code stringlengths 2.5k 150k | kind stringclasses 1 value |
|---|---|
```
import warnings
warnings.simplefilter('ignore', FutureWarning)
import matplotlib
matplotlib.rcParams['axes.grid'] = True # show gridlines by default
%matplotlib inline
import pandas as pd
```
## Getting Comtrade data into your notebook
In this exercise, you will practice loading data from Comtrade into a pandas dataframe and getting it into a form where you can start to work with it.
The following steps and code are an example. Your task for this exercise is stated at the end, after the example.
The data is obtained from the [United Nations Comtrade](http://comtrade.un.org/data/) website, by selecting the following configuration:
- Type of Product: goods
- Frequency: monthly
- Periods: all of 2020
- Reporter: Kenya
- Partners: all
- Flows: imports and exports
- HS (as reported) commodity codes: 0401 (Milk and cream, neither concentrated nor sweetened) and 0402 (Milk and cream, concentrated or sweetened)
Clicking on 'Preview' results in a message that the data exceeds 500 rows. Data was downloaded using the *Download CSV* button and the download file renamed appropriately.
```
LOCATION ='comtrade_milk_kenya_monthly_2020.csv'
```
Load the data in from the specified location, ensuring that the various codes are read as strings. Preview the first few rows of the dataset.
```
milk = pd.read_csv(LOCATION, dtype={'Commodity Code':str, 'Reporter Code':str})
milk.head(5)
milk.tail(5)
#limit the columns
COLUMNS = ['Year', 'Period','Trade Flow','Reporter', 'Partner', 'Commodity','Commodity Code','Trade Value (US$)']
milk = milk[COLUMNS]
milk
```
Derive two new dataframes that separate out the 'World' partner data and the data for individual partner countries.
```
milk_world = milk[milk['Partner'] == 'World']
milk_countries = milk[milk['Partner'] != 'World']
#store as csv
milk_countries.to_csv('kenyamilk.csv',index=False)
```
To load the data back in:
```
load_test= pd.read_csv('kenyamilk.csv', dtype={'Commodity Code':str,'Reporter Code':str})
load_test.head(3)
```
### Subsetting Your Data
For large or heterogenous datasets, it is often convenient to create subsets of the data. To further separate out the imports:
```
milk_imports = milk[milk['Trade Flow'] == 'Imports']
milk_countries_imports = milk_countries[milk_countries['Trade Flow'] == 'Imports']
milk_world_imports=milk_world[milk_world['Trade Flow'] == 'Imports']
```
### Sorting the data
Having loaded in the data, find the most valuable partners in terms of import trade flow during a particular month by sorting the data by *decreasing* trade value and then selecting the top few rows.
```
milkImportsInJanuary2020 = milk_countries_imports[milk_countries_imports['Period'] == 202001]
milkImportsInJanuary2020.sort_values('Trade Value (US$)',ascending=False).head(10)
```
### Grouping the data
Split the data into two different subsets of data (imports and exports), by grouping on trade flow.
```
groups = milk_countries.groupby('Trade Flow')
groups.get_group('Imports').head()
```
As well as grouping on a single term, you can create groups based on multiple columns by passing in several column names as a list. For example, generate groups based on commodity code and trade flow, and then preview the keys used to define the groups
```
GROUPING_COMMFLOW = ['Commodity Code','Trade Flow']
groups = milk_countries.groupby(GROUPING_COMMFLOW)
groups.groups.keys()
```
Retrieve a group based on multiple group levels by passing in a tuple that specifies a value for each index column. For example, if a grouping is based on the 'Partner' and 'Trade Flow' columns, the argument of get_group has to be a partner/flow pair, like ('Uganda', 'Import') to get all rows associated with imports from Uganda.
```
GROUPING_PARTNERFLOW = ['Partner','Trade Flow']
groups = milk_countries.groupby(GROUPING_PARTNERFLOW)
GROUP_PARTNERFLOW= ('Uganda','Imports')
groups.get_group( GROUP_PARTNERFLOW )
```
To find the leading partner for a particular commodity, group by commodity, get the desired group, and then sort the result.
```
groups = milk_countries.groupby(['Commodity Code'])
groups.get_group('0402').sort_values("Trade Value (US$)", ascending=False).head()
```
| github_jupyter |
```
import requests
import json
url = "https://microsoft-computer-vision3.p.rapidapi.com/analyze"
querystring = {"language":"en","descriptionExclude":"Celebrities","visualFeatures":"ImageType,Categories,Description","details":"Celebrities"}
payload = "{\r\n \"url\": \"https://neilpatel.com/wp-content/uploads/2017/09/image-editing-tools.jpg\"\r\n}"
headers = {
'content-type': "application/json",
'x-rapidapi-key': "dacaae5850mshbcab4ca9a7b2a4dp13a15bjsnd9aaa23a2ee4",
'x-rapidapi-host': "microsoft-computer-vision3.p.rapidapi.com"
}
response = requests.request("POST", url, data=payload, headers=headers, params=querystring)
result = response.text
print(result)
type(response)
type(result)
data = json.loads(result)
print(data)
type(data)
data['description']['captions'][0]['text']
test = {
"book": [
{
"id":"01",
"language": "Java",
"edition": "third",
"author": "Herbert Schildt"
},
{
"id":"07",
"language": "C++",
"edition": "second",
"author": "E.Balagurusamy"
}
]
}
print(test)
type(test)
winner_record = {'marks':97,'name':'Winner','distinction':True}
winner_record['distinction']
records = {
'Maths':[
{'marks':97,'name':'Winner','distinction':True},
{'marks':99,'name':'Emeto','distinction':False}
],
'English':[
{'marks':97,'name':'Winner','distinction':True},
{'marks':99,'name':'Emeto','distinction':False}
]
}
age = [22,'Winner',2,76,12,16]
records['Maths'][1]['distinction']
for element in records['Maths']:
if element['name'] == 'Emeto':
print(element['distinction'])
else:
print('wrong record!')
{
"$schema": "http://json-schema.org/draft-04/schema#",
"title": "Product",
"description": "A product from Acme's catalog",
"type": "object",
"properties": {
"id": {
"description": "The unique identifier for a product",
"type": "integer"
},
"name": {
"description": "Name of the product",
"type": "string"
},
"price": {
"type": "number",
"minimum": 0,
"exclusiveMinimum": true
}
},
"required": ["id", "name", "price"]
}
'Maths':[
{'marks':97,'name':'Winner','distinction':True},
{'marks':99,'name':'Emeto','distinction':False}
[
{
"id": 2,
"name": "An ice sculpture",
"price": 12.50,
},
{
"id": 3,
"name": "A blue mouse",
"price": 25.50,
}
]
d = {}
d['Name'] = 'Winner Emeto'
d['Country'] = 'Nigeria'
var = json.dumps(d,ensure_ascii=False)
print(var)
type(var)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/unburied/DS-Unit-1-Sprint-3-Statistical-Tests-and-Experiments/blob/master/LS_DS_142_Sampling_Confidence_Intervals_and_Hypothesis_Testing.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Lambda School Data Science Module 142
## Sampling, Confidence Intervals, and Hypothesis Testing
## Prepare - examine other available hypothesis tests
If you had to pick a single hypothesis test in your toolbox, t-test would probably be the best choice - but the good news is you don't have to pick just one! Here's some of the others to be aware of:
```
import numpy as np
from scipy.stats import chisquare # One-way chi square test
# Chi square can take any crosstab/table and test the independence of rows/cols
# The null hypothesis is that the rows/cols are independent -> low chi square
# The alternative is that there is a dependence -> high chi square
# Be aware! Chi square does *not* tell you direction/causation
ind_obs = np.array([[1, 1], [2, 2]]).T
print(ind_obs)
print(chisquare(ind_obs, axis=None))
dep_obs = np.array([[16, 18, 16, 14, 12, 12], [32, 24, 16, 28, 20, 24]]).T
print(dep_obs)
print(chisquare(dep_obs, axis=None))
# Distribution tests:
# We often assume that something is normal, but it can be important to *check*
# For example, later on with predictive modeling, a typical assumption is that
# residuals (prediction errors) are normal - checking is a good diagnostic
from scipy.stats import normaltest
# Poisson models arrival times and is related to the binomial (coinflip)
sample = np.random.poisson(5, 1000)
print(normaltest(sample)) # Pretty clearly not normal
# Kruskal-Wallis H-test - compare the median rank between 2+ groups
# Can be applied to ranking decisions/outcomes/recommendations
# The underlying math comes from chi-square distribution, and is best for n>5
from scipy.stats import kruskal
x1 = [1, 3, 5, 7, 9]
y1 = [2, 4, 6, 8, 10]
print(kruskal(x1, y1)) # x1 is a little better, but not "significantly" so
x2 = [1, 1, 1]
y2 = [2, 2, 2]
z = [2, 2] # Hey, a third group, and of different size!
print(kruskal(x2, y2, z)) # x clearly dominates
```
And there's many more! `scipy.stats` is fairly comprehensive, though there are even more available if you delve into the extended world of statistics packages. As tests get increasingly obscure and specialized, the importance of knowing them by heart becomes small - but being able to look them up and figure them out when they *are* relevant is still important.
## Live Lecture - let's explore some more of scipy.stats
Candidate topics to explore:
- `scipy.stats.chi2` - the Chi-squared distribution, which we can use to reproduce the Chi-squared test
- Calculate the Chi-Squared test statistic "by hand" (with code), and feed it into `chi2`
- Build a confidence interval with `stats.t.ppf`, the t-distribution percentile point function (the inverse of the CDF) - we can write a function to return a tuple of `(mean, lower bound, upper bound)` that you can then use for the assignment (visualizing confidence intervals)
```
# Taking requests! Come to lecture with a topic or problem and we'll try it.
```
## Assignment - Build a confidence interval
A confidence interval refers to a neighborhood around some point estimate, the size of which is determined by the desired p-value. For instance, we might say that 52% of Americans prefer tacos to burritos, with a 95% confidence interval of +/- 5%.
52% (0.52) is the point estimate, and +/- 5% (the interval $[0.47, 0.57]$) is the confidence interval. "95% confidence" means a p-value $\leq 1 - 0.95 = 0.05$.
In this case, the confidence interval includes $0.5$ - which is the natural null hypothesis (that half of Americans prefer tacos and half burritos, thus there is no clear favorite). So in this case, we could use the confidence interval to report that we've failed to reject the null hypothesis.
But providing the full analysis with a confidence interval, including a graphical representation of it, can be a helpful and powerful way to tell your story. Done well, it is also more intuitive to a layperson than simply saying "fail to reject the null hypothesis" - it shows that in fact the data does *not* give a single clear result (the point estimate) but a whole range of possibilities.
How is a confidence interval built, and how should it be interpreted? It does *not* mean that 95% of the data lies in that interval - instead, the frequentist interpretation is "if we were to repeat this experiment 100 times, we would expect the average result to lie in this interval ~95 times."
For a 95% confidence interval and a normal(-ish) distribution, you can simply remember that +/-2 standard deviations contains 95% of the probability mass, and so the 95% confidence interval based on a given sample is centered at the mean (point estimate) and has a range of +/- 2 (or technically 1.96) standard deviations.
Different distributions/assumptions (90% confidence, 99% confidence) will require different math, but the overall process and interpretation (with a frequentist approach) will be the same.
Your assignment - using the data from the prior module ([congressional voting records](https://archive.ics.uci.edu/ml/datasets/Congressional+Voting+Records)):
1. Generate and numerically represent a confidence interval
2. Graphically (with a plot) represent the confidence interval
3. Interpret the confidence interval - what does it tell you about the data and its distribution?
Stretch goals:
1. Write a summary of your findings, mixing prose and math/code/results. *Note* - yes, this is by definition a political topic. It is challenging but important to keep your writing voice *neutral* and stick to the facts of the data. Data science often involves considering controversial issues, so it's important to be sensitive about them (especially if you want to publish).
2. Apply the techniques you learned today to your project data or other data of your choice, and write/discuss your findings here.
3. Refactor your code so it is elegant, readable, and can be easily run for all issues.
## Resources
- [Interactive visualize the Chi-Squared test](https://homepage.divms.uiowa.edu/~mbognar/applets/chisq.html)
- [Calculation of Chi-Squared test statistic](https://en.wikipedia.org/wiki/Pearson%27s_chi-squared_test)
- [Visualization of a confidence interval generated by R code](https://commons.wikimedia.org/wiki/File:Confidence-interval.svg)
- [Expected value of a squared standard normal](https://math.stackexchange.com/questions/264061/expected-value-calculation-for-squared-normal-distribution) (it's 1 - which is why the expected value of a Chi-Squared with $n$ degrees of freedom is $n$, as it's the sum of $n$ squared standard normals)
```
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
columns = ['party', 'handicapped_infants', 'water_project_cost_sharing',
'adoption_of_the_budget_resolution', 'physician_fee_freeze',
'el_salvador_aid', 'religious_groups_in_schools',
'anti_satellite_test_ban', 'aid_to_nicaraguan_contras',
'mx_missile', 'immigration', 'synfuels_corporation_cutback',
'education_spending', 'superfund_right_to_sue' , 'crime' ,
'duty_free_exports', 'export_administration_act_south_africa']
house_votes = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/voting-records/house-votes-84.data',
header = None, names = columns)
house_votes. head()
#return number of vote to fillNA based on party and feature
def vote_counts(party, feature):
#get series of based off party and feature
subset = house_votes[house_votes.party == party]
#divide values from the series
values = subset[feature].value_counts().to_dict()
yays = values['y']
nays = values['n']
abstained = values['?']
#convert above values to create NA replacement values
#based on ratio of current votes
vote_yay = int((yays / (yays + nays)) * abstained)
vote_nay = int((nays / (yays + nays)) * abstained)
#ensure new values equal current NA sum
if (vote_yay - vote_nay) > 0:
while (vote_yay + vote_nay) < abstained:
vote_yay += 1
else:
while (vote_yay + vote_nay) < abstained:
vote_nay += 1
return vote_yay, vote_nay
#Assign NA values based on the ratio in party votes
def clean(features):
#get vote counts to replace NAN values
r_yays, r_nays = vote_counts('republican', features)
d_yays, d_nays = vote_counts('democrat' , features)
#filter down to '?' for current feature and replace the top number of rows
#based on vote counts. Check to ensure vote counts are greater than zero
if r_yays > 0:
(house_votes.loc[(house_votes.party == 'republican') &
(house_votes[features] == '?'), features])[:r_yays] = 'y'
(house_votes.loc[(house_votes.party == 'republican') &
(house_votes[features] == '?'), features]) = 'n'
else:
(house_votes.loc[(house_votes.party == 'republican') &
(house_votes[features] == '?'), features])[:r_nays] = 'n'
(house_votes.loc[(house_votes.party == 'republican') &
(house_votes[features] == '?'), features]) = 'y'
if d_yays > 0:
(house_votes.loc[(house_votes.party == 'democrat') &
(house_votes[features] == '?'), features])[:d_yays] = 'y'
(house_votes.loc[(house_votes.party == 'democrat') &
(house_votes[features] == '?'), features]) = 'n'
else:
(house_votes.loc[(house_votes.party == 'democrat') &
(house_votes[features] == '?'), features])[:d_nays] = 'n'
(house_votes.loc[(house_votes.party == 'democrat') &
(house_votes[features] == '?'), features]) = 'y'
#CLean all '?' values in dataframe based on ratio of party votes
columns.pop(0)
for col in columns:
clean(col)
house_votes.head()
for col in columns:
house_votes[col] = np.where(house_votes[col] == 'y', 1, 0)
house_votes['crime'].value_counts()
from scipy.stats import ttest_ind, ttest_ind_from_stats, ttest_rel
#find a closely contested issue
for col in columns:
i = house_votes[col].value_counts()[0]
j = house_votes[col].value_counts()[1]
if abs(i - j) < 15:
print(col)
blues = house_votes[house_votes.party == 'democrat']['el_salvador_aid']
reds = house_votes[house_votes.party == 'republican']['el_salvador_aid']
stat, pval = ttest_ind(reds, blues, equal_var = False)
stat, pval
blues.value_counts(),reds.value_counts()
tidy = house_votes.copy()
for col in columns:
tidy[col] = np.where(tidy[col] == 1, 'y', 'n')
tidy.head()
table = pd.crosstab(tidy.party, columns = tidy['el_salvador_aid'], normalize = 'all')
table
ax = table.plot.bar(yerr = .05, color = ['b','r']);
ax.legend(loc = 10, labels = ['Yes', 'No'])
ax.set_title('Votes on El Salvador Aid with a confidence interval of 95%');
plt.xticks(rotation = 360);
```
| github_jupyter |
# Netflix Shows and Movies - Exploratory Analysis
The dataset consists of meta details about the movies and tv shows such as the title, director, and cast of the shows / movies. Details such as the release year, the rating, duration etc. As the first step, let's load the dataset, create some new features. In this kernel, I have analysed this dataset to find top insights and findings.
## Dataset Preparation
```
import plotly.graph_objects as go
from plotly.offline import init_notebook_mode, iplot
import pandas as pd
df = pd.read_csv("../input/netflix-shows/netflix_titles_nov_2019.csv")
## add new features in the dataset
df["date_added"] = pd.to_datetime(df['date_added'])
df['year_added'] = df['date_added'].dt.year
df['month_added'] = df['date_added'].dt.month
df['season_count'] = df.apply(lambda x : x['duration'].split(" ")[0] if "Season" in x['duration'] else "", axis = 1)
df['duration'] = df.apply(lambda x : x['duration'].split(" ")[0] if "Season" not in x['duration'] else "", axis = 1)
df.head()
```
## 1. Content Type on Netflix
```
col = "type"
grouped = df[col].value_counts().reset_index()
grouped = grouped.rename(columns = {col : "count", "index" : col})
## plot
trace = go.Pie(labels=grouped[col], values=grouped['count'], pull=[0.05, 0], marker=dict(colors=["#6ad49b", "#a678de"]))
layout = go.Layout(title="", height=400, legend=dict(x=0.1, y=1.1))
fig = go.Figure(data = [trace], layout = layout)
iplot(fig)
```
- 2/3rd of the content on netflix is movies and remaining 33% of them are TV Shows.
## 2. Growth in content over the years
```
d1 = df[df["type"] == "TV Show"]
d2 = df[df["type"] == "Movie"]
col = "year_added"
vc1 = d1[col].value_counts().reset_index()
vc1 = vc1.rename(columns = {col : "count", "index" : col})
vc1['percent'] = vc1['count'].apply(lambda x : 100*x/sum(vc1['count']))
vc1 = vc1.sort_values(col)
vc2 = d2[col].value_counts().reset_index()
vc2 = vc2.rename(columns = {col : "count", "index" : col})
vc2['percent'] = vc2['count'].apply(lambda x : 100*x/sum(vc2['count']))
vc2 = vc2.sort_values(col)
trace1 = go.Scatter(x=vc1[col], y=vc1["count"], name="TV Shows", marker=dict(color="#a678de"))
trace2 = go.Scatter(x=vc2[col], y=vc2["count"], name="Movies", marker=dict(color="#6ad49b"))
data = [trace1, trace2]
layout = go.Layout(title="Content added over the years", legend=dict(x=0.1, y=1.1, orientation="h"))
fig = go.Figure(data, layout=layout)
fig.show()
```
- The growth in number of movies on netflix is much higher than that od TV shows. About 1300 new movies were added in both 2018 and 2019. The growth in content started from 2013. Netflix kept on adding different movies and tv shows on its platform over the years. This content was of different variety - content from different countries, content which was released over the years.
## 3. Original Release Year of the movies
```
col = "release_year"
vc1 = d1[col].value_counts().reset_index()
vc1 = vc1.rename(columns = {col : "count", "index" : col})
vc1['percent'] = vc1['count'].apply(lambda x : 100*x/sum(vc1['count']))
vc1 = vc1.sort_values(col)
vc2 = d2[col].value_counts().reset_index()
vc2 = vc2.rename(columns = {col : "count", "index" : col})
vc2['percent'] = vc2['count'].apply(lambda x : 100*x/sum(vc2['count']))
vc2 = vc2.sort_values(col)
trace1 = go.Bar(x=vc1[col], y=vc1["count"], name="TV Shows", marker=dict(color="#a678de"))
trace2 = go.Bar(x=vc2[col], y=vc2["count"], name="Movies", marker=dict(color="#6ad49b"))
data = [trace1, trace2]
layout = go.Layout(title="Content added over the years", legend=dict(x=0.1, y=1.1, orientation="h"))
fig = go.Figure(data, layout=layout)
fig.show()
col = 'month_added'
vc1 = d1[col].value_counts().reset_index()
vc1 = vc1.rename(columns = {col : "count", "index" : col})
vc1['percent'] = vc1['count'].apply(lambda x : 100*x/sum(vc1['count']))
vc1 = vc1.sort_values(col)
trace1 = go.Bar(x=vc1[col], y=vc1["count"], name="TV Shows", marker=dict(color="#a678de"))
data = [trace1]
layout = go.Layout(title="In which month, the conent is added the most?", legend=dict(x=0.1, y=1.1, orientation="h"))
fig = go.Figure(data, layout=layout)
fig.show()
```
Some of the oldest movies on Netflix
```
small = df.sort_values("release_year", ascending = True)
small = small[small['duration'] != ""]
small[['title', "release_year"]][:15]
```
Some of the oldest TV Shows on Netflix
```
small = df.sort_values("release_year", ascending = True)
small = small[small['season_count'] != ""]
small[['title', "release_year"]][:15]
```
There are movies / shows on the platform which were released way back in 1930s and 40s.
## 4. Content from different Countries
```
country_codes = {'afghanistan': 'AFG',
'albania': 'ALB',
'algeria': 'DZA',
'american samoa': 'ASM',
'andorra': 'AND',
'angola': 'AGO',
'anguilla': 'AIA',
'antigua and barbuda': 'ATG',
'argentina': 'ARG',
'armenia': 'ARM',
'aruba': 'ABW',
'australia': 'AUS',
'austria': 'AUT',
'azerbaijan': 'AZE',
'bahamas': 'BHM',
'bahrain': 'BHR',
'bangladesh': 'BGD',
'barbados': 'BRB',
'belarus': 'BLR',
'belgium': 'BEL',
'belize': 'BLZ',
'benin': 'BEN',
'bermuda': 'BMU',
'bhutan': 'BTN',
'bolivia': 'BOL',
'bosnia and herzegovina': 'BIH',
'botswana': 'BWA',
'brazil': 'BRA',
'british virgin islands': 'VGB',
'brunei': 'BRN',
'bulgaria': 'BGR',
'burkina faso': 'BFA',
'burma': 'MMR',
'burundi': 'BDI',
'cabo verde': 'CPV',
'cambodia': 'KHM',
'cameroon': 'CMR',
'canada': 'CAN',
'cayman islands': 'CYM',
'central african republic': 'CAF',
'chad': 'TCD',
'chile': 'CHL',
'china': 'CHN',
'colombia': 'COL',
'comoros': 'COM',
'congo democratic': 'COD',
'Congo republic': 'COG',
'cook islands': 'COK',
'costa rica': 'CRI',
"cote d'ivoire": 'CIV',
'croatia': 'HRV',
'cuba': 'CUB',
'curacao': 'CUW',
'cyprus': 'CYP',
'czech republic': 'CZE',
'denmark': 'DNK',
'djibouti': 'DJI',
'dominica': 'DMA',
'dominican republic': 'DOM',
'ecuador': 'ECU',
'egypt': 'EGY',
'el salvador': 'SLV',
'equatorial guinea': 'GNQ',
'eritrea': 'ERI',
'estonia': 'EST',
'ethiopia': 'ETH',
'falkland islands': 'FLK',
'faroe islands': 'FRO',
'fiji': 'FJI',
'finland': 'FIN',
'france': 'FRA',
'french polynesia': 'PYF',
'gabon': 'GAB',
'gambia, the': 'GMB',
'georgia': 'GEO',
'germany': 'DEU',
'ghana': 'GHA',
'gibraltar': 'GIB',
'greece': 'GRC',
'greenland': 'GRL',
'grenada': 'GRD',
'guam': 'GUM',
'guatemala': 'GTM',
'guernsey': 'GGY',
'guinea-bissau': 'GNB',
'guinea': 'GIN',
'guyana': 'GUY',
'haiti': 'HTI',
'honduras': 'HND',
'hong kong': 'HKG',
'hungary': 'HUN',
'iceland': 'ISL',
'india': 'IND',
'indonesia': 'IDN',
'iran': 'IRN',
'iraq': 'IRQ',
'ireland': 'IRL',
'isle of man': 'IMN',
'israel': 'ISR',
'italy': 'ITA',
'jamaica': 'JAM',
'japan': 'JPN',
'jersey': 'JEY',
'jordan': 'JOR',
'kazakhstan': 'KAZ',
'kenya': 'KEN',
'kiribati': 'KIR',
'north korea': 'PRK',
'south korea': 'KOR',
'kosovo': 'KSV',
'kuwait': 'KWT',
'kyrgyzstan': 'KGZ',
'laos': 'LAO',
'latvia': 'LVA',
'lebanon': 'LBN',
'lesotho': 'LSO',
'liberia': 'LBR',
'libya': 'LBY',
'liechtenstein': 'LIE',
'lithuania': 'LTU',
'luxembourg': 'LUX',
'macau': 'MAC',
'macedonia': 'MKD',
'madagascar': 'MDG',
'malawi': 'MWI',
'malaysia': 'MYS',
'maldives': 'MDV',
'mali': 'MLI',
'malta': 'MLT',
'marshall islands': 'MHL',
'mauritania': 'MRT',
'mauritius': 'MUS',
'mexico': 'MEX',
'micronesia': 'FSM',
'moldova': 'MDA',
'monaco': 'MCO',
'mongolia': 'MNG',
'montenegro': 'MNE',
'morocco': 'MAR',
'mozambique': 'MOZ',
'namibia': 'NAM',
'nepal': 'NPL',
'netherlands': 'NLD',
'new caledonia': 'NCL',
'new zealand': 'NZL',
'nicaragua': 'NIC',
'nigeria': 'NGA',
'niger': 'NER',
'niue': 'NIU',
'northern mariana islands': 'MNP',
'norway': 'NOR',
'oman': 'OMN',
'pakistan': 'PAK',
'palau': 'PLW',
'panama': 'PAN',
'papua new guinea': 'PNG',
'paraguay': 'PRY',
'peru': 'PER',
'philippines': 'PHL',
'poland': 'POL',
'portugal': 'PRT',
'puerto rico': 'PRI',
'qatar': 'QAT',
'romania': 'ROU',
'russia': 'RUS',
'rwanda': 'RWA',
'saint kitts and nevis': 'KNA',
'saint lucia': 'LCA',
'saint martin': 'MAF',
'saint pierre and miquelon': 'SPM',
'saint vincent and the grenadines': 'VCT',
'samoa': 'WSM',
'san marino': 'SMR',
'sao tome and principe': 'STP',
'saudi arabia': 'SAU',
'senegal': 'SEN',
'serbia': 'SRB',
'seychelles': 'SYC',
'sierra leone': 'SLE',
'singapore': 'SGP',
'sint maarten': 'SXM',
'slovakia': 'SVK',
'slovenia': 'SVN',
'solomon islands': 'SLB',
'somalia': 'SOM',
'south africa': 'ZAF',
'south sudan': 'SSD',
'spain': 'ESP',
'sri lanka': 'LKA',
'sudan': 'SDN',
'suriname': 'SUR',
'swaziland': 'SWZ',
'sweden': 'SWE',
'switzerland': 'CHE',
'syria': 'SYR',
'taiwan': 'TWN',
'tajikistan': 'TJK',
'tanzania': 'TZA',
'thailand': 'THA',
'timor-leste': 'TLS',
'togo': 'TGO',
'tonga': 'TON',
'trinidad and tobago': 'TTO',
'tunisia': 'TUN',
'turkey': 'TUR',
'turkmenistan': 'TKM',
'tuvalu': 'TUV',
'uganda': 'UGA',
'ukraine': 'UKR',
'united arab emirates': 'ARE',
'united kingdom': 'GBR',
'united states': 'USA',
'uruguay': 'URY',
'uzbekistan': 'UZB',
'vanuatu': 'VUT',
'venezuela': 'VEN',
'vietnam': 'VNM',
'virgin islands': 'VGB',
'west bank': 'WBG',
'yemen': 'YEM',
'zambia': 'ZMB',
'zimbabwe': 'ZWE'}
## countries
from collections import Counter
colorscale = ["#f7fbff", "#ebf3fb", "#deebf7", "#d2e3f3", "#c6dbef", "#b3d2e9", "#9ecae1",
"#85bcdb", "#6baed6", "#57a0ce", "#4292c6", "#3082be", "#2171b5", "#1361a9",
"#08519c", "#0b4083", "#08306b"
]
def geoplot(ddf):
country_with_code, country = {}, {}
shows_countries = ", ".join(ddf['country'].dropna()).split(", ")
for c,v in dict(Counter(shows_countries)).items():
code = ""
if c.lower() in country_codes:
code = country_codes[c.lower()]
country_with_code[code] = v
country[c] = v
data = [dict(
type = 'choropleth',
locations = list(country_with_code.keys()),
z = list(country_with_code.values()),
colorscale = [[0,"rgb(5, 10, 172)"],[0.65,"rgb(40, 60, 190)"],[0.75,"rgb(70, 100, 245)"],\
[0.80,"rgb(90, 120, 245)"],[0.9,"rgb(106, 137, 247)"],[1,"rgb(220, 220, 220)"]],
autocolorscale = False,
reversescale = True,
marker = dict(
line = dict (
color = 'gray',
width = 0.5
) ),
colorbar = dict(
autotick = False,
title = ''),
) ]
layout = dict(
title = '',
geo = dict(
showframe = False,
showcoastlines = False,
projection = dict(
type = 'Mercator'
)
)
)
fig = dict( data=data, layout=layout )
iplot( fig, validate=False, filename='d3-world-map' )
return country
country_vals = geoplot(df)
tabs = Counter(country_vals).most_common(25)
labels = [_[0] for _ in tabs][::-1]
values = [_[1] for _ in tabs][::-1]
trace1 = go.Bar(y=labels, x=values, orientation="h", name="", marker=dict(color="#a678de"))
data = [trace1]
layout = go.Layout(title="Countries with most content", height=700, legend=dict(x=0.1, y=1.1, orientation="h"))
fig = go.Figure(data, layout=layout)
fig.show()
```
## 5. Distribution of Movie Duration
```
import plotly.figure_factory as ff
x1 = d2['duration'].fillna(0.0).astype(float)
fig = ff.create_distplot([x1], ['a'], bin_size=0.7, curve_type='normal', colors=["#6ad49b"])
fig.update_layout(title_text='Distplot with Normal Distribution')
fig.show()
```
## 6. TV Shows with many seasons
```
col = 'season_count'
vc1 = d1[col].value_counts().reset_index()
vc1 = vc1.rename(columns = {col : "count", "index" : col})
vc1['percent'] = vc1['count'].apply(lambda x : 100*x/sum(vc1['count']))
vc1 = vc1.sort_values(col)
trace1 = go.Bar(x=vc1[col], y=vc1["count"], name="TV Shows", marker=dict(color="#a678de"))
data = [trace1]
layout = go.Layout(title="Seasons", legend=dict(x=0.1, y=1.1, orientation="h"))
fig = go.Figure(data, layout=layout)
fig.show()
```
## 7. The ratings of the content ?
```
col = "rating"
vc1 = d1[col].value_counts().reset_index()
vc1 = vc1.rename(columns = {col : "count", "index" : col})
vc1['percent'] = vc1['count'].apply(lambda x : 100*x/sum(vc1['count']))
vc1 = vc1.sort_values(col)
vc2 = d2[col].value_counts().reset_index()
vc2 = vc2.rename(columns = {col : "count", "index" : col})
vc2['percent'] = vc2['count'].apply(lambda x : 100*x/sum(vc2['count']))
vc2 = vc2.sort_values(col)
trace1 = go.Bar(x=vc1[col], y=vc1["count"], name="TV Shows", marker=dict(color="#a678de"))
trace2 = go.Bar(x=vc2[col], y=vc2["count"], name="Movies", marker=dict(color="#6ad49b"))
data = [trace1, trace2]
layout = go.Layout(title="Content added over the years", legend=dict(x=0.1, y=1.1, orientation="h"))
fig = go.Figure(data, layout=layout)
fig.show()
```
## 8. What are the top Categories ?
```
col = "listed_in"
categories = ", ".join(d2['listed_in']).split(", ")
counter_list = Counter(categories).most_common(50)
labels = [_[0] for _ in counter_list][::-1]
values = [_[1] for _ in counter_list][::-1]
trace1 = go.Bar(y=labels, x=values, orientation="h", name="TV Shows", marker=dict(color="#a678de"))
data = [trace1]
layout = go.Layout(title="Content added over the years", legend=dict(x=0.1, y=1.1, orientation="h"))
fig = go.Figure(data, layout=layout)
fig.show()
```
## 9. Top Actors on Netflix with Most Movies
```
def country_trace(country, flag = "movie"):
df["from_us"] = df['country'].fillna("").apply(lambda x : 1 if country.lower() in x.lower() else 0)
small = df[df["from_us"] == 1]
if flag == "movie":
small = small[small["duration"] != ""]
else:
small = small[small["season_count"] != ""]
cast = ", ".join(small['cast'].fillna("")).split(", ")
tags = Counter(cast).most_common(25)
tags = [_ for _ in tags if "" != _[0]]
labels, values = [_[0]+" " for _ in tags], [_[1] for _ in tags]
trace = go.Bar(y=labels[::-1], x=values[::-1], orientation="h", name="", marker=dict(color="#a678de"))
return trace
from plotly.subplots import make_subplots
traces = []
titles = ["United States", "","India","", "United Kingdom", "Canada","", "Spain","", "Japan"]
for title in titles:
if title != "":
traces.append(country_trace(title))
fig = make_subplots(rows=2, cols=5, subplot_titles=titles)
fig.add_trace(traces[0], 1,1)
fig.add_trace(traces[1], 1,3)
fig.add_trace(traces[2], 1,5)
fig.add_trace(traces[3], 2,1)
fig.add_trace(traces[4], 2,3)
fig.add_trace(traces[5], 2,5)
fig.update_layout(height=1200, showlegend=False)
fig.show()
```
## 9. Top Actors on Netflix with Most TV Shows
```
traces = []
titles = ["United States","", "United Kingdom"]
for title in titles:
if title != "":
traces.append(country_trace(title, flag="tv_shows"))
fig = make_subplots(rows=1, cols=3, subplot_titles=titles)
fig.add_trace(traces[0], 1,1)
fig.add_trace(traces[1], 1,3)
fig.update_layout(height=600, showlegend=False)
fig.show()
small = df[df["type"] == "Movie"]
small = small[small["country"] == "India"]
col = "director"
categories = ", ".join(small[col].fillna("")).split(", ")
counter_list = Counter(categories).most_common(12)
counter_list = [_ for _ in counter_list if _[0] != ""]
labels = [_[0] for _ in counter_list][::-1]
values = [_[1] for _ in counter_list][::-1]
trace1 = go.Bar(y=labels, x=values, orientation="h", name="TV Shows", marker=dict(color="orange"))
data = [trace1]
layout = go.Layout(title="Movie Directors from India with most content", legend=dict(x=0.1, y=1.1, orientation="h"))
fig = go.Figure(data, layout=layout)
fig.show()
small = df[df["type"] == "Movie"]
small = small[small["country"] == "United States"]
col = "director"
categories = ", ".join(small[col].fillna("")).split(", ")
counter_list = Counter(categories).most_common(12)
counter_list = [_ for _ in counter_list if _[0] != ""]
labels = [_[0] for _ in counter_list][::-1]
values = [_[1] for _ in counter_list][::-1]
trace1 = go.Bar(y=labels, x=values, orientation="h", name="TV Shows", marker=dict(color="orange"))
data = [trace1]
layout = go.Layout(title="Movie Directors from US with most content", legend=dict(x=0.1, y=1.1, orientation="h"))
fig = go.Figure(data, layout=layout)
fig.show()
```
Standup Comedies by Jay Karas
```
tag = "jay karas"
df["relevant"] = df['director'].fillna("").apply(lambda x : 1 if tag in x.lower() else 0)
small = df[df["relevant"] == 1]
small[['title', 'release_year', 'listed_in']]
```
## 10. StandUp Comedies on Netflix
- United States
```
tag = "Stand-Up Comedy"
df["relevant"] = df['listed_in'].fillna("").apply(lambda x : 1 if tag.lower() in x.lower() else 0)
small = df[df["relevant"] == 1]
small[small["country"] == "United States"][["title", "country","release_year"]].head(10)
```
- India
```
tag = "Stand-Up Comedy"
df["relevant"] = df['listed_in'].fillna("").apply(lambda x : 1 if tag.lower() in x.lower() else 0)
small = df[df["relevant"] == 1]
small[small["country"] == "India"][["title", "country","release_year"]].head(10)
```
## More Work in Progress
| github_jupyter |
```
import pandas as pd
import os
import numpy as np
from string_search import *
data_dir = r'C:\Users\ozano\Desktop\senet'
data_path = os.path.join(data_dir, 'results_me.csv')
df = pd.read_csv(data_path, sep = ';')
cols_to_use = ['AD', 'ADRES']
df = df[cols_to_use]
df.shape
"""df = pd.concat([df for _ in range(2300)], axis = 0)
df.shape"""
```
## Preprocess
```
for col in df.columns:
df[col] = df[col].apply(lambda x: preprocess(x))
"""df.loc[df.index == 0, 'AD'] = 'zeynel'
df.loc[df.index == 2, 'AD'] = 'zeynel abakayli'
df.loc[df.index == 3, 'AD'] = 'zeynel bakaysdsfdlik ad'"""
df.head(10)
```
## Get N-Grams
```
df_ngram = pd.DataFrame()
for col in df.columns:
df_ngram[col] = df[col].apply(lambda x: get_n_grams(x))
df_ngram.head()
```
## Create Index
```
#%%timeit 85.8 ms
import math
def get_n_gram_length(x):
return max(1, len(x))
n_index_tokens = np.array([4, 7, 10, 15, 20, 30, 50, 70])
labels = ['AD', 'ADRES']
index_data = {label: {'index':[], 'vocabulary':[], 'lengths': []} for label in labels}
for n_index_token in n_index_tokens:
for label in labels:
index, vocab, lengths = create_ngram_index_sparse(df_ngram[label].values, first_n_tokens = n_index_token)
index_data[label]['index'].append(index)
index_data[label]['vocabulary'].append(vocab)
index_data[label]['lengths'].append(lengths)
```
## Search
```
s = 'mehmet caliskan ahmet doger'
input_n_gram_set = reduce_n_grams(input_n_grams)
len([t for t in input_n_gram_set if t in reduce_n_grams(get_n_grams(s))]) / len(input_n_gram_set)
input_string = 'mehmet calik'
search_label = 'AD'
search_person = True
# mehmet kocamanoglu erkan calik
# mehmet caliskan ahmet doger
# reneva otomotiv insaat gida turizm tasimacilik sanayi ve tic
# metin aydinhusamettin aydin
#search_person = False # Person: True, Company: False, Address: False
input_n_grams = get_n_grams(input_string)
n_ngrams = len(input_n_grams)
#index_size = np.argmin(np.abs(n_index_tokens - n_ngrams))
index_size = len(n_index_tokens) - 1
if not search_person:
for i, n_index_token in enumerate(n_index_tokens[:-1]):
if n_ngrams <= n_index_token:
index_size = i
break
print(f'Input n_gram count: {n_ngrams}')
print(f'Searching with index size: {n_index_tokens[index_size]}')
search_data = (index_data[search_label]['index'][index_size], index_data[search_label]['vocabulary'][index_size], index_data[search_label]['lengths'][-1])
match_groups = search_groups_vectorized(input_n_grams, search_data[0], search_data[1], search_data[2],
get_sorted = True, search_person = search_person)
search_count = 0
while len(match_groups[1.0]) == 0 and len(match_groups[0.8]) == 0 and index_size < len(n_index_tokens) - 1 and not search_person:
index_size = index_size + 1
print(f'Searching with index size: {n_index_tokens[index_size]}')
search_data = (index_data[search_label]['index'][index_size], index_data[search_label]['vocabulary'][index_size], index_data[search_label]['lengths'][-1])
match_groups = search_groups_vectorized(input_n_grams, search_data[0], search_data[1], search_data[2],
get_sorted = True, search_person = search_person)
match_values = get_match_values(match_groups, df[search_label].values)
```
## Get values
```
match_values[1.0]
match_values[0.8]
match_values[0.6]
match_values[0.4]
match_values[0.0][:25]
```
| github_jupyter |
```
import feather
import os
import re
import pickle
import time
import datetime
import numpy as np
import pandas as pd
from numba import jit
from sklearn.metrics import roc_auc_score
from sklearn.cross_validation import StratifiedKFold
from sklearn.metrics import matthews_corrcoef
import seaborn as sns
import matplotlib.pyplot as plt
from scipy.sparse import csr_matrix, hstack
from ml_toolbox.xgboostmonitor_utils import *
import ml_toolbox.xgboostmonitor_utils as xgbm
%matplotlib inline
import xgboost as xgb
import subprocess
from hyperopt import fmin, tpe, hp, STATUS_OK, Trials
# Custom modules
import const
import func
```
## Load data
```
y = func.read_last_column(os.path.join(const.BASE_PATH,const.TRAIN_FILES[0]+'.csv'))
print y.head(3)
y = y.Response.values
# Load columns name
num_cols = func.get_columns_csv(os.path.join(const.BASE_PATH, const.TRAIN_FILES[0]))[:200]
train_stack = feather.read_dataframe('divers/tr_stack1.feather')
#test_stack = feather.read_dataframe('divers/te_stack1.feather')
#tr_lauren = feather.read_dataframe('../input/tr_lauren.feather')
#te_lauren = feather.read_dataframe('../input/te_lauren.feather')
#leak = pd.read_csv('../input/leak_feature.csv')
tr_feather_set1 = feather.read_dataframe('divers/train.feather')
#te_feather_set1 = pd.read_csv('divers/test_eng.csv')
tr_feather_set1.columns = [x + '_v2' for x in tr_feather_set1.columns]
train = pd.concat([train_stack,tr_feather_set1],axis = 1)
set(train_stack.columns) & set(tr_feather_set1.columns)
features = list(train.columns)
features.remove("Y")
#features.remove("Id")
#features.remove("Id")
features.remove("Response")
#features.remove("tdeltadevrel_block1a")
features.remove("cluster_n500")
features.remove("unique_path")
features.remove('magic3')
features.remove('magic4')
X = train[features]
del train_stack,tr_feather_set1,train
import gc
gc.collect()
print('X_num_raw: {}'.format(X.shape))
print const.CV
with open(const.CV, 'rb') as f:
cv = pickle.load(f)
n_cv = len(cv)
n_cv
x_train = xgb.DMatrix(X,
label=y)
```
## Train simple model
```
def score_xgboost_full(params):
global counter
#print ('Params testing %d: %s' % (counter, params))
counter += 1
print('Predicting XGBoost score with ({}):'.format(counter))
print('\t {} samples'.format(x_train.num_row()))
print('\t {} features'.format(x_train.num_col()))
print('\t {} parameters'.format(params))
preds_val = np.zeros(y.shape)
for (itrain, ival) in cv:
x_tr = x_train.slice(itrain)
x_va = x_train.slice(ival)
watchlist = [ (x_tr, 'train'), (x_va, 'eval')]
eval_result = {}
bst = xgb.train(params,
x_tr,
num_boost_round=params['num_round'],
evals=watchlist,
evals_result=eval_result,
early_stopping_rounds=params['early_stopping'],
verbose_eval=5)
#print('\t score: {}'.format(roc_auc_score(y_val, y_pred_val)))
train_score = eval_result['train']['auc'][bst.best_iteration]
val_score = eval_result['eval']['auc'][bst.best_iteration]
# pick the best threshold based on oof predictions
preds_val[ival] = bst.predict(x_va, ntree_limit=bst.best_ntree_limit)
thresholds = np.linspace(0.01, 0.99, 50)
mcc = np.array([matthews_corrcoef(y[ival], preds_val[ival]>thr) for thr in thresholds])
th_val = thresholds[mcc.argmax()]
mcc_val = mcc.max()
print train_score
print val_score
print th_val
print mcc_val
return preds_val
def score_xgboost(params):
global counter
#print ('Params testing %d: %s' % (counter, params))
counter += 1
print('Predicting XGBoost score with ({}):'.format(counter))
print('\t {} samples'.format(x_train.num_row()))
print('\t {} features'.format(x_train.num_col()))
print('\t {} parameters'.format(params))
(itrain, ival) = cv[3]
x_tr = x_train.slice(itrain)
x_va = x_train.slice(ival)
watchlist = [ (x_tr, 'train'), (x_va, 'eval')]
eval_result = {}
bst = xgb.train(params,
x_tr,
num_boost_round=params['num_round'],
evals=watchlist,
evals_result=eval_result,
early_stopping_rounds=params['early_stopping'],
verbose_eval=5)
#print('\t score: {}'.format(roc_auc_score(y_val, y_pred_val)))
train_score = eval_result['train']['auc'][bst.best_iteration]
val_score = eval_result['eval']['auc'][bst.best_iteration]
# pick the best threshold based on oof predictions
preds_val = bst.predict(x_va, ntree_limit=bst.best_ntree_limit)
thresholds = np.linspace(0.01, 0.99, 50)
mcc = np.array([matthews_corrcoef(y[ival], preds_val>thr) for thr in thresholds])
th_val = thresholds[mcc.argmax()]
mcc_val = mcc.max()
print train_score
print val_score
print th_val
print mcc_val
return {'loss': 1-val_score,
'status': STATUS_OK,
'train_score': train_score,
'best_iter': bst.best_iteration,
'mcc': mcc_val,
'threshold': th_val}
params = {'max_depth': 7, 'eta':0.1, 'silent':1, 'objective':'binary:logistic' }
#param['nthread'] = 1
params['eval_metric'] = 'auc'
params['subsample'] = 0.9
params['colsample_bytree']= 0.8
params['min_child_weight'] = 12
params['booster'] = "gbtree"
params['seed'] = 1712
params['num_round'] = 200
params['early_stopping'] = 100
df = score_xgboost_full(params)
params = {'max_depth': 7, 'eta':0.1, 'silent':1, 'objective':'binary:logistic' }
#param['nthread'] = 1
params['eval_metric'] = 'auc'
params['subsample'] = hp.uniform('subsample', 0.7, 0.9) #,0.86
params['colsample_bytree']= hp.uniform('colsample_bytree', 0.7, 0.9) #0.92
params['min_child_weight'] = hp.choice('min_child_weight', range(50))
params['booster'] = "gbtree"
params['seed'] = 1712
params['num_round'] = 200
params['early_stopping'] = 30
# Hyperopt
trials = Trials()
counter = 0
best = fmin(score_xgboost,
params,
algo=tpe.suggest,
max_evals=200,
trials=trials)
par_values = {'max_depth': range(8,21)}
parameters = trials.trials[0]['misc']['vals'].keys()
f, axes = plt.subplots(nrows=2, ncols=2, figsize=(16,16))
cmap = plt.cm.Dark2
par_best_score = {}
df = pd.DataFrame(columns=parameters + ['train_auc','val_auc'])
for i, val in enumerate(parameters):
xs = np.array([t['misc']['vals'][val] for t in trials.trials if 'loss' in t['result']]).ravel()
val_auc = [1-t['result']['loss'] for t in trials.trials if 'loss' in t['result']]
train_auc = [t['result']['train_score'] for t in trials.trials if 'train_score' in t['result']]
best_iter = [t['result']['best_iter'] for t in trials.trials if 'best_iter' in t['result']]
mcc = [t['result']['mcc'] for t in trials.trials if 'mcc' in t['result']]
tr = [t['result']['threshold'] for t in trials.trials if 'threshold' in t['result']]
df[val] = xs
df['val_auc'] = val_auc
df['train_auc'] = train_auc
df['best_iter'] = best_iter
df['threshold'] = tr
df['mcc'] = mcc
par_best_score[val] = xs[val_auc.index(min(val_auc))]
#print trials.trials[ys.index(max(ys))]
#print i, val, max(ys)
#xs, ys = zip(sorted(xs), sorted(ys))
#ys = np.array(ys)
axes[i/2,i%2].scatter(xs, mcc, s=20, linewidth=0.01, alpha=0.5, c=cmap(float(i)/len(parameters)))
axes[i/2,i%2].set_title(val)
print par_best_score
df['diffs'] = df['train_auc'] - df['val_auc']
ax = df.plot.scatter('threshold','mcc')
#ax.set_xlim([0.921, 0.926])
ax = df.plot.scatter('val_auc','mcc')
ax.set_xlim([0.924, 0.928])
ax = df.plot.scatter('subsample','diffs')
#ax.set_xlim([0.924, 0.928])
ax = df.plot.scatter('colsample_bytree','diffs')
ax = df.plot.scatter('min_child_weight','diffs')
df.sort_values('mcc', ascending=False)
#df.drop(['gamma'], axis=1, inplace=True)
#df.to_csv('./data/xgboost_hyperopt_1fold_100iter.csv', index=False)
df['colsample_bytree'] = df['colsample_bytree'].round(2)
df.sort_values('val_auc', ascending=False)
df.head()
df['subsample'] = df['subsample'].round(2)
df['colsample_bytree'] = df['colsample_bytree'].round(2)
def plot_scores_for_pars(par):
f, ax = plt.subplots(1,3, figsize=(16,6), sharex=True)
df.groupby(par)['val_auc'].mean().plot(ax=ax[0])
df.groupby(par)['train_auc'].mean().plot(ax=ax[1])
df.groupby(par)['diffs'].mean().plot(ax=ax[2])
ax[0].set_ylabel('Test auc')
ax[1].set_ylabel('Train auc')
ax[2].set_ylabel('Difference')
ax[0].set_xlabel(par)
ax[1].set_xlabel(par)
ax[2].set_xlabel(par)
plot_scores_for_pars('subsample')
plot_scores_for_pars('colsample_bytree')
plot_scores_for_pars('min_child_weight')
plot_scores_for_pars('gamma')
plot_scores_for_pars('gamma')
df.groupby('sub_r')['val_auc'].mean().plot()
df.groupby('sub_r')['train_auc'].mean().plot()
df.groupby('colt_r')['val_auc'].mean().plot()
df.groupby('colt_r')['train_auc'].mean().plot()
df.groupby('coll_r')['val_auc'].mean().plot()
df.groupby('coll_r')['train_auc'].mean().plot()
df.plot('train_auc', 'val_auc',kind='scatter', ylim=[0.918, 0.922])
df.plot('val_auc', 'diffs', kind='scatter', xlim=[0.918, 0.922])
df.plot('gamma', 'diffs',kind='scatter')
df.plot.scatter('colsample_bytree', 'val_auc', by='max_depth')
```
| github_jupyter |
```
import gzip
import os
import pickle
import numpy as np
from sklearn import svm
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import SGDClassifier
scaled_balanced_fit_1_test = np.load(gzip.open('scaled_balanced_fit_1_test.npy.gz', 'rb'))
scaled_balanced_fit_10_test = np.load(gzip.open('scaled_balanced_fit_10_test.npy.gz', 'rb'))
scaled_balanced_fit_10_test = np.load(gzip.open('scaled_balanced_fit_10_test.npy.gz', 'rb'))
scaled_balanced_fit_cv = np.load(gzip.open('scaled_balanced_fit_cv.npy.gz', 'rb'))
ordered_features = pickle.load(open('ordered_features', 'rb'))
train_1_X = scaled_balanced_fit_1_test[:,:-1]
train_1_Y = scaled_balanced_fit_1_test[:,-1]
train_10_X = scaled_balanced_fit_10_test[:,:-1]
train_10_Y = scaled_balanced_fit_10_test[:,-1]
train_100_X = scaled_balanced_fit_10_test[:,:-1]
train_100_Y = scaled_balanced_fit_10_test[:,-1]
cv_X = scaled_balanced_fit_cv[:,:-1]
cv_Y = scaled_balanced_fit_cv[:,-1]
my_svm_1 = svm.SVC(kernel='linear')
my_svm_1.fit(train_1_X, train_1_Y)
my_svm_1.score(cv_X, cv_Y)
my_svm_10 = svm.SVC(kernel='linear')
my_svm_10.fit(train_10_X, train_10_Y)
my_svm_10.score(cv_X, cv_Y)
my_svm_1_rbf = svm.SVC(C=10,kernel='rbf')
my_svm_1_rbf.fit(train_1_X, train_1_Y)
my_svm_1_rbf.score(cv_X, cv_Y)
predictions = my_svm_10_rbf.predict(cv_X)
join = np.array([predictions, cv_Y])
my_svm_10_C = svm.SVC(C=10, kernel='rbf')
my_svm_10_C.fit(train_10_X, train_10_Y)
my_svm_10_C.score(cv_X, cv_Y)
parameters = {'kernel': ['linear', 'rbf'], 'C':[0.1, 1, 10]}
svc = svm.SVC()
clf = GridSearchCV(estimator=svc, param_grid=parameters, cv=5, n_jobs=8)
clf.fit(train_1_X, train_1_Y)
clf.best_score_ clf.best_params_
parameters = {'kernel': ['poly'], 'C':[10, 100], 'degree': [2, 3, 4, 5]}
svc2 = svm.SVC()
clf2 = GridSearchCV(estimator=svc2, param_grid=parameters, cv=5, n_jobs=8)
clf2.fit(train_1_X, train_1_Y)
clf2.best_params_, clf2.best_score_
clf2.best_params_, clf2.best_score_
parameters = {'kernel': ['sigmoid'], 'C':[0.1, 1, 10]}
svc3 = svm.SVC()
clf3 = GridSearchCV(estimator=svc3, param_grid=parameters, cv=5, n_jobs=8)
clf3.fit(train_1_X, train_1_Y)
clf3.best_score_, clf3.best_params_
scaled_fit = np.load(gzip.open('scaled_fit.npy.gz', 'rb'))
scaled_X = scaled_fit[:,:-1]
scaled_Y = scaled_fit[:,-1]
my_svm_1_rbf.score(scaled_X, scaled_Y)
sgd = SGDClassifier()
parameters = {'alpha': [0.00001, 0.0001, 0.001], 'loss':['hinge'], 'tol': [1e-3]}
csgd = GridSearchCV(estimator=sgd, param_grid=parameters, cv=5, n_jobs=3)
csgd.fit(scaled_X, scaled_Y)
csgd.best_score_, csgd.best_params_
sgd.score(scaled_X, scaled_Y)
from sklearn.kernel_approximation import RBFSampler
rbf_feature = RBFSampler(gamma=1, random_state=1)
X_features = rbf_feature.fit_transform(train_100_X)
clfxx = SGDClassifier()
clfxx.fit(X_features, train_100_Y)
fff = rbf_feature.transform(scaled_X[:200000])
clfxx.score(fff, scaled_Y[:200000])
from sklearn.ensemble import RandomForestClassifier
forest_estimator = RandomForestClassifier(n_estimators=100)
forest_fit = forest_estimator.fit(train_10_X, train_10_Y)
forest_fit.score(scaled_X, scaled_Y)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/pgautam8601/Accelerated_Computer_Science_Fundamentals_Specialization/blob/master/Content_Based_Recommender_System.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#EDA (Exploratory Data Analysis)
import pandas as pd
import gdown
import os
import numpy as np
import warnings
warnings.filterwarnings('ignore')
import matplotlib.pyplot as plt
import datetime
%matplotlib inline
pip install plotly_express==0.4.0
import plotly_express as px
books = pd.read_csv("books.csv")
tags_data = pd.read_csv("book_tags.csv")
book_tags = pd.read_csv("tags.csv")
ratings_data=pd.read_csv('ratings.csv')
ratings_data = ratings_data.sort_values("user_id")
ratings_data.drop_duplicates(subset =["user_id","book_id"], keep = False, inplace = True)
books= books.drop(columns=['book_id', 'best_book_id', 'work_id', 'isbn', 'isbn13', 'title','work_ratings_count',
'work_text_reviews_count', 'ratings_1', 'ratings_2', 'ratings_3', 'ratings_4', 'ratings_5',
'image_url','small_image_url'])
books = books.dropna()
books.drop_duplicates(subset='original_title',keep=False,inplace=True)
top_rated = books.sort_values('average_rating', ascending=False)
tf_top_rated = top_rated[:15]
fig = px.bar(tf_top_rated, x="average_rating", y="original_title", title='Top Rated Books and Their Ratings',
orientation='h', color='original_title', width=1500, height=700)
fig.show()
#distribution of average ratings of all the 10000 books
import matplotlib.pyplot as plt
plt.figure(figsize=(12,6))
plt.title("Distribution of Average Ratings")
books["average_rating"].hist()
display()
books2 = pd.read_csv('books.csv')
books2 = books2.dropna()
books_filter = pd.DataFrame(books2, columns=['book_id', 'authors', 'original_title', 'average_rating'])
books_filter = books_filter.sort_values('average_rating', ascending=False)
books_filter.head(15)
top_author_counts = books['authors'].value_counts().reset_index()
top_author_counts.columns = ['value', 'count']
top_author_counts['value'] = top_author_counts['value']
top_author_counts = top_author_counts.sort_values('count')
fig = px.bar(top_author_counts.tail(10), x="count", y="value", title='Top Authors', orientation='h', color='value',
width=1000, height=700)
fig.show()
ratings_data.drop_duplicates(subset =["user_id","book_id"], keep = False, inplace = True)
books.drop_duplicates(subset='original_title',keep=False,inplace=True)
books2= books2.drop(columns=[ 'best_book_id', 'work_id', 'isbn13', 'title','work_ratings_count',
'work_text_reviews_count', 'ratings_1', 'ratings_2', 'ratings_3', 'ratings_4', 'ratings_5',
'image_url','small_image_url'])
merge_data2 = pd.merge(books2, ratings_data, on='book_id')
merge_data2 = merge_data2.sort_values('book_id', ascending=True)
merge_data2.head()
years= books2['original_publication_year'].value_counts().reset_index()
years.columns = ['year', 'count']
years['year'] = years['year']
years = years.sort_values('count')
fig = px.bar(years.tail(50), x="count", y="year", title='Publication Year', orientation='h', color='count',
width=1000, height=700)
fig.show()
lang= books2['language_code'].value_counts().reset_index()
lang.columns = ['value', 'count']
lang['value'] = lang['value']
lang = lang.sort_values('count')
fig = px.bar(lang.tail(10), x="count", y="value", title='Languages', orientation='h', color='count',
width=1000, height=700)
fig.show()
# Mean rating per user
MRPU = ratings_data.groupby(['user_id']).mean().reset_index()
MRPU['mean_rating'] = MRPU['rating']
MRPU.drop(['book_id','rating'],axis=1, inplace=True)
MRPU.head(15)
rating_data = pd.merge(ratings_data, MRPU, on=['user_id', 'user_id'])
rating_data.head(200)
ratings_data[ratings_data['user_id']== 11141 ].head(10)
ratings_data['user_id'].unique()[-1] # Total number of users
book_tags.tail()
#Data Preprocessing
import seaborn as sns
import matplotlib.pyplot as plt
import re
import matplotlib.style as style
import os
books = pd.read_csv("books.csv")
book_tags = pd.read_csv("book_tags.csv")
tags = pd.read_csv("tags.csv")
ratings = pd.read_csv("ratings.csv")
tags.head()
book_tags.head()
#Joining book_tags and tags dataframe by Left join.
book_tags = pd.merge(book_tags,tags,on='tag_id',how='left')
book_tags.drop(book_tags[book_tags.duplicated()].index, inplace = True)
book_tags
books.head()
#Removing unnecessary Columns
books.drop(columns=['id', 'best_book_id', 'work_id', 'isbn', 'isbn13', 'title','work_ratings_count','ratings_count','work_text_reviews_count', 'ratings_1', 'ratings_2', 'ratings_3','ratings_4', 'ratings_5', 'image_url','small_image_url'], inplace= True)
#Renaming Columns
books.rename(columns={'original_publication_year':'pub_year', 'original_title':'title', 'language_code':'language', 'average_rating':'rating'}, inplace=True)
books.isnull().sum()
#Dropping the Null values
books.dropna(inplace= True)
#Using Split-String function for creating a list of authors
books['authors'] = books.authors.str.split(',')
books
book_authors = books.copy()
#For every row in the dataframe, iterating through the list of authors and placing 1 in the corresponding column
for index, row in books.iterrows():
for author in row['authors']:
book_authors.at[index, author] = 1
#Filling in the NaN values with 0 to show that a book isn't written by that author
book_authors = book_authors.fillna(0)
book_authors.head()
#Generalising the format of author names for simplicity
book_authors.columns = [c.lower().strip().replace(' ', '_') for c in book_authors.columns]
#Setting book_id as index of the dataframe
book_authors = book_authors.set_index(book_authors['book_id'])
#Dropping unnecessary columns
book_authors.drop(columns= {'book_id','pub_year','title','rating','books_count', 'authors','language'}, inplace=True)
#FINAL book_authors:-
book_authors.head()
#Content-based Recommendation System
#Creating an input user to recommend books to:-
user_1 = pd.DataFrame([{'book_id':2767052, 'rating':5.0},{'book_id':3, 'rating':4.0}, {'book_id':41865, 'rating':4.5},{'book_id':15613, 'rating':3.0},{'book_id':2657, 'rating':2.5}])
user_1
user_authors = book_authors[book_authors.index.isin(user_1['book_id'].tolist())].reset_index(drop=True)
user_authors
user_1.rating
#Dot Product to get Weights
userProfile = user_authors.transpose().dot(user_1['rating'])
#The User Profile
userProfile
recommendation = (((book_authors*userProfile).sum(axis=1))/(userProfile.sum())).sort_values(ascending=False)
#Top 20 recommendations
recommendation.head(20)
#The final recommendation table:
books.loc[books['book_id'].isin(recommendation.head(20).keys())].reset_index()
```
| github_jupyter |
# Can fingerprint distances discriminate DFG conformations?
The `kissim` fingerprint encodes the pocket residues' spatial distance to four centers—the pocket centroid, hinge region, DFG region and front pocket—and should therefore discriminate between two structures in different conformations; when we compare two structures in *different* conformations the fingerprint distance should be higher than for two structures in *similar* conformations.
Let's check if this is true using DFG conformations from KLIFS. Plot distribution of fingerprint distances grouped by in/in, out/out, and in/out pairs.
- Use fingerprint distances for structure pairs between all kinases
- Use fingerprint distances for structure pairs between the same kinase
```
%load_ext autoreload
%autoreload 2
from pathlib import Path
import pandas as pd
import matplotlib.pyplot as plt
from IPython.display import display, Markdown
from opencadd.databases.klifs import setup_remote
from kissim.comparison import FingerprintDistanceGenerator
from src.definitions import COVERAGE_CUTOFF
from src.paths import PATH_RESULTS
HERE = Path(_dh[-1]) # noqa: F821
DATA = PATH_RESULTS / "all"
plt.style.use("seaborn")
```
## Load fingerprint distances with sufficient coverage
Choose fingerprint distances that are based on spatial distances only (**weighting scheme: 010**) and that are based on a sufficient pairwise fingerprint bit coverage (default: `COVERAGE_CUTOFF`).
```
COVERAGE_CUTOFF
# Set path
fingerprint_distance_file = DATA / "fingerprint_distances_010.csv.bz2"
# Load data
fingerprint_distance_generator = FingerprintDistanceGenerator.from_csv(fingerprint_distance_file)
print(f"Number of kinases: {len(fingerprint_distance_generator.kinase_ids)}")
print(f"Number of structures: {len(fingerprint_distance_generator.structure_ids)}")
structure_distances = fingerprint_distance_generator.data
print(f"Number of structure pairs: {structure_distances.shape[0]}")
structure_distances = structure_distances[
structure_distances["bit_coverage"] >= COVERAGE_CUTOFF
].reset_index(drop=True)
print(f"Number of structure pairs: {structure_distances.shape[0]}")
structure_distances.head()
```
## Add DFG conformation
Add DFG conformation from KLIFS to each structure pair.
```
def get_dfg(dfg, structure_klifs_id):
try:
return dfg[structure_klifs_id]
except KeyError:
return None
%%time
klifs_session = setup_remote()
structures = klifs_session.structures.all_structures()
dfg = structures.set_index("structure.klifs_id")["structure.dfg"]
structure_distances["dfg.1"] = structure_distances["structure.1"].apply(lambda x: get_dfg(dfg, x))
structure_distances["dfg.2"] = structure_distances["structure.2"].apply(lambda x: get_dfg(dfg, x))
structure_distances.head()
```
## Plot DFG conformation pairs
Group the structure pairs by DFG conformation pairs—in/in, out/out, in/out—and plot their fingerprint distance distributions.
```
def structure_distances_by_dfg_conformation_pairs(structure_distances):
"""Distances for all, in/in, out/out, and in/out structure pairs."""
dfg_in_in = structure_distances[
(structure_distances["dfg.1"] == "in") & (structure_distances["dfg.2"] == "in")
]["distance"]
dfg_out_out = structure_distances[
(structure_distances["dfg.1"] == "out") & (structure_distances["dfg.2"] == "out")
]["distance"]
dfg_in_out = structure_distances[
((structure_distances["dfg.1"] == "in") & (structure_distances["dfg.2"] == "out"))
| ((structure_distances["dfg.1"] == "out") & (structure_distances["dfg.2"] == "in"))
]["distance"]
structure_distances_dfg = pd.DataFrame(
{"in/in": dfg_in_in, "out/out": dfg_out_out, "in/out": dfg_in_out}
)
structure_distances_dfg = pd.DataFrame(structure_distances_dfg)
return structure_distances_dfg
def plot_structure_distances_by_dfg_conformation_pairs(structure_distances, kinase):
"""Plot distribution of structure distances per DFG conformation pair."""
# Data
structure_distances_dfg = structure_distances_by_dfg_conformation_pairs(structure_distances)
print("Number of structure pairs per conformation pair:")
print(structure_distances_dfg.notna().sum())
# Boxplot
fig, ax = plt.subplots(1, 1, figsize=(3.33, 3.33))
structure_distances_dfg.plot(kind="box", ax=ax)
ax.set_xlabel(
"Type of DFG conformation pairs"
if kinase is None
else f"Type of DFG conformation pairs ({kinase})"
)
ax.set_ylabel("Fingerprint distances (spatial features only)")
if kinase is None:
fig.savefig(
HERE / "../figures/kissim_discriminates_dfg.png",
dpi=300,
bbox_inches="tight",
)
else:
fig.savefig(
HERE / f"../figures/kissim_discriminates_dfg_{kinase}.png",
dpi=300,
bbox_inches="tight",
)
plt.show()
# Stats
display(structure_distances_dfg.describe())
```
### All structures
Use fingerprint distances for structure pairs between all kinases.
```
plot_structure_distances_by_dfg_conformation_pairs(structure_distances, kinase=None)
```
<div class="alert alert-block alert-info">
When including all kinases at the same time, the distribution of fingerprint distances is similar for structure pairs with the same DFG conformations (in/in and out/out) and different DFG conformations (in/out).
The fingerprint seems not to discriminate DFG-conformations on a kinome-wide level, maybe because the encoded spatial information is not restricted to only DFG conformation features. We may see a disciminative effect when comparing structures for a single kinase.
</div>
### Structures for one kinase
Use fingerprint distances for structure pairs between the same kinase; use only kinases that have a sufficient number of structures in DFG-in and DFG-out conformations (default: 10).
```
def kinases_with_high_dfg_in_out_coverage(structure_distances, dfg_structure_coverage_cutoff=10):
"""Given a dataset, get kinases with a threshold DFG in/out coverage."""
# Get structure KLIFS IDs in our dataset
structure_klifs_ids = (
pd.concat(
[
structure_distances["structure.1"].drop_duplicates(),
structure_distances["structure.2"].drop_duplicates(),
]
)
.drop_duplicates()
.to_list()
)
print(f"Number of structures: {len(structure_klifs_ids)}")
# Get structural metadata
klifs_session = setup_remote()
structures = klifs_session.structures.all_structures()
structures = structures[structures["structure.klifs_id"].isin(structure_klifs_ids)]
# Count number of structures per kinase and conformation
dfg_by_kinase = structures.groupby("kinase.klifs_name").apply(
lambda x: x["structure.dfg"].value_counts()
)
dfg_by_kinase = dfg_by_kinase.reset_index()
dfg_by_kinase.columns = ["kinase", "dfg", "n_structures"]
# Keep only in/out rows
dfg_by_kinase = dfg_by_kinase[(dfg_by_kinase["dfg"] == "in") | (dfg_by_kinase["dfg"] == "out")]
# Keep only rows with at least xxx structures
dfg_by_kinase = dfg_by_kinase[dfg_by_kinase["n_structures"] >= dfg_structure_coverage_cutoff]
# Keep only kinases with both in/out conformations
n_conformations_by_kinase = dfg_by_kinase.groupby("kinase").size()
dfg_by_kinase = dfg_by_kinase[
dfg_by_kinase["kinase"].isin(
n_conformations_by_kinase[n_conformations_by_kinase == 2].index
)
]
return dfg_by_kinase.set_index(["kinase", "dfg"])
dfg_by_kinase = kinases_with_high_dfg_in_out_coverage(
structure_distances, dfg_structure_coverage_cutoff=10
)
dfg_by_kinase
for kinase, dfg in dfg_by_kinase.reset_index().groupby("kinase"):
display(Markdown(f"#### {kinase}"))
dfg = dfg.set_index("dfg")
n_dfg_in = dfg.loc["in", "n_structures"]
n_dfg_out = dfg.loc["out", "n_structures"]
print(f"Number of DFG-in structures: {n_dfg_in}")
print(f"Number of DFG-out structures: {n_dfg_out}")
dfg_in_percentage = round(n_dfg_in / (n_dfg_in + n_dfg_out) * 100, 2)
print(f"Percentage of DFG-in: {dfg_in_percentage}%")
structure_distances_by_kinase = structure_distances[
(structure_distances["kinase.1"] == kinase) & (structure_distances["kinase.2"] == kinase)
].reset_index(drop=True)
plot_structure_distances_by_dfg_conformation_pairs(structure_distances_by_kinase, kinase)
```
<div class="alert alert-block alert-info">
We compare here only fingerprint distances for pairs of structures that describe the same kinase. We observe two interesting shifts:
1. The distribution for out/out pairs is overall lower than for in/in pairs. Potential explanations: definitions for DFG-out are stricter than for DFG-in; "real" diversity of DFG-out structures could be still unknown due to the lower number of structures for DFG-out than for DFG-in.
2. The distribution of different DFG conformations (in/out) is overall higher than for equal DFG conformations (in/in and out/out). The fingerprint can discriminate DFG conformations of the same kinase.
</div>
| github_jupyter |
# ML Pipeline Preparation
Follow the instructions below to help you create your ML pipeline.
### 1. Import libraries and load data from database.
- Import Python libraries
- Load dataset from database with [`read_sql_table`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_sql_table.html)
- Define feature and target variables X and Y
```
# import libraries
#import sys
#sqlalchemy_utils.__version__
#from distutils.sysconfig import get_python_lib
#print(get_python_lib())
from time import time
import numpy as np
import pandas as pd
from sqlalchemy import create_engine
from sqlalchemy_utils import database_exists
import pickle
import nltk
nltk.download(['punkt', 'wordnet', 'averaged_perceptron_tagger', 'stopwords'])
from nltk.tokenize import word_tokenize
from nltk.stem import WordNetLemmatizer
from nltk.corpus import stopwords
from nltk.tokenize import RegexpTokenizer
import string
from sklearn.pipeline import Pipeline
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sklearn.multioutput import MultiOutputClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report, accuracy_score
from sklearn.svm import SVC
```
## Inspect data
```
database_filepath = "data/disaster_response.db"
database_exists(f'sqlite:///{database_filepath}')
engine = create_engine(f'sqlite:///{database_filepath}')
connection = engine.connect()
df = pd.read_sql_table("messages_categories", con=connection)
df.head()
df.columns
for col in df.iloc[:, 3:]:
print(df[col].unique())
def load_data(database_filepath):
'''
Input:
database_filename(str): Filepath of the database.
Output:
X(numpy.ndarray): Array of input features.
y(numpy.ndarray): Output labels, classes.
'''
try:
database_exists(f'sqlite:///{database_filepath}')
engine = create_engine(f'sqlite:///{database_filepath}')
connection = engine.connect()
df = pd.read_sql_table("messages_categories", con=connection)
labels = df.iloc[:,4:].columns
X = df["message"].values
y = df.iloc[:,4:].values
connection.close()
return X, y, labels
except:
print("Database does not exist! Check your database_filepath!")
```
### 2. Write a tokenization function to process your text data
```
def tokenize(text):
''' Normalize, lemmantize and tokenize text messages.
Input:
text(str): Text messages.
Output:
clean_tokens(str): Normalize, lemmantize and tokenize text messages.
'''
stop_words = set(stopwords.words('english'))
# normalize text
normalized_text = text.lower().strip()
# tokenize text
tokens = word_tokenize(normalized_text)
# lemmantize text and remove stop words and non alpha numericals
clean_tokens = []
for token in tokens:
lemmatizer = WordNetLemmatizer()
clean_token = lemmatizer.lemmatize(token)
if clean_token not in stop_words and clean_token.isalpha():
clean_tokens.append(clean_token)
return clean_tokens
```
### 3. Build a machine learning pipeline
This machine pipeline should take in the `message` column as input and output classification results on the other 36 categories in the dataset. You may find the [MultiOutputClassifier](http://scikit-learn.org/stable/modules/generated/sklearn.multioutput.MultiOutputClassifier.html) helpful for predicting multiple target variables.
```
#https://machinelearningmastery.com/prepare-text-data-machine-learning-scikit-learn/
X, y, labels = load_data("data/disaster_response.db")
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
X_train
y_train
labels
for data in [X_train, X_test, y_train, y_test]:
print(data.shape, type(data))
type(df)
from sklearn.utils.multiclass import type_of_target
(type_of_target(y_test))
```
### 4. Train pipeline
- Split data into train and test sets
- Train pipeline
```
def build_model():
'''Build a Machine Learning pipeline using TfidfTransformer, RandomForestClassifier and GridSearchCV
Input:
None
Output:
cv(sklearn.model_selection._search.GridSearchCV): Results of GridSearchCV
'''
text_clf = Pipeline([
('vect', CountVectorizer(tokenizer=partial(tokenize))),
('tfidf', TfidfTransformer()),
('clf', MultiOutputClassifier(
estimator=RandomForestClassifier()))
])
parameters = {
'clf__estimator__max_depth': [4, 6, 10, 12],
'clf__estimator__n_estimators': [20, 40, 100],
}
grid_fit = GridSearchCV(
estimator=text_clf,
param_grid=parameters,
verbose=3,
cv=2,
n_jobs=-1)
return grid_fit
from sklearn.utils import parallel_backend
from functools import partial
with parallel_backend('multiprocessing'):
model = build_model() # stop_words='english'
model.fit(X_train,y_train)
```
### 5. Test your model
Report the f1 score, precision and recall for each output category of the dataset. You can do this by iterating through the columns and calling sklearn's `classification_report` on each.
```
def evaluate_model(model, X_test, y_test, labels):
""" Function that will predict on X_test messages using build_model() function that
transforms messages, extract features and trains a classifer.
Input:
model(sklearn.model_selection._search.GridSearchCV):
X_test(numpy.ndarray): Numpy array of messages that based on which trained model will predict.
y_test(numpy.ndarray): Numpy array of classes that will be used to validate model predictions.
labels(pandas.core.indexes.base.Index): Target labels for a multiclass prediction.
Output:
df(pandas.core.frame.DataFrame): Dataframe that contains report showing the main classification metrics.
"""
y_pred = model.predict(X_test)
df = pd.DataFrame(classification_report(y_test, y_pred, target_names=labels, output_dict=True)).T.reset_index()
df = df.rename(columns = {"index": "labels"})
return df
model.best_score_
model.best_estimator_
X_train.shape, X_test.shape, y_train.shape, y_test.shape, len(labels)
with parallel_backend('multiprocessing'):
df_evaluation = evaluate_model(model, X_test, y_test, labels)
df_evaluation
df_evaluation[["labels", "precision"]].plot(x="labels", y = "precision", kind="bar", rot=90);
df_evaluation[["labels", "recall"]].plot(x="labels", y = "recall", kind="bar", rot=90);
from sklearn.metrics import confusion_matrix
import seaborn as sns
%matplotlib inline
pred = best_clf.predict(X_test)
sns.heatmap(confusion_matrix(y_test, pred), annot = True, fmt = '')
```
### 6.Default model
Use grid search to find better parameters.
```
def build_model():
'''Build a Machine Learning pipeline using TfidfTransformer, RandomForestClassifier and GridSearchCV
Input:
None
Output:
cv(sklearn.model_selection._search.GridSearchCV): Results of GridSearchCV
'''
text_clf = Pipeline([
('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer()),
('clf', MultiOutputClassifier(estimator=RandomForestClassifier()))
])
return text_clf
model = build_model()
model.fit(X_train,y_train)
df_evaluation_no_grid = evaluate_model(model, X_test, y_test, labels)
df_evaluation_no_grid
df_evaluation_no_grid[["labels", "precision"]].plot(x="labels", y = "precision", kind="bar", rot=90);
```
### 7. Test your model
Show the accuracy, precision, and recall of the tuned model.
Since this project focuses on code quality, process, and pipelines, there is no minimum performance metric needed to pass. However, make sure to fine tune your models for accuracy, precision and recall to make your project stand out - especially for your portfolio!
### 8. Try improving your model further. Here are a few ideas:
* try other machine learning algorithms
* add other features besides the TF-IDF
### 9. Export your model as a pickle file
```
def save_model(model, filepath):
'''Saves the model to defined filepath
Input
model(sklearn.model_selection._search.GridSearchCV): The model to be saved.
model_filepath(str): Filepath where the model will be saved.
Output
This function will save the model as a pickle file on the defined filepath.
'''
temporary_pickle = open(filepath, 'wb')
pickle.dump(model, temporary_pickle)
temporary_pickle.close()
print("Model has been succesfully saved!")
save_model(model, "models/model.pkl")
```
### 10. Use this notebook to complete `train.py`
Use the template file attached in the Resources folder to write a script that runs the steps above to create a database and export a model based on a new dataset specified by the user.
```
# Feature importance
# Import a supervised learning model that has 'feature_importances_'
#from sklearn.tree import DecisionTreeClassifier
# Train the supervised model on the training set using .fit(X_train, y_train)
#model = DecisionTreeClassifier()
#model.fit(X_train, y_train)
# Extract the feature importances using .feature_importances_
#importances = model.feature_importances_
# Plot
#vs.feature_plot(importances, X_train, y_train)
```
| github_jupyter |
### Imports
```
# !pip install --upgrade category_encoders rich catboost
from rich.console import Console
console = Console()
print = console.print
from wrangling import X, y
import time
import math
import pandas as pd
import numpy as np
import seaborn as sns
import sys
import matplotlib.pyplot as plt
import yellowbrick as yb
from yellowbrick.features import Rank1D
from yellowbrick.regressor import AlphaSelection, PredictionError, ResidualsPlot
from yellowbrick.datasets import load_energy
from yellowbrick.model_selection import ValidationCurve
from yellowbrick.style import set_palette
from sklearn.metrics import mean_squared_error
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import r2_score
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split
from sklearn.model_selection import RepeatedStratifiedKFold
from sklearn.model_selection import GridSearchCV, RandomizedSearchCV
from sklearn.gaussian_process import GaussianProcessRegressor
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import Ridge
from sklearn.linear_model import Lars
from sklearn.linear_model import TheilSenRegressor
from sklearn.linear_model import HuberRegressor
from sklearn.linear_model import PassiveAggressiveRegressor
from sklearn.linear_model import ARDRegression
from sklearn.linear_model import BayesianRidge
from sklearn.linear_model import ElasticNet
from sklearn.linear_model import OrthogonalMatchingPursuit
from sklearn.svm import SVR
from sklearn.svm import NuSVR
from sklearn.svm import LinearSVR
from sklearn.preprocessing import StandardScaler
from sklearn.ensemble import RandomForestRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.kernel_ridge import KernelRidge
from sklearn.isotonic import IsotonicRegression
import xgboost as xgb
from xgboost import XGBRegressor
import lightgbm as lgb
import catboost as ctb
from hyperopt import STATUS_OK, STATUS_FAIL, Trials, fmin, hp, tpe
np.set_printoptions(precision=3, suppress=True)
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
import warnings
warnings.filterwarnings('ignore')
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size=0.30,random_state=21)
plt.style.context('dark_background')
set_palette('sns_bright')
cm = sns.color_palette("blend:white,#00ff77", as_cmap=True)
def headd(i):
return i.style.background_gradient(cmap = cm,axis=None)
```
### Visuals
```
## Ranking the features
fig, ax = plt.subplots(1, figsize=(10, 35))
vzr = Rank1D(ax=ax, color='#00ff77')
vzr.fit(X_train, y_train)
vzr.transform(X_train)
sns.despine(left=True, bottom=True)
vzr.poof();
# Showing the Residuals, differences between observed and predicted values of data
# the 'delta' between the actual target value and the fitted value. Residual is a crucial concept in regression problems
model = Ridge()
visualizer = ResidualsPlot(
model,
hist=False,
qqplot=True,
size=(600, 200),
train_color="indigo",
test_color="#00ff77",
)
visualizer.fit(X_train, y_train)
visualizer.score(X_test, y_test)
# visualizer.score(X_test, y_test)
g = visualizer.poof();
X_outliers = pd.DataFrame(index=X.columns, columns=['outliers', 'outliers%'])
for col in X.columns:
if any(x in str(X[col].dtype)for x in ['int', 'float', 'int64', 'uint8']):
X_outliers.loc[col, 'count'] = len(X)
X_outliers.loc[col, 'q1'] = X[col].quantile(0.25)
X_outliers.loc[col, 'q3'] = X[col].quantile(0.75)
X_outliers.loc[col, 'iqr'] = X_outliers.loc[col, 'q3'] - X_outliers.loc[col, 'q1']
X_outliers.loc[col, 'lower'] = X_outliers.loc[col, 'q1'] - (3 * X_outliers.loc[col, 'iqr'])
X_outliers.loc[col, 'upper'] = X_outliers.loc[col, 'q3'] + (3 * X_outliers.loc[col, 'iqr'])
X_outliers.loc[col, 'min'] = X[col].min()
X_outliers.loc[col, 'max'] = X[col].max()
X_outliers.loc[col, 'outliers'] = ((X[col] < X_outliers.loc[col, 'lower']) | (X[col] > X_outliers.loc[col,'upper'])).sum()
X_outliers.loc[col, 'outliers%'] = np.round(X_outliers.loc[col,
'outliers'] / len(X) *100)
# headd(X_outliers.head(10))
#Distribution of price
%matplotlib inline
fig, axs = plt.subplots(ncols=2, figsize=(14, 4))
fig.suptitle('Distribution of max guests (before and after removing large listings > 10)', weight='bold', fontsize=12)
# Before cleaning
x_axis=X['numberOfGuests'].dropna()
sns.distplot(pd.Series(x_axis, name='Max guests (before cleaning)'), ax=axs[0])
# Remove where price > 1000
condition = X[X['numberOfGuests'] > 400]
rows_to_drop = condition.index
print("You dropped {} rows.".format(condition.shape[0]))
X = X.drop(rows_to_drop, axis=0)
print("Dataset has {} rows, {} columns.".format(*X.shape))
#After cleaning
x_axis=X['numberOfGuests'].dropna()
sns.distplot(pd.Series(x_axis, name='Max guests (after cleaning)'), ax=axs[1]);
## Adding est. Annual Revenue
print("Dataset has {} rows, {} before engineering.".format(*X.shape))
avg_occupancy_per_week = 4
X['yield'] = avg_occupancy_per_week * y * 52
# cols_to_drop = ['cleaning_fee']
# df = df.drop(cols_to_drop, axis = 1)
print("Dataset has {} rows, {} columns.".format(*X.shape))
```
### Linear Regression
```
# scaler = StandardScaler()
# X_train = scaler.fit_transform(X_train)
# X_test = scaler.transform(X_test)
model = LinearRegression()
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
fig, ax = plt.subplots()
ax.scatter(y_pred, y_test, edgecolors=(0, 0, 1))
ax.plot([y_test.min(), y_test.max()], [y_test.min(), y_test.max()], 'r--', lw=3)
ax.set_xlabel('Predicted')
ax.set_ylabel('Actual')
plt.show();
# model evaluation for testing set
mae = mean_absolute_error(y_test, y_pred)
mse = mean_squared_error(y_test, y_pred)
rmse = math.sqrt(mean_squared_error(y_test, y_pred))
r2 = r2_score(y_test, y_pred)
print("The model performance for testing set")
print("--------------------------------------")
print('MAE: {}'.format(round(mae)))
print('MSE: {}'.format(round(mse)))
print('RMSE: {}'.format(round(rmse)))
print('R2: {}'.format(round(r2, 3)))
```
### Multiple Models
```
regressors = {
"XGBRegressor": XGBRegressor(),
"RandomForestRegressor": RandomForestRegressor(),
"DecisionTreeRegressor": DecisionTreeRegressor(),
"SVR": SVR(),
"NuSVR": NuSVR(),
"LinearSVR": LinearSVR(),
"KernelRidge": KernelRidge(),
"LinearRegression": LinearRegression(),
"Ridge":Ridge(),
"HuberRegressor": HuberRegressor(),
"PassiveAggressiveRegressor": PassiveAggressiveRegressor(),
"ARDRegression": ARDRegression(),
"BayesianRidge": BayesianRidge(),
"ElasticNet": ElasticNet(),
"OrthogonalMatchingPursuit": OrthogonalMatchingPursuit(),
}
df_models = pd.DataFrame(columns=['Model', 'Run_Time', 'MAE', 'MSE', 'R2', 'RMSE', 'RMSE_CV'])
for key in regressors:
print('✓',key)
start_time = time.time()
regressor = regressors[key]
model = regressor.fit(X_train, y_train)
y_pred = model.predict(X_test)
scores = cross_val_score(model,
X_train,
y_train,
scoring="neg_mean_squared_error",
cv=10)
row = {'Model': key,
'Run_Time': format(round((time.time() - start_time)/60,2)),
'MAE': round(mean_absolute_error(y_test, y_pred)),
'MSE': round(mean_squared_error(y_test, y_pred)),
'R2': round(r2_score(y_test, y_pred), 3),
'RMSE': round(np.sqrt(mean_squared_error(y_test, y_pred))),
'RMSE_CV': round(np.mean(np.sqrt(-scores)))
}
df_models = df_models.append(row, ignore_index=True)
df_models
df_models.sort_values(by='RMSE_CV', ascending=True)
```
### Focusing on XGB
```
hyperparameter_grid = {
'n_estimators': [100],
'max_depth': [2, 3, 5],
'learning_rate': [.001,.01]
}
random_cv = RandomizedSearchCV(
estimator=XGBRegressor(),
param_distributions=hyperparameter_grid,
cv=3,
n_iter=30,
scoring = 'neg_mean_absolute_error',
n_jobs = -1,
verbose = 0,
return_train_score = True,
random_state=13
)
random_cv.fit(X_train,y_train)
random_cv.best_estimator_
regressor = random_cv.best_estimator_
regressor.fit(X_train,y_train)
y_pred = regressor.predict(X_test)
# model evaluation for testing set
mae = mean_absolute_error(y_test, y_pred)
mse = mean_squared_error(y_test, y_pred)
rmse = math.sqrt(mean_squared_error(y_test, y_pred))
r2 = r2_score(y_test, y_pred)
print("The model performance for testing set")
print("--------------------------------------")
print('MAE: {}'.format(round(mae)))
print('MSE: {}'.format(round(mse)))
print('RMSE: {}'.format(round(rmse)))
print('R2: {}'.format(round(r2, 3)))
df = pd.DataFrame({'Actual': y_test, 'Predicted': np.around(y_pred)})
df
```
| github_jupyter |
```
import datetime as dt
import numpy as np
import pandas as pd
import panel as pn
pn.extension('tabulator')
```
The ``Tabulator`` widget allows displaying and editing a pandas DataFrame. The `Tabulator` is a largely backward compatible replacement for the [`DataFrame`](./DataFrame.ipynb) widget and will eventually replace it. It is built on the [Tabulator](http://tabulator.info/) library, which provides for a wide range of features.
For more information about listening to widget events and laying out widgets refer to the [widgets user guide](../../user_guide/Widgets.ipynb). Alternatively you can learn how to build GUIs by declaring parameters independently of any specific widgets in the [param user guide](../../user_guide/Param.ipynb). To express interactivity entirely using Javascript without the need for a Python server take a look at the [links user guide](../../user_guide/Param.ipynb).
#### Parameters:
For layout and styling related parameters see the [customization user guide](../../user_guide/Customization.ipynb).
##### Core
* **``aggregators``** (``dict``): A dictionary mapping from index name to an aggregator to be used for `hierarchical` multi-indexes (valid aggregators include 'min', 'max', 'mean' and 'sum'). If separate aggregators for different columns are required the dictionary may be nested as `{index_name: {column_name: aggregator}}`
* **``configuration``** (``dict``): A dictionary mapping used to specify tabulator options not explicitly exposed by panel.
* **``editors``** (``dict``): A dictionary mapping from column name to a bokeh `CellEditor` instance or tabulator editor specification.
* **``embed_content``** (``boolean``): Whether to embed the `row_content` or to dynamically fetch it when a row is expanded.
* **``expanded``** (``list``): The currently expanded rows as a list of integer indexes.
* **``filters``** (``list``): A list of client-side filter definitions that are applied to the table.
* **``formatters``** (``dict``): A dictionary mapping from column name to a bokeh `CellFormatter` instance or tabulator formatter specification.
* **``groupby``** (`list`): Groups rows in the table by one or more columns.
* **``header_align``** (``dict`` or ``str``): A mapping from column name to header alignment or a fixed header alignment, which should be one of `'left'`, `'center'`, `'right'`.
* **``header_filters``** (``boolean``/``dict``): A boolean enabling filters in the column headers or a dictionary providing filter definitions for specific columns.
* **``hierarchical``** (boolean, default=False): Whether to render multi-indexes as hierarchical index (note hierarchical must be enabled during instantiation and cannot be modified later)
* **``hidden_columns``** (`list`): List of columns to hide.
* **``layout``** (``str``, `default='fit_data_table'`): Describes the column layout mode with one of the following options `'fit_columns'`, `'fit_data'`, `'fit_data_stretch'`, `'fit_data_fill'`, `'fit_data_table'`.
* **``frozen_columns``** (`list`): List of columns to freeze, preventing them from scrolling out of frame. Column can be specified by name or index.
* **``frozen_rows``**: (`list`): List of rows to freeze, preventing them from scrolling out of frame. Rows can be specified by positive or negative index.
* **``page``** (``int``, `default=1`): Current page, if pagination is enabled.
* **``page_size``** (``int``, `default=20`): Number of rows on each page, if pagination is enabled.
* **``pagination``** (`str`, `default=None`): Set to `'local` or `'remote'` to enable pagination; by default pagination is disabled with the value set to `None`.
* **``row_content``** (``callable``): A function that receives the expanded row as input and should return a Panel object to render into the expanded region below the row.
* **``row_height``** (``int``, `default=30`): The height of each table row.
* **``selection``** (``list``): The currently selected rows as a list of integer indexes.
* **``selectable``** (`boolean` or `str` or `int`, `default=True`): Defines the selection mode:
* `True`
Selects rows on click. To select multiple use Ctrl-select, to select a range use Shift-select
* `False`
Disables selection
* `'checkbox'`
Adds a column of checkboxes to toggle selections
* `'checkbox-single'`
Same as 'checkbox' but header does not alllow select/deselect all
* `'toggle'`
Selection toggles when clicked
* `int`
The maximum number of selectable rows.
* **``selectable_rows``** (`callable`): A function that should return a list of integer indexes given a DataFrame indicating which rows may be selected.
* **``show_index``** (``boolean``, `default=True`): Whether to show the index column.
* **``text_align``** (``dict`` or ``str``): A mapping from column name to alignment or a fixed column alignment, which should be one of `'left'`, `'center'`, `'right'`.
* **`theme`** (``str``, `default='simple'`): The CSS theme to apply (note that changing the theme will restyle all tables on the page), which should be one of `'default'`, `'site'`, `'simple'`, `'midnight'`, `'modern'`, `'bootstrap'`, `'bootstrap4'`, `'materialize'`, `'bulma'`, `'semantic-ui'`, or `'fast'`.
* **``titles``** (``dict``): A mapping from column name to a title to override the name with.
* **``value``** (``pd.DataFrame``): The pandas DataFrame to display and edit
* **``widths``** (``dict``): A dictionary mapping from column name to column width in the rendered table.
##### Display
* **``disabled``** (``boolean``): Whether the widget is editable
* **``name``** (``str``): The title of the widget
##### Properties
* **``current_view``** (``DataFrame``): The current view of the table that is displayed, i.e. after sorting and filtering are applied
* **``selected_dataframe``** (``DataFrame``): A DataFrame reflecting the currently selected rows.
___
The ``Tabulator`` widget renders a DataFrame using an interactive grid, which allows directly editing the contents of the dataframe in place, with any changes being synced with Python. The `Tabulator` will usually determine the appropriate formatter appropriately based on the type of the data:
```
df = pd.DataFrame({
'int': [1, 2, 3],
'float': [3.14, 6.28, 9.42],
'str': ['A', 'B', 'C'],
'bool': [True, False, True],
'date': [dt.date(2019, 1, 1), dt.date(2020, 1, 1), dt.date(2020, 1, 10)],
'datetime': [dt.datetime(2019, 1, 1, 10), dt.datetime(2020, 1, 1, 12), dt.datetime(2020, 1, 10, 13)]
}, index=[1, 2, 3])
df_widget = pn.widgets.Tabulator(df)
df_widget
```
## Formatters
By default the widget will pick bokeh ``CellFormatter`` and ``CellEditor`` types appropriate to the dtype of the column. These may be overriden by explicit dictionaries mapping from the column name to the editor or formatter instance. For example below we create a ``SelectEditor`` instance to pick from four options in the ``str`` column and a ``NumberFormatter`` to customize the formatting of the float values:
```
from bokeh.models.widgets.tables import NumberFormatter, BooleanFormatter
bokeh_formatters = {
'float': NumberFormatter(format='0.00000'),
'bool': BooleanFormatter(),
}
pn.widgets.Tabulator(df, formatters=bokeh_formatters)
```
The list of valid Bokeh formatters includes:
* [BooleanFormatter](https://docs.bokeh.org/en/latest/docs/reference/models/widgets.tables.html#bokeh.models.widgets.tables.BooleanFormatter)
* [DateFormatter](https://docs.bokeh.org/en/latest/docs/reference/models/widgets.tables.html#bokeh.models.widgets.tables.DateFormatter)
* [NumberFormatter](https://docs.bokeh.org/en/latest/docs/reference/models/widgets.tables.html#bokeh.models.widgets.tables.NumberFormatter)
* [HTMLTemplateFormatter](https://docs.bokeh.org/en/latest/docs/reference/models/widgets.tables.html#bokeh.models.widgets.tables.HTMLTemplateFormatter)
* [StringFormatter](https://docs.bokeh.org/en/latest/docs/reference/models/widgets.tables.html#bokeh.models.widgets.tables.StringFormatter)
* [ScientificFormatter](https://docs.bokeh.org/en/latest/docs/reference/models/widgets.tables.html#bokeh.models.widgets.tables.ScientificFormatter)
However in addition to the formatters exposed by Bokeh it is also possible to provide valid formatters built into the Tabulator library. These may be defined either as a string or as a dictionary declaring the 'type' and other arguments, which are passed to Tabulator as the `formatterParams`:
```
tabulator_formatters = {
'float': {'type': 'progress', 'max': 10},
'bool': {'type': 'tickCross'}
}
pn.widgets.Tabulator(df, formatters=tabulator_formatters)
```
The list of valid Tabulator formatters can be found in the [Tabulator documentation](http://tabulator.info/docs/4.9/format#format-builtin).
## Editors
Just like the formatters, the `Tabulator` will natively understand the Bokeh `Editor` types. However, in the background it will replace most of them with equivalent editors natively supported by the tabulator library:
```
from bokeh.models.widgets.tables import CheckboxEditor, NumberEditor, SelectEditor, DateEditor, TimeEditor
bokeh_editors = {
'float': NumberEditor(),
'bool': CheckboxEditor(),
'str': SelectEditor(options=['A', 'B', 'C', 'D']),
}
pn.widgets.Tabulator(df[['float', 'bool', 'str']], editors=bokeh_editors)
```
Therefore it is often preferable to use one of the [Tabulator editors](http://tabulator.info/docs/5.0/edit#edit) directly. Note that in addition to the standard Tabulator editors the Tabulator widget also supports `'date'` and `'datetime'` editors:
```
from bokeh.models.widgets.tables import CheckboxEditor, NumberEditor, SelectEditor
bokeh_editors = {
'float': {'type': 'number', 'max': 10, 'step': 0.1},
'bool': {'type': 'tickCross', 'tristate': True, 'indeterminateValue': None},
'str': {'type': 'autocomplete', 'values': True},
'date': 'date',
'datetime': 'datetime'
}
edit_table = pn.widgets.Tabulator(df, editors=bokeh_editors)
edit_table
```
When editing a cell the data stored on the `Tabulator.value` is updated and you can listen to any changes using the usual `.param.watch(callback, 'value')` mechanism. However if you need to know precisely which cell was changed you may also attach an `on_edit` callback which will be passed a `TableEditEvent` containing the:
- `column`: Name of the edited column
- `row`: Integer index of the edited row
- `value`: The updated value
```
edit_table.on_edit(lambda e: print(e.column, e.row, e.value))
```
### Column layouts
By default the DataFrame widget will adjust the sizes of both the columns and the table based on the contents, reflecting the default value of the parameter: `layout="fit_data_table"`. Alternative modes allow manually specifying the widths of the columns, giving each column equal widths, or adjusting just the size of the columns.
#### Manual column widths
To manually adjust column widths provide explicit `widths` for each of the columns:
```
custom_df = pd._testing.makeMixedDataFrame()
pn.widgets.Tabulator(custom_df, widths={'index': 70, 'A': 50, 'B': 50, 'C': 70, 'D': 130})
```
You can also declare a single width for all columns this way:
```
pn.widgets.Tabulator(custom_df, widths=130)
```
#### Autosize columns
To automatically adjust the columns dependending on their content set `layout='fit_data'`:
```
pn.widgets.Tabulator(custom_df, layout='fit_data', width=400)
```
To ensure that the table fits all the data but also stretches to fill all the available space, set `layout='fit_data_stretch'`:
```
pn.widgets.Tabulator(custom_df, layout='fit_data_stretch', width=400)
```
The `'fit_data_fill'` option on the other hand won't stretch the last column but still fill the space:
```
pn.widgets.Tabulator(custom_df, layout='fit_data_fill', width=400)
```
Perhaps the most useful of these options is `layout='fit_data_table'` (and therefore the default) since this will automatically size both the columns and the table:
```
pn.widgets.Tabulator(custom_df, layout='fit_data_table')
```
#### Equal size
The simplest option is simply to allocate each column equal amount of size:
```
pn.widgets.Tabulator(custom_df, layout='fit_columns', width=650)
```
## Alignment
The content of a column or its header can be horizontally aligned with `text_align` and `header_align`. These two parameters accept either a string that globally defines the alignment or a dictionnary that declares which particular columns are meant to be aligned and how.
```
pn.widgets.Tabulator(df, header_align='center', text_align={'str': 'right', 'bool': 'center'}, widths=200)
```
## Styling
The ability to style the contents of a table based on its content and other considerations is very important. Thankfully pandas provides a powerful [styling API](https://pandas.pydata.org/pandas-docs/stable/user_guide/style.html), which can be used in conjunction with the `Tabulator` widget. Specifically the `Tabulator` widget exposes a `.style` attribute just like a `pandas.DataFrame` which lets the user apply custom styling using methods like `.apply` and `.applymap`. For a detailed guide to styling see the [Pandas documentation](https://pandas.pydata.org/pandas-docs/stable/user_guide/style.html).
Here we will demonstrate with a simple example, starting with a basic table:
```
style_df = pd.DataFrame(np.random.randn(10, 5), columns=list('ABCDE'))
styled = pn.widgets.Tabulator(style_df)
```
Next we define two functions which apply styling cell-wise (`color_negative_red`) and column-wise (`highlight_max`), which we then apply to the `Tabulator` using the `.style` API and then display the `styled` table:
```
def color_negative_red(val):
"""
Takes a scalar and returns a string with
the css property `'color: red'` for negative
strings, black otherwise.
"""
color = 'red' if val < 0 else 'black'
return 'color: %s' % color
def highlight_max(s):
'''
highlight the maximum in a Series yellow.
'''
is_max = s == s.max()
return ['background-color: yellow' if v else '' for v in is_max]
styled.style.applymap(color_negative_red).apply(highlight_max)
styled
```
## Theming
The Tabulator library ships with a number of themes, which are defined as CSS stylesheets. For that reason changing the theme on one table will affect all Tables on the page and it will usually be preferable to see the theme once at the class level like this:
```python
pn.widgets.Tabulator.theme = 'default'
```
For a full list of themes see the [Tabulator documentation](http://tabulator.info/docs/4.9/theme), however the default themes include:
- `'simple'`
- `'default'`
- `'midnight'`
- `'site'`
- `'modern'`
- `'bootstrap'`
- `'bootstrap4'`
- `'materialize'`
- `'semantic-ui'`
- `'bulma'`
## Selection
The `selection` parameter controls which rows in the table are selected and can be set from Python and updated by selecting rows on the frontend:
```
sel_df = pd.DataFrame(np.random.randn(10, 5), columns=list('ABCDE'))
select_table = pn.widgets.Tabulator(sel_df, selection=[0, 3, 7])
select_table
```
Once initialized, the ``selection`` parameter will return the integer indexes of the selected rows, while the ``selected_dataframe`` property will return a new DataFrame containing just the selected rows:
```
select_table.selection = [1, 4, 9]
select_table.selected_dataframe
```
The `selectable` parameter declares how the selections work.
- `True`: Selects rows on click. To select multiple use Ctrl-select, to select a range use Shift-select
- `False`: Disables selection
- `'checkbox'`: Adds a column of checkboxes to toggle selections
- `'checkbox-single'`: Same as `'checkbox'` but disables (de)select-all in the header
- `'toggle'`: Selection toggles when clicked
- Any positive `int`: A number that sets the maximum number of selectable rows
```
pn.widgets.Tabulator(sel_df, selection=[0, 3, 7], selectable='checkbox')
```
Additionally we can also disable selection for specific rows by providing a `selectable_rows` function. The function must accept a DataFrame and return a list of integer indexes indicating which rows are selectable, e.g. here we disable selection for every second row:
```
pn.widgets.Tabulator(sel_df, selectable_rows=lambda df: list(range(0, len(df), 2)))
```
### Freezing rows and columns
Sometimes your table will be larger than can be displayed in a single viewport, in which case scroll bars will be enabled. In such cases, you might want to make sure that certain information is always visible. This is where the `frozen_columns` and `frozen_rows` options come in.
#### Frozen columns
When you have a large number of columns and can't fit them all on the screen you might still want to make sure that certain columns do not scroll out of view. The `frozen_columns` option makes this possible by specifying a list of columns that should be frozen, e.g. `frozen_columns=['index']` will freeze the index column:
```
wide_df = pd._testing.makeCustomDataframe(10, 10, r_idx_names=['index'])
pn.widgets.Tabulator(wide_df, frozen_columns=['index'], width=400)
```
#### Frozen rows
Another common scenario is when you have certain rows with special meaning, e.g. aggregates that summarize the information in the rest of the table. In this case you may want to freeze those rows so they do not scroll out of view. You can achieve this by setting a list of `frozen_rows` by integer index (which can be positive or negative, where negative values are relative to the end of the table):
```
date_df = pd._testing.makeTimeDataFrame().iloc[:10]
agg_df = pd.concat([date_df, date_df.median().to_frame('Median').T, date_df.mean().to_frame('Mean').T])
agg_df.index= agg_df.index.map(str)
pn.widgets.Tabulator(agg_df, frozen_rows=[-2, -1], width=400)
```
## Row contents
A table can only display so much information without becoming difficult to scan. We may want to render additional information to a table row to provide additional context. To make this possible you can provide a `row_content` function which is given the table row as an argument and should return a panel object that will be rendered into an expanding region below the row. By default the contents are fetched dynamically whenever a row is expanded, however using the `embed_content` parameter we can embed all the content.
Below we create a periodic table of elements where the Wikipedia page for each element will be rendered into the expanded region:
```
from bokeh.sampledata.periodic_table import elements
periodic_df = elements[['atomic number', 'name', 'atomic mass', 'metal', 'year discovered']].set_index('atomic number')
content_fn = lambda row: pn.pane.HTML(
f'<iframe src="http://en.wikipedia.org/wiki/{row["name"]}?printable=yes" width="100%" height="300px"></iframe>',
sizing_mode='stretch_width'
)
periodic_table = pn.widgets.Tabulator(
periodic_df, height=500, layout='fit_columns', sizing_mode='stretch_width',
row_content=content_fn, embed_content=True
)
periodic_table
```
The currently expanded rows can be accessed (and set) on the `expanded` parameter:
```
periodic_table.expanded
```
## Grouping
Another useful option is the ability to group specific rows together, which can be achieved using `groups` parameter. The `groups` parameter should be composed of a dictionary mapping from the group titles to the column names:
```
pn.widgets.Tabulator(date_df, groups={'Group 1': ['A', 'B'], 'Group 2': ['C', 'D']})
```
## Groupby
In addition to grouping columns we can also group rows by the values along one or more columns:
```
from bokeh.sampledata.autompg import autompg
pn.widgets.Tabulator(autompg, groupby=['yr', 'origin'], height=240)
```
### Hierarchical Multi-index
The `Tabulator` widget can also render a hierarchical multi-index and aggregate over specific categories. If a DataFrame with a hierarchical multi-index is supplied and the `hierarchical` is enabled the widget will group data by the categories in the order they are defined in. Additionally for each group in the multi-index an aggregator may be provided which will aggregate over the values in that category.
For example we may load population data for locations around the world broken down by sex and age-group. If we specify aggregators over the 'AgeGrp' and 'Sex' indexes we can see the aggregated values for each of those groups (note that we do not have to specify an aggregator for the outer index since we specify the aggregators over the subgroups in this case the 'Sex'):
```
from bokeh.sampledata.population import data as population_data
pop_df = population_data[population_data.Year == 2020].set_index(['Location', 'AgeGrp', 'Sex'])[['Value']]
pn.widgets.Tabulator(value=pop_df, hierarchical=True, aggregators={'Sex': 'sum', 'AgeGrp': 'sum'}, height=400)
```
## Pagination
When working with large tables we sometimes can't send all the data to the browser at once. In these scenarios we can enable pagination, which will fetch only the currently viewed data from the server backend. This may be enabled by setting `pagination='remote'` and the size of each page can be set using the `page_size` option:
```
large_df = pd._testing.makeCustomDataframe(100000, 5)
%%time
paginated_table = pn.widgets.Tabulator(large_df, pagination='remote', page_size=10)
paginated_table
```
Contrary to the `'remote'` option, `'local'` pagination entirely loads the data but still allows to display it on multiple pages.
```
%%time
medium_df = pd._testing.makeCustomDataframe(10000, 5)
paginated_table = pn.widgets.Tabulator(large_df, pagination='local', page_size=10)
paginated_table
```
## Filtering
A very common scenario is that you want to attach a number of filters to a table in order to view just a subset of the data. You can achieve this through callbacks or other reactive approaches but the `.add_filter` method makes it much easier.
#### Constant and Widget filters
The simplest approach to filtering is to select along a column with a constant or dynamic value. The `.add_filter` method allows passing in constant values, widgets and parameters. If a widget or parameter is provided the table will watch the object for changes in the value and update the data in response. The filtering will depend on the type of the constant or dynamic value:
- scalar: Filters by checking for equality
- `tuple`: A tuple will be interpreted as range.
- `list`/`set`: A list or set will be interpreted as a set of discrete scalars and the filter will check if the values in the column match any of the items in the list.
As an example we will create a DataFrame with some data of mixed types:
```
mixed_df = pd._testing.makeMixedDataFrame()
filter_table = pn.widgets.Tabulator(mixed_df)
filter_table
```
Now we will start adding filters one-by-one, e.g. to start with we add a filter for the `'A'` column, selecting a range from 0 to 3:
```
filter_table.add_filter((0, 3), 'A')
```
Next we add dynamic widget based filter, a `RangeSlider` which allows us to further narrow down the data along the `'A'` column:
```
slider = pn.widgets.RangeSlider(start=0, end=3, name='A Filter')
filter_table.add_filter(slider, 'A')
```
Lastly we will add a `MultiSelect` filter along the `'C'` column:
```
select = pn.widgets.MultiSelect(options=['foo1', 'foo2', 'foo3', 'foo4', 'foo5'], name='C Filter')
filter_table.add_filter(select, 'C')
```
Now let's display the table alongside the widget based filters:
```
pn.Row(
pn.Column(slider, select),
filter_table
)
```
After filtering you can inspect the current view with the `current_view` property:
```
filter_table.current_view
```
#### Function based filtering
For more complex filtering tasks you can supply a function that should accept the DataFrame to be filtered as the first argument and must return a filtered copy of the data. Let's start by loading some data.
```
import sqlite3
from bokeh.sampledata.movies_data import movie_path
con = sqlite3.Connection(movie_path)
movies_df = pd.read_sql('SELECT Title, Year, Genre, Director, Writer, imdbRating from omdb', con)
movies_df = movies_df[~movies_df.Director.isna()]
movies_table = pn.widgets.Tabulator(movies_df, pagination='remote', layout='fit_columns', width=800)
```
By using the `pn.bind` function, which binds widget and parameter values to a function, complex filtering can be achieved. E.g. here we will add a filter function that uses tests whether the string or regex is contained in the 'Director' column of a listing of thousands of movies:
```
director_filter = pn.widgets.TextInput(name='Director filter', value='Chaplin')
def contains_filter(df, pattern, column):
if not pattern:
return df
return df[df[column].str.contains(pattern)]
movies_table.add_filter(pn.bind(contains_filter, pattern=director_filter, column='Director'))
pn.Row(director_filter, movies_table)
```
### Client-side filtering
In addition to the Python API the Tabulator widget also offers a client-side filtering API, which can be exposed through `header_filters` or by manually adding filters to the rendered Bokeh model. The API for declaring header filters is almost identical to the API for defining [Editors](#Editors). The `header_filters` can either be enabled by setting it to `True` or by manually supplying filter types for each column. The types of filters supports all the same options as the editors, in fact if you do not declare explicit `header_filters` the tabulator will simply use the defined `editors` to determine the correct filter type:
```
bokeh_editors = {
'float': {'type': 'number', 'max': 10, 'step': 0.1},
'bool': {'type': 'tickCross', 'tristate': True, 'indeterminateValue': None},
'str': {'type': 'autocomplete', 'values': True}
}
header_filter_table = pn.widgets.Tabulator(
df[['float', 'bool', 'str']], height=140, width=400, layout='fit_columns',
editors=bokeh_editors, header_filters=True
)
header_filter_table
```
When a filter is applied client-side the `filters` parameter is synced with Python. The definition of `filters` looks something like this:
```
[{'field': 'Director', 'type': '=', 'value': 'Steven Spielberg'}]
```
Try applying a filter and then inspect the `filters` parameter:
```
header_filter_table.filters
```
For all supported filtering types see the [Tabulator Filtering documentation](http://tabulator.info/docs/4.9/filter).
If we want to change the filter type for the `header_filters` we can do so in the definition by supplying a dictionary indexed by the column names and then either providing a dictionary which may define the `'type'`, a comparison `'func'`, a `'placeholder'` and any additional keywords supported by the particular filter type.
```
movie_filters = {
'Title': {'type': 'input', 'func': 'like', 'placeholder': 'Enter title'},
'Year': {'placeholder': 'Enter year'},
'Genre': {'type': 'input', 'func': 'like', 'placeholder': 'Enter genre'},
'Director': {'type': 'input', 'func': 'like', 'placeholder': 'Enter director'},
'Writer': {'type': 'input', 'func': 'like', 'placeholder': 'Enter writer'},
'imdbRating': {'type': 'number', 'func': '>=', 'placeholder': 'Enter minimum rating'}
}
filter_table = pn.widgets.Tabulator(
movies_df, pagination='remote', layout='fit_columns', page_size=10, sizing_mode='stretch_width',
header_filters=movie_filters
)
filter_table
```
## Downloading
The `Tabulator` also supports triggering a download of the data as a CSV or JSON file dependending on the filename. The download can be triggered with the `.download()` method, which optionally accepts the filename as the first argument.
To trigger the download client-side (i.e. without involving the server) you can use the `.download_menu` method which creates a `TextInput` and `Button` widget, which allow setting the filename and triggering the download respectively:
```
download_df = pd.DataFrame(np.random.randn(10, 5), columns=list('ABCDE'))
download_table = pn.widgets.Tabulator(download_df)
filename, button = download_table.download_menu(
text_kwargs={'name': 'Enter filename', 'value': 'default.csv'},
button_kwargs={'name': 'Download table'}
)
pn.Row(
pn.Column(filename, button),
download_table
)
```
## Streaming
When we are monitoring some source of data that updates over time, we may want to update the table with the newly arriving data. However, we do not want to transmit the entire dataset each time. To handle efficient transfer of just the latest data, we can use the `.stream` method on the `Tabulator` object:
```
stream_df = pd.DataFrame(np.random.randn(10, 5), columns=list('ABCDE'))
stream_table = pn.widgets.Tabulator(stream_df, layout='fit_columns', width=450)
stream_table
```
As example, we will schedule a periodic callback that streams new data every 1000ms (i.e. 1s) five times in a row:
```
def stream_data(follow=True):
stream_df = pd.DataFrame(np.random.randn(10, 5), columns=list('ABCDE'))
stream_table.stream(stream_df, follow=follow)
pn.state.add_periodic_callback(stream_data, period=1000, count=5)
```
If you are viewing this example with a live Python kernel you will be able to watch the table update and scroll along. If we want to disable the scrolling behavior, we can set `follow=False`:
```
stream_data(follow=False)
```
## Patching
In certain cases we don't want to update the table with new data but just patch existing data.
```
patch_table = pn.widgets.Tabulator(df[['int', 'float', 'str', 'bool']])
patch_table
```
The easiest way to patch the data is by supplying a dictionary as the patch value. The dictionary should have the following structure:
```python
{
column: [
(index: int or slice, value),
...
],
...
}
```
As an example, below we will patch the 'bool' and 'int' columns. On the `'bool'` column we will replace the 0th and 2nd row and on the `'int'` column we replace the first two rows:
```
patch_table.patch({
'bool': [
(0, False),
(2, False)
],
'int': [
(slice(0, 2), [3, 2])
]
})
```
## Static Configuration
Panel does not expose all options available from Tabulator, if a desired option is not natively supported, it can be set via the `configuration` argument.
This dictionary can be seen as a base dictionary which the tabulator object fills and passes to the Tabulator javascript-library.
As an example, we can turn off sorting and resizing of columns by disabling the `headerSort` and `resizable` options.
```
df = pd.DataFrame({
'int': [1, 2, 3],
'float': [3.14, 6.28, 9.42],
'str': ['A', 'B', 'C'],
'bool': [True, False, True],
'date': [dt.date(2019, 1, 1), dt.date(2020, 1, 1), dt.date(2020, 1, 10)]
}, index=[1, 2, 3])
df_widget = pn.widgets.Tabulator(df, configuration={'columnDefaults': {
'resizable': False,
'headerSort': True
}})
df_widget.servable()
```
These and other available tabulator options are listed at http://tabulator.info/docs/4.9/options.
Obviously not all options will work though, especially any settable callbacks and options which are set by the internal panel tabulator module (for example the `columns` option).
Additionally it should be noted that the configuration parameter is not responsive so it can only be set at instantiation time.
| github_jupyter |
```
# Copyright 2022 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
<table align="left">
<td>
<a href="https://colab.research.google.com/github/GoogleCloudPlatform/vertex-ai-samples/blob/main/vertex-ai-samples/notebooks/community/feature_store/mobile_gaming_feature_store.ipynb">
<img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
</a>
</td>
<td>
<a href="https://github.com/inardini/vertex-ai-samples/blob/main/vertex-ai-samples/notebooks/community/feature_store/mobile_gaming_feature_store.ipynb">
<img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
View on GitHub
</a>
</td>
</table>
## Overview
Imagine you are a member of the Data Science team working on the same Mobile Gaming application reported in the [Churn prediction for game developers using Google Analytics 4 (GA4) and BigQuery ML](https://cloud.google.com/blog/topics/developers-practitioners/churn-prediction-game-developers-using-google-analytics-4-ga4-and-bigquery-ml) blog post.
Your team successfully implemented a model that determines the likelihood of specific users returning to your app and consumes that insight to drive marketing incentives. As a result, the company consolidates its user base.
Now, businesses want to use that information in real-time to monetize it by implementing a conditional ads system. In particular, each time a user plays with the app, they want to display ads depending on the customer demographic, behavioral information and the resulting propensity of return. Of course, the new application should work with a minimum impact on the user experience.
Given the business challenge, the team is required to design and build a possible serving system which needs to minimize real-time prediction serving latency.
The assumptions are:
1. Predictions would be delivered synchronously
2. Scalability, support for multiple ML frameworks and security are essential.
3. Only demographic features (country, operating system and language) passed in real time.
2. The system would be able to handle behavioral features as static reference features calculated each 24h (offline batch feature engineering job).
3. It has to migrate training serving skew by a timestamp data model, a point-in-time lookups to avoid data leakage and a feature distribution monitoring service.
Based on those assumptions, low read-latency lookup data store and a performing serving engine are needed. Indeed, about the data store, even if you can implement governance on BigQuery, it is still not optimized for singleton lookup operations. Also, the solution need a low overhead serving system that can seamlessly scale up and down based on requests.
Last year, Google Cloud announced Vertex AI, a managed machine learning (ML) platform that allows data science teams to accelerate the deployment and maintenance of ML models. The platform is composed of several building blocks and two of them are Vertex AI Feature store and Vertex AI prediction.
With Vertex AI Feature store, you have a managed service for low latency scalable feature serving. It also provides a centralized feature repository with easy APIs to search & discover features and feature monitoring capabilities to track drift and other quality issues. With Vertex AI Prediction, you will deploy models into production more easily with online serving via HTTP or batch prediction for bulk scoring. It offers a unified scalable framework to deploy custom models trained in TensorFlow, scikit or XGB, as well as BigQuery ML and AutoML models, and on a broad range of machine types and GPUs.
Below the high level picture puts together once the team decides to go with Google Cloud:
<img src="./assets/solution_overview_final.png"/>
In order:
1. Once you create historical features, they are ingested into Vertex AI Feature store
2. Then you can train and deploy the model using BigQuery (or AutoML)
3. Once the model is deployed, the ML serving engine will receive a prediction request passing entity ID and demographic attributes.
4. Features related to a specific entity will be retrieved from the Vertex AI Feature store and passed them as inputs to the model for online prediction.
5. The predictions will be returned back to the activation layer.
### Dataset
The dataset is the public sample export data from an actual mobile game app called "Flood It!" (Android, iOS)
### Objective
In the following notebook, you will learn the role of Vertex AI Feature Store in a scenario when the user's activities within the first 24 hours of first user engagement and the gaming platform would consume in order to offer conditional ads.
**Notice that we assume that already know how to set up a Vertex AI Feature store. In case you are not, please check out [this detailed notebook](https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/main/notebooks/official/feature_store/gapic-feature-store.ipynb).**
At the end, you will more confident about how Vertex AI Feature store
1. Provide a centralized feature repository with easy APIs to search & discover features and fetch them for training/serving.
