime = pd.to_datetime(t_change.ch_createdTime)
t_change.ch_updatedTime = pd.to_datetime(t_change.ch_updatedTime)
```
## Import CSV into local in-memory sqlite server
```
# use sqlalchemy to create a local sqlite engine and connect it to the notebook.
from sqlalchemy import create_engine
engine = create_engine('sqlite://', echo=False)
engine.connect
# use pandas to_sql function to convert the dataframe into a sql table
t_file.to_sql("t_file", con= engine, if_exists='replace', index=False)
t_people.to_sql("t_people", con= engine, if_exists='replace', index=False)
t_change.to_sql("t_change", con= engine, if_exists='replace', index=False)
t_history.to_sql("t_history", con= engine, if_exists='replace', index=False)
t_revision.to_sql("t_revision", con= engine, if_exists='replace', index=False)
```
#### tables in the database
```
# get all the table names in the engine
engine.table_names()
```
#### Schema
```
# display the schema diagram
from IPython.display import Image
Image(url='https://image.slidesharecdn.com/msr16yang-160515235151/95/msr-2016-data-showcase-mining-code-review-repositories-11-638.jpg?cb=1463356751')
```
#### Query syntax
```
use pandas sql method to read sql queries on the engine.
pd.read_sql(
"""
SELECT * FROM t_revision
;
""", con= engine).head(2)
# note that the output is a pandas dataframe and can be object referenced
# t_revision_sql_result = pd.read_sql(
# """
# SELECT * FROM t_revision
# ;
# """, con= engine).head(2)
```
## What percentage of code reviews gets merged?
```
merged_ = pd.read_sql("""
SELECT
ch_project,
count(*) as total_changes,
sum(case when ch_status = 'MERGED' then 1 else 0 end) as Total_Merged,
sum(case when ch_status = 'ABANDONED' then 1 else 0 end) as Total_abandoned
from (SELECT distinct
ch_project
, ch_branch
, ch_topic
, ch_authorAccountId
, ch_createdTime
, ch_updatedTime
, ch_status
FROM t_change)
group by 1;
""", con = engine)
merged_['pct_merged'] = merged_['Total_Merged'] / merged_['total_changes'] * 100
merged_['pct_abandoned'] = merged_['Total_abandoned'] / merged_['total_changes'] * 100
import matplotlib.pyplot as plt
import seaborn as sns
merged_[merged_['total_changes'] > 20].pct_merged.hist(bins = 50
, color = 'b'
, figsize = (5,3)
, legend = True)
plt.show()
merged_[merged_['total_changes'] > 20].pct_abandoned.hist(bins = 50
, color = 'r'
, figsize = (5,3)
, legend = True)
plt.show()
```
### Result
```
print("Average percent merged " + str(round(merged_[merged_['total_changes'] > 20].pct_merged.mean(), 2)) + "%")
print("Median percent merged " + str(round(merged_[merged_['total_changes'] > 20].pct_merged.median(), 2)) + "%")
print("------------------------")
print("Average percent abandoned " + str(round(merged_[merged_['total_changes'] > 20].pct_abandoned.mean(), 2)) + "%")
print("Median percent abandoned " + str(round(merged_[merged_['total_changes'] > 20].pct_abandoned.median(), 2)) + "%")
