code stringlengths 2.5k 150k | kind stringclasses 1 value |
|---|---|
# Fixpoint Quantization and Overflow #
This notebook shows how number can be represented in binary format and how to (re-)quantize signals. This is also shown practically in Python using ``pyfda_fix_lib`` (numpy based).
Most images don't show in the github HTML at the moment (05/2020). Running the notebook locally works as well as the Notebook Viewer at https://nbviewer.jupyter.org/. Simply copy and paste the URL of this notebook.
```
import os, sys
import time
module_path = os.path.abspath(os.path.join('..')) # append directory one level up to import path
if module_path not in sys.path: # ... if it hasn't been appended already
sys.path.append(module_path)
import dsp_nmigen.pyfda_fix_lib as fx
%matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('script.mplstyle')
import numpy as np
import scipy.signal as sig
figsize = {"figsize":(13,7)}
```
## Theory ##
Unsigned integers are
The next figure shows how signed integer numbers are represented in two's complement format. The MSB is the sign bit which can be interpreted as the negative value $-2^{W-1}$.
<figure>
<center>
<img src='img/twos_complement_signed_int.png' alt='Signed integers in twos complement' width='40%'/>
<figcaption><b>Fig. x:</b> Signed integers in twos complement</figcaption>
</center>
</figure>
The same is possible for signed fractional values, although it is important to realize that the *binary point only exists in the developer's head*! All arithmetic stuff can be implemented with "integer thinking" (and some designers just do that). IMHO, thinking in fractional numbers makes it easier to track the real world value and to separate the range and the resolution.
<figure>
<center>
<img src='img/twos_complement_signed_frac.png' alt='Signed fractional values in twos complement' width='50%'/>
<figcaption><b>Fig. x:</b> Signed fractional values in twos complement</figcaption>
</center>
</figure>
### Increasing the wordlength
Before adding two fixpoint numbers their binary points need to be aligned, i.e. they need to have the same number of integer and fractional bits. This can be achieved by extending integer and / or fractional part.
The value of a fixpoint number doesn't change when zeros are appended to the fractional part (for positive and negative numbers) so that part is easy.
Prepending zeros to the integer part would change the sign of a negative number. Instead, the integer part is extended with copies of the sign bit, also called **sign extension**. VHDL has the function ``SEXT(std_log_vect, int)`` for achieving this.
<img src="img/requant_extension.png" alt="Extnd integer and fractional part" width="40%"/>
### Reduce the number of fractional bits
The problem of reducing the number of fractional places is well known from the decimal system. Different methods have been developed to achieve this like rounding, truncation, see e.g. https://www.eetimes.com/an-introduction-to-different-rounding-algorithms for more details than you'll ever need.
<figure>
<img src='img/requant_reduce_fractional.png' alt='Reduce fractional word length' width='80%'/>
<figcaption>Fig. x: Reduction of fractional wordlength</figcaption>
</figure>
## pyfda_fix_lib
This section uses and describes the fixpoint library `pyfda_fix_lib.py`. A quantizer is constructed as an instance of the class `Fixed()`, its properties are configured with a quantization dict `Q = fx.Fixed(q_dict)`. The quantization dict (in this case `q_dict`) sets the format `QI.QF` at the output of the quantizer and its quantization and overflow behaviour with the following keys:
- **'WI'** : number of integer bits (integer)
- **'WF'** : number of fractional bits (integer)
- **'quant'**: requantization behaviour (**'floor'**, 'round', 'fix', 'ceil', 'rint', 'none')
- **'ovfl'** : overflow behaviour (**'wrap'**, 'sat', 'none')
- **'frmt'** : number base / format for output (**'float'**, 'dec', 'bin', 'hex', 'csd'). Non-floats are scaled with 'scale'
- **'scale'**: float or a keyword; the factor between the fixpoint integer representation (FXP) and the real world value (RWV), RWV = FXP / scale. By default, scale = 1 << WI. If ``scale`` is a float, this value is used.
**Examples:**
```
WI.WF = 3.0, FXP = "b0110." = 6, scale = 8 -> RWV = 6 / 8 = 0.75
WI.WF = 1.2, FXP = "b01.10" = 1.5, scale = 2 -> RWV = 1.5 / 2 = 0.75
```
Alternatively, if:
- ``q_obj['scale'] == 'int'``: `scale = 1 << self.WF`
- ``q_obj['scale'] == 'norm'``: `scale = 2.**(-self.WI)`
`?fx.Fixed` shows the available options.
### Example
In the following example a quantizer is defined with an output format of 0 integer bits and 3 fractional bits, overflows are wrapped around in two's complement style and additional fractional bits are simply truncated ("floor").
```
q_dict = {'WI':0, 'WF': 3, # number of integer and fractional bits
'quant':'floor', 'ovfl': 'wrap'} # quantization and overflow behaviour
Q = fx.Fixed(q_dict) # instance of fixpoint class Fixed()
for i in np.arange(12)/10: # i = 0, 0.1, 0.2, ...
print("q<{0:>3.2f}> = {1:>5.3f}".format(i, Q.fixp(i))) # quantize i
# uncomment to show documentation
# ?fx.Fixed
```
### Signal Quantization
A sine signal $s(t)$ is quantized in the code below. The plot shows $s(t)$, the quantized signal $s_Q(t)$ the difference between both signals, the quantization error $\epsilon(t)$.
```
N = 10000; f_a = 1
t = np.linspace(0, 1, N, endpoint=False)
s = 1.1 * np.sin(2 * np.pi * f_a * t)
#
q_dict = {'WI':0, 'WF': 4, 'quant':'fix', 'ovfl': 'wrap'} # also try 'round' ; 'sat'
Q = fx.Fixed(q_dict) # quantizer instance with parameters defined above
t_cpu = time.perf_counter()
sq = Q.fixp(s) # quantize s
print('Overflows:\t{0}'.format(Q.N_over))
print('Run time:\t{0:.3g} ms for {1} quantizations\n'.format((time.perf_counter()-t_cpu)*1000, Q.N))
#
fig1, ax1 = plt.subplots(**figsize)
ax1.set_title('Quantized Signal $s_Q$({0}.{1}) with Quantizer Settings "{2}", "{3}"'.format(Q.WI, Q.WF, Q.ovfl, Q.quant))
ax1.plot(t, s, label = r'$s(t)$', lw=2)
ax1.step(t, sq, where = 'post', label = r'$s_Q(t)$', lw=2)
ax1.plot(t, s-sq, label = r'$\epsilon(t) = s(t) - s_Q(t)$', lw=2)
ax1.legend(fontsize = 14)
ax1.grid(True)
ax1.set_xlabel(r'$t \rightarrow$'); ax1.set_ylabel(r'$s \rightarrow$');
#
```
### Transfer Function of the Quantizer
The transfer function of the quantizer shows the quantized signal $s_Q(t)$ over the input signal $s(t)$ (a ramp). The advantage ove reusing the sine from the previous cell is that the input range can be taylored more easily and that the step size is constant.
```
Q.resetN() # reset overflow counter
x = np.linspace(-2, 2, N, endpoint=False) # generate ramp signal
xq = Q.fixp(x) # quantize x
print('Overflows:\t{0}'.format(Q.N_over))
fig2, ax2 = plt.subplots(**figsize); ax2.grid(True)
ax2.set_title('Quantization Transfer Function')
ax2.step(x,xq, where = 'post')
ax2.set_xlabel(r'$x \rightarrow$'); ax2.set_ylabel(r'$x_Q \rightarrow$');
```
### Number bases and formats
Quantized values can be printed in different number bases, e.g. as a binary string:
```
q_dict_f = {'WI':0, 'WF': 8, # number of ingeger and fractional bits
'quant':'floor', 'ovfl': 'wrap', # quantization and overflow behaviour
'frmt':'bin'} # output format
Q1 = fx.Fixed(q_dict_f) # instance of fixpoint class Fixed()
for i in np.arange(12)/10: # i = 0, 0.1, 0.2, ...
print("q<{0:>3.2f}> = {1}".format(i, Q1.float2frmt(Q1.fixp(i)))) # quantize i + display it in the wanted format
```
### Format Conversion
Numbers can also be converted back to floating with the method `frmt2float()`:
```
Q0 = fx.Fixed({'Q':'3.8', 'quant':'round', 'ovfl':'wrap', 'frmt':'bin'}) # direct setting of quantization options, use 'Q' instead of
Q1_dict = Q0.q_obj # read out Q0 quantization dict
Q1_dict.update({'WI':1, 'WF':5}) # update dict
Q1 = fx.Fixed(Q1_dict) # and create a new quantizer instance with it
frmt_str = "0.011"
x_org = Q0.frmt2float(frmt_str)
x_q = Q1.frmt2float(frmt_str)
print("q<b{0}> = {1} -> b{2} = {3}".format(frmt_str, x_org, Q1.float2frmt(x_q), x_q))
```
Bug ?: Illegal characters in the string to be formatted (e.g. frmt2float("0.0x11") do not raise an error but are ignored.
## FIR Filters
The following cell designs an equiripple filter and plots its impulse response (same as the coefficients) and its magnitude frequency response.
```
numtaps = 50 # filter order
N_FFT = 2000 # number of frequency bins per half plane
b = sig.remez(numtaps,[0,0.1,0.12, 0.5], [1,0], [1,10]) # frequency bands, target amplitude, weights
w, h = sig.freqz(b, [1], worN=N_FFT)
f = w / (2*np.pi)
fig, (ax1, ax2) = plt.subplots(2, **figsize); ax1.grid(True); ax2.grid(True)
ax1.set_title('Equiripple Lowpass (FIR) Filter: Impulse Response')
ax1.stem(np.arange(numtaps), b, use_line_collection=True)
ax1.set_xlabel(r'$n \rightarrow$'); ax1.set_ylabel(r'$b_n = h[n] \rightarrow$')
ax2.set_title('Magnitude Frequency Response')
ax2.plot(f,np.abs(h))
ax2.set_xlabel(r'$F \rightarrow$'); ax2.set_ylabel(r'$|H(F)| \rightarrow$')
fig.set_tight_layout(True)
```
Fixpoint filters process quantized input data (quantizer $Q_X$ in the image below), have quantized coeffients, a maximum accumulator width ($Q_A$) and a quantized output ($Q_Y$). Fig xx shows the topology of a direct form FIR filter.
<figure>
<center>
<img src='img/fir_df.png' alt='Direct Form FIR Filter' width='30%'/>
<figcaption><b>Fig. x:</b> Direct Form FIR Filter</figcaption>
</center>
</figure>
Due to the non-linear effects of quantization, fixpoint filters can only be simulated in the time domain, taking the filter topology into account. It can make a large difference for IIR filters in which order recursive and transversal part of the filter are calculated (direct form 1 vs. direct form 2) which is not the case for ideal systems.
Fixpoint filters process quantized input data (quantizer $Q_X$ in the image above), have quantized coeffients, a maximum accumulutator width ($Q_A$) and a quantized output ($Q_Y$).
```
class FIX_FIR_DF(fx.Fixed):
"""
Usage:
Q = FIX_FIR_DF(q_mul, q_acc) # Instantiate fixpoint filter object
x_bq = self.Q_mul.fxp_filt(x[k:k + len(bq)] * bq)
The fixpoint object has two different quantizers:
- q_mul describes requanitization after coefficient multiplication
- q_acc describes requantization after each summation in the accumulator
(resp. in the common summation point)
"""
def __init__(self, q_mul, q_acc):
"""
Initialize fixed object with q_obj
"""
# test if all passed keys of quantizer object are known
self.Q_mul = fx.Fixed(q_mul)
self.Q_mul.resetN() # reset overflow counter of Q_mul
self.Q_acc = fx.Fixed(q_acc)
self.Q_acc.resetN() # reset overflow counter of Q_acc
self.resetN() # reset filter overflow-counter
def fxp_filt_df(self, x, bq, verbose = True):
"""
Calculate filter (direct form) response via difference equation with
quantization
Parameters
----------
x : scalar or array-like
input value(s)
bq : array-like
filter coefficients
Returns
-------
yq : ndarray
The quantized input value(s) as an ndarray with np.float64. If this is
not what you want, see examples.
"""
# Initialize vectors (also speeds up calculation)
yq = accu_q = np.zeros(len(x))
x_bq = np.zeros(len(bq))
for k in range(len(x) - len(bq)):
# weighted state-vector x at time k:
x_bq = self.Q_mul.fixp(x[k:k + len(bq)] * bq)
# sum up x_bq to get accu[k]
accu_q[k] = self.Q_acc.fixp(sum(x_bq))
yq = accu_q # scaling at the output of the accumulator
if (self.Q_mul.N_over and verbose): print('Overflows in Multiplier: ',
Fixed.Q_mul.N_over)
if (self.Q_acc.N_over and verbose): print('Overflows in Accumulator: ',
self.Q_acc.N_over)
self.N_over = self.Q_mul.N_over + self.Q_acc.N_over
return yq
# nested loop would be much slower!
# for k in range(Nx - len(bq)):
# for i in len(bq):
# accu_q[k] = fixed(q_acc, (accu_q[k] + fixed(q_mul, x[k+i]*bq[i+1])))
q_bxy = {'WI':0, 'WF': 7, 'quant':'floor', 'ovfl': 'sat'} # quantization dict for x, q and coefficients
q_accu = {'WI':0, 'WF': 15, 'quant':'floor', 'ovfl': 'wrap'} # ... for accumulator
Q_X = fx.Fixed(q_bxy); Q_Y = fx.Fixed(q_bxy); Q_b = fx.Fixed(q_bxy); Q_accu = fx.Fixed(q_accu)
fil_q = FIX_FIR_DF(q_accu, q_accu)
x = np.zeros(1000); x[0] = 1
xq = Q_X.fixp(x); bq = Q_b.fixp(b)
yq = fil_q.fxp_filt_df(xq,bq)
fig, (ax1, ax2) = plt.subplots(2, **figsize); ax1.grid(True); ax2.grid(True)
ax1.set_title('Equiripple Lowpass (FIR) Filter')
ax1.stem(np.arange(numtaps), yq[:numtaps], use_line_collection=True)
ax1.set_xlabel(r'$n \rightarrow$'); ax1.set_ylabel(r'$b_n = h[n] \rightarrow$')
ax2.plot(f,np.abs(h), label="ideal system")
ax2.plot(f, np.abs(np.fft.rfft(yq, 2*N_FFT)[:-1]), label = "quantized system")
ax2.set_xlabel(r'$F \rightarrow$'); ax2.set_ylabel(r'$|H(F)| \rightarrow$')
ax2.legend()
fig.set_tight_layout(True)
```
## IIR Filters
Unlike FIR filters, IIR filters cannot be implemented with the same elegance and efficiency of array mathematics as each sample depends on the output sample as well. The following code cell designs an elliptic low pass filter and plots its magnitude frequency response.
```
b,a = sig.ellip(4,1,40, 2*0.1) # order, pass band ripple, stop band ripple, corner frequency w.r.t. f_S/2
w, h = sig.freqz(b,a, worN=2000)
fig, ax = plt.subplots(1, **figsize); ax.grid(True)
ax.set_title('Elliptic Lowpass (IIR) Filter')
ax.plot(w / (2*np.pi),20*np.log10(np.abs(h)))
ax.set_xlabel(r'$F \rightarrow$'); ax.set_ylabel(r'$|H(F)| \rightarrow$')
print("b=", b)
print("a=", a)
```
<figure>
<center>
<img src='img/iir_df1_df2.png' alt='Direct Form IIR Filter' width='60%'/>
<figcaption><b>Fig. x:</b> Direct Form FIR Filter Type 1 and 2</figcaption>
</center>
</figure>
The following two examples show how to implement the most simple recursive filters (no transversal path, i.e. "all-pole filters").
```
def IIR1(Q_ACCU, x, a):
"""
Rekursives Filter mit y[i] = Q< x[i-1] + a y[i-1] >
"""
y = np.zeros(len(x))
for i in range(0,len(x)-1):
y[i+1] = Q_ACCU.fixp(x[i] + a * y[i])
return y
def IIR2(Q_ACCU, x, a):
"""
Rekursives Filter mit y[i] = Q< x[i-2] + y[i-1] - a y[i-2] >
"""
y = np.zeros(len(x))
for i in range(len(x)-2):
y[i+2] = Q_ACCU.fixp(x[i] + y[i+1] - y[i]*a)
return y
alpha = 0.93 # coefficient
N_sim = 200 # number of simulation steps
x = np.zeros(N_sim); x[0] = 1.0 # x is dirac pulse with weight 1
q_x = {'WI':0,'WF':3,'quant':'round','ovfl':'sat'} # fixpoint quantization for stimulus
q_coeff = {'WI':0,'WF':3,'quant':'round','ovfl':'wrap'} # coefficient quantization dict
# quantizer settings for accumulator
#q_accu = {'WI':0,'WF':4,'quant':'fix','ovfl':'sat'} # saturation and round towards zero -> no limit cycles
q_accu = {'Q':0.8,'quant':'floor','ovfl':'wrap'} # große Grenzzyklen bei QI = 0
# kleine Grenzzyklen mit round / floor, abhängig von alpha:
# q_accu = {'WI':0,'WF':4,'quant':'floor','ovfl':'wrap'}
# Keine Quantisierung -> Werte für I, F beliebig
q_ideal = {'WI':0,'WF':0,'quant':'none','ovfl':'none'}
Q_coeff = fx.Fixed(q_coeff) # Fixpoint Object mit Parametern "q_coeff"
Q_ideal = fx.Fixed(q_ideal) # Fixpoint-Objekt ohne Quantisierung und Overflow
Q_accu = fx.Fixed(q_accu) # Fixpoint-Objekt mit Parametern "q_accu"
n = np.arange(N_sim)
t1 = time.perf_counter()
alpha_q = Q_ideal.fixp(alpha)
y = IIR1(Q_ideal, x, alpha_q) # ohne Quantisierung
#yq = IIR2(fx_IIR, x, alpha_q)
yq = IIR1(Q_accu, x, alpha_q)
```
***
### Copyright
(c) 2016 - 2020 Christian Münker
This notebook is provided as [Open Educational Resource](https://en.wikipedia.org/wiki/Open_educational_resources) , feel free to use it for your own purposes. Please attribute the work as follows: *Christian Münker, dsp_migen documentation*.
| github_jupyter |
# Recurrent Neural Nets - Fake News
The RNN (LSTM) architechture that we are using is shown below, a many to one RNN.

<img src='https://media.giphy.com/media/l0Iyau7QcKtKUYIda/giphy.gif'>
We achieve 87% accuracy in a test set. However, the article in Second reference claims to have 93% accuracy. The main difference is that they seem to use a Bag of Words Model, which loses the order of words when sending into the ML algorithm. Also
## References:
1. Data: https://github.com/GeorgeMcIntire/fake_real_news_dataset
2. Classification using Scikit Learn: https://blog.kjamistan.com/comparing-scikit-learn-text-classifiers-on-a-fake-news-dataset/
3. Glove vectors: https://nlp.stanford.edu/projects/glove/
```
!pip install tqdm
!conda install -y Pillow
%matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import re
from keras.models import Sequential
from keras.layers import Activation, Dropout, Flatten, Dense, BatchNormalization, LSTM, Embedding, Reshape
from keras.models import load_model, model_from_json
from sklearn.model_selection import train_test_split
import os
import urllib
from urllib.request import urlretrieve
from os import mkdir, makedirs, remove, listdir
from collections import Counter
from utilties import *
folder_path = 'data'
file = './data/fakenews.zip'
url = 'https://github.com/GeorgeMcIntire/fake_real_news_dataset/raw/master/fake_or_real_news.csv.zip'
downloadData(file, url)
#################################
# Download GLOVE vector dataset
#################################
file = './data/glove.6B.zip'
url = 'http://nlp.stanford.edu/data/glove.6B.zip'
downloadData(file, url)
with open('./data/glove.6B.50d.txt','rb') as f:
lines = f.readlines()
glove_weights = np.zeros((len(lines), 50))
words = []
for i, line in enumerate(lines):
word_weights = line.split()
words.append(word_weights[0])
weight = word_weights[1:]
glove_weights[i] = np.array([float(w) for w in weight])
word_vocab = [w.decode("utf-8") for w in words]
word2glove = dict(zip(word_vocab, glove_weights))
```
Preprocessing steps: lower case, remove urls, some punctuations etc.
```
from keras.engine.topology import Layer
import keras.backend as K
from keras import initializers
import numpy as np
class Embedding2(Layer):
def __init__(self, input_dim, output_dim, fixed_weights, embeddings_initializer='uniform',
input_length=None, **kwargs):
kwargs['dtype'] = 'int32'
if 'input_shape' not in kwargs:
if input_length:
kwargs['input_shape'] = (input_length,)
else:
kwargs['input_shape'] = (None,)
super(Embedding2, self).__init__(**kwargs)
self.input_dim = input_dim
self.output_dim = output_dim
self.embeddings_initializer = embeddings_initializer
self.fixed_weights = fixed_weights
self.num_trainable = input_dim - len(fixed_weights)
self.input_length = input_length
w_mean = fixed_weights.mean(axis=0)
w_std = fixed_weights.std(axis=0)
self.variable_weights = w_mean + w_std*np.random.randn(self.num_trainable, output_dim)
def build(self, input_shape, name='embeddings'):
fixed_weight = K.variable(self.fixed_weights, name=name+'_fixed')
variable_weight = K.variable(self.variable_weights, name=name+'_var')
self._trainable_weights.append(variable_weight)
self._non_trainable_weights.append(fixed_weight)
self.embeddings = K.concatenate([fixed_weight, variable_weight], axis=0)
self.built = True
def call(self, inputs):
if K.dtype(inputs) != 'int32':
inputs = K.cast(inputs, 'int32')
out = K.gather(self.embeddings, inputs)
return out
def compute_output_shape(self, input_shape):
if not self.input_length:
input_length = input_shape[1]
else:
input_length = self.input_length
return (input_shape[0], input_length, self.output_dim)
df = pd.read_csv('data/fake_or_real_news.csv')
df.drop('Unnamed: 0', axis=1, inplace=True)
df.title = df.title.str.lower()
df.text = df.text.str.lower()
df.title = df.title.str.replace(r'http[\w:/\.]+','<URL>') # remove urls
df.text = df.text.str.replace(r'http[\w:/\.]+','<URL>') # remove urls
df.title = df.title.str.replace(r'[^\.\w\s]','') #remove everything but characters and punctuation
df.text = df.text.str.replace(r'[^\.\w\s]','') #remove everything but characters and punctuation
df.title = df.title.str.replace(r'\.\.+','.') #replace multple periods with a single one
df.text = df.text.str.replace(r'\.\.+','.') #replace multple periods with a single one
df.title = df.title.str.replace(r'\.',' . ') #replace periods with a single one
df.text = df.text.str.replace(r'\.',' . ') #replace multple periods with a single one
df.title = df.title.str.replace(r'\s\s+',' ') #replace multple white space with a single one
df.text = df.text.str.replace(r'\s\s+',' ') #replace multple white space with a single one
df.title = df.title.str.strip()
df.text = df.text.str.strip()
print(df.shape)
df.head()
```
Get all the unique words. We will only consider words that have been used more than 5 times. Finally from this we create a dictionary mapping words to integers.
Once this is done we will create a list of reviews where the words are converted to ints.
```
all_text = ' '.join(df.text.values)
words = all_text.split()
u_words = Counter(words).most_common()
u_words_counter = u_words
u_words_frequent = [word[0] for word in u_words if word[1]>5] # we will only consider words that have been used more than 5 times
u_words_total = [k for k,v in u_words_counter]
word_vocab = dict(zip(word_vocab, range(len(word_vocab))))
word_in_glove = np.array([w in word_vocab for w in u_words_total])
words_in_glove = [w for w,is_true in zip(u_words_total,word_in_glove) if is_true]
words_not_in_glove = [w for w,is_true in zip(u_words_total,word_in_glove) if not is_true]
print('Fraction of unique words in glove vectors: ', sum(word_in_glove)/len(word_in_glove))
# # create the dictionary
word2num = dict(zip(words_in_glove,range(len(words_in_glove))))
len_glove_words = len(word2num)
freq_words_not_glove = [w for w in words_not_in_glove if w in u_words_frequent]
b = dict(zip(freq_words_not_glove,range(len(word2num), len(word2num)+len(freq_words_not_glove))))
word2num = dict(**word2num, **b)
word2num['<Other>'] = len(word2num)
num2word = dict(zip(word2num.values(), word2num.keys()))
int_text = [[word2num[word] if word in word2num else word2num['<Other>']
for word in content.split()] for content in df.text.values]
print('The number of unique words are: ', len(u_words))
print('The first review looks like this: ')
print(int_text[0][:20])
print('And once this is converted back to words, it looks like: ')
print(' '.join([num2word[i] for i in int_text[0][:20]]))
plt.hist([len(t) for t in int_text],50)
plt.show()
print('The number of articles greater than 500 in length is: ', np.sum(np.array([len(t)>500 for t in int_text])))
print('The number of articles less than 50 in length is: ', np.sum(np.array([len(t)<50 for t in int_text])))
```
You cannot pass differing lengths of sentences to the algorithm. Hence we shall prepad the sentence with `<PAD>`. Sequences less than 500 in length will be prepadded and sequences that are longer than 500 will be truncated. It is assumed that the sentiment of the review can be asserted from the first 500 words.
```
num2word[len(word2num)] = '<PAD>'
word2num['<PAD>'] = len(word2num)
for i, t in enumerate(int_text):
if len(t)<500:
int_text[i] = [word2num['<PAD>']]*(500-len(t)) + t
elif len(t)>500:
int_text[i] = t[:500]
else:
continue
x = np.array(int_text)
y = (df.label.values=='REAL').astype('int')
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size=0.1, random_state=42)
```
A real news article:
```
df[df.label=='REAL'].text.values[0]
```
A fake news article:
```
df[df.label=='FAKE'].text.values[0]
```
## Many to One LSTM
### Basic Method:
This method is no different to the method utilised in the sentiment analysis lesson.
```
model = Sequential()
model.add(Embedding(len(word2num), 50)) # , batch_size=batch_size
model.add(LSTM(64))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='rmsprop', metrics=['accuracy'])
model.summary()
batch_size = 128
epochs = 5
model.fit(X_train, y_train, batch_size=batch_size, epochs=1, validation_data=(X_test, y_test))
```
### Method 2: Fixed Embeddings
This is where we use the `Embedding2` class to which we give a set of weights which remain the same through training. Note especially the number of trainable parameters in the summary.
```
model = Sequential()
model.add(Embedding2(len(word2num), 50,
fixed_weights=np.array([word2glove[w] for w in words_in_glove]))) # , batch_size=batch_size
model.add(LSTM(64))
model.add(Dense(1, activation='sigmoid'))
# rmsprop = keras.optimizers.RMSprop(lr=1e-4)
model.compile(loss='binary_crossentropy', optimizer='rmsprop', metrics=['accuracy'])
model.summary()
```
I may heave cheated and run the following block 3 times. Good thing about Keras is that it remembers the last learning rate and goes from there.
```
batch_size = 128
model.fit(X_train, y_train, batch_size=batch_size, epochs=15, validation_data=(X_test, y_test))
sentence = "North korea is testing out missiles on americans living overseas .".lower()
sentence_num = [word2num[w] if w in word2num else word2num['<Other>'] for w in sentence.split()]
sentence_num = [word2num['<PAD>']]*(500-len(sentence_num)) + sentence_num
sentence_num = np.array(sentence_num)
model.predict(sentence_num[None,:])
' '.join([num2word[w] for w in sentence_num])
sentence = "The chemicals in the water is turning the freaking frogs gay says cnn . ".lower()
sentence_num = [word2num[w] if w in word2num else word2num['<Other>'] for w in sentence.split()]
sentence_num = [word2num['<PAD>']]*(500-len(sentence_num)) + sentence_num
sentence_num = np.array(sentence_num)
model.predict(sentence_num[None,:])
sentence = "President Trump is the greatest president of all time period .".lower()
sentence_num = [word2num[w] if w in word2num else word2num['<Other>'] for w in sentence.split()]
sentence_num = [word2num['<PAD>']]*(0) + sentence_num
sentence_num = np.array(sentence_num)
model.predict(sentence_num[None,:])
model.evaluate(X_test, y_test)
```
| github_jupyter |
```
ls
#imports
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import plotly.express as px
from datetime import datetime
```
## Background: Customer Lifetime Value
In marketing, customer lifetime value (CLV or often CLTV), lifetime customer value (LCV), or life-time value (LTV) is a prediction of the net profit attributed to the entire future relationship with a customer. Customer lifetime value can also be defined as the dollar value of a customer relationship, based on the present value of the projected future cash flows from the customer relationship. Customer lifetime value is an important concept in that it encourages firms to shift their focus from quarterly profits to the long-term health of their customer relationships. Customer lifetime value is an important number because it represents an upper limit on spending to acquire new customers. For this reason it is an important element in calculating payback of advertising spent in marketing mix modeling.
```
df = pd.read_csv("HW1 clv_transactions.csv", date_parser=True)
df.head()
```
## Step 1 - About the Dataset
The dataset available is in CSV format and contains 4,200 transactions records. Each row in the dataset represent a single transaction. There are four columns in the dataset.
- The TransactionID is a unique identifier of individual transactions.
- The TransactionDate is the date of the transaction.
- The CustomerID is the identifier of the customer who made the transaction.
- And the Amount is the recorded amount of transaction in US dollars.
```
df.isna().sum() # 1. There are no missing values in the dataset
print("Min. Transaction Date: {}".format(df.TransactionDate.apply(lambda x:datetime.strptime(x, '%m/%d/%Y')).min())) # 2. Min Transaction Date
print("Max. Transaction Date: {}".format(df.TransactionDate.apply(lambda x:datetime.strptime(x, '%m/%d/%Y')).max())) # 2. Max Transaction Date
print("Count of unique customers: {}".format(len(set(df.CustomerID)))) # 3. Count of unique customers
print("*Descriptive Stats*:\nMin: {}, Max: {}, Var: {:.2f}, Std: {:.2f}".format(df.Amount.min(),df.Amount.max(), df.Amount.var(), df.Amount.std()))
df["Trx_Year"] = df.TransactionDate.apply(lambda x:datetime.strptime(x, '%m/%d/%Y').year)
plt.figure(figsize=(16,8))
plt.scatter(x='Trx_Year', y='Amount', data=df)
plt.xlabel("Transaction Year")
plt.ylabel("Transaction Amount ($)")
plt.title("Transactions over time")
plt.show()
print("### Overall there doesn't seem to be any relationship of amounts over time. Will explore this again after removing outliers")
df.groupby("Trx_Year")[["Amount"]].sum().reset_index() # Extracting year
x = df.groupby("Trx_Year")[["Amount"]].sum()
x.plot(figsize=(16,8))
plt.xlabel("Transaction Year")
plt.ylabel("Total Transaction Amount ($)")
plt.title("Transactions over time")
plt.show()
```
Other that 2013, there has been a general increase in transaction amount over the years
## Step 2 - Further exploring the data
```
# 1. Are there any outliers?
plt.figure(figsize=(16,8))
plt.boxplot(df['Amount'])
plt.grid()
plt.title("Boxplot of Transaction Amount")
plt.show()
```
Yes, there are a few outliers in the dataset
```
df.Amount.describe()
plt.figure(figsize=(16,8))
plt.hist(x='Amount', data=df, bins =50)
plt.xlabel("Transaction Amount")
plt.ylabel("Frequency")
plt.title("Histogram of Transaction Amount")
```
Becuase of the extremely low frequency of the outliers, they are not visible
Amounts that are 3 SDs away from the mean can be treated as outlier and negative values can be treated as anomalies. For the purpose of elimination, lets group them together as 'Outlier'
```
#Labeling the outliers and error
df['Outlier'] = np.logical_or(np.logical_or( (df['Amount'] <= (df['Amount'].mean() - (df['Amount'].std() * 3))), # 3SDs less
(df['Amount'] >= (df['Amount'].mean() + (df['Amount'].std() * 3))) ), # 3SDs more
df['Amount']<0) # negative
#There are 5 outliers in the dataset
print("Transaction Outliers:")
df[df["Outlier"]==True]
# 2. If so how would you treat them?
# We will filter out all the outliers
df = df[df['Outlier'] == False]
# New Boxplot
plt.figure(figsize=(16,8))
plt.title("Boxplot of Amount after getting rid of Outliers ( >3 sd away from mean)")
plt.boxplot(df['Amount'])
plt.grid()
plt.show()
plt.figure(figsize=(16,8))
plt.hist(x='Amount', data=df, bins =50,rwidth=0.7)
plt.xlabel("Transaction Amount")
plt.ylabel("Frequency")
plt.title("Histogram of Transaction Amount after Outlier removal")
plt.grid()
plt.figure(figsize=(18,12))
ax = sns.boxplot(x='Trx_Year', y='Amount', data=df,boxprops={'facecolor':'None'}, showfliers=False,whiskerprops={'linewidth':0})
ax = sns.swarmplot(x='Trx_Year', y='Amount', data=df)
plt.xlabel("Transaction Year")
plt.ylabel("Transaction Amount ($)")
plt.title("Transactions over time")
plt.show()
```
The mean transaction amounts appear to have stayed constant over the years but the counts have increased as seen in the above swarm plot
## Step 3 - Determining when the customers were first acquired
```
# Similar to a partition by function in SQL -- gets the minimum Trx_Year for each CustomerID
df["Origin Year"] = df.groupby('CustomerID').Trx_Year.transform(np.min)
```
Displaying the origin year for a random customer
```
df.head(10)
```
#### Finding the Origin year for a random customer
```
df.loc[df["CustomerID"] == np.random.randint(100)]
```
## Step 4 - Calculating cumulative transaction amounts
```
df_cohort = df.groupby(['Origin Year', 'Trx_Year']) \
.agg(Trx_amount=('Amount', 'sum')) \
.reset_index(drop=False)
df_cohort["period_since_aquisition"]= (df_cohort["Trx_Year"]-df_cohort["Origin Year"]+1)*12
df_cohort["Cum_Trx_amount"] = df_cohort.groupby('Origin Year').Trx_amount.transform(np.cumsum)
cohort_table = df_cohort.pivot_table(index="Origin Year", columns="period_since_aquisition", values="Cum_Trx_amount").reset_index()
cohort_table["Origin Year"] = cohort_table["Origin Year"].apply(lambda x: str(x)+"-01-01 - " + str(x)+"-12-31", )
cohort_table.rename_axis("", axis=1).fillna("")
```
## Step 5 - New Customers Cohort Analysis
```
a = df.groupby(['Origin Year', 'Trx_Year']) \
.agg(Unique_customers=('CustomerID', 'nunique')) \
.reset_index()
a["x"] = a["Unique_customers"] * np.floor(a["Origin Year"] / a["Trx_Year"]).astype("int")
a["x"] = a["x"].astype("int")
a["Unique_customers"] = a.groupby(['Origin Year']).x.transform(max)
del a["x"]
a["period_since_aquisition"]= (a["Trx_Year"]-a["Origin Year"]+1)*12
a = a.pivot_table(index="Origin Year", columns="period_since_aquisition", values="Unique_customers").reset_index()
a["Origin Year"] = a["Origin Year"].apply(lambda x: str(x)+"-01-01 - " + str(x)+"-12-31", )
a = a.rename_axis("", axis=1).fillna("")
a
```
## Step 6 - Historic CLV
```
clv_table = ((cohort_table.set_index("Origin Year"))/(a.set_index("Origin Year"))).fillna("")
clv_table
#Plotting historic CLV
plt.figure(figsize=(16,10))
cohort1, = plt.plot(clv_table.iloc[0], marker="o")
cohort2, = plt.plot(clv_table.iloc[1 ,0:5], marker= "o")
cohort3, = plt.plot(clv_table.iloc[2 ,0:4], marker= "o")
cohort4, = plt.plot(clv_table.iloc[3 ,0:3], marker= "o")
cohort5, = plt.plot(clv_table.iloc[4 ,0:2], marker= "o")
cohort6, = plt.plot(clv_table.iloc[5 ,0:1], marker= "o")
plt.title("Historic CLV ")
plt.xlabel('Age')
plt.ylabel('Historic CLV')
plt.grid()
plt.legend([cohort1, cohort2, cohort3, cohort4, cohort5, cohort6], clv_table.index,
loc='upper left', prop={'size': 14})
plt.show()
```
## Step 7 - Interpretting Results
```
pd.DataFrame(clv_table.replace("", np.nan).mean(axis = 0, skipna = True).round(2), columns=["HistoricCLV"]).reset_index().rename(columns={"period_since_aquisition":"Age"})
```
- As we can see, customers aquired in 2011 have spent $ 7862.24
- Each cohort seems to exhibit similar spending pattern. The business implication is that customers will continue to give incremental and consitant business year-over-year and therefore efforts should be made to retain them.
- Based on the Historic CLV, it can also be observed that older customers spend less money after 2 years than they did initially. This trend is worth noting and can be taking into consideration while planning markieting budgets for older customers.
- We also can notice that customers aquired in later years have been spending less on average in their first year. This could be due to the drop in product price but is again worth noting.
| github_jupyter |
# Leave-One-Patient-Out classification of individual volumes
Here, we train a classifier for each patient, based on the data of all the other patients except the current one (Leave One Out Cross-Validation). To this end, we treat each volume as an independent observation, so we have a very large sample of volumes which are used for training; and later, we do not classify the patient as a whole, but the classifier makes a decision for each of the held-out patient's 200 volumes. Therefore, at this stage, we have not made a decision on the patient level, but only at the volume-as-unit-of-observation level.
### import modules
```
import os
import pickle
import numpy as np
import pandas as pd
from sklearn import svm, preprocessing, metrics
from PIL import Image
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('ticks')
sns.set_context('poster')
sns.set_context('poster')
# after converstion to .py, we can use __file__ to get the module folder
try:
thisDir = os.path.realpath(__file__)
# in notebook form, we take the current working directory (we need to be in 'notebooks/' for this!)
except:
thisDir = '.'
# convert relative path into absolute path, so this will work with notebooks and py modules
supDir = os.path.abspath(os.path.join(os.path.dirname(thisDir), '..'))
supDir
```
### get meta df
We need this e.g. to get information about conclusiveness
```
data_df = pd.read_csv(
'../data/interim/csv/info_epi_zscored_zdiff_summarymaps_2dpredclean_corr_df.csv',
index_col=[0, 1],
header=0)
data_df.tail()
```
#### conclusiveness filters
```
is_conclusive = data_df.loc[:, 'pred'] != 'inconclusive'
is_conclusive.sum()
```
### get data
```
def make_group_df(data_df,metric='corr_df'):
'''load correlation data of all patients'''
group_df = pd.DataFrame()
for p in data_df.index:
# get data
filename = data_df.loc[p, metric]
this_df = pd.read_csv(filename, index_col=[0], header=0)
# add patient infos to index
this_df.index = [[p[0]], [p[1]]]
group_df = pd.concat([group_df, this_df])
# reorder the colums and make sure volumes are integer values
group_df.columns = group_df.columns.astype(int)
# sort across rows, then across columns, to make sure that volumes
# are in the right order
group_df = group_df.sort_index(axis=0)
group_df = group_df.sort_index(axis=1)
assert all(group_df.columns == range(200)), 'wrong order of volumes'
return group_df
group_df = make_group_df(data_df)
group_df.tail()
```
#### filter data
```
# only conclusive cases
conclusive_df = group_df[is_conclusive]
# only inconclusive cases
inconclusive_df = group_df[is_conclusive == False]
# all cases unfiltered
withinconclusive_df = group_df.copy()
print(conclusive_df.shape, inconclusive_df.shape, withinconclusive_df.shape)
```
### get design
```
conds_file = os.path.join(supDir,'models','conds.p')
with open(conds_file, 'rb') as f:
conds = pickle.load(f)
print(conds)
```
### get colors
```
with open('../models/colors.p', 'rb') as f:
color_dict = pickle.load(f)
my_cols = {}
for i, j in zip(['red', 'blue', 'yellow'], ['left', 'right', 'bilateral']):
my_cols[j] = color_dict[i]
```
### invert the resting timepoints
```
inv_df = conclusive_df*conds
inv_df.tail()
```
### train the classifier
```
stack_df = pd.DataFrame(inv_df.stack())
stack_df.tail()
stack_df.shape
my_groups = ['left','bilateral','right']
dynamite_df = stack_df.copy()
dynamite_df.columns = ['correlation']
dynamite_df['group'] = dynamite_df.index.get_level_values(0)
sns.catplot(data=dynamite_df,y='group',x='correlation',kind='bar',orient='h',palette=my_cols,order=my_groups,aspect=1)
plt.axvline(0,color='k',linewidth=3)
plt.xlim(0.05,-0.05,-0.01)
sns.despine(left=True,trim=True)
plt.ylabel('')
plt.savefig('../reports/figures/10-dynamite-plot.png',dpi=300,bbox_inches='tight')
plt.show()
from scipy import stats
t,p = stats.ttest_ind(dynamite_df.loc['bilateral','correlation'],dynamite_df.loc['left','correlation'])
print('\nt=%.2f,p=%.64f'%(t,p))
t,p = stats.ttest_ind(dynamite_df.loc['bilateral','correlation'],dynamite_df.loc['right','correlation'])
print('\nt=%.2f,p=%.38f'%(t,p))
t,p = stats.ttest_ind(dynamite_df.loc['left','correlation'],dynamite_df.loc['right','correlation'])
print('\nt=%.2f,p=%.248f'%(t,p))
```
### as histogram
```
fig,ax = plt.subplots(1,1,figsize=(8,5))
for group in my_groups:
sns.distplot(stack_df.loc[group,:],color=my_cols[group],label=group,ax=ax)
plt.legend()
plt.xlim(0.4,-0.4,-0.2)
sns.despine()
plt.show()
```
### set up the classifier
```
clf = svm.SVC(kernel='linear',C=1.0,probability=False,class_weight='balanced')
def scale_features(X):
'''z-transform the features before applying a SVC.
The scaler is also stored so it can later be re-used on test data'''
my_scaler = preprocessing.StandardScaler()
my_scaler.fit(X)
X_scaled = my_scaler.transform(X)
return X_scaled,my_scaler
def encode_labels(y):
'''get from number labels to strings and back'''
my_labeler = preprocessing.LabelEncoder()
my_labeler.fit(np.unique(y))
y_labels = my_labeler.transform(y)
return y_labels, my_labeler
def train_classifier(df):
'''get features and labels
* scale the features
* transform the labels
* apply the classifier
'''
X = df.values
y = df.index.get_level_values(0)
X_scaled,my_scaler = scale_features(X)
y_labels, my_labeler = encode_labels(y)
clf.fit(X_scaled,y_labels)
return clf,my_scaler,my_labeler
example_clf, example_scaler, example_labeler = train_classifier(stack_df)
example_clf
example_scaler
example_labeler.classes_
def get_boundaries(clf,my_scaler):
'''find the point where the classifier changes its prediction;
this is an ugly brute-force approach and probably there is a much
easier way to do this
'''
d = {}
for i in np.linspace(-1,1,10000):
this_val = my_scaler.transform(np.array([i]).reshape(1,-1))
this_predict = clf.predict(this_val)
d[i] = this_predict[-1]
df = pd.DataFrame(d,index=['pred']).T
return df[(df-df.shift(1))!=0].dropna().index[1:]
from datetime import datetime
```
### get class boundaries of all folds
```
import tqdm
def get_all_boundaries(stack_df):
'''for each fold, get the boundaries, by
training on everybody but the held-out patient
and storing the boundaries'''
all_boundaries = {}
conclusive_pats = np.unique(stack_df.index.get_level_values(1))
for p in tqdm.tqdm(conclusive_pats):
# in the current fold, we drop one patient
df = stack_df.drop(p,level=1)
# train on this fold's data
clf,my_scaler,my_labeler = train_classifier(df)
# get the classifier boundaries
boundaries = get_boundaries(clf,my_scaler)
all_boundaries[p] = boundaries
return all_boundaries
```
Compute the boundaries and store them for later re-use:
```
all_boundaries = get_all_boundaries(stack_df)
bound_df = pd.DataFrame(all_boundaries).T
bound_df.tail()
bound_df.to_csv('../data/processed/csv/bound_df.csv')
```
To make things faster, we can re-load the computed boundaries here:
```
bound_df = pd.read_csv('../data/processed/csv/bound_df.csv',index_col=[0],header=0)
bound_df.tail()
```
rename so boundaries have meaningful descriptions:
```
bound_df = bound_df.rename(columns={'0':'B/R','1':'L/B'})
bound_df.tail()
bound_df.describe()
```
#### show the class boundaries overlaid on the data distribution
```
fig,ax = plt.subplots(1,1,figsize=(8,5))
for group in my_groups:
sns.distplot(stack_df.loc[group,:],color=my_cols[group],label=group,ax=ax)
for b in bound_df.values.flatten():
plt.axvline(b,alpha=0.1,color=color_dict['black'])
plt.legend()
plt.xlabel('correlation')
plt.ylabel('density')
plt.xlim(0.4,-0.4,-0.2)
plt.ylim(0,8)
plt.legend(loc=(0.65,0.65))
sns.despine(trim=True,offset=5)
plt.savefig('../reports/figures/10-distribution-plot.png',dpi=300,bbox_inches='tight')
plt.show()
```
#### make swarm/factorplot with boundary values
```
sns_df = pd.DataFrame(bound_df.stack())
sns_df.columns = ['correlation']
sns_df.loc[:,'boundary'] = sns_df.index.get_level_values(1)
sns_df.loc[:,'dummy'] = 0
sns_df.tail()
fig,ax = plt.subplots(1,1,figsize=(4,5))
sns.swarmplot(data=sns_df,
x='correlation',
y='dummy',
hue='boundary',
orient='h',
palette={'L/B':my_cols['left'],'B/R':my_cols['right']},
size=4,
alpha=0.9,
ax=ax
)
plt.xlim(0.04,-0.02,-0.02)
ax.set_ylabel('')
ax.set_yticks([])
sns.despine(left=True,trim=True)
plt.savefig('../reports/figures/10-boundary-swarm-plot.png',dpi=300,bbox_inches='tight')
plt.show()
```
### combine above into one plot
```
sns.set_style('dark')
fig = plt.figure(figsize=(16,6))
ax1 = fig.add_axes([0.36, .999, 1, .7], xticklabels=[], yticklabels=[])
ax1.imshow(Image.open('../reports/figures/10-dynamite-plot.png'))
ax2 = fig.add_axes([0, 1, 1, 0.8], xticklabels=[], yticklabels=[])
ax2.imshow(Image.open('../reports/figures/10-distribution-plot.png'))
ax3 = fig.add_axes([0.65, 1, 1, 0.8], xticklabels=[], yticklabels=[])
ax3.imshow(Image.open('../reports/figures/10-boundary-swarm-plot.png'))
plt.text(0,1, 'A',transform=ax2.transAxes, fontsize=32)
plt.text(1.04,1, 'B',transform=ax2.transAxes, fontsize=32)
plt.text(1.63,1, 'C',transform=ax2.transAxes, fontsize=32)
plt.savefig('../reports/figures/10-training-overview.png',dpi=300,bbox_inches='tight')
plt.show()
```
### make predictions for all patients (conc and inconc)
#### invert
```
all_inv_df = group_df*conds
all_inv_df.tail()
def make_preds(this_df,clf,my_scaler,my_labeler):
'''apply fitted classifier to the held-out patient;
based on what has been done during training, we
* scale the features using the stored scaler
* transform the labels using the stored labeler
* apply the classifier using the stored classfier
'''
scaled_features = my_scaler.transform(this_df.T)
predictions = clf.predict(scaled_features)
labeled_predictions = my_labeler.inverse_transform(predictions)
counts = pd.Series(labeled_predictions).value_counts()
counts_df = pd.DataFrame(counts).T
counts_df.index = pd.MultiIndex.from_tuples(this_df.index)
return counts_df
```
Example:
```
make_preds(all_inv_df.iloc[[-1]],example_clf, example_scaler, example_labeler)
import warnings
# this is necessary to get rid of https://github.com/scikit-learn/scikit-learn/issues/10449
with warnings.catch_warnings():
warnings.filterwarnings("ignore",category=DeprecationWarning)
for p in tqdm.tqdm(all_inv_df.index):
# get data in leave-one-out fashion
this_df = all_inv_df.loc[[p],:]
other_df = stack_df.drop(p[-1],level=1)
# train on this fold's data
clf,my_scaler,my_labeler = train_classifier(other_df)
# make predictions
p_df = make_preds(this_df,clf,my_scaler,my_labeler)
out_name = '../data/processed/csv/%s_counts_df.csv' % p[-1]
p_df.to_csv(out_name)
data_df.loc[p,'counts_df'] = out_name
data_df.to_csv('../data/processed/csv/info_epi_zscored_zdiff_summarymaps_2dpredclean_corr_counts_df.csv')
```
### train classifier once on all data and store
We store a classifer trained on all data as a pickle file so we can re-use it in the future on new data
```
clf,my_scaler,my_labeler = train_classifier(stack_df)
d = {'clf':clf,'scaler':my_scaler,'labeler':my_labeler}
with open('../models/volume_clf.p','wb') as f:
pickle.dump(d,f)
```
#### toolbox model
The toolbox assumes that a dataset used as input is a new dataset and was not part of this study
```
#clf_file = os.path.join(supDir,'models','volume_clf.p')
#with open(clf_file,'rb') as f:
# clf_dict = pickle.load(f)
#
#clf = clf_dict['clf']
#my_scaler = clf_dict['scaler']
#my_labeler = clf_dict['labeler']
#def make_p(pFolder,pName,clf=clf,my_scaler=my_scaler,my_labeler=my_labeler):
#
# filename = os.path.join(pFolder, ''.join([ pName, '_corr_df.csv']))
# this_df = pd.read_csv(filename, index_col=[0], header=0)
# this_df.index = [['correlations'],[pName]]
# inv_df = this_df*conds
# counts_df = make_preds(inv_df,clf,my_scaler,my_labeler)
#
# out_name = os.path.join(pFolder, ''.join([ pName, '_counts_df.csv']))
# counts_df.to_csv(out_name)
#
# return out_name
```
### summary
For each patient, a classfier has been developed based on all the other patient (Leave-One-Out) and applied to the 200 volumes of that patient. There are now 200 decisions for each patient, as many as there are volumes. These data are stored in csv files which we can now access to make a prediction on the level of the patient.