2. Simplify deployments of models for Online Prediction, via low latency scalable feature serving.
3. Mitigate training serving skew and data leakage by performing point in time lookups to fetch historical data for training.
### Costs
This tutorial uses billable components of Google Cloud:
* Vertex AI
* BigQuery
* Cloud Storage
Learn about [Vertex AI
pricing](https://cloud.google.com/vertex-ai/pricing) and [Cloud Storage
pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
Calculator](https://cloud.google.com/products/calculator/)
to generate a cost estimate based on your projected usage.
### Set up your local development environment
**If you are using Colab or Google Cloud Notebooks**, your environment already meets
all the requirements to run this notebook. You can skip this step.
**Otherwise**, make sure your environment meets this notebook's requirements.
You need the following:
* The Google Cloud SDK
* Git
* Python 3
* virtualenv
* Jupyter notebook running in a virtual environment with Python 3
The Google Cloud guide to [Setting up a Python development
environment](https://cloud.google.com/python/setup) and the [Jupyter
installation guide](https://jupyter.org/install) provide detailed instructions
for meeting these requirements. The following steps provide a condensed set of
instructions:
1. [Install and initialize the Cloud SDK.](https://cloud.google.com/sdk/docs/)
1. [Install Python 3.](https://cloud.google.com/python/setup#installing_python)
1. [Install
virtualenv](https://cloud.google.com/python/setup#installing_and_using_virtualenv)
and create a virtual environment that uses Python 3. Activate the virtual environment.
1. To install Jupyter, run `pip3 install jupyter` on the
command-line in a terminal shell.
1. To launch Jupyter, run `jupyter notebook` on the command-line in a terminal shell.
1. Open this notebook in the Jupyter Notebook Dashboard.
### Install additional packages
Install additional package dependencies not installed in your notebook environment, such as {XGBoost, AdaNet, or TensorFlow Hub TODO: Replace with relevant packages for the tutorial}. Use the latest major GA version of each package.
```
import os
# The Google Cloud Notebook product has specific requirements
IS_GOOGLE_CLOUD_NOTEBOOK = os.path.exists("/opt/deeplearning/metadata/env_version")
# Google Cloud Notebook requires dependencies to be installed with '--user'
USER_FLAG = ""
if IS_GOOGLE_CLOUD_NOTEBOOK:
USER_FLAG = "--user"
! pip3 install --upgrade pip
! pip3 install {USER_FLAG} --upgrade git+https://github.com/googleapis/python-aiplatform.git@main -q --no-warn-conflicts
! pip3 install {USER_FLAG} --upgrade pandas==1.3.5 -q --no-warn-conflicts
! pip3 install {USER_FLAG} --upgrade google-cloud-bigquery==2.24.0 -q --no-warn-conflicts
! pip3 install {USER_FLAG} --upgrade tensorflow==2.8.0 -q --no-warn-conflicts
```
### Restart the kernel
After you install the additional packages, you need to restart the notebook kernel so it can find the packages.
```
# Automatically restart kernel after installs
import os
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
```
## Before you begin
### Set up your Google Cloud project
**The following steps are required, regardless of your notebook environment.**
1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
1. [Make sure that billing is enabled for your project](https://cloud.google.com/billing/docs/how-to/modify-project).
1. [Enable the Vertex AI API and Compute Engine API](https://console.cloud.google.com/flows/enableapi?apiid=aiplatform.googleapis.com,compute_component).
1. If you are running this notebook locally, you will need to install the [Cloud SDK](https://cloud.google.com/sdk).
1. Enter your project ID in the cell below. Then run the cell to make sure the
Cloud SDK uses the right project for all the commands in this notebook.
**Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
#### Set your project ID
**If you don't know your project ID**, you may be able to get your project ID using `gcloud`.
```
import os
PROJECT_ID = ""
# Get your Google Cloud project ID from gcloud
if not os.getenv("IS_TESTING"):
shell_output = !gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID: ", PROJECT_ID)
```
Otherwise, set your project ID here.
```
if PROJECT_ID == "" or PROJECT_ID is None:
PROJECT_ID = "" # @param {type:"string"}
!gcloud config set project '' #change it
```
#### Timestamp
If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append it onto the name of resources you create in this tutorial.
```
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
```
### Authenticate your Google Cloud account
**If you are using Google Cloud Notebooks**, your environment is already
authenticated. Skip this step.
**If you are using Colab**, run the cell below and follow the instructions
when prompted to authenticate your account via oAuth.
**Otherwise**, follow these steps:
1. In the Cloud Console, go to the [**Create service account key**
page](https://console.cloud.google.com/apis/credentials/serviceaccountkey).
2. Click **Create service account**.
3. In the **Service account name** field, enter a name, and
click **Create**.
4. In the **Grant this service account access to project** section, click the **Role** drop-down list. Type "Vertex AI"
into the filter box, and select
**Vertex AI Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
5. Click *Create*. A JSON file that contains your key downloads to your
local environment.
6. Enter the path to your service account key as the
`GOOGLE_APPLICATION_CREDENTIALS` variable in the cell below and run the cell.
```
import os
import sys
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# The Google Cloud Notebook product has specific requirements
IS_GOOGLE_CLOUD_NOTEBOOK = os.path.exists("/opt/deeplearning/metadata/env_version")
# If on Google Cloud Notebooks, then don't execute this code
if not IS_GOOGLE_CLOUD_NOTEBOOK:
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
%env GOOGLE_APPLICATION_CREDENTIALS ''
```
### Create a Cloud Storage bucket
**The following steps are required, regardless of your notebook environment.**
Set the name of your Cloud Storage bucket below. It must be unique across all
Cloud Storage buckets.
You may also change the `REGION` variable, which is used for operations
throughout the rest of this notebook. Make sure to [choose a region where Vertex AI services are
available](https://cloud.google.com/vertex-ai/docs/general/locations#available_regions). You may
not use a Multi-Regional Storage bucket for training with Vertex AI.
```
BUCKET_URI = "" # @param {type:"string"}
REGION = "[your-region]" # @param {type:"string"}
if BUCKET_URI == "" or BUCKET_URI is None or BUCKET_URI == "gs://[your-bucket-name]":
BUCKET_URI = "gs://" + PROJECT_ID + "-aip-" + TIMESTAMP
if REGION == "[your-region]":
REGION = "us-central1"
```
**Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
```
! gsutil mb -l $REGION -p $PROJECT_ID $BUCKET_URI
```
Run the following cell to grant access to your Cloud Storage resources from Vertex AI Feature store
```
! gsutil uniformbucketlevelaccess set on $BUCKET_URI
```
Finally, validate access to your Cloud Storage bucket by examining its contents:
```
! gsutil ls -al $BUCKET_URI
```
### Create a Bigquery dataset
```
BQ_DATASET = "Mobile_Gaming" # @param {type:"string"}
LOCATION = "US"
!bq mk --location=$LOCATION --dataset $PROJECT_ID:$BQ_DATASET
```
### Import libraries
```
# General
import os
import sys
import time
# Data Engineering
import pandas as pd
# Vertex AI and its Feature Store
from google.cloud import aiplatform as vertex_ai
from google.cloud import bigquery
# EntityType
from google.cloud.aiplatform import Feature, Featurestore
```
### Define constants
```
# Data Engineering and Feature Engineering
FEATURES_TABLE = "wide_features_table" # @param {type:"string"}
MIN_DATE = "2018-10-03"
MAX_DATE = "2018-10-04"
FEATURES_TABLE_DAY_ONE = f"wide_features_table_{MIN_DATE}"
FEATURES_TABLE_DAY_TWO = f"wide_features_table_{MAX_DATE}"
FEATURESTORE_ID = "mobile_gaming" # @param {type:"string"}
ENTITY_TYPE_ID = "user"
# BQ Model Training and Deployment
MODEL_NAME = f"churn_logit_classifier_{TIMESTAMP}"
MODEL_TYPE = "LOGISTIC_REG"
AUTO_CLASS_WEIGHTS = "TRUE"
MAX_ITERATIONS = "50"
INPUT_LABEL_COLS = "churned"
JOB_ID = f"extract_{MODEL_NAME}_{TIMESTAMP}"
MODEL_SOURCE = bigquery.model.ModelReference.from_api_repr(
{"projectId": PROJECT_ID, "datasetId": BQ_DATASET, "modelId": MODEL_NAME}
)
SERVING_DIR = "serving_dir"
DESTINATION_URI = f"{BUCKET_URI}/model"
EXTRACT_JOB_CONFIG = bigquery.ExtractJobConfig(destination_format="ML_TF_SAVED_MODEL")
VERSION = "v1"
SERVING_CONTAINER_IMAGE_URI = (
"us-docker.pkg.dev/vertex-ai/prediction/tf2-cpu.2-7:latest"
)
ENDPOINT_NAME = "mobile_gaming_churn"
DEPLOYED_MODEL_NAME = f"churn_logistic_classifier_{VERSION}"
# Vertex AI Feature store
ONLINE_STORE_NODES_COUNT = 3
ENTITY_ID = "user"
API_ENDPOINT = f"{REGION}-aiplatform.googleapis.com"
FEATURE_TIME = "user_first_engagement"
ENTITY_ID_FIELD = "user_pseudo_id"
BQ_SOURCE_URI_DAY_ONE = f"bq://{PROJECT_ID}.{BQ_DATASET}.{FEATURES_TABLE_DAY_ONE}"
BQ_SOURCE_URI_DAY_TWO = f"bq://{PROJECT_ID}.{BQ_DATASET}.{FEATURES_TABLE_DAY_TWO}"
BQ_DESTINATION_OUTPUT_URI = f"bq://{PROJECT_ID}.{BQ_DATASET}.train_snapshot_{TIMESTAMP}"
SERVING_FEATURE_IDS = {"customer": ["*"]}
READ_INSTANCES_TABLE = f"ground_truth_{TIMESTAMP}"
READ_INSTANCES_URI = f"bq://{PROJECT_ID}.{BQ_DATASET}.{READ_INSTANCES_TABLE}"
# Vertex AI AutoML model
DATASET_NAME = f"churn_mobile_gaming_{TIMESTAMP}"
AUTOML_TRAIN_JOB_NAME = f"automl_classifier_training_{TIMESTAMP}"
AUTOML_MODEL_NAME = f"churn_automl_classifier_{TIMESTAMP}"
MODEL_DEPLOYED_NAME = "churn_automl_classifier_v1"
SERVING_MACHINE_TYPE = "n1-highcpu-4"
MIN_NODES = 1
MAX_NODES = 1
```
### Helpers
```
def run_bq_query(query: str):
"""
An helper function to run a BigQuery job
Args:
query: a formatted SQL query
Returns:
None
"""
try:
job = bq_client.query(query)
_ = job.result()
except RuntimeError as error:
print(error)
def upload_model(
display_name: str,
serving_container_image_uri: str,
artifact_uri: str,
sync: bool = True,
) -> vertex_ai.Model:
"""
Args:
display_name: The name of Vertex AI Model artefact
serving_container_image_uri: The uri of the serving image
artifact_uri: The uri of artefact to import
sync:
Returns: Vertex AI Model
"""
model = vertex_ai.Model.upload(
display_name=display_name,
artifact_uri=artifact_uri,
serving_container_image_uri=serving_container_image_uri,
sync=sync,
)
model.wait()
print(model.display_name)
print(model.resource_name)
return model
def create_endpoint(display_name: str) -> vertex_ai.Endpoint:
"""
An utility to create a Vertex AI Endpoint
Args:
display_name: The name of Endpoint
Returns: Vertex AI Endpoint
"""
endpoint = vertex_ai.Endpoint.create(display_name=display_name)
print(endpoint.display_name)
print(endpoint.resource_name)
return endpoint
def deploy_model(
model: vertex_ai.Model,
machine_type: str,
endpoint: vertex_ai.Endpoint = None,
deployed_model_display_name: str = None,
min_replica_count: int = 1,
max_replica_count: int = 1,
sync: bool = True,
) -> vertex_ai.Model:
"""
An helper function to deploy a Vertex AI Endpoint
Args:
model: A Vertex AI Model
machine_type: The type of machine to serve the model
endpoint: An Vertex AI Endpoint
deployed_model_display_name: The name of the model
min_replica_count: Minimum number of serving replicas
max_replica_count: Max number of serving replicas
sync: Whether to execute method synchronously
Returns: vertex_ai.Model
"""
model_deployed = model.deploy(
endpoint=endpoint,
deployed_model_display_name=deployed_model_display_name,
machine_type=machine_type,
min_replica_count=min_replica_count,
max_replica_count=max_replica_count,
sync=sync,
)
model_deployed.wait()
print(model_deployed.display_name)
print(model_deployed.resource_name)
return model_deployed
def endpoint_predict_sample(
instances: list, endpoint: vertex_ai.Endpoint
) -> vertex_ai.models.Prediction:
"""
An helper function to get prediction from Vertex AI Endpoint
Args:
instances: The list of instances to score
endpoint: An Vertex AI Endpoint
Returns:
vertex_ai.models.Prediction
"""
prediction = endpoint.predict(instances=instances)
print(prediction)
return prediction
def simulate_prediction(
endpoint: vertex_ai.Endpoint, online_sample: dict
) -> vertex_ai.models.Prediction:
"""
An helper function to simulate online prediction with customer entity type
- format entities for prediction
- retrive static features with a singleton lookup operations from Vertex AI Feature store
- run the prediction request and get back the result
Args:
endpoint:
online_sample:
Returns:
vertex_ai.models.Prediction
"""
online_features = pd.DataFrame.from_dict(online_sample)
entity_ids = online_features["entity_id"].tolist()
customer_aggregated_features = customer_entity_type.read(
entity_ids=entity_ids,
feature_ids=[
"cnt_user_engagement",
"cnt_level_start_quickplay",
"cnt_level_end_quickplay",
"cnt_level_complete_quickplay",
"cnt_level_reset_quickplay",
"cnt_post_score",
"cnt_spend_virtual_currency",
"cnt_ad_reward",
"cnt_challenge_a_friend",
"cnt_completed_5_levels",
"cnt_use_extra_steps",
],
)
prediction_sample_df = pd.merge(
customer_aggregated_features.set_index("entity_id"),
online_features.set_index("entity_id"),
left_index=True,
right_index=True,
).reset_index(drop=True)
prediction_sample = prediction_sample_df.to_dict("records")
prediction = endpoint.predict(prediction_sample)
return prediction
```
# Setting the Online (real-time) prediction scenario
As we mentioned at the beginning, this section would simulate the original but this time we introduce Vertex AI for online (real-time) serving. In particular, we will
1. Create static features including demographic and behavioral attibutes
2. Training a simple BQML model
3. Export and deploy the model to Vertex AI endpoint
<img src="./assets/data_processing.png"/>
## Initiate clients
```
bq_client = bigquery.Client(project=PROJECT_ID, location=LOCATION)
vertex_ai.init(project=PROJECT_ID, location=REGION, staging_bucket=BUCKET_URI)
```
## Data and Feature Engineering
The original dataset contains raw event data we cannot ingest in the feature store as they are.
In this section, we will pre-process the raw data into an appropriate format.
**Notice we simulate those transformations in different point of time (day one and day two).**
### Label, Demographic and Behavioral Transformations
This section is based on the [Churn prediction for game developers using Google Analytics 4 (GA4) and BigQuery ML](https://cloud.google.com/blog/topics/developers-practitioners/churn-prediction-game-developers-using-google-analytics-4-ga4-and-bigquery-ml?utm_source=linkedin&utm_medium=unpaidsoc&utm_campaign=FY21-Q2-Google-Cloud-Tech-Blog&utm_content=google-analytics-4&utm_term=-) blog article by Minhaz Kazi and Polong Lin
```
preprocess_sql_query = f"""
CREATE OR REPLACE TABLE
`{PROJECT_ID}.{BQ_DATASET}.{FEATURES_TABLE}` AS
WITH
# query to create label --------------------------------------------------------------------------------
get_label AS (
SELECT
user_pseudo_id,
user_first_engagement,
user_last_engagement,
# EXTRACT(MONTH from TIMESTAMP_MICROS(user_first_engagement)) as month,
# EXTRACT(DAYOFYEAR from TIMESTAMP_MICROS(user_first_engagement)) as julianday,
# EXTRACT(DAYOFWEEK from TIMESTAMP_MICROS(user_first_engagement)) as dayofweek,
#add 24 hr to user's first touch
(user_first_engagement + 86400000000) AS ts_24hr_after_first_engagement,
#churned = 1 if last_touch within 24 hr of app installation, else 0
IF (user_last_engagement < (user_first_engagement + 86400000000),
1,
0 ) AS churned,
#bounced = 1 if last_touch within 10 min, else 0
IF (user_last_engagement <= (user_first_engagement + 600000000),
1,
0 ) AS bounced,
FROM
(
SELECT
user_pseudo_id,
MIN(event_timestamp) AS user_first_engagement,
MAX(event_timestamp) AS user_last_engagement
FROM
`firebase-public-project.analytics_153293282.events_*`
WHERE event_name="user_engagement"
GROUP BY
user_pseudo_id
)
GROUP BY 1,2,3),
# query to create class weights --------------------------------------------------------------------------------
get_class_weights AS (
SELECT
CAST(COUNT(*) / (2*(COUNT(*) - SUM(churned))) AS STRING) AS class_weight_zero,
CAST(COUNT(*) / (2*SUM(churned)) AS STRING) AS class_weight_one,
FROM
get_label
),
# query to extract demographic data for each user ---------------------------------------------------------
get_demographic_data AS (
SELECT * EXCEPT (row_num)
FROM (
SELECT
user_pseudo_id,
geo.country as country,
device.operating_system as operating_system,
device.language as language,
ROW_NUMBER() OVER (PARTITION BY user_pseudo_id ORDER BY event_timestamp DESC) AS row_num
FROM `firebase-public-project.analytics_153293282.events_*`
WHERE event_name="user_engagement")
WHERE row_num = 1),
# query to extract behavioral data for each user ----------------------------------------------------------
get_behavioral_data AS (
SELECT
user_pseudo_id,
SUM(IF(event_name = 'user_engagement', 1, 0)) AS cnt_user_engagement,
SUM(IF(event_name = 'level_start_quickplay', 1, 0)) AS cnt_level_start_quickplay,
SUM(IF(event_name = 'level_end_quickplay', 1, 0)) AS cnt_level_end_quickplay,
SUM(IF(event_name = 'level_complete_quickplay', 1, 0)) AS cnt_level_complete_quickplay,
SUM(IF(event_name = 'level_reset_quickplay', 1, 0)) AS cnt_level_reset_quickplay,
SUM(IF(event_name = 'post_score', 1, 0)) AS cnt_post_score,
SUM(IF(event_name = 'spend_virtual_currency', 1, 0)) AS cnt_spend_virtual_currency,
SUM(IF(event_name = 'ad_reward', 1, 0)) AS cnt_ad_reward,
SUM(IF(event_name = 'challenge_a_friend', 1, 0)) AS cnt_challenge_a_friend,
SUM(IF(event_name = 'completed_5_levels', 1, 0)) AS cnt_completed_5_levels,
SUM(IF(event_name = 'use_extra_steps', 1, 0)) AS cnt_use_extra_steps,
FROM (
SELECT
e.*
FROM
`firebase-public-project.analytics_153293282.events_*` e
JOIN
get_label r
ON
e.user_pseudo_id = r.user_pseudo_id
WHERE
e.event_timestamp <= r.ts_24hr_after_first_engagement
)
GROUP BY 1)
SELECT
PARSE_TIMESTAMP('%Y-%m-%d %H:%M:%S', FORMAT_TIMESTAMP('%Y-%m-%d %H:%M:%S', TIMESTAMP_MICROS(ret.user_first_engagement))) AS user_first_engagement,
# ret.month,
# ret.julianday,
# ret.dayofweek,
dem.*,
CAST(IFNULL(beh.cnt_user_engagement, 0) AS FLOAT64) AS cnt_user_engagement,
CAST(IFNULL(beh.cnt_level_start_quickplay, 0) AS FLOAT64) AS cnt_level_start_quickplay,
CAST(IFNULL(beh.cnt_level_end_quickplay, 0) AS FLOAT64) AS cnt_level_end_quickplay,
CAST(IFNULL(beh.cnt_level_complete_quickplay, 0) AS FLOAT64) AS cnt_level_complete_quickplay,
CAST(IFNULL(beh.cnt_level_reset_quickplay, 0) AS FLOAT64) AS cnt_level_reset_quickplay,
CAST(IFNULL(beh.cnt_post_score, 0) AS FLOAT64) AS cnt_post_score,
CAST(IFNULL(beh.cnt_spend_virtual_currency, 0) AS FLOAT64) AS cnt_spend_virtual_currency,
CAST(IFNULL(beh.cnt_ad_reward, 0) AS FLOAT64) AS cnt_ad_reward,
CAST(IFNULL(beh.cnt_challenge_a_friend, 0) AS FLOAT64) AS cnt_challenge_a_friend,
CAST(IFNULL(beh.cnt_completed_5_levels, 0) AS FLOAT64) AS cnt_completed_5_levels,
CAST(IFNULL(beh.cnt_use_extra_steps, 0) AS FLOAT64) AS cnt_use_extra_steps,
ret.churned as churned,
CASE
WHEN churned = 0 THEN ( SELECT class_weight_zero FROM get_class_weights)
ELSE ( SELECT class_weight_one
FROM get_class_weights)
END AS class_weights
FROM
get_label ret
LEFT OUTER JOIN
get_demographic_data dem
ON
ret.user_pseudo_id = dem.user_pseudo_id
LEFT OUTER JOIN
get_behavioral_data beh
ON
ret.user_pseudo_id = beh.user_pseudo_id
WHERE ret.bounced = 0
"""
run_bq_query(preprocess_sql_query)
```
### Create table to update entities
```
processed_sql_query_day_one = f"""
CREATE OR REPLACE TABLE
`{PROJECT_ID}.{BQ_DATASET}.{FEATURES_TABLE_DAY_ONE}` AS
SELECT
*
FROM
`{PROJECT_ID}.{BQ_DATASET}.{FEATURES_TABLE}`
WHERE
user_first_engagement < '{MAX_DATE}'
"""
processed_sql_query_day_two = f"""
CREATE OR REPLACE TABLE
`{PROJECT_ID}.{BQ_DATASET}.{FEATURES_TABLE_DAY_TWO}` AS
SELECT
*
FROM
`{PROJECT_ID}.{BQ_DATASET}.{FEATURES_TABLE}`
WHERE
user_first_engagement >= '{MAX_DATE}'
"""
queries = processed_sql_query_day_one, processed_sql_query_day_two
for query in queries:
run_bq_query(query)
```
## Model Training
We created demographic and aggregate behavioral features. It is time to train our BQML model.
#### Train an Logistic classifier model
```
train_model_query = f"""
CREATE OR REPLACE MODEL `{PROJECT_ID}.{BQ_DATASET}.{MODEL_NAME}`
OPTIONS(MODEL_TYPE='{MODEL_TYPE}',
AUTO_CLASS_WEIGHTS={AUTO_CLASS_WEIGHTS},
MAX_ITERATIONS={MAX_ITERATIONS},
INPUT_LABEL_COLS=['{INPUT_LABEL_COLS}'])
AS SELECT * EXCEPT(user_first_engagement, user_pseudo_id, class_weights)
FROM `{PROJECT_ID}.{BQ_DATASET}.{FEATURES_TABLE_DAY_ONE}`;
"""
run_bq_query(train_model_query)
```
## Model Deployment
Once we get the model, you can export it and deploy to an Vertex AI Endpoint.
This is just one of the 5 ways to use BigQuery and Vertex AI together. [Check](https://cloud.google.com/blog/products/ai-machine-learning/five-integrations-between-vertex-ai-and-bigquery) this article to know more about them.
#### Export the model
```
model_extract_job = bigquery.ExtractJob(
client=bq_client,
job_id=JOB_ID,
source=MODEL_SOURCE,
destination_uris=[DESTINATION_URI],
job_config=EXTRACT_JOB_CONFIG,
)
try:
job = model_extract_job.result()
except job.error_result as error:
print(error)
```
#### (Locally) Check the SavedModel format
```
%%bash -s "$SERVING_DIR" "$DESTINATION_URI"
mkdir -p -m 777 $1
gsutil cp -r $2 $1
%%bash -s "$SERVING_DIR"
saved_model_cli show --dir $1/model/ --all
```
#### Upload and Deploy Model on Vertex AI Endpoint
```
bq_model = upload_model(
display_name=MODEL_NAME,
serving_container_image_uri=SERVING_CONTAINER_IMAGE_URI,
artifact_uri=DESTINATION_URI,
)
endpoint = create_endpoint(display_name=ENDPOINT_NAME)
deployed_model = deploy_model(
model=bq_model,
machine_type="n1-highcpu-4",
endpoint=endpoint,
deployed_model_display_name=DEPLOYED_MODEL_NAME,
min_replica_count=1,
max_replica_count=1,
sync=True,
)
```
#### Test predictions
```
instance = {
"cnt_ad_reward": 0,
"cnt_challenge_a_friend": 0,
"cnt_completed_5_levels": 0,
"cnt_level_complete_quickplay": 0,
"cnt_level_end_quickplay": 0,
"cnt_level_reset_quickplay": 0,
"cnt_level_start_quickplay": 0,
"cnt_post_score": 0,
"cnt_spend_virtual_currency": 0,
"cnt_use_extra_steps": 0,
"cnt_user_engagement": 14,
"country": "United States",
"language": "en-us",
"operating_system": "ANDROID",
}
bqml_predictions = endpoint_predict_sample(instances=[instance], endpoint=endpoint)
```
# Serve ML features at scale with low latency
At that point, **we deploy our simple model which would requires fetching aggregated attributes as input features in real time**.
That's why **we need a datastore optimized for singleton lookup operations** which would be able to scale and serve those aggregated feature online in low latency.
In other terms, we need to introduce Vertex AI Feature Store. Again, we assume you already know how to set up and work with a Vertex AI Feature store.
## Feature store for features management
In this section, we explore all Feature store management activities from create a Featurestore resource all way down to read feature values online.
Below you can see the feature store data model and a plain representation of how the data will be organized.
<img src="./assets/data_model_3.png"/>
### Create featurestore, ```mobile_gaming```
```
print(f"Listing all featurestores in {PROJECT_ID}")
feature_store_list = Featurestore.list()
if len(list(feature_store_list)) == 0:
print(f"The {PROJECT_ID} is empty!")
else:
for fs in feature_store_list:
print("Found featurestore: {}".format(fs.resource_name))
try:
mobile_gaming_feature_store = Featurestore.create(
featurestore_id=FEATURESTORE_ID,
online_store_fixed_node_count=ONLINE_STORE_NODES_COUNT,
labels={"team": "dataoffice", "app": "mobile_gaming"},
sync=True,
)
except RuntimeError as error:
print(error)
else:
FEATURESTORE_RESOURCE_NAME = mobile_gaming_feature_store.resource_name
print(f"Feature store created: {FEATURESTORE_RESOURCE_NAME}")
```
### Create the ```User``` entity type and its features
```
try:
user_entity_type = mobile_gaming_feature_store.create_entity_type(
entity_type_id=ENTITY_ID, description="User Entity", sync=True
)
except RuntimeError as error:
print(error)
else:
USER_ENTITY_RESOURCE_NAME = user_entity_type.resource_name
print("Entity type name is", USER_ENTITY_RESOURCE_NAME)
```
### Set Feature Monitoring
Feature [monitoring](https://cloud.google.com/vertex-ai/docs/featurestore/monitoring) is in preview, so you need to use v1beta1 Python which is a lower-level API than the one we've used so far in this notebook.
The easiest way to set this for now is using [console UI](https://console.cloud.google.com/vertex-ai/features). For completeness, below is example to do this using v1beta1 SDK.
```
from google.cloud.aiplatform_v1beta1 import \
FeaturestoreServiceClient as v1beta1_FeaturestoreServiceClient
from google.cloud.aiplatform_v1beta1.types import \
entity_type as v1beta1_entity_type_pb2
from google.cloud.aiplatform_v1beta1.types import \
featurestore_monitoring as v1beta1_featurestore_monitoring_pb2
from google.cloud.aiplatform_v1beta1.types import \
featurestore_service as v1beta1_featurestore_service_pb2
from google.protobuf.duration_pb2 import Duration
v1beta1_admin_client = v1beta1_FeaturestoreServiceClient(
client_options={"api_endpoint": API_ENDPOINT}
)
v1beta1_admin_client.update_entity_type(
v1beta1_featurestore_service_pb2.UpdateEntityTypeRequest(
entity_type=v1beta1_entity_type_pb2.EntityType(
name=v1beta1_admin_client.entity_type_path(
PROJECT_ID, REGION, FEATURESTORE_ID, ENTITY_ID
),
monitoring_config=v1beta1_featurestore_monitoring_pb2.FeaturestoreMonitoringConfig(
snapshot_analysis=v1beta1_featurestore_monitoring_pb2.FeaturestoreMonitoringConfig.SnapshotAnalysis(
monitoring_interval=Duration(seconds=86400), # 1 day
),
),
),
)
)
```
### Create features
#### Create Feature configuration
For simplicity, I created the configuration in a declarative way. Of course, we can create an helper function to built it from Bigquery schema.
Also notice that we want to pass some feature on-fly. In this case, it country, operating system and language looks perfect for that.
```
feature_configs = {
"country": {
"value_type": "STRING",
"description": "The country of customer",
"labels": {"status": "passed"},
},
"operating_system": {
"value_type": "STRING",
"description": "The operating system of device",
"labels": {"status": "passed"},
},
"language": {
"value_type": "STRING",
"description": "The language of device",
"labels": {"status": "passed"},
},
"cnt_user_engagement": {
"value_type": "DOUBLE",
"description": "A variable of user engagement level",
"labels": {"status": "passed"},
},
"cnt_level_start_quickplay": {
"value_type": "DOUBLE",
"description": "A variable of user engagement with start level",
"labels": {"status": "passed"},
},
"cnt_level_end_quickplay": {
"value_type": "DOUBLE",
"description": "A variable of user engagement with end level",
"labels": {"status": "passed"},
},
"cnt_level_complete_quickplay": {
"value_type": "DOUBLE",
"description": "A variable of user engagement with complete status",
"labels": {"status": "passed"},
},
"cnt_level_reset_quickplay": {
"value_type": "DOUBLE",
"description": "A variable of user engagement with reset status",
"labels": {"status": "passed"},
},
"cnt_post_score": {
"value_type": "DOUBLE",
"description": "A variable of user score",
"labels": {"status": "passed"},
},
"cnt_spend_virtual_currency": {
"value_type": "DOUBLE",
"description": "A variable of user virtual amount",
"labels": {"status": "passed"},
},
"cnt_ad_reward": {
"value_type": "DOUBLE",
"description": "A variable of user reward",
"labels": {"status": "passed"},
},
"cnt_challenge_a_friend": {
"value_type": "DOUBLE",
"description": "A variable of user challenges with friends",
"labels": {"status": "passed"},
},
"cnt_completed_5_levels": {
"value_type": "DOUBLE",
"description": "A variable of user level 5 completed",
"labels": {"status": "passed"},
},
"cnt_use_extra_steps": {
"value_type": "DOUBLE",
"description": "A variable of user extra steps",
"labels": {"status": "passed"},
},
"churned": {
"value_type": "INT64",
"description": "A variable of user extra steps",
"labels": {"status": "passed"},
},
"class_weights": {
"value_type": "STRING",
"description": "A variable of class weights",
"labels": {"status": "passed"},
},
}
```
#### Create features using `batch_create_features` method
```
try:
user_entity_type.batch_create_features(feature_configs=feature_configs, sync=True)
except RuntimeError as error:
print(error)
else:
for feature in user_entity_type.list_features():
print("")
print(f"The resource name of {feature.name} feature is", feature.resource_name)
```
### Search features
```
feature_query = "feature_id:cnt_user_engagement"
searched_features = Feature.search(query=feature_query)
searched_features
```
### Import ```User``` feature values using ```ingest_from_bq``` method
You need to import feature values before you can use them for online/offline serving.
```
FEATURES_IDS = [feature.name for feature in user_entity_type.list_features()]
try:
user_entity_type.ingest_from_bq(
feature_ids=FEATURES_IDS,
feature_time=FEATURE_TIME,
bq_source_uri=BQ_SOURCE_URI_DAY_ONE,
entity_id_field=ENTITY_ID_FIELD,
disable_online_serving=False,
worker_count=20,
sync=True,
)
except RuntimeError as error:
print(error)
```
**Comment: How does Vertex AI Feature Store mitigate training serving skew?**
Let's just think about what is happening for a second.
We just ingest customer behavioral features we engineered before when we trained the model. And we are now going to serve the same features for online prediction.
But, what if those attributes on the incoming prediction requests would differ with respect to the one calculated during the model training? In particular, what if the correct attributes have different characteristics as the data the model was trained on? At that point, you should start perceiving this idea of **skew** between training and serving data. So what? Imagine now that the mobile gaming app go trending and users start challenging friends more frequently. This would change the distribution of the `cnt_challenge_a_friend`. But the model, which estimates your churn probability, was trained on a different distribution. And if we assume that type and frequency of ads depend on those predictions, it would happen that you target wrong users with wrong ads with an expected frequency because this offline-online feature inconsistency.
**Vertex AI Feature store** addresses those skew by an ingest-one and and re-used many logic. Indeed, once the feature is computed, the same features would be available both in training and serving.
## Simulate online prediction requests
```
online_sample = {
"entity_id": ["DE346CDD4A6F13969F749EA8047F282A"],
"country": ["United States"],
"operating_system": ["IOS"],
"language": ["en"],
}
prediction = simulate_prediction(endpoint=endpoint, online_sample=online_sample)
print(prediction)
```
# Train a new churn ML model using Vertex AI AutoML
Now assume that you have a meeting with the team and you decide to use Vertex AI AutoML to train a new version of the model.
But while you were discussing about that, new data where ingested into the feature store.
## Ingest new data in the feature store
```
try:
user_entity_type.ingest_from_bq(
feature_ids=FEATURES_IDS,
feature_time=FEATURE_TIME,
bq_source_uri=BQ_SOURCE_URI_DAY_TWO,
entity_id_field=ENTITY_ID_FIELD,
disable_online_serving=False,
worker_count=1,
sync=True,
)
except RuntimeError as error:
print(error)
```
## Avoid data leakage with point-in-time lookup to fetch training data
Now, without a datastore with a timestamp data model, some data leakage would happen and you would end by training the new model on a different dataset. As a consequence, you cannot compare those models. In order to avoid that, **you need to be able to train model on the same data at same specific point in time we use in the previous version of the model**.
<center><img src="./assets/point_in_time_2.png"/><center/>
**With the Vertex AI Feature store, you can fetch feature values corresponding to a particular timestamp thanks to point-in-time lookup capability.** In terms of SDK, you need to define a `read instances` object which is a list of entity id / timestamp pairs, where the entity id is the `user_pseudo_id` and `user_first_engagement` indicates we want to read the latest information available about that user. In this way, we will be able to reproduce the exact same training sample you need for the new model.
Let's see how to do that.
### Define query for reading instances at a specific point in time
```
# WHERE ABS(MOD(FARM_FINGERPRINT(STRING(user_first_engagement, 'UTC')), 10)) < 8
read_instances_query = f"""
CREATE OR REPLACE TABLE
`{PROJECT_ID}.{BQ_DATASET}.{READ_INSTANCES_TABLE}` AS
SELECT
user_pseudo_id as customer,
TIMESTAMP_TRUNC(CURRENT_TIMESTAMP(), SECOND, "UTC") as timestamp
FROM
`{BQ_DATASET}.{FEATURES_TABLE_DAY_ONE}` AS e
ORDER BY
user_first_engagement
"""
```
### Create the BigQuery instances table
```
run_bq_query(read_instances_query)
```
### Serve features for batch training
```
mobile_gaming_feature_store.batch_serve_to_bq(
bq_destination_output_uri=BQ_DESTINATION_OUTPUT_URI,
serving_feature_ids=SERVING_FEATURE_IDS,
read_instances_uri=READ_INSTANCES_URI,
)
```
## Train and Deploy AutoML model on Vertex AI
Now that we reproduce the training sample, we use the Vertex AI SDK to train an new version of the model using Vertex AI AutoML.
### Create the Managed Tabular Dataset from a CSV
```
dataset = vertex_ai.TabularDataset.create(
display_name=DATASET_NAME,
bq_source=BQ_DESTINATION_OUTPUT_URI,
)
dataset.resource_name
```
### Create and Launch the Training Job to build the Model
```
automl_training_job = vertex_ai.AutoMLTabularTrainingJob(
display_name=AUTOML_TRAIN_JOB_NAME,
optimization_prediction_type="classification",
optimization_objective="maximize-au-roc",
column_transformations=[
{"categorical": {"column_name": "country"}},
{"categorical": {"column_name": "operating_system"}},
{"categorical": {"column_name": "language"}},
{"numeric": {"column_name": "cnt_user_engagement"}},
{"numeric": {"column_name": "cnt_level_start_quickplay"}},
{"numeric": {"column_name": "cnt_level_end_quickplay"}},
{"numeric": {"column_name": "cnt_level_complete_quickplay"}},
{"numeric": {"column_name": "cnt_level_reset_quickplay"}},
{"numeric": {"column_name": "cnt_post_score"}},
{"numeric": {"column_name": "cnt_spend_virtual_currency"}},
{"numeric": {"column_name": "cnt_ad_reward"}},
{"numeric": {"column_name": "cnt_challenge_a_friend"}},
{"numeric": {"column_name": "cnt_completed_5_levels"}},
{"numeric": {"column_name": "cnt_use_extra_steps"}},
],
)
# This will take around an 2 hours to run
automl_model = automl_training_job.run(
dataset=dataset,
target_column=INPUT_LABEL_COLS,
training_fraction_split=0.8,
validation_fraction_split=0.1,
test_fraction_split=0.1,
weight_column="class_weights",
model_display_name=AUTOML_MODEL_NAME,
disable_early_stopping=False,
)
```
### Deploy Model to the same Endpoint with Traffic Splitting
Vertex AI Endpoint provides a managed traffic splitting service. All you need to do is to define the splitting policy and then the service will deal it for you.
Be sure that both models have the same serving function. In our case both BQML Logistic classifier and Vertex AI AutoML support same prediction format.
```
model_deployed_id = endpoint.list_models()[0].id
RETRAIN_TRAFFIC_SPLIT = {"0": 50, model_deployed_id: 50}
endpoint.deploy(
automl_model,
deployed_model_display_name=MODEL_DEPLOYED_NAME,
traffic_split=RETRAIN_TRAFFIC_SPLIT,
machine_type=SERVING_MACHINE_TYPE,
accelerator_count=0,
min_replica_count=MIN_NODES,
max_replica_count=MAX_NODES,
)
```
## Time to simulate online predictions
```
for i in range(2000):
simulate_prediction(endpoint=endpoint, online_sample=online_sample)
time.sleep(1)
```
Below the Vertex AI Endpoint UI result you would able to see after the online prediction simulation ends
<img src="./assets/prediction_results.jpg"/>
## Cleaning up
To clean up all Google Cloud resources used in this project, you can [delete the Google Cloud
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
Otherwise, you can delete the individual resources you created in this tutorial
```
# delete feature store
mobile_gaming_feature_store.delete(sync=True, force=True)
# delete Vertex AI resources
endpoint.undeploy_all()
bq_model.delete()
automl_model.delete()
%%bash -s "$SERVING_DIR"
rm -Rf $1
# Warning: Setting this to true will delete everything in your bucket
delete_bucket = False
if delete_bucket and "BUCKET_URI" in globals():
! gsutil -m rm -r $BUCKET_URI
# Delete the BigQuery Dataset
!bq rm -r -f -d $PROJECT_ID:$BQ_DATASET
```
| github_jupyter |
#Author : Jay Shukla
##Task 3 : Prediction using Decision Tree Algorithm
##GRIP @ The Sparks Foundation
Decision Trees are versatile Machine Learning algorithms that can perform both classification and regression tasks, and even multioutput tasks.For the given ‘Iris’ dataset, I created the Decision Tree classifier and visualized it graphically. The purpose of this task is if we feed any new data to this classifier, it would be able to predict the right class accordingly.
## Importing Libraries
```
# Importing libraries in Python
from sklearn.datasets import load_iris
from sklearn.tree import DecisionTreeClassifier, export_graphviz
from sklearn.model_selection import train_test_split
import sklearn.metrics as sm
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import pydot
from IPython.display import Image
```
## Loading Dataset
```
iris = load_iris()
X=iris.data[:,:]
y=iris.target
```
### Data Preprocessing
```
data=pd.DataFrame(iris['data'],columns=["Petal length","Petal Width","Sepal Length","Sepal Width"])
data['Species']=iris['target']
data['Species']=data['Species'].apply(lambda x: iris['target_names'][x])
data.head()
data.describe()
```
## Data Visualisation
```
sns.pairplot(data)
# Scatter plot of data based on Sepal Length and Width features
sns.FacetGrid(data,hue='Species').map(plt.scatter,'Sepal Length','Sepal Width').add_legend()
plt.show()
# Scatter plot of data based on Petal Length and Width features
sns.FacetGrid(data,hue='Species').map(plt.scatter,'Petal length','Petal Width').add_legend()
plt.show()
```
## Creating the Model
```
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.1, random_state=1)
tree_classifier = DecisionTreeClassifier()
tree_classifier.fit(X_train,y_train)
print("Training Complete.")
y_pred = tree_classifier.predict(X_test)
```
### Making a dataframe consisting True & Predicted Data
```
df = pd.DataFrame({'Actual': y_test, 'Predicted': y_pred})
df
```
## Visualising the Decision Tree Model
```
export_graphviz(
tree_classifier,
out_file="img\desision_tree.dot",
feature_names=iris.feature_names[:],
class_names=iris.target_names,
rounded=True,
filled=True
)
(graph,) = pydot.graph_from_dot_file('img\desision_tree.dot')
graph.write_png('img\desision_tree.png')
Image(filename='img\desision_tree.png')
```
## Evaluating the Model
```
sm.accuracy_score(df['Actual'],df['Predicted'])
```
## Conclusion
As we can see the Decision Tree model predicts the data with 100% accuracy. Thus here I can say that the task - 3 is fulfilled which consisted **Training**,**Visualising** & **Evaluating** the model.
| github_jupyter |
# VGGNet in Keras
In this notebook, we fit a model inspired by the "very deep" convolutional network [VGGNet](https://arxiv.org/pdf/1409.1556.pdf) to classify flowers into the 17 categories of the Oxford Flowers data set. Derived from [these](https://github.com/the-deep-learners/TensorFlow-LiveLessons/blob/master/notebooks/old/L3-3c__TFLearn_VGGNet.ipynb) [two](https://github.com/the-deep-learners/TensorFlow-LiveLessons/blob/master/notebooks/alexnet_in_keras.ipynb) earlier notebooks.
#### Set seed for reproducibility
```
import numpy as np
np.random.seed(42)
```
#### Load dependencies
```
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPooling2D
from keras.layers.normalization import BatchNormalization
from keras.callbacks import TensorBoard # for part 3.5 on TensorBoard
```
#### Load *and preprocess* data
```
import tflearn.datasets.oxflower17 as oxflower17
X, Y = oxflower17.load_data(one_hot=True)
```
#### Design neural network architecture
```
model = Sequential()
model.add(Conv2D(64, 3, activation='relu', input_shape=(224, 224, 3)))
model.add(Conv2D(64, 3, activation='relu'))
model.add(MaxPooling2D(2, 2))
model.add(BatchNormalization())
model.add(Conv2D(128, 3, activation='relu'))
model.add(Conv2D(128, 3, activation='relu'))
model.add(MaxPooling2D(2, 2))
model.add(BatchNormalization())
model.add(Conv2D(256, 3, activation='relu'))
model.add(Conv2D(256, 3, activation='relu'))
model.add(Conv2D(256, 3, activation='relu'))
model.add(MaxPooling2D(2, 2))
model.add(BatchNormalization())
model.add(Conv2D(512, 3, activation='relu'))
model.add(Conv2D(512, 3, activation='relu'))
model.add(Conv2D(512, 3, activation='relu'))
model.add(MaxPooling2D(2, 2))
model.add(BatchNormalization())
model.add(Conv2D(512, 3, activation='relu'))
model.add(Conv2D(512, 3, activation='relu'))
model.add(Conv2D(512, 3, activation='relu'))
model.add(MaxPooling2D(2, 2))
model.add(BatchNormalization())
model.add(Flatten())
model.add(Dense(4096, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(4096, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(17, activation='softmax'))
model.summary()
```
#### Configure model
```
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
```
#### Configure TensorBoard (for part 5 of lesson 3)
```
tensorbrd = TensorBoard('logs/vggnet')
```
#### Train!
```
model.fit(X, Y, batch_size=64, epochs=16, verbose=1, validation_split=0.1, shuffle=True,
callbacks=[tensorbrd])
```
| github_jupyter |
```
import os
import pandas as pd
import numpy as np
from multiprocessing import Pool, cpu_count
valid_ids = np.load("../results/misc/glacier_ids_valid.npy")
valid_ids
static_features = pd.read_csv("../hackathon_2018/data/raw/RGI-Asia/rgi60_Asia.csv")
static_features.columns
tsl_store = pd.read_hdf("../data/FIT_forcing/tsl/TSL-filtered-noWinterMax_SLAthres.h5", mode="r")
meteo_path = "../data/FIT_forcing/meteo/"
def read_tsl_minmax(rgi_id, store=tsl_store):
df = store[store['RGI_ID']==rgi_id][["SC_median", "LS_DATE"]]
df.index = pd.to_datetime(df["LS_DATE"])
tsl = pd.DataFrame(df["SC_median"]) # previously TSL_normalized
tsl_min = tsl.min()[0]
tsl_max = tsl.max()[0]
return tsl_min, tsl_max
def read_meteo(rgi_id, path=meteo_path):
meteo = pd.read_hdf(f"{meteo_path}{rgi_id}.h5")
return meteo
read_tsl_minmax("RGI60-13.00014")
def MM_rescaler(Xsc, min_, max_):
X = Xsc * (max_ - min_) + min_
return X
def basin_wise(rgi_id, freq="M", subset_jjas=False):
if freq == "M":
freq_prefix = "monthly"
elif freq == "W":
freq_prefix = "weekly"
if subset_jjas:
subset_prefix = "JJAS"
else:
subset_prefix = "full"
source_dir = f"../results/data4ml/{freq_prefix}_{subset_prefix}/"
data = pd.read_csv(f"{source_dir}{rgi_id}.csv", compression="gzip")
static_features_slice = static_features[static_features["RGIId"]==rgi_id].copy()
static_features_slice = static_features_slice[['CenLon', 'CenLat', 'Area', 'Zmin', 'Zmax', 'Zmed',
'Slope', 'Aspect', 'Lmax']].copy()
for c in static_features_slice.columns:
data[c] = static_features_slice[c].values[0]
tsl_min, tsl_max = read_tsl_minmax(rgi_id)
data["TSL"] = MM_rescaler(data["TSL_normalized"].values, tsl_min, tsl_max)
data = data.drop("TSL_normalized", axis=1)
return data
f = basin_wise("RGI60-13.00014")
f.head()
output_file = "../results/data4ml/domain/monthly_full.csv"
def combine_for_domain(output_f, freq="M", subset_jjas=False):
ids_valid = valid_ids
for idx in ids_valid:
chunk = basin_wise(idx, freq, subset_jjas)
chunk.to_csv(output_f, mode="a", index=False, header=False)
# monthly full
combine_for_domain(output_file, freq="M", subset_jjas=False)
%%time
# weekly full
combine_for_domain("../results/data4ml/domain/weekly_full.csv", freq="W", subset_jjas=False)
# monthly JJAS
combine_for_domain("../results/data4ml/domain/monthly_JJAS.csv", freq="M", subset_jjas=True)
# weekly JJAS
combine_for_domain("../results/data4ml/domain/weekly_JJAS.csv", freq="W", subset_jjas=True)
f = basin_wise("RGI60-13.00014")
f.head()
ff = pd.read_pickle("../hackathon_2018/data/for_training/RGI60-13.00014.pkl")
ff.columns[-40:]
ff[["TSL_ELEV"]].describe()
f[["TSL_normalized"]].describe()
```
| github_jupyter |
# UAS Collected Traffic Data Analysis (UAS4T)
The scope of the competition is to evaluate the accuracy of statistical or CI methods in transportation-related
detection problems with specific reference in queue formation in urban arterials. The focus will be on obtaining
results as close as possible to the real data.
As per the requirements of the competition, we tried to develop an algorithm to estimate the maximum length of the queues per lane that
are formed for different approaches of an intersection and roads (to be specified) during the monitoring duration.
This algorithm outputs following components:
i. Maximum length of queue
ii. Lane the maximum length occurred
iii. Coordinates of the start and end of the maximum queue
iv. Timestamp of the maximum queue occurrence
v. Whether, when and where a spillback is formed
```
import numpy as np
import cv2
import csv
import json
import os
from shapely.geometry import Point
from shapely.geometry.polygon import Polygon
from matplotlib import pyplot as plt
#DISP : Display Flag for Vehicle Trajectory,Lanes and Spillback
DISP = 1
#Color Indications of all the vehicles from the given data
obj_types = {' Car': {'color': (255, 70, 70),
'width': 5,
'height': 5, },
' Medium Vehicle': {'color': (70, 255, 70),
'width': 4,
'height': 4, },
' Motorcycle': {'color': (70, 70, 255),
'width': 3,
'height': 3, },
' Heavy Vehicle': {'color': (255, 255, 0),
'width': 6,
'height': 6, },
' Bus': {'color': (70, 100, 255),
'width': 6,
'height': 6, },
' Taxi': {'color': (255, 0, 255),
'width': 4,
'height': 4, }, }
#Properties of the trajectory
traj_props = {}
traj_props['lat_min'] = 1000000
traj_props['lat_max'] = 0
traj_props['lon_min'] = 1000000
traj_props['lon_max'] = 0
traj_props['lon_diff'] = 0
traj_props['lat_diff'] = 0
traj_props['max_time'] = 813
traj_props['min_time'] = 0
traj_props['img_height'] = 0
traj_props['img_width'] = 0
traj_props['scale_trajectory'] = 200000
traj_props['longitude_km'] = 111.2
traj_props['lattitude_km'] = 127.2
#Route information initialization
routes_information={}
routes_names = ['LeofAlexandras_tw_28isOktovriou', 'OktovriouIs28_tw_LeofAlexandras', 'OktovriouIs28_tw_South']
for route in routes_names:
route_information={}
route_information['direction']=0
route_information['orientation_range'] = []
route_information['max_queue']={}
route_information['max_queue']['length'] = 0
route_information['max_queue']['time'] = 0.00
route_information['max_queue']['points'] = []
route_information['max_queue']['n_vehicles'] = []
routes_information[route]= route_information
routes_information['LeofAlexandras_tw_28isOktovriou']['direction'] = 0
routes_information['OktovriouIs28_tw_LeofAlexandras']['direction'] = 225
routes_information['OktovriouIs28_tw_South']['direction'] = 90
routes_information['LeofAlexandras_tw_28isOktovriou']['orientation_range'] = [337.5, 22.5]
routes_information['OktovriouIs28_tw_LeofAlexandras']['orientation_range'] = [202.5, 247.5]
routes_information['OktovriouIs28_tw_South']['orientation_range'] = [67.5, 112.5]
routes_information['LeofAlexandras_tw_28isOktovriou']['lane_axis'] = [0, 0] # 0 col 1 row
routes_information['OktovriouIs28_tw_LeofAlexandras']['lane_axis'] = [0, 1]
routes_information['OktovriouIs28_tw_South']['lane_axis'] = [1, 1]
```
### Locating the Lane Areas
Location of the lane areas is determined by anchor drawing concept. Proposed approach is modified / improved version of edge drawing algorithm and called as anchor drawing. In anchor drawing, we will draw a line passing through the maximum peak pixels. We get continuous line in anchor drawing compared to edge drawing.
```
class AnchorDrawing:
"""
A class to draw the anchor line to locate the lane.
...
Attributes
----------
Methods
-------
moveUp_(x, y):
Compute next peak value towards up direction
moveDown_(x, y):
Compute next peak value towards down direction
moveRight_(x,y):
Compute next peak value towards right direction
moveLeft_(x, y):
Compute next peak value towards left direction
moveon_peak_points(x, y, direct_next):
Compute the next peaks
compute_anchors(image):
Computing initial peak or anchor points list
Draw(image):
Entry point of the class
"""
# initiation
def __init__(self):
self.anchorThreshold_ = 50
# dimension of image
self.height = 0
self.width = 0
self.Degree = np.array([])
self.anchor_image = np.array([])
self.Img = np.array([])
self.horizontal_move = 1
self.vertical_move = -1
self.left_move = -1
self.right_move = 1
self.up_move = -1
self.down_move = 1
def moveUp_(self, x, y):
'''
Compute next peak value towards up direction
Input Parameters:
x: Row index
y: Column index
Output Parameters:
list_points: Segment of peak points
direct_next: Search direction of left side similart to right, up and down
'''
list_points = [] # array to store peak points
direct_next = None # search direction of left side similart to right, up and down
while x > 0 and self.Img[x, y] > 0 and not self.anchor_image[x, y]:
next_y = [max(0, y - 1), y, min(self.width - 1, y + 1)] # search in a valid area
list_points.append((x, y)) # extend line segments
if self.Degree[x, y] == self.vertical_move:
self.anchor_image[x, y] = True # mark as anchor peak
y_last = y # record parent pixel
x, y = x - 1, next_y[np.argmax(self.Img[x - 1, next_y])] # walk to next pixel with max gradient
else:
direct_next = y - y_last # change direction to continue search
break # stop and proceed to next search
return list_points, direct_next
def moveDown_(self, x, y):
'''
Compute next peak value towards down direction
Input Parameters:
x: Row index
y: Column index
Output Parameters:
list_points: Segment of peak points
direct_next: Search direction of left side similart to right, up and down
'''
list_points = []
direct_next = None
while x < self.height - 1 and self.Img[x, y] > 0 and not self.anchor_image[x, y]:
next_y = [max(0, y - 1), y, min(self.width - 1, y + 1)]
list_points.append((x, y))
if self.Degree[x, y] == self.vertical_move:
self.anchor_image[x, y] = True
y_last = y
x, y = x + 1, next_y[np.argmax(self.Img[x + 1, next_y])]
else:
direct_next = y - y_last
break
return list_points, direct_next
def moveRight_(self, x, y):
'''
Compute next peak value towards right direction
Input Parameters:
x: Row index
y: Column index
Output Parameters:
list_points: Segment of peak points
direct_next: Search direction of left side similart to right, up and down
'''
list_points = []
direct_next = None
while y < self.width - 1 and self.Img[x, y] > 0 and not self.anchor_image[x, y]:
next_x = [max(0, x - 1), x, min(self.height - 1, x + 1)]
list_points.append((x, y))
if self.Degree[x, y] == self.horizontal_move:
self.anchor_image[x, y] = True
x_last = x
x, y = next_x[np.argmax(self.Img[next_x, y + 1])], y + 1
else:
direct_next = x - x_last
break
return list_points, direct_next
def moveLeft_(self, x, y):
'''
Compute next peak value towards left direction
Input Parameters:
x: Row index
y: Column index
Output Parameters:
list_points: Segment of peak points
direct_next: Search direction of left side similart to right, up and down
'''
list_points = []
direct_next = None
while y > 0 and self.Img[x, y] > 0 and not self.anchor_image[x, y]:
next_x = [max(0, x - 1), x, min(self.height - 1, x + 1)]
list_points.append((x, y))
if self.Degree[x, y] == self.horizontal_move:
self.anchor_image[x, y] = True
x_last = x
x, y = next_x[np.argmax(self.Img[next_x, y - 1])], y - 1
else:
direct_next = x - x_last
break
return list_points, direct_next
def moveon_peak_points(self, x, y, direct_next):
'''
Returns the
Input Parameters:
x: Row index
y: Column index
direct_next: Search direction of left side similart to right, up and down
Output Parameters:
list_points: Segment of peak points
'''
list_points = [(x, y)]
while direct_next is not None:
x, y = list_points[-1][0], list_points[-1][1]
# if the last point is towords horizontal, search horizontally
if self.Degree[x, y] == self.horizontal_move:
# get points sequence
if direct_next == self.left_move:
s, direct_next = self.moveLeft_(x, y)
elif direct_next == self.right_move:
s, direct_next = self.moveRight_(x, y)
else:
break
elif self.Degree[x, y] == self.vertical_move: # search vertically
if direct_next == self.up_move:
s, direct_next = self.moveUp_(x, y)
elif direct_next == self.down_move:
s, direct_next = self.moveDown_(x, y)
else:
break
else: # invalid point found
break
if len(s) > 1:
list_points.extend(s[1:])
return list_points
# find list of anchors
def compute_anchors(self, image):
'''
Computing initial peak or anchor points list
Input Parameters:
image: Input accumulated image with vehicle trajectories
Output Parameters:
anchor_list: List of anchor points
'''
# detect the anchor points
anchor_list = []
self.Degree = np.zeros(image.shape, np.float64)
for row in range(1, self.height - 1):
for col in range(1, self.width - 1):
if (image[row, col] > self.anchorThreshold_):
if ((image[row - 1, col] < image[row, col] and image[row + 1, col] < image[row, col]) or \
(image[row, col - 1] < image[row, col] and image[row, col + 1] < image[row, col]) or\
(image[row - 1, col - 1] < image[row, col] and image[row + 1, col + 1] < image[row, col]) or\
(image[row - 1, col + 1] < image[row, col] and image[row + 1, col - 1] < image[row, col])):
anchor_list.append((row, col))
ysum = int(image[row-1, col])+image[row+1, col] + image[row-1, col-1]+image[row+1, col-1] + image[row-1, col+1]+image[row+1, col+1]
xsum = int(image[row, col-1])+image[row, col+1] + image[row-1, col-1]+image[row-1, col+1] + image[row+1, col-1]+image[row+1, col+1]
if (ysum > xsum):
self.Degree[row, col] = -1
else:
self.Degree[row, col] = 1
return anchor_list
def Draw(self, image):
'''
Entry point of the class
Input Parameters:
image: Input accumulated image with vehicle trajectories
Output Parameters:
anchor_line: List of anchor points
'''
self.height = image.shape[0]
self.width = image.shape[1]
self.Img = image.copy()
# compute anchor points list
anchor_list = self.compute_anchors(image)
anchor_line = []
self.anchor_image = np.zeros(self.Img.shape, dtype=bool)
for anchor in anchor_list:
if not self.anchor_image[anchor]: # if not mark as anchor peak
# serch for next peak point in direction 1
point_list1 = self.moveon_peak_points(anchor[0], anchor[1], 1)
self.anchor_image[anchor] = False
# serch for next peak point in direction -1
point_list2 = self.moveon_peak_points(anchor[0], anchor[1], -1)
# concat two point lists
if len(point_list1[::-1] + point_list2) > 0:
anchor_line.append(point_list1[::-1] + point_list2[1:])
return anchor_line
def readdata(file_name):
'''
Reading the data from given input csv
Input Parameters:
file_name: csv filename/path
Output Parameters:
data: Dictionary of all trajectory points along with vehicle information
'''
csv_file = open(file_name, 'r')
lines = csv_file.readlines()
num_lines = len(lines)
object_list = []
for row in range(1, num_lines):
# all_lines.append(row)
line = lines[row]
line_parts = line.split(';')
object_prop = {}
object_prop['trajectory'] = {}
object_prop['trajectory']['lat'] = []
object_prop['trajectory']['lon'] = []
object_prop['trajectory']['speed'] = []
object_prop['trajectory']['lon_acc'] = []
object_prop['trajectory']['lat_acc'] = []
object_prop['trajectory']['time'] = []
object_prop['trajectory']['x'] = []
object_prop['trajectory']['y'] = []
object_prop['track_id'] = int(line_parts[0])
object_prop['type'] = line_parts[1]
object_prop['traveled_d'] = float(line_parts[2])
object_prop['avg_speed'] = float(line_parts[3])
for step in range(4, len(line_parts) - 6, 6):
latitude = float(line_parts[step])
longitude = float(line_parts[step + 1])
speed_v = float(line_parts[step + 2])
latitude_acc = float(line_parts[step + 3])
longitude_acc = float(line_parts[step + 4])
time_stamp = float(line_parts[step + 5])
object_prop['trajectory']['lat'].append(latitude)
object_prop['trajectory']['lon'].append(longitude)
object_prop['trajectory']['speed'].append(speed_v)
object_prop['trajectory']['lon_acc'].append(longitude_acc)
object_prop['trajectory']['lat_acc'].append(latitude_acc)
object_prop['trajectory']['time'].append(time_stamp)
if (traj_props['lon_max'] < longitude):
traj_props['lon_max'] = longitude
if (traj_props['lat_max'] < latitude):
traj_props['lat_max'] = latitude
if (traj_props['lon_min'] > longitude):
traj_props['lon_min'] = longitude
if (traj_props['lat_min'] > latitude):
traj_props['lat_min'] = latitude
if (traj_props['min_time'] > time_stamp):
traj_props['min_time'] = time_stamp
if (traj_props['max_time'] < time_stamp):
traj_props['max_time'] = time_stamp
object_list.append(object_prop)
traj_props['lon_diff'] = traj_props['lon_max'] - traj_props['lon_min']
traj_props['lat_diff'] = traj_props['lat_max'] - traj_props['lat_min']
traj_props['img_height'] = int(round(traj_props['lat_diff'] * traj_props['scale_trajectory']))
traj_props['img_width'] = int(round(traj_props['lon_diff'] * traj_props['scale_trajectory']))
data = {}
data['object_list'] = object_list
data['traj_props'] = traj_props
return data
def get_line(point1, point2):
'''
Computes list of line points between two points
Input Parameters:
point1:(Col, Row)
point2:(Col, Row)
Output Parameters:
points: list of line points from point1 to point2
'''
points = []
issteep = abs(point2[1] - point1[1]) > abs(point2[0] - point1[0])
if issteep:
point1[0], point1[1] = point1[1], point1[0]
point2[0], point2[1] = point2[1], point2[0]
rev = False
if point1[0] > point2[0]:
point1[0], point2[0] = point2[0], point1[0]
point1[1], point2[1] = point2[1], point1[1]
rev = True
deltax = point2[0] - point1[0]
deltay = abs(point2[1] - point1[1])
error = int(deltax / 2)
y = point1[1]
if point1[1] < point2[1]:
ystep = 1
else:
ystep = -1
for x in range(point1[0], point2[0] + 1):
if issteep:
points.append((y, x))
else:
points.append((x, y))
error -= deltay
if error < 0:
y += ystep
error += deltax
# Reverse the list if the coordinates were reversed
if rev:
points.reverse()
return points
def find_ang(p1, p2):
'''
Computes the angle of line with respect to horizontal axis
Input Parameters:
p1: (Col, Row)
p2: (Col, Row)
Output Parameters:
angle_in_degrees: Angle of the line(0-360)
'''
# ********90********
# **45********135**
# 0**************180
# **-45*******-135*
# *******-90*******
angle_in_degrees = np.arctan2(p1[1] - p2[1], p1[0] - p2[0]) * 180 / np.pi
if (angle_in_degrees < 0):
angle_in_degrees += 360
return angle_in_degrees
def normalizeimg(accum):
'''
Linear normalization of accumated array with vehicle trajectories
Input Parameters:
accum: accumulated array
Output Parameters:
accum: normalized accumulated array(0-255)
'''
min = 0
try:
min = accum[accum != 0].min()
except:
print()
if (min != 0):
accum[accum != 0] = (accum[accum != 0] - min) * (255 / float(accum.max() - min + 0.000000001))
else:
accum = (accum - accum.min()) * (255 / float(accum.max() - accum.min() + 0.000000001))
return accum
def getLanePoints(img_side,direction=0, rRad = 3 ,n_lanes = 3, start_point = None):
'''
Computes lane areas using anchor drawing algorithm
Input Parameters:
img_side: Normalized accumulated array
direction: Direction of the route
rRad: Road radius approximate value
n_lanes: Number of lanes default
start_point: Starting point of lane (Col, Row)
Output Parameters:
max_length_lanes: Returns lane information of the route with lane points, polygons
'''
img_height = img_side.shape[0]
img_width = img_side.shape[1]
ad = AnchorDrawing()
edges = ad.Draw(img_side)
len_indices = []
[len_indices.append(len(item)) for item in edges]
areas_index = np.argsort(np.array(len_indices))
max_length_lanes = []
for lNum in range(-1, -(n_lanes+1), -1):
lane_info={}
lpoints = edges[areas_index[lNum]]
if direction ==0:
if (lpoints[0][1] > lpoints[-1][1]):
lpoints.reverse()
elif direction == 90:
if (lpoints[0][0] > lpoints[-1][0]):
lpoints.reverse()
if start_point:
row_vector= [p[0] for p in lpoints]
if start_point[1] in row_vector:
idx = row_vector.index(start_point[1])
lpoints = lpoints[idx:]
elif direction == 225:
if (lpoints[0][1] < lpoints[-1][1]):
lpoints.reverse()
lane_info['lane_points'] = [(p[1], p[0]) for p in lpoints]
line1=[]
line2=[]
ang = 0
# Compute polygon area using lane points and road radius
for pnum in range(0, len(lpoints)):
if (pnum < len(lpoints)-1):
ang = find_ang((lpoints[pnum][1],lpoints[pnum][0]), (lpoints[pnum+1][1], lpoints[pnum+1][0]))
if (ang == 45.0 ):# Push in the form of x,y
line1.append(( max(0,lpoints[pnum][1]-rRad),min(img_height, lpoints[pnum][0]+rRad)))
line2.append((min(img_width, lpoints[pnum][1] + rRad) , max(0, lpoints[pnum][0] - rRad)) )
elif(ang == 225.0):
line1.append((min(img_width, lpoints[pnum][1] + rRad), max(0, lpoints[pnum][0] - rRad)) )
line2.append((max(0, lpoints[pnum][1] - rRad), min(img_height, lpoints[pnum][0] + rRad)))
elif(ang == 0.0):
line1.append((lpoints[pnum][1] , min(img_height, lpoints[pnum][0] + rRad)))
line2.append((lpoints[pnum][1] , max(0, lpoints[pnum][0] - rRad)))
elif (ang == 180.0):
line1.append((lpoints[pnum][1], max(0, lpoints[pnum][0] - rRad)) )
line2.append((lpoints[pnum][1], min(img_height, lpoints[pnum][0] + rRad)))
elif(ang == 135.0):
line1.append((max(0, lpoints[pnum][1] - rRad), max(0, lpoints[pnum][0] - rRad)))
line2.append((min(img_width, lpoints[pnum][1] + rRad), min(img_height, lpoints[pnum][0] + rRad)))
elif (ang == 315.0):
line1.append((min(img_width, lpoints[pnum][1] + rRad), min(img_height, lpoints[pnum][0] + rRad)) )
line2.append((max(0, lpoints[pnum][1] - rRad), max(0, lpoints[pnum][0] - rRad)))
elif(ang == 90.0):
line1.append((max(0, lpoints[pnum][1] - rRad), lpoints[pnum][0]))
line2.append((min(img_width, lpoints[pnum][1] + rRad), lpoints[pnum][0]))
elif (ang == 270.0):
line1.append((min(img_width, lpoints[pnum][1] + rRad), lpoints[pnum][0]) )
line2.append((max(0, lpoints[pnum][1] - rRad), lpoints[pnum][0]))
line2.reverse()
poly_lane = line1+line2
poly_lane.append(line1[0])
polygon = Polygon(poly_lane)
lane_info['poly'] = polygon
lane_info['vertices'] = poly_lane
max_length_lanes.append(lane_info)
return max_length_lanes
def get_accumulator(trajectory_data, padd, routes_information):
'''
Computes accumulated array with all the vehicle trajectories
Input Parameters:
trajectory_data: Data contains vehicle trajectory points and properties
padd: Padding value for both rows and columns
routes_information: Contains route direction, orientation ranges
Output Parameters:
accum: Dictionary with accumulated array for all the routes
'''
adj_ang = -15 # Angle adjustment to keep routes aligned with clear direction
point_step = 30 # point step to get smooth line points
ang_step = 2 # angle step to find angle/direction of vehicle in trajectory
accum_weights = [[1, 1, 1],
[1, 2, 1],
[1, 1, 1]] # accumulator weights array highlights hehicle path
accum_weights = np.array(accum_weights, np.uint8) * 2
object_list = trajectory_data['object_list']
traj_props = trajectory_data['traj_props']
padd_x = padd
padd_y = padd
acc_w =traj_props['img_width']+ padd+padd+1
acc_h = traj_props['img_height']+ padd+padd+1
routes_names = list(routes_information.keys())
accum = {}
for route_name in routes_names:
accum[route_name] = np.zeros([acc_h, acc_w], np.uint32)
num_obj = len(object_list)
for id_num in range(0, num_obj):
length = len(object_list[id_num]['trajectory']['time'])
for idx in range(0, length):
data['object_list'][id_num]['trajectory']['x'].append(
data['traj_props']['scale_trajectory'] * (object_list[id_num]['trajectory']['lon'][idx] - traj_props['lon_min']))
data['object_list'][id_num]['trajectory']['y'].append(
(data['traj_props']['scale_trajectory'] * (object_list[id_num]['trajectory']['lat'][idx] - traj_props['lat_min'])))
comp_line_points = []
xx1 = round(data['object_list'][id_num]['trajectory']['x'][0]) + padd_x
yy1 = round(data['object_list'][id_num]['trajectory']['y'][0]) + padd_y
start_point = min(point_step, length)
for idx in range(start_point, length - point_step, point_step):
xx2 = round(data['object_list'][id_num]['trajectory']['x'][idx]) + padd_x
yy2 = round(data['object_list'][id_num]['trajectory']['y'][idx]) + padd_y
points = get_line([xx1, yy1], [xx2, yy2])
[comp_line_points.append(p) for p in points if p not in comp_line_points]
xx1 = xx2
yy1 = yy2
# Compute angle of each point in line
list_ang = []
length = len(comp_line_points)
for idx in range(0, length):
ang = 0
if (length - ang_step > idx):
ang = find_ang(comp_line_points[idx], comp_line_points[idx + ang_step])
list_ang.append(ang)
for idx in range(ang_step, length):
for route_name in routes_names:
if (routes_information[route_name]['orientation_range'][1] < routes_information[route_name]['orientation_range'][0] ):
if (list_ang[idx] <= routes_information[route_name]['orientation_range'][1] + adj_ang or list_ang[idx] > routes_information[route_name]['orientation_range'][0] + adj_ang): # WW
accum[route_name][comp_line_points[idx][1] - padd:comp_line_points[idx][1] + padd + 1,
comp_line_points[idx][0] - padd: comp_line_points[idx][0] + padd + 1] += accum_weights # vec_acc_filter[6]
else:
if (list_ang[idx] <= routes_information[route_name]['orientation_range'][1] + adj_ang and list_ang[idx] > routes_information[route_name]['orientation_range'][0] + adj_ang): # WW
accum[route_name][comp_line_points[idx][1] - padd:comp_line_points[idx][1] + padd + 1,
comp_line_points[idx][0] - padd: comp_line_points[idx][0] + padd + 1] += accum_weights # vec_acc_filter[6]
return accum
def swap_data(lane_data, vehi_num1, vehi_num2):
'''
Swaps vehicle information in lanedata
Input Parameters:
lane_data: Contains vehciles information(points, ids, speed)
vehi_num1: Vehicle index in lane
vehi_num2: Vehicle index in lane
Output Parameters:
'''
# Swap vehicle properties
temp = lane_data['ID'][vehi_num1]
lane_data['ID'][vehi_num1] = lane_data['ID'][vehi_num2]
lane_data['ID'][vehi_num2] = temp
temp = lane_data['x'][vehi_num1]
lane_data['x'][vehi_num1] = lane_data['x'][vehi_num2]
lane_data['x'][vehi_num2] = temp
temp = lane_data['y'][vehi_num1]
lane_data['y'][vehi_num1] = lane_data['y'][vehi_num2]
lane_data['y'][vehi_num2] = temp
temp = lane_data['speed'][vehi_num1]
lane_data['speed'][vehi_num1] = lane_data['speed'][vehi_num2]
lane_data['speed'][vehi_num2] = temp
def sort_ids(routes_data , lane_name, _direction):
'''
Sorts vehicle IDs using position and direction of lane
Input Parameters:
routes_data: Contains route information along with vehicle data
lane_name: Route name
_direction: Direction of the route
Output Parameters:
'''
# Sort vehicle IDs using position and direction of lane
for lane_num in range(0, len(routes_data[lane_name])):
for vehi_num1 in range(0 , len(routes_data[lane_name][lane_num]['ID'])):
for vehi_num2 in range(vehi_num1+1 , len(routes_data[lane_name][lane_num]['ID'])):
if _direction == 0:
if(routes_data[lane_name][lane_num]['x'][vehi_num1] > routes_data[lane_name][lane_num]['x'][vehi_num2]):
swap_data(routes_data[lane_name][lane_num], vehi_num1, vehi_num2)
if _direction == 90:
if(routes_data[lane_name][lane_num]['y'][vehi_num1] > routes_data[lane_name][lane_num]['y'][vehi_num2]):
swap_data(routes_data[lane_name][lane_num], vehi_num1, vehi_num2)
if _direction == 225:
if (routes_data[lane_name][lane_num]['x'][vehi_num1] < routes_data[lane_name][lane_num]['x'][vehi_num2]):
swap_data(routes_data[lane_name][lane_num], vehi_num1, vehi_num2)
def finddist(p1, p2):
'''
Finds distance between two points
Input Parameters:
p1: (Col, Row)
p2: (Col, Row)
Output Parameters:
'''
return ((p1[0]-p2[0])*(p1[0]-p2[0]) + (p1[1]-p2[1])*(p1[1]-p2[1]) ) ** 0.5
def find_queue_info(routes_data, laneInformation, temporal_data, route_name, axis_vec ):
'''
Computes vehicles inqueue with respect to lane number in the route,
Estimates spillback positions according to queue information
Input Parameters:
routes_data: Contains route information along with vehicle data
laneInformation: Lane area information
temporal_data: Previous timeframe routes data
route_name: Name of the route
axis_vec: Axis of start point, end point of the lane
Output Parameters:
queue_spillback_data: Returns array of queues, spillback points with respect to lane, routes
'''
n_lanes = len(routes_data[route_name])
speed_threshold = 7 # vehicle speed threshold (if less consider as queue vehicle)
gap_bw_vehicles = 2 # maximum number of vehicles non stationary of vehicles
max_dist_gap = 30 # max distance gap in queue
threshold_spill_to_lane_startpoint = 20 # if spill happen at start position of lane avoide it
num_temporal_frams = 3 # number of temporal frames to be considered
out_queue_info = []
for l_num in range(0, n_lanes):
out_queue_info.append([])
spillback_data = {}
spillback_data['exist'] = False
queue_data = {}
queue_data['exist'] = False
new_ids_in_current_frame_q = []
route_queue_indices = []
for l_num in range(0, n_lanes): # iterate through lane numbers
list_vehicle_details = {}
list_vehicle_details['point'] = []
list_vehicle_details['index'] = []
# find stationary vehicles in lane
for v_num in range(0, len(routes_data[route_name][l_num]['ID'])): # iterate through vehicles in lane
if (routes_data[route_name][l_num]['speed'][v_num] < speed_threshold ):
list_vehicle_details['point'].append( [ routes_data[route_name][l_num]['x'][v_num] , routes_data[route_name][l_num]['y'][v_num]])
list_vehicle_details['index'].append(v_num)
_queue_list = []
_indices_list = []
_queue = []
_indices= []
# split queues in lane
for idx in range(0, len(list_vehicle_details['index'])-1):
if (list_vehicle_details['index'][idx+1] - list_vehicle_details['index'][idx] <= gap_bw_vehicles and
finddist(list_vehicle_details['point'][idx+1] ,list_vehicle_details['point'][idx]) < max_dist_gap):
if list_vehicle_details['point'][idx] not in _queue:
_queue.append(list_vehicle_details['point'][idx])
_indices.append(list_vehicle_details['index'][idx])
if list_vehicle_details['point'][idx+1] not in _queue:
_queue.append(list_vehicle_details['point'][idx+1])
_indices.append(list_vehicle_details['index'][idx+1])
if (idx == len(list_vehicle_details['index'])-2 ):
if (len(_queue) >1):
_queue_list.append(_queue)
_indices_list.append(_indices)
else:
if (len(_queue) > 1):
_queue_list.append(_queue)
_indices_list.append(_indices)
_queue = []
_indices = []
route_queue_indices.append(_indices_list)
# get queue vehicle information and points
if (len(_queue_list) > 0):
for qnum in range(0, len(_queue_list)):
if (len(_queue_list[qnum]) > 1):
queue_data['exist'] = True
list_axis1_values = [p[axis_vec[0]] for p in laneInformation[route_name][l_num]['lane_points']]
list_axis2_values = [p[axis_vec[1]] for p in laneInformation[route_name][l_num]['lane_points']]
near_val1 = list_axis1_values[min(range(len(list_axis1_values)), key=lambda i: abs(list_axis1_values[i] - _queue_list[qnum][0][axis_vec[0]]))]
near_val2 = list_axis2_values[min(range(len(list_axis2_values)), key=lambda i: abs(list_axis2_values[i] - _queue_list[qnum][-1][axis_vec[1]]))]
s_idx = list_axis1_values.index(near_val1)
e_idx = list_axis2_values.index(near_val2)
if (s_idx> e_idx):
temp = e_idx
e_idx = s_idx
s_idx = temp
qdetails = {}
q_points = []
for _q_line_point in range(s_idx, e_idx+1):
q_points.append(laneInformation[route_name][l_num]['lane_points'][_q_line_point])
qdetails['points'] = q_points
qdetails['n_vehicles'] = _indices_list[qnum]
out_queue_info[l_num].append(qdetails)
# Observing whether spill_back happening
if (temporal_data.count(0) == 0):
spillback_data['ID'] = []
spillback_data['points'] = []
previus_ids = []
for n_frame in range(-1, -num_temporal_frams, -1):
for l_num in range(0, n_lanes):
previus_ids.extend(temporal_data[n_frame][route_name][l_num]['ID'])
for l_num in range(0, n_lanes):
for current_id in routes_data[route_name][l_num]['ID']:
if (current_id not in previus_ids):
new_id_index = routes_data[route_name][l_num]['ID'].index(current_id)
for qind in range(0, len(route_queue_indices[l_num])):
if (qind > 0):
if new_id_index <= route_queue_indices[l_num][qind][0] and new_id_index >= route_queue_indices[l_num][qind-1][0-1]:
spill_point = [routes_data[route_name][l_num]['x'][new_id_index], routes_data[route_name][l_num]['y'][new_id_index]]
dist_from_start_point = finddist(laneInformation[route_name][l_num]['lane_points'][0], spill_point)
if (dist_from_start_point > threshold_spill_to_lane_startpoint):
#spill_back_ids.append(new_id_index)
spillback_data['exist'] = True
spillback_data['ID'].append(routes_data[route_name][l_num]['ID'][new_id_index])
spillback_data['points'].append(spill_point)
elif (qind ==0):
if new_id_index <= route_queue_indices[l_num][qind][0]:
spill_point = [routes_data[route_name][l_num]['x'][new_id_index], routes_data[route_name][l_num]['y'][new_id_index]]
dist_from_start_point = finddist(laneInformation[route_name][l_num]['lane_points'][0], spill_point)
if (dist_from_start_point > threshold_spill_to_lane_startpoint):
spillback_data['exist'] = True
#spill_back_ids.append(new_id_index)
spillback_data['ID'].append(routes_data[route_name][l_num]['ID'][new_id_index])
spillback_data['points'].append([routes_data[route_name][l_num]['x'][new_id_index], routes_data[route_name][l_num]['y'][new_id_index]])
queue_data['points'] = out_queue_info
queue_spillback_data = {}
queue_spillback_data['queue'] = queue_data
queue_spillback_data['spillback'] = spillback_data
return queue_spillback_data
def init_route_info(lane_names):
'''
Initializes route information
Input Parameters:
lane_names: Name of the route
Output Parameters:
routes_data: Initial route information
'''
routes_data={}
for route_name in lane_names:
routes_data[route_name] = []
for l_num in range(0,3):
lane_info = {}
lane_info['ID'] = []
lane_info['speed'] = []
lane_info['x'] = []
lane_info['y'] = []
routes_data[route_name].append(lane_info)
return routes_data
if __name__ == '__main__':
file_spill_back = open('spillback.txt', 'w') # Create results file for spill back posistions
max_length_parameter = 'points' # n_vehicles input parameter to compute maximum queue length points: trajectory length, n_vehicles: number of vehicles
file_name = 'competition_dataset.csv'
data = readdata(file_name)
traj_props = data['traj_props']
# Method 1
padd = 1
routes_accumulater_data = get_accumulator(data, padd, routes_information)
Lane_information = {}# Init lane information
route_names = list(routes_information.keys())
lane_array = {}
for route_name in route_names:
lane_array[route_name] = normalizeimg(routes_accumulater_data[route_name][padd:-padd-1, padd:-padd-1] )
Lane_information['LeofAlexandras_tw_28isOktovriou'] = getLanePoints(lane_array['LeofAlexandras_tw_28isOktovriou'], direction=0)
Lane_information['OktovriouIs28_tw_LeofAlexandras'] = getLanePoints(lane_array['OktovriouIs28_tw_LeofAlexandras'], direction=225)
# selecting OktovriouIs28_tw_South start point based on LeofAlexandras_tw_28isOktovriou start point
Lane_information['OktovriouIs28_tw_South'] = getLanePoints(lane_array['OktovriouIs28_tw_South'], direction=90,
start_point=Lane_information['LeofAlexandras_tw_28isOktovriou'][0]['lane_points'][0])
if DISP:
font = cv2.FONT_HERSHEY_SIMPLEX # Font style for display text
fontScale = 0.5 # fontscale for display text
color = (255, 0, 0) # font color for display text
thickness = 2 # font thickness for display text
legend = np.ones((250, 250, 3), np.uint8) * 255 # vehicle type legend initiolization
keys = list(obj_types.keys()) # object list
gap = 3 # gap pixes between each legend parameters
siz = 15 # size of each legend color box
for key in range(0, len(keys)):# drow legend on image
cv2.rectangle(legend, (gap, gap + (key * siz)), ((siz - gap), (siz - gap) + (key * siz)), obj_types[keys[key]]['color'], -1)
cv2.putText(legend, keys[key], (gap + siz, (siz - gap) + (key * siz)), font, fontScale, obj_types[keys[key]]['color'], 1, cv2.LINE_AA)
ui_img = np.ones((traj_props['img_height'], traj_props['img_width'] , 3), np.uint8) * 255 # Initiolize image to dispay trajectory
legend = np.flipud(legend) # flip virtically legend because world coordinate(Trajectory coordinates) to image coordinates
ui_img[-legend.shape[0]: , -legend.shape[1]:] = legend # add legend to initial trajectory image
# DROW LANE polygon lines
lane_colors = [(255, 200, 200), (200, 255, 200), (200, 200, 255)] # lane colors
for route in route_names:
for nLane in range(0, len(Lane_information[route])):
vert = Lane_information[route][nLane]['vertices'] # vertices
pts = np.array(vert, np.int32)
cv2.polylines(ui_img, [pts], False, lane_colors[nLane], 1)
start_time = 0
end_time = 813
time_step = 0.04
num_samples = int((end_time - start_time) / time_step)
final_sample = int(end_time / time_step)
vec_time = [int((tim_itr * 0.04 * 100) + 0.5) / 100.0 for tim_itr in range(0, num_samples)]
object_list = data['object_list']
temporal_info = [0] * 15 # initiolize buffer for temporal information
spill_back_data_list = []
for time_stamp in vec_time:
if DISP:
image = np.copy(ui_img)
routes_data = init_route_info(route_names)
for obj_id in range(0, len(object_list)):
if time_stamp in object_list[obj_id]['trajectory']['time']:
time_index = object_list[obj_id]['trajectory']['time'].index(time_stamp)
col_range = object_list[obj_id]['trajectory']['x'][time_index]
row_range = object_list[obj_id]['trajectory']['y'][time_index]
traj_point = Point(col_range, row_range) # logitude lattitude
if DISP:
cv2.rectangle(image, (round(col_range) - 1, round(row_range) - 1), (round(col_range) + 1, round(row_range) + 1), obj_types[object_list[obj_id]['type']]['color'], 1)
obj_allocate = True
for route in route_names:
if (obj_allocate):
for nLane in range(0, len(Lane_information[route])):
if(obj_allocate):
if (Lane_information[route][nLane]['poly'].contains(traj_point)):
routes_data[route][nLane]['ID'].append(object_list[obj_id]['track_id'] ) #
routes_data[route][nLane]['speed'].append(object_list[obj_id]['trajectory']['speed'][time_index])
routes_data[route][nLane]['x'].append(object_list[obj_id]['trajectory']['x'][time_index])
routes_data[route][nLane]['y'].append(object_list[obj_id]['trajectory']['y'][time_index])
obj_allocate = False
break
for route_name in route_names:
# Sort vehicle data in lane according to te position from intersection point
sort_ids(routes_data, route_name, routes_information[route_name]['direction'])
# Finding queue points if exists for all routes considering all vehicle speeds are 0 in queue
_queue_spillback_info = find_queue_info(routes_data,Lane_information,temporal_info ,route_name, routes_information[route_name]['lane_axis'] )
if _queue_spillback_info['queue']['exist']:
for l_num in range(0, len(_queue_spillback_info['queue']['points'])):
for _queue in _queue_spillback_info['queue']['points'][l_num]:
if (len(_queue[max_length_parameter]) > routes_information[route_name]['max_queue']['length']):
routes_information[route_name]['max_queue']['length'] = len(_queue['points'])
routes_information[route_name]['max_queue']['points'] = _queue['points']
routes_information[route_name]['max_queue']['time'] = time_stamp
routes_information[route_name]['max_queue']['n_vehicles'] =len(_queue['n_vehicles'])
if DISP:
pts = np.array(_queue['points'], np.int32)
cv2.polylines(image, [pts], False, (255,100,70), 2)
if _queue_spillback_info['spillback']['exist']:
for n_spill in range(0,len(_queue_spillback_info['spillback']['points'])):
spill_point = _queue_spillback_info['spillback']['points'][n_spill]
spill_id = _queue_spillback_info['spillback']['ID'][n_spill]
spill_index = object_list[int(spill_id) - 1]['trajectory']['time'].index(time_stamp)
spill_data = {}
spill_data['Longitude'] = str(object_list[int(spill_id)-1]['trajectory']['lon'][spill_index])
spill_data['Latitude'] = str(object_list[int(spill_id) - 1]['trajectory']['lon'][spill_index])
spill_data['Object_ID'] = spill_id
spill_data['Time'] = time_stamp
spill_data['SpillBack_Route_area'] = route_name
spill_back_data_list.append(spill_data)
if DISP:
spill_point = _queue_spillback_info['spillback']['points'][n_spill]
cv2.circle(image, (int(spill_point[0]), int(spill_point[1])), 4, (0,0,255), 2)
#exist1, slippback_data = find_spillback_info(temporal_info)
temporal_info.pop(0)
temporal_info.append(routes_data)
if DISP:
image = np.flipud(image)
image = np.array(image)
cv2.putText(image, 'Time: ' + str(time_stamp), (image.shape[1] - 400, 30), font, 0.7, (255, 100, 100), 1, cv2.LINE_AA)
cv2.imshow("UAS Trajectory View",image)
cv2.waitKey(1)
# Make a list for temporal information
with open('results.csv', mode='w') as results_file:
writer_q = csv.DictWriter(results_file, fieldnames = ['Route_name','Time_stamp','Maximum_queue_length_points','Maximum_queue_length_meters', 'Num_vehicles','Coordinates'])
writer_q.writeheader()
for route_name in route_names:
if (routes_information[route_name]['max_queue']['length'] > 0):
coordinates = []
for pnum in range(0, len(routes_information[route_name]['max_queue']['points'])):
coordinates.append((((routes_information[route_name]['max_queue']['points'][pnum][0] / float(data['traj_props']['scale_trajectory'])) + traj_props['lon_min']),
((routes_information[route_name]['max_queue']['points'][pnum][1] / float(data['traj_props']['scale_trajectory'])) + traj_props['lat_min'])))
distance = finddist(coordinates[0], coordinates[-1])
dist_m = distance * 112 * 1000 # world coordinates to meters
writer_q.writerow({'Route_name':route_name,
'Time_stamp': str(routes_information[route_name]['max_queue']['time']),
'Maximum_queue_length_points':routes_information[route_name]['max_queue']['length'],
'Maximum_queue_length_meters' :dist_m,
'Num_vehicles':routes_information[route_name]['max_queue']['n_vehicles'],
'Coordinates': coordinates}
)
if (len(spill_back_data_list) > 0 ):
spill_back_fields = list(spill_back_data_list[0].keys())
writer = csv.DictWriter(results_file, fieldnames=spill_back_fields)
writer.writeheader()
for spill in spill_back_data_list:
writer.writerow(spill)
```
| github_jupyter |
```
from pymorphit.pymorphit import Morphit
import re
import itertools
import random as rd
```
## Obiettivo: generare un set di frasi del tipo "dove si trovano le mie bollette"
### Scrivo delle liste-vocabolario immaginando possibili alternative lessicali
```
mia = ["mia", ""]
bolletta = ["bolletta", "bollette"]
essere = ["sono", "stare", "si* trova"] # l'asterisco permette di segnalare a pymorphit che la parola precedente è invariante
```
### Posso interrogare wordnet italiano se non mi vengono in mente sinonimi
```
def synonyms(lemma):
from nltk.corpus import wordnet as wn
synmorph = {}
wnet_lemmas = wn.lemmas(lemma, lang="ita")
for index, wnet_lemma in enumerate(wnet_lemmas):
syns = wnet_lemma.synset()
m = re.findall("\'([^']*)\'", str(syns))
synmorph.update({m[0]: syns.lemma_names(lang="ita")})
return synmorph
synonyms("bolletta")
```
### "fattura" per esempio è un buon sinonimo di 'bolletta', lo aggiungo alla mia variabile
```
bolletta.append("fattura")
```
## Le mie liste-vocabolario sono un po' piccole, ma vediamo quante combinazioni posso generare a partire da queste. Tante sono le combinazioni, tante le frasi uniche che posso generare!
### Scrivo una semplice funzione con itertools, che prende in argomento le mie variabili (nell'ordine naturale della frase)
```
def combo(*args):
slots = [i for i in args]
return [combo for combo in itertools.product(*slots)]
combos1 = combo(essere, mia, bolletta)
print(f'Le possibili combinazioni sono in tutto {len(combos1)}')
print()
combos1
```
### La lista 'bolletta' contiene le parole destinate a essere la testa della frase, ossia la parola alla quale tutte le altre si devono accordare per generare una frase morfologicamente corretta
### Qui entra in gioco Pymorphit: per ogni combinazione viene inizializzato l'oggetto Morphit corrispondente alla testa. Per ragioni di efficienza, le parole già viste da Morphit vengono stoccate (funzione store)
```
seen = {}
def store(w, seen=seen):
if w not in seen.keys():
seen[w] = Morphit(w, "NOUN")
return seen[w]
```
## Adesso ciclo sulla lista di combinazioni e genero le frasi, tutte correttamente concordate con i metodi di Pymorphit
```
gen_sents = []
for essere, mia, bill in combos1:
# h sta per head: la parola a cui si accordano le altri parti del discorso
h = store(bill)
# i verbi di tipo "essere" di accordano al soggetto "bolletta" e cosivvia.
gen_sents.append(f'dove {h.agr(essere)} {h.article()} {h.agr(mia)} {h.word}?')
gen_sents
```
### Ci sono un po' di doppi spazi. Puliamo le frasi con una funzione di formattazione
```
def formatt(frase):
punkt1 = re.sub(r'([A-z])( )([,.?!"])',r"\1\3", frase)
punkt2 = re.sub(r'(\')( )([A-z])',r"\1\3", punkt1)
formatted = re.sub(r"(^|[.?!])\s*([a-zA-Zè])", lambda p: p.group(0).upper(), punkt2)
return re.sub(" +"," ", formatted)
[formatt(i) for i in gen_sents]
```
## Adesso qualcosa di più elaborato...
```
non = ["dove","non"]
trovo = ["trovo", "vedo", "posso vedere"]
mia = ["mia", ""]
bolletta = ["bolletta", "bollette", "fatture"]
su = ["in", "su"]
area = ["area", "spazio"]
clienti = ['riservato', 'clienti*', "personale"]
combos2 = combo(non, trovo, mia, bolletta, su, area, clienti)
print(f'Le possibili combinazioni sono in tutto {len(combos2)}')
gen_sents = []
for non, trovo, mia, bolletta, su, area, clienti in combos2:
# qui abbiamo due teste, perché il secondo sintagma si accorda alla variabile "area"
h1 = store(bolletta)
h2 = store(area)
gen_sents.append(f'{non} {trovo} {h1.article()} {h1.agr(mia)} {h1.word} {h2.preposition(su)} {h2.word} {h2.agr(clienti)}')
# formatto
gen_sents_format = [formatt(i) for i in gen_sents]
```
### Anche se sono tutte diverse, la ripetitività di queste frasi è ovviamente molto alta, perché il vocabolario di riferimento è piccolo. Potrei volerne prendere solo un subset di 100 frasi
```
rd.sample(gen_sents_format, 100)
```
| github_jupyter |
```
import torch
import torch.nn as nn
class CopyGeneratorLoss(nn.Module):
"""Copy generator criterion."""
def __init__(self, vocab_size, force_copy, unk_index=-1,
ignore_index=-100, eps=1e-20):
super(CopyGeneratorLoss, self).__init__()
self.force_copy = force_copy
self.eps = eps
self.vocab_size = vocab_size
self.ignore_index = ignore_index
self.unk_index = unk_index
def forward(self, scores, align, target):
"""
Args:
scores (FloatTensor): ``(batch_size*tgt_len)`` x dynamic vocab size
whose sum along dim 1 is less than or equal to 1, i.e. cols
softmaxed.
align (LongTensor): ``(batch_size x tgt_len)``
target (LongTensor): ``(batch_size x tgt_len)``
"""
# probabilities assigned by the model to the gold indices vocabulary tokens
vocab_probs = scores.gather(1, target.unsqueeze(1)).squeeze(1)
print(vocab_probs)
# probability of tokens copied from source
# offset the indices by vocabulary size.
copy_ix = align.unsqueeze(1) + self.vocab_size
print(copy_ix)
copy_tok_probs = scores.gather(1, copy_ix).squeeze(1)
print(copy_tok_probs)
# Set scores for unk to 0 and add eps
# (those that should not be copied)
copy_tok_probs[align == self.unk_index] = 0
copy_tok_probs += self.eps # to avoid -inf logs
# find the indices in which you do not use the copy mechanism
non_copy = align == self.unk_index # tensor([-1, 1, 2, -1, -1, -1, -1])
print(non_copy)
print(self.unk_index)
# If copy then use copy probs
# If non-copy then use vocab probs
probs = torch.where(
non_copy, copy_tok_probs + vocab_probs, copy_tok_probs
)
print(probs)
loss = -probs.log() # just NLLLoss; can the module be incorporated?
# Drop padding.
loss[target == self.ignore_index] = 0
return loss
vocab_size = 4 # special tokens
batch_size = 1
tgt_len = 7
copy_size = 3 # input entity embed
my_scores = torch.zeros(batch_size * tgt_len, vocab_size + copy_size)
for i in range(batch_size * tgt_len):
my_scores[i,:] = torch.arange(7)
my_scores
tgt_plan = torch.tensor([-2, 1, 2, -1, -1, -3, -4])
my_target = torch.randint(low=0, high=vocab_size - 1, size=(batch_size * tgt_len,)).long().view(-1)
my_target = torch.where(tgt_plan < 0, tgt_plan, 0) + 4
my_target
my_align = torch.randint(low=-1, high=copy_size - 1, size=(batch_size * tgt_len,)).long().view(-1)
my_align = torch.where(tgt_plan >= 0, tgt_plan, -1)
my_align
loss = CopyGeneratorLoss(vocab_size, force_copy=False)
loss(my_scores, my_align, my_target)
class CopyGenerator(nn.Module):
"""An implementation of pointer-generator networks
:cite:`DBLP:journals/corr/SeeLM17`.
These networks consider copying words
directly from the source sequence.
The copy generator is an extended version of the standard
generator that computes three values.
* :math:`p_{softmax}` the standard softmax over `tgt_dict`
* :math:`p(z)` the probability of copying a word from
the source
* :math:`p_{copy}` the probility of copying a particular word.
taken from the attention distribution directly.
The model returns a distribution over the extend dictionary,
computed as
:math:`p(w) = p(z=1) p_{copy}(w) + p(z=0) p_{softmax}(w)`
Args:
input_size (int): size of input representation
output_size (int): size of output vocabulary
pad_idx (int)
"""
def __init__(self, input_size, output_size, pad_idx):
super(CopyGenerator, self).__init__()
self.linear = nn.Linear(input_size, output_size)
self.linear_copy = nn.Linear(input_size, 1)
self.pad_idx = pad_idx
def forward(self, hidden, attn, src_map):
"""
Compute a distribution over the target dictionary
extended by the dynamic dictionary implied by copying
source words.
Args:
hidden (FloatTensor): hidden outputs ``(batch x tlen, input_size)``
attn (FloatTensor): attn for each ``(batch x tlen, input_size)``
src_map (FloatTensor):
A sparse indicator matrix mapping each source word to
its index in the "extended" vocab containing.
``(src_len, batch, extra_words)``
"""
# CHECKS
batch_by_tlen, _ = hidden.size()
batch_by_tlen_, slen = attn.size()
slen_, batch, cvocab = src_map.size()
aeq(batch_by_tlen, batch_by_tlen_)
aeq(slen, slen_)
# Original probabilities.
logits = self.linear(hidden)
logits[:, self.pad_idx] = -float('inf')
prob = torch.softmax(logits, 1)
# Probability of copying p(z=1) batch.
p_copy = torch.sigmoid(self.linear_copy(hidden))
# Probability of not copying: p_{word}(w) * (1 - p(z))
out_prob = torch.mul(prob, 1 - p_copy)
mul_attn = torch.mul(attn, p_copy)
copy_prob = torch.bmm(
mul_attn.view(-1, batch, slen).transpose(0, 1),
src_map.transpose(0, 1)
).transpose(0, 1)
copy_prob = copy_prob.contiguous().view(-1, cvocab)
return torch.cat([out_prob, copy_prob], 1)
vocab_size = 4 # doc_start, edu_end, doc_end, pad
pad_idx = -4
input_size = 5
batch_size = 2
tlen = 3 # sentence length
copy_generator = CopyGenerator(input_size, vocab_size, pad_idx)
input_emb = torch.zeros(batch_size * tgt_len, input_size)
for i in range(batch_size * tgt_len):
input_emb[i,:] = torch.rand(input_size)
input_emb
attn = torch.zeros(batch_size * tgt_len, input_size)
for i in range(batch_size * tgt_len):
attn[i,:] = torch.rand(input_size)
attn
copy_generator(input_emb, attn)
```
| github_jupyter |
# Experiment Size
We can use the knowledge of our desired practical significance boundary to plan out our experiment. By knowing how many observations we need in order to detect our desired effect to our desired level of reliability, we can see how long we would need to run our experiment and whether or not it is feasible.
Let's use the example from the video, where we have a baseline click-through rate of 10% and want to see a manipulation increase this baseline to 12%. How many observations would we need in each group in order to detect this change with power $1-\beta = .80$ (i.e. detect the 2% absolute increase 80% of the time), at a Type I error rate of $\alpha = .05$?
```
# import packages
import numpy as np
import scipy.stats as stats
import matplotlib.pyplot as plt
%matplotlib inline
```
## Method 1: Trial and Error
One way we could solve this is through trial and error. Every sample size will have a level of power associated with it; testing multiple sample sizes will gradually allow us to narrow down the minimum sample size required to obtain our desired power level. This isn't a particularly efficient method, but it can provide an intuition for how experiment sizing works.
Fill in the `power()` function below following these steps:
1. Under the null hypothesis, we should have a critical value for which the Type I error rate is at our desired alpha level.
- `se_null`: Compute the standard deviation for the difference in proportions under the null hypothesis for our two groups. The base probability is given by `p_null`. Remember that the variance of the difference distribution is the sum of the variances for the individual distributions, and that _each_ group is assigned `n` observations.
- `null_dist`: To assist in re-use, this should be a [scipy norm object](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.norm.html). Specify the center and standard deviation of the normal distribution using the "loc" and "scale" arguments, respectively.
- `p_crit`: Compute the critical value of the distribution that would cause us to reject the null hypothesis. One of the methods of the `null_dist` object will help you obtain this value (passing in some function of our desired error rate `alpha`).
2. The power is the proportion of the distribution under the alternative hypothesis that is past that previously-obtained critical value.
- `se_alt`: Now it's time to make computations in the other direction. This will be standard deviation of differences under the desired detectable difference. Note that the individual distributions will have different variances now: one with `p_null` probability of success, and the other with `p_alt` probability of success.
- `alt_dist`: This will be a scipy norm object like above. Be careful of the "loc" argument in this one. The way the `power` function is set up, it expects `p_alt` to be greater than `p_null`, for a positive difference.
- `beta`: Beta is the probability of a Type-II error, or the probability of failing to reject the null for a particular non-null state. That means you should make use of `alt_dist` and `p_crit` here!
The second half of the function has already been completed for you, which creates a visualization of the distribution of differences for the null case and for the desired detectable difference. Use the cells that follow to run the function and observe the visualizations, and to test your code against a few assertion statements. Check the following page if you need help coming up with the solution.
```
def power(p_null, p_alt, n, alpha = .05, plot = True):
"""
Compute the power of detecting the difference in two populations with
different proportion parameters, given a desired alpha rate.
Input parameters:
p_null: base success rate under null hypothesis
p_alt : desired success rate to be detected, must be larger than
p_null
n : number of observations made in each group
alpha : Type-I error rate
plot : boolean for whether or not a plot of distributions will be
created
Output value:
power : Power to detect the desired difference, under the null.
"""
# Compute the power
se_null = np.sqrt((p_null * (1-p_null) + p_null * (1-p_null)) / n)
null_dist = stats.norm(loc = 0, scale = se_null)
p_crit = null_dist.ppf(1 - alpha)
se_alt = np.sqrt((p_null * (1-p_null) + p_alt * (1-p_alt) ) / n)
alt_dist = stats.norm(loc = p_alt - p_null, scale = se_alt)
beta = alt_dist.cdf(p_crit)
if plot:
# Compute distribution heights
low_bound = null_dist.ppf(.01)
high_bound = alt_dist.ppf(.99)
x = np.linspace(low_bound, high_bound, 201)
y_null = null_dist.pdf(x)
y_alt = alt_dist.pdf(x)
# Plot the distributions
plt.plot(x, y_null)
plt.plot(x, y_alt)
plt.vlines(p_crit, 0, np.amax([null_dist.pdf(p_crit), alt_dist.pdf(p_crit)]),
linestyles = '--')
plt.fill_between(x, y_null, 0, where = (x >= p_crit), alpha = .5)
plt.fill_between(x, y_alt , 0, where = (x <= p_crit), alpha = .5)
plt.legend(['null','alt'])
plt.xlabel('difference')
plt.ylabel('density')
plt.show()
# return power
return (1 - beta)
power(.1, .12, 1000)
assert np.isclose(power(.1, .12, 1000, plot = False), 0.4412, atol = 1e-4)
assert np.isclose(power(.1, .12, 3000, plot = False), 0.8157, atol = 1e-4)
assert np.isclose(power(.1, .12, 5000, plot = False), 0.9474, atol = 1e-4)
print('You should see this message if all the assertions passed!')
```
## Method 2: Analytic Solution
Now that we've got some intuition for power by using trial and error, we can now approach a closed-form solution for computing a minimum experiment size. The key point to notice is that, for an $\alpha$ and $\beta$ both < .5, the critical value for determining statistical significance will fall between our null click-through rate and our alternative, desired click-through rate. So, the difference between $p_0$ and $p_1$ can be subdivided into the distance from $p_0$ to the critical value $p^*$ and the distance from $p^*$ to $p_1$.
<img src= 'images/ExpSize_Power.png'>
Those subdivisions can be expressed in terms of the standard error and the z-scores:
$$p^* - p_0 = z_{1-\alpha} SE_{0},$$
$$p_1 - p^* = -z_{\beta} SE_{1};$$
$$p_1 - p_0 = z_{1-\alpha} SE_{0} - z_{\beta} SE_{1}$$
In turn, the standard errors can be expressed in terms of the standard deviations of the distributions, divided by the square root of the number of samples in each group:
$$SE_{0} = \frac{s_{0}}{\sqrt{n}},$$
$$SE_{1} = \frac{s_{1}}{\sqrt{n}}$$
Substituting these values in and solving for $n$ will give us a formula for computing a minimum sample size to detect a specified difference, at the desired level of power:
$$n = \lceil \big(\frac{z_{\alpha} s_{0} - z_{\beta} s_{1}}{p_1 - p_0}\big)^2 \rceil$$
where $\lceil ... \rceil$ represents the ceiling function, rounding up decimal values to the next-higher integer. Implement the necessary variables in the function below, and test them with the cells that follow.
```
def experiment_size(p_null, p_alt, alpha = .05, beta = .20):
"""
Compute the minimum number of samples needed to achieve a desired power
level for a given effect size.
Input parameters:
p_null: base success rate under null hypothesis
p_alt : desired success rate to be detected
alpha : Type-I error rate
beta : Type-II error rate
Output value:
n : Number of samples required for each group to obtain desired power
"""
# Get necessary z-scores and standard deviations (@ 1 obs per group)
z_null = stats.norm.ppf(1 - alpha)
z_alt = stats.norm.ppf(beta)
sd_null = np.sqrt(p_null * (1-p_null) + p_null * (1-p_null))
sd_alt = np.sqrt(p_null * (1-p_null) + p_alt * (1-p_alt) )
# Compute and return minimum sample size
p_diff = p_alt - p_null
n = ((z_null*sd_null - z_alt*sd_alt) / p_diff) ** 2
return np.ceil(n)
experiment_size(.1, .12)
assert np.isclose(experiment_size(.1, .12), 2863)
print('You should see this message if the assertion passed!')
```
## Notes on Interpretation
The example explored above is a one-tailed test, with the alternative value greater than the null. The power computations performed in the first part will _not_ work if the alternative proportion is greater than the null, e.g. detecting a proportion parameter of 0.88 against a null of 0.9. You might want to try to rewrite the code to handle that case! The same issue should not show up for the second approach, where we directly compute the sample size.
If you find that you need to do a two-tailed test, you should pay attention to two main things. First of all, the "alpha" parameter needs to account for the fact that the rejection region is divided into two areas. Secondly, you should perform the computation based on the worst-case scenario, the alternative case with the highest variability. Since, for the binomial, variance is highest when $p = .5$, decreasing as $p$ approaches 0 or 1, you should choose the alternative value that is closest to .5 as your reference when computing the necessary sample size.
Note as well that the above methods only perform sizing for _statistical significance_, and do not take into account _practical significance_. One thing to realize is that if the true size of the experimental effect is the same as the desired practical significance level, then it's a coin flip whether the mean will be above or below the practical significance bound. This also doesn't even consider how a confidence interval might interact with that bound. In a way, experiment sizing is a way of checking on whether or not you'll be able to get what you _want_ from running an experiment, rather than checking if you'll get what you _need_.
## Alternative Approaches
There are also tools and Python packages that can also help with sample sizing decisions, so you don't need to solve for every case on your own. The sample size calculator [here](http://www.evanmiller.org/ab-testing/sample-size.html) is applicable for proportions, and provides the same results as the methods explored above. (Note that the calculator assumes a two-tailed test, however.) Python package "statsmodels" has a number of functions in its [`power` module](https://www.statsmodels.org/stable/stats.html#power-and-sample-size-calculations) that perform power and sample size calculations. Unlike previously shown methods, differences between null and alternative are parameterized as an effect size (standardized difference between group means divided by the standard deviation). Thus, we can use these functions for more than just tests of proportions. If we want to do the same tests as before, the [`proportion_effectsize`](http://www.statsmodels.org/stable/generated/statsmodels.stats.proportion.proportion_effectsize.html) function computes [Cohen's h](https://en.wikipedia.org/wiki/Cohen%27s_h) as a measure of effect size. As a result, the output of the statsmodel functions will be different from the result expected above. This shouldn't be a major concern since in most cases, you're not going to be stopping based on an exact number of observations. You'll just use the value to make general design decisions.
```
# example of using statsmodels for sample size calculation
from statsmodels.stats.power import NormalIndPower
from statsmodels.stats.proportion import proportion_effectsize
# leave out the "nobs" parameter to solve for it
NormalIndPower().solve_power(effect_size = proportion_effectsize(.12, .1), alpha = .05, power = 0.8,
alternative = 'larger')
```
| github_jupyter |
```
# Copyright 2021 Google LLC
# Use of this source code is governed by an MIT-style
# license that can be found in the LICENSE file or at
# https://opensource.org/licenses/MIT.
# Notebook authors: Kevin P. Murphy (murphyk@gmail.com)
# and Mahmoud Soliman (mjs@aucegypt.edu)
# This notebook reproduces figures for chapter 15 from the book
# "Probabilistic Machine Learning: An Introduction"
# by Kevin Murphy (MIT Press, 2021).
# Book pdf is available from http://probml.ai
```
<a href="https://opensource.org/licenses/MIT" target="_parent"><img src="https://img.shields.io/github/license/probml/pyprobml"/></a>
<a href="https://colab.research.google.com/github/probml/pml-book/blob/main/pml1/figure_notebooks/chapter15_neural_networks_for_sequences_figures.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Figure 15.1:<a name='15.1'></a> <a name='rnn'></a>
Recurrent neural network (RNN) for generating a variable length output sequence $ \bm y _ 1:T $ given an optional fixed length input vector $ \bm x $
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.1.png" width="256"/>
## Figure 15.2:<a name='15.2'></a> <a name='rnnTimeMachine'></a>
Example output of length 500 generated from a character level RNN when given the prefix ``the''. We use greedy decoding, in which the most likely character at each step is computed, and then fed back into the model. The model is trained on the book \em The Time Machine by H. G. Wells.
To reproduce this figure, click the open in colab button: <a href="https://colab.research.google.com/github/probml/probml-notebooks/blob/master/notebooks-d2l/rnn_torch.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
## Figure 15.3:<a name='15.3'></a> <a name='imageCaptioning'></a>
Illustration of a CNN-RNN model for image captioning. The pink boxes labeled ``LSTM'' refer to a specific kind of RNN that we discuss in \cref sec:LSTM . The pink boxes labeled $W_ \text emb $ refer to embedding matrices for the (sampled) one-hot tokens, so that the input to the model is a real-valued vector. From https://bit.ly/2FKnqHm . Used with kind permission of Yunjey Choi
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.3.png" width="256"/>
## Figure 15.4:<a name='15.4'></a> <a name='rnnBiPool'></a>
(a) RNN for sequence classification. (b) Bi-directional RNN for sequence classification
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.4_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.4_B.png" width="256"/>
## Figure 15.5:<a name='15.5'></a> <a name='biRNN'></a>
(a) RNN for transforming a sequence to another, aligned sequence. (b) Bi-directional RNN for the same task
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.5_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.5_B.png" width="256"/>
## Figure 15.6:<a name='15.6'></a> <a name='deepRNN'></a>
Illustration of a deep RNN. Adapted from Figure 9.3.1 of <a href='#dive'>[Zha+20]</a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.6.png" width="256"/>
## Figure 15.7:<a name='15.7'></a> <a name='seq2seq'></a>
Encoder-decoder RNN architecture for mapping sequence $ \bm x _ 1:T $ to sequence $ \bm y _ 1:T' $
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.7.png" width="256"/>
## Figure 15.8:<a name='15.8'></a> <a name='NMT'></a>
(a) Illustration of a seq2seq model for translating English to French. The - character represents the end of a sentence. From Figure 2.4 of <a href='#Luong2016thesis'>[Luo16]</a> . Used with kind permission of Minh-Thang Luong. (b) Illustration of greedy decoding. The most likely French word at each step is highlighted in green, and then fed in as input to the next step of the decoder. From Figure 2.5 of <a href='#Luong2016thesis'>[Luo16]</a> . Used with kind permission of Minh-Thang Luong
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.8_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.8_B.png" width="256"/>
## Figure 15.9:<a name='15.9'></a> <a name='BPTT'></a>
An RNN unrolled (vertically) for 3 time steps, with the target output sequence and loss node shown explicitly. From Figure 8.7.2 of <a href='#dive'>[Zha+20]</a> . Used with kind permission of Aston Zhang
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.9.png" width="256"/>
## Figure 15.10:<a name='15.10'></a> <a name='GRU'></a>
Illustration of a GRU. Adapted from Figure 9.1.3 of <a href='#dive'>[Zha+20]</a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.10.png" width="256"/>
## Figure 15.11:<a name='15.11'></a> <a name='LSTM'></a>
Illustration of an LSTM. Adapted from Figure 9.2.4 of <a href='#dive'>[Zha+20]</a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.11.png" width="256"/>
## Figure 15.12:<a name='15.12'></a> <a name='stsProb'></a>
Conditional probabilities of generating each token at each step for two different sequences. From Figures 9.8.1--9.8.2 of <a href='#dive'>[Zha+20]</a> . Used with kind permission of Aston Zhang
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.12_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.12_B.png" width="256"/>
## Figure 15.13:<a name='15.13'></a> <a name='beamSearch'></a>
Illustration of beam search using a beam of size $K=2$. The vocabulary is $\mathcal Y = \ A,B,C,D,E\ $, with size $V=5$. We assume the top 2 symbols at step 1 are A,C. At step 2, we evaluate $p(y_1=A,y_2=y)$ and $p(y_1=C,y_2=y)$ for each $y \in \mathcal Y $. This takes $O(K V)$ time. We then pick the top 2 partial paths, which are $(y_1=A,y_2=B)$ and $(y_1=C,y_2=E)$, and continue in the obvious way. Adapted from Figure 9.8.3 of <a href='#dive'>[Zha+20]</a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.13.png" width="256"/>
## Figure 15.14:<a name='15.14'></a> <a name='textCNN'></a>
Illustration of the TextCNN model for binary sentiment classification. Adapted from Figure 15.3.5 of <a href='#dive'>[Zha+20]</a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.14.png" width="256"/>
## Figure 15.15:<a name='15.15'></a> <a name='wavenet'></a>
Illustration of the wavenet model using dilated (atrous) convolutions, with dilation factors of 1, 2, 4 and 8. From Figure 3 of <a href='#wavenet'>[Aar+16]</a> . Used with kind permission of Aaron van den Oord
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.15.png" width="256"/>
## Figure 15.16:<a name='15.16'></a> <a name='attention'></a>
Attention computes a weighted average of a set of values, where the weights are derived by comparing the query vector to a set of keys. From Figure 10.3.1 of <a href='#dive'>[Zha+20]</a> . Used with kind permission of Aston Zhang
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.16.pdf" width="256"/>
## Figure 15.17:<a name='15.17'></a> <a name='attenRegression'></a>
Kernel regression in 1d. (a) Kernel weight matrix. (b) Resulting predictions on a dense grid of test points.
To reproduce this figure, click the open in colab button: <a href="https://colab.research.google.com/github/probml/probml-notebooks/blob/master/notebooks/kernel_regression_attention.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.17_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.17_B.png" width="256"/>
## Figure 15.18:<a name='15.18'></a> <a name='seq2seqAttn'></a>
Illustration of seq2seq with attention for English to French translation. Used with kind permission of Minh-Thang Luong
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.18.png" width="256"/>
## Figure 15.19:<a name='15.19'></a> <a name='translationHeatmap'></a>
Illustration of the attention heatmaps generated while translating two sentences from Spanish to English. (a) Input is ``hace mucho frio aqui.'', output is ``it is very cold here.''. (b) Input is ``¿todavia estan en casa?'', output is ``are you still at home?''. Note that when generating the output token ``home'', the model should attend to the input token ``casa'', but in fact it seems to attend to the input token ``?''. Adapted from https://www.tensorflow.org/tutorials/text/nmt_with_attention
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.19_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.19_B.png" width="256"/>
## Figure 15.20:<a name='15.20'></a> <a name='EHR'></a>
Example of an electronic health record. In this example, 24h after admission to the hospital, the RNN classifier predicts the risk of death as 19.9\%; the patient ultimately died 10 days after admission. The ``relevant'' keywords from the input clinical notes are shown in red, as identified by an attention mechanism. From Figure 3 of <a href='#Rajkomar2018'>[Alv+18]</a> . Used with kind permission of Alvin Rakomar
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.20.png" width="256"/>
## Figure 15.21:<a name='15.21'></a> <a name='SNLI'></a>
Illustration of sentence pair entailment classification using an MLP with attention to align the premise (``I do need sleep'') with the hypothesis (``I am tired''). White squares denote active attention weights, blue squares are inactive. (We are assuming hard 0/1 attention for simplicity.) From Figure 15.5.2 of <a href='#dive'>[Zha+20]</a> . Used with kind permission of Aston Zhang
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.21.png" width="256"/>
## Figure 15.22:<a name='15.22'></a> <a name='showAttendTell'></a>
Image captioning using attention. (a) Soft attention. Generates ``a woman is throwing a frisbee in a park''. (b) Hard attention. Generates ``a man and a woman playing frisbee in a field''. From Figure 6 of <a href='#showAttendTell'>[Kel+15]</a> . Used with kind permission of Kelvin Xu
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.22_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.22_B.png" width="256"/>
## Figure 15.23:<a name='15.23'></a> <a name='transformerTranslation'></a>
Illustration of how encoder self-attention for the word ``it'' differs depending on the input context. From https://ai.googleblog.com/2017/08/transformer-novel-neural-network.html . Used with kind permission of Jakob Uszkoreit
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.23.png" width="256"/>
## Figure 15.24:<a name='15.24'></a> <a name='multiHeadAttn'></a>
Multi-head attention. Adapted from Figure 9.3.3 of <a href='#dive'>[Zha+20]</a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.24.png" width="256"/>
## Figure 15.25:<a name='15.25'></a> <a name='positionalEncodingSinusoids'></a>
(a) Positional encoding matrix for a sequence of length $n=60$ and an embedding dimension of size $d=32$. (b) Basis functions for columsn 6 to 9.