```
## How long does it take to get a code review merged?
```
merged_time_ = pd.read_sql("""
SELECT
ch_project
, ch_branch
, ch_topic
, case when ch_status = 'MERGED' then min(ch_createdTime) else 0 end as first_ch_createdTime
, case when ch_status = 'MERGED' then max(ch_updatedTime) else 0 end as last_ch_updatedTime
, count(*) as total_changes
from (SELECT distinct
ch_project
, ch_branch
, ch_topic
, ch_authorAccountId
, ch_createdTime
, ch_updatedTime
, ch_status
FROM t_change)
group by 1, 2, 3;
""", con = engine)
merged_time_['first_ch_createdTime'] = pd.to_datetime(merged_time_.first_ch_createdTime)
merged_time_['last_ch_updatedTime'] = pd.to_datetime(merged_time_.last_ch_updatedTime)
merged_time_['time_to_merge'] = (merged_time_.last_ch_updatedTime - merged_time_.first_ch_createdTime).dt.total_seconds() / 60 / 60 / 24
merged_time_[merged_time_['total_changes'] > 5].time_to_merge.hist(bins = 5
, color = 'y'
, figsize = (5,3)
, legend = True)
plt.show()
```
### Result
### Where total project changes are <= 10
```
print("Average time to get merged "
+ str(round(merged_time_[merged_time_['total_changes'] <= 10].time_to_merge.mean(), 2))
+ " days")
print("Median time to get merged "
+ str(round(merged_time_[merged_time_['total_changes'] <= 10].time_to_merge.median(), 2))
+ " days")
```
### Where total project changes are > 10
```
print("Average time to get merged "
+ str(round(merged_time_[merged_time_['total_changes'] > 10].time_to_merge.mean(), 2))
+ " days")
print("Median time to get merged "
+ str(round(merged_time_[merged_time_['total_changes'] > 10].time_to_merge.median(), 2))
+ " days")
```
| github_jupyter |
<a href="https://colab.research.google.com/github/Meng-MiamiOH/MTH231/blob/main/exercise_machine_learning_and_coding.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Exercise for Machine Learning and Coding Session
For this exercise, you will be using the Breast Cancer Wisconsin (Diagnostic) Database to create classifiers that can help diagnose patients. First, read through the description of the dataset (below).
```
import numpy as np
import pandas as pd
from sklearn.datasets import load_breast_cancer
cancer = load_breast_cancer()
print(cancer.DESCR) # Print the data set description
```
The object returned by load_breast_cancer() is a scikit-learn Bunch object, which is similar to a dictionary.
```
cancer.keys()
```
### 1. Convert the sklearn.dataset `cancer['data']` to a `(569, 30)` DataFrame `X` and convert the `cancer['target']` to a `(569, 1)` Series `y`.
```
X = pd.DataFrame(data=cancer['data'], columns=cancer['feature_names'])
y = pd.Series(cancer['target'])
print(X)
print(y)
```
### 2. What is the class distribution? (i.e. how many instances of `malignant` (encoded 0) and how many `benign` (encoded 1)?)
```
len(y[y == 0])
len(y[y == 1])
```
### 4. Using `train_test_split`, split `X` and `y` into training and test sets `X_train`, `X_test`, `y_train`, and `y_test`.
* `X_train` *has shape* `(426, 30)`
* `X_test` *has shape* `(143, 30)`
* `y_train` *has shape* `(426,)`
* `y_test` *has shape* `(143,)`
```
# your code
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y)
```
### 5. Using KNeighborsClassifier, fit a k-nearest neighbors (knn) classifier with `X_train`, `y_train` and using one nearest neighbor (`n_neighbors = 1`).
```
# your code
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=1)
knn.fit(X_train,y_train)
```
### 6. Using your knn classifier, predict the class label using the mean value for each feature.
```
mean = X_train.mean().values.reshape(1, -1)
mean
# your code
knn.predict(mean)
```
### 7. Use the knn classifer your created for Question 5 to predict the class labels for the test set `X_text`.
*The result should be a numpy array with shape `(143,)` and values either `0.0` or `1.0`.*
```
# your code
knn.predict(X_test)
print(y_test)
```
### 8. Find the score (mean accuracy) of your knn classifier using `X_test` and `y_test`.
*The calculated score should be a float between 0 and 1*
```
# your code
knn.score(X_test,y_test)