**************
< [Previous](09-mw-correlations-with-template.ipynb) | [Contents](00-mw-overview-notebook.ipynb) | [Next >](11-mw-logistic-regression.ipynb)
| github_jupyter |
```
import numpy as np
import tensorflow as tf
assert tf.__version__.startswith('2')
from tensorflow_examples.lite.model_maker.core.data_util.image_dataloader import ImageClassifierDataLoader
from tensorflow_examples.lite.model_maker.core.task import image_classifier
from tensorflow_examples.lite.model_maker.core.task.model_spec import efficientnet_lite2_spec, efficientnet_lite4_spec
from tensorflow_examples.lite.model_maker.core.task.model_spec import ImageModelSpec
import matplotlib.pyplot as plt
# from matplotlib.pyplot import specgram
# import librosa
# import librosa.display
from sklearn.metrics import classification_report, accuracy_score, confusion_matrix
import pandas as pd
tf.config.list_physical_devices('GPU')
def train_valid_fold_images(fold):
fold = str(fold)
base_path = "../downsampled/imagenet_structure/"
return ImageClassifierDataLoader.from_folder(base_path + folder + "/train"), ImageClassifierDataLoader.from_folder(base_path + fold + "/valid")
train_data, valid_data = train_valid_fold_images('1')
# Customize the pre-trained TensorFlow model
model = image_classifier.create(train_data, model_spec=efficientnet_lite2_spec)
# Evaluate the model
loss, accuracy = model.evaluate(valid_data)
```
# Lite4
```
# Customize the pre-trained TensorFlow model
model_lite4 = image_classifier.create(train_data,
model_spec=efficientnet_lite4_spec, warmup_steps = 100)
loss, accuracy = model_lite4.evaluate(valid_data)
model_lite4.export('UrbanEfficientNetLite4_out1_e5_noshuffle.tflite', 'urban_label_lite4.txt')
model_lite4 = image_classifier.create(train_data,
model_spec=efficientnet_lite4_spec,
shuffle = True,
batch_size = 24,
warmup_steps = 100,
epochs = 8)
loss, accuracy = model_lite4.evaluate(valid_data)
model_path_prefix = './models/UrbanEfficientNetLite4_val1_e8_shuffle'
model_lite4.export(model_path_prefix+'.tflite', model_path_prefix+'_labels.txt')
ds_test = model_lite4._gen_dataset(valid_data, 24, is_training=False)
ds_test
model_lite4.model.predict_classes
import inspect
print(inspect.getsource(model_lite4.model.predict_classes))
valid_predicts = model_lite4.predict_top_k(valid_data)
valid_label = [valid_data.index_to_label[label.numpy()] for i, (image, label) in enumerate(valid_data.dataset.take(len(predicts)))]
valid_predict_label = [i[0][0] for i in valid_predicts]
from sklearn.metrics import classification_report, accuracy_score, confusion_matrix
print(classification_report(y_true = valid_true_id, y_pred = valid_predict_id, labels = valid_data.index_to_label))
import pandas as pd
conf = confusion_matrix(y_true = valid_label, y_pred = valid_predict_label, labels = valid_data.index_to_label)
conf = pd.DataFrame(conf, columns = valid_data.index_to_label)
conf['true_row_label'] = valid_data.index_to_label
conf.set_index('true_row_label', drop = True, inplace = True)
conf_perc = round(conf.div(conf.sum(axis=1), axis=0),2)
conf_perc
def predict_class(data):
predict = model_lite4.predict_top_k(data)
return [i[0][0] for i in predict]
#e12
print(classification_report(y_true = valid_label, y_pred = valid_predict_label))
model_lite4 = image_classifier.create(train_data,
model_spec=efficientnet_lite4_spec,
shuffle = True,
batch_size = 24,
warmup_steps = 100,
epochs = 20)
loss, accuracy = model_lite4.evaluate(valid_data)
model_path_prefix = './models/UrbanEfficientNetLite4_val1_e20_shuffle'
model_lite4.export(model_path_prefix+'.tflite', model_path_prefix+'_labels.txt')
valid_predicts = model_lite4.predict_top_k(valid_data)
valid_label = [valid_data.index_to_label[label.numpy()] for i, (image, label) in enumerate(valid_data.dataset.take(len(valid_predicts)))]
valid_predict_label = [i[0][0] for i in valid_predicts]
#e20
print(classification_report(y_true = valid_label, y_pred = valid_predict_label))
conf = confusion_matrix(y_true = valid_label, y_pred = valid_predict_label, labels = valid_data.index_to_label)
conf = pd.DataFrame(conf, columns = valid_data.index_to_label)
conf['true_row_label'] = valid_data.index_to_label
conf.set_index('true_row_label', drop = True, inplace = True)
conf_perc = round(conf.div(conf.sum(axis=1), axis=0),2)
conf_perc
```
# Multiple Models
```
def train_valid_folder_images(folder):
folder = str(folder)
base_path = "../downsampled/imagenet_structure/"
return ImageClassifierDataLoader.from_folder(base_path + folder + "/train"), ImageClassifierDataLoader.from_folder(base_path + folder + "/valid")
def create_fit_submodel(image_folder_substring, epochs = 10, warmup_steps = 100, batch_size = 24):
train_data, valid_data = train_valid_folder_images(image_folder_substring)
return image_classifier.create(train_data,
model_spec=efficientnet_lite4_spec,
shuffle = True,
epochs = epochs,
batch_size = batch_size,
warmup_steps = warmup_steps,
validation_data = valid_data)
np.setdiff1d(labels,submodels['engine-air-other'],True).tolist()
import os
import glob
import shutil
from pathlib import Path
"/path"+os.listdir()
data_path = Path('../downsampled/imagenet_structure/ensemble/')
submodels = {'motors-other': ['air_conditioner', 'engine_idling','drilling', 'jackhammer'],}
labels = ['air_conditioner','car_horn','children_playing',
'dog_bark','drilling','engine_idling','gun_shot','jackhammer','siren','street_music']
def move_submodel_files(submodel_folder_name, submodel_class_list):
for d in ['train', 'valid']:
if not os.path.exists(data_path/submodel_folder_name/d/'motors'):
os.mkdir(data_path/submodel_folder_name/d/'motors')
for c in submodel_class_list: #np.setdiff1d(labels,submodel_class_list,True).tolist():
png_files = list(Path(data_path/submodel_folder_name/d/c).glob('*.png'))
for f in png_files:
shutil.move(str(f), str(data_path/submodel_folder_name/d/'motors'))
os.rmdir(data_path/submodel_folder_name/d/c)
for k, v in submodels.items():
print(k)
move_submodel_files(k,v)
```
## Drill-Jackhammer
```
model_drill_jackhammer-other = create_fit_submodel('ensemble/drilling-jackhammer-other', epochs = 20)
model_drill_jackhammer.model.save('models/ensemble/drill_jackhammer_e15')
loss, accuracy = model_drill_jackhammer.evaluate(valid_data)
model_path_prefix = './models/UrbanDrillJackhammerEfficientNet'
model_drill_jackhammer.export(model_path_prefix+'.tflite', model_path_prefix+'_labels.txt')
valid_predicts = model_drill_jackhammer.predict_top_k(valid_data)
valid_label = [valid_data.index_to_label[label.numpy()] for i, (image, label) in enumerate(valid_data.dataset.take(len(valid_predicts)))]
valid_predict_label = [i[0][0] for i in valid_predicts]
#e20
print(classification_report(y_true = valid_label, y_pred = valid_predict_label))
```
## Engine-Air
```
train_data, valid_data = train_valid_folder_images('ensemble/engine-air')
model_engine_air = image_classifier.create(train_data,
model_spec=efficientnet_lite4_spec,
shuffle = True,
batch_size = 24,
warmup_steps = 100,
epochs = 10)
loss, accuracy = model_engine_air.evaluate(valid_data)
model_path_prefix = './models/UrbanEngineAirEfficientNet'
model_engine_air.export(model_path_prefix+'.tflite', model_path_prefix+'_labels.txt')
valid_predicts = model_engine_air.predict_top_k(valid_data)
valid_label = [valid_data.index_to_label[label.numpy()] for i, (image, label) in enumerate(valid_data.dataset.take(len(valid_predicts)))]
valid_predict_label = [i[0][0] for i in valid_predicts]
#e20
print(classification_report(y_true = valid_label, y_pred = valid_predict_label))
```
## Other Classes
```
train_data, valid_data = train_valid_folder_images('ensemble/other')
model_other = image_classifier.create(train_data,
model_spec=efficientnet_lite4_spec,
shuffle = True,
batch_size = 24,
warmup_steps = 100,
epochs = 10)
loss, accuracy = model_other.evaluate(valid_data)
model_path_prefix = './models/UrbanOtherEfficientNet'
model_other.export(model_path_prefix+'.tflite', model_path_prefix+'_labels.txt')
valid_predicts = model_other.predict_top_k(valid_data)
valid_label = [valid_data.index_to_label[label.numpy()] for i, (image, label) in enumerate(valid_data.dataset.take(len(valid_predicts)))]
valid_predict_label = [i[0][0] for i in valid_predicts]
#e20
print(classification_report(y_true = valid_label, y_pred = valid_predict_label))
```
# Import Tflite models & Ensemble
```
from PIL import Image
class urban_ensemble():
def __init__(tflite_path_dict):
self.path_dict = tflite_path_dict
def _load_tflite_model(model_path, label_path):
with tf.io.gfile.GFile('model_path', 'rb') as f:
model_content = f.read()
with tf.io.gfile.GFile('label_path', 'r') as f:
label_names = f.read().split('\n')
interpreter = tf.lite.Interpreter(model_content = model_content)
input_index = interpreter.get_input_details()[0]['index']
output = interpreter.tensor(interpreter.get_output_details()[0]["index"])
imtest = Image.open('7061-6-0-0.png')
imtest = np.asarray(imtest)/255
tf.lite.
def load_labels(path):
with open(path, 'r') as f:
return {i: line.strip() for i, line in enumerate(f.readlines())}
def set_input_tensor(interpreter, image):
tensor_index = interpreter.get_input_details()[0]['index']
input_tensor = interpreter.tensor(tensor_index)()[0]
input_tensor[:, :] = image
def classify_image(interpreter, image_array, top_k=1):
"""Returns a sorted array of classification results."""
interpreter.set_tensor(input_details[0]['index'], input_array)
interpreter.invoke()
output_details = interpreter.get_output_details()[0]
output = np.squeeze(interpreter.get_tensor(output_details['index']))
# If the model is quantized (uint8 data), then dequantize the results
if output_details['dtype'] == np.uint8:
scale, zero_point = output_details['quantization']
output = scale * (output - zero_point)
ordered = np.argpartition(-output, top_k)
return [(i, output[i]) for i in ordered[:top_k]]
##################################################
class UrbanInterpreter():
def __init__(model_dict):
self.model_files_dict = model_files_dict
def _read_tflite_model(model_path):
with tf.io.gfile.GFile(model_path, 'rb') as f:
return f.read()
def _read_tflite_labels(label_path)
with tf.io.gfile.GFile(label_path, 'r') as f:
return f.read().split('\n')
def _initialize_interpreter(model_files, label_names):
model_content = _read_tflite_model(model_files(['tflite_file']))
label_names = _read_tflite_labels(model_files(['labels']))
interpreter = tf.lite.Interpreter(model_content= model_content)
interpreter.allocate_tensors()
input_index = interpreter.get_input_details()[0]['index']
output_index = interpreter.get_output_details()[0]["index"]
return interpreter
def urban_ensemble_predict(model_dict, image_path):
labels= load_labels(labels)
interpreter = tf.lite.Interpreter(model)
interpreter.allocate_tensors()
_, height, width, _ = interpreter.get_input_details()[0]['shape']
image = np.asarray(Image.open(image_path).resize((width, height)))/255
start_time = time.time()
results = classify_image(interpreter, image)
label_id, prob = results[0]
finally:
camera.stop_preview()
# Read TensorFlow Lite model from TensorFlow Lite file.
with tf.io.gfile.GFile('flower_classifier.tflite', 'rb') as f:
model_content = f.read()
# Read label names from label file.
with tf.io.gfile.GFile('flower_labels.txt', 'r') as f:
label_names = f.read().split('\n')
# Initialze TensorFlow Lite inpterpreter.
interpreter = tf.lite.Interpreter(model_content=model_content)
interpreter.allocate_tensors()
input_index = interpreter.get_input_details()[0]['index']
output = interpreter.tensor(interpreter.get_output_details()[0]["index"])
# Run predictions on each test image data and calculate accuracy.
accurate_count = 0
for i, (image, label) in enumerate(test_data.dataset):
# Pre-processing should remain the same. Currently, just normalize each pixel value and resize image according to the model's specification.
image, _ = model.preprocess(image, label)
# Add batch dimension and convert to float32 to match with the model's input
# data format.
image = tf.expand_dims(image, 0).numpy()
# Run inference.
interpreter.set_tensor(input_index, image)
interpreter.invoke()
# Post-processing: remove batch dimension and find the label with highest
# probability.
predict_label = np.argmax(output()[0])
# Get label name with label index.
predict_label_name = label_names[predict_label]
accurate_count += (predict_label == label.numpy())
accuracy = accurate_count * 1.0 / test_data.size
print('TensorFlow Lite model accuracy = %.4f' % accuracy)
```
| github_jupyter |
# BOW Featurization and modeling
```
import pandas as pd
import numpy as np
import re
from tqdm import tqdm
import warnings
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.metrics import accuracy_score, confusion_matrix, classification_report
from sklearn.naive_bayes import MultinomialNB
from sklearn.svm import SVC
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from joblib import dump
from sklearn.model_selection import train_test_split
warnings.filterwarnings("ignore")
data = pd.read_csv("AFFR_preprocessed_100k.csv")
data.tail(3)
def text_splitter(text):
return text.split()
# max_features = 20000 means we want only most useful(most occured) 20000 features not all
vectorizer = CountVectorizer(tokenizer = text_splitter,ngram_range=(1, 3),max_features=20000,min_df=5, max_df=0.7)
review_vector = vectorizer.fit_transform(data['Reviews'].values.astype(str))
dump(vectorizer,"AFFR_vectorizer.pkl")
review_vector.shape
# Getting labels seperate
y_label = data["rating"]
x_train, x_test, y_train, y_test = train_test_split(review_vector, y_label, test_size = 0.20)
```
### SVM Classifier with RBF kernel
```
%%time
svmclassifier = SVC(kernel='rbf',verbose=True,gamma="auto")
svmclassifier.fit(x_train, y_train)
y_pred = svmclassifier.predict(x_test)
print("Confusion matrix: \n",confusion_matrix(y_test,y_pred))
print("Classification report: \n",classification_report(y_test,y_pred))
print("Accuracy score is: ",accuracy_score(y_test,y_pred))
print("Model Saving ...")
dump(svmclassifier,"AFFR_SVM_model.pkl")
```
### Naive Bayes
```
%%time
# Here Gaussian Naive Bayes does not taking Sparse matrix it requires dense
NB_classifier = MultinomialNB()
NB_classifier.fit(x_train, y_train)
y_pred = NB_classifier.predict(x_test)
print("Confusion matrix: \n",confusion_matrix(y_test,y_pred))
print("Classification report: \n",classification_report(y_test,y_pred))
print("Accuracy score is: ",accuracy_score(y_test,y_pred))
print("Model Saving ...")
dump(NB_classifier,"AFFR_NB_model.pkl")
%%time
LR_classifier = LogisticRegression(n_jobs=-1)
LR_classifier.fit(x_train, y_train)
y_pred = LR_classifier.predict(x_test)
print("Confusion matrix: \n",confusion_matrix(y_test,y_pred))
print("Classification report: \n",classification_report(y_test,y_pred))
print("Accuracy score is: ",accuracy_score(y_test,y_pred))
print("Model Saving ...")
dump(LR_classifier,"AFFR_LR_model.pkl")
%%time
KNN_classifier = KNeighborsClassifier(n_jobs=-1)
KNN_classifier.fit(x_train, y_train)
y_pred = KNN_classifier.predict(x_test)
print("Confusion matrix: \n",confusion_matrix(y_test,y_pred))
print("Classification report: \n",classification_report(y_test,y_pred))
print("Accuracy score is: ",accuracy_score(y_test,y_pred))
print("Model Saving ...")
dump(KNN_classifier,"AFFR_KNN_model.pkl")
%%time
RF_classifier = RandomForestClassifier(n_jobs=-1)
RF_classifier.fit(x_train, y_train)
y_pred = RF_classifier.predict(x_test)
print("Confusion matrix: \n",confusion_matrix(y_test,y_pred))
print("Classification report: \n",classification_report(y_test,y_pred))
print("Accuracy score is: ",accuracy_score(y_test,y_pred))
print("Model Saving ...")
dump(RF_classifier,"AFFR_RF_model.pkl")
%%time
DT_classifier = DecisionTreeClassifier()
DT_classifier.fit(x_train, y_train)
y_pred = DT_classifier.predict(x_test)
print("Confusion matrix: \n",confusion_matrix(y_test,y_pred))
print("Classification report: \n",classification_report(y_test,y_pred))
print("Accuracy score is: ",accuracy_score(y_test,y_pred))
print("Model Saving ...")
dump(DT_classifier,"AFFR_DT_model.pkl")
```
| github_jupyter |
```
#import the packages here
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import emcee
import math
import george
from george import kernels
from pylab import *
import corner
#load data and model here
spectra = pd.read_csv('/home/ade/Documents/Research/comparemodel/model_spectra_all_200.csv')
##this part will be substituted by the new code developed in
##tutorial_create many tables in a single fits file
#start of wavelength range
print(str(spectra.w_obj[min(range(len(spectra.w_obj)), key=lambda i: abs(spectra.w_obj[i]-5770.))]))
print(min(range(len(spectra.w_obj)), key=lambda i: abs(spectra.w_obj[i]-5770.)))
nearest_index_max_start_wav = min(range(len(spectra.w_obj)), key=lambda i: abs(spectra.w_obj[i]-5770))
if spectra.w_obj[nearest_index_max_start_wav] < 5770.:
nearest_index_max_start_wav = nearest_index_max_start_wav + 1
print(spectra.w_obj[nearest_index_max_start_wav])
print(nearest_index_max_start_wav)
#end of wavelength range
print(str(spectra.w_obj[min(range(len(spectra.w_obj)), key=lambda i: abs(spectra.w_obj[i]-5790.))]))
print(min(range(len(spectra.w_obj)), key=lambda i: abs(spectra.w_obj[i]-5790.)))
nearest_index_min_end_wav = min(range(len(spectra.w_obj)), key=lambda i: abs(spectra.w_obj[i]-5790.))
if spectra.w_obj[nearest_index_min_end_wav] > 5790.:
nearest_index_min_end_wav = nearest_index_min_end_wav - 1
print(spectra.w_obj[nearest_index_min_end_wav])
print(nearest_index_min_end_wav)
#the spectra
spectra = spectra.iloc[nearest_index_max_start_wav: nearest_index_min_end_wav]
spectra
#propagation of uncertainty to acquire error of f_obj/f_mean
def y_error(y_obj, e_obj, y_mod, e_mod):
return (e_obj/y_obj)+(e_mod/y_mod) * (y_obj/y_mod)
#yerr = y_error(spectra.f_obj, spectra.e_obj, spectra.f_mean, spectra.f_stddev)
yerr = spectra.e_obj
plt.figure(figsize = (10,10))
plt.errorbar(spectra.w_obj,spectra.f_obj/spectra.f_mean, yerr = yerr, fmt = 'k.', capsize=0)
#plt.scatter(spectra.w_obj,spectra.f_obj/spectra.f_mean)
#plt.xlim(5775, 5785)
plt.ylim(0, 1.5)
plt.xlabel("Wavelength (nm)")
plt.ylabel("Norm. Flux")
plt.show()
#collection of function: prior and likelihood
#def model(w, A, sigma, mu): #w = wavelength
# return (1-(A/(2*np.pi*(sigma**2))**0.5) * (np.exp(-np.power(w - mu, 2.) / (2 * np.power(sigma, 2.)))))
##this version is the updated model
##merged with asymmetry introduction
def model(p,x):
a,s,m,asym=p
#return 1-((a/(2*np.pi*(s**2))**0.5) * exp(-(x-m)**2/(2*s**2)))
ns = (2*s)/(1+np.exp(asym*(x-m))) #ns = new s = new sigma after asymmetry introduction
return (1-(a/(2*np.pi*(ns**2))**0.5) * (np.exp(-np.power(x - m, 2.) / (2 * np.power(ns, 2.)))))
#return 1-a * exp(-(x-m)**2/(2*s**2)
def log_likelihood(theta, w, difference, error): #difference = f_obj/f_mean, error = error propagation of f_obj/f_mean
A, sigma, mu, asym = theta
ns = (2*sigma)/(1+np.exp(asym*(w-mu)))
model = (1-(A/(2*np.pi*(ns**2))**0.5) * (np.exp(-np.power(w - mu, 2.) / (2 * np.power(ns, 2.)))))
return -0.5*np.sum((difference-model)**2/error**2) #+ np.log((A/(2*np.pi*(sigma**2))**0.5))
def log_prior(theta):
A, sigma, mu, asym = theta
#if 0 < A < 5 and 0.01 < sigma < 5.5 and (5780.5 - 3) < mu < (5780.5 + 3):
# return 0.0
#return -np.inf
lnp=0.0
#Flat prior on A
if 0 <= A <= 1:
lnp-=0.0
#elif 0.2 < A <= 0.4:
# lnp+=np.log(np.log(-5 * A + 2))
else :
return -np.inf
#Flat prior on asymmetry
if -0.75 <= asym <= 0.75:
lnp-=0.0
#elif 0.2 < A <= 0.4:
# lnp+=np.log(np.log(-5 * A + 2))
else :
return -np.inf
#Skewed Gaussian prior on sigma
m_sigma = 0.2
s_sigma = 0.5
gamma = 2
if 0.00 < sigma < 1.5:
lnp += np.log(1./(np.sqrt(2*np.pi*(s_sigma**2)))) + (-0.5*(sigma-m_sigma)**2/s_sigma**2) + np.log(1 + math.erf((gamma*(sigma-m_sigma))/s_sigma*(2**0.5)))
else :
return -np.inf
#Gaussian prior on mu
m_mu = i_mu
s_mu = 1.
if 5778 < mu < 5782:
lnp += np.log(1.0/(np.sqrt(2*np.pi)*s_mu))-(0.5*(mu-m_mu)**2/s_mu**2)
else :
return -np.inf
return lnp
def log_probability(theta, w, difference, error):
lp = log_prior(theta)
#print (lp)
if not np.isfinite(lp):
return -np.inf
return lp + log_likelihood(theta, w, difference, error)
#distribution of walkers
i_A, i_sigma, i_mu, i_asym = 0.6, 1.0, 5780, 0.0 #initial A, sigma, mu. Or simply the mean
i_b, i_a, i_c, i_d = 0.05, 0.05, 2, 0.1 #the spread, or mu of Gaussian distribution
pos_A = np.random.normal(i_A, i_b, 100)
pos_sigma = np.random.normal(i_sigma, i_a, 100)
pos_mu = np.random.normal(i_mu, i_c, 100)
pos_asym = np.random.normal(i_asym, i_d, 100)
#the vertical slide
import lmfit
def residual(pars, y_obj, y_model):
"""Model a decaying sine wave and subtract data."""
vals = pars.valuesdict()
const = vals['const']
return y_obj - (const*y_model)
from lmfit import Parameters, fit_report, minimize
fit_params = Parameters()
fit_params.add('const', value=1.0)
out = minimize(residual, fit_params, args=(spectra['f_obj'], spectra['f_mean']))
spectra.f_mean = spectra.f_mean*out.params['const'].value
gp=george.GP(5*np.var(spectra.f_obj/spectra.f_mean)*kernels.Matern32Kernel(1.05), mean=0)
gp.compute(spectra.w_obj, yerr)
pos = np.array([pos_A, pos_sigma, pos_mu, pos_asym]).T #transpose the array of A, sigma, and mu
nwalkers, ndim = pos.shape
sampler = emcee.EnsembleSampler(nwalkers, ndim, log_probability, args=(spectra.w_obj, spectra.f_obj/spectra.f_mean, yerr), threads = 8)
position, _ , _ = sampler.run_mcmc(pos, 500, progress=True);
flat_samples = sampler.get_chain(discard = 0, flat=True)[sampler.flatlnprobability>np.percentile(sampler.flatlnprobability,20)]
labels = [r"A", r"\sigma", r"\mu", r"asym"]
fig, axes = plt.subplots(4, figsize=(10, 7), sharex=True)
samples = sampler.get_chain()
for j in range(ndim):
ax = axes[j]
ax.plot(samples[:, :, j], "k", alpha=0.3)
ax.set_xlim(0, len(samples))
ax.set_ylabel(labels[j])
ax.yaxis.set_label_coords(-0.1, 0.5)
s=[np.median(samples[:,0]), np.median(samples[:,1]), np.median(samples[:,2]), np.median(samples[:,3])]
labels = ["A", r"$\sigma$", r"$\mu$", "asym"]
names = gp.get_parameter_names()
inds = [0, 1, 2, 3]
corner.corner(sampler.flatchain[:, inds], labels=labels, quantiles=[0.16, 0.5, 0.84], show_titles=True, title_kwargs={"fontsize": 14});
xx=np.linspace(5770,5790,1000)
s=[np.median(flat_samples[:,0]), np.median(flat_samples[:,1]), np.median(flat_samples[:,2]), np.median(samples[:,3])]
m = gp.sample_conditional(spectra.f_obj/spectra.f_mean - model(s, spectra.w_obj), xx) + model(s, xx) #correlated noise + model
mm = model(s, xx) #model only
mu = gp.sample_conditional(spectra.f_obj/spectra.f_mean - model(s, spectra.w_obj), xx)
figure(1, figsize = (15,10))
errorbar(spectra.w_obj, spectra.f_obj/spectra.f_mean, yerr=yerr, fmt=".k", capsize=0)
plot(xx,mm,'r-', label = 'model')
plot(xx,m,'b-', label = 'model + correlated noise')
plot(xx,mu,'k-', label = 'correlated noise')
legend()
samples = sampler.get_chain(discard = 0, flat=True)[sampler.flatlnprobability>np.percentile(sampler.flatlnprobability,20)]
figure(1, figsize = (15,10))
errorbar(spectra.w_obj, spectra.f_obj/spectra.f_mean, yerr=yerr, fmt=".k", capsize=0)
ylim(0.5, 1.3)
for s in samples[np.random.randint(len(samples), size=10)]:
#gp.set_parameter_vector(s)
#print(s)
mu = gp.sample_conditional(spectra.f_obj/spectra.f_mean - model(s, spectra.w_obj), xx) + model(s, xx)
#mu = gp.sample_conditional(spectra.f_obj/spectra.f_mean - model(spectra.w_obj, s[s][0], s[s][1], s[s][2]), xx) + model(xx, s[s][0], s[s][1], s[s][2])
#mu_val.append(mu)
m = gp.sample_conditional(spectra.f_obj/spectra.f_mean - model(s, spectra.w_obj), xx) + 1
mm = model(s, xx)
plot(xx,mm,'r-', label = 'model')
plot(xx,m,'k-', label = 'correlated noise')
plt.plot(xx, mu, color="#4682b4", alpha=0.3)
```
| github_jupyter |
# DataFrames
We can think of a DataFrame as a bunch of Series objects put together to share the same index. Let's use pandas to explore this topic!
```
import pandas as pd
import numpy as np
from numpy.random import randn
np.random.seed(101)
df = pd.DataFrame(randn(5,4),index='A B C D E'.split(),columns='W X Y Z'.split())
df
```
## Selection and Indexing
Let's learn the various methods to grab data from a DataFrame
```
df['W']
# Pass a list of column names
df[['W','Z']]
# SQL Syntax (NOT RECOMMENDED!)
df.W
```
DataFrame Columns are just Series
```
type(df['W'])
```
**Creating a new column:**
```
df['new'] = df['W'] + df['Y']
df
```
** Removing Columns**
```
df.drop('new',axis=1)
# Not inplace unless specified!
df
df.drop('new',axis=1,inplace=True)
df
```
Can also drop rows this way:
```
df.drop('E',axis=0)
```
** Selecting Rows**
```
df.loc['A']
```
Or select based off of position instead of label
```
df.iloc[2]
```
** Selecting subset of rows and columns **
```
df.loc['B','Y']
df.loc[['A','B'],['W','Y']]
```
### Conditional Selection
An important feature of pandas is conditional selection using bracket notation, very similar to numpy:
```
df
df>0
df[df>0]
df[df['W']>0]
df[df['W']>0]['Y']
df[df['W']>0][['Y','X']]
```
For two conditions you can use | and & with parenthesis:
```
df[(df['W']>0) & (df['Y'] > 1)]
```
## More Index Details
Let's discuss some more features of indexing, including resetting the index or setting it something else. We'll also talk about index hierarchy!
```
df
# Reset to default 0,1...n index
df.reset_index()
newind = 'CA NY WY OR CO'.split()
df['States'] = newind
df
df.set_index('States')
df
df.set_index('States',inplace=True)
df
```
## Multi-Index and Index Hierarchy
Let us go over how to work with Multi-Index, first we'll create a quick example of what a Multi-Indexed DataFrame would look like:
```
# Index Levels
outside = ['G1','G1','G1','G2','G2','G2']
inside = [1,2,3,1,2,3]
hier_index = list(zip(outside,inside))
hier_index = pd.MultiIndex.from_tuples(hier_index)
hier_index
df = pd.DataFrame(np.random.randn(6,2),index=hier_index,columns=['A','B'])
df
```
Now let's show how to index this! For index hierarchy we use df.loc[], if this was on the columns axis, you would just use normal bracket notation df[]. Calling one level of the index returns the sub-dataframe:
```
df.loc['G1']
df.loc['G1'].loc[1]
df.index.names
df.index.names = ['Group','Num']
df
df.xs('G1')
df.xs(['G1',1])
df.xs(1,level='Num')
```
| github_jupyter |
# Problem set 5: Writing your own algorithms
[<img src="https://mybinder.org/badge_logo.svg">](https://mybinder.org/v2/gh/NumEconCopenhagen/exercises-2020/master?urlpath=lab/tree/PS5/problem_set_5.ipynb)
This problem set has no tasks, but only problems of increasing complexity. See how far you can get :)
```
import math
```
# Factorial
Remember that the factorial of $n$ is
$$
n\cdot(n-1)\cdot(n-2)\cdot...\cdot 1
$$
**Problem:** Correct the following function so that it returns the factorial of n using *functional recursion*.
```
def factorial(n):
if n == 1:
return 1
else:
return n # + missing code
print(factorial(5))
```
**Answer:**
```
def factorial(n):
if n == 1:
return 1
else:
return n*factorial(n-1)
for n in [1,2,3,4,5]:
y = factorial(n)
print(f'the factorial of {n} is {y}')
assert(y == math.factorial(n))
```
# Descending bubble sort
**Problem:** Sort a list of numbers in-place descending (from high to low).
**Inputs:** List of numbers.
**Outputs:** None.
**Algorithm:**
```
L = [54, 26, 93, 17, 77, 31, 44, 55, 20] # test list
# write your code here (hint: use the bubble_sort() algorithm from the lectures)
```
**Answer:**
```
def swap(L,i,j):
temp = L[i] # save value at i
L[i] = L[j] # overwrite value at i with value at j
L[j] = temp # write original value at i to value at j
def bubble_sort(L):
for k in range(len(L)-1,0,-1):
for i in range(k):
if L[i] < L[i+1]:
swap(L,i,i+1)
bubble_sort(L)
print('sorted',L)
```
# Linear search for index
**Problem:** Consider a number `x` and a sorted list of numbers `L`. Assume `L[0] <= x < L[-1]`. Find the index `i` such that `L[i] <= x < L[i+1]` using a linear search.
**Inputs:** A sorted list of numbers `L` and a number `x`.
**Outputs:** Integer.
```
L = [0, 1, 2, 8, 13, 17, 19, 32, 42] # test list
# write your code here (hint: use the linear_seach() algorithm from the lecture)
```
**Answer:**
```
def linear_search(L,x):
# a. prep
i = 0
N = len(L)
found = False
# b. main
while i < N-1 and not found:
if x >= L[i] and x < L[i+1]: # comparison
found = True
else:
i += 1 # increment
# c. return
return i
# test
for x in [3,7,13,18,32]:
i = linear_search(L,x)
print(f'{x} gives the index {i}')
assert(x >= L[i] and x < L[i+1]),(x,i,L[i])
```
# Bisection
**Problem:** Find an (apporximate) solution to $f(x) = 0$ in the interval $[a,b]$ where $f(a)f(b) < 0$ (i.e. one is positive and the other is negative).
> If $f$ is a *continuous* function then the intermediate value theorem ensures that a solution exists.
**Inputs:** Function $f$, float interval $[a,b]$, float tolerance $\epsilon > 0$.
**Outputs:** Float.
**Algorithm:** `bisection()`
1. Set $a_0 = a$ and $b_0 = b$.
2. Compute $f(m_0)$ where $m_0 = (a_0 + b_0)/2$ is the midpoint
3. Determine the next sub-interval $[a_1,b_1]$:
i. If $f(a_0)f(m_0) < 0$ then $a_1 = a_0$ and $b_1 = m_0$
ii. If $f(m_0)f(b_0) < 0$ then $a_1 = m_0$ and $b_1 = b_0$
4. Repeat step 2 and step 3 until $|f(m_n)| < \epsilon$
```
f = lambda x: (2.1*x-1.7)*(x-3.3) # test function
def bisection(f,a,b,tau):
pass
# write your code here
result = bisection(f,0,1,1e-8)
print(result)
```
**Answer:**
```
def bisection(f,a,b,tol=1e-8):
""" bisection
Solve equation f(x) = 0 for a <= x <= b.
Args:
f (function): function
a (float): left bound
b (float): right bound
tol (float): tolerance on solution
Returns:
"""
# test inputs
if f(a)*f(b) >= 0:
print("bisection method fails.")
return None
# step 1: initialize
a_n = a
b_n = b
# step 2-4:
while True:
# step 2: midpoint and associated value
m_n = (a_n+b_n)/2
f_m_n = f(m_n)
# step 3: determine sub-interval
if abs(f_m_n) < tol:
return m_n
elif f(a_n)*f_m_n < 0:
a_n = a_n
b_n = m_n
elif f(b_n)*f_m_n < 0:
a_n = m_n
b_n = b_n
else:
print("bisection method fails.")
return None
return (a_n + b_n)/2
result = bisection(f,0,1,1e-8)
print(f'result is {result:.3f} with f({result:.3f}) = {f(result):.16f}')
```
# Find prime numbers (hard)
**Goal:** Implement a function in Python for the sieve of Eratosthenes.
The **sieve of Eratosthenes** is a simple algorithm for finding all prime numbers up to a specified integer. It was created by the ancient Greek mathematician Eratosthenes. The algorithm to find all the prime numbers less than or equal to a given integer $n$.
**Algorithm:** `sieve_()`
1. Create a list of integers from $2$ to $n$: $2, 3, 4, ..., n$ (all potentially primes)
`primes = list(range(2,n+1))`
2. Start with a counter $i$ set to $2$, i.e. the first prime number
3. Starting from $i+i$, count up by $i$ and remove those numbers from the list, i.e. $2i$, $3i$, $4i$ etc.
`primes.remove(i)`
4. Find the first number of the list following $i$. This is the next prime number.
5. Set $i$ to the number found in the previous step.
6. Repeat steps 3-5 until $i$ is greater than $\sqrt {n}$.
7. All the numbers, which are still in the list, are prime numbers.
**A more detailed explanation:** See this [video](https://www.youtube.com/watch?v=klcIklsWzrY&feature=youtu.be)
```
def sieve(n):
pass # write your code here
print(sieve(100))
```
**Answer:**
```
def sieve(n):
""" sieve of Eratosthenes
Return all primes between 2 and n.
Args:
n (integer): maximum number to consider
"""
# a. step 1: create list of potential primes
primes = list(range(2,n+1))
# b. step 2: initialize i
index = 0
i = primes[index]
# c. step 3-6
while i < math.sqrt(n):
# step 3: remove
k = i
while i <= n:
i += k
if i in primes:
primes.remove(i)
# step 4: next number
index += 1
# step 5: set i
i = primes[index]
return primes
print('primes from 2 to 100:',sieve(100))
```
# More Problems
See [Project Euler](https://projecteuler.net/about).
| github_jupyter |
## Better retrieval via "Dense Passage Retrieval"
### Importance of Retrievers
The Retriever has a huge impact on the performance of our overall search pipeline.
### Different types of Retrievers
#### Sparse
Family of algorithms based on counting the occurences of words (bag-of-words) resulting in very sparse vectors with length = vocab size.
Examples: BM25, TF-IDF
Pros: Simple, fast, well explainable
Cons: Relies on exact keyword matches between query and text
#### Dense
These retrievers use neural network models to create "dense" embedding vectors. Within this family there are two different approaches:
a) Single encoder: Use a **single model** to embed both query and passage.
b) Dual-encoder: Use **two models**, one to embed the query and one to embed the passage
Recent work suggests that dual encoders work better, likely because they can deal better with the different nature of query and passage (length, style, syntax ...).
Examples: REALM, DPR, Sentence-Transformers ...
Pros: Captures semantinc similarity instead of "word matches" (e.g. synonyms, related topics ...)
Cons: Computationally more heavy, initial training of model
### "Dense Passage Retrieval"
In this Tutorial, we want to highlight one "Dense Dual-Encoder" called Dense Passage Retriever.
It was introdoced by Karpukhin et al. (2020, https://arxiv.org/abs/2004.04906.
Original Abstract:
_"Open-domain question answering relies on efficient passage retrieval to select candidate contexts, where traditional sparse vector space models, such as TF-IDF or BM25, are the de facto method. In this work, we show that retrieval can be practically implemented using dense representations alone, where embeddings are learned from a small number of questions and passages by a simple dual-encoder framework. When evaluated on a wide range of open-domain QA datasets, our dense retriever outperforms a strong Lucene-BM25 system largely by 9%-19% absolute in terms of top-20 passage retrieval accuracy, and helps our end-to-end QA system establish new state-of-the-art on multiple open-domain QA benchmarks."_
Paper: https://arxiv.org/abs/2004.04906
Original Code: https://fburl.com/qa-dpr
*Use this [link](https://colab.research.google.com/github/deepset-ai/haystack/blob/master/tutorials/Tutorial6_Better_Retrieval_via_DPR.ipynb) to open the notebook in Google Colab.*
## Prepare environment
### Colab: Enable the GPU runtime
Make sure you enable the GPU runtime to experience decent speed in this tutorial.
**Runtime -> Change Runtime type -> Hardware accelerator -> GPU**
<img src="https://raw.githubusercontent.com/deepset-ai/haystack/master/docs/img/colab_gpu_runtime.jpg">
```
# Make sure you have a GPU running
!nvidia-smi
# Install the latest release of Haystack in your own environment
#! pip install farm-haystack
# Install the latest master of Haystack and install the version of torch that works with the colab GPUs
!pip install git+https://github.com/deepset-ai/haystack.git
!pip install torch==1.5.1+cu101 torchvision==0.6.1+cu101 -f https://download.pytorch.org/whl/torch_stable.html
from haystack import Finder
from haystack.indexing.cleaning import clean_wiki_text
from haystack.indexing.utils import convert_files_to_dicts, fetch_archive_from_http
from haystack.reader.farm import FARMReader
from haystack.reader.transformers import TransformersReader
from haystack.utils import print_answers
```
## Document Store
FAISS is a library for efficient similarity search on a cluster of dense vectors.
The `FAISSDocumentStore` uses a SQL(SQLite in-memory be default) database under-the-hood
to store the document text and other meta data. The vector embeddings of the text are
indexed on a FAISS Index that later is queried for searching answers.
```
from haystack.database.faiss import FAISSDocumentStore
document_store = FAISSDocumentStore()
```
## Cleaning & indexing documents
Similarly to the previous tutorials, we download, convert and index some Game of Thrones articles to our DocumentStore
```
# Let's first get some files that we want to use
doc_dir = "data/article_txt_got"
s3_url = "https://s3.eu-central-1.amazonaws.com/deepset.ai-farm-qa/datasets/documents/wiki_gameofthrones_txt.zip"
fetch_archive_from_http(url=s3_url, output_dir=doc_dir)
# Convert files to dicts
dicts = convert_files_to_dicts(dir_path=doc_dir, clean_func=clean_wiki_text, split_paragraphs=True)
# Now, let's write the dicts containing documents to our DB.
document_store.write_documents(dicts)
```
## Initalize Retriever, Reader, & Finder
### Retriever
**Here:** We use a `DensePassageRetriever`
**Alternatives:**
- The `ElasticsearchRetriever`with custom queries (e.g. boosting) and filters
- Use `EmbeddingRetriever` to find candidate documents based on the similarity of embeddings (e.g. created via Sentence-BERT)
- Use `TfidfRetriever` in combination with a SQL or InMemory Document store for simple prototyping and debugging
```
from haystack.retriever.dense import DensePassageRetriever
retriever = DensePassageRetriever(document_store=document_store, embedding_model="dpr-bert-base-nq",
do_lower_case=True, use_gpu=True, embed_title=True)
# Important:
# Now that after we have the DPR initialized, we need to call update_embeddings() to iterate over all
# previously indexed documents and update their embedding representation.
# While this can be a time consuming operation (depending on corpus size), it only needs to be done once.
# At query time, we only need to embed the query and compare it the existing doc embeddings which is very fast.
document_store.update_embeddings(retriever)
```
### Reader
Similar to previous Tutorials we now initalize our reader.
Here we use a FARMReader with the *deepset/roberta-base-squad2* model (see: https://huggingface.co/deepset/roberta-base-squad2)
#### FARMReader
```
# Load a local model or any of the QA models on
# Hugging Face's model hub (https://huggingface.co/models)
reader = FARMReader(model_name_or_path="deepset/roberta-base-squad2", use_gpu=True)
```
### Finder
The Finder sticks together reader and retriever in a pipeline to answer our actual questions.
```
finder = Finder(reader, retriever)
```
## Voilà! Ask a question!
```
# You can configure how many candidates the reader and retriever shall return
# The higher top_k_retriever, the better (but also the slower) your answers.
prediction = finder.get_answers(question="Who created the Dothraki vocabulary?", top_k_retriever=10, top_k_reader=5)
#prediction = finder.get_answers(question="Who is the father of Arya Stark?", top_k_retriever=10, top_k_reader=5)
#prediction = finder.get_answers(question="Who is the sister of Sansa?", top_k_retriever=10, top_k_reader=5)
print_answers(prediction, details="minimal")
```
| github_jupyter |
# Segundo parcial tema A
__U.N.L.Z. - Facultad de Ingeniería__
__Electrotecnia__
__Alumno:__ Daniel Antonio Lorenzo
<mark><strong>(Resolución en python3)</strong></mark>
<a href="https://colab.research.google.com/github/daniel-lorenzo/Electrotecnia/blob/master/Ejercitacion/2do_parc_tema_A.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
## Pregunta 1
<img src="img/2doparc_A-01.png">
### Resolución
__Datos:__
$\left\{
\begin{array}{l}
S_N = 11000 \, \mathrm{VA} \\
U_{N1} = 400 \, \mathrm{V} \\
U_{N2} = 200 \, \mathrm{V}
\end{array}
\right. \qquad \qquad$
__Ensayo de cortocircuito:__
$\left\{
\begin{array}{l}
P_{cc} = 300 \, \mathrm{W} \\
U_{cc} = 20 \, \mathrm{V} \\
I_{cc} = 27,5 \, \mathrm{A}
\end{array}
\right.$
__Corrientes nominales:__
$$ I_{N1} = \frac{S_N}{U_{N1}} \qquad ; \qquad I_{N2} = \frac{S_N}{U_{N2}} $$
__Relación de transformación:__
$$ \mathrm{a} = \frac{U_{N1}}{U_{N2}} $$
__Ensayo de corto circuito:__
$$ Z_{eq1} = \frac{U_{cc}}{I_{cc}} $$
$$ P_{cc} = I_{cc} U_{cc} \cos \varphi_{cc} \quad \rightarrow \quad \varphi_{cc} = \arccos \left( \frac{P_{cc}}{I_{cc} U_{cc}} \right) $$
__Parámetros longitudinales:__
$$\begin{array}{|l|l|}
\hline
\mbox{Primario} & \mbox{Secundario} \\
\hline
R_{eq1} = Z_{eq1} \cos \varphi_{cc} & R_{eq2} = R_{eq1}/\mathrm{a}^2 \\
X_{eq1} = Z_{eq1} \sin \varphi_{cc} & X_{eq2} = X_{eq1}/\mathrm{a}^2 \\
\hline
\end{array}$$
```
import numpy as np # importa biblioteca numpy
# Datos:
SN = 11000 # [VA] Potencia nominal aparente
UN1 = 400 # [V] Voltaje nominal primario
UN2 = 200 # [V] Voltaje nominal secundario
Pcc = 300 # [W] Potencia de cortocircuito
Ucc = 20 # [V] Voltaje de cortocircuito
Icc = 27.5 # [A] Corriente de cortocircuito
# Cálculos:
IN1 = SN/UN1 # [A] Corriente nominal 1
IN2 = SN/UN2 # [A] Corriente nominal 2
a = UN1/UN2 # Relación de transformación
# Ensayo de cortocircuito:
Zeq1 = Ucc/Icc
phi_cc = np.arccos(Pcc/(Icc*Ucc))
# Parámetros longitudinales
# Primario # Secundario
Req1 = Zeq1*np.cos(phi_cc) ; Req2 = Req1/a**2
Xeq1 = Zeq1*np.sin(phi_cc) ; Xeq2 = Xeq1/a**2
print('Resultado:')
print('Req2 = %.3f Ohm ; Xeq2 = %.3f Ohm'%(Req2,Xeq2))
```
## Pregunta 2
<img src="img/2doparc_A-02.png">
### Resolución
__Ensayo de vacío:__
$\left\{
\begin{array}{l}
P_0 = 110 \, \mathrm{W} \\
U_0 = 200 \, \mathrm{V} \\
I_0 = 2 \, \mathrm{A}
\end{array}
\right.$
__Cálculo de $\varphi_0$__
$$ P_0 = I_0 U_0 \cos \varphi_0 \qquad \rightarrow \qquad \varphi_0 = \arccos \left( \frac{P_0}{I_0 U_0} \right) $$
$\begin{array}{l}
I_p = I_0 \cos \varphi_0 \\
I_m = I_0 \sin \varphi_0
\end{array}$
$$\begin{array}{|l|l|}
\hline
\mbox{Secundario (BT)} & \mbox{Primario (AT)} \\
\hline
R_{p2} = U_0 / I_p & R_{p1} = R_{p2} \cdot \mathrm{a}^2 \\
X_{m2} = U_0 / I_m & X_{eq2} = X_{m1} \cdot \mathrm{a}^2 \\
\hline
\end{array}$$
```
# Datos
Po = 110 # [W]
Uo = 200 # [V]
Io = 2 # [A]
# Ensayo de vacío
phi_o = np.arccos(Po/(Io*Uo))
Ip = Io*np.cos(phi_o)
Im = Io*np.sin(phi_o)
# Secundario (BT) # Primario (AT)
Rp2 = Uo/Ip ; Rp1 = Rp2*a**2
Xm2 = Uo/Im ; Xm1 = Xm2*a**2
print('Resultado:')
print('Rp2 = %.2f Ohm ; Xm2 = %.2f Ohm'%(Rp2,Xm2))
```
## Pregunta 3
<img src="img/2doparc_A-03.png">
### Resolución
__Datos:__
$\left\{
\begin{array}{l}
\mathrm{fp_{reg}} = 0,7 \\
\mathrm{fp_{rend} = 0,7}
\end{array}
\right.$
__Regulación:__
$$\begin{array}{lcl}
\mathrm{fp_{reg}} = \cos \varphi_\mathrm{reg} & \quad \Rightarrow \quad & \varphi_\mathrm{reg} = \arccos \mathrm{fp_{reg}} \\
I_\mathrm{2reg} = \mathrm{fp_{reg}} I_{N2} & \quad \wedge \quad & U_{20} = U_0
\end{array}$$
__Tensión de salida o aplicada a la carga:__
$$ U_\mathrm{2reg,ind} = U_{20} - I_\mathrm{2reg} (R_{eq2} \cos \varphi_\mathrm{reg} + X_{eq2} \sin \varphi_\mathrm{reg}) $$
__Regulación:__
$$ \mathrm{reg_{ind}} = \mathrm{ \frac{U_{20} - U_{2reg,ind}}{U_{20}} \times 100} $$
```
# Datos:
fp_reg = 0.7
fp_rend = 0.7
# Cálculos de regulación
phi_reg = np.arccos(fp_reg)
I2_reg = fp_reg*IN2 ; U20 = Uo
# Tensión de salida o aplicada a la carga:
U2reg_ind = U20 - I2_reg*(Req2*np.cos(phi_reg) + Xeq2*np.sin(phi_reg))
# Regulación
reg_ind = (U20 - U2reg_ind)/U20*100
print('Resultado:')
print('reg_ind = %.2f'%reg_ind)
```
## Pregunta 4
<img src="img/2doparc_A-04.png">
### Resolución
__Datos:__
$\left\{
\begin{array}{l}
\mathrm{fp_{reg}} = 0,7 \\
\mathrm{fp_{rend} = 0,8}
\end{array}
\right.$
__Regulación:__
$$\begin{array}{lcl}
\mathrm{fp_{reg}} = \cos \varphi_\mathrm{reg} & \quad \Rightarrow \quad & \varphi_\mathrm{reg} = \arccos \mathrm{fp_{reg}} \\
I_\mathrm{2reg} = \mathrm{fp_{reg}} I_{N2} & \quad \wedge \quad & U_{20} = U_0
\end{array}$$
__Tensión de salida o aplicada a la carga:__
$$ U_\mathrm{2reg,ind} = U_{20} - I_\mathrm{2reg} (R_{eq2} \cos \varphi_\mathrm{reg} + X_{eq2} \sin \varphi_\mathrm{reg}) $$
__Regulación:__
$$ \mathrm{reg_{ind}} = \mathrm{ \frac{U_{20} - U_{2reg,ind}}{U_{20}} \times 100} $$
__Cálculos de rendimiento__
$$ \mathrm{fp_{rend}} = \cos \mathrm{\varphi_{rend}} \qquad \rightarrow \qquad \mathrm{\varphi_{rend} = \arccos fp_{rend}} $$
$I_\mathrm{2rend} = I_{N2}$
$P_{cu} = I_\mathrm{2rend}^2 R_{eq2}$
__Rendimiento para el caso inductivo__
$$ U_\mathrm{2rend,ind} = U_{20} - I_\mathrm{2rend} (R_{eq2} \cos \varphi_\mathrm{rend} + X_{eq2} \sin \varphi_\mathrm{rend}) $$
$$ \eta_\mathrm{ind} = \frac{U_\mathrm{2rend,ind} I_\mathrm{2rend} \cos \varphi_\mathrm{rend}}{U_\mathrm{2rend,ind} I_\mathrm{2rend} \cos \varphi_\mathrm{rend} + P_{cu} + P_0} $$
```
# Datos:
fp_reg = 0.7
fp_rend = 0.8
# Cálculos de regulación
phi_reg = np.arccos(fp_reg)
I2_reg = fp_reg*IN2 ; U20 = Uo
# Tensión de salida o aplicada a la carga:
U2reg_ind = U20 - I2_reg*(Req2*np.cos(phi_reg) + Xeq2*np.sin(phi_reg))
# Regulación
reg_ind = (U20 - U2reg_ind)/U20*100
# Cálculos de rendimiento
phi_rend = np.arccos(fp_rend)
I2_rend = IN2
Pcu = I2_rend**2*Req2
# Rendimiento para el caso inductivo:
U2rend_ind = U20 - I2_rend*(Req2*np.cos(phi_rend) + Xeq2*np.sin(phi_rend))
n_ind = (U2rend_ind*I2_rend*np.cos(phi_rend))/(U2rend_ind*I2_rend*np.cos(phi_rend) + Pcu + Po)
print('Resultado:')
print('n_ind = %.2f'%n_ind)
```
## Pregunta 5
<img src="img/2doparc_A-05.png">
### Resolución
__Dato:__ $\quad \rightarrow \quad \mathrm{fp_{max} = 1}$
$$ \mathrm{fp_{max} = \cos \varphi_{max}} \qquad \rightarrow \qquad \varphi_\mathrm{max} = \arccos \mathrm{fp_{max}} $$
$$ I_\mathrm{2max} = \sqrt{ \frac{P_0}{R_{eq2}} } $$
$P_\mathrm{cu,max} = I_\mathrm{2max}^2 R_{eq2}$
$$ U_\mathrm{2,max} = U_{N2} - I_\mathrm{2max} (R_{eq2} \cos \varphi_\mathrm{max} + X_\mathrm{eq2} \underbrace{ \sin \varphi_\mathrm{max} }_{\rightarrow \, 0}) $$
__Rendimiento máximo:__
$$ \eta_\mathrm{max} = \frac{U_\mathrm{2max} I_\mathrm{2max} \mathrm{fp_{max}}}{U_\mathrm{2max} I_\mathrm{2max} \mathrm{fp_{max}} + P_\mathrm{cu,max} + P_0} $$
```
# Dato:
fp_max = 1 # factor de potencia máximo, (resistivo puro)
phi_max = np.arccos(fp_max)
I2_max = np.sqrt(Po/Req2)
Pcu_max = I2_max**2*Req2
U2_max = UN2 - I2_max*(Req2*np.cos(phi_max) + Xeq2*np.sin(phi_max))
n_max = (U2_max*I2_max*fp_max)/(U2_max*I2_max*fp_max + Pcu_max + Po)
print('Resulatado:')
print('n_max = %.3f'%n_max)
```
## Pregunta 6
<img src="img/2doparc_A-06.png">
## Pregunta 7
<img src="img/2doparc_A-07.png">
$$ S = \sqrt{3} \cdot U_\mathrm{Linea} I_\mathrm{Linea} $$
## Pregunta 8
<img src="img/2doparc_A-08.png">
## Pregunta 9
<img src="img/2doparc_A-09.png">
## Pregunta 10
<img src="img/2doparc_A-10.png">
### Resolución
$$ S_t^2 = (P_r + P_s + P_t)^2 + (Q_r + Q_s + Q_t)^2 $$
$$ 100^2 = (11 + 11 + P_t)^2 + (11 + 11 + 11)^2 $$
$$ 100^2 = (22 + P_t)^2 + (33)^2 $$
$$ P_t = \sqrt{(100)^2 - (33)^2} - 22 $$
$$ P_t = 72,398 \, \mathrm{kW} $$
```
Pt = np.sqrt(100**2 - 33**2) - 22
print('Resultado:')
print('Pt = %.3f kW'%(Pt))
```
----------
<a href="https://colab.research.google.com/github/daniel-lorenzo/Electrotecnia/blob/master/Ejercitacion/2do_parc_tema_A.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
| github_jupyter |
# Initialization
```
import re
from collections import Counter
def words(text): return re.findall(r'\w+', text.lower())
WORDS = Counter(words(open('../input/twitterdataset2/big.txt').read()))
def P(word, N=sum(WORDS.values())):
"Probability of `word`."