To reproduce this figure, click the open in colab button: <a href="https://colab.research.google.com/github/probml/probml-notebooks/blob/master/notebooks-d2l/positional_encoding.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.25_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.25_B.png" width="256"/>
## Figure 15.26:<a name='15.26'></a> <a name='transformer'></a>
The transformer. From <a href='#Weng2018attention'>[Lil18]</a> . Used with kind permission of Lilian Weng. Adapted from Figures 1--2 of <a href='#Vaswani2017'>[Ash+17]</a>
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.26.png" width="256"/>
## Figure 15.27:<a name='15.27'></a> <a name='attentionBakeoff'></a>
Comparison of (1d) CNNs, RNNs and self-attention models. From Figure 10.6.1 of <a href='#dive'>[Zha+20]</a> . Used with kind permission of Aston Zhang
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.27.png" width="256"/>
## Figure 15.28:<a name='15.28'></a> <a name='VIT'></a>
The Vision Transformer (ViT) model. This treats an image as a set of input patches. The input is prepended with the special CLASS embedding vector (denoted by *) in location 0. The class label for the image is derived by applying softmax to the final ouput encoding at location 0. From Figure 1 of <a href='#ViT'>[Ale+21]</a> . Used with kind permission of Alexey Dosovitski
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.28.png" width="256"/>
## Figure 15.29:<a name='15.29'></a> <a name='transformers_taxonomy'></a>
Venn diagram presenting the taxonomy of different efficient transformer architectures. From <a href='#Tay2020transformers'>[Yi+20]</a> . Used with kind permission of Yi Tay
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.29.pdf" width="256"/>
## Figure 15.30:<a name='15.30'></a> <a name='rand_for_fast_atten'></a>
Attention matrix $\mathbf A $ rewritten as a product of two lower rank matrices $\mathbf Q ^ \prime $ and $(\mathbf K ^ \prime )^ \mkern -1.5mu\mathsf T $ with random feature maps $\boldsymbol \phi ( \bm q _i) \in \mathbb R ^M$ and $\boldsymbol \phi ( \bm v _k) \in \mathbb R ^M$ for the corresponding queries/keys stored in the rows/columns. Used with kind permission of Krzysztof Choromanski
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.30.png" width="256"/>
## Figure 15.31:<a name='15.31'></a> <a name='fatten'></a>
Decomposition of the attention matrix $\mathbf A $ can be leveraged to improve attention computations via matrix associativity property. To compute $\mathbf AV $, we first calculate $\mathbf G =( \bm k ^ \prime )^ \mkern -1.5mu\mathsf T \mathbf V $ and then $ \bm q ^ \prime \mathbf G $, resulting in linear in $N$ space and time complexity. Used with kind permission of Krzysztof Choromanski
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.31.png" width="256"/>
## Figure 15.32:<a name='15.32'></a> <a name='elmo'></a>
Illustration of ELMo bidrectional language model. Here $y_t=x_ t+1 $ when acting as the target for the forwards LSTM, and $y_t = x_ t-1 $ for the backwards LSTM. (We add \text \em bos \xspace and \text \em eos \xspace sentinels to handle the edge cases.) From <a href='#Weng2019LM'>[Lil19]</a> . Used with kind permission of Lilian Weng
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.32.png" width="256"/>
## Figure 15.33:<a name='15.33'></a> <a name='GPT'></a>
Illustration of (a) BERT and (b) GPT. $E_t$ is the embedding vector for the input token at location $t$, and $T_t$ is the output target to be predicted. From Figure 3 of <a href='#bert'>[Jac+19]</a> . Used with kind permission of Ming-Wei Chang
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.33_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.33_B.png" width="256"/>
## Figure 15.34:<a name='15.34'></a> <a name='bertEmbedding'></a>
Illustration of how a pair of input sequences, denoted A and B, are encoded before feeding to BERT. From Figure 14.8.2 of <a href='#dive'>[Zha+20]</a> . Used with kind permission of Aston Zhang
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.34.png" width="256"/>
## Figure 15.35:<a name='15.35'></a> <a name='bert-tasks'></a>
Illustration of how BERT can be used for different kinds of supervised NLP tasks. (a) Single sentence classification (e.g., sentiment analysis); (b) Sentence-pair classification (e.g., textual entailment); (d) Single sentence tagging (e.g., shallow parsing); (d) Question answering. From Figure 4 of <a href='#bert'>[Jac+19]</a> . Used with kind permission of Ming-Wei Chang
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.35_A.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.35_B.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.35_C.png" width="256"/>
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.35_D.png" width="256"/>
## Figure 15.36:<a name='15.36'></a> <a name='T5'></a>
Illustration of how the T5 model (``Text-to-text Transfer Transformer'') can be used to perform multiple NLP tasks, such as translating English to German; determining if a sentence is linguistic valid or not ( \bf CoLA stands for ``Corpus of Linguistic Acceptability''); determining the degree of semantic similarity ( \bf STSB stands for ``Semantic Textual Similarity Benchmark''); and abstractive summarization. From Figure 1 of <a href='#T5'>[Col+19]</a> . Used with kind permission of Colin Raffel
```
#@title Click me to run setup { display-mode: "form" }
try:
if PYPROBML_SETUP_ALREADY_RUN:
print('skipping setup')
except:
PYPROBML_SETUP_ALREADY_RUN = True
print('running setup...')
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
%reload_ext autoreload
%autoreload 2
!pip install superimport deimport -qqq
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport)
except Exception as e:
print(e)
print('finished!')
```
<img src="https://raw.githubusercontent.com/probml/pml-book/main/pml1/figures/images/Figure_15.36.png" width="256"/>
## References:
<a name='wavenet'>[Aar+16]</a> V. Aaron, D. Sander, Z. Heiga, S. Karen, V. Oriol, G. Alex, K. Nal, S. Andrew and K. Koray. "WaveNet: A Generative Model for Raw Audio". abs/1609.03499 (2016). arXiv: 1609.03499
<a name='ViT'>[Ale+21]</a> D. Alexey, B. Lucas, K. A. Dirk, Z. Xiaohua, U. T. Mostafa, M. Matthias, H. G. Sylvain, U. Jakob and H. Neil. "An Image is Worth 16x16 Words: Transformers for ImageRecognition at Scale". (2021).
<a name='Rajkomar2018'>[Alv+18]</a> R. Alvin, O. Eyal, C. Kai, D. A. Nissan, H. Michaela, L. PeterJ, L. LiuXiaobing, M. Jake, S. Mimi, S. Patrik, Y. Hector, Z. Kun, Z. Yi, F. Gerardo, D. GavinE, I. Jamie, L. Quoc, L. K. Alexander, T. Justin, W. De, W. James, W. Jimbo, L. Dana, V. L, C. Katherine, P. Michael, M. MadabushiSrinivasan, S. NigamH, B. AtulJ, H. D, C. Claire, C. GregS and D. Jeffrey. "Scalable and accurate deep learning with electronic healthrecords". In: NPJ Digit Med (2018).
<a name='Vaswani2017'>[Ash+17]</a> V. Ashish, S. Noam, P. Niki, U. Jakob, J. Llion, G. AidanN, K. KaiserLukasz and P. Illia. "Attention Is All You Need". (2017).
<a name='T5'>[Col+19]</a> R. Colin, S. Noam, R. Adam, L. LeeKatherine, N. Sharan, M. Michael, Z. ZhouYanqi, L. Wei and L. PeterJ. "Exploring the Limits of Transfer Learning with a UnifiedText-to-Text Transformer". abs/1910.10683 (2019). arXiv: 1910.10683
<a name='bert'>[Jac+19]</a> D. Jacob, C. Ming-Wei, L. Kenton and T. ToutanovaKristina. "BERT: Pre-training of Deep Bidirectional Transformers forLanguage Understanding". (2019).
<a name='showAttendTell'>[Kel+15]</a> X. Kelvin, B. JimmyLei, K. Ryan, C. K. Aaron, S. Ruslan, Z. S and B. Yoshua. "Show, Attend and Tell: Neural Image Caption Generation withVisual Attention". (2015).
<a name='Weng2018attention'>[Lil18]</a> W. Lilian "Attention? Attention!". In: lilianweng.github.io/lil-log (2018).
<a name='Weng2019LM'>[Lil19]</a> W. Lilian "Generalized Language Models". In: lilianweng.github.io/lil-log (2019).
<a name='Luong2016thesis'>[Luo16]</a> M. Luong "Neural machine translation". (2016).
<a name='Tay2020transformers'>[Yi+20]</a> T. Yi, D. Mostafa, B. Dara and M. MetzlerDonald. "Efficient Transformers: A Survey". abs/2009.06732 (2020). arXiv: 2009.06732
<a name='dive'>[Zha+20]</a> A. Zhang, Z. Lipton, M. Li and A. Smola. "Dive into deep learning". (2020).
| github_jupyter |
<center>

## [mlcourse.ai](https://mlcourse.ai/) Open Machine learning course
<center> **Author:Natalia Domozhirova, slack: @ndomozhirova** <center>
# <center>Tutorial</center>
# <center>KERAS : easy way to construct the Neural Networks</center>
<center>

## Introduction
Keras is a high-level neural networks API, written in Python.
Major Keras features:
- its capable of running on top of TensorFlow, CNTK, or Theano.
- Keras allows for easy and fast prototyping and supports both Perceptrons, Convolutional networks and Recurrent networks (including LSTM), as well as their combinations.
- Keras is compatible with: Python 2.7-3.6.
To make the process more interesting let's consider the classification example from the real life.
## Example description
Let's take the task from one Hakaton, organized by some polypropylene producer this year. So, let’s consider the production of the polypropylene granules by the extruder. Extruder is a kind of “meat grinder” which has the knives at the end of the process which are cutting the output product onto the granules.
The problem is that sometimes the production mass has an irregular consistency and sticks to the knives. When there is a lot of stuck mass - knives can no longer function. In this case it is necessary to stop production process, which is very expensive. If we would catch the very beginning of such sticking process - there is a method to very quickly and painless clean the knives and continue production without stopping.
So, the task is to send the stop signal to operator a bit in advance (let’s say not later then 15 minutes before such event) – so that he would have a time for necessary manipulations.
<center>
<img src="http://i68.tinypic.com/2rr2glg.jpg" style="height:250px">
Now we have already preprocessed normalized dataset with the vectors of the system sensors' values (5,160 features) and 0/1 targets. It is already devided into the [train](https://drive.google.com/open?id=1TMlClLguxcXTOAJt8VKe-iLrndJuFShl) and [test](https://drive.google.com/open?id=1JonMu0wmMbUqcbSd17Qr2A3AhVF3nutZ).
Let's download and prepare to work our datasets. In the datasets there are targets in zero column and the timestamps -in the 1st column.So, let's extract our train and test matrix as well as our targets. Also we'll transform the targets to categorical -so to have as a result our targets as 2-dimentional vectors, i.e. the vectors of probabilities of 0 and 1.
```
import numpy as np
import pandas as pd
from keras.utils import np_utils
df_train = pd.read_csv("train2.tsv", sep="\t", header=None)
df_test = pd.read_csv("test2.tsv", sep="\t", header=None)
Y_train = np.array(df_train[0].values.astype(int))
Y_test = np.array(df_test[0].values.astype(int))
X_train = np.array(df_train.iloc[:, 2:].values.astype(float))
X_test = np.array(df_test.iloc[:, 2:].values.astype(float))
Y_train = Y_train.astype(np.int)
Y_test = Y_test.astype(np.int)
Y_train = np_utils.to_categorical(Y_train, 2)
Y_test = np_utils.to_categorical(Y_test, 2)
print(X_train.shape)
print(Y_train.shape)
print(X_test.shape)
print(Y_test.shape)
```
## The Neural Network construction
Let's consider how the simple Newral Network(NN), Multilayer Perceptron (MLP), with 3 hidden layers (as a baseline), constructed by Keras, could help us to solve this problem.
As we have hidden layers - this would be a Deep Neural Network.
Also, we can see, that we need to have 5160 neurons in the input layer, as this is the size of our vector X and 2 neurons in the last layer - as this is the size of our target (vs. the picture below, where there are 4 neurons on the output layer).
You can read, for example, [here](https://en.wikipedia.org/wiki/Multilayer_perceptron) or [here](https://towardsdatascience.com/meet-artificial-neural-networks-ae5939b1dd3a) some more information about MLP structure.
<center>
<img src="http://i66.tinypic.com/2d6tsm.jpg" style="height:250px">
The core data structure of Keras is a **_model_** - a way to organize layers. The simplest type of model is the **_Sequential_** model, a linear stack of layers, which is appropriate for MLP construction (for more complex architectures, you should use the Keras functional API, which allows to build arbitrary graphs of layers).
```
import keras
from keras import Sequential
model1 = Sequential()
```
After the type of model is defined, we need to consistently add layers as **_Dense_**. Stacking layers is as easy as **_.add()_**.
While adding the layer we need to define the **number of neurons** and ***Activation*** **functions** which we can tune afterwards. For the fist layer we also need to add the dimention of X vectors (***input_dim***). In our case this is 5,160. The last layer consists on 2 neurons exactly as our target vestors Y_train and Y_test do.
The **number of layers** can also be tuned.
```
from keras.layers import Activation, Dense
model1.add(Dense(64, input_dim=5160))
model1.add(Activation("relu"))
model1.add(Dense(64))
model1.add(Activation("sigmoid"))
model1.add(Dense(128))
model1.add(Activation("tanh"))
model1.add(Dense(2))
model1.add(Activation("softmax"))
```
Once our model looks good, we need to configure its learning process with **_.compile()_**.
Here we should also describe the **_loss_** **function** and **_metrics_** we want to use as well as **_optimizer_** (the type of the Gradient descent to be used) which seem appropriate in each particular case.
```
model1.compile(loss="categorical_crossentropy", optimizer="adam", metrics=["accuracy"])
```
Now we can iterate on our training data in batches with the ***batch_size*** we want, where X_train and y_train are Numpy arrays just like in the Scikit-Learn API.
Also we can define the number of ***epochs*** (i.e. the max number of the full cycles of model's training).
***Verbose=1*** just lets us see the summary of the current stage of calcualtions.
We can also printing our model parameters using ***model.summary()***.
It is also can be useful to see the **shapes** of X_train, y_train,X_test,y_test
Also, we can save **the best model version** during the trainig process via the ***callback_save*** option.
And there is a ***callback_early stop*** option to stop the training process when we don't have significant improvement(defined by the ***min_delta***) during the certain number of epochs (***patience***).
Now our first model is ready:
```
from keras.callbacks import EarlyStopping, ModelCheckpoint
model1.summary()
print(X_train.shape)
print(Y_train.shape)
print(X_test.shape)
print(Y_test.shape)
callback_save = ModelCheckpoint(
"best_model1.model1", monitor="val_acc", verbose=1, save_best_only=True, mode="auto"
)
callback_earlystop = EarlyStopping(
monitor="val_loss", min_delta=0, patience=10, verbose=1, mode="auto"
)
model1.fit(
X_train,
Y_train,
batch_size=20,
epochs=10000,
verbose=1,
validation_data=(X_test, Y_test),
callbacks=[callback_save, callback_earlystop],
)
```
**So, we got a baseline with Accuracy= 0.79**. It looks cool, as we even didn't tune anything yet!
Let's try to improve this result. For example, we can introduce **Dropout** - this is a kind of regularization for the Neral Networks.
The **level of drop out** (in the brackets, along with a ***seed***) is a probability of the exclusion from the certain layer the random choice neuron during the current calculations. So, drop outs help to prevent the NN overfitting.
Let's create the new model:
```
from keras.layers import Dropout
model2 = Sequential()
model2.add(Dense(64, input_dim=5160))
model2.add(Activation("relu"))
model2.add(Dropout(0.3, seed=123))
model2.add(Dense(64))
model2.add(Activation("sigmoid"))
model2.add(Dropout(0.4, seed=123))
model2.add(Dense(128))
model2.add(Activation("tanh"))
model2.add(Dropout(0.5, seed=123))
model2.add(Dense(2))
model2.add(Activation("softmax"))
model2.compile(loss="categorical_crossentropy", optimizer="adam", metrics=["accuracy"])
model2.summary()
print(X_train.shape)
print(Y_train.shape)
print(X_test.shape)
print(Y_test.shape)
callback_save = ModelCheckpoint(
"best_model2.model2", monitor="val_acc", verbose=1, save_best_only=True, mode="auto"
)
callback_earlystop = EarlyStopping(
monitor="val_loss", min_delta=0, patience=10, verbose=1, mode="auto"
)
model2.fit(
X_train,
Y_train,
batch_size=20,
epochs=10000,
verbose=1,
validation_data=(X_test, Y_test),
callbacks=[callback_save, callback_earlystop],
)
```
Thus, adding the drop-outs we've **increased Accuracy on the test up to 0.86830**
We can also **tune all gyper-parameters** like the **number of layers**, the **levels of drop-outs**, **activation functions**, **optimizer**, **the number of neurons** etc.
For this purposes we can use, for example, another very friendly and easy-to-apply - Hyperas library. The description with examples you can find [here](https://github.com/maxpumperla/hyperas).
As a result of such tuning we've got the following model configuration:
```
model3 = Sequential()
model3.add(Dense(64, input_dim=5160))
model3.add(Activation("relu"))
model3.add(Dropout(0.11729755246044238, seed=123))
model3.add(Dense(256))
model3.add(Activation("relu"))
model3.add(Dropout(0.8444244099007299, seed=123))
model3.add(Dense(1024))
model3.add(Activation("linear"))
model3.add(Dropout(0.41266207281071243, seed=123))
model3.add(Dense(256))
model3.add(Activation("relu"))
model3.add(Dropout(0.4844455237320119, seed=123))
model3.add(Dense(2))
model3.add(Activation("softmax"))
model3.compile(
loss="categorical_crossentropy", optimizer="rmsprop", metrics=["accuracy"]
)
model3.summary()
print(X_train.shape)
print(Y_train.shape)
print(X_test.shape)
print(Y_test.shape)
callback_save = ModelCheckpoint(
"best_model3.model3", monitor="val_acc", verbose=1, save_best_only=True, mode="auto"
)
callback_earlystop = EarlyStopping(
monitor="val_loss", min_delta=0, patience=10, verbose=1, mode="auto"
)
model3.fit(
X_train,
Y_train,
batch_size=60,
epochs=10000,
verbose=1,
validation_data=(X_test, Y_test),
callbacks=[callback_save, callback_earlystop],
)
```
**Now, with tunned parameters, we've managed to imporove Accuracy up to 0.88073**
With Keras it is also possible to use **L1/L2 weight regularizations** which allow to apply penalties on layer parameters or layer activity during optimization. These penalties are incorporated in the loss function that the network optimizers.Let's add some regularization on to the 1st layer.
```
from keras import regularizers
model4 = Sequential()
model4.add(
Dense(
64,
input_dim=5160,
kernel_regularizer=regularizers.l2(0.0015),
activity_regularizer=regularizers.l1(0.0015),
)
)
model3.add(Activation("relu"))
model4.add(Dropout(0.11729755246044238, seed=123))
model4.add(Dense(256))
model4.add(Activation("relu"))
model4.add(Dropout(0.8444244099007299, seed=123))
model4.add(Dense(1024))
model4.add(Activation("linear"))
model4.add(Dropout(0.41266207281071243, seed=123))
model4.add(Dense(256))
model4.add(Activation("relu"))
model4.add(Dropout(0.4844455237320119, seed=123))
model4.add(Dense(2))
model4.add(Activation("softmax"))
model4.compile(
loss="categorical_crossentropy", optimizer="rmsprop", metrics=["accuracy"]
)
model4.summary()
print(X_train.shape)
print(Y_train.shape)
print(X_test.shape)
print(Y_test.shape)
callback_save = ModelCheckpoint(
"best_model4.model4", monitor="val_acc", verbose=1, save_best_only=True, mode="auto"
)
callback_earlystop = EarlyStopping(
monitor="val_loss", min_delta=0, patience=10, verbose=1, mode="auto"
)
model4.fit(
X_train,
Y_train,
batch_size=60,
epochs=10000,
verbose=1,
validation_data=(X_test, Y_test),
callbacks=[callback_save, callback_earlystop],
)
```
So, we can see, that adding regualrization with the current coeffitients to the firs layer we've got just **Accuracy of 0.84421** which didn't improve the result. This means, that, as usual, they should be carefully tuned :)
When we **want to use the best trained model** we got, we can just download previously (automatically) saved the best one (via ***load_model***) and apply to the data needed.
Let's see what we'll get on the test set:
```
from keras.models import load_model
model = load_model("best_model3.model3")
result = model.predict_on_batch(X_test)
result[:5]
```
You may also be interested to get a **list of all weight tensors** of the model, as Numpy arrays via ***get_weights***:
```
weights = model.get_weights()
weights[:1]
```
Besides, you would propbably like to get the **model config** to re-use it in the future. This can be done via ***get_config***:
```
config = model.get_config()
config
```
So, the model can be **reinstantiated** from its config via ***from_config***:
```
model3 = Sequential.from_config(config)
```
For more model tuning options proposed by Keras pls see [here](https://keras.io/)
## What about the other types of the Neural Networks?
Yes, you can use the similar approach re the layers' construction principles for LSTM, CNN and some other types of the Deep Neural Networks. For more details pls see [here](https://keras.io/).
## References:
1. https://keras.io/
2. https://towardsdatascience.com/meet-artificial-neural-networks-ae5939b1dd3a
3. https://www.quantinsti.com/blog/installing-keras-python-r
4. https://livebook.manning.com/#!/book/deep-learning-with-python/chapter-7
5. https://github.com/hyperopt/hyperopt
| github_jupyter |
<i>Copyright (c) Microsoft Corporation. All rights reserved.</i>
<i>Licensed under the MIT License.</i>
# Pretraining word and entity embeddings
This notebook trains word embeddings and entity embeddings for DKN initializations.
```
from gensim.test.utils import common_texts, get_tmpfile
from gensim.models import Word2Vec
import time
from utils.general import *
import numpy as np
import pickle
from utils.task_helper import *
class MySentenceCollection:
def __init__(self, filename):
self.filename = filename
self.rd = None
def __iter__(self):
self.rd = open(self.filename, 'r', encoding='utf-8', newline='\r\n')
return self
def __next__(self):
line = self.rd.readline()
if line:
return list(line.strip('\r\n').split(' '))
else:
self.rd.close()
raise StopIteration
InFile_dir = 'data_folder/my'
OutFile_dir = 'data_folder/my/pretrained-embeddings'
OutFile_dir_KG = 'data_folder/my/KG'
OutFile_dir_DKN = 'data_folder/my/DKN-training-folder'
```
Wrod2vec [4] can learn high-quality distributed vector representations that capture a large number of precise syntactic and semantic word relationships. We use word2vec algorithm implemented in Gensim [5] to generate word embeddings.
<img src="https://recodatasets.z20.web.core.windows.net/kdd2020/images%2Fword2vec.JPG" width="300">
```
def train_word2vec(Path_sentences, OutFile_dir):
OutFile_word2vec = os.path.join(OutFile_dir, r'word2vec.model')
OutFile_word2vec_txt = os.path.join(OutFile_dir, r'word2vec.txt')
create_dir(OutFile_dir)
print('start to train word embedding...', end=' ')
my_sentences = MySentenceCollection(Path_sentences)
model = Word2Vec(my_sentences, size=32, window=5, min_count=1, workers=8, iter=10) # user more epochs for better accuracy
model.save(OutFile_word2vec)
model.wv.save_word2vec_format(OutFile_word2vec_txt, binary=False)
print('\tdone . ')
Path_sentences = os.path.join(InFile_dir, 'sentence.txt')
t0 = time.time()
train_word2vec(Path_sentences, OutFile_dir)
t1 = time.time()
print('time elapses: {0:.1f}s'.format(t1 - t0))
```
We leverage a graph embedding model to encode entities into embedding vectors.
<img src="https://recodatasets.z20.web.core.windows.net/kdd2020/images%2Fkg-embedding-math.JPG" width="600">
<img src="https://recodatasets.z20.web.core.windows.net/kdd2020/images%2Fkg-embedding.JPG" width="600">
We use an open-source implementation of TransE (https://github.com/thunlp/Fast-TransX) for generating knowledge graph embeddings:
```
!bash ./run_transE.sh
```
DKN take considerations of both the entity embeddings and its context embeddings.
<img src="https://recodatasets.z20.web.core.windows.net/kdd2020/images/context-embedding.JPG" width="600">
```
##### build context embedding
EMBEDDING_LENGTH = 32
entity_file = os.path.join(OutFile_dir_KG, 'entity2vec.vec')
context_file = os.path.join(OutFile_dir_KG, 'context2vec.vec')
kg_file = os.path.join(OutFile_dir_KG, 'train2id.txt')
gen_context_embedding(entity_file, context_file, kg_file, dim=EMBEDDING_LENGTH)
load_np_from_txt(
os.path.join(OutFile_dir_KG, 'entity2vec.vec'),
os.path.join(OutFile_dir_DKN, 'entity_embedding.npy'),
)
load_np_from_txt(
os.path.join(OutFile_dir_KG, 'context2vec.vec'),
os.path.join(OutFile_dir_DKN, 'context_embedding.npy'),
)
format_word_embeddings(
os.path.join(OutFile_dir, 'word2vec.txt'),
os.path.join(InFile_dir, 'word2idx.pkl'),
os.path.join(OutFile_dir_DKN, 'word_embedding.npy')
)
```
## Reference
\[1\] Wang, Hongwei, et al. "DKN: Deep Knowledge-Aware Network for News Recommendation." Proceedings of the 2018 World Wide Web Conference on World Wide Web. International World Wide Web Conferences Steering Committee, 2018.<br>
\[2\] Knowledge Graph Embeddings including TransE, TransH, TransR and PTransE. https://github.com/thunlp/KB2E <br>
of the 58th Annual Meeting of the Association for Computational Linguistics. https://msnews.github.io/competition.html <br>
\[3\] GloVe: Global Vectors for Word Representation. https://nlp.stanford.edu/projects/glove/ <br>
\[4\] Tomas Mikolov, Ilya Sutskever, Kai Chen, Greg Corrado, and Jeffrey Dean. 2013. Distributed representations of words and phrases and their compositionality. In Proceedings of the 26th International Conference on Neural Information Processing Systems - Volume 2 (NIPS’13). Curran Associates Inc., Red Hook, NY, USA, 3111–3119. <br>
\[5\] Gensim Word2vec embeddings : https://radimrehurek.com/gensim/models/word2vec.html <br>
| github_jupyter |
<table style="width:100%; background-color: #D9EDF7">
<tr>
<td style="border: 1px solid #CFCFCF">
<b>Renewable power plants: Download and process Notebook</b>
<ul>
<li><a href="main.ipynb">Main Notebook</a></li>
<li>Download and process Notebook</li>
<li><a href="validation_and_output.ipynb">Validation and output Notebook</a></li>
</ul>
<br>This Notebook is part of the <a href="http://data.open-power-system-data.org/renewable_power_plants"> Renewable power plants Data Package</a> of <a href="http://open-power-system-data.org">Open Power System Data</a>.
</td>
</tr>
</table>
This script downlads and extracts the original data of renewable power plant lists from the data sources, processes and merges them. It subsequently adds the geolocation for each power plant. Finally it saves the DataFrames as pickle-files. Make sure you run the download and process Notebook before the validation and output Notebook.
# Table of contents
* [1. Script setup](#1.-Script-setup)
* [2. Settings](#2.-Settings)
* [2.1 Choose download option](#2.1-Choose-download-option)
* [2.2 Download function](#2.2-Download-function)
* [2.3 Setup translation dictionaries](#2.3-Setup-translation-dictionaries)
* [3. Download and process per country](#3.-Download-and-process-per-country)
* [3.1 Germany DE](#3.1-Germany-DE)
* [3.1.1 Download and read](#3.1.1-Download-and-read)
* [3.1.2 Translate column names](#3.1.2-Translate-column-names)
* [3.1.3 Add information and choose columns](#3.1.3-Add-information-and-choose-columns)
* [3.1.4 Merge DataFrames](#3.1.4-Merge-DataFrames)
* [3.1.5 Translate values and harmonize energy source](#3.1.5-Translate-values-and-harmonize-energy-source)
* [3.1.6 Transform electrical_capacity from kW to MW](#3.1.6-Transform-electrical_capacity-from-kW-to-MW)
* [3.1.7 Georeferencing](#3.1.7-Georeferencing)
* [3.1.8 Save](#3.1.8-Save)
* [3.2 Denmark DK](#3.2-Denmark-DK)
* [3.2.1 Download and read](#3.2.1-Download-and-read)
* [3.2.2 Translate column names](#3.2.2-Translate-column-names)
* [3.2.3 Add data source and missing information](#3.2.3-Add-data-source-and-missing-information)
* [3.2.4 Translate values and harmonize energy source](#3.2.4-Translate-values-and-harmonize-energy-source)
* [3.2.5 Georeferencing](#3.1.5-Georeferencing)
* [3.2.6 Merge DataFrames and choose columns](#3.2.6-Merge-DataFrames-and-choose-columns)
* [3.1.7 Transform electrical_capacity from kW to MW](#3.1.7-Transform-electrical_capacity-from-kW-to-MW)
* [3.2.8 Save](#3.1.8-Save)
* [3.3 France FR](#3.3-France-FR)
* [3.3.1 Download and read](#3.3.1-Download-and-read)
* [3.3.2 Rearrange columns and translate column names](#3.3.2-Rearragne-columns-and-translate-column-names)
* [3.3.3 Add data source](#3.3.3-Add-data-source)
* [3.3.4 Translate values and harmonize energy source](#3.3.4-Translate-values-and-harmonize-energy-source)
* [3.3.5 Georeferencing](#3.3.5-Georeferencing)
* [3.3.6 Save](#3.3.6-Save)
* [3.4 Poland PL](#3.4-Poland-PL)
* [3.4.1 Download and read](#3.4.1-Download-and-read)
* [3.4.2 Rearrange data from rtf-file](#3.4.2-Rearrange-data-from-rtf-file)
* [3.4.3 Add data source](#3.4.3-Add-data-source)
* [3.4.4 Translate values and harmonize energy source](#3.4.4-Translate-values-and-harmonize-energy-source)
* [3.4.5 Georeferencing -_work in progress_](#3.4.6-Georeferencing---work-in-progress)
* [3.4.6 Save](#3.4.7-Save)
* [Part 2: Validation and output](validation_and_output.ipynb)
# 1. Script setup
```
# importing all necessary Python libraries for this Script
from collections import OrderedDict
import io
import json
import os
import subprocess
import zipfile
import posixpath
import urllib.parse
import urllib.request
import numpy as np
import pandas as pd
import requests
import sqlite3
import logging
import getpass
import utm # for transforming geoinformation in the utm-format
import re # provides regular expression matching operations
# Starting from ipython 4.3.0 logging is not directing its ouput to the out cell. It might be operating system related but
# until the issue is fixed, we are going to use print().
# Issue on GitHub: https://github.com/ipython/ipykernel/issues/111
# Set up a log
logging.basicConfig(handlers=[logging.StreamHandler()])
logger = logging.getLogger('notebook')
logger.setLevel('INFO')
nb_root_logger = logging.getLogger()
formatter = logging.Formatter('%(asctime)s - %(name)s - %(levelname)s'\
'- %(message)s',datefmt='%d %b %Y %H:%M:%S')
# Create input and output folders if they don't exist
os.makedirs('input/original_data', exist_ok=True)
os.makedirs('output', exist_ok=True)
os.makedirs('output/renewable_power_plants', exist_ok=True)
```
# 2. Settings
## 2.1 Choose download option
The original data can either be downloaded from the original data sources as specified below or from the opsd-Server. Default option is to download from the original sources as the aim of the project is to stay as close to original sources as possible. However, if problems with downloads e.g. due to changing urls occur, you can still run the script with the original data from the opsd_server.
```
download_from = 'original_sources'
# download_from = 'opsd_server'
if download_from == 'opsd_server':
# While OPSD is in beta, we need to supply authentication
password = getpass.getpass('Please enter the beta user password:')
session = requests.session()
session.auth = ('beta', password)
# Specify direction to original_data folder on the opsd data server
url_opsd = 'http://data.open-power-system-data.org/renewables_power_plants/'
version = '2016-09-30'
folder = '/original_data'
```
## 2.2 Download function
```
def download_and_cache(url, session=None):
"""This function downloads a file into a folder called
original_data and returns the local filepath."""
path = urllib.parse.urlsplit(url).path
filename = posixpath.basename(path)
filepath = "input/original_data/" + filename
print(url)
# check if file exists, if not download it
filepath = "input/original_data/" + filename
print(filepath)
if not os.path.exists(filepath):
if not session:
print('No session')
session = requests.session()
print("Downloading file: ", filename)
r = session.get(url, stream=True)
chuncksize = 1024
with open(filepath, 'wb') as file:
for chunck in r.iter_content(chuncksize):
file.write(chunck)
else:
print("Using local file from", filepath)
filepath = '' + filepath
return filepath
```
## 2.3 Setup translation dictionaries
Column and value names of the original data sources will be translated to English and standardized across different sources. Standardized column names, e.g. "electrical_capacity" are required to merge data in one DataFrame.<br>
The column and the value translation lists are provided in the input folder of the Data Package.
```
# Get column translation list
columnnames = pd.read_csv('input/column_translation_list.csv')
# Get value translation list
valuenames = pd.read_csv('input/value_translation_list.csv')
```
# 3. Download and process per country
For one country after the other, the original data is downloaded, read, processed, translated, eventually georeferenced and saved. If respective files are already in the local folder, these will be utilized.
To process the provided data [pandas DataFrame](http://pandas.pydata.org/pandas-docs/stable/dsintro.html#dataframe) is applied.<br>
## 3.1 Germany DE
### 3.1.1 Download and read
The data which will be processed below is provided by the following data sources:
**[Netztransparenz.de](https://www.netztransparenz.de/de/Anlagenstammdaten.htm)** - Official grid transparency platform from the German TSOs (50Hertz, Amprion, TenneT and TransnetBW).
**Bundesnetzagentur (BNetzA)** - German Federal Network Agency for Electricity, Gas, Telecommunications, Posts and Railway (Data for [roof-mounted PV power plants](http://www.bundesnetzagentur.de/cln_1422/DE/Sachgebiete/ElektrizitaetundGas/Unternehmen_Institutionen/ErneuerbareEnergien/Photovoltaik/DatenMeldgn_EEG-VergSaetze/DatenMeldgn_EEG-VergSaetze_node.html) and for [all other renewable energy power plants](http://www.bundesnetzagentur.de/cln_1412/DE/Sachgebiete/ElektrizitaetundGas/Unternehmen_Institutionen/ErneuerbareEnergien/Anlagenregister/Anlagenregister_Veroeffentlichung/Anlagenregister_Veroeffentlichungen_node.html))
```
# point URLs to original data depending on the chosen download option
if download_from == 'original_sources':
url_netztransparenz ='https://www.netztransparenz.de/de/file/Anlagenstammdaten_2015_final.zip'
url_bnetza ='http://www.bundesnetzagentur.de/SharedDocs/Downloads/DE/Sachgebiete/Energie/Unternehmen_Institutionen/ErneuerbareEnergien/Anlagenregister/VOeFF_Anlagenregister/2016_06_Veroeff_AnlReg.xls?__blob=publicationFile&v=1'
url_bnetza_pv = 'https://www.bundesnetzagentur.de/SharedDocs/Downloads/DE/Sachgebiete/Energie/Unternehmen_Institutionen/ErneuerbareEnergien/Photovoltaik/Datenmeldungen/Meldungen_Aug-Mai2016.xls?__blob=publicationFile&v=2'
elif download_from == 'opsd_server':
url_netztransparenz = (url_opsd + version + folder + '/DE/Netztransparenz/' + 'Anlagenstammdaten_2015_final.zip')
url_bnetza = (url_opsd + version + folder + '/DE/BNetzA/' + '2016_06_Veroeff_AnlReg.xls')
url_bnetza_pv = (url_opsd + version + folder + '/DE/BNetzA/' + 'Meldungen_Aug-Mai2016.xls')
# Download all data sets before processing.
if download_from == 'original_sources':
netztransparenz_zip = %time zipfile.ZipFile(download_and_cache(url_netztransparenz))
bnetza_xls = %time download_and_cache(url_bnetza)
bnetza_pv_xls = %time download_and_cache(url_bnetza_pv)
elif download_from == 'opsd_server':
# Check if the user is offline
# if offline, do not give a session as parameter.
try:
online = True
r = session.get('http://data.open-power-system-data.org/renewables_power_plants/')
except requests.ConnectionError:
logger.warning('The user is offline. Proceeding with the script!')
try:
netztransparenz_zip = %time zipfile.ZipFile(download_and_cache(url_netztransparenz, session))
bnetza_xls = %time download_and_cache(url_bnetza, session)
bnetza_pv_xls = %time download_and_cache(url_bnetza_pv, session)
except zipfile.BadZipFile:
raise FileNotFoundError('One of the Zip File is corrupted! Delete them \
Also, check your opsd password!')
# Read TSO data from zip file
print('Reading Amprion_Anlagenstammdaten_2015.csv')
amprion_df = pd.read_csv(netztransparenz_zip.open('Amprion_Anlagenstammdaten_2015.csv'),
sep=';',
thousands='.',
decimal=',',
header=0,
parse_dates=[11, 12, 13, 14],
encoding='cp1252',
dayfirst=True)
print('Reading 50Hertz_Anlagenstammdaten_2015.csv')
hertz_df = pd.read_csv(netztransparenz_zip.open('50Hertz_Anlagenstammdaten_2015.csv'),
sep=';',
thousands='.',
decimal=',',
header=0,
parse_dates=[11, 12, 13, 14],
encoding='cp1252',
dayfirst=True)
print('Reading TenneT_Anlagenstammdaten_2015.csv')
tennet_df = pd.read_csv(netztransparenz_zip.open('TenneT_Anlagenstammdaten_2015.csv'),
sep=';',
thousands='.',
decimal=',',
header=0,
parse_dates=[11, 12, 13, 14],
encoding='cp1252',
dayfirst=True)
print('Reading TransnetBW_Anlagenstammdaten_2015.csv')
transnetbw_df = pd.read_csv(netztransparenz_zip.open('TransnetBW_Anlagenstammdaten_2015.csv'),
sep=';',
thousands='.',
decimal=',',
header=0,
parse_dates=[11, 12, 13, 14],
encoding='cp1252',
dayfirst=True,
low_memory=False)
# Read BNetzA register
print('Reading bnetza - 2016_06_Veroeff_AnlReg.xls')
bnetza_df = pd.read_excel(bnetza_xls,
sheetname='Gesamtübersicht',
header=0,
converters={'4.9 Postleit-zahl': str,
'Gemeinde-Schlüssel': str})
# Read BNetzA-PV register
print('Reading bnetza_pv - Meldungen_Aug-Mai2016.xls')
bnetza_pv = pd.ExcelFile(bnetza_pv_xls)
# Combine all PV BNetzA sheets into one DataFrame
print('Concatenating bnetza_pv')
bnetza_pv_df = pd.concat(bnetza_pv.parse(sheet, skiprows=10,
converters={'Anlage \nPLZ': str}
) for sheet in bnetza_pv.sheet_names)
# Drop not needed NULL "Unnamed:" column
bnetza_pv_df = bnetza_pv_df.drop(bnetza_pv_df.columns[[7]], axis=1)
```
### 3.1.2 Translate column names
To standardise the DataFrame the original column names from the German TSOs and the BNetzA wil be translated and new english column names wil be assigned to the DataFrame. The unique column names are required to merge the DataFrame.<br>
The column_translation_list is provided here as csv in the input folder. It is loaded in _2.3 Setup of translation dictionaries_.
```
# Choose the translation terms for Germany, create dictionary and show dictionary
idx_DE = columnnames[columnnames['country'] == 'DE'].index
column_dict_DE = columnnames.loc[idx_DE].set_index('original_name')['opsd_name'].to_dict()
column_dict_DE
print('Translation')
amprion_df.rename(columns=column_dict_DE, inplace=True)
hertz_df.rename(columns=column_dict_DE, inplace=True)
tennet_df.rename(columns=column_dict_DE, inplace=True)
transnetbw_df.rename(columns=column_dict_DE, inplace=True)
bnetza_df.rename(columns=column_dict_DE, inplace=True)
bnetza_pv_df.rename(columns=column_dict_DE, inplace=True)
```
### 3.1.3 Add information and choose columns
All data source names and (for the BNetzA-PV data) the energy source will is added.
```
# Add data source names to the DataFrames
transnetbw_df['data_source'] = 'TransnetBW'
tennet_df['data_source'] = 'TenneT'
amprion_df['data_source'] = 'Amprion'
hertz_df['data_source'] = '50Hertz'
bnetza_df['data_source'] = 'BNetzA'
bnetza_pv_df['data_source'] = 'BNetzA_PV'
# Add for the BNetzA PV data the energy source
bnetza_pv_df['energy_source'] = 'Photovoltaics'
# Correct datetime-format
def decom_fkt(x):
x = str(x)
if x == 'nan':
x = ''
else:
x = x[0:10]
return x
bnetza_df['decommissioning_date'] = bnetza_df['decommissioning_date'].apply(
decom_fkt)
# Just some of all the columns of this DataFrame are utilized further
bnetza_df = bnetza_df.loc[:,('commissioning_date','decommissioning_date','notification_reason',
'energy_source',
'electrical_capacity_kW','thermal_capacity_kW',
'voltage_level','dso','eeg_id','bnetza_id',
'federal_state','postcode','municipality_code','municipality',
'address','address_number',
'utm_zone','utm_east','utm_north',
'data_source')]
```
### 3.1.4 Merge DataFrames
The individual DataFrames from the TSOs (Netztransparenz.de) and BNetzA are merged.
```
dataframes = [transnetbw_df, tennet_df, amprion_df, hertz_df, bnetza_pv_df, bnetza_df]
DE_renewables = pd.concat(dataframes)
# Make sure the decommissioning_column has the right dtype
DE_renewables['decommissioning_date'] = pd.to_datetime(DE_renewables['decommissioning_date'])
DE_renewables.reset_index(drop=True, inplace=True)
```
**First look at DataFrame structure and format**
```
DE_renewables.info()
```
### 3.1.5 Translate values and harmonize energy source
Different German terms for energy source, energy source subtypes and voltage levels are translated and harmonized across the individual data sources. The value_translation_list is provided here as csv in the input folder. It is loaded in _2.3 Setup of translation dictionaries_.
```
# Choose the translation terms for Germany, create dictionary and show dictionary
idx_DE = valuenames[valuenames['country'] == 'DE'].index
value_dict_DE = valuenames.loc[idx_DE].set_index('original_name')['opsd_name'].to_dict()
value_dict_DE
print('replacing..')
# Running time: some minutes. %time prints the time your computer required for this step
%time DE_renewables.replace(value_dict_DE, inplace=True)
```
**Separate and assign energy source and subtypes**
```
# Create dctionnary in order to assign energy_source to its subtype
energy_source_dict_DE = valuenames.loc[idx_DE].set_index('opsd_name')['energy_source'].to_dict()
# Column energy_source partly contains subtype information, thus this column is copied
# to new column for energy_source_subtype...
DE_renewables['energy_source_subtype'] = DE_renewables['energy_source']
# ...and the energy source subtype values in the energy_source column are replaced by
# the higher level classification
DE_renewables['energy_source'].replace(energy_source_dict_DE, inplace=True)
# Overview of dictionary
energy_source_dict_DE
```
**Summary of DataFrame**
```
# Electrical capacity per energy_source (in MW)
DE_renewables.groupby(['energy_source'])['electrical_capacity_kW'].sum() / 1000
# Electrical capacity per energy_source_subtype (in MW)
DE_renewables.groupby(['energy_source_subtype'])['electrical_capacity_kW'].sum() / 1000
```
### 3.1.6 Transform electrical_capacity from kW to MW
```
# kW to MW
DE_renewables[['electrical_capacity_kW','thermal_capacity_kW']] /= 1000
# adapt column name
DE_renewables.rename(columns={'electrical_capacity_kW' : 'electrical_capacity',
'thermal_capacity_kW' : 'thermal_capacity'},inplace=True)
```
### 3.1.7 Georeferencing
#### Get coordinates by postcode
*(for data with no existing geocoordinates)*
The available post code in the original data provides a first approximation for the geocoordinates of the RE power plants.<br>
The BNetzA data provides the full zip code whereas due to data privacy the TSOs only report the first three digits of the power plant's post code (e.g. 024xx) and no address. Subsequently a centroid of the post code region polygon is used to find the coordinates.
With data from
* http://www.suche-postleitzahl.org/downloads?download=plz-gebiete.shp.zip
* http://www.suche-postleitzahl.org/downloads?download_file=plz-3stellig.shp.zip
* http://www.suche-postleitzahl.org/downloads
a CSV-file for all existing German post codes with matching geocoordinates has been compiled. The latitude and longitude coordinates were generated by running a PostgreSQL + PostGIS database. Additionally the respective TSO has been added to each post code. *(A Link to the SQL script will follow here later)*
*(License: http://www.suche-postleitzahl.org/downloads, Open Database Licence for free use. Source of data: © OpenStreetMap contributors)*
```
# Read generated postcode/location file
postcode = pd.read_csv('input/de_tso_postcode_gps.csv',
sep=';',
header=0)
# Drop possible duplicates in postcodes
postcode.drop_duplicates('postcode', keep='last',inplace=True)
# Show first entries
postcode.head()
```
** Merge geometry information by using the postcode**
```
# Take postcode and longitude/latitude informations
postcode = postcode[[0,3,4]]
DE_renewables = DE_renewables.merge(postcode, on=['postcode'], how='left')
```
#### Transform geoinformation
*(for data with already existing geoinformation)*
In this section the existing geoinformation (in UTM-format) will be transformed into latidude and longitude coordiates as a uniform standard for geoinformation.
The BNetzA data set offers UTM Geoinformation with the columns *utm_zone (UTM-Zonenwert)*, *utm_east* and *utm_north*. Most of utm_east-values include the utm_zone-value **32** at the beginning of the number. In order to properly standardize and transform this geoinformation into latitude and longitude it is necessary to remove this utm_zone value. For all UTM entries the utm_zone 32 is used by the BNetzA.
|utm_zone| utm_east| utm_north| comment|
|---|---|---| ----|
|32| 413151.72| 6027467.73| proper coordinates|
|32| **32**912159.6008| 5692423.9664| caused error by 32|
**How many different utm_zone values are in the data set?**
```
DE_renewables.groupby(['utm_zone'])['utm_zone'].count()
```
**Remove the utm_zone "32" from the utm_east value**
```
# Find entries with 32 value at the beginning
ix_32 = (DE_renewables['utm_east'].astype(str).str[:2] == '32')
ix_notnull = DE_renewables['utm_east'].notnull()
# Remove 32 from utm_east entries
DE_renewables.loc[ix_32,'utm_east'] = DE_renewables.loc[ix_32,'utm_east'].astype(str).str[2:].astype(float)
```
**Conversion UTM to lat/lon**
```
# Convert from UTM values to latitude and longitude coordinates
try:
DE_renewables['lonlat'] = DE_renewables.loc[ix_notnull, ['utm_east', 'utm_north', 'utm_zone']].apply(
lambda x: utm.to_latlon(x[0], x[1], x[2], 'U'),
axis=1) \
.astype(str)
except:
DE_renewables['lonlat'] = np.NaN
lat = []
lon = []
for row in DE_renewables['lonlat']:
try:
# Split tuple format into the column lat and lon
row = row.lstrip('(').rstrip(')')
lat.append(row.split(',')[0])
lon.append(row.split(',')[1])
except:
# set NaN
lat.append(np.NaN)
lon.append(np.NaN)
DE_renewables['latitude'] = pd.to_numeric(lat)
DE_renewables['longitude'] = pd.to_numeric(lon)
# Add new values to DataFrame lon and lat
DE_renewables['lat'] = DE_renewables[['lat', 'latitude']].apply(
lambda x: x[1] if pd.isnull(x[0]) else x[0],
axis=1)
DE_renewables['lon'] = DE_renewables[['lon', 'longitude']].apply(
lambda x: x[1] if pd.isnull(x[0]) else x[0],
axis=1)
```
**Check: missing coordinates by data source and type**
```
print('Missing Coordinates ', DE_renewables.lat.isnull().sum())
DE_renewables[DE_renewables.lat.isnull()].groupby(['energy_source',
'data_source']
)['data_source'].count()
```
**Remove temporary columns**
```
# drop lonlat column that contains both, latitute and longitude
DE_renewables.drop(['lonlat','longitude','latitude'], axis=1, inplace=True)
```
### 3.1.8 Save
The merged, translated, cleaned, DataFrame will be saved temporily as a pickle file, which stores a Python object fast.
```
DE_renewables.to_pickle('DE_renewables.pickle')
```
## 3.2 Denmark DK
### 3.2.1 Download and read
The data which will be processed below is provided by the following data sources:
** [Energistyrelsen (ens) / Danish Energy Agency](http://www.ens.dk/info/tal-kort/statistik-noegletal/oversigt-energisektoren/stamdataregister-vindmoller)** - The wind turbines register is released by the Danish Energy Agency.
** [Energinet.dk](http://www.energinet.dk/DA/El/Engrosmarked/Udtraek-af-markedsdata/Sider/Statistik.aspx)** - The data of solar power plants are released by the leading transmission network operator Denmark.
```
# point URLs to original data depending on the chosen download option
if download_from == 'original_sources':
url_DK_ens = 'https://ens.dk/sites/ens.dk/files/Statistik/anlaegprodtilnettet_0.xls'
url_DK_energinet = 'http://www.energinet.dk/SiteCollectionDocuments/Danske%20dokumenter/El/SolcelleGraf.xlsx'
url_DK_geo = 'http://download.geonames.org/export/zip/DK.zip'
elif download_from == 'opsd_server':
url_DK_ens = (url_opsd + version + folder + '/DK/anlaegprodtilnettet.xls')
url_DK_energinet = (url_opsd + version + folder + '/DK/SolcelleGraf.xlsx')
url_DK_geo = (url_opsd + version + folder + 'DK/DK.zip')
# Get wind turbines data
DK_wind_df = pd.read_excel(download_and_cache(url_DK_ens),
sheetname='IkkeAfmeldte-Existing turbines',
thousands='.',
header=17,
skipfooter=3,
parse_cols=16,
converters={'Møllenummer (GSRN)': str,
'Kommune-nr': str,
'Postnr': str}
)
# Get photovoltaic data
DK_solar_df = pd.read_excel(download_and_cache(url_DK_energinet),
sheetname='Data',
converters={'Postnr': str}
)
DK_wind_df.head(2)
DK_solar_df.head(2)
```
### 3.2.2 Translate column names
```
# Choose the translation terms for Denmark, create dictionary and show dictionary
idx_DK = columnnames[columnnames['country'] == 'DK'].index
column_dict_DK = columnnames.loc[idx_DK].set_index('original_name')['opsd_name'].to_dict()
column_dict_DK
# Translate columns by list
DK_wind_df.rename(columns = column_dict_DK, inplace=True)
DK_solar_df.rename(columns = column_dict_DK, inplace=True)
```
### 3.2.3 Add data source and missing information
```
# Add names of the data sources to the DataFrames
DK_wind_df['data_source'] = 'Energistyrelsen'
DK_solar_df['data_source'] = 'Energinet.dk'
# Add energy_source for each of the two DataFrames
DK_wind_df['energy_source'] = 'Wind'
DK_solar_df['energy_source'] = 'Solar'
DK_solar_df['energy_source_subtype'] = 'Photovoltaics'
```
### 3.2.4 Translate values and harmonize energy source
```
idx_DK = valuenames[valuenames['country'] == 'DK'].index
value_dict_DK = valuenames.loc[idx_DK].set_index('original_name')['opsd_name'].to_dict()
value_dict_DK
DK_wind_df.replace(value_dict_DK, inplace=True)
```
### 3.2.5 Georeferencing
**UTM32 to lat/lon** *(Data from Energistyrelsen)*
The Energistyrelsen data set offers UTM Geoinformation with the columns utm_east and utm_north belonging to the UTM zone 32. In this section the existing geoinformation (in UTM-format) will be transformed into latidude and longitude coordiates as a uniform standard for geoinformation.
```
# Index for all values with utm information
idx_notnull= DK_wind_df['utm_east'].notnull()
# Convert from UTM values to latitude and longitude coordinates
DK_wind_df['lonlat'] = DK_wind_df.loc[idx_notnull,['utm_east','utm_north']
].apply(lambda x: utm.to_latlon(x[0],
x[1],32,'U'), axis=1).astype(str)
# Split latitude and longitude in two columns
lat = []
lon = []
for row in DK_wind_df['lonlat']:
try:
# Split tuple format
# into the column lat and lon
row = row.lstrip('(').rstrip(')')
lat.append(row.split(',')[0])
lon.append(row.split(',')[1])
except:
# set NAN
lat.append(np.NaN)
lon.append(np.NaN)
DK_wind_df['lat'] = pd.to_numeric(lat)
DK_wind_df['lon'] = pd.to_numeric(lon)
# drop lonlat column that contains both, latitute and longitude
DK_wind_df.drop('lonlat', axis=1, inplace=True)
```
**Postcode to lat/lon (WGS84)**
*(for data from Energinet.dk)*
The available post code in the original data provides an approximation for the geocoordinates of the solar power plants.<br>
The postcode will be assigned to latitude and longitude coordinates with the help of the postcode table.
** [geonames.org](http://download.geonames.org/export/zip/?C=N;O=D)** The postcode data from Denmark is provided by Geonames and licensed under a [Creative Commons Attribution 3.0 license](http://creativecommons.org/licenses/by/3.0/).
```
# Get geo-information
zip_DK_geo = zipfile.ZipFile(download_and_cache(url_DK_geo))
# Read generated postcode/location file
DK_geo = pd.read_csv(zip_DK_geo.open('DK.txt'), sep='\t', header=-1)
# add column names as defined in associated readme file
DK_geo.columns = ['country_code','postcode','place_name','admin_name1',
'admin_code1','admin_name2','admin_code2','admin_name3',
'admin_code3','lat','lon','accuracy']
# Drop rows of possible duplicate postal_code
DK_geo.drop_duplicates('postcode', keep='last',inplace=True)
DK_geo['postcode'] = DK_geo['postcode'].astype(str)
# Add longitude/latitude infomation assigned by postcode (for Energinet.dk data)
DK_solar_df = DK_solar_df.merge(DK_geo[['postcode','lon','lat']],
on=['postcode'],
how='left')
print('Missing Coordinates DK_wind ',DK_wind_df.lat.isnull().sum())
print('Missing Coordinates DK_solar ',DK_solar_df.lat.isnull().sum())
```
### 3.2.6 Merge DataFrames and choose columns
```
dataframes = [DK_wind_df, DK_solar_df]
DK_renewables = pd.concat(dataframes)
DK_renewables = DK_renewables.reset_index()
# Only these columns will be kept for the renewable power plant list output
column_interest = ['commissioning_date', 'energy_source','energy_source_subtype',
'electrical_capacity_kW', 'dso','gsrn_id', 'postcode',
'municipality_code','municipality','address', 'address_number',
'utm_east', 'utm_north', 'lon','lat','hub_height',
'rotor_diameter', 'manufacturer', 'model', 'data_source']
# Clean DataFrame from columns other than specified above
DK_renewables = DK_renewables.loc[:, column_interest]
DK_renewables.reset_index(drop=True, inplace=True)
```
### 3.2.7 Transform electrical_capacity from kW to MW
```
# kW to MW
DK_renewables['electrical_capacity_kW'] /= 1000
# adapt column name
DK_renewables.rename(columns={'electrical_capacity_kW': 'electrical_capacity'},
inplace=True)
DK_renewables.head(2)
```
### 3.2.8 Save
```
DK_renewables.to_pickle('DK_renewables.pickle')
```
## 3.3 France FR
### 3.3.1 Download and read
The data which will be processed below is provided by the following data source:
** [Ministery of the Environment, Energy and the Sea](http://www.statistiques.developpement-durable.gouv.fr/energie-climat/r/energies-renouvelables.html?tx_ttnews%5Btt_news%5D=24638&cHash=d237bf9985fdca39d7d8c5dc84fb95f9)** - Number of installations and installed capacity of the different renewable source for every municipality in France. Service of observation and statistics, survey, date of last update: 15/12/2015. Data until 31/12/2014.
```
# point URLs to original data depending on the chosen download option
if download_from == 'original_sources':
url_FR_gouv = "http://www.statistiques.developpement-durable.gouv.fr/fileadmin/documents/Themes/Energies_et_climat/Les_differentes_energies/Energies_renouvelables/donnees_locales/2014/electricite-renouvelable-par-commune-2014.xls"
url_FR_geo = 'http://public.opendatasoft.com/explore/dataset/code-postal-code-insee-2015/download/?format=csv&timezone=Europe/Berlin&use_labels_for_header=true'
else:
url_FR_gouv = (url_opsd + version + folder + '/FR/electricite-renouvelable-par-commune-2014.xls')
url_FR_geo = (url_opsd + version + folder + 'FR/code-postal-code-insee-2015.csv')
# Get data of renewables per municipality
FR_re_df = pd.read_excel(download_and_cache(url_FR_gouv),
sheetname='Commune',
encoding = 'UTF8',
thousands='.',
decimals=',',
header=[2, 3],
skipfooter=9, # contains summarized values
index_col=[0, 1], # required for MultiIndex
converters={'Code officiel géographique':str})
```
### 3.3.2 Rearrange columns and translate column names
The French data source contains number of installations and sum of installed capacity per energy source per municipality. The structure is adapted to the power plant list of other countries. The list is limited to the plants which are covered by article 10 of february 2000 by an agreement to a purchase commitment.
```
# Rearrange data
FR_re_df.index.rename(['insee_com', 'municipality'], inplace=True)
FR_re_df.columns.rename(['energy_source', None], inplace=True)
FR_re_df = (FR_re_df
.stack(level='energy_source', dropna=False)
.reset_index(drop = False))
# Choose the translation terms for France, create dictionary and show dictionary
idx_FR = columnnames[columnnames['country'] == 'FR'].index
column_dict_FR = columnnames.loc[idx_FR].set_index('original_name')['opsd_name'].to_dict()
column_dict_FR
# Translate columnnames
FR_re_df.rename(columns = column_dict_FR, inplace=True)
# Drop all rows that just contain NA
FR_re_df = FR_re_df.dropna()
FR_re_df.head()
```
### 3.3.3 Add data source
```
FR_re_df['data_source'] = 'gouv.fr'
FR_re_df.info()
```
### 3.3.4 Translate values and harmonize energy source
** Kept secret if number of installations < 3**
If the number of installations is less than 3, it is marked with an _s_ instead of the number 1 or 2 due to statistical confidentiality ([further explanation by the data provider](http://www.statistiques.developpement-durable.gouv.fr/fileadmin/documents/Themes/Energies_et_climat/Les_differentes_energies/Energies_renouvelables/donnees_locales/2014/methodo-donnees-locales-electricte-renouvelable-12-2015-b.pdf)). Here, the _s_ is changed to _< 3_. This is done in the same step as the other value translations of the energy sources.
```
idx_FR = valuenames[valuenames['country'] == 'FR'].index
value_dict_FR = valuenames.loc[idx_FR].set_index('original_name')['opsd_name'].to_dict()
value_dict_FR
FR_re_df.replace(value_dict_FR, inplace=True)
```
**Separate and assign energy source and subtypes**
```
# Create dictionnary in order to assign energy_source to its subtype
energy_source_dict_FR = valuenames.loc[idx_FR].set_index('opsd_name')['energy_source'].to_dict()
# Column energy_source partly contains subtype information, thus this column is copied
# to new column for energy_source_subtype...
FR_re_df['energy_source_subtype'] = FR_re_df['energy_source']
# ...and the energy source subtype values in the energy_source column are replaced by
# the higher level classification
FR_re_df['energy_source'].replace(energy_source_dict_FR, inplace=True)
FR_re_df.reset_index(drop=True, inplace=True)
```
### 3.3.5 Georeferencing
#### Municipality (INSEE) code to lon/lat
The available INSEE code in the original data provides a first approximation for the geocoordinates of the renewable power plants. The following data source is utilized for assigning INSEE code to coordinates of the municipalities:
** [OpenDataSoft](http://public.opendatasoft.com/explore/dataset/code-postal-code-insee-2015/information/)** publishes a list of French INSEE codes and corresponding coordinates is published under the [Licence Ouverte (Etalab)](https://www.etalab.gouv.fr/licence-ouverte-open-licence).
```
# Downlad French geo-information. As download_and_cache_function is not working
# properly yet, thus other way of downloading
filename = 'code-postal-insee-2015.csv'
filepath = "input/original_data/" + filename
if not os.path.exists(filepath):
print("Downloading file: ", filename)
FR_geo_csv = urllib.request.urlretrieve(url_FR_geo, filepath)
else:
print("Using local file from", filepath)
# Read INSEE Code Data
FR_geo = pd.read_csv('input/original_data/code-postal-insee-2015.csv',
sep=';',
header=0,
converters={'Code_postal':str})
# Drop possible duplicates of the same INSEE code
FR_geo.drop_duplicates('INSEE_COM', keep='last',inplace=True)
# create columns for latitude/longitude
lat = []
lon = []
# split in latitude/longitude
for row in FR_geo['Geo Point']:
try:
# Split tuple format
# into the column lat and lon
row = row.lstrip('(').rstrip(')')
lat.append(row.split(',')[0])
lon.append(row.split(',')[1])
except:
# set NAN
lat.append(np.NaN)
lon.append(np.NaN)
# add these columns to the INSEE DataFrame
FR_geo['lat'] = pd.to_numeric(lat)
FR_geo['lon'] = pd.to_numeric(lon)
# Column names of merge key have to be named identically
FR_re_df.rename(columns={'municipality_code': 'INSEE_COM'}, inplace=True)
# Merge longitude and latitude columns by the Code INSEE
FR_re_df = FR_re_df.merge(FR_geo[['INSEE_COM','lat','lon']],
on=['INSEE_COM'],
how='left')
# Translate Code INSEE column back to municipality_code
FR_re_df.rename(columns={'INSEE_COM': 'municipality_code'}, inplace=True)
FR_re_df.head(2)
```
### 3.3.6 Save
```
FR_re_df.to_pickle('FR_renewables.pickle')
```
## 3.4 Poland PL
### 3.4.1 Download and read
The data which will be processed below is provided by the following data source:
** [Urzad Regulacji Energetyki (URE) / Energy Regulatory Office](http://www.ure.gov.pl/uremapoze/mapa.html)** - Number of installations and installed capacity per energy source of renewable energy. Summed per powiat (districts) .
#### The Polish data has to be downloaded manually
if you have not chosen download_from = opsd_server.
- Go to http://www.ure.gov.pl/uremapoze/mapa.html
- Click on the British flag in the lower right corner for Englisch version
- Set detail to highest (to the right) in the upper right corner
- Click on the printer symbol in the lower left corner
- 'Generate', then the rtf-file simple.rtf will be downloaded
- Put it in the folder input/original_data on your computer
- If the download is temporarily not working choose download_from == 'opsd_server'
```
if download_from == 'opsd_server':
url_PL_ure = (url_opsd + version + folder + '/PL/simple.rtf')
download_and_cache(url_PL_ure)
# read rtf-file to string with the correct encoding
with open('input/original_data/simple.rtf', 'r') as rtf:
file_content = rtf.read()
file_content = file_content.encode('utf-8').decode('iso-8859-2')
```
### 3.4.2 Rearrange data from rft-file
The rtf file has one table for each district in the rtf-file which needs to be separated from each and other and restructured to get all plants in one DataFrame with the information: district, energy_source, number_of_installations, installed_capacity. Thus in the following, the separating items are defined, the district tables split in parts, all put in one list and afterwards transferred to a pandas DataFrame.
```
# a new line is separating all parts
sep_split_into_parts = r'{\fs12 \f1 \line }'
# separates the table rows of each table
sep_data_parts = r'\trql'
reg_exp_district = r'(?<=Powiat:).*(?=})'
reg_exp_installation_type = (
r'(?<=\\fs12 \\f1 \\pard \\intbl \\ql \\cbpat[2|3|4] \{\\fs12 \\f1 ).*(?=\})')
reg_exp_installation_value = (
r'(?<=\\fs12 \\f1 \\pard \\intbl \\qr \\cbpat[3|4] \{\\fs12 \\f1 ).*(?=})')
# split file into parts
parts = file_content.split(sep_split_into_parts)
# list containing the data
data_set = []
for part in parts:
# match district
district = re.findall(reg_exp_district, part)
if len(district) == 0:
pass
else:
district = district[0].lstrip()
# separate each part
data_parts = part.split(sep_data_parts)
# data structure: data_row = {'district': '', 'install_type': '', 'quantity': '', 'power': ''}
for data_rows in data_parts:
wrapper_list = []
# match each installation type
installation_type = re.findall(reg_exp_installation_type, data_rows)
for inst_type in installation_type:
wrapper_list.append({'district': district, 'energy_source_subtype': inst_type})
# match data - contains twice as many entries as installation type (quantity, power vs. install type)
data_values = re.findall(reg_exp_installation_value, data_rows)
if len(data_values) == 0:
#log.debug('data values empty')
pass
else:
# connect data
for i, _ in enumerate(wrapper_list):
wrapper_list[i]['number_of_installations'] = data_values[(i * 2)]
wrapper_list[i]['electrical_capacity'] = data_values[(i * 2) + 1]
# prepare to write to file
for data in wrapper_list:
data_set.append(data)
# mapping of malformed unicode which appear in the Polish district names
polish_truncated_unicode_map = {
r'\uc0\u322': 'ł',
r'\uc0\u380': 'ż',
r'\uc0\u243': 'ó',
r'\uc0\u347': 'ś',
r'\uc0\u324': 'ń',
r'\uc0\u261': 'ą',
r'\uc0\u281': 'ę',
r'\uc0\u263': 'ć',
r'\uc0\u321': 'Ł',
r'\uc0\u378': 'ź',
r'\uc0\u346': 'Ś',
r'\uc0\u379': 'Ż'
}
# changing malformed unicode
for entry in data_set:
while r'\u' in entry['district']:
index = entry['district'].index(r'\u')
offset = index + 9
to_be_replaced = entry['district'][index:offset]
if to_be_replaced in polish_truncated_unicode_map.keys():
# offset + 1 because there is a trailing whitespace
entry['district'] = entry['district'].replace(entry['district'][index:offset + 1],
polish_truncated_unicode_map[to_be_replaced])
else:
break
# Create pandas DataFrame with similar structure as the other countries
PL_re_df = pd.DataFrame(data_set)
```
### 3.4.3 Add data source
```
PL_re_df['data_source'] = 'Urzad Regulacji Energetyki'
```
### 3.4.4 Translate values and harmonize energy source
```
idx_PL = valuenames[valuenames['country'] == 'PL'].index
value_dict_PL = valuenames.loc[idx_PL].set_index('original_name')['opsd_name'].to_dict()
value_dict_PL
PL_re_df.head()
# Replace install_type descriptions with energy_source subtype
PL_re_df.energy_source_subtype.replace(value_dict_PL, inplace=True)
```
**Assign energy_source_subtype to energy_source**
```
# Create dictionnary in order to assign energy_source to its subtype
energy_source_dict_PL = valuenames.loc[idx_PL].set_index('opsd_name')['energy_source'].to_dict()
# Create new column for energy_source
PL_re_df['energy_source'] = PL_re_df.energy_source_subtype
# Fill this with the energy source instead of subtype information
PL_re_df.energy_source.replace(energy_source_dict_PL, inplace=True)
energy_source_dict_PL
```
** Adjust datatype of numeric columns**
```
# change type to numeric
PL_re_df['electrical_capacity'] = pd.to_numeric(PL_re_df['electrical_capacity'])
# Additionally commas are deleted
PL_re_df['number_of_installations'] = pd.to_numeric(
PL_re_df['number_of_installations'].str.replace(',',''))
```
**Aggregate**
For entries/rows of the same district and energy_source_subtype, electrical capacity and number of installations are aggregaated.
```
PL_re_df = PL_re_df.groupby(['district','energy_source','energy_source_subtype'],
as_index = False
).agg({'electrical_capacity': sum,
'number_of_installations': sum,
'data_source': 'first'})
```
### 3.4.5 Georeferencing - _work in progress_
```
# ToDo: GeoReferencing
# to get GEOINFO
# NTS 4 - powiats and cities with powiat status (314 + 66 units)
# http://stat.gov.pl/en/regional-statistics/nomenclature-nts-161/
# http://forum.geonames.org/gforum/posts/list/795.page
```
### 3.4.6 Save
```
PL_re_df.to_pickle('PL_renewables.pickle')
```
Check and validation of the renewable power plants list as well as the creation of CSV/XLSX/SQLite files can be found in Part 2 of this script. It also generates a daily time series of cumulated installed capacities by energy source.
| github_jupyter |
```
import numpy as np
import pandas as pd
import unidecode
from nltk.corpus import stopwords
from nltk.stem.snowball import SnowballStemmer
from nltk.tokenize import RegexpTokenizer
import nltk
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.model_selection import train_test_split
from sklearn.dummy import DummyClassifier
from sklearn.metrics import mean_absolute_error
from sklearn.ensemble import GradientBoostingRegressor
from nlp_tweets.preprocess import clean_text
from nlp_tweets.eda import get_categorial_confusion_matrix, plot_roc_curve_mean
PATH_RAW = '../data/raw/'
PATH_PROCESSED = '../data/processed/'
TRAIN_COLUMNS = ['id', 'text', 'target']
TEST_COLUMNS = ['id', 'text']
```
# Loading
```
tweets_df = pd.read_csv(PATH_RAW + 'train.csv')
tweets_df = tweets_df[TRAIN_COLUMNS]
tweets_df.head()
tweets_test_df = pd.read_csv(PATH_RAW + 'test.csv')
tweets_test_df = tweets_test_df[TEST_COLUMNS]
```
# Preprocessing
```
to_remove_caracters = ['/', '-', '[', ']', ',', ':', '&', '>', '<', '\\', '#', '.', '?', '!', "'"]
stops = list(stopwords.words("english")) + list(stopwords.words("french"))
stops = [unidecode.unidecode(stop_words) for stop_words in stops]
stop_words = ['', 'may', 'sent', ]
stop_words += stops
tweets_df['text'] = tweets_df['text'].apply(clean_text(to_remove_caracters,
stop_words))
tweets_test_df['text'] = tweets_test_df['text'].apply(clean_text(to_remove_caracters,
stop_words))
stemmer = SnowballStemmer('english')
tokenizer = RegexpTokenizer(r'[a-zA-Z\']+')
def tokenize(text):
words = [stemmer.stem(word) for word in tokenizer.tokenize(text.lower())]
return words
vectorizer = TfidfVectorizer(tokenizer = tokenize, max_features = 1000)
X = vectorizer.fit_transform(tweets_df['text'].values)
X_to_pred = vectorizer.transform(tweets_test_df['text'].values)
words = vectorizer.get_feature_names()
```
# Train test split
```
X_train, X_test, y_train, y_test = train_test_split(X, tweets_df['target'],
stratify=tweets_df['target'], test_size=0.2)
print(X_train.shape, X_test.shape)
```
# Create Model
## Baseline
```
baseline_model = DummyClassifier(strategy="most_frequent")
```
### Train
```
baseline_model.fit(X_train, y_train)
```
### Test
```
baseline_pre_on_test = baseline_model.predict(X_test)
baseline_mae = mean_absolute_error(y_test, baseline_pre_on_test)
print(f'The mae for the baseline on test set is {baseline_mae}')
get_categorial_confusion_matrix(y_test, baseline_pre_on_test, display=True)
```
# Random forest
```
xgb_model = GradientBoostingRegressor(n_estimators=500)
```
### Train
```
xgb_model.fit(X_train, y_train)
```
### Finding the best threshold
```
pred_proba_train = xgb_model.predict(X_train)
pred_proba_test = xgb_model.predict(X_test)
threshold = plot_roc_curve_mean(y_train=y_train.values, y_train_pred_proba=pred_proba_train,
y_test=y_test.values, y_test_pred_proba=pred_proba_test)
```
### Test
```
model_pred_on_test = xgb_model.predict(X_test)
model_pred_on_test = (model_pred_on_test > threshold).astype(int)
xgb_mae = mean_absolute_error(y_test, model_pred_on_test)
print(f'The mae for the baseline on test set is {xgb_mae}')
get_categorial_confusion_matrix(y_test, model_pred_on_test, display=True)
```
# Predict
```
predictions = xgb_model.predict(X_to_pred)
predictions = (predictions > threshold).astype(int)
output = pd.read_csv(PATH_RAW + 'sample_submission.csv')
output['target'] = predictions
output.to_csv(PATH_PROCESSED + 'to_submit.csv', index=False)
```
| github_jupyter |
```
theBoard = {'7': ' ' , '8': ' ' , '9': ' ' ,
'4': ' ' , '5': ' ' , '6': ' ' ,
'1': ' ' , '2': ' ' , '3': ' ' }
board_keys = []
for key in theBoard:
board_keys.append(key)
''' We will have to print the updated board after every move in the game and
thus we will make a function in which we'll define the printBoard function
so that we can easily print the board everytime by calling this function. '''
def printBoard(board):
print(board['7'] + '|' + board['8'] + '|' + board['9'])
print('-+-+-')
print(board['4'] + '|' + board['5'] + '|' + board['6'])
print('-+-+-')
print(board['1'] + '|' + board['2'] + '|' + board['3'])
# Now we'll write the main function which has all the gameplay functionality.
def game():
possibleMoves=[1,2,3,4,5,6,7,8,9]
turn = 'X'
count = 0
for i in range(10):
printBoard(theBoard)
print("It's your turn," + turn + " Move to which place?")
move = input()
while int(move) not in possibleMoves:
print("That place is not one of possible moves:", possibleMoves,"\n move to which place?")
move = input()
else:
if theBoard[move] == ' ':
theBoard[move] = turn
possibleMoves.remove(int(move))
count += 1
else:
print("That place is already filled.\nMove to which place?")
continue
# Now we will check if player X or O has won,for every move after 5 moves.
if count >= 5:
if theBoard['7'] == theBoard['8'] == theBoard['9'] != ' ': # across the top
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['4'] == theBoard['5'] == theBoard['6'] != ' ': # across the middle
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['1'] == theBoard['2'] == theBoard['3'] != ' ': # across the bottom
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['1'] == theBoard['4'] == theBoard['7'] != ' ': # down the left side
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['2'] == theBoard['5'] == theBoard['8'] != ' ': # down the middle
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['3'] == theBoard['6'] == theBoard['9'] != ' ': # down the right side
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['7'] == theBoard['5'] == theBoard['3'] != ' ': # diagonal
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
elif theBoard['1'] == theBoard['5'] == theBoard['9'] != ' ': # diagonal
printBoard(theBoard)
print("\nGame Over.\n")
print(" **** " +turn + " won. ****")
break
# If neither X nor O wins and the board is full, we'll declare the result as 'tie'.
if count == 9:
print("\nGame Over.\n")
print("It's a Tie!!")
# Now we have to change the player after every move.
if turn =='X':
turn = 'O'
else:
turn = 'X'
# Now we will ask if player wants to restart the game or not.
restart = input("Do want to play Again?(y/n)")
if restart == "y" or restart == "Y":
for key in board_keys:
theBoard[key] = " "
game()
if __name__ == "__main__":
game()
```
| github_jupyter |
##Mounting Drive
```
from google.colab import drive
drive.mount('/content/drive')
```
## Installing and Importing required Libraries
```
!pip install -q transformers
import numpy as np
import pandas as pd
from sklearn import metrics
import transformers
import torch
from torch.utils.data import Dataset, DataLoader, RandomSampler, SequentialSampler
from transformers import RobertaTokenizer, RobertaModel, RobertaConfig
from tqdm.notebook import tqdm
from transformers import get_linear_schedule_with_warmup
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
n_gpu = torch.cuda.device_count()
torch.cuda.get_device_name(0)
```
## Data Preprocessing
```
df = pd.read_csv("/content/drive/My Drive/bert-multilabel/train.csv")
df['list'] = df[df.columns[3:]].values.tolist()
new_df = df[['TITLE', 'ABSTRACT', 'list']].copy()
new_df.head()
```
## Model Configurations
```
# Defining some key variables that will be used later on in the training
MAX_LEN = 512
TRAIN_BATCH_SIZE = 8
VALID_BATCH_SIZE = 4
EPOCHS = 3
LEARNING_RATE = 1e-05
tokenizer = RobertaTokenizer.from_pretrained('roberta-base')
```
## Creating Custom Dataset class
```
class CustomDataset(Dataset):
def __init__(self, dataframe, tokenizer, max_len):
self.tokenizer = tokenizer
self.data = dataframe
self.abstract = dataframe.ABSTRACT
self.title = dataframe.TITLE
self.targets = self.data.list
self.max_len = max_len
self.max_len_title = 200
def __len__(self):
return len(self.abstract)
def __getitem__(self, index):
abstract = str(self.abstract[index])
title = str(self.title[index])
abstract = " ".join(abstract.split())
title = " ".join(abstract.split())
inputs_abstract = self.tokenizer.encode_plus(
abstract,
None,
add_special_tokens = True,
max_length = self.max_len,
pad_to_max_length = True,
truncation = True
)
inputs_title = self.tokenizer.encode_plus(
title,
None,
add_special_tokens = True,
max_length = self.max_len_title,
pad_to_max_length = True,
truncation = True
)
ids_abstract = inputs_abstract['input_ids']
mask_abstract = inputs_abstract['attention_mask']
ids_title = inputs_title['input_ids']
mask_title = inputs_title['attention_mask']
return{
'ids_abstract': torch.tensor(ids_abstract, dtype=torch.long),
'mask_abstract': torch.tensor(mask_abstract, dtype=torch.long),
'ids_title': torch.tensor(ids_title, dtype=torch.long),
'mask_title': torch.tensor(mask_title, dtype=torch.long),
'targets': torch.tensor(self.targets[index], dtype=torch.float)
}
train_size = 0.8
train_dataset=new_df.sample(frac=train_size,random_state=200)
test_dataset=new_df.drop(train_dataset.index).reset_index(drop=True)
train_dataset = train_dataset.reset_index(drop=True)
print("FULL Dataset: {}".format(new_df.shape))
print("TRAIN Dataset: {}".format(train_dataset.shape))
print("TEST Dataset: {}".format(test_dataset.shape))
training_set = CustomDataset(train_dataset, tokenizer, MAX_LEN)
testing_set = CustomDataset(test_dataset, tokenizer, MAX_LEN)
train_params = {'batch_size': TRAIN_BATCH_SIZE,
'shuffle': True,
'num_workers': 0
}
test_params = {'batch_size': VALID_BATCH_SIZE,
'shuffle': True,
'num_workers': 0
}
training_loader = DataLoader(training_set, **train_params)
testing_loader = DataLoader(testing_set, **test_params)
```
## Roberta Model
```
# Creating the customized model, by adding a drop out and a dense layer on top of roberta to get the final output for the model.
class RobertaMultiheadClass(torch.nn.Module):
def __init__(self):
super(RobertaMultiheadClass, self).__init__()
self.roberta = transformers.RobertaModel.from_pretrained('roberta-base')
self.drop = torch.nn.Dropout(0.3)
self.linear_1 = torch.nn.Linear(1536, 768)
self.linear_2 = torch.nn.Linear(768, 6)
def forward(self, ids_1, mask_1, ids_2, mask_2):
_, output_1= self.roberta(ids_1, attention_mask = mask_1)
_, output_2= self.roberta(ids_2, attention_mask = mask_2)
output = torch.cat((output_1, output_2), dim = 1)
output = self.drop(output)
output = self.linear_1(output)
output = self.drop(output)
output = self.linear_2(output)
return output
model = RobertaMultiheadClass()
model.to(device)
```
## Hyperparameters & Loss function
```
def loss_fn(outputs, targets):
return torch.nn.BCEWithLogitsLoss()(outputs, targets)
param_optimizer = list(model.named_parameters())
no_decay = ["bias", "LayerNorm.bias", "LayerNorm.weight"]
optimizer_parameters = [
{
"params": [
p for n, p in param_optimizer if not any(nd in n for nd in no_decay)
],
"weight_decay": 0.001,
},
{
"params": [
p for n, p in param_optimizer if any(nd in n for nd in no_decay)
],
"weight_decay": 0.0,
},
]
optimizer = torch.optim.AdamW(optimizer_parameters, lr=3e-5)
num_training_steps = int(len(train_dataset) / TRAIN_BATCH_SIZE * EPOCHS)
scheduler = get_linear_schedule_with_warmup(
optimizer,
num_warmup_steps = 0,
num_training_steps = num_training_steps
)
```
## Train & Eval Functions
```
def train(epoch):
model.train()
for _,data in tqdm(enumerate(training_loader, 0), total=len(training_loader)):
ids_1 = data['ids_abstract'].to(device, dtype = torch.long)
mask_1 = data['mask_abstract'].to(device, dtype = torch.long)
ids_2 = data['ids_title'].to(device, dtype = torch.long)
mask_2 = data['mask_title'].to(device, dtype = torch.long)
targets = data['targets'].to(device, dtype = torch.float)
outputs = model(ids_1, mask_1, ids_2, mask_2)
optimizer.zero_grad()
loss = loss_fn(outputs, targets)
if _%1000==0:
print(f'Epoch: {epoch}, Loss: {loss.item()}')
optimizer.zero_grad()
loss.backward()
optimizer.step()
scheduler.step()
def validation(epoch):
model.eval()
fin_targets=[]
fin_outputs=[]
with torch.no_grad():
for _, data in tqdm(enumerate(testing_loader, 0), total=len(testing_loader)):
ids_1 = data['ids_abstract'].to(device, dtype = torch.long)
mask_1 = data['mask_abstract'].to(device, dtype = torch.long)
ids_2 = data['ids_title'].to(device, dtype = torch.long)
mask_2 = data['mask_title'].to(device, dtype = torch.long)
targets = data['targets'].to(device, dtype = torch.float)
outputs = model(ids_1, mask_1, ids_2, mask_2)
fin_targets.extend(targets.cpu().detach().numpy().tolist())
fin_outputs.extend(torch.sigmoid(outputs).cpu().detach().numpy().tolist())
return fin_outputs, fin_targets
```
## Training Model
```
MODEL_PATH = "/content/drive/My Drive/roberta-multilabel/model.bin"
best_micro = 0
for epoch in range(EPOCHS):
train(epoch)
outputs, targets = validation(epoch)
outputs = np.array(outputs) >= 0.5
accuracy = metrics.accuracy_score(targets, outputs)
f1_score_micro = metrics.f1_score(targets, outputs, average='micro')
f1_score_macro = metrics.f1_score(targets, outputs, average='macro')
print(f"Accuracy Score = {accuracy}")
print(f"F1 Score (Micro) = {f1_score_micro}")
print(f"F1 Score (Macro) = {f1_score_macro}")
if f1_score_micro > best_micro:
torch.save(model.state_dict(), MODEL_PATH)
best_micro = f1_score_micro
```
## Predictions
```
PATH = "/content/drive/My Drive/roberta-multilabel/model.bin"
model = RobertaMultiheadClass()
model.load_state_dict(torch.load(PATH))
model.to(device)
model.eval()
def predict(id, abstract, title):
MAX_LENGTH = 512
inputs_abstract = tokenizer.encode_plus(
abstract,
None,
add_special_tokens=True,
max_length=512,
pad_to_max_length=True,
return_token_type_ids=True,
truncation = True
)
inputs_title = tokenizer.encode_plus(
title,
None,
add_special_tokens=True,
max_length=200,
pad_to_max_length=True,
return_token_type_ids=True,
truncation = True
)
ids_1 = inputs_abstract['input_ids']
mask_1 = inputs_abstract['attention_mask']
ids_1 = torch.tensor(ids_1, dtype=torch.long).unsqueeze(0)
mask_1 = torch.tensor(mask_1, dtype=torch.long).unsqueeze(0)
ids_2 = inputs_title['input_ids']
mask_2 = inputs_title['attention_mask']
ids_2 = torch.tensor(ids_2, dtype=torch.long).unsqueeze(0)
mask_2 = torch.tensor(mask_2, dtype=torch.long).unsqueeze(0)
ids_1 = ids_1.to(device)
mask_1 = mask_1.to(device)
ids_2 = ids_2.to(device)
mask_2 = mask_2.to(device)
with torch.no_grad():
outputs = model(ids_1, mask_1, ids_2, mask_2)
outputs = torch.sigmoid(outputs).squeeze()
outputs = np.round(outputs.cpu().numpy())
out = np.insert(outputs, 0, id)
return out
def submit():
test_df = pd.read_csv('/content/drive/My Drive/bert-multilabel/test.csv')
sample_submission = pd.read_csv('/content/drive/My Drive/bert-multilabel/sample_submission_UVKGLZE.csv')
y = []
for id, abstract, title in tqdm(zip(test_df['ID'], test_df['ABSTRACT'], test_df['TITLE']),
total=len(test_df)):
out = predict(id, abstract, title)
y.append(out)
y = np.array(y)
submission = pd.DataFrame(y, columns=sample_submission.columns).astype(int)
return submission
submission = submit()
submission
submission.to_csv("roberta_baseline.csv", index=False)
```
| github_jupyter |
```
a = 2
b = 5
if a > b:
print('an is greater')
else:
print('b is greater')
a = 2
b = 5
if a > b:
print('a is greater')
else:
print('b is greater')
print("Done")
a = 5
b = 2
if a>b:
print('a is grearter')
elif b>a:
print('b is greater')
else:
print('they both are equal')
a = 5
b = 2
if a>b:
print('a is greater')
print('Value of a is',a)
print('And value of b is',b)
elif b>a:
print('b is greater')
print('Value of a is',a)
print('And value of b is',b)
else:
print('they both are equal')
a = 10
b = 50
if a>b:
print('a is greater')
print('Value of a is',a)
print('And value of b is',b)
elif b>a:
print('b is greater')
print('Value of a is',a)
print('And value of b is',b)
else:
print('they are both equal')
a = 15
b = 15
if a>b:
print('a is greater')
print('Value of a is',a)
print('And value of b is',b)
elif b>a:
print('b is greater')
print('Value of a is',a)
print('And value of b is',b)
else:
print('they are both equal')
a = 5
b = 10
c = 20
if a>b and a>c:
print('a is the greatest')
elif b>a and b>c:
print('b is the greatest')
elif c>a and c>b:
print('c is the greatest')
else:
print('There is a tie somewhere')
a = 10
b = 20
if a!=b:
if a>b:
print("a is greater")
else:
print("b is greater")
else:
print("a and b are equal")
a = int(input("Please enter value for a:"))
b = int(input("Please enter value for b:"))
if a!=b:
if a>b:
print("a is greater")
else:
print("b is greater")
else:
print(" a and b are equal")
a = input("Enter a number: ")
b = input("Enter the second number: ")
if a.replace('.','',1).isdigit() and b.replace('.','',1).isdigit():
a = float(a)
b = float(b)
if a!=b:
if a > b:
print('a is greater')
else:
print('b is greater')
else:
print("a and b are equal")
else:
print("Intergers were not entered")
```
# Excersises
# 1.1
```
a = int(input("Enter a number: "))
if (a % 2) == 0:
print("is Even")
else:
print('is odd')
```
# 1.2
```
a = int(input("Enter a number: "))
if a!=0:
if a > 0:
print("a is positive")
else:
print("a is negative")
else:
print('a is equal to 0')
```
# 1.3
```
grade = int(input("Enter grade here: "))
if grade<25:
print('F')
elif grade>=25 and grade<=45:
print('E')
elif grade>=45 and grade<=60:
print('D')
elif grade>=60 and grade<=75:
print('C')
elif grade>=75 and grade<=90:
print('B')
elif grade>=90:
print('A')
a = input("Please enter an interger: ")
if a.isdigit():
a = int(a)
if a == 0:
print('a is zero')
elif a%2 ==0:
print('a is even')
else:
print('a is not even')
else:
print("Input was not an interger")
```
| github_jupyter |
# Kafka brokerの構築手順
```
set -o pipefail
```
## Ansibleの設定
ansibleをつかってKafka brokerクラスタ(とzookeeperクラスタ)を構築する。
ansibleのインベントリファイルを作成する。
```
cat >inventory.yml <<EOF
all:
children:
kafka:
hosts:
server1.example.jp:
#ここにKafkaをうごかすホストを羅列する。行末のコロンを忘れずに
vars:
ansible_user: piyo #実行ユーザは変更する
ansible_ssh_private_key_file: ~/.ssh/id_rsa
ansible_python_interpreter: /usr/bin/python3
zookeeper:
hosts:
server1.example.jp:
#ここにZookeeperをうごかすホストを羅列する。行末のコロンを忘れずに
vars:
ansible_user: piyo #実行ユーザは変更する
ansible_ssh_private_key_file: ~/.ssh/id_rsa
ansible_python_interpreter: /usr/bin/python3
EOF
```
ansibleの設定ファイルを作成する。
```
cat >ansible.cfg <<EOF
[defaults]
command_warnings = False
inventory = ./inventory.yml
EOF
```
ansibleを通じてzookeeperとkafkaを実行するホストにアクセスできるのを確認する。
```
ansible all -m ping
```
Dockerがインストールされているのを確認する。
```
ansible all -m command -a "docker version"
```
## ZooKeeperクラスタの構築
zookeeperのdockerイメージ名とポート番号を設定する。
PPORT,LPORT,CPORTはほかのサービスのポート番号とぶつかっているのでなければ変更する必要はない。
zookeeperの仕様によりCPORTは変更できない。
```
DOCKER_IMAGE="zookeeper"
ZK_PPORT=12888 # peer
ZK_LPORT=13888 # leader
ZK_CPORT=2181 # client
```
zookeeperを起動するスクリプトを生成する。
```
LIST_ZOOKEEPER_HOSTS="$(ansible-inventory --list | jq -r '.zookeeper.hosts|.[]')"
list_zookeeper_hosts() {
echo "$LIST_ZOOKEEPER_HOSTS"
}
print_servers() {
local MYID="$1"
local HOST
local ID=1
local SERVER
list_zookeeper_hosts | while read HOST; do
if [ "$ID" = "$MYID" ]; then
local ANYADDR="0.0.0.0"
HOST="$ANYADDR"
fi
printf "server.$ID=$HOST:$ZK_PPORT:$ZK_LPORT "
ID=$((ID + 1))
done
printf "\n"
}
print_docker_run() {
local DIR="$1"
local ID=1
list_zookeeper_hosts | while read HOST; do
#local NAME="sinetstream-zookeeper-$ID"
local NAME="sinetstream-zookeeper"
local SERVERS="$(print_servers "$ID")"
{
printf "docker run --rm --detach --name '$NAME' --env 'ZOO_MY_ID=$ID' --env 'ZOO_SERVERS=$SERVERS' --publish $ZK_PPORT:$ZK_PPORT --publish $ZK_LPORT:$ZK_LPORT --publish $ZK_CPORT:$ZK_CPORT $DOCKER_IMAGE"
} > "$DIR/zookeeper-docker_run-${HOST}.sh"
ID=$((ID + 1))
done
}
mkdir -p tmp &&
rm -f tmp/*.sh &&
print_docker_run tmp &&
ls -l tmp/*.sh
```
ansibleをつかってzookeeperサーバーを起動する。
```
ansible zookeeper -m script -a 'tmp/zookeeper-docker_run-{{inventory_hostname}}.sh'
ansible zookeeper -m command -a 'docker ps --filter "name=sinetstream-zookeeper"'
```
## Kafkaクラスタ
公式のKafka一式をダウンロードする。
手元でダウンロードしてから各ホストにコピーする。
```
KAFKA="kafka_2.12-2.4.1"
wget --mirror http://ftp.kddilabs.jp/infosystems/apache/kafka/2.4.1/$KAFKA.tgz
ansible kafka -m command -a "mkdir -p \$PWD/sinetstream-kafka"
ansible kafka -m copy -a "src=$KAFKA.tgz dest=\$PWD/sinetstream-kafka/"
```
### KafkaブローカーをうごかすCentOSのコンテナを作成
認証方法をどれかひとつ選択する。
```
#KAFKA_AUTH=SSL # SSL/TLS認証(クライアント認証); 通信にTLSをつかい、認証に証明書をつかう
KAFKA_AUTH=SASL_SSL_SCRAM # SCRAM認証/TLS; 通信にTLSをつかい、認証にSCRAM(パスワード)をつかう
#KAFKA_AUTH=SASL_SSL_PLAIN # パスワード認証/TLS; 通信にTLSをつかい、認証に平文パスワードをつかう
#KAFKA_AUTH=PLAINTEXT # 通信は暗号化されず、認証もない ※つかってはいけない
```
truststore/keystoreを保護するためのパスワードを設定する。パスワードは適当に強度の高い文字列を指定する。
```
TRUSTSTORE_PASSWORD="trust-pass-00"
KEYSTORE_PASSWORD="key-pass-00"
```
SCAM認証やパスワード認証を使う場合には、ユーザーのリストと各ユーザのパスワードを設定する。
SSL/TLS認証を使う場合はパスワードは設定しなくてよく、ユーザのリストだけを設定する。SSL/TLS認証でのユーザ名は証明書のCommon Nameである。
ユーザ `admin` はkafkaブローカ間の通信につかう特別なユーザなので消してはいけない。
パスワードは十分な強度をもったものに変更すべきである。
```
USER_LIST="user01 user02 user03 CN=client0,C=JP"
PASSWORD_admin="admin-pass"
PASSWORD_user01="user01-pass"
PASSWORD_user02="user02-pass"
PASSWORD_user03="user03-pass"
```
認可(ACL)の設定。
```
KAFKA_ACL_DEFAULT_TO_ALLOW="false" # trueに設定するとACLが設定されていないユーザはアクセスが許可される。
ACL_user01="readwrite"
ACL_user02="write"
ACL_user03="read"
ACL_CN_client0_C_JP="readwrite" # 英数字以外は _ に置き換えて
```
```
KAFKA_PORT_SSL=9093
KAFKA_PORT_SASL_SSL=9093
```
認証方法の詳細なパラメータを設定する。
```
SCRAM_MECHANISM="SCRAM-SHA-256"
```
Kafkaブローカを動かすコンテナを作る。
```
ansible kafka -m command -a "docker run \
--detach \
--interactive \
--net host \
--name sinetstream-kafka \
--volume \$PWD/sinetstream-kafka:/sinetstream-kafka \
centos:7"
ansible kafka -m command -a "docker exec sinetstream-kafka true"
```
コンテナにKafkaの実行に必要なソフトウェアをインストールする。
```
ansible kafka -m command -a "docker exec sinetstream-kafka yum update -y"
ansible kafka -m command -a "docker exec sinetstream-kafka yum install -y java-1.8.0-openjdk openssl"
ansible kafka -m command -a "docker exec sinetstream-kafka tar xf /sinetstream-kafka/$KAFKA.tgz" &&
ansible kafka -m command -a "docker exec sinetstream-kafka ln -s /$KAFKA /kafka"
```
### Kafkaブローカの設定
kafkaブローカの設定ファイルを生成する。
```
LIST_KAFKA_HOSTS="$(ansible-inventory --list | jq -r '.kafka.hosts|.[]')"
list_kafka_hosts() {
echo "$LIST_KAFKA_HOSTS"
}
print_server_properties() {
local HOST="$1"
local ID="$2"
echo "broker.id=${ID}"
local ZKHOST
printf "zookeeper.connect="
list_zookeeper_hosts | sed "s/\$/:${ZK_CPORT}/" | paste -s -d,
printf "listeners="
{
case "$KAFKA_AUTH" in
PLAINTEXT) echo "PLAINTEXT://:${KAFKA_PORT_PLAINTEXT}" ;;
SSL) echo "SSL://:${KAFKA_PORT_SSL}" ;;
SASL_SSL*) echo "SASL_SSL://:${KAFKA_PORT_SASL_SSL}"
echo "SSL://:$((KAFKA_PORT_SASL_SSL+1))" ;;
esac
} | paste -s -d,
printf "advertised.listeners="
{
case "$KAFKA_AUTH" in
PLAINTEXT) echo "PLAINTEXT://${HOST}:${KAFKA_PORT_PLAINTEXT}" ;;
SSL) echo "SSL://${HOST}:${KAFKA_PORT_SSL}" ;;
SASL_SSL*) echo "SASL_SSL://${HOST}:${KAFKA_PORT_SASL_SSL}"
echo "SSL://${HOST}:$((KAFKA_PORT_SASL_SSL+1))" ;; # for inter-broker
esac
} | paste -s -d,
# CA証明書の設定
echo "ssl.truststore.location=/sinetstream-kafka/truststore.p12"
echo "ssl.truststore.password=${TRUSTSTORE_PASSWORD}"
echo "ssl.truststore.type=pkcs12"
# サーバー秘密鍵の設定
echo "ssl.keystore.location=/sinetstream-kafka/keystore.p12"
echo "ssl.keystore.password=${KEYSTORE_PASSWORD}"
echo "ssl.keystore.type=pkcs12"
case "$KAFKA_AUTH" in
SSL)
# SSL/TLS認証(クライアント認証)
echo "ssl.client.auth=required"
echo "security.inter.broker.protocol=SSL"
;;
SASL_SSL_SCRAM)
# SCRAM認証/TLS
echo "ssl.client.auth=required"
echo "security.inter.broker.protocol=SSL"
echo "sasl.enabled.mechanisms=${SCRAM_MECHANISM}"
#echo "sasl.mechanism.inter.broker.protocol=${SCRAM_MECHANISM}"
local scram_mechanism="$(echo "${SCRAM_MECHANISM}" | tr '[A-Z]' '[a-z]')"
echo "listener.name.sasl_ssl.${scram_mechanism}.sasl.jaas.config=org.apache.kafka.common.security.scram.ScramLoginModule required \\"
echo " username=admin password=${PASSWORD_admin};"
;;
SASL_SSL_PLAIN)
# パスワード認証/TLS
echo "ssl.client.auth=required"
echo "security.inter.broker.protocol=SSL"
echo "sasl.enabled.mechanisms=PLAIN"
#echo "sasl.mechanism.inter.broker.protocol=PLAIN"
echo "listener.name.sasl_ssl.plain.sasl.jaas.config=org.apache.kafka.common.security.plain.PlainLoginModule required \\"
echo " username=admin password=${PASSWORD_admin} \\"
local USER PASSWORD
for USER in ${USER_LIST}; do
eval PASSWORD=\$PASSWORD_${USER}
echo " user_${USER}=\"${PASSWORD}\" \\"
done
echo " ;"
;;
esac
# 認可
echo "authorizer.class.name=kafka.security.auth.SimpleAclAuthorizer" # ZooKeeperに記録されているACL設定による認可
echo "allow.everyone.if.no.acl.found=${KAFKA_ACL_DEFAULT_TO_ALLOW}"
echo "super.users=User:admin" # adminには特権を与える
}
ID=1
tar x -f $KAFKA.tgz --to-stdout $KAFKA/config/server.properties >server.properties &&
mkdir -p tmp &&
rm -f tmp/*.properties &&
list_kafka_hosts | while read HOST; do
{
cat server.properties
print_server_properties "$HOST" "$ID"
} >"tmp/server-${HOST}.properties"
ID=$((ID + 1))
done
ls -l tmp/server-*.properties
```
kafkaブローカの設定ファイルを各ホストにコピーする。
```
ansible kafka -m copy -a "src=tmp/server-{{inventory_hostname}}.properties dest=\$PWD/sinetstream-kafka/server.properties"
```
### SSL/TLSのための証明書を設定
opensslをつかってPEM形式の証明書をkafkaブローカが扱えるPKCS#12(p12)形式に変換する。
CA証明書・サーバ秘密鍵・サーバ証明書をkafkaブローカの動かすコンテナ内にコピーする。
自己署名CA証明書の場合はCA秘密鍵もコピーする。
```
CA_CERT_PATH=./cacert.pem
CA_KEY_PATH=NONE
CA_KEY_PATH=./cakey.pem # CA証明書が自己署名の場合はCA秘密鍵も指定する
BROKER_CERT_PATH=./broker.crt
BROKER_KEY_PATH=./broker.key
# 以下、変更しなくてよい
CA_CERT_FILE=$(basename "${CA_CERT_PATH}")
BROKER_CERT_FILE=$(basename "${BROKER_CERT_PATH}")
BROKER_KEY_FILE=$(basename "${BROKER_KEY_PATH}")
if [ "x$CA_KEY_PATH" != "xNONE" ]; then
CA_KEY_FILE=$(basename "${CA_KEY_PATH}")
else
CA_KEY_FILE=""
fi
ansible kafka -m copy -a "src=${CA_CERT_PATH} dest=\$PWD/sinetstream-kafka/${CA_CERT_FILE}" &&
ansible kafka -m copy -a "src=${BROKER_CERT_PATH} dest=\$PWD/sinetstream-kafka/${BROKER_CERT_FILE}" &&
ansible kafka -m copy -a "src=${BROKER_KEY_PATH} dest=\$PWD/sinetstream-kafka/${BROKER_KEY_FILE}" &&
if [ -n "${CA_KEY_FILE}" ]; then
ansible kafka -m copy -a "src=${CA_KEY_PATH} dest=\$PWD/sinetstream-kafka/${CA_KEY_FILE}"
fi
```
CA証明書を変換してtruststoreに登録する。
```
ansible kafka -m command -a "docker exec sinetstream-kafka \
openssl pkcs12 -export \
-in sinetstream-kafka/${CA_CERT_FILE} \
${CA_KEY_FILE:+-inkey sinetstream-kafka/${CA_KEY_FILE}} \
-name private-ca \
-CAfile sinetstream-kafka/${CA_CERT_FILE}\
-caname private-ca \
-out sinetstream-kafka/truststore.p12 \
-passout pass:${TRUSTSTORE_PASSWORD}" &&
ansible kafka -m command -a "docker exec sinetstream-kafka \
openssl pkcs12 -in sinetstream-kafka/truststore.p12 -passin pass:${TRUSTSTORE_PASSWORD} -info -noout"
```
サーバ秘密鍵・サーバ証明書・CA証明書を変換してkeystoreに登録する。
```
ansible kafka -m command -a "docker exec sinetstream-kafka \
openssl pkcs12 -export \
-in sinetstream-kafka/${BROKER_CERT_FILE} \
-inkey sinetstream-kafka/${BROKER_KEY_FILE} \
-name broker \
-CAfile sinetstream-kafka/${CA_CERT_FILE} \
-caname private-ca \
-out sinetstream-kafka/keystore.p12 \
-passout pass:${KEYSTORE_PASSWORD}" &&
ansible kafka -m command -a "docker exec sinetstream-kafka \
openssl pkcs12 -in sinetstream-kafka/keystore.p12 -passin pass:${KEYSTORE_PASSWORD} -info -noout"
```
### SCRAM認証の設定
パスワードをzookeeperに保存する。
```
if [ "x$KAFKA_AUTH" = "xSASL_SSL_SCRAM" ]; then
ZK1="$(list_zookeeper_hosts | head -1)"
KAFKA1="$(list_kafka_hosts | head -1)"
for USER in admin ${USER_LIST}; do
eval PASSWORD=\$PASSWORD_${USER}
ansible kafka --limit="${KAFKA1}" -m command -a "docker exec sinetstream-kafka \
/kafka/bin/kafka-configs.sh --zookeeper ${ZK1}:${ZK_CPORT} --alter \
--entity-type users \
--entity-name ${USER} \
--add-config 'SCRAM-SHA-256=[iterations=8192,password=${PASSWORD}]'"
done &&
ansible kafka -m command -a "docker exec sinetstream-kafka \
/kafka/bin/kafka-configs.sh --zookeeper ${ZK1}:${ZK_CPORT} --describe --entity-type users"
fi
```
### 認可(ACL)の設定
ブローカがつかっているサーバ証明書のCommon Nameを設定する。ブローカ間通信の認可で必要となる。
```
ADMIN_USER="CN=server1.example.jp,C=JP"
ZK1="$(list_zookeeper_hosts | head -1)"
KAFKA1="$(list_kafka_hosts | head -1)"
ansible kafka --limit="${KAFKA1}" -m command -a "docker exec sinetstream-kafka \
/kafka/bin/kafka-acls.sh --authorizer-properties zookeeper.connect=${ZK1}:${ZK_CPORT} \
--add --allow-principal User:${ADMIN_USER} --cluster --operation All" &&
for USER in ${USER_LIST}; do
USER1=$(echo "$USER" | sed 's/[^[:alnum:]]/_/g') # サニタイズ
eval ACL=\$ACL_${USER1}
case "${ACL}" in
*write*)
ansible kafka --limit="${KAFKA1}" -m command -a "docker exec sinetstream-kafka \
/kafka/bin/kafka-acls.sh --authorizer-properties zookeeper.connect=${ZK1}:${ZK_CPORT} \
--add --allow-principal User:${USER} \
--producer --topic '*'"
;;
esac
case "${ACL}" in
*read*)
ansible kafka --limit="${KAFKA1}" -m command -a "docker exec sinetstream-kafka \
/kafka/bin/kafka-acls.sh --authorizer-properties zookeeper.connect=${ZK1}:${ZK_CPORT} \
--add --allow-principal User:${USER} \
--consumer --topic '*' --group '*'"
;;
esac
done
ansible kafka --limit="${KAFKA1}" -m command -a "docker exec sinetstream-kafka \
/kafka/bin/kafka-acls.sh --authorizer-properties zookeeper.connect=${ZK1}:${ZK_CPORT} \
--list"
```
### Kafkaブローカー起動
```
ansible kafka -m command -a "docker exec --detach sinetstream-kafka \
/kafka/bin/kafka-server-start.sh /sinetstream-kafka/server.properties"
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
dataset = pd.read_csv('../data/2016-11-22-reimbursements.xz',
dtype={'document_id': np.str,
'congressperson_id': np.str,
'congressperson_document': np.str,
'term_id': np.str,
'cnpj_cpf': np.str,
'reimbursement_number': np.str},
low_memory=False)
dataset = dataset[dataset['year']==2016]
dataset.head()
```
# Find spends: congress person per month
```
def find_spends_by_month(df, applicant_id):
'''
Return a dataframe with the sum of values of spends by month
of the congress person of "applicant_id"
Parameters:
- df: pandas dataframe to be sliced
- applicant_id: unique id of the congress person
Ex: find_spends_by_month(df, 731)
Result dataframe contains:
- 1/Jan sum
- 2/Feb sum
- 3/Mar sum
- 4/Apr sum
- 5/May sum
- 6/Jun sum
- 7/Jul sum
- 8/Aug sum
- 9/Sep sum
- 10/Oct sum
- 11/Nov sum
- 12/Dec sum
- name
'''
months={1:"Jan",
2:"Feb",
3:"Mar",
4:"Apr",
5:"May",
6:"Jun",
7:"Jul",
8:"Aug",
9:"Sep",
10:"Oct",
11:"Nov",
12:"Dec"}
df_applicant = df[df.applicant_id == applicant_id]
result = {
"name":df_applicant["congressperson_name"].unique()
}
for m in months.keys():
data = df_applicant[df.month == m]
result["{:>02}".format(m) + "/" + months[m]] = data.total_net_value.sum()
df_final = pd.DataFrame([result])
ax = df_final.plot(kind='bar', title ="Congress Person Spends by Month", figsize=(25, 20), legend=True, fontsize=12)
ax.set_xlabel("Month", fontsize=12)
ax.set_ylabel("Value", fontsize=12)
plt.show()
return pd.DataFrame([result])
find_spends_by_month(dataset, 731)
```
# Find spends: Congress Person per Subquotas
```
def find_spends_by_subquota(df, applicant_id):
'''
Return a dataframe with the sum of values of spends by subquotas
of the congress person of "applicant_id"
Parameters:
- df: pandas dataframe to be sliced
- applicant_id: unique id of the congress person
'''
df_applicant = df[df.applicant_id == applicant_id]
result = {
"name":df_applicant["congressperson_name"].unique()
}
for c in df["subquota_description"].unique():
data = df_applicant[df.subquota_description == c]
result[c] = data.total_net_value.sum()
df_final = pd.DataFrame([result])
ax = df_final.plot(kind='bar', title ="Congress Person Spends by Subquotas", figsize=(25, 20), legend=True, fontsize=12)
ax.set_xlabel("Subquotas", fontsize=12)
ax.set_ylabel("Value", fontsize=12)
plt.show()
return pd.DataFrame([result])
find_spends_by_subquota(dataset, 731)
def find_spends_by_subquota(df, applicant_id, month=None):
'''
Return a dataframe with the sum of values of spends by subquotas
of the congress person of "applicant_id" and month "month"
Parameters:
- df: pandas dataframe to be sliced
- applicant_id: unique id of the congress person
'''
df_applicant = df[df.applicant_id == applicant_id]
result = {
"name":df_applicant["congressperson_name"].unique(),
"total": 0
}
if month != None:
df_applicant = df_applicant[df_applicant.month==month]
for c in df["subquota_description"].unique():
data = df_applicant[df.subquota_description == c]
result[c] = data.total_net_value.sum()
result["total"] += result[c]
df_final = pd.DataFrame([result])
ax = df_final.plot(kind='bar', title ="Congress Person", figsize=(25, 20), legend=True, fontsize=12)
ax.set_xlabel("Name", fontsize=12)
ax.set_ylabel("Value", fontsize=12)
plt.show()
return pd.DataFrame([result])
find_spends_by_subquota(dataset, 731, 3)
```
# Find spends: all congress people
```
def find_sum_of_values(df, aggregator, property):
'''
Return a dataframe with the statistics of values from "property"
aggregated by unique values from the column "aggregator"
Parameters:
- df: pandas dataframe to be sliced
- aggregator: dataframe column that will be
filtered by unique values
- property: dataframe column containing values to be summed
Ex: find_sum_of_values(data, 'congressperson_name', 'net_value')
Result dataframe contains (for each aggregator unit):
- property sum
- property mean value
- property max value
- property mean value
- number of occurences in total
'''
total_label = '{}_total'.format(property)
max_label = '{}_max'.format(property)
mean_label = '{}_mean'.format(property)
min_label = '{}_min'.format(property)
result = {
'occurences': [],
aggregator: df[aggregator].unique(),
max_label: [],
mean_label: [],
min_label: [],
total_label: [],
}
for item in result[aggregator]:
if isinstance(df[aggregator].iloc[0], str):
item = str(item)
values = df[df[aggregator] == item]
property_total = int(values[property].sum())
occurences = int(values[property].count())
result[total_label].append(property_total)
result['occurences'].append(occurences)
result[mean_label].append(property_total/occurences)
result[max_label].append(np.max(values[property]))
result[min_label].append(np.min(values[property]))
return pd.DataFrame(result).sort_values(by=aggregator)
df = find_sum_of_values(dataset, "congressperson_name", "total_net_value")
df[:10]
```
# Finding congress people that spent more than 500 thousand per year
```
df = df[df.total_net_value_total > 500000]
df
ax = df[["total_net_value_total"]].plot(kind='bar', title ="Congress Person Spends", figsize=(15, 10), legend=True, fontsize=12)
ax.set_xlabel("Congress Person", fontsize=12)
ax.set_ylabel("Value", fontsize=12)
plt.show()
```
| github_jupyter |
```
from lxml import etree
root = etree.Element("root")
print(root.tag)
root.append(etree.Element('child1'))
child2 = etree.SubElement(root, "child2")
child3 = etree.SubElement(root, "child3")
print(etree.tostring(root, pretty_print=True).decode('utf-8'))
child = root[0]
print(child.tag)
print(len(root))
root.index(root[1])
children = list(root)
children
for child in root:
print(child.tag)
root.insert(0, etree.Element("child0"))
print(etree.tostring(root, pretty_print=True).decode('utf-8'))
start = root[:1]
end = root[-1:]
print(start[0].tag)
print(end[0].tag)
if root: # this no longer works!
print("The root element has children")
etree.iselement(root)
if len(root):
print("The root element has children")
print(etree.tostring(root, pretty_print=True).decode('utf-8'))
root[0] = root[-1]
print(etree.tostring(root, pretty_print=True).decode('utf-8'))
root is root[0].getparent()
from copy import deepcopy
element = etree.Element("neu")
element.append( deepcopy(root[1]) )
print(element[0].tag)
print([c.tag for c in root])
row = [1,2,3,4,5,6]
iterrow = iter(row)
[(i,j) for i,j in (iterrow(),iterrow())]
print("str",1)
foldername = os.curdir
filename = 'test.xml'
imgSize = (800, 1000)
from lxml.etree import ElementTree, Element, SubElement
from lxml import etree
pointlist = [(1,2),
(3,4),
(5,6),
(7,8),
(9,10)]
top = Element('annotation')
for i, each_object in enumerate(pointlist):
point = SubElement(top, "point"+str(i).zfill(3))
X = SubElement(point,"X")
X.text = str(each_object[0])
Y = SubElement(point,"Y")
Y.text = str(each_object[1])
print(etree.tostring(top, pretty_print=True).decode('utf-8'))
for each_object in self.shapelist:
object_item = SubElement(top, 'object')
name = SubElement(object_item, 'name')
try:
name.text = unicode(each_object['name'])
except NameError:
# Py3: NameError: name 'unicode' is not defined
name.text = each_object['name']
pose = SubElement(object_item, 'pose')
pose.text = "Unspecified"
truncated = SubElement(object_item, 'truncated')
if int(each_object['ymax']) == int(self.imgSize[0]) or (int(each_object['ymin'])== 1):
truncated.text = "1" # max == height or min
elif (int(each_object['xmax'])==int(self.imgSize[1])) or (int(each_object['xmin'])== 1):
truncated.text = "1" # max == width or min
else:
truncated.text = "0"
difficult = SubElement(object_item, 'difficult')
difficult.text = str( bool(each_object['difficult']) & 1 )
bndbox = SubElement(object_item, 'bndbox')
xmin = SubElement(bndbox, 'xmin')
xmin.text = str(each_object['xmin'])
ymin = SubElement(bndbox, 'ymin')
ymin.text = str(each_object['ymin'])
xmax = SubElement(bndbox, 'xmax')
xmax.text = str(each_object['xmax'])
ymax = SubElement(bndbox, 'ymax')
ymax.text = str(each_object['ymax'])
from PyQt5.QtGui import QImageReader
formats = ['*.%s' % fmt.data().decode("ascii").lower() for fmt in QImageReader.supportedImageFormats()]
formats
class cache(object):
def __init__(self, folderpath, filename, num = 1000):
#
self.folder = path = os.path.dirname(ustr(self.filePath)) if self.filePath else '.'
self.total
pass
def update(self):
def openFile(self, _value=False):
formats = ['*.%s' % fmt.data().decode("ascii").lower() for fmt in QImageReader.supportedImageFormats()]
filters = "Image & Label files (%s)" % ' '.join(formats + ['*%s' % LabelFile.suffix])
filename = QFileDialog.getOpenFileName(self, '%s - Choose Image or Label file' % __appname__, path, filters)
if filename:
if isinstance(filename, (tuple, list)):
filename = filename[0]
self.loadFile(filename)
def prev(self, _value=False):
# Proceding prev image without dialog if having any label
if self.autoSaving.isChecked():
if self.defaultSaveDir is not None:
if self.dirty is True:
self.saveFile()
else:
self.changeSavedirDialog()
return
if not self.mayContinue():
return
if len(self.mImgList) <= 0:
return
if self.filePath is None:
return
currIndex = self.mImgList.index(self.filePath)
if currIndex - 1 >= 0:
filename = self.mImgList[currIndex - 1]
if filename:
self.loadFile(filename)
def openNextImg(self, _value=False):
# Proceding prev image without dialog if having any label
if self.autoSaving.isChecked():
if self.defaultSaveDir is not None:
if self.dirty is True:
self.saveFile()
else:
self.changeSavedirDialog()
return
if not self.mayContinue():
return
if len(self.mImgList) <= 0:
return
filename = None
if self.filePath is None:
filename = self.mImgList[0]
else:
currIndex = self.mImgList.index(self.filePath)
if currIndex + 1 < len(self.mImgList):
filename = self.mImgList[currIndex + 1]
if filename:
self.loadFile(filename)
import math
math.floor(-0.1)
from PyQt5.QtGui import QImage
image = QImage()
image.load("/Volumes/Jerry's/zhouhan/420V5/420V10Q225/420V10Q225_0000.jpg")
image
alist = [1,2,3,4,5]
flag = alist.index(0)
flag
from PyQt5.QtGui import QImage
filename = "/Volumes/Jerry's/zhouhan/420V5/420V10Q225/420V10Q225_0000.jpg"
with open(filename, 'rb') as f:
imageData = f.read()
image = QImage.fromData(imageData)
image.format()
image.Format_RGB32
image.width()
image.height()
image.depth()
import qimage2ndarray
import numpy as np
import cv2
import matplotlib.pyplot as plt
# cv2.
plt.imshow((qimage2ndarray.rgb_view(image)).reshape(800,1072,3))
plt.show()
import cv2
filename = "/Volumes/Jerry's/zhouhan/420V5/420V10Q225/420V10Q225_0000.jpg"
cvimg = cv2.imread(filename)
cvimg.shape
import os
import glob
import cv2
import matplotlib.pyplot as plt
from PyQt5.QtCore import pyqtSignal
import numpy as np
# background subtraction and other preprossing procedure
FILENAME = "background.jpg"
class Preprocess:
preprocessed = pyqtSignal()
def __init__(self, dirname, length=10):
if os.path.isdir(dirname):
self.dirname = dirname
else:
return None
self.imgPathList = glob.glob(os.path.join(self.dirname,'*.jpg'))
tmp_img = cv2.imread(self.imgPathList[0])
height, width, depth = tmp.img
self.bg = np.zeros((height, width, depth))
self.imgList = []
self.length = length
def __call__(self):
img_path = os.path.join(self.dirname, FILENAME)
# if we process it before and there exists backgrounnd image.
if os.path.exists(img_path):
self.bg = cv2.imread(img_path)
else:
self.imgPathList
for index, f in enumerate(paths):
if index >= self.length:
self.bg /= self.length
break
img = cv2.imread(f)
self.bg = np.add(self.bg, img)
plt.imshow(bgimg_gray)
def qimage2numpyarray(image):
height = image.height()
width = image.width()
return (qimage2ndarray.rgb_view(image)).reshape(height, width, 3)
arr = qimage2numpyarray(image)
arr
def numpyarray2qimage(arr):
# height = self.numpyarray.shape[1]
# width = self.numpyarray.shape[0]
numpyarray = np.transpose(arr, (1,0,2))
return QImage(arr.tobytes(), 800, 1072, 6400, QImage.Format_RGB888)
qimage2numpyarray(numpyarray2qimage(arr))
def getCurrent(self):
if self.subtractBackGround:
return self.numpyarray2qimage(\
np.subtract(self.qimage2numpyarray(self.data[self.cursor]) \
,self.background))
else:
return self.data[self.cursor]
def qimage2numpyarray(self, imgdata):
height = self.data[0].height()
width = self.data[0].width()
return (qimage2ndarray.rgb_view(imgdata)).reshape(height, width, 3)
def numpyarray2qimage(self, numpyarray):
# height = self.numpyarray.shape[1]
# width = self.numpyarray.shape[0]
numpyarray = np.transpose(numpyarray, (1,0,2))
return QImage(numpyarray.tobytes(), numpyarray.shape[1], numpyarray.shape[0], QImage.Format_RGB888)
ims = []
cal = []
cal_gray = []
paths = glob.glob('data/600V350Q401/*.jpg')
step = 1
image = cv.imread(paths[0])
bgimg_gray = np.zeros((800, 1072))
# print(type(bgimg_gray))
# print(bgimg_gray.shape)
for index, f in zip(range(len(paths)-1), paths[::step]):
if index >= step + 1000:
bgimg_gray /= index
break
image = cv.imread(f)
image = cv.cvtColor(image, cv.COLOR_BGR2RGB)
image_gray = cv.cvtColor(image, cv.COLOR_RGB2GRAY)
bgimg_gray = np.add(bgimg_gray, image_gray)
plt.imshow(bgimg_gray, cmap='gray')
from PyQt5.QtCore import QThread, pyqtSignal
class BackendThread(QThread):
update_cache = pyqtSignal(int)
def run(self):
class LabelsList():
def __init__(self, length):
self.labelslist = [None for _ in range(length)]
def __len__(self):
return len(self.labelslist)
def update(self,i, shapes):
self.labelslist[i] = shapes
def __getitem__(self, n):
try:
self.labelslist[n]
except IndexError:
return None
else:
return self.labelslist[n]
def __setitem__(self, n, value):
self.labelslist[n] = value
def __delitem__(self, n):
todelete = self.labelslist[n]
self.labelslist[n] = None
del(todelete)
ll = LabelsList(999)
ll[1] = 2
len(ll)
ll[1]
ll[98] is None
l = [1,2,3,4]
for i1, i2 in enumerate(l, start=1):
print(i1, i2)
a = slice(0,5,1)
a.indices(4)
ls = [None for _ in range(20)]
ls1, ls[1] = ls[1], None
del ls1
len(ls)
class lsmanager:
def __init__(self):
self.ls = [1,2,3]
def __delitem__(self, i):
self.ls[i], tmp = None, self.ls[i]
del tmp
def __getitem__(self, i):
return self.ls[i]
def show(self,i):
print(self[i])
ls = lsmanager()
ls
ls.show(1)
del ls[1]
ls.ls
del(ls[2])
ls.ls
import sys
class _const:
class ConstError(TypeError):
pass
class ConstCaseError(ConstError):
pass
def __setattr__(self, name, value):
if name in self.__dict__.keys():
raise(self.ConstError, "Can't change const. ")
if not name.isupper():
raise(ConstCaseError, \
'const name "{}" is not all uppercase'.format(name))
self.__dict__[name] = value
sys.modules["const"]=_const()
# sys.modules
import const
const.A = 1
const.A
from PyQt5.QtCore import QPointF
from PyQt5.QtGui import QColor
import os
# import glob
import cv2
import matplotlib.pyplot as plt
import numpy as np
from PyQt5.QtCore import pyqtSignal, QPointF, QThread
# background subtraction and other preprossing procedure
BACKGROUND_FILENAME = "background.jpg"
class PreprocessThread(QThread):
def __init__(self, imgList, length=10, imread_format=cv2.IMREAD_GRAYSCALE):
super(PreprocessThread, self).__init__()
self.imgPathList = imgList
self.backgroundFilePath = os.path.join(os.path.dirname(self.imgPathList[0]), BACKGROUND_FILENAME)
self.background = None
self.length = length
self.imread_format = imread_format
tmp_img = cv2.imread(self.imgPathList[0], self.imread_format)
if len(tmp_img.shape) == 2:
self.height, self.width, self.depth = *tmp_img.shape, 1
elif len(tmp_img.shape) == 3:
self.height, self.width, self.depth = tmp_img.shape
def generateBackground(self):
if self.depth == 1:
background = np.zeros((self.height, self.width))
elif self.detph == 3:
background = np.zeros((self.height, self.width, self.depth))
print("self.height, self.width, self.depth", self.height, self.width, self.depth)
for index, f in zip(range(self.length), self.imgPathList):
print("enter iter", index)
img = cv2.imread(f, self.imread_format)
background = np.add(background, img)
background /= min(self.length, len(self.imgPathList))
cv2.imwrite(self.backgroundFilePath,background)
return background
def run(self):
print("enter run")
if os.path.exists(self.backgroundFilePath):
print("enter self.backgroundFilePath")
self.background = cv2.imread(self.backgroundFilePath, self.imread_format)
print(self.background, 1)
if self.background is None:
print(self.background, 2)
self.background = self.generateBackground()
print(self.background, 3)
def __getitem__(self,i):
ori = self.load_origin(i)
result = self.backgroundSubtraction(ori)
result = self.equalizeHist(result)
return result
def load_origin(self, i):
return cv2.imread(self.imgPathList[i], self.imread_format)
def backgroundSubtraction(self, img, beta = 0.85):
background = self.background if self.background is not None else self.generateBackground()
result = np.subtract(img, beta*self.background)
result = abs(result)
return result
def equalizeHist(self, img):
return cv2.equalizeHist(np.array(img, dtype=np.uint8))
imgList = [ "/Volumes/Jerry's/zhouhan/420V5/420V10Q225/420V10Q225_0000.jpg",
"/Volumes/Jerry's/zhouhan/420V5/420V10Q225/420V10Q225_0001.jpg",
"/Volumes/Jerry's/zhouhan/420V5/420V10Q225/420V10Q225_0002.jpg",
"/Volumes/Jerry's/zhouhan/420V5/420V10Q225/420V10Q225_0003.jpg",
"/Volumes/Jerry's/zhouhan/420V5/420V10Q225/420V10Q225_0004.jpg",
"/Volumes/Jerry's/zhouhan/420V5/420V10Q225/420V10Q225_0005.jpg",
"/Volumes/Jerry's/zhouhan/420V5/420V10Q225/420V10Q225_0006.jpg",
"/Volumes/Jerry's/zhouhan/420V5/420V10Q225/420V10Q225_0007.jpg",
"/Volumes/Jerry's/zhouhan/420V5/420V10Q225/420V10Q225_0008.jpg",
"/Volumes/Jerry's/zhouhan/420V5/420V10Q225/420V10Q225_0009.jpg",
"/Volumes/Jerry's/zhouhan/420V5/420V10Q225/420V10Q225_0010.jpg"]
backend = PreprocessThread(imgList)
backend.start()
backend[1]
type(backend[1])
backend[1].min()
plt.imshow(backend[1],cmap='gray')
plt.show()
cv2.imshow("img",cv2.fromarray(backend[1]))
filename = "/Volumes/Jerry's/zhouhan/420V5/420V10Q225/420V10Q225_0000.jpg"
os.path.basename(filename)
os.path.dirname(filename)
np.subtract()
import numpy as np
from PyQt5.QtGui import QImage
import cv2
```
[将np.array 转成 QImage 格式](https://www.aliyun.com/jiaocheng/433464.html?spm=5176.100033.2.9.34cb4d61JspVQl)
```
def ndarray2qimage(ndarray):
# qimage = QImage()
if len(ndarray.shape) == 2:
height, width = ndarray.shape
elif len(ndarray.shape) == 3:
height, width, depth = ndarray.shape
if ndarray.dtype == np.uint8:
bytesPerComponent = 8
bytesPerLine = bytesPerComponent * width
# image = cv2.cvtColor(ndarray, cv2.COLOR_GRAY2RGB)
qimage = QImage(ndarray.data, width, height, bytesPerLine, QImage.Format_RGB888)
print(qimage)
def numpyarray2qimage(arr):
# height = self.numpyarray.shape[1]
# width = self.numpyarray.shape[0]
numpyarray = np.transpose(arr, (1,0,2))
return QImage(arr.tobytes(), 800, 1072, 6400, QImage.Format_RGB888)
# class mycsms(QWidget, Ui_csms, Ui_MainWindow):
# def __init__(self):
# super(mycsms, self).__init__()
# self.setupUi(self)
# self.image = QImage()
# self.device = cv2.VideoCapture(0)
# self.playTimer = Timer("updatePlay()")
# self.connect(self.playTimer, SIGNAL("updatePlay()"), self.showCamer)
# # 读摄像头
# def showCamer(self):
# if self.device.isOpened():
# ret, frame = self.device.read()
# else:
# ret = False
# # 读写磁盘方式
# # cv2.imwrite("2.png", frame)
# # self.image.load("2.png")
# height, width, bytesPerComponent = frame.shape
# bytesPerLine = bytesPerComponent * width
# # 变换彩色空间顺序
# cv2.cvtColor(frame, cv2.COLOR_BGR2RGB,frame)
# # 转为QImage对象
# self.image = QImage(frame.data, width, height, bytesPerLine, QImage.Format_RGB888)
# self.view.setPixmap(QPixmap.fromImage(self.image))
img = cv2.imread(filename, cv2.IMREAD_GRAYSCALE)
ndarray2qimage(img)
import os
os.remove()
cv2.imwrite()
float("11a")
dct = dict()
dct.update({"1":2})
dct.update({"1":3})
dct
dct.update(dict(('1',2)))
import pandas as pd
df = pd.DataFrame(columns=['dir','subdir', 'img', 'index', 'radius'])
df.update([1,2,3,4,5])
df
df.append([1,2,3,4,5])
df = df.append([{'dir':1,'subdir':2, 'img':3, 'index':4, 'radius':5}], ignore_index=True)
df
df['radius'].mean()
values, bins = np.histogram(df['radius'])
bins = [(bins[i], bins[i+1]) for i in range(len(bins)-1)]
bins
distribution= ''
for bin_, value in zip(bins, values):
distribution += '({0[0]:.2f}, {0[0]:.2f}) : {1: d}\n'.format(bin_, value)
print(distribution)
import os
filename = "/Volumes/Jerry's/zhouhan/420V5/420V10Q225/420V10Q225_0000.jpg"
os.path.basename(filename)
os.path.basename(os.path.dirname(filename))
os.path.dirname(os.path.dirname(filename))
xmlfile = os.path.basename(filename)
subdir = os.path.basename(os.path.dirname(filename))
dir_ = os.path.dirname(os.path.dirname(filename))
os.path.join(xmlfile, subdir,dir_)
from math import sqrt
def getRadius(pts):
def distance(p1, p2):
x1, y1 = p1
x2, y2 = p2
return sqrt( (x2-x1)**2 + (y2-y1)**2 )
p1, p2, p3, p4 = pts
return sqrt(distance(p1, p2) * distance(p3, p4))
pts = [(0,0),
(0,1),
(1,2),
(1,3)]
getRadius(pts)
lst = [1,2,3,4]
lst.index(1)
from PyQt5.QtCore import QPointF
csvfilename = "/Volumes/Jerry's/zhouhan/420V5/420V10Q225/track.csv"
import pandas as pd
df = pd.read_csv(csvfilename,encoding='utf-8')
df
bool(df.difficult[0])
dir_ = df.get_value(0,'dir')
subdir = df.get_value(0,'subdir')
dirPath = os.path.join(dir_, subdir)
for i, row in df.iterrows():
imgfile = row.imgfile
label = row.label
shapeType = row.shapeType
points = eval(row.points)
difficult = bool(row.difficult)
type(df.difficult[0])
from PyQt5.QtGui import QPainterPath
from PyQt5.QtCore import QPoint
path = QPainterPath(self.points[0])
for p in self.points[1:]:
path.lineTo(p)
from collections import OrderedDict
d = OrderedDict()
d[1] = []
d[2] = []
OrderedDict[0]
d[3]
for i, j in d.items():
print(i,j)
d[1].append(1)
d
x=1
y=2
(x,y) in [(1,2)]
from collections import namedtuple
node = namedtuple("node",["i_img", "data"])
nodes = [node(0, 0),
node(1, 1),
node(2, 2),
node(2, 3),
node(3, 4),
node(3, 5),
node(3, 6),
node(4, 7),
node(4, 8),
node(4, 9),
node(4, 10),
node(4, 11),
node(5, 12),
node(6, 13)]
tree = Tree(1)
tree.addKid(tree.root, 2)
tree.addKid(tree.root, 3)
[i for i in filter(None,[None, None,None,None])]
from functools import reduce
reduce(lambda x,y: x and y, [True, True, False])
if reduce(lambda x,y: x or y, [Node(shape_obj) in for shape_obj]):
continue
{"hello":"world", "say":"hi"}.get("hello")
reduce(lambda x,y: x and y, [True, True, False])
import unittest
class TestStringMethods(unittest.TestCase):
def test_upper(self):
self.assertEqual('foo'.upper(), 'FOO')
def test_isupper(self):
self.assertTrue('FOO'.isupper())
self.assertFalse('Foo'.isupper())
def test_split(self):
s = 'hello world'
self.assertEqual(s.split(), ['hello', 'world'])
# check that s.split fails when the separator is not a string
with self.assertRaises(TypeError):
s.split(2)
if __name__ == '__main__':
unittest.main()
list(reversed([1,2,3,4]))
ls = ["C", "A", "A 1", "A_2 ", "A 3", "B", "B 1", "B 2", "b 3"]
sorted(ls)
ls.sort()
ls
list(map(lambda s: s.lower().strip(), ls))
from collections import OrderedDict
list.sort()
def groupByList(ls, sep=" "):
ls = list(map(lambda s: s.lower().strip(), ls))
ls.sort()
lists = OrderedDict()
for label in ls:
key = " "
if " " not in label:
key = label
lists[key] = []
if label.split(sep, 1)[0] in lists.keys():
lists[label.split(sep, 1)[0]].append(label)
return lists
lists = groupByList(ls)
lists
for l in lists:
print(l)
"1 2 3".split(" ", maxsplit=1)
str.split()
ls = [1,2,3,4,5,6]
for _,j,k in zip(*(iter(ls),) * 3):
print(j,k)
dict1 = dict([(1, [1,1]),(2,[2,2])])
dict2 = dict([(1, [2]),(3,[3,3])])
dict1.update(dict2)
dict1
import pandas as pd
df = pd.DataFrame([(0,1,"x"),(2,3,"y"),(4,5,"y")], columns= ['x','y','z'])
df
df["x"][1]
for i, row in df.iterrows():
print(row)
print(row[0])
print(row[[0,1]])
print(tuple(row[["x","y"]]))
print(row.x)
print(type(row))
print(tuple(row))
print()
break
ls = [4,2,3,5]
ls.sort()
ls
```
| github_jupyter |
```
# default_exp visrectrans
```
# VisRecTrans
> A class for creating a custom [Vision Transformer (ViT)](https://arxiv.org/abs/2010.11929) model for visual recognition
```
#export
#hide
from nbdev.showdoc import *
from fastai.vision.all import *
import timm
import math
import warnings
#export
#hide
class EmbedBlock (Module) :
def __init__ (self, num_patches, embed_dim) :
self.cls_tokens = nn.Parameter(torch.zeros(1, 1, embed_dim))
self.pos_embeds = nn.Parameter(torch.zeros(1, num_patches + 1, embed_dim))
def forward (self, x) :
B = x.shape[0]
cls_tokens = self.cls_tokens.expand(B, -1, -1)
x = torch.cat((cls_tokens, x), dim = 1)
x = x + self.pos_embeds
return x
#export
#hide
class Header (Module) :
def __init__ (self, ni, num_classes) :
self.head = nn.Linear(ni, num_classes)
def forward (self, x) :
x = x[:, 0] # Extracting the clsass token, which is used for the classification task.
x = self.head(x)
return x
#export
#hide
def custom_ViT (timm_model_name, num_patches, embed_dim, ni, num_classes, pretrained = True) :
model = timm.create_model(timm_model_name, pretrained)
module_layers = list(model.children())
return nn.Sequential(
module_layers[0],
EmbedBlock(num_patches, embed_dim),
nn.Sequential(*module_layers[1:-1]),
Header(ni, num_classes)
)
#export
#hide
# The function below is heavily inspired by "https://github.com/rwightman/pytorch-image-models/blob/5f9aff395c224492e9e44248b15f44b5cc095d9c/timm/models/vision_transformer.py"
def _no_grad_trunc_normal_(tensor, mean, std, a, b):
# Cut & paste from PyTorch official master until it's in a few official releases - RW
# Method based on https://people.sc.fsu.edu/~jburkardt/presentations/truncated_normal.pdf
def norm_cdf(x):
# Computes standard normal cumulative distribution function
return (1. + math.erf(x / math.sqrt(2.))) / 2.
if (mean < a - 2 * std) or (mean > b + 2 * std):
warnings.warn("mean is more than 2 std from [a, b] in nn.init.trunc_normal_. "
"The distribution of values may be incorrect.",
stacklevel=2)
with torch.no_grad():
# Values are generated by using a truncated uniform distribution and
# then using the inverse CDF for the normal distribution.
# Get upper and lower cdf values
l = norm_cdf((a - mean) / std)
u = norm_cdf((b - mean) / std)
# Uniformly fill tensor with values from [l, u], then translate to
# [2l-1, 2u-1].
tensor.uniform_(2 * l - 1, 2 * u - 1)
# Use inverse cdf transform for normal distribution to get truncated
# standard normal
tensor.erfinv_()
# Transform to proper mean, std
tensor.mul_(std * math.sqrt(2.))
tensor.add_(mean)
# Clamp to ensure it's in the proper range
tensor.clamp_(min=a, max=b)
return tensor
def trunc_normal_(layer, param, mean=0., std=1., a=-2., b=2.):
# type : (Tensor, float, float, float, float) -> Tensor
"""Fills the input Tensor with values drawn from a truncated
normal distribution. The values are effectively drawn from the
normal distribution :math:`\mathcal{N}(\text{mean}, \text{std}^2)`
with values outside :math:`[a, b]` redrawn until they are within
the bounds. The method used for generating the random values works
best when :math:`a \leq \text{mean} \leq b`.