```
### 9. Try to train kNN classifier with different k (1, 3, 5, 10) and compare the performance.
```
# your code
knn = KNeighborsClassifier(n_neighbors=1)
knn.fit(X_train,y_train)
print(knn.score(X_test,y_test))
knn = KNeighborsClassifier(n_neighbors=3)
knn.fit(X_train,y_train)
print(knn.score(X_test,y_test))
knn = KNeighborsClassifier(n_neighbors=5)
knn.fit(X_train,y_train)
print(knn.score(X_test,y_test))
knn = KNeighborsClassifier(n_neighbors=10)
knn.fit(X_train,y_train)
print(knn.score(X_test,y_test))
```
### 10. Use logistic regression model to create a classifier for the data.
```
# your code
from sklearn.linear_model import LogisticRegression
lc = LogisticRegression().fit(X_train, y_train)
```
### 11. Evaluate the logistic classifier and compare it with kNN classifier.
```
# your code
lc.score(X_test,y_test)
```
### 12. Use cross-validation to calculate accuracy
```
import numpy as ny
# your code
```
### 13. Use cross-validation to calculate f1
```
# your code
```
| github_jupyter |
```
import pandas as pd
listings = pd.read_csv('/Users/mattmastin/Desktop/airbnb/listings.csv')
listings_summary = pd.read_csv('/Users/mattmastin/Desktop/airbnb/listings_summary.csv')
hoods = pd.read_csv('/Users/mattmastin/Desktop/airbnb/neighbourhoods.csv')
reviews_summary = pd.read_csv('/Users/mattmastin/Desktop/airbnb/reviews_summary.csv')
print(listings.shape)
listings.head()
listings.describe(exclude='number')
pd.set_option('display.max_columns', 500)
print(listings_summary.shape)
listings_summary.head()
listings_summary.describe(exclude='number')
pd.set_option('display.max_rows', 500)
listings_summary.isnull().sum()
print(hoods.shape)
hoods.head()
listings.isnull().sum()
listings_summary.describe()
listings_summary = listings_summary.drop(columns=['id', 'host_name', 'latitude', 'longitude', 'room type', 'price'])
df = pd.concat([listings, listings_summary], axis=1)
print(df.shape)
df.head()
listings_summary.isnull().sum()
df = listings_summmary.drop(['space', 'notes', 'access', 'interaction', 'house_rules',
'thumbnail_url', 'medium_url', 'xl_picture_url', 'host_about',
'host_response_time', 'host_response_rate', 'host_acceptance_rate',
'host_neighbourhood', 'host_listings_count', 'host_total_listings_count',
'host_verifications', 'host_identity_verified', 'is_location_exact',
'first_review', 'last_review'], axis=1)
df.isnull().sum()
df.shape
df.head()
df = df.drop(columns=['listing_url', 'experiences_offered', 'picture_url', 'host_name',
'host_location', 'host_is_superhost', 'neighbourhood_cleansed',
'city', 'market', 'instant_bookable', 'is_business_travel_ready',
'require_guest_phone_verification', 'smart_location',
'calendar_updated', 'has_availability', 'availability_30',
'availability_60', 'availability_90', 'availability_365',
'calculated_host_listings_count', 'review_scores_cleanliness',
'review_scores_checkin', 'review_scores_communication'], axis=1)
print(df.shape)
df.head()
df = df.drop(columns=['host_since', 'street'], axis=1)
df['price'] = df['price'].str.replace('$', '')
df['price'] = df['price'].str.replace(',', '')
df['price'] = df['price'].astype(float)
df['security_deposit'] = df['security_deposit'].str.replace('$', '')
df['security_deposit'] = df['security_deposit'].str.replace(',', '')
df['security_deposit'] = df['security_deposit'].astype(float)
df['cleaning_fee'] = df['cleaning_fee'].str.replace('$', '')
df['cleaning_fee'] = df['cleaning_fee'].str.replace(',', '')
df['cleaning_fee'] = df['cleaning_fee'].astype(float)
df.head()
df.shape
df['extra_people'] = df['extra_people'].str.replace('$', '')
df['extra_people'] = df['extra_people'].str.replace(',', '')
df['extra_people'] = df['extra_people'].astype(float)
df.dtypes
df.isnull().sum()
df['security_deposit'] = df['security_deposit'].fillna(0)
df['cleaning_fee'] = df['cleaning_fee'].fillna(0)
df.describe(exclude='number')
df = df.drop(columns=['summary'], axis=1)
df.head()
df = df.drop(columns=['neighbourhood'], axis=1)
df.describe(exclude='number')
df.shape
df.to_csv('listings_summary_clean.csv', index=False)
df2 = pd.read_csv('listings_summary_clean.csv')
print(df2.shape)
df2.head()
```
| github_jupyter |
This notebook was prepared by [Donne Martin](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/system-design-primer).