return WORDS[word] / N
def correction(word):
"Most probable spelling correction for word."
return max(candidates(word), key=P)
def candidates(word):
"Generate possible spelling corrections for word."
return (known([word]) or known(edits1(word)) or known(edits2(word)) or [word])
def known(words):
"The subset of `words` that appear in the dictionary of WORDS."
return set(w for w in words if w in WORDS)
def edits1(word):
"All edits that are one edit away from `word`."
letters = 'abcdefghijklmnopqrstuvwxyz'
splits = [(word[:i], word[i:]) for i in range(len(word) + 1)]
deletes = [L + R[1:] for L, R in splits if R]
transposes = [L + R[1] + R[0] + R[2:] for L, R in splits if len(R)>1]
replaces = [L + c + R[1:] for L, R in splits if R for c in letters]
inserts = [L + c + R for L, R in splits for c in letters]
return set(deletes + transposes + replaces + inserts)
def edits2(word):
"All edits that are two edits away from `word`."
return (e2 for e1 in edits1(word) for e2 in edits1(e1))
#Naive Bayes
from nltk.corpus import words as wsss
import pandas as pd
from nltk.tokenize import WordPunctTokenizer
import re
from bs4 import BeautifulSoup
from nltk.stem import PorterStemmer
from nltk.tokenize import sent_tokenize, word_tokenize
from sklearn.tree import DecisionTreeClassifier
from nltk.corpus import stopwords
import nltk.classify.util
from nltk.classify import NaiveBayesClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.svm import LinearSVC
from sklearn.neighbors import KNeighborsClassifier
from nltk.classify import SklearnClassifier
from nltk.corpus import names
import numpy as np
import time
import matplotlib.pyplot as plt
from pandas import *
wordss=[]
for i in wsss.words():
wordss.append(i)
start = time.time()
num=0
df = read_csv("../input/twitterdataset/traindata.csv")
df1 = df['tag']
df2 = df['content']
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
f= open("processed_dataset.csv","w")
def clean(s):
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
b=tweet_cleaner(s)
test_result.append(b)
words = word_tokenize(tweet_cleaner(s))
#print(words)
wordsFiltered=[]
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
f.write(',')
for i in tmp:
f.write(i+' ')
f.write('\n')
#print(li)
return tmp
#return words
p=[]
s=[]
n=[]
c=[]
counterz = 0
for t in range(0,len(df2)):
if counterz%100==0:
print(counterz)
counterz=counterz+1
if df1[t]=='politics':
sen=df2[t]
f.write('0')
p=p+clean(sen)
elif df1[t]=='sports':
sen=df2[t]
f.write('1')
s=s+clean(sen)
elif df1[t]=='natural':
sen=df2[t]
f.write('2')
n=n+clean(sen)
elif df1[t]=='crime':
sen=df2[t]
f.write('3')
c=c+clean(sen)
f.close()
#print(p)
#print(s)
#print(n)
#print(c)
normal_vocab=[]
def word_feats(words):
return dict([(words, True)])
for a in p:
if a in p and a in s and a in n and a in c:
normal_vocab=normal_vocab+list(a);
p.remove(a)
s.remove(a)
n.remove(a)
c.remove(a)
if a in p and a in s:
normal_vocab=normal_vocab+list(a);
p.remove(a)
s.remove(a)
if a in s and a in n:
normal_vocab=normal_vocab+list(a);
n.remove(a)
s.remove(a)
if a in n and a in c:
normal_vocab=normal_vocab+list(a);
n.remove(a)
c.remove(a)
if a in p and a in c:
normal_vocab=normal_vocab+list(a);
p.remove(a)
c.remove(a)
if a in s and a in c:
normal_vocab=normal_vocab+list(a);
c.remove(a)
s.remove(a)
if a in p and a in n:
normal_vocab=normal_vocab+list(a);
p.remove(a)
n.remove(a)
if a in p and a in s and a in n:
normal_vocab=normal_vocab+list(a);
p.remove(a)
s.remove(a)
n.remove(a)
if a in s and a in n and a in c:
normal_vocab=normal_vocab+list(a);
c.remove(a)
s.remove(a)
n.remove(a)
if a in p and a in n and a in c:
normal_vocab=normal_vocab+list(a);
p.remove(a)
c.remove(a)
n.remove(a)
if a in p and a in s and a in c:
normal_vocab=normal_vocab+list(a);
p.remove(a)
s.remove(a)
c.remove(a)
politics_vocab=p
#print(politics_vocab)
sports_vocab=s
natural_vocab=n
crime_vocab=c
politics_features = [(word_feats(pol), 'pol') for pol in politics_vocab]
sports_features = [(word_feats(spo), 'spo') for spo in sports_vocab]
natural_features = [(word_feats(nat), 'nat') for nat in natural_vocab]
crime_features = [(word_feats(cri), 'cri') for cri in crime_vocab]
normal_features = [(word_feats(nor), 'nor') for nor in normal_vocab]
train_set = politics_features + sports_features + natural_features+crime_features
#print(train_set)
```
# Naive Bayes
```
classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
#classifier = SklearnClassifier(LinearSVC(tol=1e-6, max_iter=20000, class_weight='balanced', C=100, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
#classifier = SklearnClassifier(RandomForestClassifier(n_estimators=5, max_depth=10, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/test.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# SVM
```
#classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
classifier = SklearnClassifier(LinearSVC(tol=1e-6, max_iter=20000, class_weight='balanced', C=100, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
#classifier = SklearnClassifier(RandomForestClassifier(n_estimators=5, max_depth=10, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/testv2.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2v2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# SVM, tol=10e-5
```
#classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
classifier = SklearnClassifier(LinearSVC(tol=1e-5, max_iter=20000, class_weight='balanced', C=100, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
#classifier = SklearnClassifier(RandomForestClassifier(n_estimators=5, max_depth=10, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/testv2.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2v2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# SVM, tol=10e-7
```
#classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
classifier = SklearnClassifier(LinearSVC(tol=1e-7, max_iter=20000, class_weight='balanced', C=100, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
#classifier = SklearnClassifier(RandomForestClassifier(n_estimators=5, max_depth=10, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/testv2.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2v2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# SVM, max_iter=10000
```
#classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
classifier = SklearnClassifier(LinearSVC(tol=1e-6, max_iter=10000, class_weight='balanced', C=100, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
#classifier = SklearnClassifier(RandomForestClassifier(n_estimators=5, max_depth=10, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/testv2.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2v2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# SVM, max_iter=30000
```
#classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
classifier = SklearnClassifier(LinearSVC(tol=1e-6, max_iter=30000, class_weight='balanced', C=100, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
#classifier = SklearnClassifier(RandomForestClassifier(n_estimators=5, max_depth=10, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/testv2.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2v2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# SVM, no class weight
```
#classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
classifier = SklearnClassifier(LinearSVC(tol=1e-6, max_iter=20000, C=100, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
#classifier = SklearnClassifier(RandomForestClassifier(n_estimators=5, max_depth=10, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/testv2.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2v2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# SVM, C=10
```
#classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
classifier = SklearnClassifier(LinearSVC(tol=1e-6, max_iter=20000, class_weight='balanced', C=10, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
#classifier = SklearnClassifier(RandomForestClassifier(n_estimators=5, max_depth=10, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/testv2.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2v2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# SVM, C=1000
```
#classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
classifier = SklearnClassifier(LinearSVC(tol=1e-6, max_iter=20000, class_weight='balanced', C=1000, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
#classifier = SklearnClassifier(RandomForestClassifier(n_estimators=5, max_depth=10, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/testv2.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2v2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# SVM, no loss
```
#classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
classifier = SklearnClassifier(LinearSVC(tol=1e-6, max_iter=20000, class_weight='balanced', C=100, multi_class='ovr', random_state=0), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
#classifier = SklearnClassifier(RandomForestClassifier(n_estimators=5, max_depth=10, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/testv2.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2v2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# Logistic Regression
```
#classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
#classifier = SklearnClassifier(LinearSVC(tol=1e-6, max_iter=20000, class_weight='balanced', C=100, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
#classifier = SklearnClassifier(RandomForestClassifier(n_estimators=5, max_depth=10, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/test.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# Decision Tree
```
#classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
#classifier = SklearnClassifier(LinearSVC(tol=1e-6, max_iter=20000, class_weight='balanced', C=100, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
#classifier = SklearnClassifier(RandomForestClassifier(n_estimators=5, max_depth=10, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/testv2.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2v2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# Random Forest
```
#classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
#classifier = SklearnClassifier(LinearSVC(tol=1e-6, max_iter=20000, class_weight='balanced', C=100, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
classifier = SklearnClassifier(RandomForestClassifier(n_estimators=5, max_depth=10, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/test.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# Random Forest, n_estimator=3
```
#classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
#classifier = SklearnClassifier(LinearSVC(tol=1e-6, max_iter=20000, class_weight='balanced', C=100, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
classifier = SklearnClassifier(RandomForestClassifier(n_estimators=3, max_depth=10, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/test.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# Random Forest, n_estimators=7
```
#classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
#classifier = SklearnClassifier(LinearSVC(tol=1e-6, max_iter=20000, class_weight='balanced', C=100, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
classifier = SklearnClassifier(RandomForestClassifier(n_estimators=7, max_depth=10, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/test.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# Random Forest, max_depth=5
```
#classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
#classifier = SklearnClassifier(LinearSVC(tol=1e-6, max_iter=20000, class_weight='balanced', C=100, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
classifier = SklearnClassifier(RandomForestClassifier(n_estimators=5, max_depth=5, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/test.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# Random Forest, max_depth=15
```
#classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
#classifier = SklearnClassifier(LinearSVC(tol=1e-6, max_iter=20000, class_weight='balanced', C=100, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
classifier = SklearnClassifier(RandomForestClassifier(n_estimators=5, max_depth=15, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/test.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# KNN
```
#classifier = NaiveBayesClassifier.train(train_set)
##classifier = SklearnClassifier(LinearSVC(random_state=0), sparse=False).train(train_set)
#classifier = SklearnClassifier(LinearSVC(tol=1e-6, max_iter=20000, class_weight='balanced', C=100, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
#classifier = SklearnClassifier(RandomForestClassifier(n_estimators=5, max_depth=10, random_state=0, criterion="entropy"), sparse=False).train(train_set)
classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/testv2.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2v2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
#print(words)
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
#print(wordsFiltered)
for word in wordsFiltered:
l=ps.stem(word)
li.append(l)
#print(li)
tmp=[]
for i in li:
if i not in tmp:
if i!='b':
if len(i)==1:
continue
elif len(i)<3:
if i in wordss:
tmp.append(correction(i))
else:
tmp.append(correction(i))
sentence = tmp
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
# SVM [18]
```
import re
from collections import Counter
def words(text): return re.findall(r'\w+', text.lower())
WORDS = Counter(words(open('../input/twitterdataset2/big.txt').read()))
def P(word, N=sum(WORDS.values())):
"Probability of `word`."
return WORDS[word] / N
def correction(word):
"Most probable spelling correction for word."
return max(candidates(word), key=P)
def candidates(word):
"Generate possible spelling corrections for word."
return (known([word]) or known(edits1(word)) or known(edits2(word)) or [word])
def known(words):
"The subset of `words` that appear in the dictionary of WORDS."
return set(w for w in words if w in WORDS)
def edits1(word):
"All edits that are one edit away from `word`."
letters = 'abcdefghijklmnopqrstuvwxyz'
splits = [(word[:i], word[i:]) for i in range(len(word) + 1)]
deletes = [L + R[1:] for L, R in splits if R]
transposes = [L + R[1] + R[0] + R[2:] for L, R in splits if len(R)>1]
replaces = [L + c + R[1:] for L, R in splits if R for c in letters]
inserts = [L + c + R for L, R in splits for c in letters]
return set(deletes + transposes + replaces + inserts)
def edits2(word):
"All edits that are two edits away from `word`."
return (e2 for e1 in edits1(word) for e2 in edits1(e1))
#Naive Bayes
from nltk.corpus import words as wsss
import pandas as pd
from nltk.tokenize import WordPunctTokenizer
import re
from bs4 import BeautifulSoup
from nltk.stem import PorterStemmer
from nltk.tokenize import sent_tokenize, word_tokenize
from sklearn.tree import DecisionTreeClassifier
from nltk.corpus import stopwords
import nltk.classify.util
from nltk.classify import NaiveBayesClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.svm import LinearSVC
from sklearn.neighbors import KNeighborsClassifier
from nltk.classify import SklearnClassifier
from nltk.corpus import names
import numpy as np
import time
import matplotlib.pyplot as plt
from pandas import *
wordss=[]
for i in wsss.words():
wordss.append(i)
start = time.time()
num=0
df = read_csv("../input/twitterdataset/traindata.csv")
df1 = df['tag']
df2 = df['content']
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
f= open("processed_dataset.csv","w")
def clean(s):
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
b=tweet_cleaner(s)
test_result.append(b)
words = word_tokenize(tweet_cleaner(s))
#print(words)
return words
p=[]
s=[]
n=[]
c=[]
counterz = 0
for t in range(0,len(df2)):
counterz=counterz+1
if df1[t]=='politics':
sen=df2[t]
f.write('0')
p=p+clean(sen)
elif df1[t]=='sports':
sen=df2[t]
f.write('1')
s=s+clean(sen)
elif df1[t]=='natural':
sen=df2[t]
f.write('2')
n=n+clean(sen)
elif df1[t]=='crime':
sen=df2[t]
f.write('3')
c=c+clean(sen)
f.close()
#print(p)
#print(s)
#print(n)
#print(c)
normal_vocab=[]
def word_feats(words):
return dict([(words, True)])
for a in p:
if a in p and a in s and a in n and a in c:
normal_vocab=normal_vocab+list(a);
p.remove(a)
s.remove(a)
n.remove(a)
c.remove(a)
if a in p and a in s:
normal_vocab=normal_vocab+list(a);
p.remove(a)
s.remove(a)
if a in s and a in n:
normal_vocab=normal_vocab+list(a);
n.remove(a)
s.remove(a)
if a in n and a in c:
normal_vocab=normal_vocab+list(a);
n.remove(a)
c.remove(a)
if a in p and a in c:
normal_vocab=normal_vocab+list(a);
p.remove(a)
c.remove(a)
if a in s and a in c:
normal_vocab=normal_vocab+list(a);
c.remove(a)
s.remove(a)
if a in p and a in n:
normal_vocab=normal_vocab+list(a);
p.remove(a)
n.remove(a)
if a in p and a in s and a in n:
normal_vocab=normal_vocab+list(a);
p.remove(a)
s.remove(a)
n.remove(a)
if a in s and a in n and a in c:
normal_vocab=normal_vocab+list(a);
c.remove(a)
s.remove(a)
n.remove(a)
if a in p and a in n and a in c:
normal_vocab=normal_vocab+list(a);
p.remove(a)
c.remove(a)
n.remove(a)
if a in p and a in s and a in c:
normal_vocab=normal_vocab+list(a);
p.remove(a)
s.remove(a)
c.remove(a)
politics_vocab=p
#print(politics_vocab)
sports_vocab=s
natural_vocab=n
crime_vocab=c
politics_features = [(word_feats(pol), 'pol') for pol in politics_vocab]
sports_features = [(word_feats(spo), 'spo') for spo in sports_vocab]
natural_features = [(word_feats(nat), 'nat') for nat in natural_vocab]
crime_features = [(word_feats(cri), 'cri') for cri in crime_vocab]
normal_features = [(word_feats(nor), 'nor') for nor in normal_vocab]
train_set = politics_features + sports_features + natural_features+crime_features
#print(train_set)
#classifier = NaiveBayesClassifier.train(train_set)
classifier = SklearnClassifier(LinearSVC(max_iter=20000), sparse=False).train(train_set)
#classifier = SklearnClassifier(LinearSVC(tol=1e-6, max_iter=20000, class_weight='balanced', C=100, multi_class='ovr', random_state=0, loss='hinge'), sparse=False).train(train_set)
#classifier = SklearnClassifier(LogisticRegression(), sparse=False).train(train_set)
#classifier = SklearnClassifier(DecisionTreeClassifier(), sparse=False).train(train_set)
#classifier = SklearnClassifier(RandomForestClassifier(n_estimators=5, max_depth=10, random_state=0, criterion="entropy"), sparse=False).train(train_set)
#classifier = SklearnClassifier(KNeighborsClassifier(), sparse=False).train(train_set)
cols = ['text']
dt = pd.read_csv("../input/twitterdataset/test.csv",header=None, names=cols)
dtm = read_csv("../input/twitterdataset/testmatch2.csv")
dtm1 = dtm['tag']
dtm2 = dtm['content']
#print(classifier)
# Predict
tok = WordPunctTokenizer()
pat1 = r'@[A-Za-z0-9]+'
pat2 = r'https?://[A-Za-z0-9./]+'
combined_pat = r'|'.join((pat1, pat2))
def tweet_cleaner(text):
soup = BeautifulSoup(text, 'lxml')
souped = soup.get_text()
stripped = re.sub(combined_pat, '', souped)
try:
clean = stripped.decode("utf-8-sig").replace(u"\ufffd", "?")
except:
clean = stripped
letters_only = re.sub("[^a-zA-Z]", " ", clean)
lower_case = letters_only.lower()
words = tok.tokenize(lower_case)
return (" ".join(words)).strip()
testing = dt.text[:]
stopWords = set(stopwords.words('english'))
ps = PorterStemmer()
li=[]
test_result = []
wordsFiltered = []
c=0
n=0
po=0
sp=0
cr=0
na=0
poli=0
spor=0
natu=0
crim=0
o=0
counterz=0
import numpy as np
actual = []
predicted = []
for t in testing:
if counterz%20==0:
print(counterz)
counterz=counterz+1
pol=0
cri=0
nat=0
spo=0
pol2=0
cri2=0
nat2=0
spo2=0
#print(t)
#print(wd)
b=tweet_cleaner(t)
#print(b)
wd = word_tokenize(b)
test_result.append(b)
words = word_tokenize(tweet_cleaner(t))
sentence = words
#print(sentence)
#sentence = sentence.lower()
#words = sentence.split(' ')
w=sentence
wordsFiltered=[]
for word in w:
classResult = classifier.classify( word_feats(word))
if classResult == 'pol':
pol = pol + 1
if classResult == 'spo':
spo = spo + 1
if classResult == 'nat':
nat = nat + 1
if classResult == 'cri':
cri = cri + 1
#print(neg)
#print(pos)
#print(li)
#print(len(words))
pol2=float(pol)/(len(wd))
spo2=float(spo)/(len(wd))
nat2=float(nat)/(len(wd))
cri2=float(cri)/(len(wd))
#print('Politics: ' + str(pol2))
#print('Sports: ' + str(spo2))
#print('Natural: ' + str(nat2))
#print('Crime: ' + str(cri2))
if dtm1[o]=='politics':
actual.append(0)
elif dtm1[o]=='sports':
actual.append(1)
elif dtm1[o]=='natural':
actual.append(2)
elif dtm1[o]=='crime':
actual.append(3)
if pol2>=spo2 and pol2>=nat2 and pol2>=cri2:
po=po+1
predicted.append(0)
if dtm1[o]=='politics':
poli=poli+1
#print('politics')
elif spo2>=pol2 and spo2>=nat2 and spo2>=cri2:
sp=sp+1
predicted.append(1)
if dtm1[o]=='sports':
spor=spor+1
#print('sports')
elif nat2>=spo2 and nat2>=pol2 and nat2>=cri2:
na=na+1
predicted.append(2)
if dtm1[o]=='natural':
natu=natu+1
#print('natural')
elif cri2>=spo2 and cri2>=nat2 and cri2>=pol2:
cr=cr+1
predicted.append(3)
if dtm1[o]=='crime':
crim=crim+1
#print('crime')
#print(li)
while len(li)>0:
li.pop()
c=c+float(pol2+spo2+nat2+cri2)
n=n+1
o=o+1
#np.save("actual.npy", np.array(actual))
#np.save("SVMpredicted.npy", np.array(predicted))
print(actual)
print(predicted)
politics_count=0
sports_count=0
natural_count=0
crime_count=0
for i in range(0, len(testing)):
if dtm1[i]=='politics':
politics_count=politics_count+1
elif dtm1[i]=='sports':
sports_count=sports_count+1
elif dtm1[i]=='natural':
natural_count=natural_count+1
elif dtm1[i]=='crime':
crime_count=crime_count+1
print('Politics Count: ' + str(politics_count))
print('Sports Count: ' + str(sports_count))
print('Natural Count: ' + str(natural_count))
print('Crime Count: ' + str(crime_count))
print()
ac=poli+spor+crim+natu
print('Politics Predicted: ' + str(po))
print('Sports Predicted: ' + str(sp))
print('Natural Predicted: ' + str(na))
print('Crime Predicted: ' + str(cr))
print()
print('Politics Correct: ' + str(poli))
print('Sports Correct: ' + str(spor))
print('Natural Correct: ' + str(natu))
print('Crime Correct: ' + str(crim))
print()
print('Test Data: ' + str(n))
print('Accuracy: ' + str(ac/n))
end = time.time()
print('Run Time: ' +str(end - start))# -*- coding: utf-8 -*-
```
| github_jupyter |
```
import json
import os
import sys
import warnings
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from datetime import datetime
from imblearn.over_sampling import SMOTE
from IPython.display import clear_output
from pprint import pprint
from sklearn.preprocessing import StandardScaler
from sklearn.impute import SimpleImputer as Imputer
from sklearn.pipeline import Pipeline
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import accuracy_score, log_loss, f1_score, recall_score, precision_score
from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score
warnings.filterwarnings('ignore')
sys.path.append(os.path.join('..', 'src'))
import importlib
import utils, model, params, s05_2_feature_engineering
importlib.reload(model)
from model import get_model_params, timer, measure_prediction_time, apply_ml_model, save_model_parameters, save_model_metrics
importlib.reload(s05_2_feature_engineering)
from s05_2_feature_engineering import build_polynomials, transform_label, treat_skewness
```
# define functions
notice that I had to build 2 functions for xgboost. The second one applies hyperparameter tuning.
There are some other refs for a different approach:
* https://www.kaggle.com/kabure/extensive-eda-and-modeling-xgb-hyperopt
```
def apply_ml_model(alg, X_train_set, y_train_set, cols,
encoding='ordinal', treat_collinearity = False, build_polynomals_method=None, do_transform_label=False,
do_treat_skewness=False, smote=False,
useTrainCV=True, cv_folds=5, early_stopping_rounds=50, verbosity=False):
'''
\n
'''
start_time = datetime.now()
X_train_set = X_train_set[cols]
if smote:
oversampling = SMOTE(sampling_strategy = 0.5, k_neighbors=5, random_state=42)
X_train_set, y_train_set = oversampling.fit_resample(X_train_set, y_train_set)
X_train_set = pd.DataFrame(X_train_set, columns = cols)
if build_polynomals_method:
X_train_set = build_polynomials(X_train_set, ProjectParameters().numerical_cols, method = build_polynomals_method)
if do_transform_label:
y_train_set = transform_label(y_train_set, do_transform_label)
if do_treat_skewness:
X_train_set = treat_skewness(X_train_set, set_name)
predictors = X_train_set.columns.to_list()
if useTrainCV:
xgb_param = alg.get_xgb_params()
xgtrain = xgb.DMatrix(X_train_set[predictors].values, label=y_train_set)
# metrics = ['rmse']
metrics = ['logloss']
cvresult = xgb.cv(xgb_param, xgtrain, num_boost_round=alg.get_params()['n_estimators'], nfold=cv_folds,
metrics=metrics, early_stopping_rounds=early_stopping_rounds, verbose_eval=verbosity)
alg.set_params(n_estimators=cvresult.shape[0])
#Fit the algorithm on the data
alg.fit(X_train_set[predictors], y_train_set,eval_metric='auc')
#Predict training set:
X_train_set_predictions = alg.predict(X_train_set[predictors])
# best_score = mean_squared_error(y_train_set, X_train_set_predictions)
# best_score = accuracy_score(y_train_set, X_train_set_predictions)
best_score = f1_score(y_train_set, X_train_set_predictions)
#Print model report:
print ("\nModel Report")
print ("Score : %.4g" % best_score)
# print ("R2 : %.4g" % r2_score(y_train_set, X_train_set_predictions))
print ("f1 : %.4g" % f1_score(y_train_set, X_train_set_predictions))
train_time = round(timer(start_time),9)
prediction_time = measure_prediction_time(alg, X_train_set)
return alg, best_score, train_time, prediction_time, X_train_set
def gridsearch_and_save(model_type, models, cols, build_polynomals_method=False,
do_treat_skewness=False, smote=False):
ml_dict = {model_type: {}}
grid_results = {}
count = 0
for k,model in models.items():
# print(model)
# report step
count += 1
clear_output(wait=True)
print('grid {} of {}'.format(count, len(models)))
# apply model with parameter from grid step
grid_results[k] = {'best_score': {}, 'train_time': {}, 'prediction_time': {}, 'best_params': {}}
(clf,
grid_results[k]['best_score'],
grid_results[k]['train_time'],
grid_results[k]['prediction_time'],
data_transformed) = apply_ml_model(model, X_train, y_train, columns, encoding='ordinal',
build_polynomals_method=build_polynomals_method,
do_treat_skewness=do_treat_skewness, smote=smote
)
grid_results[k]['best_params'] = clf.get_params()
ml_dict[model_type]['columns'] = columns
# search for best score on grid and save data and parameters
best_score = 0
for k,v in grid_results.items():
score = v['best_score']
if score >= best_score:
best_score = score
ml_dict[model_type]['best_score'] = v['best_score']
ml_dict[model_type]['train_time'] = v['train_time']
ml_dict[model_type]['prediction_time'] = v['prediction_time']
ml_dict[model_type]['best_params'] = v['best_params']
save_model_parameters(models_reports, model_type, ml_dict[model_type]['best_params'])
save_model_metrics(model_outputs, model_type, ml_dict)
# print('Model Report')
# print('score:', ml_dict[model_type]['best_score'])
return clf
```
# Define paths
```
inputs = os.path.join('..', 'data', '03_processed')
outputs = os.path.join('..', 'data', '03_processed')
models_reports = os.path.join('..', 'data', '04_models')
model_outputs = os.path.join('..', 'data', '05_model_output')
reports = os.path.join('..', 'data', '06_reporting')
```
# Data capture
As dataset is small, we might export data to a pandas dataframe.
```
inputs = os.path.join('..', 'data', '03_processed')
models_reports = os.path.join('..', 'data', '04_models')
model_outputs = os.path.join('..', 'data', '05_model_output')
reports = os.path.join('..', 'data', '06_reporting')
# data_list = ['X_train', 'X_train_oh', 'y_train']
# dfs_dict = build_data_dict(inputs, data_list)
X_train = pd.read_csv(os.path.join(inputs, 'X_train.csv'), index_col='id')
X_train_onehot = pd.read_csv(os.path.join(inputs, 'X_train_onehot.csv'), index_col='id')
y_train = pd.read_csv(os.path.join(inputs, 'y_train.csv'), index_col='id')
data_list = [X_train, X_train_onehot, y_train]
for df in data_list:
print(df.shape)
```
# Machine Learning
```
import xgboost as xgb
from xgboost.sklearn import XGBClassifier
import itertools
params = {
'learning_rate': 0.1,
'min_child_weight': 1,
'scale_pos_weight': 1,
'random_state': 42,
'nthread': 8
}
grid_params = {
'learning_rate': [0.1, 0.01], # [0.01, 0.1]
'n_estimators': [50, 10], # [10, 50, 100, 300], standard is 10
'max_depth': [5, 3], # [3, 5, 10], standard is 5
'gamma': [0], # 0, 0.01
'subsample': [0.8, 0.6], # [0.5, 0.8]
'colsample_bytree': [0.8, 0.6] # [0.5, 0.8]
}
# key = 'standard'
parameter_groups = {}
keys, values = zip(*grid_params.items())
permutations_dicts = [dict(zip(keys, v)) for v in itertools.product(*values)]
for i, param_group in enumerate(permutations_dicts):
objective = 'binary:logistic'
model = XGBClassifier()
for k,v in params.items():
setattr(model, k, v)
for k,v in param_group.items():
setattr(model, k, v)
setattr(model, 'objective', objective)
parameter_groups[i] = model
```
Notice that we have a binary target. If target type was 'regression', we would have to run the following code for the param group loop:
```python
from xgboost.sklearn import XGBRegressor
objective = 'reg:squarederror'
model = XGBRegressor()
# do_transform_label=None
```
# model with all features
```
model_name = 'tree_xgb'
columns = X_train.columns.to_list()
clf = gridsearch_and_save(model_name, parameter_groups, columns)
```
### feature importances
```
feature_importances = pd.Series(clf.get_booster().get_score(importance_type='total_gain')).sort_values(ascending=False)
feature_importances.head(20).plot(kind='bar', title='Feature Importances (total_gain)')
plt.ylabel('Feature Importance Score');
plt.savefig(os.path.join(reports, 'xgboost_importances_gain.jpg'), bbox_inches='tight')
feature_importances = pd.Series(clf.get_booster().get_score(importance_type='weight')).sort_values(ascending=False)
feature_importances.head(20).plot(kind='bar', title='Feature Importances (weight)')
plt.ylabel('Feature Importance Score');
plt.savefig(os.path.join(reports, 'xgboost_importances_weight.jpg'), bbox_inches='tight')
```
# model with numerical features and cycle
### feature importances
# model with numerical features, cycle, and smote
### feature importances
# model with numerical features, presets, cycle, and smote
# model with numerical features
### feature importances
| github_jupyter |
# CSC421 Assignment 2 - Part II First-Order Logic (5 points) #
### Author: George Tzanetakis
This notebook is based on the supporting material for topics covered in **Chapter 7 - Logical Agents** from the book *Artificial Intelligence: A Modern Approach.* You can consult and modify the code provided in logic.py and logic.ipynb for completing the assignment questions. This part does rely on the provided code.
```
Birds can fly, unless they are penguins and ostriches, or if they happen
to be dead, or have broken wings, or are confined to cages, or have their
feet stuck in cement, or have undergone experiences so dreadful as to render
them psychologically incapable of flight
Marvin Minsky
```
# Introduction - First-Order Logic and knowledge engineering
In this assignment we explore First-Order Logic (FOL) using the implementation of knowledge base and first-order inference provided by the textbook authors. We also look into matching a limited form of unification.
**NOTE THAT THE GRADING IN THIS ASSIGNMENT IS DIFFERENT FOR GRADUATE STUDENTS AND THEY HAVE TO DO EXTRA WORK FOR FULL MARKS**
# Question 2A (Minimum) (CSC421 - 1 point, CSC581C - 0 points)
Consider the following propositional logic knowledge base.
* It is not sunny this afternoon and it is colder than yesterday.
* We will go swimming only if it is sunny.
* If we do not go swimming then we will take a canoe trip.
* If we take a canoe trip, then we will be home by sunset.
Denote:
* p = It is sunny this afternoon
* q = it is colder than yesterday
* r = We will go swimming
* s= we will take a canoe trip
* t= We will be home by sunset
Express this knowledge base using propositional logic using the expression syntax used in logic.ipynb. You can incoprorate any code you need from logic.ipynb and logic.py. In order to access the associated code the easiest way is to place your notebook in the same folder as the aima_python source code. Using both model checking and theorem proving inference (you can use the implementations provided) show that this knowledge base entails the sentence if it is not sunny this afternoon then we will be home by sunset.
```
from utils import *
from logic import *
from notebook import psource
# YOUR CODE GOES HERE
knowledge_base = PropKB()
(P, Q, R, S, T) = symbols("P, Q, R, S, T")
sentence1 = expr('R ==> P')
sentence2 = expr('~R ==> S')
sentence3 = expr('S ==> T')
sentence4 = expr('~P & Q')
knowledge_base.tell(sentence1)
knowledge_base.tell(sentence2)
knowledge_base.tell(sentence3)
knowledge_base.tell(sentence4)
#knowledge_base.clauses
knowledge_base.ask_if_true(expr('~P ==> T'))
```
# Question 2B (Minimum) (CSC421 - 1 point, CSC581C - 0 point)
Encode the kindship domain described in section 8.3.2 of the textbook using FOL and FolKB implementation in logic.ipynb and encode as facts the relationships between the members of the Simpsons family from the popular TV show:
https://en.wikipedia.org/wiki/Simpson_family
Show how the following queries can be answered using the KB:
* Who are the children of Homer ?
* Who are the parents of Bart ?
* Are Lisa and Homer siblings ?
* Are Lisa and Bart siblings ?
```
# YOUR CODE GOES HERE
'''
Parent(x,y): x is a parent to y
Child(x,y): x is a child of y
Sibling(x,y): x and y are siblings
'''
# Define Clauses
clauses = []
# 1.
clauses.append(expr("Female(p) & Parent(p, c) ==> Child(c,p)"))
clauses.append(expr("Mother(p) ==> Female(p)"))
# 2.
clauses.append(expr("Male(m) & Spouse(m, f) ==> Husband(m,f)"))
clauses.append(expr("Spouse(m, f) ==> Male(m)"))
clauses.append(expr("Spouse(m, f) ==> Female(f)"))
# 3.
# 4.
clauses.append(expr("Parent(p, c) ==> Child(c,p)"))
clauses.append(expr("Child(c, p) ==> Parent(p,c)"))
# 5.
clauses.append(expr("(Parent(p,x) & Parent(p,y)) ==> Sibling(x, y)"))
clauses.append(expr("(Sibling(x, y)) ==> (Sibling(y,x))"))
# Init First Order Logic KB
fam_kb = FolKB(clauses)
# Tell KB who the parents are
fam_kb.tell(expr("Parent(Homer, Bart)"))
fam_kb.tell(expr("Parent(Marge, Bart)"))
fam_kb.tell(expr("Parent(Homer, Lisa)"))
# Ask KB Question
homers_children = fol_fc_ask(fam_kb, expr('Child(x, Homer)'))
print("Homers Children: ",list(homers_children))
barts_parents = fol_fc_ask(fam_kb, expr('Parent(x, Bart)'))
print("Barts Parents: ",list(barts_parents))
siblings = fol_fc_ask(fam_kb, expr('Sibling(Lisa, Homer)'))
print("Lisa - Homer Sibs? ",bool(list(siblings))) # Check if list empty
siblings = fol_fc_ask(fam_kb, expr('Sibling(Lisa, Bart)'))
print("Lisa - Bart Sibs? ",bool(list(siblings))) # Check if list empty
```
# Question 2C (Expected) 1 point
Encode the electronic circuit domain described in section 8.4.2 of your textbook using the FolKB implementation in logics.ipynb. Encode the general knowledge of the domain as well as the specific problem instance shown in Figure 8.6. Post the same queries described by the book to the inference procedure.
```
# YOUR CODE GOES HERE
'''
Terms:
Terminal(t1)
Connected(t1, t2)
SignalOn(t)
SignalOff(t)
Gate(g)
4 Gates
'''
clauses = []
# 1
#clauses.append(expr("Out(x,g) ==> Terminal(Out(x,g))"))
#clauses.append(expr("In(x,g) ==> Terminal(x,g)"))
clauses.append(expr("Terminal(x, g) & Terminal(y, g) & Connected(x, y) ==> SameSignal(x, y)"))
clauses.append(expr("SameSignal(x,y) ==> SameSignal(y, x)"))
clauses.append(expr("SameSignal(x,y) & Signal(x, 1) ==> Signal(y, 1)"))
#clauses.append(expr("SameSignal(x,y) & Signal(x, 1) ==> Signal(y, 1)"))
# 2 - Signal at each terminal is either on or off...
clauses.append(expr("Signal(t,0) ==> Terminal(t)"))
clauses.append(expr("Signal(t,1) ==> Terminal(t)"))
# 3
clauses.append(expr("Connected(t1, t2) ==> Connected(t2,t1)"))
# 4 - There are 4 types of gates
clauses.append(expr("OR(g) ==> Gate(g)"))
clauses.append(expr("XOR(g) ==> Gate(g)"))
clauses.append(expr("AND(g) ==> Gate(g)"))
clauses.append(expr("NOT(g) ==> Gate(g)"))
# 5 AND Gate
clauses.append(expr("(Gate(g) & AND(g) & Signal(In(1,g), 1) & Signal(In(2,g), 1)) ==> Signal(Out(g), 1)"))
clauses.append(expr("(Gate(g) & AND(g) & Signal(In(1,g), 1) & Signal(In(2,g), 0)) ==> Signal(Out(g), 0)"))
clauses.append(expr("(Gate(g) & AND(g) & Signal(In(1,g), 0) & Signal(In(2,g), 1)) ==> Signal(Out(g), 0)"))
clauses.append(expr("(Gate(g) & AND(g) & Signal(In(1,g), 0) & Signal(In(2,g), 0)) ==> Signal(Out(g), 0)"))
# 6 OR Gate
clauses.append(expr("(Gate(g) & OR(g) & Signal(In(1,g), 1)) ==> Signal(Out(g), 1)"))
clauses.append(expr("(Gate(g) & OR(g) & Signal(In(2,g), 1)) ==> Signal(Out(g), 1)"))
clauses.append(expr("(Gate(g) & OR(g) & Signal(In(1,g), 0)) & Signal(In(2,g), 0) ==> Signal(Out(g), 0)"))
# 7 XOR
clauses.append(expr("(Gate(g) & XOR(g) & Signal(In(1,g), 1) & Signal(In(2,g), 1)) ==> Signal(Out(g), 0)"))
clauses.append(expr("(Gate(g) & XOR(g) & Signal(In(1,g), 1) & Signal(In(2,g), 0)) ==> Signal(Out(g), 1)"))
clauses.append(expr("(Gate(g) & XOR(g) & Signal(In(1,g), 0) & Signal(In(2,g), 1)) ==> Signal(Out(g), 1)"))
clauses.append(expr("(Gate(g) & XOR(g) & Signal(In(1,g), 0) & Signal(In(2,g), 0)) ==> Signal(Out(g), 0)"))
# 8 NOT Gate
clauses.append(expr("(Gate(g) & NOT(g) & Signal(In(1,g), 0)) ==> Signal(Out(g), 1)"))
clauses.append(expr("(Gate(g) & NOT(g) & Signal(In(1,g), 1)) ==> Signal(Out(g), 0)"))
# 9
clauses.append(expr("NOT(g) ==> Arity(g, 1, 1)"))
clauses.append(expr("AND(g) ==> Arity(g, 2, 1)"))
clauses.append(expr("XOR(g) ==> Arity(g, 2, 1)"))
clauses.append(expr("OR(g) ==> Arity(g, 2, 1)"))
# 10
clauses.append(expr("Out(g) ==> Arity(g, 2, 1)"))
circuit_kb = FolKB(clauses)
circuit_kb.tell(expr("XOR(X1)"))
circuit_kb.tell(expr("XOR(X2)"))
circuit_kb.tell(expr("AND(A1)"))
circuit_kb.tell(expr("AND(A2)"))
circuit_kb.tell(expr("OR(O1)"))
circuit_kb.tell(expr("OR(O1)"))
circuit_kb.tell(expr("Connected(Out(A, X1), In(A, X2))"))
circuit_kb.tell(expr("Connected(Out(A, X1), In(B, A2))"))
circuit_kb.tell(expr("Connected(Out(A, A2), In(A, O1))"))
circuit_kb.tell(expr("Connected(Out(A, A1), In(B, O1))"))
#circuit_kb.clauses
list(fol_fc_ask(circuit_kb, expr('XOR(x)')))
```
# QUESTION 1D (EXPECTED) 1 point
In this question we explore Prolog which is a programming language based on logic. We won't go into details but just wanted to give you a flavor of the syntax and how it connects to what we have learned. For this question you
will NOT be using the notebook so your answer should just be the source code. We will use http://tau-prolog.org/ which is a Prolog implementation that can run in a browser. When you access the webpage there is a text window labeled try it for entering your knowledge base and under it there is a text entry field for entering your query.
For example type in the Try it window and press enter:
```Prolog
likes(sam, salad).
likes(dean, pie).
likes(sam, apples).
likes(dean, whiskey).
```
Then enter the query:
```Prolog
likes(sam,X).
```
When you press Enter once you will get X=apples. and X=salad. Note the periods at the end of each statement.
Encode the kinship domain from question 2B in Prolog and answer the queries from 2B. Notice that in Prolog the constants start with lower case letters and the variables start with upper case letters.
Provide your code for the KB and queries using markup. See the syntax for Prolog of this cell by double clicking for editing.
# YOUR CODE GOES HERE
```Prolog
:- use_module(library(lists)).
child(X,Y) :-
parent(Y,X).
parent(A,B) :-
child(B,A)
sibling(A,B) :-
parent(X,A),
parent(X,B).
sibling(A,B) :-
sibling(B,A).
spouse(A,B) :-
spouse(B,A).
husband(H,W) :-
male(H),
spouse(H,W).
parent(marge, bart).
parent(marge, lisa).
parent(homer, bart).
parent(homer, lisa).
```
#### Goals and responses
```Prolog
child(X,homer). -> X = bart; X = lisa .
parent(X,bart). -> X = marge ; X = homer.
sibling(lisa,homer). -> false.
siblings(lisa, bart). -> true.
```
# QUESTION 1E (ADVANCED) 1 point
Implement exercise 8.26 using the code in logic.ipynb as well as the KB you wrote for the circuit domain.
```
# YOUR CODE GOES HERE
```
# QUESTION 1F (ADVANCED) (CSC421 - 0 points, CSC581C - 2 points)
This question explores the automatic constructions of a first-order logic knowledge base from a web resource and is more open ended than the other ones. The website https://www.songfacts.com/ contains a large variety of facts about music. Check the https://www.songfacts.com/categories link for some categories. Using selenium Python bindings https://selenium-python.readthedocs.io/ access the webpage and scrape at least three categories. Your code should scrape the information from the pages and convert it into relationships and facts in first-order logic using the syntax of expressions in logic.ipynb. Once you build your knowledge-base then write 4 non-trivial queries that show-case the expressiveness of FOL. These queries should not be possible to be answered easily using the web interface i.e they should have some logical connectives, more than one possible answer etc.
The translation of the song facts from the web page to FOL should NOT be done by hand but using the web scraping tool you develop. You can use multiple cells in your answer.
```
# YOUR CODE GOES HERE
```
| github_jupyter |
# Imports
```
from pathlib import Path
import mlflow
import mlflow.spark
import pandas as pd
from pyspark.ml.feature import StringIndexer, OneHotEncoder, VectorAssembler
from pyspark.ml.regression import LinearRegression, RandomForestRegressor
from pyspark.ml.evaluation import RegressionEvaluator
from pyspark.ml import Pipeline, PipelineModel
from pyspark.ml.tuning import CrossValidator, ParamGridBuilder
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('ML in Spark').getOrCreate()
spark
DATA_PATH = Path('data')
!ls {DATA_PATH}
```
# Data
```
df = (spark
.read
.parquet((DATA_PATH / 'sf-airbnb-clean.parquet').as_posix()))
df.count()
df.columns
df.select("neighbourhood_cleansed", "room_type", "bedrooms", "bathrooms",
"number_of_reviews", "price").show(5)
train_df, test_df = df.randomSplit([.8, .2], seed=42)
train_df.count(), test_df.count()
train_df.dtypes
```
# Linear Regression
```
cat_cols = [field for (field, dtype) in train_df.dtypes if dtype == 'string']
num_cols = [field for (field, dtype) in train_df.dtypes if dtype != 'string' and field != 'price']
print(f'Categorical fields:\n{cat_cols}\n')
print(f'Numerical fields:\n{num_cols}')
indexed_cols = [col + '_indexed' for col in cat_cols]
encoded_cols = [col + '_encoded' for col in cat_cols]
indexer = (StringIndexer()
.setInputCols(cat_cols)
.setOutputCols(indexed_cols)
.setHandleInvalid('skip'))
ohe = (OneHotEncoder()
.setInputCols(indexed_cols)
.setOutputCols(encoded_cols))
vector_assembler = (VectorAssembler()
.setInputCols(encoded_cols)
.setOutputCol('features'))
lr = LinearRegression(featuresCol='features', labelCol='price')
lr_pipeline = Pipeline(stages=[indexer, ohe, vector_assembler, lr])
lr_pipeline_model = lr_pipeline.fit(train_df)
pred_df = lr_pipeline_model.transform(test_df)
pred_df.select('features', 'price', 'prediction').show(5)
reg_eval = RegressionEvaluator(predictionCol='prediction', labelCol='price', metricName='rmse')
reg_eval.evaluate(pred_df)
reg_eval.setMetricName('r2').evaluate(pred_df)
lr_pipeline_model.write().overwrite().save('linear_reg_pip')
lr_pipeline_model = PipelineModel.load('linear_reg_pip')
reg_eval.setMetricName('r2').evaluate(lr_pipeline_model.transform(test_df))
```
# RandomForest
```
indexed_cols = [col + '_indexed' for col in cat_cols]
indexer = (StringIndexer()
.setInputCols(cat_cols)
.setOutputCols(indexed_cols)
.setHandleInvalid('skip'))
vector_assembler = (VectorAssembler()
.setInputCols(indexed_cols)
.setOutputCol('features'))
rf = RandomForestRegressor(featuresCol='features', labelCol='price', maxBins=40, seed=42)
rf_pipeline = Pipeline(stages=[indexer, vector_assembler, rf])
rf_pipeline_model = rf_pipeline.fit(train_df)
pred_df = rf_pipeline_model.transform(test_df)
pred_df.select('features', 'price', 'prediction').show(5)
reg_eval = RegressionEvaluator(predictionCol='prediction', labelCol='price', metricName='rmse')
reg_eval.evaluate(pred_df)
reg_eval.setMetricName('r2').evaluate(pred_df)
(pd.DataFrame(
list(
zip(rf_pipeline_model.stages[-2].getInputCols(), rf_pipeline_model.stages[-1].featureImportances))
, columns=['feature', 'importance'])
.sort_values(by='importance', ascending=False))
```
# Hyperparameter Tuning
```
indexed_cols = [col + '_indexed' for col in cat_cols]
indexer = (StringIndexer()
.setInputCols(cat_cols)
.setOutputCols(indexed_cols)
.setHandleInvalid('skip'))
vector_assembler = (VectorAssembler()
.setInputCols(indexed_cols)
.setOutputCol('features'))
rf = RandomForestRegressor(featuresCol='features', labelCol='price', maxBins=40, seed=42)
rf_pipeline = Pipeline(stages=[indexer, vector_assembler, rf])
evaluator = RegressionEvaluator(labelCol='price', predictionCol='prediction', metricName='rmse')
param_grid = (ParamGridBuilder()
.addGrid(rf.maxDepth, [2, 4, 6])
.addGrid(rf.numTrees, [10, 100])
.build())
cv = (CrossValidator()
.setEstimator(rf_pipeline)
.setEvaluator(evaluator)
.setEstimatorParamMaps(param_grid)
.setNumFolds(3)
.setSeed(42))
%time cv.fit(train_df)
cv = (CrossValidator()
.setEstimator(rf_pipeline)
.setEvaluator(evaluator)
.setEstimatorParamMaps(param_grid)
.setNumFolds(3)
.setParallelism(4)
.setSeed(42))
%time cv.fit(train_df)
cv = (CrossValidator()
.setEstimator(rf)
.setEvaluator(evaluator)
.setEstimatorParamMaps(param_grid)
.setParallelism(4)
.setNumFolds(3)
.setSeed(42))
rf_pipeline = Pipeline(stages=[indexer, vector_assembler, cv])
%time rf_pipeline.fit(train_df)
```
| github_jupyter |
# Name:- Parshwa Shah
# Experiment No.:- 2
# Roll No.- 34
# UID:- 2019230071
# Batch:- B
<h2>Aim:- To implement and use Principal Component Analysis using Python Platform </h2>
<center> <h1> Principal Component Analysis </h1><center>
<h3>Importing the libraries </h3>
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
```
<h3> Importing the dataset </h3>
```
! wget https://archive.ics.uci.edu/ml/machine-learning-databases/autos/imports-85.data imports-85.data
dataset = pd.read_csv('imports-85.data')
dataset
```
Attribute Information:
Attribute: Attribute Range
1. symboling: -3, -2, -1, 0, 1, 2, 3.