Args:
tensor: an n-dimensional `torch.Tensor`
mean: the mean of the normal distribution
std: the standard deviation of the normal distribution
a: the minimum cutoff value
b: the maximum cutoff value
Examples:
>>> w = torch.empty(3, 5)
>>> nn.init.trunc_normal_(w)
"""
tensor = layer.get_parameter(param)
return _no_grad_trunc_normal_(tensor, mean, std, a, b)
#export
class VisRecTrans :
"""Class for setting up a vision transformer for visual recognition.
Returns a pretrained custom ViT model for the given `model_name` and `num_classes`, by default, or, with randomly initialized parameters, if `pretrained`
is set to False.
"""
models_list = ['vit_large_patch16_224', 'vit_large_patch16_224_in21k', 'vit_huge_patch14_224_in21k', 'vit_small_patch16_224', 'vit_small_patch16_224_in21k']
# Two tasks : (1) Generalize the assignments of num_path (2) (3) ()
def __init__(self, model_name, num_classes, pretrained = True) :
self.model_name = model_name
self.num_classes = num_classes
self.pretrained = pretrained
if self.model_name == 'vit_small_patch16_224' :
self.num_patches = 196
self.embed_dim = 384
self.ni = 384
elif self.model_name == 'vit_small_patch16_224_in21k' :
self.num_patches = 196
self.embed_dim = 384
self.ni = 384
elif self.model_name == 'vit_large_patch16_224' :
self.num_patches = 196
self.embed_dim = 1024
self.ni = 1024
elif self.model_name == 'vit_large_patch16_224_in21k' :
self.num_patches = 196
self.embed_dim = 1024
self.ni = 1024
elif self.model_name == 'vit_huge_patch14_224_in21k' :
self.num_patches = 256
self.embed_dim = 1280
self.ni = 1280
def create_model (self) :
"""Method for creating the model.
"""
return custom_ViT(self.model_name, self.num_patches, self.embed_dim, self.ni, self.num_classes, self.pretrained)
def initialize (self, model) :
"""Mthod for initializing the given `model`. This method uses truncated normal distribution for
initializing the position embedding as well as the class token, and, the head of the model is
initialized using He initialization.
"""
trunc_normal_(model[1], 'cls_tokens')
trunc_normal_(model[1], 'pos_embeds')
apply_init(model[3], nn.init.kaiming_normal_)
def get_callback (self) :
"""Method for getting the callback to train the embedding block of the `model`. It is highly recommended
to use the callback, returned by this method, while training a ViT model.
"""
class TrainEmbedCallback(Callback) :
def before_train(self) :
self.model[1].training = True
self.model[1].requires_grad_(True)
def before_validation(self) :
self.model[1].training = False
self.model[1].requires_grad_(False)
return TrainEmbedCallback()
show_doc(VisRecTrans.create_model)
show_doc(VisRecTrans.initialize)
show_doc(VisRecTrans.get_callback)
```
Let's see if this class is working well :
```
vis_rec_ob = VisRecTrans('vit_small_patch16_224', 10, False)
model_test = vis_rec_ob.create_model()
vis_rec_ob.initialize(model_test)
assert isinstance(model_test, nn.Sequential)
```
As we see, the model is a sequential list of layers, and can be used with the `Learner` class of [fastai](https://docs.fast.ai), as we use any other model.
#### The list of models supported by the `VisRecTrans` class :
```
VisRecTrans.models_list
```
| github_jupyter |
```
%matplotlib notebook
import numpy as np
import matplotlib.pyplot as plt
import tessreduce as tr
from scipy.optimize import minimize
from scipy import signal
from astropy.convolution import Gaussian2DKernel
from scipy.optimize import minimize
def Delta_basis(Size = 13):
kernal = np.zeros((Size,Size))
x,y = np.where(kernal==0)
middle = int(len(x)/2)
basis = []
for i in range(len(x)):
b = kernal.copy()
if (x[i] == x[middle]) & (y[i] == y[middle]):
b[x[i],y[i]] = 1
else:
b[x[i],y[i]] = 1
b[x[middle],y[middle]] = -1
basis += [b]
basis = np.array(basis)
coeff = np.ones(len(basis))
return basis, coeff
ra = 189.1385817
dec = 11.2316535
#ra = 64.526125
#dec = -63.61506944
tpf = tr.Get_TESS(ra,dec)
mask = tr.Make_mask(tpf,)
bkg = tr.New_background(tpf,mask,)
flux = tpf.flux.value - bkg
ref = flux[100]
offset = tr.Centroids_DAO(flux,ref,TPF=tpf,parallel=False)
cor = tr.Shift_images(offset,flux)
def Delta_kernal(Scene,Image,Size=13):
Basis, coeff_0 = Delta_basis(Size)
bds = []
for i in range(len(coeff_0)):
bds += [(0,1)]
coeff_0 *= 0.01
res = minimize(optimize_delta, coeff_0, args=(Basis,Scene,Image),
bounds=bds)
k = np.nansum(res.x[:,np.newaxis,np.newaxis]*Basis,axis=0)
return k
def optimize_delta(Coeff, Basis, Scene, Image):
Kernal = np.nansum(Coeff[:,np.newaxis,np.newaxis]*Basis,axis=0)
template = signal.fftconvolve(Scene, Kernal, mode='same')
im = Image.copy()
res = np.nansum(abs(im-template))
#print(res)
return res
thing = Delta_kernal(cor[100],cor[1200],Size=7)
def Make_temps(image, ref,size=7):
k = Delta_kernal(ref,image,Size=size)
template = signal.fftconvolve(ref, k, mode='same')
return template
ref = cor[100]
temps = np.zeros_like(cor)
for i in range(cor.shape[0]):
k = Delta_kernal(ref,cor[i],Size=7)
template = signal.fftconvolve(ref, k, mode='same')
print(i)
import multiprocessing
from joblib import Parallel, delayed
from tqdm import tqdm
num_cores = multiprocessing.cpu_count()
tmps = Parallel(n_jobs=num_cores)(delayed(Make_temps)(flux[i],ref) for i in tqdm(range(cor.shape[0])))
templates = np.array(tmps)
templates.shape
sub = cor - templates
sub2 = f - templates
#lc1, sky1 = tr.diff_lc(sub,tpf=tpf,ra=ra,dec=dec,tar_ap=3,sky_in=3,sky_out=5)
lc1, sky1 = tr.diff_lc(cor,tpf=tpf,ra=ra,dec=dec)
lc2, sky2 = tr.diff_lc(cor,tpf=tpf,x=45,y=50)
lc1[1] = lc1[1] - lc2[1]
plt.figure()
plt.fill_between(lc1[0],lc1[1]-lc1[2],lc1[1]+lc1[2],alpha=0.2)
plt.plot(lc1[0],lc1[1],'.')
plt.plot(sky1[0],sky1[1])
plt.fill_between(lc2[0],lc2[1]-lc2[2],lc2[1]+lc2[2],alpha=0.2,color='C2')
plt.plot(lc2[0],lc2[1],'C2')
plt.figure()
plt.subplot(121)
plt.imshow(cor[1000]-ref,vmin=-10,vmax=10)
plt.colorbar()
plt.subplot(122)
plt.imshow(sub[1000],vmin=-10,vmax=10)
plt.colorbar()
test = np.zeros_like(cor[100])
test[45,45] = 1000
test = cor[100]
basis, coeff_0 = Delta_basis(7)
bds = []
for i in range(len(coeff_0)):
bds += [(0,1)]
coeff_0 *= 0.01
res = minimize(optimize_delta, coeff_0, args=(basis,test,cor[1200]),bounds=bds)
res
from scipy.signal import convolve
k = np.nansum(res.x[:,np.newaxis,np.newaxis]*basis,axis=0)
template = signal.fftconvolve(test, k, mode='same')
np.sum(Kernel)
plt.figure()
plt.imshow(Kernel)
plt.colorbar()
np.nansum(template)
np.nansum(test)
plt.figure()
plt.subplot(121)
plt.imshow(cor[1000]-template,vmin=-10,vmax=10)
plt.colorbar()
plt.subplot(122)
plt.imshow(cor[1000]-cor[100],vmin=-10,vmax=10)
plt.colorbar()
im = cor[1000].copy()
template = signal.fftconvolve(cor[100], Kernel, mode='same')
im[im< 10] = np.nan
res = np.nansum(abs(np.log10(im - np.log10(template))))
a = im/cor[100]-1
m = sigma_clip(a,sigma=5).mask
a[m] = np.nan
plt.figure()
plt.imshow(a)
mask
np.nansum(a)
sigma_clip(im/cor[100]-1)
from astropy.stats import sigma_clip
```
| github_jupyter |
```
from __future__ import print_function
# !pip install tensorflow-gpu
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import matplotlib.pyplot as plt
%matplotlib inline
from keras.models import Sequential
from keras.layers import Dense , Dropout , Lambda, Flatten, Conv2D, MaxPool2D, BatchNormalization, Input,Concatenate
from keras.optimizers import SGD
from sklearn.model_selection import train_test_split
from keras.models import model_from_json
import cv2
import glob
import os
import pickle
#set global parameters
img_rows = 224
img_cols = 224
max_files = -1
read_from_cache = False
!cd ~/sharedfolder/
!git pull
%cd day04/
!ls
Path = '/root/sharedfolder/day04/'
filelist = glob.glob(Path)
filelist.sort()
filelist
def read_image(path,img_rows,img_cols):
img = cv2.imread(path)
return cv2.resize(img, (img_cols, img_rows))
def read_train(path,img_rows,img_cols,max_files):
# img_rows & img_cols set the size of the image in the output
# max files is the maximal number of images to read from each category
# use max_files=-1 to read all images within the train subfolders
X_train = []
y_train = []
counter = 0
print('Read train images')
files = glob.glob(path+'*.JPG')
for fl in files:
flbase = os.path.basename(fl)
img = read_image(fl, img_rows, img_cols)
X_train.append(np.asarray(img))
# y_train.append(j)
counter+=1
if (counter>=max_files)&(max_files>0):
break
return np.array(X_train)#, np.array(y_train)
def cache_data(data, path):
# this is a helper function used to cache data once it was read and preprocessed
if os.path.isdir(os.path.dirname(path)):
file = open(path, 'wb')
pickle.dump(data, file)
file.close()
else:
print('Directory doesnt exists')
def restore_data(path):
# this is a helper function used to restore cached data
data = dict()
if os.path.isfile(path):
file = open(path, 'rb')
data = pickle.load(file)
return data
def save_model(model,filename):
# this is a helper function used to save a keras NN model architecture and weights
json_string = model.to_json()
if not os.path.isdir('cache'):
os.mkdir('cache')
open(os.path.join('cache', filename+'_architecture.json'), 'w').write(json_string)
model.save_weights(os.path.join('cache', filename+'_model_weights.h5'), overwrite=True)
def read_model(filename):
# this is a helper function used to restore a keras NN model architecture and weights
model = model_from_json(open(os.path.join('cache', filename+'_architecture.json')).read())
model.load_weights(os.path.join('cache', filename+'_model_weights.h5'))
return model
tr_data = read_train(Path,224,224,-1)
y_train = pd.read_csv(Path+'card_files_labels.csv')
from keras.utils.np_utils import to_categorical
ids = y_train.card_file
y_train.drop('card_file',inplace=True,axis=1)
OHE_y_train = (y_train)
tr_data.shape
y_train
OHE_y_train.shape
plt.imshow(tr_data[1])
model= Sequential()
model.add(Conv2D(16,(2,2),activation='relu',input_shape=(img_rows,img_cols,3)))
model.add(Conv2D(32,(2,2),activation='relu'))
model.add(MaxPool2D(pool_size=(2,2),padding='valid'))
model.add(Conv2D(32,(2,2),activation='relu'))
model.add(Conv2D(16,(2,2),activation='relu'))
model.add(MaxPool2D(pool_size=(2,2),padding='valid'))
model.add(Flatten())
model.add(Dense(57, activation='sigmoid'))
model.summary()
model.compile(optimizer='adadelta', loss='binary_crossentropy', metrics=['accuracy'])
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import ReduceLROnPlateau
rlrop = ReduceLROnPlateau(factor=0.3)
datagen = ImageDataGenerator(
horizontal_flip=True,
rescale=0.5,
shear_range=0.1,
zoom_range=0.4,
rotation_range=360,
width_shift_range=0.1,
height_shift_range=0.1
)
model.fit_generator(datagen.flow(tr_data, OHE_y_train, batch_size=8,save_to_dir='/root/sharedfolder/double/gen_imgs/'),callbacks=[rlrop],
validation_data=datagen.flow(tr_data,OHE_y_train),
steps_per_epoch=50,validation_steps = 10, epochs=4,verbose=2)
cd /root/sharedfolder/double/single\ images/
ls
single_tr_data = read_train('/root/sharedfolder/double/single images/',80,80,-1)
sngl_imgs = [x.split('/')[-1].split('.')[0] for x in glob.glob('/root/sharedfolder/double/single images/'+'*.JPG')]
# sngl_imgs
np.setdiff1d(np.array(y_train.columns),sngl_imgs)
plt.imshow(single_tr_data[0])
from PIL import Image
background = Image.open('/root/sharedfolder/double/gen_imgs/blank.jpg', 'r')
img = Image.open('/root/sharedfolder/double/single images/cat.JPG', 'r')
img_w, img_h = img.size
bg_w, bg_h = background.size
offset = ((bg_w - img_w) / 2, (bg_h - img_h) / 2)
# img = img.rotate(45,resample=Image.NEAREST)
background.paste(img, offset)
background.save('/root/sharedfolder/double/gen_imgs/out.png')
gen = cv2.imread('/root/sharedfolder/double/gen_imgs/out.png')
plt.imshow(gen)
ls /root/sharedfolder/double/gen_imgs/
rm /root/sharedfolder/double/gen_imgs/*.png
img_rows,img_cols = 80,80
model= Sequential()
model.add(Conv2D(16,(2,2),activation='relu',input_shape=(img_rows,img_cols,3)))
model.add(Flatten())
model.add(Dense(57, activation='sigmoid'))
model.summary()
model.compile(optimizer='adadelta', loss='binary_crossentropy', metrics=['accuracy'])
singles_datagen = ImageDataGenerator(
horizontal_flip=True,
rotation_range=360,
width_shift_range=0.1,
height_shift_range=0.1
)
model.fit_generator(singles_datagen.flow(single_tr_data[:55], OHE_y_train, batch_size=8,save_to_dir='/root/sharedfolder/double/gen_imgs/'),callbacks=[rlrop],
validation_data=singles_datagen.flow(single_tr_data[:55],OHE_y_train),
steps_per_epoch=50,validation_steps = 10, epochs=4,verbose=2)
ls
gen = cv2.imread('/root/sharedfolder/double/gen_imgs/_1_9894.png')
plt.imshow(gen)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/kipsangchepesa/Python-Roulette/blob/master/How_does_land_surface_cover_affect_surface_temperature.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Defining the question.
How does land surface cover affect surface temperature?
##Dataset provided is for Reading between 1962 and 2010 with two land covers:
• 100% Broad leaf trees
• 100% Bare soil
and it can be downloaded from the following link:
a).[Broaad leaf trees](https:///www.met.reading.ac.uk/~swrmethn/python_teaching/assignment_data/baresoil_temperature_1962_2010.nc )
b).[Bare soil](https://www.met.reading.ac.uk/~swrmethn/python_teaching/assignment_data/broadleaftree_temperature_1962_2010.nc)
##Metrics for success
1. To explore the interaction between climate, land cover and surface temperature using a land surface model by using output from the JULES land surface model.
2. To make sure that the configuration of JULES that produced the results we are using computes the energy and water balances at the land surface, based on prescribed weather and land cover.
##Importing the libraries
```
pip install rioxarray
pip install nc_time_axis
pip install cftime
import numpy as np
import pandas as pd
import cftime
import nc_time_axis
import rioxarray
import xarray
import matplotlib.pyplot as plt
```
##Task 1
Once you have downloaded the data, write a series of functions for reading, plotting and comparing time series of surface temperature for each land cover type. How does surface temperature vary with land cover? You should consider displaying your results in a variety of ways – for example histograms, time series and some simple statistics.
Present your results as a short description and interpretation (maximum one page) plus four plots. You should also submit your Python code file for assessment.
```
#fh becomes the file handle of the open netCDF file,
#and the ‘r’ denotes that we want to open the file in read only mode.
baresoil = '/content/baresoil_temperature_1962_2010.nc'
broadleaf ='/content/broadleaftree_temperature_1962_2010.nc'
def ncreader(data):
xr=rioxarray.open_rasterio(data)
xr.attrs['units']='mm'
xr_dimens=xr.dims
print(xr)
print("The dimensions are\n"+str(xr_dimens))
#calling the function to read the baresoil nc file land surface
ncreader(baresoil)
#Reading broadleaf using our user define function
ncreader(broadleaf)
print(xr[:, :10, :10])
```
##plotting
```
xr[:200, :10, :10].plot()
xr[:400, :15, :15].plot()
xr = xr.where(xr != xr.attrs['missing_value'])
xr[120, :20, :20].plot()
xr = xr.where(xr != xr.attrs['missing_value'])
xr[100, :, :].plot()
```
##Task 2
Write a set of functions that will calculate and compare the mean seasonal cycle of temperature for each land cover type (i.e., the average over the years of temperature for each calendar month). Describe your results using plots and brief text.
Present your results in the form of a short description and interpretation (no more than one page) plus no more than four plots. You should also submit your code file for assessment.
```
xr.mean()
```
To get the actual value we’ll need to multiple by the scale factor.
```
print(xr.attrs['scale_factor'])
#It will be the same since scale factor in baresoil data is 1,0 as seen above
xr.mean()* xr.attrs['scale_factor']
```
##Calculating and Plotting Statistics
```
xr.mean(dim='y').plot()
```
Total annual temperature
```
(xr.sum(dim='y') * xr.attrs['scale_factor']).plot()
```
| github_jupyter |
# Домашнее задание. Титаник
В этом домашнем задании вам предлагается решить задачу предсказания выживших пассажиров Титаника. Эту задачу машинного обучения рано или поздно решает, наверное, любой, кто обучается анализу данных. Информация о датасете доступна по <a href=https://www.kaggle.com/c/titanic.>ссылке</a>.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
```
## Загрузка данных
Загрузим датасет Titanic из приложенных файлов. В первом файле находится информация о пассажирах, во втором -- информация о том, выжил пассажир или нет.
Если вы запускаете ноутбук на Google Colab, самым простым способом загрузить данные в ноутбук будет примонтировать к Colab ваш Google Drive. Для этого сначала положите в свою корневую директорию Google Drive файлы, приложенные к этому ноутбуку, а затем выполните три закоментированные строчки в клетке ниже. <a href=https://www.machinelearningmastery.ru/downloading-datasets-into-google-drive-via-google-colab-bcb1b30b0166/>Подробная инструкция по работе с Google Drive + Google Colab</a>
```
#from google.colab import drive
#drive.mount('/content/gdrive/')
#data = pd.read_csv('/content/gdrive/My Drive/titanic_data.csv, index_col='PassengerId')
#Если запускаете ноутбук локально:
data = pd.read_csv('/content/titanic_data.csv', index_col='PassengerId')
basic_features = data.columns
y = pd.read_csv('titanic_surv.csv')
y.index = data.index
print(f'Всего {len(data)} пассажиров в выборке')
data.head()
y.head()
data = data.join(y)
data.head()
```
## Исследование датасета
### Задание 1 (1 балл)
Опишите датасет. Сколько в нём мужчин, сколько женщин? Посчитайте распределение по классам пассажиров. Используйте функцию ``pd.Series.value_counts``. <a href=https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.value_counts.html>Пример использования</a>
Посчитайте долю выживших мужчин к общему количеству мужчин, а также женщин к общему количеству женщин. Сделайте выводы.
Проведите схожую аналитику по возрасту выживших и по классу выживших.
Вам поможет функция ``plt.hist()``. <a href=https://matplotlib.org/3.3.1/gallery/pyplots/pyplot_text.html#sphx-glr-gallery-pyplots-pyplot-text-py> Пример использования </a>
```
import seaborn as sns
gender = data["Sex"].value_counts()
plt.figure()
sns.barplot(gender.index, gender.values, alpha=0.8)
plt.title('Gender distribution')
plt.ylabel('Number', fontsize=12)
plt.xlabel('Gender', fontsize=12)
plt.show()
Pclass = data["Pclass"].value_counts()
plt.figure()
sns.barplot(Pclass.index, Pclass.values, alpha=0.8)
plt.title('Class distribution')
plt.ylabel('Number', fontsize=12)
plt.xlabel('Class', fontsize=12)
plt.show()
cross = pd.crosstab(data["Sex"], data["Survived"])
print(f"Ratio survived female : {cross[1].values[0]/(cross[0].values[0] + cross[1].values[0])}")
print(f"Ratio survived male : {cross[1].values[1]/(cross[0].values[1] + cross[1].values[1])}")
cross
sns.countplot(x="Sex", hue="Survived", data=data)
cross_class = pd.crosstab(data["Pclass"], data["Survived"])
print(f"Ratio survived Pclass 1 : {cross_class[1].values[0]/(cross_class[0].values[0] + cross_class[1].values[0])}")
print(f"Ratio survived Pclass 2 : {cross_class[1].values[1]/(cross_class[0].values[1] + cross_class[1].values[1])}")
print(f"Ratio survived Pclass 3 : {cross_class[1].values[2]/(cross_class[0].values[2] + cross_class[1].values[2])}")
cross_class
sns.countplot(x="Pclass", hue="Survived", data=data)
```
## Заполнение пропусков в данных
В данных имеются пропуски в трёх колонках:
```
data.columns[data.isna().any()].tolist()
```
Для простоты заполним все пропуски в категориальных колонках новым классом "0", а в числовой колонке Age --- медианным значением.
```
data.loc[:, ['Cabin', 'Embarked']] = data.loc[:, ['Cabin', 'Embarked']].fillna('0')
data['Age'] = data['Age'].fillna(data['Age'].median())
```
## Подготовка плана тестирования
### Задание 2. (0 баллов)
Разделите выборку на обучающее и тестовое множество в соотношении 70:30.
```
from sklearn.model_selection import train_test_split
data_train, data_test = train_test_split(data, test_size=0.3)
```
## Выделение новых признаков
Теперь отложим тестовые данные и приступим к моделированию. Внимательно изучите данные. Можете ли вы выделить признаки, которые не указаны явно в таблице?
### Задание 3 (1 балл)
Сформируйте по крайней мере один новый признак и объясните ваш выбор. Пример признака сформирован за вас.
Указания:
- Пассажиров можно поделить на несколько классов по их именам.
- Различных номеров кают слишком много. Но буквы в номерах кают указывают на их местоположение. (***Этот признак сформирован в примере***)
- Возможно, имеет смысл отделить мальчиков от мужчин. Отделять девочек от женщин может быть не так важно
- Другие идеи для признаков можно почерпнуть на форуме в обсуждении задачи на kaggle.com.
```
data
data['Cabin']
def get_cabin_letter(row):
return row['Cabin'][0]
data_train['cabin_type'] = data.apply(get_cabin_letter, axis=1)
```
Сюда добавьте описание вашего нового признака
```
def title(row): #rename this function appropriately
return row['Name'].split(",")[1].split(".")[0]
data_train['title'] = data_train.apply(title, axis=1)
```
## Кодирование категориальных признаков и обработка данных
Все признаки сейчас делятся на числовые, бинарные, категориальные и текстовые. К текстовым признакам можно отнести имя пассажира, номер билета и номер каюты: значения этих признаков уникальны почти для всех пассажиров, и простого способа использовать их в модели не существует.
### Задание 4 (0 баллов)
Пока используя только изначальные признаки, перекодируйте категориальные признаки функцией ``pd.get_dummies``. Естественно, имена пассажиров и номера билетов и кают кодировать не стоит, поскольку все значения этих признаков уникальные.
```
sex_dummies = pd.get_dummies(data_train.Sex, prefix='Sex')
embarked_dummies = pd.get_dummies(data_train.Embarked, prefix='Embarked')
class_dummies = pd.get_dummies(data_train.Pclass, prefix='PClass')
cabin_dummies = pd.get_dummies(data_train.cabin_type, prefix='cabin_type')
title_dummies = pd.get_dummies(data_train.title, prefix='title')
df_train = pd.concat([data_train, sex_dummies, class_dummies, cabin_dummies, title_dummies, embarked_dummies], axis=1)
df_train.head(5)
y = df_train["Survived"]
df_train.drop(columns=["Pclass", "Survived", "Name", "Sex",
"Ticket", "Cabin", "title", "Embarked",
"cabin_type"],
inplace=True)
df_train.head(5)
X_train, X_test, y_train, y_test = train_test_split(df_train, y, test_size=0.3)
```
## Обучение baseline-модели
### Задание 5 (1 балл)
Обучите одну из простых известных вам моделей. Измерьте качество полученной модели на кросс-валидации (используйте только обучающую выборку!)
```
from xgboost import XGBClassifier
from sklearn.metrics import f1_score
train_data_baseline = X_train[["Age", "SibSp", "Fare"]]
test_data_baseline = X_test[["Age", "SibSp", "Fare"]]
modelXGB = XGBClassifier()
modelXGB.fit(train_data_baseline, y_train)
preds = modelXGB.predict(test_data_baseline)
f1 = f1_score(y_test, preds)
print(f"Score is {f1}")
```
### Задание 6 (1 балл)
Добавьте в модель два новых сгенерированных признака. Если нужно, добавьте признаки, сгенерированные ohe-hot кодированием. Обучите ту же самую модель на расширенном множестве признаков. Улучшилось ли качество предсказания на кросс-валидации?
```
train_data_baseline = X_train[["Age", "SibSp", "Fare", "Sex_female", "Sex_male"]]
test_data_baseline = X_test[["Age", "SibSp", "Fare", "Sex_female", "Sex_male"]]
modelXGB = XGBClassifier()
modelXGB.fit(train_data_baseline, y_train)
preds = modelXGB.predict(test_data_baseline)
f1 = f1_score(y_test, preds)
print(f"Score is {f1}. It is higher!")
```
## Дополнительное моделирование
### Задание 7 (2 балла)
Теперь более серьёзно подойдём к моделированию. Попробуйте несколько алгоритмов из тех, что мы проходили в курсе. Вам помогут ноутбуки с линейными алгоритмами и выбором модели. Хотя бы для одного алгоритма проведите подбор оптимального гиперпараметра.
```
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import roc_auc_score, roc_curve
def search(X, y, model, param_name, grid, draw=True):
parameters = {param_name: grid}
CV_model = GridSearchCV(estimator=model, param_grid=parameters, scoring='roc_auc', n_jobs=-1)
CV_model.fit(X, y)
means = CV_model.cv_results_['mean_test_score']
error = CV_model.cv_results_['std_test_score']
if draw:
plt.figure(figsize=(15,8))
plt.title('choose ' + param_name)
plt.plot(grid, means, label='mean values of score')
plt.fill_between(grid, means - 2 * error, means + 2 * error, color='green', label='deviation area between errors')
plt.legend()
plt.xlabel('parameter')
plt.ylabel('roc_auc')
plt.xticks(np.arange(4, 25, step=1))
plt.show()
return means, error
models = [KNeighborsClassifier(n_jobs=-1)]
param_names = ['n_neighbors']
grids = [np.array(np.linspace(4, 25, 10), dtype='int')]
for model, param_name, grid, param_scale in zip(models,
param_names,
grids,
param_scales):
search(X_train, y_train, model, param_name, grid, param_scale)
```
## Результаты моделирования
### Задание 8 (1 балл)
Измерьте качество итоговой модели на кросс-валидации. Выполните предсказание на тестовом множестве и сохраните их в переменную ``y_test``. Измерьте итоговое качество на тестовом множестве.
```
def search(X, y, model, param_name, grid, draw=True):
parameters = {param_name: grid}
CV_model = GridSearchCV(estimator=model, param_grid=parameters, cv=5, scoring='roc_auc', n_jobs=-1)
CV_model.fit(X, y)
means = CV_model.cv_results_['mean_test_score']
error = CV_model.cv_results_['std_test_score']
if draw:
plt.figure(figsize=(15,8))
plt.title('choose ' + param_name)
plt.plot(grid, means, label='mean values of score')
plt.fill_between(grid, means - 2 * error, means + 2 * error, color='green', label='deviation area between errors')
plt.legend()
plt.xlabel('parameter')
plt.ylabel('roc_auc')
plt.xticks(np.arange(4, 25, step=1))
plt.show()
return means, error
models = [KNeighborsClassifier(n_jobs=-1)]
param_names = ['n_neighbors']
grids = [np.array(np.linspace(4, 25, 10), dtype='int')]
param_scales = ['ordinary']
for model, param_name, grid, param_scale in zip(models,
param_names,
grids,
param_scales):
search(X_train, y_train, model, param_name, grid, param_scale)
def plot_roc_curve(model, X_train, X_test, y_train, y_test):
y_train_proba = model.predict_proba(X_train)[:, 1]
y_test_proba = model.predict_proba(X_test)[:, 1]
plt.figure(figsize=(12,10))
print(f'Train roc-auc: {roc_auc_score(y_train, y_train_proba)}')
print(f'Test roc-auc: {roc_auc_score(y_test, y_test_proba)}')
plt.plot(*roc_curve(y_train, y_train_proba)[:2], label='train roc-curve')
plt.plot(*roc_curve(y_test, y_test_proba)[:2], label='test roc-curve')
plt.plot([0,1], [0,1], linestyle='--', color='black')
plt.grid(True)
plt.legend()
plt.show()
model = KNeighborsClassifier(n_neighbors=13, n_jobs=-1).fit(X_train, y_train)
plot_roc_curve(model, X_train, X_test, y_train, y_test)
```
## Выводы
### Задание 9 (3 балла)
Сделайте выводы. Какие из идей сработали? Какие оказались лишними?
Gender is one of the most important feature, which improves predictions significantly, also Age gives increment as well. But some features do not contain any usefull information such as Name and Ticket.
```
```
| github_jupyter |
# T5 for Cross-Language Plagiarism Detection
Author: João Phillipe Cardenuto
In this notebook we implement a model regarding the Detailed Analysis of the CLPD.
# Import Libraries
```
! pip install -q pytorch-lightning
! pip install -q transformers
# Mount drive
from google.colab import drive
drive.mount('/content/drive',force_remount=True)
# Comum libraries
import os
import random
from typing import Dict
from typing import List
import numpy as np
import pandas as pd
import re
from argparse import Namespace
from tqdm.notebook import trange, tqdm_notebook
# Dataset
import sys
sys.path.insert(0, "/work/src/DataloaderCLPD/")
from LoadDataset import *
# Torch
import torch
from torch.utils.data import DataLoader
from torch.utils.data import Dataset
import torch.nn.functional as F
from torch.optim.lr_scheduler import StepLR
# HugginFace
from transformers import T5Tokenizer, T5ForConditionalGeneration
# Sklearn
from sklearn.metrics import accuracy_score, f1_score
from sklearn.model_selection import train_test_split
# https://scikit-learn.org/stable/modules/generated/sklearn.metrics.confusion_matrix.html
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
# Tersorflow
import tensorboard
%load_ext tensorboard
# Lightning
import pytorch_lightning as pl
from pytorch_lightning.callbacks import EarlyStopping, ModelCheckpoint
from pytorch_lightning.loggers import TensorBoardLogger
# Setup seeds
seed = 0
np.random.seed(seed)
torch.manual_seed(seed)
random.seed(seed)
if torch.cuda.is_available():
dev = "cuda:0"
else:
dev = "cpu"
device = torch.device(dev)
print("Device",dev)
if "cuda" in dev:
print("GPU: ", torch.cuda.get_device_name(0))
# Loading T5 Tokenizer with portuguese chars
port_tokenizer = T5Tokenizer.from_pretrained('t5-base')
extra_tokens = ['Ç','ç','À' , 'È' , 'Ì' , 'Ò' , 'Ù' , 'à' , 'è' , 'ì' , 'ò' , 'ù' , 'Á' , 'É' , 'Í' , 'Ó' , 'Ú' , 'á' , 'é' , 'í' , 'ó' , 'ú' , 'Â' , 'Ê' , 'Î' , 'Ô' , 'Û' , 'â' , 'ê' , 'î' , 'ô' , 'û' , 'Ã' ,'Ẽ', 'Õ' , 'ã', 'ẽ','õ' , 'Ë', 'ä' , 'ë' , 'ï' , 'ö' , 'ü']
new_tokens = {}
for i in extra_tokens: print(f'({i},{port_tokenizer.decode(port_tokenizer.encode(i))})',end=", ")
print("\n","*-"*10,"New Tokens","*-"*10)
for i in extra_tokens:
# Add_tokens return 0 if token alredy exist, and 1 if It doesnt.
if port_tokenizer.add_tokens(i):
print(f"{i},{port_tokenizer.encode(text=i,add_special_tokens=False,)}", end=" |")
new_tokens[i] = port_tokenizer.encode(text=i,add_special_tokens=False)[0]
```
## Loading Data
Using LoadDataset module to load capes dataset
```
# We using sample size of x it represent x*(2*N_negatives + 2) = x*(ENG + PT + ENG_NEGATIVE_1 ... + ENG_NEGATIVE_N +
# PT_NEGATIVE_1 + ... + PT_NEGATIVE_N)
capes_dataset = CLPDDataset(name='capes',data_type='train',sample_size=1000,val_size=0.2,n_negatives=1)
train_capes, val_capes = capes_dataset.get_organized_data(tokenizer=port_tokenizer,tokenizer_type='t5')
# Samples
print(train_capes[0])
print("Number of Samples:", len(train_capes.pairs))
dataloader_debug = DataLoader(train_capes, batch_size=10, shuffle=True,
num_workers=0)
token_ids, attention_mask, token_type_ids, labels, _ = next(iter(dataloader_debug))
print('token_ids:\n', token_ids)
print('token_type_ids:\n', token_type_ids)
print('attention_mask:\n', attention_mask)
print('labels:\n', labels)
print('token_ids.shape:', token_ids.shape)
print('token_type_ids.shape:', token_type_ids.shape)
print('attention_mask.shape:', attention_mask.shape)
print('labels.shape:', labels.shape)
batch_size = 2
# train_dataloader = DataLoader(dataset_train, batch_size=batch_size,
# shuffle=True, num_workers=4)
val_dataloader = DataLoader(val_capes, batch_size=batch_size, shuffle=True,
num_workers=4)
test_dataloader = DataLoader(val_capes, batch_size=batch_size, shuffle=False,
num_workers=4)
# test_dataloader = DataLoader(dataset_test, batch_size=batch_size,
# shuffle=False, num_workers=4)
label
port_tokenizer.decode(a[0])
port_tokenizer.encode(f"{0} {port_tokenizer.eos_token}",max_length=3, pad_to_max_length=True)
[valid_prediction(a[index],label[index])
for index in range(len(a))]
a = mode2l(input_ids=input_ids.to(device), attention_mask=attention_mask.to(device),
lm_labels=labels.to(device))[0]
```
## T5-Model with Pytorch Lightning
```
def valid_prediction(pred,label):
"""
Decode prediction and label.
Return ( prediction, label) if decode(pred) in {0,1},
otherwise return (not label, label)
"""
text_result = port_tokenizer.decode(pred)
label = port_tokenizer.decode(label)
# Check if string is numeric
if text_result.replace('.','',1).isnumeric():
value = float(text_result)
if value == 1 or value == 0:
return (int(value) , int(label))
# Return a different number from the label
return (int(not int(label)), int(label))
class T5Finetuner(pl.LightningModule):
def __init__(self, hparams,train_dataloader,val_dataloader,test_dataloader):
super(T5Finetuner, self).__init__()
#Hiperparameters
self.hparams = hparams
self.experiment_name = f"{self.hparams.experiment_name}_{self.hparams.version}"
# Dataloaders
self._train_dataloader = train_dataloader
self._val_dataloader = val_dataloader
self._test_dataloader = test_dataloader
# Learnning Rate and Loss Function
self.learning_rate = hparams.learning_rate
self.lossfunc = torch.nn.CrossEntropyLoss()
# Optimizer
self.optimizer = self.hparams.optimizer
self.target_max_length = self.hparams.target_max_length
# Retrieve model from Huggingface
self.model = T5ForConditionalGeneration.from_pretrained('t5-base').to(device)
def forward(self, input_ids, attention_mask, labels=None):
# If labels are None, It will return a loss and a logit
# Else it return the predicted logits for each sentence
if self.training:
# Ref https://huggingface.co/transformers/model_doc/t5.html#training
loss = self.model(input_ids=input_ids,
attention_mask=attention_mask,
lm_labels=labels)[0]
return loss
else:
# REF https://huggingface.co/transformers/main_classes/model.html?highlight=generate#transformers.PreTrainedModel.generate
predicted_token_ids = self.model.generate(
input_ids=input_ids,
max_length=self.target_max_length,
do_sample=False,
)
return predicted_token_ids
def training_step(self, batch, batch_nb):
# batch
input_ids, attention_mask, _, label,_ = batch
# fwd
loss = self(input_ids=input_ids.to(device),
attention_mask=attention_mask.to(device),
labels=label.to(device))
# logs
tensorboard_logs = {'train_loss': loss.item()}
return {'loss': loss, 'log': tensorboard_logs}
def validation_step(self, batch, batch_nb):
# batch
input_ids, attention_mask, _, labels,_ = batch
# fwd
predicted_token_ids = self(input_ids.to(device), attention_mask=None,)
pred_true_decoded = [valid_prediction(predicted_token_ids[index],labels[index])
for index in range(len(predicted_token_ids))]
y_pred = [y[0] for y in pred_true_decoded]
y_true = [y[1] for y in pred_true_decoded]
return {'y_pred': y_pred, 'y_true': y_true}
def validation_epoch_end(self, outputs):
y_true = np.array([ y for x in outputs for y in x['y_true'] ])
y_pred = np.array([ y for x in outputs for y in x['y_pred'] ])
val_f1 = f1_score(y_pred=y_pred, y_true=y_true)
val_f1 = torch.tensor(val_f1)
tensorboard_logs = {'val_f1': val_f1 }
return {'val_f1': val_f1,
'progress_bar': tensorboard_logs, "log": tensorboard_logs}
def test_step(self, batch, batch_nb):
input_ids, attention_mask, _ , labels, pairs = batch
predicted_token_ids = self(input_ids.to(device), attention_mask=None)
pred_true_decoded = [valid_prediction(predicted_token_ids[index],labels[index])
for index in range(len(predicted_token_ids))]
y_pred = [y[0] for y in pred_true_decoded]
y_true = [y[1] for y in pred_true_decoded]
return {'pairs': pairs, 'y_true': y_true, 'y_pred':y_pred }
def test_epoch_end(self, outputs):
pairs = [pair for x in outputs for pair in x['pairs']]
y_true = np.array([ y for x in outputs for y in x['y_true'] ])
y_pred = np.array([ y for x in outputs for y in x['y_pred'] ])
# Write failure on file
with open (f"FAILURE_{self.experiment_name}.txt", 'w') as file:
for index,pair in enumerate(pairs):
if y_true[index] != y_pred[index]:
file.write("="*50+f"\n[Y_TRUE={y_true[index]} != Y_PRED={y_pred[index]}]\n"+pair \
+'\n'+"="*50+'\n')
print("CONFUSION MATRIX:")
print(confusion_matrix(y_true=y_true, y_pred=y_pred))
print("SKLEARN REPORT")
print(classification_report(y_true=y_true, y_pred=y_pred))
test_f1 = f1_score(y_pred=y_pred, y_true=y_true)
tensorboard_logs = {'test_f1': test_f1}
return {'test_f1': test_f1, 'log': tensorboard_logs,
'progress_bar': tensorboard_logs}
def configure_optimizers(self):
optimizer = self.optimizer(
[p for p in self.parameters() if p.requires_grad],
lr=self.learning_rate)
scheduler = StepLR(optimizer, step_size=self.hparams.steplr_epochs, gamma=self.hparams.scheduling_factor)
return [optimizer], [scheduler]
def train_dataloader(self):
return self._train_dataloader
def val_dataloader(self):
return self._val_dataloader
def test_dataloader(self):
return self._test_dataloader
hyperparameters = {"experiment_name": "T5-DEBUG",
"max_epochs": 2,
"patience": 4,
"optimizer": torch.optim.Adam,
"target_max_length": 10,
"scheduling_factor": 0.8,
"learning_rate": 1e-5,
"steplr_epochs":4,
}
model = T5Finetuner(hparams=Namespace(**hyperparameters),
train_dataloader=val_dataloader,
val_dataloader=test_dataloader,
test_dataloader=test_dataloader)
```
## Number of Parameter
```
sum([torch.tensor(x.size()).prod() for x in model.parameters() if x.requires_grad]) # trainable parameters
```
## Fast dev run
```
trainer = pl.Trainer(gpus=1,
logger=False,
checkpoint_callback=False, # Disable checkpoint saving.
fast_dev_run=True,
amp_level='O2', use_amp=False
)
trainer.fit(model)
trainer.test(model)
del model
```
## Overfit on a Batch
We notice that easily the model can overfit on a batch
```
hyperparameters = {"experiment_name": "T5CLPD",
"optimizer": torch.optim.Adam,
"target_max_length": 3,
"max_epochs": 5,
"patience": 4,
"steplr_epochs":5,
"scheduling_factor": 0.95,
"learning_rate": 6e-5,
"max_length": 200
}
trainer = pl.Trainer(gpus=1,
logger=False,
max_epochs=hyperparameters['max_epochs'],
check_val_every_n_epoch=5,
checkpoint_callback=False, # Disable checkpoint saving
overfit_pct=0.5,
amp_level='O2', use_amp=False)
model = T5Finetuner(hparams=Namespace(**hyperparameters),
train_dataloader=val_dataloader,
val_dataloader=test_dataloader,
test_dataloader=test_dataloader)
trainer.fit(model)
trainer.test(model)
# del model
# del trainer
train_capes[0]
```
## Training
```
# Training will perform a cross-dataset.
max_length = 200
scielo_dataset = CLPDDataset(name='scielo',data_type='train',sample_size=100000,val_size=0.2,max_length=200,n_negatives=1)
scielo_test = CLPDDataset(name='scielo',data_type='test',n_negatives=1,max_length=200)
# T5 Tokenizer with portuguese chars
tokenizer = port_tokenizer
# Traning data
train_scielo, val_scielo = scielo_dataset.get_organized_data(tokenizer=tokenizer,tokenizer_type='t5')
test_scielo = scielo_test.get_organized_data(tokenizer=tokenizer,tokenizer_type='t5')
len(test_scielo)
#------tester-----------#
# DataLoaders #
#-------------------#
batch_size = 32
train_dataloader = DataLoader(train_scielo, batch_size=batch_size,
shuffle=True, num_workers=4)
val_dataloader = DataLoader(val_scielo, batch_size=batch_size, shuffle=False,
num_workers=4)
test_dataloader = DataLoader(test_scielo, batch_size=batch_size,
shuffle=False, num_workers=4)
# Hiperparameters
hyperparameters = {"experiment_name": "T5-SCIELO",
"optimizer": torch.optim.Adam,
"target_max_length": 3,
"max_epochs": 3,
"patience": 4,
"steplr_epochs":1,
"scheduling_factor": 0.95,
"learning_rate": 6e-5,
"max_length": max_length,
'batch_size': batch_size
}
#------------------------------#
# Checkpoints #
#------------------------------#
log_path = 'logs'
ckpt_path = os.path.join(log_path, hyperparameters["experiment_name"], "-{epoch}-{val_f1:.2f}")
checkpoint_callback = ModelCheckpoint(prefix=hyperparameters["experiment_name"], # prefixo para nome do checkpoint
filepath=ckpt_path, # path onde será salvo o checkpoint
monitor="val_f1",
mode="max",
save_top_k=2)
# Hard coded
# resume_from_checkpoint = '/content/drive/My Drive/P_IA376E_2020S1/Class-8 BERT/TASK/logs/Electra-400/Electra-400-epoch=37-val_loss=0.18.ckpt'
resume_from_checkpoint= False
# Configuração do Early Stop
early_stop = EarlyStopping(monitor="val_loss",
patience=hyperparameters["patience"],
verbose=False,
mode='min'
)
logger = TensorBoardLogger(hyperparameters["experiment_name"],name='T5' ,version="NEGATIVE_1")
# Lighting Trainer
trainer = pl.Trainer(gpus=1,
logger=logger,
max_epochs=hyperparameters["max_epochs"],
check_val_every_n_epoch=1,
accumulate_grad_batches=5,
checkpoint_callback=checkpoint_callback,
# resume_from_checkpoint=resume_from_checkpoint,
amp_level='O2', use_amp=False)
hparams = Namespace(**hyperparameters)
model = T5Finetuner(hparams=hparams,train_dataloader=train_dataloader,val_dataloader=val_dataloader,test_dataloader=test_dataloader)
# Train
trainer.fit(model)
```
## Test model on Capes dataset
```
trainer.test(model)
```
-----
# Test model on CAPES dataset
```
# T5 tokenizer with portuguese chars
tokenizer = port_tokenizer
max_length = 200
batch_size = 128
scielo_dataset = CLPDDataset(name='capes',data_type='test',n_negatives=1,max_length=200)
scielo_dataset = scielo_dataset.get_organized_data(tokenizer=tokenizer,tokenizer_type='t5')
scielo_dataloader = DataLoader(scielo_dataset, batch_size=batch_size,
shuffle=False, num_workers=4)
# Hiperparameters
hyperparameters = {"experiment_name": "T5-SCIELO",
"version": 'TEST-ON-CAPES',
"optimizer": torch.optim.Adam,
"target_max_length": 3,
"max_epochs": 3,
"patience": 4,
"steplr_epochs":1,
"scheduling_factor": 0.95,
"learning_rate": 6e-5,
"max_length": max_length,
'batch_size': batch_size
}
#------------------------------#
# Checkpoints #
#------------------------------#
log_path = 'logs'
ckpt_path = os.path.join(log_path, hyperparameters["experiment_name"], "-{epoch}-{val_loss:.2f}")
# Resume from checkpoint Hard coded
resume_from_checkpoint= '/work/src/T5/logs/T5-SCIELO/T5-SCIELO-epoch=1-val_f1=0.98.ckpt'
# Logger
logger = TensorBoardLogger(hyperparameters["experiment_name"], name='T5' ,version=hyperparameters['version'])
# Lighting Tester
tester = pl.Trainer(gpus=1,
logger=logger,
resume_from_checkpoint=resume_from_checkpoint,
amp_level='O2', use_amp=False)
hparams = Namespace(**hyperparameters)
model = T5Finetuner(hparams=hparams,train_dataloader=None,val_dataloader=None,test_dataloader=scielo_dataloader)
tester.test(model)
```
-----
## Test on books dataset
```
tokenizer = port_tokenizer
max_length = 200
batch_size = 300
books_dataset = CLPDDataset(name='books',data_type='test')
books_dataset = books_dataset.get_organized_data(tokenizer=tokenizer,tokenizer_type='t5')
books_dataloader = DataLoader(books_dataset, batch_size=batch_size,
shuffle=False, num_workers=4)
# Hiperparameters
hyperparameters = {"experiment_name": "T5-SCIELO",
"version": 'TEST-ON-BOOKS',
"optimizer": torch.optim.Adam,
"target_max_length": 3,
"max_epochs": 3,
"patience": 4,
"steplr_epochs":1,
"scheduling_factor": 0.95,
"learning_rate": 6e-5,
"max_length": max_length,
'batch_size': batch_size
}
#------------------------------#
# Checkpoints #
#------------------------------#
# Resume from checkpoint Hard coded
resume_from_checkpoint= '/work/src/T5/logs/T5-SCIELO/T5-SCIELO-epoch=1-val_f1=0.98.ckpt'
# Logger
logger = TensorBoardLogger(hyperparameters["experiment_name"], name='T5' ,version=hyperparameters['version'])
# Lighting Tester
tester = pl.Trainer(gpus=1,
logger=logger,
resume_from_checkpoint=resume_from_checkpoint,
amp_level='O2', use_amp=False)
hparams = Namespace(**hyperparameters)
model = T5Finetuner(hparams=hparams,train_dataloader=None,val_dataloader=None,test_dataloader=books_dataloader)
tester.test(model)
```
| github_jupyter |
# Bubble Sort
```
def bubble_sort(l):
n = len(l)
# print(n)
# Outer loop. Goes over the whole thing 'n' times
# (because each time, one 'highest' will have moved to the end)
for i in range(n):
swapped = False
# try to bubble the highest one up
for j in range(0, (n-i)-1):
# compare pairs, move higher one up (the highest will always reach the end this way!)
if l[j] > l[j+1]:
l[j], l[j+1] = l[j+1], l[j]
swapped = True
if not swapped:
break
l = [1, 2, 4, 1, 2, 5, 5, 6, 1, 110, 15]
bubble_sort(l)
print(l)
```
# Insertion Sort
```
def insertion_sort(l):
# Go through all elements (except first).
# Call it 'Key'.
# Each time, the key would be 'inserted' in its place.
# At each iteration, stuff less than i would be sorted already.
for i in range(1, len(l)):
key = l[i] # hold this key
# start comparing keys to things on its left!
# stop when less or equal value found (or we reach left end)
j =i-1
while j >= 0 and key < l[j]:
l[j+1] = l[j] # move this to right. Slot left on j
j -= 1
l[j+1] = key # Place key in free slot..(j+1 because we decrement j above)
l = [1, 2, 4, 1, 2, 5, 5, 6, 1, 110, 15]
insertion_sort(l)
print(l)
```
# Selection Sort
```
def selection_sort(l):
n = len(l)
# for each element in the list (starting from left)
for i in range(n):
min_idx = i # find the minimum ...
# ... in the *rest* of the list
for j in range(i+1, n):
if l[j] < l[min_idx]:
min_idx = j
# swap the minimum with current element, now we have (sorted stuff till i)
l[i] , l[min_idx] = l[min_idx], l[i]
l = [1, 2, 4, 1, 2, 5, 5, 6, 1, 110, 15]
selection_sort(l)
print(l)
```
# Qucik Sort
```
import random
def qsort(l, fst, lst):
if fst >= lst: return
i, j = fst, lst
pivot = l[random.randint(fst, lst)]
while i <= j:
while l[i] < pivot: i += 1
while l[j] > pivot: j -= 1
if i <= j:
l[i], l[j] = l[j], l[i]
i, j = i + 1, j-1
qsort(l, fst, j)
qsort(l, i, lst)
l = [1, 2, 4, 1, 2, 5, 5, 6, 1, 110, 15]
qsort(l, 0, len(l)-1)
print(l)
```
# Sorting in the Rel World
# Descending Sorts
```
def selection_sort(l):
n = len(l)
# for each element in the list (starting from left)
for i in range(n):
idx = i # find the replacement ...
# ... in the *rest* of the list
for j in range(i+1, n):
if l[j] > l[idx]: # change to > for descending
idx = j
# swap the replacement with current element, now we have ( sorted stuff till i)
l[i] , l[idx] = l[idx], l[i]
l = [1, 2, 4, 1, 2, 5, 5, 6, 1, 110, 15]
selection_sort(l)
print(l)
```
But there's better way!
```
def less_than(a, b):
return a < b
def selection_sort(l, compare_with):
n = len(l)
# for each element in the list (starting from left)
for i in range(n):
min_idx = i # find the minimum ...
# ... in the *rest* of the list
for j in range(i+1, n):
if compare_with(l[j], l[min_idx]): # now the "comparison" is out-sourced !
min_idx = j
# swap the minimum with current element, now we have ( sorted stuff till i)
l[i] , l[min_idx] = l[min_idx], l[i]
l = [1, 2, 4, 1, 2, 5, 5, 6, 1, 110, 15]
selection_sort(l, less_than)
print(l)
def greater_than(a, b):
return a > b
l = [1, 2, 4, 1, 2, 5, 5, 6, 1, 110, 15]
selection_sort(l, greater_than)
print(l)
```
# Taking it Even Futher
Now we can do all sorts of stuff with this without making a single change to our selcetion sort code.
```
all_tuples = [
(24, 25),
(1, 2),
(2, 4),
(3, 5),
]
def tuple_less_than(a, b):
return sum(a) < sum(b)
def tuple_greater_than(a, b):
return sum(a) > sum(b)
print("Ascending:\t", end="")
selection_sort(all_tuples, tuple_less_than)
print(all_tuples)
print("Descending:\t", end="")
selection_sort(all_tuples, tuple_greater_than)
print(all_tuples)
```
# Sorting in Python
If you have a list of dictionaries -- each representing a student, for instance.
```
d = [
{ 'name': 'Khalid', 'age': 5},
{ 'name': 'Usman', 'age': 12},
{ 'name': 'Ali', 'age': 7},
{ 'name': 'Farooq', 'age': 3},
]
def d_less_than(a, b):
return a['age'] < b['age']
selection_sort(d, d_less_than)
print(d)
def student_age(a):
return a['age']
print(d)
d.sort(key=student_age, reverse=True)
print(d)
```
# Sorting Objects of Custom Classes
```
class Student:
def __init__(self, name, age):
self.name = name
self.age = age
def __str__(self):
return self.name + ': ' + str(self.age)
s1 = Student('Wajid', 5)
s2 = Student('Usman', 7)
s3 = Student('Ali', 3)
s = [s1, s2, s3]
```
You don't even have to give the function a name -- just use an anonymous function like so:
```
for i in s:
print(i)
```
# Anonymous Function
```
s.sort(key=lambda x: x.age) # reverse
for i in s:
print(i)
```
| github_jupyter |
# Lockman SWIRE master catalogue
## Preparation of Spitzer datafusion SERVS data
The Spitzer catalogues were produced by the datafusion team are available in `dmu0_DataFusion-Spitzer`.
Lucia told that the magnitudes are aperture corrected.
In the catalouge, we keep:
- The internal identifier (this one is only in HeDaM data);
- The position;
- The fluxes in aperture 2 (1.9 arcsec);
- The “auto” flux (which seems to be the Kron flux);
- The stellarity in each band
A query of the position in the Spitzer heritage archive show that the SERVS-ELAIS-N1 images were observed in 2009. Let's take this as epoch.
```
from herschelhelp_internal import git_version
print("This notebook was run with herschelhelp_internal version: \n{}".format(git_version()))
%matplotlib inline
#%config InlineBackend.figure_format = 'svg'
import matplotlib.pyplot as plt
plt.rc('figure', figsize=(10, 6))
from collections import OrderedDict
import os
from astropy import units as u
from astropy.coordinates import SkyCoord
from astropy.table import Column, Table
import numpy as np
from herschelhelp_internal.flagging import gaia_flag_column
from herschelhelp_internal.masterlist import nb_astcor_diag_plot, remove_duplicates
from herschelhelp_internal.utils import astrometric_correction, flux_to_mag
OUT_DIR = os.environ.get('TMP_DIR', "./data_tmp")
try:
os.makedirs(OUT_DIR)
except FileExistsError:
pass
RA_COL = "servs_ra"
DEC_COL = "servs_dec"
```
## I - Column selection
```
imported_columns = OrderedDict({
'internal_id': "servs_intid",
'ra_12': "servs_ra",
'dec_12': "servs_dec",
'flux_aper_2_1': "f_ap_servs_irac1",
'fluxerr_aper_2_1': "ferr_ap_servs_irac1",
'flux_auto_1': "f_servs_irac1",
'fluxerr_auto_1': "ferr_servs_irac1",
'class_star_1': "servs_stellarity_irac1",
'flux_aper_2_2': "f_ap_servs_irac2",
'fluxerr_aper_2_2': "ferr_ap_servs_irac2",
'flux_auto_2': "f_servs_irac2",
'fluxerr_auto_2': "ferr_servs_irac2",
'class_star_2': "servs_stellarity_irac2",
})
catalogue = Table.read("../../dmu0/dmu0_DataFusion-Spitzer/data/DF-SERVS_Lockman-SWIRE.fits")[list(imported_columns)]
for column in imported_columns:
catalogue[column].name = imported_columns[column]
epoch = 2009
# Clean table metadata
catalogue.meta = None
# Adding magnitude and band-flag columns
for col in catalogue.colnames:
if col.startswith('f_'):
errcol = "ferr{}".format(col[1:])
magnitude, error = flux_to_mag(
np.array(catalogue[col])/1.e6, np.array(catalogue[errcol])/1.e6)
# Note that some fluxes are 0.
catalogue.add_column(Column(magnitude, name="m{}".format(col[1:])))
catalogue.add_column(Column(error, name="m{}".format(errcol[1:])))
# Band-flag column
if "ap" not in col:
catalogue.add_column(Column(np.zeros(len(catalogue), dtype=bool), name="flag{}".format(col[1:])))
catalogue[:10].show_in_notebook()
```
## II - Removal of duplicated sources
We remove duplicated objects from the input catalogues.
```
SORT_COLS = ['ferr_ap_servs_irac1', 'ferr_ap_servs_irac2']
FLAG_NAME = "servs_flag_cleaned"
nb_orig_sources = len(catalogue)
catalogue = remove_duplicates(catalogue, RA_COL, DEC_COL, sort_col=SORT_COLS,flag_name=FLAG_NAME)
nb_sources = len(catalogue)
print("The initial catalogue had {} sources.".format(nb_orig_sources))
print("The cleaned catalogue has {} sources ({} removed).".format(nb_sources, nb_orig_sources - nb_sources))
print("The cleaned catalogue has {} sources flagged as having been cleaned".format(np.sum(catalogue[FLAG_NAME])))
```
## III - Astrometry correction
We match the astrometry to the Gaia one. We limit the Gaia catalogue to sources with a g band flux between the 30th and the 70th percentile. Some quick tests show that this give the lower dispersion in the results.
```
gaia = Table.read("../../dmu0/dmu0_GAIA/data/GAIA_Lockman-SWIRE.fits")
gaia_coords = SkyCoord(gaia['ra'], gaia['dec'])
nb_astcor_diag_plot(catalogue[RA_COL], catalogue[DEC_COL],
gaia_coords.ra, gaia_coords.dec)
delta_ra, delta_dec = astrometric_correction(
SkyCoord(catalogue[RA_COL], catalogue[DEC_COL]),
gaia_coords
)
print("RA correction: {}".format(delta_ra))
print("Dec correction: {}".format(delta_dec))
catalogue[RA_COL] += delta_ra.to(u.deg)
catalogue[DEC_COL] += delta_dec.to(u.deg)
nb_astcor_diag_plot(catalogue[RA_COL], catalogue[DEC_COL],
gaia_coords.ra, gaia_coords.dec)
```
## IV - Flagging Gaia objects
```
catalogue.add_column(
gaia_flag_column(SkyCoord(catalogue[RA_COL], catalogue[DEC_COL]), epoch, gaia)
)
GAIA_FLAG_NAME = "servs_flag_gaia"
catalogue['flag_gaia'].name = GAIA_FLAG_NAME
print("{} sources flagged.".format(np.sum(catalogue[GAIA_FLAG_NAME] > 0)))
```
## V - Flagging objects near bright stars
## VI - Saving to disk
```
catalogue.write("{}/SERVS.fits".format(OUT_DIR), overwrite=True)
```
| github_jupyter |
---
_You are currently looking at **version 1.0** of this notebook. To download notebooks and datafiles, as well as get help on Jupyter notebooks in the Coursera platform, visit the [Jupyter Notebook FAQ](https://www.coursera.org/learn/python-data-analysis/resources/0dhYG) course resource._
---
# Merging Dataframes
```
import pandas as pd
df = pd.DataFrame([{'Name': 'Chris', 'Item Purchased': 'Sponge', 'Cost': 22.50},
{'Name': 'Kevyn', 'Item Purchased': 'Kitty Litter', 'Cost': 2.50},
{'Name': 'Filip', 'Item Purchased': 'Spoon', 'Cost': 5.00}],
index=['Store 1', 'Store 1', 'Store 2'])
df
df['Date'] = ['December 1', 'January 1', 'mid-May']
df
df['Delivered'] = True
df
df['Feedback'] = ['Positive', None, 'Negative']
df
adf = df.reset_index()
adf['Date'] = pd.Series({0: 'December 1', 2: 'mid-May'})
adf
staff_df = pd.DataFrame([{'Name': 'Kelly', 'Role': 'Director of HR'},
{'Name': 'Sally', 'Role': 'Course liasion'},
{'Name': 'James', 'Role': 'Grader'}])
staff_df = staff_df.set_index('Name')
student_df = pd.DataFrame([{'Name': 'James', 'School': 'Business'},
{'Name': 'Mike', 'School': 'Law'},
{'Name': 'Sally', 'School': 'Engineering'}])
student_df = student_df.set_index('Name')
print(staff_df.head())
print()
print(student_df.head())
pd.merge(staff_df, student_df, how='outer', left_index=True, right_index=True)
pd.merge(staff_df, student_df, how='inner', left_index=True, right_index=True)
pd.merge(staff_df, student_df, how='left', left_index=True, right_index=True)
pd.merge(staff_df, student_df, how='right', left_index=True, right_index=True)
staff_df = staff_df.reset_index()
student_df = student_df.reset_index()
pd.merge(staff_df, student_df, how='left', left_on='Name', right_on='Name')
staff_df = pd.DataFrame([{'Name': 'Kelly', 'Role': 'Director of HR', 'Location': 'State Street'},
{'Name': 'Sally', 'Role': 'Course liasion', 'Location': 'Washington Avenue'},
{'Name': 'James', 'Role': 'Grader', 'Location': 'Washington Avenue'}])
student_df = pd.DataFrame([{'Name': 'James', 'School': 'Business', 'Location': '1024 Billiard Avenue'},
{'Name': 'Mike', 'School': 'Law', 'Location': 'Fraternity House #22'},
{'Name': 'Sally', 'School': 'Engineering', 'Location': '512 Wilson Crescent'}])
pd.merge(staff_df, student_df, how='left', left_on='Name', right_on='Name')
staff_df = pd.DataFrame([{'First Name': 'Kelly', 'Last Name': 'Desjardins', 'Role': 'Director of HR'},
{'First Name': 'Sally', 'Last Name': 'Brooks', 'Role': 'Course liasion'},
{'First Name': 'James', 'Last Name': 'Wilde', 'Role': 'Grader'}])
student_df = pd.DataFrame([{'First Name': 'James', 'Last Name': 'Hammond', 'School': 'Business'},
{'First Name': 'Mike', 'Last Name': 'Smith', 'School': 'Law'},
{'First Name': 'Sally', 'Last Name': 'Brooks', 'School': 'Engineering'}])
staff_df
student_df
pd.merge(staff_df, student_df, how='inner', left_on=['First Name','Last Name'], right_on=['First Name','Last Name'])
```
# Idiomatic Pandas: Making Code Pandorable
```
import pandas as pd
df = pd.read_csv('census.csv')
df
df11=(df.query('SUMLEV==50 & STNAME =="Alabama"')
.dropna()
.set_index(['STNAME','CTYNAME'])
.rename(columns={'ESTIMATESBASE2010': 'Estimates Base 2010'}))
df11
df = df[df['SUMLEV']==50]
df.set_index(['STNAME','CTYNAME'], inplace=True)
df.rename(columns={'ESTIMATESBASE2010': 'Estimates Base 2010'})
import numpy as np
def min_max(row):
data = row[['POPESTIMATE2010',
'POPESTIMATE2011',
'POPESTIMATE2012',
'POPESTIMATE2013',
'POPESTIMATE2014',
'POPESTIMATE2015']]
return pd.Series({'min': np.min(data), 'max': np.max(data)})
df.apply(min_max, axis=1)
import numpy as np
def min_max(row):
data = row[['POPESTIMATE2010',
'POPESTIMATE2011',
'POPESTIMATE2012',
'POPESTIMATE2013',
'POPESTIMATE2014',
'POPESTIMATE2015']]
row['max'] = np.max(data)
row['min'] = np.min(data)
return row
df.apply(min_max, axis=1)
rows = ['POPESTIMATE2010',
'POPESTIMATE2011',
'POPESTIMATE2012',
'POPESTIMATE2013',
'POPESTIMATE2014',
'POPESTIMATE2015']
df.apply(lambda x: np.max(x[rows]), axis=1)
```
# Group by
```
import pandas as pd
import numpy as np
df = pd.read_csv('census.csv')
df = df[df['SUMLEV']==50]
df
%%timeit -n 10
for state in df['STNAME'].unique():
avg = np.average(df['CENSUS2010POP'])
print('Counties in state ' + state + ' have an average population of ' + str(avg))
%%timeit -n 10
for group, frame in df.groupby('STNAME'):
avg = np.average(frame['CENSUS2010POP'])
print('Counties in state ' + group + ' have an average population of ' + str(avg))
df.head()
df = df.set_index('STNAME')
def fun(item):
if item[0]<'M':
return 0
if item[0]<'Q':
return 1
return 2
for group, frame in df.groupby(fun):
print('There are ' + str(len(frame)) + ' records in group ' + str(group) + ' for processing.')
df = pd.read_csv('census.csv')
df = df[df['SUMLEV']==50]
df.groupby('STNAME').agg({'CENSUS2010POP': np.average})
print(type(df.groupby(level=0)['POPESTIMATE2010','POPESTIMATE2011']))
print(type(df.groupby(level=0)['POPESTIMATE2010']))
(df.set_index('STNAME').groupby(level=0)['CENSUS2010POP']
.agg({'size':np.size,'avg': np.average, 'sum': np.sum, 'std':np.std}))
(df.set_index('STNAME').groupby(level=0)['POPESTIMATE2010','POPESTIMATE2011']
.agg({'avg': np.average, 'sum': np.sum}))
(df.set_index('STNAME').groupby(level=0)['POPESTIMATE2010']
.agg({'POPESTIMATE2010': np.average, 'POPESTIMATE2010': np.sum}))
```
# Scales
```
df = pd.DataFrame(['A+', 'A', 'A-', 'B+', 'B', 'B-', 'C+', 'C', 'C-', 'D+', 'D'],
index=['excellent', 'excellent', 'excellent', 'good', 'good', 'good', 'ok', 'ok', 'ok', 'poor', 'poor'])
df.rename(columns={0: 'Grades'}, inplace=True)
df
df['Grades'].astype('category').head()
grades = df['Grades'].astype('category',
categories=['D', 'D+', 'C-', 'C', 'C+', 'B-', 'B', 'B+', 'A-', 'A', 'A+'],
ordered=True)
grades.head()
grades > 'C'
df = pd.read_csv('census.csv')
df = df[df['SUMLEV']==50]
df = df.set_index('STNAME').groupby(level=0)['CENSUS2010POP'].agg({'avg': np.average})
pd.cut(df['avg'],10)
```
# Pivot Tables
```
#http://open.canada.ca/data/en/dataset/98f1a129-f628-4ce4-b24d-6f16bf24dd64
df = pd.read_csv('cars.csv')
df.head()
df.pivot_table(values='(kW)', index='YEAR', columns='Make', aggfunc=np.mean)
df.pivot_table(values='(kW)', index='YEAR', columns='Make', aggfunc=[np.mean,np.min], margins=True)
```
# Date Functionality in Pandas
```
import pandas as pd
import numpy as np
```
### Timestamp
```
pd.Timestamp('9/1/2016 10:05AM')
```
### Period
```
pd.Period('1/2016')
pd.Period('3/5/2016')
```
### DatetimeIndex
```
t1 = pd.Series(list('abc'), [pd.Timestamp('2016-09-01'), pd.Timestamp('2016-09-02'), pd.Timestamp('2016-09-03')])
t1
type(t1.index)
```
### PeriodIndex
```
t2 = pd.Series(list('def'), [pd.Period('2016-09'), pd.Period('2016-10'), pd.Period('2016-11')])
t2
type(t2.index)
```
### Converting to Datetime
```
d1 = ['2 June 2013', 'Aug 29, 2014', '2015-06-26', '7/12/16']
ts3 = pd.DataFrame(np.random.randint(10, 100, (4,2)), index=d1, columns=list('ab'))
ts3
ts3.index = pd.to_datetime(ts3.index)
ts3
pd.to_datetime('4.7.12', dayfirst=True)
```
### Timedeltas
```
pd.Timestamp('9/3/2016')-pd.Timestamp('9/1/2016')
pd.Timestamp('9/2/2016 8:10AM') + pd.Timedelta('12D 3H')
```
### Working with Dates in a Dataframe
```
dates = pd.date_range('10-01-2016', periods=9, freq='2W-SUN')
dates
df = pd.DataFrame({'Count 1': 100 + np.random.randint(-5, 10, 9).cumsum(),
'Count 2': 120 + np.random.randint(-5, 10, 9)}, index=dates)
df
df.index.weekday_name
df.diff()
df.resample('M').mean()
df['2017']
df['2016-12']
df['2016-12':]
df.asfreq('W', method='ffill')
import matplotlib.pyplot as plt
%matplotlib inline
df.plot()
```
| github_jupyter |
<h2 align=center> Principal Component Analysis</h2>
### Task 2: Load the Data and Libraries
---
```
%matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
plt.style.use("ggplot")
plt.rcParams["figure.figsize"] = (12,8)
# data URL: https://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data
iris = pd.read_csv('dataset/iris.data', header=None)
iris.head()
iris.columns = ['sepal_length','sepal_width','petal_length','petal_width','species']
iris.dropna(how='all', inplace=True)
iris.head()
iris.info()
```
### Task 3: Visualize the Data
---
```
sns.scatterplot(x=iris.sepal_length, y=iris.sepal_width, hue=iris.species, style=iris.species)
```
### Task 4: Standardize the Data
---
```
x = iris.iloc[:,0:4].values
y = iris.species.values
from sklearn.preprocessing import StandardScaler
x = StandardScaler().fit_transform(x)
```
### Task 5: Compute the Eigenvectors and Eigenvalues
---
Covariance: $\sigma_{jk} = \frac{1}{n-1}\sum_{i=1}^{N}(x_{ij}-\bar{x_j})(x_{ik}-\bar{x_k})$
Coviance matrix: $Σ = \frac{1}{n-1}((X-\bar{x})^T(X-\bar{x}))$
```
covariance_matrix = np.cov(x.T)
print(covariance_matrix)
```
We can prove this by looking at the covariance matrix. It has the property that it is symmetric. We also constrain the each of the columns (eigenvectors) such that the values sum to one. Thus, they are orthonormal to each other.
Eigendecomposition of the covriance matrix: $Σ = W\wedge W^{-1}$
```
eigen_values, eigen_vectors = np.linalg.eig(covariance_matrix)
print('Eigenvectors: \n', eigen_vectors)
print('Eigenvalues:\n', eigen_values )
```
### Task 6: Singular Value Decomposition (SVD)
---
```
eigen_vec_svd, s,v = np.linalg.svd(x.T)
eigen_vec_svd
```
### Task 7: Picking Principal Components Using the Explained Variance
---
```
for val in eigen_values:
print(val)
variance_explained = [(i/sum(eigen_values))*100 for i in eigen_values]
variance_explained
cumulative_variance_explained = np.cumsum(variance_explained)
cumulative_variance_explained
sns.lineplot(x=[1,2,3,4], y=cumulative_variance_explained)
plt.xlabel('No. of components')
plt.ylabel('Cumulative explained variance')
plt.title('Explained variance vs. no. of components')
plt.show()
```
### Task 8: Project Data Onto Lower-Dimensional Linear Subspace
---
X_pca = X.W
```
eigen_vectors
projection_matrix = (eigen_vectors.T[:][:])[:2].T
print('Projection Matrix:')
print(projection_matrix)
x_pca = x.dot(projection_matrix)
for species in ('Iris-setosa','Iris-versicolor','Iris-virginica'):
sns.scatterplot(x_pca[y==species,0], x_pca[y==species,1])
```
| github_jupyter |
```
from libraries.dynamics import spread_zombie_dynamics as szd
from libraries.dynamics import graph_by_default
from networkx.algorithms.approximation.vertex_cover import min_weighted_vertex_cover
import datetime as dt
import tqdm
import networkx as nx
import dwave_networkx as dnx
import dimod
import scipy.linalg as sc
import copy
import matplotlib.pyplot as plt
import numpy as np
G, pos = graph_by_default(nodes = 210)
ini_date = dt.datetime(year = 2019, month = 8, day = 18)
dynamic = szd(graph = G, INTIAL_DATE = ini_date)
dynamic.graph_pos = pos
graph_2months = 1
for epoch in tqdm.tqdm(range(61)): # Just 20 epochs
dynamic.step() # Run one step in dynamic procedure
if epoch == 60 : graph_2months = dynamic.graph
print(dynamic) # See basic statistics at each iteration
```
# Betweenness Centrality
```
# from collections import OrderedDict
# bet_centrality = nx.betweenness_centrality(graph_2months, normalized = True,
# endpoints = False)
# G is the Karate Social Graph, parameters normalized
# and endpoints ensure whether we normalize the value
# and consider the endpoints respectively.
# d_descending = OrderedDict(sorted(bet_centrality.items(),
# key=lambda kv: kv[1], reverse=True))
# print(dict(d_descending))
```
# Nuclear bombs
```
graph_path = './graph/without_see_nodes_graph_ini_2.gexf'
G = nx.readwrite.gexf.read_gexf(graph_path)
ini_date = dt.datetime(year = 2019, month = 8, day = 18)
dynamic = szd(graph = G, INTIAL_DATE = ini_date)
dynamic.graph_pos = {G.nodes[n]['node_id']:(eval(n)[1],-eval(n)[0]) for n in G.nodes()}
dynamic.load_checkpoint('./checkpoints/szd_18-10-2019.dyn')
fig, ax = plt.subplots(figsize = (5,5))
dynamic.plot_graph(ax = ax)
G = copy.deepcopy(dynamic.graph)
zombiesquantity = nx.classes.function.get_node_attributes(G, 'zombie_pop')
Numberzombies = {i:v for i,v in zombiesquantity.items() if v > 0}
b = {i:v for i,v in Numberzombies.items() if v >= max(Numberzombies.values()) * 0.8}
hist, bins = np.histogram(list(Numberzombies.values()), bins =100)
hist = np.cumsum(hist)
hist
Numberzombies
import plotly.graph_objects as go
#plt.hist(nodes_zombies.values())
fig = go.Figure()
l = list(Numberzombies.values())
fig.add_trace(go.Histogram(x=l, cumulative_enabled=True))
fig.show()
def cells_bombs(G, beta=0.9):
zombiesquantity = nx.classes.function.get_node_attributes(G, 'zombie_pop')
numberzombies = {i:v for i,v in zombiesquantity.items() if v > 0}
hist, bins = np.histogram(list(numberzombies.values()), bins =100)
hist = np.cumsum(hist)
zombies = np.argmin(np.abs(hist - ((hist[-1] - hist[0]) * beta+ hist[0])))
b = {i:v for i,v in numberzombies.items() if v >= bins[zombies]}
return b
cells_bombs(G)
fig, axs = plt.subplots(nrows = 2, ncols = 2, figsize = (12,8))
ax_info = {
"18-10-2019": axs[0,0], "30-10-2019": axs[0,1],
"15-11-2019": axs[1,0], "30-11-2019": axs[1,1],
}
for epoch in tqdm.tqdm(range(61)): # 2 months
current_date = "{0:%d-%m-%Y}".format(dynamic.current_date)
zombiesquantity = nx.classes.function.get_node_attributes(dynamic.graph, 'zombie_pop')
Numberzombies = {i:v for i,v in zombiesquantity.items() if v > 0}
bombcells = {i:v for i,v in Numberzombies.items() if v >= max(Numberzombies.values()) * 0.8}
if current_date in ax_info.keys():
dynamic.plot_graph(ax = ax_info[current_date])
dynamic.save_checkpoint()
print(dynamic) # See basic statistics at each iteration
if current_date == "01-11-2019": break
dynamic.step() # Run one step in dynamic procedure
```
# Highest degree for immunization
```
import time
import operator
G = copy.deepcopy(graph_2months)
T = 4000
#RECORD START TIME
start_time=time.time()
end_time=start_time
delta_time=end_time-start_time
times=[] #list of times when solution is found, tuple=(VC size,delta_time)
# INITIALIZE SOLUTION VC SETS AND FRONTIER SET TO EMPTY SET
OptVC = []
CurVC = []
Frontier = []
neighbor = []
# ESTABLISH INITIAL UPPER BOUND
UpperBound = G.number_of_nodes()
print('Initial UpperBound:', UpperBound)
CurG = G.copy() # make a copy of G
# sort dictionary of degree of nodes to find node with highest degree
v = find_maxdeg(CurG)
#v=(1,0)
# APPEND (V,1,(parent,state)) and (V,0,(parent,state)) TO FRONTIER
Frontier.append((v[0], 0, (-1, -1))) # tuples of node,state,(parent vertex,parent vertex state)
Frontier.append((v[0], 1, (-1, -1)))
# print(Frontier)
while Frontier!=[] and delta_time<T:
(vi,state,parent)=Frontier.pop() #set current node to last element in Frontier
#print('New Iteration(vi,state,parent):', vi, state, parent)
backtrack = False
#print(parent[0])
# print('Neigh',vi,neighbor)
# print('Remaining no of edges',CurG.number_of_edges())
if state == 0: # if vi is not selected, state of all neighbors=1
neighbor = CurG.neighbors(vi) # store all neighbors of vi
for node in list(neighbor):
CurVC.append((node, 1))
CurG.remove_node(node) # node is in VC, remove neighbors from CurG
elif state == 1: # if vi is selected, state of all neighbors=0
# print('curg',CurG.nodes())
CurG.remove_node(vi) # vi is in VC,remove node from G
#print('new curG',CurG.edges())
else:
pass
CurVC.append((vi, state))
CurVC_size = VC_Size(CurVC)
#print('CurVC Size', CurVC_size)
# print(CurG.number_of_edges())
# print(CurG.edges())
# print('no of edges',CurG.number_of_edges())
if CurG.number_of_edges() == 0: # end of exploring, solution found
#print('In FIRST IF STATEMENT')
if CurVC_size < UpperBound:
OptVC = CurVC.copy()
#print('OPTIMUM:', OptVC)
print('Current Opt VC size', CurVC_size)
UpperBound = CurVC_size
#print('New VC:',OptVC)
times.append((CurVC_size,time.time()-start_time))
backtrack = True
#print('First backtrack-vertex-',vi)
else: #partial solution
#maxnode, maxdegree = find_maxdeg(CurG)
CurLB = Lowerbound(CurG) + CurVC_size
#print(CurLB)
#CurLB=297
if CurLB < UpperBound: # worth exploring
# print('upper',UpperBound)
vj = find_maxdeg(CurG)
Frontier.append((vj[0], 0, (vi, state)))#(vi,state) is parent of vj
Frontier.append((vj[0], 1, (vi, state)))
# print('Frontier',Frontier)
else:
# end of path, will result in worse solution,backtrack to parent
backtrack=True
#print('Second backtrack-vertex-',vi)
if backtrack==True:
#print('Hello. CurNode:',vi,state)
if Frontier != []: #otherwise no more candidates to process
nextnode_parent = Frontier[-1][2] #parent of last element in Frontier (tuple of (vertex,state))
#print(nextnode_parent)
# backtrack to the level of nextnode_parent
if nextnode_parent in CurVC:
id = CurVC.index(nextnode_parent) + 1
while id < len(CurVC): #undo changes from end of CurVC back up to parent node
mynode, mystate = CurVC.pop() #undo the addition to CurVC
CurG.add_node(mynode) #undo the deletion from CurG
# find all the edges connected to vi in Graph G
# or the edges that connected to the nodes that not in current VC set.
curVC_nodes = list(map(lambda t:t[0], CurVC))
for nd in G.neighbors(mynode):
if (nd in CurG.nodes()) and (nd not in curVC_nodes):
CurG.add_edge(nd, mynode) #this adds edges of vi back to CurG that were possibly deleted
elif nextnode_parent == (-1, -1):
# backtrack to the root node
CurVC.clear()
CurG = G.copy()
else:
print('error in backtracking step')
end_time=time.time()
delta_time=end_time-start_time
if delta_time>T:
print('Cutoff time reached')
#TO FIND THE VERTEX WITH MAXIMUM DEGREE IN REMAINING GRAPH
def find_maxdeg(g):
deglist = dict(g.degree())
deglist_sorted = sorted(deglist.items(), reverse=True, key=operator.itemgetter(1)) # sort in descending order of node degree
v = deglist_sorted[0] # tuple - (node,degree)
return v
#EXTIMATE LOWERBOUND
def Lowerbound(graph):
lb=graph.number_of_edges() / find_maxdeg(graph)[1]
lb=ceil(lb)
return lb
def ceil(d):
"""
return the minimum integer that is bigger than d
"""
if d > int(d):
return int(d) + 1
else:
return int(d)
#CALCULATE SIZE OF VERTEX COVER (NUMBER OF NODES WITH STATE=1)
def VC_Size(VC):
# VC is a tuple list, where each tuple = (node_ID, state, (node_ID, state)) vc_size is the number of nodes which has state == 1
vc_size = 0
for element in VC:
vc_size = vc_size + element[1]
return vc_size
deglist = dict(cells.degree())
deglist_sorted = sorted(deglist.items(), reverse=True, key=operator.itemgetter(1)) # sort in descending order of node degree
```
| github_jupyter |
## Univariate Analysis
```
import pandas as pd
# load the csv file of data
df_suicide = pd.read_csv('suicide_data.csv')
# display the head() of the dataset
df_suicide.head()
df_suicide.info()
df_suicide.isnull().sum()
```
### Consider the variable 'gdp_per_capita' for univariate analysis
#### In our data, 'gdp_per_capita' is a numerical variable. Thus, we can use different visualization techniques (histogram, boxplot, violinplot, and so on) to study the distribution of the variable.
### Histogram - [Kindly Provide the Inferences]
```
# Plot the histogram
import matplotlib.pyplot as plt
plt.hist(df_suicide['gdp_per_capita ($)'],bins=100)
```
### Inferences: The histogram appears right-skewed.
```
df_suicide['gdp_per_capita ($)'].describe()
df_suicide['gdp_per_capita ($)'].mode()
df_suicide['gdp_per_capita ($)'].median()
df_suicide['gdp_per_capita ($)'].skew()
```
### Distribution plot: Use distplot() to plot a kernel density estimator (KDE) along with the histogram to study the distribution of the data
```
import seaborn as sns
# Plot the distribution plot
sns.distplot(df_suicide['gdp_per_capita ($)'])
```
### Violin plot: It is similar to a boxplot which shows the distribution of the quantitative variable
```
# plot the violin plot
sns.violinplot(df_suicide['gdp_per_capita ($)'])
```
### Boxplot: Boxplot can be used to visualize the presence of outliers in the data. In boxplot, the observation will be an outlier, if it is outside the specific range (1.5 times IQR above the third quartile and below the first quartile)
```
#Plot the Box plot
sns.boxplot(df_suicide['gdp_per_capita ($)'])
```
# Multivariate Analysis
### Grouped boxplot : It is used to compare the distribution of different categories in the categorical variable. Plot a grouped boxplot to check the extreme values for the population in the different generations
```
# Plot a Groped Box plot
fig, ax = plt.subplots(figsize=(20,7))
sns.boxplot(ax=ax, x="population",y="generation",data=df_suicide, linewidth=1.5 ,palette="Set3")
```
## Heat Map
```
#visualize the correlation matrix using the heatmap.
df_suicide_copy=df_suicide.copy().drop(columns=["HDI for year","country","age","country-year"])
list(df_suicide_copy.columns)
df_suicide_copy['sex'] = df_suicide_copy['sex'].replace('male',0).replace('female',1).astype('int32')
df_suicide_copy['generation'] = df_suicide_copy['generation'].replace('Generation X',0).replace('Silent',1).replace('G.I. Generation',2).replace('Boomers',3).replace('Millenials',4).replace('Generation Z',5).astype('int32')
df_suicide_copy.head()
df_suicide_copy[' gdp_for_year ($) '] = df_suicide_copy[' gdp_for_year ($) '].str.replace(',','').astype('float64')
fig, ax = plt.subplots(figsize=(8,6))
ax=sns.heatmap(df_suicide_copy.corr(), annot=True, vmin=-1, vmax=1, center= 0, cmap= 'coolwarm')
ax.set_ylim(8.0, 0)
```
| github_jupyter |
## P7: Résolvez des problèmes en utilisant des algorithmes en Python #1
Repartir des essais de découverte précédent pour améliorer la calcul brute force au mieux.
Si la contrainte est toujours du style budget maximum, il est inutile de conserver les combinaisons dont le cout est > BUDGET.
Cela n'en devient pas pour autant de la programmation dynamique, réservée à l'algo suivant ;
la programmation dynamique évitant de recalculer des sous-arbres déja parcourus.
* Caculer simultanément à l'explosion combinatoire.
Dans les essais de découverte, j'ai dans un 1er temps explosé toutes les combinaisons possibles puis ensuite seulement calculé et limité au sous-ensemble de budget <= BUDGET.
Un calcul simultané permettrait la vérification de contrainte non atteinte et la réduction de l'ensemble des solutions possibles.
* Structure de donnée.
Pour "porter" les combinaisons une liste est envisageable avec pour chercher les données (cout, profit) de chaque action un dictionnaire semble efficace.
### 1. Préparation des données
```
import csv as csv
import re as re
import time
try:
import big_o
except ModuleNotFoundError:
print('cherche',ModuleNotFoundError)
!pip install --upgrade pip
!pip install big_o
import big_o
# constant
#FILE = "data/p7-20-shares.csv"
#FIELDNAMES = ['name', 'cost', 'profit_share', 'profit']
FILE = "data/dataset1_Python+P7"
FIELDNAMES = ['name', 'cost', 'profit']
BUDGET = 50000
```
### En complément de Big_O, mesure du temps
Si la mesure du temps d'execution est dépendante du type de PC, des processus en cours d'execution etc., elle permet une approximation de l'efficacité des algorithme choisis complémentaire à celle de la notation big_o.
Suivant le conseil de [Marina Mele](http://www.marinamele.com/author/marina-melegmail-com) dans [7 tips to Time Python scripts and control Memory & CPU usage](https://www.marinamele.com/7-tips-to-time-python-scripts-and-control-memory-and-cpu-usage)
```
def fn_timer(function):
# @wraps(function)
def function_timer(*args, **kwargs):
t0 = time.perf_counter_ns()
result = function(*args, **kwargs)
t1 = time.perf_counter_ns()
print ("Total time running %s: %s nanoseconds" %
(function.__name__, str(t1-t0))
)
return result
return function_timer
# nettoyer les données des fichiers en entrée
def clean_char(texte: str) -> str:
""" on ne conserve que les caractères lisibles
les lettres, chiffres, ponctuations décimales et signes
les valeurs negatives sont acceptées, du point de vue profit.
"""
texte_propre = re.sub(r"[^a-zA-Z0-9\-\.\,\+]", "", texte.replace(',','.'))
return texte_propre
```
### Amélioration des données
Sur python le travail avec float est plus couteux qu'en entier.
Une solution est alors de multiplier par 100 cout, budget et profit car cela ne change pas
le résultat.
```
""" lecture, nettoyage et chargement en dict.
les non valeurs NaN sont rejetées.
"""
action_dict = {}
try:
with open(FILE, "r", newline='', encoding='utf-8') as file:
csv_reader = csv.DictReader(file, fieldnames=FIELDNAMES,
delimiter=';', doublequote=False)
# skip the header
next(csv_reader)
for idx, line in enumerate(csv_reader):
clean_data = True
if line[FIELDNAMES[0]] != "":
cle = clean_char(line[FIELDNAMES[0]])
else:
print(f" line {idx} had missing share name; dropped.")
clean_data = False
if line[FIELDNAMES[1]] != "":
cout = 100 * int(clean_char(line[FIELDNAMES[1]]))
else:
print(f" line {idx} had missing cost data; dropped.")
clean_data = False
if line[FIELDNAMES[3]] != "":
gain = int(100 * float(clean_char(line[FIELDNAMES[3]])))
else:
print(f" line {idx} had missing profit data; dropped.")
clean_data = False
if (gain < 0) or (cout < 0):
print(f" line {idx} had negative value; accepted but pls check.")
if (clean_data):
action_dict[cle] = (cout, gain)
except FileNotFoundError:
print(f" fichier non trouvé, Merci de vérifier son nom {file_name} : {FileNotFoundError}")
except IOError:
print(f" une erreur est survenue à l'écriture du fichier {file_name} : {IOError}")
action_dict
```
### 2. Calcul brute force.
$
{n \choose k}={\frac {n!}{k!(n-k)!}}.
### 2.1 Estimation simple du nombre de calcul maximum.
entendre : Non réduit à la contrainte BUDGET.
```
def number_combi(n: int, k: int) -> int:
def factorielle(x:int) -> int:
if x <=1:
return 1
else:
return (x * factorielle(x-1))
top = factorielle(n)
bot = factorielle(k) * factorielle(n-k)
if bot > 0:
return top/bot
else:
raise Error
denombrement = False
if denombrement:
N = len(action_dict.keys())
number_of_combi = 0
for K in range(N):
number_of_combi += number_combi(N, K)
print(number_of_combi)
```
Nous sommes prévenu, il y a environ 1 million de combinaison d'un portefeuille de 20 actions!
Le calcul des combinaisons va être long!!
#### 2.3 Fonction combinatoire proposée.
```
# initialize
actions_list = [([x],y[0],y[1]) for x,y in action_dict.items()]
actions_list
def explode_iter(elements, length):
""" global action_dict to fetch cost & profit
elements would be list(tuple(list, cost, profit))
"""
# pour le 1er élément de la combinaison, il peut prendre toute valeur de la liste
for i in range(len(elements)):
# singelton -> on rapporte des t-uple de 1 avec le 1er élément lui-même
if length == 1:
yield (elements[i][0] , elements[i][1], elements[i][2])
else:
# on prend la liste réduite à partir de l'élément suivant i et
# on en cherche les combinaisons de longueur -1
#print('sous-liste:',elements[i+1:len(elements)])
#print('à i:',i)
for next1 in explode_iter(elements[i+1:len(elements)], length-1):
#print('next1:',next1)
#print('elements[0][1]',elements[i][1])
#print('next1[0]:',next1[0])
# et on retourne la liste précédent plus la liste de l'appel suivant
cout = elements[i][1] + next1[1]
profit = elements[i][2] + next1[2]
# optimisation
if cout <= BUDGET:
yield (elements[i][0] + next1[0],
cout,
profit)
# toutes les combinaisons de k éléments de la liste l
list(explode_iter(actions_list, 1))
@fn_timer
def algo_force_brute(portefeuille: list) -> list:
liste_output = []
for largeur in range(1,len(portefeuille)+1):
liste_output.extend(list(explode_iter(portefeuille, largeur)))
return liste_output
explosion_force_brute = algo_force_brute(actions_list)
#explosion_force_brute
```
Résultat
Total time running algo_force_brute: 13138267100 nanoseconds
Le print de l'explosion combinatoire est lourd en ressource.
`IOPub data rate exceeded.
`The Jupyter server will temporarily stop sending output
`to the client in order to avoid crashing it.
`To change this limit, set the config variable
`--ServerApp.iopub_data_rate_limit`.
Current values:
ServerApp.iopub_data_rate_limit=1000000.0 (bytes/sec)
ServerApp.rate_limit_window=3.0 (secs)
#### Etape 3 déterminer l'élément à plus haut profit pour un budget <= 50000
Un tri puis une recherche dichotomique?
ou
Une recherche non dichotomique dans une liste non préalablement triée??
ou le parcours complet de la liste une fois
```
## tri par profit puis cout décroissant
explosion_force_brute.sort(key = lambda sub : sub[2] ,reverse = True)
explosion_force_brute[0]
meilleur_profit = 0
meilleur_budget = 0
meilleur_combinaison = []
for courant in explosion_force_brute:
if (courant[2] > meilleur_profit) & (courant[1] < BUDGET):
meilleur_profit = courant[2]
meilleur_budget = courant[1]
meilleur_combinaison = courant[0]
print(meilleur_profit, meilleur_budget, meilleur_combinaison)
```
9907 49800 ['Action-4', 'Action-5', 'Action-6', 'Action-8', 'Action-10', 'Action-11', 'Action-13', 'Action-18', 'Action-19', 'Action-20']
| github_jupyter |
```
%pylab inline
from scipy.interpolate import interpn
from helpFunctions import surfacePlot
from constant import *
from multiprocessing import Pool
from functools import partial
import warnings
import math
warnings.filterwarnings("ignore")
np.printoptions(precision=2)
# load the policy funciton
Vgrid_renting = np.load("Vgrid_renting.npy")
cgrid_renting = np.load("cgrid_renting.npy")
bgrid_renting = np.load("bgrid_renting.npy")
kgrid_renting = np.load("kgrid_renting.npy")
hgrid_renting = np.load("hgrid_renting.npy")
Mgrid_renting = np.load("Mgrid_renting.npy")
Hgrid_renting = np.load("Hgrid_renting.npy")
V1000 = np.load("Vgrid1000.npy")
V1500 = np.load("Vgrid1500.npy")
V2000 = np.load("Vgrid2000.npy")
V750 = np.load("Vgrid750.npy")
cgrid1000 = np.load("cgrid1000.npy")
bgrid1000 = np.load("bgrid1000.npy")
kgrid1000 = np.load("kgrid1000.npy")
qgrid1000 = np.load("qgrid1000.npy")
cgrid1500 = np.load("cgrid1500.npy")
bgrid1500 = np.load("bgrid1500.npy")
kgrid1500 = np.load("kgrid1500.npy")
qgrid1500 = np.load("qgrid1500.npy")
cgrid2000 = np.load("cgrid2000.npy")
bgrid2000 = np.load("bgrid2000.npy")
kgrid2000 = np.load("kgrid2000.npy")
qgrid2000 = np.load("qgrid2000.npy")
cgrid750 = np.load("cgrid750.npy")
bgrid750 = np.load("bgrid750.npy")
kgrid750 = np.load("kgrid750.npy")
qgrid750 = np.load("qgrid750.npy")
indexH = [750, 1000, 1500, 2000]
cgrid = [cgrid750, cgrid1000, cgrid1500, cgrid2000]
bgrid = [bgrid750, bgrid1000, bgrid1500, bgrid2000]
kgrid = [kgrid750, kgrid1000, kgrid1500, kgrid2000]
qgrid = [qgrid750, qgrid1000, qgrid1500, qgrid2000]
import quantecon as qe
import timeit
mc = qe.MarkovChain(Ps)
H_options = [0, 750, 1000, 1500, 2000]
M_options = [0.2, 0.5, 0.8]
# wealth discretization
ws = np.array([10,25,50,75,100,125,150,175,200,250,500,750,1000,1500,3000])
w_grid_size = len(ws)
# 401k amount discretization
ns = np.array([1, 5, 10, 15, 25, 50, 100, 150, 400, 1000])
n_grid_size = len(ns)
def closest(lst, K):
'''
Find the closest value of K in a list lst
'''
lst = np.asarray(lst)
idx = (np.abs(lst - K)).argmin()
return lst[idx]
def action_rent(t, x):
w, n, e, s, z = x
points = (ws, ns)
c = interpn(points, cgrid_renting[:,:,e,s,z,t], x[:2], method = "nearest", bounds_error = False, fill_value = None)[0]
b = interpn(points, bgrid_renting[:,:,e,s,z,t], x[:2], method = "nearest", bounds_error = False, fill_value = None)[0]
k = interpn(points, kgrid_renting[:,:,e,s,z,t], x[:2], method = "nearest", bounds_error = False, fill_value = None)[0]
if e == 1:
H = interpn(points, Hgrid_renting[:,:,e,s,z,t], x[:2], method = "nearest", bounds_error = False, fill_value = None)[0]
# decide to buy or not to buy a house
H = H_options[np.sum(H>=np.array(H_options))-1]
if H == 0:
h = interpn(points, hgrid_renting[:,:,e,s,z,t], x[:2], method = "nearest", bounds_error = False, fill_value = None)[0]
return (c,b,k,h)
else:
M = interpn(points, Mgrid_renting[:,:,e,s,z,t], x[:2], method = "nearest", bounds_error = False, fill_value = None)[0]
M = closest(H*M_options, M)
return (c,b,k,M,H)
else:
h = interpn(points, hgrid_renting[:,:,e,s,z,t], x[:2], method = "nearest", bounds_error = False, fill_value = None)[0]
return (c,b,k,h)
def action_own(t, x):
w, n, M, e, s, z, H = x
# Mortgage amount, * 0.25 is the housing price per unit
Ms = np.array([0.01*H,0.05*H,0.1*H,0.2*H,0.3*H,0.4*H,0.5*H,0.8*H]) * pt
points = (ws,ns,Ms)
c = interpn(points, cgrid[indexH.index(H)][:,:,:,e,s,z,t], x[:3], method = "nearest", bounds_error = False, fill_value = None)[0]
b = interpn(points, bgrid[indexH.index(H)][:,:,:,e,s,z,t], x[:3], method = "nearest", bounds_error = False, fill_value = None)[0]
k = interpn(points, kgrid[indexH.index(H)][:,:,:,e,s,z,t], x[:3], method = "nearest", bounds_error = False, fill_value = None)[0]
q = interpn(points, qgrid[indexH.index(H)][:,:,:,e,s,z,t], x[:3], method = "nearest", bounds_error = False, fill_value = None)[0]
if q <= 0.75:
q = 0.5
else:
q = 1
return (c,b,k,q)
def transition_to_rent(x,a,t,s_next):
'''
imput a is np array constains all possible actions
from x = [w, n, e, s, z] to x = [w, n, e, s, z]
'''
w, n, e, s, z = x
c, b, k, h = a
s = int(s)
s_next = int(s_next)
# transition of z
if z == 1:
z_next = 1
else:
if k == 0:
z_next = 0
else:
z_next = 1
# variables used to collect possible states and probabilities
x_next = []
w_next = b*(1+r_b[s]) + k*(1+r_k[s_next])
n_next = gn(t, n, x, (r_b[s]+r_k[s_next])/2)
if t >= T_R:
return [w_next, n_next, 0, s_next, z_next]
else:
if e == 1:
for e_next in [0,1]:
x_next.append([w_next, n_next, e_next, s_next, z_next])
prob_next = [Pe[s,e], 1 - Pe[s,e]]
else:
for e_next in [0,1]:
x_next.append([w_next, n_next, e_next, s_next, z_next])
prob_next = [1-Pe[s,e], Pe[s,e]]
return x_next[np.random.choice(len(prob_next), 1, p = prob_next)[0]]
def transition_to_own(x,a,t,s_next):
'''
imput a is np array constains all possible actions
from x = [w, n, e, s, z] to x = [w, n, M, e, s, z, H]
'''
w, n, e, s, z = x
# variables used to collect possible states and probabilities
x_next = []
c, b, k, M, H = a
s = int(s)
s_next = int(s_next)
# transition of z
if z == 1:
z_next = 1
else:
if k == 0:
z_next = 0
else:
z_next = 1
M_next = M*(1+rh)
w_next = b*(1+r_b[s]) + k*(1+r_k[s_next])
n_next = gn(t, n, x, (r_b[s]+r_k[s_next])/2)
if t >= T_R:
return [w_next, n_next, M_next, 0, s_next, z_next, H]
else:
if e == 1:
for e_next in [0,1]:
x_next.append([w_next, n_next, M_next, e_next, s_next, z_next, H])
prob_next = [Pe[s,e], 1 - Pe[s,e]]
else:
for e_next in [0,1]:
x_next.append([w_next, n_next, M_next, e_next, s_next, z_next, H])
prob_next = [1-Pe[s,e], Pe[s,e]]
return x_next[np.random.choice(len(prob_next), 1, p = prob_next)[0]]
def transition_after_own(x, a, t, s_next):
'''
Input: state and action and time
Output: possible future states and corresponding probability
'''
w, n, M, e, s, z, H = x
c,b,k,q = a
# variables used to collect possible states and probabilities
x_next = []
m = M/D[T_max-t]
M_next = M*(1+rh) - m
# transition of z
if z == 1:
z_next = 1
else:
if k == 0:
z_next = 0
else:
z_next = 1
n_next = gn(t, n, x, (r_b[s]+r_k[s_next])/2)
w_next = b*(1+r_b[int(s)]) + k*(1+r_k[s_next])
if t >= T_R:
return [w_next, n_next, M_next, 0, s_next, z_next, H]
else:
if e == 1:
for e_next in [0,1]:
x_next.append([w_next, n_next, M_next, e_next, s_next, z_next, H])
prob_next = [Pe[s,e], 1 - Pe[s,e]]
else:
for e_next in [0,1]:
x_next.append([w_next, n_next, M_next, e_next, s_next, z_next, H])
prob_next = [1-Pe[s,e], Pe[s,e]]
return x_next[np.random.choice(len(prob_next), 1, p = prob_next)[0]]
'''
Start with renting:
w = 5
n = 0
e = 1
s = 1
1000 agents for 1 economy, 500 economies.