# Design a call center
## Constraints and assumptions
* What levels of employees are in the call center?
* Operator, supervisor, director
* Can we assume operators always get the initial calls?
* Yes
* If there is no available operators or the operator can't handle the call, does the call go to the supervisors?
* Yes
* If there is no available supervisors or the supervisor can't handle the call, does the call go to the directors?
* Yes
* Can we assume the directors can handle all calls?
* Yes
* What happens if nobody can answer the call?
* It gets queued
* Do we need to handle 'VIP' calls where we put someone to the front of the line?
* No
* Can we assume inputs are valid or do we have to validate them?
* Assume they're valid
## Solution
```
%%writefile call_center.py
from abc import ABCMeta, abstractmethod
from collections import deque
from enum import Enum
class Rank(Enum):
OPERATOR = 0
SUPERVISOR = 1
DIRECTOR = 2
class Employee(metaclass=ABCMeta):
def __init__(self, employee_id, name, rank, call_center):
self.employee_id = employee_id
self.name = name
self.rank = rank
self.call = None
self.call_center = call_center
def take_call(self, call):
"""Assume the employee will always successfully take the call."""
self.call = call
self.call.employee = self
self.call.state = CallState.IN_PROGRESS
def complete_call(self):
self.call.state = CallState.COMPLETE
self.call_center.notify_call_completed(self.call)
@abstractmethod
def escalate_call(self):
pass
def _escalate_call(self):
self.call.state = CallState.READY
call = self.call
self.call = None
self.call_center.notify_call_escalated(call)
class Operator(Employee):
def __init__(self, employee_id, name):
super(Operator, self).__init__(employee_id, name, Rank.OPERATOR)
def escalate_call(self):
self.call.level = Rank.SUPERVISOR
self._escalate_call()
class Supervisor(Employee):
def __init__(self, employee_id, name):
super(Operator, self).__init__(employee_id, name, Rank.SUPERVISOR)
def escalate_call(self):
self.call.level = Rank.DIRECTOR
self._escalate_call()
class Director(Employee):
def __init__(self, employee_id, name):
super(Operator, self).__init__(employee_id, name, Rank.DIRECTOR)
def escalate_call(self):
raise NotImplemented('Directors must be able to handle any call')
class CallState(Enum):
READY = 0
IN_PROGRESS = 1
COMPLETE = 2
class Call(object):
def __init__(self, rank):
self.state = CallState.READY
self.rank = rank
self.employee = None
class CallCenter(object):
def __init__(self, operators, supervisors, directors):
self.operators = operators
self.supervisors = supervisors
self.directors = directors
self.queued_calls = deque()
def dispatch_call(self, call):
if call.rank not in (Rank.OPERATOR, Rank.SUPERVISOR, Rank.DIRECTOR):
raise ValueError('Invalid call rank: {}'.format(call.rank))
employee = None
if call.rank == Rank.OPERATOR:
employee = self._dispatch_call(call, self.operators)
if call.rank == Rank.SUPERVISOR or employee is None:
employee = self._dispatch_call(call, self.supervisors)
if call.rank == Rank.DIRECTOR or employee is None:
employee = self._dispatch_call(call, self.directors)
if employee is None:
self.queued_calls.append(call)
def _dispatch_call(self, call, employees):
for employee in employees:
if employee.call is None:
employee.take_call(call)
return employee
return None
def notify_call_escalated(self, call): # ...
def notify_call_completed(self, call): # ...
def dispatch_queued_call_to_newly_freed_employee(self, call, employee): # ...
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.