2. normalized-losses: continuous from 65 to 256.
3. make:
alfa-romero, audi, bmw, chevrolet, dodge, honda,
isuzu, jaguar, mazda, mercedes-benz, mercury,
mitsubishi, nissan, peugot, plymouth, porsche,
renault, saab, subaru, toyota, volkswagen, volvo
4. fuel-type: diesel, gas.
5. aspiration: std, turbo.
6. num-of-doors: four, two.
7. body-style: hardtop, wagon, sedan, hatchback, convertible.
8. drive-wheels: 4wd, fwd, rwd.
9. engine-location: front, rear.
10. wheel-base: continuous from 86.6 120.9.
11. length: continuous from 141.1 to 208.1.
12. width: continuous from 60.3 to 72.3.
13. height: continuous from 47.8 to 59.8.
14. curb-weight: continuous from 1488 to 4066.
15. engine-type: dohc, dohcv, l, ohc, ohcf, ohcv, rotor.
16. num-of-cylinders: eight, five, four, six, three, twelve, two.
17. engine-size: continuous from 61 to 326.
18. fuel-system: 1bbl, 2bbl, 4bbl, idi, mfi, mpfi, spdi, spfi.
19. bore: continuous from 2.54 to 3.94.
20. stroke: continuous from 2.07 to 4.17.
21. compression-ratio: continuous from 7 to 23.
22. horsepower: continuous from 48 to 288.
23. peak-rpm: continuous from 4150 to 6600.
24. city-mpg: continuous from 13 to 49.
25. highway-mpg: continuous from 16 to 54.
26. price: continuous from 5118 to 45400.
<h3> Finalize the column names </h3>
```
dataset.columns=['symboling','normalized-losses','make','fuel-type','aspiration','num-of-doors','body-style','drive-wheels','engine-location',
'wheel-base','length','width','height','curb-weight','engine-type','num-of-cylinders','engine-size','fuel-system','bore','stroke','compression-ratio','horsepower','peak-rpm','city-mpg','highway-mpg','price']
dataset
```
<h3>Drop the symboling column </h3>
```
dataset= dataset.drop('symboling', axis = 1)
dataset
```
<h3> Check for NaN values in dataset </h3>
```
dataset.isna().sum().plot(kind = 'bar')
```
<h3> Perform the Exploratory Data Analysis </h3>
```
dataset['normalized-losses'].value_counts().plot(kind = 'bar')
dataset['price'].value_counts()
```
<h3> Some columns had "?" values</h3>
```
#'?' in dataset[''].values
for col in dataset.columns:
if '?' in dataset[col].values:
print("col=",col)
```
<h3> Check count of "?" in all columns </h3>
```
dataset['bore'].value_counts().plot(kind = 'barh')
dataset['stroke'].value_counts().plot(kind = 'barh')
dataset['horsepower'].value_counts().plot(kind = 'barh')
dataset['peak-rpm'].value_counts().plot(kind = 'barh')
dataset['num-of-doors'].value_counts()
```
<h3> Convert "?" to NaN to get replaced with mean </h3>
```
dataset = dataset.replace('?',np.NAN)
dataset.isnull().sum()
```
<h3> Replace "?" with mean of column values </h3>
```
cols = ['normalized-losses', 'bore', 'stroke', 'horsepower', 'peak-rpm','price']
for col in cols:
dataset[col]=pd.to_numeric(dataset[col])
dataset[col].fillna(dataset[col].mean(), inplace=True)
dataset.head()
```
<h3> Now, no "?" or NaN values in dataset </h3>
```
"?" in dataset.values
dataset
```
<h3> Perform data visualization of some columns with 'price' column </h3>
```
# price range of automobile based on the maker
plt.figure(figsize = (12,5))
sns.boxplot(x='make',y='price',data=dataset);
plt.xticks(rotation = 90)
plt.show()
sns.boxplot(x='fuel-type',y='price',data=dataset)
sns.boxplot(x='aspiration',y='price',data=dataset)
sns.boxplot(x='body-style',y='price',data=dataset)
sns.boxplot(x='num-of-doors',y='price',data=dataset)
sns.boxplot(x='drive-wheels',y='price',data=dataset)
sns.boxplot(x='engine-location',y='price',data=dataset)
sns.boxplot(x='engine-type',y='price',data=dataset)
sns.boxplot(x='num-of-cylinders',y='price',data=dataset)
sns.boxplot(x='fuel-system',y='price',data=dataset)
plt.figure(figsize=(8, 5))
plt.scatter(x='engine-size',y='price',data=dataset)
plt.xlabel('engine-size')
plt.ylabel('Price');
plt.figure(figsize=(8, 5))
plt.scatter(x='peak-rpm',y='price',data=dataset)
plt.xlabel('peak-rpm')
plt.ylabel('Price');
plt.figure(figsize=(8, 5))
plt.scatter(x='stroke',y='price',data=dataset)
plt.xlabel('stroke')
plt.ylabel('Price');
dataset
```
<h3> Label Encode the Categorical Data </h3>
```
from sklearn import preprocessing
label_encoder = preprocessing.LabelEncoder()
dataset['make']= label_encoder.fit_transform(dataset['make'])
dataset['fuel-type']= label_encoder.fit_transform(dataset['fuel-type'])
dataset['aspiration']= label_encoder.fit_transform(dataset['aspiration'])
dataset['num-of-doors']= label_encoder.fit_transform(dataset['num-of-doors'])
dataset['body-style']= label_encoder.fit_transform(dataset['body-style'])
dataset['drive-wheels']= label_encoder.fit_transform(dataset['drive-wheels'])
dataset['engine-location']= label_encoder.fit_transform(dataset['engine-location'])
dataset['engine-type']= label_encoder.fit_transform(dataset['engine-type'])
dataset['num-of-cylinders']= label_encoder.fit_transform(dataset['num-of-cylinders'])
dataset['fuel-system']= label_encoder.fit_transform(dataset['fuel-system'])
dataset
```
<h3> Plot the correlation graph </h3>
```
# plt.imshow(dataset, cmap='hot', interpolation='nearest')
# plt.show()
plt.figure(figsize = (14,9))
hm = sns.heatmap(dataset.corr(),annot=True,cmap='Blues')
# displaying the plotted heatmap
plt.show()
```
<h3> Plot the covariance matrix </h3>
```
covMatrix = pd.DataFrame.cov(dataset)
(covMatrix)
```
<h3> Divide the dataset into independent and dependent columns </h3>
```
X = dataset.iloc[:,:24]
X
y = dataset.iloc[:,24:]
y
```
<h3> Perform MinMax Normalization </h3>
```
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
X=scaler.fit_transform(X)
#y=scaler.fit_transform(y)
X
y
```
<h3> Split the dataset into 80% training set and 20% in test set </h3>
```
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.2,random_state=66)
X_train.shape
y_train
```
<h3> Train the Linear Regression model and print the summary and results </h3>
```
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_absolute_error,mean_squared_error,r2_score
regressor = LinearRegression()
regressor.fit(X_train,y_train)
y_pred = regressor.predict(X_test)
print("MAE=",mean_absolute_error(y_test,y_pred))
print("MSE=",mean_squared_error(y_test,y_pred))
print("R2 Score=",r2_score(y_test,y_pred))
```
<h3> Do parameter tuning and get the results of PCA on regression model for different number of components </h3>
```
from sklearn.decomposition import PCA
i_value=[]
r2_value=[]
mae_value=[]
for i in range(1,23):
# Applying PCA
pca = PCA(n_components = i)
X_train1 = pca.fit_transform(X_train)
X_test1 = pca.transform(X_test)
comp_results = pca.components_
print(f"PCA for {i} Components is \n {pca.components_}")
# Training And Testing the model
regressor = LinearRegression()
regressor.fit(X_train1,y_train)
y_pred = regressor.predict(X_test1)
mae_value.append(mean_absolute_error(y_test,y_pred))
r2_value.append(r2_score(y_pred,y_test).round(4))
i_value.append(i)
# Plotting the r2 Score with each number of component for PCA
plt.plot(i_value,r2_value,marker='o',mfc='red',mec='red',color='blue')
plt.xlabel('Number of Components')
plt.ylabel('R2_Score')
plt.title('No_of_Components VS R2_Score')
plt.grid(b=None)
plt.show()
# Plotting the r2 Score with different number of components for PCA
plt.plot(i_value,mae_value,marker='o',mfc='red',mec='red',color='blue')
plt.xlabel('Number of Components')
plt.ylabel('MAE')
plt.title('No_of_Components VS MAE')
plt.grid(b=None)
plt.show()
```
<h3> When components are 16, R2 score is the highest and MAE is the lowest. </h3>
<h3> Conclusion: Hence, from this experiment, I understood the concept of PCA and how dimensionality reduction can help in improvising the model. Also, I performed PCA on Automobile dataset and did dimensionality reduction.</h3>
| github_jupyter |
# Stroop Effect: Test a Perceptual Phenomenon
###### by Maneesh Divana for Udacity's Machine Learning Foundation Nanodegree
----------
### Stroop Effect:
In a Stroop task, participants are presented with a list of words, with each word displayed in a color of ink. The participant’s task is to say out loud the color of the ink in which the word is printed.
The task has two conditions:
1. **Congruent words condition** - words being displayed are color words whose names match the colors in which they are printed: for example <span style="color:red"><em>RED</em></span>, <span style="color:blue"><em>BLUE</em></span>.
2. **Incongruent words condition** - words being displayed are color words whose names do not match the colors in which they are printed: for example <span style="color:orange"><em>PURPLE</em></span>, <span style="color:purple"><em>ORANGE</em></span>.
In each case, we measure the time it takes to name the ink colors in equally-sized lists. Each participant will go through and record a time from each condition.
----------------
### Investigation:
------------------------
#### Q1. What is our independent variable? What is our dependent variable?
- **Independent Variable:** It is the variable which is manipulated in the experiment. In this experiment the word-color congruency (Does the name of word corresponds to it's color?) is the independent variable.
- **Dependent Variable:** It is the variable which is measured in the experiment. In this experiment the time it takes to identify the word-color congruency/incongruency is the dependent variable.
-------------------
#### Q2. What is an appropriate set of hypotheses for this task? What kind of statistical test do you expect to perform? Justify your choices.**
With null hypothesis(H<sub>0</sub>) we assume that there will be no change. So the null hypothesis here is that the time taken to identify inconguent words is less than or equal to the time taken to identify congruent words.
With Alternative hypothesis(H<sub>A</sub>) we write what we think is the actual result of the experiment. So the alternative hypothesis here is that the time taken to identify inconguent words more than the time taken to identify congruent words.
So,
***H<sub>0</sub>: μ<sub>i</sub> ≤ μ<sub>c</sub>***
***H<sub>A</sub>: μ<sub>i</sub> > μ<sub>c</sub>***
(where, μ<sub>i</sub> => population mean of the time taken for incongruent task & μ<sub>c</sub> => population mean of the time taken for congruent task.)
**Statistical Test:** *<span style="color:green">Dependent Sample One Tailed t-Test</span>*
**Reasons:**
- Since the population mean and standard deviation is unknown z-test cannot be used here.
- Also since the size of the sample is less than 30, t-test is suitable for this experiment.
- This is dependent sample t-test because the same participant is subjected to both the conditions.
- It is a one tailed t-test because we clearly mention the direction of the test in the hypothesis.
---------------------
```
# Prepare Dataset
%matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# Loading dataset
stroop_data = pd.read_csv("./stroopdata.csv")
count = stroop_data.count()
print(f"Number of rows in dataframe:\n{count}")
# Finding statistics
congruent_series = stroop_data["Congruent"]
incongruent_series = stroop_data["Incongruent"]
print(f"Congruent Sample Mean : {congruent_series.mean()}")
print(f"Congruent Sample Median : {congruent_series.median()}")
print(f"Congruent Sample Mode : {congruent_series.mode()[0]}")
print(f"Congruent Sample Variance : {congruent_series.var()}")
print(f"Congruent Sample Stdev : {congruent_series.std()}") # Normalized by N-1 by default
print("")
print(f"Incongruent Sample Mean : {incongruent_series.mean()}")
print(f"Incongruent Sample Median : {incongruent_series.median()}")
print(f"Incongruent Sample Mode : {incongruent_series.mode()[0]}")
print(f"Incongruent Sample Variance : {incongruent_series.var()}")
print(f"Incongruent Sample Stdev : {incongruent_series.std()}") # Normalized by N-1 by default
```
-------------
#### Q3. Report some descriptive statistics regarding this dataset. Include at least one measure of central tendency and at least one measure of variability.
***Mesaures of Central Tendency:***
- Congruent Condition Sample Median = 14.357
- Incongruent Condition Sample Median = 21.018
- Congruent Condition Sample Mean (**x̄<sub>c</sub>**) = 14.051
- Incongruent Condition Sample Mean (**x̄<sub>i</sub>**) = 22.016
***Measures of Variability:***
- Congruent Condition Sample Variance = 12.669
- Incongruent Condition Sample Variance = 23.012
- Congruent Condition Sample Standard Deviation (**S<sub>c</sub>**) = 3.559
- Incongruent Condition Sample Standard Deviation (**S<sub>i</sub>**) = 4.797
***Sample Size = 24***
--------------------
#### 4. Provide one or two visualizations that show the distribution of the sample data. Write one or two sentences noting what you observe about the plot or plots.
***Observations:***
- From the box-plot, we can see that there are 2 outliers which would skew the true mean of incongruent values hence moving the distribtion slightly to right.
- From the histograms, we can see that both the distributions are almost positively skewed, the mean is close to the mode and median indicating a normal distribution. Also the outliers can be seen in the incongruent condition histogram which might affect the sampling mean and skew it a little to right.
- Hence, sampling mean would be almost equal to the population mean.
- From the line graph we can see that the times for incongruent condition are larger than congruent condtions for the same person.
Below are the histograms and box-plot of the congruent and incongruent sample data.
*Histograms:*
```
# set up figure & axes
fig, axes = plt.subplots(1, 2)
# drop sharex, sharey, layout & add ax=axes
axes[0].set_title('Congruent', ha='center')
axes[1].set_title('Incongruent', ha='center')
axes[0].set_xlabel("Time Taken")
axes[1].set_xlabel("Time Taken")
congruent_series.hist(ax=axes[0], figsize=[15,5])
incongruent_series.hist(ax=axes[1], figsize=[15,5])
# set title
plt.suptitle('Histogram of Sample Data', x=0.5, y=1.05, ha='center', fontsize='xx-large')
# Box Plot of the data
stroop_data.boxplot(figsize=[7,5])
xticks = [i for i in range(1, 26)]
yticks = [i for i in range(0, 36, 5) ]
axes = stroop_data.plot.line(figsize=[15,5],
title='Time Graph for Congruent vs Incongurent',
xticks=xticks,
yticks=yticks)
axes.set_xlabel("Participant")
axes.set_ylabel("Time Taken")
```
---------------------------
#### 5. Now, perform the statistical test and report your results. What is your confidence level and your critical statistic value? Do you reject the null hypothesis or fail to reject it? Come to a conclusion in terms of the experiment task. Did the results match up with your expectations?
*Lets now perform our one-tailed dependent t-test:*
- Congruent Condition Sample Mean (**X̄<sub>c</sub>**) = 14.051
- Incongruent Condition Sample Mean (**X̄<sub>i</sub>**) = 22.016
- Congruent Condition Sample Standard Deviation (**S<sub>c</sub>**) = 3.559
- Incongruent Condition Sample Standard Deviation (**S<sub>i</sub>**) = 4.797
- **n = 24**
We will calculate **t<sub>critical</sub>** & **t<sub>statistic</sub>** for a Confidence Level of 95%.
So, **$\alpha$ = 0.05** with **Degree of freedom (df) = $n - 1$ = 23**
- **t<sub>critical</sub> = 1.714** for $\alpha$=0.05 and df=23
- Sampling Mean Difference (**X̄<sub>D</sub>**) = (**X̄<sub>i</sub>**) - (**X̄<sub>c</sub>**) = 22.016 - 14.051 = **7.965**
- Standard Deviation of Mean Difference (**S<sub>D</sub>**) = **4.865**
- Standard Error of Mean Difference (**SEM**) = **$\frac{S_{\text{D}}}{\sqrt{n}}$** = **$\frac{4.865}{\sqrt{24}}$** = **$\frac{4.865}{4.9}$** = **0.993**
- **t<sub>statistic</sub>** = **$\frac{X_{\text{D}}}{\frac{S_{\text{D}}}{\sqrt{n}}}$** = **$\frac{X_{\text{D}}}{SEM}$** = **$\frac{7.965}{0.993}$** = **8.021**
**Result:**
**Since the t<sub>statistic</sub> (=8.0210) is greater than our t<sub>critical</sub> (=1.714), we reject the null hypothesis and our our alternative hypothesis hold true.**
**Hence we can conclude that participants take significantly more time to respond in the Incongruent condition task.**
**The p-value is < .00001.**
--------
***Calculations***
```
new_stroop_data = stroop_data.copy()
new_stroop_data["Difference"] = new_stroop_data["Incongruent"] - new_stroop_data["Congruent"]
new_stroop_data["Mean Difference"] = np.nan
new_stroop_data["Stdev Difference"] = np.nan
new_stroop_data.at[0, "Mean Difference"] = new_stroop_data["Difference"].mean()
new_stroop_data.at[0, "Stdev Difference"] = new_stroop_data["Difference"].std()
new_stroop_data
```
----------
#### 6. Optional: What do you think is responsible for the effects observed? Can you think of an alternative or similar task that would result in a similar effect? Some research about the problem will be helpful for thinking about these two questions!
**Reason for the effects observed:** One possible reason might be that the brain develops a pattern from the congruent task where the name of the color and the word are the same and then the brain tries to find the same pattern and take more time to process the word-color mismatch in the incongruent task.
**Alternative/Similar Task:** An experiment to measure the speed of typing using QUERTY keyboard & typing using DVORAK keyboard. This will test the explanation that people (their brain to be precise) tend to develop a pattern to do certain tasks and in this case it's people typing in QWERTY keyboards. When this pattern is broken by making them use a DVORAK keyboard, they would take more time to type a given paragraph.
----------
### References
+ [Quick Steps and Explanations of topics in Statistics](http://www.statisticshowto.com/)
+ [Socscistatistics for calculating p-value](http://www.socscistatistics.com/pvalues/tdistribution.aspx)
+ [Jupyter Notebook Markdown & LaTeX Documnetation](https://jupyter-notebook.readthedocs.io/en/stable/examples/Notebook/Working%20With%20Markdown%20Cells.html)
+ [Pandas Documentation](https://pandas.pydata.org/pandas-docs/stable/)
----------------------
| github_jupyter |
<!--BOOK_INFORMATION-->
<img align="left" style="padding-right:10px;" src="fig/cover-small.jpg">
*This notebook contains an excerpt from the [Whirlwind Tour of Python](http://www.oreilly.com/programming/free/a-whirlwind-tour-of-python.csp) by Jake VanderPlas; the content is available [on GitHub](https://github.com/jakevdp/WhirlwindTourOfPython).*
*The text and code are released under the [CC0](https://github.com/jakevdp/WhirlwindTourOfPython/blob/master/LICENSE) license; see also the companion project, the [Python Data Science Handbook](https://github.com/jakevdp/PythonDataScienceHandbook).*
<!--NAVIGATION-->
< [Basic Python Semantics: Variables and Objects](03-Semantics-Variables.ipynb) | [Contents](Index.ipynb) | [Built-In Types: Simple Values](05-Built-in-Scalar-Types.ipynb) >
# Basic Python Semantics: Operators
In the previous section, we began to look at the semantics of Python variables and objects; here we'll dig into the semantics of the various *operators* included in the language.
By the end of this section, you'll have the basic tools to begin comparing and operating on data in Python.
## Arithmetic Operations
Python implements seven basic binary arithmetic operators, two of which can double as unary operators.
They are summarized in the following table:
| Operator | Name | Description |
|--------------|----------------|--------------------------------------------------------|
| ``a + b`` | Addition | Sum of ``a`` and ``b`` |
| ``a - b`` | Subtraction | Difference of ``a`` and ``b`` |
| ``a * b`` | Multiplication | Product of ``a`` and ``b`` |
| ``a / b`` | True division | Quotient of ``a`` and ``b`` |
| ``a // b`` | Floor division | Quotient of ``a`` and ``b``, removing fractional parts |
| ``a % b`` | Modulus | Integer remainder after division of ``a`` by ``b`` |
| ``a ** b`` | Exponentiation | ``a`` raised to the power of ``b`` |
| ``-a`` | Negation | The negative of ``a`` |
| ``+a`` | Unary plus | ``a`` unchanged (rarely used) |
These operators can be used and combined in intuitive ways, using standard parentheses to group operations.
For example:
```
# addition, subtraction, multiplication
(4 + 8) * (6.5 - 3)
```
Floor division is true division with fractional parts truncated:
```
# True division
print(11 / 2)
# Floor division
print(11 // 2)
```
The floor division operator was added in Python 3; you should be aware if working in Python 2 that the standard division operator (``/``) acts like floor division for integers and like true division for floating-point numbers.
Finally, I'll mention an eighth arithmetic operator that was added in Python 3.5: the ``a @ b`` operator, which is meant to indicate the *matrix product* of ``a`` and ``b``, for use in various linear algebra packages.
## Bitwise Operations
In addition to the standard numerical operations, Python includes operators to perform bitwise logical operations on integers.
These are much less commonly used than the standard arithmetic operations, but it's useful to know that they exist.
The six bitwise operators are summarized in the following table:
| Operator | Name | Description |
|--------------|-----------------|---------------------------------------------|
| ``a & b`` | Bitwise AND | Bits defined in both ``a`` and ``b`` |
| <code>a | b</code>| Bitwise OR | Bits defined in ``a`` or ``b`` or both |
| ``a ^ b`` | Bitwise XOR | Bits defined in ``a`` or ``b`` but not both |
| ``a << b`` | Bit shift left | Shift bits of ``a`` left by ``b`` units |
| ``a >> b`` | Bit shift right | Shift bits of ``a`` right by ``b`` units |
| ``~a`` | Bitwise NOT | Bitwise negation of ``a`` |
These bitwise operators only make sense in terms of the binary representation of numbers, which you can see using the built-in ``bin`` function:
```
bin(10)
```
The result is prefixed with ``'0b'``, which indicates a binary representation.
The rest of the digits indicate that the number 10 is expressed as the sum $1 \cdot 2^3 + 0 \cdot 2^2 + 1 \cdot 2^1 + 0 \cdot 2^0$.
Similarly, we can write:
```
bin(4)
```
Now, using bitwise OR, we can find the number which combines the bits of 4 and 10:
```
4 | 10
bin(4 | 10)
```
These bitwise operators are not as immediately useful as the standard arithmetic operators, but it's helpful to see them at least once to understand what class of operation they perform.
In particular, users from other languages are sometimes tempted to use XOR (i.e., ``a ^ b``) when they really mean exponentiation (i.e., ``a ** b``).
## Assignment Operations
We've seen that variables can be assigned with the "``=``" operator, and the values stored for later use. For example:
```
a = 24
print(a)
```
We can use these variables in expressions with any of the operators mentioned earlier.
For example, to add 2 to ``a`` we write:
```
a + 2
```
We might want to update the variable ``a`` with this new value; in this case, we could combine the addition and the assignment and write ``a = a + 2``.
Because this type of combined operation and assignment is so common, Python includes built-in update operators for all of the arithmetic operations:
```
a += 2 # equivalent to a = a + 2
print(a)
```
There is an augmented assignment operator corresponding to each of the binary operators listed earlier; in brief, they are:
|||||
|-|-|
|``a += b``| ``a -= b``|``a *= b``| ``a /= b``|
|``a //= b``| ``a %= b``|``a **= b``|``a &= b``|
|<code>a |= b</code>| ``a ^= b``|``a <<= b``| ``a >>= b``|
Each one is equivalent to the corresponding operation followed by assignment: that is, for any operator "``■``", the expression ``a ■= b`` is equivalent to ``a = a ■ b``, with a slight catch.
For mutable objects like lists, arrays, or DataFrames, these augmented assignment operations are actually subtly different than their more verbose counterparts: they modify the contents of the original object rather than creating a new object to store the result.
## Comparison Operations
Another type of operation which can be very useful is comparison of different values.
For this, Python implements standard comparison operators, which return Boolean values ``True`` and ``False``.
The comparison operations are listed in the following table:
| Operation | Description || Operation | Description |
|---------------|-----------------------------------||---------------|--------------------------------------|
| ``a == b`` | ``a`` equal to ``b`` || ``a != b`` | ``a`` not equal to ``b`` |
| ``a < b`` | ``a`` less than ``b`` || ``a > b`` | ``a`` greater than ``b`` |
| ``a <= b`` | ``a`` less than or equal to ``b`` || ``a >= b`` | ``a`` greater than or equal to ``b`` |
These comparison operators can be combined with the arithmetic and bitwise operators to express a virtually limitless range of tests for the numbers.
For example, we can check if a number is odd by checking that the modulus with 2 returns 1:
```
# 25 is odd
25 % 2 == 1
# 66 is odd
66 % 2 == 1
```
We can string-together multiple comparisons to check more complicated relationships:
```
# check if a is between 15 and 30
a = 25
15 < a < 30
```
And, just to make your head hurt a bit, take a look at this comparison:
```
-1 == ~0
```
Recall that ``~`` is the bit-flip operator, and evidently when you flip all the bits of zero you end up with -1.
If you're curious as to why this is, look up the *two's complement* integer encoding scheme, which is what Python uses to encode signed integers, and think about what happens when you start flipping all the bits of integers encoded this way.
## Boolean Operations
When working with Boolean values, Python provides operators to combine the values using the standard concepts of "and", "or", and "not".
Predictably, these operators are expressed using the words ``and``, ``or``, and ``not``:
```
x = 4
(x < 6) and (x > 2)
(x > 10) or (x % 2 == 0)
not (x < 6)
```
Boolean algebra aficionados might notice that the XOR operator is not included; this can of course be constructed in several ways from a compound statement of the other operators.
Otherwise, a clever trick you can use for XOR of Boolean values is the following:
```
# (x > 1) xor (x < 10)
(x > 1) != (x < 10)
```
These sorts of Boolean operations will become extremely useful when we begin discussing *control flow statements* such as conditionals and loops.
One sometimes confusing thing about the language is when to use Boolean operators (``and``, ``or``, ``not``), and when to use bitwise operations (``&``, ``|``, ``~``).
The answer lies in their names: Boolean operators should be used when you want to compute *Boolean values (i.e., truth or falsehood) of entire statements*.
Bitwise operations should be used when you want to *operate on individual bits or components of the objects in question*.
## Identity and Membership Operators
Like ``and``, ``or``, and ``not``, Python also contains prose-like operators to check for identity and membership.
They are the following:
| Operator | Description |
|---------------|---------------------------------------------------|
| ``a is b`` | True if ``a`` and ``b`` are identical objects |
| ``a is not b``| True if ``a`` and ``b`` are not identical objects |
| ``a in b`` | True if ``a`` is a member of ``b`` |
| ``a not in b``| True if ``a`` is not a member of ``b`` |
### Identity Operators: "``is``" and "``is not``"
The identity operators, "``is``" and "``is not``" check for *object identity*.
Object identity is different than equality, as we can see here:
```
a = [1, 2, 3]
b = [1, 2, 3]
a == b
a is b
a is not b
```
What do identical objects look like? Here is an example:
```
a = [1, 2, 3]
b = a
a is b
```
The difference between the two cases here is that in the first, ``a`` and ``b`` point to *different objects*, while in the second they point to the *same object*.
As we saw in the previous section, Python variables are pointers. The "``is``" operator checks whether the two variables are pointing to the same container (object), rather than referring to what the container contains.
With this in mind, in most cases that a beginner is tempted to use "``is``" what they really mean is ``==``.
### Membership operators
Membership operators check for membership within compound objects.
So, for example, we can write:
```
1 in [1, 2, 3]
2 not in [1, 2, 3]
```
These membership operations are an example of what makes Python so easy to use compared to lower-level languages such as C.
In C, membership would generally be determined by manually constructing a loop over the list and checking for equality of each value.
In Python, you just type what you want to know, in a manner reminiscent of straightforward English prose.
<!--NAVIGATION-->
< [Basic Python Semantics: Variables and Objects](03-Semantics-Variables.ipynb) | [Contents](Index.ipynb) | [Built-In Types: Simple Values](05-Built-in-Scalar-Types.ipynb) >
| github_jupyter |
# Pandas 101
> almost every operation doesn't effect the original object returns new DataFrame object
---
### Loading Data
```
import pandas as pd
df = pd.read_csv('pokemon_data.csv')
# data with a delimiter
# df = pd.read_csv('pokemon_data.txt', delimiter='\t')
```
#### Head & Tail
```
df.head(10)
# returns first 10 rows as DataFrame
# returns last 5 rows
df.tail()
```
### Reading the data
```
# Reading headers
df.columns
# Read one column
df['Name'][:10]
# df[['Name']][:10] # this one also works
# Read multiple columns
df[['Name', 'Type 1', 'HP']]
df.head(5)
# specific indexed row
# iloc: integer location
df.iloc[2]
df.iloc[1:4]
# Specific cell value
df.iloc[2, 1]
df.iloc[2]['Name']
# iter through rows
for index, row in df.iterrows():
print(f'{index}: {row}')
if index == 4: break
# get the specific rows based on conditions
df.loc[df['HP'] > 150]
# > type(df.loc[df['HP'] > 150])
# > pandas.core.frame.DataFrame
```
### Describing Data
```
df.describe()
# all numerical/statistical data description
```
### Sorting Data
#### Single Column
```
# ascending sort
df_name = df.sort_values('Name')
df_name.head(10)
# descending sort
df_name = df.sort_values('Name', ascending=False)
df_name.head(10)
```
#### Multiple Columns all ascending
```
df.head(10)
# sort by type and then HP ascending
df_type_hp = df.sort_values(['Type 1', 'HP'])
df_type_hp.head(10)
```
⬆ Both **`Type 1`** and **`HP`** columns are sorted ascending
#### Multiple Columns mixed sorting
Sort **`Type 1`** ascending and **`HP`** descending
```
df_type_hp_mixed = df.sort_values(['Type 1', 'HP'], ascending=[True, False])
# ^ Both statements are same
# 1: True
# 0: False
# df_type_hp_mixed = df.sort_values(['Type 1', 'HP'], ascending=[1, 0])
df_type_hp_mixed.head(10)
```
⬆ **`Type 1`** sorted ascending and **`HP`** sorted descending
### Making changes to data
#### Adding new column
```
df['Total'] = df['HP'] + df['Attack'] + df['Defense'] + df['Speed']
```
**New column `Total` is added**
```
df.head(3)
```
**Adding a column using `iloc`**
```
# df['SpTotal'] = df['Sp. Atk'] + df['Sp. Def']
df['SpTotal'] = df.iloc[:, 7:9].sum(axis=1)
```
* **axis=0** : Vertical/Column
* **axis=1** : Horizontal/Row
```py
df['SpTotal'] # New column to create
df.iloc[:, 7:9].sum(axis=1) # df.iloc[all_rows/row_range, 7 and 8 columns] sum horizontally
```
```
df.head()
df['Spam'] = df.iloc[0:5, 5:7].sum(axis=1)
df.head(10)
```
**`NaN` for column `Spam` from 5th row**
#### Drop a column(s)
```
# it doesn't drop in df instered it returns new df
df = df.drop('Total', axis=1)
df.head()
```
| github_jupyter |
# Jupyter Notebook Fundamentals
A **notebook** is a collection **cells**. These cells are run to execute code, render formatted text or display graphical visualizations.
## Understanding Code Cells and Markdown Cells
The following cell (with the gray text area) is a code cell.
```
# This is a code cell
# By default, a new cell added in a notebook is a code cell
1 + 1
```
This notebook is written in Python. Because of this, you need to select the appropriate **Kernel** that you use to run the cells of this notebook.
To select your Kernel:
1. In the notebook toolbar, select the **Kernel** dropdown.
2. From the drop-down, select **Python 3**.

The code cell above has not run yet, so the expressions of `1 + 1` has not been evaluated. To run the code cell, select the cell by placing your cursor within the cell text area and do any of the following:
- Press `F5` to run the cell
- Use the cell Run icon to the left of the cell

The following cell is another example of a code cell. Run it to see its output.
```
# This is also a code cell
print("Welcome to your SQL Server 2019 Big Data cluster!")
```
The following cell, which displays its output as formatted text is a text cell that supports [markdown](https://en.wikipedia.org/wiki/Markdown) format.
This is a *text* cell.
To create a text cell, select the cell command menu on the upper-right (...). In the context menu, select **Insert Text Before** to add a text cell above this one, or **Insert Text After** to add one after this cell.

Double click on the above cell and notice how the cell changes to an editable code cell.
A preview of the markdown is displayed below the cell. To finish editing, simply click somewhere outside of the cell or press `Esc`.
### Understanding cell output
By default, a notebook cell will output the value of evaluating the last line the cell.
Run the following cell. Observe that the entire cell is echoed in the output because the cell contains only one line.
```
"Hello SQL world!"
```
Next, examine the following cell. What do you expect the output to be? Run the cell and confirm your understanding.
```
"Hello SQL world!"
"And, hello Jupyter notebook!"
```
If you want to ensure your output displays something, use the `print` method.
```
print("Hello SQL world!")
print("And, hello Jupyter notebook!")
```
Not all code lines return a value to be output. Run the following cell to see one such an example.
```
text_variable = "Hello, hello!"
```
## Running multiple notebook cells
It is not uncommon to need to run (or re-run) a all notebook cells in top to bottom order.
To do this, select **Run Cells** in the toolbar above the notebook. This runs all cells starting from the first.
## Adding code cells
You can add new code cells in the same way you add text cells.
To do this, select the cell command menu on the upper-right (...). In the context menu, select **Insert Code Before** to add a code cell above this one, or **Insert Code After** to add one after this cell.

You can also use this command menu to delete a cell.
## Understanding notebook state
When you execute notebook cells, their execution is backed by a process running on a cluster or locally, depending on the Kernel you select. The state of your notebook, such as the values of variables, is maintained in the process. All variables default to a global scope (unless you author your code so it has nested scopes) and this global state can be a little confusing at first when you re-run cells.
Run the following two cells in order and take note of the value ouput for the variable `y`:
```
x = 10
y = x + 1
y
```
Next, run the following cell.
```
x = 100
```
Now select the cell that has the lines `y = x + 1` and `y`. And re-run that cell. Did the value of `y` meet your expectation?
The value of `y` should now be `101`. This is because it is not the actual order of the cells that determines the value, but the order in which they are run and how that affects the underlying state itself. To understand this, realize that when the code `x = 100` was run, this changed the value of `x`, and then when you re-ran the cell containing `y = x + 1` this evaluation used the current value of x which is 100. This resulted in `y` having a value of `101` and not `11`.
### Clearing results
You can use the **Clear Results** toolbar item above the notebook to clear all displayed output from underneath code cells.
You typically do this when you want to cleanly re-run a notebook you have been working on and eliminate any accidental changes to the state that may have occured while you were authoring the notebook.
| github_jupyter |
# Logistic Regression with a Neural Network mindset
Welcome to your first (required) programming assignment! You will build a logistic regression classifier to recognize cats. This assignment will step you through how to do this with a Neural Network mindset, and so will also hone your intuitions about deep learning.
**Instructions:**
- Do not use loops (for/while) in your code, unless the instructions explicitly ask you to do so.
**You will learn to:**
- Build the general architecture of a learning algorithm, including:
- Initializing parameters
- Calculating the cost function and its gradient
- Using an optimization algorithm (gradient descent)
- Gather all three functions above into a main model function, in the right order.
## 1 - Packages ##
First, let's run the cell below to import all the packages that you will need during this assignment.
- [numpy](www.numpy.org) is the fundamental package for scientific computing with Python.
- [h5py](http://www.h5py.org) is a common package to interact with a dataset that is stored on an H5 file.
- [matplotlib](http://matplotlib.org) is a famous library to plot graphs in Python.
- [PIL](http://www.pythonware.com/products/pil/) and [scipy](https://www.scipy.org/) are used here to test your model with your own picture at the end.
```
import numpy as np
import matplotlib.pyplot as plt
import h5py
import scipy
from PIL import Image
from scipy import ndimage
from lr_utils import load_dataset
%matplotlib inline
```
## 2 - Overview of the Problem set ##
**Problem Statement**: You are given a dataset ("data.h5") containing:
- a training set of m_train images labeled as cat (y=1) or non-cat (y=0)
- a test set of m_test images labeled as cat or non-cat
- each image is of shape (num_px, num_px, 3) where 3 is for the 3 channels (RGB). Thus, each image is square (height = num_px) and (width = num_px).
You will build a simple image-recognition algorithm that can correctly classify pictures as cat or non-cat.
Let's get more familiar with the dataset. Load the data by running the following code.
```
# Loading the data (cat/non-cat)
train_set_x_orig, train_set_y, test_set_x_orig, test_set_y, classes = load_dataset()
```
We added "_orig" at the end of image datasets (train and test) because we are going to preprocess them. After preprocessing, we will end up with train_set_x and test_set_x (the labels train_set_y and test_set_y don't need any preprocessing).
Each line of your train_set_x_orig and test_set_x_orig is an array representing an image. You can visualize an example by running the following code. Feel free also to change the `index` value and re-run to see other images.
```
# Example of a picture
index = 90
plt.imshow(train_set_x_orig[index])
print ("y = " + str(train_set_y[:, index]) + ", it's a '" + classes[np.squeeze(train_set_y[:, index])].decode("utf-8") + "' picture.")
```
Many software bugs in deep learning come from having matrix/vector dimensions that don't fit. If you can keep your matrix/vector dimensions straight you will go a long way toward eliminating many bugs.
**Exercise:** Find the values for:
- m_train (number of training examples)
- m_test (number of test examples)
- num_px (= height = width of a training image)
Remember that `train_set_x_orig` is a numpy-array of shape (m_train, num_px, num_px, 3). For instance, you can access `m_train` by writing `train_set_x_orig.shape[0]`.
```
### START CODE HERE ### (≈ 3 lines of code)
m_train = train_set_x_orig.shape[0]
m_test = test_set_x_orig.shape[0]
num_px = train_set_x_orig.shape[1]
### END CODE HERE ###
print ("Number of training examples: m_train = " + str(m_train))
print ("Number of testing examples: m_test = " + str(m_test))
print ("Height/Width of each image: num_px = " + str(num_px))
print ("Each image is of size: (" + str(num_px) + ", " + str(num_px) + ", 3)")
print ("train_set_x shape: " + str(train_set_x_orig.shape))
print ("train_set_y shape: " + str(train_set_y.shape))
print ("test_set_x shape: " + str(test_set_x_orig.shape))
print ("test_set_y shape: " + str(test_set_y.shape))
```
**Expected Output for m_train, m_test and num_px**:
<table style="width:15%">
<tr>
<td>**m_train**</td>
<td> 209 </td>
</tr>
<tr>
<td>**m_test**</td>
<td> 50 </td>
</tr>
<tr>
<td>**num_px**</td>
<td> 64 </td>
</tr>
</table>
For convenience, you should now reshape images of shape (num_px, num_px, 3) in a numpy-array of shape (num_px $*$ num_px $*$ 3, 1). After this, our training (and test) dataset is a numpy-array where each column represents a flattened image. There should be m_train (respectively m_test) columns.
**Exercise:** Reshape the training and test data sets so that images of size (num_px, num_px, 3) are flattened into single vectors of shape (num\_px $*$ num\_px $*$ 3, 1).
A trick when you want to flatten a matrix X of shape (a,b,c,d) to a matrix X_flatten of shape (b$*$c$*$d, a) is to use:
```python
X_flatten = X.reshape(X.shape[0], -1).T # X.T is the transpose of X
```
```
# Reshape the training and test examples
### START CODE HERE ### (≈ 2 lines of code)
train_set_x_flatten = train_set_x_orig.reshape(train_set_x_orig.shape[0],-1).T
test_set_x_flatten = test_set_x_orig.reshape(test_set_x_orig.shape[0],-1).T
### END CODE HERE ###
print ("train_set_x_flatten shape: " + str(train_set_x_flatten.shape))
print ("train_set_y shape: " + str(train_set_y.shape))
print ("test_set_x_flatten shape: " + str(test_set_x_flatten.shape))
print ("test_set_y shape: " + str(test_set_y.shape))
print ("sanity check after reshaping: " + str(train_set_x_flatten[0:5,0]))
```
**Expected Output**:
<table style="width:35%">
<tr>
<td>**train_set_x_flatten shape**</td>
<td> (12288, 209)</td>
</tr>
<tr>
<td>**train_set_y shape**</td>
<td>(1, 209)</td>
</tr>
<tr>
<td>**test_set_x_flatten shape**</td>
<td>(12288, 50)</td>
</tr>
<tr>
<td>**test_set_y shape**</td>
<td>(1, 50)</td>
</tr>
<tr>
<td>**sanity check after reshaping**</td>
<td>[17 31 56 22 33]</td>
</tr>
</table>
To represent color images, the red, green and blue channels (RGB) must be specified for each pixel, and so the pixel value is actually a vector of three numbers ranging from 0 to 255.
One common preprocessing step in machine learning is to center and standardize your dataset, meaning that you substract the mean of the whole numpy array from each example, and then divide each example by the standard deviation of the whole numpy array. But for picture datasets, it is simpler and more convenient and works almost as well to just divide every row of the dataset by 255 (the maximum value of a pixel channel).
<!-- During the training of your model, you're going to multiply weights and add biases to some initial inputs in order to observe neuron activations. Then you backpropogate with the gradients to train the model. But, it is extremely important for each feature to have a similar range such that our gradients don't explode. You will see that more in detail later in the lectures. !-->
Let's standardize our dataset.
```
train_set_x = train_set_x_flatten/255.
test_set_x = test_set_x_flatten/255.
```
<font color='blue'>
**What you need to remember:**
Common steps for pre-processing a new dataset are:
- Figure out the dimensions and shapes of the problem (m_train, m_test, num_px, ...)
- Reshape the datasets such that each example is now a vector of size (num_px \* num_px \* 3, 1)
- "Standardize" the data
## 3 - General Architecture of the learning algorithm ##
It's time to design a simple algorithm to distinguish cat images from non-cat images.
You will build a Logistic Regression, using a Neural Network mindset. The following Figure explains why **Logistic Regression is actually a very simple Neural Network!**
<img src="images/LogReg_kiank.png" style="width:650px;height:400px;">
**Mathematical expression of the algorithm**:
For one example $x^{(i)}$:
$$z^{(i)} = w^T x^{(i)} + b \tag{1}$$
$$\hat{y}^{(i)} = a^{(i)} = sigmoid(z^{(i)})\tag{2}$$
$$ \mathcal{L}(a^{(i)}, y^{(i)}) = - y^{(i)} \log(a^{(i)}) - (1-y^{(i)} ) \log(1-a^{(i)})\tag{3}$$
The cost is then computed by summing over all training examples:
$$ J = \frac{1}{m} \sum_{i=1}^m \mathcal{L}(a^{(i)}, y^{(i)})\tag{6}$$
**Key steps**:
In this exercise, you will carry out the following steps:
- Initialize the parameters of the model
- Learn the parameters for the model by minimizing the cost
- Use the learned parameters to make predictions (on the test set)
- Analyse the results and conclude
## 4 - Building the parts of our algorithm ##
The main steps for building a Neural Network are:
1. Define the model structure (such as number of input features)
2. Initialize the model's parameters
3. Loop:
- Calculate current loss (forward propagation)
- Calculate current gradient (backward propagation)
- Update parameters (gradient descent)
You often build 1-3 separately and integrate them into one function we call `model()`.
### 4.1 - Helper functions
**Exercise**: Using your code from "Python Basics", implement `sigmoid()`. As you've seen in the figure above, you need to compute $sigmoid( w^T x + b) = \frac{1}{1 + e^{-(w^T x + b)}}$ to make predictions. Use np.exp().
```
# GRADED FUNCTION: sigmoid
def sigmoid(z):
"""
Compute the sigmoid of z
Arguments:
z -- A scalar or numpy array of any size.
Return:
s -- sigmoid(z)
"""
### START CODE HERE ### (≈ 1 line of code)
s = 1 / (1 + np.exp(-z))
### END CODE HERE ###
return s
print ("sigmoid([0, 2]) = " + str(sigmoid(np.array([0,2]))))
```
**Expected Output**:
<table>
<tr>
<td>**sigmoid([0, 2])**</td>
<td> [ 0.5 0.88079708]</td>
</tr>
</table>
### 4.2 - Initializing parameters
**Exercise:** Implement parameter initialization in the cell below. You have to initialize w as a vector of zeros. If you don't know what numpy function to use, look up np.zeros() in the Numpy library's documentation.
```
# GRADED FUNCTION: initialize_with_zeros
def initialize_with_zeros(dim):
"""
This function creates a vector of zeros of shape (dim, 1) for w and initializes b to 0.
Argument:
dim -- size of the w vector we want (or number of parameters in this case)
Returns:
w -- initialized vector of shape (dim, 1)
b -- initialized scalar (corresponds to the bias)
"""
### START CODE HERE ### (≈ 1 line of code)
w = np.zeros((dim,1))
b = 0
### END CODE HERE ###
assert(w.shape == (dim, 1))
assert(isinstance(b, float) or isinstance(b, int))
return w, b
dim = 2
w, b = initialize_with_zeros(dim)
print ("w = " + str(w))
print ("b = " + str(b))
```
**Expected Output**:
<table style="width:15%">
<tr>
<td> ** w ** </td>
<td> [[ 0.]
[ 0.]] </td>
</tr>
<tr>
<td> ** b ** </td>
<td> 0 </td>
</tr>
</table>
For image inputs, w will be of shape (num_px $\times$ num_px $\times$ 3, 1).
### 4.3 - Forward and Backward propagation
Now that your parameters are initialized, you can do the "forward" and "backward" propagation steps for learning the parameters.