'''
names = ['w', 'n', 'M', 'e', 's', 'z', 'H',
'yt', 'c', 'b', 'k', 'q', 'h','live']
x0 = [5, 0, 1, 1, 0]
numAgents = 1000
numEcons = 500
import quantecon as qe
import random as rd
mc = qe.MarkovChain(Ps)
EconStates = [mc.simulate(ts_length=T_max - T_min, init=0) for _ in range(numEcons)]
def simulation(i):
track = np.zeros((T_max - T_min,len(names)))
econState = EconStates[i//numAgents]
alive = True
x = x0
for t in range(1,len(econState)-1):
if rd.random() > Pa[t]:
alive = False
if alive:
track[t, 13] = 1
s_next = econState[t+1]
# if is still renting
if (len(x) == 5):
a = action_rent(t,x)
# continue to rent
if (len(a) == 4):
# x = [w,n,e,s,z]
# a = [c,b,k,h]
track[t, 0] = x[0]
track[t, 1] = x[1]
track[t, 2] = 0
track[t, 3] = x[2]
track[t, 4] = x[3]
track[t, 5] = x[4]
track[t, 6] = 0
track[t, 7] = y(t,x)
track[t, 8] = a[0]
track[t, 9] = a[1]
track[t, 10] = a[2]
track[t, 11] = 0
track[t, 12] = a[3]
x = transition_to_rent(x,a,t,s_next)
# switch to own
else:
# a = [c,b,k,M,H]
track[t, 0] = x[0]
track[t, 1] = x[1]
track[t, 2] = a[3]
track[t, 3] = x[2]
track[t, 4] = x[3]
track[t, 5] = x[4]
track[t, 6] = a[4]
track[t, 7] = y(t,x)
track[t, 8] = a[0]
track[t, 9] = a[1]
track[t, 10] = a[2]
track[t, 11] = 0
track[t, 12] = a[4]*(1+kappa)
x = transition_to_own(x,a,t,s_next)
# if owning a house already
else:
a = action_own(t, x)
# x = [w,n,M,e,s,z,H]
# a = [c,b,k,q]
track[t, 0] = x[0]
track[t, 1] = x[1]
track[t, 2] = x[2]
track[t, 3] = x[3]
track[t, 4] = x[4]
track[t, 5] = x[5]
track[t, 6] = x[6]
track[t, 7] = y(t,x)
track[t, 8] = a[0]
track[t, 9] = a[1]
track[t, 10] = a[2]
track[t, 11] = a[3]
# calculate housing consumption
if a[3] == 1:
Vh = (1+kappa)*x[6]
else:
Vh = (1-kappa)*(x[6]/2)
track[t, 12] = Vh
x = transition_after_own(x, a, t, s_next)
return track
%%time
pool = Pool()
agents = pool.map(simulation, list(range(numAgents*numEcons)))
pool.close()
ww = np.zeros((T_max-T_min, numAgents*numEcons))
nn = np.zeros((T_max-T_min, numAgents*numEcons))
MM = np.zeros((T_max-T_min, numAgents*numEcons))
ee = np.zeros((T_max-T_min, numAgents*numEcons))
ss = np.zeros((T_max-T_min, numAgents*numEcons))
ZZ = np.zeros((T_max-T_min, numAgents*numEcons))
HH = np.zeros((T_max-T_min, numAgents*numEcons))
ytyt = np.zeros((T_max-T_min, numAgents*numEcons))
cc = np.zeros((T_max-T_min, numAgents*numEcons))
bb = np.zeros((T_max-T_min, numAgents*numEcons))
kk = np.zeros((T_max-T_min, numAgents*numEcons))
qq = np.zeros((T_max-T_min, numAgents*numEcons))
hh = np.zeros((T_max-T_min, numAgents*numEcons))
live = np.zeros((T_max-T_min, numAgents*numEcons))
def separateAttributes(agents):
for i in range(numAgents*numEcons):
ww[:,i] = agents[i][:,0]
nn[:,i] = agents[i][:,1]
MM[:,i] = agents[i][:,2]
ee[:,i] = agents[i][:,3]
ss[:,i] = agents[i][:,4]
ZZ[:,i] = agents[i][:,5]
HH[:,i] = agents[i][:,6]
ytyt[:,i] = agents[i][:,7]
cc[:,i] = agents[i][:,8]
bb[:,i] = agents[i][:,9]
kk[:,i] = agents[i][:,10]
qq[:,i] = agents[i][:,11]
hh[:,i] = agents[i][:,12]
live[:,i] = agents[i][:,13]
separateAttributes(agents)
def quantileForPeopleWholive(attribute, quantiles = [0.25, 0.5, 0.75]):
qList = []
for i in range(T_max):
if len(np.where(live[i,:] == 1)[0]) == 0:
qList.append(np.array([0] * len(quantiles)))
else:
qList.append(np.quantile(attribute[i, np.where(live[i,:] == 1)], q = quantiles))
return np.array(qList)
def meanForPeopleWholive(attribute):
means = []
for i in range(T_max):
if len(np.where(live[i,:] == 1)[0]) == 0:
means.append(np.array([0]))
else:
means.append(np.mean(attribute[i, np.where(live[i,:] == 1)]))
return np.array(means)
# Population during the entire simulation period
plt.plot(np.mean(live,axis = 1))
plt.plot(quantileForPeopleWholive(ww,[0.25,0.5,0.75,0.99]))
plt.plot(quantileForPeopleWholive(nn))
plt.plot(quantileForPeopleWholive(MM))
plt.plot(meanForPeopleWholive(ee))
plt.plot(quantileForPeopleWholive(ss))
plt.plot(quantileForPeopleWholive(ZZ))
plt.plot(quantileForPeopleWholive(HH))
plt.plot(quantileForPeopleWholive(ytyt))
plt.plot(quantileForPeopleWholive(cc))
plt.plot(quantileForPeopleWholive(bb))
plt.plot(quantileForPeopleWholive(kk))
plt.plot(quantileForPeopleWholive(qq))
plt.plot(quantileForPeopleWholive(hh))
# mean value of the key variables
plt.figure(figsize = [14,8])
plt.plot(meanForPeopleWholive(ww), label = "wealth")
plt.plot(meanForPeopleWholive(cc), label = "Consumption")
plt.plot(meanForPeopleWholive(bb), label = "Bond")
plt.plot(meanForPeopleWholive(kk), label = "Stock")
plt.legend()
plt.plot(meanForPeopleWholive(nn), label = "401k")
Hgrid_renting.shape
# Downpayment.
for age in range(30):
print(Hgrid_renting[:,9,1,10,1,10] - Mgrid_renting[:,9,1,10,1,10])
alpha
```
| github_jupyter |
```
import sys, time
import numpy as np
import scipy.signal as sig
import astropy.io.fits as fits
import pyfftw
import pyfftw.interfaces.numpy_fft as fft
pyfftw.interfaces.cache.enable()
pyfftw.interfaces.cache.set_keepalive_time(1.)
class Galaxy:
# This class acts as a data structure for holding galaxy data
# such as surface brightness, distance, etc.
def __init__(self, ID):
self.ID = ID
self.data = {'ID': ID, 'T': None, 'D': None, 'sma': [], 'ell': None, 'muFUV': [], 'muNUV': []}
def add_data(self, data):
for tpl in data:
dt, dp = tpl
if dt in self.data:
if dt in ['muFUV', 'muNUV', 'sma']:
# If the data is one of these types then it should be appended
# TODO: It's still unclear how the surface brightness is formatted.
self.data[dt].append(dp)
else:
# If not, then just fill in the corresponding entry
self.data[dt] = dp
else:
# Data types that are not already included can be added
self.data[dt] = dp
return self.data
def get_data(self):
return self.data
class Galaxy_box:
# This class holds a bunch of galaxies, and can return one at random or by specifying a galaxy id
# A distance range can be specified
def __init__(self, Dmin=0, Dmax=1000):
self.galaxies = {}
# Read galaxy data into a dictionary
# Galaxy data from Bouquin+, 2018 catalog
# Paper: https://iopscience.iop.org/article/10.3847/1538-4365/aaa384/pdf
# Catalog: https://bit.ly/2SmJWHV
# TODO: It's still unclear how the surface brightness is formatted.
with open('table1.dat') as file:
for line in file:
splitline = line.split()
if Dmin < float(splitline[4]) < Dmax:
gal_id = splitline[0]
tmp_gal = Galaxy(gal_id)
tmp_gal.add_data([('T', float(splitline[3])),('D', float(splitline[4]))])
self.galaxies[gal_id] = tmp_gal
with open('table3.dat') as file:
for line in file:
splitline = line.split()
gal_id = splitline[0]
if gal_id in self.galaxies:
self.galaxies[gal_id].add_data([('sma', int(splitline[1])), ('ell', float(splitline[2])),
('muFUV', float(splitline[3])), ('muNUV', float(splitline[7]))])
def get_galaxy(self, method = 'random', ID = None):
# Grab a galaxy out of the box at random or using an ID
if method == 'random':
return self.galaxies[np.random.choice(list(self.galaxies.keys()))]
elif method=='ID':
if ID:
if ID not in self.galaxies:
print('Invalid galaxy ID')
return None
else:
return self.galaxies[ID]
else:
print('No ID provided')
return None
else:
print('Invalid method')
return None
def py_zogy(N, R, P_N, P_R, S_N, S_R, SN, SR, dx=0.25, dy=0.25):
'''Python implementation of ZOGY image subtraction algorithm. Copied from https://github.com/cenko/ZOGY.
Modified so that inputs are arrays rather than external files.
Assumes images have been aligned, background subtracted, and gain-matched.
Arguments:
N: New image
R: Reference image
P_N: PSF of New image
P_R: PSF or Reference image
S_N: 2D Uncertainty (sigma) of New image
S_R: 2D Uncertainty (sigma) of Reference image
SN: Average uncertainty (sigma) of New image
SR: Average uncertainty (sigma) of Reference image
dx: Astrometric uncertainty (sigma) in x coordinate
dy: Astrometric uncertainty (sigma) in y coordinate
Returns:
D: Subtracted image
P_D: PSF of subtracted image
S_corr: Corrected subtracted image
'''
# Load the PSFs into memory
P_N_small = P_N
P_R_small = P_R
# Place PSF at center of image with same size as new / reference
P_N = np.zeros(N.shape)
P_R = np.zeros(R.shape)
idx = [slice(N.shape[0]/2 - P_N_small.shape[0]/2,
N.shape[0]/2 + P_N_small.shape[0]/2 + 1),
slice(N.shape[1]/2 - P_N_small.shape[1]/2,
N.shape[1]/2 + P_N_small.shape[1]/2 + 1)]
P_N[idx] = P_N_small
P_R[idx] = P_R_small
# Shift the PSF to the origin so it will not introduce a shift
P_N = fft.fftshift(P_N)
P_R = fft.fftshift(P_R)
# Take all the Fourier Transforms
N_hat = fft.fft2(N)
R_hat = fft.fft2(R)
P_N_hat = fft.fft2(P_N)
P_R_hat = fft.fft2(P_R)
# Fourier Transform of Difference Image (Equation 13)
D_hat_num = (P_R_hat * N_hat - P_N_hat * R_hat)
D_hat_den = np.sqrt(SN**2 * np.abs(P_R_hat**2) + SR**2 * np.abs(P_N_hat**2))
D_hat = D_hat_num / D_hat_den
# Flux-based zero point (Equation 15)
FD = 1. / np.sqrt(SN**2 + SR**2)
# Difference Image
# TODO: Why is the FD normalization in there?
D = np.real(fft.ifft2(D_hat)) / FD
# Fourier Transform of PSF of Subtraction Image (Equation 14)
P_D_hat = P_R_hat * P_N_hat / FD / D_hat_den
# PSF of Subtraction Image
P_D = np.real(fft.ifft2(P_D_hat))
P_D = fft.ifftshift(P_D)
P_D = P_D[idx]
# Fourier Transform of Score Image (Equation 17)
S_hat = FD * D_hat * np.conj(P_D_hat)
# Score Image
S = np.real(fft.ifft2(S_hat))
# Now start calculating Scorr matrix (including all noise terms)
# Start out with source noise
# Sigma to variance
V_N = S_N**2
V_R = S_R**2
# Fourier Transform of variance images
V_N_hat = fft.fft2(V_N)
V_R_hat = fft.fft2(V_R)
# Equation 28
kr_hat = np.conj(P_R_hat) * np.abs(P_N_hat**2) / (D_hat_den**2)
kr = np.real(fft.ifft2(kr_hat))
# Equation 29
kn_hat = np.conj(P_N_hat) * np.abs(P_R_hat**2) / (D_hat_den**2)
kn = np.real(fft.ifft2(kn_hat))
# Noise in New Image: Equation 26
V_S_N = np.real(fft.ifft2(V_N_hat * fft.fft2(kn**2)))
# Noise in Reference Image: Equation 27
V_S_R = np.real(fft.ifft2(V_R_hat * fft.fft2(kr**2)))
# Astrometric Noise
# Equation 31
# TODO: Check axis (0/1) vs x/y coordinates
S_N = np.real(fft.ifft2(kn_hat * N_hat))
dSNdx = S_N - np.roll(S_N, 1, axis=1)
dSNdy = S_N - np.roll(S_N, 1, axis=0)
# Equation 30
V_ast_S_N = dx**2 * dSNdx**2 + dy**2 * dSNdy**2
# Equation 33
S_R = np.real(fft.ifft2(kr_hat * R_hat))
dSRdx = S_R - np.roll(S_R, 1, axis=1)
dSRdy = S_R - np.roll(S_R, 1, axis=0)
# Equation 32
V_ast_S_R = dx**2 * dSRdx**2 + dy**2 * dSRdy**2
# Calculate Scorr
S_corr = S / np.sqrt(V_S_N + V_S_R + V_ast_S_N + V_ast_S_R)
return D, P_D, S_corr
```
| github_jupyter |
```
%%javascript
if(IPython.tab_as_tab_everywhere)IPython.tab_as_tab_everywhere()
import matplotlib.pyplot as plt
import os, sys, json
import pandas as pd
import numpy as np
from tqdm import tqdm
import hashlib
# from tf.keras.models import Sequential # This does not work!
from tensorflow.python.keras.models import Sequential
from tensorflow.python.keras.layers import InputLayer, Input
from tensorflow.python.keras.layers import Reshape, MaxPooling2D,Dropout
from tensorflow.python.keras.layers import Conv2D, Dense, Flatten
from tensorflow.python.keras.models import Model
from tensorflow.python.keras.optimizers import Adam
from sklearn.metrics import roc_auc_score
pd.set_option('display.max_rows', 50)
pd.set_option('display.max_columns', 200)
train = pd.read_csv('data/application_train.csv', encoding='ISO-8859-1').sample(frac=1)
train['INVERSE_TARGET'] = 1 - train['TARGET']
train.loc[train['DAYS_EMPLOYED'] == 365243,'DAYS_EMPLOYED'] = 0
train['DAYS_EMPLOYED'] = -train['DAYS_EMPLOYED']
train['DAYS_BIRTH'] = -train['DAYS_BIRTH']
train['DAYS_REGISTRATION'] = -train['DAYS_REGISTRATION']
df = pd.DataFrame()
df['CNT_CHILDREN'] = train['CNT_CHILDREN'].clip(0,4)
df['DAYS_BIRTH'] = train['DAYS_BIRTH']
df['DAYS_EMPLOYED'] = train['DAYS_EMPLOYED']
df['DAYS_REGISTRATION'] = train['DAYS_REGISTRATION']
#documents
df['documents'] = 1
for id_ in range(2,22):
df['documents'] += train['FLAG_DOCUMENT_' + str(id_)]
#objects
preprocess = {}
for col in train:
if train[col].dtype.kind in 'bif' and not col in ['TARGET','INVERSE_TARGET','SK_ID_CURR'] and not col in df.columns:
noNaN = train[col].count() == len(train)
if noNaN:
df[col] = train[col]
else:
df[col] = train[col].fillna(0)
df[col + '_nan'] = train[col].isna()
preprocess[col] ={
'type':'num',
'noNaN': int(noNaN)
}
if train[col].dtype != 'O': continue
train[col] = train[col].fillna('none')
dic = train.groupby([col])['TARGET'].mean().to_dict()
preprocess[col] = {
'type':'dic',
'dic':dic
}
df[col] = train[col].map(dic)
use_cols = pd.DataFrame(columns=['col','cor'])
for col in df:
cor = np.corrcoef(train['TARGET'],df[col])[0][1]
use_cols = use_cols.append({
'col': col,
'cor': cor,
'abs': abs(cor)
},ignore_index=True)
use = use_cols.sort_values(by=['abs'],ascending=False).head(150)['col'].tolist()
df_use = df[use]
df_norm = pd.DataFrame()
lims = {}
for col in df_use:
column = df_use[col]
d_min = float(column.min())
d_max = float(column.max())
dif = d_max - d_min
lims[col] = {
'a': d_min,
'b': dif,
}
df_norm[col] = (df_use[col] - d_min) / dif
columns = len(df_use.columns)
train_percent = 0.9
train_num = int(len(train) * train_percent)
test_num = len(train) - train_num
train_x = df_norm.head(train_num).as_matrix()
train_y = train[['TARGET','INVERSE_TARGET']].head(train_num).as_matrix()
test_x = df_norm.tail(test_num).as_matrix()
test_y = train[['TARGET','INVERSE_TARGET']].tail(test_num).as_matrix()
hashlib.md5(str.encode(str(df_use.columns))).hexdigest()
# Create an input layer which is similar to a feed_dict in TensorFlow.
# Note that the input-shape must be a tuple containing the image-size.
inputs = Input(shape=(columns,))
# Variable used for building the Neural Network.
net = inputs
# First fully-connected / dense layer with ReLU-activation.
w = 300
h = 4
for _ in range(h):
net = Dense(w, activation='relu')(net)
net = Dropout(0.5)(net)
net = Dense(2, activation='softmax')(net)
# Output of the Neural Network.
outputs = net
model = Model(inputs=inputs, outputs=outputs)
model.compile(optimizer='sgd', loss='categorical_crossentropy')
for i in range(1):
model.fit(x=train_x, y=train_y,validation_split=0.2,epochs=10, batch_size=128)
print('roc' + str(i),roc_auc_score(test_y[:,0],model.predict(test_x)[:,0]))
model.save('credit.keras')
json.dump({
'preprocess': preprocess,
'scale': lims,
'use': use
},open('credit.json', 'w'))
roc_auc_score(train_y[:,0],model.predict(train_x)[:,0])
```
| github_jupyter |
[](https://www.pythonista.io)
# *Marsmallow*.
https://marshmallow.readthedocs.io
## El proyecto *Marshmallow*.
```
!pip install marshmallow
```
### La clase ```marshmallow.Schema```.
### Campos.
https://marshmallow.readthedocs.io/en/stable/marshmallow.fields.html
### Validadores.
https://marshmallow.readthedocs.io/en/stable/marshmallow.validate.html
**Ejemplo:**
```
from marshmallow import Schema
from marshmallow.fields import String, Int, Float, Bool
from marshmallow.validate import Length, OneOf, Range
from json import dumps
from data import carreras
class AlumnoSchema(Schema):
cuenta = Int(required=True, validate=Range(min=1000000, max=9999999))
nombre = String(required=True, validate=Length(min=2, max=50))
primer_apellido = String(required=True, validate=Length(min=2, max=50))
segundo_apellido = String(required=False, validate=Length(min=2, max=50))
carrera = String(required=True, validate=OneOf(carreras))
semestre = Int(required=True, validate=Range(min=1, max=50))
promedio = Float(required=True, validate=Range(min=1, max=50))
al_corriente = Bool(required=True)
AlumnoSchema().load({
'cuenta': 1231221,
'al_corriente': False,
'carrera': 'Arquitectura',
'nombre': 'Pedro',
'primer_apellido': 'Solis',
'promedio': 7.8,
'semestre': 3,
'segundo_apellido': 'Cabañas'})
AlumnoSchema().load({
'cuenta': 1232210,
'al_corriente': False,
'carrera': 'Arquitectura',
'nombre': 'Pedro',
'primer_apellido': 'Solis',
'promedio': 7.8,
'semestre': 3,})
AlumnoSchema().load({
'cuenta': 1232210,
'al_corriente': False,
'carrera': 'Veterinaria',
'nombre': 'Pedro',
'primer_apellido': 'Solis',
'promedio': 7.8,
'semestre': 3,})
AlumnoSchema().load({
'cuenta': 1232210,
'al_corriente': False,
'carrera': 'Derecho',
'nombre': 'Pedro',
'primer_apellido': 'Solis',
'promedio': 7.8,
'semestre': 3,
'Género': 'M'})
from dataclasses import dataclass
@dataclass
class Alumno:
cuenta: int
nombre: str
primer_apellido: str
segundo_apellido: str
carrera: str
semestre: int
promedio: float
al_corriente: bool
alumno = Alumno(cuenta=1231221,
al_corriente=False,
carrera='Arquitectura',
nombre='Pedro',
primer_apellido='Solis',
promedio=7.8,
semestre=3,
segundo_apellido='Cabañas')
AlumnoSchema().dump(alumno)
```
## La clase ```marshmallow.Schema.Meta```.
https://marshmallow.readthedocs.io/en/stable/extending.html#custom-class-meta-options
## *Marshmallow-SQLAlchemy*.
https://marshmallow-sqlalchemy.readthedocs.io/en/latest/
### La clase ```marshmallow_sqlalchemy.SQLAlchemySchema```.
```
import sqlalchemy as sa
from sqlalchemy.ext.declarative import declarative_base
from sqlalchemy.orm import scoped_session, sessionmaker, relationship, backref
engine = sa.create_engine("sqlite:///data/alumnosql.sqlite")
session = scoped_session(sessionmaker(bind=engine))
Base = declarative_base()
class AlumnosSQL(Base):
__tablename__ = 'alumno'
cuenta = sa.Column(sa.Integer, primary_key=True)
nombre = sa.Column(sa.String(50))
primer_apellido = sa.Column(sa.String(50))
segundo_apellido = sa.Column(sa.String(50))
carrera = sa.Column(sa.String(50))
semestre = sa.Column(sa.Integer)
promedio = sa.Column(sa.Float)
al_corriente = sa.Column(sa.Boolean)
Base.metadata.create_all(engine)
from marshmallow_sqlalchemy import SQLAlchemySchema, auto_field
class AlumnoSchema(SQLAlchemySchema):
class Meta:
model = AlumnosSQL
load_instance = True
cuenta = auto_field()
nombre = auto_field()
primer_apellido = auto_field()
segundo_apellido = auto_field()
carrera = auto_field()
semestre = auto_field()
promedio = auto_field()
al_corriente = auto_field()
alumno = AlumnosSQL(cuenta=12347)
session.add(alumno)
session.commit()
alumno_schema = AlumnoSchema()
alumno_schema.dump(alumno)
nuevo_alumno = alumno_schema.load({
'cuenta': 12345,
'al_corriente': False,
'carrera': 'Derecho',
'nombre': 'Pedro',
'primer_apellido': 'Solis',
'promedio': 7.8,
'semestre': 3,},
session=session)
nuevo_alumno
session.add(nuevo_alumno)
session.commit()
%load_ext sql
%sql sqlite:///data/alumnosql.sqlite
%%sql
select * from alumno
```
### La clase ```marshmallow_sqlalchemy.SQLAlchemyAutoSchema```.
```
from marshmallow_sqlalchemy import SQLAlchemyAutoSchema
class AlumnoSchema2(SQLAlchemyAutoSchema):
class Meta:
model = AlumnosSQL
include_relationships = True
load_instance = True
alumno_schema2 = AlumnoSchema2()
alumno_schema2.dump(nuevo_alumno)
```
<p style="text-align: center"><a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Licencia Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />Esta obra está bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Licencia Creative Commons Atribución 4.0 Internacional</a>.</p>
<p style="text-align: center">© José Luis Chiquete Valdivieso. 2021.</p>
| github_jupyter |
```
# coding: utf-8
import os
import sys
from math import log
import numpy as np
from collections import Counter
incrementer = 0.000000001
def getFileContents(filename):
data = None
with open(filename, 'r') as f:
data = f.readlines()
return data
def getFileFromCommandLine():
filename = sys.argv[1]
return getFileContents(filename)
def splitWordTag(word_tag_pair):
splitted = word_tag_pair.split('/')
tag = splitted[-1]
word = '/'.join(splitted[:-1])
return word, tag
def getUniqueTags(tagged_data):
tags = {}
for line in tagged_data:
word_tag_pairs = line.strip().split(' ')
for word_tag_pair in word_tag_pairs:
word, tag = splitWordTag(word_tag_pair)
if tag in tags.keys():
tags[tag] += 1
else:
tags[tag] = 1
return tags
def getOpenProbabilities(tagged_data, all_tags_dict):
global incrementer
sentences_count = len(tagged_data)
open_tag_count_dict = {}
for line in tagged_data:
first_word_tag_pairs = line.strip().split(' ')[0]
word, tag = splitWordTag(first_word_tag_pairs)
if tag in open_tag_count_dict.keys():
open_tag_count_dict[tag] += 1
else:
open_tag_count_dict[tag] = 1
#increment all existing tags count to one
open_tag_count_dict.update((tag, occurances + incrementer) for tag, occurances in open_tag_count_dict.items())
sentences_count += (sentences_count*incrementer)
#add one to non-opening tags
for tag in all_tags_dict.keys():
try:
val = open_tag_count_dict[tag]
except KeyError as e:
open_tag_count_dict[tag] = incrementer
sentences_count += incrementer
open_tag_count_dict.update((tag, (occurances*1.0)/sentences_count) for tag, occurances in open_tag_count_dict.items())
return open_tag_count_dict
def getCloseProbabilities(tagged_data, all_tags_dict):
global incrementer
sentences_count = len(tagged_data)
close_tag_count_dict = {}
for line in tagged_data:
last_word_tag_pairs = line.strip().split(' ')[-1]
word, tag = splitWordTag(last_word_tag_pairs)
if tag in close_tag_count_dict.keys():
close_tag_count_dict[tag] += 1
else:
close_tag_count_dict[tag] = 1
#increment all existing tags count by one
close_tag_count_dict.update((tag, occurances + incrementer) for tag, occurances in close_tag_count_dict.items())
sentences_count += (sentences_count*incrementer)
#add one to non-closing tags
for tag in all_tags_dict.keys():
try:
val = close_tag_count_dict[tag]
except KeyError as e:
close_tag_count_dict[tag] = incrementer
sentences_count += incrementer
close_tag_count_dict.update((tag, (occurances*1.0)/sentences_count) for tag, occurances in close_tag_count_dict.items())
return close_tag_count_dict
def buildTransitionMatrix(tagged_data, tags_dict):
global incrementer
tags = tags_dict.keys()
tags.sort()
tags_index_dict = {}
tags_index_dict_reverse = {}
for index, tag in enumerate(tags):
tags_index_dict[tag] = index
tags_index_dict_reverse[index] = tag
tag_count = len(tags)
feature_tags = {'PAGE_SEP' : [], 'URLS' : [], 'NUMERICS' : []}
feature_counts = {'PAGE_SEP' : 0, 'URLS' : 0, 'NUMERICS' : 0}
#Change this line to np.ones for add 1 smoothing
transition_matrix = np.zeros(shape=(tag_count, tag_count))
for line in tagged_data:
prev_tag = None
word_tag_pairs = line.strip().split(' ')
for word_tag_pair in word_tag_pairs:
word, tag = splitWordTag(word_tag_pair)
if word.count('=') > 10 or word.count('_') > 10 or word.count('*') > 10 or word.count('-') > 10 or word.count('+') > 10:
feature_tags['PAGE_SEP'].append(tag)
feature_counts['PAGE_SEP'] += 1
elif any(word.lower().endswith(last) for last in ('.com', '.net', '.org', '.edu')) or word.startswith('http') or word.startswith('www.'):
feature_tags['URLS'].append(tag)
feature_counts['URLS'] += 1
elif [char.isdigit() for char in word].count(True) * 1.0 > len(word) * 0.4:
feature_tags['NUMERICS'].append(tag)
feature_counts['NUMERICS'] += 1
else:
pass
if prev_tag is not None:
transition_matrix[tags_index_dict[prev_tag]][tags_index_dict[tag]] += 1
prev_tag = tag
new_feature_tags = { 'PAGE_SEP' : 'xAMITx', 'URLS' : 'xAMITx', 'NUMERICS' : 'xAMITx' }
try:
for feature in feature_tags:
possible_tags = feature_tags[feature]
possible_tags_counter = Counter(possible_tags)
most_common_tags = possible_tags_counter.most_common(1)
if len(most_common_tags) > 0:
best_possible_tag, tag_count = most_common_tags[0]
if tag_count > feature_counts[feature] * 0.35:
new_feature_tags[feature] = best_possible_tag
except:
pass
transition_matrix = transition_matrix + incrementer
probability_transition_matrix = transition_matrix/transition_matrix.sum(axis=1, keepdims=True)
# print "Transition Values aree NaN : ", np.argwhere(np.isnan(probability_transition_matrix))
# probability_transition_matrix[np.isnan(probability_transition_matrix)] = incrementer
# probability_transition_matrix = np.log(probability_transition_matrix)
return probability_transition_matrix.tolist(), tags_index_dict, tags_index_dict_reverse, new_feature_tags
def getUniqueWords(tagged_data):
words = []
for line in tagged_data:
word_tag_pairs = line.strip().split(' ')
for word_tag_pair in word_tag_pairs:
word, tag = splitWordTag(word_tag_pair)
words.append(word)
return list(set(words))
def computeEmissionProbabilities(tagged_data, tags_dict):
global incrementer
tags = tags_dict.keys()
tags.sort()
words = getUniqueWords(tagged_data)
words.sort()
tags_index_dict = {}
for index, tag in enumerate(tags):
tags_index_dict[tag] = index
words_index_dict = {}
words_index_dict_reverse = {}
for index, word in enumerate(words):
words_index_dict[word] = index
words_index_dict_reverse[index] = word
tag_count = len(tags)
word_count = len(words)
# word_count + 1 => Last column for unseen words
emission_matrix = np.zeros(shape=(tag_count, word_count + 1))
for line in tagged_data:
prev_tag = None
word_tag_pairs = line.strip().split(' ')
for word_tag_pair in word_tag_pairs:
word, tag = splitWordTag(word_tag_pair)
emission_matrix[tags_index_dict[tag]][words_index_dict[word]] += 1
prev_tag = tag
#increment 1 in all the elements so that the last col for unseen words have non zero values
# emission_matrix = emission_matrix + incrementer
probability_emission_matrix = emission_matrix/emission_matrix.sum(axis=1, keepdims=True)
# print "Emission Values are NaN : ", np.argwhere(np.isnan(probability_emission_matrix))
probability_emission_matrix[np.isnan(probability_emission_matrix)] = incrementer
# probability_emission_matrix = np.log(probability_emission_matrix)
return probability_emission_matrix.tolist(), tags_index_dict, words_index_dict, words_index_dict_reverse
def printEmissionProbabilities(count):
counter = 0
global probability_emission_matrix, tags_index_dict, words_index_dict
word_count = len(words_index_dict.keys())
tag_count = len(tags_index_dict.keys())
for word, word_index in words_index_dict.iteritems():
for tag, tag_index in tags_index_dict.iteritems():
if probability_emission_matrix[tag_index][word_index] != 0:
print tag, " => ", word, ' => ', probability_emission_matrix[tag_index][word_index]
counter += 1
if counter > count:
return
def writeModelToFile(probability_transition_matrix, opening_probabilities, closing_probabilities, probability_emission_matrix, tags_index_dict, words_index_dict, new_feature_tags):
total_tags = len(tags_index_dict.keys())
total_words = len(words_index_dict.keys())
lineCounter = 7
text = ''
text += '---------------------TransitionMatrix---------------------' + '\n'
lineCounter += 1
tr_start_line_number = lineCounter
tr_end_line_number = tr_start_line_number
for row in range(len(probability_transition_matrix)):
row_text = ''
for col in range(len(probability_transition_matrix[0])):
row_text += str(probability_transition_matrix[row][col]) + '\t'
row_text = row_text.strip()
text += row_text + '\n'
tr_end_line_number += 1
text += '---------------------EmissionMatrix---------------------' + '\n'
em_start_line_number = tr_end_line_number + 1
em_end_line_number = em_start_line_number
for row in range(len(probability_emission_matrix)):
row_text = ''
for col in range(len(probability_emission_matrix[0])):
row_text += str(probability_emission_matrix[row][col]) + '\t'
row_text = row_text.strip()
text += row_text + '\n'
em_end_line_number += 1
text += '---------------------OpeningClosingProbabilities---------------------' + '\n'
oc_start_line_number = em_end_line_number + 1
oc_end_line_number = oc_start_line_number
for tag in opening_probabilities:
tag_details = tag + '\t' + str(opening_probabilities[tag]) + '\t' + str(closing_probabilities[tag]) + '\t' + str(tags_index_dict[tag]) + '\n'
text += tag_details
oc_end_line_number += 1
text += '---------------------Words---------------------' + '\n'
wi_start_line_number = oc_end_line_number + 1
wi_end_line_number = wi_start_line_number
for word in words_index_dict:
word_details = word + '\t' + str(words_index_dict[word]) + '\n'
text += word_details
wi_end_line_number += 1
text += '---------------------AdditionalFeatures---------------------' + '\n'
af_start_line_number = wi_end_line_number + 1
af_end_line_number = af_start_line_number
for feature_name in new_feature_tags.keys():
text += feature_name + '\t' + new_feature_tags[feature_name] + '\n'
af_end_line_number += 1
header = ''
header += 'total_tags:' + str(total_tags) + '\n'
header += 'total_words:' + str(total_words) + '\n'
header += 'tranistion_matrix:' + str(tr_start_line_number) + ':' + str(tr_end_line_number) + '\n'
header += 'emission_matrix:' + str(em_start_line_number) + ':' + str(em_end_line_number) + '\n'
header += 'open_close_probabilities:' + str(oc_start_line_number) + ':' + str(oc_end_line_number) + '\n'
header += 'word_indexes:' + str(wi_start_line_number) + ':' + str(wi_end_line_number) + '\n'
header += 'additional_features:' + str(af_start_line_number) + ':' + str(af_end_line_number) + '\n'
text = header + text
filename = 'hmmmodel.txt'
with open(filename, 'w') as output_file:
output_file.write(text)
if __name__ == '__main__':
tagged_data = getFileFromCommandLine()
# tagged_data = getFileContents('data/zh_train_tagged.txt')
tags_dict = getUniqueTags(tagged_data)
opening_probabilities = getOpenProbabilities(tagged_data, tags_dict)
closing_probabilities = getCloseProbabilities(tagged_data, tags_dict)
probability_transition_matrix, tags_index_dict, tags_index_dict_reverse, new_feature_tags = buildTransitionMatrix(tagged_data, tags_dict)
probability_emission_matrix, tags_index_dict, words_index_dict, words_index_dict_reverse = computeEmissionProbabilities(tagged_data, tags_dict)
writeModelToFile(probability_transition_matrix, opening_probabilities, closing_probabilities, probability_emission_matrix, tags_index_dict, words_index_dict, new_feature_tags)
print "Done"
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
!git clone https://github.com/AIZOOTech/FaceMaskDetection.git
import os
import sys
import numpy as np
import pandas as pd
import torch
import torchvision
import torch.nn as nn
import torch.nn.functional as F
import xml.etree.ElementTree as ET
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
import torch.optim as optim
import cv2
from tqdm import tqdm
from skimage import io
sys.path.append('/content/FaceMaskDetection/models')
class MaskDataset(Dataset):
def __init__(self, img_root_dir, xml_root_dir, transform = None):
self.ids = [x[:-4] for x in os.listdir(img_root_dir)]
self.labels = []
for xml_file in os.listdir(xml_root_dir):
label = ET.parse(os.path.join(xml_root_dir , xml_file)).getroot().find('object').find('name').text
if label == 'with_mask':
self.labels.append(1)
else:
self.labels.append(0)
self.img_root_dir = img_root_dir
self.xml_root_dir = xml_root_dir
self.transform = transform
def __len__(self):
return len(self.ids)
def __getitem__(self, index):
img_path = os.path.join(r"/content/drive/Shareddrives/deepmask/DeepMask/FaceMaskDataset/archive_2/images", self.ids[index] + '.png')
img = cv2.imread(img_path)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
height, width, _ = img.shape
image_resized = cv2.resize(img, (360,360))
image_np = image_resized / 255.0 # 归一化到0~1
image_exp = np.expand_dims(image_np, axis=0)
image_transposed = image_exp.transpose((0, 3, 1, 2))
input_img = torch.tensor(image_transposed).float()
#skimage.color.convert_colorspace()
y_label = torch.tensor(float(self.labels[index]))
return (input_img, y_label)
class Net(nn.Module):
def __init__(self, pre_model):
super().__init__()
self.pre_model = pre_model
def forward(self, x):
x = x.squeeze()
x = self.pre_model(x)
#print(x.shape())
x = F.sigmoid(nn.Linear(self.pre_model.fc.in_features, 2)(x))
return x
def load_data():
dataset = MaskDataset(img_root_dir = '/content/drive/Shareddrives/deepmask/DeepMask/FaceMaskDataset/archive_2/images',
xml_root_dir = '/content/drive/Shareddrives/deepmask/DeepMask/FaceMaskDataset/archive_2/annotations',
transform = None)#transforms.Compose([])#transforms.ToPILImage(),
#transforms.Resize((256, 256)),
#transforms.Grayscale(),
#transforms.ToTensor()]))
train_size = int(0.8 * len(dataset))
test_size = len(dataset) - train_size
train_dataset, test_dataset = torch.utils.data.random_split(dataset, [train_size, test_size])
train_loader = DataLoader(train_dataset, batch_size=64, shuffle=True)
test_loader = DataLoader(test_dataset, batch_size=64, shuffle=True)
return train_loader, test_loader
def train(model, train_loader, optimizer, epoch):
model.train()
correct = 0
for batch_idx, (data, taget) in tqdm(enumerate(train_loader)):
optimizer.zero_grad()
output = model(data)
loss = F.cross_entropy(output, target)
loss.backward()
optimizer.step()
if batch_idx % 100 == 0:
print('Train Epoch: {}\\tLoss: {:.6f}'.format(epoch, loss.item()))
accuracy = test(model, train_loader)
return accuracy
def test(model, test_loader):
model.eval()
correct = 0
with torch.no_grad():
for data, target in test_loader:
output = model(data)
pred = output.argmax(dim=1, keepdim=True)
correct += pred.eq(target.view_as(pred)).sum().item()
accuracy = 100. * correct / len(test_loader.dataset)
return accuracy
def main():
#num_epochs = 3
#learning_rate = 0.001
#train_loader, test_loader = load_data()
#model_path = '/content/FaceMaskDetection/models/model360.pth'
#model = torch.load(model_path)
#net = Net(model)
#optimizer = optim.SGD(net.parameters(), lr=learning_rate)
#train_acc_arr, test_acc_arr = [], []
#for epoch in range(num_epochs):
# train_acc = train(net, train_loader, optimizer, epoch)
## print('\nTrain set Accuracy: {:.0f}%\n'.format(train_acc))
# test_acc = test(net, test_loader)
# print('\nTrain set Accuracy: {:.0f}%\n'.format(test_acc))
# train_acc_arr.append(train_acc)
# test_acc_arr.append(test_acc)
#torch.save(net.state.dict(), "deepMaskModel.pt")
if __name__ == '__main__':
main()
import torch
import sys
import cv2
import numpy as np
import copy
import os
sys.path.append(r'C:\Users\shrey\hackUmass\FaceMaskDetection\models')
model_path = '/content/FaceMaskDetection/models/model360.pth'
model = torch.load(model_path)
model.eval()
for i in os.listdir(r"/content/drive/Shareddrives/deepmask/DeepMask/FaceMaskDataset/archive_2/images"):
img_path = os.path.join(r"/content/drive/Shareddrives/deepmask/DeepMask/FaceMaskDataset/archive_2/images", i)
img = cv2.imread(img_path)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
height, width, _ = img.shape
image_resized = cv2.resize(img, (360,360))
image_np = image_resized / 255.0 # 归一化到0~1
image_exp = np.expand_dims(image_np, axis=0)
image_transposed = image_exp.transpose((0, 3, 1, 2))
input_img = torch.tensor(image_transposed).float()
print(input_img.shape)
a,b = model.forward(input_img)
a = a.detach()
b = b.detach()
```
| github_jupyter |
```
import pyodbc
import pandas as pd
import config as cfg
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn.metrics import accuracy_score
from sklearn.cluster import KMeans
%matplotlib inline
import seaborn as sns
import matplotlib.pyplot as plt
cnxn = pyodbc.connect( 'DRIVER={ODBC Driver 13 for SQL Server};SERVER=' + cfg.mssql['server'] + ';DATABASE='
+ cfg.mssql['database'] + ';UID=' + cfg.mssql['username'] + ';PWD=' + cfg.mssql['password'] )
query = "SELECT * FROM BankView WHERE [State]='TX';"
data = pd.read_sql(query, cnxn, index_col='BankID')
data.head()
data['CrimeRate1000'].mean()
data['Population'].mean()
data.isnull().sum()
values = {'CrimeRate1000': data['CrimeRate1000'].mean(), 'Population': data['Population'].mean(), 'AvgRating' : data['AvgRating'].mean()}
data.fillna(value=values, inplace=True)
data.shape
```
## KMeans
```
feature_cols = [ 'Take', 'PDistance', 'Officers1000', 'FFLCount', 'AvgRating', 'CrimeRate1000']
X = data[feature_cols]
# calculate SC for K=2 through K=19
k_range = range(2, 20)
scores = []
for k in k_range:
km = KMeans(n_clusters=k, random_state=1)
km.fit(X)
scores.append(metrics.silhouette_score(X, km.labels_))
```
The silhouette score measures how close various clusters created are. A higher silhouette score is better as it means that we dont have too many overlapping clusters. This measure has a range of [-1, 1].
```
scores
# plot the results
plt.plot(k_range, scores)
plt.xlabel('Number of clusters')
plt.ylabel('Silhouette Coefficient')
plt.grid(True)
```
## Try K-Means with 2 clusters
```
# K-means with 2 clusters is overfit and NOT the best for this data, so try 6
km = KMeans(n_clusters=2, random_state=1)
km.fit(X)
data['cluster'] = km.labels_
data.sort_values('cluster')
data_X = X.copy()
data_X['cluster'] = km.labels_
centers = data_X.groupby('cluster').mean()
centers
data[data.cluster == 1].cluster.count()
# K-means cluster 6
km = KMeans(n_clusters=6, random_state=1)
km.fit(X)
data['cluster'] = km.labels_
data.groupby('cluster').Name.count()
# K-means cluster 10
km = KMeans(n_clusters=10, random_state=1)
km.fit(X)
data['cluster'] = km.labels_
data.groupby('cluster').Name.count()
```
## KMeans Centers
```
data_X = X.copy()
data_X['cluster'] = km.labels_
centers = data_X.groupby('cluster').mean()
centers
import numpy as np
colors = np.array(['red', 'green', 'blue', 'black', 'gray', 'purple'])
pd.scatter_matrix(X, c=colors[data.cluster], figsize=(10,10), s=100);
```
| github_jupyter |
# Effect of the sample size in cross-validation
In the previous notebook, we presented the general cross-validation framework
and how to assess if a predictive model is underfitting, overfitting, or
generalizing. Besides these aspects, it is also important to understand how
the different errors are influenced by the number of samples available.
In this notebook, we will show this aspect by looking a the variability of
the different errors.
Let's first load the data and create the same model as in the previous
notebook.
```
from sklearn.datasets import fetch_california_housing
housing = fetch_california_housing(as_frame=True)
data, target = housing.data, housing.target
target *= 100 # rescale the target in k$
```
<div class="admonition note alert alert-info">
<p class="first admonition-title" style="font-weight: bold;">Note</p>
<p class="last">If you want a deeper overview regarding this dataset, you can refer to the
Appendix - Datasets description section at the end of this MOOC.</p>
</div>
```
from sklearn.tree import DecisionTreeRegressor
regressor = DecisionTreeRegressor()
```
## Learning curve
To understand the impact of the number of samples available for training on
the generalization performance of a predictive model, it is possible to
synthetically reduce the number of samples used to train the predictive model
and check the training and testing errors.
Therefore, we can vary the number of samples in the training set and repeat
the experiment. The training and testing scores can be plotted similarly to
the validation curve, but instead of varying a hyperparameter, we vary the
number of training samples. This curve is called the **learning curve**.
It gives information regarding the benefit of adding new training samples
to improve a model's generalization performance.
Let's compute the learning curve for a decision tree and vary the
proportion of the training set from 10% to 100%.
```
import numpy as np
train_sizes = np.linspace(0.1, 1.0, num=5, endpoint=True)
train_sizes
```
We will use a `ShuffleSplit` cross-validation to assess our predictive model.
```
from sklearn.model_selection import ShuffleSplit
cv = ShuffleSplit(n_splits=30, test_size=0.2)
```
Now, we are all set to carry out the experiment.
```
from sklearn.model_selection import learning_curve
results = learning_curve(
regressor, data, target, train_sizes=train_sizes, cv=cv,
scoring="neg_mean_absolute_error", n_jobs=2)
train_size, train_scores, test_scores = results[:3]
# Convert the scores into errors
train_errors, test_errors = -train_scores, -test_scores
```
Now, we can plot the curve.
```
import matplotlib.pyplot as plt
plt.errorbar(train_size, train_errors.mean(axis=1),
yerr=train_errors.std(axis=1), label="Training error")
plt.errorbar(train_size, test_errors.mean(axis=1),
yerr=test_errors.std(axis=1), label="Testing error")
plt.legend()
plt.xscale("log")
plt.xlabel("Number of samples in the training set")
plt.ylabel("Mean absolute error (k$)")
_ = plt.title("Learning curve for decision tree")
```
Looking at the training error alone, we see that we get an error of 0 k$. It
means that the trained model (i.e. decision tree) is clearly overfitting the
training data.
Looking at the testing error alone, we observe that the more samples are
added into the training set, the lower the testing error becomes. Also, we
are searching for the plateau of the testing error for which there is no
benefit to adding samples anymore or assessing the potential gain of adding
more samples into the training set.
If we achieve a plateau and adding new samples in the training set does not
reduce the testing error, we might have reach the Bayes error rate using the
available model. Using a more complex model might be the only possibility to
reduce the testing error further.
## Summary
In the notebook, we learnt:
* the influence of the number of samples in a dataset, especially on the
variability of the errors reported when running the cross-validation;
* about the learning curve, which is a visual representation of the capacity
of a model to improve by adding new samples.
| github_jupyter |
<center>
<img src="https://gitlab.com/ibm/skills-network/courses/placeholder101/-/raw/master/labs/module%201/images/IDSNlogo.png" width="300" alt="cognitiveclass.ai logo" />
</center>
<h1 align=center><font size = 5>Assignment: SQL Notebook for Peer Assignment</font></h1>
Estimated time needed: **60** minutes.
## Introduction
Using this Python notebook you will:
1. Understand the Spacex DataSet
2. Load the dataset into the corresponding table in a Db2 database
3. Execute SQL queries to answer assignment questions
## Overview of the DataSet
SpaceX has gained worldwide attention for a series of historic milestones.
It is the only private company ever to return a spacecraft from low-earth orbit, which it first accomplished in December 2010.
SpaceX advertises Falcon 9 rocket launches on its website with a cost of 62 million dollars wheras other providers cost upward of 165 million dollars each, much of the savings is because Space X can reuse the first stage.
Therefore if we can determine if the first stage will land, we can determine the cost of a launch.
This information can be used if an alternate company wants to bid against SpaceX for a rocket launch.
This dataset includes a record for each payload carried during a SpaceX mission into outer space.
### Download the datasets
This assignment requires you to load the spacex dataset.
In many cases the dataset to be analyzed is available as a .CSV (comma separated values) file, perhaps on the internet. Click on the link below to download and save the dataset (.CSV file):
<a href="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBM-DS0321EN-SkillsNetwork/labs/module_2/data/Spacex.csv?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01" target="_blank">Spacex DataSet</a>
### Store the dataset in database table
**it is highly recommended to manually load the table using the database console LOAD tool in DB2**.
<img src = "https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBM-DS0321EN-SkillsNetwork/labs/module_2/images/spacexload.png">
Now open the Db2 console, open the LOAD tool, Select / Drag the .CSV file for the dataset, Next create a New Table, and then follow the steps on-screen instructions to load the data. Name the new table as follows:
**SPACEXDATASET**
**Follow these steps while using old DB2 UI which is having Open Console Screen**
**Note:While loading Spacex dataset, ensure that detect datatypes is disabled. Later click on the pencil icon(edit option).**
1. Change the Date Format by manually typing DD-MM-YYYY and timestamp format as DD-MM-YYYY HH\:MM:SS
2. Change the PAYLOAD_MASS\_\_KG\_ datatype to INTEGER.
<img src = "https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBM-DS0321EN-SkillsNetwork/labs/module_2/images/spacexload2.png">
**Changes to be considered when having DB2 instance with the new UI having Go to UI screen**
* Refer to this insruction in this <a href="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-DB0201EN-SkillsNetwork/labs/Labs_Coursera_V5/labs/Lab%20-%20Sign%20up%20for%20IBM%20Cloud%20-%20Create%20Db2%20service%20instance%20-%20Get%20started%20with%20the%20Db2%20console/instructional-labs.md.html?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01">link</a> for viewing the new Go to UI screen.
* Later click on **Data link(below SQL)** in the Go to UI screen and click on **Load Data** tab.
* Later browse for the downloaded spacex file.
<img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBM-DS0321EN-SkillsNetwork/labs/module_2/images/browsefile.png" width="800"/>
* Once done select the schema andload the file.
<img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBM-DS0321EN-SkillsNetwork/labs/module_2/images/spacexload3.png" width="800"/>
```
!pip install sqlalchemy==1.3.9
!pip install ibm_db_sa
!pip install ipython-sql
```
### Connect to the database
Let us first load the SQL extension and establish a connection with the database
```
%load_ext sql
```
**DB2 magic in case of old UI service credentials.**
In the next cell enter your db2 connection string. Recall you created Service Credentials for your Db2 instance before. From the **uri** field of your Db2 service credentials copy everything after db2:// (except the double quote at the end) and paste it in the cell below after ibm_db_sa://
<img src ="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-DB0201EN-SkillsNetwork/labs/FinalModule_edX/images/URI.jpg">
in the following format
**%sql ibm_db_sa://my-username:my-password@my-hostname:my-port/my-db-name**
**DB2 magic in case of new UI service credentials.**
<img src ="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBM-DS0321EN-SkillsNetwork/labs/module_2/images/servicecredentials.png" width=600>
* Use the following format.
* Add security=SSL at the end
**%sql ibm_db_sa://my-username:my-password@my-hostname:my-port/my-db-name?security=SSL**
```
# The code was removed by Watson Studio for sharing.
```
## Tasks
Now write and execute SQL queries to solve the assignment tasks.
### Task 1
##### Display the names of the unique launch sites in the space mission
```
%sql select unique LAUNCH_SITE from SPACEXTBL
```
### Task 2
##### Display 5 records where launch sites begin with the string 'CCA'
```
%sql select * from SPACEXTBL where LAUNCH_SITE like 'CCA%' limit 5
```
### Task 3
##### Display the total payload mass carried by boosters launched by NASA (CRS)
```
%sql select sum(payload_mass__kg_) from SPACEXTBL where customer = 'NASA (CRS)'
```
### Task 4
##### Display average payload mass carried by booster version F9 v1.1
```
%sql select avg(payload_mass__kg_) from SPACEXTBL where booster_version = 'F9 v1.1'
```
### Task 5
##### List the date when the first succesful landing outcome in ground pad was acheived.
*Hint:Use min function*
```
%sql select min(date) from SPACEXTBL where landing__outcome = 'Success (ground pad)'
```
### Task 6
##### List the names of the boosters which have success in drone ship and have payload mass greater than 4000 but less than 6000
```
%sql select booster_version from SPACEXTBL where landing__outcome = 'Success (drone ship)' and payload_mass__kg_ > 4000 and payload_mass__kg_ < 6000
```
### Task 7
##### List the total number of successful and failure mission outcomes
```
%sql select mission_outcome, count(*) from SPACEXTBL group by mission_outcome
```
### Task 8
##### List the names of the booster_versions which have carried the maximum payload mass. Use a subquery
```
%sql select booster_version from SPACEXTBL where payload_mass__kg_ = (select max(payload_mass__kg_) from SPACEXTBL)
```
### Task 9
##### List the records which will display the month names, failure landing_outcomes in drone ship ,booster versions, launch_site for the months in year 2015
```
%sql select monthname(date), landing__outcome, booster_version, launch_site from SPACEXTBL where year(date) = 2015
```
### Task 10
##### Rank the count of successful landing_outcomes between the date 2010-06-04 and 2017-03-20 in descending order.
```
%sql select landing__outcome, count(landing__outcome) as landing_outcome_count from SPACEXTBL where date between '2010-06-04' and '2017-03-20' group by landing__outcome order by count(landing__outcome) desc
```
### Reference Links
* <a href ="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-DB0201EN-SkillsNetwork/labs/Labs_Coursera_V5/labs/Lab%20-%20String%20Patterns%20-%20Sorting%20-%20Grouping/instructional-labs.md.html?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01&origin=www.coursera.org">Hands-on Lab : String Patterns, Sorting and Grouping</a>
* <a href="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-DB0201EN-SkillsNetwork/labs/Labs_Coursera_V5/labs/Lab%20-%20Built-in%20functions%20/Hands-on_Lab__Built-in_Functions.md.html?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01&origin=www.coursera.org">Hands-on Lab: Built-in functions</a>
* <a href="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-DB0201EN-SkillsNetwork/labs/Labs_Coursera_V5/labs/Lab%20-%20Sub-queries%20and%20Nested%20SELECTs%20/instructional-labs.md.html?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01&origin=www.coursera.org">Hands-on Lab : Sub-queries and Nested SELECT Statements</a>
* <a href="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-DB0201EN-SkillsNetwork/labs/Module%205/DB0201EN-Week3-1-3-SQLmagic.ipynb?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01">Hands-on Tutorial: Accessing Databases with SQL magic</a>
* <a href= "https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-DB0201EN-SkillsNetwork/labs/Module%205/DB0201EN-Week3-1-4-Analyzing.ipynb?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01">Hands-on Lab: Analyzing a real World Data Set</a>
## Author(s)
<h4> Lakshmi Holla </h4>
## Other Contributors
<h4> Rav Ahuja </h4>
## Change log
| Date | Version | Changed by | Change Description |
|------|--------|--------|---------|
| 2021-07-09 | 0.2 |Lakshmi Holla | Changes made in magic sql|
| 2021-05-20 | 0.1 |Lakshmi Holla | Created Initial Version |
## <h3 align="center"> © IBM Corporation 2021. All rights reserved. <h3/>
| github_jupyter |
## Some Math
Let's assume all objects are always centered at $x=0$ to simplify the FFT handling.
We need a few relations to understand the math.
1. The Fourier transform of a function like $x^2W(x)$ is $F[x^2W(x)] \propto \frac{d^2\hat{W}(k)}{dk^2}$.
2. The Fourier transform of a Gaussian is a Gaussian, which we can write generically as $\exp(-\alpha^2 k^2)$. Here $\alpha$ is related to the real-space FWHM of the profile via some constants we won't bother with.
3. A convolution in real-space is a product in Fourier space.
4. A weighted sum over a profile in real-space can be written as an integral in Fourier space.
This last relation is worth discussin in detail. Suppose we have an image $I(x)$, a weight function $W(x)$, and we want to compute the integral $f = \int dx I(x) W(x)$. This integral is actuall the value of the convolution of $I(x)$ with $W(x)$ at $x=0$,
$$
f = \int dx I(x) W(x) = \left. \int dx I(x) W(x - y)\right|_{y = 0}
$$
In Fourier space we can write this relation as
$$
f \propto \left.\int dk \hat{I}(k)\hat{W}(k) \exp(-iky)\right|_{y=0} = \int dk \hat{I}(k)\hat{W}(k)
$$
So this property combined item 1 above means we can write the weighted moments of an object in real-space as integrals in Fourier space over the weight function and its derivatives
$$
f \propto \int dk \hat{I}(k)\hat{W}(k)
$$
$$
<x^2> \propto \int dk \hat{I}(k)\frac{d^{2}\hat{W}(k)}{dk_x^2}
$$
$$
<xy> \propto \int dk \hat{I}(k)\frac{d^2\hat{W}(k)}{dk_x dk_y}
$$
$$
<y^2> \propto \int dk \hat{I}(k)\frac{d^2\hat{W}(k)}{dk_y^2}
$$
## What about the PSF?
So now let's assume we have an object, a PSF, and a weight function. Further, let's assume that the weight function is always bigger than the PSF and that the weight function is Gaussian.
In this case, we can immediately see that all of the derivatives of the weight function in Fourier space can be written as a product of some polynomial and the weight function iself. The constraint that the weight function be larger than the PSF means that $\alpha_{psf} < \alpha_{w}$. Finally, we have some object with $\alpha_g$.
In terms of the profile of $k$ we have the following situation illustrated in the plot below.
```
import proplot as plot
import numpy as np
def prof(k, a):
return np.log10(np.exp(-(a*k)**2))
k = np.logspace(-1, 1.5, 100)
apsf = 1
aw = 1.5
ag = 0.25
fig, axs = plot.subplots(figsize=(4, 4))
axs.semilogx(k, prof(k, np.sqrt(ag**2+apsf**2)), label='gal+psf')
axs.semilogx(k, prof(k, apsf), label='psf')
axs.semilogx(k, prof(k, ag), label='gal')
axs.semilogx(k, prof(k, aw), label='wgt')
axs.format(xlabel='log10[k]', ylabel='log10[f(k)]')
axs.legend()
```
From this plot you can see that even for real-space moments, as long as the Fourier transforms of the moment kernels are broader than PSF, we remove modes suppressed by the PSF. Thus we can set these suppressed modes (where the PSF amplitude cannot be deconvolved) in Fourier space to zero without harm.
| github_jupyter |
# Neural Networks
## 1. Neural Networks
In this section, we will implement backpropagation algorithm to learn the parameters for the neural network.
### 1.1 Visualizing the data
The data is the same as assignment 3, 5000 training examples, each contains a 20 pixel by 20 pixel grayscale image of the digit.
```
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
%matplotlib inline
from scipy.io import loadmat
data = loadmat('ex3data1.mat')
X = data["X"] # 5000x400 np array
y = data["y"] # 5000x1 np array (2d)
y = y.flatten() # change to (5000,) 1d array and
y[y==10] = 0 # in original data, 10 is used to represent 0
def displayData(X):
""" displays the 100 rows of digit image data stored in X in a nice grid.
It returns the figure handle fig, ax
"""
# form the big 10 x 10 matrix containing all 100 images data
# padding between 2 images
pad = 1
# initialize matrix with -1 (black)
wholeimage = -np.ones((20*10+9, 20*10+9))
# fill values
for i in range(10):
for j in range(10):
wholeimage[j*21:j*21+20, i*21:i*21+20] = X[10*i+j, :].reshape((20, 20))
fig, ax = plt.subplots(figsize=(6, 6))
ax.imshow(wholeimage.T, cmap=plt.cm.gray, vmin=-1, vmax=1)
ax.axis('off')
return fig, ax
# randomly select 100 data points to display
rand_indices = np.random.randint(0, 5000, size=100)
sel = X[rand_indices, :]
# display images
fig, ax = displayData(sel)
```
### 1.2 Model representation
Our neural network is shown in the following figure. It has 3 layers: an input layer, a hidden layer and an output layer. The neural network used contains 25 units in the 2nd layer and 10 output units (corresponding to 10 digit classes).

### 1.3 Feedforward and cost function
Recall that the cost function for the neural network (without regularization) is:
$$ J(\theta)=\frac{1}{m}\sum_{i=1}^{m} \sum_{k=1}^{K}[-y^{(i)}log((h_\theta(x^{(i)}))_k)-(1-y^{(i)})log(1-(h_\theta(x^{(i)}))_k)]$$
where $h_\theta(x^{(i)})$ is computed as shown in the above figure and K=10 is the total number of possible labels. Note that $h_\theta(x^{(i)})_k = a_k^{(3)}$ is the activation of the k-th output unit. Also, remember that whereas the original labels (in the variable y) were 0, 1, ..., 9, for the purpose of training a neural network, we need to recode the labels as vectors containing only values 0 or 1, so:
$$ y = \left[\matrix{1\\ 0\\ 0\\ \vdots\\ 0}\right], \left[\matrix{0\\ 1\\ 0\\ \vdots\\ 0}\right], ..., or \left[\matrix{0\\ 0\\ 0\\ \vdots\\ 1}\right] $$
#### Vectorization
Matrix dimensions:
$X_{wb}$: 5000 x 401
$\Theta^{(1)}$: 25 x 401
$\Theta^{(2)}$: 10 x 26
$a^{(2)}$: 5000 x 25 or 5000 x 26 after adding intercept terms
$a^{(3)} or H_\theta(x)$: 5000 x 10
$Y$: 5000 x 10
$$a^{(2)} = g(X_{wb}\Theta^{(1)^T})$$
$$ H_\theta(x) = a^{(3)} = g(a^{(2)}_{wb}\Theta^{(2)^T})$$
$$ H_\theta(x) = \left[\matrix{-(h_\theta(x^{(1)}))^T-\\ -(h_\theta(x^{(2)}))^T-\\ \vdots\\ -(h_\theta(x^{(m)}))^T-}\right] $$
$$ Y = \left[\matrix{-(y^{(1)})^T-\\ -(y^{(2)})^T-\\ \vdots\\ -(y^{(m)})^T-}\right] $$
Therefore, cost is:
$$ J(\theta)=\frac{1}{m} \sum_{matrix-elements} (-Y .* log(H_\theta(x))-(1-Y) .* log(1-H_\theta(x))) $$
Note the element wise multiplication (.*) and sum of all matrix elements in the above equation.
### 1.4 Regularized cost function
The cost function for neural networks with regularization is given by:
$$ J(\theta)=\frac{1}{m}\sum_{i=1}^{m} \sum_{k=1}^{K}[-y^{(i)}log((h_\theta(x^{(i)}))_k)-(1-y^{(i)})log(1-(h_\theta(x^{(i)}))_k)] + \frac{\lambda}{2m}\left[\sum_{j=1}^{25}\sum_{k=1}^{400}(\Theta_{j, k}^{(1)})^2 + \sum_{j=1}^{10}\sum_{k=1}^{25}(\Theta_{j, k}^{(2)})^2\right]$$
Note that even though the additional regularization term seems complicated with all the cascaded Sigma symbols, it is actually just the sum of all elements (after taking square) in the $\Theta$ matrix, one of them is 25 by 400, the other is 10 by 25 (recall that bias term is by convention not included in regularization). If your regularization parameter $\lambda$ is very very large, then all your $\Theta$ will converge to zero.
#### Vectorization
For the regularization term, there's actually nothing much to vectorize. Using elementwise self-multiplication then sum all elements in the result will do it:
$$ J(\theta)=\frac{1}{m} \sum_{matrix-elements} (-Y .* log(H_\theta(x))-(1-Y) .* log(1-H_\theta(x))) + \frac{\lambda}{2m} \left[\sum_{matrix-elements}(\Theta_{j, k}^{(1)} .* \Theta_{j, k}^{(1)})+\sum_{matrix-elements}(\Theta_{j, k}^{(2)} .* \Theta_{j, k}^{(2)})\right]$$
```
def sigmoid(z):
""" sigmoid(z) computes the sigmoid of z. z can be a number,
vector, or matrix.
"""
g = 1 / (1 + np.exp(-z))
return g
def nnCostFunction(nn_params, input_lsize, hidden_lsize, num_labels, X, y, lmd):
""" computes the cost and gradient of the neural network. The
parameters for the neural network are "unrolled" into the vector
nn_params and need to be converted back into the weight matrices.
The returned parameter grad should be a "unrolled" vector of the
partial derivatives of the neural network.
X should already include bias terms
Y is a 2d matrix
"""
# number of training samples
m, n = X.shape
# restore Theta1 and Theta2 from nn_params
Theta1 = nn_params[:hidden_lsize*(input_lsize+1)].reshape((hidden_lsize, input_lsize+1))
Theta2 = nn_params[hidden_lsize*(input_lsize+1):].reshape((num_labels, hidden_lsize+1))
# forward propagation
a2 = sigmoid(X @ Theta1.T)
a2_wb = np.concatenate((np.ones((m, 1)), a2), axis=1)
a3 = sigmoid(a2_wb @ Theta2.T) # i.e. H_theta
# Calculate cost
temp1 = -y * np.log(a3) - (1-y) * np.log(1-a3)
temp2 = np.sum((Theta1**2).flatten()) + np.sum((Theta2**2).flatten())
J = np.sum(temp1.flatten()) / m + lmd * temp2 / (2*m)
return J
# define input_lsize, hidden_lsize and numb_labels
input_lsize = 400
hidden_lsize = 25
num_labels = 10
m = len(y) # number of samples
# add bias terms to X
X_wb = np.concatenate((np.ones((m, 1)), X), axis=1)
# convert y to 2d matrix Y, 5000 by 10
# each row represents a sample, containing 0 or 1
Y = np.zeros((m, num_labels))
for i, v in enumerate(y):
# # NOTE: v=0 maps to position 9
# if v != 0:
# Y[i, v-1] = 1
# else:
# Y[i, 9] = 1
#print(Y[:100, :])
# using Python's zero-indexing convention
Y[i, v] = 1
# Load pre-calculated nn_params Theta1 and Theta2
# In ex4weights are 2 parameters:
# Theta1: 25 by 401
# Theta2: 10 by 26
# from scipy.io import loadmat
data = loadmat('ex3weights.mat')
Theta1 = data["Theta1"]
Theta2 = data["Theta2"]
# unroll Theta1 and Theta2 into nn_params
# NOTE: ndarray.flatten() will unroll by row, which does not match the A(:) behavior in MATLAB (by column)
# However, since the flattened data will be reshaped by ndarray,reshape(), which by default
# reshape by row, so you will actually get the original Theta1 and Theta2 back
# In summary, your flatten() and reshape() function should use the same order
# either both by numpy default, or both by 'F' order
nn_params = np.concatenate((Theta1.flatten(), Theta2.flatten()))
print(nn_params.shape) # should be (10285,)
# Regularization factor
lmd = 0
# Test nnCostFunction()
J = nnCostFunction(nn_params, input_lsize, hidden_lsize, num_labels, X_wb, Y, lmd)
print(J)
print("Expected ~0.287629")
# test cost function with reularization
lmd = 1
J = nnCostFunction(nn_params, input_lsize, hidden_lsize, num_labels, X_wb, Y, lmd)
print(J)
print("Expected around 0.383770")
```
## 2. Backpropagation
In this part, we implement the backpropagation algo to compute the gradient for the neural network cost function. Once this is done, we will be able to train the neural network by minimizing the cost function using an optimizer.
### 2.1 Sigmoid gradient
The gradient for the sigmoid function can be computed as:
$$ g'(z)=\frac{d}{dz}g(z)=g(z)(1-g(z))$$
where
$$g(z)=\frac{1}{1+e^{-z}}$$
For large values (both positive and negative) of z, the gradient should be close to 0. When z = 0, the gradient should be exactly 0.25.
```
def sigmoidGradient(z):
""" computes the gradient of the sigmoid function
evaluated at z. This should work regardless if z is a matrix or a
vector. In particular, if z is a vector or matrix, you should return
the gradient for each element.
"""
return sigmoid(z) * (1 - sigmoid(z))
# test sigmoidGradient(z)
z = np.array([-10, 0, 10])
print(sigmoidGradient(z))
```
### 2.2 Random initialization
When training neural networks, it is important to randomly initialize the parameters for symmetry breaking. Otherwise, the units in hidden layers will be identical to each other.
One effective strategy for random initialization is to randomly select values for $\Theta^{(l)}$ uniformly in the range $[-\epsilon_{init}, \epsilon_{init}]$. You should use $\epsilon_{init}=0.12$. This range of values ensures that the parameters are kept small and makes the learning more efficient.
```
def randInitializeWeights(L_in, L_out):
""" randomly initializes the weights of a layer with
L_in incoming connections and L_out outgoing connections.
Note that return variable W should be set to a matrix of size(L_out, 1 + L_in) as
the first column of W handles the "bias" terms.
"""
epsilon_init = 0.12
W = np.random.rand(L_out, 1+L_in) * 2 * epsilon_init - epsilon_init
return W
```
### 2.3 Backpropagation

Recall that the intuition behind the backpropagation algorithm is as follows. Given a training example (x(t); y(t)), we will first run a "forward pass" to compute all the activations throughout the network, including the output value of the hypothesis $h_\theta(x)$. Then, for each node $j$ in layer $l$, we would like to compute an "error term" $\delta_j^{(l)}$ that measures how much that node was "responsible" for any errors in the output.
For an output node, we can directly measure the difference between the network's activation and the true target value, and use that to define $\delta_j^{(3)}$(since layer 3 is the output layer). For the hidden units, you will compute
$\delta_j^{(l)}$ based on a weighted average of the error terms of the nodes in layer $(l + 1)$.
Detailed steps are as follows:
1) Perform a feedforward pass, computing the activations for Layers 2 and 3
2) For each output unit k in Layer 3 (the output layer), set
$$\delta_k^{(3)}=a_k^{(3)}-y_k$$
where $y_k\in[0,1]$ indicates whether the current training example belongs to class k or not.
3) For Layer 2, set
$$\delta^{(2)} = (\Theta^{(2)})^T\delta^{(3)}.*g'(z^{(2)})$$
4) Accumulate the gradient from this example using the following formula. Note that you should skip or remove $\delta_0^{(2)}$:
$$\Delta^{(l)}=\Delta^{(l)}+\delta^{(l+1)}(a^{(l)})^T$$
Do this for all training examples.
5) Obtain the gradient by dividing the accumulated gradients by m:
$$\frac{\partial}{\partial\Theta_{ij}^{(l)}}J(\Theta) = D_{ij}^{(l)} = \frac{1}{m}\Delta_{ij}^{(l)}$$
#### Vectorization
Here, we still use the full vectorization form that we used above, so we have:
$$\delta^{(3)}=a^{(3)}-y$$
$$\delta^{(2)} = \delta^{(3)}\Theta^{(2)}.*g'(z^{(2)})$$
$$\Delta^{(l)}=(\delta^{(l+1)})^Ta^{(l)}$$
where the matrix dimensions are as follows:
$X_{wb}, a^{(1)}$: 5000 x 401 with intercept terms
$a^{(2)}, \delta^{(2)}, z^{(2)}$: 5000 x 25, without intercept terms
$a^{(3)}, y, \delta^{(3)}$: 5000 x 10
$\Theta^{(1)}$: 25 x 401 (but intercept terms will remain unchanged in gradient descent)
$\Theta^{(2)}$: 10 x 26 (but intercept terms will remain unchanged in gradient descent)
### 2.4 Regularized Neural Networks
To account for regularization, we can add an additional term after computing the gradient using backpropagation.
The formula are as follows:
$$\frac{\partial}{\partial\Theta_{ij}^{(l)}}J(\Theta) = D_{ij}^{(l)} = \frac{1}{m}\Delta_{ij}^{(l)}\qquad for\; j=0$$
$$\frac{\partial}{\partial\Theta_{ij}^{(l)}}J(\Theta) = D_{ij}^{(l)} = \frac{1}{m}\Delta_{ij}^{(l)}+\frac{\lambda}{m}\Theta_{ij}^{(l)}\qquad for\; j=1$$
Note that you should not regularize the first column of $\Theta$.
```
def nnCostFunction2(nn_params, input_lsize, hidden_lsize, num_labels, X, y, lmd):
""" computes the cost and gradient of the neural network. The
parameters for the neural network are "unrolled" into the vector
nn_params and need to be converted back into the weight matrices.
The returned parameter grad should be a "unrolled" vector of the
partial derivatives of the neural network.
X should already include bias terms
Y is a 2d matrix
"""
# number of training samples
m, n = X.shape
# restore Theta1 and Theta2 from nn_params
Theta1 = nn_params[:hidden_lsize*(input_lsize+1)].reshape((hidden_lsize, input_lsize+1))
Theta2 = nn_params[hidden_lsize*(input_lsize+1):].reshape((num_labels, hidden_lsize+1))
# forward propagation
z2 = X @ Theta1.T
a2 = sigmoid(z2)
a2_wb = np.concatenate((np.ones((m, 1)), a2), axis=1)
a3 = sigmoid(a2_wb @ Theta2.T) # i.e. H_theta
# Calculate cost
temp1 = -y * np.log(a3) - (1-y) * np.log(1-a3)
temp2 = np.sum((Theta1**2).flatten()) + np.sum((Theta2**2).flatten())
J = np.sum(temp1.flatten()) / m + lmd * temp2 / (2*m)
# Calculate gradient
delta3 = a3 - y # 5000x10
delta2 = delta3 @ Theta2[:, 1:] * sigmoidGradient(z2) # 5000x25
DT2 = delta3.T @ a2_wb # 10x26
DT1 = delta2.T @ X # 25x401, X is a1
Theta1_grad = DT1 / m
Theta2_grad = DT2 / m
# print("Theta1.shape is {}".format(Theta1.shape))
# print("Theta2.shape is {}".format(Theta2.shape))
# print("Theta1_grad.shape is {}".format(Theta1_grad.shape))
# print("Theta2_grad.shape is {}".format(Theta2_grad.shape))
# adding regularization
Theta1_grad[:, 1:] += lmd * Theta1[:, 1:] / m
Theta2_grad[:, 1:] += lmd * Theta2[:, 1:] / m
# unroll gradients (note in numpy, default order is by row first)
grad = np.concatenate((Theta1_grad.flatten(), Theta2_grad.flatten()))
return J, grad
# test gradient without regularization
lmd = 0
debug_J, debug_grad = nnCostFunction2(nn_params, input_lsize, hidden_lsize, num_labels, X_wb, Y, lmd)
print(debug_grad[:10])
print("Expected: [ 6.18712766e-05 0.00000000e+00 0.00000000e+00 4.15336892e-09 \n" +
"-5.29868773e-08 1.42184272e-07 1.59715308e-06 -8.89999550e-07 \n" +
"-1.45513067e-06 -4.08953470e-07]")
# test gradient with regularization
lmd = 3
debug_J, debug_grad = nnCostFunction2(nn_params, input_lsize, hidden_lsize, num_labels, X_wb, Y, lmd)
print(debug_grad[:10])
print("Expected: [ 6.18712766e-05 -6.33744979e-12 1.31648811e-12 2.87621717e-14 \n" +
"3.09854983e-10 -3.45710507e-09 -2.85907272e-08 -1.54564033e-08 \n" +
"2.10275154e-08 1.92242492e-08]")
```
### 2.6 Learning parameters using 'minimize' function
```
from scipy.optimize import minimize
# initial conidition, 1d array
init_Theta1 = randInitializeWeights(input_lsize, hidden_lsize)
init_Theta2 = randInitializeWeights(hidden_lsize, num_labels)
init_nn_params = np.concatenate((init_Theta1.flatten(), init_Theta2.flatten()))
# run optimization
result = minimize(nnCostFunction2, init_nn_params, args=(input_lsize, hidden_lsize, num_labels, X_wb, Y, lmd),
method='TNC', jac=True, options={'disp': True})
print(result.x)
# Obtain Theta1 and Theta2 from result.x
nn_params = result.x
Theta1 = nn_params[:hidden_lsize*(input_lsize+1)].reshape((hidden_lsize, input_lsize+1))
Theta2 = nn_params[hidden_lsize*(input_lsize+1):].reshape((num_labels, hidden_lsize+1))
def predict(X, Theta1, Theta2):
""" predicts output given network parameters Theta1 and Theta2 in Theta.
The prediction from the neural network will be the label that has the largest output.
"""
a2 = sigmoid(X @ Theta1.T)
# add intercept terms to a2
m, n = a2.shape
a2_wb = np.concatenate((np.ones((m, 1)), a2), axis=1)
a3 = sigmoid(a2_wb @ Theta2.T)
# print(a3[:10, :])
# apply np.argmax to the output matrix to find the predicted label
# for that training sample
p = np.argmax(a3, axis=1)
# p[p==10] = 0
return p # this is a 1d array
# prediction accuracy
pred = predict(X_wb, Theta1, Theta2)
print(pred.shape)
accuracy = np.sum((pred==y).astype(int))/m*100
print('Training accuracy is {:.2f}%'.format(accuracy))
# randomly show 10 images and corresponding results
# randomly select 10 data points to display
rand_indices = np.random.randint(0, 5000, size=10)
sel = X[rand_indices, :]
for i in range(10):
# Display predicted digit
print("Predicted {} for this image: ".format(pred[rand_indices[i]]))
# display image
fig, ax = plt.subplots(figsize=(2, 2))
ax.imshow(sel[i, :].reshape(20, 20).T, cmap=plt.cm.gray, vmin=-1, vmax=1)
ax.axis('off')
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/SainiManisha/convnet-tutorial/blob/master/CNN_MNIST.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Convolutional Neural Networks
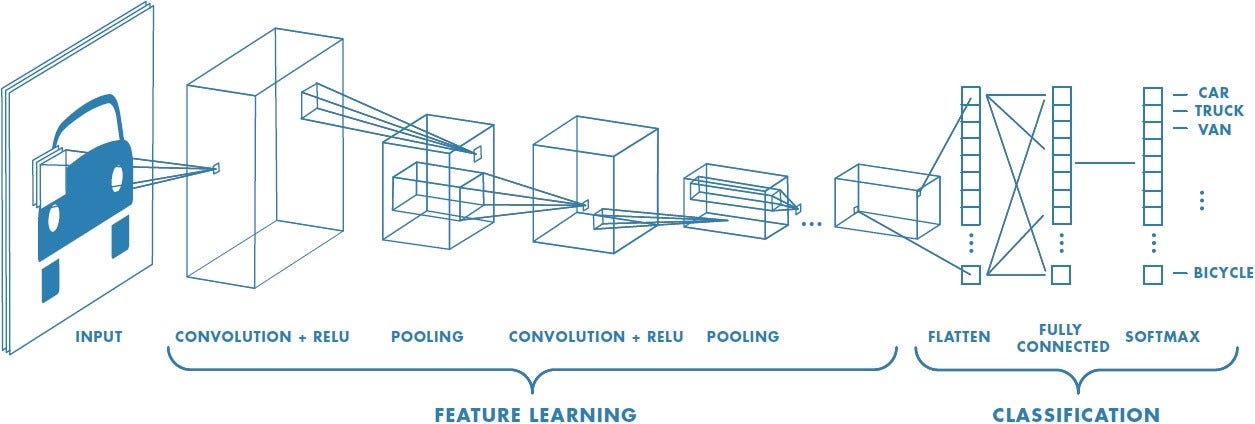
## Convolution
 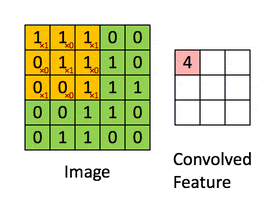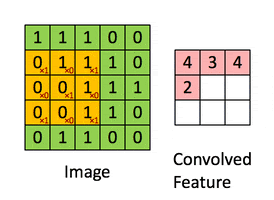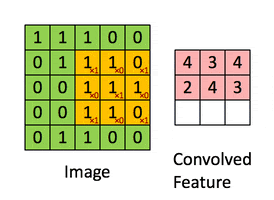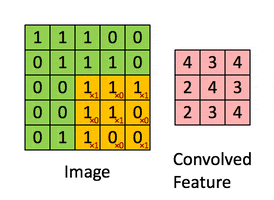
## Max Pooling

**Import the Library and Packages**
```
from matplotlib import pyplot as plt
import numpy as np
```
**Import Dataset**
```
from tensorflow.keras.datasets import mnist
from tensorflow.keras.utils import to_categorical
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
```
**Normalize the Data**
```
train_images = train_images.reshape((60000, 28, 28, 1))
train_images = train_images.astype('float32') / 255
test_images = test_images.reshape((10000, 28, 28, 1))
test_images = test_images.astype('float32') / 255
print('train_images shape',train_images.shape)
print('train_labels shape',train_labels.shape)
print('test_images shape',test_images.shape)
print('test_labels shape',test_labels.shape)
```
**Change the class Label using one hot encoding**
```
train_labels = to_categorical(train_labels)
test_labels = to_categorical(test_labels)
```
**Build the model for feature extraction**
```
#@title
from tensorflow.keras import layers
from tensorflow.keras import models
model = models.Sequential()
model.add(layers.Conv2D(32, (3, 3),
activation='relu',
name='C1',
input_shape=(28, 28, 1)))
model.add(layers.MaxPooling2D((2, 2),
name='M2'))
model.add(layers.Conv2D(64, (3, 3),
name='C3', activation='relu'))
model.add(layers.MaxPooling2D((2, 2), name='M4'))
model.add(layers.Conv2D(64, (3, 3),
activation='relu', name='C5'))
model.summary()
```

**Adding a classifier on top of the convnet**
```
model.add(layers.Flatten(name='F6'))
model.add(layers.Dense(64, activation='relu', name='FC7'))
model.add(layers.Dense(10, activation='softmax', name='FC8'))
model.summary()
```
**Compile and train the Model**
```
model.compile(optimizer='sgd',
loss='categorical_crossentropy',
metrics=['accuracy'])
history = model.fit(
train_images, train_labels,
epochs=10, batch_size=180,
validation_split=0.3)
```
**Evaluate the Model**
```
test_loss, test_acc = model.evaluate(test_images, test_labels, verbose=0)
test_acc
```
**Plot the Graph of Accuracy and Loss in case of Model**
```
import matplotlib.pyplot as plt
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.xlabel('Epoch')
plt.ylabel('Accuracy')
plt.title('Model Accuracy')
plt.legend(['Train', 'Val'],loc='lower right')
plt.show()
import matplotlib.pyplot as plt
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.title('Model loss')
plt.legend(['Train','val'], loc='lower right')
plt.show()
```
**Save the Model for reuse**
```
model.save('mnist_model.h5')
```
**Re-use the saved Model**
```
from tensorflow.keras.models import load_model
model = load_model('mnist_model.h5')
model.summary()
```
**Predict the class label of the test Images**
```
import matplotlib.pyplot as plt
import numpy as np
test_images_ss = test_images[:25]
predicted = model.predict(test_images_ss)
i = 0
plt.figure(figsize=[8, 8])
for (image, label) in zip(test_images_ss, predicted):
label = np.argmax(label)
image = image.reshape((28,28))
plt.subplot(5, 5, i + 1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(image, cmap=plt.cm.binary)
plt.xlabel(label)
i += 1
plt.suptitle("Predicted Labels")
plt.show()
```
**Visualizing the features learned by ConvNets**
```
from tensorflow.keras import models
layer_outputs = [layer.output for layer in model.layers[:5]]
activation_model = models.Model(inputs=model.input, outputs=layer_outputs)
activation_model.outputs
sample_test_image = np.expand_dims(train_images[10], axis=0)
sample_test_image.shape
activations = activation_model.predict(sample_test_image)
print(activations)
len(activations)
layer_names = [layer.name for layer in model.layers[:5]]
layer_names
col_size = 16
for layer_name, activation in zip(layer_names, activations):
max_val = activation.max()
activation /= max_val
num_filters = activation.shape[-1]
row_size = num_filters / col_size
print("\n" * 2)
print(layer_name)
plt.figure(figsize=[col_size * 2, row_size * 2])
for index in range(num_filters):
plt.subplot(row_size, col_size, index + 1)
plt.imshow(activation[0, :, :, index])
plt.axis("off")
plt.show()
```
**Confusion Matrix**
```
!pip install scikit-plot
from scikitplot.metrics import plot_confusion_matrix
logits = model.predict(test_images)
predicted = np.argmax(logits, axis=-1)
labels = np.argmax(test_labels, axis=-1)
plot_confusion_matrix(labels, predicted)
```
| github_jupyter |
# Tensorflow training
In this tutorial, you will train a mnist model in TensorFlow.
## Prerequisites
* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning (AML)
* Go through the [configuration notebook](../../../configuration.ipynb) to:
* install the AML SDK
* create a workspace and its configuration file (`config.json`)
[//]: # * Review the [tutorial](../train-hyperparameter-tune-deploy-with-tensorflow/train-hyperparameter-tune-deploy-with-tensorflow.ipynb) on single-node TensorFlow training using the SDK
```
# Check core SDK version number
import azureml.core
print("SDK version:", azureml.core.VERSION)
```
## Initialize workspace
Initialize a [Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace) object from the existing workspace you created in the Prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`.
```
from azureml.core.workspace import Workspace
ws = Workspace.from_config()
print('Workspace name: ' + ws.name,
'Azure region: ' + ws.location,
'Subscription id: ' + ws.subscription_id,
'Resource group: ' + ws.resource_group, sep='\n')
```
## Specify existing Kubernetes Compute
```
from azureml.core.compute import ComputeTarget, KubernetesCompute
from azureml.core.compute_target import ComputeTargetException
# choose a name for your Kubernetes compute
compute_name = 'gpucluster-1x'
compute_target = ComputeTarget(workspace=ws, name=compute_name)
compute_target
```
## Create a Dataset for Files
A Dataset can reference single or multiple files in your datastores or public urls. The files can be of any format. Dataset provides you with the ability to download or mount the files to your compute. By creating a dataset, you create a reference to the data source location. If you applied any subsetting transformations to the dataset, they will be stored in the dataset as well. The data remains in its existing location, so no extra storage cost is incurred. [Learn More](https://aka.ms/azureml/howto/createdatasets)
```
#initialize file dataset
from azureml.core.dataset import Dataset
web_paths = ['http://yann.lecun.com/exdb/mnist/train-images-idx3-ubyte.gz',
'http://yann.lecun.com/exdb/mnist/train-labels-idx1-ubyte.gz',
'http://yann.lecun.com/exdb/mnist/t10k-images-idx3-ubyte.gz',
'http://yann.lecun.com/exdb/mnist/t10k-labels-idx1-ubyte.gz'
]
dataset = Dataset.File.from_files(path = web_paths)
```
you may want to register datasets using the register() method to your workspace so they can be shared with others, reused across various experiments, and referred to by name in your training script.
```
#register dataset to workspace
dataset = dataset.register(workspace = ws,
name = 'mnist dataset for Arc',
description='training and test dataset',
create_new_version=True)
# list the files referenced by dataset
dataset
```
## Train model on the Kubernetes compute
### Create a project directory
Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script, and any additional files your training script depends on.
```
import os
script_folder = './tf-resume-training'
os.makedirs(script_folder, exist_ok=True)
```
Copy the training script `tf_mnist_with_checkpoint.py` into this project directory.
```
import shutil
# the training logic is in the tf_mnist_with_checkpoint.py file.
shutil.copy('./tf_mnist_with_checkpoint.py', script_folder)
shutil.copy('./utils.py', script_folder)
```
### Create an experiment
Create an [Experiment](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#experiment) to track all the runs in your workspace for this distributed TensorFlow tutorial.
```
from azureml.core import Experiment
experiment_name = 'akse-arc-tf-training1'
experiment = Experiment(ws, name=experiment_name)
```
### Create ScriptRun
```
from azureml.core.environment import Environment
from azureml.core.conda_dependencies import CondaDependencies
env = Environment("env-tf")
cd = CondaDependencies.create(pip_packages=['azureml-dataset-runtime[pandas,fuse]', 'azureml-defaults','tensorflow==1.13.1','horovod==0.16.1'])
env.docker.base_image='mcr.microsoft.com/azureml/openmpi4.1.0-cuda11.0.3-cudnn8-ubuntu18.04'
env.python.conda_dependencies = cd
# Register environment to re-use later
env.register(workspace = ws)
from azureml.core import ScriptRunConfig
args=['--data-folder', dataset.as_named_input('mnist').as_mount()]
src = ScriptRunConfig(source_directory=script_folder,
script='tf_mnist_with_checkpoint.py',
compute_target=compute_target,
environment=env,
arguments=args)
```
In the above code, we passed our training data reference `ds_data` to our script's `--data-folder` argument. This will 1) mount our datastore on the remote compute and 2) provide the path to the data zip file on our datastore.
### Submit job
### Run your experiment . Note that this call is asynchronous.
```
run = experiment.submit(src)
print(run)
run
```
### Monitor your run
You can monitor the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes.
| github_jupyter |
<img src="http://cfs22.simplicdn.net/ice9/new_logo.svgz "/>
# Assignment: Health Insurance Cost
*The comments/sections provided are your cues to perform the assignment. You don't need to limit yourself to the number of rows/cells provided. You can add additional rows in each section to add more lines of code.*
*If at any point in time you need help on solving this assignment, view our demo video to understand the different steps of the code.*
**Happy coding!**
* * *
## Health Insurance Cost
__DESCRIPTION__
Health insurance has become an indispensable part of our lives in recent years, and people are paying for it so that they are covered in the event of an accident or other unpredicted factors.
You are provided with medical costs dataset that has features such as Age, Cost, BMI.
__Objective:__
Determine the factors that contribute the most in the calculation of insurance costs.
Predict the health Insurance Cost.
__Actions to Perform:__
Find the correlation of every pair of features (and the outcome variable).
Visualize the correlations using a heatmap.
Normalize your inputs.
Use the test data to find out the accuracy of the model.
Visualize how your model uses the different features and which features have a greater effect.
```
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
%matplotlib inline
from sklearn.linear_model import LogisticRegression
import warnings
warnings.filterwarnings('ignore')
insuranceDF = pd.read_csv('insurance2.csv')
print(insuranceDF.head())
```
**Independent variables**
1. age : age of policyholder
2. sex: gender of policy holder (female=0, male=1)
3. bmi: Body mass index, ideally 18.5 to 25
4. children: number of children / dependents of policyholder
5. smoker: smoking state of policyholder (non-smoke=0;smoker=1)
6. region: the residential area of policyholder in the US (northeast=0, northwest=1, southeast=2, southwest=3)
7. charges: individual medical costs billed by health insurance
**Target variable**
1. insuranceclaim - categorical variable (0,1)
```
insuranceDF.info()
```
Let's start by finding correlation of every pair of features (and the outcome variable), and visualizing the correlations using a heatmap.
```
corr = insuranceDF.corr()
print(corr)
sns.heatmap(corr,
xticklabels=corr.columns,
yticklabels=corr.columns)
```
The dataset consists the records of 1338 patients in total. Using 1000 records for training and 300 records for testing, and the last 38 records to cross check your model.
```
dfTrain = insuranceDF[:1000]
dfTest = insuranceDF[1000:1300]
dfCheck = insuranceDF[1300:]
trainLabel = np.asarray(dfTrain['insuranceclaim'])
trainData = np.asarray(dfTrain.drop('insuranceclaim',1))
testLabel = np.asarray(dfTest['insuranceclaim'])
testData = np.asarray(dfTest.drop('insuranceclaim',1))
```
Before using machine learning, normalize the inputs. Machine Learning models often benefit substantially from input normalization. It also makes it easier to understand the importance of each feature later, when looking at the model weights. Normalize the data such that each variable has 0 mean and standard deviation of 1.
```
means = np.mean(trainData, axis=0)
stds = np.std(trainData, axis=0)
trainData = (trainData - means)/stds
testData = (testData - means)/stds
insuranceCheck = LogisticRegression()
insuranceCheck.fit(trainData, trainLabel)
```
Now, use test data to find out accuracy of the model.
```
accuracy = insuranceCheck.score(testData, testLabel)
print("accuracy = ", accuracy * 100, "%")
```
To get a better sense of what is going on inside the logistic regression model, visualize how your model uses the different features and which features have greater effect.
```
coeff = list(insuranceCheck.coef_[0])
labels = list(dfTrain.drop('insuranceclaim',1).columns)
features = pd.DataFrame()
features['Features'] = labels
features['importance'] = coeff
features.sort_values(by=['importance'], ascending=True, inplace=True)
features['positive'] = features['importance'] > 0
features.set_index('Features', inplace=True)
features.importance.plot(kind='barh', figsize=(11, 6),color = features.positive.map({True: 'blue', False: 'red'}))
plt.xlabel('Importance')
```
From the above figure,
1. BMI, Smoker have significant influence on the model, specially BMI.
2. Children have a negative influence on the prediction, i.e. higher number children / dependents are correlated with a policy holder who has not taken insurance claim.
3. Although age was more correlated than BMI to the output variables, the model relies more on BMI. This can happen for several reasons, including the fact that the correlation captured by age is also captured by some other variable, whereas the information captured by BMI is not captured by other variables.
Note that this above interpretations require that your input data is normalized. Without that, you can't claim that importance is proportional to weights.
| github_jupyter |
# **Assignment 5 Solutions** #
### **Q1. Implement a Union-Find Data Structure** ###
In lecture 6 we discussed Union-Find Data structures. The lecture was based on these [slides](https://www.cs.princeton.edu/courses/archive/spr09/cos226/lectures/01UnionFind.pdf). The slides contain Java code fo the Union-Find operations. For this assignment you should implement a Python class that implements a Union-Find Data Structure that uses weighting and path compression. Essentially this amounts to translating the code in the slides using the same specifications. Make sure you review the material and understand what the code does, and how it works.
```
# your implementation goes here
class UnionFind:
def __init__(self, n):
self.id = list(range(n))
self.sz = [1]*n
def root(self,i):
while (i != self.id[i]):
id[i] = id[id[i]]; #this is for path compression
i = self.id[i]
return i
def find(self,p,q):
return (self.root(p) == self.root(q))
def unite(self,p,q):
i = self.root(p)
j = self.root(q)
# weighted version
if self.sz[i]<self.sz[j]:
self.id[i] = j
self.sz[j] = self.sz[j]+self.sz[i]
else:
self.id[j] = i
self.sz[i] = self.sz[i]+self.sz[j]
S = UnionFind(5)
print(S.id)
print(S.sz)
print(S.find(3,2))
S.unite(3,2)
print(S.id)
print(S.sz)
```
### **Q2. Random Permutation**
Implement a function *randperm* that takes as input a number $n$, and returns a random permutation of the numbers [0...n-1]. This was covered in lecture 7. Your implementation should use $O(1)$ space in addition to the space needed for the output. (Note: you can use any random number generator functions from Python's *random* module, but you have to give you own implementation for randperm)
```
# your implementation goes here
import random
def randperm(n):
prm = list(range(n))
for j in range(n):
# random number in (0,n-j)
k = random.randrange(n-j)
tmp = prm[k]
prm[k] = prm[n-1-j]
prm[n-1-j] = tmp
return prm
randperm(5)
```
### **Q3. Adjacency matrices, powers, numpy** <br>
(this exercise should be useful for your mini-project)

Consider the above graph [(also here)](https://drive.google.com/file/d/1tIyXRGiQvMv-1EcJxzkQS2MpMNL9hUOA/view?usp=sharing).
The following exercise should be **repeated twice**. For the given directed graph and for the same graph where all edges have no directions.
**(a)** Create a numpy array containing the adjacency matrix $A$ for the graph.
**(b)** A sequence of nodes $v_1,v_2,...,v_k$ is called a walk on graph $G$, if $(v_i,v_{i+1})$ is an edge in $G$. The length of a walk with $k$ vertices is defined to be $k-1$. In the above graph pick a pair of nodes $(i,j)$, and report all different walks of length 3 from $i$ to $j$. (That is find, all the ways of going from $i$ to $j$ in 3 steps).
**(c)** Using numpy, calculate $A^3$, the third power of the adjacency matrix. Read the entry $(i,j)$ of this matrix. What do you observe?
```
# the following demonstrates the answer for the directed graph
# your code goes here
import numpy as np
A = np.zeros([10,10])
A[0,2] = 1
A[1,9] = 1
A[2,7] = 1
A[3,0] = 1
A[3,5] = 1
A[4,1] = 1
A[4,2] = 1
A[5,4] = 1
A[6,4] = 1
A[7,1] = 1
A[7,8] = 1
A[8,5] = 1
A[8,9] = 1
A[9,6] = 1
A[9,8] = 1
A3 = np.linalg.matrix_power(A,3)
# A3(i,j) is equal to the number of walks of length 3 between (i,j)
print(A3)
# the following demonstrates the answer for the undirected graph
# to avoid re-typing all opposite edges, I will use the matrix transpose
# transpose(A) is the array with the edges flipped
# then summing up the two matrices gives all edges
# because edge [8,9] has already both directions, its weight has to be halved in A_u
A_u = A + np.transpose(A)
A_u[8,9] = 1
A_u[9,8] = 1
A3_u = np.linalg.matrix_power(A_u,3)
print(A3_u)
```
### **Q4. A theoretical question**
Suppose a Python module contains an implementation of a function *maxSpanningTree(G)* that takes as input the adjacency list of graph $G$, with **positive** edge weights, and returns the edges of a maximum weight spanning tree. Further suppose that you can run this function, but you cannot access the code.
Explain how to use *maxSpanningTree(G)* in order to implement a *minSpanningTree(G)* function.
Make a copy $G'$ of $G$. Take all weights of $G'$, find the largest weight $w$, and then: negate all weights of $G'$ and add $w+1$. Running *maxSpanningTree(G)* on $G'$ will now give a tree $T'$, which contains the edges of a min spanning tree of $G$.
| github_jupyter |
```
import pandas as pd
from pathlib import Path
from sklearn.ensemble import GradientBoostingRegressor
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
from sklearn.model_selection import learning_curve,RepeatedKFold
from sklearn.pipeline import make_pipeline
from yellowbrick.model_selection import LearningCurve
from yellowbrick.regressor import ResidualsPlot
from yellowbrick.regressor import PredictionError
from sklearn.metrics import mean_squared_error,mean_absolute_error,r2_score
from sklearn import metrics
from sklearn.externals import joblib
from sklearn.feature_selection import VarianceThreshold
from sklearn.model_selection import GridSearchCV, KFold
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from scipy import stats
from scipy.special import boxcox1p
from sklearn.linear_model import Lasso
from sklearn.feature_selection import SelectFromModel
from sklearn.compose import TransformedTargetRegressor
from sklearn.preprocessing import QuantileTransformer,PowerTransformer
from sklearn.preprocessing import RobustScaler,MinMaxScaler,StandardScaler
from sklearn.manifold import TSNE
%matplotlib inline
#dataframe final
df_final = pd.read_csv("../data/DF_train15noChangeContact_skempiAB_modeller_final.csv",index_col=0)
pdb_names = df_final.index
features_names = df_final.drop('ddG_exp',axis=1).columns
df_final.shape
df_final["ddG_exp"].max() - df_final["ddG_exp"].min()
f, ax = plt.subplots(figsize=(10, 7))
sns.distplot(df_final['ddG_exp']);
plt.savefig("Train15_Distribution.png",dpi=300,bbox_inches="tight")
# Split train and independent test data
X_train, X_test, y_train, y_test = train_test_split(df_final.drop('ddG_exp',axis=1), df_final['ddG_exp'],
test_size=0.2, random_state=13)
f, ax = plt.subplots(figsize=(10, 7))
sns.distplot(y_train, color="red", label="ddG_exp_train");
sns.distplot(y_test, color="skyblue", label="ddG_exp_test");
sns.distplot(y_train, fit=stats.norm);
# Get the fitted parameters used by the function
(mu, sigma) = stats.norm.fit(y_train)
print( '\n mu = {:.2f} and sigma = {:.2f}\n'.format(mu, sigma))
#Now plot the distribution
plt.legend(['Normal dist. ($\mu=$ {:.2f} and $\sigma=$ {:.2f} )'.format(mu, sigma)],
loc='best')
plt.ylabel('Frequency')
plt.title('ddG distribution')
#Get also the QQ-plot
fig = plt.figure()
res = stats.probplot(y_train, plot=plt)
plt.show()
```
# probably need to transform target variable
## Correlation
```
# join train data for Exploratory analisis of training data
train = X_train.join(y_train)
sns.set(font_scale=0.6)
#correlation matrix
corrmat = train.corr()
f, ax = plt.subplots(figsize=(14, 11))
sns.heatmap(corrmat, square=True,cbar_kws={"shrink": .8});
#plt.savefig("Train15_initCorr.png",dpi=300,bbox_inches="tight")
sns.set(font_scale=1.2)
#top 10. correlation matrix
k = 15 #number of variables for heatmap
cols = corrmat.nlargest(k, 'ddG_exp')['ddG_exp'].index
cm = np.corrcoef(train[cols].values.T)
f, ax = plt.subplots(figsize=(10, 7))
sns.heatmap(cm, cbar=True, annot=True, square=True, fmt='.2f', annot_kws={'size': 10}, yticklabels=cols.values, xticklabels=cols.values);
#plt.savefig("Train15_initCorrTOP15.png",dpi=300,bbox_inches="tight")
sns.set(font_scale=1)
plt.subplots(figsize=(15, 5))
plt.subplot(1, 2, 1)
g = sns.regplot(x=train['van_der_waals_change'], y=train['ddG_exp'], fit_reg=False).set_title("Antes")
# Delete outliers
plt.subplot(1, 2, 2)
train = train.drop(train[(train['van_der_waals_change']>3)].index)
g = sns.regplot(x=train['van_der_waals_change'], y=train['ddG_exp'], fit_reg=False).set_title("Despues")
#plt.savefig("Train15_outlierVDWchange.png",dpi=600,bbox_inches="tight")
sns.set(font_scale=1)
plt.subplots(figsize=(15, 5))
plt.subplot(1, 2, 1)
g = sns.regplot(x=train['dg_change'], y=train['ddG_exp'], fit_reg=False).set_title("Antes")
# Delete outliers
plt.subplot(1, 2, 2)
train = train.drop(train[(train['dg_change'].abs()>8)].index)
g = sns.regplot(x=train['dg_change'], y=train['ddG_exp'], fit_reg=False).set_title("Despues")
#plt.savefig("Train15_outlierDgchange.png",dpi=600,bbox_inches="tight")
```
### NO missing values, skip this dataprocess
##
### Feature engeenering, checking interaction of sift contact with the highest correlated energetic feature
In order to treat this dataset, first I will check if adding interactions betwen some features improve corr, next i will check for skewess features. Finally i will write a custom transform class for every step.
```
y_train = train['ddG_exp']
X_train = train.drop('ddG_exp',axis=1)
```
### Check corr of new features
```
features_interaction_contactVDW = X_train.iloc[:,:15].mul(X_train["van_der_waals_change"],axis=0)# funciona mucho mejor
features_interaction_contactVDW.columns = features_interaction_contactVDW.columns.str[:]+"_vdw_change_interaction"
corr_matrix = features_interaction_contactVDW.corrwith(y_train,axis=0)#.abs()
#the matrix is symmetric so we need to extract upper triangle matrix without diagonal (k = 1)
print(corr_matrix.sort_values(ascending=False).round(6))
```
### Check skewness
```
skew_features = X_train.skew().sort_values(ascending=False)
skew_features
print(skew_features.to_csv())
```
### Check features by percetange of zero values
```
overfit = []
for i in X_train.columns:
counts = X_train[i].value_counts()
zeros = counts.iloc[0]
if zeros / len(X_train) * 100 >90.:
overfit.append(i)
print(overfit)
```
### make custom transformer for preprocess in pipeline
```
from sklearn.base import BaseEstimator, TransformerMixin
class FeaturesInteractions(BaseEstimator, TransformerMixin):
#Class constructor method that takes ..
def __init__(self, interaction1, interaction2 ):
self.interaction1 = interaction1
self.interaction2 = interaction2
#Return self nothing else to do here
def fit( self, X, y = None ):
return self
def transform(self, X , y=None ):
X_interactions = X.loc[:,self.interaction1].mul(X[self.interaction2],axis=0)
X_interactions.columns = X_interactions.columns.values+'/{}'.format(self.interaction2)
# set columns names
X = X.join(X_interactions)
return X
class SkewTransformer(BaseEstimator, TransformerMixin):
def __init__(self, threshold=0.6, method='quantile'):
self.threshold = threshold
self.method = method
#Return self nothing else to do here
def fit(self, X, y = None ):
skewes_ = X.skew().sort_values(ascending=False)
self.skew_features = skewes_[skewes_.abs() > self.threshold]
if self.method == 'quantile':
self.t = QuantileTransformer(output_distribution="normal",random_state=13)
self.t.fit(X[self.skew_features.index])
return self
def transform(self, X, y=None):
X[self.skew_features.index] = self.t.transform(X[self.skew_features.index])
return X
class ZeroThreshold(BaseEstimator, TransformerMixin):
def __init__(self, threshold=90.):
self.threshold = threshold
def fit(self, X, y = None ):
self.feature_names = X.columns
self.overfit = []
for i in X.columns:
counts = X[i].value_counts()
zeros = counts.iloc[0]
if zeros / len(X) * 100 >self.threshold:
self.overfit.append(i)
return self
def transform(self, X, y=None):
X.drop(self.overfit,axis=1,inplace=True)
return X
```
# Modeling
```
X_train.shape, y_train.shape, X_test.shape, y_test.shape
#1) ORIGINAL
## Pipeline preprocessing
interactions = FeaturesInteractions(interaction1=X_train.columns[:15].tolist(),interaction2="van_der_waals_change")
skewness = SkewTransformer(threshold=0.6,method='quantile')
zeroth = ZeroThreshold(threshold=90.)
#2)
rf_model = GradientBoostingRegressor(random_state=13)
#3) Crear pipeline
#pipeline1 = make_pipeline(interactions,skewness, zeroth, rf_model)
pipeline1 = make_pipeline(interactions,skewness,zeroth, rf_model)
# Use transformed target regressor
# regr_trans = TransformedTargetRegressor(regressor=pipeline1,
# transformer=PowerTransformer(output_distribution='normal',random_state=13))
# # grid params
# param_grid = {
# 'regressor__gradientboostingregressor__max_depth': [9],
# 'regressor__gradientboostingregressor__max_features': ['sqrt'],
# 'regressor__gradientboostingregressor__min_samples_leaf': [21],
# 'regressor__gradientboostingregressor__min_samples_split': [2],
# 'regressor__gradientboostingregressor__n_estimators': [200],
# 'regressor__gradientboostingregressor__subsample':[0.7],
# 'regressor__gradientboostingregressor__learning_rate':[0.05],
# 'regressor__gradientboostingregressor__loss':["huber"],
# 'regressor__gradientboostingregressor__alpha':[0.4]}
param_grid = {
'gradientboostingregressor__max_depth': [6],
'gradientboostingregressor__max_features': ['sqrt'],
'gradientboostingregressor__min_samples_leaf': [30],
'gradientboostingregressor__min_samples_split': [2],
'gradientboostingregressor__n_estimators': [100],
'gradientboostingregressor__subsample':[0.8],
'gradientboostingregressor__learning_rate':[0.05],
'gradientboostingregressor__loss':["huber"],
'gradientboostingregressor__alpha':[0.9]}
cv = RepeatedKFold(n_splits=10,n_repeats=10,random_state=13)
# Instantiate the grid search model
grid1 = GridSearchCV(pipeline1, param_grid, verbose=5, n_jobs=-1,cv=cv,scoring=['neg_mean_squared_error','r2'],
refit='neg_mean_squared_error',return_train_score=True)
grid1.fit(X_train,y_train)
# index of best scores
rmse_bestCV_test_index = grid1.cv_results_['mean_test_neg_mean_squared_error'].argmax()
rmse_bestCV_train_index = grid1.cv_results_['mean_train_neg_mean_squared_error'].argmax()
r2_bestCV_test_index = grid1.cv_results_['mean_test_r2'].argmax()
r2_bestCV_train_index = grid1.cv_results_['mean_train_r2'].argmax()
# scores
rmse_bestCV_test_score = grid1.cv_results_['mean_test_neg_mean_squared_error'][rmse_bestCV_test_index]
rmse_bestCV_test_std = grid1.cv_results_['std_test_neg_mean_squared_error'][rmse_bestCV_test_index]
rmse_bestCV_train_score = grid1.cv_results_['mean_train_neg_mean_squared_error'][rmse_bestCV_train_index]
rmse_bestCV_train_std = grid1.cv_results_['std_train_neg_mean_squared_error'][rmse_bestCV_train_index]
r2_bestCV_test_score = grid1.cv_results_['mean_test_r2'][r2_bestCV_test_index]
r2_bestCV_test_std = grid1.cv_results_['std_test_r2'][r2_bestCV_test_index]
r2_bestCV_train_score = grid1.cv_results_['mean_train_r2'][r2_bestCV_train_index]
r2_bestCV_train_std = grid1.cv_results_['std_train_r2'][r2_bestCV_train_index]
print('CV test RMSE {:f} +/- {:f}'.format(np.sqrt(-rmse_bestCV_test_score),np.sqrt(rmse_bestCV_test_std)))
print('CV train RMSE {:f} +/- {:f}'.format(np.sqrt(-rmse_bestCV_train_score),np.sqrt(rmse_bestCV_train_std)))
print('DIFF RMSE {}'.format(np.sqrt(-rmse_bestCV_test_score)-np.sqrt(-rmse_bestCV_train_score)))
print('CV test r2 {:f} +/- {:f}'.format(r2_bestCV_test_score,r2_bestCV_test_std))
print('CV train r2 {:f} +/- {:f}'.format(r2_bestCV_train_score,r2_bestCV_train_std))
print(r2_bestCV_train_score-r2_bestCV_test_score)
print("",grid1.best_params_)
y_test_pred = grid1.best_estimator_.predict(X_test)
y_train_pred = grid1.best_estimator_.predict(X_train)
print("\nRMSE for test dataset: {}".format(np.round(np.sqrt(mean_squared_error(y_test, y_test_pred)), 2)))
print("RMSE for train dataset: {}".format(np.round(np.sqrt(mean_squared_error(y_train, y_train_pred)), 2)))
print("pearson corr test {:f}".format(np.corrcoef(y_test_pred,y_test.values.ravel())[0][1]))
print("pearson corr train {:f}".format(np.corrcoef(y_train_pred,y_train.values.ravel())[0][1]))
print('R2 test',r2_score(y_test,y_test_pred))
print('R2 train',r2_score(y_train,y_train_pred))
CV test RMSE 1.629430 +/- 0.799343
CV train RMSE 1.278502 +/- 0.225925
DIFF RMSE 0.35092797677433385
CV test r2 0.243458 +/- 0.079707
CV train r2 0.537599 +/- 0.009211
0.29414037095192286
{'gradientboostingregressor__alpha': 0.9, 'gradientboostingregressor__learning_rate': 0.05, 'gradientboostingregressor__loss': 'huber', 'gradientboostingregressor__max_depth': 6, 'gradientboostingregressor__max_features': 'sqrt', 'gradientboostingregressor__min_samples_leaf': 30, 'gradientboostingregressor__min_samples_split': 2, 'gradientboostingregressor__n_estimators': 100, 'gradientboostingregressor__subsample': 0.8}
RMSE for test dataset: 1.64
RMSE for train dataset: 1.29
pearson corr 0.591408
R2 test 0.3178301850116666
R2 train 0.5310086844583202
visualizer = ResidualsPlot(grid1.best_estimator_,title='Residuos para GradientBoostingRegressor',hist=False)
visualizer.fit(X_train, y_train.values.ravel()) # Fit the training data to the model
visualizer.score(X_test, y_test.values.ravel()) # Evaluate the model on the test data
visualizer.finalize()
visualizer.ax.set_xlabel('Valor Predicho')
visualizer.ax.set_ylabel('Residuos')
plt.savefig("GBT_R2_train15.png",dpi=600,bbox_inches="tight")
#visualizer.poof() # Draw/show/poof the data
perror = PredictionError(grid1.best_estimator_, title='Error de Entrenamiento para GradientBoostingRegressor')
perror.fit(X_train, y_train.values.ravel()) # Fit the training data to the visualizer
perror.score(X_train, y_train.values.ravel()) # Evaluate the model on the test data
perror.finalize()
plt.savefig("GBT_TrainingError_train15.png",dpi=300,bbox_inches="tight")
perror = PredictionError(grid1.best_estimator_, title='Error de Predicción para GradientBoostingRegressor')
perror.fit(X_train, y_train.values.ravel()) # Fit the training data to the visualizer
perror.score(X_test, y_test.values.ravel()) # Evaluate the model on the test data
perror.finalize()
plt.savefig("GBT_PredictionError_train15.png",dpi=600,bbox_inches="tight")
#g = perror.poof()
full_data = pd.concat([X_train, X_test])
y_full = pd.concat([y_train, y_test])
viz = LearningCurve(grid1.best_estimator_, cv=cv, n_jobs=-1,scoring='neg_mean_squared_error',
train_sizes=np.linspace(0.2, 1.0, 10),title='Curva de aprendizaje para GradientBoostingRegressor')
viz.fit(full_data, y_full)
viz.finalize()
viz.ax.set_xlabel('Muestras de entrenamiento')
viz.ax.set_ylabel('Score')
plt.savefig("GBT_LearningCurve_train15.png",dpi=600,bbox_inches="tight")
#viz.poof()
print("RMSE CV Train {}".format(np.sqrt(-viz.train_scores_mean_[-1])))
print("RMSE CV Test {}".format(np.sqrt(-viz.test_scores_mean_[-1])))
np.sqrt(viz.test_scores_std_)
final_gbt = grid1.best_estimator_.fit(full_data,y_full)
# save final model
joblib.dump(final_gbt, 'GBTmodel_train15skempiAB_FINAL.overf.pkl')
feature_importance = final_gbt.named_steps['gradientboostingregressor'].feature_importances_
#feature_importance = feature_importance * 100.0 # * (feature_importance / feature_importance.max())
idx_features = feature_importance.argsort()[::-1]
fnames = final_gbt.named_steps.zerothreshold.feature_names.drop(final_gbt.named_steps.zerothreshold.overfit)
plt.figure(figsize=(15,4))
plt.bar(np.arange(len(fnames)), feature_importance[idx_features])
plt.xticks(range(len(fnames)),fnames[idx_features],rotation=90)
plt.autoscale(enable=True, axis='x', tight=True)
plt.xlabel(u"Importancia de caracteristicas")
plt.savefig("GBT_featureImportance.png",dpi=600,bbox_inches="tight")
importances = list(final_gbt.named_steps['gradientboostingregressor'].feature_importances_)
feature_list = fnames
# List of tuples with variable and importance
feature_importances = [(feature, round(importance, 4)) for feature, importance in zip(feature_list, importances)]
# Sort the feature importances by most important first
feature_importances = sorted(feature_importances, key = lambda x: x[1], reverse = True)
# Print out the feature and importances
[print('Variable: {:20} Importance: {}'.format(*pair)) for pair in feature_importances]
```
## Salvar modelo final, entrenado con el total de lso datos
```
full_prediction = final_gbt.predict(full_data)
full_pred_bin = np.where(np.abs(full_prediction) > 0.5,1,0)
full_true_bin = np.where(y_full > 0.5,1,0)
from sklearn.metrics import accuracy_score,classification_report,roc_auc_score,confusion_matrix
print(classification_report(full_true_bin,full_pred_bin))
rmse_test = np.round(np.sqrt(mean_squared_error(y_test, y_test_pred)), 3)
df_pred = pd.DataFrame({"Predicted ddG(kcal/mol)": y_test_pred, "Actual ddG(kcal/mol)": y_test.values.ravel()})
pearsonr_test = round(df_pred.corr().iloc[0,1],3)
g = sns.regplot(x="Actual ddG(kcal/mol)", y="Predicted ddG(kcal/mol)",data=df_pred)
plt.title("Predicted vs Experimental ddG (Independent set: 123 complexes)")
plt.text(-2,3,"pearsonr = %s" %pearsonr_test)
plt.text(4.5,-0.5,"RMSE = %s" %rmse_test)
#plt.savefig("RFmodel_300_testfit.png",dpi=600)
df_train_pred = pd.DataFrame({"Predicted ddG(kcal/mol)": y_train.values.ravel(), "Actual ddG(kcal/mol)": y_train_pred})
pearsonr_train = round(df_train_pred.corr().iloc[0,1],3)
rmse_train = np.round(np.sqrt(mean_squared_error(y_train.values.ravel(), y_train_pred)), 3)
g = sns.regplot(x="Actual ddG(kcal/mol)", y="Predicted ddG(kcal/mol)",data=df_train_pred)
plt.text(-0.4,6.5,"pearsonr = %s" %pearsonr_train)
plt.text(3.5,-2.5,"RMSE = %s" %rmse_train)
plt.title("Predicted vs Experimental ddG (Train set: 492 complexes)")
#plt.savefig("RFmodel_300_trainfit.png",dpi=600)
rf_model = grid1.best_estimator_.named_steps["randomforestregressor"]
importances = list(rf_model.feature_importances_)
feature_list = df_final.columns
# List of tuples with variable and importance
feature_importances = [(feature, round(importance, 4)) for feature, importance in zip(feature_list, importances)]
# Sort the feature importances by most important first
feature_importances = sorted(feature_importances, key = lambda x: x[1], reverse = True)
# Print out the feature and importances
[print('Variable: {:20} Importance: {}'.format(*pair)) for pair in feature_importances]
RepeatedKFold?
# Algorithms used for modeling
from sklearn.linear_model import ElasticNetCV, LassoCV, BayesianRidge, RidgeCV
from sklearn.ensemble import RandomForestRegressor, GradientBoostingRegressor, ExtraTreesRegressor
from sklearn.kernel_ridge import KernelRidge
from sklearn.model_selection import ShuffleSplit
from sklearn.svm import SVR
import xgboost as xgb
e_alphas = [0.0001, 0.0002, 0.0003, 0.0004, 0.0005, 0.0006, 0.0007]
e_l1ratio = [0.8, 0.85, 0.9, 0.95, 0.99, 1]
alphas_alt = [14.5, 14.6, 14.7, 14.8, 14.9, 15, 15.1, 15.2, 15.3, 15.4, 15.5]
alphas2 = [5e-05, 0.0001, 0.0002, 0.0003, 0.0004, 0.0005, 0.0006, 0.0007, 0.0008]
ridge = make_pipeline(MinMaxScaler(), RidgeCV(alphas=alphas_alt, cv=cv))
lasso = make_pipeline(MinMaxScaler(), LassoCV(max_iter=1e7, alphas=alphas2, random_state=42, cv=cv))
elasticnet = make_pipeline(MinMaxScaler(), ElasticNetCV(max_iter=1e7, alphas=e_alphas, cv=cv, l1_ratio=e_l1ratio))
svr = make_pipeline(MinMaxScaler(), SVR(C= 20, epsilon= 0.008, gamma=0.0003,))
gb = make_pipeline(GradientBoostingRegressor())
bayesianridge = make_pipeline(MinMaxScaler(),BayesianRidge())
rf = make_pipeline(RandomForestRegressor())
xgbr = make_pipeline(xgb.XGBRegressor())
#Machine Learning Algorithm (MLA) Selection and Initialization
models = [ridge, elasticnet, lasso, gb, bayesianridge, rf, xgbr]
# First I will use ShuffleSplit as a way of randomising the cross validation samples.
cvr = RepeatedKFold(n_splits=10,n_repeats=5,random_state=13)
#create table to compare MLA metrics
columns = ['Name', 'Parameters', 'Train Accuracy Mean', 'Test Accuracy']
before_model_compare = pd.DataFrame(columns = columns)
#index through models and save performance to table
row_index = 0
for alg in models:
#set name and parameters
model_name = alg.__class__.__name__
before_model_compare.loc[row_index, 'Name'] = model_name
before_model_compare.loc[row_index, 'Parameters'] = str(alg.get_params())
alg.fit(X_train, y_train.values.ravel())
#score model with cross validation: http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_validate.html#sklearn.model_selection.cross_validate
training_results = np.sqrt((-cross_val_score(alg, X_train, y_train.values.ravel(), cv = cvr, scoring= 'neg_mean_squared_error')).mean())
#training_results = cross_val_score(alg, X_train, y_train, cv = shuff, scoring= 'r2').mean()
test_results = np.sqrt(((y_test.values.ravel()-alg.predict(X_test))**2).mean())
#test_results = r2_score(y_pred=alg.predict(X_test),y_true=y_test)
before_model_compare.loc[row_index, 'Train Accuracy Mean'] = (training_results)#*100
before_model_compare.loc[row_index, 'Test Accuracy'] = (test_results)#*100
row_index+=1
print(row_index, alg.__class__.__name__, 'trained...')
decimals = 3
before_model_compare['Train Accuracy Mean'] = before_model_compare['Train Accuracy Mean'].apply(lambda x: round(x, decimals))
before_model_compare['Test Accuracy'] = before_model_compare['Test Accuracy'].apply(lambda x: round(x, decimals))
before_model_compare
```
| github_jupyter |
```
import identification_py2 as ob
import matplotlib.pyplot as plt
import glob #for returning files having the specified path extension
import os #checking for empty file
%pylab inline
```
###### Passing all the data into arrays
```
task_null = sorted(glob.glob('step_log_new/*/*task*.log')) #corresponds to .log files that has data related to the first position
control_null = sorted(glob.glob('step_log_new/*/*control*.log'))
task_remaining = sorted(glob.glob('step_log_new/*/*task*.log.*')) #corresponds to remaining log.'n' files
control_remaining = sorted(glob.glob('step_log_new/*/*control*.log.*'))
task_v = sorted(task_null + task_remaining) #set of all task_velocity logs
control_o = sorted(control_null + control_remaining) #set of all control logs
observations = len(task_null) #total number of experiments conducted/observations taken
positions = int(len(task_v) / observations) #number of points in the given task space
task_full = [] #A task_velocity list whose each element is a list of similar log files i.e from the same position
control_full = [] #A control_output list whose each element is a list of similar log files i.e from the same position
for i in range(0, positions):
task_full.append([])
control_full.append([])
for j in range(0, observations):
task_full[i].append(task_v[i + (j * positions)])
control_full[i].append(control_o[i + (j * positions)])
count = 0 #counter that returns the number of empty files
for i in range(0, positions):
for j in range(0, observations):
if os.stat(task_full[i][j]).st_size == 0:
count = count + 1
for i in range(0, positions):
for j in range(0, observations-count):
if os.stat(task_full[i][j]).st_size == 0:
del(task_full[i][j])
del(control_full[i][j])
# Reading all the data into a dataframe array
df_ist_soll = []
for i in range(0, positions):
df_ist_soll.append([])
for j in range(0, observations):
try:
df_ist_soll[i].append(ob.batch_read_data(control_full[i][j], task_full[i][j]))
except:
continue
```
###### Displaying all the observations
```
# The first try except code avoids the errors arising due to the already existing Overview directory.
# The second try except code avoids the errors resulting from the plotting of the empty data file
try:
os.makedirs('View_Data/')
except OSError, e:
if e.errno != os.errno.EEXIST:
raise
pass
for i in range(0, positions):
fig = plt.figure(figsize = (10,30))
fig.suptitle('Position %s'%(i + 1), fontsize = 20, fontweight = 'bold')
for j in range(0, observations):
try:
ax = fig.add_subplot(observations, 1, j + 1)
ax.set_title('Observation %s'%(j + 1))
plt.tight_layout(rect=[0, 0.03, 1, 0.95])
plt.plot(df_ist_soll[i][j])
except:
pass
plt.savefig('View_Data/Position %s.png'%(i + 1))
```
| github_jupyter |
```
def secondsconvert(seconds):
hours = seconds // 3600
minutes = (seconds - hours * 3600) // 60
remaining_seconds = seconds - hours * 3600 - minutes * 60
return hours, minutes, remaining_seconds
hours, minutes, remaining_seconds = secondsconvert(46790)
print(hours, minutes, remaining_seconds)
print(secondsconvert(1))
print(secondsconvert(890000))
def sec(s):
hours = s // 3600
mins = (s - hours * 3600) // 60
secs_remaining = s - hours * 3600 - mins * 60
return hours, mins, secs_remaining
print(sec(50000))
print(sec(50000))
def luckynumber(name):
number = len(name)*10
print("Hello " + name + ". Your lucky number is " + str (number))
luckynumber("andre")
luckynumber("Lisa")
def mytest (name):
number = len(name)*100
print("Hi " + name + ". My lucky number is " + str(number))
mytest("lisa")
def numbertest (seconds):
hours = seconds//3600
return hours
numbertest(56000)
def luckynum(name):
number = len(name)*10
print("Hi " + name + ". My lucky number is " + str(number))
luckynum("andre")
luckynum("andre")
def triangle_area(base, height):
return base*height/2
triangle_area(3,2)
def convert_seconds(seconds):
hrs = seconds // 3600
minutes = (seconds - hrs * 3600) // 60
remaining_secs = seconds - hrs * 3600 - minutes * 60
return hrs, minutes, remaining_secs
convert_seconds(4500)
convert_seconds(4800)
result = convert_seconds(400003)
result
def lnber (name):
number = len(name)*2
print("Hi "+ name + " my lucky number is " + str(number))
lnber("andre")
def l(name):
n = len("anthony")
print("Hi " + "i am "+ str(n) + " year old")
l("andre")
def n(name):
n = len(name)+2
return n +5
n("lisa")
print(1 > 0)
print(1 = 1)
print(1 == 1)
print(1 != 1)
print(1 + 1)
```
in python uppercase letters are sorted alphabetically before lowercase letters
```
def usernamelength (username):
if len(username) < 5 :
print("""not enough characters""")
else:
print("good to go")
usernamelength(andre)
usernamelength("andre")
usernamelength("faf")
unl_john = usernamelength("john")
def is_positivenumber (number):
if (number < 0):
return False
else:
return True
is_positivenumber(-1)
is_positivenumber(1)
is_positivenumber(-1)
is_positivenumber(9)
is_positivenumber(-5)
is_positivenumber(-9)
is_positivenumber(5)
is_positivenumber(5)
def namelenth(name):
if name > 5:
print("too short")
else:
print("good")
namelenth(andre)
namelenth("andre")
def namelenth(name):
if len(name) > 5:
return "greater than 5"
else:
return 5 + len(name)
lisa = namelenth("lisa")
lisa
def username(name):
if len(name) < 3:
print("username is too short")
else:
print("username is ok")
username("macdonald")
username("to")
def is_even(number):
if number % 2 == 0:
return True
return False
is_even(26)
is_even(23)
```
modulo operator used above
```
def iseven (number):
if number % 2 == 0:
return "Even"
else:
return "Odd"
iseven(6)
```
if i receive error about /.git/index.lock then remove the file by entering the following statement in the terminal
rm -f ./.git/index.lock
```
def iseven2(number):
if number % 2 == 1:
return "even"
elif number / 3 == 2:
return "what is this"
else:
return "odd"
iseven2(6)
iseven2(7)
iseven2(6)
iseven2(6)
```
IMPORTANT COMPARISON OPERATORS:
a == b: a is equal to b
a != b: a is different than b
a < b: a is smaller than b
a <= b: a is smaller or equal to b
a > b: a is bigger than b
a >= b: a is bigger or equal to b
```
def exam_score (score):
if score > 60:
grade = "pass"
elif score < 60 and score >50:
grade = "fail"
else:
grade = "do it over"
return grade
exam_score(45)
exam_score(55)
secondsconvert(3600)
def secondsconvert(seconds):
hours = seconds // 3600
minutes = (seconds - hours * 3600) // 60
remaining_seconds = seconds - hours * 3600 - minutes * 60
return hours, minutes, remaining_seconds
secondsconvert(6200)
secondsconvert(5400)
secondsconvert(5000)
```
| github_jupyter |
<a href="http://cocl.us/pytorch_link_top">
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/Pytochtop.png" width="750" alt="IBM Product " />
</a>
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/cc-logo-square.png" width="200" alt="cognitiveclass.ai logo" />
<h1>Differentiation in PyTorch</h1>
<h2>Objective</h2><ul><li> How to perform differentiation in pytorch.</li></ul>
<h2>Table of Contents</h2>
<p>In this lab, you will learn the basics of differentiation.</p>
<ul>
<li><a href="#Derivative">Derivatives</a></li>
<li><a href="#Partial_Derivative">Partial Derivatives</a></li>
</ul>
<p>Estimated Time Needed: <strong>25 min</strong></p>
<hr>
<h2>Preparation</h2>
The following are the libraries we are going to use for this lab.