**Exercise:** Implement a function `propagate()` that computes the cost function and its gradient.
**Hints**:
Forward Propagation:
- You get X
- You compute $A = \sigma(w^T X + b) = (a^{(0)}, a^{(1)}, ..., a^{(m-1)}, a^{(m)})$
- You calculate the cost function: $J = -\frac{1}{m}\sum_{i=1}^{m}y^{(i)}\log(a^{(i)})+(1-y^{(i)})\log(1-a^{(i)})$
Here are the two formulas you will be using:
$$ \frac{\partial J}{\partial w} = \frac{1}{m}X(A-Y)^T\tag{7}$$
$$ \frac{\partial J}{\partial b} = \frac{1}{m} \sum_{i=1}^m (a^{(i)}-y^{(i)})\tag{8}$$
```
# GRADED FUNCTION: propagate
def propagate(w, b, X, Y):
"""
Implement the cost function and its gradient for the propagation explained above
Arguments:
w -- weights, a numpy array of size (num_px * num_px * 3, 1)
b -- bias, a scalar
X -- data of size (num_px * num_px * 3, number of examples)
Y -- true "label" vector (containing 0 if non-cat, 1 if cat) of size (1, number of examples)
Return:
cost -- negative log-likelihood cost for logistic regression
dw -- gradient of the loss with respect to w, thus same shape as w
db -- gradient of the loss with respect to b, thus same shape as b
Tips:
- Write your code step by step for the propagation. np.log(), np.dot()
"""
m = X.shape[1]
# FORWARD PROPAGATION (FROM X TO COST)
### START CODE HERE ### (≈ 2 lines of code)
A = sigmoid(np.dot(w.T,X)+b) # compute activation
#print(A.shape)
#print(Y.shape)
cost = -1 / m * np.sum(Y*np.log(A)+(1-Y)*np.log(1-A)) # compute cost
### END CODE HERE ###
# BACKWARD PROPAGATION (TO FIND GRAD)
### START CODE HERE ### (≈ 2 lines of code)
dw = 1 / m * np.dot(X,(A-Y).T)
db = 1 / m * np.sum(A-Y)
### END CODE HERE ###
assert(dw.shape == w.shape)
assert(db.dtype == float)
cost = np.squeeze(cost)
assert(cost.shape == ())
grads = {"dw": dw,
"db": db}
return grads, cost
w, b, X, Y = np.array([[1],[2]]), 2, np.array([[1,2],[3,4]]), np.array([[1,0]])
grads, cost = propagate(w, b, X, Y)
print ("dw = " + str(grads["dw"]))
print ("db = " + str(grads["db"]))
print ("cost = " + str(cost))
```
**Expected Output**:
<table style="width:50%">
<tr>
<td> ** dw ** </td>
<td> [[ 0.99993216]
[ 1.99980262]]</td>
</tr>
<tr>
<td> ** db ** </td>
<td> 0.499935230625 </td>
</tr>
<tr>
<td> ** cost ** </td>
<td> 6.000064773192205</td>
</tr>
</table>
### d) Optimization
- You have initialized your parameters.
- You are also able to compute a cost function and its gradient.
- Now, you want to update the parameters using gradient descent.
**Exercise:** Write down the optimization function. The goal is to learn $w$ and $b$ by minimizing the cost function $J$. For a parameter $\theta$, the update rule is $ \theta = \theta - \alpha \text{ } d\theta$, where $\alpha$ is the learning rate.
```
# GRADED FUNCTION: optimize
def optimize(w, b, X, Y, num_iterations, learning_rate, print_cost = False):
"""
This function optimizes w and b by running a gradient descent algorithm
Arguments:
w -- weights, a numpy array of size (num_px * num_px * 3, 1)
b -- bias, a scalar
X -- data of shape (num_px * num_px * 3, number of examples)
Y -- true "label" vector (containing 0 if non-cat, 1 if cat), of shape (1, number of examples)
num_iterations -- number of iterations of the optimization loop
learning_rate -- learning rate of the gradient descent update rule
print_cost -- True to print the loss every 100 steps
Returns:
params -- dictionary containing the weights w and bias b
grads -- dictionary containing the gradients of the weights and bias with respect to the cost function
costs -- list of all the costs computed during the optimization, this will be used to plot the learning curve.
Tips:
You basically need to write down two steps and iterate through them:
1) Calculate the cost and the gradient for the current parameters. Use propagate().
2) Update the parameters using gradient descent rule for w and b.
"""
costs = []
for i in range(num_iterations):
# Cost and gradient calculation (≈ 1-4 lines of code)
### START CODE HERE ###
grads, cost = propagate(w, b, X, Y)
### END CODE HERE ###
# Retrieve derivatives from grads
dw = grads["dw"]
db = grads["db"]
# update rule (≈ 2 lines of code)
### START CODE HERE ###
w = w - dw *learning_rate
b = b - db *learning_rate
### END CODE HERE ###
# Record the costs
if i % 100 == 0:
costs.append(cost)
# Print the cost every 100 training examples
if print_cost and i % 100 == 0:
print ("Cost after iteration %i: %f" %(i, cost))
params = {"w": w,
"b": b}
grads = {"dw": dw,
"db": db}
return params, grads, costs
params, grads, costs = optimize(w, b, X, Y, num_iterations= 100, learning_rate = 0.009, print_cost = False)
print ("w = " + str(params["w"]))
print ("b = " + str(params["b"]))
print ("dw = " + str(grads["dw"]))
print ("db = " + str(grads["db"]))
```
**Expected Output**:
<table style="width:40%">
<tr>
<td> **w** </td>
<td>[[ 0.1124579 ]
[ 0.23106775]] </td>
</tr>
<tr>
<td> **b** </td>
<td> 1.55930492484 </td>
</tr>
<tr>
<td> **dw** </td>
<td> [[ 0.90158428]
[ 1.76250842]] </td>
</tr>
<tr>
<td> **db** </td>
<td> 0.430462071679 </td>
</tr>
</table>
**Exercise:** The previous function will output the learned w and b. We are able to use w and b to predict the labels for a dataset X. Implement the `predict()` function. There is two steps to computing predictions:
1. Calculate $\hat{Y} = A = \sigma(w^T X + b)$
2. Convert the entries of a into 0 (if activation <= 0.5) or 1 (if activation > 0.5), stores the predictions in a vector `Y_prediction`. If you wish, you can use an `if`/`else` statement in a `for` loop (though there is also a way to vectorize this).
```
# GRADED FUNCTION: predict
def predict(w, b, X):
'''
Predict whether the label is 0 or 1 using learned logistic regression parameters (w, b)
Arguments:
w -- weights, a numpy array of size (num_px * num_px * 3, 1)
b -- bias, a scalar
X -- data of size (num_px * num_px * 3, number of examples)
Returns:
Y_prediction -- a numpy array (vector) containing all predictions (0/1) for the examples in X
'''
m = X.shape[1]
Y_prediction = np.zeros((1,m))
w = w.reshape(X.shape[0], 1)
# Compute vector "A" predicting the probabilities of a cat being present in the picture
### START CODE HERE ### (≈ 1 line of code)
A = sigmoid(np.dot(w.T,X)+b)
### END CODE HERE ###
for i in range(A.shape[1]):
# Convert probabilities A[0,i] to actual predictions p[0,i]
### START CODE HERE ### (≈ 4 lines of code)
if A[0,i]>0.5:
Y_prediction[0,i] =1
else:
Y_prediction[0,i] =0
### END CODE HERE ###
assert(Y_prediction.shape == (1, m))
return Y_prediction
print ("predictions = " + str(predict(w, b, X)))
```
**Expected Output**:
<table style="width:30%">
<tr>
<td>
**predictions**
</td>
<td>
[[ 1. 1.]]
</td>
</tr>
</table>
<font color='blue'>
**What to remember:**
You've implemented several functions that:
- Initialize (w,b)
- Optimize the loss iteratively to learn parameters (w,b):
- computing the cost and its gradient
- updating the parameters using gradient descent
- Use the learned (w,b) to predict the labels for a given set of examples
## 5 - Merge all functions into a model ##
You will now see how the overall model is structured by putting together all the building blocks (functions implemented in the previous parts) together, in the right order.
**Exercise:** Implement the model function. Use the following notation:
- Y_prediction for your predictions on the test set
- Y_prediction_train for your predictions on the train set
- w, costs, grads for the outputs of optimize()
```
# GRADED FUNCTION: model
def model(X_train, Y_train, X_test, Y_test, num_iterations = 2000, learning_rate = 0.5, print_cost = False):
"""
Builds the logistic regression model by calling the function you've implemented previously
Arguments:
X_train -- training set represented by a numpy array of shape (num_px * num_px * 3, m_train)
Y_train -- training labels represented by a numpy array (vector) of shape (1, m_train)
X_test -- test set represented by a numpy array of shape (num_px * num_px * 3, m_test)
Y_test -- test labels represented by a numpy array (vector) of shape (1, m_test)
num_iterations -- hyperparameter representing the number of iterations to optimize the parameters
learning_rate -- hyperparameter representing the learning rate used in the update rule of optimize()
print_cost -- Set to true to print the cost every 100 iterations
Returns:
d -- dictionary containing information about the model.
"""
### START CODE HERE ###
# initialize parameters with zeros (≈ 1 line of code)
w, b = initialize_with_zeros(X_train.shape[0])
# Gradient descent (≈ 1 line of code)
parameters, grads, costs = optimize(w, b, X_train, Y_train, num_iterations, learning_rate, print_cost = False)
# Retrieve parameters w and b from dictionary "parameters"
w = parameters["w"]
b = parameters["b"]
# Predict test/train set examples (≈ 2 lines of code)
Y_prediction_test = predict(w, b, X_test)
Y_prediction_train = predict(w, b, X_train)
### END CODE HERE ###
# Print train/test Errors
print("train accuracy: {} %".format(100 - np.mean(np.abs(Y_prediction_train - Y_train)) * 100))
print("test accuracy: {} %".format(100 - np.mean(np.abs(Y_prediction_test - Y_test)) * 100))
d = {"costs": costs,
"Y_prediction_test": Y_prediction_test,
"Y_prediction_train" : Y_prediction_train,
"w" : w,
"b" : b,
"learning_rate" : learning_rate,
"num_iterations": num_iterations}
return d
```
Run the following cell to train your model.
```
d = model(train_set_x, train_set_y, test_set_x, test_set_y, num_iterations = 20000, learning_rate = 0.005, print_cost = True)
```
**Expected Output**:
<table style="width:40%">
<tr>
<td> **Train Accuracy** </td>
<td> 99.04306220095694 % </td>
</tr>
<tr>
<td>**Test Accuracy** </td>
<td> 70.0 % </td>
</tr>
</table>
**Comment**: Training accuracy is close to 100%. This is a good sanity check: your model is working and has high enough capacity to fit the training data. Test error is 68%. It is actually not bad for this simple model, given the small dataset we used and that logistic regression is a linear classifier. But no worries, you'll build an even better classifier next week!
Also, you see that the model is clearly overfitting the training data. Later in this specialization you will learn how to reduce overfitting, for example by using regularization. Using the code below (and changing the `index` variable) you can look at predictions on pictures of the test set.
```
# Example of a picture that was wrongly classified.
index = 30
plt.imshow(test_set_x[:,index].reshape((num_px, num_px, 3)))
print ("y = " + str(test_set_y[0,index]) + ", you predicted that it is a \"" + classes[d["Y_prediction_test"][0,index]].decode("utf-8") + "\" picture.")
```
Let's also plot the cost function and the gradients.
```
# Plot learning curve (with costs)
costs = np.squeeze(d['costs'])
plt.plot(costs)
plt.ylabel('cost')
plt.xlabel('iterations (per hundreds)')
plt.title("Learning rate =" + str(d["learning_rate"]))
plt.show()
```
**Interpretation**:
You can see the cost decreasing. It shows that the parameters are being learned. However, you see that you could train the model even more on the training set. Try to increase the number of iterations in the cell above and rerun the cells. You might see that the training set accuracy goes up, but the test set accuracy goes down. This is called overfitting.
## 6 - Further analysis (optional/ungraded exercise) ##
Congratulations on building your first image classification model. Let's analyze it further, and examine possible choices for the learning rate $\alpha$.
#### Choice of learning rate ####
**Reminder**:
In order for Gradient Descent to work you must choose the learning rate wisely. The learning rate $\alpha$ determines how rapidly we update the parameters. If the learning rate is too large we may "overshoot" the optimal value. Similarly, if it is too small we will need too many iterations to converge to the best values. That's why it is crucial to use a well-tuned learning rate.
Let's compare the learning curve of our model with several choices of learning rates. Run the cell below. This should take about 1 minute. Feel free also to try different values than the three we have initialized the `learning_rates` variable to contain, and see what happens.
```
learning_rates = [0.01, 0.001, 0.0001]
models = {}
for i in learning_rates:
print ("learning rate is: " + str(i))
models[str(i)] = model(train_set_x, train_set_y, test_set_x, test_set_y, num_iterations = 1500, learning_rate = i, print_cost = False)
print ('\n' + "-------------------------------------------------------" + '\n')
for i in learning_rates:
plt.plot(np.squeeze(models[str(i)]["costs"]), label= str(models[str(i)]["learning_rate"]))
plt.ylabel('cost')
plt.xlabel('iterations')
legend = plt.legend(loc='upper center', shadow=True)
frame = legend.get_frame()
frame.set_facecolor('0.90')
plt.show()
```
**Interpretation**:
- Different learning rates give different costs and thus different predictions results.
- If the learning rate is too large (0.01), the cost may oscillate up and down. It may even diverge (though in this example, using 0.01 still eventually ends up at a good value for the cost).
- A lower cost doesn't mean a better model. You have to check if there is possibly overfitting. It happens when the training accuracy is a lot higher than the test accuracy.
- In deep learning, we usually recommend that you:
- Choose the learning rate that better minimizes the cost function.
- If your model overfits, use other techniques to reduce overfitting. (We'll talk about this in later videos.)
## 7 - Test with your own image (optional/ungraded exercise) ##
Congratulations on finishing this assignment. You can use your own image and see the output of your model. To do that:
1. Click on "File" in the upper bar of this notebook, then click "Open" to go on your Coursera Hub.
2. Add your image to this Jupyter Notebook's directory, in the "images" folder
3. Change your image's name in the following code
4. Run the code and check if the algorithm is right (1 = cat, 0 = non-cat)!
```
## START CODE HERE ## (PUT YOUR IMAGE NAME)
my_image = "mycat.jpg" # change this to the name of your image file
## END CODE HERE ##
# We preprocess the image to fit your algorithm.
fname = "images/" + my_image
image = np.array(ndimage.imread(fname, flatten=False))
my_image = scipy.misc.imresize(image, size=(num_px,num_px)).reshape((1, num_px*num_px*3)).T
my_predicted_image = predict(d["w"], d["b"], my_image)
plt.imshow(image)
print("y = " + str(np.squeeze(my_predicted_image)) + ", your algorithm predicts a \"" + classes[int(np.squeeze(my_predicted_image)),].decode("utf-8") + "\" picture.")
```
<font color='blue'>
**What to remember from this assignment:**
1. Preprocessing the dataset is important.
2. You implemented each function separately: initialize(), propagate(), optimize(). Then you built a model().
3. Tuning the learning rate (which is an example of a "hyperparameter") can make a big difference to the algorithm. You will see more examples of this later in this course!
Finally, if you'd like, we invite you to try different things on this Notebook. Make sure you submit before trying anything. Once you submit, things you can play with include:
- Play with the learning rate and the number of iterations
- Try different initialization methods and compare the results
- Test other preprocessings (center the data, or divide each row by its standard deviation)
Bibliography:
- http://www.wildml.com/2015/09/implementing-a-neural-network-from-scratch/
- https://stats.stackexchange.com/questions/211436/why-do-we-normalize-images-by-subtracting-the-datasets-image-mean-and-not-the-c
| github_jupyter |
```
import torch
import torch.nn as nn
from torchinfo import summary
def build_circle_segmenter():
circle_segmenter = nn.Sequential(
nn.Conv2d(
in_channels=1,
out_channels=8,
kernel_size=3,
stride=1,
padding=1
),
nn.ReLU(),
nn.Conv2d(
in_channels=8,
out_channels=8,
kernel_size=3,
stride=1,
padding=1
),
nn.ReLU(),
nn.MaxPool2d(
kernel_size=2,
stride=2
),
nn.Conv2d(
in_channels=8,
out_channels=16,
kernel_size=3,
stride=1,
padding=1
),
nn.ReLU(),
nn.Conv2d(
in_channels=16,
out_channels=16,
kernel_size=3,
stride=1,
padding=1
),
nn.ReLU(),
nn.MaxPool2d(
kernel_size=2,
stride=2
),
nn.Conv2d(
in_channels=16,
out_channels=32,
kernel_size=3,
stride=1,
padding=1
),
nn.ReLU(),
nn.Conv2d(
in_channels=32,
out_channels=32,
kernel_size=3,
stride=1,
padding=1
),
nn.ReLU(),
#nn.MaxPool2d(
# kernel_size=2,
# stride=2
#),
#nn.AdaptiveAvgPool2d((8, 8)),
nn.ConvTranspose2d(
in_channels=32,
out_channels=16,
kernel_size=2,
stride=2,
padding=0
),
nn.ReLU(),
nn.Conv2d(
in_channels=16,
out_channels=16,
kernel_size=3,
stride=1,
padding=1
),
nn.ReLU(),
nn.ConvTranspose2d(
in_channels=16,
out_channels=8,
kernel_size=2,
stride=2,
padding=0
),
nn.ReLU(),
nn.Conv2d(
in_channels=8,
out_channels=1,
kernel_size=3,
stride=1,
padding=1
),
nn.ReLU(),
)
return circle_segmenter
summary(build_circle_segmenter(), input_size=(2, 1, 128, 128))
import itertools
import matplotlib.pyplot as plt
import numpy as np
from PIL import Image, ImageDraw
def get_empty_image(image_width, image_height):
# mode="F" for 32-bit floating point pixels
# mode="LA" for 8-bit grayscale with alpha channel
empty_image = Image.new(
mode="F",
size=(image_width, image_height),
color=255
)
return empty_image
def draw_a_circle(target_image, circle_e1, circle_e2, circle_radius, outline_color=255):
"""
The most simple image of a circle?
"""
artist = ImageDraw.ImageDraw(target_image)
artist.arc(
(
circle_e1 - circle_radius/2,
circle_e2 - circle_radius/2,
circle_e1 + circle_radius/2,
circle_e2 + circle_radius/2
),
start=0,
end=360,
width=1,
fill=outline_color
)
return target_image
def fill_a_circle(target_image, circle_e1, circle_e2, circle_radius, circle_fill_color=255):
artist = ImageDraw.ImageDraw(target_image)
artist.ellipse(
(
circle_e1 - circle_radius/2,
circle_e2 - circle_radius/2,
circle_e1 + circle_radius/2,
circle_e2 + circle_radius/2
),
width=1,
#outline=255, # what happens without this?
fill=circle_fill_color
)
return target_image
def image_of_circles(circle_count):
"""
The most simple image of a circle?
"""
image_of_filled_circles = get_empty_image(
image_width=128,
image_height=128
)
circle_radius_list = list()
image_of_outlined_circles = get_empty_image(
image_width=128,
image_height=128
)
circle_parameters_list = list()
for circle_i in range(circle_count):
circle_radius = np.random.randint(low=10, high=40)
circle_radius_list.append(circle_radius)
circle_e1=np.random.randint(low=20, high=80)
circle_e2=np.random.randint(low=20, high=80)
circle_fill_color = np.random.randint(low=100, high=200)
circle_parameters_list.append(
{
"circle_radius": circle_radius,
"circle_e1": circle_e1,
"circle_e2": circle_e2,
"circle_fill_color": circle_fill_color
}
)
for circle_parameters in circle_parameters_list:
# these are the input training images
fill_a_circle(
target_image=image_of_filled_circles,
**circle_parameters
#circle_e1=circle_e1,
#circle_e2=circle_e2,
#circle_radius=circle_radius,
)
# these are the "target" training images
fill_a_circle(
target_image=image_of_outlined_circles,
**circle_parameters
#circle_e1=circle_e1,
#circle_e2=circle_e2,
#circle_radius=circle_radius,
)
# draw outlines on the "target" training images
for circle_parameters in circle_parameters_list:
circle_parameters.pop("circle_fill_color")
draw_a_circle(
target_image=image_of_outlined_circles,
outline_color=0,
**circle_parameters,
#circle_e1=circle_e1,
#circle_e2=circle_e2,
#circle_radius=circle_radius,
)
return (circle_radius_list, image_of_filled_circles, image_of_outlined_circles)
import matplotlib.pyplot as plt
# generate 4 sets of input/output
circle_radiuses_list = list()
input_images = list()
output_images = list()
for _ in range(4):
circle_count = np.random.randint(low=1, high=5)
circle_radiuses, input_circles_image, output_circles_image = image_of_circles(circle_count=circle_count)
circle_radiuses_list.append(circle_radiuses)
input_images.append(input_circles_image)
output_images.append(output_circles_image)
fig, axs = plt.subplots(nrows=2, ncols=2)
for i, (r, c) in enumerate(itertools.product(range(2), range(2))):
print('circle radiuses: {}'.format(circle_radiuses))
axs[r][c].imshow(input_images[i], origin="lower")
#print(np.array(im))
fig, axs = plt.subplots(nrows=2, ncols=2)
for i, (r, c) in enumerate(itertools.product(range(2), range(2))):
print('circle radiuses: {}'.format(circle_radiuses))
axs[r][c].imshow(output_images[i], origin="lower")
#print(np.array(im))
# a class to interact with DataLoaders
class CircleImageDataset:
def __init__(self, circle_image_count):
self.circle_image_list = list()
for i in range(circle_image_count):
circle_count = np.random.randint(low=1, high=5)
circle_radius_list, input_circles_image, target_circles_image = image_of_circles(
circle_count=circle_count
)
# sort the circle radiuses in descending order
# otherwise the training data is a little ambiguous?
#sorted_circle_radius_list = sorted(circle_radius_list, reverse=True)
# the network output is a 8-element array of circle radiuses
#circle_radiuses = np.zeros((8, ), dtype=np.float32)
#circle_radiuses[:circle_count] = sorted_circle_radius_list
self.circle_image_list.append(
(
# get the right type here - single precision floating point
# this depends on how the optimization is handled
# but I want to get it right here
#circle_radiuses,
# the PIL image is converted to a 2D numpy array here
# in addition an extra dimension is inserted for 'channel'
# which PyTorch convolutional networks expect
np.expand_dims(
np.array(target_circles_image),
axis=0
),
# the PIL image is converted to a 2D numpy array here
# in addition an extra dimension is inserted for 'channel'
# which PyTorch convolutional networks expect
np.expand_dims(
np.array(input_circles_image),
axis=0
)
)
)
def __getitem__(self, index):
# self.circle_image_list looks like
# [ (radius_0, radius_1, ...), image_0), (radius_0, radius_1, ...), image_1), ...]
# this dataset returns only (radius, image)
return self.circle_image_list[index]
def __len__(self):
return len(self.circle_image_list)
def test_circle_image_dataset():
circle_image_dataset = CircleImageDataset(100)
print(f"len(circle_image_dataset): {len(circle_image_dataset)}")
target_circle_image, input_circle_image = circle_image_dataset[99]
print(f"target image.shape : {target_circle_image.shape}")
print(f"input image.shape : {input_circle_image.shape}")
test_circle_image_dataset()
from torch.utils.data import DataLoader
def test_circle_image_dataloader():
circle_image_dataloader = DataLoader(CircleImageDataset(circle_image_count=100), batch_size=10)
for batch in circle_image_dataloader:
print(f"len(batch): {len(batch)}")
print(f"len(batch[0]): {len(batch[0])}")
print(f"batch[0].shape: {batch[0].shape}")
print(f"len(batch[1]): {len(batch[1])}")
print(f"batch[1].shape: {batch[1].shape}")
target_circle_images, input_circle_images = batch
# note correct_radii.shape does not match predicted_radii.shape
print(f"target_circle_images.shape: {target_circle_images.shape}")
print(f"target_circle_images.dtype: {target_circle_images.dtype}")
print(f"input_circle_images.shape: {input_circle_images.shape}")
test_circle_segmenter = build_circle_segmenter()
predicted_circle_images = test_circle_segmenter(input_circle_images)
print(f"predicted_circle_images.shape: {predicted_circle_images.shape}")
print(f"predicted_circle_images.dtype: {predicted_circle_images.dtype}")
break
test_circle_image_dataloader()
# 100,000, no shuffle, works, 20 epochs is ok but sometimes training does not progress
# 100,000 with shuffling has more stable training
# 10,000, no shuffle, works, 50 epochs is ok
train_circle_image_loader = DataLoader(
CircleImageDataset(circle_image_count=10000),
batch_size=100,
shuffle=True
)
test_circle_image_loader = DataLoader(
CircleImageDataset(circle_image_count=1000),
batch_size=100
)
#validate_circle_image_loader = DataLoader(CircleImageDataset(circle_image_count=1000), batch_size=100)
len(train_circle_image_loader.dataset)
def train(
circle_segmenter_model,
optimizer,
loss_function,
train_dataloader,
test_dataloader,
epoch_count
):
if torch.cuda.is_available():
device = torch.device("cuda")
else:
device = torch.device("cpu")
circle_segmenter_model.to(device)
for epoch_i in range(epoch_count):
training_loss = 0.0
circle_segmenter_model.train()
for correct_segmented_circle_images, circle_images in train_dataloader:
optimizer.zero_grad()
# torch calls circle_images 'inputs'
circle_images = circle_images.to(device)
# make the correct_radii array match predicted_radii.shape
#correct_radii = torch.unsqueeze(correct_radii, 1)
correct_segmented_circle_images = correct_segmented_circle_images.to(device)
predicted_radii = circle_segmenter_model(circle_images)
loss = loss_function(predicted_radii, correct_segmented_circle_images)
loss.backward()
optimizer.step()
training_loss += loss.data.item()
training_loss /= len(train_circle_image_loader.dataset)
test_loss = 0.0
circle_segmenter_model.eval()
for correct_segmented_circle_images, circle_images in test_dataloader:
# torch calls circle_images 'inputs'
circle_images = circle_images.to(device)
#inputs = inputs.to(device)
# make correct_radii have the same shape as predicted_radii
#correct_radii = torch.unsqueeze(correct_radii, 1)
correct_segmented_circle_images = correct_segmented_circle_images.to(device)
predicted_radii = circle_segmenter_model(circle_images)
loss = loss_function(predicted_radii, correct_segmented_circle_images)
test_loss += loss.data.item()
test_loss /= len(test_dataloader.dataset)
print(
#'Epoch: {}, Training Loss: {:.2f}, Test Loss: {:.2f}, percent_wrong = {}'.format(
'Epoch: {}, Training Loss: {:.2f}, Test Loss: {:.2f}'.format(
epoch_i, training_loss, test_loss
)
)
import torch.optim
circle_segmenter = build_circle_segmenter()
train(
circle_segmenter,
torch.optim.Adam(circle_segmenter.parameters()),
torch.nn.MSELoss(),
train_circle_image_loader,
test_circle_image_loader,
epoch_count=100
)
# try out the circle segmenter
if torch.cuda.is_available():
device = torch.device("cuda")
else:
device = torch.device("cpu")
circle_segmenter.eval()
a_circle_image_dataloader = DataLoader(CircleImageDataset(10), batch_size=1)
for a_circle_target_image, a_circle_input_image in a_circle_image_dataloader:
print(f"input shape : {a_circle_input_image.shape}")
a_segmented_circle_tensor = circle_segmenter(a_circle_input_image.to(device))
a_segmented_circle_image = a_segmented_circle_tensor.cpu().detach().numpy()
print(f"output shape: {a_segmented_circle_image.shape}")
fig, axs = plt.subplots(nrows=1, ncols=2)
#for i, (r, c) in enumerate(itertools.product(range(1), range(2))):
#print('circle radiuses: {}'.format(circle_radiuses))
axs[0].imshow(a_circle_input_image[0, 0, :, :], origin="lower")
axs[1].imshow(a_segmented_circle_image[0, 0, :, :], origin="lower")
#print(np.array(im))
```
| github_jupyter |
# 机器学习纳米学位
## 非监督学习
## 项目 3: 创建用户分类
欢迎来到机器学习工程师纳米学位的第三个项目!在这个 notebook 文件中,有些模板代码已经提供给你,但你还需要实现更多的功能来完成这个项目。除非有明确要求,你无须修改任何已给出的代码。以**'练习'**开始的标题表示接下来的代码部分中有你必须要实现的功能。每一部分都会有详细的指导,需要实现的部分也会在注释中以 **'TODO'** 标出。请仔细阅读所有的提示!
除了实现代码外,你还**必须**回答一些与项目和你的实现有关的问题。每一个需要你回答的问题都会以**'问题 X'**为标题。请仔细阅读每个问题,并且在问题后的**'回答'**文字框中写出完整的答案。我们将根据你对问题的回答和撰写代码所实现的功能来对你提交的项目进行评分。
>**提示:**Code 和 Markdown 区域可通过 **Shift + Enter** 快捷键运行。此外,Markdown 可以通过双击进入编辑模式。
## 开始
在这个项目中,你将分析一个数据集的内在结构,这个数据集包含很多客户真对不同类型产品的年度采购额(用**金额**表示)。这个项目的任务之一是如何最好地描述一个批发商不同种类顾客之间的差异。这样做将能够使得批发商能够更好的组织他们的物流服务以满足每个客户的需求。
这个项目的数据集能够在[UCI机器学习信息库](https://archive.ics.uci.edu/ml/datasets/Wholesale+customers)中找到.因为这个项目的目的,分析将不会包括 'Channel' 和 'Region' 这两个特征——重点集中在6个记录的客户购买的产品类别上。
运行下面的的代码单元以载入整个客户数据集和一些这个项目需要的 Python 库。如果你的数据集载入成功,你将看到后面输出数据集的大小。
```
# 检查你的Python版本
from sys import version_info
if version_info.major != 3:
raise Exception('请使用Python 3.x 来完成此项目')
# 引入这个项目需要的库
import numpy as np
import pandas as pd
import visuals as vs
from IPython.display import display # 使得我们可以对DataFrame使用display()函数
# 设置以内联的形式显示matplotlib绘制的图片(在notebook中显示更美观)
%matplotlib inline
# 高分辨率显示
# %config InlineBackend.figure_format='retina'
# 载入整个客户数据集
try:
data = pd.read_csv("customers.csv")
data.drop(['Region', 'Channel'], axis = 1, inplace = True)
print("Wholesale customers dataset has {} samples with {} features each.".format(*data.shape))
except:
print("Dataset could not be loaded. Is the dataset missing?")
```
## 分析数据
在这部分,你将开始分析数据,通过可视化和代码来理解每一个特征和其他特征的联系。你会看到关于数据集的统计描述,考虑每一个属性的相关性,然后从数据集中选择若干个样本数据点,你将在整个项目中一直跟踪研究这几个数据点。
运行下面的代码单元给出数据集的一个统计描述。注意这个数据集包含了6个重要的产品类型:**'Fresh'**, **'Milk'**, **'Grocery'**, **'Frozen'**, **'Detergents_Paper'**和 **'Delicatessen'**。想一下这里每一个类型代表你会购买什么样的产品。
```
# 显示数据集的一个描述
display(data.describe())
```
### 练习: 选择样本
为了对客户有一个更好的了解,并且了解代表他们的数据将会在这个分析过程中如何变换。最好是选择几个样本数据点,并且更为详细地分析它们。在下面的代码单元中,选择**三个**索引加入到索引列表`indices`中,这三个索引代表你要追踪的客户。我们建议你不断尝试,直到找到三个明显不同的客户。
```
# TODO:从数据集中选择三个你希望抽样的数据点的索引
indices = [29,85,128]
# 为选择的样本建立一个DataFrame
samples = pd.DataFrame(data.loc[indices], columns = data.keys()).reset_index(drop = True)
print("Chosen samples of wholesale customers dataset:")
display(samples)
```
### 问题 1
在你看来你选择的这三个样本点分别代表什么类型的企业(客户)?对每一个你选择的样本客户,通过它在每一种产品类型上的花费与数据集的统计描述进行比较,给出你做上述判断的理由。
**提示:** 企业的类型包括超市、咖啡馆、零售商以及其他。注意不要使用具体企业的名字,比如说在描述一个餐饮业客户时,你不能使用麦当劳。
**回答:**
* 29 餐馆 Fresh值远大于75%的样本
* 85 超市 多项数值较高,特别是Grocery是所有样本中的最大值,应该是一家比较大的超市
* 128 咖啡馆 Milk值大于75%的样本,其他项的值在样本中都不高
### 练习: 特征相关性
一个有趣的想法是,考虑这六个类别中的一个(或者多个)产品类别,是否对于理解客户的购买行为具有实际的相关性。也就是说,当用户购买了一定数量的某一类产品,我们是否能够确定他们必然会成比例地购买另一种类的产品。有一个简单的方法可以检测相关性:我们用移除了某一个特征之后的数据集来构建一个监督学习(回归)模型,然后用这个模型去预测那个被移除的特征,再对这个预测结果进行评分,看看预测结果如何。
在下面的代码单元中,你需要实现以下的功能:
- 使用 `DataFrame.drop` 函数移除数据集中你选择的不需要的特征,并将移除后的结果赋值给 `new_data` 。
- 使用 `sklearn.model_selection.train_test_split` 将数据集分割成训练集和测试集。
- 使用移除的特征作为你的目标标签。设置 `test_size` 为 `0.25` 并设置一个 `random_state` 。
- 导入一个 DecisionTreeRegressor (决策树回归器),设置一个 `random_state`,然后用训练集训练它。
- 使用回归器的 `score` 函数输出模型在测试集上的预测得分。
```
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeRegressor
# TODO:为DataFrame创建一个副本,用'drop'函数丢弃一个特征# TODO:
new_data = data.drop('Fresh', axis=1)
# TODO:使用给定的特征作为目标,将数据分割成训练集和测试集
X_train, X_test, y_train, y_test = train_test_split(new_data, data['Fresh'], test_size=0.25, random_state=42)
# TODO:创建一个DecisionTreeRegressor(决策树回归器)并在训练集上训练它
regressor = DecisionTreeRegressor(random_state=42)
regressor.fit(X_train, y_train)
# TODO:输出在测试集上的预测得分
score = regressor.score(X_test, y_test)
print(score)
```
### 问题 2
你尝试预测哪一个特征?预测的得分是多少?这个特征对于区分用户的消费习惯来说必要吗?为什么?
**提示:** 决定系数(coefficient of determination),$R^2$ 结果在0到1之间,1表示完美拟合,一个负的 $R^2$ 表示模型不能够拟合数据。
**回答:**
* 尝试预测 Fresh, 预测的得分为-0.3857,这个得分表明模型不能够拟合数据,说明这个特征有必要用来区分用户。
### 可视化特征分布
为了能够对这个数据集有一个更好的理解,我们可以对数据集中的每一个产品特征构建一个散布矩阵(scatter matrix)。如果你发现你在上面尝试预测的特征对于区分一个特定的用户来说是必须的,那么这个特征和其它的特征可能不会在下面的散射矩阵中显示任何关系。相反的,如果你认为这个特征对于识别一个特定的客户是没有作用的,那么通过散布矩阵可以看出在这个数据特征和其它特征中有关联性。运行下面的代码以创建一个散布矩阵。
```
# 对于数据中的每一对特征构造一个散布矩阵
pd.plotting.scatter_matrix(data, alpha = 0.3, figsize = (40,20), diagonal = 'kde');
```
### 问题 3
这里是否存在一些特征他们彼此之间存在一定程度相关性?如果有请列出。这个结果是验证了还是否认了你尝试预测的那个特征的相关性?这些特征的数据是怎么分布的?
**提示:** 这些数据是正态分布(normally distributed)的吗?大多数的数据点分布在哪?
**回答:**
* 数据是正偏态分布的,大多数数据分布在均值左边。
* Milk 和 Grocery, Milk 和 Detergents_Paper, Grocery 和 Detergents_Paper 这3对数据从散布矩阵图中可以看出是呈线性相关的。
* Fresh与其他特征的分布图数据点比较分散,说明相关性弱,验证了前面的猜测。
## 数据预处理
在这个部分,你将通过在数据上做一个合适的缩放,并检测异常点(你可以选择性移除)将数据预处理成一个更好的代表客户的形式。预处理数据是保证你在分析中能够得到显著且有意义的结果的重要环节。
### 练习: 特征缩放
如果数据不是正态分布的,尤其是数据的平均数和中位数相差很大的时候(表示数据非常歪斜)。这时候通常用一个[非线性的缩放](https://github.com/czcbangkai/translations/blob/master/use_of_logarithms_in_economics/use_of_logarithms_in_economics.pdf)是很合适的,[(英文原文)](http://econbrowser.com/archives/2014/02/use-of-logarithms-in-economics) — 尤其是对于金融数据。一种实现这个缩放的方法是使用 [Box-Cox 变换](http://scipy.github.io/devdocs/generated/scipy.stats.boxcox.html),这个方法能够计算出能够最佳减小数据倾斜的指数变换方法。一个比较简单的并且在大多数情况下都适用的方法是使用自然对数。
在下面的代码单元中,你将需要实现以下功能:
- 使用 `np.log` 函数在数据 `data` 上做一个对数缩放,然后将它的副本(不改变原始data的值)赋值给 `log_data`。
- 使用 `np.log` 函数在样本数据 `samples` 上做一个对数缩放,然后将它的副本赋值给 `log_samples`。
```
# TODO:使用自然对数缩放数据
log_data = np.log(data)
# TODO:使用自然对数缩放样本数据
log_samples = np.log(samples)
# 为每一对新产生的特征制作一个散射矩阵
pd.plotting.scatter_matrix(log_data, alpha = 0.3, figsize = (14,8), diagonal = 'kde');
```
### 观察
在使用了一个自然对数的缩放之后,数据的各个特征会显得更加的正态分布。对于任意的你以前发现有相关关系的特征对,观察他们的相关关系是否还是存在的(并且尝试观察,他们的相关关系相比原来是变强了还是变弱了)。
运行下面的代码以观察样本数据在进行了自然对数转换之后如何改变了。
```
# 展示经过对数变换后的样本数据
display(log_samples)
```
### 练习: 异常值检测
对于任何的分析,在数据预处理的过程中检测数据中的异常值都是非常重要的一步。异常值的出现会使得把这些值考虑进去后结果出现倾斜。这里有很多关于怎样定义什么是数据集中的异常值的经验法则。这里我们将使用[ Tukey 的定义异常值的方法](http://datapigtechnologies.com/blog/index.php/highlighting-outliers-in-your-data-with-the-tukey-method/):一个异常阶(outlier step)被定义成1.5倍的四分位距(interquartile range,IQR)。一个数据点如果某个特征包含在该特征的 IQR 之外的特征,那么该数据点被认定为异常点。
在下面的代码单元中,你需要完成下面的功能:
- 将指定特征的 25th 分位点的值分配给 `Q1` 。使用 `np.percentile` 来完成这个功能。
- 将指定特征的 75th 分位点的值分配给 `Q3` 。同样的,使用 `np.percentile` 来完成这个功能。
- 将指定特征的异常阶的计算结果赋值给 `step`。
- 选择性地通过将索引添加到 `outliers` 列表中,以移除异常值。
**注意:** 如果你选择移除异常值,请保证你选择的样本点不在这些移除的点当中!
一旦你完成了这些功能,数据集将存储在 `good_data` 中。
```
# 对于每一个特征,找到值异常高或者是异常低的数据点
indexDict = dict()
for feature in log_data.keys():
# TODO: 计算给定特征的Q1(数据的25th分位点)
Q1 = np.percentile(log_data[feature],25)
# TODO: 计算给定特征的Q3(数据的75th分位点)
Q3 = np.percentile(log_data[feature],75)
# TODO: 使用四分位范围计算异常阶(1.5倍的四分位距)
step = 1.5*(Q3-Q1)
# 显示异常点
print("Data points considered outliers for the feature '{}':".format(feature))
display(log_data[~((log_data[feature] >= Q1 - step) & (log_data[feature] <= Q3 + step))])
outliers = log_data[~((log_data[feature] >= Q1 - step) & (log_data[feature] <= Q3 + step))].index.tolist()
for index in outliers:
if index not in indexDict:
indexDict[index] = 0
indexDict[index] += 1
# TODO(可选): 选择你希望移除的数据点的索引
outliers = [index for index in indexDict.keys() if indexDict[index] > 1]
outliers.sort()
print(outliers)
# 以下代码会移除outliers中索引的数据点, 并储存在good_data中
good_data = log_data.drop(log_data.index[outliers]).reset_index(drop = True)
```
### 问题 4
请列出所有在多于一个特征下被看作是异常的数据点。这些点应该被从数据集中移除吗?为什么?把你认为需要移除的数据点全部加入到到 `outliers` 变量中。
**回答:**
* [65, 66, 75, 128, 154]
* 应该从数据集中移除,因为主成分分析是考虑所有值的协方差,对异常值敏感。
## 特征转换
在这个部分中你将使用主成分分析(PCA)来分析批发商客户数据的内在结构。由于使用PCA在一个数据集上会计算出最大化方差的维度,我们将找出哪一个特征组合能够最好的描绘客户。
### 练习: 主成分分析(PCA)
既然数据被缩放到一个更加正态分布的范围中并且我们也移除了需要移除的异常点,我们现在就能够在 `good_data` 上使用PCA算法以发现数据的哪一个维度能够最大化特征的方差。除了找到这些维度,PCA 也将报告每一个维度的解释方差比(explained variance ratio)--这个数据有多少方差能够用这个单独的维度来解释。注意 PCA 的一个组成部分(维度)能够被看做这个空间中的一个新的“特征”,但是它是原来数据中的特征构成的。
在下面的代码单元中,你将要实现下面的功能:
- 导入 `sklearn.decomposition.PCA` 并且将 `good_data` 用 PCA 并且使用6个维度进行拟合后的结果保存到 `pca` 中。
- 使用 `pca.transform` 将 `log_samples` 进行转换,并将结果存储到 `pca_samples` 中。
```
# TODO:通过在good data上进行PCA,将其转换成6个维度
from sklearn.decomposition import PCA
pca = PCA(n_components=6, random_state=42)
pca.fit(good_data)
# TODO:使用上面的PCA拟合将变换施加在log_samples上
pca_samples = pca.transform(log_samples)
# 生成PCA的结果图
pca_results = vs.pca_results(good_data, pca)
```
### 问题 5
数据的第一个和第二个主成分**总共**表示了多少的方差? 前四个主成分呢?使用上面提供的可视化图像,从用户花费的角度来讨论前四个主要成分中每个主成分代表的消费行为并给出你做出判断的理由。
**提示:**
* 对每个主成分中的特征分析权重的正负和大小。
* 结合每个主成分权重的正负讨论消费行为。
* 某一特定维度上的正向增长对应正权特征的增长和负权特征的减少。增长和减少的速率和每个特征的权重相关。[参考资料:Interpretation of the Principal Components](https://onlinecourses.science.psu.edu/stat505/node/54)
**回答:**
* 第一个和第二个主成分总共表示了70.68%的方差,前四个总共93.11%
* 第一个主成分与Detergents_Paper,Milk,Grocery呈负相关,这3个组成中1个减少其他两个也随之减少,这个视为衡量客户规模。
* 第二个主成分与Fresh,Frozen,Delicatessen呈负相关,这个组成可以视为衡量用户的食品需求程度,餐馆,零售商。
* 第三个主成分与Fresh呈负相关,Delicatessen呈正相关,这个组成可以视为客户不同类型的食品需求,可能是不同类型的餐馆
* 第四个主成分与Frozen呈正相关,Delicatessen呈负相关,这个组成表明一般Frozen需求多的客户,Delicatessen会减少,这个也可能是不同类型的餐馆。
### 观察
运行下面的代码,查看经过对数转换的样本数据在进行一个6个维度的主成分分析(PCA)之后会如何改变。观察样本数据的前四个维度的数值。考虑这和你初始对样本点的解释是否一致。
```
# 展示经过PCA转换的sample log-data
display(pd.DataFrame(np.round(pca_samples, 4), columns = pca_results.index.values))
```
### 练习:降维
当使用主成分分析的时候,一个主要的目的是减少数据的维度,这实际上降低了问题的复杂度。当然降维也是需要一定代价的:更少的维度能够表示的数据中的总方差更少。因为这个,**累计解释方差比(cumulative explained variance ratio)**对于我们确定这个问题需要多少维度非常重要。另外,如果大部分的方差都能够通过两个或者是三个维度进行表示的话,降维之后的数据能够被可视化。
在下面的代码单元中,你将实现下面的功能:
- 将 `good_data` 用两个维度的PCA进行拟合,并将结果存储到 `pca` 中去。
- 使用 `pca.transform` 将 `good_data` 进行转换,并将结果存储在 `reduced_data` 中。
- 使用 `pca.transform` 将 `log_samples` 进行转换,并将结果存储在 `pca_samples` 中。
```
# TODO:通过在good data上进行PCA,将其转换成两个维度
pca = PCA(n_components=2, random_state=42)
pca.fit(good_data)
# TODO:使用上面训练的PCA将good data进行转换
reduced_data = pca.transform(good_data)
# TODO:使用上面训练的PCA将log_samples进行转换
pca_samples = pca.transform(log_samples)
# 为降维后的数据创建一个DataFrame
reduced_data = pd.DataFrame(reduced_data, columns = ['Dimension 1', 'Dimension 2'])
```
### 观察
运行以下代码观察当仅仅使用两个维度进行 PCA 转换后,这个对数样本数据将怎样变化。观察这里的结果与一个使用六个维度的 PCA 转换相比较时,前两维的数值是保持不变的。
```
# 展示经过两个维度的PCA转换之后的样本log-data
display(pd.DataFrame(np.round(pca_samples, 4), columns = ['Dimension 1', 'Dimension 2']))
```
## 可视化一个双标图(Biplot)
双标图是一个散点图,每个数据点的位置由它所在主成分的分数确定。坐标系是主成分(这里是 `Dimension 1` 和 `Dimension 2`)。此外,双标图还展示出初始特征在主成分上的投影。一个双标图可以帮助我们理解降维后的数据,发现主成分和初始特征之间的关系。
运行下面的代码来创建一个降维后数据的双标图。
```
# 可视化双标图
vs.biplot(good_data, reduced_data, pca)
```
### 观察
一旦我们有了原始特征的投影(红色箭头),就能更加容易的理解散点图每个数据点的相对位置。
在这个双标图中,哪些初始特征与第一个主成分有强关联?哪些初始特征与第二个主成分相关联?你观察到的是否与之前得到的 pca_results 图相符?
## 聚类
在这个部分,你讲选择使用 K-Means 聚类算法或者是高斯混合模型聚类算法以发现数据中隐藏的客户分类。然后,你将从簇中恢复一些特定的关键数据点,通过将它们转换回原始的维度和规模,从而理解他们的含义。
### 问题 6
使用 K-Means 聚类算法的优点是什么?使用高斯混合模型聚类算法的优点是什么?基于你现在对客户数据的观察结果,你选用了这两个算法中的哪一个,为什么?
**回答:**
* K-Means算法的优点是简单,迭代快速,对数据量有一定的伸缩性。
* 高斯混合聚类模型的优点是可以得出在各个分类的概率。
* 这个问题我们选用GMM,特征数据经过对数缩放后,在各维度更加接近正太分布,且散点图中我们看不出明显的分类,GMM能输出更多的信息。
### 练习: 创建聚类
针对不同情况,有些问题你需要的聚类数目可能是已知的。但是在聚类数目不作为一个**先验**知道的情况下,我们并不能够保证某个聚类的数目对这个数据是最优的,因为我们对于数据的结构(如果存在的话)是不清楚的。但是,我们可以通过计算每一个簇中点的**轮廓系数**来衡量聚类的质量。数据点的[轮廓系数](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.silhouette_score.html)衡量了它与分配给他的簇的相似度,这个值范围在-1(不相似)到1(相似)。**平均**轮廓系数为我们提供了一种简单地度量聚类质量的方法。
在接下来的代码单元中,你将实现下列功能:
- 在 `reduced_data` 上使用一个聚类算法,并将结果赋值到 `clusterer`,需要设置 `random_state` 使得结果可以复现。
- 使用 `clusterer.predict` 预测 `reduced_data` 中的每一个点的簇,并将结果赋值到 `preds`。
- 使用算法的某个属性值找到聚类中心,并将它们赋值到 `centers`。
- 预测 `pca_samples` 中的每一个样本点的类别并将结果赋值到 `sample_preds`。
- 导入 `sklearn.metrics.silhouette_score` 包并计算 `reduced_data` 相对于 `preds` 的轮廓系数。
- 将轮廓系数赋值给 `score` 并输出结果。
```
# TODO:在降维后的数据上使用你选择的聚类算法
from sklearn.metrics import silhouette_score
from sklearn.mixture import GaussianMixture
k = 2
clusterer = GaussianMixture(n_components=k,random_state=42)
clusterer.fit(reduced_data)
# TODO:预测每一个点的簇
preds = clusterer.predict(reduced_data)
# TODO:找到聚类中心
centers = clusterer.means_
# TODO:预测在每一个转换后的样本点的类
sample_preds = clusterer.predict(pca_samples)
# TODO:计算选择的类别的平均轮廓系数(mean silhouette coefficient)
score = silhouette_score(reduced_data, preds, random_state=42)
print("聚类数:{0} 平均轮廓系数{1}" .format(k, score))
```
### 问题 7
汇报你尝试的不同的聚类数对应的轮廓系数。在这些当中哪一个聚类的数目能够得到最佳的轮廓系数?
*回答*
* 聚类数:2 平均轮廓系数0.42191684646261485
* 聚类数:3 平均轮廓系数0.4042487382407879
* 聚类数:4 平均轮廓系数0.2932695648465841
* 聚类数:5 平均轮廓系数0.3004563887252593
* 聚类数:6 平均轮廓系数0.3261394504711576
* 聚类数:7 平均轮廓系数0.3242272053843606
* 聚类数:8 平均轮廓系数0.29647665639724213
* 聚类数:9 平均轮廓系数0.3071874795794863
当聚类数目为2时,得到最佳的轮廓系数
### 聚类可视化
一旦你选好了通过上面的评价函数得到的算法的最佳聚类数目,你就能够通过使用下面的代码块可视化来得到的结果。作为实验,你可以试着调整你的聚类算法的聚类的数量来看一下不同的可视化结果。但是你提供的最终的可视化图像必须和你选择的最优聚类数目一致。
```
# 从已有的实现中展示聚类的结果
vs.cluster_results(reduced_data, preds, centers, pca_samples)
```
### 练习: 数据恢复
上面的可视化图像中提供的每一个聚类都有一个中心点。这些中心(或者叫平均点)并不是数据中真实存在的点,但是是所有预测在这个簇中的数据点的平均。对于创建客户分类的问题,一个簇的中心对应于那个分类的平均用户。因为这个数据现在进行了降维并缩放到一定的范围,我们可以通过施加一个反向的转换恢复这个点所代表的用户的花费。
在下面的代码单元中,你将实现下列的功能:
- 使用 `pca.inverse_transform` 将 `centers` 反向转换,并将结果存储在 `log_centers` 中。
- 使用 `np.log` 的反函数 `np.exp` 反向转换 `log_centers` 并将结果存储到 `true_centers` 中。
```
# TODO:反向转换中心点
log_centers = pca.inverse_transform(centers)
# TODO:对中心点做指数转换
true_centers = np.exp(log_centers)
# 显示真实的中心点
segments = ['Segment {}'.format(i) for i in range(0,len(centers))]
true_centers = pd.DataFrame(np.round(true_centers), columns = data.keys())
true_centers.index = segments
display(true_centers)
```
### 问题 8
考虑上面的代表性数据点在每一个产品类型的花费总数,你认为这些客户分类代表了哪类客户?为什么?需要参考在项目最开始得到的统计值来给出理由。
**提示:** 一个被分到`'Cluster X'`的客户最好被用 `'Segment X'`中的特征集来标识的企业类型表示。
**回答:**
* Cluster 0代表餐馆类客户,因为这类客户中Milk,Grocery,Detergents_Paper都远低于样本均值,大部分需求是食品类商品。
* Cluster 1代表超市类客户,这类客户Milk,Grocery,Detergents_Paper都高于样本均值,这些数据符合常规超市的商品种类。
### 问题 9
对于每一个样本点**问题 8 **中的哪一个分类能够最好的表示它?你之前对样本的预测和现在的结果相符吗?
运行下面的代码单元以找到每一个样本点被预测到哪一个簇中去。
```
# 显示预测结果
for i, pred in enumerate(sample_preds):
print("Sample point", i, "predicted to be in Cluster", pred)
```
**回答:**
样本29,85与之前的预测相符,样本128与之前的预测不符,我觉得原因是前面预测时没有考虑到数据的相关性。
## 结论
在最后一部分中,你要学习如何使用已经被分类的数据。首先,你要考虑不同组的客户**客户分类**,针对不同的派送策略受到的影响会有什么不同。其次,你要考虑到,每一个客户都被打上了标签(客户属于哪一个分类)可以给客户数据提供一个多一个特征。最后,你会把客户分类与一个数据中的隐藏变量做比较,看一下这个分类是否辨识了特定的关系。
### 问题 10
在对他们的服务或者是产品做细微的改变的时候,公司经常会使用 [A/B tests ](https://en.wikipedia.org/wiki/A/B_testing)以确定这些改变会对客户产生积极作用还是消极作用。这个批发商希望考虑将他的派送服务从每周5天变为每周3天,但是他只会对他客户当中对此有积极反馈的客户采用。这个批发商应该如何利用客户分类来知道哪些客户对它的这个派送策略的改变有积极的反馈,如果有的话?你需要给出在这个情形下A/B 测试具体的实现方法,以及最终得出结论的依据是什么?
**提示:** 我们能假设这个改变对所有的客户影响都一致吗?我们怎样才能够确定它对于哪个类型的客户影响最大?
**回答:**
我们可以利用销售额来确定配送策略改变是否有效。
* 将客户分为A1,A2,B1,B2,其中A1、B1是前面分类为Cluster 0的客户,A2、B2是前面分类为Cluster 1的客户。
* 对A1、A2采用每周5天,对B1、B2采用每周3天的派送服务。
* 统计A1、A2、B1、B2的销售额。
* 如果B1销售额大于A1,则派送策略改变对Cluster 0客户有效; 如果B2销售额大于A2,则派送策略改变对Cluster 1客户有效
* 通过不同类客户的不同派送策略的增长率,我们可以得出影响最大的那一类客户。
### 问题 11
通过聚类技术,我们能够将原有的没有标记的数据集中的附加结构分析出来。因为每一个客户都有一个最佳的划分(取决于你选择使用的聚类算法),我们可以把用户分类作为数据的一个[工程特征](https://en.wikipedia.org/wiki/Feature_learning#Unsupervised_feature_learning)。假设批发商最近迎来十位新顾客,并且他已经为每位顾客每个产品类别年度采购额进行了预估。进行了这些估算之后,批发商该如何运用它的预估和非监督学习的结果来对这十个新的客户进行更好的预测?
**提示**:在下面的代码单元中,我们提供了一个已经做好聚类的数据(聚类结果为数据中的cluster属性),我们将在这个数据集上做一个小实验。尝试运行下面的代码看看我们尝试预测‘Region’的时候,如果存在聚类特征'cluster'与不存在相比对最终的得分会有什么影响?这对你有什么启发?
```
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
# 读取包含聚类结果的数据
cluster_data = pd.read_csv("cluster.csv")
y = cluster_data['Region']
X = cluster_data.drop(['Region'], axis = 1)
# 划分训练集测试集
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=24)
clf = RandomForestClassifier(random_state=24)
clf.fit(X_train, y_train)
score_with_cluster = clf.score(X_test, y_test)
# 移除cluster特征
X_train = X_train.copy()
X_train.drop(['cluster'], axis=1, inplace=True)
X_test = X_test.copy()
X_test.drop(['cluster'], axis=1, inplace=True)
clf.fit(X_train, y_train)
score_no_cluster = clf.score(X_test, y_test)
print("不使用cluster特征的得分: %.4f"%score_no_cluster)
print("使用cluster特征的得分: %.4f"%score_with_cluster)
```
**回答:**
* 使用cluster特征后得分更高,我们以后在监督学习过程中,可以先使用聚类技术来给数据添加新的分类特征,这可能会提供预测准确性。
### 可视化内在的分布
在这个项目的开始,我们讨论了从数据集中移除 `'Channel'` 和 `'Region'` 特征,这样在分析过程中我们就会着重分析用户产品类别。通过重新引入 `Channel` 这个特征到数据集中,并施加和原来数据集同样的 PCA 变换的时候我们将能够发现数据集产生一个有趣的结构。
运行下面的代码单元以查看哪一个数据点在降维的空间中被标记为 `'HoReCa'` (旅馆/餐馆/咖啡厅)或者 `'Retail'`。另外,你将发现样本点在图中被圈了出来,用以显示他们的标签。
```
# 根据‘Channel‘数据显示聚类的结果
vs.channel_results(reduced_data, outliers, pca_samples)
```
### 问题 12
你选择的聚类算法和聚类点的数目,与内在的旅馆/餐馆/咖啡店和零售商的分布相比,有足够好吗?根据这个分布有没有哪个簇能够刚好划分成'零售商'或者是'旅馆/饭店/咖啡馆'?你觉得这个分类和前面你对于用户分类的定义是一致的吗?
**回答:**
* 这个分布和我们之前的聚类算法得出的结果基本一致。
* 数据之间有交叉重叠,没有簇能刚好划分数据。
* 这个分类和我对用户分类定义相反。
> **注意**: 当你写完了所有的代码,并且回答了所有的问题。你就可以把你的 iPython Notebook 导出成 HTML 文件。你可以在菜单栏,这样导出**File -> Download as -> HTML (.html)**把这个 HTML 和这个 iPython notebook 一起做为你的作业提交。
| github_jupyter |
# BRIL Work Suite
<hr>
A commandline toolkit for CMS Beam Radiation Instrumentation and Luminosity (BRIL).
The BRIL Work Suite works on top of the python virtual environment brilconda which is a custom build of the <a href="https://store.continuum.io/cshop/anaconda/">Anaconda Scientific Python</a> distribution. The toolkit is compatible with python2 and python3.
As prerequisite, the python virtual environment should be centrally or privately installed.
And there is NO requirement on the PYTHONPATH nor LD_LIBRARY_PATH to envoke the software.
The virtual environment brilconda has much longer release cycle than the bril toolkit.
</hr>
##### Documentation for release 3.6.x
<p>
**Table Of Contents:**
1. [Prerequisite for running and installing brilws](#prerequisite)
2. [Install brilws](#installation)
3. [Install brilconda virtual environment](#brilcondainstallation)
4. [Command Structure](#commandstructure)
5. [brilcalc](#brilcalc)
* [lumi](#brilcalclumi)
* [beam](#brilcalcbeam)
* [trg](#brilcalctrg)
6. [briltag](#briltag)
* [listiov](#briltaglistiov)
* [insertiov](#briltaginsertiov)
* [listdata](#briltaglistdata)
* [insertdata](#briltaginsertdata)
7. [Appendix A : trigger rules](#triggerrules)
<a id='top'></a>
<a id='prerequisite'></a>
## Prerequisite for running and installing brilws
1. work with centrally installed virtual environment:
*python2*
```
export PATH=$HOME/.local/bin:/cvmfs/cms-bril.cern.ch/brilconda/bin:$PATH (bash)
```
*python3*
```
export PATH=$HOME/.local/bin:/cvmfs/cms-bril.cern.ch/brilconda3/bin:$PATH (bash)
```
2. work with central installationon bril online cluster in .cms
*python2*
```
export PATH=$HOME/.local/bin:/nfshome0/lumipro/brilconda/bin:$PATH (bash)
```
*python3*
```
export PATH=$HOME/.local/bin:/nfshome0/lumipro/brilconda3/bin:$PATH (bash)
```
3. privately installation of virtual environment
```
export PATH=<brilcondabasedir>/bin:$PATH (bash)
```
Note: the **\$PATH** requirement must be fullfilled for the **shell** from which the brilws and pip commands run. Be aware that external scripts, e.g. cmsenv.sh, alter the **\$PATH** environment, users should make sure the environment is as above **at the moment of** running the brilws and pip commands.
<a id='installation'></a>
## Installation of brilws
**Please do not attempt this if the prerequisite condition is not satisfied**
### Install brilws
#### cern
```
pip install --user brilws
```
#### online (on bril host)
```
pip install --no-index --find-links=file:///nfshome0/lumipro/installers/linux-64 --user brilws
```
#### private virtual environment brilconda
```
pip install brilws
```
### Install a specific version
Specify the version of the package with == operator: e.g. brilws==3.6.2
### Uninstall brilws
**Please do not attempt this if prerequisite condition is not satisfied**
```
pip uninstall brilws
```
### Check most recent version (Please check prerequisite condition is satisfied before anything)
```
pip show brilws
```
<a id='brilcondainstallation'></a>
## Install virtual environment brilconda (Optional)
<p>
The virtual environment is centrally installed in the following locations:
#### cvmfs : /cvmfs/cms-bril.cern.ch/brilconda (python2), /cvmfs/cms-bril.cern.ch/brilconda3 (python3)
Note: the cvmfs mount is on-demand. It does not appear unless specifically accessed.
#### bril online cluster : /nfshome0/lumipro/brilconda (python2), /nfshome0/lumipro/brilconda3 (python3)
If local resource permits, users can choose to install the brilconda environment privately for more control over the working environment. Install brilconda on different platforms:
##### Linux-x86_64 with installer
###### python2
```
wget https://cern.ch/cmslumisw/installers/linux-64/Brilconda-1.1.7-Linux-x86_64.sh
bash Brilconda-1.1.7-Linux-x86_64.sh -b -p <localbrilcondabase>
```
###### python3
```
wget https://cern.ch/cmslumisw/installers/linux-64/Brilconda-3.2.16-Linux-x86_64.sh
bash Brilconda-3.2.16-Linux-x86_64.sh -b -p <localbrilcondabase>
```
##### MacOS-64 with installer
###### python2
```
wget https://cern.ch/cmslumisw/installers/macos-64/Brilconda-1.1.7-MacOSX-x86_64.sh
bash Brilconda-1.1.7-MacOSX-x86_64.sh -b -p <localbrilcondabase>
```
###### python3
```
wget https://cern.ch/cmslumisw/installers/macos-64/Brilconda-3.2.16-MacOSX-x86_64.sh
bash Brilconda-3.2.16-MacOSX-x86_64.sh -b -p <localbrilcondabase>
```
Regardless the location of the installation, the virtual environment is activated whenever its bin directory is in the PATH, and disactivated when the bin directory is not present in the PATH.
<p>
<a id='offsite access'></a>
### Off-site access
<p>
BRIL tools analyse data in the database server at CERN which is closed to the outside. Therefore the most convienient way is to run the toolkit on hosts (private or public) at CERN. If you must use the software installed outside the CERN intranet, a ssh tunnel to the database server at CERN has to be established first. Since the tunneling procedure requires a valid cern computer account, a complete unregistered person will not be able to access the BRIL data in any case.
<p>
The following instruction assumes the easiest setup: you have **two** open sessions on the same off-site host, e.g. cmslpc32.fnal.gov, one for the ssh tunnel and another for execution. It is also assumed that all the software are installed and the $PATH variable set correctly on the execution shell.
<p>
#### ssh tunneling session
```
ssh -N -L 10121:itrac5117-v.cern.ch:10121 <cernusername>@lxplus.cern.ch
```
Provide you cern account password and the tunnel will kept open without getting back the shell prompt.
#### open an execution session on the same host that has the ssh tunnel
```
brilcalc lumi -c offsite -r 281636
```
<p>
Run bril tools with connection string **-c offsite**.
<p>
<a id='web cache access'></a>
### web-cache access
<p>
In addition to online, offline, on-site and off-site access to bril database server, indirect access
via web cache is also supported for bril tools (read-only). Specificlly, web cache refers to the
frontier/squid infrastructure available for all CMS sites. (It is also possible to have your own private setup.)
</p>
<p>
Using web cache access allows queries and results to be cached close to you therefore reducing the overall load on the database server at CERN. It is much faster to get already cached results. Another convenience is that accessing data from outside the CERN firewall does not require ssh tunnels in this case.
</p>
<p>
However, keep in mind that web cached are not real time data. The caches are reset every about 5 hours. In another word, the most recent results are not necessarily correct. For example, querying last run from the cache might show 5 lumi sections, while there were actually 200 lumi sections in the database. You get 5 because the cache close to you was already populated by yourself or other peoples since the last cache reset.
Any update operation on the database server will not be reflected by the cache for a few hours.
</p>
<p>
Weigh carefully advantage and disadvantage and use your own judgement when choosing between direct and indirect accesses.
</p>
#### use the cache server at CERN
Option ** -c web ** directs you to the web cache server at CERN ** cmsfrontier.cern.ch:8000/LumiCalc **. e.g.
```
brilcalc lumi -c web -r 284077
```
#### use any cache server
Option **```-c <config file>```** allows you to use the web cache most conveniently located and configured.
The configuration file can be found at each CMS site, it is normally called as "site-local-config.xml".
You can use it as it is or use it as a starting example to build your own configuration:
```
brilcalc lumi -r 284077 -c /afs/cern.ch/cms/SITECONF/CERN/JobConfig/site-local-config.xml
```
To build custom cache config xml, only contents enclosed in the ```<frontier-connect>``` tag are required by the bril tool, all other tags are ignored: e.g.
```
more /home/me/mycache.xml
<frontier-connect>
<proxy url="http://cmst0frontier.cern.ch:3128"/>
<proxy url="http://cmst0frontier.cern.ch:3128"/>
<proxy url="http://cmst0frontier1.cern.ch:3128"/>
<proxy url="http://cmst0frontier2.cern.ch:3128"/>
<server url="http://cmsfrontier.cern.ch:8000/FrontierInt"/>
<server url="http://cmsfrontier.cern.ch:8000/FrontierInt"/>
<server url="http://cmsfrontier1.cern.ch:8000/FrontierInt"/>
<server url="http://cmsfrontier2.cern.ch:8000/FrontierInt"/>
<server url="http://cmsfrontier3.cern.ch:8000/FrontierInt"/>
<server url="http://cmsfrontier4.cern.ch:8000/FrontierInt"/>
</frontier-connect>
brilcalc lumi -r 284077 -c /home/me/mycache.xml
```
Note: if you are working in the .cms network, you can not use the web cache. Anyway there is no need for that.
<p style="text-align:center">[Back to Top](#top)
<a id='commandstructure'></a>
## Commands
<p>
**Please do not attempt anything bellow if prerequisite condition is not satisfied**
The commandlines in the work suite are made of main and sub commands. Each subcommand have its independent help menus. For example, brilcalc lumi -h, where brilcalc is the main command, lumi is the subcommand.
```
brilcalc -h
```
```
briltag -h
```
```
brilcalc lumi -h
```
```
brilcalc beam -h
```
```
brilcalc trg -h
```
<a id='brilcalc'></a>
## brilcalc
<p>
<a id='common options'></a>
##### common command options
<p>There are four categories of common command options: selection, filter, output and display, database connection
* Selections: select on input lumi section, run, fill number or begin/end time in terms of UTC time or fill/run number. These selections can be overlapping.
* Filters: Filter on the result
* Output and Display
* Database connection
<p>Reasonable default parameters are given, readonly users do not need to specify these options.
</p>
<a id='brilcalclumi'></a>
##### lumi
<p>
Subcommand to show luminosity measured by BRIL. By default, i.e. without --type or --normtag options, the summary of the best known luminosity from online is shown. Switches and options specific to the subcommand are explained below.
<p>
###### --filedata FILEDATA
<p>
Raw data can be fetched from files and combined with other data from database to produce calibrated luminosity results. When using input from files, database connection parameters, implicitly or explicitly given, are still required. This option is an extention rather than a replacement of the general database connection options.
</p>
<p>Results will have the same granularity as that of the input. If inputs from files are in lumi-section, then the output is per LS, if inputs are in NB4, then output is on NB4. Since the NB4 granularity is 16 times larger than the LS granularity, this option should be used on small data sets, for example, those on VDM scans.
</p>
<p>
The input parameter can be either a directory or a single file. The input files must be in hdf5 format and contain at least the BRIL "tcds" table.
</p>
<p>
The file input option works only together with --byls or --byls --xing. And the output luminosity unit is fixed hz/ub. Using option -u will not change the output unit.
</p>
<p>
In this case, because the output is instantaneous luminosity and the integration time is unknown a priori by the software, the summary sum table cannot be calculated therefore will not be displayed at the end with this option.
</p>
###### --byls switch
Show luminosity and average pileup by lumi section.
###### --xing switch
Show luminosity by lumi section and per bunch crossing. The bxidx field of the output has the range of [1,3564]. More often not all 3564 possible bunches are interesting. Three filters can be specified with this switch to reduce the total output. The filters can be applied accumulatively.
* Bunch filters specific to --xing switch
###### --normtag NORMTAG
Apply calibration/correction function defined by a tag. Each norm tag is bound to a specific luminometer. By using the tag, you have automatically selected a luminometer.
When selecting data from a combination of luminometers, a json format file or string is accepted (specified below). In this case, both -i and --normtag options can select run and lumi sections. When both given, the **intersection** of the two are used. Also, it is required that --normtag selection is a superset of the -i selection. On failing this check, warning messages will be printed but the program continues.
The composite selection is a list of lists as described below:
<pre>
[
[normtag,{run:[[startls,stopls],[startls,stopls]...]} | run, run, ... ],
[normtag,{run:[[startls,stopls],[startls,stopls]...]} | run, run, ... ],
...
]
</pre>
Run, ls selection matrices follow each normtag in which the chosen lumi sections are valid.
Here "|" means "or". Flat format of the inner list [normtag, run, run,...] is a shortcut to indicate that all lumi sections in the run are selected. Internally each run number string is translated to {run:[[1,999999]]}.
Example below:
<pre>
[ ["bcm1fv1",{"251118":[[1,20]]}],
["hfocv1",{"251118":[[21,21]]}],
["bcm1fv1",{"251118":[[22,24]]}],
["bcm1fv1","251119","251131"],
["hfocv1","251134"],
["bcm1fv1","251167"],
["hfocv1","251168"]
]
</pre>
When this type of selection file is given --type argument has no more effects.
The same rules apply for quoting as with -i : when in file, **key** fields and **string** fields must be quoted. While as string input, the entire string should be quoted.
###### -u UNIT
Show luminosity in the specified unit and scale the output value accordingly. Recognised unit strings are
<pre>
/kb , 1e21/cm2 , hz/kb , 1e21/cm2s
/b , 1e24/cm2 , hz/b , 1e24/cm2s
/mb , 1e27/cm2 , hz/mb , 1e27/cm2s
/ub , 1e30/cm2 , hz/ub , 1e30/cm2s
/nb , 1e33/cm2 , hz/nb , 1e33/cm2s
/pb , 1e36/cm2 , hz/pb , 1e36/cm2s
/fb , 1e39/cm2 , hz/fb , 1e39/cm2s
/ab , 1e42/cm2 , hz/ab , 1e42/cm2s
</pre>
###### --type LUMITYPE
Show results from the selected luminometer. Current available choices are hfoc,hfet,bcm1f,bcm1fsi,bcm1futca,pltzero,pltslink,dt,pxl,ramses,radmon.
This option can be skipped if --normtag is specified because every norm tag is bound to a specific luminometer. If option --normtag is present, then --type is ignored.
Option --type is **required** if **--without-correction** switch is specified.
###### --without-correction
Show raw data taken by a specific luminometer. --type LUMITYPE must be provided if this switch is specified.
###### --datatag DATATAG
A specific version of lumi and beam data. The initial datatag "online" is guaranteed to exist. Other data tags are the patches to its previous version. If not specified, the most recent datatag, i.e. the best version of data, is used.
Trigger related data are summary copies of trigger databases, they are not versioned.
###### --precision PRECISION
Define the luminosity value output format and precision. The format string can be "[0-9]f" or "[0-9]e". The integer number specifies the precision and "f" stands for fixed floating point format, "e" for the scientific notation. Without this option, the default luminosity value output is "9f". Choose the precision and format according to your need. Too high precision output slows down the program and bloat the output file on disk. In addition, the luminosity number output precision can be reduced by using -u UNIT or -n SCALEFACTOR options.
###### --hltpath HLTPATH
**HLTPATH** is hlt path name or pattern. The string pattern uses the file system convention fnmatch. \*, ? ,\[seq], \[!seq] operators are recognised. In order not to confuse with the file system, please double quote the hltpath string if there are special char \*, ?, \[] \!
Show nominal luminosity scaled down by HLT and L1 prescale factors also taking into account L1 bit masks. Nominal luminosity means that taken during CMS running.
This scaling filter has effect on all lumi switches or without, e.g. --byls, --xing. The output format could be different from the default one: one or two fields are removed and replaced with the long HLT path string.
Any of the following conditions cause the rejection of a lumisection in the selected path: hlt prescale=0, relavent l1 bits prescale=0 or relavent l1 bits masked.
Each output unit is grouped by hlt path name meaning if the output is one per lumi section, then for each of them, all possible hltpaths are shown before changing to the next unit.
Since release >=3.3.0 Sum delivered, Sum recorded fields in the Summary section show the sum luminosity of all groups above. Their meaning depends on the user's context.
###### --ignore-mask switch
Switch off the effect of L1 bit masks. This should be used only for debugging purpose. The masks should be considered for the result to be correct.
###### --without-checkjson switch
Switch off cross-checking with the json selection.
###### --minBiasXsec MINBIASXSEC
Specify minimum bias cross section (ub) to use when calculating the average pileup column for the --byls output. This option should be used together with --type or --normtag. The program does not accept this option if neither --type nor --normtag is found.
**Note:** the average pileup per lumi section is recalculated only if both --byls --normtag or --byls --type are specified. In other cases, the result shown is what calculated online during data acquisition time and cannot be changed.
Reference minbias cross section table:
Single Beam E (Gev) | Accelerator mode | minBiasXsec (ub)
--- | --- | ---
6500 | pp | 80000
2510 | pp | 65000
6370 | pbpb | 7800000
3500 | pp | 71500
4000 | pp | 73000
5000 | ppb | 2061000
8160 | ppb | 2135000
2720 | xexe | 5650000
<a id='brilcalcbeam'></a>
##### beam
Subcommand to show beam information (timeinfo, single beam energy, beam intensities, ncolliding bunches) per lumi section.
<p>
###### --xing switch
Show beam intensities per bunch crossing. The bxidx field of the output has the range of [1,3564]
<a id='brilcalctrg'></a>
##### trg
Subcommand to show L1, HLT bit/path and prescale information per run/per lumi section. The hlt path shown by this subcommand is constraint to only those potentially relevant to luminosity. Paths containing substring such as "Calibration","Random","DQM","Physics","Activity" are ignored.
<p>
Without any filter options the summary map of the hltconfig to run number is shown. Switches and filter options specific to this subcommand are explained below.
###### --pathinfo switch
Show HLT path and its L1 seed expression as well as the logical relationship between the l1 bits in the expression. Complicated L1seed logics with no simple scaling relationship between L1 bits are excluded. Filter options --hltconfig , -r, --hltpath reduce the output size.
###### --prescale switch
If no --hltpath filter is specified, show prescale index change during the given run. If --hltpath filter is given, show the real HLT and L1 prescale values of the selected run on the selected hltpath(s).
Note: if the selected path is not effective , e.g. prescale=0 or all its seed bits are masked or having prescale=0, no result will be shown. Only truely effective hltpath are shown. You can use the --ignore-mask option to relax the requirement on l1bit masks. But keeping in mind that for the correctness of the result L1 mask should not be ignored, it is the same as prescale value=0.
###### trg subcommand specific selection options
Since L1 and HLT information are not associated with LHC fill numbers, this subcommand supports only one common time selection option -r RUN.
<p style="text-align:center">[Back to Top](#top)
<a id='briltag'></a>
## briltag
<p>
Subcommand to manage norm and data tags. Tags are used for versioning.
Calibration and correction factors can be applied offline and having different values at different time period. Norm tag describes the function and parameters applied to the raw data. Each correction has its validity in time( run number ).
Sometimes the raw data need to be patched. Data tag is the version of the raw data. For the moment, there is only one version "online". And the tool is not fully developed for handling data tags.
##### common command options
<p> Database connection parameters -c, -p are common. However, one should distinguish readonly and readwrite operations. listxxx commands are readonly therefore -c, -p option values work as normal. insertxxx commands involve database write operations, only BRIL operators can perform it using specific -c, -p parameters.
<a id='briltaglistiov'></a>
##### listiov
Show norm tag info of each luminometer. By default, the summary is shown. With **--name** it shows the detail of the selected tag.
<p> Some predefined functions shown below.
function | description | parammeters
--- | --- | ---
**poly1d** | polynomial | coefs: array of coefficients
**inversepoly1d** | 1/polynomial | coefs: array of coefficients
**afterglow** | afterglow correction by bx threshold | afterglowthresholds: array of (nbx threshold,factor)
**poly1dWafterflow** | poly1d followed by afterglow correction | coefs, afterglowthresholds
**inversepoly1dWafterglow** | product of inverse poly1d and afterglow | coefs, afterglowthresholds
**poly2dlL** | polynomial2d c[i,j]\*x^i*y^j where x=per bunch lumi , y=totallumi | coefs: array of coefficients
**poly2dlLWafterglow** | poly2dlL followed by afterglow correction | coefs, afterglowthresholds
**afterglowWpoly2dlL** | afterglow correction followed by poly2dlL | coefs, afterglowthresholds
**Note** :
The correction function is on the **single bunch instantaneous luminosity (SBIL)**. Its effect on the luminosity integrated over all orbits is calculated and applied automatically by the software based on the assumption that the bunches are equally populated.
<a id='briltaginsertiov'></a>
##### insertiov
To insert normtag data into database. Data to insert must be defined in an input file of yaml format and given to the -y option.
<p>
<a id='briltaglistdata'></a>
##### listdata
List all data tags
<p>
<a id='briltaginsertdata'></a>
##### insertdata
Creat a new data tag. lumi beam data are versioned, trigger data are not. The condition to open a new datatag is if there are old data under the most recent tag for the lumi sections to load (patch). Loading new data to the most recent tag does not require opening a new one. For example, if loading run=1,ls=1 to tag v1 would cause a duplicated id error, then you need to create a data tag v2.
<p>
<p style="text-align:center">[Back to Top](#top)
<a id='triggerrules'></a>
## Appendix A : Trigger Rules
<p>
Effective luminosity in a given HLT path is : total_luminosity/(HLT_prescale*L1_prescale).
Rules on deducing L1 contribution to the total prescale factor :
1. Do not consider any masked L1 bit in the seed expression, unless --ignore-mask switch is specified.
2. If the path is single seeded, take the seed bit's prescale value.
3. If (A.1 OR A.2 OR A.3...) , ignore the triggers with prescale 0, take the minimum of the remaining prescale values.
4. If (A AND B AND C...), take the product of the prescale values.
5. If NOT, or a combination of AND/OR/NOT, reject this path.
6. For HLT_L1seed logic with more than one logical operators, users are considered taking their own risk and should use own judgement.
Note: above rules are effective since release 2.1.x.
Two suggested by not implemented rules:
1. (NOT IMPLEMENTED) If (A.1 OR A.2 OR A.3...) AND (B.1 OR B.2 OR B.3...) AND ... , for each group, take the minimum of the non-zero prescal, or zero if they are all zero. Then the product of the prescales of each group. Note: not implemented means such path are rejected by brilcalc.
2. (NOT IMPLEMENTED) In some cases, L1 masks and prescale should be ignored. Note: not implemented means users are expected know about such cases and use --ignore-mask by themselves.
<p style="text-align:center">[Back to Top](#top)
```
from IPython.core.display import HTML
def css_styling():
styles = open('./style/custom.css','r').read()
return HTML(styles)
css_styling()
```
| github_jupyter |
```
import pandas as pd
all_muts_df = pd.read_pickle("./data/2_5_df.pkl")
display(all_muts_df.shape, all_muts_df.head())
cog_df = pd.read_pickle("./data/COG_df.pkl")
TU_objects_df = pd.read_pickle("./data/TU_objects_df.pkl")
operon_df = pd.read_pickle("./data/operon_df.pkl")
TU_df = pd.read_pickle("./data/TU_df.pkl")
gene_synonym_df = pd.read_pickle("./data/gene_synonym_df.pkl")
```
!!! Currently using lists instead of sets for COG hits because I want to know the count of items per COG affected with each mutation, where a mutation can affect multiple unique genomic targets that belong to the same COG. If I used a set, would lose the magnitude of hits to a COG per mutations.
THIS SHOULD GET FIXED WITH COG LINK DICTS, WHERE THE NUMBER OF TIMES AN OPERON SHOWS UP CAN IN THE LINKING LIST CAN BE THE NUMBER OF GENES INVOVLED IN MUTATING THE COG.
```
from util.gene import get_gene_bnum
# This is currently not attributing gene without COG to "Function unknown"
# Last time checking this, it wasn't necessary to add the addition "Function unknown" entries here
# though these extra entries were required with the link dictionaries
def _get_COGs(gene_id):
annots = []
bnum = get_gene_bnum(gene_id, gene_synonym_df)
gene_cog_df = cog_df[cog_df["locus"] == bnum]
if len(gene_cog_df) > 0:
for COG in gene_cog_df["COG description"]:
d = {"name": COG}
annots.append(d)
return annots
def get_COGs(mut_row):
annots = []
for feat_d in mut_row["genomic features"]:
if feat_d["genetic"]:
annots += _get_COGs(feat_d["RegulonDB ID"])
# This will catch whether no features exist in the feat_ID_set
# or if those features are just integenic regions.
# Get operon genetic feature link dict through overlapping mutation and operon ranges.
# All genetic features are annotated within both the "genetic features" and "genetic feature links" column.
# If no integenic genomic feature is hit by a mutation, use the genetic feature range in an overlap analysis with operons.
else:
TU_IDs = set(TU_objects_df[TU_objects_df["TU_OBJECT_ID"] == feat_d["RegulonDB ID"]]["TRANSCRIPTION_UNIT_ID"])
for TU_ID in TU_IDs:
subset_TU_genes_df = TU_objects_df[(TU_objects_df["TRANSCRIPTION_UNIT_ID"] == TU_ID) & (
TU_objects_df["TU_OBJECT_CLASS"] == 'GN')]
for gene_id in subset_TU_genes_df["TU_OBJECT_ID"].drop_duplicates():
# RegulonDB can return 'nan' or '' gene names
if str(gene_id) != 'nan' and gene_id != '':
annots += _get_COGs(gene_id)
return annots
test_mut_row_d = {
"exp": "test exp",
"genomic features":
[
{'name': 'rph-pyrE attenuator terminator',
'RegulonDB ID': 'ECK125144791',
'range': (3815799, 3815828),
'genetic': False,
'feature type': 'attenuator terminator',
'operon': 'rph-pyrE',
'mutation set count': 5,
'significant': True},
{'name': 'rph',
'RegulonDB ID': 'ECK120000854',
'range': (3815863, 3816549),
'genetic': True,
'feature type': 'gene',
'operon': 'rph-pyrE',
'mutation set count': 5,
'significant': True}
]}
assert(get_COGs(test_mut_row_d) == [{'name': 'Nucleotide transport and metabolism'}])
all_muts_df["COGs"] = all_muts_df.apply(lambda r: get_COGs(r), axis=1)
all_muts_df.head()
from util.operon import get_operon_name_set, get_operon_ID_set
def _add_links_to_d(bnum, d, op_ID_set):
gene_cog_df = cog_df[cog_df["locus"]==bnum]
if len(gene_cog_df) > 0:
for cog in gene_cog_df["COG description"]:
if cog not in d.keys(): d[cog] = []
d[cog] += list(op_ID_set)
# else:
# if "Function unknown" not in d.keys(): d["Function unknown"] = []
# d["Function unknown"] = list(op_ID_set)
def get_COG_operon_links(mut_row):
COG_op_link_d = dict()
for feat_d in mut_row["genomic features"]:
# link_exists = False
if feat_d["genetic"]:
op_ID_set = get_operon_ID_set(feat_d["RegulonDB ID"], TU_objects_df, TU_df, operon_df)
if str(feat_d["RegulonDB ID"]) != 'nan' and feat_d["RegulonDB ID"] != '':
bnum = get_gene_bnum(feat_d["RegulonDB ID"], gene_synonym_df)
_add_links_to_d(bnum, COG_op_link_d, op_ID_set)
# link_exists = True
else:
TU_IDs = set(TU_objects_df[TU_objects_df["TU_OBJECT_ID"]==feat_d["RegulonDB ID"]]["TRANSCRIPTION_UNIT_ID"])
for TU_ID in TU_IDs:
subset_TU_genes_df = TU_objects_df[(TU_objects_df["TRANSCRIPTION_UNIT_ID"]==TU_ID) & (TU_objects_df["TU_OBJECT_CLASS"]=='GN')]
op_ID_set = set()
for _, TU_gene in subset_TU_genes_df.iterrows():
op_ID_set = op_ID_set | get_operon_ID_set(TU_gene["TU_OBJECT_ID"], TU_objects_df, TU_df, operon_df)
for gene_id in subset_TU_genes_df["TU_OBJECT_ID"].drop_duplicates():
if str(gene_id) != 'nan' and gene_id != '': # RegulonDB can return 'nan' or '' gene names
bnum = get_gene_bnum(gene_id, gene_synonym_df)
_add_links_to_d(bnum, COG_op_link_d, op_ID_set)
# link_exists = True
# Per mutation feature
# Need to be looking at previously populated level of annotations depdending on how the current level is being populated
# for regulon, it just means that if there are no operons, go immediately to genomic features.
# if link_exists == False:
# if "Function unknown" not in COG_op_link_d.keys(): COG_op_link_d["Function unknown"] = []
# if len(operon_name_set) > 0: # need to always check the nearest annotation type for defined features
# COG_op_link_d["Function unknown"] += list(operon_name_set)
# else:
# COG_op_link_d["Function unknown"].append(feat_d["RegulonDB ID"])
return COG_op_link_d
all_muts_df["COG links"] = all_muts_df.apply(get_COG_operon_links, axis=1)
all_muts_df.head()
all_muts_df.to_pickle("./data/2_6_df.pkl")
```
| github_jupyter |
```
import sys
sys.path.append('../python_packages_static/')
import pyemu
import os
import glob
import shutil
```
# We want to set up for global sensitivity analysis.
### NOTE: Make sure `run_ensemble` is set appropriately - If `run_ensemble` is set to `True`, local runs are performed. If `run_ensemble` set to `False`results from the journal article are used.
```
run_ensemble=False
pst_root = 'prior_mc_wide'
if run_ensemble==True:
pst = pyemu.Pst(f'../noptmax0_testing/{pst_root}.pst')
else:
pst = pyemu.Pst(f'../output/noptmax0/{pst_root}.pst')
output_dir = '../run_data'
```
### set `tie_by_group` to `True`. Also update the lower bound for CHD parameters to nearly 1.0 (because many of these values are at or near the bottom of model cells and if sampling sets them below the bottom of the cell, MODFLOW6 will not run). Also unfix the CHD parameters so they will be evaluated. All other defailts. for `pestpp-sen` will be accepted.
```
pst.pestpp_options['tie_by_group'] = True
pst.parameter_data.loc[pst.parameter_data.pargp=='chd', 'parlbnd'] = 0.999999
pst.parameter_data.partrans = 'log'
pst.parameter_data.partrans.unique()
pst.write(f'../run_data/{pst_root}_sens.pst')
```
## If `run_ensemble=True` the cell below will run a local `prior_mc_wide_sens` global sensitivity analysis
* **NOTE: must have the `pestpp-sen` executable in system path or in `../run_data/`**
* same process as in notebooks 1.0 and 1.3 for parallel run
* for this `pestpp-sen`run, the total number of model runs is 64, which is = # of parameter groups * 4
* will run in parallel locally using the number of cores specified below by `num_workers`
* creates a new directory called `"../master_sen/"` which is a copy of run_data
* while running generates worker directories that are removed when run is complete
* results moved to `"../run_data/"`
```
if run_ensemble==True:
# set some variables for starting a group of PEST++ workers on the local machine
# MAKE SURE THAT PESTPP-IES and MF6 executables are in your system path or are in '../run_data'
num_workers = 5 # number of local workers -- VERY IMPORTANT, DO NOT MAKE TOO BIG
if sys.platform == 'win32':
pst_exe = 'pestpp-sen.exe'
else:
pst_exe = 'pestpp-sen'
template_ws = '../run_data' # template_directory
m_d = '../master_sen'
pyemu.os_utils.start_workers(worker_dir=template_ws,
exe_rel_path=pst_exe,
pst_rel_path=f'{pst_root}_sens.pst',
num_workers=num_workers,
master_dir=m_d
)
if run_ensemble==True:
# move results into run_data and clean up
move_result_files = glob.glob(os.path.join(m_d, f'{pst_root}_sens*'))
move_result_files = [f for f in move_result_files if 'pst' not in f]
[shutil.copy(os.path.join(m_d, file), output_dir) for file in move_result_files]
# Remove master dir.
shutil.rmtree(m_d)
```
| github_jupyter |
# Consulting Project
## Recommender Systems
Your final result should be in the form of a function that can take in a Spark DataFrame of a single customer's ratings for various meals and output their top 3 suggested meals. For example:
```
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
plt.style.use('seaborn')
%matplotlib inline
df = pd.read_csv('movielens_ratings.csv')
df.describe().transpose()
df.corr()
sns.heatmap(df.corr(), cmap='coolwarm')
import numpy as np
df['mealskew'] = df['movieId'].apply(lambda id: np.nan if id > 31 else id)
df.describe().transpose()
mealmap = { 2. : "Chicken Curry",
3. : "Spicy Chicken Nuggest",
5. : "Hamburger",
9. : "Taco Surprise",
11. : "Meatloaf",
12. : "Ceaser Salad",
15. : "BBQ Ribs",
17. : "Sushi Plate",
19. : "Cheesesteak Sandwhich",
21. : "Lasagna",
23. : "Orange Chicken",
26. : "Spicy Beef Plate",
27. : "Salmon with Mashed Potatoes",
28. : "Penne Tomatoe Pasta",
29. : "Pork Sliders",
30. : "Vietnamese Sandwich",
31. : "Chicken Wrap",
np.nan: "Cowboy Burger",
4. : "Pretzels and Cheese Plate",
6. : "Spicy Pork Sliders",
13. : "Mandarin Chicken PLate",
14. : "Kung Pao Chicken",
16. : "Fried Rice Plate",
8. : "Chicken Chow Mein",
10. : "Roasted Eggplant ",
18. : "Pepperoni Pizza",
22. : "Pulled Pork Plate",
0. : "Cheese Pizza",
1. : "Burrito",
7. : "Nachos",
24. : "Chili",
20. : "Southwest Salad",
25.: "Roast Beef Sandwich"}
df['meal_name'] = df['mealskew'].map(mealmap)
df.head()
df.to_csv('Meal_Info.csv',index=False)
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('recconsulting').getOrCreate()
from pyspark.ml.evaluation import RegressionEvaluator
from pyspark.ml.recommendation import ALS
data = spark.read.csv('Meal_Info.csv',inferSchema=True,header=True)
data.printSchema()
data.describe().show()
data.show()
data = data.na.drop(how='any')
(training, test) = data.randomSplit([0.8, 0.2])
# Build the recommendation model using ALS on the training data
als = ALS(maxIter=5, regParam=0.01, userCol="userId", itemCol="mealskew", ratingCol="rating")
model = als.fit(training)
# Evaluate the model by computing the RMSE on the test data
predictions = model.transform(test)
predictions.show()
evaluator = RegressionEvaluator(metricName="rmse", labelCol="rating",predictionCol="prediction")
rmse = evaluator.evaluate(predictions)
print("Root-mean-square error = " + str(rmse))
# Single_user
single_user = test.filter(test['userId'] ==19).select(['movieId', 'userId', 'mealskew'])
single_user.show()
recommendations = model.transform(single_user)
recommendations.orderBy('prediction', ascending=False).show()
```
| github_jupyter |
```
data_paths = {
'Micro': "../data/runs/2018-11-10-micro-rad-khype1e6/",
'NNOrig': "../data/runs/2018-11-09-model188-equilibriation-penalty",
'NN': "../data/runs/2018-11-10-model188-khyp1e6-rerun/"
}
ng_path = "../data/processed/training.nc"
from toolz import valmap
from src.data.sam import SAMRun
from uwnet.thermo import compute_q2
import xarray as xr
import holoviews as hv
hv.extension('bokeh')
%opts Image[width=600, height=400, colorbar=True](cmap='viridis')
%opts Curve[width=400]
def plot_xy(da, dynamic=True):
return hv.Dataset(da).to.image(["x", "y"], dynamic=dynamic)
data_paths = {
'Micro': "../data/runs/2018-11-10-micro-rad-khype1e6/",
'NNOrig': "../data/runs/2018-11-09-model188-equilibriation-penalty",
'NN': "../data/runs/2018-11-10-model188-khyp1e6-rerun/"
}
ng_path = "../data/processed/training.nc"
ngaqua = xr.open_dataset(ng_path).isel(step=0)
runs = valmap(SAMRun, data_paths)
ngaqua = xr.open_dataset(ng_path).isel(step=0)
```
# Net Precip
## Spinup
```
def get_first_steps(ds):
first_step = ds.isel(time=slice(1,4))
first_step['time'] = (first_step.time-first_step.time[0])*86400
first_step.time.attrs['units'] = 's'
return first_step
ds = runs['NN'].data_3d
first_step = get_first_steps(ds)
first_step.FQTNN.mean(['x']).plot(col='time')
```
Here is a comparision of P-E over these first few time steps:
```
net_precip_nn = -(ngaqua.layer_mass * ds.FQTNN).sum('z')/1000
net_precip_nn['time'] = (net_precip_nn.time -net_precip_nn.time[0])*24
net_precip_nn[[1, 5, 6, 7]].mean('x').plot(hue='time')
plt.title("Zonal mean P-E (time in hours)");
```
## Comparison to NG-Aqua
Let's open the NGAqua data and plot Q2
```
from uwnet.thermo import compute_q2
q2 = compute_q2(ngaqua.isel(time=slice(0, 2))).dropna('time')
```
Here is thea actual mean Q2 in the first tiem step
```
q2.mean('x').plot()
```
Here is a comparision of the zonally averaged net precipitation.
```
q2_int = -(ngaqua.layer_mass * q2).sum('z')/1000
fqtnn_int = -(ngaqua.layer_mass * first_step.FQTNN).sum('z')/1000
q2_int.mean(['x']).plot(label='NGaqua')
fqtnn_int[0].mean('x').plot(label='NN-prediction')
plt.title("Zonal mean of Net Precip")
plt.legend()
```
There is some systematic difference, but overally it is too noisy to tell from the first time step
# Spin up of vertical velocity
```
plot_data = runs['NN'].data_3d.W[4::3, 8]
dmap = plot_xy(plot_data, dynamic=False)\
.relabel(f"W at z={float(plot_data.z)}")
dmap
```
| github_jupyter |
# Deep Neural Network for Image Classification: Application
When you finish this, you will have finished the last programming assignment of Week 4, and also the last programming assignment of this course!
You will use the functions you'd implemented in the previous assignment to build a deep network, and apply it to cat vs non-cat classification. Hopefully, you will see an improvement in accuracy relative to your previous logistic regression implementation.
**After this assignment you will be able to:**
- Build and apply a deep neural network to supervised learning.
Let's get started!
## 1 - Packages
Let's first import all the packages that you will need during this assignment.
- [numpy](https://www.numpy.org/) is the fundamental package for scientific computing with Python.
- [matplotlib](http://matplotlib.org) is a library to plot graphs in Python.
- [h5py](http://www.h5py.org) is a common package to interact with a dataset that is stored on an H5 file.
- [PIL](http://www.pythonware.com/products/pil/) and [scipy](https://www.scipy.org/) are used here to test your model with your own picture at the end.
- dnn_app_utils provides the functions implemented in the "Building your Deep Neural Network: Step by Step" assignment to this notebook.
- np.random.seed(1) is used to keep all the random function calls consistent. It will help us grade your work.
```
import time
import numpy as np
import h5py
import matplotlib.pyplot as plt
import scipy
from PIL import Image
from scipy import ndimage
from dnn_app_utils_v3 import *
%matplotlib inline
plt.rcParams['figure.figsize'] = (5.0, 4.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
%load_ext autoreload
%autoreload 2
np.random.seed(1)
```
## 2 - Dataset
You will use the same "Cat vs non-Cat" dataset as in "Logistic Regression as a Neural Network" (Assignment 2). The model you had built had 70% test accuracy on classifying cats vs non-cats images. Hopefully, your new model will perform a better!
**Problem Statement**: You are given a dataset ("data.h5") containing:
- a training set of m_train images labelled as cat (1) or non-cat (0)
- a test set of m_test images labelled as cat and non-cat
- each image is of shape (num_px, num_px, 3) where 3 is for the 3 channels (RGB).
Let's get more familiar with the dataset. Load the data by running the cell below.
```
train_x_orig, train_y, test_x_orig, test_y, classes = load_data()
```
The following code will show you an image in the dataset. Feel free to change the index and re-run the cell multiple times to see other images.
```
# Example of a picture
index = 25
plt.imshow(train_x_orig[index])
print ("y = " + str(train_y[0,index]) + ". It's a " + classes[train_y[0,index]].decode("utf-8") + " picture.")
# Explore your dataset
m_train = train_x_orig.shape[0]
num_px = train_x_orig.shape[1]
m_test = test_x_orig.shape[0]
print ("Number of training examples: " + str(m_train))
print ("Number of testing examples: " + str(m_test))
print ("Each image is of size: (" + str(num_px) + ", " + str(num_px) + ", 3)")
print ("train_x_orig shape: " + str(train_x_orig.shape))
print ("train_y shape: " + str(train_y.shape))
print ("test_x_orig shape: " + str(test_x_orig.shape))
print ("test_y shape: " + str(test_y.shape))
```
As usual, you reshape and standardize the images before feeding them to the network. The code is given in the cell below.
<img src="images/imvectorkiank.png" style="width:450px;height:300px;">
<caption><center> <u>Figure 1</u>: Image to vector conversion. <br> </center></caption>
```
# Reshape the training and test examples
train_x_flatten = train_x_orig.reshape(train_x_orig.shape[0], -1).T # The "-1" makes reshape flatten the remaining dimensions
test_x_flatten = test_x_orig.reshape(test_x_orig.shape[0], -1).T
# Standardize data to have feature values between 0 and 1.
train_x = train_x_flatten/255
test_x = test_x_flatten/255
print ("train_x's shape: " + str(train_x.shape))
print ("test_x's shape: " + str(test_x.shape))
```
$12,288$ equals $64 \times 64 \times 3$ which is the size of one reshaped image vector.
## 3 - Architecture of your model
Now that you are familiar with the dataset, it is time to build a deep neural network to distinguish cat images from non-cat images.
You will build two different models:
- A 2-layer neural network
- An L-layer deep neural network
You will then compare the performance of these models, and also try out different values for $L$.
Let's look at the two architectures.
### 3.1 - 2-layer neural network
<img src="images/2layerNN_kiank.png" style="width:650px;height:400px;">
<caption><center> <u>Figure 2</u>: 2-layer neural network. <br> The model can be summarized as: ***INPUT -> LINEAR -> RELU -> LINEAR -> SIGMOID -> OUTPUT***. </center></caption>
<u>Detailed Architecture of figure 2</u>:
- The input is a (64,64,3) image which is flattened to a vector of size $(12288,1)$.
- The corresponding vector: $[x_0,x_1,...,x_{12287}]^T$ is then multiplied by the weight matrix $W^{[1]}$ of size $(n^{[1]}, 12288)$.
- You then add a bias term and take its relu to get the following vector: $[a_0^{[1]}, a_1^{[1]},..., a_{n^{[1]}-1}^{[1]}]^T$.
- You then repeat the same process.
- You multiply the resulting vector by $W^{[2]}$ and add your intercept (bias).
- Finally, you take the sigmoid of the result. If it is greater than 0.5, you classify it to be a cat.
### 3.2 - L-layer deep neural network
It is hard to represent an L-layer deep neural network with the above representation. However, here is a simplified network representation:
<img src="images/LlayerNN_kiank.png" style="width:650px;height:400px;">
<caption><center> <u>Figure 3</u>: L-layer neural network. <br> The model can be summarized as: ***[LINEAR -> RELU] $\times$ (L-1) -> LINEAR -> SIGMOID***</center></caption>
<u>Detailed Architecture of figure 3</u>:
- The input is a (64,64,3) image which is flattened to a vector of size (12288,1).
- The corresponding vector: $[x_0,x_1,...,x_{12287}]^T$ is then multiplied by the weight matrix $W^{[1]}$ and then you add the intercept $b^{[1]}$. The result is called the linear unit.
- Next, you take the relu of the linear unit. This process could be repeated several times for each $(W^{[l]}, b^{[l]})$ depending on the model architecture.
- Finally, you take the sigmoid of the final linear unit. If it is greater than 0.5, you classify it to be a cat.
### 3.3 - General methodology
As usual you will follow the Deep Learning methodology to build the model:
1. Initialize parameters / Define hyperparameters
2. Loop for num_iterations:
a. Forward propagation
b. Compute cost function
c. Backward propagation
d. Update parameters (using parameters, and grads from backprop)
4. Use trained parameters to predict labels
Let's now implement those two models!