```
# These are the libraries will be useing for this lab.
import torch
import matplotlib.pylab as plt
```
<!--Empty Space for separating topics-->
<h2 id="Derivative">Derivatives</h2>
Let us create the tensor <code>x</code> and set the parameter <code>requires_grad</code> to true because you are going to take the derivative of the tensor.
```
# Create a tensor x
x = torch.tensor(2.0, requires_grad = True)
print("The tensor x: ", x)
```
Then let us create a tensor according to the equation $ y=x^2 $.
```
# Create a tensor y according to y = x^2
y = x ** 2
print("The result of y = x^2: ", y)
```
Then let us take the derivative with respect x at x = 2
```
# Take the derivative. Try to print out the derivative at the value x = 2
y.backward()
print("The dervative at x = 2: ", x.grad)
```
The preceding lines perform the following operation:
$\\frac{\\mathrm{dy(x)}}{\\mathrm{dx}}=2x$
$\\frac{\\mathrm{dy(x=2)}}{\\mathrm{dx}}=2(2)=4$
```
print('data:',x.data)
print('grad_fn:',x.grad_fn)
print('grad:',x.grad)
print("is_leaf:",x.is_leaf)
print("requires_grad:",x.requires_grad)
print('data:',y.data)
print('grad_fn:',y.grad_fn)
print('grad:',y.grad)
print("is_leaf:",y.is_leaf)
print("requires_grad:",y.requires_grad)
```
Let us try to calculate the derivative for a more complicated function.
```
# Calculate the y = x^2 + 2x + 1, then find the derivative
x = torch.tensor(2.0, requires_grad = True)
y = x ** 2 + 2 * x + 1
print("The result of y = x^2 + 2x + 1: ", y)
y.backward()
print("The dervative at x = 2: ", x.grad)
```
The function is in the following form:
$y=x^{2}+2x+1$
The derivative is given by:
$\\frac{\\mathrm{dy(x)}}{\\mathrm{dx}}=2x+2$
$\\frac{\\mathrm{dy(x=2)}}{\\mathrm{dx}}=2(2)+2=6$
<!--Empty Space for separating topics-->
<h3>Practice</h3>
Determine the derivative of $ y = 2x^3+x $ at $x=1$
```
# Practice: Calculate the derivative of y = 2x^3 + x at x = 1
# Type your code here
```
Double-click <b>here</b> for the solution.
<!--
x = torch.tensor(1.0, requires_grad=True)
y = 2 * x ** 3 + x
y.backward()
print("The derivative result: ", x.grad)
-->
<!--Empty Space for separating topics-->
We can implement our own custom autograd Functions by subclassing
torch.autograd.Function and implementing the forward and backward passes
which operate on Tensors
```
class SQ(torch.autograd.Function):
@staticmethod
def forward(ctx,i):
"""
In the forward pass we receive a Tensor containing the input and return
a Tensor containing the output. ctx is a context object that can be used
to stash information for backward computation. You can cache arbitrary
objects for use in the backward pass using the ctx.save_for_backward method.
"""
result=i**2
ctx.save_for_backward(i)
return result
@staticmethod
def backward(ctx, grad_output):
"""
In the backward pass we receive a Tensor containing the gradient of the loss
with respect to the output, and we need to compute the gradient of the loss
with respect to the input.
"""
i, = ctx.saved_tensors
grad_output = 2*i
return grad_output
```
We can apply it the function
```
x=torch.tensor(2.0,requires_grad=True )
sq=SQ.apply
y=sq(x)
y
print(y.grad_fn)
y.backward()
x.grad
```
<h2 id="Partial_Derivative">Partial Derivatives</h2>
We can also calculate <b>Partial Derivatives</b>. Consider the function: $f(u,v)=vu+u^{2}$
Let us create <code>u</code> tensor, <code>v</code> tensor and <code>f</code> tensor
```
# Calculate f(u, v) = v * u + u^2 at u = 1, v = 2
u = torch.tensor(1.0,requires_grad=True)
v = torch.tensor(2.0,requires_grad=True)
f = u * v + u ** 2
print("The result of v * u + u^2: ", f)
```
This is equivalent to the following:
$f(u=1,v=2)=(2)(1)+1^{2}=3$
<!--Empty Space for separating topics-->
Now let us take the derivative with respect to <code>u</code>:
```
# Calculate the derivative with respect to u
f.backward()
print("The partial derivative with respect to u: ", u.grad)
```
the expression is given by:
$\\frac{\\mathrm{\\partial f(u,v)}}{\\partial {u}}=v+2u$
$\\frac{\\mathrm{\\partial f(u=1,v=2)}}{\\partial {u}}=2+2(1)=4$
<!--Empty Space for separating topics-->
Now, take the derivative with respect to <code>v</code>:
```
# Calculate the derivative with respect to v
print("The partial derivative with respect to u: ", v.grad)
```
The equation is given by:
$\\frac{\\mathrm{\\partial f(u,v)}}{\\partial {v}}=u$
$\\frac{\\mathrm{\\partial f(u=1,v=2)}}{\\partial {v}}=1$
<!--Empty Space for separating topics-->
Calculate the derivative with respect to a function with multiple values as follows. You use the sum trick to produce a scalar valued function and then take the gradient:
```
# Calculate the derivative with multiple values
x = torch.linspace(-10, 10, 10, requires_grad = True)
Y = x ** 2
y = torch.sum(x ** 2)
```
We can plot the function and its derivative
```
# Take the derivative with respect to multiple value. Plot out the function and its derivative
y.backward()
plt.plot(x.detach().numpy(), Y.detach().numpy(), label = 'function')
plt.plot(x.detach().numpy(), x.grad.detach().numpy(), label = 'derivative')
plt.xlabel('x')
plt.legend()
plt.show()
```
The orange line is the slope of the blue line at the intersection point, which is the derivative of the blue line.
The method <code> detach()</code> excludes further tracking of operations in the graph, and therefore the subgraph will not record operations. This allows us to then convert the tensor to a numpy array. To understand the sum operation <a href="https://pytorch.org/tutorials/beginner/blitz/autograd_tutorial.html">Click Here</a>
<!--Empty Space for separating topics-->
The <b>relu</b> activation function is an essential function in neural networks. We can take the derivative as follows:
```
# Take the derivative of Relu with respect to multiple value. Plot out the function and its derivative
x = torch.linspace(-10, 10, 1000, requires_grad = True)
Y = torch.relu(x)
y = Y.sum()
y.backward()
plt.plot(x.detach().numpy(), Y.detach().numpy(), label = 'function')
plt.plot(x.detach().numpy(), x.grad.detach().numpy(), label = 'derivative')
plt.xlabel('x')
plt.legend()
plt.show()
```
<!--Empty Space for separating topics-->
```
y.grad_fn
```
<h3>Practice</h3>
Try to determine partial derivative $u$ of the following function where $u=2$ and $v=1$: $ f=uv+(uv)^2$
```
# Practice: Calculate the derivative of f = u * v + (u * v) ** 2 at u = 2, v = 1
# Type the code here
```
Double-click **here** for the solution.
<!--
u = torch.tensor(2.0, requires_grad = True)
v = torch.tensor(1.0, requires_grad = True)
f = u * v + (u * v) ** 2
f.backward()
print("The result is ", u.grad)
-->
<!--Empty Space for separating topics-->
<a href="http://cocl.us/pytorch_link_bottom">
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/notebook_bottom%20.png" width="750" alt="PyTorch Bottom" />
</a>
<h2>About the Authors:</h2>
<a href="https://www.linkedin.com/in/joseph-s-50398b136/">Joseph Santarcangelo</a> has a PhD in Electrical Engineering, his research focused on using machine learning, signal processing, and computer vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.
Other contributors: <a href="https://www.linkedin.com/in/michelleccarey/">Michelle Carey</a>, <a href="www.linkedin.com/in/jiahui-mavis-zhou-a4537814a">Mavis Zhou</a>
## Change Log
| Date (YYYY-MM-DD) | Version | Changed By | Change Description |
| ----------------- | ------- | ---------- | ----------------------------------------------------------- |
| 2020-09-21 | 2.0 | Shubham | Migrated Lab to Markdown and added to course repo in GitLab |
<hr>
Copyright © 2018 <a href="cognitiveclass.ai?utm_source=bducopyrightlink&utm_medium=dswb&utm_campaign=bdu">cognitiveclass.ai</a>. This notebook and its source code are released under the terms of the <a href="https://bigdatauniversity.com/mit-license/">MIT License</a>.
| github_jupyter |
# Nieve bays implementation on simple dataset
```
from pprint import pprint
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import math
import operator
from math import sqrt, exp
from matplotlib import pyplot as plt
import os
#read in file and then give it feature headers
os.path.isfile('/Users/16786/Desktop/wine.data')
df= pd.read_csv('/Users/16786/Desktop/wine.data')
df.columns = [ 'Alcohol', 'Malic acid', 'Ash', 'alcalinity of ash', 'Magnesium', 'Total phenols', 'Flavanoids', 'Nonflavanoid phenols',
'Proanthocyanins', 'olor intensity', 'Hue', 'OD280', 'Proline', 'isTrain']
#df.columns = ['sepal length', 'sepal width', ' petal length', 'petal width', 'FlowerClass']
#seperate aproximently 25% out to be in the test set with which sample goes in each set being random
df['isTrain'] = np.random.uniform(0,1,len(df)) <= .75
df.head(100)
#adjusting the alcohol classes to start at 0 instead of 1.
for x in range (0,len(df)):
if df.iloc[x,0] == 1:
df.iloc[x,0] = 0
if df.iloc[x,0] == 2:
df.iloc[x,0] = 1
if df.iloc[x,0] == 3:
df.iloc[x,0] = 2
df.head(10)
features = df.columns[1:13]
#seperating train and test into two different dataframes for time effecency
train, test = df[df['isTrain']==True], df[df['isTrain']==False]
#lengths of each set this will change with every run
print('length of train set ' + str(len(train)))
print('length of test set '+ str(len(test)))
print((train[0:0+1]).iloc[0,4])
df.head(10)
#extracting mean, SD and variance of each columns contents based on its flowerclass for the P(Xi|y) claculation later
meanlist0 = np.mean(df.loc[df['Alcohol'] == 0])
meanlist1 = np.mean(df.loc[df['Alcohol'] == 1])
meanlist2 = np.mean(df.loc[df['Alcohol'] == 2])
meanlistAll = [meanlist0,meanlist1,meanlist2]
SDlist0 = np.std(df.loc[df['Alcohol'] == 0])
SDlist1 = np.std(df.loc[df['Alcohol'] == 1])
SDlist2 = np.std(df.loc[df['Alcohol'] == 2])
SDlistAll = [SDlist0,SDlist1,SDlist2]
varlist0 = SDlist0 * SDlist0
varlist1 = SDlist1 * SDlist1
varlist2 = SDlist2 * SDlist2
varlistAll = [varlist0,varlist1,varlist2]
#populate list of classifications
test_set_classifications = [0]
for n in range (0, len(test)-1):
test_set_classifications.append(-1)
#preforms a classification for each member in the test set and store the results in test_set_classifications
for x in range (0, len(test)-1):
productlist = np.ones(3)
#tests P(Xi|C)(P(C))/P(X) for each class C in the dataset and stores the resulting propablitys in the list productlist
for i in range(0, 3):
product = 1
#loop to calculate P(Xi|C) for each feature in the dataset and store the product in product
for j in range(1, len(features)):
#calculating term before the exponent in the Gaussian Naive Bayes P(Xi|C) equation
component1 = 1/sqrt(2*3.14*varlistAll[i][j])
#calculating the exponent in the Gaussian Naive Bayes P(Xi|C) equation
component2 = exp(-0.5 * pow(((test[x:x+1].iloc[0,j]) - meanlistAll[i][j]),2)/varlistAll[i][j])
product = product * component1 * component2
#multiplying by the prior probobility
product = product * (len(train['Alcohol'] == i)/len(train))
productlist[i] = product
#finds the index of the maximum value in the probability list then store that index in a list containing the classification of each test set sample the index represents class
test_set_classifications[x] = np.argmax(productlist)
correct = 0
print(test[0:1].iloc[0,0])
print(test_set_classifications[0])
#computes the accuracy of the model
for x in range (0, len(test)):
a = 0
if test_set_classifications[x] == test[x:x+1].iloc[0,0]:
correct += 1
# print(correct)
accuracy = correct/len(test)
print("The accuracy is ")
print(accuracy)
```
| github_jupyter |
# Part I. ETL Pipeline for Pre-Processing the Files
## PLEASE RUN THE FOLLOWING CODE FOR PRE-PROCESSING THE FILES
#### Import Python packages
```
# Import Python packages
import pandas as pd
import cassandra
import re
import os
import glob
import numpy as np
import json
import csv
from prettytable import PrettyTable
```
#### Creating list of filepaths to process original event csv data files
```
# checking your current working directory
print(os.getcwd())
# Get your current folder and subfolder event data
filepath = os.getcwd() + '/event_data'
# Create a for loop to create a list of files and collect each filepath
for root, dirs, files in os.walk(filepath):
# join the file path and roots with the subdirectories using glob
file_path_list = glob.glob(os.path.join(root,'*'))
print(file_path_list)
```
#### Processing the files to create the data file csv that will be used for Apache Casssandra tables
```
# initiating an empty list of rows that will be generated from each file
full_data_rows_list = []
# for every filepath in the file path list
for f in file_path_list:
# reading csv file
with open(f, 'r', encoding = 'utf8', newline='') as csvfile:
# creating a csv reader object
csvreader = csv.reader(csvfile)
next(csvreader)
# extracting each data row one by one and append it
for line in csvreader:
#print(line)
full_data_rows_list.append(line)
# uncomment the code below if you would like to get total number of rows
print(len(full_data_rows_list))
# uncomment the code below if you would like to check to see what the list of event data rows will look like
# print(full_data_rows_list)
# creating a smaller event data csv file called event_datafile_full csv that will be used to insert data into the \
# Apache Cassandra tables
csv.register_dialect('myDialect', quoting=csv.QUOTE_ALL, skipinitialspace=True)
with open('event_datafile_new.csv', 'w', encoding = 'utf8', newline='') as f:
writer = csv.writer(f, dialect='myDialect')
writer.writerow(['artist','firstName','gender','itemInSession','lastName','length',\
'level','location','sessionId','song','userId'])
for row in full_data_rows_list:
if (row[0] == ''):
continue
writer.writerow((row[0], row[2], row[3], row[4], row[5], row[6], row[7], row[8], row[12], row[13], row[16]))
# check the number of rows in your csv file
with open('event_datafile_new.csv', 'r', encoding = 'utf8') as f:
print(sum(1 for line in f))
```
# Part II. Complete the Apache Cassandra coding portion of your project.
## Now you are ready to work with the CSV file titled <font color=red>event_datafile_new.csv</font>, located within the Workspace directory. The event_datafile_new.csv contains the following columns:
- artist
- firstName of user
- gender of user
- item number in session
- last name of user
- length of the song
- level (paid or free song)
- location of the user
- sessionId
- song title
- userId
The image below is a screenshot of what the denormalized data should appear like in the <font color=red>**event_datafile_new.csv**</font> after the code above is run:<br>
<img src="images/image_event_datafile_new.jpg">
## Begin writing your Apache Cassandra code in the cells below
#### Creating a Cluster
```
# This should make a connection to a Cassandra instance your local machine
# (127.0.0.1)
host = '127.0.0.1'
from cassandra.cluster import Cluster
try:
cluster = Cluster([host]) #If you have a locally installed Apache Cassandra instance
# To establish connection and begin executing queries, need a session
session = cluster.connect()
except Exception as e:
print(e)
```
#### Create Keyspace
```
# Create a keyspace if it does not already exist
try:
session.execute("""
CREATE KEYSPACE IF NOT EXISTS btproject2
WITH REPLICATION =
{ 'class' : 'SimpleStrategy', 'replication_factor' : 1 }
"""
)
except Exception as e:
print(e)
```
#### Set Keyspace
```
try:
session.set_keyspace("btproject2")
except Exception as e:
print(e)
```
### Now we need to create tables to run the following queries. Remember, with Apache Cassandra you model the database tables on the queries you want to run.
## Create queries to ask the following three questions of the data
### 1. Give me the artist, song title and song's length in the music app history that was heard during sessionId = 338, and itemInSession = 4
### 2. Give me only the following: name of artist, song (sorted by itemInSession) and user (first and last name) for userid = 10, sessionid = 182
### 3. Give me every user name (first and last) in my music app history who listened to the song 'All Hands Against His Own'
```
# Creating table for Query 1
create_table_qry1 = "create table if not exists song_play_library "
create_table_qry1 = create_table_qry1 + "(session_id int, item_in_session int, artist_name text, song_title text, \
song_length float, primary key (session_id, item_in_session))"
try:
session.execute(create_table_qry1)
except Exception as e:
print(e)
# Confirm that the table is created by querying metadata
desc_qry = ("select * from system_schema.columns where keyspace_name = 'btproject2' and table_name = 'song_play_library'")
try:
rows = session.execute(desc_qry)
pt = PrettyTable(['Keyspace', 'Table', 'Column', 'Clustering Order', 'Kind', 'Position', 'Type'])
for row in rows:
pt.add_row([row.keyspace_name, row.table_name, row.column_name, row.clustering_order, row.kind, row.position, row.type])
print(pt)
except Exception as e:
print(e)
# Insert data into 'song_play_library' table from the combined CSV file picking the necessary columns
file = 'event_datafile_new.csv'
with open(file, encoding = 'utf8') as f:
csvreader = csv.reader(f)
next(csvreader) # skip header
for line in csvreader:
# print(type(line[8]), type(line[3]), type(line[0]), type(line[9]), type(line[5]))
insert_table_qry1 = "insert into song_play_library (session_id, item_in_session, artist_name, song_title, song_length) "
insert_table_qry1 = insert_table_qry1 + "values (%s, %s, %s, %s, %s)"
session.execute(insert_table_qry1, (int(line[8]), int(line[3]), line[0], line[9], float(line[5])))
```
#### Do a SELECT to verify that the data have been inserted into each table
```
# Query 1: Select the artist, song title and song's length in the music app history
# that was heard during sessionId = 338, and itemInSession = 4
select_table_qry1 = "select session_id, item_in_session, artist_name, song_title, song_length \
from song_play_library \
WHERE session_id = 338 \
and item_in_session = 4"
try:
rows = session.execute(select_table_qry1)
except Exception as e:
print(e)
pt = PrettyTable(['Session ID', 'Item in Session', 'Artist', 'Song', 'Length'])
for row in rows:
pt.add_row([row.session_id, row.item_in_session, row.artist_name, row.song_title, row.song_length])
print(pt)
```
### COPY AND REPEAT THE ABOVE THREE CELLS FOR EACH OF THE THREE QUESTIONS
```
# Creating table for Query 2
create_table_qry2 = "create table if not exists song_playlist_session "
create_table_qry2 = create_table_qry2 + "(user_id int, session_id int, item_in_session int, artist_name text, song_title text, \
user_fname text, user_lname text, primary key ((user_id, session_id), item_in_session))"
try:
session.execute(create_table_qry2)
except Exception as e:
print(e)
# Confirm that the table is created by querying metadata
desc_qry = ("select * from system_schema.columns where keyspace_name = 'btproject2' and table_name = 'song_playlist_session'")
try:
rows = session.execute(desc_qry)
pt = PrettyTable(['Keyspace', 'Table', 'Column', 'Clustering Order', 'Kind', 'Position', 'Type'])
for row in rows:
pt.add_row([row.keyspace_name, row.table_name, row.column_name, row.clustering_order, row.kind, row.position, row.type])
print(pt)
except Exception as e:
print(e)
# Insert data into 'song_playlist_session' table from the combined CSV file picking the necessary columns
file = 'event_datafile_new.csv'
with open(file, encoding = 'utf8') as f:
csvreader = csv.reader(f)
next(csvreader) # skip header
for line in csvreader:
## TO-DO: Assign the INSERT statements into the `query` variable
# print(type(line[10]), type(line[8]), type(line[3]), type(line[0]), type(line[9]), type(line[1]), type(line[4]))
insert_table_qry2 = "insert into song_playlist_session (user_id, session_id, item_in_session, artist_name, song_title, user_fname, user_lname) "
insert_table_qry2 = insert_table_qry2 + "values (%s, %s, %s, %s, %s, %s, %s)"
## TO-DO: Assign which column element should be assigned for each column in the INSERT statement.
## For e.g., to INSERT artist_name and user first_name, you would change the code below to `line[0], line[1]`
session.execute(insert_table_qry2, (int(line[10]), int(line[8]), int(line[3]), line[0], line[9], line[1], line[4]))
# Query 2: Select only the following: name of artist, song (sorted by itemInSession) and user (first and last name)
# for userid = 10, sessionid = 182
select_table_qry2 = "select artist_name,song_title, user_fname, user_lname from song_playlist_session WHERE user_id = 10 and session_id = 182"
try:
rows = session.execute(select_table_qry2)
except Exception as e:
print(e)
pt = PrettyTable(['Artist', 'Song', 'User First Name', 'User Last Name'])
for row in rows:
pt.add_row([row.artist_name, row.song_title, row.user_fname, row.user_lname])
print(pt)
# Create table for Query 3
create_table_qry3 = "create table if not exists user_play_library "
create_table_qry3 = create_table_qry3 + "(song_title text, user_id int, user_fname text, user_lname text, primary key (song_title, user_id))"
try:
session.execute(create_table_qry3)
except Exception as e:
print(e)
# Confirm that the table is created by querying metadata
desc_qry = ("select * from system_schema.columns where keyspace_name = 'btproject2' and table_name = 'user_play_library'")
try:
rows = session.execute(desc_qry)
pt = PrettyTable(['Keyspace', 'Table', 'Column', 'Clustering Order', 'Kind', 'Position', 'Type'])
for row in rows:
pt.add_row([row.keyspace_name, row.table_name, row.column_name, row.clustering_order, row.kind, row.position, row.type])
print(pt)
except Exception as e:
print(e)
# Insert data into 'user_play_library' table from the combined CSV file picking the necessary columns
file = 'event_datafile_new.csv'
with open(file, encoding = 'utf8') as f:
csvreader = csv.reader(f)
next(csvreader) # skip header
for line in csvreader:
## TO-DO: Assign the INSERT statements into the `query` variable
# print(type(line[10]), type(line[8]), type(line[3]), type(line[0]), type(line[9]), type(line[1]), type(line[4]))
insert_table_qry3 = "insert into user_play_library (song_title, user_id, user_fname, user_lname) "
insert_table_qry3 = insert_table_qry3 + "values (%s, %s, %s, %s)"
## TO-DO: Assign which column element should be assigned for each column in the INSERT statement.
## For e.g., to INSERT artist_name and user first_name, you would change the code below to `line[0], line[1]`
session.execute(insert_table_qry3, (line[9], int(line[10]), line[1], line[4]))
# Query 3: Select every user name (first and last) in the music app history who listened to the song 'All Hands Against His Own'
select_table_qry3 = "select user_fname, user_lname from user_play_library WHERE song_title = 'All Hands Against His Own'"
try:
rows = session.execute(select_table_qry3)
except Exception as e:
print(e)
pt = PrettyTable(['User First Name', 'User Last Name'])
for row in rows:
pt.add_row([row.user_fname, row.user_lname])
print(pt)
```
### Drop the tables before closing out the sessions
```
## TO-DO: Drop the table before closing out the sessions
table_list = ['song_play_library', 'user_play_library', 'song_playlist_session']
for table_name in table_list:
drop_table_qry = "drop table if exists " + table_name
try:
rows = session.execute(drop_table_qry)
except Exception as e:
print(e)
```
### Close the session and cluster connection¶
```
session.shutdown()
cluster.shutdown()
```
| github_jupyter |
## Image Cleaner Widget
fastai offers several widgets to support the workflow of a deep learning practitioner. The purpose of the widgets are to help you organize, clean, and prepare your data for your model. Widgets are separated by data type.
```
from fastai.vision import *
from fastai.widgets import DatasetFormatter, ImageCleaner, ImageDownloader, download_google_images
from fastai.gen_doc.nbdoc import *
%reload_ext autoreload
%autoreload 2
path = untar_data(URLs.MNIST_SAMPLE)
data = ImageDataBunch.from_folder(path)
learn = cnn_learner(data, models.resnet18, metrics=error_rate)
learn.fit_one_cycle(2)
learn.save('stage-1')
```
We create a databunch with all the data in the training set and no validation set (DatasetFormatter uses only the training set)
```
db = (ImageList.from_folder(path)
.split_none()
.label_from_folder()
.databunch())
learn = cnn_learner(db, models.resnet18, metrics=[accuracy])
learn.load('stage-1');
show_doc(DatasetFormatter)
```
The [`DatasetFormatter`](/widgets.image_cleaner.html#DatasetFormatter) class prepares your image dataset for widgets by returning a formatted [`DatasetTfm`](/vision.data.html#DatasetTfm) based on the [`DatasetType`](/basic_data.html#DatasetType) specified. Use `from_toplosses` to grab the most problematic images directly from your learner. Optionally, you can restrict the formatted dataset returned to `n_imgs`.
```
show_doc(DatasetFormatter.from_similars)
from fastai.gen_doc.nbdoc import *
from fastai.widgets.image_cleaner import *
show_doc(DatasetFormatter.from_toplosses)
show_doc(ImageCleaner)
```
[`ImageCleaner`](/widgets.image_cleaner.html#ImageCleaner) is for cleaning up images that don't belong in your dataset. It renders images in a row and gives you the opportunity to delete the file from your file system. To use [`ImageCleaner`](/widgets.image_cleaner.html#ImageCleaner) we must first use `DatasetFormatter().from_toplosses` to get the suggested indices for misclassified images.
```
ds, idxs = DatasetFormatter().from_toplosses(learn)
ImageCleaner(ds, idxs, path)
```
[`ImageCleaner`](/widgets.image_cleaner.html#ImageCleaner) does not change anything on disk (neither labels or existence of images). Instead, it creates a 'cleaned.csv' file in your data path from which you need to load your new databunch for the files to changes to be applied.
```
df = pd.read_csv(path/'cleaned.csv', header='infer')
# We create a databunch from our csv. We include the data in the training set and we don't use a validation set (DatasetFormatter uses only the training set)
np.random.seed(42)
db = (ImageList.from_df(df, path)
.split_none()
.label_from_df()
.databunch(bs=64))
learn = cnn_learner(db, models.resnet18, metrics=error_rate)
learn = learn.load('stage-1')
```
You can then use [`ImageCleaner`](/widgets.image_cleaner.html#ImageCleaner) again to find duplicates in the dataset. To do this, you can specify `duplicates=True` while calling ImageCleaner after getting the indices and dataset from `.from_similars`. Note that if you are using a layer's output which has dimensions <code>(n_batches, n_features, 1, 1)</code> then you don't need any pooling (this is the case with the last layer). The suggested use of `.from_similars()` with resnets is using the last layer and no pooling, like in the following cell.
```
ds, idxs = DatasetFormatter().from_similars(learn, layer_ls=[0,7,1], pool=None)
ImageCleaner(ds, idxs, path, duplicates=True)
show_doc(ImageDownloader)
```
[`ImageDownloader`](/widgets.image_downloader.html#ImageDownloader) widget gives you a way to quickly bootstrap your image dataset without leaving the notebook. It searches and downloads images that match the search criteria and resolution / quality requirements and stores them on your filesystem within the provided `path`.
Images for each search query (or label) are stored in a separate folder within `path`. For example, if you pupulate `tiger` with a `path` setup to `./data`, you'll get a folder `./data/tiger/` with the tiger images in it.
[`ImageDownloader`](/widgets.image_downloader.html#ImageDownloader) will automatically clean up and verify the downloaded images with [`verify_images()`](/vision.data.html#verify_images) after downloading them.
```
path = Config.data_path()/'image_downloader'
os.makedirs(path, exist_ok=True)
ImageDownloader(path)
```
#### Downloading images in python scripts outside Jupyter notebooks
```
path = Config.data_path()/'image_downloader'
files = download_google_images(path, 'aussie shepherd', size='>1024*768', n_images=30)
len(files)
show_doc(download_google_images)
```
After populating images with [`ImageDownloader`](/widgets.image_downloader.html#ImageDownloader), you can get a an [`ImageDataBunch`](/vision.data.html#ImageDataBunch) by calling `ImageDataBunch.from_folder(path, size=size)`, or using the data block API.
```
# Setup path and labels to search for
path = Config.data_path()/'image_downloader'
labels = ['boston terrier', 'french bulldog']
# Download images
for label in labels:
download_google_images(path, label, size='>400*300', n_images=50)
# Build a databunch and train!
src = (ImageList.from_folder(path)
.split_by_rand_pct()
.label_from_folder()
.transform(get_transforms(), size=224))
db = src.databunch(bs=16, num_workers=0)
learn = cnn_learner(db, models.resnet34, metrics=[accuracy])
learn.fit_one_cycle(3)
```
#### Downloading more than a hundred images
To fetch more than a hundred images, [`ImageDownloader`](/widgets.image_downloader.html#ImageDownloader) uses `selenium` and `chromedriver` to scroll through the Google Images search results page and scrape image URLs. They're not required as dependencies by default. If you don't have them installed on your system, the widget will show you an error message.
To install `selenium`, just `pip install selenium` in your fastai environment.
**On a mac**, you can install `chromedriver` with `brew cask install chromedriver`.
**On Ubuntu**
Take a look at the latest Chromedriver version available, then something like:
```
wget https://chromedriver.storage.googleapis.com/2.45/chromedriver_linux64.zip
unzip chromedriver_linux64.zip
```
Note that downloading under 100 images doesn't require any dependencies other than fastai itself, however downloading more than a hundred images [uses `selenium` and `chromedriver`](/widgets.image_cleaner.html#Downloading-more-than-a-hundred-images).
`size` can be one of:
```
'>400*300'
'>640*480'
'>800*600'
'>1024*768'
'>2MP'
'>4MP'
'>6MP'
'>8MP'
'>10MP'
'>12MP'
'>15MP'
'>20MP'
'>40MP'
'>70MP'
```
## Methods
## Undocumented Methods - Methods moved below this line will intentionally be hidden
```
show_doc(ImageCleaner.make_dropdown_widget)
show_doc(ImageCleaner.next_batch)
show_doc(DatasetFormatter.sort_idxs)
show_doc(ImageCleaner.make_vertical_box)
show_doc(ImageCleaner.relabel)
show_doc(DatasetFormatter.largest_indices)
show_doc(ImageCleaner.delete_image)
show_doc(ImageCleaner.empty)
show_doc(ImageCleaner.empty_batch)
show_doc(DatasetFormatter.comb_similarity)
show_doc(ImageCleaner.get_widgets)
show_doc(ImageCleaner.write_csv)
show_doc(ImageCleaner.create_image_list)
show_doc(ImageCleaner.render)
show_doc(DatasetFormatter.get_similars_idxs)
show_doc(ImageCleaner.on_delete)
show_doc(ImageCleaner.make_button_widget)
show_doc(ImageCleaner.make_img_widget)
show_doc(DatasetFormatter.get_actns)
show_doc(ImageCleaner.batch_contains_deleted)
show_doc(ImageCleaner.make_horizontal_box)
show_doc(DatasetFormatter.get_toplosses_idxs)
show_doc(DatasetFormatter.padded_ds)
```
## New Methods - Please document or move to the undocumented section
| github_jupyter |
```
from datascience import *
path_data = '../../data/'
import numpy as np
import matplotlib
matplotlib.use('Agg')
%matplotlib inline
import matplotlib.pyplot as plots
plots.style.use('fivethirtyeight')
```
# Prediction Intervals
One of the primary uses of regression is to make predictions for a new individual who was not part of our original sample but is similar to the sampled individuals. In the language of the model, we want to estimate $y$ for a new value of $x$.
Our estimate is the height of the true line at $x$. Of course, we don't know the true line. What we have as a substitute is the regression line through our sample of points.
The **fitted value** at a given value of $x$ is the regression estimate of $y$ based on that value of $x$. In other words, the fitted value at a given value of $x$ is the height of the regression line at that $x$.
```
baby = Table.read_table(path_data + 'baby.csv')
def standard_units(any_numbers):
"Convert any array of numbers to standard units."
return (any_numbers - np.mean(any_numbers))/np.std(any_numbers)
def correlation(t, x, y):
return np.mean(standard_units(t.column(x))*standard_units(t.column(y)))
def slope(table, x, y):
r = correlation(table, x, y)
return r * np.std(table.column(y))/np.std(table.column(x))
def intercept(table, x, y):
a = slope(table, x, y)
return np.mean(table.column(y)) - a * np.mean(table.column(x))
def fit(table, x, y):
a = slope(table, x, y)
b = intercept(table, x, y)
return a * table.column(x) + b
def residual(table, x, y):
return table.column(y) - fit(table, x, y)
def scatter_fit(table, x, y):
plots.scatter(table.column(x), table.column(y), s=20)
plots.plot(table.column(x), fit(table, x, y), lw=2, color='gold')
plots.xlabel(x)
plots.ylabel(y)
```
Suppose we try to predict a baby's birth weight based on the number of gestational days. As we saw in the previous section, the data fit the regression model fairly well and a 95% confidence interval for the slope of the true line doesn't contain 0. So it seems reasonable to carry out our prediction.
The figure below shows where the prediction lies on the regression line. The red line is at $x = 300$.
```
scatter_fit(baby, 'Gestational Days', 'Birth Weight')
s = slope(baby, 'Gestational Days', 'Birth Weight')
i = intercept(baby, 'Gestational Days', 'Birth Weight')
fit_300 = s*300 + i
plots.scatter(300, fit_300, color='red', s=20)
plots.plot([300,300], [0, fit_300], color='red', lw=2)
plots.ylim([0, 200]);
```
The height of the point where the red line hits the regression line is the fitted value at 300 gestational days.
The function `fitted_value` computes this height. Like the functions `correlation`, `slope`, and `intercept`, its arguments include the name of the table and the labels of the $x$ and $y$ columns. But it also requires a fourth argument, which is the value of $x$ at which the estimate will be made.
```
def fitted_value(table, x, y, given_x):
a = slope(table, x, y)
b = intercept(table, x, y)
return a * given_x + b
```
The fitted value at 300 gestational days is about 129.2 ounces. In other words, for a pregnancy that has a duration of 300 gestational days, our estimate for the baby's weight is about 129.2 ounces.
```
fit_300 = fitted_value(baby, 'Gestational Days', 'Birth Weight', 300)
fit_300
```
## The Variability of the Prediction
We have developed a method making one prediction of a new baby's birth weight based on the number of gestational days, using the data in our sample. But as data scientists, we know that the sample might have been different. Had the sample been different, the regression line would have been different too, and so would our prediction. To see how good our prediction is, we must get a sense of how variable the prediction can be.
To do this, we must generate new samples. We can do that by bootstrapping the scatter plot as in the previous section. We will then fit the regression line to the scatter plot in each replication, and make a prediction based on each line. The figure below shows 10 such lines, and the corresponding predicted birth weight at 300 gestational days.
```
x = 300
lines = Table(['slope','intercept'])
for i in range(10):
rep = baby.sample(with_replacement=True)
a = slope(rep, 'Gestational Days', 'Birth Weight')
b = intercept(rep, 'Gestational Days', 'Birth Weight')
lines.append([a, b])
lines['prediction at x='+str(x)] = lines.column('slope')*x + lines.column('intercept')
xlims = np.array([291, 309])
left = xlims[0]*lines[0] + lines[1]
right = xlims[1]*lines[0] + lines[1]
fit_x = x*lines['slope'] + lines['intercept']
for i in range(10):
plots.plot(xlims, np.array([left[i], right[i]]), lw=1)
plots.scatter(x, fit_x[i], s=30)
```
The predictions vary from one line to the next. The table below shows the slope and intercept of each of the 10 lines, along with the prediction.
```
lines
```
## Bootstrap Prediction Interval
If we increase the number of repetitions of the resampling process, we can generate an empirical histogram of the predictions. This will allow us to create an interval of predictions, using the same percentile method that we used create a bootstrap confidence interval for the slope.
Let us define a function called ``bootstrap_prediction`` to do this. The function takes five arguments:
- the name of the table
- the column labels of the predictor and response variables, in that order
- the value of $x$ at which to make the prediction
- the desired number of bootstrap repetitions
In each repetition, the function bootstraps the original scatter plot and finds the predicted value of $y$ based on the specified value of $x$. Specifically, it calls the function `fitted_value` that we defined earlier in this section to find the fitted value at the specified $x$.
Finally, it draws the empirical histogram of all the predicted values, and prints the interval consisting of the "middle 95%" of the predicted values. It also prints the predicted value based on the regression line through the original scatter plot.
```
# Bootstrap prediction of variable y at new_x
# Data contained in table; prediction by regression of y based on x
# repetitions = number of bootstrap replications of the original scatter plot
def bootstrap_prediction(table, x, y, new_x, repetitions):
# For each repetition:
# Bootstrap the scatter;
# get the regression prediction at new_x;
# augment the predictions list
predictions = make_array()
for i in np.arange(repetitions):
bootstrap_sample = table.sample()
bootstrap_prediction = fitted_value(bootstrap_sample, x, y, new_x)
predictions = np.append(predictions, bootstrap_prediction)
# Find the ends of the approximate 95% prediction interval
left = percentile(2.5, predictions)
right = percentile(97.5, predictions)
# Prediction based on original sample
original = fitted_value(table, x, y, new_x)
# Display results
Table().with_column('Prediction', predictions).hist(bins=20)
plots.xlabel('predictions at x='+str(new_x))
plots.plot(make_array(left, right), make_array(0, 0), color='yellow', lw=8);
print('Height of regression line at x='+str(new_x)+':', original)
print('Approximate 95%-confidence interval:')
print(left, right)
bootstrap_prediction(baby, 'Gestational Days', 'Birth Weight', 300, 5000)
```
The figure above shows a bootstrap empirical histogram of the predicted birth weight of a baby at 300 gestational days, based on 5,000 repetitions of the bootstrap process. The empirical distribution is roughly normal.
An approximate 95% prediction interval of scores has been constructed by taking the "middle 95%" of the predictions, that is, the interval from the 2.5th percentile to the 97.5th percentile of the predictions. The interval ranges from about 127 to about 131. The prediction based on the original sample was about 129, which is close to the center of the interval.
## The Effect of Changing the Value of the Predictor
The figure below shows the histogram of 5,000 bootstrap predictions at 285 gestational days. The prediction based on the original sample is about 122 ounces, and the interval ranges from about 121 ounces to about 123 ounces.
```
bootstrap_prediction(baby, 'Gestational Days', 'Birth Weight', 285, 5000)
```
Notice that this interval is narrower than the prediction interval at 300 gestational days. Let us investigate the reason for this.
The mean number of gestational days is about 279 days:
```
np.mean(baby.column('Gestational Days'))
```
So 285 is nearer to the center of the distribution than 300 is. Typically, the regression lines based on the bootstrap samples are closer to each other near the center of the distribution of the predictor variable. Therefore all of the predicted values are closer together as well. This explains the narrower width of the prediction interval.
You can see this in the figure below, which shows predictions at $x = 285$ and $x = 300$ for each of ten bootstrap replications. Typically, the lines are farther apart at $x = 300$ than at $x = 285$, and therefore the predictions at $x = 300$ are more variable.
```
x1 = 300
x2 = 285
lines = Table(['slope','intercept'])
for i in range(10):
rep = baby.sample(with_replacement=True)
a = slope(rep, 'Gestational Days', 'Birth Weight')
b = intercept(rep, 'Gestational Days', 'Birth Weight')
lines.append([a, b])
xlims = np.array([260, 310])
left = xlims[0]*lines[0] + lines[1]
right = xlims[1]*lines[0] + lines[1]
fit_x1 = x1*lines['slope'] + lines['intercept']
fit_x2 = x2*lines['slope'] + lines['intercept']
plots.xlim(xlims)
for i in range(10):
plots.plot(xlims, np.array([left[i], right[i]]), lw=1)
plots.scatter(x1, fit_x1[i], s=30)
plots.scatter(x2, fit_x2[i], s=30)
```
### Words of caution ###
All of the predictions and tests that we have performed in this chapter assume that the regression model holds. Specifically, the methods assume that the scatter plot resembles points generated by starting with points that are on a straight line and then pushing them off the line by adding random normal noise.
If the scatter plot does not look like that, then perhaps the model does not hold for the data. If the model does not hold, then calculations that assume the model to be true are not valid.
Therefore, we must first decide whether the regression model holds for our data, before we start making predictions based on the model or testing hypotheses about parameters of the model. A simple way is to do what we did in this section, which is to draw the scatter diagram of the two variables and see whether it looks roughly linear and evenly spread out around a line. We should also run the diagnostics we developed in the previous section using the residual plot.
| github_jupyter |
```
import os
import pandas as pd
import numpy as np
import cv2
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
%matplotlib inline
# Loading the annotations
df_anno = pd.read_csv('data/TABLES/data/MISC/truecover_tables.csv')
# Renaming the columns as the original labels aren't correct
df_anno.columns = ['name','xmin','ymin','w','h','label']
df_anno.head()
df_anno['xmin']=df_anno['xmin'].apply(lambda x:x-5)
df_anno['ymin']=df_anno['ymin'].apply(lambda x:x-5)
df_anno['w']=df_anno['w'].apply(lambda x:x+5)
df_anno['h']=df_anno['h'].apply(lambda x:x+5)
# creating data for test
# Define training Labels; here 1 being table class is defined
training_labels = [1]
PATH_OF_FOLDER='data/TABLES/data'
def create_annotations(PATH_OF_FOLDER,df_anno,training_labels,type_of_anno='train'):
f=open(PATH_OF_FOLDER+'/'+type_of_anno+'_anno.txt','w')
for i in df_anno.iterrows():
name = i[1]['name'].split('.')[0]
xmin = i[1]['xmin']
ymin = i[1]['ymin']
width = i[1]['w']
height = i[1]['h']
xmax = xmin + width
ymax = ymin + height
label = i[1]['label']
# All annotations for these files are going to be stored there
path=os.path.join(PATH_OF_FOLDER,'Annotations', name + '.txt')
# print(path)
if label in training_labels:
data=str(xmin)+';'+str(ymin)+';'+str(xmax)+';'+str(ymax)+';'+str(label)+'\n'
with open(path,'a') as f1:
f1.write(data) #write the annotation in the Annotation folder file
f.write(name+';'+data) #write the annotation in the complete list of training files
f1.close()
else:
print (label)
f.close()
def create_imagesets(PATH_OF_FOLDER,df_train,type_of_anno='train'):
train_list=open(os.path.join(PATH_OF_FOLDER,'ImageSets/'+type_of_anno+'.txt'),'a')
train_files=np.unique(df_train['name'])
for t in train_files:
train_list.write(t.split('.')[0]+'\n') #creates the list of files used for final training updated in ImageSets/train
train_list.close()
create_annotations(PATH_OF_FOLDER,df_anno,training_labels,type_of_anno='client_test')
create_imagesets(PATH_OF_FOLDER,df_anno,type_of_anno='test')
```
# MISC
```
# Converting the path
images = os.listdir('data/TABLES/data/MISC/table_img/')
for i in images:
print('data/TABLES/data/Images/'+'.'.join(i.split('.')[:-1])+'.jpg')
img = cv2.imread('data/TABLES/data/MISC/table_img/'+i)
cv2.imwrite('data/TABLES/data/Images/'+'.'.join(i.split('.')[:-1])+'.jpg',img)
```
| github_jupyter |
# MaterialsCoord benchmarking – symmetry of bonding algorithms
Several near neighbor methods do not produce symmetrical bonding. For example, if site A is bonded to site B, it is not guaranteed that site B will be bonded to site A. In the MaterialsCoord benchmark we enforce symmetrical bonding for all algorithms. In this notebook, we assess how unsymmetrical the bonding is for each near neighbor method.
*Written using:*
- MaterialsCoord==0.2.0
*Authors: Alex Ganose (05/20/20)*
---
First, lets initialize the near neighbor methods we are interested in.
```
from pymatgen.analysis.local_env import BrunnerNN_reciprocal, EconNN, JmolNN, \
MinimumDistanceNN, MinimumOKeeffeNN, MinimumVIRENN, \
VoronoiNN, CrystalNN
nn_methods = [
MinimumDistanceNN(), MinimumOKeeffeNN(), MinimumVIRENN(), JmolNN(),
EconNN(tol=0.5), BrunnerNN_reciprocal(), VoronoiNN(tol=0.5), CrystalNN()
]
```
Next, import the benchmark and choose which structures we are interested in.
```
from materialscoord.core import Benchmark
structure_groups = ["common_binaries", "elemental", "A2BX4", "ABX3", "ABX4"]
bm = Benchmark.from_structure_group(structure_groups)
```
Enforcing symmetry always increases the number of assigned bonds. To assess the symmetry, we therefore calculate the number of additional bonds resulting from enforcing symmetrical bonding. Calculating the coordination number from a `StructureGraph` object (as returned by `NearNeighbors.get_bonded_structure()`) always enforces symmetry. In contrast, calculating the coordination number directly from the `NearNeighbors.get_cn()` method does not enforce symmetry.
```
import numpy as np
from tqdm.auto import tqdm
symmetry_results = []
no_symmetry_results = []
for nn_method in tqdm(nn_methods):
nn_symmetry_cns = []
nn_no_symmetry_cns = []
for structure in bm.structures.values():
bs = nn_method.get_bonded_structure(structure)
for site_idx in range(len(structure)):
nn_symmetry_cns.append(bs.get_coordination_of_site(site_idx))
nn_no_symmetry_cns.append(nn_method.get_cn(structure, site_idx))
symmetry_results.append(nn_symmetry_cns)
no_symmetry_results.append(nn_no_symmetry_cns)
symmetry_results = np.array(symmetry_results)
no_symmetry_results = np.array(no_symmetry_results)
import pandas as pd
symmetry_totals = symmetry_results.sum(axis=1)
no_symmetry_totals = no_symmetry_results.sum(axis=1)
no_symmetry_norm = no_symmetry_totals / symmetry_totals
symmetry_extra = 1 - no_symmetry_norm
symmetry_df = pd.DataFrame(
columns=[n.__class__.__name__ for n in nn_methods],
data=[no_symmetry_norm, symmetry_extra],
index=["without symmetry", "with symmetry"]
)
import seaborn as sns
import matplotlib.pyplot as plt
from pathlib import Path
sns.set(font="Helvetica", font_scale=1.3, rc={"figure.figsize": (7, 7)})
sns.set_style("white", {"axes.edgecolor": "black", "axes.linewidth": 1.3})
plt.style.use({"mathtext.fontset": "custom", "mathtext.rm": "Arial", "axes.grid.axis": "x"})
symmetry_df = symmetry_df.rename(columns={"BrunnerNN_reciprocal": "BrunnerNN"})
ax = symmetry_df.T.plot(kind='bar', stacked=True)
ax.set_xticklabels(symmetry_df.columns, rotation=60)
ax.legend(frameon=False, loc="upper left", bbox_to_anchor=(1, 1))
ax.set(ylabel="Fraction of bonds assigned", xlabel="", ylim=(0, 1))
ax.tick_params(axis='y', which='major', size=10, width=1, color='k', left=True, direction="in")
plt.savefig(Path("plots/symmetry.pdf"), bbox_inches="tight")
plt.show()
! open .
```
| github_jupyter |
Import Modules
```
import requests
import pandas as pd
website_url = requests.get("https://en.wikipedia.org/wiki/List_of_postal_codes_of_Canada:_M").text
```
get website data
```
from bs4 import BeautifulSoup
soup = BeautifulSoup(website_url,"lxml")
```
iterate through table rows
```
pcs = []
prevP = ""
currN = ""
prevB = ""
for table_row in soup.select("table.wikitable tr"):
cells = table_row.findAll('td')
if len(cells) > 0:
pc = cells[0].text.strip()
b = cells[1].text.strip()
n = cells[2].text.strip()
if ((b == "Not assigned" and n=="Not assigned" or b == "blank")):
a="skipping"
else:
if (n =="Not Assigned"):
n = b; #neighbourhood becomes borough
if (pc == prevP):
currN = currN + "," + n #seperate if more than one with comma
else:
if (prevP != ""):
pcs.append([prevP,prevB,currN])
prevP = pc
prevB = b
currN = n
pcs.append([prevP,prevB,currN])
df = pd.DataFrame(pcs,columns=['Postal Code','Borough','Neighborhood']).sort_values(by=['Postal Code'])
```
show first 5 enties
```
df.head()
df.shape
import pandas as pd
import requests
import csv
import pandas as pd
import io
import requests
url="http://cocl.us/Geospatial_data"
s=requests.get(url).content
df2=pd.read_csv(io.StringIO(s.decode('utf-8')))
df2.head()
df3 = pd.merge(df, df2, on="Postal Code")
import folium
import numpy as np
from sklearn.cluster import KMeans
import matplotlib.cm as cm
import matplotlib.colors as colors
from geopy.geocoders import Nominatim
import datetime
from pandas.io.json import json_normalize
%matplotlib inline
data = df3[df3['Borough'].str.contains('Toronto', regex = False)].reset_index(drop=True)
data.head()
col_names = ['Postal Code','Borough','Neighborhood','Latitude','Longitude']
toronto_neigh = pd.DataFrame(columns = col_names)
#toronto_neigh
for i in range(data.shape[0]):
postcode = data.loc[i, 'Postal Code']
borough = data.loc[i, 'Borough']
lat = data.loc[i, 'Latitude'].astype(float)
lng = data.loc[i, 'Longitude'].astype(float)
neigh = data.loc[i,'Neighborhood'].split(", ")
for j in range(len(neigh)):
toronto_neigh = toronto_neigh.append(pd.DataFrame(np.array([[postcode, borough, neigh[j], lat, lng]]), columns = col_names))
toronto_neigh = toronto_neigh.reset_index(drop = True)
toronto_neigh.head()
# create folium map
toronto_map = folium.Map(location = [lat, lng], zoom_start = 11
)
for lat, lng, borough, neighborhood in zip(toronto_neigh['Latitude'], toronto_neigh['Longitude'], toronto_neigh['Borough'],toronto_neigh['Neighborhood']):
label = '{}, {} ({}, {})'.format(neighborhood, borough, lat, lng)
label = folium.Popup(label, parse_html= True)
folium.CircleMarker([float(lat),float(lng)],
radius = 3,
popup = label,
color = 'red',
fill = True,
fill_color = '#a72920',
fill_opacity = 0.5,
parse_html = False).add_to(toronto_map)
display(toronto_map)
```
api call data
```
now = datetime.datetime.now()
date = "%4d%02d%02d" % (now.year, now.month, now.day)
CLIENT_ID = '3ECJQTXHODVLXC0PN5LT5NM2ABWKXK4YORSKACOYAQ1RBOU1' # Foursquare ID
CLIENT_SECRET = '0XCMHV3VM5B3MDYANVU20ARUNNHL2LOPJ0DZNQYOYSJWTZ41' # Foursquare Secret
print('Your credentails:')
print('CLIENT_ID: ' + CLIENT_ID)
print('CLIENT_SECRET:' + CLIENT_SECRET)
VERSION = date
i = 0
latitude = toronto_neigh.loc[i, 'Latitude'] # neighborhood latitude value
longitude = toronto_neigh.loc[i, 'Longitude'] # neighborhood longitude value
neighborhood_name = toronto_neigh.loc[i, 'Neighborhood'] # neighborhood name
radius = 500
limit = 100
url = "https://api.foursquare.com/v2/venues/search?client_id={}&client_secret={}&ll={},{}&v={}&radius={}&limit={}".format(
CLIENT_ID,
CLIENT_SECRET,
latitude,
longitude,
VERSION,
radius,
limit)
results = requests.get(url).json()
def get_category_type(row):
try:
categories_list = row['categories']
except:
categories_list = row['venue.categories']
if len(categories_list) == 0:
return None
else:
return categories_list[0]['name']
```
call for venue data from neighborhoods
```
venues = results['response']['venues']
nearby_venues = json_normalize(venues)
# filter columns
filtered_columns = ['name', 'categories', 'location.lat', 'location.lng']
nearby_venues =nearby_venues.loc[:, filtered_columns]
nearby_venues['categories'] = nearby_venues.apply(get_category_type, axis=1)
nearby_venues.columns = [col.split(".")[-1] for col in nearby_venues.columns]
def getNearbyVenues(names, latitudes, longitudes, radius=500, LIMIT = 100):
venues_list=[]
for name, lat, lng in zip(names, latitudes, longitudes):
print(name)
url = 'https://api.foursquare.com/v2/venues/explore?&client_id={}&client_secret={}&v={}&ll={},{}&radius={}&limit={}'.format(
CLIENT_ID,
CLIENT_SECRET,
VERSION,
lat,
lng,
radius,
LIMIT)
results = requests.get(url).json()["response"]['groups'][0]['items']
# return only relevant information for each nearby venue
venues_list.append([(
name,
lat,
lng,
v['venue']['name'],
v['venue']['location']['lat'],
v['venue']['location']['lng'],
v['venue']['categories'][0]['name']) for v in results])
nearby_venues = pd.DataFrame([item for venue_list in venues_list for item in venue_list])
nearby_venues.columns = ['Neighborhood',
'Neighborhood Latitude',
'Neighborhood Longitude',
'Venue',
'Venue Latitude',
'Venue Longitude',
'Venue Category']
return(nearby_venues)
```
get venue data
```
toronto_venues = getNearbyVenues(names=toronto_neigh['Neighborhood'],
latitudes=toronto_neigh['Latitude'],
longitudes=toronto_neigh['Longitude']
)
toronto_onehot = pd.get_dummies(toronto_venues[['Venue Category']], prefix="", prefix_sep="")
toronto_onehot['Neighborhood'] = toronto_venues['Neighborhood']
#neighborhood column to the first column
fixed_columns = [toronto_onehot.columns[-1]] + list(toronto_onehot.columns[:-1])
toronto_onehot = toronto_onehot[fixed_columns]
toronto_grouped = toronto_onehot.groupby(['Neighborhood']).mean().reset_index()
def return_most_common_venues(row, num_top_venues):
row_categories = row.iloc[1:]
row_categories_sorted = row_categories.sort_values(ascending=False)
return row_categories_sorted.index.values[0:num_top_venues]
```
pre process data ready for kcluster
```
num_top_venues = 5
indicators = ['st', 'nd', 'rd']
# create columns according to number of top venues
columns = []
for ind in np.arange(num_top_venues):
try:
columns.append('{}{} Most Common Venue'.format(ind+1, indicators[ind]))
except:
columns.append('{}th Most Common Venue'.format(ind+1))
# create a new dataframe
neighborhoods_venues_sorted = pd.DataFrame(columns=columns)
neighborhoods_venues_sorted['Neighborhood'] = toronto_grouped['Neighborhood']
fixed_columns = [neighborhoods_venues_sorted.columns[-1]] + list(neighborhoods_venues_sorted.columns[:-1])
neighborhoods_venues_sorted = neighborhoods_venues_sorted[fixed_columns]
for ind in np.arange(toronto_grouped.shape[0]):
neighborhoods_venues_sorted.iloc[ind, 1:] = return_most_common_venues(toronto_grouped.iloc[ind, :], num_top_venues)
# set number of clusters
kclusters = 5
toronto_grouped_clustering = toronto_grouped.drop('Neighborhood', 1)
#cluster
kmeans = KMeans(n_clusters=kclusters, random_state=0).fit(toronto_grouped_clustering)
neighborhoods_venues_sorted.insert(0, 'Cluster Labels', kmeans.labels_)
toronto_merged = toronto_neigh
# merge toronto_grouped with toronto_data to add latitude/longitude for each neighborhood
toronto_merged = toronto_merged.join(neighborhoods_venues_sorted.set_index('Neighborhood'), on='Neighborhood')
```
map data with clusters shown
```
map_clusters = folium.Map(location=[latitude, longitude], zoom_start=11)
x = np.arange(kclusters)
ys = [i + x + (i*x)**2 for i in range(kclusters)]
colors_array = cm.rainbow(np.linspace(0, 1, len(ys)))
rainbow = [colors.rgb2hex(i) for i in colors_array]
markers_colors = []
for lat, lon, poi, cluster in zip(toronto_merged['Latitude'], toronto_merged['Longitude'], toronto_merged['Neighborhood'], toronto_merged['Cluster Labels']):
label = folium.Popup(str(poi) + ' Cluster ' + str(cluster), parse_html=True)
folium.CircleMarker(
[float(lat), float(lon)],
radius=5,
popup=label,
color=rainbow[cluster-1],
fill=True,
fill_color=rainbow[cluster-1],
fill_opacity=0.7).add_to(map_clusters)
map_clusters
```
| github_jupyter |
```
#Program Name: Daily-XChart-datapane_v1_single_multi_on_gsdr_synergy
#Purpose: Chart scatter line box aggr table data
#Author: Greg Turmel, Director, Data Governance
#Date: 2020.08.30 - 2021.06.30
#Errata: 0.1 Improvements can be made to script using for/looping through the databases
import os, sys, argparse, csv, pyodbc, sql, time, datetime
from datetime import datetime, timedelta
import sqlalchemy as db
from dotenv import load_dotenv # add this line
from matplotlib.backends.backend_pdf import PdfPages
import pandas as pd
import altair as alt
from altair import Chart, X, Y, Axis, SortField
from vega_datasets import data as vega_data
import datapane as dp
#import datatable as dt
import numpy as np
import matplotlib # notebook
import matplotlib.pyplot as plt
import seaborn as sns
#import chart_studio.plotly as py
import plotly.graph_objects as go
load_dotenv() # add this line
user = os.getenv('MySQLeUser')
password = os.getenv('MySQLeUserPass')
host = os.getenv('MySQLeHOST')
db = os.getenv('MySQLeDB')
%matplotlib inline
%load_ext sql
conn = pyodbc.connect('Driver={SQL Server};'
'Server=DEVODSSQL;'
'Database=Greg;'
'Trusted_Connection=yes;')
sql_query01 = pd.read_sql_query('''
select DISTINCT TableName
FROM [dbo].[tableRowCountGSDR_Synergy]
ORDER BY TableName;
'''
,conn) # Load the list of distinct tables to graph
sql_query02 = pd.read_sql_query('''
select *
FROM [dbo].[tableRowCountGSDR_Synergy]
ORDER BY TableName;
'''
,conn) # Load the list of distinct tables to graph
df = pd.DataFrame(sql_query02)
#df = df.astype({'TodaysDate':np.int64,'RecordCount':np.int64})
df['TodaysDate'] = pd.to_datetime(df['TodaysDate'].astype(str), format='%Y%m%d')
df = df.sort_values(by=['TableName','SchemaName','DatabaseName','TodaysDate'], ascending=[True,True,True,True])
df = df.reset_index(drop=True)
dfgsdrsyn0001 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'dbo') & (df['TableName'] == 'ELLPGM_SASI')]
dfgsdrsyn0002 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'dbo') & (df['TableName'] == 'ELLPGM_SYN')]
dfgsdrsyn0003 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'dbo') & (df['TableName'] == 'HOME_LANG_NEW')]
dfgsdrsyn0004 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'dbo') & (df['TableName'] == 'HOME_LANG_OLD')]
dfgsdrsyn0005 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'dbo') & (df['TableName'] == 'sysssislog')]
dfgsdrsyn0006 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_A504_SYNERGY')]
dfgsdrsyn0007 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AATC_SYNERGY')]
dfgsdrsyn0008 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AATD_SYNERGY')]
dfgsdrsyn0009 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AATO_ROT_SYNERGY')]
dfgsdrsyn0010 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AATP_SYNERGY')]
dfgsdrsyn0011 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AATT_SYNERGY')]
dfgsdrsyn0012 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ACHS_SYNERGY')]
dfgsdrsyn0013 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ACHS_SYNERGY2')]
dfgsdrsyn0014 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ACLH_SYNERGY')]
dfgsdrsyn0015 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ACLS_NEXT_SYNERGY')]
dfgsdrsyn0016 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ACLS_SYNERGY')]
dfgsdrsyn0017 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ACLS_SYNERGY_BCK')]
dfgsdrsyn0018 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ACNF_SYNERGY')]
dfgsdrsyn0019 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ACNR_SYNERGY')]
dfgsdrsyn0020 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ACRS_SYNERGY')]
dfgsdrsyn0021 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ADDITIONAL_TEACHER_SYNERGY')]
dfgsdrsyn0022 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ADIS')]
dfgsdrsyn0023 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ADIS_MISSING')]
dfgsdrsyn0024 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ADIS_SYNERGY')]
dfgsdrsyn0025 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ADIS_SYNERGY_KS')]
dfgsdrsyn0026 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ADPO')]
dfgsdrsyn0027 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ADPO_MISSING')]
dfgsdrsyn0028 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ADPO_SYNERGY')]
dfgsdrsyn0029 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ADPO_SYNERGY_KS')]
dfgsdrsyn0030 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ADST')]
dfgsdrsyn0031 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ADST_NEXT_SYNERGY')]
dfgsdrsyn0032 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ADST_SYNERGY')]
dfgsdrsyn0033 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AELT_SYNERGY')]
dfgsdrsyn0034 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AEMG_SYNERGY')]
dfgsdrsyn0035 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AENR')]
dfgsdrsyn0036 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AENR_C')]
dfgsdrsyn0037 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AENR_NON_ADA_ADM_SYNERGY')]
dfgsdrsyn0038 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AENR_SYNERGY')]
dfgsdrsyn0039 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AFEE_SYNERGY')]
dfgsdrsyn0040 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AFID_SYNERGY')]
dfgsdrsyn0041 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AHLT_SYNERGY')]
dfgsdrsyn0042 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ALKR_SYNERGY')]
dfgsdrsyn0043 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ALRT_SYNERGY')]
dfgsdrsyn0044 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AMST_ADDL_STF_STU')]
dfgsdrsyn0045 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AMST_NEXT_SYNERGY')]
dfgsdrsyn0046 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AMST_SECT_STF')]
dfgsdrsyn0047 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AMST_SECT_STF_HIS')]
dfgsdrsyn0048 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AMST_SYNERGY')]
dfgsdrsyn0049 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_AONL_SYNERGY')]
dfgsdrsyn0050 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_APGD_Synergy')]
dfgsdrsyn0051 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_APGL_Synergy')]
dfgsdrsyn0052 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_APGM_SYNERGY')]
dfgsdrsyn0053 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_APRN_SYNERGY')]
dfgsdrsyn0054 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ASCH_SYNERGY')]
dfgsdrsyn0055 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ASGR_SYNERGY')]
dfgsdrsyn0056 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ASTI_SYNERGY')]
dfgsdrsyn0057 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ASTU_DIPLOMA_SEALS_SYNERGY')]
dfgsdrsyn0058 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ASTU_GPA')]
dfgsdrsyn0059 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ASTU_NEXT_SYNERGY')]
dfgsdrsyn0060 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ASTU_NON_ADA_ADM_SYNERGY')]
dfgsdrsyn0061 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ASTU_SYNERGY')]
dfgsdrsyn0062 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ASTU_SYNERGY_DEL')]
dfgsdrsyn0063 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ASTU_SYNERGY_KERI')]
dfgsdrsyn0064 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ASTU_SYNERGY2')]
dfgsdrsyn0065 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ATBL_SYNERGY')]
dfgsdrsyn0066 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ATCH_NEXT_SYNERGY')]
dfgsdrsyn0067 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ATCH_SYNERGY')]
dfgsdrsyn0068 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ATRM_NEXT_SYNERGY')]
dfgsdrsyn0069 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ATRM_SYNERGY')]
dfgsdrsyn0070 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_ATTD_SYNERGY')]
dfgsdrsyn0071 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_PARAPRO_SYNERGY')]
dfgsdrsyn0072 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_PHLOTE')]
dfgsdrsyn0073 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_RUN_CONTROL')]
dfgsdrsyn0074 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_RUN_CONTROL_SCH')]
dfgsdrsyn0075 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_SMST_SYNERGY')]
dfgsdrsyn0076 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_SMST_SYNERGY_STAGE')]
dfgsdrsyn0077 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_UABN_SYNERGY')]
dfgsdrsyn0078 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_UATR_SYNERGY')]
dfgsdrsyn0079 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_UCES_SYNERGY')]
dfgsdrsyn0080 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_UCMS_SYNERGY')]
dfgsdrsyn0081 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_UGFT_SYNERGY')]
dfgsdrsyn0082 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'SASI_UPSR_SYNERGY')]
dfgsdrsyn0083 = df[(df['DatabaseName'] == 'GSDR_Synergy') & (df['SchemaName'] == 'GEMS') & (df['TableName'] == 'TEST_STU_ACC_OOD')]
#alt.data_transformers.enable("json", urlpath="/user-redirect/files/")
alt.data_transformers.enable('json')
selection = alt.selection_multi();
alt.Chart(df).mark_point(filled=True).encode(
alt.X('TodaysDate', scale=alt.Scale(zero=False)),
alt.Y('RecordCount', scale=alt.Scale(zero=False)),
alt.Size('RecordCount:Q'),
alt.OpacityValue(0.5),
alt.Order('RecordCount:Q', sort='descending'),
tooltip = [alt.Tooltip('TodaysDate:T'),
alt.Tooltip('RecordCount:Q'),
alt.Tooltip('DatabaseName:N'),
alt.Tooltip('SchemaName:N'),
alt.Tooltip('TableName:N'),
],
color=alt.condition(selection, 'cluster:N', alt.value('grey'))
).add_selection(selection).properties(
title='GSDR_Synergy All Tables'
).properties(
width=750,
height=300
).interactive()
hover = alt.selection_multi(on='mouseover', nearest=True, empty='none')
base = alt.Chart(df, title='GSDR_Synergy All Tables').encode(
alt.X('TodaysDate:T')
#title.configure_title(fontSize=24)
).properties(
width=750, height=300,
)
points = base.mark_point().add_selection(
hover
)
line_A = base.mark_line(color='#5276A7').encode(
alt.Y('average(RecordCount):Q', axis=alt.Axis(titleColor='#5276A7'))
)
line_B = base.mark_line(color='#F18727').encode(
alt.Y('average(Ddifference):Q', axis=alt.Axis(titleColor='#F18727'))
)
alt.layer(line_A, line_B).resolve_scale(y='independent').interactive()
#Individual Tables - graphed with datapane and altair
#alt.data_transformers.enable("json", urlpath="/user-redirect/files/")
alt.data_transformers.enable('json')
selection = alt.selection_multi();
alt.Chart(dfgsdrsyn0006).mark_point(filled=True).encode(
alt.X('TodaysDate', scale=alt.Scale(zero=False)),
alt.Y('RecordCount', scale=alt.Scale(zero=False)),
alt.Size('RecordCount:Q'),
alt.OpacityValue(0.5),
alt.Order('RecordCount:Q', sort='descending'),
tooltip = [alt.Tooltip('TodaysDate:T'),
alt.Tooltip('RecordCount:Q'),
alt.Tooltip('DatabaseName:N'),
alt.Tooltip('SchemaName:N'),
alt.Tooltip('TableName:N'),
],
color=alt.condition(selection, 'cluster:N', alt.value('grey'))
).add_selection(selection).properties(
title='SASI_A504_SYNERGY'
).properties(
width=750,
height=300
).interactive()
hover = alt.selection_multi(on='mouseover', nearest=True, empty='none')
base = alt.Chart(dfgsdrsyn0006, title='SASI_A504_SYNERGY').encode(
alt.X('TodaysDate:T')
#title.configure_title(fontSize=24)
).properties(
width=750, height=300,
)
points = base.mark_point().add_selection(
hover
)
line_A = base.mark_line(color='#5276A7').encode(
alt.Y('average(RecordCount):Q', axis=alt.Axis(titleColor='#5276A7'))
)
line_B = base.mark_line(color='#F18727').encode(
alt.Y('average(Ddifference):Q', axis=alt.Axis(titleColor='#F18727'))
)
alt.layer(line_A, line_B).resolve_scale(y='independent').interactive()
#alt.data_transformers.enable("json", urlpath="/user-redirect/files/")
alt.data_transformers.enable('json')
selection = alt.selection_multi();
alt.Chart(dfgsdrsyn0008).mark_point(filled=True).encode(
alt.X('TodaysDate', scale=alt.Scale(zero=False)),
alt.Y('RecordCount', scale=alt.Scale(zero=False)),
alt.Size('RecordCount:Q'),
alt.OpacityValue(0.5),
alt.Order('RecordCount:Q', sort='descending'),
tooltip = [alt.Tooltip('TodaysDate:T'),
alt.Tooltip('RecordCount:Q'),
alt.Tooltip('DatabaseName:N'),
alt.Tooltip('SchemaName:N'),
alt.Tooltip('TableName:N'),
],
color=alt.condition(selection, 'cluster:N', alt.value('grey'))
).add_selection(selection).properties(
title='SASI_AATD_SYNERGY'
).properties(
width=750,
height=300
).interactive()
hover = alt.selection_multi(on='mouseover', nearest=True, empty='none')
base = alt.Chart(dfgsdrsyn0008, title='SASI_AATD_SYNERGY').encode(
alt.X('TodaysDate:T')
#title.configure_title(fontSize=24)
).properties(
width=750, height=300,
)
points = base.mark_point().add_selection(
hover
)
line_A = base.mark_line(color='#5276A7').encode(
alt.Y('average(RecordCount):Q', axis=alt.Axis(titleColor='#5276A7'))
)
line_B = base.mark_line(color='#F18727').encode(
alt.Y('average(Ddifference):Q', axis=alt.Axis(titleColor='#F18727'))
)
alt.layer(line_A, line_B).resolve_scale(y='independent').interactive()
#alt.data_transformers.enable("json", urlpath="/user-redirect/files/")
alt.data_transformers.enable('json')
selection = alt.selection_multi();
alt.Chart(dfgsdrsyn0011).mark_point(filled=True).encode(
alt.X('TodaysDate', scale=alt.Scale(zero=False)),
alt.Y('RecordCount', scale=alt.Scale(zero=False)),
alt.Size('RecordCount:Q'),
alt.OpacityValue(0.5),
alt.Order('RecordCount:Q', sort='descending'),
tooltip = [alt.Tooltip('TodaysDate:T'),
alt.Tooltip('RecordCount:Q'),
alt.Tooltip('DatabaseName:N'),
alt.Tooltip('SchemaName:N'),
alt.Tooltip('TableName:N'),
],
color=alt.condition(selection, 'cluster:N', alt.value('grey'))
).add_selection(selection).properties(
title='SASI_AATT_SYNERGY'
).properties(
width=750,
height=300
).interactive()
hover = alt.selection_multi(on='mouseover', nearest=True, empty='none')
base = alt.Chart(dfgsdrsyn0011, title='SASI_AATT_SYNERGY').encode(
alt.X('TodaysDate:T')
#title.configure_title(fontSize=24)
).properties(
width=750, height=300,
)
points = base.mark_point().add_selection(
hover
)
line_A = base.mark_line(color='#5276A7').encode(
alt.Y('average(RecordCount):Q', axis=alt.Axis(titleColor='#5276A7'))
)
line_B = base.mark_line(color='#F18727').encode(
alt.Y('average(Ddifference):Q', axis=alt.Axis(titleColor='#F18727'))
)
alt.layer(line_A, line_B).resolve_scale(y='independent').interactive()
#alt.data_transformers.enable("json", urlpath="/user-redirect/files/")
alt.data_transformers.enable('json')
selection = alt.selection_multi();
alt.Chart(dfgsdrsyn0012).mark_point(filled=True).encode(
alt.X('TodaysDate', scale=alt.Scale(zero=False)),
alt.Y('RecordCount', scale=alt.Scale(zero=False)),
alt.Size('RecordCount:Q'),
alt.OpacityValue(0.5),
alt.Order('RecordCount:Q', sort='descending'),
tooltip = [alt.Tooltip('TodaysDate:T'),
alt.Tooltip('RecordCount:Q'),
alt.Tooltip('DatabaseName:N'),
alt.Tooltip('SchemaName:N'),
alt.Tooltip('TableName:N'),
],
color=alt.condition(selection, 'cluster:N', alt.value('grey'))
).add_selection(selection).properties(
title='SASI_ACHS_SYNERGY'
).properties(
width=750,
height=300
).interactive()
hover = alt.selection_multi(on='mouseover', nearest=True, empty='none')
base = alt.Chart(dfgsdrsyn0012, title='SASI_ACHS_SYNERGY').encode(
alt.X('TodaysDate:T')
#title.configure_title(fontSize=24)
).properties(
width=750, height=300,
)
points = base.mark_point().add_selection(
hover
)
line_A = base.mark_line(color='#5276A7').encode(
alt.Y('average(RecordCount):Q', axis=alt.Axis(titleColor='#5276A7'))
)
line_B = base.mark_line(color='#F18727').encode(
alt.Y('average(Ddifference):Q', axis=alt.Axis(titleColor='#F18727'))
)
alt.layer(line_A, line_B).resolve_scale(y='independent').interactive()
#alt.data_transformers.enable("json", urlpath="/user-redirect/files/")
alt.data_transformers.enable('json')
selection = alt.selection_multi();
alt.Chart(dfgsdrsyn0014).mark_point(filled=True).encode(
alt.X('TodaysDate', scale=alt.Scale(zero=False)),
alt.Y('RecordCount', scale=alt.Scale(zero=False)),
alt.Size('RecordCount:Q'),
alt.OpacityValue(0.5),
alt.Order('RecordCount:Q', sort='descending'),
tooltip = [alt.Tooltip('TodaysDate:T'),
alt.Tooltip('RecordCount:Q'),
alt.Tooltip('DatabaseName:N'),
alt.Tooltip('SchemaName:N'),
alt.Tooltip('TableName:N'),
],
color=alt.condition(selection, 'cluster:N', alt.value('grey'))
).add_selection(selection).properties(
title='SASI_ACLH_SYNERGY'
).properties(
width=750,
height=300
).interactive()
hover = alt.selection_multi(on='mouseover', nearest=True, empty='none')
base = alt.Chart(dfgsdrsyn0014, title='SASI_ACLH_SYNERGY').encode(
alt.X('TodaysDate:T')
#title.configure_title(fontSize=24)
).properties(
width=750, height=300,
)
points = base.mark_point().add_selection(
hover
)
line_A = base.mark_line(color='#5276A7').encode(
alt.Y('average(RecordCount):Q', axis=alt.Axis(titleColor='#5276A7'))
)
line_B = base.mark_line(color='#F18727').encode(
alt.Y('average(Ddifference):Q', axis=alt.Axis(titleColor='#F18727'))
)
alt.layer(line_A, line_B).resolve_scale(y='independent').interactive()
#alt.data_transformers.enable("json", urlpath="/user-redirect/files/")
alt.data_transformers.enable('json')
selection = alt.selection_multi();
alt.Chart(dfgsdrsyn0015).mark_point(filled=True).encode(
alt.X('TodaysDate', scale=alt.Scale(zero=False)),
alt.Y('RecordCount', scale=alt.Scale(zero=False)),
alt.Size('RecordCount:Q'),
alt.OpacityValue(0.5),
alt.Order('RecordCount:Q', sort='descending'),
tooltip = [alt.Tooltip('TodaysDate:T'),
alt.Tooltip('RecordCount:Q'),
alt.Tooltip('DatabaseName:N'),
alt.Tooltip('SchemaName:N'),
alt.Tooltip('TableName:N'),
],
color=alt.condition(selection, 'cluster:N', alt.value('grey'))
).add_selection(selection).properties(
title='SASI_ACLS_NEXT_SYNERGY'
).properties(
width=750,
height=300
).interactive()
hover = alt.selection_multi(on='mouseover', nearest=True, empty='none')
base = alt.Chart(dfgsdrsyn0015, title='SASI_ACLS_NEXT_SYNERGY').encode(
alt.X('TodaysDate:T')
#title.configure_title(fontSize=24)
).properties(
width=750, height=300,
)
points = base.mark_point().add_selection(
hover
)
line_A = base.mark_line(color='#5276A7').encode(
alt.Y('average(RecordCount):Q', axis=alt.Axis(titleColor='#5276A7'))
)
line_B = base.mark_line(color='#F18727').encode(
alt.Y('average(Ddifference):Q', axis=alt.Axis(titleColor='#F18727'))
)
alt.layer(line_A, line_B).resolve_scale(y='independent').interactive()
#alt.data_transformers.enable("json", urlpath="/user-redirect/files/")
alt.data_transformers.enable('json')
selection = alt.selection_multi();
alt.Chart(dfgsdrsyn0021).mark_point(filled=True).encode(
alt.X('TodaysDate', scale=alt.Scale(zero=False)),
alt.Y('RecordCount', scale=alt.Scale(zero=False)),
alt.Size('RecordCount:Q'),
alt.OpacityValue(0.5),
alt.Order('RecordCount:Q', sort='descending'),
tooltip = [alt.Tooltip('TodaysDate:T'),
alt.Tooltip('RecordCount:Q'),
alt.Tooltip('DatabaseName:N'),
alt.Tooltip('SchemaName:N'),
alt.Tooltip('TableName:N'),
],
color=alt.condition(selection, 'cluster:N', alt.value('grey'))
).add_selection(selection).properties(
title='SASI_ADDITIONAL_TEACHER_SYNERGY'
).properties(
width=750,
height=300
).interactive()
hover = alt.selection_multi(on='mouseover', nearest=True, empty='none')
base = alt.Chart(dfgsdrsyn0021, title='SASI_ADDITIONAL_TEACHER_SYNERGY').encode(
alt.X('TodaysDate:T')
#title.configure_title(fontSize=24)
).properties(
width=750, height=300,
)
points = base.mark_point().add_selection(
hover
)
line_A = base.mark_line(color='#5276A7').encode(
alt.Y('average(RecordCount):Q', axis=alt.Axis(titleColor='#5276A7'))
)
line_B = base.mark_line(color='#F18727').encode(
alt.Y('average(Ddifference):Q', axis=alt.Axis(titleColor='#F18727'))
)
alt.layer(line_A, line_B).resolve_scale(y='independent').interactive()
```
| github_jupyter |
<p><img src="https://i.imgur.com/fWVJjgs.jpg" style="float: left; margin: 20px; height: 200px"></p>
### Original notebook from Kevin Markham at [Data School](http://www.dataschool.io/)
**Related:** [GitHub repository](https://github.com/justmarkham/python-reference) and [blog post](http://www.dataschool.io/python-quick-reference/)
## Table of contents
1. <a href="#1.-Imports">Imports</a>
2. <a href="#2.-Data-Types">Data Types</a>
3. <a href="#3.-Math">Math</a>
4. <a href="#4.-Comparisons-and-Boolean-Operations">Comparisons and Boolean Operations</a>
5. <a href="#5.-Conditional-Statements">Conditional Statements</a>
6. <a href="#6.-Lists">Lists</a>
7. <a href="#7.-Tuples">Tuples</a>
8. <a href="#8.-Strings">Strings</a>
9. <a href="#9.-Dictionaries">Dictionaries</a>
10. <a href="#10.-Sets">Sets</a>
11. <a href="#11.-Defining-Functions">Defining Functions</a>
12. <a href="#12.-Anonymous-%28Lambda%29-Functions">Anonymous (Lambda) Functions</a>
13. <a href="#13.-For-Loops-and-While-Loops">For Loops and While Loops</a>
14. <a href="#14.-Comprehensions">Comprehensions</a>
15. <a href="#15.-Map-and-Filter">Map and Filter</a>
## 1. Imports
```
# 'generic import' of math module
import math
math.sqrt(25)
# import a function
from math import sqrt
sqrt(25) # no longer have to reference the module
# import multiple functions at once
from math import cos, floor
# import all functions in a module (generally discouraged)
from csv import *
# define an alias
import datetime as dt
# show all functions in math module
print(dir(math))
```
[<a href="#Python-Quick-Reference-by-Data-School">Back to top</a>]
## 2. Data Types
**Determine the type of an object:**
```
type(2)
type(2.0)
type('two')
type(True)
type(None)
```
**Check if an object is of a given type:**
```
isinstance(2.0, int)
isinstance(2.0, (int, float))
```
**Convert an object to a given type:**
```
float(2)
int(2.9)
str(2.9)
```
**Zero, `None`, and empty containers are converted to `False`:**
```
bool(0)
bool(None)
bool('') # empty string
bool([]) # empty list
bool({}) # empty dictionary
```
**Non-empty containers and non-zeros are converted to `True`:**
```
bool(2)
bool('two')
bool([2])
```
[<a href="#Python-Quick-Reference-by-Data-School">Back to top</a>]
## 3. Math
```
10 + 4
10 - 4
10 * 4
10 ** 4 # exponent
5 % 4 # modulo - computes the remainder
# Python 2: returns 2 (because both types are 'int')
# Python 3: returns 2.5
10 / 4
10 / float(4)
# force '/' in Python 2 to perform 'true division' (unnecessary in Python 3)
from __future__ import division
10 / 4 # true division
10 // 4 # floor division
```
[<a href="#Python-Quick-Reference-by-Data-School">Back to top</a>]
## 4. Comparisons and Boolean Operations
**Assignment statement:**
```
x = 5
```
**Comparisons:**
```
x > 3
x >= 3
x != 3
x == 5
```
**Boolean operations:**
```
5 > 3 and 6 > 3
5 > 3 or 5 < 3
not False
False or not False and True # evaluation order: not, and, or
```
[<a href="#Python-Quick-Reference-by-Data-School">Back to top</a>]
## 5. Conditional Statements
```
# if statement
if x > 0:
print('positive')
# if/else statement
if x > 0:
print('positive')
else:
print('zero or negative')
# if/elif/else statement
if x > 0:
print('positive')
elif x == 0:
print('zero')
else:
print('negative')
# single-line if statement (sometimes discouraged)
if x > 0: print('positive')
# single-line if/else statement (sometimes discouraged), known as a 'ternary operator'
'positive' if x > 0 else 'zero or negative'
```
[<a href="#Python-Quick-Reference-by-Data-School">Back to top</a>]
## 6. Lists
- **List properties:** ordered, iterable, mutable, can contain multiple data types
```
# create an empty list (two ways)
empty_list = []
empty_list = list()
# create a list
simpsons = ['homer', 'marge', 'bart']
```
**Examine a list:**
```
# print element 0
simpsons[0]
len(simpsons)
```
**Modify a list (does not return the list):**
```
# append element to end
simpsons.append('lisa')
simpsons
# append multiple elements to end
simpsons.extend(['itchy', 'scratchy'])
simpsons
# insert element at index 0 (shifts everything right)
simpsons.insert(0, 'maggie')
simpsons
# search for first instance and remove it
simpsons.remove('bart')
simpsons
# remove element 0 and return it
simpsons.pop(0)
# remove element 0 (does not return it)
del simpsons[0]
simpsons
# replace element 0
simpsons[0] = 'krusty'
simpsons
# concatenate lists (slower than 'extend' method)
neighbors = simpsons + ['ned', 'rod', 'todd']
neighbors
```
**Find elements in a list:**
```
# counts the number of instances
simpsons.count('lisa')
# returns index of first instance
simpsons.index('itchy')
```
**List slicing:**
```
weekdays = ['mon', 'tues', 'wed', 'thurs', 'fri']
# element 0
weekdays[0]
# elements 0 (inclusive) to 3 (exclusive)
weekdays[0:3]
# starting point is implied to be 0
weekdays[:3]
# elements 3 (inclusive) through the end
weekdays[3:]
# last element
weekdays[-1]
# every 2nd element (step by 2)
weekdays[::2]
# backwards (step by -1)
weekdays[::-1]
# alternative method for returning the list backwards
list(reversed(weekdays))
```
**Sort a list in place (modifies but does not return the list):**
```
simpsons.sort()
simpsons
# sort in reverse
simpsons.sort(reverse=True)
simpsons
# sort by a key
simpsons.sort(key=len)
simpsons
```
**Return a sorted list (does not modify the original list):**
```
sorted(simpsons)
sorted(simpsons, reverse=True)
sorted(simpsons, key=len)
```
**Insert into an already sorted list, and keep it sorted:**
```
num = [10, 20, 40, 50]
from bisect import insort
insort(num, 30)
num
```
**Object references and copies:**
```
# create a second reference to the same list
same_num = num
# modifies both 'num' and 'same_num'
same_num[0] = 0
print(num)
print(same_num)
# copy a list (two ways)
new_num = num[:]
new_num = list(num)
```
**Examine objects:**
```
num is same_num # checks whether they are the same object
num is new_num
num == same_num # checks whether they have the same contents
num == new_num
```
[<a href="#Python-Quick-Reference-by-Data-School">Back to top</a>]
## 7. Tuples
- **Tuple properties:** ordered, iterable, immutable, can contain multiple data types
- Like lists, but they don't change size
```
# create a tuple directly
digits = (0, 1, 'two')
# create a tuple from a list
digits = tuple([0, 1, 'two'])
# trailing comma is required to indicate it's a tuple
zero = (0,)
```
**Examine a tuple:**
```
digits[2]
len(digits)
# counts the number of instances of that value
digits.count(0)
# returns the index of the first instance of that value
digits.index(1)
```
**Modify a tuple:**
```
# elements of a tuple cannot be modified (this would throw an error)
# digits[2] = 2
# concatenate tuples
digits = digits + (3, 4)
digits
```
**Other tuple operations:**
```
# create a single tuple with elements repeated (also works with lists)
(3, 4) * 2
# sort a list of tuples
tens = [(20, 60), (10, 40), (20, 30)]
sorted(tens) # sorts by first element in tuple, then second element
# tuple unpacking
bart = ('male', 10, 'simpson') # create a tuple
(sex, age, surname) = bart # assign three values at once
print(sex)
print(age)
print(surname)
```
[<a href="#Python-Quick-Reference-by-Data-School">Back to top</a>]
## 8. Strings
- **String properties:** iterable, immutable
```
# convert another data type into a string
s = str(42)
s
# create a string directly
s = 'I like you'
```
**Examine a string:**
```
s[0]
len(s)
```
**String slicing is like list slicing:**
```
s[:6]
s[7:]
s[-1]
```
**Basic string methods (does not modify the original string):**
```
s.lower()
s.upper()
s.startswith('I')
s.endswith('you')
# checks whether every character in the string is a digit
s.isdigit()
# returns index of first occurrence, but doesn't support regex
s.find('like')
# returns -1 since not found
s.find('hate')
# replaces all instances of 'like' with 'love'
s.replace('like', 'love')
```
**Split a string:**
```
# split a string into a list of substrings separated by a delimiter
s.split(' ')
# equivalent (since space is the default delimiter)
s.split()
s2 = 'a, an, the'
s2.split(',')
```
**Join or concatenate strings:**
```
# join a list of strings into one string using a delimiter
stooges = ['larry', 'curly', 'moe']
' '.join(stooges)
# concatenate strings
s3 = 'The meaning of life is'
s4 = '42'
s3 + ' ' + s4
```
**Remove whitespace from the start and end of a string:**
```
s5 = ' ham and cheese '
s5.strip()
```
**String substitutions:**
```
# old way
'raining %s and %s' % ('cats', 'dogs')
# new way
'raining {} and {}'.format('cats', 'dogs')
# new way (using named arguments)
'raining {arg1} and {arg2}'.format(arg1='cats', arg2='dogs')
```
**String formatting ([more examples](https://mkaz.tech/python-string-format.html)):**
```
# use 2 decimal places
'pi is {:.2f}'.format(3.14159)
```
**Normal strings versus raw strings:**
```
# normal strings allow for escaped characters
print('first line\nsecond line')
# raw strings treat backslashes as literal characters
print(r'first line\nfirst line')
```
[<a href="#Python-Quick-Reference-by-Data-School">Back to top</a>]
## 9. Dictionaries
- **Dictionary properties:** unordered, iterable, mutable, can contain multiple data types
- Made of key-value pairs
- Keys must be unique, and can be strings, numbers, or tuples
- Values can be any type
```
# create an empty dictionary (two ways)
empty_dict = {}
empty_dict = dict()
# create a dictionary (two ways)
family = {'dad':'homer', 'mom':'marge', 'size':6}
family = dict(dad='homer', mom='marge', size=6)
family
# convert a list of tuples into a dictionary
list_of_tuples = [('dad', 'homer'), ('mom', 'marge'), ('size', 6)]
family = dict(list_of_tuples)
family
```
**Examine a dictionary:**
```
# pass a key to return its value
family['dad']
# return the number of key-value pairs
len(family)
# check if key exists in dictionary
'mom' in family
# dictionary values are not checked
'marge' in family
# returns a list of keys (Python 2) or an iterable view (Python 3)
family.keys()
# returns a list of values (Python 2) or an iterable view (Python 3)
family.values()
# returns a list of key-value pairs (Python 2) or an iterable view (Python 3)
family.items()
```
**Modify a dictionary (does not return the dictionary):**
```
# add a new entry
family['cat'] = 'snowball'
family
# edit an existing entry
family['cat'] = 'snowball ii'
family
# delete an entry
del family['cat']
family
# dictionary value can be a list
family['kids'] = ['bart', 'lisa']
family
# remove an entry and return the value
family.pop('dad')
# add multiple entries
family.update({'baby':'maggie', 'grandpa':'abe'})
family
```
**Access values more safely with `get`:**
```
family['mom']
# equivalent to a dictionary lookup
family.get('mom')
# this would throw an error since the key does not exist
# family['grandma']
# return None if not found
family.get('grandma')
# provide a default return value if not found
family.get('grandma', 'not found')
```
**Access a list element within a dictionary:**
```
family['kids'][0]
family['kids'].remove('lisa')
family
```
**String substitution using a dictionary:**
```
'youngest child is %(baby)s' % family
```
[<a href="#Python-Quick-Reference-by-Data-School">Back to top</a>]
## 10. Sets
- **Set properties:** unordered, iterable, mutable, can contain multiple data types
- Made of unique elements (strings, numbers, or tuples)
- Like dictionaries, but with keys only (no values)
```
# create an empty set
empty_set = set()
# create a set directly
languages = {'python', 'r', 'java'}
# create a set from a list
snakes = set(['cobra', 'viper', 'python'])
```
**Examine a set:**
```
len(languages)
'python' in languages
```
**Set operations:**
```
# intersection
languages & snakes
# union
languages | snakes
# set difference
languages - snakes
# set difference
snakes - languages
```
**Modify a set (does not return the set):**
```
# add a new element
languages.add('sql')
languages
# try to add an existing element (ignored, no error)
languages.add('r')
languages
# remove an element
languages.remove('java')
languages
# try to remove a non-existing element (this would throw an error)
# languages.remove('c')
# remove an element if present, but ignored otherwise
languages.discard('c')
languages
# remove and return an arbitrary element
languages.pop()
# remove all elements
languages.clear()
languages
# add multiple elements (can also pass a set)
languages.update(['go', 'spark'])
languages
```
**Get a sorted list of unique elements from a list:**
```
sorted(set([9, 0, 2, 1, 0]))
```
[<a href="#Python-Quick-Reference-by-Data-School">Back to top</a>]
## 11. Defining Functions
**Define a function with no arguments and no return values:**
```
def print_text():
print('this is text')
# call the function
print_text()
```
**Define a function with one argument and no return values:**
```
def print_this(x):
print(x)
# call the function
print_this(3)
# prints 3, but doesn't assign 3 to n because the function has no return statement
n = print_this(3)
```
**Define a function with one argument and one return value:**
```
def square_this(x):
return x**2
# include an optional docstring to describe the effect of a function
def square_this(x):
"""Return the square of a number."""
return x**2
# call the function
square_this(3)
# assigns 9 to var, but does not print 9
var = square_this(3)
```
**Define a function with two 'positional arguments' (no default values) and one 'keyword argument' (has a default value):**
```
def calc(a, b, op='add'):
if op == 'add':
return a + b
elif op == 'sub':
return a - b
else:
print('valid operations are add and sub')
# call the function
calc(10, 4, op='add')
# unnamed arguments are inferred by position
calc(10, 4, 'add')
# default for 'op' is 'add'
calc(10, 4)
calc(10, 4, 'sub')
calc(10, 4, 'div')
```
**Use `pass` as a placeholder if you haven't written the function body:**
```
def stub():
pass
```
**Return two values from a single function:**
```
def min_max(nums):
return min(nums), max(nums)
# return values can be assigned to a single variable as a tuple
nums = [1, 2, 3]
min_max_num = min_max(nums)
min_max_num
# return values can be assigned into multiple variables using tuple unpacking
min_num, max_num = min_max(nums)
print(min_num)
print(max_num)
```
[<a href="#Python-Quick-Reference-by-Data-School">Back to top</a>]
## 12. Anonymous (Lambda) Functions
- Primarily used to temporarily define a function for use by another function
```
# define a function the "usual" way
def squared(x):
return x**2
# define an identical function using lambda
squared = lambda x: x**2
```
**Sort a list of strings by the last letter:**
```
# without using lambda
simpsons = ['homer', 'marge', 'bart']
def last_letter(word):
return word[-1]
sorted(simpsons, key=last_letter)
# using lambda
sorted(simpsons, key=lambda word: word[-1])
```
[<a href="#Python-Quick-Reference-by-Data-School">Back to top</a>]
## 13. For Loops and While Loops
**`range` returns a list of integers (Python 2) or a sequence (Python 3):**
```
# includes the start value but excludes the stop value
range(0, 3)
# default start value is 0
range(3)
# third argument is the step value
range(0, 5, 2)
# Python 2 only: use xrange to create a sequence rather than a list (saves memory)
xrange(100, 100000, 5)
```
**`for` loops:**
```
# not the recommended style
fruits = ['apple', 'banana', 'cherry']
for i in range(len(fruits)):
print(fruits[i].upper())
# recommended style
for fruit in fruits:
print(fruit.upper())
# iterate through two things at once (using tuple unpacking)
family = {'dad':'homer', 'mom':'marge', 'size':6}
for key, value in family.items():
print(key, value)
# use enumerate if you need to access the index value within the loop
for index, fruit in enumerate(fruits):
print(index, fruit)
```
**`for`/`else` loop:**
```
for fruit in fruits:
if fruit == 'banana':
print('Found the banana!')
break # exit the loop and skip the 'else' block
else:
# this block executes ONLY if the for loop completes without hitting 'break'
print("Can't find the banana")
```
**`while` loop:**
```
count = 0
while count < 5:
print('This will print 5 times')
count += 1 # equivalent to 'count = count + 1'
```
[<a href="#Python-Quick-Reference-by-Data-School">Back to top</a>]
## 14. Comprehensions
**List comprehension:**
```
# for loop to create a list of cubes
nums = [1, 2, 3, 4, 5]
cubes = []
for num in nums:
cubes.append(num**3)
cubes
# equivalent list comprehension
cubes = [num**3 for num in nums]
cubes
# for loop to create a list of cubes of even numbers
cubes_of_even = []
for num in nums:
if num % 2 == 0:
cubes_of_even.append(num**3)
cubes_of_even
# equivalent list comprehension
# syntax: [expression for variable in iterable if condition]
cubes_of_even = [num**3 for num in nums if num % 2 == 0]
cubes_of_even
# for loop to cube even numbers and square odd numbers
cubes_and_squares = []
for num in nums:
if num % 2 == 0:
cubes_and_squares.append(num**3)
else:
cubes_and_squares.append(num**2)
cubes_and_squares
# equivalent list comprehension (using a ternary expression)
# syntax: [true_condition if condition else false_condition for variable in iterable]
cubes_and_squares = [num**3 if num % 2 == 0 else num**2 for num in nums]
cubes_and_squares
# for loop to flatten a 2d-matrix
matrix = [[1, 2], [3, 4]]
items = []
for row in matrix:
for item in row:
items.append(item)
items
# equivalent list comprehension
items = [item for row in matrix
for item in row]
items
```
**Set comprehension:**
```
fruits = ['apple', 'banana', 'cherry']
unique_lengths = {len(fruit) for fruit in fruits}
unique_lengths
```
**Dictionary comprehension:**
```
fruit_lengths = {fruit:len(fruit) for fruit in fruits}
fruit_lengths
fruit_indices = {fruit:index for index, fruit in enumerate(fruits)}
fruit_indices
```
[<a href="#Python-Quick-Reference-by-Data-School">Back to top</a>]
## 15. Map and Filter
**`map` applies a function to every element of a sequence and returns a list (Python 2) or iterator (Python 3):**
```
simpsons = ['homer', 'marge', 'bart']
map(len, simpsons)
# equivalent list comprehension
[len(word) for word in simpsons]
map(lambda word: word[-1], simpsons)
# equivalent list comprehension
[word[-1] for word in simpsons]
```
**`filter` returns a list (Python 2) or iterator (Python 3) containing the elements from a sequence for which a condition is `True`:**
```
nums = range(5)
filter(lambda x: x % 2 == 0, nums)
# equivalent list comprehension
[num for num in nums if num % 2 == 0]
```
[<a href="#Python-Quick-Reference-by-Data-School">Back to top</a>]
| github_jupyter |
# Python Data Analytics
<img src="images/pandas_logo.png" alt="pandas" style="width: 400px;"/>
Pandas is a numerical package used extensively in data science. You can call the install the ``pandas`` package by
```
pip install pandas
```
Like ``numpy``, the underlying routines are written in C with improved performance
<a href="https://colab.research.google.com/github/ryan-leung/PHYS4650_Python_Tutorial/blob/master/notebooks/04-Introduction-to-Pandas.ipynb"><img align="right" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory">
</a>
```
import pandas
pandas.__version__
import pandas as pd
import numpy as np
```
# Built-In Documentation in jupyter
For example, to display all the contents of the pandas namespace, you can type
```ipython
In [3]: pd.<TAB>
```
And to display Pandas's built-in documentation, you can use this:
```ipython
In [4]: pd?
```
# The Pandas Series Object
A Pandas Series is a one-dimensional array of indexed data.
```
data = pd.Series([1., 2., 3., 4.])
data
data = pd.Series([1, 2, 3, 4])
data
```
To retrieve back the underlying numpy array, we have the values attribute
```
data.values
```
The ``index`` is an array-like object of type ``pd.Index``.
```
data.index
```
Slicing and indexing just like Python standard ``list``
```
data[1]
data[1:3]
```
# The Pandas Index
The index is useful to denote each record, the datatypes of the index can be varied. You can think of another numpy array binded to the data array.
```
data = pd.Series([1, 2, 3, 4],
index=['a', 'b', 'c', 'd'])
data
```
If we supply a dictionary to the series, it will be constructed with an index.
By default, a ``Series`` will be created where the index is drawn from the sorted keys.
```
location = {
'Berlin': (52.5170365, 13.3888599),
'London': (51.5073219, -0.1276474),
'Sydney': (-33.8548157, 151.2164539),
'Tokyo': (34.2255804, 139.294774527387),
'Paris': (48.8566101, 2.3514992),
'Moscow': (46.7323875, -117.0001651)
}
location = pd.Series(location)
location
location['Berlin']
```
Unlike a dictionary, though, the Series also supports array-style operations such as slicing
```
location['London':'Paris']
```
# The Pandas DataFrame Object
The pandas dataframe object is a very powerful ``table`` like object.
```
location = {
'Berlin': (52.5170365, 13.3888599),
'London': (51.5073219, -0.1276474),
'Sydney': (-33.8548157, 151.2164539),
'Tokyo': (34.2255804, 139.294774527387),
'Paris': (48.8566101, 2.3514992),
'Moscow': (46.7323875, -117.0001651)
}
location = pd.DataFrame(location)
location
# Switching rows to columns is as easy as a transpose
location.T
# Change the columns by .columns attribute
location = location.T
location.columns = ['lat', 'lon']
location
location.index
location.columns
```
# Read Data
pandas has built-in data readers, you can type ``pd.read<TAB>`` to see what data format does it support:

we will focus in csv file which is widely used
We have some data downloaded from airbnb, you can find it in the folder, you may also download the file by executing the following code:
```
import urllib.request
urllib.request.urlretrieve(
'http://data.insideairbnb.com/taiwan/northern-taiwan/taipei/2018-11-27/visualisations/listings.csv',
'airbnb_taiwan_listing.csv'
)
urllib.request.urlretrieve(
'http://data.insideairbnb.com/china/hk/hong-kong/2018-11-12/visualisations/listings.csv',
'airbnb_hongkong_listing.csv'
)
```
# Read CSV files
```
airbnb_taiwan = pd.read_csv('airbnb_taiwan_listing.csv')
airbnb_taiwan
airbnb_hongkong = pd.read_csv('airbnb_hongkong_listing.csv')
airbnb_hongkong
```
# Filter data
```
mask = airbnb_hongkong['price'] > 1000
airbnb_hongkong[mask]
# In one line :
airbnb_taiwan[airbnb_taiwan['price'] > 4000]
```
# Missing Data in Pandas
Missing data is very important in pandas dataframe/series operations. Pandas do element-to-element operations based on index. If the index does not match, it will produce a not-a-number (NaN) results.
```
A = pd.Series([2, 4, 6], index=[0, 1, 2])
B = pd.Series([1, 3, 5], index=[1, 2, 3])
A + B
A.add(B, fill_value=0)
```
The following table lists the upcasting conventions in Pandas when NA values are introduced:
|Typeclass | Conversion When Storing NAs | NA Sentinel Value |
|--------------|-----------------------------|------------------------|
| ``floating`` | No change | ``np.nan`` |
| ``object`` | No change | ``None`` or ``np.nan`` |
| ``integer`` | Cast to ``float64`` | ``np.nan`` |
| ``boolean`` | Cast to ``object`` | ``None`` or ``np.nan`` |
Pandas treats ``None`` and ``NaN`` as essentially interchangeable for indicating missing or null values. They are convention functions to replace and find these values:
- ``isnull()``: Generate a boolean mask indicating missing values
- ``notnull()``: Opposite of ``isnull()``
- ``dropna()``: Return a filtered version of the data
- ``fillna()``: Return a copy of the data with missing values filled or imputed
```
# Fill Zero
(A + B).fillna(0)
# forward-fill
(A + B).fillna(method='ffill')
# back-fill
(A + B).fillna(method='bfill')
```
# Data Aggregations
we will use back the airbnb data to demonstrate data aggerations
```
airbnb_hongkong['price'].describe()
```
The following table summarizes some other built-in Pandas aggregations:
| Aggregation | Description |
|--------------------------|---------------------------------|
| ``count()`` | Total number of items |
| ``first()``, ``last()`` | First and last item |
| ``mean()``, ``median()`` | Mean and median |
| ``min()``, ``max()`` | Minimum and maximum |
| ``std()``, ``var()`` | Standard deviation and variance |
| ``mad()`` | Mean absolute deviation |
| ``prod()`` | Product of all items |
| ``sum()`` | Sum of all items |
```
data_grouped = airbnb_hongkong.groupby(['neighbourhood'])
data_mean = data_grouped['price'].mean()
data_mean
data_mean = airbnb_taiwan.groupby(['neighbourhood'])['price'].mean()
data_mean
airbnb_taiwan.groupby(['room_type']).id.count()
airbnb_hongkong.groupby(['room_type']).id.count()
airbnb_taiwan.groupby(['room_type'])['price'].describe()
airbnb_hongkong.groupby(['room_type'])['price'].describe()
```
# Combining Two or more dataframe
```
airbnb = pd.concat([airbnb_taiwan, airbnb_hongkong], keys=['taiwan', 'hongkong'])
airbnb
airbnb.index
airbnb.index = airbnb.index.droplevel(level=1)
airbnb.index
airbnb.groupby(['room_type', airbnb.index])['price'].describe()
```
# Easy Plotting in pandas
```
airbnb_taiwan.groupby(['room_type']).id.count()
%matplotlib inline
c = airbnb_taiwan.groupby(['room_type']).id.count()
c.plot.bar()
c = airbnb_taiwan.groupby(['room_type']).id.count().rename("count")
d = airbnb_taiwan.id.count()
(c / d * 100).plot.bar()
```
# Time series data
Time seies data refers to metrics that has a time dimensions, such as stocks data and weather. In this example, we will look at some random time-series data:
```
import numpy as np
ts = pd.Series(np.random.randn(1000), index=pd.date_range('2016-01-01', periods=1000))
ts.plot()
ts = ts.cumsum()
ts.plot()
```
# Datetime index filtering
```
ts.index
ts['2016-02-01':'2016-05-01'].plot()
```
# Summary
Pandas is a very helpful packages in data science, it helps you check and visualize data very quickly. This files contains only a very small portion of the pandas function. please read other materials for more informations.
| github_jupyter |
# Effective Data Visualization
## PyCon 2020
## Husni Almoubayyed [https://husni.space]
## Intro on packages:
- **Matplotlib and Seaborn**: Main plotting package in python is called matplotlib. Matplotlib is the base for another package which builds on top of it called Seaborn. We will use Seaborn when possible as it makes most things a lot more easier and allows us to achieve plots with sensible choices and significantly less lines of code. We will still use matplotlib for some things and it is important to understand every time Seaborn creates a plot it is calling Matplotlib in the background (it is also sometimes calling other things like statsmodels in the background to do some statistical calculations)
Matplotlib and Seaborn syntax is usually used as follows: plt. or sns.*typeofgraph*(arguments)
arguments are usually X and Y coordinates (or names of X and Y columns in a dataframe), colors, sizes, etc.
- **Pandas** is a library that handles [Pan]el [Da]ta. Basically it allows us to manipulate data in tables a lot more easily.
- **Numpy** is a python library that contains all the standard numerical operations you might wanna do
- **Sci-Kit Learn (sklearn)** is a widely used library that you can use to do most common non-deep machine learning methods.
## Intro to datasets:
We will use a few standard datasets throughout this tutorial. These can be imported from seaborn as will be shown later:
- **diamonds**: data on diamonds with prices, carats, color, clarity, cut, etc.
- **flights**: number of passengers in each month for each year for a few years in the ~50s
- **iris**: famous biology dataset that quantifies the morphologic variation of Iris flowers of three related species
- **titanic**: data on all titanic passengers including survival, age, ticket price paid, etc.
- **anscombe**: this is compiled of 4 different datasets that have the same first and second moments but look dramatically different
- **digits**: handwritten data of digits, used widely in machine learning
Other datasets that are not directly imported from seaborn:
- **financial data**: this will be requested in real time from yahoo finance using pandas.
- **CoViD-19 data**: https://usafactsstatic.blob.core.windows.net/public/data/covid-19/covid_confirmed_usafacts.csv, credits to usafacts.org
## Installation Instructions
Install pip https://pip.pypa.io/en/stable/installing/
In the command line: $pip install notebook
or
Install conda https://www.anaconda.com/distribution/ (for Python >3.7)
and then run:
```
!pip install --upgrade matplotlib numpy scipy sklearn pandas seaborn plotly plotly-geo pandas_datareader
```
You might need to restart the kernel at this point to use any newly installed packages
Alternatively, you can go to bit.ly/PyConViz2020 to use a Colab hosted version of this notebook.
```
# import numpy and matplotlib and setup inline plots by running:
%pylab inline
import seaborn as sns
import pandas as pd
sns.set_style('darkgrid')
sns.set_context('notebook', font_scale=1.5)
sns.set_palette('colorblind')
# set the matplotlib backend to a higher-resolution option, on macOS, this is:
%config InlineBackend.figure_format = 'retina'
# set larger figure size for the rest of this notebook
matplotlib.rcParams['figure.figsize'] = 12, 8
```
## Data Exploration
```
anscombe = sns.load_dataset('anscombe')
for dataset in ['I','II','III','IV']:
print(anscombe[anscombe['dataset']==dataset].describe())
sns.lmplot(x='x', y='y', data=anscombe[anscombe['dataset']=='I'], height=8)
iris = sns.load_dataset('iris')
iris.head()
sns.scatterplot('petal_length', 'petal_width', hue='species', data=iris)
```
## Exercise:
On the same plot, fit 3 linear models for the 3 different iris species with the same x and y axes
```
sns.jointplot('petal_length', 'petal_width', data=iris, height=8, kind='kde')
sns.pairplot(iris, height=8, hue='species')
```
How about categorical data?
We can make boxplots and violin plots simply by running:
```
sns.catplot()
```
**Exercise:** Load up the flights dataset, plot a linear model of the passengers number as a function of year, one for each month of the year.
**Exercise:** Load up the diamonds dataset from seaborn. Plot the price as a function of carat, with different color grades colored differently. choose a small marker size and change the transparency (alpha agrument) to a smaller value than 1. Add some jitter to the x values to make them clearer.
**Exercise:** Load up the Titanic dataset from seaborn. Make a boxplot of the fare of the ticket paid against whether a person survived or not.
## Polar coordinates
```
plt.quiver??
X = np.random.uniform(0, 10, 100)
Y = np.random.uniform(0, 1, 100)
U = np.ones_like(X)
V = np.ones_like(Y)
f = plt.figure()
ax = f.add_subplot(111)
ax.quiver(X, Y, U, V, headlength=0, headaxislength=0, color='steelblue')
theta = np.linspace(0,2*np.pi,100)
r = np.linspace(0, 1, 100)
dr = 1
dt = 0
U = dr * cos(theta) - dt * sin (theta)
V = dr * sin(theta) + dt * cos(theta)
f = plt.figure()
ax = f.add_subplot(111, polar=True)
ax.quiver(theta, r, U, V, headlength=0, headaxislength=0, color='steelblue')
theta = np.linspace(0,2*np.pi,100)
r = np.random.uniform(0, 1, 100)
U = dr * cos(theta)
V = dr * sin(theta)
f = plt.figure()
ax = f.add_subplot(111, polar=True)
ax.quiver(theta, r, U, V, headlength=0, headaxislength=0, color='steelblue')
```
**Exercise 1:** radial plot with all sticks starting at a radius of 1
**Exercise 2:** all sticks are horizontal
**Exercise 3:** Use a 'mollweide' projection using the projection argument of add_subplot(). Use horizontal sticks now but make sure your sticks span the entire space.
# 2. Density Estimation
Often when we are making plots, we are trying to estimate the underlying distribution from which it was randomly drawn, this is known as Density Estimation in statistics. The simplest density estimator that does not make particular assumptions on the distribution of the data (we call this nonparametric) is the histogram.
## Histograms
```
# import out first dataset, an example from biology
iris = sns.load_dataset('iris')
iris.head()
data = iris['sepal_length']
plt.
data = iris['sepal_length']
f, (ax1, ax2) = plt.subplots(1, 2)
ax1.
ax2.
```
Formally, The histogram estimator is $$ \hat{p}(x) = \frac{\hat{\theta_j}}{h} $$ where $$ \hat{\theta_j} = \frac{1}{n} \sum_{i=1}^n I(X_i \in B_j ) $$
We can calculate the mean squared error, which is a metric that tells us how well our estimator is, it turns out to be: $$MSE(x) = bias^2(x) + Var(x) = Ch^2 + \frac{C}{nh} $$
minimized by choosing $h = (\frac{C}{n})^{1/3}$, resulting in a risk (the expected value of the MSE) of:
$$ R = \mathcal{O}(\frac{1}{n})^{2/3}$$
This means that
- There is a bias-variance tradeoff when it comes to choosing the width of the bins, lower width ($h$), means more bias and less variance. There is no choice of $h$ that optimizes both.
- The risk goes down at a pretty slow rate as the number of datapoints increases, which begs the question, is there a better estimator that converges more quickly? The answer is yes, this is achieved by:
## Kernel Density Estimation
Kernels follow the conditions:
$$ K(x) \geq 0, \int K(x) dx = 1, \int x K(x) dx = 0$$
```
sns.
```
So how is this better than the histogram?
We can again calculate the MSE, which turns out to be:
$$MSE(x) = bias^2(x) + Var(x) = C_1h^4 + \frac{C_2}{nh}$$
minimized by choosing $ h = (\frac{C_1}{4nC_2})^{1/5} $, giving a risk of:
$$ R_{KDE} = \mathcal{O}(\frac{1}{n})^{4/5} < R_{histogram}$$
This still has a bias-variance tradeoff, but the estimator converges faster than in the case of histograms. Can we do even better? The answer is no, due to something in statistics called the minimax theorem.
**Exercise**: Instead of using just petal length, consider a 2D distribution with the two axes being petal length and petal width. Plot the distribution, the Histogram of the distribution and the KDE of the distribution. Make sure you play around with bin numbers and bandwidth to get a reasonably satisfying plot
```
data=iris[['petal_length', 'petal_width']]
sns.scatterplot('petal_length', 'petal_width', data=iris)
sns.distplot(iris['petal_length'])
```
# 3. Visualizing High Dimensional Datasets
```
from sklearn.decomposition import PCA
from sklearn.datasets import load_digits
from sklearn.datasets import make_swiss_roll
import mpl_toolkits.mplot3d as p3
from sklearn.cluster import AgglomerativeClustering
from sklearn.neighbors import kneighbors_graph
from sklearn.manifold import TSNE
digits = load_digits()
shape(digits['data'])
```
## Principal Component Analysis
PCA computes the linear projections of greatest variance from the top eigenvectors of the data covariance matrix
Check out some more cool visualization of PCA at https://setosa.io/ev/principal-component-analysis/ and read more about the math and applications at https://www.cs.cmu.edu/~bapoczos/other_presentations/PCA_24_10_2009.pdf
**Exercise:** Use PCA to reduce the dimensionality of the digits dataset. Plot them color-coded by the different classes of digits.
### Failures of PCA
```
X, t = make_swiss_roll(1000, 0.05)
ward = AgglomerativeClustering(n_clusters=5,
connectivity=kneighbors_graph(X, n_neighbors=5, include_self=False),
linkage='ward').fit(X)
labels = ward.labels_
fig = plt.figure()
ax = p3.Axes3D(fig)
ax.view_init(7, -80)
for label in np.unique(labels):
ax.scatter(X[labels == label, 0], X[labels == label, 1], X[labels == label, 2])
pca = PCA(2)
projected = pca.fit_transform(X)
for label in np.unique(labels):
sns.scatterplot(projected[labels == label, 0], projected[labels == label, 1],
color=plt.cm.jet(float(label) / np.max(labels + 1)), marker='.')
```
## t-Distributed Stochastic Neighbor Embedding
Converts similarities between data points to joint probabilities and tries to minimize the Kullback-Leibler divergence between the joint probabilities of the low-dimensional embedding and the high-dimensional data.
First, t-SNE constructs a probability distribution over pairs of high-dimensional objects in such a way that similar objects have a high probability of being picked while dissimilar points have an extremely small probability of being picked. Second, t-SNE defines a similar probability distribution over the points in the low-dimensional map, and it minimizes the Kullback–Leibler divergence (KL divergence ) between the two distributions with respect to the locations of the points in the map.
For more details on t-SNE, check out the original paper http://www.jmlr.org/papers/volume9/vandermaaten08a/vandermaaten08a.pdf
```
tSNE = TSNE(learning_rate=10,
perplexity=30)
projected = tSNE.fit_transform(X)
plt.scatter(projected[:, 0], projected[:, 1],
c=labels, alpha=0.3,
cmap=plt.cm.get_cmap('Paired', 5))
#plt.colorbar()
```
**Exercise:** Do this again for the digits dataset. Does this look better than PCA?
# 4. Interactive Visualization
```
# import libraries we're gonna use
import pandas_datareader.data as web
import datetime
import plotly.figure_factory as ff
import plotly.graph_objs as go
start = datetime.datetime(2008, 1, 1)
end = datetime.datetime(2018, 1, 1)
# This fetches the stock prices for the S%P 500 for the dates we selected from Yahoo Finance.
spy_df =
data = go.Scatter(x=spy_df.Date, y=spy_df.Close)
go.Figure(data)
```
**Exercise:** A candlestick chart is a powerful chart in finance that shows the starting price, closing price, highest price and lowerst price of a trading day. Create a aandlestick chart of the first 90 days of the data. You can find Candlestick in the 'go' module.
**Exercise:** It's hard to compare AAPL to SPY when viewed as is. Can you plot this again in a way that makes the returns of AAPL more easily comparable to the returns of the benchmark SPY?