## 4 - Two-layer neural network
**Question**: Use the helper functions you have implemented in the previous assignment to build a 2-layer neural network with the following structure: *LINEAR -> RELU -> LINEAR -> SIGMOID*. The functions you may need and their inputs are:
```python
def initialize_parameters(n_x, n_h, n_y):
...
return parameters
def linear_activation_forward(A_prev, W, b, activation):
...
return A, cache
def compute_cost(AL, Y):
...
return cost
def linear_activation_backward(dA, cache, activation):
...
return dA_prev, dW, db
def update_parameters(parameters, grads, learning_rate):
...
return parameters
```
```
### CONSTANTS DEFINING THE MODEL ####
n_x = 12288 # num_px * num_px * 3
n_h = 7
n_y = 1
layers_dims = (n_x, n_h, n_y)
# GRADED FUNCTION: two_layer_model
def two_layer_model(X, Y, layers_dims, learning_rate = 0.0075, num_iterations = 3000, print_cost=False):
"""
Implements a two-layer neural network: LINEAR->RELU->LINEAR->SIGMOID.
Arguments:
X -- input data, of shape (n_x, number of examples)
Y -- true "label" vector (containing 1 if cat, 0 if non-cat), of shape (1, number of examples)
layers_dims -- dimensions of the layers (n_x, n_h, n_y)
num_iterations -- number of iterations of the optimization loop
learning_rate -- learning rate of the gradient descent update rule
print_cost -- If set to True, this will print the cost every 100 iterations
Returns:
parameters -- a dictionary containing W1, W2, b1, and b2
"""
np.random.seed(1)
grads = {}
costs = [] # to keep track of the cost
m = X.shape[1] # number of examples
(n_x, n_h, n_y) = layers_dims
# Initialize parameters dictionary, by calling one of the functions you'd previously implemented
### START CODE HERE ### (≈ 1 line of code)
parameters = initialize_parameters(n_x, n_h, n_y)
### END CODE HERE ###
# Get W1, b1, W2 and b2 from the dictionary parameters.
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
# Loop (gradient descent)
for i in range(0, num_iterations):
# Forward propagation: LINEAR -> RELU -> LINEAR -> SIGMOID. Inputs: "X, W1, b1, W2, b2". Output: "A1, cache1, A2, cache2".
### START CODE HERE ### (≈ 2 lines of code)
A1, cache1 = linear_activation_forward(X, W1, b1, activation="relu")
A2, cache2 = linear_activation_forward(A1, W2, b2, activation="sigmoid")
### END CODE HERE ###
# Compute cost
### START CODE HERE ### (≈ 1 line of code)
cost = compute_cost(A2, Y)
### END CODE HERE ###
# Initializing backward propagation
dA2 = - (np.divide(Y, A2) - np.divide(1 - Y, 1 - A2))
# Backward propagation. Inputs: "dA2, cache2, cache1". Outputs: "dA1, dW2, db2; also dA0 (not used), dW1, db1".
### START CODE HERE ### (≈ 2 lines of code)
dA1, dW2, db2 = linear_activation_backward(dA2, cache2, activation="sigmoid")
dA0, dW1, db1 = linear_activation_backward(dA1, cache1, activation="relu")
### END CODE HERE ###
# Set grads['dWl'] to dW1, grads['db1'] to db1, grads['dW2'] to dW2, grads['db2'] to db2
grads['dW1'] = dW1
grads['db1'] = db1
grads['dW2'] = dW2
grads['db2'] = db2
# Update parameters.
### START CODE HERE ### (approx. 1 line of code)
parameters = update_parameters(parameters, grads, learning_rate)
### END CODE HERE ###
# Retrieve W1, b1, W2, b2 from parameters
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
# Print the cost every 100 training example
if print_cost and i % 100 == 0:
print("Cost after iteration {}: {}".format(i, np.squeeze(cost)))
if print_cost and i % 100 == 0:
costs.append(cost)
# plot the cost
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per hundreds)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
return parameters
```
Run the cell below to train your parameters. See if your model runs. The cost should be decreasing. It may take up to 5 minutes to run 2500 iterations. Check if the "Cost after iteration 0" matches the expected output below, if not click on the square (⬛) on the upper bar of the notebook to stop the cell and try to find your error.
```
parameters = two_layer_model(train_x, train_y, layers_dims = (n_x, n_h, n_y), num_iterations = 2500, print_cost=True)
```
**Expected Output**:
<table>
<tr>
<td> **Cost after iteration 0**</td>
<td> 0.6930497356599888 </td>
</tr>
<tr>
<td> **Cost after iteration 100**</td>
<td> 0.6464320953428849 </td>
</tr>
<tr>
<td> **...**</td>
<td> ... </td>
</tr>
<tr>
<td> **Cost after iteration 2400**</td>
<td> 0.048554785628770226 </td>
</tr>
</table>
Good thing you built a vectorized implementation! Otherwise it might have taken 10 times longer to train this.
Now, you can use the trained parameters to classify images from the dataset. To see your predictions on the training and test sets, run the cell below.
```
predictions_train = predict(train_x, train_y, parameters)
```
**Expected Output**:
<table>
<tr>
<td> **Accuracy**</td>
<td> 1.0 </td>
</tr>
</table>
```
predictions_test = predict(test_x, test_y, parameters)
```
**Expected Output**:
<table>
<tr>
<td> **Accuracy**</td>
<td> 0.72 </td>
</tr>
</table>
**Note**: You may notice that running the model on fewer iterations (say 1500) gives better accuracy on the test set. This is called "early stopping" and we will talk about it in the next course. Early stopping is a way to prevent overfitting.
Congratulations! It seems that your 2-layer neural network has better performance (72%) than the logistic regression implementation (70%, assignment week 2). Let's see if you can do even better with an $L$-layer model.
## 5 - L-layer Neural Network
**Question**: Use the helper functions you have implemented previously to build an $L$-layer neural network with the following structure: *[LINEAR -> RELU]$\times$(L-1) -> LINEAR -> SIGMOID*. The functions you may need and their inputs are:
```python
def initialize_parameters_deep(layers_dims):
...
return parameters
def L_model_forward(X, parameters):
...
return AL, caches
def compute_cost(AL, Y):
...
return cost
def L_model_backward(AL, Y, caches):
...
return grads
def update_parameters(parameters, grads, learning_rate):
...
return parameters
```
```
### CONSTANTS ###
layers_dims = [12288, 20, 7, 5, 1] # 4-layer model
# GRADED FUNCTION: L_layer_model
def L_layer_model(X, Y, layers_dims, learning_rate = 0.0075, num_iterations = 3000, print_cost=False):#lr was 0.009
"""
Implements a L-layer neural network: [LINEAR->RELU]*(L-1)->LINEAR->SIGMOID.
Arguments:
X -- data, numpy array of shape (num_px * num_px * 3, number of examples)
Y -- true "label" vector (containing 0 if cat, 1 if non-cat), of shape (1, number of examples)
layers_dims -- list containing the input size and each layer size, of length (number of layers + 1).
learning_rate -- learning rate of the gradient descent update rule
num_iterations -- number of iterations of the optimization loop
print_cost -- if True, it prints the cost every 100 steps
Returns:
parameters -- parameters learnt by the model. They can then be used to predict.
"""
np.random.seed(1)
costs = [] # keep track of cost
# Parameters initialization. (≈ 1 line of code)
### START CODE HERE ###
parameters = initialize_parameters_deep(layers_dims)
### END CODE HERE ###
# Loop (gradient descent)
for i in range(0, num_iterations):
# Forward propagation: [LINEAR -> RELU]*(L-1) -> LINEAR -> SIGMOID.
### START CODE HERE ### (≈ 1 line of code)
AL, caches = L_model_forward(X, parameters)
### END CODE HERE ###
# Compute cost.
### START CODE HERE ### (≈ 1 line of code)
cost = compute_cost(AL, Y)
### END CODE HERE ###
# Backward propagation.
### START CODE HERE ### (≈ 1 line of code)
grads = L_model_backward(AL, Y, caches)
### END CODE HERE ###
# Update parameters.
### START CODE HERE ### (≈ 1 line of code)
parameters = update_parameters(parameters, grads, learning_rate)
### END CODE HERE ###
# Print the cost every 100 training example
if print_cost and i % 100 == 0:
print ("Cost after iteration %i: %f" %(i, cost))
if print_cost and i % 100 == 0:
costs.append(cost)
# plot the cost
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per hundreds)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
return parameters
```
You will now train the model as a 4-layer neural network.
Run the cell below to train your model. The cost should decrease on every iteration. It may take up to 5 minutes to run 2500 iterations. Check if the "Cost after iteration 0" matches the expected output below, if not click on the square (⬛) on the upper bar of the notebook to stop the cell and try to find your error.
```
parameters = L_layer_model(train_x, train_y, layers_dims, num_iterations = 2500, print_cost = True)
```
**Expected Output**:
<table>
<tr>
<td> **Cost after iteration 0**</td>
<td> 0.771749 </td>
</tr>
<tr>
<td> **Cost after iteration 100**</td>
<td> 0.672053 </td>
</tr>
<tr>
<td> **...**</td>
<td> ... </td>
</tr>
<tr>
<td> **Cost after iteration 2400**</td>
<td> 0.092878 </td>
</tr>
</table>
```
pred_train = predict(train_x, train_y, parameters)
```
<table>
<tr>
<td>
**Train Accuracy**
</td>
<td>
0.985645933014
</td>
</tr>
</table>
```
pred_test = predict(test_x, test_y, parameters)
```
**Expected Output**:
<table>
<tr>
<td> **Test Accuracy**</td>
<td> 0.8 </td>
</tr>
</table>
Congrats! It seems that your 4-layer neural network has better performance (80%) than your 2-layer neural network (72%) on the same test set.
This is good performance for this task. Nice job!
Though in the next course on "Improving deep neural networks" you will learn how to obtain even higher accuracy by systematically searching for better hyperparameters (learning_rate, layers_dims, num_iterations, and others you'll also learn in the next course).
## 6) Results Analysis
First, let's take a look at some images the L-layer model labeled incorrectly. This will show a few mislabeled images.
```
print_mislabeled_images(classes, test_x, test_y, pred_test)
```
**A few types of images the model tends to do poorly on include:**
- Cat body in an unusual position
- Cat appears against a background of a similar color
- Unusual cat color and species
- Camera Angle
- Brightness of the picture
- Scale variation (cat is very large or small in image)
## 7) Test with your own image (optional/ungraded exercise) ##
Congratulations on finishing this assignment. You can use your own image and see the output of your model. To do that:
1. Click on "File" in the upper bar of this notebook, then click "Open" to go on your Coursera Hub.
2. Add your image to this Jupyter Notebook's directory, in the "images" folder
3. Change your image's name in the following code
4. Run the code and check if the algorithm is right (1 = cat, 0 = non-cat)!
```
from PIL import Image
fileImage = Image.open("test.png").convert("RGB").resize([num_px,num_px],Image.ANTIALIAS)
my_label_y = [1] # the true class of your image (1 -> cat, 0 -> non-cat)
image = np.array(fileImage)
my_image = image.reshape(num_px*num_px*3,1)
my_image = my_image/255.
my_predicted_image = predict(my_image, my_label_y, parameters)
plt.imshow(image)
print ("y = " + str(np.squeeze(my_predicted_image)) + ", your L-layer model predicts a \"" + classes[int(np.squeeze(my_predicted_image)),].decode("utf-8") + "\" picture.")
```
**References**:
- for auto-reloading external module: http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
```
import pickle # For storing an array in a '.pickle' file for later use
# If you want to save multiple variables, put them into an array
my_content = [train_x_orig, train_y, test_x_orig, test_y, classes]
# Saving variables in an array
with open("trainingDataset.pickle",'wb') as fileToBeWritten:
# For compatibility we use open(filename, 'wb') for non-text files and open(filename, 'w') for text files
pickle.dump(my_content,fileToBeWritten)
# Loading Variables
with open('trainingDatasetL-LayerNN.pickle','rb') as fileToBeRead:
ttrain_x_orig, train_y, test_x_orig, test_y, classes = pickle.load(fileToBeRead)
```
| github_jupyter |
<h2 align="center">Predicting Titanic Passenger Survival
<h2 align="center">with Supervised Machine Learning, by Logistic Regression (Classification)
</h2>
__1) The Goal:__ Build a machine learning model that is able to predict which of the passengers of the Titanic has survived or not.
__2) Get the Data:__ The data is given.
__3) Split the Data:__ The data is already split: "train.csv" and "test.csv".
__4) Exploratory Data Analysis (EDA)__
__5)-9) Feature Engineering (FE), Train Model, Optimize Hyperparameters/Cross-Validation__
- Train a Logistic Regression classification model (alternative models: Decision Trees, Random Forest)
- Create features using one-hot encoding
- Calculate the train and test accuracy
- Calculate a cross-validation score
- Train a Random Forest classification model
__10) Calculate Test Score:__ Use the test data set aside to calculate the test score.
__11) Deploy and Monitor:__ Submit your prediction to Kaggle.
<h2> <center>Exploratory Data Analysis</h2> </center>
```
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
from sklearn.preprocessing import KBinsDiscretizer
from sklearn.preprocessing import OneHotEncoder
from sklearn.model_selection import train_test_split
from sklearn.dummy import DummyClassifier
from sklearn.linear_model import LogisticRegression
import statsmodels.discrete.discrete_model as sm
```
#### Step 1: Read the file train.csv into Python and print a few rows.
```
train = pd.read_csv('./raw data/train.csv')
train.head()
```
#### Step 2: Calculate the number of surviving/non-surviving passengers and display it as a bar plot.
```
i=sum(train['Survived'])
labels_step2 = ['surviving', 'non-surviving']
x = np.arange(len(labels_step2))
y = [sum(train['Survived']), (train['Survived'].count()-sum(train['Survived']))]
plt.figure(figsize=(10,4))
plt.bar(x, y)
plt.xticks(x, labels_step2, size=16)
plt.title(f"Only {i} passangers survived.\n\
(i.e. {round(100*train['Survived'].sum()/len(train))}% of the passangers in the train data)\n\
{round(100*(len(train)-train['Survived'].sum())/len(train))}% of the passangers died.", fontsize=18)
plt.ylabel('Number of passangers', fontsize=15)
plt.show()
```
#### Step 3: Calculate the proportion of surviving 1st class passengers with regards to the total number of 1st class passengers.
```
print(f"{train[(train['Pclass']==1) & (train['Survived']==1)]['Survived'].count()} first class passengers survived,\n\
i.e. {100*round(train[(train['Pclass']==1) & (train['Survived']==1)]['Survived'].count()/sum(train['Pclass']==1),2)}% of the 1st class passengers\n\
and {100*round(train[(train['Pclass']==1) & (train['Survived']==1)]['Survived'].count()/len(train),2)}% of all passengers.")
```
#### Step 4: Create a bar plot with separate bars for male/female passengers and 1st/2nd/3rd class passengers.
```
ax = sns.countplot(x="Pclass", hue="Sex", data=train)
```
#### Step 5: Create a histogram showing the age distribution of passengers. Compare surviving/non-surviving passengers.
```
ax = sns.histplot(x="Age", hue="Survived", data=train, multiple="dodge", shrink = 0.7)
```
#### Step 6: Calculate the average age for survived and drowned passengers separately.
```
print('The average age of the survived passengers is {}, and the average age of the drowned passengers is {}.'.format(round(train.groupby('Survived')['Age'].mean().tolist()[0]),round(train.groupby('Survived')['Age'].mean().tolist()[1])))
```
#### Step 7: Replace missing age values by the mean age.
```
train['Age_mean_imputed'] = train['Age'].fillna(round(np.float64(train['Age'].mean()),1))
```
#### Step 8: Create a table counting the number of surviving/dead passengers separately for 1st/2nd/3rd class and male/female.
```
pd.crosstab(train['Pclass'], [train['Sex'], train['Survived']], rownames=['Pclass'], colnames=['Sex', 'Survived'])
```
<h2> <center>Feature Engineering</h2> </center>
```
train.fillna(0.0, inplace=True) # little crutch to make the data work
```
### Binning: a numerical column is converted to categories
```
# transform a numerical column
kbins = KBinsDiscretizer(n_bins=4, encode='onehot', strategy='uniform')
columns = train[['Age_mean_imputed']]
kbins.fit(columns)
t = kbins.transform(columns)
t = t.todense() # before this, t is a sparse matrix data type
# create nice labels
edges = kbins.bin_edges_[0].astype('int32')
labels = []
for i in range(len(edges)-1):
edge1 = edges[i]
edge2 = edges[i+1]
labels.append(f"{edge1} to {edge2}")
print(t.shape)
age_binned = pd.DataFrame(t, columns=labels)
plt.figure(figsize=(15,4))
ax = sns.barplot(data=age_binned)
# create a DataFrame
age_binned = pd.DataFrame(t, columns=labels)
plt.figure(figsize=(15,4))
ax = sns.barplot(data=age_binned)
train = pd.concat([train, age_binned], axis=1)
train.head(5)
```
<h2> <center>Train the Model</h2> </center>
```
X = train.drop(columns=['Survived', 'Name', 'Sex', 'Ticket', 'Cabin', 'Embarked'])
y = train['Survived']
X.head(2)
test = pd.read_csv('./raw data/test.csv')
test['Age_mean_imputed'] = train['Age'].fillna(round(np.float64(train['Age'].mean()),1))
test.fillna(0.0, inplace=True) # little crutch to make the data work
# transform a numerical column
kbins = KBinsDiscretizer(n_bins=4, encode='onehot', strategy='uniform')
columns = test[['Age_mean_imputed']]
kbins.fit(columns)
t = kbins.transform(columns)
t = t.todense() # before this, t is a sparse matrix data type
# create nice labels
edges = kbins.bin_edges_[0].astype('int32')
labels = []
for i in range(len(edges)-1):
edge1 = edges[i]
edge2 = edges[i+1]
labels.append(f"{edge1} to {edge2}")
age_binned = pd.DataFrame(t, columns=labels)
test = pd.concat([test, age_binned], axis=1)
X_new = test.drop(columns=['Name', 'Sex', 'Ticket', 'Cabin', 'Embarked'])
X_new.head(2)
m = LogisticRegression()
m.fit(X, y)
print(m.score(X,y), m.predict(X_new))
logit = sm.Logit(y, X)
f = logit.fit()
print(f.params)
print(f.summary())
```
<h2> <center>Optimize Hyperparameters/Cross-Validation</h2> </center>
Train Model, Optimize Hyperparameters/Cross-Validation
- Train a Logistic Regression classification model (alternative models: Decision Trees, Random Forest)
- Create features using one-hot encoding
- Calculate the train and test accuracy
- Calculate a cross-validation score
- Train a Random Forest classification model
- Submit the predictions to Kaggle
- Give a 5-minute lightning talk by the end of the week
#### 10) Calculate Test Score
- Use the test data set aside to calculate the test score.
#### 11) Deploy and Monitor
<h2> <center>Calculate Test Score</h2> </center>
<h2> <center>Evaluate the Model using the Test Data</h2> </center>
- accuracy
- confussion matrix: precission, recall, F1-score
<h2> <center>Make Predictions</h2> </center>
| github_jupyter |
# Breadth First Search
```
graph={'a':['b','c'],'b':['a','d'],'c':['a','e'],'d':['b','d','i'],
'e':['c','d','f','j'],'f':['e','g','i','j'],'g':['f','h'],'h':['g','i'],
'i':['d','f','h'],'j':['e','f']}
graph_quality={'a':None,'b':None,'c':None,'d':None,'e':None,'f':None,'g':'Mango','h':None,'i':None,'j':None}
start = 'a'
desired_quality='Mango'
# we prefer all connections that are closer so fewest steps taken
def check_node(node,desired_quality,graph_quality=None):
if graph_quality==None:
return False
return desired_quality == graph_quality[node]
def breadth_first(start,graph,desired_quality,graph_quality=None):
queue=[start]
found_result = False
queue_int=0
parent={start:[[]]}
checked=set()
while found_result==False:
node=queue[queue_int]
if node not in checked:
if check_node(node,desired_quality,graph_quality) or desired_quality==node:
found_result=True
break
else:
queue+=graph[node]
for sub_node in graph[node]:
if sub_node not in parent.keys():
parent[sub_node]=[[node]+x for x in parent[node]]
else:
parent[sub_node]+=[[node]+x for x in parent[node]]
checked.add(node)
queue_int+=1
else:
queue_int+=1
if len(checked)==len(graph.keys()):
print('No: {0} in graph'.format(desired_quality))
return None
print('{0} found'.format(desired_quality))
node_routes = {len(route): route for route in parent[node]}
shortest_route = node_routes[min(node_routes.keys())][::-1] + [node]
print('shortest route: {0}'.format('-'.join(shortest_route)))
return shortest_route
parent=breadth_first(start,graph,'d')
```
# Dijkstra
```
def lowest_cost(costs,processed):
lowest=float("inf")
current_node = None
for node in costs.keys():
cost=costs[node]
if cost < lowest and node not in processed:
current_node = node
lowest = cost
return current_node
lowest_cost({'b':2,'c':3,'d':5,'e':7,'a':8},[])
def dijkstra(start,end,graph):
"""Implementation of dijkstra algorithm
Parameters
-----------
start : string
start point for the graph
end : string
end point for the graph
graph : dictionary
Dictionary should contain the
graph, with each dictonary element
containing the weighted elements
example:
{'a':{'b':2,'c':3},'b':{'a':2,'d':4},'c':{'a':3,'d':3},'d':{'b':4,'c':3}}
Returns
----------
cost : int/float
cost of getting to end node
route : list of strings
the nodes to get to end node
"""
# establish initial parents/costs
parents={x:start for x in graph[start].keys()}
costs=graph[start]
processed=[]
# establish cost of all nodes as infinite to start
for node in graph.keys():
if node not in costs.keys():
costs[node]=float('inf')
current_node=lowest_cost(costs, processed)
# Main controlflow loop
while current_node is not None:
cost=costs[current_node]
neighbours=graph[current_node]
# loop through each node's neighbours
for n in neighbours.keys():
new_cost=cost+neighbours[n]
if new_cost < costs[n]:
costs[n]=new_cost
parents[n]=current_node
# add current node to processed, so not re-run
processed.append(current_node)
current_node=lowest_cost(costs,processed)
# Calculate the route that got from start to end
last=end
route=[end]
while last!=start:
last=parents[last]
route+=[last]
# reverse the route so goes from start - end
route=route[::-1]
print('Cost to get from {0} to {1} is: {2}.'.format(start,end,costs[end]))
print('Nodes to result: {0}'.format(route))
return costs[end], route
dijkstra('a','d',{'a':{'b':2,'c':3},'b':{'a':2,'d':4},'c':{'a':3,'d':3},'d':{'b':4,'c':3}})
```
| github_jupyter |
```
!pip install -U -q PyDrive
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
# 1. Authenticate and create the PyDrive client.
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
file_id = '1Nl6SYOBwJT9F1GmtI5U9BSacr6EdsrVc'
downloaded = drive.CreateFile({'id': file_id})
downloaded.GetContentFile('ThoraricSurgery.csv')
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
import numpy as np
import tensorflow as tf
np.random.seed(3)
tf.random.set_seed(3)
Data_set = np.loadtxt('ThoraricSurgery.csv', delimiter=",")
X = Data_set[:, 0:17]
Y = Data_set[:, 17]
model = Sequential()
model.add(Dense(30, input_dim=17, activation='relu'))
model.add(Dense(1, activation='relu'))
model.compile(loss='mean_squared_error', optimizer='adam', metrics=['accuracy'])
model.fit(X, Y, epochs=100, batch_size=10)
file_id = '1Oq9LWJuz0ZdL9yiUVDamNC6XWDS_6-6_'
downloaded = drive.CreateFile({'id': file_id})
downloaded.GetContentFile('pima-indians-diabetes.csv')
import pandas as pd
df = pd.read_csv('pima-indians-diabetes.csv', names=['pregnant', 'plasma', 'pressure', 'thickness', 'insulin', 'BMI', 'pedigree', 'age', 'class'])
print(df.head())
df.info()
df.describe()
df[['pregnant', 'class']]
print(df[['pregnant', 'class']].groupby(['pregnant'], as_index=False).mean().sort_values(by='pregnant', ascending=True))
import matplotlib.pyplot as plt
import seaborn as sns
plt.figure(figsize=(12, 12))
sns.heatmap(df.corr(), linewidth=0.1, vmax=0.5, cmap=plt.cm.gist_heat, linecolor='white', annot=True)
plt.show()
grid = sns.FacetGrid(df, col='class')
grid.map(plt.hist, 'plasma', bins=10)
plt.show()
np.random.seed(3)
tf.random.set_seed(3)
dataset = np.loadtxt('pima-indians-diabetes.csv', delimiter=',')
X = dataset[:, 0:8]
Y = dataset[:,8]
model = Sequential()
model.add(Dense(12, input_dim=8, activation='relu'))
model.add(Dense(8, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
model.fit(X, Y, epochs=200, batch_size=10)
print('\n Accuracy:%.4f' % (model.evaluate(X, Y)[1]))
!pip install -U -q PyDrive
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
# 1. Authenticate and create the PyDrive client.
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
file_id = '1sxjj3p8_9g41_uxd1O9KHYv191_E8h7e'
downloaded1 = drive.CreateFile({'id': file_id})
downloaded1.GetContentFile('iris.csv')
import pandas as pd
df = pd.read_csv('iris.csv', names=['sepal_length', 'sepal_width', 'petal_length', 'petal_width', 'species'])
print(df.head())
import seaborn
import matplotlib.pyplot as plt
sns.pairplot(df, hue='species')
plt.show()
dataset = df.values
X = dataset[:, 0:4].astype(float)
Y_obj = dataset[:, 4]
from sklearn.preprocessing import LabelEncoder
e = LabelEncoder()
e.fit(Y_obj)
Y = e.transform(Y_obj)
from keras.utils import np_utils
Y_encoded = tf.keras.utils.to_categorical(Y)
model = Sequential()
model.add(Dense(16, input_dim=4, activation='relu'))
model.add(Dense(3, activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
model.fit(X, Y_encoded, epochs=50, batch_size=1)
print('\n Accuracy:%.4f' % (model.evaluate(X, Y_encoded)[1]))
file_id = '1fB7IcP3uIG0L-hLzm65UyrPnvx4MnifP'
downloaded2 = drive.CreateFile({'id': file_id})
downloaded2.GetContentFile('sonar.csv')
import pandas as pd
df = pd.read_csv('sonar.csv', header=None)
print(df.info())
df.head()
from keras.models import Sequential
from keras.layers.core import Dense
from sklearn.preprocessing import LabelEncoder
import pandas as pd
import numpy as np
import tensorflow as tf
np.random.seed(3)
tf.random.set_seed(3)
df = pd.read_csv('sonar.csv', header=None)
dataset = df.values
X = dataset[:, 0:60]
Y_obj = dataset[:, 60]
e = LabelEncoder()
e.fit(Y_obj)
Y = e.transform(Y_obj)
model = Sequential()
model.add(Dense(24, input_dim=60, activation='relu'))
model.add(Dense(10, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='mean_squared_error', optimizer='adam', metrics=['accuracy'])
model.fit(X, Y, epochs=200, batch_size=5)
print('\n Accuracy:%.4f'%(model.evaluate(X, Y)[1]))
from keras.models import Sequential
from keras.layers.core import Dense
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split, StratifiedKFold
import pandas as pd
import numpy as np
import tensorflow as tf
seed = 0
np.random.seed(seed)
tf.random.set_seed(seed)
df = pd.read_csv('sonar.csv', header=None)
dataset = df.values
X = dataset[:, 0:60]
Y_obj = dataset[:, 60]
e = LabelEncoder()
e.fit(Y_obj)
Y = e.transform(Y_obj)
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.3, random_state=seed)
model = Sequential()
model.add(Dense(24, input_dim=60, activation='relu'))
model.add(Dense(10, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='mean_squared_error', optimizer='adam', metrics=['accuracy'])
model.fit(X_train, Y_train, epochs=130, batch_size=5)
print('\n Test Accuracy:%.4f'%(model.evaluate(X_test, Y_test)[1]))
from keras.models import Sequential
from keras.layers.core import Dense
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import StratifiedKFold
import pandas as pd
import numpy as np
import tensorflow as tf
seed = 0
np.random.seed(seed)
tf.random.set_seed(seed)
df = pd.read_csv('sonar.csv', header=None)
dataset = df.values
X = dataset[:, 0:60]
Y_obj = dataset[:, 60]
e = LabelEncoder()
e.fit(Y_obj)
Y = e.transform(Y_obj)
n_fold = 10
skf = StratifiedKFold(n_splits=n_fold, shuffle=True, random_state=seed)
accuracy = []
for train, test in skf.split(X, Y):
model = Sequential()
model.add(Dense(24, input_dim=60, activation='relu'))
model.add(Dense(10, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='mean_squared_error', optimizer='adam', metrics=['accuracy'])
model.fit(X[train], Y[train], epochs=100, batch_size=5)
k_accuracy = '%.4f'%(model.evaluate(X[test], Y[test])[1])
accuracy.append(k_accuracy)
print('\n %.f fold Accuracy:'%n_fold, accuracy)
file_id = '1lzDJerOjrYCntwKNTh4_7mCp5c5T0Y4D'
downloaded3 = drive.CreateFile({'id': file_id})
downloaded3.GetContentFile('wine.csv')
df_pre = pd.read_csv('wine.csv', header=None)
df = df_pre.sample(frac=1)
df.head(5)
df.info
from keras.models import Sequential
from keras.layers import Dense
from keras.callbacks import ModelCheckpoint, EarlyStopping
import pandas as pd
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
seed = 0
np.random.seed(seed)
tf.random.set_seed(3)
df_pre = pd.read_csv('wine.csv', header=None)
df = df_pre.sample(frac=1)
dataset = df.values
X = dataset[:, 0:12]
Y = dataset[:, 12]
model = Sequential()
model.add(Dense(30, input_dim=12, activation='relu'))
model.add(Dense(12, activation='relu'))
model.add(Dense(8, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
model.fit(X, Y, epochs=200, batch_size=200)
print('\n Accuracy:%.4f'%(model.evaluate(X, Y)[1]))
from keras.models import Sequential
from keras.layers import Dense
from keras.callbacks import ModelCheckpoint
import pandas as pd
import numpy
import os
import matplotlib.pyplot as plt
import tensorflow as tf
seed = 0
numpy.random.seed(seed)
tf.random.set_seed(seed)
df_pre = pd.read_csv('wine.csv', header=None)
df = df_pre.sample(frac=0.15)
dataset = df.values
X = dataset[:,0:12]
Y = dataset[:,12]
model = Sequential()
model.add(Dense(30, input_dim=12, activation='relu'))
model.add(Dense(12, activation='relu'))
model.add(Dense(8, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
MODEL_DIR = './model/'
if not os.path.exists(MODEL_DIR):
os.mkdir(MODEL_DIR)
modelpath="./model/{epoch:02d}-{val_loss:.4f}.hdf5"
checkpointer = ModelCheckpoint(filepath=modelpath, monitor='val_loss', verbose=1, save_best_only=True)
history = model.fit(X, Y, validation_split=0.33, epochs=1000, batch_size=500)
y_vloss=history.history['val_loss']
y_acc=history.history['acc']
x_len = numpy.arange(len(y_acc))
plt.plot(x_len, y_vloss, "o", c="red", markersize=3)
plt.plot(x_len, y_acc, "o", c="blue", markersize=3)
plt.show()
from keras.callbacks import EarlyStopping
early_stopping_callback = EarlyStopping(monitor='val_loss', patience=100)
model.fit(X, Y, validation_split=0.33, epochs=2000, batch_size=500, callbacks=[early_stopping_callback])
!pip install -U -q PyDrive
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
# 1. Authenticate and create the PyDrive client.
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
file_id = '1QZecsECqfJO4dXD0rTYAu6GB35aop1fn'
downloaded4 = drive.CreateFile({'id': file_id})
downloaded4.GetContentFile('housing.csv')
import pandas as pd
df = pd.read_csv('housing.csv', delim_whitespace=True, header=None)
df.info
df.head()
from keras.models import Sequential
from keras.layers import Dense
from sklearn.model_selection import train_test_split
import numpy as np
import pandas as pd
import tensorflow as tf
seed = 0
np.random.seed(seed)
tf.random.set_seed(3)
df = pd.read_csv('housing.csv', delim_whitespace=True, header=None)
dataset = df.values
X = dataset[:, 0:13]
Y = dataset[:, 13]
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.3, random_state=seed)
model = Sequential()
model.add(Dense(30, input_dim=13, activation='relu'))
model.add(Dense(6, activation='relu'))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
model.fit(X_train, Y_train, epochs=200, batch_size=10)
Y_prediction = model.predict(X_test).flatten()
for i in range(10):
label = Y_test[i]
prediction = Y_prediction[i]
print('real:{} predict:{}'.format(label, prediction))
```
| github_jupyter |
# Docstrings
A [docstring](https://www.python.org/dev/peps/pep-0257/#id4) is a string embedded in a piece of code for the purpose of documenting it.
Docstrings in Sabueso are written in [NumPy style](). Why? Well, Sabueso's main developers know there are other docstring styles, but the NumPy's one is clear, easy to be read, and widely extended.
```{admonition} Note
:class: note
When Sabueso's web documentation is compiled, [Sphinx](https://www.sphinx-doc.org/en/master/index.html) handles the docstrings with the assistance of [the Napoleon extension](https://www.sphinx-doc.org/en/master/usage/extensions/napoleon.html). Have a look to the [sphinx section in this guide]().
```
```{admonition} It is suggested...
:class: tip, dropdown
Although you can find some guidelines to write Sabueso's docstrings below, if you are not familiar with the docstrings and the NumPy style you may also check the following webs:
- [PEP 257 -- Docstring Conventions](https://www.python.org/dev/peps/pep-0257/)
- [numpydoc's documentation](https://numpydoc.readthedocs.io/en/latest/format.html)
- [Example in numpydoc's documentation](https://numpydoc.readthedocs.io/en/latest/example.html)
- [Example in Napoleon's documentation](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_numpy.html)
```
## Documenting a class
```python
class ClassName():
"""The summary line for a class docstring should fit on one line.
If the class has public attributes, they may be documented here
in an ``Attributes`` section and follow the same formatting as a
function's ``Args`` section. Alternatively, attributes may be documented
inline with the attribute's declaration (see __init__ method below).
Properties created with the ``@property`` decorator should be documented
in the property's getter method.
Attributes
----------
attr1 : str
Description of `attr1`.
attr2 : :obj:`int`, optional
Description of `attr2`.
"""
def __init__(a, b=14):
"""Example of docstring on the __init__ method.
The __init__ method may be documented in either the class level
docstring, or as a docstring on the __init__ method itself.
Either form is acceptable, but the two should not be mixed. Choose one
convention to document the __init__ method and be consistent with it.
Note
----
Do not include the `self` parameter in the ``Parameters`` section.
Parameters
----------
a : obj:`list` of :obj:`str`
Description of `A`. Multiple
lines are supported.
b : :obj:`int`, default=14
Description of `B`.
"""
```
```{admonition} See also
:class: attention
Before writting your own docstrings, have a look to [Sabueso's code](https://github.com/uibcdf/Sabueso/tree/main/sabueso) and the [API documentation](https://uibcdf.org/Sabueso/api.html). There you can find useful examples on how the docstrings are written and rendered, and what are the usual conventions adopted in Sabueso's docstrings.
```
## Documenting a method
Some advices need to be followed to write a methods documentation. But first let's see an example of the docstring to document a method in Sabueso's code:
```python
def method_name(a, b=True):
"""A short one line description without variable names or the method name
Paragraph with detailed explanation.
Parameters
----------
a : int
Description of parameter `a`.
b : bool, default=True
Description of input argument `b` (the default is True, which implies ...).
Returns
-------
int
Description of the output int value.
Examples
--------
First example comment.
>>> 2+2
4
Comment explaining the second example.
>>> import numpy as np
>>> np.add([[1, 2], [3, 4]],
... [[5, 6], [7, 8]])
array([[ 6, 8],
[10, 12]])
See Also
--------
:func:`sabueso.get`, :func:`sabueso.select`
Notes
-----
Section to include notes.
Todo
----
Section to include a todo message.
Warning
-------
Section to include a warning message.
"""
pass
```
### About working with magnitudes
...
```{admonition} See also
:class: attention
Before writting your own docstrings, have a look to [Sabueso's code](https://github.com/uibcdf/Sabueso/tree/main/sabueso) and the [API documentation](https://uibcdf.org/Sabueso/api.html). There you can find useful examples on how the docstrings are written and rendered, and what are the usual conventions adopted in Sabueso's docstrings.
```
| github_jupyter |
**This notebook is an exercise in the [Intermediate Machine Learning](https://www.kaggle.com/learn/intermediate-machine-learning) course. You can reference the tutorial at [this link](https://www.kaggle.com/alexisbcook/data-leakage).**
---
Most people find target leakage very tricky until they've thought about it for a long time.
So, before trying to think about leakage in the housing price example, we'll go through a few examples in other applications. Things will feel more familiar once you come back to a question about house prices.
# Setup
The questions below will give you feedback on your answers. Run the following cell to set up the feedback system.
```
# Set up code checking
from learntools.core import binder
binder.bind(globals())
from learntools.ml_intermediate.ex7 import *
print("Setup Complete")
```
# Step 1: The Data Science of Shoelaces
Nike has hired you as a data science consultant to help them save money on shoe materials. Your first assignment is to review a model one of their employees built to predict how many shoelaces they'll need each month. The features going into the machine learning model include:
- The current month (January, February, etc)
- Advertising expenditures in the previous month
- Various macroeconomic features (like the unemployment rate) as of the beginning of the current month
- The amount of leather they ended up using in the current month
The results show the model is almost perfectly accurate if you include the feature about how much leather they used. But it is only moderately accurate if you leave that feature out. You realize this is because the amount of leather they use is a perfect indicator of how many shoes they produce, which in turn tells you how many shoelaces they need.
Do you think the _leather used_ feature constitutes a source of data leakage? If your answer is "it depends," what does it depend on?
After you have thought about your answer, check it against the solution below.
```
# Check your answer (Run this code cell to receive credit!)
q_1.check()
```
# Step 2: Return of the Shoelaces
You have a new idea. You could use the amount of leather Nike ordered (rather than the amount they actually used) leading up to a given month as a predictor in your shoelace model.
Does this change your answer about whether there is a leakage problem? If you answer "it depends," what does it depend on?
```
# Check your answer (Run this code cell to receive credit!)
q_2.check()
```
# 3. Getting Rich With Cryptocurrencies?
You saved Nike so much money that they gave you a bonus. Congratulations.
Your friend, who is also a data scientist, says he has built a model that will let you turn your bonus into millions of dollars. Specifically, his model predicts the price of a new cryptocurrency (like Bitcoin, but a newer one) one day ahead of the moment of prediction. His plan is to purchase the cryptocurrency whenever the model says the price of the currency (in dollars) is about to go up.
The most important features in his model are:
- Current price of the currency
- Amount of the currency sold in the last 24 hours
- Change in the currency price in the last 24 hours
- Change in the currency price in the last 1 hour
- Number of new tweets in the last 24 hours that mention the currency
The value of the cryptocurrency in dollars has fluctuated up and down by over $\$$100 in the last year, and yet his model's average error is less than $\$$1. He says this is proof his model is accurate, and you should invest with him, buying the currency whenever the model says it is about to go up.
Is he right? If there is a problem with his model, what is it?
```
# Check your answer (Run this code cell to receive credit!)
q_3.check()
```
# Step 4: Preventing Infections
An agency that provides healthcare wants to predict which patients from a rare surgery are at risk of infection, so it can alert the nurses to be especially careful when following up with those patients.
You want to build a model. Each row in the modeling dataset will be a single patient who received the surgery, and the prediction target will be whether they got an infection.
Some surgeons may do the procedure in a manner that raises or lowers the risk of infection. But how can you best incorporate the surgeon information into the model?
You have a clever idea.
1. Take all surgeries by each surgeon and calculate the infection rate among those surgeons.
2. For each patient in the data, find out who the surgeon was and plug in that surgeon's average infection rate as a feature.
Does this pose any target leakage issues?
Does it pose any train-test contamination issues?
```
# Check your answer (Run this code cell to receive credit!)
q_4.check()
```
# Step 5: Housing Prices
You will build a model to predict housing prices. The model will be deployed on an ongoing basis, to predict the price of a new house when a description is added to a website. Here are four features that could be used as predictors.
1. Size of the house (in square meters)
2. Average sales price of homes in the same neighborhood
3. Latitude and longitude of the house
4. Whether the house has a basement
You have historic data to train and validate the model.
Which of the features is most likely to be a source of leakage?
```
# Fill in the line below with one of 1, 2, 3 or 4.
potential_leakage_feature = 2
# Check your answer
q_5.check()
#q_5.hint()
#q_5.solution()
```
# Conclusion
Leakage is a hard and subtle issue. You should be proud if you picked up on the issues in these examples.
Now you have the tools to make highly accurate models, and pick up on the most difficult practical problems that arise with applying these models to solve real problems.
There is still a lot of room to build knowledge and experience. Try out a [Competition](https://www.kaggle.com/competitions) or look through our [Datasets](https://kaggle.com/datasets) to practice your new skills.
Again, Congratulations!
---
*Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum/161289) to chat with other Learners.*
| github_jupyter |
```
import numpy as np
np.random.seed(42)
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.metrics import roc_auc_score
from keras.models import Model
from keras.layers import Input, Embedding, Dense, Conv2D, MaxPool2D
from keras.layers import Reshape, Flatten, Concatenate, Dropout, SpatialDropout1D
from keras.preprocessing import text, sequence
from keras.callbacks import Callback
import warnings
warnings.filterwarnings('ignore')
import os
os.environ['OMP_NUM_THREADS'] = '4'
EMBEDDING_FILE = 'fasttext-crawl-300d-2M.vec'
train = df2
final_out=pd.read_csv('../input/yangutu/Devex_submission_format.csv', low_memory=False, encoding='latin1')
test = pd.read_csv("../input/yangutu/Devex_test_questions.csv",low_memory=False, encoding='latin1')
X_train = train["Text"].fillna("fillna").values
y_train=train[[i for i in train.columns if i not in ["Text","ID"]]]
X_test = test["Text"].fillna("fillna").values
max_features = 3000
maxlen = 100
embed_size = 300
tokenizer = text.Tokenizer(num_words=max_features)
tokenizer.fit_on_texts(list(X_train) + list(X_test))
X_train = tokenizer.texts_to_sequences(X_train)
X_test = tokenizer.texts_to_sequences(X_test)
x_train = sequence.pad_sequences(X_train, maxlen=maxlen)
x_test = sequence.pad_sequences(X_test, maxlen=maxlen)
def get_coefs(word, *arr): return word, np.asarray(arr, dtype='float32')
embeddings_index = dict(get_coefs(*o.rstrip().rsplit(' ')) for o in open(EMBEDDING_FILE))
word_index = tokenizer.word_index
nb_words = min(max_features, len(word_index))
embedding_matrix = np.zeros((nb_words, embed_size))
for word, i in word_index.items():
if i >= max_features: continue
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None: embedding_matrix[i] = embedding_vector
class RocAucEvaluation(Callback):
def __init__(self, validation_data=(), interval=1):
super(Callback, self).__init__()
self.interval = interval
self.X_val, self.y_val = validation_data
def on_epoch_end(self, epoch, logs={}):
if epoch % self.interval == 0:
y_pred = self.model.predict(self.X_val, verbose=0)
score = roc_auc_score(self.y_val, y_pred)
print("\n ROC-AUC - epoch: %d - score: %.6f \n" % (epoch+1, score))
filter_sizes = [1,2,3,5]
num_filters = 32
#used this on kaggle gpu
class RocAucEvaluation(Callback):
def __init__(self, validation_data=(), interval=1):
super(Callback, self).__init__()
self.interval = interval
self.X_val, self.y_val = validation_data
def on_epoch_end(self, epoch, logs={}):
if epoch % self.interval == 0:
y_pred = self.model.predict(self.X_val, verbose=0)
self.y_val = np.array([0, 0, 0, 0])
y_pred = np.array([1, 0, 0, 0])
try:
roc_auc_score(self.y_val, y_pred)
except ValueError:
pass
# print("\n ROC-AUC - epoch: %d - score: %.6f \n" % (epoch+1, score))
filter_sizes = [1,2,3,5]
num_filters = 32
def get_model():
inp = Input(shape=(maxlen, ))
x = Embedding(max_features, embed_size, weights=[embedding_matrix])(inp)
x = SpatialDropout1D(0.4)(x)
x = Reshape((maxlen, embed_size, 1))(x)
conv_0 = Conv2D(num_filters, kernel_size=(filter_sizes[0], embed_size), kernel_initializer='normal',
activation='elu')(x)
conv_1 = Conv2D(num_filters, kernel_size=(filter_sizes[1], embed_size), kernel_initializer='normal',
activation='elu')(x)
conv_2 = Conv2D(num_filters, kernel_size=(filter_sizes[2], embed_size), kernel_initializer='normal',
activation='elu')(x)
conv_3 = Conv2D(num_filters, kernel_size=(filter_sizes[3], embed_size), kernel_initializer='normal',
activation='elu')(x)
maxpool_0 = MaxPool2D(pool_size=(maxlen - filter_sizes[0] + 1, 1))(conv_0)
maxpool_1 = MaxPool2D(pool_size=(maxlen - filter_sizes[1] + 1, 1))(conv_1)
maxpool_2 = MaxPool2D(pool_size=(maxlen - filter_sizes[2] + 1, 1))(conv_2)
maxpool_3 = MaxPool2D(pool_size=(maxlen - filter_sizes[3] + 1, 1))(conv_3)
z = Concatenate(axis=1)([maxpool_0, maxpool_1, maxpool_2, maxpool_3])
z = Flatten()(z)
z = Dropout(0.1)(z)
outp = Dense(27, activation="sigmoid")(z)
model = Model(inputs=inp, outputs=outp)
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
return model
model = get_model()
batch_size = 32
epochs = 10
X_tra, X_val, y_tra, y_val = train_test_split(x_train, y_train, train_size=0.95, random_state=233)
RocAuc = RocAucEvaluation(validation_data=(X_val, y_val), interval=1)
hist = model.fit(X_tra, y_tra, batch_size=batch_size, epochs=epochs, validation_data=(X_val, y_val),
callbacks=[RocAuc], verbose=2)
y_pred = model.predict(x_test, batch_size=32)
```
| github_jupyter |
# Signals and Sampling
In this notebook, we will be exploring how signals look, how they are processed and sampled. We will be using the healthy cough sound to explore these properties of signals.
This script is based on the Standford MIR project found [here](https://github.com/stevetjoa/stanford-mir).
Use [librosa.load](https://librosa.github.io/librosa/generated/librosa.core.load.html#librosa.core.load) to load an audio file into an audio array. Return both the audio array as well as the sample rate:
```
import librosa
fileName = '/home/shakes/Dev/workspace/trunk/courses/ELEC3004/Signal_sample/HealthyCoughs.wav'
x, sr = librosa.load(fileName)
```
If you receive an error with librosa.load, you may need to [install ffmpeg](https://librosa.github.io/librosa/install.html#ffmpeg).
Display the length of the audio array and sample rate:
```
print x.shape
print sr
```
### Visualizing Audio
In order to display plots inside the Jupyter notebook, run the following commands, preferably at the top of your notebook:
```
%matplotlib inline
import seaborn # optional
import matplotlib.pyplot as plt
import librosa.display
```
Plot the audio array using [librosa.display.waveplot](https://librosa.github.io/librosa/generated/librosa.display.waveplot.html#librosa.display.waveplot):
```
plt.figure(figsize=(12, 4))
librosa.display.waveplot(x, sr=sr)
```
Display a spectrogram using [librosa.display.specshow](https://librosa.github.io/librosa/generated/librosa.display.specshow.html):
```
X = librosa.stft(x)
Xdb = librosa.amplitude_to_db(X)
plt.figure(figsize=(12, 5))
librosa.display.specshow(Xdb, sr=sr, x_axis='time', y_axis='hz')
```
### Playing and Writing Audio
Using [IPython.display.Audio](http://ipython.org/ipython-doc/2/api/generated/IPython.lib.display.html#IPython.lib.display.Audio), you can play an audio file:
```
import IPython.display as ipd
ipd.Audio(fileName) # load a local WAV file
```
Audio can also accept a NumPy array. Let's synthesize a pure tone at 440 Hz:
```
import numpy
sr = 22050 # sample rate
T = 2.0 # seconds
t = numpy.linspace(0, T, int(T*sr), endpoint=False) # time variable
tone = 0.5*numpy.sin(2*numpy.pi*440*t) # pure sine wave at 440 Hz
```
Listen to the audio array:
```
ipd.Audio(tone, rate=sr) # load a NumPy array
```
Let's do some basic processing to the cough audio signal. We will do a low pass filter of the signal the old fashioned way
```
X = numpy.fft.rfft(x)
#Determine the frequencies of the FFT
W = numpy.fft.fftfreq(X.size, d=1.0/sr)
# If our original signal time was in seconds, this is now in Hz
# low pass signal
cut_X = X.copy()
cut_X[(W<100)] = 0
cut_signal = numpy.fft.irfft(cut_X)
```
Plot the audio array using [librosa.display.waveplot](https://librosa.github.io/librosa/generated/librosa.display.waveplot.html#librosa.display.waveplot) or [librosa.display.specshow](https://librosa.github.io/librosa/generated/librosa.display.specshow.html):
```
#plt.figure(figsize=(12, 4))
#librosa.display.waveplot(cut_signal, sr=sr)
X = librosa.stft(cut_signal)
Xdb = librosa.amplitude_to_db(X)
plt.figure(figsize=(12, 5))
librosa.display.specshow(Xdb, sr=sr, x_axis='time', y_axis='hz')
```
Play the result
```
ipd.Audio(cut_signal, rate=sr) # load a NumPy array
```
librosa.output.write_wav saves a NumPy array to a WAV file.
```
outFileName = '/home/shakes/Dev/workspace/trunk/courses/ELEC3004/Signal_sample/HealthyCough_lpass.wav'
librosa.output.write_wav(outFileName, x, sr)
```
How would you do high pass filtering?
| github_jupyter |
This example notebook uses the averaging functions found ins the diff_classifier msd module to find average msd profiles over input msd datasets using precision-weighted averaging. Precision is the inverse of the standard squared error. This increases the contribution of videos that have many particles and more homogeneous datasets to the final calculated MSD.