```
covidf = pd.read_csv('~/Downloads/covid_confirmed_usafacts.csv',
dtype={"countyFIPS": str})
covidf.head()
values=covidf['4/5/20']
colorscale = ["#f7fbff","#deebf7","#c6dbef","#9ecae1",
"#6baed6","#4292c6","#2171b5","#08519c","#08306b"]
endpts = list(np.logspace(1, 5, len(colorscale) - 1))
fig = ff.create_choropleth(
fips=covidf['countyFIPS'], values=covidf['4/9/20'],# scope=['usa'],
binning_endpoints=endpts, colorscale=colorscale,
title_text = 'CoViD-19 Confirmed cases as of 4/9/20',
legend_title = '# of cases'
)
go.Figure(fig)
```
Many more types of plotly charts are available with examples here https://plotly.com/python/
# Effective Communication through Plotting
```
image = [[i for i in range(100)]]*10
sns.heatmap(image, cmap='jet', square=True)
```
## Color
```
# code snippet from Jake Vandeplas https://jakevdp.github.io/blog/2014/10/16/how-bad-is-your-colormap/
def grayify_cmap(cmap):
"""Return a grayscale version of the colormap"""
cmap = plt.cm.get_cmap(cmap)
colors = cmap(np.arange(cmap.N))
# convert RGBA to perceived greyscale luminance
# cf. http://alienryderflex.com/hsp.html
RGB_weight = [0.299, 0.587, 0.114]
luminance = np.sqrt(np.dot(colors[:, :3] ** 2, RGB_weight))
colors[:, :3] = luminance[:, np.newaxis]
return cmap.from_list(cmap.name + "_grayscale", colors, cmap.N)
flights = sns.load_dataset("flights").pivot("month", "year", "passengers")
sns.heatmap(flights, cmap='jet')
sns.heatmap(flights, cmap=grayify_cmap('jet'))
```
## 3 Types of Viable Color palettes/colormaps:
### 1. Perceptually uniform sequential
```
sns.heatmap(flights, cmap='viridis')
sns.heatmap(flights, cmap='Purples')
```
## 2. Diverging
```
import pandas as pd
pit_climate_df = pd.DataFrame(
dict(Month = ['Jan', 'Feb', 'Mar', 'Apr', 'May', 'Jun',
'Jul', 'Aug', 'Sep', 'Oct', 'Nov', 'Dec'],
High = [2, 3, 10, 16, 22, 27, 28, 28, 24, 17, 10, 5],
Low = [-7, -5, 0, 5, 11, 16, 18, 18, 14,7, 3, -2])
)
pit_climate_df.head()
sns.heatmap(pit_climate_df[['High', 'Low']].T,
cmap='coolwarm',
center=0,#np.mean(pit_climate_df[['High', 'Low']].mean().mean()),
square=True,
xticklabels=pit_climate_df['Month'])
```
## 3. Categorical
example from before:
```
plt.scatter(projected[:, 0], projected[:, 1],
c=digits.target, alpha=0.3,
cmap=plt.cm.get_cmap('Paired', 10))
plt.colorbar()
from IPython.display import Image
Image('Resources/51417489_2006270206137719_6713863014199590912_n.png')
Image('Resources/50283372_1999138550184218_5288878489854803968_o.png')
```
You can also specify a color palette to use for the rest of a notebook or script by running
Other things to consider:
* Use salient marker types, full list at https://matplotlib.org/3.2.1/api/markers_api.html
```
d1 = np.random.uniform(-2.5, 2.5, (100, 100))
d2 = np.random.randn(5,5)
sns.scatterplot(d1[:,0], d1[:,1], marker='+', color='steelblue')
sns.scatterplot(d2[:,0], d2[:,1], color='steelblue')
sns.lmplot('petal_length', 'petal_width', iris,
height=10,
hue='species',
markers=['1','2','3'],
fit_reg=False)
sns.scatterplot(d1[:,0], d1[:,1], marker='+', color='steelblue')
```
There are more than 2 axes on a 2-dimensional screen. Can you think of ways to include more axes?
We can use each of the following to map to an axis:
- color
- size (for numerical data)
- shape (for categorical data)
- literally making a 3D plot (as in the swiss roll, useful in the case of 3 spatial dimensions)
```
sns.set_palette('colorblind')
```
Read more on choosing colors at:
* https://seaborn.pydata.org/tutorial/color_palettes.html
* https://matplotlib.org/3.1.0/tutorials/colors/colormaps.html
One of my favorite resources on clarity in plotting:
* http://blogs.nature.com/methagora/2013/07/data-visualization-points-of-view.html
New interesting package that we don't have time for today but is definitely worth mentioning. Makes visualization more intuitive by making it declarative is Altair https://altair-viz.github.io
| github_jupyter |
```
# Import the dependencies.(6.5.2)
import pandas as pd
import gmaps
import requests
# Import the API key.
from config import g_key
# Store the CSV you saved created in part one into a DataFrame.(6.5.2)
city_data_df = pd.read_csv("weather_data/cities.csv")
city_data_df.head()
# Configure gmaps to use your Google API key.(6.5.2)
gmaps.configure(api_key=g_key)
# Ask the customer to add a minimum and maximum temperature value.(6.5.3)
min_temp = float(input("What is the minimum temperature you would like for your trip? "))
max_temp = float(input("What is the maximum temperature you would like for your trip? "))
# Filter the dataset to find the cities that fit the criteria.(6.5.3)
preferred_cities_df = city_data_df.loc[(city_data_df["Max Temp"] <= max_temp) & \
(city_data_df["Max Temp"] >= min_temp)]
preferred_cities_df.head(10)
preferred_cities_df.count()
# Create DataFrame called hotel_df to store hotel names along with city, country, max temp, and coordinates.(6.5.4)
hotel_df = preferred_cities_df[["City", "Country", "Max Temp", "Lat", "Lng"]].copy()
hotel_df["Hotel Name"] = ""
hotel_df.head(10)
# Set parameters to search for a hotel.(6.5.4)
params = {
"radius": 5000,
"type": "lodging",
"key": g_key
}
for index, row in hotel_df.iterrows():
# get lat, lng from df
lat = row["Lat"]
lng = row["Lng"]
params["location"] = f"{lat}, {lng}"
base_url = "https://maps.googleapis.com/maps/api/place/nearbysearch/json"
# Make request and get the JSON data from the search.
hotels = requests.get(base_url, params=params).json()
try:
hotel_df.loc[index, "Hotel Name"] = hotels["results"][0]["name"]
except:
print("Hotel not found.")
hotel_df
# Add a heatmap of temperature for the vacation spots.(6.5.4)
locations = hotel_df[["Lat", "Lng"]]
max_temp = hotel_df["Max Temp"]
fig = gmaps.figure(center=(30.0, 31.0), zoom_level=1.5)
heat_layer = gmaps.heatmap_layer(locations, weights=max_temp, dissipating=False,
max_intensity=300, point_radius=4)
fig.add_layer(heat_layer)
# Call the figure to plot the data.
fig
# Add a heatmap of temperature for the vacation spots and marker for each city.(6.5.4)
locations = hotel_df[["Lat", "Lng"]]
max_temp = hotel_df["Max Temp"]
fig = gmaps.figure(center=(30.0, 31.0), zoom_level=1.5)
heat_layer = gmaps.heatmap_layer(locations, weights=max_temp,
dissipating=False, max_intensity=300, point_radius=4)
marker_layer = gmaps.marker_layer(locations)
fig.add_layer(heat_layer)
fig.add_layer(marker_layer)
# Call the figure to plot the data.
fig
info_box_template = """
<dl>
<dt>Hotel Name</dt><dd>{Hotel Name}</dd>
<dt>City</dt><dd>{City}</dd>
<dt>Country</dt><dd>{Country}</dd>
<dt>Max Temp</dt><dd>{Max Temp} °F</dd>
</dl>
"""
# Store the DataFrame Row.
hotel_info = [info_box_template.format(**row) for index, row in hotel_df.iterrows()]
# Add a heatmap of temperature for the vacation spots and a pop-up marker for each city.
locations = hotel_df[["Lat", "Lng"]]
max_temp = hotel_df["Max Temp"]
fig = gmaps.figure(center=(30.0, 31.0), zoom_level=1.5)
heat_layer = gmaps.heatmap_layer(locations, weights=max_temp,dissipating=False,
max_intensity=300, point_radius=4)
marker_layer = gmaps.marker_layer(locations, info_box_content=hotel_info)
fig.add_layer(heat_layer)
fig.add_layer(marker_layer)
# Call the figure to plot the data.
fig
```
| github_jupyter |
# Mask R-CNN - Train on Shapes Dataset
This notebook shows how to train Mask R-CNN on your own dataset. To keep things simple we use a synthetic dataset of shapes (squares, triangles, and circles) which enables fast training. You'd still need a GPU, though, because the network backbone is a Resnet101, which would be too slow to train on a CPU. On a GPU, you can start to get okay-ish results in a few minutes, and good results in less than an hour.
The code of the *Shapes* dataset is included below. It generates images on the fly, so it doesn't require downloading any data. And it can generate images of any size, so we pick a small image size to train faster.
```
import os
import sys
import random
import math
import re
import time
import numpy as np
import cv2
import matplotlib
import matplotlib.pyplot as plt
from config import Config
import utils
import model as modellib
import visualize
from model import log
%matplotlib inline
# Root directory of the project
ROOT_DIR = os.getcwd()
# Directory to save logs and trained model
MODEL_DIR = os.path.join(ROOT_DIR, "logs")
# Local path to trained weights file
COCO_MODEL_PATH = os.path.join(ROOT_DIR, "mask_rcnn_coco.h5")
# Download COCO trained weights from Releases if needed
if not os.path.exists(COCO_MODEL_PATH):
utils.download_trained_weights(COCO_MODEL_PATH)
```
## Configurations
```
class ShapesConfig(Config):
"""Configuration for training on the toy shapes dataset.
Derives from the base Config class and overrides values specific
to the toy shapes dataset.
"""
# Give the configuration a recognizable name
NAME = "shapes"
# Train on 1 GPU and 8 images per GPU. We can put multiple images on each
# GPU because the images are small. Batch size is 8 (GPUs * images/GPU).
GPU_COUNT = 1
IMAGES_PER_GPU = 8
# Number of classes (including background)
NUM_CLASSES = 1 + 3 # background + 3 shapes
# Use small images for faster training. Set the limits of the small side
# the large side, and that determines the image shape.
IMAGE_MIN_DIM = 128
IMAGE_MAX_DIM = 128
# Use smaller anchors because our image and objects are small
RPN_ANCHOR_SCALES = (8, 16, 32, 64, 128) # anchor side in pixels
# Reduce training ROIs per image because the images are small and have
# few objects. Aim to allow ROI sampling to pick 33% positive ROIs.
TRAIN_ROIS_PER_IMAGE = 32
# Use a small epoch since the data is simple
STEPS_PER_EPOCH = 100
# use small validation steps since the epoch is small
VALIDATION_STEPS = 5
config = ShapesConfig()
config.display()
```
## Notebook Preferences
```
def get_ax(rows=1, cols=1, size=8):
"""Return a Matplotlib Axes array to be used in
all visualizations in the notebook. Provide a
central point to control graph sizes.
Change the default size attribute to control the size
of rendered images
"""
_, ax = plt.subplots(rows, cols, figsize=(size*cols, size*rows))
return ax
```
## Dataset
Create a synthetic dataset
Extend the Dataset class and add a method to load the shapes dataset, `load_shapes()`, and override the following methods:
* load_image()
* load_mask()
* image_reference()
```
class ShapesDataset(utils.Dataset):
"""Generates the shapes synthetic dataset. The dataset consists of simple
shapes (triangles, squares, circles) placed randomly on a blank surface.
The images are generated on the fly. No file access required.
"""
def load_shapes(self, count, height, width):
"""Generate the requested number of synthetic images.
count: number of images to generate.
height, width: the size of the generated images.
"""
# Add classes
self.add_class("shapes", 1, "square")
self.add_class("shapes", 2, "circle")
self.add_class("shapes", 3, "triangle")
# Add images
# Generate random specifications of images (i.e. color and
# list of shapes sizes and locations). This is more compact than
# actual images. Images are generated on the fly in load_image().
for i in range(count):
bg_color, shapes = self.random_image(height, width)
self.add_image("shapes", image_id=i, path=None,
width=width, height=height,
bg_color=bg_color, shapes=shapes)
def load_image(self, image_id):
"""Generate an image from the specs of the given image ID.
Typically this function loads the image from a file, but
in this case it generates the image on the fly from the
specs in image_info.
"""
info = self.image_info[image_id]
bg_color = np.array(info['bg_color']).reshape([1, 1, 3])
image = np.ones([info['height'], info['width'], 3], dtype=np.uint8)
image = image * bg_color.astype(np.uint8)
for shape, color, dims in info['shapes']:
image = self.draw_shape(image, shape, dims, color)
return image
def image_reference(self, image_id):
"""Return the shapes data of the image."""
info = self.image_info[image_id]
if info["source"] == "shapes":
return info["shapes"]
else:
super(self.__class__).image_reference(self, image_id)
def load_mask(self, image_id):
"""Generate instance masks for shapes of the given image ID.
"""
info = self.image_info[image_id]
shapes = info['shapes']
count = len(shapes)
mask = np.zeros([info['height'], info['width'], count], dtype=np.uint8)
for i, (shape, _, dims) in enumerate(info['shapes']):
mask[:, :, i:i+1] = self.draw_shape(mask[:, :, i:i+1].copy(),
shape, dims, 1)
# Handle occlusions
occlusion = np.logical_not(mask[:, :, -1]).astype(np.uint8)
for i in range(count-2, -1, -1):
mask[:, :, i] = mask[:, :, i] * occlusion
occlusion = np.logical_and(occlusion, np.logical_not(mask[:, :, i]))
# Map class names to class IDs.
class_ids = np.array([self.class_names.index(s[0]) for s in shapes])
return mask, class_ids.astype(np.int32)
def draw_shape(self, image, shape, dims, color):
"""Draws a shape from the given specs."""
# Get the center x, y and the size s
x, y, s = dims
if shape == 'square':
cv2.rectangle(image, (x-s, y-s), (x+s, y+s), color, -1)
elif shape == "circle":
cv2.circle(image, (x, y), s, color, -1)
elif shape == "triangle":
points = np.array([[(x, y-s),
(x-s/math.sin(math.radians(60)), y+s),
(x+s/math.sin(math.radians(60)), y+s),
]], dtype=np.int32)
cv2.fillPoly(image, points, color)
return image
def random_shape(self, height, width):
"""Generates specifications of a random shape that lies within
the given height and width boundaries.
Returns a tuple of three valus:
* The shape name (square, circle, ...)
* Shape color: a tuple of 3 values, RGB.
* Shape dimensions: A tuple of values that define the shape size
and location. Differs per shape type.
"""
# Shape
shape = random.choice(["square", "circle", "triangle"])
# Color
color = tuple([random.randint(0, 255) for _ in range(3)])
# Center x, y
buffer = 20
y = random.randint(buffer, height - buffer - 1)
x = random.randint(buffer, width - buffer - 1)
# Size
s = random.randint(buffer, height//4)
return shape, color, (x, y, s)
def random_image(self, height, width):
"""Creates random specifications of an image with multiple shapes.
Returns the background color of the image and a list of shape
specifications that can be used to draw the image.
"""
# Pick random background color
bg_color = np.array([random.randint(0, 255) for _ in range(3)])
# Generate a few random shapes and record their
# bounding boxes
shapes = []
boxes = []
N = random.randint(1, 4)
for _ in range(N):
shape, color, dims = self.random_shape(height, width)
shapes.append((shape, color, dims))
x, y, s = dims
boxes.append([y-s, x-s, y+s, x+s])
# Apply non-max suppression wit 0.3 threshold to avoid
# shapes covering each other
keep_ixs = utils.non_max_suppression(np.array(boxes), np.arange(N), 0.3)
shapes = [s for i, s in enumerate(shapes) if i in keep_ixs]
return bg_color, shapes
# Training dataset
dataset_train = ShapesDataset()
dataset_train.load_shapes(500, config.IMAGE_SHAPE[0], config.IMAGE_SHAPE[1])
dataset_train.prepare()
# Validation dataset
dataset_val = ShapesDataset()
dataset_val.load_shapes(50, config.IMAGE_SHAPE[0], config.IMAGE_SHAPE[1])
dataset_val.prepare()
# Load and display random samples
image_ids = np.random.choice(dataset_train.image_ids, 4)
for image_id in image_ids:
image = dataset_train.load_image(image_id)
mask, class_ids = dataset_train.load_mask(image_id)
visualize.display_top_masks(image, mask, class_ids, dataset_train.class_names)
```
## Ceate Model
```
# Create model in training mode
model = modellib.MaskRCNN(mode="training", config=config,
model_dir=MODEL_DIR)
# Which weights to start with?
init_with = "coco" # imagenet, coco, or last
if init_with == "imagenet":
model.load_weights(model.get_imagenet_weights(), by_name=True)
elif init_with == "coco":
# Load weights trained on MS COCO, but skip layers that
# are different due to the different number of classes
# See README for instructions to download the COCO weights
model.load_weights(COCO_MODEL_PATH, by_name=True,
exclude=["mrcnn_class_logits", "mrcnn_bbox_fc",
"mrcnn_bbox", "mrcnn_mask"])
elif init_with == "last":
# Load the last model you trained and continue training
model.load_weights(model.find_last()[1], by_name=True)
```
## Training
Train in two stages:
1. Only the heads. Here we're freezing all the backbone layers and training only the randomly initialized layers (i.e. the ones that we didn't use pre-trained weights from MS COCO). To train only the head layers, pass `layers='heads'` to the `train()` function.
2. Fine-tune all layers. For this simple example it's not necessary, but we're including it to show the process. Simply pass `layers="all` to train all layers.
```
# Train the head branches
# Passing layers="heads" freezes all layers except the head
# layers. You can also pass a regular expression to select
# which layers to train by name pattern.
model.train(dataset_train, dataset_val,
learning_rate=config.LEARNING_RATE,
epochs=1,
layers='heads')
# Fine tune all layers
# Passing layers="all" trains all layers. You can also
# pass a regular expression to select which layers to
# train by name pattern.
model.train(dataset_train, dataset_val,
learning_rate=config.LEARNING_RATE / 10,
epochs=2,
layers="all")
# Save weights
# Typically not needed because callbacks save after every epoch
# Uncomment to save manually
# model_path = os.path.join(MODEL_DIR, "mask_rcnn_shapes.h5")
# model.keras_model.save_weights(model_path)
```
## Detection
```
class InferenceConfig(ShapesConfig):
GPU_COUNT = 1
IMAGES_PER_GPU = 1
inference_config = InferenceConfig()
# Recreate the model in inference mode
model = modellib.MaskRCNN(mode="inference",
config=inference_config,
model_dir=MODEL_DIR)
# Get path to saved weights
# Either set a specific path or find last trained weights
# model_path = os.path.join(ROOT_DIR, ".h5 file name here")
model_path = model.find_last()[1]
# Load trained weights (fill in path to trained weights here)
assert model_path != "", "Provide path to trained weights"
print("Loading weights from ", model_path)
model.load_weights(model_path, by_name=True)
# Test on a random image
image_id = random.choice(dataset_val.image_ids)
original_image, image_meta, gt_class_id, gt_bbox, gt_mask =\
modellib.load_image_gt(dataset_val, inference_config,
image_id, use_mini_mask=False)
log("original_image", original_image)
log("image_meta", image_meta)
log("gt_class_id", gt_class_id)
log("gt_bbox", gt_bbox)
log("gt_mask", gt_mask)
visualize.display_instances(original_image, gt_bbox, gt_mask, gt_class_id,
dataset_train.class_names, figsize=(8, 8))
results = model.detect([original_image], verbose=1)
r = results[0]
visualize.display_instances(original_image, r['rois'], r['masks'], r['class_ids'],
dataset_val.class_names, r['scores'], ax=get_ax())
```
## Evaluation
```
# Compute VOC-Style mAP @ IoU=0.5
# Running on 10 images. Increase for better accuracy.
image_ids = np.random.choice(dataset_val.image_ids, 10)
APs = []
for image_id in image_ids:
# Load image and ground truth data
image, image_meta, gt_class_id, gt_bbox, gt_mask =\
modellib.load_image_gt(dataset_val, inference_config,
image_id, use_mini_mask=False)
molded_images = np.expand_dims(modellib.mold_image(image, inference_config), 0)
# Run object detection
results = model.detect([image], verbose=0)
r = results[0]
# Compute AP
AP, precisions, recalls, overlaps =\
utils.compute_ap(gt_bbox, gt_class_id,
r["rois"], r["class_ids"], r["scores"])
APs.append(AP)
print("mAP: ", np.mean(APs))
```
| github_jupyter |
___
<a href='http://www.pieriandata.com'> <img src='../Pierian_Data_Logo.png' /></a>
___
# Matplotlib Exercises
Welcome to the exercises for reviewing matplotlib! Take your time with these, Matplotlib can be tricky to understand at first. These are relatively simple plots, but they can be hard if this is your first time with matplotlib, feel free to reference the solutions as you go along.
Also don't worry if you find the matplotlib syntax frustrating, we actually won't be using it that often throughout the course, we will switch to using seaborn and pandas built-in visualization capabilities. But, those are built-off of matplotlib, which is why it is still important to get exposure to it!
**NOTE: ALL THE COMMANDS FOR PLOTTING A FIGURE SHOULD ALL GO IN THE SAME CELL. SEPARATING THEM OUT INTO MULTIPLE CELLS MAY CAUSE NOTHING TO SHOW UP.**
# Exercises
**We will focus on two commons tasks, plotting a known relationship from an equation and plotting raw data points.**
Follow the instructions to complete the tasks to recreate the plots using this data:
----
----
### Task One: Creating data from an equation
It is important to be able to directly translate a real equation into a plot. Your first task actually is pure numpy, then we will explore how to plot it out with Matplotlib. The [world famous equation](https://en.wikipedia.org/wiki/Mass%E2%80%93energy_equivalence) from Einstein:
$$E=mc^2$$
Use your knowledge of Numpy to create two arrays: E and m , where **m** is simply 11 evenly spaced values representing 0 grams to 10 grams. E should be the equivalent energy for the mass. You will need to figure out what to provide for **c** for the units m/s, a quick google search will easily give you the answer (we'll use the close approximation in our solutions).
**NOTE: If this confuses you, then hop over to the solutions video for a guided walkthrough.**
```
# CODE HERE
# CODE HERE
```
### Part Two: Plotting E=mc^2
Now that we have the arrays E and m, we can plot this to see the relationship between Energy and Mass.
**TASK: Import what you need from Matplotlib to plot out graphs:**
**TASK: Recreate the plot shown below which maps out E=mc^2 using the arrays we created in the previous task. Note the labels, titles, color, and axis limits. You don't need to match perfectly, but you should attempt to re-create each major component.**
```
# CODE HERE
# DON'T RUN THE CELL BELOW< THAT WILL ERASE THE PLOT!
```
### Part Three (BONUS)
**Can you figure out how to plot this on a logarthimic scale on the y axis? Place a grid along the y axis ticks as well. We didn't show this in the videos, but you should be able to figure this out by referencing Google, StackOverflow, Matplotlib Docs, or even our "Additional Matplotlib Commands" notebook. The plot we show here only required two more lines of code for the changes.**
```
# CODE HERE
# DONT RUN THE CELL BELOW! THAT WILL ERASE THE PLOT!
```
---
---
## Task Two: Creating plots from data points
In finance, the yield curve is a curve showing several yields to maturity or interest rates across different contract lengths (2 month, 2 year, 20 year, etc. ...) for a similar debt contract. The curve shows the relation between the (level of the) interest rate (or cost of borrowing) and the time to maturity, known as the "term", of the debt for a given borrower in a given currency.
The U.S. dollar interest rates paid on U.S. Treasury securities for various maturities are closely watched by many traders, and are commonly plotted on a graph such as the one on the right, which is informally called "the yield curve".
**For this exercise, we will give you the data for the yield curves at two separate points in time. Then we will ask you to create some plots from this data.**
## Part One: Yield Curve Data
**We've obtained some yeild curve data for you from the [US Treasury Dept.](https://www.treasury.gov/resource-center/data-chart-center/interest-rates/pages/textview.aspx?data=yield). The data shows the interest paid for a US Treasury bond for a certain contract length. The labels list shows the corresponding contract length per index position.**
**TASK: Run the cell below to create the lists for plotting.**
```
labels = ['1 Mo','3 Mo','6 Mo','1 Yr','2 Yr','3 Yr','5 Yr','7 Yr','10 Yr','20 Yr','30 Yr']
july16_2007 =[4.75,4.98,5.08,5.01,4.89,4.89,4.95,4.99,5.05,5.21,5.14]
july16_2020 = [0.12,0.11,0.13,0.14,0.16,0.17,0.28,0.46,0.62,1.09,1.31]
```
**TASK: Figure out how to plot both curves on the same Figure. Add a legend to show which curve corresponds to a certain year.**
```
# CODE HERE
# DONT RUN THE CELL BELOW! IT WILL ERASE THE PLOT!
```
**TASK: The legend in the plot above looks a little strange in the middle of the curves. While it is not blocking anything, it would be nicer if it were *outside* the plot. Figure out how to move the legend outside the main Figure plot.**
```
# CODE HERE
# DONT RUN THE CELL BELOW! IT WILL ERASE THE PLOT!
```
**TASK: While the plot above clearly shows how rates fell from 2007 to 2020, putting these on the same plot makes it difficult to discern the rate differences within the same year. Use .suplots() to create the plot figure below, which shows each year's yield curve.**
```
# CODE HERE
# DONT RUN THE CELL BELOW! IT WILL ERASE THE PLOT!
```
**BONUS CHALLENGE TASK: Try to recreate the plot below that uses twin axes. While this plot may actually be more confusing than helpful, its a good exercise in Matplotlib control.**
```
# CODE HERE
# DONT RUN THE CELL BELOW! IT ERASES THE PLOT!
```
-----
| github_jupyter |
# CME 193: Introduction to Scientific Python
## Spring 2018
## Lecture 1
# Lecture 1 Contents
* Course Outline
* Introduction
* Python Basics
* Installing Python
---
# Quick Poll
## Who has written one line of code?
## ...a for loop?
## ...a function?
## Who has heard of recursion?
## ...object oriented programming?
## ...unit testing?
## ... generators ?
## Who has programmed in Python before?
## Who has programmed in R or Stata?
## Anyone written tensorflow?
# Course Outline
## Instructor
## Jacob Perricone
##### jacobp2@stanford.edu
* 2nd year MS Student in ICME
* Raised in New York City
* BSE in Operations Research and Financial Engineering with Certificates in Computer Science, Statistics and Machine Learning from Princeton University
* Python is almost always my language of choice (I use matlab for their solvers, C if i really care about speed).
## Remmelt Ammerlaan
##### remmelt@stanford.edu
* First yeat MS student in the data science track at ICME.
* Born in the Netherlands did my Bachelor’s at McGill in Math and Physics.
* My first introduction to coding was in Python and I still use it today for most of my work.
# Course Content
- Variables Functions Data types
- Strings, Lists, Tuples, Dictionaries
- File input and output (I/O)
- Classes, object oriented programming
- Exception handling, Recursion
- Numpy, Scipy, Pandas and Scikit-learn
- Jupyter, Matplotlib and Seaborn
- Unit tests, multithreading
- List subject to change depending on time, interests
# Course setup
- 8 total sessions
- 45 min lecture
- 5 min break
- 30 min interactive - demos and exercises
- First part of class will be traditional lecture, second part will give you time to work on exercises in class
- **Required deliverables:**
- Exercises
- Two homework assignments
# More abstract setup of course
- My job is to show and explain to you the possibilities and resources of python
- Ultimately, though, it is your job to teach yourself Python
- In order for it to stick, you will need to put in considerable effort
# Exercises
We will work on exercises second half of the class. **Try to finish as much as possible during class time**. They will help you understand topics we just talked about. If you do not finish in class, please try to look at and understand them prior to the next class meeting.
<br>
Feel free (or: you are strongly encouraged) to work in pairs on the exercises. It’s acceptable to hand in the same copy of code for your exercise solutions if you work in pairs, but you must mention your partner.
# Exercises
#### (continued)
- At the end of the course, you will be required to hand in your solutions for the exercises you attempted.
- This is to show your active participation in class.
- You are expected to do at least 70% of the assigned exercises. Feel free to skip some problems, for example if you lack some required math background knowledge.
- Don’t worry about this now, just save all the code you write.
# Feedback
- If you have comments or would like things to be done differently, please let me know as you think of them. Can tell me in person, via email or Canvas.
- Questionnaires at the end of the quarter are nice, but they won’t help you.
# Workload
- The only way to learn Python, is by writing a lot of Python.
- Good news: Python is fun to write. Put in the effort these 4 weeks and reap the rewards.
- From past experience: If you are new to programming, consider this a hard 3 unit class where you will have to figure out quite a bit on your own. However, if you have a solid background in another language, this class should be pretty easy.
# To new programmers
- If you have never programmed before, be warned that this will be difficult.
- The problem: 4 weeks, 8 lectures, 1 unit. We will simply go too fast.
- Alternative: spend some time learning on your own (Codecademy / Udacity etc). There are so many excellent resources online these days. We offer this class every quarter.
# Important course information
- **Website**: https://web.stanford.edu/~jacobp2/src/html/cme193.html
- **Canvas**: Use Canvas for discussing problems, turning in homework, etc. Also, I will view participation of Canvas discussion as participation on the course.
- **Office hours**: Directly after class, or by appointment in Huang basement
# References
- The internet is an excellent source, and Google is a perfect starting point.
- Stackoverflow is the most popular online community of coding QA.
- The official documentation is also good: https://docs.python.org/3/.
- Course website and Canvas has a list of useful references - I will also try to update them with specific material with each lecture.
# Last words before we get to it
- Do the work - utilize the class time
- Make friends
- Fail often
- Fail gently
- Be resourceful
# Python Versions
- The most commonly used versions of Python are 2.7 and 3.4 (or 3.6)
- The first version of Python 3 was released 10 years ago in 2008
- The final version of Python 2.X (2.7) came out mid-2010.
- The 2.x branch will see no new major releases
- 3.x is under active development and has already seen over five years of stable releases, with 3.6 in 2016
- All improvements to the standard library are only available in Python 3
# So what's the deal here ?
## Which one to use?
- The release of Python 3.0 caused a ruckus in the developer world since Python 3 is not backward compatable.
- For a long time many large libraries did not support Python 3, so people continued on in Python 2.7
# ... That was 9 years ago
- Many companies and I'm sure some courses at Stanford still use 2.7 (I still use it too), but I guarentee that all of them, including me, want to move to Python 3.
- Python 3 is the future, and so we will be using it (release 3.6)
# Introduction

```
print("Hello, world!")
```
# Python Basics
## Values
- A value is the fundamental thing that a program manipulates.
- Values can be ```“Hello, world!”, 42, 12.34, True```
- Values have types. . .
## Types
1. Numeric Types:
- Integer Types: ``` 92, 12, 0, 1 ```
- Floats (Floating point numbers): ``` 3.1415```
- Complex Numbers: ``` a + b*i ``` (composed of real and imaginary component, both of which are floats ``` a + b*i ```)
- Booleans: ```True/False ``` are a subtype of integers (0 is false, 1 is true)
2. Sequence types:
- Lists : ``` [1,2,3,4,5]```
- Tuples: ``` (1, 2) ```
- range objects: More on this later
3. Strings:
- ``` "Hello World"``` (strings in python are actually immutable sequences but more on this later)
4. There are more...
## Types continued
- Use type to find out the type of a variable, as in
- ``` python
type("Hello, Word")
```
which returns ``` <class 'str'>```
<br>
- Unlike C/C++ and Java, variables can change types. Python keeps track of the type internally (strongly-typed).
## Variables
- One of the most basic and powerful concepts is that of a variable.
- A variable assigns a name to a value.
- Variables are nothing more than reserved memory locations that store values.
- Python variables does not need explicit declaration to reserve memory.
```
message = "Hello, world!"
n = 42
e = 2.71
# note we can print variables:
print(n) # yields 42
# note: everything after pound sign is a comment
```
## Variables
- Almost always preferred to use variables over values:
- Easier to update code
- Easier to understand code (useful naming)
- What does the following code do:
``` python
print(4.2 * 3.5)
```
```
length = 4.2
height = 3.5
area = length * height
print(area)
```
## Keywords
- Not allowed to use keywords for naming, they define structure and rules of a language.
- Python has 29 keywords, they include:
- ```True```
- ```False```
- ```continue```
- ```def```
- ```for```
- ```and ```
- ```return ```
- ```is```
- ```in ```
- ```class```
## Integers
- Basic Operators for integers:
- ```a + b```: addition of a and b
- ```a - b```: subtraction of a and b
- ```a * b```: multiplication of a and b
- ``` a / b```: division of and b
- ``` a % b ```: Modulus (divides a by b and returns the remainder)
- ```a**b```: a to the power of b
- ``` a // b``` : Floor division. Divides a by b and truncates the decimals. If ``` a/b ``` is negative, rounds away from zero toward negative infinity
- Note: ```a / b``` differs in versions of python:
- Python 3 : ```5 / 2``` yields ```2.5```
- Python 2 : ```5 / 2``` yields ```2```
- In Python 2 if one of the operands is a float, the return value is a float: ```5 / 2.0``` yields ```2.5```
- Note: Python automatically uses long integers for very large integers.
- Bitwise operators on integer types:
- ``` x | y ``` bitwise or of x and y
- ``` x ^ y ``` exclusive or of x and y
- ``` x & y ``` bitwise and of x and y
- ``` x << n ``` x shifted to left by n bits
- ``` x >> n ``` x shifted to right by n bits
## Floats
- A floating point number approximates a real number. Note: only finite precision, and finite range (overflow)!
- Basic Operators for floats:
- ```a + b```: addition of a and b
- ```a - b```: subtraction of a and b
- ```a * b```: multiplication of a and b
- ``` a / b```: division of and b
- ``` a % b ```: Modulus (divides a by b and returns the remainder)
- ```a**b```: a to the power of b
- ``` a // b``` : Floor division. Divides a by b and truncates the decimals. If ``` a/b ``` is negative, rounds away
```
import sys
print((sys.float_info))
```
## Comparison Operators
#### Comparisons:
1. Equals: ``` == ```
- ``` 5 == 5 ``` yields ```True```
2. Does not equal: ```!=```
- ``` 5 != 5``` yields ```False```
3. Comparison of object identity: ``` is ```
- ``` x is y ``` yields True if x and y are the same object
4. Negated objected identity: ``` is not ```
- ``` x is not y ``` yields False if x and y are the same object
5. Greater than: ``` > ```
- ```5 > 4 ``` yields ```True```
6. Greater than or equal to: ``` >= ``` (greater than or equal to)
- ```5 >= 5``` yields ```True```
7. Similarly, we have ```<``` and ```<=```.
#### Logical Operators:
There are three logical boolean operations that are used to compare values. They evaluate expressions down to Boolean values, returning either ``` True ``` or ``` False ```
- ``` x or y ```: if x is false, then y, else x (only evaluates y if x is false)
- ``` x and y ```: if x is false, then x, else y (only evaluates second if x is true)
- ``` not x ```: if x is false, then True, else False
Typically used to evaluate whether two or more expressions are true: ``` (5 < 7) and (7 <= 10)```, which is equivalent to ``` 5 < 7 <= 10 ```
## Statements, expressions and operators
- A statement is an instruction that Python can execute, such as
``` x=3 ```
- Python also supports statements such as
* ```x += 3```
- x = x + 3
* ```x -= 3```
- x = x - 3
* ```x *= 3```
- x = x*3
* ```x /= 3```
- x = x/3
* ``` x //= 3```
- x = x//3
- **Note**: the statements above combine an operation and assignment, and thereby only work if the variable x is defined (i.e. ``` x += 3 ``` will fail unless it is preceeded by ``` x = some value ```)
- Operators are special symbols that represent computations, like addition, the values they *operate* on are called operands
- An expression is a combination of values, variable and operators, like ``` x+3 ```
## Modules
- Not all functionality available comes automatically when starting Python, and with good reasons.
- We can add extra functionality by importing modules:
```import math```
- Then we can use things like ```math.pi```
- Useful modules: math, os, random, and as we will see later numpy, scipy and matplotlib.
- More on modules later!
```
import math
math.pi
```
## Control-flow
## Control statements
Control statements allow you to do more complicated tasks:
- ```if```
- ```for```
- ```while```
```
traffic_light = 'green'
```
Using if, we can execute part of a program conditional on some statement being true.
```python
if traffic_light == 'green':
move()
```
## Indentation
In Python, blocks of code are defined using indentation. The indentation within the block needs to be consistent.
This means that everything indented after an if statement is only executed if the statement is True.
If the statement is False, the program skips all indented code and resumes at the first line of unindented code
```
statement = True # Changing this to False will change the control flow!
if statement:
# if statement is True, then all code here
# gets executed but not if statement is False
print("The statement is true")
print("Else, this would not be printed")
# the next lines get executed either way
print("Hello, world,")
print("Bye, world!")
```
## Indentation
- Whitespace is meaningful in Python: especially indentation and placement of newlines.
- Use a newline to end a line of code.
- Use a backslash when must go to next line prematurely.
- No braces to mark blocks of code in Python...
- Use consistent indentation instead.
- The first line with less indentation is outside of the block.
- The first line with more indentation starts a nested block
- Often a colon appears at the start of a new block. (E.g. for function and class definitions.)
- The preferred method of indendation is spaces (not tabs). Python 3 now support inconsistent use of spaces and tabs, but stick to one.
## ```if-else``` statement
We can add more conditions to the ```if``` statement using ```else``` and ```elif``` (short for else if).
Consider the following example
```
# Skipping the cell below as to not introduce functions yet.
def drive():
print("Drive")
def accelerate():
print("Accelerate")
def stop():
print("Stop!")
x = 5
print(x)
traffic_light = 'green'
if traffic_light == 'green':
drive()
elif traffic_light == 'orange':
accelerate()
else:
stop()
traffic_light = 'orange'
if traffic_light == 'green':
drive()
elif traffic_light == 'orange':
accelerate()
else:
stop()
traffic_light = 'blue'
if traffic_light == 'green':
drive()
elif traffic_light == 'orange':
accelerate()
else:
stop()
```
## ```for``` loops
- Very often, one wants to repeat some action. This can be achieved by a for loop
```
for i in range(5):
print(i**2, end='')
print("hello world")
print("hello world", end = ' ')
print("I am on the same line")
```
## ```for``` loops
- ```range(n)``` yields an immutable ```sequence``` of integers $ 0, . . . , n − 1$. More on this later!
## ```while``` loops
- When we do not know how many iterations are needed, we can use while.
```
i = 1
while i < 100:
print(i, end=' ')
i += i**2 # a += b is short for a = a + b
```
## ```continue```
- The keyword ```continue``` continues with the next iteration of the smallest enclosing loop.
```
for num in range(2, 10):
if num % 2 == 0:
print("Found an even number", num)
continue
print("Found an odd number", num)
```
## ```break```
- The keyword ```break``` allows us to jump out of the smallest enclosing for or while loop.
```
max_n = 10
for n in range(2, max_n):
is_prime = True
for x in range(2, n):
if n % x == 0: # n divisible by x
is_prime = False
print(n, 'equals', x, '*', n/x)
break
# executed if no break in for loop
# loop fell through without finding a factor
if is_prime:
print(n, 'is a prime number')
```
## ```pass```
- The pass statement does nothing, which can come in handy when you are working on something and want to implement some part of your code later.
```
traffic_light = 'green'
if traffic_light == 'green':
pass
else:
stop()
```
# Installing Python
# Anaconda
* Mac OS and Windows
* Anaconda Python is a Python distribution maintained by Continuum Analytics
* The distribution contains the most widely used Python implementation and
up-to-date versions of the most important modules for data science, analytics,
and scientific computing.
* Anaconda is easy to download and install and is free to use in personal,
academic, and commercial environments.
* By default, Anaconda Python installs into a user local directory and usually
does not impact a system provided version of Python. In some cases, having
multiple version of Python installed causes problems.
* Check out this link to download python 3.6 through conda https://anaconda.org/anaconda/python
* I'd recommend setting up an environment for your python using https://conda.io/docs/user-guide/tasks/manage-python.html. Come see me if you need some help
# Jupyter Notebook
```python
# this is a code cell
print("hello from Jupyter notebook")
# hit ctrl-return to execute code block
# hit shift-return to execute code block and move to next cell
```
## This is a Markdown cell
* Markdown is a light weight way to annotate text for nice presentation on the
web or in PDF
* For example the `*` will create a bulletted list
* We can easily do *italics*
## Resources
* <http://jupyter.org/>
* <https://try.jupyter.org/>
```
x = 5
print(x)
```
## This is a markdown cell
# Using the Python interpreter
* An *interpreter* is a program that reads and executes commands
* It is also sometimes called a REPL or read-evaluate-print-loop
* One way to interact with Python is to use the interpreter
* This is useful for interactive work, learning, and simple testing
## Start the Python interpreter on Mac OS
* Open `Terminal.app`. This is located at
`/Applications/Utilities/Terminal.app` or may be found using Spotlight Search.
* This is the Bash prompt where commands are entered after `$`
* Type `python` and hit enter to start the interpreter (The default for Mac is python 2.7, if you see this come see me)
* This a great way to experiment and learn Python
* To exit the interpreter and return to bash:
* Enter `>>> exit()`
* Use the keyboard command `ctrl-d`
## Start the Python interpreter from Jupyter
It is possible to access a Python interpreter from inside of the Jupyter
notebook. This can be a very quick and handy way to experiment with small bits
of Python code.
* From the Jupyter home screen, select the "Terminal" from the "New" dropdown
menu.
# Scripting model
* A Python script is a text file containing Python code
* Python script file names typically end in `.py`
## Let's create our first script
1. Create a text file named `firstscript.py` with your favorite text editor
2. Insert the following Python code into `firstscript.py`:
```python
print("Hello from Python.")
print("I am your first script!")
```
3. Open your favorite terminal emulator (`Terminal.app` on Mac OS)
4. Navigate to the directory containing `firstscript.py` with the `cd` command.
5. Execute the command `$ python firstscript.py`
## Why scripts?
Let's write a simple Python script to compute the first `n` numbers in the
Fibonacci series. As a reminder, each number in the Fibonacci series is the sum
of the two previous numbers. Let `F(i)` be the `i`th number in the series. We
define `F(0) = 0` and `F(1) = 1`, then `F(i) = F(i-1) + F(i-2)` for `i >= 2`.
Numbers `F(0)` to `F(n)` can be computed with the following Python code:
```python
n = 10
if n >= 0:
fn2 = 0
print(fn2,end=',')
if n >= 1:
fn1 = 1
print(fn1,end=',')
for i in range(2,n+1):
fn = fn1 + fn2
print(fn,end=',')
fn2 = fn1
fn1 = fn
print()
```
**Note, the above code is a preview of Python syntax that we will review in this
course.**
## Fibonacci (continued)
Now, paste this code into a file named `fib.py`. Execute the file with
the command `$ python fib.py`. The result should like:
```
$ python fib.py
0,1,1,2,3,5,8,13,21,34,55,
```
To see the utility of scripts, we need to add a bit more code. Change the first
line of `fib.py` to be:
```
import sys
n = int(sys.argv[1])
```
This will instruct the script to obtain the value of `n` from the command line:
```
$ python fib.py 0
0,
$ python fib.py 5
0,1,1,2,3,5,
$ python fib.py 21
0,1,1,2,3,5,8,13,21,34,55,89,144,233,377,610,987,1597,2584,4181,6765,10946,
```
We have increased the utility of our program by making it simple to run from the
command line with different input arguments.
# Fibonacci (continue):
1. When one types ``` python my_script.py ``` all code at indentation level 0 gets run
2. Unlike other languages, there's no ``` main()``` function that gets run automatically
2. However, the interpreter will define a few special variables. If the script is being run directly, the interpreter will set the variable ``` __name__ = '__main__' ```. Now, the proper way to write the fib.py script includes material that will be presented in subsequent slides.
```python
import sys
def fib(n):
if n >= 0:
fn2 = 0
print(fn2,end=',')
if n >= 1:
fn1 = 1
print(fn1,end=',')
for i in range(2,n+1):
fn = fn1 + fn2
print(fn,end=',')
fn2 = fn1
fn1 = fn
if __name__ == '__main__':
n = int(sys.argv[1])
fib(n) ```
| github_jupyter |
```
! nvidia-smi
```
# Introduction
This notebook holds the code for the [Involution](https://arxiv.org/abs/2103.06255) layer in tesorflow. The idea behind this layer is to invert the inherent properties of Convolution. Where convolution is spatial-agnostic and channel-specific, involution is spatial-specific and channel-agnostic.
# Imports
```
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
```
# Convolution
To understand involution we need to first understand convolution. Let us consider $X\in\mathbb{R}^{H\times W\times C_{inp}}$ denote the input feature map where $H, W$ represent its height and width and $C_{inp}$ be its channel size. A collection of $C_{out}$ number of convolution filters with fixed kernel size of $K \times K$ is denoted as $\mathcal{F}\in\mathbb{R}^{C_{out}\times C_{inp}\times K\times K}$.
The filters perform a Multiply-Add operation on the input feature map in a sliding window manner to yeild the output feature map $Y\in \mathbb{R}^{H\times W\times C_{out}}$.
# Involution
Involution kernels $\mathcal{H}\in \mathbb{R}^{H\times W\times K\times K\times G}$ are devised to oprate in a symettrically oppposite manner as that of the convolution kernels. Observing the shape of the involution kernels we observe the following things:
- Each pixel of the input feature map is entitled to get its own involution kernel.
- Each kernel is of the shape of $K\times K\times G$.
- The output $Y$ will be of the same shape as that of the input feature map $X$.
The problem with involution is that we cannot define a fixed shaped kernel, that would hurt resolution independence in the neural network. This thought led the researchers to conceptualize a generation function $\phi$ that generates the involution kernels conditioned on the original input tensor.
$$
\mathcal{H}_{ij}=\phi{(X_{ij})}\\
\mathcal{H}_{ij}=W_{1}\sigma{(W_{0}X_{ij})}\\
$$
```
class Involution(tf.keras.layers.Layer):
def __init__(self, channel, group_number, kernel_size, stride, reduction_ratio):
super().__init__()
# The assert makes sure that the user knows about the
# reduction size. We cannot have 0 filters in Conv2D.
assert reduction_ratio <= channel, print("Reduction ration must be less than or equal to channel size")
self.channel = channel
self.group_number = group_number
self.kernel_size = kernel_size
self.stride = stride
self.reduction_ratio = reduction_ratio
self.o_weights = tf.keras.layers.AveragePooling2D(
pool_size=self.stride,
strides=self.stride,
padding="same") if self.stride > 1 else tf.identity
self.kernel_gen = tf.keras.Sequential([
tf.keras.layers.Conv2D(
filters=self.channel//self.reduction_ratio,
kernel_size=1),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.ReLU(),
tf.keras.layers.Conv2D(
filters=self.kernel_size*self.kernel_size*self.group_number,
kernel_size=1)
])
def call(self, x):
_, H, W, C = x.shape
H = H//self.stride
W = W//self.stride
# Extract input feature blocks
unfolded_x = tf.image.extract_patches(
images=x,
sizes=[1,self.kernel_size,self.kernel_size,1],
strides=[1,self.stride,self.stride,1],
rates=[1,1,1,1],
padding="SAME") # B, H, W, K*K*C
unfolded_x = tf.keras.layers.Reshape(
target_shape=(H,
W,
self.kernel_size*self.kernel_size,
C//self.group_number,
self.group_number)
)(unfolded_x) # B, H, W, K*K, C//G, G
# generate the kernel
kernel_inp = self.o_weights(x)
kernel = self.kernel_gen(kernel_inp) # B, H, W, K*K*G
kernel = tf.keras.layers.Reshape(
target_shape=(H,
W,
self.kernel_size*self.kernel_size,
1,
self.group_number)
)(kernel) # B, H, W, K*K, 1, G
# Multiply-Add op
out = tf.math.multiply(kernel, unfolded_x) # B, H, W, K*K, C//G, G
out = tf.math.reduce_sum(out, axis=3) # B, H, W, C//G, G
out = tf.keras.layers.Reshape(
target_shape=(H,
W,
C)
)(out) # B, H, W, C
return out
```
# Comparison
In this section we will try and emulate [TensorFlow's tutorial on CIFAR classification](https://www.tensorflow.org/tutorials/images/cnn). Here we build one model with convolutional layers while the other will be based on involuitonal layers.
```
(train_images, train_labels), (test_images, test_labels) = tf.keras.datasets.cifar10.load_data()
# Normalize pixel values to be between 0 and 1
train_images, test_images = train_images / 255.0, test_images / 255.0
train_ds = tf.data.Dataset.from_tensor_slices((train_images, train_labels)).shuffle(256).batch(256)
test_ds = tf.data.Dataset.from_tensor_slices( (test_images, test_labels)).batch(256)
class_names = ['airplane', 'automobile', 'bird', 'cat', 'deer',
'dog', 'frog', 'horse', 'ship', 'truck']
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i])
# The CIFAR labels happen to be arrays,
# which is why you need the extra index
plt.xlabel(class_names[train_labels[i][0]])
plt.show()
```
## Convolutional Neural Network
```
convolution_model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(32, (3, 3), activation='relu', input_shape=(32, 32, 3), padding="same"),
tf.keras.layers.MaxPooling2D((2, 2)),
tf.keras.layers.Conv2D(64, (3, 3), activation='relu', padding="same"),
tf.keras.layers.MaxPooling2D((2, 2)),
tf.keras.layers.Conv2D(64, (3, 3), activation='relu', padding="same"),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(10),
])
convolution_model.summary()
convolution_model.compile(
optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy']
)
conv_history = convolution_model.fit(
train_ds,
epochs=10,
validation_data=test_ds
)
```
### Loss plot
```
plt.plot(conv_history.history["loss"], label="loss")
plt.plot(conv_history.history["val_loss"], label="val_loss")
plt.legend()
plt.show()
```
### Accuracy plot
```
plt.plot(conv_history.history["accuracy"], label="acc")
plt.plot(conv_history.history["val_accuracy"], label="val_acc")
plt.legend()
plt.show()
```
# Involutional Neural Network
```
involution_model = tf.keras.models.Sequential([
Involution(channel=3,group_number=1,kernel_size=3,stride=1,reduction_ratio=2),
tf.keras.layers.ReLU(name="relu1"),
tf.keras.layers.MaxPooling2D((2, 2)),
Involution(channel=3,group_number=1,kernel_size=3,stride=1,reduction_ratio=2),
tf.keras.layers.ReLU(name="relu2"),
tf.keras.layers.MaxPooling2D((2, 2)),
Involution(channel=3,group_number=1,kernel_size=3,stride=1,reduction_ratio=2),
tf.keras.layers.ReLU(name="relu3"),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(10),
])
involution_model.compile(
optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy']
)
inv_history = involution_model.fit(
train_ds,
epochs=10,
validation_data=test_ds
)
involution_model.summary()
```
### Loss Plot
```
plt.plot(inv_history.history["loss"], label="loss")
plt.plot(inv_history.history["val_loss"], label="val_loss")
plt.legend()
plt.show()
```
### Accuracy Plot
```
plt.plot(inv_history.history["accuracy"], label="acc")
plt.plot(inv_history.history["val_accuracy"], label="val_acc")
plt.legend()
plt.show()
```
### Observation
A fun little experiment is to see the activation maps of the involution kernel.
```
layer_names = ["relu1","relu2","relu3"]
outputs = [involution_model.get_layer(name).output for name in layer_names]
vis_model = tf.keras.Model(involution_model.input, outputs)
fig, axes = plt.subplots(nrows=10, ncols=4, figsize=(10, 20))
[ax.axis("off") for ax in np.ravel(axes)]
for (ax_orig, ax_relu1, ax_relu2, ax_relu3), test_image in zip(axes, test_images[:10]):
relu_images_list = vis_model.predict(tf.expand_dims(test_image,0))
ax_orig.imshow(tf.clip_by_value(test_image, 0, 1))
ax_orig.set_title("Input Image")
ax_relu1.imshow(tf.clip_by_value(relu_images_list[0].squeeze(), 0, 1))
ax_relu1.set_title("ReLU 1")
ax_relu2.imshow(tf.clip_by_value(relu_images_list[1].squeeze(), 0, 1))
ax_relu2.set_title("ReLU 2")
ax_relu3.imshow(tf.clip_by_value(relu_images_list[2].squeeze(), 0, 1))
ax_relu3.set_title("ReLU 3")
```
| github_jupyter |
# Making Predictions with Time Series Data
## Author:
### Aaron Washington Chen [GitHub](https://github.com/AaronWChen)
## Executive Summary
This project sought the 5 best zipcodes for short-term (<5 year turnaround) real estate investments based off of Zillow sales data.
The primary suggestion at minimum viable product (MVP) delivery in September 2019 was to [avoid](https://www.investopedia.com/investing/next-housing-recession-2020-predicts-zillow/) investing in real estate due to market conditions (over valued) and market predictions ([likely recession](http://zillow.mediaroom.com/2018-05-22-Experts-Predict-Next-Recession-Will-Begin-in-2020)).
If the client insisted on investing in real estate, the 5 zipcodes/areas recommended with estimated conservative returns at the end of 2021 were:
1. Benson, NC 27504 ($~50k on ~$150k purchase)
2. Centerville, TN 37033 ($~20k on ~$85k purchase)
3. Austin, TX 78758 (~$53k on ~$300k purchase)
4. Cedar Creek, TX 78612 (~$72k on ~$212k purchase)
5. Charlotte, NC 28208 (~$32k on ~$125k purchase)
[Summary Presentation](https://drive.google.com/open?id=1NprmQa0j-SuBJF2Hp1D3n5trHvHh7ka2OdE29vGjBuE)
## Project Information
As a consultant for a real estate investment firm, I was asked to answer the following question in 5 days:
> What are the top 5 best zipcodes for us to invest in?
The original data came from [Zillow Research](https://www.zillow.com/research/data/) but has been included in this repo in the data folder as ```zillow_data.csv```
Because of the large number of records to potentially analyze (~15,000 zipcodes), it was logistically impossible to do a complete and accurate prediction for each zipcode in the time window.
Instead, I looked at the work and comments of experts who have a lot more data than I do. Using several [other](https://www.zillow.com/research/local-market-reports/) [Zillow](https://www.zillow.com/research/home-searches-potential-moves-13192/) [articles](http://zillow.mediaroom.com/2018-01-09-San-Jose-and-Raleigh-are-Zillows-Hottest-Housing-Markets-for-2018), I narrowed down the search to areas that have been generating a lot of search and interest requests on Zillow's own servers. This reduced the zipcodes of interest down to a few hundred.
To perform the predictions, I used Facebook's Prophet library inside Jupyter Notebooks to display data tables and generate visualizations faster and more intuitively than with scripts.
A short turnaround time for investment was chosen because the predictions became too unreliable much further into the future.
## Improvements and Next Steps
With a few metropolitan areas and several hundred zipcodes predicted, some next steps could involve refactoring the code to use individualized Auto Regressive Integrated Moving Average (ARIMA) models to perform the predictions. However, each zipcode would require a custom-tuned ARIMA model and this amount of time investment may not be worth it, depending on client wishes.
## Running the Code
If you are looking to run and/or work on this project yourself, you will need to:
1. Install Python 3 (I prefer and recommend [Anaconda](https://www.anaconda.com/distribution/))
2. [Install PyStan](https://pystan.readthedocs.io/en/latest/installation_beginner.html)
3. [Install Facebook Prophet](https://facebook.github.io/prophet/docs/installation.html)
4. Clone [this repo](https://github.com/AaronWChen/time_series_zillow)
5. Install the packages in ```requirements.txt``` via pip (```pip install -r requirements.txt``` from the command line)
If you want to see the high level summary, the presentation slides and presenter notes are all that are necessary. However, feel free to peruse the Jupyter Notebook files generated for each metropolitan area inside the python directory.
If you are looking to make changes to the code, I recommend using Visual Studio Code to open the files and edit.
| github_jupyter |
LAB 2 - Milica miskovic
First, let us import all packages
```
import pandas as pd
import numpy as np
import matplotlib as plt
from sklearn.decomposition import PCA
```
Then we read the data file into data frame
```
ratings = pd.read_csv('ratings.dat', sep="::", header=None, engine='python')
ratings.columns = ['UserID', 'MovieID', 'Rating', 'Timestamp']
ratings.drop(['Timestamp'], axis=1, inplace=True)
print(ratings[:5])
```
1. Create m x u matrix with movies as row and users as column
```
ratings_pivot = ratings.pivot_table('Rating', index='MovieID', columns='UserID')
print(ratings_pivot.iloc[0:10, 0:10])
```
2. Normalize the matrix (we have to fill NaNs with 0 first, and call it matrix a)
```
a = ratings_pivot.fillna(value=0, inplace=True)
a = StandardScaler().fit_transform(ratings_pivot)
print(a.shape)
np.set_printoptions(precision=2)
print(a[0:10, 0:10])
print(type(ratings_pivot))
ratings_pivot.index.values
```
3. Compute SVD to get U, S and V
```
U, Sigma, V = np.linalg.svd(a)
```
4. From your V.T select 50 components.
```
V_T = V[:50]
```
Shape of U ,S and V after reduction:
```
print(U.shape)
print(Sigma.shape)
print(V.shape)
print(V_T.shape)
```
New, reduced matrix a we can recover by multiplying U ,S and V after reduction:
```
a_reduced = V_T.dot(a.T)
a_reduced = a_reduced.T #we have to transpose in order to get movies as rows and components as columns
print(a_reduced.shape)
```
In order to have movie names, we have to import movies.dat file.
```
movies = pd.read_csv('movies.dat', sep="::", header=None, engine='python')
movies.columns = ['MovieID', 'Title', 'Genres']
movies.drop(['Genres'], axis=1, inplace=True)
print(movies[:5])
movies.loc[1, 'Title'] #the second row is the movie with id 2, but not necessarily
```
The function:
```
def recommend(movie_id, matrix):
# first we have to check if input is valid:
if isinstance(movie_id, int) and isinstance(movie_id, np.int) and movie_id > 0 and movie_id <= matrix.shape[0]:
# initialize cos vector:
cos = np.zeros(matrix.shape[0])
# find which row of a matrix is our movie:
movie_loc = ratings_pivot.index.get_loc(movie_id)
# calculate cos sim between our movie and every other:
for i in range(0, matrix.shape[0]):
a = np.dot(matrix[movie_loc], matrix[i])
b = np.linalg.norm(matrix[movie_loc]) * np.linalg.norm(matrix[i])
cos[i] = abs(a/b)
# sort in descending order:
order = np.argsort(-cos)
# find ten best fits (the first one is going to be our movie):
ten = order[1:11]
print('Recommended movies are:')
print(ten)
ten_most_similar = movies.loc[movies['MovieID'].isin(ten)]
else:
print('Invalid movie ID.')
return(ten_most_similar)
```
Let us try with movie ID 2, 'Jumanji (1995)':
```
recommend(2, a_reduced)
```
6. Repeat the same process except now instead of using SVD you will use PCA to get the eigenvectors.
7. You will require co-variance matrix as an input to your eig function.
Matrix a is already standardized and NaNs are filled with 0.
Eigenvectors and eigenvalues:
```
cov_mat = np.cov(a.T)
print(cov_mat.shape)
eig_vals, eig_vecs = np.linalg.eig(cov_mat)
eig_pairs = [(np.abs(eig_vals[i]), eig_vecs[:,i]) for i in range(len(eig_vals))]
eig_pairs_sorted = sorted(eig_pairs, key=lambda x: x[0])
eig_vec_reduced = np.stack([vec for val, vec in eig_pairs[:50]], axis=1)
print(eig_vec_reduced.shape)
```
8. Use that same steps after that to get 50 components and use cosine similarity to get the results.
```
a_reduced_PCA = np.matmul(a, eig_vec_reduced)
print(a.shape)
print(eig_vec_reduced.shape)
print(a_reduced_PCA.shape)
recommend(2, a_reduced_PCA)
```
9. Compare the results for SVD and PCA.
```
print('10 recommended movies for the movie with id 2, Jumanji (1995) using SVD:')
print(recommend(2, a_reduced))
print('10 recommended movies for the movie with id 2, Jumanji (1995) using PCA:')
print(recommend(2, a_reduced_PCA))
```
As we can see, both lists are the same.
| github_jupyter |
# Section 4.3 : CYCLICAL MOMENTUM
## Summary
- Learning rate and momentum are closely dependent, and both must be optimised
- Momentum should be set as high as possible without causing instabilities in training
- Momentum cannot be optimised in a similar way to LR, by using a momentum finder
- Optimum settings found to be use of cyclical LR (initially increasing) and cyclical momentum (initially decreasing)
- If constant LR is used, a large (but not too large), constant momentum should be used
- Too large a constant momentum results in instabilities, which are visible in early training
## Momentum in SGD
SGD parameter updates:
$\theta_{iter+1} = \theta_{iter}− \epsilon\delta L(F(x,\theta),\theta),$
where $\theta$ are the parameters, $\epsilon$ is the learning rate, and $L(F(x,\theta),\theta)$ is the gradient.
Momentum modifies the update rule to:
$\nu_{iter+1} = \alpha\nu_{iter}− \epsilon\delta L(F(x,\theta),\theta)$
$\theta_{iter+1} = \theta_{iter}+\nu_{iter},$
where $\nu$ is velocity, and $\alpha$ is the momentum coefficient, i.e. the effect of $\alpha$ on the update is of the same scale as $\epsilon$.
## Cyclical momentum Example
Let's take the same model and train a few different configurations fo learning rate and momentum:
```
%matplotlib inline
from __future__ import division
import sys
import os
sys.path.append('../')
from Modules.Basics import *
from Modules.Class_Basics import *
data, features = importData()
nFolds = 5
preprocParams = {'normIn':True, 'pca':False}
compileArgs = {'loss':'binary_crossentropy', 'optimizer':'sgd', 'depth':3, 'width':128, 'lr':5e2}
trainParams = {'epochs':20, 'batch_size':256, 'verbose':0}
```
### Constant LR, Constant Momentum
```
from pathlib import Path
import os
results_ConstLR_ConstMom85, history_ConstLR_ConstMom85 = cvTrainClassifier(data, features, nFolds, preprocParams,
{'version':'modelRelu', 'nIn':len(features),
'compileArgs':{**compileArgs, 'momentum':0.85}},
trainParams, useEarlyStop=False, plot=False)
results_ConstLR_ConstMom90, history_ConstLR_ConstMom90 = cvTrainClassifier(data, features, nFolds, preprocParams,
{'version':'modelRelu', 'nIn':len(features),
'compileArgs':{**compileArgs, 'momentum':0.90}},
trainParams, useEarlyStop=False, plot=False)
results_ConstLR_ConstMom95, history_ConstLR_ConstMom95 = cvTrainClassifier(data, features, nFolds, preprocParams,
{'version':'modelRelu', 'nIn':len(features),
'compileArgs':{**compileArgs, 'momentum':0.95}},
trainParams, useEarlyStop=False, plot=False)
results_ConstLR_ConstMom99, history_ConstLR_ConstMom99 = cvTrainClassifier(data, features, nFolds, preprocParams,
{'version':'modelRelu', 'nIn':len(features),
'compileArgs':{**compileArgs, 'momentum':0.99}},
trainParams, useEarlyStop=False, plot=False)
getModelHistoryComparisonPlot([history_ConstLR_ConstMom85, history_ConstLR_ConstMom90, history_ConstLR_ConstMom95, history_ConstLR_ConstMom99],
['LR=500, Mom=0.85', 'LR=500, Mom=0.90', 'LR=500, Mom=0.95', 'LR=500, Mom=0.99'], cv=True)
```
Similar to the paper, we see that using a constant learning rate requires high values of momentum to converge quickly: as the coefficient is increased, the networks reach their minima in fewer and fewer epochs. At very high momenta (<span style="color:red">red</span>), the network eventually overfits and starts diverging. However it shows slight instability in it's early stages of training, which (as the paper suggests) could be used to catch the eventual overfitting early, and adjust the coefficient.
### Constant LR, Cyclical Momentum
```
stepScale = 4
results_ConstLR_CycMom95_85, history_ConstLR_CycMom95_85 = cvTrainClassifier(data, features, nFolds, preprocParams,
{'version':'modelRelu', 'nIn':len(features),
'compileArgs':compileArgs},
trainParams, useEarlyStop=False, plot=False,
useCallbacks={'LinearCMom':{'maxMom':0.95,'minMom':0.85,
'scale':stepScale, 'plotMom':False}})
results_ConstLR_CycMom99_90, history_ConstLR_CycMom99_90 = cvTrainClassifier(data, features, nFolds, preprocParams,
{'version':'modelRelu', 'nIn':len(features),
'compileArgs':compileArgs},
trainParams, useEarlyStop=False, plot=False,
useCallbacks={'LinearCMom':{'maxMom':0.99,'minMom':0.90,
'scale':stepScale, 'plotMom':False}})
results_ConstLR_CycMom99_95, history_ConstLR_CycMom99_95 = cvTrainClassifier(data, features, nFolds, preprocParams,
{'version':'modelRelu', 'nIn':len(features),
'compileArgs':compileArgs},
trainParams, useEarlyStop=False, plot=False,
useCallbacks={'LinearCMom':{'maxMom':0.99,'minMom':0.95,
'scale':stepScale, 'plotMom':False}})
getModelHistoryComparisonPlot([history_ConstLR_CycMom95_85, history_ConstLR_CycMom99_90, history_ConstLR_CycMom99_95, history_ConstLR_ConstMom99],
['LR=500, Cyclical mom [0.95-0.85]', 'LR=500, Cyclical mom [0.99-0.90]', 'LR=500, Cyclical mom [0.99-0.95]', 'LR=500, Mom=0.99'], cv=True)
```
Here we can see that using a cyclical momentum schedule can be quite unstable (loss fluctuates, possibly an artifact of stepisize), but does provide some resistance to overfitting (late test loss is slow to rise).
Comparing to a constant momentum of 0.99 (<span style="color:red">red</span>) to a cyclical momentum between 0.99 and 0.95 <span style="color:green">green</span>, we can see that the cycling supresses the rise in test loss in late training, and achieves better minima in loss. Initial training is also better, however the artifacts of the scheduling cause mild divergence around epochs 7 and 15, preventing the network from convereging earlier than might otherwise be possible.
As the width of the cycle is increased (<span style="color:green">green</span> to <span style="color:orange">orange</span>), these artifacts become more apparent as the mild diveregnces become sharp spikes.
### Cyclical LR, Constant Momentum
```
stepScale = 4
results_CycLR_ConstMom85, history_CycLR_ConstMom85 = cvTrainClassifier(data, features, nFolds, preprocParams,
{'version':'modelRelu', 'nIn':len(features),
'compileArgs':{**compileArgs, 'momentum':0.85}},
trainParams, useEarlyStop=False, plot=False,
useCallbacks={'LinearCLR':{'maxLR':5e2,'minLR':5e1,
'scale':stepScale, 'plotLR':False}})
results_CycLR_ConstMom90, history_CycLR_ConstMom90 = cvTrainClassifier(data, features, nFolds, preprocParams,
{'version':'modelRelu', 'nIn':len(features),
'compileArgs':{**compileArgs, 'momentum':0.90}},
trainParams, useEarlyStop=False, plot=False,
useCallbacks={'LinearCLR':{'maxLR':5e2,'minLR':5e1,
'scale':stepScale, 'plotLR':False}})
results_CycLR_ConstMom95, history_CycLR_ConstMom95 = cvTrainClassifier(data, features, nFolds, preprocParams,
{'version':'modelRelu', 'nIn':len(features),
'compileArgs':{**compileArgs, 'momentum':0.95}},
trainParams, useEarlyStop=False, plot=False,
useCallbacks={'LinearCLR':{'maxLR':5e2,'minLR':5e1,
'scale':stepScale, 'plotLR':False}})
results_CycLR_ConstMom99, history_CycLR_ConstMom99 = cvTrainClassifier(data, features, nFolds, preprocParams,
{'version':'modelRelu', 'nIn':len(features),
'compileArgs':{**compileArgs, 'momentum':0.99}},
trainParams, useEarlyStop=False, plot=False,
useCallbacks={'LinearCLR':{'maxLR':5e2,'minLR':5e1,
'scale':stepScale, 'plotLR':False}})
getModelHistoryComparisonPlot([history_CycLR_ConstMom85, history_CycLR_ConstMom90, history_CycLR_ConstMom95, history_CycLR_ConstMom99, history_ConstLR_ConstMom99],
['Cyclical LR [50-500], mom=0.85', 'Cyclical LR [50-500], mom=0.90', 'Cyclical LR [50-500], mom=0.95', 'Cyclical LR [50-500], mom=0.99', 'LR=500, Mom=0.99'], cv=True)
```
Here we see that moving to a cyclical LR schedule might help reduce the instability of using very high momenta. Comparing <span style="color:red">red</span> to <span style="color:purple">purple</span>, we find that the cyclical LR gives a slightly smoother loss evolution, reaches a better loss, and supresses the late-stage overfitting.
### Cyclical LR, cyclical Momentum
```
stepScale = 4
results_CycLR_CycMom95_85, history_CycLR_CycMom95_85 = cvTrainClassifier(data, features, nFolds, preprocParams,
{'version':'modelRelu', 'nIn':len(features),
'compileArgs':compileArgs},
trainParams, useEarlyStop=False, plot=False,
useCallbacks={'LinearCLR':{'maxLR':5e2,'minLR':5e1,
'scale':stepScale, 'plotLR':False},
'LinearCMom':{'maxMom':0.95,'minMom':0.85,
'scale':stepScale, 'plotMom':False}})
results_CycLR_CycMom99_90, history_CycLR_CycMom99_90 = cvTrainClassifier(data, features, nFolds, preprocParams,
{'version':'modelRelu', 'nIn':len(features),
'compileArgs':compileArgs},
trainParams, useEarlyStop=False, plot=False,
useCallbacks={'LinearCLR':{'maxLR':5e2,'minLR':5e1,
'scale':stepScale, 'plotLR':False},
'LinearCMom':{'maxMom':0.99,'minMom':0.90,
'scale':stepScale, 'plotMom':False}})
results_CycLR_CycMom99_95, history_CycLR_CycMom99_95 = cvTrainClassifier(data, features, nFolds, preprocParams,
{'version':'modelRelu', 'nIn':len(features),
'compileArgs':compileArgs},
trainParams, useEarlyStop=False, plot=False,
useCallbacks={'LinearCLR':{'maxLR':5e2,'minLR':5e1,
'scale':stepScale, 'plotLR':False},
'LinearCMom':{'maxMom':0.99,'minMom':0.95,
'scale':stepScale, 'plotMom':False}})
getModelHistoryComparisonPlot([history_CycLR_CycMom95_85, history_CycLR_CycMom99_90, history_CycLR_CycMom99_95, history_ConstLR_CycMom99_95],
['Cyclical LR [50-500], Cyclical Mom [0.95-0.85]', 'Cyclical LR [50-500], Cyclical Mom [0.99-0.90]', 'Cyclical LR [50-500], Cyclical Mom [0.99-0.95]', 'LR=500, Cyclical Mom [0.99-0.95]'], cv=True)
```
Comparing the best CLR+CM setup (<span style="color:green">green</span>) to the fixed LR+CM setup (<span style="color:red">red</span>) it seems that cycling the lR degrades the performance of the network (best loss is higher), however the network never overfits; unlink the <span style="color:red">red</span> line, it reaches it's minima after 7 epochs and then plateus. It's possible that the stability might actually be a consequence of underfitting, in which case the learning rate could perhaps be increased.
### Comparison
```
getModelHistoryComparisonPlot([history_ConstLR_ConstMom99, history_ConstLR_CycMom99_95, history_CycLR_ConstMom99, history_CycLR_CycMom99_95],
['LR=500, Mom=0.99', 'LR=500, Cyclical Mom [0.99-0.95]', 'Cyclical LR [50-500], Mom=0.99', 'Cyclical LR [50-500], Cyclical LR [0.99-0.95]'], cv=True)
```
Comparing the best performing setups from each sechudle configuration it seems that of the hyperparameters tested, for this dataset and architecture, a cycled LR with a constant momentum (<span style="color:green">green</span>) provides the lowest loss, but eventually overfits.
Cycling the momentum and keeping the LR constant (<span style="color:orange">orange</span>) reaches almost as good a loss, but after 40% more epochs, and although it later provides less overfitting, it does suffer from regular peaks and troughs due to the cycling.
Cycling both the LR and the momentum (<span style="color:red">red</span>) causes convergence in the same number of epochs as <span style="color:green">green</span>, but at a higher loss. Having reached its minimum, the test loss then remains flat, possibly indicating that with further adjustments of the hyperparameters it might provide superior performance to <span style="color:green">green</span>.
| github_jupyter |
```
import torch
from torch.autograd import Variable
import torch.nn as nn
import captcha_setting
import operator
import torchvision.transforms as transforms
from PIL import Image
import cv2 as cv
import os
from matplotlib import pyplot as plt
import numpy as np
import copy
#for eachimg in filter_containor:
class CNN(nn.Module):
def __init__(self):
super(CNN, self).__init__()
self.layer1 = nn.Sequential(
nn.Conv2d(1, 32, kernel_size=3, padding=1),
nn.BatchNorm2d(32),
nn.Dropout(0.1), # drop 50% of the neuron
nn.ReLU(),
nn.MaxPool2d(2))
self.layer2 = nn.Sequential(
nn.Conv2d(32, 64, kernel_size=3, padding=1),
nn.BatchNorm2d(64),
nn.Dropout(0.1), # drop 50% of the neuron
nn.ReLU(),
nn.MaxPool2d(2))
self.layer3 = nn.Sequential(
nn.Conv2d(64, 64, kernel_size=3, padding=1),
nn.BatchNorm2d(64),
nn.Dropout(0.1), # drop 50% of the neuron
nn.ReLU(),
nn.MaxPool2d(2))
self.fc = nn.Sequential(
nn.Linear((captcha_setting.IMAGE_WIDTH//8)*(captcha_setting.IMAGE_HEIGHT//8)*64, 1024),
nn.Dropout(0.1), # drop 50% of the neuron
nn.ReLU())
self.rfc = nn.Sequential(
nn.Linear(1024, 256),#captcha_setting.MAX_CAPTCHA*captcha_setting.ALL_CHAR_SET_LEN),
nn.ReLU()
)
self.rfc2 = nn.Sequential(
nn.Linear(256, captcha_setting.MAX_CAPTCHA*captcha_setting.ALL_CHAR_SET_LEN),
)
def forward(self, x):
out = self.layer1(x)
out = self.layer2(out)
out = self.layer3(out)
out = out.view(out.size(0), -1)
out = self.fc(out)
#print(out.shape)
out = self.rfc(out)
out = self.rfc2(out)
#out = out.view(out.size(0), -1)
#print(out.shape)
return out
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# cnn = CNN()
cnn = CNN()
cnn.eval()
cnn.load_state_dict(torch.load('model_final_mix_2.pkl'))
cnn.to(device)
transform = transforms.Compose([
# transforms.ColorJitter(),
transforms.Grayscale(),
transforms.ToTensor(),
# transforms.Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225])
])
root_path = '/home/ning_a/Desktop/CAPTCHA/dark_web_captcha/mania_data/'
img_list = os.listdir(root_path)
correct = 0
total = 0
for img_cur_path in img_list:
total += 1
label_predicted = ''
img = cv.imread(root_path+img_cur_path)
img_temp = cv.imread(root_path+img_cur_path)
n_img = np.zeros((img.shape[0],img.shape[1]))
img_aft = cv.normalize(img, n_img, 0,255,cv.NORM_MINMAX)
gray = cv.cvtColor(img_aft,cv.COLOR_BGR2GRAY)
ret, thresh = cv.threshold(gray,0,255,cv.THRESH_BINARY_INV+cv.THRESH_OTSU)
im2,contours,hierarchy = cv.findContours(thresh,cv.RETR_EXTERNAL,cv.CHAIN_APPROX_SIMPLE)
filter_containor = []
temp_img = copy.deepcopy(img)
max_x = 160
max_y = 60
max_x2 = 0
max_y2 = 0
for i in range(0,len(contours)):
x, y, w, h = cv.boundingRect(contours[i])
newimage=img[y:y+h,x:x+w] # 先用y确定高,再用x确定宽
nrootdir=("cut_image/")
if h<5 or w<5:
continue
filter_containor.append([x, y, w, h])
cv.rectangle(temp_img, (x,y), (x+w,y+h), (153,153,0), 1)
if not os.path.isdir(nrootdir):
os.makedirs(nrootdir)
cv.imwrite( nrootdir+str(i)+".jpg",newimage)
if(x<max_x):
max_x = x
if(y<max_y):
max_y = y
if(x+w>max_x2):
max_x2 = x+w
if(y+h>max_y2):
max_y2 = y+h
cv.imwrite( "temp3.png",img_temp[max_y:max_y2, max_x:max_x2])
seg_img = cv.imread("temp3.png")
filter_containor = []
w = (max_x2-max_x)//6
# print(max_x, max_x2)
# print(max_y, max_y2)
# print(img_temp[9:45,27:130].shape)
for i in range(6):
# print(seg_img[:,i*w:w+i*w].shape)
cv.imwrite( "temp3.png",seg_img[:,i*w:w+i*w])
filter_containor.append(Image.open("temp3.png"))
# print(img_temp.shape)
#>>>>>>>>>>>>>>>>>>>>>>>>>>>>>
for eachimg in filter_containor:
#print(eachimg)
fix_size = (30, 60)
eachimg = eachimg.resize(fix_size)
image = transform(eachimg).unsqueeze(0)
plt.imshow(eachimg)
plt.show()
print(image.shape)
image = torch.tensor(image, device=device).float()
image = Variable(image).to(device)
#print(image.shape)
#image, labels = image.to(device), labels.to(device)
# vimage = generator(image)
predict_label = cnn(image)
#labels = labels.cpu()
predict_label = predict_label.cpu()
_, predicted = torch.max(predict_label, 1)
label_predicted+= captcha_setting.ALL_CHAR_SET[predicted]
# print(captcha_setting.ALL_CHAR_SET[predicted])
# print(captcha_setting.ALL_CHAR_SET[predicted])
parsed_label = ''
if('_' in img_cur_path):
parsed_label = img_cur_path.split('_')[0]
else:
parsed_label = img_cur_path.split('.')[0]
if(parsed_label == label_predicted):
correct +=1
print(label_predicted, parsed_label)
# break
print(correct/total)
```
| github_jupyter |
# DeepLab Demo
This demo will demostrate the steps to run deeplab semantic segmentation model on sample input images.
```
#@title Imports
import os
from io import BytesIO
import tarfile
import tempfile
from six.moves import urllib
from matplotlib import gridspec
from matplotlib import pyplot as plt
import numpy as np
from PIL import Image
import tensorflow as tf
import sys
sys.path.append("/home/jhpark/dohai90/workspaces/models/research")
#@title Helper methods
class DeepLabModel(object):
"""Class to load deeplab model and run inference."""
INPUT_TENSOR_NAME = 'ImageTensor:0'
OUTPUT_TENSOR_NAME = 'SemanticPredictions:0'
INPUT_SIZE = 513
FROZEN_GRAPH_NAME = 'frozen_inference_graph'
def __init__(self, tarball_path):
"""Creates and loads pretrained deeplab model."""
self.graph = tf.Graph()
graph_def = None
# Extract frozen graph from tar archive.
tar_file = tarfile.open(tarball_path)
for tar_info in tar_file.getmembers():
if self.FROZEN_GRAPH_NAME in os.path.basename(tar_info.name):
file_handle = tar_file.extractfile(tar_info)
graph_def = tf.GraphDef.FromString(file_handle.read())
break
tar_file.close()
if graph_def is None:
raise RuntimeError('Cannot find inference graph in tar archive.')
with self.graph.as_default():
tf.import_graph_def(graph_def, name='')
self.sess = tf.Session(graph=self.graph)
def run(self, image):
"""Runs inference on a single image.
Args:
image: A PIL.Image object, raw input image.
Returns:
resized_image: RGB image resized from original input image.
seg_map: Segmentation map of `resized_image`.
"""
width, height = image.size
resize_ratio = 1.0 * self.INPUT_SIZE / max(width, height)
target_size = (int(resize_ratio * width), int(resize_ratio * height))
resized_image = image.convert('RGB').resize(target_size, Image.ANTIALIAS)
batch_seg_map = self.sess.run(
self.OUTPUT_TENSOR_NAME,
feed_dict={self.INPUT_TENSOR_NAME: [np.asarray(resized_image)]})
seg_map = batch_seg_map[0]
return resized_image, seg_map
def create_pascal_label_colormap():
"""Creates a label colormap used in PASCAL VOC segmentation benchmark.
Returns:
A Colormap for visualizing segmentation results.
"""
colormap = np.zeros((256, 3), dtype=int)
ind = np.arange(256, dtype=int)
for shift in reversed(range(8)):
for channel in range(3):
colormap[:, channel] |= ((ind >> channel) & 1) << shift
ind >>= 3
return colormap
def label_to_color_image(label):
"""Adds color defined by the dataset colormap to the label.
Args:
label: A 2D array with integer type, storing the segmentation label.
Returns:
result: A 2D array with floating type. The element of the array
is the color indexed by the corresponding element in the input label
to the PASCAL color map.
Raises:
ValueError: If label is not of rank 2 or its value is larger than color
map maximum entry.
"""
if label.ndim != 2:
raise ValueError('Expect 2-D input label')
colormap = create_pascal_label_colormap()
if np.max(label) >= len(colormap):
raise ValueError('label value too large.')
return colormap[label]
def vis_segmentation(image, seg_map):
"""Visualizes input image, segmentation map and overlay view."""
plt.figure(figsize=(15, 5))
grid_spec = gridspec.GridSpec(1, 4, width_ratios=[6, 6, 6, 1])
plt.subplot(grid_spec[0])
plt.imshow(image)
plt.axis('off')
plt.title('input image')
plt.subplot(grid_spec[1])
seg_image = label_to_color_image(seg_map).astype(np.uint8)
plt.imshow(seg_image)
plt.axis('off')
plt.title('segmentation map')
plt.subplot(grid_spec[2])
plt.imshow(image)
plt.imshow(seg_image, alpha=0.7)
plt.axis('off')
plt.title('segmentation overlay')
unique_labels = np.unique(seg_map)
ax = plt.subplot(grid_spec[3])
plt.imshow(
FULL_COLOR_MAP[unique_labels].astype(np.uint8), interpolation='nearest')
ax.yaxis.tick_right()
plt.yticks(range(len(unique_labels)), LABEL_NAMES[unique_labels])
plt.xticks([], [])
ax.tick_params(width=0.0)
plt.grid('off')
plt.show()
LABEL_NAMES = np.asarray([
'background', 'aeroplane', 'bicycle', 'bird', 'boat', 'bottle', 'bus',
'car', 'cat', 'chair', 'cow', 'diningtable', 'dog', 'horse', 'motorbike',
'person', 'pottedplant', 'sheep', 'sofa', 'train', 'tv'
])
FULL_LABEL_MAP = np.arange(len(LABEL_NAMES)).reshape(len(LABEL_NAMES), 1)
FULL_COLOR_MAP = label_to_color_image(FULL_LABEL_MAP)
#@title Select and download models {display-mode: "form"}
MODEL_NAME = 'xception_coco_voctrainaug' # @param ['mobilenetv2_coco_voctrainaug', 'mobilenetv2_coco_voctrainval', 'xception_coco_voctrainaug', 'xception_coco_voctrainval']
_DOWNLOAD_URL_PREFIX = 'http://download.tensorflow.org/models/'
_MODEL_URLS = {
'mobilenetv2_coco_voctrainaug':
'deeplabv3_mnv2_pascal_train_aug_2018_01_29.tar.gz',
'mobilenetv2_coco_voctrainval':
'deeplabv3_mnv2_pascal_trainval_2018_01_29.tar.gz',
'xception_coco_voctrainaug':
'deeplabv3_pascal_train_aug_2018_01_04.tar.gz',
'xception_coco_voctrainval':
'deeplabv3_pascal_trainval_2018_01_04.tar.gz',
}
_TARBALL_NAME = 'deeplab_model.tar.gz'
model_dir = tempfile.mkdtemp()
tf.gfile.MakeDirs(model_dir)
download_path = os.path.join(model_dir, _TARBALL_NAME)
print('downloading model, this might take a while...')
urllib.request.urlretrieve(_DOWNLOAD_URL_PREFIX + _MODEL_URLS[MODEL_NAME],
download_path)
print('download completed! loading DeepLab model...')
MODEL = DeepLabModel(download_path)
print('model loaded successfully!')
download_path
```
## Run on sample images
Select one of sample images (leave `IMAGE_URL` empty) or feed any internet image
url for inference.
Note that we are using single scale inference in the demo for fast computation,
so the results may slightly differ from the visualizations in
[README](https://github.com/tensorflow/models/blob/master/research/deeplab/README.md),
which uses multi-scale and left-right flipped inputs.
```
#@title Run on sample images {display-mode: "form"}
SAMPLE_IMAGE = 'image1' # @param ['image1', 'image2', 'image3']
IMAGE_URL = '' #@param {type:"string"}
_SAMPLE_URL = ('https://github.com/tensorflow/models/blob/master/research/'
'deeplab/g3doc/img/%s.jpg?raw=true')
def run_visualization(url):
"""Inferences DeepLab model and visualizes result."""
try:
f = urllib.request.urlopen(url)
jpeg_str = f.read()
original_im = Image.open(BytesIO(jpeg_str))
except IOError:
print('Cannot retrieve image. Please check url: ' + url)
return
print('running deeplab on image %s...' % url)
resized_im, seg_map = MODEL.run(original_im)
vis_segmentation(resized_im, seg_map)
image_url = IMAGE_URL or _SAMPLE_URL % SAMPLE_IMAGE
run_visualization(image_url)
```
| github_jupyter |
# YATClient Twitter Project
## Background Info
* The 2019 European parliamentary elections took place on 24 – 26 May.
* As a result of the election, there were eight political parties in Germany that held two or more seats in parliament.
* Our project analyzes the twitter data in the three weeks leading up to the election to look for insights related to the outcome.
|Party|2009|2014|2019|$ \Delta $| % |
|---|---|---|---|---|---|
|CDU/CSU | 42 | 34| 29| <span style="color:red">-5 |<span style="color:red"> -14.71 </span>|
|SPD | 23 | 27| 16| <span style="color:red">-11 | <span style="color:red">-40.74 </span>|
|die Grünen | 14 | 11| 21| <span style="color:green">-10 |<span style="color:green"> 90.91 </span>|
|die Linke | 8 | 7| 5| <span style="color:red">-2 | <span style="color:red">-28.57 </span>|
|AfD | 0 | 7| 11| <span style="color:green"> 4 | <span style="color:green">57.14 </span>|
|FDP | 12 | 3| 5| <span style="color:green"> 2 | <span style="color:green">66.67 </span>|
|die Partei | 0 | 1| 2| <span style="color:green"> 1 | <span style="color:green">100.00 </span>|
|FW | 0 | 1| 2| <span style="color:green"> 1 | <span style="color:green">100.00 </span>|
## Obtain Data and Install
```
#This will install the package in the actual jupyter notebook Kernel. [with_Jupyter] is obsolete if you start it out of jupyter, but for demo purpose left inside.
import sys
!{sys.executable} --version
!{sys.executable} -m pip install -e "."[with_jupyter] #"." describes the path to the package, in this case its the same folder.
```
## Example imports:
import yatclient as yat
from yatclient import TweetAnalyzer,TwitterClient
from yatclient import * #uses __all__ list in __init__.py filename must be declared in this case eg.: twitter_client.TwitterClient
### Enter your twitter Creds here:
```
#Access
CONSUMER_KEY = ""
CONSUMER_SECRET = ""
ACCESS_TOKEN = "-"
ACCESS_TOKEN_SECRET = ""
```
### This will download the twitter data of major parties three weeks before European Parliament Election.
- download data for all parties
```
%%time
import yatclient as yat
import datetime
Parties = ["spdde", "fdp","die_Gruenen","afd","dieLinke","fwlandtag","diepartei","cdu","csu"]
EuropawahlDate = datetime.date(2019, 5, 24)
ThreeWeeksBeforeDate = EuropawahlDate - datetime.timedelta(weeks=3)
today = datetime.date.today()
twitter_client = yat.TwitterClient(CONSUMER_KEY, CONSUMER_SECRET, ACCESS_TOKEN, ACCESS_TOKEN_SECRET, Parties)
tweets = twitter_client.get_user_timeline_tweets(start_date = str(ThreeWeeksBeforeDate), end_date = str(EuropawahlDate), retweets = True)
analyzer_load = yat.TweetAnalyzer(tweets)
analyzer_load.write_to_csv("tweets{}.csv".format(today), encoding = "utf-8",index = False)
print("saved data to tweets{}.csv\n".format(today))
df = analyzer_load.get_dataframe()
df.head(2)
print("Timeframe from:\n",df.groupby(by=["author"]).date.min(),"\n\nto date:\n",df.groupby(by=["author"]).date.max(),"\n")
#Wenn Daten bereits vorhanden:
import yatclient as yat
#
analyzer = yat.TweetAnalyzer()
analyzer.read_from_csv("tweets2019-07-05.csv")
dfr = analyzer.get_dataframe()
dfr = dfr.drop(columns=['Unnamed: 0'])
dfr= analyzer_load.get_dataframe()
df_SPDEuropa = dfr.loc[dfr["author"] == "SPDEuropa"].copy()
df_fdp = dfr.loc[dfr["author"] == "fdp"].copy()
df_Die_Gruenen = dfr.loc[dfr["author"] == "Die_Gruenen"].copy()
df_AfD = dfr.loc[dfr["author"] == "AfD"].copy()
df_dieLinke = dfr.loc[dfr["author"] == "dieLinke"].copy()
df_fwlandtag = dfr.loc[dfr["author"] == "fwlandtag"].copy()
df_DiePARTEI = dfr.loc[dfr["author"] == "DiePARTEI"].copy()
df_CDU = dfr.loc[dfr["author"] == "CDU"].copy()
df_CSU = dfr.loc[dfr["author"] == "CSU"].copy()
analyzer_SPDEuropa = yat.TweetAnalyzer(df = df_SPDEuropa)
analyzer_fdp = yat.TweetAnalyzer(df = df_fdp)
analyzer_Die_Gruenen = yat.TweetAnalyzer(df = df_Die_Gruenen)
analyzer_AfD = yat.TweetAnalyzer(df = df_AfD)
analyzer_dieLinke = yat.TweetAnalyzer(df = df_dieLinke)
analyzer_fwlandtag = yat.TweetAnalyzer(df = df_fwlandtag)
analyzer_DiePARTEI = yat.TweetAnalyzer(df = df_DiePARTEI)
analyzer_CDU = yat.TweetAnalyzer(df = df_CDU)
analyzer_CSU = yat.TweetAnalyzer(df = df_CSU)
dfr.columns
# Pie
import matplotlib.pyplot as plt
# Pie chart, where the slices will be ordered and plotted counter-clockwise:
labels = 'AfD', 'Die PARTEI', 'Die Gruenen'
sizes = [22, 7, 5]
explode = (0, 0, 0.1)
fig1, ax1 = plt.subplots()
ax1.pie(sizes, explode=explode, labels=labels, autopct='%1.1f%%',
shadow=True, startangle=90)
ax1.axis('equal') # Equal aspect ratio ensures that pie is drawn as a circle.
plt.show()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
plt.style.use('seaborn')
%matplotlib inline
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
from IPython.core.display import SVG
import math
```
# Evaluate the definite integral
$\int_{a}^{b}f(x)dx = F(b) - F(a)$
$\int_{-3}^{5}4dx$
${(4x)}' = 4$
$\int_{-3}^{5}4dx = 4x|_{3}^{5} = 4(5) - 4(-3) = 20 + 12 = 32$
$\int_{-1}^{3} 7x^2dx$
${(\frac{7x^3}{3})}' = 7x^2$
$\int_{-1}^{3}4dx = \frac{7x^3}{3}|_{-1}^{3} = \frac{7(3)^3}{3} - \frac{7(-1)^3}{3}$
$\int_{1}^{6} 2dx$
${(2x)}' = 2$
$\int_{1}^{6}4dx = 2x|_{1}^{6} = 12 - 2 = 10$
_('cause F has no slope - rectangular area, so u can just `end - begin * C`)_
$\int_{-1}^{1} (9x^2 + 6x - 3) dx$
$\left[ \frac{9x^3}{3} + \frac{6x^2}{2} - 3x \right]_{-1}^{1}$
$\left[ 3x^3 + 3x^2 - 3x \right]_{-1}^{1}$
$3(1)^3 + 3(1)^2 - 3(1) - (3(-1)^3 + 3(-1)^2 - 3(-1)) = 0$
# $\int\limits_{0}^{1} -12e^x dx = \left[ -12e^x \right]^{0}_{1} = -12e^1 - (-12e^0) = -12e + 12$
# $\int\limits_{\pi}^{2\pi} -cos(x) dx = \left[ -sin(x) \right]_{\pi}^{2\pi} = -sin(2\pi) + sin(\pi) = 0$
$\int\limits_{5}^{9} \left( \frac{x - 4}{x^2} \right) dx$
$\int\limits_{5}^{9} \left(\frac{1}{x} - 4x^{-2} \right) dx$
$\left[ ln(x) + 4x^{-1} \right]^{5}_{9}$
---
$(ln(9) + 4(9)^{-1}) - (ln(5) + 4(5)^{-1})$
$(ln(9) + \frac{4}{9}) - (ln(5) + \frac{4}{5})$
$ln(9) + \frac{4}{9} - ln(5) - \frac{4}{5}$
$ln(9) + \frac{20}{45} - ln(5) - \frac{36}{45}$
$ln(9) - \frac{16}{45} - ln(5)$
$ln\left(\frac{9}{5}\right) - \frac{16}{45}$
$\int\limits_{9}^{1} \left( -15\sqrt{x} \right) dx$
$\int\limits_{9}^{1} \left( -15\sqrt{x} \right) dx$
$\int\limits_{9}^{1} \left( -15x^{1\over2} \right) dx$
$\left[ -15\frac{2}{3}x^{3\over2} \right]^{1}_{9}$
$\left[ -10x^{3\over2} \right]^{1}_{9}$
$-10(1)^{3\over2} + 10(9)^{3\over2}$
$-10 + 10(9^3)^{1\over2}$
$-10 + 10(9\cdot9\cdot9)^{1\over2}$
$-10 + 10(3\cdot3\cdot3) = -10 + 270 = 260$
$f(x) = \left\{\begin{matrix}
x^2 - 3x & x \gt 2 \\
2x - 6 & x \le 2
\end{matrix}\right.$
$\int\limits_{0}^{3} f(x) dx = \int\limits_{0}^{2} (2x - 6)dx + \int\limits_{2}^{3} (x^2 - 3x)dx$
$\int\limits_{0}^{2} (2x - 6)dx = \left[ x^2 - 6x \right]_{0}^{2} = ((2)^2 - 6(2)) - (0 - 0) = -8$
$\int\limits_{2}^{3} (x^2 - 3x)dx =
\left[ \frac{1}{3}x^3 - \frac{3}{2}x^2 \right]_{2}^{3} =
(\frac{1}{3}(3)^3 - \frac{3}{2}(3)^2) - (\frac{1}{3}(2)^3 - \frac{3}{2}(2)^2)
= -\frac{7}{6}
$
$\int\limits_{0}^{3} f(x) dx = -8 - \frac{7}{6} = -\frac{48}{6} - \frac{7}{6} = -\frac{55}{6}$
$f(x) = \left\{\begin{matrix}
-3\sqrt{x} & x \gt 1 \\
x - 3 & x \le 1
\end{matrix}\right.$
$\int\limits_{-4}^{4} f(x) dx = \int\limits_{-4}^{1} (x-3) dx + \int\limits_{1}^{4} (-3\sqrt{x}) dx
= -\frac{45}{2} - 14 = -\frac{73}{2}$
$\int\limits_{-4}^{1} (x-3) dx =
\left[ \frac{1}{2}x^2 - 3x \right]_{-4}^{1} =
\left[ \frac{1}{2}(1)^2 - 3(1) \right] - \left[ \frac{1}{2}(-4)^2 - 3(-4) \right] =
\frac{-45}{2}
$
$\int\limits_{1}^{4} (-3\sqrt{x}) dx =
-3\int\limits_{1}^{4} x^{1\over2}dx =
\left[ -3\frac{2}{3}x^{\frac{3}{2}} \right]_{1}^{4} =
\left[ -2x^{\frac{3}{2}} \right]_{1}^{4} =
\left[ -2\sqrt{x^3} \right]_{1}^{4} =
\left[ -2\sqrt{(4)^3} \right] - \left[ -2\sqrt{(1)^3} \right] =
(-2 \cdot 8) + 2 = -14
$
$f(x) = \left\{\begin{matrix}
-6x + 6 & x \lt 1 \\
sin(\pi \cdot x) & x \ge 1
\end{matrix}\right.$
$\int\limits_{0}^{3} f(x) dx = \int\limits_{0}^{1} (-6x + 6) dx + \int\limits_{1}^{3} sin(x\pi) dx = 3 + 0 = 3$
$\int\limits_{0}^{1} (-6x + 6) dx =
\left[ -3x^2 + 6x \right]_{0}^{1} =
\left[ -3(1)^2 + 6(1) \right] - \left[ -3(0)^2 + 6(0) \right] = 3
$
$\int\limits_{1}^{3} sin(x\pi) dx =
\left[ -\frac{1}{\pi}cos(x\pi) \right]_{1}^{3} =
\left[ -\frac{1}{\pi}cos(3\pi) \right] - \left[ -\frac{1}{\pi}cos(\pi) \right] =
-\frac{1}{\pi}cos(3\pi) + \frac{1}{\pi}cos(\pi) =
\frac{1}{\pi} - \frac{1}{\pi} = 0
$
$\int\limits_{0}^{4} \left| 2x - 6 \right| dx
$
$f(x) = \left\{\begin{matrix}
-(2x - 6) & x \lt 3 \\
2x - 6 & x \ge 3
\end{matrix}\right.$
$\int\limits_{0}^{4} f(x) dx = \int\limits_{0}^{3} (-2x + 6) dx + \int\limits_{3}^{4} (2x - 6) dx = 9 + 1 = 10$
$\int\limits_{0}^{3} (-2x + 6) dx =
\left[ -x^2 + 6x \right]_{0}^{3} =
\left[ -(3)^2 + 6(3) \right] - \left[ -(0)^2 + 6(0) \right] = 9
$
$\int\limits_{3}^{4} (2x - 6) dx =
\left[ x^2 - 6x \right]_{3}^{4} =
\left[ (4)^2 - 6(4) \right] - \left[ (3)^2 - 6(3) \right] =
16 - 24 - 9 + 18 = 1
$
| github_jupyter |
```
__author__ = 'Tilii: https://kaggle.com/tilii7'
import warnings
with warnings.catch_warnings():
warnings.filterwarnings('ignore')
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
import matplotlib.cm as cm
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
```
Simply loading the files without any transformation. If you wish to manipulate the data in any way, it should be done here before doing dimensionality reduction in subsequent steps.
```
print('\nLoading files ...')
train = pd.read_csv('../input/train.csv')
test = pd.read_csv('../input/test.csv')
X = train.drop(['id', 'target'], axis=1).values
y = train['target'].values.astype(np.int8)
target_names = np.unique(y)
print('\nThere are %d unique target valuess in this dataset:' % (len(target_names)), target_names)
```
Principal Component Analysis (**[PCA](https://en.wikipedia.org/wiki/Principal_component_analysis)**) identifies the combination of components (directions in the feature space) that account for the most variance in the data.
```
n_comp = 20
# PCA
print('\nRunning PCA ...')
pca = PCA(n_components=n_comp, svd_solver='full', random_state=1001)
X_pca = pca.fit_transform(X)
print('Explained variance: %.4f' % pca.explained_variance_ratio_.sum())
print('Individual variance contributions:')
for j in range(n_comp):
print(pca.explained_variance_ratio_[j])
```
Better than 90% of the data is explained by a single principal component. Just a shade under 99% of variance is explained by 15 components, which means that this dataset can be safely reduced to ~15 features.
Here we plot our 0/1 samples on the first two principal components.
```
colors = ['blue', 'red']
plt.figure(1, figsize=(10, 10))
for color, i, target_name in zip(colors, [0, 1], target_names):
plt.scatter(X_pca[y == i, 0], X_pca[y == i, 1], color=color, s=1,
alpha=.8, label=target_name, marker='.')
plt.legend(loc='best', shadow=False, scatterpoints=3)
plt.title(
"Scatter plot of the training data projected on the 1st "
"and 2nd principal components")
plt.xlabel("Principal axis 1 - Explains %.1f %% of the variance" % (
pca.explained_variance_ratio_[0] * 100.0))
plt.ylabel("Principal axis 2 - Explains %.1f %% of the variance" % (
pca.explained_variance_ratio_[1] * 100.0))
plt.savefig('pca-porto-01.png', dpi=150)
plt.show()
```
There is a nice separation between various groups of customers, but not so between 0/1 categories within each group. This is somewhat exaggerated by the fact that "0" points (blue) are plotted first and "1" points (red) are plotted last. There seems to be more red than blue in that image, even though there are >25x "0" points in reality. I'd be grateful if someone knows how to plot this in a way that would not create this misleading impression.
Regardless, 0/1 points are not separated well at all. That means that they will not be easy to classify, which we all know by now.
**[t-SNE](https://lvdmaaten.github.io/tsne/)** could potentially lead to better data separation/visualization, because unlike PCA it preserves the local structure of data points. The problem with sklearn implementation of t-SNE is its lack of memory optimization. I am pretty sure that the t-SNE code at the very bottom will lead to memory errors on most personal computers, but I leave it commented out if anyone wants to try.
Instead, I ran t-SNE using a much faster and more memory-friendly commandline version, which can be found at the link above.
Here is the output of that exercise:

Again, we can see clear separation between different groups of customers. Some groups even have a nice "coffee bean" structure where two subgroups can be identified (gender?). Alas, there is no clear separation between 0/1 categories.
In strictly technical terms, we are screwed :D
```
# tsne = TSNE(n_components=2, init='pca', random_state=1001, perplexity=30, method='barnes_hut', n_iter=1000, verbose=1)
# X_tsne = tsne.fit_transform(X) # this will either fail or take a while (most likely overnight)
# plt.figure(2, figsize=(10, 10))
# for color, i, target_name in zip(colors, [0, 1], target_names):
# plt.scatter(X_tsne[y == i, 0], X_tsne[y == i, 1], color=color, s=1,
# alpha=.8, label=target_name, marker='.')
# plt.legend(loc='best', shadow=False, scatterpoints=3)
# plt.title('Scatter plot of t-SNE embedding')
# plt.xlabel('X')
# plt.ylabel('Y')
# plt.savefig('t-SNE-porto-01.png', dpi=150)
# plt.show()
```
It was kindly brought up to me that a strange-looking PCA plot above is probably because of categorical variables in this dataset. I leave the original plot up there for posterity.
Let's encode the categorical variables and try again.
```
from sklearn.preprocessing import MinMaxScaler
def scale_data(X, scaler=None):
if not scaler:
scaler = MinMaxScaler(feature_range=(-1, 1))
scaler.fit(X)
X = scaler.transform(X)
return X, scaler
X = train.drop(['id', 'target'], axis=1)
test.drop(['id'], axis=1, inplace=True)
n_train = X.shape[0]
train_test = pd.concat((X, test)).reset_index(drop=True)
col_to_drop = X.columns[X.columns.str.endswith('_cat')]
col_to_dummify = X.columns[X.columns.str.endswith('_cat')].astype(str).tolist()
for col in col_to_dummify:
dummy = pd.get_dummies(train_test[col].astype('category'))
columns = dummy.columns.astype(str).tolist()
columns = [col + '_' + w for w in columns]
dummy.columns = columns
train_test = pd.concat((train_test, dummy), axis=1)
train_test.drop(col_to_dummify, axis=1, inplace=True)
train_test_scaled, scaler = scale_data(train_test)
X = np.array(train_test_scaled[:n_train, :])
test = np.array(train_test_scaled[n_train:, :])
print('\n Shape of processed train data:', X.shape)
print(' Shape of processed test data:', test.shape)
```
Repeating PCA and making another plot of the first two principal components.
```
print('\nRunning PCA again ...')
pca = PCA(n_components=n_comp, svd_solver='full', random_state=1001)
X_pca = pca.fit_transform(X)
print('Explained variance: %.4f' % pca.explained_variance_ratio_.sum())
print('Individual variance contributions:')
for j in range(n_comp):
print(pca.explained_variance_ratio_[j])
plt.figure(1, figsize=(10, 10))
for color, i, target_name in zip(colors, [0, 1], target_names):
plt.scatter(X_pca[y == i, 0], X_pca[y == i, 1], color=color, s=1,
alpha=.8, label=target_name, marker='.')
plt.legend(loc='best', shadow=False, scatterpoints=3)
plt.title(
"Scatter plot of the training data projected on the 1st "
"and 2nd principal components")
plt.xlabel("Principal axis 1 - Explains %.1f %% of the variance" % (
pca.explained_variance_ratio_[0] * 100.0))
plt.ylabel("Principal axis 2 - Explains %.1f %% of the variance" % (
pca.explained_variance_ratio_[1] * 100.0))
plt.savefig('pca-porto-02.png', dpi=150)
plt.show()
```
I think that's a better plot visually and there is a good number of well-defined clusters, but still no clear separation between 0/1 points.
We can red-do the t-SNE plot as well using modified dataset. **Don't try this at home** - it takes 24+ hours using a commandline version of bh_tsne.
Anyway, here is the new t-SNE plot:

Again, lots of interesting clusters, but blue and red dots overlap for the most part.
This just happens to be a difficult classification classification problem, so maybe it is not a big surprise that raw data does not contain enough info for t-SNE to distinguish clearly between the classes.
Unfortunately, it is not much better even after training. Below is a t-SNE plot of activations from the last hidden layer (3rd) of a neural network that was trained on this dataset for 80 epochs. If you download the full version (it is roughly 10.5 x 10.5 inches), you may be able to see better that lots of red dots are concetrated in the lower left quadrat (6-9 on a clock dial), and that there are clearly fewer red dots in the upper right quadrant (0-3 on a clock dial). So the network has succeded somewhat in sequestering the red dots, but they still overlap quite a bit with blue ones.

Later I will have more t-SNE plots from neural network activations in [__this kernel__](https://www.kaggle.com/tilii7/keras-averaging-runs-gini-early-stopping).
| github_jupyter |
```
import torch
import pandas as pd
import matplotlib.pyplot as plt
import os
import subprocess
import numpy as np
os.chdir("/home/jok120/sml/proj/attention-is-all-you-need-pytorch/")
basic_train_cmd = "/home/jok120/build/anaconda3/envs/pytorch_src2/bin/python " +\
"~/sml/proj/attention-is-all-you-need-pytorch/train.py " +\
"{data} {name} -e 1000 -b 1 -nws {warmup} -cl " +\
"-dm {dm} -dih {dih} --early_stopping 20 --train_only --combined_loss"
# params = {"warmup": [1, 10, 100, 250, 500, 1000 ],
# "dm": [128, 256, 512, 1024, 2048],
# "dih": [512, 1024, 2048],
# "dwv": [8, 20, 24, 48, 128, 256]}
params_repeat = {"warmup": [5, 100, 500, 1000],
"dm": [128, 256, 512, 1024, 2048],
"dih": [512, 1024, 2048]}
new_params = {"warmup": [500, 1000, 2000, 4000],
"dm": [56, 128, 256, 512, 1024, 2048],
"dih": [512, 1024, 2048]}
# data_path = "data/data_190529_multi_helix_turns.tch"
data_path = "data/data_190530_3p7k.tch"
name = "0530-3p7k-{:03}"
for name, param in zip(["0530-sh-msea-{:03}", "0530-sh-mseb-{:03}"], [params_repeat, new_params]):
i = 0
cmds = []
for dih in param["dih"]:
for dm in param["dm"]:
for warmup in param["warmup"]:
cmd = basic_train_cmd.format(data=data_path, name=name.format(i),
warmup=warmup, dm=dm, dih=dih)
cmds.append(cmd)
print(cmd[164:] + f" logs/{name}.log".format(i))
with open("logs/" + name.format(i) + ".log", "w") as log:
subprocess.call(cmd, stdout=log, shell=True)
i += 1
```
# Analysis
```
from glob import glob
import seaborn as sns
sns.set(style="darkgrid")
result_files = sorted(glob("logs/0530-3p7k*.train"))
dfs = [pd.read_csv(f) for f in result_files]
titles = [f[5:-6] for f in result_files]
dfes = []
for df in dfs:
df["rmse"] = np.sqrt(df["mse"])
dfes.append(df[df["is_end_of_epoch"]].reset_index())
def do_plot(df, title):
sns.lineplot(x=df.index, y="drmsd", data=df, label="drmsd")
sns.lineplot(x=df.index, y="rmsd", data=df, label="rmsd")
sns.lineplot(x=df.index, y="rmse", data=df, label="rmse")
sns.lineplot(x=df.index, y="combined", data=df, label="drmsd+mse")
plt.ylabel("Loss Value")
plt.xlabel("Epoch")
plt.legend()
plt.title("{} Training".format(title))
do_plot(dfes[0], titles[0])
mins = []
for df, title in zip(dfes, titles):
row = df[df["combined"] == df["combined"].min()]
row["title"] = title[:]
mins.append(row)
mins_df = pd.concat(mins)
mins_df.sort_values("combined", inplace=True)
do_plot(dfes[232], titles[232])
mins_df
names = [int(t[-3:]) for t in mins_df["title"][:10]]
print(names)
for n in names:
do_plot(dfes[n], titles[n])
plt.show()
train_logs = [f"logs/{t}.log" for t in mins_df["title"][:10]]
train_logs
def get_arg(namespace, arg):
pattern = f"{arg}=(.+?),"
return re.search(pattern, namespace).group(1)
for row in mins:
t = row["title"].item()
with open(f"logs/{t}.log", "r") as f:
args = f.readline()
for a in ["d_model", "n_warmup_steps", "d_inner_hid"]:
row[a] = get_arg(args, a)
mins = pd.concat(mins)
mins.sort_values("combined", inplace=True)
mins
def do_loss_plots_on_var(d, var):
sns.boxplot(x=var, y="combined", data=d)
sns.swarmplot(x=var, y="combined", data=d, color="black")
plt.figure()
sns.boxplot(x=var, y="rmsd", data=d)
sns.swarmplot(x=var, y="rmsd", data=d, color="black")
plt.figure()
sns.boxplot(x=var, y="rmse", data=d)
sns.swarmplot(x=var, y="rmse", data=d, color="black")
do_loss_plots_on_var(mins, "d_model")
do_loss_plots_on_var(mins, "n_warmup_steps")
do_loss_plots_on_var(mins, "d_inner_hid")
summary = mins[["title", "combined", "drmsd", "rmsd", "rmse", "d_model", "n_warmup_steps", "d_inner_hid"]]
" ".join(summary["title"][:10].to_list())
summary
```
| github_jupyter |
# Code Coverage
In the [previous chapter](Fuzzer.ipynb), we introduced _basic fuzzing_ – that is, generating random inputs to test programs. How do we measure the effectiveness of these tests? One way would be to check the number (and seriousness) of bugs found; but if bugs are scarce, we need a _proxy for the likelihood of a test to uncover a bug._ In this chapter, we introduce the concept of *code coverage*, measuring which parts of a program are actually executed during a test run. Measuring such coverage is also crucial for test generators that attempt to cover as much code as possible.
**Prerequisites**
* You need some understanding of how a program is executed.
* You should have learned about basic fuzzing in the [previous chapter](Fuzzer.ipynb).
## Synopsis
<!-- Automatically generated. Do not edit. -->
To [use the code provided in this chapter](Importing.ipynb), write
```python
>>> from fuzzingbook.Coverage import <identifier>
```
and then make use of the following features.
This chapter introduces a `Coverage` class allowing you to measure coverage for Python programs. Its typical usage is in conjunction with a `with` clause:
```python
>>> with Coverage() as cov:
>>> cgi_decode("a+b")
```
The `trace()` method returns the coverage as a list of locations covered. Each location comes as a pair (`function name`, `line`).
```python
>>> print(cov.trace())
```
The `coverage()` method returns the set of locations executed at least once:
```python
>>> print(cov.coverage())
```
## A CGI Decoder
We start by introducing a simple Python function that decodes a CGI-encoded string. CGI encoding is used in URLs (i.e., Web addresses) to encode characters that would be invalid in a URL, such as blanks and certain punctuation:
* Blanks are replaced by `'+'`
* Other invalid characters are replaced by '`%xx`', where `xx` is the two-digit hexadecimal equivalent.
In CGI encoding, the string `"Hello, world!"` would thus become `"Hello%2c+world%21"` where `2c` and `21` are the hexadecimal equivalents of `','` and `'!'`, respectively.
The function `cgi_decode()` takes such an encoded string and decodes it back to its original form. Our implementation replicates the code from \cite{Pezze2008}. (It even includes its bugs – but we won't reveal them at this point.)
```
def cgi_decode(s):
"""Decode the CGI-encoded string `s`:
* replace "+" by " "
* replace "%xx" by the character with hex number xx.
Return the decoded string. Raise `ValueError` for invalid inputs."""
# Mapping of hex digits to their integer values
hex_values = {
'0': 0, '1': 1, '2': 2, '3': 3, '4': 4,
'5': 5, '6': 6, '7': 7, '8': 8, '9': 9,
'a': 10, 'b': 11, 'c': 12, 'd': 13, 'e': 14, 'f': 15,
'A': 10, 'B': 11, 'C': 12, 'D': 13, 'E': 14, 'F': 15,
}
t = ""
i = 0
while i < len(s):
c = s[i]
if c == '+':
t += ' '
elif c == '%':
digit_high, digit_low = s[i + 1], s[i + 2]
i += 2
if digit_high in hex_values and digit_low in hex_values:
v = hex_values[digit_high] * 16 + hex_values[digit_low]
t += chr(v)
else:
raise ValueError("Invalid encoding")
else:
t += c
i += 1
return t
```
Here is an example of how `cgi_decode()` works:
```
cgi_decode("Hello+world")
```
If we want to systematically test `cgi_decode()`, how would we proceed?
The testing literature distinguishes two ways of deriving tests: _Black-box testing_ and _White-box testing._
## Black-Box Testing
The idea of *black-box testing* is to derive tests from the _specification_. In the above case, we thus would have to test `cgi_decode()` by the features specified and documented, including
* testing for correct replacement of `'+'`;
* testing for correct replacement of `"%xx"`;
* testing for non-replacement of other characters; and
* testing for recognition of illegal inputs.
Here are four assertions (tests) that cover these four features. We can see that they all pass:
```
assert cgi_decode('+') == ' '
assert cgi_decode('%20') == ' '
assert cgi_decode('abc') == 'abc'
try:
cgi_decode('%?a')
assert False
except ValueError:
pass
```
The advantage of black-box testing is that it finds errors in the _specified_ behavior. It is independent from a given implementation, and thus allows to create test even before implementation. The downside is that _implemented_ behavior typically covers more ground than _specified_ behavior, and thus tests based on specification alone typically do not cover all implementation details.
## White-Box Testing
In contrast to black-box testing, *white-box testing* derives tests from the _implementation_, notably the internal structure. White-Box testing is closely tied to the concept of _covering_ structural features of the code. If a statement in the code is not executed during testing, for instance, this means that an error in this statement cannot be triggered either. White-Box testing thus introduces a number of *coverage criteria* that have to be fulfilled before the test can be said to be sufficient. The most frequently used coverage criteria are
* *Statement coverage* – each statement in the code must be executed by at least one test input.
* *Branch coverage* – each branch in the code must be taken by at least one test input. (This translates to each `if` and `while` decision once being true, and once being false.)
Besides these, there are far more coverage criteria, including sequences of branches taken, loop iterations taken (zero, one, many), data flows between variable definitions and usages, and many more; \cite{Pezze2008} has a great overview.
Let us consider `cgi_decode()`, above, and reason what we have to do such that each statement of the code is executed at least once. We'd have to cover
* The block following `if c == '+'`
* The two blocks following `if c == '%'` (one for valid input, one for invalid)
* The final `else` case for all other characters.
This results in the same conditions as with black-box testing, above; again, the assertions above indeed would cover every statement in the code. Such a correspondence is actually pretty common, since programmers tend to implement different behaviors in different code locations; and thus, covering these locations will lead to test cases that cover the different (specified) behaviors.
The advantage of white-box testing is that it finds errors in _implemented_ behavior. It can be conducted even in cases where the specification does not provide sufficient details; actually, it helps in identifying (and thus specifying) corner cases in the specification. The downside is that it may miss _non-implemented_ behavior: If some specified functionality is missing, white-box testing will not find it.
## Tracing Executions
One nice feature of white-box testing is that one can actually automatically assess whether some program feature was covered. To this end, one _instruments_ the execution of the program such that during execution, a special functionality keeps track of which code was executed. After testing, this information can be passed to the programmer, who can then focus on writing tests that cover the yet uncovered code.
In most programming languages, it is rather difficult to set up programs such that one can trace their execution. Not so in Python. The function `sys.settrace(f)` allows to define a *tracing function* `f()` that is called for each and every line executed. Even better, it gets access to the current function and its name, current variable contents, and more. It is thus an ideal tool for *dynamic analysis* – that is, the analysis of what actually happens during an execution.
To illustrate how this works, let us again look into a specific execution of `cgi_decode()`.
```
cgi_decode("a+b")
```
To track how the execution proceeds through `cgi_decode()`, we make use of `sys.settrace()`. First, we define the _tracing function_ that will be called for each line. It has three parameters:
* The `frame` parameter gets you the current _frame_, allowing access to the current location and variables:
* `frame.f_code` is the currently executed code with `frame.f_code.co_name` being the function name;
* `frame.f_lineno` holds the current line number; and
* `frame.f_locals` holds the current local variables and arguments.
* The `event` parameter is a string with values including `"line"` (a new line has been reached) or `"call"` (a function is being called).
* The `arg` parameter is an additional _argument_ for some events; for `"return"` events, for instance, `arg` holds the value being returned.
We use the tracing function for simply reporting the current line executed, which we access through the `frame` argument.
```
coverage = []
def traceit(frame, event, arg):
if event == "line":
global coverage
function_name = frame.f_code.co_name
lineno = frame.f_lineno
coverage.append(lineno)
return traceit
```
We can switch tracing on and off with `sys.settrace()`:
```
import sys
def cgi_decode_traced(s):
global coverage
coverage = []
sys.settrace(traceit) # Turn on
cgi_decode(s)
sys.settrace(None) # Turn off
```
When we compute `cgi_decode("a+b")`, we can now see how the execution progresses through `cgi_decode()`. After the initialization of `hex_values`, `t`, and `i`, we see that the `while` loop is taken three times – one for every character in the input.
```
cgi_decode_traced("a+b")
print(coverage)
```
Which lines are these, actually? To this end, we get the source code of `cgi_decode_code` and encode it into an array `cgi_decode_lines`, which we will then annotate with coverage information. First, let us get the source code of `cgi_encode`:
```
import inspect
cgi_decode_code = inspect.getsource(cgi_decode)
```
`cgi_decode_code` is a string holding the source code. We can print it out with Python syntax highlighting:
```
from bookutils import print_content, print_file
print_content(cgi_decode_code[:300] + "...", ".py")
```
Using `splitlines()`, we split the code into an array of lines, indexed by line number.
```
cgi_decode_lines = [""] + cgi_decode_code.splitlines()
```
`cgi_decode_lines[L]` is line L of the source code.
```
cgi_decode_lines[1]
```
We see that the first line (9) executed is actually the initialization of `hex_values`...
```
cgi_decode_lines[9:13]
```
... followed by the initialization of `t`:
```
cgi_decode_lines[15]
```
To see which lines actually have been covered at least once, we can convert `coverage` into a set:
```
covered_lines = set(coverage)
print(covered_lines)
```
Let us print out the full code, annotating lines not covered with '#':
```
for lineno in range(1, len(cgi_decode_lines)):
if lineno not in covered_lines:
print("# ", end="")
else:
print(" ", end="")
print("%2d " % lineno, end="")
print_content(cgi_decode_lines[lineno], '.py')
```
We see that a number of lines (notably comments) have not been executed, simply because they are not executable. However, we also see that the lines under `if c == '%'` have _not_ been executed yet. If `"a+b"` were our only test case so far, this missing coverage would now encourage us to create another test case that actually covers these lines.
## A Coverage Class
In this book, we will make use of coverage again and again – to _measure_ the effectiveness of different test generation techniques, but also to _guide_ test generation towards code coverage. Our previous implementation with a global `coverage` variable is a bit cumbersome for that. We therefore implement some functionality that will help us measuring coverage easily.
The key idea of getting coverage is to make use of the Python `with` statement. The general form
```python
with OBJECT [as VARIABLE]:
BODY
```
executes `BODY` with `OBJECT` being defined (and stored in `VARIABLE`). The interesting thing is that at the beginning and end of `BODY`, the special methods `OBJECT.__enter__()` and `OBJECT.__exit__()` are automatically invoked; even if `BODY` raises an exception. This allows us to define a `Coverage` object where `Coverage.__enter__()` automatically turns on tracing and `Coverage.__exit__()` automatically turns off tracing again. After tracing, we can make use of special methods to access the coverage. This is what this looks like during usage:
```python
with Coverage() as cov:
function_to_be_traced()
c = cov.coverage()
```
Here, tracing is automatically turned on during `function_to_be_traced()` and turned off again after the `with` block; afterwards, we can access the set of lines executed.
Here's the full implementation with all its bells and whistles. You don't have to get everything; it suffices that you know how to use it:
```
class Coverage(object):
# Trace function
def traceit(self, frame, event, arg):
if self.original_trace_function is not None:
self.original_trace_function(frame, event, arg)
if event == "line":
function_name = frame.f_code.co_name
lineno = frame.f_lineno
self._trace.append((function_name, lineno))
return self.traceit
def __init__(self):
self._trace = []
# Start of `with` block
def __enter__(self):
self.original_trace_function = sys.gettrace()
sys.settrace(self.traceit)
return self
# End of `with` block
def __exit__(self, exc_type, exc_value, tb):
sys.settrace(self.original_trace_function)
def trace(self):
"""The list of executed lines, as (function_name, line_number) pairs"""
return self._trace
def coverage(self):
"""The set of executed lines, as (function_name, line_number) pairs"""
return set(self.trace())
```
Let us put this to use:
```
with Coverage() as cov:
cgi_decode("a+b")
print(cov.coverage())
```
As you can see, the `Coverage()` class not only keeps track of lines executed, but also of function names. This is useful if you have a program that spans multiple files.
## Comparing Coverage
Since we represent coverage as a set of executed lines, we can also apply _set operations_ on these. For instance, we can find out which lines are covered by individual test cases, but not others:
```
with Coverage() as cov_plus:
cgi_decode("a+b")
with Coverage() as cov_standard:
cgi_decode("abc")
cov_plus.coverage() - cov_standard.coverage()
```
This is the single line in the code that is executed only in the `'a+b'` input.
We can also compare sets to find out which lines still need to be covered. Let us define `cov_max` as the maximum coverage we can achieve. (Here, we do this by executing the "good" test cases we already have. In practice, one would statically analyze code structure, which we introduce in [the chapter on symbolic testing](SymbolicFuzzer.ipynb).)
```
import bookutils
with Coverage() as cov_max:
cgi_decode('+')
cgi_decode('%20')
cgi_decode('abc')
try:
cgi_decode('%?a')
except:
pass
```
Then, we can easily see which lines are _not_ yet covered by a test case:
```
cov_max.coverage() - cov_plus.coverage()
```
Again, these would be the lines handling `"%xx"`, which we have not yet had in the input.
## Coverage of Basic Fuzzing
We can now use our coverage tracing to assess the _effectiveness_ of testing methods – in particular, of course, test _generation_ methods. Our challenge is to achieve maximum coverage in `cgi_decode()` just with random inputs. In principle, we should _eventually_ get there, as eventually, we will have produced every possible string in the universe – but exactly how long is this? To this end, let us run just one fuzzing iteration on `cgi_decode()`:
```
from Fuzzer import fuzzer
sample = fuzzer()
sample
```
Here's the invocation and the coverage we achieve. We wrap `cgi_decode()` in a `try...except` block such that we can ignore `ValueError` exceptions raised by illegal `%xx` formats.
```
with Coverage() as cov_fuzz:
try:
cgi_decode(sample)
except:
pass
cov_fuzz.coverage()
```
Is this already the maximum coverage? Apparently, there are still lines missing:
```
cov_max.coverage() - cov_fuzz.coverage()
```
Let us try again, increasing coverage over 100 random inputs. We use an array `cumulative_coverage` to store the coverage achieved over time; `cumulative_coverage[0]` is the total number of lines covered after input 1,
`cumulative_coverage[1]` is the number of lines covered after inputs 1–2, and so on.
```
trials = 100
def population_coverage(population, function):
cumulative_coverage = []
all_coverage = set()
for s in population:
with Coverage() as cov:
try:
function(s)
except:
pass
all_coverage |= cov.coverage()
cumulative_coverage.append(len(all_coverage))
return all_coverage, cumulative_coverage
```
Let us create a hundred inputs to determine how coverage increases:
```
def hundred_inputs():
population = []
for i in range(trials):
population.append(fuzzer())
return population
```
Here's how the coverage increases with each input:
```
all_coverage, cumulative_coverage = population_coverage(
hundred_inputs(), cgi_decode)
%matplotlib inline
import matplotlib.pyplot as plt # type: ignore
plt.plot(cumulative_coverage)
plt.title('Coverage of cgi_decode() with random inputs')
plt.xlabel('# of inputs')
plt.ylabel('lines covered')
```
This is just _one_ run, of course; so let's repeat this a number of times and plot the averages.
```
runs = 100
# Create an array with TRIALS elements, all zero
sum_coverage = [0] * trials
for run in range(runs):
all_coverage, coverage = population_coverage(hundred_inputs(), cgi_decode)
assert len(coverage) == trials
for i in range(trials):
sum_coverage[i] += coverage[i]
average_coverage = []
for i in range(trials):
average_coverage.append(sum_coverage[i] / runs)
plt.plot(average_coverage)
plt.title('Average coverage of cgi_decode() with random inputs')
plt.xlabel('# of inputs')
plt.ylabel('lines covered')
```
We see that on average, we get full coverage after 40–60 fuzzing inputs.
## Getting Coverage from External Programs
Of course, not all the world is programming in Python. The good news is that the problem of obtaining coverage is ubiquitous, and almost every programming language has some facility to measure coverage. Just as an example, let us therefore demonstrate how to obtain coverage for a C program.
Our C program (again) implements `cgi_decode`; this time as a program to be executed from the command line:
```shell
$ ./cgi_decode 'Hello+World'
Hello World
```
Here comes the C code, first as a Python string. We start with the usual C includes:
```
cgi_c_code = """
/* CGI decoding as C program */
#include <stdlib.h>
#include <string.h>
#include <stdio.h>
"""
```
Here comes the initialization of `hex_values`:
```
cgi_c_code += r"""
int hex_values[256];
void init_hex_values() {
for (int i = 0; i < sizeof(hex_values) / sizeof(int); i++) {
hex_values[i] = -1;
}
hex_values['0'] = 0; hex_values['1'] = 1; hex_values['2'] = 2; hex_values['3'] = 3;
hex_values['4'] = 4; hex_values['5'] = 5; hex_values['6'] = 6; hex_values['7'] = 7;
hex_values['8'] = 8; hex_values['9'] = 9;
hex_values['a'] = 10; hex_values['b'] = 11; hex_values['c'] = 12; hex_values['d'] = 13;
hex_values['e'] = 14; hex_values['f'] = 15;
hex_values['A'] = 10; hex_values['B'] = 11; hex_values['C'] = 12; hex_values['D'] = 13;
hex_values['E'] = 14; hex_values['F'] = 15;
}
"""
```
Here's the actual implementation of `cgi_decode()`, using pointers for input source (`s`) and output target (`t`):
```
cgi_c_code += r"""
int cgi_decode(char *s, char *t) {
while (*s != '\0') {
if (*s == '+')
*t++ = ' ';
else if (*s == '%') {
int digit_high = *++s;
int digit_low = *++s;
if (hex_values[digit_high] >= 0 && hex_values[digit_low] >= 0) {
*t++ = hex_values[digit_high] * 16 + hex_values[digit_low];
}
else
return -1;
}
else
*t++ = *s;
s++;
}
*t = '\0';
return 0;
}
"""
```
Finally, here's a driver which takes the first argument and invokes `cgi_decode` with it:
```
cgi_c_code += r"""
int main(int argc, char *argv[]) {
init_hex_values();
if (argc >= 2) {
char *s = argv[1];
char *t = malloc(strlen(s) + 1); /* output is at most as long as input */
int ret = cgi_decode(s, t);
printf("%s\n", t);
return ret;
}
else
{
printf("cgi_decode: usage: cgi_decode STRING\n");
return 1;
}
}
"""
```
Let us create the C source code: (Note that the following commands will overwrite the file `cgi_decode.c`, if it already exists in the current working directory. Be aware of this, if you downloaded the notebooks and are working locally.)
```
with open("cgi_decode.c", "w") as f:
f.write(cgi_c_code)
```
And here we have the C code with its syntax highlighted:
```
from bookutils import print_file
print_file("cgi_decode.c")
```
We can now compile the C code into an executable. The `--coverage` option instructs the C compiler to instrument the code such that at runtime, coverage information will be collected. (The exact options vary from compiler to compiler.)
```
!cc --coverage -o cgi_decode cgi_decode.c
```
When we now execute the program, coverage information will automatically be collected and stored in auxiliary files:
```
!./cgi_decode 'Send+mail+to+me%40fuzzingbook.org'
```
The coverage information is collected by the `gcov` program. For every source file given, it produces a new `.gcov` file with coverage information.
```
!gcov cgi_decode.c
```
In the `.gcov` file, each line is prefixed with the number of times it was called (`-` stands for a non-executable line, `#####` stands for zero) as well as the line number. We can take a look at `cgi_decode()`, for instance, and see that the only code not executed yet is the `return -1` for an illegal input.
```
lines = open('cgi_decode.c.gcov').readlines()
for i in range(30, 50):
print(lines[i], end='')
```
Let us read in this file to obtain a coverage set:
```
def read_gcov_coverage(c_file):
gcov_file = c_file + ".gcov"
coverage = set()
with open(gcov_file) as file:
for line in file.readlines():
elems = line.split(':')
covered = elems[0].strip()
line_number = int(elems[1].strip())
if covered.startswith('-') or covered.startswith('#'):
continue
coverage.add((c_file, line_number))
return coverage
coverage = read_gcov_coverage('cgi_decode.c')
list(coverage)[:5]
```
With this set, we can now do the same coverage computations as with our Python programs.
## Finding Errors with Basic Fuzzing
Given sufficient time, we can indeed cover each and every line within `cgi_decode()`, whatever the programming language would be. This does not mean that they would be error-free, though. Since we do not check the result of `cgi_decode()`, the function could return any value without us checking or noticing. To catch such errors, we would have to set up a *results checker* (commonly called an *oracle*) that would verify test results. In our case, we could compare the C and Python implementations of `cgi_decode()` and see whether both produce the same results.
Where fuzzing is great at, though, is in finding _internal errors_ that can be detected even without checking the result. Actually, if one runs our `fuzzer()` on `cgi_decode()`, one quickly finds such an error, as the following code shows:
```
from ExpectError import ExpectError
with ExpectError():
for i in range(trials):
try:
s = fuzzer()
cgi_decode(s)
except ValueError:
pass
```
So, it is possible to cause `cgi_decode()` to crash. Why is that? Let's take a look at its input:
```
s
```
The problem here is at the end of the string. After a `'%'` character, our implementation will always attempt to access two more (hexadecimal) characters, but if these are not there, we will get an `IndexError` exception.
This problem is also present in our C variant, which inherits it from the original implementation \cite{Pezze2008}:
```c
int digit_high = *++s;
int digit_low = *++s;
```
Here, `s` is a pointer to the character to be read; `++` increments it by one character.
In the C implementation, the problem is actually much worse. If the `'%'` character is at the end of the string, the above code will first read a terminating character (`'\0'` in C strings) and then the following character, which may be any memory content after the string, and which thus may cause the program to fail uncontrollably. The somewhat good news is that `'\0'` is not a valid hexadecimal character, and thus, the C version will "only" read one character beyond the end of the string.
Interestingly enough, none of the manual tests we had designed earlier would trigger this bug. Actually, neither statement nor branch coverage, nor any of the coverage criteria commonly discussed in literature would find it. However, a simple fuzzing run can identify the error with a few runs – _if_ appropriate run-time checks are in place that find such overflows. This definitely calls for more fuzzing!
## Synopsis
This chapter introduces a `Coverage` class allowing you to measure coverage for Python programs. Its typical usage is in conjunction with a `with` clause:
```
with Coverage() as cov:
cgi_decode("a+b")
```
The `trace()` method returns the coverage as a list of locations covered. Each location comes as a pair (`function name`, `line`).
```
print(cov.trace())
```
The `coverage()` method returns the set of locations executed at least once:
```
print(cov.coverage())
```
## Lessons Learned
* Coverage metrics are a simple and fully automated means to approximate how much functionality of a program is actually executed during a test run.
* A number of coverage metrics exist, the most important ones being statement coverage and branch coverage.
* In Python, it is very easy to access the program state during execution, including the currently executed code.
At the end of the day, let's clean up: (Note that the following commands will delete all files in the current working directory that fit the pattern `cgi_decode.*`. Be aware of this, if you downloaded the notebooks and are working locally.)
```
import os
import glob
for file in glob.glob("cgi_decode") + glob.glob("cgi_decode.*"):
os.remove(file)
```
## Next Steps
Coverage is not only a tool to _measure_ test effectiveness, but also a great tool to _guide_ test generation towards specific goals – in particular uncovered code. We use coverage to
* [guide _mutations_ of existing inputs towards better coverage in the chapter on mutation fuzzing](MutationFuzzer.ipynb)
## Background
Coverage is a central concept in systematic software testing. For discussions, see the books in the [Introduction to Testing](Intro_Testing.ipynb).
## Exercises
### Exercise 1: Fixing cgi_decode
Create an appropriate test to reproduce the `IndexError` discussed above. Fix `cgi_decode()` to prevent the bug. Show that your test (and additional `fuzzer()` runs) no longer expose the bug. Do the same for the C variant.
**Solution.** Here's a test case:
```
with ExpectError():
assert cgi_decode('%') == '%'
with ExpectError():
assert cgi_decode('%4') == '%4'
assert cgi_decode('%40') == '@'
```
Here's a fix:
```
def fixed_cgi_decode(s):
"""Decode the CGI-encoded string `s`:
* replace "+" by " "
* replace "%xx" by the character with hex number xx.
Return the decoded string. Raise `ValueError` for invalid inputs."""
# Mapping of hex digits to their integer values
hex_values = {
'0': 0, '1': 1, '2': 2, '3': 3, '4': 4,
'5': 5, '6': 6, '7': 7, '8': 8, '9': 9,
'a': 10, 'b': 11, 'c': 12, 'd': 13, 'e': 14, 'f': 15,
'A': 10, 'B': 11, 'C': 12, 'D': 13, 'E': 14, 'F': 15,
}
t = ""
i = 0
while i < len(s):
c = s[i]
if c == '+':
t += ' '
elif c == '%' and i + 2 < len(s): # <--- *** FIX ***
digit_high, digit_low = s[i + 1], s[i + 2]
i += 2
if digit_high in hex_values and digit_low in hex_values:
v = hex_values[digit_high] * 16 + hex_values[digit_low]
t += chr(v)
else:
raise ValueError("Invalid encoding")
else:
t += c
i += 1
return t
assert fixed_cgi_decode('%') == '%'
assert fixed_cgi_decode('%4') == '%4'
assert fixed_cgi_decode('%40') == '@'
```
Here's the test:
```
for i in range(trials):
try:
s = fuzzer()
fixed_cgi_decode(s)
except ValueError:
pass
```
For the C variant, the following will do:
```
cgi_c_code = cgi_c_code.replace(
r"if (*s == '%')", # old code
r"if (*s == '%' && s[1] != '\0' && s[2] != '\0')" # new code
)
```
Go back to the above compilation commands and recompile `cgi_decode`.
### Exercise 2: Branch Coverage
Besides statement coverage, _branch coverage_ is one of the most frequently used criteria to determine the quality of a test. In a nutshell, branch coverage measures how many different _control decisions_ are made in code. In the statement
```python
if CONDITION:
do_a()
else:
do_b()
```
for instance, both the cases where `CONDITION` is true (branching to `do_a()`) and where `CONDITION` is false (branching to `do_b()`) have to be covered. This holds for all control statements with a condition (`if`, `while`, etc.).
How is branch coverage different from statement coverage? In the above example, there is actually no difference. In this one, though, there is:
```python
if CONDITION:
do_a()
something_else()
```
Using statement coverage, a single test case where `CONDITION` is true suffices to cover the call to `do_a()`. Using branch coverage, however, we would also have to create a test case where `do_a()` is _not_ invoked.
Using our `Coverage` infrastructure, we can simulate branch coverage by considering _pairs of subsequent lines executed_. The `trace()` method gives us the list of lines executed one after the other:
```
with Coverage() as cov:
cgi_decode("a+b")
trace = cov.trace()
trace[:5]
```
#### Part 1: Compute branch coverage
Define a function `branch_coverage()` that takes a trace and returns the set of pairs of subsequent lines in a trace – in the above example, this would be
```python
set(
(('cgi_decode', 9), ('cgi_decode', 10)),
(('cgi_decode', 10), ('cgi_decode', 11)),
# more_pairs
)
```
Bonus for advanced Python programmers: Define `BranchCoverage` as a subclass of `Coverage` and make `branch_coverage()` as above a `coverage()` method of `BranchCoverage`.
**Solution.** Here's a simple definition of `branch_coverage()`:
```
def branch_coverage(trace):
coverage = set()
past_line = None
for line in trace:
if past_line is not None:
coverage.add((past_line, line))
past_line = line
return coverage
branch_coverage(trace)
```
Here's a definition as a class:
```
class BranchCoverage(Coverage):
def coverage(self):
"""The set of executed line pairs"""
coverage = set()
past_line = None
for line in self.trace():
if past_line is not None:
coverage.add((past_line, line))
past_line = line
return coverage
```
#### Part 2: Comparing statement coverage and branch coverage
Use `branch_coverage()` to repeat the experiments in this chapter with branch coverage rather than statement coverage. Do the manually written test cases cover all branches?
**Solution.** Let's repeat the above experiments with `BranchCoverage`:
```
with BranchCoverage() as cov:
cgi_decode("a+b")
print(cov.coverage())
with BranchCoverage() as cov_plus:
cgi_decode("a+b")
with BranchCoverage() as cov_standard:
cgi_decode("abc")
cov_plus.coverage() - cov_standard.coverage()
with BranchCoverage() as cov_max:
cgi_decode('+')
cgi_decode('%20')
cgi_decode('abc')
try:
cgi_decode('%?a')
except:
pass
cov_max.coverage() - cov_plus.coverage()
sample
with BranchCoverage() as cov_fuzz:
try:
cgi_decode(s)
except:
pass
cov_fuzz.coverage()
cov_max.coverage() - cov_fuzz.coverage()
def population_branch_coverage(population, function):
cumulative_coverage = []
all_coverage = set()
for s in population:
with BranchCoverage() as cov:
try:
function(s)
except:
pass
all_coverage |= cov.coverage()
cumulative_coverage.append(len(all_coverage))
return all_coverage, cumulative_coverage
all_branch_coverage, cumulative_branch_coverage = population_branch_coverage(
hundred_inputs(), cgi_decode)
plt.plot(cumulative_branch_coverage)
plt.title('Branch coverage of cgi_decode() with random inputs')
plt.xlabel('# of inputs')
plt.ylabel('line pairs covered')
len(cov_max.coverage())
all_branch_coverage - cov_max.coverage()
```
The additional coverage comes from the exception raised via an illegal input (say, `%g`).
```
cov_max.coverage() - all_branch_coverage
```
This is an artefact coming from the subsequent execution of `cgi_decode()` when computing `cov_max`.
#### Part 3: Average coverage
Again, repeat the above experiments with branch coverage. Does `fuzzer()` cover all branches, and if so, how many tests does it take on average?
**Solution.** We repeat the experiments we ran with line coverage with branch coverage.
```
runs = 100
# Create an array with TRIALS elements, all zero
sum_coverage = [0] * trials
for run in range(runs):
all_branch_coverage, coverage = population_branch_coverage(
hundred_inputs(), cgi_decode)
assert len(coverage) == trials
for i in range(trials):
sum_coverage[i] += coverage[i]
average_coverage = []
for i in range(trials):
average_coverage.append(sum_coverage[i] / runs)
plt.plot(average_coverage)
plt.title('Average branch coverage of cgi_decode() with random inputs')
plt.xlabel('# of inputs')
plt.ylabel('line pairs covered')
```
We see that achieving branch coverage takes longer than statement coverage; it simply is a more difficult criterion to satisfy with random inputs.
| github_jupyter |
# Mission pilot: GTR EDA
In this pilot, we are developing indicators to evaluate the impact of innovation missions.
We are doing this with Gateway to Research and Horizon 2020 data, and with two two missions in mind. They are:
* Use of AI to diagnose, prevent and treat chronic diseases
* Improve quality of life for an aging population
We want to write a pipeline that more or less automatically takes keywords related to missions, queries the data and generates indicators and visualisations.
The two datasets we are working with have similar structures (*projects linked to organisations and outcomes*) so it would be nice to write a pipeline that allows us to move between sources seamlessly.
The output will be a methodology and preliminary findings for a EURITO working paper. Emerging findings will also be relevant for the project mapping innovation in Scotland.
## Tasks
1. Load and explore GtR data
* Link between projects and outcomes
* Coverage of outcomes
2. Process GtR data
* Classify projects into disciplines
* Label projects with TRLs (this will come from other team-members)
3. Create mission fields
* Generate mission-related keywords
* Query the data
4. Generate indicators
* Level of activity per mission
* Evolution of activity per mission (absolute and normalised)
* Network structure of mission
* Interdisciplinary activity and its evolution
* Communities involved
* % of relevant fields involved in mission (eg how much of total AI activity is about chronic diseases; how much of aging is about quality of life and well-being)?
* Technological maturity of fields
* Explore distribution of outcomes by mission area
* Generate lists of DOIs (papers) for cross-ref analysis
5. Report
### 0. Preamble
#### Imports
```
%matplotlib inline
import matplotlib.pyplot as plt
# Functions
def missing_count(df,ax):
'''
Creates a barchart with share of missing values by variable in the data
'''
df.apply(lambda x: pd.isnull(x),axis=1).mean().sort_values(ascending=False).plot.bar(ax=ax,color='blue')
def get_year(date):
'''
Extracts years from GTR data. These are strings so we just get the first number
'''
if pd.isnull(date)==False:
y = date.split('-')[0]
return(int(y))
else:
return(np.nan)
```
### 1. Load GtR data
We will load the data in a different Notebook (we need a special environment to use the data getters)
```
my_tables = ['_projects','_organisations','_products','_intellectualproperties','_publications',
'technicalproducts',
'_spinouts','_link_table']
all_tables = os.listdir('../data/external/gtr/2019-01-24/')
all_tables = os.listdir('../data/external/gtr/2019-01-24/')
#A dict where every item is a df with the key = table name. We do a bit of string manipulation to remove dates etc.
my_data = {file.split('_')[-1][:-4]:pd.read_csv('../data/external/gtr/2019-01-24/'+file).iloc[:,1:] for file in all_tables if any(x in file for x in my_tables)}
my_data.keys()
#Load the linking file here because we will use it throughout
link = my_data['table']
link.head()
```
### Projects
```
projects = my_data['projects']
projects.head()
```
Some rapid observations:
* what does start mean?
* What does created mean?
* abstractText seems to be the main textual description
```
fig,ax = plt.subplots()
missing_count(projects,ax)
ax.set_title('Missing values in Project GTR data')
projects['year_created'] = projects['created'].apply(lambda x: get_year(x))
projects['year_created'].value_counts()
```
Created is a data collection variable
```
projects['year_started'] = projects['start'].apply(lambda x: get_year(x))
projects['year_started'].value_counts().loc[np.arange(2006,2020)].plot.bar(color='blue',title='Started year')
```
Not sure about what `year_stared` means but it doesn't cover many records
```
projects['leadFunder'].value_counts().plot.bar(color='blue',title='funder_distribution')
```
EPSRC and Innovate UK are the most active organisations in the data
```
projects['grantCategory'].value_counts(normalize=True)[:10].plot.bar(color='blue',title='project types')
```
Research grants are the main category with almost half of the observations
##### Does the fund data contain the actual start and end date for the project?
```
fund = pd.read_csv('../data/external/gtr/2019-01-24/gtr_funds.csv').iloc[:,1:]
fund.head()
fund.shape
```
One fund per project. Bodes well
```
fund['year'] = fund['start'].apply(lambda x: get_year(x))
fund['year'].value_counts().loc[np.arange(2000,2020)].plot.bar(color='blue',title='Start date (funded data)')
```
The dataset begins in 2016. What is the bump in 2017?
```
pd.crosstab(fund['category'],fund['year'])[np.arange(2006,2020)].T.plot.bar(stacked=True,title='funding by category')
```
Most of the funding levels in the data reflect income
#### Merge funding with projects to get dates and funding by project
`projects_f = projects merged w/ funding`
```
#Merge everything. It works
projects_f = pd.merge(
pd.merge(fund,link,left_on='id',right_on='id'),
projects,
left_on='project_id',right_on='id')
projects_f.head()
```
Lots of guff there. I should tidy it up later
#### Let's also check the Topics: does this refer to project keywords?
```
topics = pd.read_csv('../data/external/gtr/2019-01-24/gtr_topic.csv').iloc[:,1:]
topics.head()
topics['topic_type'].value_counts()
```
Yes. What are their keys in the linked_table?
```
link['table_name'].value_counts()
```
We can loop over the projects and allocate them topics
Create a research activity lookup and a research topic lookup.
```
res_activity_lookup,res_topic_lookup = [{this_id:text for this_id,text in zip(topics.loc[topics['topic_type']==topic_type,'id'],
topics.loc[topics['topic_type']==topic_type,'text'])} for
topic_type in ['researchActivity','researchTopic']]
list(res_activity_lookup.values())[:10]
```
`researchActivity` is a medical set of subjects
```
list(res_topic_lookup.values())[:10]
```
`researchTopics` is generic
### Organisations
```
orgs = my_data['organisations']
orgs.head()
orgs.shape
```
This will be interesting to look at collaboration networks between organisations.
Note that this is likely to over-estimate collaboration given that departments (which might be disconnected) will be subsumed under organisations. Maybe we could use the persons data to unpick that?
### Outputs
Brief exploration of the data and standardisation with a single schema (for merging with projects)
#### Products
```
products = my_data['products']
products.head()
products.shape
```
They seem to be primarily medical products
```
products.stage.value_counts()
products.type.value_counts()
```
Yes - this is clearly a medical database. We might use it for our two missions given their health focus
#### Intellectual Property
```
ip = my_data['intellectualproperties']
ip.head()
ip.shape
ip.protection.value_counts().plot.bar(color='blue',title='ip')
```
Around 3000 patents and a bunch of random labels. We can use it perhaps to look at field maturity
#### Technical products
```
technical = my_data['softwareandtechnicalproducts']
technical.head()
technical.shape
technical['type'].value_counts().plot.bar(color='blue',title='Technical outputs')
```
Perhaps we could create a 'practical output' dummy for the field...
Or query the application databases with project names once we have identified them?
```
np.sum(['github' in x for x in technical.supportingUrl if pd.isnull(x)==False])
```
1686 projects in GitHUb
#### Spinouts
```
spinouts = my_data['spinouts']
spinouts.head()
spinouts.yearEstablished.value_counts().loc[np.arange(2006,2020)].plot.bar(color='blue',title='spinout_year')
```
#### Publications
```
pubs = my_data['publications']
pubs.head()
np.random.randint(0,len(pubs),40000)
#Get missing values for a random sample (10%) of the publications
fig,ax = plt.subplots()
pub_sample = pubs.loc[list(np.random.randint(0,len(pubs),70000)),:]
missing_count(pub_sample,ax)
```
80% have DOI
```
pubs.type.value_counts().plot.bar(color='blue',title='Type of publication')
pubs['year'] = pubs['datePublished'].apply(get_year)
pubs['year'].value_counts().loc[np.arange(2006,2020)].plot.bar(color='blue',title='publication years')
pubs['journalTitle'].value_counts()[:10]
```
### Discipline coverage check
Here we run outputs vs projects to get who were the funders. We want to check how much cross-organisation coverage there is in the data
We also identify projects that appear in different 'impact' databases
```
#Containers for the data
impactful_projects= []
impact_funders = [projects['leadFunder'].value_counts()]
impact_names = ['prods','ip','tech','spin','pubs']
for name,data in zip(impact_names,[products,ip,technical,spinouts,pubs]):
#Merges outputs and projects via the product file
merged = pd.merge(
pd.merge(data,link,left_on='id',right_on='id'),
projects,left_on='project_id',right_on='id')
#number of times that a project appears in an output df
project_counts = merged['project_id'].value_counts()
project_counts.name = name
#Put it with the featured projects
impactful_projects.append(project_counts)
#Funder impact by project
funder_freqs = merged['leadFunder'].value_counts()
funder_freqs.name = name
#Merges with the outpi
impact_funders.append(funder_freqs)
fig,ax = plt.subplots(figsize=(8,5))
pd.concat(impact_funders,axis=1,sort=False).apply(lambda x: x/x.sum(),axis=0).T.plot.bar(stacked=True,ax=ax,width=0.8)
ax.legend(bbox_to_anchor=(1,1))
```
Some observations:
* No output data for innovate UK
* MRC over-represented in all outputs (not bad for the mission pilots)
* STFC over-represented in publications(physics)
```
# And the impactful projects
#Concatenate the previous outputs
project_impacts = pd.concat(impactful_projects,axis=1,sort=True).fillna(0)
#Concatenate with the projects file
projects_imp = pd.concat([projects_f.set_index('project_id'),project_impacts],axis=1)
projects_imp.shape
```
Fillnas with 0s.
Should drop innovate from follow-on analyses here given that they don't seem to be tracking the impact of their projects in the same way as research councils.
```
projects_imp[impact_names] = projects_imp[impact_names].fillna(0)
import seaborn as sns
#What is the correlation between different types of outputs...and level of funding?
projects_imp[impact_names+['amount']].corr()
```
Lots of potential confounders here - types of projects, disciplines...
#### Some observations for next steps
* Analysis focusing on 2006-2018
* We need to integrate the topic data (new link file)?
* Analyses of TRL/output by field will need to consider differences between disciplines in their outlets. To which extent are the disciplines in a mission field more or less productive than their constituent fields?
* ...
## 2. Process data
* Classify projects into disciplines.
* I need the project topics for this labelling
* Any TRL tags would go in here
Add research activities and topics to the projects data
```
topic_lookup = link.loc[link['table_name']=='gtr_topic',:]
#We simply group by projects and run the names vs the research activity and topic lookup
#Group
res_topics_by_project = topic_lookup.groupby('project_id')
#Extract lookups
activity_project_lookup,topic_project_lookup = [res_topics_by_project['id'].apply(
lambda x: [lookup[el] for el in [t for t in x] if el in lookup.keys()]).to_dict() for lookup in [res_activity_lookup,res_topic_lookup]]
#Add topics
projects_imp['research_topics'],projects_imp['research_activities']= [[
lookup[x] if x in lookup.keys() else [] for x in projects_imp.index] for lookup in [
topic_project_lookup,activity_project_lookup]]
projects_imp['has_topic'],projects_imp['has_activities'] = [[len(x)>0 for x in projects_imp[var]] for var in
['research_topics','research_activities']]
pd.crosstab(projects_imp.loc[projects_imp.grantCategory=='Research Grant','year'],
projects_imp.loc[projects_imp.grantCategory=='Research Grant','has_topic'],normalize=0).plot.bar(
stacked=True,title='Share of Research Grants with subject data')
```
Only around 80% of the projects have topics. This is quite different from previous versions of the data which had many more topics.
One to check with Joel
```
fig,ax = plt.subplots(figsize=(10,5),ncols=2,sharey=True)
pd.crosstab(projects_imp['leadFunder'],projects_imp['has_topics'],normalize=0).plot.barh(stacked=True,ax=ax[0],legend=False)
pd.crosstab(projects_imp['leadFunder'],projects_imp['has_activities'],normalize=0).plot.barh(stacked=True,ax=ax[1])
```
Almost no topics in MRC - because they label their projects with activities.
For now we will use an older dataset with better topic coverage for labelling (see notebook `02_jmg_discipline_modelling`)
### Discipline classifier
Save data to train the model in the `02_jmg`... notebook
```
gtr_for_pred = projects_imp.dropna(axis=0,subset=['abstractText'])
gtr_for_pred.to_csv(f'../data/processed/{today_str}_gtr_for_prediction.csv',index_label=False)
```
Load data after training
```
gtr_predicted = pd.read_csv('../data/processed/24_1_2019_gtr_w_predicted_labels.csv',index_col=None).iloc[:,1:].set_index('index')
```
### Create a clean dataset
```
discs = ['biological_sciences', 'physics', 'engineering_technology',
'medical_sciences', 'social_sciences', 'mathematics_computing',
'environmental_sciences', 'arts_humanities']
my_vars = ['title','year','abstractText','status','grantCategory','leadFunder','amount']
rename_vars = ['title','year','abstract','status','grant_category','funder','amount']
proj_labelled = pd.concat([projects_imp,gtr_predicted],axis=1,sort=False)[my_vars + discs + impact_names]
proj_labelled.rename(columns={x:y for x,y in zip(my_vars,rename_vars)},inplace=True)
#We will focus on projects with abstracts
proj_abst = proj_labelled.loc[gtr_predicted.index,:]
proj_abst.to_csv(f'../data/processed/{today_str}_projects_clean.csv')
```
Notebook `003` continues with the analysis
| github_jupyter |
# Balanced Parentheses Exercise
In this exercise you are going to apply what you learned about stacks with a real world problem. We will be using stacks to make sure the parentheses are balanced in mathematical expressions such as: $((3^2 + 8)*(5/2))/(2+6).$ In real life you can see this extend to many things such as text editor plugins and interactive development environments for all sorts of bracket completion checks.
Take a string as an input and return `True` if it's parentheses are balanced or `False` if it is not.
Try to code up a solution and pass the test cases.
#### Code
```
# Our Stack Class - Brought from previous concept
# No need to modify this
class Stack:
def __init__(self):
self.items = []
def size(self):
return len(self.items)
def push(self, item):
self.items.append(item)
def pop(self):
if self.size()==0:
return None
else:
return self.items.pop()
def equation_checker(equation):
"""
Check equation for balanced parentheses
Args:
equation(string): String form of equation
Returns:
bool: Return if parentheses are balanced or not
"""
# TODO: Intiate stack object
# TODO: Interate through equation checking parentheses
# TODO: Return True if balanced and False if not
pass
```
#### Test Cases
```
print ("Pass" if (equation_checker('((3^2 + 8)*(5/2))/(2+6)')) else "Fail")
print ("Pass" if not (equation_checker('((3^2 + 8)*(5/2))/(2+6))')) else "Fail")
```
<span class="graffiti-highlight graffiti-id_ky3qi6e-id_jfute45"><i></i><button>Hide Solution</button></span>
```
# Solution
# Our Stack Class
class Stack:
def __init__(self):
self.items = []
def size(self):
return len(self.items)
def push(self, item):
self.items.append(item)
def pop(self):
if self.size()==0:
return None
else:
return self.items.pop()
def equation_checker(equation):
"""
Check equation for balanced parentheses
Args:
equation(string): String form of equation
Returns:
bool: Return if parentheses are balanced or not
"""
stack = Stack()
for char in equation:
if char == "(":
stack.push(char)
elif char == ")":
if stack.pop() == None:
return False
if stack.size() == 0:
return True
else:
return False
print ("Pass" if (equation_checker('((3^2 + 8)*(5/2))/(2+6)')) else "Fail")
print ("Pass" if not (equation_checker('((3^2 + 8)*(5/2))/(2+6))')) else "Fail")
```
| github_jupyter |
# Constellation and Chain Analysis: MultiConstellation Multihop
<img src="MultipleConstellations.jpg" alt="Drawing" style="width: 500px;"/>
**Terminology**
* Node = Object in STK
* Edge = Access between two objects in STK
* Strand = The sequence of nodes and edges to complete access in a chain
**This notebook shows how to:**
* Find the shortest path between a starting and ending constellation, with many potential intermediate constellations.
* A chain will be built between each sequential pair in the constellationOrder list. Then networkx will be used build the network with the nodes coming from the constellations and the connections between the nodes coming from the chain accesses. Multiple sublists can be passed into constellationOrderList.
* To reduce the runtime on subsequent runs, the results can be saved to binary files and loaded in for later use. The strands from the chains will be saved in the SavedNodes folder, the nodes and associated time delays will be saved in the SavedNodes folder, the node positions over time are saved in the SavedPositions folder and the accesses between nodes over time are saved in the SavedEdges folder. These folders will be created as subfolders of the directory used to run the script. During the first run the files will automatically be built and saved, subsequent runs will reload these files. To make changes simply delete the associated .pkl file for any changed strands, nodes, etc. and the script will recompute the data as needed. Or force all of the data to be overridden by setting the override options to be True.
* Typical STK constraints such as range, link duration, Eb/No, etc are taken into account
* Data in the df variable can be pushed back into STK as a user supplied variable, a strand can be shown using object lines, and active objects over the analysis time period or at a time instance can be turned on.
```
import numpy as np
import pandas as pd
pd.set_option('max_colwidth', 120)
from comtypes.client import CreateObject
from comtypes.client import GetActiveObject
from comtypes.gen import STKObjects
from comtypes.gen import STKUtil
from comtypes.gen import AgSTKVgtLib
import seaborn as sns
import matplotlib.pyplot as plt
from chainPathLib2 import *
import time
import networkx as nx
folders = ['SavedNodes','SavedPositions','SavedStrands','SavedEdges','SavedNetworkData']
for folder in folders:
if not os.path.exists(folder):
os.makedirs(folder)
```
## Constellation Connection Order, Computation Time, Metric, Saved Data Options
```
constellationOrderLists = [['Targets','ObservingSatsFORs','ObservingSats','ObservingSatsTransmitters','ObservingSatsReceivers','ObservingSats','RelaySatsFORs','RelaySats','RelaySatsFORs','EndLocations'],['RelaySats','RelaySats']]
startingConstellation = 'Targets'
endingConstellation = 'EndLocations'
start = 0 # EpSec
stop = 60*10 # EpSec
metric = 'timeDelay' # 'distance' or 'timeDelay'
nodeDelays = {'ObservingSatsFORs':0.01,'ObservingSatsTransmitters':0.005,'ObservingSatsReceivers':0.005,'RelaySatsFORs':0.01,'RelaySats':0.002} # Add in time delays. Provide the constellation name in STK and the node delays
stkVersion = 12
overrideStrands = False # Override previously computed chains
overrideNodeDelaysByNode = False # Override previously built node delay dictionaries
overrideNodesTimesPos = False # Override previously built node positions
overrideNetwork = False # Override previously built node positions
# Connect to STK
stkApp = GetActiveObject('STK{}.Application'.format(stkVersion))
stkRoot = stkApp.Personality2
stkRoot.Isolate()
stkRoot.UnitPreferences.SetCurrentUnit('DateFormat','EpSec') # Units to EpSec for ease of use
stkRoot.ExecuteCommand('Units_SetConnect / Date "EpochSeconds"');
stkRoot.ExecuteCommand('VO * ObjectLine DeleteAll'); # Clean up old object lines
# Build chains and create a dict of time delays for each node
t1 = time.time()
chainNames = createDirectedChains(stkRoot,constellationOrderLists,start=start,stop=stop,color=12895232)
print(time.time()-t1)
t1 = time.time()
nodeDelaysByNode = getNodeDelaysByNode(stkRoot,nodeDelays,chainNames=chainNames,overrideData=overrideNodeDelaysByNode)
print(time.time()-t1)
```
## Compute Strands and Distances
```
# Compute strands
t1 = time.time()
strands,dfStrands = getAllStrands(stkRoot,chainNames,start,stop,overrideData=overrideStrands)
print(time.time()-t1)
dfStrands
# Compute node positions, distances and time delays
# Time resolution of distance/time computation
step = 10 # sec
t1 = time.time()
nodesTimesPos = computeNodesPosOverTime(stkRoot,strands,start,stop,step,overrideData=overrideNodesTimesPos) # Pull node position over time
t2 = time.time()
print(t2-t1)
t1 = time.time()
strandsAtTimes = getStrandsAtTimes(strands,start,stop,step) # Discretize strand intervals into times
t2 = time.time()
print(t2-t1)
t1 = time.time()
timeNodePos = computeTimeNodePos(strandsAtTimes,nodesTimesPos) # Nodes and positions at each time
t2 = time.time()
print(t2-t1)
t1 = time.time()
timesEdgesDistancesDelays = computeTimeEdgesDistancesDelays(strandsAtTimes,nodesTimesPos,nodeDelaysByNode,overrideData=True) # Edges, distances and delays at each time
t2 = time.time()
print(t2-t1)
```
## Use NX for Network Metrics and Reliability Analysis
```
# Get pairs of each starting and ending node permutation in the constellations
startingNodes = getNodesFromConstellation(stkRoot,startingConstellation)
endingNodes = getNodesFromConstellation(stkRoot,endingConstellation)
nodePairs = [(start,end) for start,end in itertools.product(startingNodes, endingNodes)] # full permutation
pd.DataFrame(nodePairs)
# Loop through each node pair and compute network metrics
# Edit computeNetworkMetrics in chainPathLibCustom to for additional metrics
for nodePair in nodePairs:
df = computeNetworkMetrics(start,stop,step,timeNodePos,timesEdgesDistancesDelays,[nodePair[0]],[nodePair[1]],metric,computeNumNodesToLoseAccessBetweenAnyPair=True,overrideData=overrideNetwork,printTime=False,diNetwork=True)
```
## Investigate Routing between Nodes
```
# Pick a starting and ending node
startingNode = 'Target/Target7'
endingNode = 'Place/Washington_DC'
# Load df
filename = 'SavedNetworkData/df{}{}.pkl'.format(startingNode.split('/')[-1],endingNode.split('/')[-1])
with open(filename,'rb') as f:
df = pickle.load(f)
df = addLightAndNodeDelays(df,timesEdgesDistancesDelays)
dfIntervals = createDfIntervals(df,stop,step)
addStrandsAsObjectLines(stkRoot,dfIntervals,color='yellow')
# Add data back into STK for reporting and plotting
t1 = time.time()
df['distance'] = df['distance']*1000 # May need to fix meter/kilometer issue
addDataToSTK(stkRoot,chainNames[0],df) # Adds data in df back into STK to the first chain under User Supplied data
print(time.time()-t1)
df
dfIntervals
# Active objects in the network over time
objPaths = list(set((item for sublist in df['strand'] for item in sublist)))
# Turn on the objects in the scenario
turnGraphicsOnOff(stkRoot,objPaths,onOrOff = 'On',parentsOnly = True)
# Turn off the objects in the sceario
turnGraphicsOnOff(stkRoot,objPaths,onOrOff = 'Off',parentsOnly = True)
```
## Investigate Instances in Time
```
# Look at an instance in time (pick a time in df)
t = 0
stkRoot.CurrentTime = t
# Look at strand order and the node delay
objPaths = df['strand'][t/step]
nodeDelaysByStrand = {node:nodeDelaysByNode[node] for node in objPaths}
pd.DataFrame([*nodeDelaysByStrand.items()],columns=['node','nodeDelay'])
#look at possible connections for the object of interest at that time
nodeInterest = objPaths[0]
possibleNodeConnections(t,nodeInterest,timesEdgesDistancesDelays)
# Turn on the objects in the scenario
turnGraphicsOnOff(stkRoot,objPaths,onOrOff = 'On',parentsOnly = False)
# Turn off the objects in the sceario
turnGraphicsOnOff(stkRoot,objPaths,onOrOff = 'Off',parentsOnly = False)
```
## Investigate Node Utilization
```
# Most frequnt node in the shortest path and the sum of their durations
strands = dfIntervals[['strand','start','stop']].values
dfNodesIntervals = getNodesIntervalsFromStrands(strands)
dfNodeActive = getActiveDuration(dfNodesIntervals,start,stop)
dfNodeActive.sort_values('sum dur',ascending=False).head(10)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import seaborn as sns
# ML Models
from sklearn.linear_model import LinearRegression
from sklearn.svm import SVR
from sklearn.ensemble import RandomForestRegressor
import itertools
```
### Loading data
```
# TODO: Test on merged data from 1st and 2nd iteration
users_data = pd.read_json("proper_data/users2.json")
deliveries_data = pd.read_json("proper_data/deliveries2.json")
events_data = pd.read_json("proper_data/sessions2.json")
products_data = pd.read_json("proper_data/products2.json")
users_data
deliveries_data
events_data
products_data.info()
```
## Preprocessing
## Extracting features
### "Indexing features"
```
events_data['year'] = events_data.timestamp.dt.year
events_data['month'] = events_data.timestamp.dt.month
years = list(events_data.year.unique())
months = list(events_data.month.unique())
user_ids = list(events_data.user_id.unique())
triplets = []
for triplet in itertools.product(years, months, user_ids):
triplets.append(triplet)
processed_data = pd.DataFrame(triplets, columns=['year', 'month', 'user_id'])
processed_data
```
### Number of all events per user per month
```
all_events = events_data.groupby(['user_id', 'year', 'month']).aggregate({"session_id": "count"}) \
.rename(columns={"session_id": "all_sessions"}) \
.reset_index()
```
### Number of buying events per user per month
```
buying_events = events_data[events_data['event_type'] == 'BUY_PRODUCT'] \
.groupby(['user_id', 'year', 'month']) \
.aggregate({"session_id": "count"}) \
.rename(columns={"session_id": "buying_sessions"}) \
.reset_index()
```
### Buying ratio
```
events_ratio = pd.merge(all_events, buying_events, how="left", on=["year", "month", "user_id"])
# all_events
# buying_events
events_ratio['buying_sessions'].fillna(0, inplace=True)
events_ratio['buying_ratio'] = round(events_ratio['buying_sessions'] / events_ratio['all_sessions'], 4)
# merging with processed_data
processed_data = pd.merge(processed_data, events_ratio.drop("all_sessions", axis=1), how='left', on=['year', 'month', 'user_id'])
processed_data['buying_sessions'].fillna(0, inplace=True)
processed_data['buying_ratio'].fillna(0, inplace=True)
processed_data
```
### Data Cleaner
```
def clean_customers(customers_data):
customers_data.info()
pass
def clean_products(products_data):
products_data.info()
pass
def clean_deliveries(deliveries_data):
info_df = deliveries_data.info()
if any([info_df[feature]['Non-Null Count'] != len(deliveries_data) for feature in info_df]):
print()
pass
def clean_events():
pass
```
### Constructing target variable - how much money the user spent per month
```
buying_sessions = events_data[events_data['event_type'] == "BUY_PRODUCT"]
deals = pd.merge(buying_sessions, products_data, how="left", on=['product_id'])
deals['final_price'] = deals['price'] * (1 - deals['offered_discount'] * 0.01)
monthly_deals = deals.groupby(['year', 'month', 'user_id']) \
.aggregate({"final_price": "sum"}) \
.rename(columns={"final_price": "money_monthly"}) \
.reset_index()
processed_data = pd.merge(processed_data, monthly_deals, how='left', on=['year', 'month', 'user_id'])
processed_data
processed_data['money_monthly'].fillna(0, inplace=True)
processed_data
```
| github_jupyter |
# <center>Value Function Approximation</center>
## <center>Part II</center>
## <center>Reference: Sutton and Barto, Chapter 9-11</center>
## <center>Table of Contents</center>
<br>
* **Batch Reinforcement Methods**<br><br>
* **Least Squares Policy Iteration(LSPI)**<br><br>
# <center>Batch Reinforcement Methods</center>
## <center>Batch Reinforcement Methods</center>
<br>
* Gradient descent is simple and appealing<br><br>
* But it is not sample efficient<br><br>
* Batch methods seek to find the best fitting value function<br><br>
* Given the agent’s experience (“training data”)<br><br>
## <center>Least Squares Prediction</center>
<center><img src="img/fa2_slides1.JPG" alt="Multi-armed Bandit" style="width: 600px;"/></center>
## <center>Stochastic Gradient Descent with Experience Replay</center>
<center><img src="img/fa2_slides3.JPG" alt="Multi-armed Bandit" style="width: 600px;"/></center>
## <center>Experience Replay in Deep Q-Networks (DQN)</center>
<center><img src="img/fa2_slides4.JPG" alt="Multi-armed Bandit" style="width: 600px;"/></center>
# <center>DQN in ATARI</center>
## The model
<center><img src="img/fa2_ex1.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## Performance
<center><img src="img/fa2_ex2.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## Benefits of Experience Replay and Double DQN
<center><img src="img/fa2_ex3.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## DQN Example and Code
<center><img src="img/ex.png" alt="Multi-armed Bandit" style="width: 300px;"/></center>
#### CartPole Example
The agent has to decide between two actions - moving the cart left or right - so that the pole attached to it stays upright.
##### State Space
State is the difference between the current screen patch and the previous one. This will allow the agent to take the velocity of the pole into account from one image.
##### Q-network
* Our model will be a convolutional neural network that takes in the difference between the current and previous screen patches.
* It has two outputs, representing Q(s,left) and Q(s,right) (where s is the input to the network).
* In effect, the network is trying to predict the quality of taking each action given the current input.
<center><img src="img/2.png" alt="Multi-armed Bandit" style="width: 600px;"/></center>
##### Replay Memory
* Experience replay memory is used for training the DQN.
* It stores the transitions that the agent observes, allowing us to reuse this data later.
* By sampling from it randomly, the transitions that build up a batch are decorrelated.
* It has been shown that this greatly stabilizes and improves the DQN training procedure.
<center><img src="img/1.png" alt="Multi-armed Bandit" style="width: 600px;"/></center>
##### Input Extraction
How do we get the crop of the cart?
<center><img src="img/3.png" alt="Multi-armed Bandit" style="width: 600px;"/></center>
##### Selecting an Action
This is done based on $\epsilon$ greedy policy.
<center><img src="img/4.png" alt="Multi-armed Bandit" style="width: 600px;"/></center>
##### Training
<center><img src="img/5.png" alt="Multi-armed Bandit" style="width: 600px;"/></center>
<center><img src="img/6.png" alt="Multi-armed Bandit" style="width: 600px;"/></center>
<center><img src="img/7.png" alt="Multi-armed Bandit" style="width: 600px;"/></center>
# <center>Linear Least Squares Prediction</center>
## <center>Linear Least Squares Prediction</center>
<br><br>
* Experience replay finds least squares solution<br><br>
* But it may take many iterations<br><br>
* Using linear value function approximation $\hat{v}(s, w) = x(s)^Tw$<br><br>
* We can solve the least squares solution directly
## <center>Linear Least Squares Prediction</center>
<center><img src="img/fa2_slides5.JPG" alt="Multi-armed Bandit" style="width: 600px;"/></center>
## <center>Linear Least Squares Prediction Algorithms</center>
<center><img src="img/fa2_slides6.JPG" alt="Multi-armed Bandit" style="width: 600px;"/></center>
## <center>Linear Least Squares Prediction Algorithms</center>
<center><img src="img/fa2_slides7.JPG" alt="Multi-armed Bandit" style="width: 600px;"/></center>
## <center>Least Squares Policy Iteration(LSPI)</center>
<center><img src="img/fa2_slides8.JPG" alt="Multi-armed Bandit" style="width: 600px;"/></center>
## <center>Least Squares Action-Value Function Approximation</center>
<center><img src="img/fa2_slides9.JPG" alt="Multi-armed Bandit" style="width: 600px;"/></center>
## <center>Least Squares Control</center>
<center><img src="img/fa2_slides10.JPG" alt="Multi-armed Bandit" style="width: 600px;"/></center>
## <center>Least Squares Q-Learning</center>
<center><img src="img/fa2_slides11.JPG" alt="Multi-armed Bandit" style="width: 600px;"/></center>
## <center>Least Squares Policy Iteration(LSPI) Algorithm</center>
<center><img src="img/fa2_slides12.JPG" alt="Multi-armed Bandit" style="width: 600px;"/></center>
| github_jupyter |
```
%pylab inline
from solveMDP_richLow import *
Vgrid = np.load("richLow.npy")
matplotlib.rcParams['figure.figsize'] = [16, 8]
plt.rcParams.update({'font.size': 15})
# wealth discretization
wealthLevel = 300
polynomialDegree = 1
ws = jnp.linspace(0, np.power(wealthLevel,1/polynomialDegree), 100)**polynomialDegree
intendedBuyer = []
ages = []
for age in range(20, 50):
for w in ws:
t = age - 20
# x = [w,ab,s,e,o,z]
x = [w, 0, 0, 1, 0, 1]
_,a = V_solve(t,Vgrid[:,:,:,:,:,:,t+1],x)
if a[4] == 1:
ages.append(age)
intendedBuyer.append(w)
break
plt.plot(ages, intendedBuyer,'*-')
intendedBuyer
'''
Policy plot:
Input:
x = [w,ab,s,e,o,z] single action
x = [0,1, 2,3,4,5]
a = [c,b,k,h,action] single state
a = [0,1,2,3,4]
'''
wealthLevel = [100, 150, 200, 250]
ageLevel = [30, 45, 60, 75]
savingsRatio = []
investmentsRatio = []
for wealth in wealthLevel:
savingR = []
investmentR = []
for age in ageLevel:
t = age - 20
x = [wealth, 0, 4, 1, 0, 1]
_,a = V_solve(t,Vgrid[:,:,:,:,:,:,t+1],x)
savingR.append((a[1]+a[2])/wealth)
investmentR.append(a[2]/(a[1]+a[2]))
savingsRatio.append(savingR)
investmentsRatio.append(investmentR)
import pandas as pd
df_saving = pd.DataFrame(np.array(savingsRatio), columns = ['age '+ str(age) for age in ageLevel], index= ['wealth ' + str(wealth) for wealth in wealthLevel])
df_investment = pd.DataFrame(np.array(investmentsRatio), columns = ['age '+ str(age) for age in ageLevel], index= ['wealth ' + str(wealth) for wealth in wealthLevel])
print("savingRatio:")
display(df_saving)
print("investmentRatio:")
display(df_investment)
```
### Simulation Part
```
%%time
# total number of agents
num = 10000
'''
x = [w,ab,s,e,o,z]
x = [5,0, 0,0,0,0]
'''
from jax import random
from quantecon import MarkovChain
# number of economies and each economy has 100 agents
numEcon = 100
numAgents = 100
mc = MarkovChain(Ps)
econStates = mc.simulate(ts_length=T_max-T_min,init=0,num_reps=numEcon)
econStates = jnp.array(econStates,dtype = int)
@partial(jit, static_argnums=(0,))
def transition_real(t,a,x, s_prime):
'''
Input:
x = [w,ab,s,e,o,z] single action
x = [0,1, 2,3,4,5]
a = [c,b,k,h,action] single state
a = [0,1,2,3,4]
Output:
w_next
ab_next
s_next
e_next
o_next
z_next
prob_next
'''
s = jnp.array(x[2], dtype = jnp.int8)
e = jnp.array(x[3], dtype = jnp.int8)
# actions taken
b = a[1]
k = a[2]
action = a[4]
w_next = ((1+r_b[s])*b + (1+r_k[s_prime])*k).repeat(nE)
ab_next = (1-x[4])*(t*(action == 1)).repeat(nE) + x[4]*(x[1]*jnp.ones(nE))
s_next = s_prime.repeat(nE)
e_next = jnp.array([e,(1-e)])
z_next = x[5]*jnp.ones(nE) + ((1-x[5]) * (k > 0)).repeat(nE)
# job status changing probability and econ state transition probability
pe = Pe[s, e]
prob_next = jnp.array([1-pe, pe])
# owner
o_next_own = (x[4] - action).repeat(nE)
# renter
o_next_rent = action.repeat(nE)
o_next = x[4] * o_next_own + (1-x[4]) * o_next_rent
return jnp.column_stack((w_next,ab_next,s_next,e_next,o_next,z_next,prob_next))
def simulation(key):
initE = random.choice(a = nE, p=E_distribution, key = key)
initS = random.choice(a = nS, p=S_distribution, key = key)
x = [5, 0, initS, initE, 0, 0]
path = []
move = []
# first 100 agents are in the 1st economy and second 100 agents are in the 2nd economy
econ = econStates[key.sum()//numAgents,:]
for t in range(T_min, T_max):
_, key = random.split(key)
if t == T_max-1:
_,a = V_solve(t,Vgrid[:,:,:,:,:,:,t],x)
else:
_,a = V_solve(t,Vgrid[:,:,:,:,:,:,t+1],x)
xp = transition_real(t,a,x, econ[t])
p = xp[:,-1]
x_next = xp[:,:-1]
path.append(x)
move.append(a)
x = x_next[random.choice(a = nE, p=p, key = key)]
path.append(x)
return jnp.array(path), jnp.array(move)
# simulation part
keys = vmap(random.PRNGKey)(jnp.arange(num))
Paths, Moves = vmap(simulation)(keys)
# x = [w,ab,s,e,o,z]
# x = [0,1, 2,3,4,5]
ws = Paths[:,:,0].T
ab = Paths[:,:,1].T
ss = Paths[:,:,2].T
es = Paths[:,:,3].T
os = Paths[:,:,4].T
zs = Paths[:,:,5].T
cs = Moves[:,:,0].T
bs = Moves[:,:,1].T
ks = Moves[:,:,2].T
hs = Moves[:,:,3].T
ms = Ms[jnp.append(jnp.array([0]),jnp.arange(T_max)).reshape(-1,1) - jnp.array(ab, dtype = jnp.int8)]*os
```
### Graph and Table
```
plt.figure(1)
plt.title("The mean values of simulation")
startAge = 20
# value of states, state has one more value, since the terminal state does not have associated action
plt.plot(range(startAge, T_max + startAge + 1),jnp.mean(ws + H*pt*os - ms,axis = 1), label = "wealth + home equity")
plt.plot(range(startAge, T_max + startAge + 1),jnp.mean(H*pt*os - ms,axis = 1), label = "home equity")
plt.plot(range(startAge, T_max + startAge + 1),jnp.mean(ws,axis = 1), label = "wealth")
# value of actions
plt.plot(range(startAge, T_max + startAge),jnp.mean(cs,axis = 1), label = "consumption")
plt.plot(range(startAge, T_max + startAge),jnp.mean(bs,axis = 1), label = "bond")
plt.plot(range(startAge, T_max + startAge),jnp.mean(ks,axis = 1), label = "stock")
plt.legend()
plt.figure(2)
plt.title("Stock Participation Ratio through Different Age Periods")
plt.plot(range(20, T_max + 21),jnp.mean(zs,axis = 1), label = "experience")
plt.legend()
plt.figure(3)
plt.title("house ownership ratio in the population")
plt.plot(range(startAge, T_max + startAge + 1),(os).mean(axis = 1), label = "ownership ratio")
plt.legend()
# agent buying time collection
agentTime = []
for t in range(30):
if ((os[t,:] == 0) & (os[t+1,:] == 1)).sum()>0:
for agentNum in jnp.where((os[t,:] == 0) & (os[t+1,:] == 1))[0]:
agentTime.append([t, agentNum])
agentTime = jnp.array(agentTime)
# agent hold time collection
agentHold = []
for t in range(30):
if ((os[t,:] == 0) & (os[t+1,:] == 0)).sum()>0:
for agentNum in jnp.where((os[t,:] == 0) & (os[t+1,:] == 0))[0]:
agentHold.append([t, agentNum])
agentHold = jnp.array(agentHold)
plt.figure(4)
plt.title("weath level for buyer, owner and renter")
www = (os*(ws+H*pt - ms)).sum(axis = 1)/(os).sum(axis = 1)
for age in range(30):
buyer = agentTime[agentTime[:,0] == age]
renter = agentHold[agentHold[:,0] == age]
bp = plt.scatter(age, ws[buyer[:,0], buyer[:,1]].mean(),color = "b")
hp = plt.scatter(age, www[age], color = "green")
rp = plt.scatter(age, ws[renter[:,0], renter[:,1]].mean(),color = "r")
plt.legend((bp,hp,rp), ("FirstTimeBuyer", "HomeOwner", "Renter"))
plt.figure(5)
plt.title("employement status for buyer and renter")
for age in range(31):
buyer = agentTime[agentTime[:,0] == age]
renter = agentHold[agentHold[:,0] == age]
bp = plt.scatter(age, es[buyer[:,0], buyer[:,1]].mean(),color = "b")
rp = plt.scatter(age, es[renter[:,0], renter[:,1]].mean(),color = "r")
plt.legend((bp, rp), ("FirstTimeBuyer", "Renter"))
# agent participate time collection
agentTimep = []
for t in range(30):
if ((zs[t,:] == 0) & (zs[t+1,:] == 1)).sum()>0:
for agentNum in jnp.where((zs[t,:] == 0) & (zs[t+1,:] == 1))[0]:
agentTimep.append([t, agentNum])
agentTimep = jnp.array(agentTimep)
# agent nonparticipate time collection
agentHoldp = []
for t in range(30):
if ((zs[t,:] == 0) & (zs[t+1,:] == 0)).sum()>0:
for agentNum in jnp.where((zs[t,:] == 0) & (zs[t+1,:] == 0))[0]:
agentHoldp.append([t, agentNum])
agentHoldp = jnp.array(agentHoldp)
plt.figure(6)
plt.title("weath level for FirstTimeTrader, ExperiencedTrader and Nonparticipant")
www = (zs*(ws+H*pt - ms)).sum(axis = 1)/(zs).sum(axis = 1)
for age in range(30):
trader = agentTimep[agentTimep[:,0] == age]
noneTrader = agentHoldp[agentHoldp[:,0] == age]
tp = plt.scatter(age, ws[trader[:,0], trader[:,1]].mean(),color = "b")
ep = plt.scatter(age, www[age], color = "green")
ip = plt.scatter(age, ws[noneTrader[:,0], noneTrader[:,1]].mean(),color = "r")
plt.legend((tp,ep,ip), ("FirstTimeTrader", "ExperiencedTrader", "Nonparticipant"))
plt.figure(7)
plt.title("employement status for FirstTimeTrader and Nonparticipant")
for age in range(30):
trader = agentTimep[agentTimep[:,0] == age]
noneTrader = agentHoldp[agentHoldp[:,0] == age]
tp = plt.scatter(age, es[trader[:,0], trader[:,1]].mean(),color = "b", label = "FirstTimeTrader")
ip = plt.scatter(age, es[noneTrader[:,0], noneTrader[:,1]].mean(),color = "r", label = "Nonparticipant")
plt.legend((tp,ip), ("FirstTimeTrader", "Nonparticipant"))
plt.figure(8)
# At every age
plt.title("Stock Investment Percentage as StockInvestmentAmount/(StockInvestmentAmount + BondInvestmentAmount)")
plt.plot((os[:T_max,:]*ks/(ks+bs)).sum(axis = 1)/os[:T_max,:].sum(axis = 1), label = "owner")
plt.plot(((1-os[:T_max,:])*ks/(ks+bs)).sum(axis = 1)/(1-os)[:T_max,:].sum(axis = 1), label = "renter")
plt.legend()
plt.figure(9)
# At every age
plt.title("Stock Investment Percentage")
plt.plot(range(startAge, startAge+T_max), (ks/(ks+bs)).mean(axis = 1), label = "ks/(ks+bs)")
plt.legend()
# # agent number, x = [w,n,m,s,e,o]
# agentNum = 35
# plt.plot(range(20, T_max + 21),(ws + os*(H*pt - ms))[:,agentNum], label = "wealth + home equity")
# plt.plot(range(20, T_max + 21),ms[:,agentNum], label = "mortgage")
# plt.plot(range(20, T_max + 20),cs[:,agentNum], label = "consumption")
# plt.plot(range(20, T_max + 20),bs[:,agentNum], label = "bond")
# plt.plot(range(20, T_max + 20),ks[:,agentNum], label = "stock")
# plt.plot(range(20, T_max + 21),os[:,agentNum]*100, label = "ownership", color = "k")
# plt.legend()
ws.mean(axis = 1)
```
| github_jupyter |
```
import pandas as pd
df = pd.read_csv("data.csv")
df
```
## Let's get a sense of what our dataframe looks like
```
df.columns
df["Total Working Population Percentage"] = df["Total Working Pop. (Age 16+) (2010)"] / (df["Population (2010)"] + df["Total Working Pop. (Age 16+) (2010)"])
df
```
### In the above cell, we obtain the ratio of the working population to the total population in each area. Let's see who has the highest....
```
df.sort_values(by="Total Working Population Percentage", ascending=False, inplace=True)
df
```
### Let's evaluate each area in Pittsburgh based on how how "athletic" their working population is.
### Let's award 10 points to those who commute to work via Taxi/Carpool/Vanpool/Other,
### 20 points to those who commute to work via Motorcycle, 40 points via Bicycle, and 60 points via walking
###
### Let's see the most "athletic" regions in Pittsburgh....
```
df["Athletic Score"] = 10 * df["Work at Home (2010)"]
+ 10 * df["Commute to Work: Other (2010)"]
+ 10 * df["Commute to Work: Taxi (2010)"]
+ 10 * df["Commute to Work: Carpool/Vanpool (2010)"]
+ 10 * df["Commute to Work: Public Transportation (2010)"]
+ 20 * df["Commute to Work: Motorcycle (2010)"]
+ 40 * df["Commute to Work: Bicycle (2010)"]
+ 60 * df["Commute to Work: Walk (2010)"]
df.sort_values(by="Athletic Score", inplace=True, ascending=False)
# df = df[["Neighborhood", "Athletic Score"]]
df.dropna(inplace=True)
df[["Neighborhood", "Athletic Score"]]
```
### Here's a quick summary of the "Athletic Score" statistic throughout Pittsburgh
```
df["Athletic Score"].plot.box()
```
### Looks like there's a clear winner
```
import geopandas
%matplotlib inline
import matplotlib.pyplot as plt
df[["Neighborhood", "Athletic Score"]]
```
### Fairywood blows everyone else out of the waters.... It isn't particularly close either.
```
# # import dataset
# steps = pd.read_csv("steps.csv")
# # filter to important info
# num_steps = steps.groupby("neighborhood").sum()['number_of_steps']
# num_steps.sort_values(ascending=False)
# # do the merge
# steps_map = neighborhoods.merge(num_steps, how='left', left_on='hood', right_on='neighborhood')
# # look at the head to confirm it merged correctly
# steps_map[['hood','number_of_steps','geometry']].head()
neighborhoods = geopandas.read_file("./Neighborhoods/Neighborhoods_.shp")
neighborhoods.plot()
# steps_map.plot(column='number_of_steps')
result = neighborhoods.merge(df, how='left', left_on='hood', right_on='Neighborhood')
result[["Neighborhood", "Athletic Score"]]
result.plot(column='Athletic Score', # set the data to be used for coloring
cmap='OrRd', # choose a color palette
edgecolor="white", # outline the districts in white
legend=True, # show the legend
legend_kwds={'label': "Number of Steps"}, # label the legend
figsize=(15, 10), # set the size
missing_kwds={"color": "lightgrey"} # set disctricts with no data to gray
)
```
| github_jupyter |
<img align="center" style="max-width: 1000px" src="banner.png">
<img align="right" style="max-width: 200px; height: auto" src="hsg_logo.png">
## Lab 05 - Convolutional Neural Networks (CNNs)
EMBA 60 W10 / EMBA 61 W5: Coding und Künstliche Intelligenz, University of St. Gallen
The lab environment of the "Coding und Künstliche Intelligenz" EMBA course at the University of St. Gallen (HSG) is based on Jupyter Notebooks (https://jupyter.org), which allow to perform a variety of statistical evaluations and data analyses.
In this lab, we will learn how to enhance vanilla Artificial Neural Networks (ANNs) using `PyTorch` to classify even more complex images. Therefore, we use a special type of deep neural network referred to **Convolutional Neural Networks (CNNs)**. CNNs encompass the ability to take advantage of the hierarchical pattern in data and assemble more complex patterns using smaller and simpler patterns. Therefore, CNNs are capable to learn a set of discriminative features 'pattern' and subsequently utilize the learned pattern to classify the content of an image.
We will again use the functionality of the `PyTorch` library to implement and train an CNN based neural network. The network will be trained on a set of tiny images to learn a model of the image content. Upon successful training, we will utilize the learned CNN model to classify so far unseen tiny images into distinct categories such as aeroplanes, cars, birds, cats, deer, dogs, frogs, horses, ships, and trucks.
The figure below illustrates a high-level view on the machine learning process we aim to establish in this lab.
<img align="center" style="max-width: 900px" src="https://raw.githubusercontent.com/HSG-AIML-Teaching/EMBA2022-Lab/main/lab_05/classification.png">
(Image of the CNN architecture created via http://alexlenail.me/)
As always, pls. don't hesitate to ask all your questions either during the lab, post them in our CANVAS (StudyNet) forum (https://learning.unisg.ch), or send us an email (using the course email).
## 1. Lab Objectives:
After today's lab, you should be able to:
> 1. Understand the basic concepts, intuitions and major building blocks of **Convolutional Neural Networks (CNNs)**.
> 2. Know how to **implement and to train a CNN** to learn a model of tiny image data.
> 3. Understand how to apply such a learned model to **classify images** images based on their content into distinct categories.
> 4. Know how to **interpret and visualize** the model's classification results.
## 2. Setup of the Jupyter Notebook Environment
Similar to the previous labs, we need to import a couple of Python libraries that allow for data analysis and data visualization. We will mostly use the `PyTorch`, `Numpy`, `Sklearn`, `Matplotlib`, `Seaborn` and a few utility libraries throughout this lab:
```
# import standard python libraries
import os, urllib, io
from datetime import datetime
import numpy as np
```
Import Python machine / deep learning libraries:
```
# import the PyTorch deep learning library
import torch, torchvision
import torch.nn.functional as F
from torch import nn, optim
from torch.autograd import Variable
```
Import the sklearn classification metrics:
```
# import sklearn classification evaluation library
from sklearn import metrics
from sklearn.metrics import classification_report, confusion_matrix
```
Import Python plotting libraries:
```
# import matplotlib, seaborn, and PIL data visualization libary
import matplotlib.pyplot as plt
import seaborn as sns
from PIL import Image
```
Enable notebook matplotlib inline plotting:
```
%matplotlib inline
```
Import Google's GDrive connector and mount your GDrive directories:
```
# import the Google Colab GDrive connector
from google.colab import drive
# mount GDrive inside the Colab notebook
drive.mount('/content/drive')
```
Create a structure of Colab Notebook sub-directories inside of GDrive to store (1) the data as well as (2) the trained neural network models:
```
# create Colab Notebooks directory
notebook_directory = '/content/drive/MyDrive/Colab Notebooks'
if not os.path.exists(notebook_directory): os.makedirs(notebook_directory)
# create data sub-directory inside the Colab Notebooks directory
data_directory = '/content/drive/MyDrive/Colab Notebooks/data_cifar10'
if not os.path.exists(data_directory): os.makedirs(data_directory)
# create models sub-directory inside the Colab Notebooks directory
models_directory = '/content/drive/MyDrive/Colab Notebooks/models_cifar10'
if not os.path.exists(models_directory): os.makedirs(models_directory)
```
Set a random `seed` value to obtain reproducible results:
```
# init deterministic seed
seed_value = 1234
np.random.seed(seed_value) # set numpy seed
torch.manual_seed(seed_value) # set pytorch seed CPU
```
Google Colab provides the use of free GPUs for running notebooks. However, if you just execute this notebook as is, it will use your device's CPU. To run the lab on a GPU, got to `Runtime` > `Change runtime type` and set the Runtime type to `GPU` in the drop-down. Running this lab on a CPU is fine, but you will find that GPU computing is faster. *CUDA* indicates that the lab is being run on GPU.
Enable GPU computing by setting the `device` flag and init a `CUDA` seed:
```
# set cpu or gpu enabled device
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu').type
# init deterministic GPU seed
torch.cuda.manual_seed(seed_value)
# log type of device enabled
print('[LOG] notebook with {} computation enabled'.format(str(device)))
```
Let's determine if we have access to a GPU provided by e.g. Google's COLab environment:
```
!nvidia-smi
```
## 3. Dataset Download and Data Assessment
The **CIFAR-10 database** (**C**anadian **I**nstitute **F**or **A**dvanced **R**esearch) is a collection of images that are commonly used to train machine learning and computer vision algorithms. The database is widely used to conduct computer vision research using machine learning and deep learning methods:
<img align="center" style="max-width: 500px; height: 500px" src="https://raw.githubusercontent.com/HSG-AIML-Teaching/EMBA2022-Lab/main/lab_05/cifar10.png">
(Source: https://www.kaggle.com/c/cifar-10)
Further details on the dataset can be obtained via: *Krizhevsky, A., 2009. "Learning Multiple Layers of Features from Tiny Images",
( https://www.cs.toronto.edu/~kriz/learning-features-2009-TR.pdf )."*
The CIFAR-10 database contains **60,000 color images** (50,000 training images and 10,000 validation images). The size of each image is 32 by 32 pixels. The collection of images encompasses 10 different classes that represent airplanes, cars, birds, cats, deer, dogs, frogs, horses, ships, and trucks. Let's define the distinct classs for further analytics:
```
cifar10_classes = ['plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
```
Thereby the dataset contains 6,000 images for each of the ten classes. The CIFAR-10 is a straightforward dataset that can be used to teach a computer how to recognize objects in images.
Let's download, transform and inspect the training images of the dataset. Therefore, we first will define the directory we aim to store the training data:
```
train_path = data_directory + '/train_cifar10'
```
Now, let's download the training data accordingly:
```
# define pytorch transformation into tensor format
transf = torchvision.transforms.Compose([torchvision.transforms.ToTensor(), torchvision.transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
# download and transform training images
cifar10_train_data = torchvision.datasets.CIFAR10(root=train_path, train=True, transform=transf, download=True)
```
Verify the volume of training images downloaded:
```
# get the length of the training data
len(cifar10_train_data)
```
Furthermore, let's investigate a couple of the training images:
```
# set (random) image id
image_id = 1800
# retrieve image exhibiting the image id
cifar10_train_data[image_id]
```
Ok, that doesn't seem easily interpretable ;) Let's first seperate the image from its label information:
```
cifar10_train_image, cifar10_train_label = cifar10_train_data[image_id]
```
Great, now we are able to visually inspect our sample image:
```
# define tensor to image transformation
trans = torchvision.transforms.ToPILImage()
# set image plot title
plt.title('Example: {}, Label: "{}"'.format(str(image_id), str(cifar10_classes[cifar10_train_label])))
# un-normalize cifar 10 image sample
cifar10_train_image_plot = cifar10_train_image / 2.0 + 0.5
# plot 10 image sample
plt.imshow(trans(cifar10_train_image_plot))
```
Fantastic, right? Let's now decide on where we want to store the evaluation data:
```
eval_path = data_directory + '/eval_cifar10'
```
And download the evaluation data accordingly:
```
# define pytorch transformation into tensor format
transf = torchvision.transforms.Compose([torchvision.transforms.ToTensor(), torchvision.transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
# download and transform validation images
cifar10_eval_data = torchvision.datasets.CIFAR10(root=eval_path, train=False, transform=transf, download=True)
```
Verify the volume of validation images downloaded:
```
# get the length of the training data
len(cifar10_eval_data)
```
## 4. Neural Network Implementation
In this section we, will implement the architecture of the **neural network** we aim to utilize to learn a model that is capable of classifying the 32x32 pixel CIFAR 10 images according to the objects contained in each image. However, before we start the implementation, let's briefly revisit the process to be established. The following cartoon provides a birds-eye view:
<img align="center" style="max-width: 900px" src="https://raw.githubusercontent.com/HSG-AIML-Teaching/EMBA2022-Lab/main/lab_05/process.png">
Our CNN, which we name 'CIFAR10Net' and aim to implement consists of two **convolutional layers** and three **fully-connected layers**. In general, convolutional layers are specifically designed to learn a set of **high-level features** ("patterns") in the processed images, e.g., tiny edges and shapes. The fully-connected layers utilize the learned features to learn **non-linear feature combinations** that allow for highly accurate classification of the image content into the different image classes of the CIFAR-10 dataset, such as, birds, aeroplanes, horses.
Let's implement the network architecture and subsequently have a more in-depth look into its architectural details:
```
# implement the CIFAR10Net network architecture
class CIFAR10Net(nn.Module):
# define the class constructor
def __init__(self):
# call super class constructor
super(CIFAR10Net, self).__init__()
# specify convolution layer 1
self.conv1 = nn.Conv2d(in_channels=3, out_channels=6, kernel_size=5, stride=1, padding=0)
# define max-pooling layer 1
self.pool1 = nn.MaxPool2d(kernel_size=2, stride=2)
# specify convolution layer 2
self.conv2 = nn.Conv2d(in_channels=6, out_channels=16, kernel_size=5, stride=1, padding=0)
# define max-pooling layer 2
self.pool2 = nn.MaxPool2d(kernel_size=2, stride=2)
# specify fc layer 1 - in 16 * 5 * 5, out 120
self.linear1 = nn.Linear(16 * 5 * 5, 120, bias=True) # the linearity W*x+b
self.relu1 = nn.ReLU(inplace=True) # the non-linearity
# specify fc layer 2 - in 120, out 84
self.linear2 = nn.Linear(120, 84, bias=True) # the linearity W*x+b
self.relu2 = nn.ReLU(inplace=True) # the non-linarity
# specify fc layer 3 - in 84, out 10
self.linear3 = nn.Linear(84, 10) # the linearity W*x+b
# add a softmax to the last layer
self.logsoftmax = nn.LogSoftmax(dim=1) # the softmax
# define network forward pass
def forward(self, images):
# high-level feature learning via convolutional layers
# define conv layer 1 forward pass
x = self.pool1(self.relu1(self.conv1(images)))
# define conv layer 2 forward pass
x = self.pool2(self.relu2(self.conv2(x)))
# feature flattening
# reshape image pixels
x = x.view(-1, 16 * 5 * 5)
# combination of feature learning via non-linear layers
# define fc layer 1 forward pass
x = self.relu1(self.linear1(x))
# define fc layer 2 forward pass
x = self.relu2(self.linear2(x))
# define layer 3 forward pass
x = self.logsoftmax(self.linear3(x))
# return forward pass result
return x
```
You may have noticed that we applied two more layers (compared to the MNIST example described in the last lab) before the fully-connected layers. These layers are referred to as **convolutional** layers and are usually comprised of three operations, (1) **convolution**, (2) **non-linearity**, and (3) **max-pooling**. Those operations are usually executed in sequential order during the forward pass through a convolutional layer.
In the following, we will have a detailed look into the functionality and number of parameters in each layer. We will start with providing images of 3x32x32 dimensions to the network, i.e., the three channels (red, green, blue) of an image each of size 32x32 pixels.
### 4.1. High-Level Feature Learning by Convolutional Layers
Let's first have a look into the convolutional layers of the network as illustrated in the following:
<img align="center" style="max-width: 600px" src="https://raw.githubusercontent.com/HSG-AIML-Teaching/EMBA2022-Lab/main/lab_05/convolutions.png">
**First Convolutional Layer**: The first convolutional layer expects three input channels and will convolve six filters each of size 3x5x5. Let's briefly revisit how we can perform a convolutional operation on a given image. For that, we need to define a kernel which is a matrix of size 5x5, for example. To perform the convolution operation, we slide the kernel along with the image horizontally and vertically and obtain the dot product of the kernel and the pixel values of the image inside the kernel ('receptive field' of the kernel).
The following illustration shows an example of a discrete convolution:
<img align="center" style="max-width: 800px" src="https://raw.githubusercontent.com/HSG-AIML-Teaching/EMBA2022-Lab/main/lab_05/convsample.png">
The left grid is called the input (an image or feature map). The middle grid, referred to as kernel, slides across the input feature map (or image). At each location, the product between each element of the kernel and the input element it overlaps is computed, and the results are summed up to obtain the output in the current location. In general, a discrete convolution is mathematically expressed by:
<center> $y(m, n) = x(m, n) * h(m, n) = \sum^{m}_{j=0} \sum^{n}_{i=0} x(i, j) * h(m-i, n-j)$, </center>
where $x$ denotes the input image or feature map, $h$ the applied kernel, and, $y$ the output.
When performing the convolution operation the 'stride' defines the number of pixels to pass at a time when sliding the kernel over the input. While 'padding' adds the number of pixels to the input image (or feature map) to ensure that the output has the same shape as the input. Let's have a look at another animated example:
<img align="center" style="max-width: 800px" src="https://raw.githubusercontent.com/HSG-AIML-Teaching/EMBA2022-Lab/main/lab_05/convsample_animated.gif">
(Source: https://towardsdatascience.com/a-comprehensive-guide-to-convolutional-neural-networks-the-eli5-way-3bd2b1164a53)
In our implementation padding is set to 0 and stride is set to 1. As a result, the output size of the convolutional layer becomes 6x28x28, since (32 input pixel - 5 kernel pixel) + 1 stride pixel = 28. This layer exhibits ((5 kernel pixel x 5 kernel pixel x 3 input channels) + 1 stride pixel) x 6 output channels = 456 parameter.
**First Max-Pooling Layer:** The max-pooling process is a sample-based discretization operation. The objective is to down-sample an input representation (image, hidden-layer output matrix, etc.), reducing its dimensionality and allowing for assumptions to be made about features contained in the sub-regions binned. To conduct such an operation, we again need to define a kernel. Max-pooling kernels are usually a tiny matrix of, e.g, of size 2x2. To perform the max-pooling operation, we slide the kernel along the image horizontally and vertically (similarly to a convolution) and compute the maximum pixel value of the image (or feature map) inside the kernel (the receptive field of the kernel).
The following illustration shows an example of a max-pooling operation:
<img align="center" style="max-width: 500px" src="https://raw.githubusercontent.com/HSG-AIML-Teaching/EMBA2022-Lab/main/lab_05/poolsample.png">
The left grid is called the input (an image or feature map). The middle grid, referred to as kernel, slides across the input feature map (or image). We use a stride of 2, meaning the step distance for stepping over our input will be 2 pixels and won't overlap regions. At each location, the max value of the region that overlaps with the elements of the kernel and the input elements it overlaps is computed, and the results are obtained in the output of the current location.
In our implementation, we do max-pooling with a 2x2 kernel and stride 2 this effectively drops the original image size from 6x28x28 to 6x14x14. Let's have a look at an exemplary visualization of 64 features learnt in the first convolutional layer on the CIFAR- 10 dataset:
<img align="center" style="max-width: 700px" src="https://raw.githubusercontent.com/HSG-AIML-Teaching/EMBA2022-Lab/main/lab_05/cnnfeatures.png">
(Source: Yu, Dingjun, Hanli Wang, Peiqiu Chen, and Zhihua Wei, **"Mixed Pooling for Convolutional Neural Networks"**, International Conference on Rough Sets and Knowledge Technology, pp. 364-375. Springer, Cham, 2014)
**Second Convolutional Layer:** The second convolutional layer expects 6 input channels and will convolve 16 filters each of size 6x5x5x. Since padding is set to 0 and stride is set 1, the output size is 16x10x10, since (14 input pixel - 5 kernel pixel) + 1 stride pixel = 10. This layer therefore has ((5 kernel pixel x 5 kernel pixel x 6 input channels) + 1 stride pixel x 16 output channels) = 2,416 parameter.
**Second Max-Pooling Layer:** The second down-sampling layer uses max-pooling with 2x2 kernel and stride set to 2. This effectively drops the size from 16x10x10 to 16x5x5.
### 4.2. Flattening of Learned Features
The output of the final-max pooling layer needs to be flattened so that we can connect it to a fully connected layer. This is achieved using the `torch.Tensor.view` method. Setting the parameter of the method to `-1` will automatically infer the number of rows required to handle the mini-batch size of the data.
### 4.3. Learning of Feature Classification
Let's now have a look into the non-linear layers of the network illustrated in the following:
<img align="center" style="max-width: 600px" src="https://raw.githubusercontent.com/HSG-AIML-Teaching/EMBA2022-Lab/main/lab_05/fullyconnected.png">
The first fully connected layer uses 'Rectified Linear Units' (ReLU) activation functions to learn potential nonlinear combinations of features. The layers are implemented similarly to the fifth lab. Therefore, we will only focus on the number of parameters of each fully-connected layer:
**First Fully-Connected Layer:** The first fully-connected layer consists of 120 neurons, thus in total exhibits ((16 input channel x 5 input pixel x 5 input pixel) + 1 bias) x 120 input neurons = 48,120 parameter.
**Second Fully-Connected Layer:** The output of the first fully-connected layer is then transferred to second fully-connected layer. The layer consists of 84 neurons equipped with ReLu activation functions, this in total exhibits (120 input dimensions + 1 bias) x 84 input neurons = 10,164 parameter.
The output of the second fully-connected layer is then transferred to the output-layer (third fully-connected layer). The output layer is equipped with a softmax (that you learned about in the previous lab 05) and is made up of ten neurons, one for each object class contained in the CIFAR-10 dataset. This layer exhibits (84 + 1) x 10 = 850 parameter.
As a result our CIFAR-10 convolutional neural exhibits a total of 456 + 2,416 + 48,120 + 10,164 + 850 = 62,006 parameter.
(Source: https://www.stefanfiott.com/machine-learning/cifar-10-classifier-using-cnn-in-pytorch/)
Now, that we have implemented our first Convolutional Neural Network we are ready to instantiate a network model to be trained:
```
model = CIFAR10Net()
```
Let's push the initialized `CIFAR10Net` model to the computing `device` that is enabled:
```
model = model.to(device)
```
Let's double check if our model was deployed to the GPU if available:
```
!nvidia-smi
```
Once the model is initialized we can visualize the model structure and review the implemented network architecture by execution of the following cell:
```
# print the initialized architectures
print('[LOG] CIFAR10Net architecture:\n\n{}\n'.format(model))
```
Looks like intended? Brilliant! Finally, let's have a look into the number of model parameters that we aim to train in the next steps of the notebook:
```
# init the number of model parameters
num_params = 0
# iterate over the distinct parameters
for param in model.parameters():
# collect number of parameters
num_params += param.numel()
# print the number of model paramters
print('[LOG] Number of to be trained CIFAR10Net model parameters: {}.'.format(num_params))
```
Ok, our "simple" CIFAR10Net model already encompasses an impressive number 62'006 model parameters to be trained.
Now that we have implemented the CIFAR10Net, we are ready to train the network. However, before starting the training, we need to define an appropriate loss function. Remember, we aim to train our model to learn a set of model parameters $\theta$ that minimize the classification error of the true class $c^{i}$ of a given CIFAR-10 image $x^{i}$ and its predicted class $\hat{c}^{i} = f_\theta(x^{i})$ as faithfully as possible.
In this lab we use (similarly to lab 05) the **'Negative Log-Likelihood (NLL)'** loss. During training the NLL loss will penalize models that result in a high classification error between the predicted class labels $\hat{c}^{i}$ and their respective true class label $c^{i}$. Now that we have implemented the CIFAR10Net, we are ready to train the network. Before starting the training, we need to define an appropriate loss function. Remember, we aim to train our model to learn a set of model parameters $\theta$ that minimize the classification error of the true class $c^{i}$ of a given CIFAR-10 image $x^{i}$ and its predicted class $\hat{c}^{i} = f_\theta(x^{i})$ as faithfully as possible.
Let's instantiate the NLL via the execution of the following PyTorch command:
```
# define the optimization criterion / loss function
nll_loss = nn.NLLLoss()
```
Let's also push the initialized `nll_loss` computation to the computing `device` that is enabled:
```
nll_loss = nll_loss.to(device)
```
Based on the loss magnitude of a certain mini-batch PyTorch automatically computes the gradients. But even better, based on the gradient, the library also helps us in the optimization and update of the network parameters $\theta$.
We will use the **Stochastic Gradient Descent (SGD) optimization** and set the `learning-rate to 0.001`. Each mini-batch step the optimizer will update the model parameters $\theta$ values according to the degree of classification error (the NLL loss).
```
# define learning rate and optimization strategy
learning_rate = 0.001
optimizer = optim.SGD(params=model.parameters(), lr=learning_rate)
```
Now that we have successfully implemented and defined the three CNN building blocks let's take some time to review the `CIFAR10Net` model definition as well as the `loss`. Please, read the above code and comments carefully and don't hesitate to let us know any questions you might have.
## 5. Neural Network Model Training
In this section, we will train our neural network model (as implemented in the section above) using the transformed images. More specifically, we will have a detailed look into the distinct training steps as well as how to monitor the training progress.
### 5.1. Preparing the Network Training
So far, we have pre-processed the dataset, implemented the CNN and defined the classification error. Let's now start to train a corresponding model for **20 epochs** and a **mini-batch size of 128** CIFAR-10 images per batch. This implies that the whole dataset will be fed to the CNN 20 times in chunks of 128 images yielding to **391 mini-batches** (50.000 training images / 128 images per mini-batch) per epoch. After the processing of each mini-batch, the parameters of the network will be updated.
```
# specify the training parameters
num_epochs = 20 # number of training epochs
mini_batch_size = 128 # size of the mini-batches
```
Furthermore, lets specifiy and instantiate a corresponding PyTorch data loader that feeds the image tensors to our neural network:
```
cifar10_train_dataloader = torch.utils.data.DataLoader(cifar10_train_data, batch_size=mini_batch_size, shuffle=True)
```
### 5.2. Running the Network Training
Finally, we start training the model. The training procedure for each mini-batch is performed as follows:
>1. do a forward pass through the CIFAR10Net network,
>2. compute the negative log-likelihood classification error $\mathcal{L}^{NLL}_{\theta}(c^{i};\hat{c}^{i})$,
>3. do a backward pass through the CIFAR10Net network, and
>4. update the parameters of the network $f_\theta(\cdot)$.
To ensure learning while training our CNN model, we will monitor whether the loss decreases with progressing training. Therefore, we obtain and evaluate the classification performance of the entire training dataset after each training epoch. Based on this evaluation, we can conclude on the training progress and whether the loss is converging (indicating that the model might not improve any further).
The following elements of the network training code below should be given particular attention:
>- `loss.backward()` computes the gradients based on the magnitude of the reconstruction loss,
>- `optimizer.step()` updates the network parameters based on the gradient.
```
# init collection of training epoch losses
train_epoch_losses = []
# set the model in training mode
model.train()
# train the CIFAR10 model
for epoch in range(num_epochs):
# init collection of mini-batch losses
train_mini_batch_losses = []
# iterate over all-mini batches
for i, (images, labels) in enumerate(cifar10_train_dataloader):
# push mini-batch data to computation device
images = images.to(device)
labels = labels.to(device)
# run forward pass through the network
output = model(images)
# reset graph gradients
model.zero_grad()
# determine classification loss
loss = nll_loss(output, labels)
# run backward pass
loss.backward()
# update network paramaters
optimizer.step()
# collect mini-batch reconstruction loss
train_mini_batch_losses.append(loss.data.item())
# determine mean min-batch loss of epoch
train_epoch_loss = np.mean(train_mini_batch_losses)
# print epoch loss
now = datetime.utcnow().strftime("%Y%m%d-%H:%M:%S")
print('[LOG {}] epoch: {} train-loss: {}'.format(str(now), str(epoch), str(train_epoch_loss)))
# set filename of actual model
model_name = 'cifar10_model_epoch_{}.pth'.format(str(epoch))
# save current model to GDrive models directory
torch.save(model.state_dict(), os.path.join(models_directory, model_name))
# determine mean min-batch loss of epoch
train_epoch_losses.append(train_epoch_loss)
```
Upon successfull training let's visualize and inspect the training loss per epoch:
```
# prepare plot
fig = plt.figure()
ax = fig.add_subplot(111)
# add grid
ax.grid(linestyle='dotted')
# plot the training epochs vs. the epochs' classification error
ax.plot(np.array(range(1, len(train_epoch_losses)+1)), train_epoch_losses, label='epoch loss (blue)')
# add axis legends
ax.set_xlabel("[training epoch $e_i$]", fontsize=10)
ax.set_ylabel("[Classification Error $\mathcal{L}^{NLL}$]", fontsize=10)
# set plot legend
plt.legend(loc="upper right", numpoints=1, fancybox=True)
# add plot title
plt.title('Training Epochs $e_i$ vs. Classification Error $L^{NLL}$', fontsize=10);
```
Ok, fantastic. The training error converges nicely. We could definitely train the network a couple more epochs until the error converges. But let's stay with the 20 training epochs for now and continue with evaluating our trained model.
## 6. Neural Network Model Evaluation
Prior to evaluating our model, let's load the best performing model. Remember, that we stored a snapshot of the model after each training epoch to our local model directory. We will now load the last snapshot saved.
```
# restore pre-trained model snapshot
best_model_name = 'https://raw.githubusercontent.com/HSG-AIML-Teaching/EMBA2022-Lab/master/lab_05/models/cifar10_model_epoch_19.pth'
# read stored model from the remote location
model_bytes = urllib.request.urlopen(best_model_name)
# load model tensor from io.BytesIO object
model_buffer = io.BytesIO(model_bytes.read())
# init pre-trained model class
best_model = CIFAR10Net()
# load pre-trained models
best_model.load_state_dict(torch.load(model_buffer, map_location=torch.device('cpu')))
```
Let's inspect if the model was loaded successfully:
```
# set model in evaluation mode
best_model.eval()
```
In order to evaluate our trained model, we need to feed the CIFAR10 images reserved for evaluation (the images that we didn't use as part of the training process) through the model. Therefore, let's again define a corresponding PyTorch data loader that feeds the image tensors to our neural network:
```
cifar10_eval_dataloader = torch.utils.data.DataLoader(cifar10_eval_data, batch_size=10000, shuffle=False)
```
We will now evaluate the trained model using the same mini-batch approach as we did when training the network and derive the mean negative log-likelihood loss of all mini-batches processed in an epoch:
```
# init collection of mini-batch losses
eval_mini_batch_losses = []
# iterate over all-mini batches
for i, (images, labels) in enumerate(cifar10_eval_dataloader):
# run forward pass through the network
output = best_model(images)
# determine classification loss
loss = nll_loss(output, labels)
# collect mini-batch reconstruction loss
eval_mini_batch_losses.append(loss.data.item())
# determine mean min-batch loss of epoch
eval_loss = np.mean(eval_mini_batch_losses)
# print epoch loss
now = datetime.utcnow().strftime("%Y%m%d-%H:%M:%S")
print('[LOG {}] eval-loss: {}'.format(str(now), str(eval_loss)))
```
Ok, great. The evaluation loss looks in-line with our training loss. Let's now inspect a few sample predictions to get an impression of the model quality. Therefore, we will again pick a random image of our evaluation dataset and retrieve its PyTorch tensor as well as the corresponding label:
```
# set (random) image id
image_id = 777
# retrieve image exhibiting the image id
cifar10_eval_image, cifar10_eval_label = cifar10_eval_data[image_id]
```
Let's now inspect the true class of the image we selected:
```
cifar10_classes[cifar10_eval_label]
```
Ok, the randomly selected image should contain a two (2). Let's inspect the image accordingly:
```
# define tensor to image transformation
trans = torchvision.transforms.ToPILImage()
# set image plot title
plt.title('Example: {}, Label: {}'.format(str(image_id), str(cifar10_classes[cifar10_eval_label])))
# un-normalize cifar 10 image sample
cifar10_eval_image_plot = cifar10_eval_image / 2.0 + 0.5
# plot cifar 10 image sample
plt.imshow(trans(cifar10_eval_image_plot))
```
Ok, let's compare the true label with the prediction of our model:
```
cifar10_eval_image.unsqueeze(0).shape
best_model(cifar10_eval_image.unsqueeze(0))
```
We can even determine the likelihood of the most probable class:
```
cifar10_classes[torch.argmax(best_model(Variable(cifar10_eval_image.unsqueeze(0))), dim=1).item()]
```
Let's now obtain the predictions for all the CIFAR-10 images of the evaluation data:
```
predictions = torch.argmax(best_model(iter(cifar10_eval_dataloader).next()[0]), dim=1)
```
Furthermore, let's obtain the overall classification accuracy:
```
metrics.accuracy_score(cifar10_eval_data.targets, predictions.detach())
```
Let's also inspect the confusion matrix of the model predictions to determine major sources of misclassification:
```
# determine classification matrix of the predicted and target classes
mat = confusion_matrix(cifar10_eval_data.targets, predictions.detach())
# initialize the plot and define size
plt.figure(figsize=(8, 8))
# plot corresponding confusion matrix
sns.heatmap(mat.T, square=True, annot=True, fmt='d', cbar=False, cmap='YlOrRd_r', xticklabels=cifar10_classes, yticklabels=cifar10_classes)
plt.tick_params(axis='both', which='major', labelsize=8, labelbottom = False, bottom=False, top = False, left = False, labeltop=True)
# set plot title
plt.title('CIFAR-10 classification matrix')
# set plot axis lables
plt.xlabel('[true label]')
plt.ylabel('[predicted label]');
```
Ok, we can easily see that our current model confuses images of cats and dogs as well as images of trucks and cars quite often. This is again not surprising since those image categories exhibit a high semantic and therefore visual similarity.
## 7. Lab Summary:
In this lab, a step by step introduction into **design, implementation, training and evaluation** of convolutional neural networks CNNs to classify tiny images of objects is presented. The code and exercises presented in this lab may serves as a starting point for developing more complex, deeper and more tailored CNNs.
| github_jupyter |
```
# What is Volcano Forecasting?
This is an introduction to understanding the data and the problem.
**Volcanoes are awesome!** both in the California way and the "instilling awe" way.

Mount Etna on a calm day. (CC-BY Kuhnmi [Flickr](https://www.flickr.com/photos/31176607@N05/25720452925))
Volcano monitoring is important for both the inhabitants on and next to volcanoes, but also globally, as seen with the [Eyjafjallajökull eruption](https://en.wikipedia.org/wiki/2010_eruptions_of_Eyjafjallaj%C3%B6kull) disrupting air travel ten years ago. Geophysics is the field that largely works with active volcanoes and their activity, measuring earthquakes, tiltmeters, changes in gravimetry, etc. Specifically, seismologists record the rumble of the Earth when the magma forces its way upward. Geochemists collect data on degassing on Volcanos. And geologists are looking how the rocks form from lava.
In the spirit of the competition, we should not find additional metadata, however, it's important to understand the kind of data we are dealing with. A Nature article ([Hall 2018](https://www.nature.com/articles/d41586-018-07420-y#ref-CR1)) describes the world's first automatic volcano forecast system on Mount Etna. The data for this is mostly of acoustic nature, specifically infrasound ([Ripepe et al. 2018](https://agupubs.onlinelibrary.wiley.com/doi/abs/10.1029/2018JB015561)). Seeing that we have 10 sensors with timeseries, it's pretty safe to make the assumption that we are dealing with a seismological problem. These tend to be the most reliable, as compared to e.g. gas analyzers. Gas analyzers have to be right on top of a vent (which tend to rebuild in the lifetime of volcanoes), whereas the seismological stations just have to listen and not run out of battery.
So let's have a look at the actual data!
# Data
Files
train.csv Metadata for the train files.
`segment_id`: ID code for the data segment. Matches the name of the associated data file.
`time_to_eruption`: The target value, the time until the next eruption.
`[train|test]/*.csv`: the data files. Each file contains ten minutes of logs from ten different sensors arrayed around a volcano. The readings have been normalized within each segment, in part to ensure that the readings fall within the range of int16 values. If you are using the Pandas library you may find that you still need to load the data as float32 due to the presence of some nulls.
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import matplotlib.pyplot as plt # plots
import seaborn as sns
from pathlib import Path
from tqdm import tqdm
random_state = 42
# You can write up to 5GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
train = pd.read_csv("../input/predict-volcanic-erueptions-ingv-oe/train.csv")
train.describe()
sequence = pd.read_csv("../input/predict-volcanic-eruptions-ingv-oe/train/1000015382.csv", dtype="Int16")
sequence.describe()
sequence.tail()
```
Each sequence is 10 minutes long with 600001 samples. The data is `int16` but contains nan's, luckily with Pandas 1.0 they introduced the [nullable integer datatype](https://pandas.pydata.org/pandas-docs/stable/user_guide/integer_na.html), just make sure to actually call it capitalized `"Int16"` so Pandas knows.
```
sequence.fillna(0).plot(subplots=True, figsize=(25, 10))
plt.tight_layout()
plt.show()
```
These stations clearly have similar data, but are shifted in time. Why?
Look at this digital elevation model from Etna from ([Bonaccorso 2011](https://agupubs.onlinelibrary.wiley.com/doi/full/10.1029/2010GC003480))

The different stations are located all around the crater. That means the further away a station is from the magma or hypocentre of an Earthquake or other accoustic signals like streets, or [humans](https://www.weforum.org/agenda/2020/07/seismic-anthropogenic-noise-lockdown-covid19/), the longer the acoustic wave has to travel, arriving later at the station. So we can assume the with regards to that one signel event above, station 9 and 10 are almost identical in how far they are from it. Sensor 8 is a bit further.
Additionally, you can see the different noise levels of the data. Sensor 5 is fantastically quiet, only responding to big events, while Sensor 6 has some extremely periodic noise on it. It'd be worth to investigate if this is always the case.
Overall, the sensors will have different lags, depending where the volcanic activity is happening. So when we build a model that uses the time series, this is something to keep in mind.
# Play With the Data
Time series are hard. Can we get away with initially playing with some aggregate statistics?
```
def agg_stats(df, idx):
df = df.agg(['sum', 'min', "mean", "std", "median", "skew", "kurtosis"])
df_flat = df.stack()
df_flat.index = df_flat.index.map('{0[1]}_{0[0]}'.format)
df_out = df_flat.to_frame().T
df_out["segment_id"] = int(idx)
return df_out
summary_stats = pd.DataFrame()
for csv in tqdm(Path("../input/predict-volcanic-eruptions-ingv-oe/train/").glob("**/*.csv"), total=4501):
df = pd.read_csv(csv)
summary_stats = summary_stats.append(agg_stats(df, csv.stem))
test_data = pd.DataFrame()
for csv in tqdm(Path("../input/predict-volcanic-eruptions-ingv-oe/test/").glob("**/*.csv"), total=4501):
df = pd.read_csv(csv)
test_data = test_data.append(agg_stats(df, csv.stem))
features = list(summary_stats.drop(["segment_id"], axis=1).columns)
target_name = ["time_to_eruption"]
summary_stats = summary_stats.merge(train, on="segment_id")
summary_stats.head()
summary_stats.describe()
```
# Train a LightGBM Regressor
Use [Cross Validation](https://scikit-learn.org/stable/modules/cross_validation.html), because if you don't [shakeup](https://www.kaggle.com/jtrotman/meta-kaggle-competition-shake-up) will not be your friend.
```
import lightgbm as lgbm
from sklearn.model_selection import KFold
import gc
n_fold = 7
folds = KFold(n_splits=n_fold, shuffle=True, random_state=random_state)
data = summary_stats
params = {
"n_estimators": 2000,
"boosting_type": "gbdt",
"metric": "mae",
"num_leaves": 66,
"learning_rate": 0.005,
"feature_fraction": 0.9,
"bagging_fraction": 0.8,
"agging_freq": 3,
"max_bins": 2048,
"verbose": 0,
"random_state": random_state,
"nthread": -1,
"device": "gpu",
}
sub_preds = np.zeros(test_data.shape[0])
feature_importance = pd.DataFrame(index=list(range(n_fold)), columns=features)
for n_fold, (trn_idx, val_idx) in enumerate(folds.split(data)):
trn_x, trn_y = data[features].iloc[trn_idx], data[target_name].iloc[trn_idx]
val_x, val_y = data[features].iloc[val_idx], data[target_name].iloc[val_idx]
model = lgbm.LGBMRegressor(**params)
model.fit(trn_x, trn_y,
eval_set= [(trn_x, trn_y), (val_x, val_y)],
eval_metric="mae", verbose=0, early_stopping_rounds=150
)
feature_importance.iloc[n_fold, :] = model.feature_importances_
sub_preds += model.predict(test_data[features], num_iteration=model.best_iteration_) / folds.n_splits
best = feature_importance.mean().sort_values(ascending=False)
best_idx = best[best > 5].index
plt.figure(figsize=(14,26))
sns.boxplot(data=feature_importance[best_idx], orient="h")
plt.title("Features Importance per Fold")
plt.tight_layout()
```
So that gives us a nice idea which data actually matters for the next iteration.
# Submit Prediction
Let's build a csv for submission.
```
submission = pd.DataFrame()
submission['segment_id'] = test_data["segment_id"]
submission['time_to_eruption'] = sub_preds
submission.to_csv('submission.csv', header=True, index=False)
```
# How Far Can One Go?
Need Inspiration?
Of course someone tried Transformers on Earthquake time series, to detect the event and different phases. I love multi tasking like that!

Seismogram tagging on full sequences with an Earthquake Transformer in Nature ([Mousavi 2020](https://www.nature.com/articles/s41467-020-17591-w)).
| github_jupyter |
**This notebook is an exercise in the [Pandas](https://www.kaggle.com/learn/pandas) course. You can reference the tutorial at [this link](https://www.kaggle.com/residentmario/summary-functions-and-maps).**
---
# Introduction
Now you are ready to get a deeper understanding of your data.
Run the following cell to load your data and some utility functions (including code to check your answers).
```
import pandas as pd
pd.set_option("display.max_rows", 5)
reviews = pd.read_csv("../input/wine-reviews/winemag-data-130k-v2.csv", index_col=0)
from learntools.core import binder; binder.bind(globals())
from learntools.pandas.summary_functions_and_maps import *
print("Setup complete.")
reviews.head()
```
# Exercises
## 1.
What is the median of the `points` column in the `reviews` DataFrame?
```
median_points = reviews.points.median()
# Check your answer
q1.check()
#q1.hint()
q1.solution()
```
## 2.
What countries are represented in the dataset? (Your answer should not include any duplicates.)
```
countries = reviews.country.unique()
# Check your answer
q2.check()
#q2.hint()
q2.solution()
```
## 3.
How often does each country appear in the dataset? Create a Series `reviews_per_country` mapping countries to the count of reviews of wines from that country.
```
reviews_per_country = reviews.country.value_counts()
# Check your answer
q3.check()
#q3.hint()
q3.solution()
```
## 4.
Create variable `centered_price` containing a version of the `price` column with the mean price subtracted.
(Note: this 'centering' transformation is a common preprocessing step before applying various machine learning algorithms.)
```
centered_price = reviews.price-reviews.price.mean()
# Check your answer
q4.check()
q4.hint()
q4.solution()
```
## 5.
I'm an economical wine buyer. Which wine is the "best bargain"? Create a variable `bargain_wine` with the title of the wine with the highest points-to-price ratio in the dataset.
```
bargain_idx = (reviews.points / reviews.price).idxmax()
bargain_wine = reviews.loc[bargain_idx, 'title']
# Check your answer
q5.check()
#q5.hint()
q5.solution()
```
## 6.
There are only so many words you can use when describing a bottle of wine. Is a wine more likely to be "tropical" or "fruity"? Create a Series `descriptor_counts` counting how many times each of these two words appears in the `description` column in the dataset.
```
n_trop = reviews.description.map(lambda desc: "tropical" in desc).sum()
n_fruity = reviews.description. map (lambda desc: "fruity" in desc).sum()
descriptor_counts = pd.Series([n_trop, n_fruity], index = ['tropical', 'fruity'])
# Check your answer
q6.check()
#q6.hint()
q6.solution()
```
## 7.
We'd like to host these wine reviews on our website, but a rating system ranging from 80 to 100 points is too hard to understand - we'd like to translate them into simple star ratings. A score of 95 or higher counts as 3 stars, a score of at least 85 but less than 95 is 2 stars. Any other score is 1 star.
Also, the Canadian Vintners Association bought a lot of ads on the site, so any wines from Canada should automatically get 3 stars, regardless of points.
Create a series `star_ratings` with the number of stars corresponding to each review in the dataset.
```
def stars (row):
if row.country == 'Canada':
return 3
elif row.points >= 95:
return 3
elif row.points >= 85:
return 2
else:
return 1
star_ratings = reviews.apply(stars, axis ='columns' )
# Check your answer
q7.check()
q7.hint()
q7.solution()
```
# Keep going
Continue to **[grouping and sorting](https://www.kaggle.com/residentmario/grouping-and-sorting)**.
---
*Have questions or comments? Visit the [course discussion forum](https://www.kaggle.com/learn/pandas/discussion) to chat with other learners.*
| github_jupyter |
# Домашняя работа. Панеш Али 193
## Формулы
### Вспомогательная функция
$
f_t(x,\mu) = tf(x, \mu) + F(x, \mu) = \\
t(\frac{1}{2}||Ax-b||_2^2+\lambda \langle 1, u\rangle) + \sum_{i=1}^n (- \ln(-(x_i - \mu_i)) - \ln(-(-x_i-\mu_i))) = \\
t(\frac{1}{2}||Ax-b||_2^2+\lambda \langle 1, u\rangle) + \sum_{i=1}^n (- \ln (\mu_i - x_i) - \ln(x_i+\mu_i))
$
### Направление
$
d_k = (d_k^x, d_k^\mu)\\
d_k^x = -[\nabla^2f_t(x, \mu)_x]^{-1}\nabla f_t(x, \mu)_x\\
d_k^\mu = -[\nabla^2f_t(x, \mu)_\mu]^{-1}\nabla f_t(x, \mu)_\mu\\
$
#### Для х:
$\\
d^xf_t(x,\mu) =
t \langle Ax - b, Adx\rangle + \sum_{i=1}^n (\frac{1}{\mu_i - x_i} - \frac{1}{\mu_i + x_i})d_{x_i}
=\\
t \langle A^T(Ax - b), dx\rangle + \sum_{i=1}^n (\frac{1}{\mu_i - x_i} - \frac{1}{\mu_i + x_i})d_{x_i}
\\
\nabla_x f_t(x, \mu)_i = (tA^T(Ax - b))_i + \frac{1}{\mu_i - x_i} - \frac{1}{\mu_i + x_i}
\\
d [f_t(x, \mu)][dx, dx] =
t \langle A^TAdx, dx\rangle + \sum_{i=1}^n (\frac{1}{(\mu_i - x_i)^2} + \frac{1}{(\mu_i + x_i)^2})d_{x_i}
\\
\nabla^2_x f_t(x, \mu)_{ii} = (tA^TA)_{ii} + \frac{1}{(\mu_i - x_i)^2} + \frac{1}{(\mu_i + x_i)^2}\\
\nabla^2_x f_t(x, \mu)_{ij, i\ne j} =
(tA^TA)_{ij, i\ne j}$
#### Для $\mu:\\$
$
d^\mu f_t(x,\mu) =
t\lambda \langle 1, d\mu\rangle + \sum_{i=1}^n (- \frac{1}{\mu_i - x_i} - \frac{1}{x_i+\mu_i})d_{\mu_i}
\\
\nabla_\mu f_t(x, \mu)_i = t\lambda - \frac{1}{\mu_i - x_i} - \frac{1}{x_i+\mu_i}
\\
d^\mu f_t(x,\mu)[d\mu d\mu] = d(\sum_{i=1}^n (- \frac{1}{\mu_i - x_i} - \frac{1}{x_i+\mu_i}))=
\sum_{i=1}^n (\frac{1}{(\mu_i - x_i)^2} + \frac{1}{(x_i+\mu_i)^2})d_{\mu_i} d_{\mu_i}
\\
\nabla^2_\mu f_t(x, \mu)_{ii} = \frac{1}{(\mu_i - x_i)^2} + \frac{1}{(x_i+\mu_i)^2}
\\
\nabla^2_\mu f_t(x, \mu)_{ij, i\ne j} = 0$
#### Смешанная часть гессиана:
$
\nabla^2_\mu f_t(x, \mu)_{i_x, i_\mu} =
d/d^{\mu_i}(\frac{1}{\mu_i - x_i} - \frac{1}{\mu_i + x_i}) =
\frac{1}{(\mu_i + x_i)^2} - \frac{1}{(\mu_i - x_i)^2}
\\
\nabla^2_\mu f_t(x, \mu)_{i_\mu, i_x} =
d/d^{x_i}(- \frac{1}{\mu_i - x_i} - \frac{1}{x_i+\mu_i}) =
\frac{1}{(x_i+\mu_i)^2} - \frac{1}{(\mu_i - x_i)^2} = \nabla^2_\mu f_t(x, \mu)_{i_x, i_\mu}
$
### Максимальное $\alpha$
$g_i(x, \mu) = \langle q, (x, \mu) \rangle$, где $(x, \mu)$ - вектор состоящий из х и $\mu$, а $q = (..., \pm 1,..., -1, ...)$ это вектор где на $i_x (=i)$ месте стоит коэфицент для $x_i$, на $i_\mu (=i + n)$ стоит коэфицент для $\mu_i$, а в остальных местах 0.
Тогда $\alpha_{\max} = \min\limits_{i\in I} \frac{\langle q_i, (x, \mu) \rangle}{\langle q_i, d \rangle} = \min\limits_{i\in I} \frac{sgn(q_{ij})x_j - \mu_j}{sgn(q_{ij})d_j-d_{j+n}}$, где $I$ - множество индексов, где знаменатель больше 0.
### Начальная точка
В данной задаче нам нужно выбрать подходящую под условия начальную точку: $x = 0_n, \mu = 1_n$
### Способ решения
Предлагаю взять для больших матриц способ `LBFGS`, чтобы наши вычисления поместились в RAM и не было потребности вычислять гессиан, а для маленьких использовать `newton`, чтобы увеличить точность.
## Эксперемент с методом барьеров
### Подготовка
```
%load_ext autoreload
import matplotlib.pyplot as plt
from IPython.display import clear_output
from itertools import product
from oracles import BarrierOracle, lasso_duality_gap
from optimization import barrier_method_lasso, newton, lbfgs
import pandas as pd
import random
import numpy as np
%autoreload 2
def set_random_seed(seed):
np.random.seed(seed)
random.seed(seed)
def test_barrier(n=500, m=5000, eps_inner=1e-3, gamma=10, lambd=1., A=None, b=None):
if A is None:
A = 50 * np.random.rand(m, n) - 100
if b is None:
b = 5 * np.random.rand(m) - 10
return barrier_method_lasso(
A, b, lambd, np.zeros(n), np.ones(n), tolerance_inner=eps_inner,
gamma=gamma, lasso_duality_gap=lasso_duality_gap, trace=True)
def plot_ldg_over_time_or_iter(df, name, list_of_elements, ax=None, is_time=True):
colors = ['g', 'r', 'b', 'y', 'm', 'c', 'k']
if ax is not None:
plt.sca(ax)
for i, elem in enumerate(list_of_elements):
rows = df.loc[(df[name] == elem)]
if len(rows.index) == 0:
continue
ldg = list(rows['duality_gap'])[0]
time = list(rows['time'])[0]
if is_time:
plt.plot(time, ldg, color=colors[i % len(colors)], linewidth=2, label=f'{name}: {elem}')
else:
plt.plot(range(len(time)), ldg, color=colors[i % len(colors)], linewidth=2, label=f'{name}: {elem}')
if is_time:
plt.xlabel('Time')
plt.title('Duality gap over time')
else:
plt.xlabel('Iteration')
plt.title('Duality gap over iterations')
plt.ylabel('Duality gap')
plt.yscale('log')
plt.legend()
def plot_ldg_over_time_and_iter(df, name, list_of_elements):
_, axs = plt.subplots(1, 2, figsize=(16, 7))
plot_ldg_over_time_or_iter(df, name, list_of_elements, axs[0])
plot_ldg_over_time_or_iter(df, name, list_of_elements, axs[1], is_time=False)
set_random_seed(107)
gamma_ = np.linspace(10, 300, 5, dtype=float)
eps_inner_ = np.logspace(-8, -1, 5)
n_ = np.linspace(100, 1000, 5, dtype=int)
m_ = np.linspace(100, 10000, 5, dtype=int)
lambdas_ = np.logspace(-2, 3, 5)
n = 500
m = 5000
A = 50 * np.random.rand(m, n) - 100
b = 5 * np.random.rand(m) - 10
df_n = pd.DataFrame(columns=['n', 'func', 'duality_gap', 'time'])
df_m = pd.DataFrame(columns=['m', 'func', 'duality_gap', 'time'])
df_gamma = pd.DataFrame(columns=['gamma', 'func', 'duality_gap', 'time'])
df_eps = pd.DataFrame(columns=['eps', 'func', 'duality_gap', 'time'])
df_lambda = pd.DataFrame(columns=['lambda', 'func', 'duality_gap', 'time'])
for i, gamma in enumerate(gamma_):
_, msg, history = test_barrier(gamma=gamma, A=A, b=b)
if msg != 'success':
print(msg)
continue
df_gamma.loc[i] = [gamma, history['func'], history['duality_gap'], history['time']]
for i, eps in enumerate(eps_inner_):
_, msg, history = test_barrier(eps_inner=eps, A=A, b=b)
if msg != 'success':
continue
df_eps.loc[i] = [eps, history['func'], history['duality_gap'], history['time']]
for i, lambd in enumerate(lambdas_):
_, msg, history = test_barrier(lambd=lambd, A=A, b=b)
if msg != 'success':
continue
df_lambda.loc[i] = [lambd, history['func'], history['duality_gap'], history['time']]
for i, m in enumerate(m_):
_, msg, history = test_barrier(m=m)
if msg != 'success':
continue
df_m.loc[i] = [m, history['func'], history['duality_gap'], history['time']]
for i, n in enumerate(n_):
_, msg, history = test_barrier(n=n)
if msg != 'success':
continue
df_n.loc[i] = [n, history['func'], history['duality_gap'], history['time']]
clear_output()
result_config = {
'eps': [df_eps, 'eps', eps_inner_],
'm': [df_m, 'm', m_],
'n': [df_n, 'n', n_],
'gamma': [df_gamma, 'gamma', gamma_],
'lambda': [df_lambda, 'lambda', lambdas_],
}
```
### Результаты
#### Размер выборки
Время сходимости от размера выборки пропорционально размеру выборки, что логично из-за меньшего количества вычислений, тогда как количество внешних итераций в целом не сильно зависит от этого параметра.
```
plot_ldg_over_time_and_iter(*result_config['m'])
```
#### Размерность
Время сходимости в заисимости пропорцианально размерности, тогда как необходимое количество итераций обратно пропорционально. Что логично, потому что на большой выборке дольше обучать Ньютона, однако она более стабильна.
```
plot_ldg_over_time_and_iter(*result_config['n'])
```
#### Gamma
При больших $\gamma$ сходимость достигается быстрее. Видимо, потому что всё большую часть занимает не добавок (F(x)), который нужен лишь для того, чтобы сделать бортики, а сама функция.
```
plot_ldg_over_time_and_iter(*result_config['gamma'])
```
#### Epsilon
Как мы видим от $\varepsilon_{inner}$ скорость вычисления результата зависит не сильно. Скорее всего это вызванно довольно быстрым и точным нахождением в Ньютоне. Однако немного зависит конечный результат.
```
plot_ldg_over_time_and_iter(*result_config['eps'])
```
#### Коэфицент регуляризации
Как можно заметить зависимость нелинейна.
```
plot_ldg_over_time_and_iter(*result_config['lambda'])
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.