**Note: The files used in this notebook analysis are acquired via the previous demo notebook `cloudknot_parallelization_demo.ipynb`.** This notebook will not be run as-is. `cloudknot_patallelization_demo.ipynb` must be run first. The bucket parameter must also be changed to the private bucket of the user, as in `cloudknot_patallelization_demo.ipynb`.
```
folder = 'test_files'
bucket = 'nancelab.publicfiles'
import numpy as np
import diff_classifier.aws as aws
import diff_classifier.msd as msd
experiment = 'test' #Used for naming purposes. Should exclude XY and well information
vids = 2
to_track = []
wells = 1
frames = 651
fps = 1
for num in range(1, vids+1):
to_track.append('{}_{}'.format(experiment, '%02d' % num)) #As per usual, use for loops to include all experiments
geomean = {}
gSEM = {}
for sample_name in to_track:
# Users can toggle between using pre-calculated geomean files and calculating new values by commenting out the relevant
# lines of code within the for loop.
aws.download_s3('{}/geomean_{}.csv'.format(folder, sample_name), 'geomean_{}.csv'.format(sample_name), bucket_name=bucket)
aws.download_s3('{}/geoSEM_{}.csv'.format(folder, sample_name), 'geoSEM_{}.csv'.format(sample_name), bucket_name=bucket)
geomean[sample_name] = np.genfromtxt('geomean_{}.csv'.format(sample_name))
gSEM[sample_name] = np.genfromtxt('geoSEM_{}.csv'.format(sample_name))
#aws.download_s3('{}/msd_{}.csv'.format(folder, sample_name), 'msd_{}.csv'.format(sample_name), bucket_name=bucket)
#geomean[sample_name], gSEM[sample_name] = msd.geomean_msdisp(sample_name, umppx=1, fps=fps,
# remote_folder=folder, bucket=bucket)
print('Done with {}'.format(sample_name))
weights, wh1 = msd.precision_weight(to_track, gSEM)
geodata = msd.precision_averaging(to_track, geomean, gSEM, weights,
bucket=bucket, folder=folder, experiment=experiment)
```
Note that in cases where two or more averaging steps are needed (for instance, if the user takes 5 videos per well with a total of four wells), averaging steps can be performed consecutively. the msd.binning function is a helpful tool by defining bins over which to average for multi-step averaging.
```
to_track.append('test')
msd.plot_all_experiments(to_track, yrange=(10**1, 10**3), bucket=bucket, folder=folder)
import pandas as pd
filename = 'features_test_01.csv'
folder = 'test_files'
aws.download_s3('{}/{}'.format(folder, filename), filename, bucket_name='nancelab.publicfiles')
fstats = pd.read_csv(filename, encoding = "ISO-8859-1", index_col='Unnamed: 0')
tgroups = {}
tgroups[0] = fstats[fstats['frames']<100]
tgroups[1] = fstats[fstats['frames']<300]
tgroups[1] = tgroups[1][tgroups[1]['frames']>100]
tgroups[2] = fstats[300<fstats['frames']]
import diff_classifier.features as ft
%matplotlib inline
ft.feature_violin(tgroups, feature='alpha', ylim=[0, 4], majorticks = np.linspace(0, 4, 11))
```
| github_jupyter |
# Setup
---
```
from graphqlclient import GraphQLClient
import pandas as pd
ENDPOINT = "https://api.thegraph.com/subgraphs/name/blocklytics/bancor"
client = GraphQLClient(ENDPOINT)
```
# Fetch data
---
```
# Results must be paginated.
# Subgraphs return a maximum of 100 rows.
limit = 100
offset = 0
fetching_results = True
converters = []
# Fetch paginated results
while fetching_results:
# This query manually removes certain converters
# See https://blocklytics.org/blog/bancor-subgraph/
QUERY = """
{{
converters(
first:{0},
skip:{1},
where: {{
id_not_in: ["0x77feb788c747a701eb65b8d3b522302aaf26b1e2", "0xcbc6a023eb975a1e2630223a7959988948e664f3", "0x11614c5f1eb215ecffe657da56d3dd12df395dc8", "0x2769eb86e3acdda921c4f36cfe6cad035d95d31b", "0x2ac0e433c3c9ad816db79852d6f933b0b117aefe", "0x37c88474b5d6c593bbd2e4ce16635c08f8215b1e", "0x445556b7215349b205997aaaf6c6dfa258eb029d", "0x46ffcdc6d8e6ed69f124d944bbfe0ac74f8fcf7f", "0x587044b74004e3d5ef2d453b7f8d198d9e4cb558"]
}}
) {{
id
smartToken {{
id
}}
tokenBalances {{
token {{
id
symbol
name
decimals
}}
balance
}}
}}
}}
""".format(limit, offset)
result = json.loads(client.execute(QUERY))
converters += result['data']['converters']
# Prepare for pagination
result_length = len(result['data']['converters'])
if limit == result_length:
offset += limit
else:
fetching_results = False
# Load data into a new df
df = pd.DataFrame()
# Iterate over converters
i = 0
for converter in converters:
# Skip empty converters
if len(converter['tokenBalances']) == 0:
continue
converter_address = converter['id']
smart_token_address = converter['smartToken']['id']
df.at[i, 'exchange'] = converter_address
# Iterate over token balances
for tokenBalance in converter['tokenBalances']:
token = tokenBalance['token']['id']
# Skip converter's smart token
# See https://blocklytics.org/blog/bancor-subgraph/
if token == smart_token_address:
continue
# Handle remaining token details
balance = tokenBalance['balance']
symbol = tokenBalance['token']['symbol']
name = tokenBalance['token']['name']
decimals = tokenBalance['token']['decimals']
try: # try/catch for missing token details
balance_converted = float(balance) / 10 ** float(decimals)
except:
print("Could not find decimals for {0}. Assumed 18".format(token))
balance_converted = float(balance) / 10 ** float(18)
# Set base token to BNT or USDB
if 'base' in df.columns and df.base.isna().iloc[i] == False:
# Base has already been set for this converter
df.at[i, 'token'] = token
df.at[i, 'tokenSymbol'] = symbol
df.at[i, 'tokenName'] = name
df.at[i, 'tokenLiquidity'] = balance_converted
else:
# No base has been set for this converter
if token == '0x1f573d6fb3f13d689ff844b4ce37794d79a7ff1c' or token == '0x309627af60f0926daa6041b8279484312f2bf060':
# Bancor converts use BNT or USDB base
df.at[i, 'base'] = token
df.at[i, 'baseSymbol'] = symbol
# df.at[i, 'baseName'] = name
df.at[i, 'baseLiquidity'] = balance_converted
else:
df.at[i, 'token'] = token
df.at[i, 'tokenSymbol'] = symbol
# df.at[i, 'tokenName'] = name
df.at[i, 'tokenLiquidity'] = balance_converted
i += 1
df['basePrice'] = df['baseLiquidity'] / df['tokenLiquidity'] # Assumes 50% weight
print(df.shape)
```
# Result
---
## USDB Converters
```
df[df.baseSymbol == "USDB"][['baseSymbol', 'baseLiquidity', 'tokenSymbol', 'tokenLiquidity', 'basePrice']]\
.sort_values(by='baseLiquidity', ascending=False)\
.reset_index(drop=True)\
.head(10)
```
## BNT Converters
```
df[df.baseSymbol == "BNT"][['baseSymbol', 'baseLiquidity', 'tokenSymbol', 'tokenLiquidity', 'basePrice']]\
.sort_values(by='baseLiquidity', ascending=False)\
.reset_index(drop=True)\
.head(10)
```
| github_jupyter |
```
#imports and settings
import pandas as pd
pd.set_option('display.expand_frame_repr', False)
import numpy as np
from matplotlib import pyplot as plt
plt.style.use('fivethirtyeight')
from patsy import dmatrices
%pylab inline
import warnings
warnings.filterwarnings('ignore')
df = pd.read_csv('../input/loan.csv', low_memory = False)
#Returns a random sample of items - 30% so the dataset is easy to work with
df_sample = df.sample(frac = 0.3)
df_sample.head(2)
#Analyzing target variable - loan_status
df_sample['loan_status'].value_counts()
#getting rid of loans with statuses we do not care about
#we do not care about current loans
#explanation of difference between charged off and default https://help.lendingclub.com/hc/en-us/articles/216127747
#we only care about those loans that are either fully paid or are
#very late
#too little examples with "does not meet the credit policy" to care about these...
mask = df_sample['loan_status'].isin(['Fully Paid','Charged Off','Default'])
df_sample = df_sample[mask]
df_sample['loan_status'].value_counts()
# now we only work with loans that are either fully paid or late > 121 days
# We create target variable with these two possible values. Positive class
# are late loans - we care about these and want to analyze in detail.
def CreateTarget(status):
if status == 'Fully Paid':
return 0
else:
return 1
df_sample['Late_Loan'] = df_sample['loan_status'].map(CreateTarget)
df_sample['Late_Loan'].value_counts()
#drop features with more than 25% missing values
features_missing_series = df_sample.isnull().sum() > len(df_sample)/10
features_missing_series = features_missing_series[features_missing_series == True]
features_missing_list = features_missing_series.index.tolist()
df_sample = df_sample.drop(features_missing_list,axis =1)
# drop features that have no or little predictive power and original target
df_sample = df_sample.drop(['id','member_id','loan_status','url','zip_code','policy_code','application_type','last_pymnt_d','last_credit_pull_d','verification_status','pymnt_plan','funded_amnt','funded_amnt_inv','sub_grade','out_prncp','out_prncp_inv','total_pymnt_inv','total_pymnt','total_pymnt_inv','total_rec_prncp','total_rec_int','total_rec_late_fee','recoveries','collection_recovery_fee','last_pymnt_amnt','initial_list_status','earliest_cr_line'],axis =1)
#replace missing values with Unknown value or mean when feature is numerical
df_sample['emp_title'].fillna('Unknown',inplace = True)
df_sample['title'].fillna('Unknown',inplace = True)
df_sample['revol_util'].fillna(df_sample['revol_util'].mean(),inplace = True)
df_sample['collections_12_mths_ex_med'].fillna(df_sample['collections_12_mths_ex_med'].mean(),inplace = True)
df_sample.isnull().sum() #there are no missing values left
#old categorical emp_length feature
df_sample['emp_length'].value_counts()
#new numerical emp_length feature
def EmpLength(year):
if year == '< 1 year':
return 0.5
elif year == 'n/a': #assuming that if not filled out employments was < 1
return 0.5
elif year == '10+ years':
return 10
else:
return float(year.rstrip(' years'))
df_sample['emp_length_num'] = df_sample['emp_length'].map(EmpLength)
df_sample = df_sample.drop('emp_length',axis =1 )
df_sample['emp_length_num'].value_counts()
#transforming to date datatype
df_sample['issue_d'] = pd.to_datetime(df_sample.issue_d)
#datatypes of features
# object = string ?
df_sample.dtypes.value_counts()
#cleaned and transformed data ready for analysis and ML
#numerical features - means
print(df_sample.select_dtypes(include=['float64']).apply(np.mean).apply(str))
# categorical variables
print(df_sample.select_dtypes(include=['object']).columns)
# target variable - boolean
print(df_sample.select_dtypes(include=['bool']).columns)
df_sample['purpose'].value_counts()
#distribution of our class/targer variable Late_Loan , True if loan is late.
plt.figure(figsize=(5,5))
df_sample['Late_Loan'].value_counts().plot(kind = 'pie',autopct='%.0f%%', startangle=100, fontsize=17)
plt.show()
Amount_By_Year = df_sample.groupby(df_sample['issue_d'].dt.year)['loan_amnt'].mean()
Amount_By_Year = pd.DataFrame(Amount_By_Year)
Amount_By_Year['YoY Change %'] = Amount_By_Year.pct_change()*100
Amount_By_Year.rename(columns = {'loan_amnt':'Average Loan Amount'})
plt.figure(figsize=(8,3))
Amount_By_Year_Status_True = df_sample.groupby([df_sample['issue_d'].dt.year,df_sample['Late_Loan'][df_sample['Late_Loan'] == True]])['loan_amnt'].mean().plot(kind = 'line', label = 'True')
Amount_By_Year_Status_False = df_sample.groupby([df_sample['issue_d'].dt.year,df_sample['Late_Loan'][df_sample['Late_Loan'] == False]])['loan_amnt'].mean().plot(kind = 'line',label = 'False')
plt.xlabel('Year')
plt.ylabel('Average Loan')
plt.legend(loc='best')
plt.show()
#This graph normalizes the purpose variables, using value counts.
#This gives us an idea of what percentage and purpose the clients are taking out loans for.
all_rows = df_sample['purpose']
pur = df_sample['purpose'].value_counts()
purp = pur/len(all_rows)
purp.plot(kind='bar')
flat = pur/len(df_sample['purpose']) * 100
print(flat)
#I'm going to look at the installment payments against the grade and term of the loan.
#This pivot table shows the installment payments by grade and term.
loan_g = pd.pivot_table(df_sample,
index= ['grade','term'],
columns= ['installment'] ,
values= 'loan_amnt',
aggfunc = sum)
loan_g.T.idxmax()
loan_g.T.idxmax().plot(kind='bar')
#This graph looks at the people who are late on their loans, renters and people paying their mortages tend to be late on payments
#A possible reason may be that the owners, others, and none may not have a financial burden.
late = df_sample[['home_ownership', 'Late_Loan']]
late_people = late['Late_Loan']== True
people = late[late_people]
sad = people['home_ownership'].value_counts().plot(kind='bar', color= 'orange')
xlabel('Living Status')
ylabel('People With Late Loans')
print(df_sample['int_rate'].mean())
mask_delinq = df_sample['delinq_2yrs'] <= 11
df_sample.groupby(df_sample['delinq_2yrs'][mask_delinq])['int_rate'].mean().plot(kind='line')
plt.show()
df_sample['delinq_2yrs'].value_counts()
mask_pub_rec = df_sample['pub_rec'] <= 6
df_sample.groupby(df_sample['pub_rec'][mask_pub_rec])['int_rate'].mean().plot(kind='line')
plt.show()
mask_pub_rec = df_sample['pub_rec'] <= 6
df_sample.groupby(df_sample['pub_rec'][mask_pub_rec])['annual_inc'].mean().plot(kind='line')
plt.show()
from patsy import dmatrices
from sklearn.model_selection import train_test_split
from sklearn import tree
from sklearn import metrics
from sklearn.model_selection import StratifiedKFold, cross_val_score
from sklearn.ensemble import RandomForestClassifier
from imblearn.over_sampling import SMOTE
df_sample['installmentAsPercent'] = df_sample['installment']/(df['annual_inc']/12)
def GradeInt(x):
if x == "A":
return 1
elif x == "B":
return 2
elif x == "C":
return 3
elif x == "D":
return 4
elif x == "E":
return 5
else:
return 6
df_sample['GradeInt'] = df_sample['grade'].map(GradeInt)
Y,X = dmatrices('Late_Loan ~ 0 + int_rate + GradeInt + loan_amnt + installment + annual_inc + dti + delinq_2yrs + inq_last_6mths + open_acc + pub_rec + revol_bal + revol_util + total_acc + collections_12_mths_ex_med + acc_now_delinq + emp_length_num + term + home_ownership + purpose + installmentAsPercent',df_sample, return_type = 'dataframe')
X_columns = X.columns
sm = SMOTE(random_state=42)
X, Y = sm.fit_sample(X, Y)
X = pd.DataFrame(X,columns=X_columns)
#distribution of our class/targer variable Late_Loan , True if loan is late.
Y_df = pd.DataFrame(Y,columns=['Late_Loan'])
plt.figure(figsize=(5,5))
Y_df['Late_Loan'].value_counts().plot(kind = 'pie',autopct='%.0f%%', startangle=100, fontsize=17)
plt.show()
X_train,X_test, Y_train, Y_test = train_test_split(X,Y,test_size=0.3)
best_model = []
for i in range(1,15):
model = tree.DecisionTreeClassifier(criterion = 'entropy', max_depth=i)
kfold = StratifiedKFold(n_splits= 10, shuffle = True)
scores = cross_val_score(model, X_train, Y_train, cv = kfold )
best_model.append(scores.mean())
plt.plot(range(1,15),best_model)
plt.show()
from sklearn.grid_search import RandomizedSearchCV
from scipy.stats import randint as sp_randint
param_dist = { "max_depth": sp_randint(6,10),
"max_features": sp_randint(3,15),
"max_leaf_nodes": [10,20,30,40,50],
"min_samples_leaf": [25,50,75,100,150,250,500],
}
random_search = RandomizedSearchCV(model,
param_distributions=param_dist,
n_iter=50)
random_search.fit(X_train, Y_train)
print(random_search.best_score_)
print(random_search.best_estimator_)
best_model = random_search.best_estimator_
best_model.fit(X_train,Y_train)
importance = sorted(zip(map(lambda x: round(x, 4), best_model.feature_importances_), X.columns),reverse=True)
y_val = []
x_val = [x[0] for x in importance]
for x in importance:
y_val.append(x[1])
pd.Series(x_val,index=y_val)[:7].plot(kind='bar')
plt.show()
from sklearn.ensemble import BaggingClassifier
bagging = BaggingClassifier(best_model, random_state=1)
bagging.fit(X,Y)
accuracy = metrics.accuracy_score(bagging.predict(X_test),Y_test)
precision = metrics.precision_score(bagging.predict(X_test),Y_test,pos_label=1)
recall = metrics.recall_score(bagging.predict(X_test),Y_test,pos_label=1)
confusion_matrix = metrics.confusion_matrix(Y_test,bagging.predict(X_test),labels=[1,0])
print(accuracy)
print(precision)
print(recall)
print(confusion_matrix)
labels = ['Late', 'Paid']
fig = plt.figure()
ax = fig.add_subplot(111)
cax = ax.matshow(confusion_matrix)
fig.colorbar(cax)
plt.title('Confusion matrix of the classifier')
ax.set_xticklabels([''] + labels)
ax.set_yticklabels([''] + labels)
plt.xlabel('Predicted')
plt.ylabel('True')
plt.show()
```
| github_jupyter |
# Tabular data
```
%reload_ext autoreload
%autoreload 2
from fastai.gen_doc.nbdoc import *
from fastai.tabular import *
from fastai import *
from fastai.docs import *
from fastai.tabular.models import *
```
[`tabular`](/tabular.html#tabular) contains all the necessary classes to deal with tabular data, across two modules:
- [`tabular.transform`](/tabular.transform.html#tabular.transform): defines the [`TabularTransform`](/tabular.transform.html#TabularTransform) class to help with preprocessing;
- [`tabular.data`](/tabular.data.html#tabular.data): defines the [`TabularDataset`](/tabular.data.html#TabularDataset) that handles that data, as well as the methods to quickly get a [`DataBunch`](/data.html#DataBunch).
To create a model, you'll need to use [`models.tabular`](/tabular.html#tabular). See below for and end-to-end example using all these modules.
## Preprocessing tabular data
Tabular data usually comes in the form of a delimited file (such as CSV) containing variables of different kinds: text/category, numbers, and perhaps some missing values. The example we'll work with in this section is a sample of the [adult dataset](https://archive.ics.uci.edu/ml/datasets/adult) which has some census information on individuals. We'll use it to train a model to predict whether salary is greater than \$50k or not.
```
untar_data(ADULT_PATH)
ADULT_PATH
df = pd.read_csv(ADULT_PATH/'adult.csv')
df.head()
```
Here all the information that will form our input is in the 14 first columns, and the dependant variable is the last column. We will split our input between two types of variables: categoricals and continuous.
- Categorical variables will be replaced by a category, a unique id that identifies them, before passing through an embedding layer.
- Continuous variables will be normalized then directly fed to the model.
Another thing we need to handle are the missing values: our model isn't going to like receiving NaNs so we should remove them in a smart way. All of this preprocessing is done by [`TabularTransform`](/tabular.transform.html#TabularTransform) objects and [`TabularDataset`](/tabular.data.html#TabularDataset).
We can define a bunch of Transforms that will be applied to our variables. Here we transform all categorical variables into categories, and we replace missing values for continuous variables by the median of the corresponding column.
```
tfms = [FillMissing, Categorify]
```
We split our data into training and validation sets.
```
train_df, valid_df = df[:-2000].copy(),df[-2000:].copy()
```
First let's split our variables between categoricals and continuous (we can ignore the dependant variable at this stage). fastai will assume all variables that aren't dependent or categorical are continuous, unless we explicitly pass a list to the `cont_names` parameter when constructing our [`DataBunch`](/data.html#DataBunch).
```
dep_var = '>=50k'
cat_names = ['workclass', 'education', 'marital-status', 'occupation', 'relationship', 'race', 'sex', 'native-country']
```
Now we're ready to pass these details to [`tabular_data_from_df`](/tabular.data.html#tabular_data_from_df) to create the [`DataBunch`](/data.html#DataBunch) that we'll use for training.
```
data = tabular_data_from_df(ADULT_PATH, train_df, valid_df, dep_var, tfms=tfms, cat_names=cat_names)
print(data.train_ds.cont_names) # `cont_names` defaults to: set(df)-set(cat_names)-{dep_var}
```
We can grab a mini-batch of data and take a look (note that [`to_np`](/torch_core.html#to_np) here converts from pytorch tensor to numpy):
```
(cat_x,cont_x),y = next(iter(data.train_dl))
for o in (cat_x, cont_x, y): print(to_np(o[:5]))
```
After being processed in [`TabularDataset`](/tabular.data.html#TabularDataset), the categorical variables are replaced by ids and the continuous variables are normalized. The codes corresponding to categorical variables are all put together, as are all the continuous variables.
## Defining a model
Once we have our data ready in a [`DataBunch`](/data.html#DataBunch), we just need to create a model to then define a [`Learner`](/basic_train.html#Learner) and start training. The fastai library has a flexible and powerful [`TabularModel`](/tabular.models.html#TabularModel) in [`models.tabular`](/tabular.html#tabular). To use that function, we just need to specify the embedding sizes for each of our categorical variables.
```
learn = get_tabular_learner(data, layers=[200,100], emb_szs={'native-country': 10}, metrics=accuracy)
learn.fit_one_cycle(1, 1e-2)
```
| github_jupyter |
# Continuous Control
---
You are welcome to use this coding environment to train your agent for the project. Follow the instructions below to get started!
### 1. Start the Environment
Run the next code cell to install a few packages. This line will take a few minutes to run!
```
!pip -q install ./python
```
The environments corresponding to both versions of the environment are already saved in the Workspace and can be accessed at the file paths provided below.
Please select one of the two options below for loading the environment.
### Install unityagents if not installed
```
# !pip install unityagents
from unityagents import UnityEnvironment
import numpy as np
# select this option to load version 1 (with a single agent) of the environment
env = UnityEnvironment(file_name='/data/Reacher_One_Linux_NoVis/Reacher_One_Linux_NoVis.x86_64')
# select this option to load version 2 (with 20 agents) of the environment
# env = UnityEnvironment(file_name='/data/Reacher_Linux_NoVis/Reacher.x86_64')
```
Environments contain **_brains_** which are responsible for deciding the actions of their associated agents. Here we check for the first brain available, and set it as the default brain we will be controlling from Python.
```
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
```
### 2. Examine the State and Action Spaces
Run the code cell below to print some information about the environment.
```
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents
num_agents = len(env_info.agents)
print('Number of agents:', num_agents)
# size of each action
action_size = brain.vector_action_space_size
print('Size of each action:', action_size)
# examine the state space
states = env_info.vector_observations
state_size = states.shape[1]
print('There are {} agents. Each observes a state with length: {}'.format(states.shape[0], state_size))
print('The state for the first agent looks like:', states[0])
```
### 3. Take Random Actions in the Environment
In the next code cell, you will learn how to use the Python API to control the agent and receive feedback from the environment.
Note that **in this coding environment, you will not be able to watch the agents while they are training**, and you should set `train_mode=True` to restart the environment.
```
# env_info = env.reset(train_mode=True)[brain_name] # reset the environment
# states = env_info.vector_observations # get the current state (for each agent)
# scores = np.zeros(num_agents) # initialize the score (for each agent)
# while True:
# actions = np.random.randn(num_agents, action_size) # select an action (for each agent)
# actions = np.clip(actions, -1, 1) # all actions between -1 and 1
# env_info = env.step(actions)[brain_name] # send all actions to tne environment
# next_states = env_info.vector_observations # get next state (for each agent)
# rewards = env_info.rewards # get reward (for each agent)
# dones = env_info.local_done # see if episode finished
# scores += env_info.rewards # update the score (for each agent)
# states = next_states # roll over states to next time step
# if np.any(dones): # exit loop if episode finished
# break
# print('Total score (averaged over agents) this episode: {}'.format(np.mean(scores)))
```
When finished, you can close the environment.
### 4. It's Your Turn!
Now it's your turn to train your own agent to solve the environment! A few **important notes**:
- When training the environment, set `train_mode=True`, so that the line for resetting the environment looks like the following:
```python
env_info = env.reset(train_mode=True)[brain_name]
```
- To structure your work, you're welcome to work directly in this Jupyter notebook, or you might like to start over with a new file! You can see the list of files in the workspace by clicking on **_Jupyter_** in the top left corner of the notebook.
- In this coding environment, you will not be able to watch the agents while they are training. However, **_after training the agents_**, you can download the saved model weights to watch the agents on your own machine!
# Importing necessary packages
```
import random
import copy
from collections import namedtuple, deque
import matplotlib.pyplot as plt
import pandas as pd
%matplotlib inline
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from dqn_agent import Agent
from model import Actor, Critic
```
## Initiating Agent
```
agent = Agent(state_size=state_size, action_size=action_size, random_seed=10)
def DDPG(episode_num=1000, max_time=10000):
"""
Params
======
:param n_episodes (int): maximum number of training episodes
:param max_t (int): maximum number of timesteps per episode
"""
scores_deque = deque(maxlen=100)
scores = []
for episode in range(1, episode_num + 1):
env_info = env.reset(train_mode=True)[brain_name]
agent.reset()
state = env_info.vector_observations[0]
score = 0
for t in range(max_time):
action = agent.act(state)
env_info = env.step(action)[brain_name]
next_state = env_info.vector_observations[0]
reward = env_info.rewards[0]
done = env_info.local_done[0]
agent.step(state, action, reward, next_state, done)
score += reward
state = next_state
if done:
break
scores_deque.append(score)
scores.append(score)
if (episode % 10 == 0):
print('\rEpisode {}\tScore: {:.2f}\tAverage Score: {:.2f}'.format(episode, score, np.mean(scores_deque)))
print('\rEpisode {}\tAverage Score: {:.2f}'.format(episode, np.mean(scores_deque)))
torch.save(agent.actor_local.state_dict(), 'checkpoint_actor.pth')
torch.save(agent.critic_local.state_dict(), 'checkpoint_critic.pth')
if np.mean(scores_deque) >= 30.0:
print(
'\nEnvironment solved in {:d} Episodes \tAverage Score: {:.2f}'.format(episode, np.mean(scores_deque)))
torch.save(agent.actor_local.state_dict(), 'checkpoint_actor.pth')
torch.save(agent.critic_local.state_dict(), 'checkpoint_critic.pth')
break
return scores
```
## Training
```
scores =DDPG(episode_num= 500)
```
# Plotting
```
pd.DataFrame(scores).to_csv("p2_scores.csv")
plt.rcParams["figure.figsize"] = [16,8]
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(1, len(scores)+1), scores)
plt.ylabel('Scores')
plt.xlabel('Episode #')
plt.show()
!
```
## Closing env
```
env.close()
```
| github_jupyter |
# Classification of Chest and Abdominal X-rays
## Classificação de radiografias de peito e abdominais
Este é um exemplo de modelo que ao receber um exame de imagem, classifica-o como radiografia do peito ou abdominal.
Code Source: Lakhani, P., Gray, D.L., Pett, C.R. et al. J Digit Imaging (2018) 31: 283. https://doi.org/10.1007/s10278-018-0079-6
The code to download and prepare dataset had been modified form the original source code.
```
# Carrega as bibliotecas necessárias para treinamento do modelo (utilizaremos o Keras)
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import applications
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras import optimizers
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dropout, Flatten, Dense, GlobalAveragePooling2D
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import Adam
!rm -rf /content/*
# Faz o Download do dataset
!wget https://github.com/kunkaweb/Hello_World_Deep_Learning/blob/master/Open_I_abd_vs_CXRs.zip?raw=true
# Renomeia arquivo
!mv Open_I_abd_vs_CXRs.zip?raw=true Open_I_abd_vs_CXRs.zip
# Mostra arquivos na pasta
!ls /content
# Descompacta arquivo
!unzip /content/Open_I_abd_vs_CXRs.zip
# Define as dimensões das imagens
img_width, img_height = 299, 299
# Diretórios de TREINAMENTO e VALIDAÇÃO das imagens
train_data_dir = 'Open_I_abd_vs_CXRs/TRAIN/'
validation_data_dir = 'Open_I_abd_vs_CXRs/VAL/'
# epochs = Numéro de épocas - vezes que passaremos sobre os dados de treinamento
# batch_size = Número de imagen processadas simultaneamente
train_samples = 65
validation_samples = 10
epochs = 20
batch_size = 5
# Define uma rede do tipo Inception V3 e use utiliza pesos de uma modelo pré-treinado no ImageNet (Transfer Learning)
# Remove o topo da rede, pois iremos acoplar uma sequência de rede que ira performar a classificação posteriormente (include_top = False)
base_model = applications.InceptionV3(weights='imagenet', include_top=False,
input_shape=(img_width, img_height,3))
# Constroi um modelo de classificação para colocar no topo da rede convolucional
# Consiste em um global average pooling layer e um fully connected layer com 256 nós
# Então aplica-se o dropout e uma função de ativação do tipo sigmóide
model_top = Sequential()
model_top.add(GlobalAveragePooling2D(input_shape=base_model.output_shape[1:],
data_format=None)),
model_top.add(Dense(256, activation='relu'))
model_top.add(Dropout(0.5))
model_top.add(Dense(1, activation='sigmoid'))
model = Model(inputs=base_model.input, outputs=model_top(base_model.output))
# Compila o modelo utilizando o otimizador Adam
# Utiliza uma taxa de aprendizado baixa (low learning rate - lr) para o transfer learning
model.compile(optimizer=Adam(learning_rate=0.0001, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=0.0),
loss='binary_crossentropy',
metrics=['accuracy'])
# Configurações extras
train_datagen = ImageDataGenerator(
rescale = 1./255, # Rescale pixel values to 0-1 to aid CNN processing
shear_range = 0.2, # 0-1 range for shearing
zoom_range = 0.2, # 0-1 range for zoom
rotation_range = 20, # 0.180 range, degrees of rotation
width_shift_range = 0.2, # 0-1 range horizontal translation
height_shift_range = 0.2, # 0-1 range vertical translation
horizontal_flip = True # set True or false
)
val_datagen = ImageDataGenerator(
rescale=1./255 # Rescale pixel values to 0-1 to aid CNN processing
)
# Class mode é definida como 'binary' pois temos um problema com duas classes (chest e abdominal)
# O gerador embaralha as imagens e envia elas em lotes para a rede - cada classe tem seu diretório próprio
train_generator = train_datagen.flow_from_directory(
train_data_dir,
target_size=(img_height, img_width),
batch_size=batch_size,
class_mode='binary'
)
validation_generator = val_datagen.flow_from_directory(
validation_data_dir,
target_size=(img_height, img_width),
batch_size=batch_size,
class_mode='binary'
)
# Ajusta (fine-tune) o modelo Inception V3 pré-treinado utilizando o data generator
# Especifica as etapas por época (número de imagens/batch_size)
history = model.fit_generator(
train_generator,
steps_per_epoch=train_samples//batch_size,
epochs=epochs,
validation_data=validation_generator,
validation_steps=validation_samples//batch_size
)
# Plota a curva de aprendizagem
import matplotlib.pyplot as plt
print(history.history.keys())
plt.figure()
plt.plot(history.history['accuracy'],'orange', label='Training accuracy')
plt.plot(history.history['val_accuracy'],'blue', label='Validation accuracy')
plt.plot(history.history['loss'],'red', label='Training loss')
plt.plot(history.history['val_loss'],'green', label='validation loss')
plt.legend()
plt.show()
import numpy as np
from keras.preprocessing import image
# Carrega, redimensiona, e mostra duas imagens no lote de TESTE
img_path = 'Open_I_abd_vs_CXRs/TEST/abd2.png'
img_path2 = 'Open_I_abd_vs_CXRs/TEST/chest2.png'
img = image.load_img(img_path, target_size=(img_width, img_height))
img2 = image.load_img(img_path2, target_size=(img_width, img_height))
plt.imshow(img)
plt.show()
# Converte a imagem em um vetor, assim o Keras consegue renderizar a predição
img = image.img_to_array(img)
# Expande o vetor de 3 dimensões (altura, largura, canais) para 4 dimensões (batch size, altura, largura, canais)
# Redimensiona os valores dos pixels para 0-1
x = np.expand_dims(img, axis=0) * 1./255
# Faz predição da imagem de teste 1
# Caso score seja menor que 0.5 = peito, senão = abdominal
score = model.predict(x)
print('Predito:', score, 'Radiografia do peito' if score < 0.5 else 'Radiografia abdominal')
# Mostra e rendereiza a predição da segunda imagem
plt.imshow(img2)
plt.show()
img2 = image.img_to_array(img2)
x = np.expand_dims(img2, axis=0) * 1./255
score = model.predict(x)
print('Predito:', score, 'Radiografia do peito' if score < 0.5 else 'Radiografia abdominal')
```
| github_jupyter |
# Recurrent Neural Network - Word Classification
## Using Special model
Implemented in TensorFlow. Using Seq2Seq to generate the sequence of letter images and recognise them by CNN.
## TODO
```
Remove random border
```
```
import numpy as np
import pandas as pd
import matplotlib as plt
import tensorflow as tf
import tensorflow.contrib.seq2seq as seq2seq
from tensorflow.python.layers import core as layers_core
from tensorflow.python.ops import math_ops
import time
import math
import unidecode
import cv2
from ocr.datahelpers import loadWordsData, correspondingShuffle, char2idx
from ocr.helpers import implt, extendImg, resize
from ocr.mlhelpers import TrainingPlot
from ocr.normalization import letterNorm, imageStandardization
from ocr.tfhelpers import Graph, create_cell
tf.reset_default_graph()
sess = tf.InteractiveSession()
print('Tensorflow', tf.__version__)
%matplotlib notebook
# Increase size of images
plt.rcParams['figure.figsize'] = (9.0, 5.0)
```
### Loading images
```
LANG = 'en'
images, labels, gaplines = loadWordsData(
['data/words2/'], loadGaplines=True)
if LANG == 'en':
for i in range(len(labels)):
labels[i] = unidecode.unidecode(labels[i])
```
## Settings
```
CHARS = 82 if LANG =='cz' else 52
PAD = 0 # Padding
EOS = 1 # End of seq
LETTER_PAD = -1.0
# Testing partialy separated dataset
sep_words = 500
num_buckets = 5
slider_size = (60, 2)
step_size = 2
N_INPUT = slider_size[0]*slider_size[1]
char_size = CHARS + 2
letter_size = 64*64
encoder_layers = 2
encoder_residual_layers = 1 # HAVE TO be smaller than encoder_layers
encoder_units = 256
decoder_layers = 2*encoder_layers # 2* is due to the bidirectional encoder
decoder_residual_layers = 2*encoder_residual_layers
decoder_units = encoder_units
wordRNN_layers = 2
wordRNN_residual_layers = 1
wordRNN_units = 128
attention_size = 256
add_output_length = 4
learning_rate = 1e-3 # 1e-4
max_gradient_norm = 5.0 # For gradient clipping
dropout = 0.4
train_per = 0.8 # Percentage of training data
TRAIN_STEPS = 1000000 # Number of training steps!
TEST_ITER = 150
LOSS_ITER = 50
SAVE_ITER = 1000
BATCH_SIZE = 20 # 64
EPOCH = 1000 # Number of batches in epoch - not accurate
save_location = 'models/word-clas/' + LANG + '/SeqRNN/Classifier3'
```
## Dataset
```
# Shuffle data for later splitting
images, labels, gaplines = correspondingShuffle([images, labels, gaplines])
# TODO: for testing, can be removed
for i in range(len(images)):
images[i] = cv2.copyMakeBorder(
images[i],
0, 0, slider_size[1]//2, slider_size[1]//2,
cv2.BORDER_CONSTANT,
value=[0, 0, 0])
labels_idx = np.empty(len(labels), dtype=object)
for i, label in enumerate(labels):
labels_idx[i] = [char2idx(c, True) for c in label]
labels_idx[i].append(EOS)
# Split data on train and test dataset
div = int(train_per * len(images))
trainImages = images[0:div]
testImages = images[div:]
trainGaplines = gaplines[0:div]
testGaplines = gaplines[div:]
trainLabels_idx = labels_idx[0:div]
testLabels_idx = labels_idx[div:]
print("Training images:", div)
print("Testing images:", len(images) - div)
def stackImage(img, a, b):
""" Add blank columns (lenght a, b) at start and end of image """
return np.concatenate(
(np.zeros((img.shape[0], a)),
np.concatenate((img, np.zeros((img.shape[0], b))), axis=1)),
axis=1)
# Dont mix train and test images
num_new_images = 0 # 2
trainImagesF = np.empty(len(trainImages) * (num_new_images+1), dtype=object)
trainGaplinesF = np.empty(len(trainImages) * (num_new_images+1), dtype=object)
trainLabelsF_idx = np.empty(len(trainImages)*(num_new_images+1), dtype=object)
for idx, img in enumerate(trainImages):
add_idx = idx*(num_new_images+1)
trainImagesF[add_idx] = img
trainGaplinesF[add_idx] = trainGaplines[idx]
trainLabelsF_idx[add_idx] = trainLabels_idx[idx]
for i in range(num_new_images):
a, b = np.random.randint(1, 16, size=2)
trainImagesF[add_idx + (i+1)] = stackImage(img, a, b)
trainGaplinesF[add_idx + (i+1)] = trainGaplines[idx] + a
trainLabelsF_idx[add_idx + (i+1)] = trainLabels_idx[idx]
print("Total train images", len(trainImagesF))
class BucketDataIterator():
""" Iterator for feeding seq2seq model during training """
def __init__(self,
images,
targets,
gaplines,
num_buckets=5,
slider=(60, 30),
slider_step=2,
train=True):
self.train = train
self.slider = slider
# PADDING of images to slider size; -(a // b) = ceil(a/b)
for i in range(len(images)):
images[i] = extendImg(
images[i],
(images[i].shape[0], max(-(-images[i].shape[1] // slider_step) * slider_step, 60)))
in_length = [(image.shape[1] + 1 - slider[1])//slider_step for image in images]
# Split images to sequence of vectors
img_seq = np.empty(len(images), dtype=object)
for i, img in enumerate(images):
img_seq[i] = [img[:, loc * slider_step: loc*slider_step + slider[1]].flatten()
for loc in range(in_length[i])]
end_letter = np.ones(letter_size) # * LETTER_PAD # End letter is full white
np.put(end_letter, [0], [1])
letter_seq = np.empty(len(images), dtype=object)
for i, img in enumerate(images):
letter_seq[i] = [imageStandardization(
letterNorm(img[:, gaplines[i][x]:gaplines[i][x+1]])).flatten()
for x in range(len(gaplines[i])-1)]
letter_seq[i].append(end_letter)
# Create pandas dataFrame and sort it by images width (length)
# letters_length is num_letter + EOS
self.dataFrame = pd.DataFrame({'in_length': in_length,
'letters_length': [len(g) for g in gaplines],
'words_length': [len(t) for t in targets],
'in_images': img_seq,
'letters': letter_seq,
'words': targets
}).sort_values('in_length').reset_index(drop=True)
bsize = int(len(images) / num_buckets)
self.num_buckets = num_buckets
# Create buckets by slicing parts by indexes
self.buckets = []
for bucket in range(num_buckets-1):
self.buckets.append(self.dataFrame.iloc[bucket * bsize: (bucket+1) * bsize])
self.buckets.append(self.dataFrame.iloc[(num_buckets-1) * bsize:])
self.buckets_size = [len(bucket) for bucket in self.buckets]
# cursor[i] will be the cursor for the ith bucket
self.cursor = np.array([0] * num_buckets)
self.bucket_order = np.random.permutation(num_buckets)
self.bucket_cursor = 0
self.shuffle()
print("Iterator created.")
def shuffle(self, idx=None):
""" Shuffle idx bucket or each bucket separately """
for i in [idx] if idx is not None else range(self.num_buckets):
self.buckets[i] = self.buckets[i].sample(frac=1).reset_index(drop=True)
self.cursor[i] = 0
def next_batch(self, batch_size):
"""
Creates next training batch of size: batch_size
Retruns: (in_images, letters, words,
in_length, letter_length, word_length)
"""
i_bucket = self.bucket_order[self.bucket_cursor]
# Increment cursor and shuffle in case of new round
self.bucket_cursor = (self.bucket_cursor + 1) % self.num_buckets
if self.bucket_cursor == 0:
self.bucket_order = np.random.permutation(self.num_buckets)
if self.cursor[i_bucket] + batch_size > self.buckets_size[i_bucket]:
self.shuffle(i_bucket)
# Handle too big batch sizes
if (batch_size > self.buckets_size[i_bucket]):
batch_size = self.buckets_size[i_bucket]
res = self.buckets[i_bucket].iloc[self.cursor[i_bucket]:
self.cursor[i_bucket]+batch_size]
self.cursor[i_bucket] += batch_size
# Check correct length of later prediction of sequences
assert np.all(res['in_length'] + add_output_length >= res['letters_length'])
input_max = max(res['in_length'])
letters_max = max(res['letters_length'])
words_max = max(res['words_length'])
input_seq = np.ones((batch_size, input_max, N_INPUT), dtype=np.float32) * LETTER_PAD
for i, img in enumerate(res['in_images']):
input_seq[i][:res['in_length'].values[i]] = img
input_seq = input_seq.swapaxes(0, 1) # Time major
letters = np.ones((batch_size, letters_max, letter_size), dtype=np.float32) * LETTER_PAD
for i, img in enumerate(res['letters']):
letters[i][:res['letters_length'].values[i]] = img
# Need to pad according to the maximum length output sequence
words = np.zeros([batch_size, words_max], dtype=np.int32)
for i, word in enumerate(res['words']):
words[i][:res['words_length'].values[i]] = word
return (input_seq, letters, words,
res['in_length'].values, res['letters_length'].values, res['words_length'].values)
def next_feed(self, size, words=True, train=None):
""" Create feed directly for model training """
if train is None:
train = self.train
(encoder_inputs_,
letter_targets_,
word_targets_,
encoder_inputs_length_,
letter_targets_length_,
word_targets_length_) = self.next_batch(size)
return {
encoder_inputs: encoder_inputs_,
encoder_inputs_length: encoder_inputs_length_,
letter_targets: letter_targets_,
letter_targets_length: letter_targets_length_,
word_targets: word_targets_,
word_targets_length: word_targets_length_,
keep_prob: (1.0 - dropout) if self.train else 1.0,
is_training: train,
is_words: words
}
# Create iterator for feeding RNN
# Create only once, it modifies: labels_idx
train_iterator = BucketDataIterator(trainImagesF,
trainLabelsF_idx,
trainGaplinesF,
num_buckets,
slider_size,
step_size,
train=True)
train_letters_iterator = BucketDataIterator(trainImagesF[:sep_words],
trainLabelsF_idx[:sep_words],
trainGaplinesF[:sep_words],
1,
slider_size,
step_size,
train=True)
test_iterator = BucketDataIterator(testImages,
testLabels_idx,
testGaplines,
num_buckets,
slider_size,
step_size,
train=False)
```
## Placeholders
```
# Only encoder inputs are time major
# Encoder inputs shape (max_seq_length, batch_size, vec_size)
encoder_inputs = tf.placeholder(shape=(None, None, N_INPUT),
dtype=tf.float32,
name='encoder_inputs')
encoder_inputs_length = tf.placeholder(shape=(None,),
dtype=tf.int32,
name='encoder_inputs_length')
# Required for letter sep. training
# Contains EOS symbol
letter_targets = tf.placeholder(shape=(None, None, letter_size),
dtype=tf.float32,
name='letter_targets')
letter_targets_length = tf.placeholder(shape=(None,),
dtype=tf.int32,
name='letter_targets_length')
# Required for word training
word_targets = tf.placeholder(shape=(None, None),
dtype=tf.int32,
name='word_targets')
word_targets_length = tf.placeholder(shape=(None,),
dtype=tf.int32,
name='word_targets_length')
# Dropout value
keep_prob = tf.placeholder(tf.float32, name='keep_prob')
# Testing control
is_training = tf.placeholder(tf.bool, shape=None, name="is_training")
is_words = tf.placeholder(tf.bool, shape=None, name="is_words")
```
### Decoder Train Feeds
```
sequence_size, batch_size, _ = tf.unstack(tf.shape(encoder_inputs)) # letter_targets
EOS_SLICE = tf.cast(tf.fill([batch_size, 1, letter_size], EOS), tf.float32)
PAD_SLICE = tf.cast(tf.fill([batch_size, 1, letter_size], LETTER_PAD), tf.float32) # PAD
# Train inputs with EOS symbol at start of seq
letter_train_inputs = tf.concat([PAD_SLICE, letter_targets], axis=1) #EOS_SLICE
letter_train_length = letter_targets_length
# Length of infer (test) letter output
# TODO: will have to make shorte...
output_length = tf.minimum(
tf.reduce_max(encoder_inputs_length) * step_size // 15 + add_output_length,
23)
```
## Encoder
```
enc_cell_fw = create_cell(encoder_units,
encoder_layers,
encoder_residual_layers,
is_dropout=True,
keep_prob=keep_prob)
enc_cell_bw = create_cell(encoder_units,
encoder_layers,
encoder_residual_layers,
is_dropout=True,
keep_prob=keep_prob)
### CNN ###
SCALE = 0.01 # 0.1
# Functions for initializing convulation and pool layers
def weights(name, shape):
return tf.get_variable(name, shape=shape,
initializer=tf.contrib.layers.xavier_initializer(),
regularizer=tf.contrib.layers.l2_regularizer(scale=SCALE))
def bias(const, shape, name=None):
return tf.Variable(tf.constant(const, shape=shape), name=name)
def conv2d2(x, W, name=None):
return tf.nn.conv2d(x, W, strides=[1, 2, 1, 1], padding='SAME', name=name)
W_conv1 = weights('W_conv1', shape=[2, 1, 1, 1])
b_conv1 = bias(0.1, shape=[1], name='b_conv1')
def CNN_1(x):
x = tf.image.per_image_standardization(x)
img = tf.reshape(x, [1, slider_size[0], slider_size[1], 1])
h_conv1 = tf.nn.relu(conv2d2(img, W_conv1) + b_conv1, name='h_conv1')
return h_conv1
inputs = tf.map_fn(
lambda seq: tf.map_fn(
lambda img:
tf.reshape(
CNN_1(tf.reshape(img, [slider_size[0], slider_size[1], 1])), [-1]),
seq),
encoder_inputs,
dtype=tf.float32)
# inputs = encoder_inputs
# Bidirectional RNN, gibe fw and bw outputs separately
enc_outputs, enc_state = tf.nn.bidirectional_dynamic_rnn(
cell_fw = enc_cell_fw,
cell_bw = enc_cell_bw,
inputs = inputs,
sequence_length = encoder_inputs_length,
dtype = tf.float32,
time_major = True)
encoder_outputs = tf.concat(enc_outputs, -1)
if encoder_layers == 1:
encoder_state = enc_state
else:
encoder_state = []
for layer_id in range(encoder_layers):
encoder_state.append(enc_state[0][layer_id]) # forward
encoder_state.append(enc_state[1][layer_id]) # backward
encoder_state = tuple(encoder_state)
```
## Decoder
```
# attention_states: size [batch_size, max_time, num_units]
attention_states = tf.transpose(encoder_outputs, [1, 0, 2])
# Create an attention mechanism
attention_mechanism = seq2seq.BahdanauAttention(
attention_size, attention_states, # decoder_units instead of attention_size
memory_sequence_length=encoder_inputs_length)
decoder_cell = create_cell(decoder_units,
decoder_layers,
decoder_residual_layers,
is_dropout=True,
keep_prob=keep_prob)
decoder_cell = seq2seq.AttentionWrapper(
decoder_cell, attention_mechanism,
attention_layer_size=attention_size)
decoder_initial_state = decoder_cell.zero_state(batch_size, tf.float32).clone(
cell_state=encoder_state)
```
#### TRAIN DECODER
```
# Helper
helper = seq2seq.TrainingHelper(
letter_train_inputs, letter_targets_length)
# Decoder
projection_layer = layers_core.Dense(
letter_size, activation=tf.tanh, use_bias=True)
decoder = seq2seq.BasicDecoder(
decoder_cell, helper, decoder_initial_state,
output_layer=projection_layer)
# Dynamic decoding
outputs, final_context_state, _ = seq2seq.dynamic_decode(
decoder)
letter_logits_train = outputs.rnn_output
letter_prediction_train = outputs.sample_id
```
#### INFERENCE DECODER
```
### CNN ###
def conv2d(x, W, name=None):
return tf.nn.conv2d(x, W, strides=[1, 1, 1, 1], padding='SAME', name=name)
def max_pool_2x2(x, name=None):
return tf.nn.max_pool(x, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='SAME', name=name)
W_conv1_ = weights('W_conv1_', shape=[16, 16, 1, 4])
b_conv1_ = bias(0.1, shape=[4], name='b_conv1_')
W_conv2_ = weights('W_conv2_', shape=[5, 5, 4, 12])
b_conv2_ = bias(0.1, shape=[12], name='b_conv2_')
W_conv3_ = weights('W_conv3_', shape=[3, 3, 12, 20])
b_conv3_ = bias(0.1, shape=[20], name='b_conv3_')
W_fc1_ = weights('W_fc2_', shape=[8*8*20, char_size])
b_fc1_ = bias(0.1, shape=[char_size], name='b_fc2_')
def CNN_2(x, clas=True):
if clas:
b_size, seq_size, _ = tf.unstack(tf.shape(x))
else:
b_size, _ = tf.unstack(tf.shape(x))
imgs = tf.reshape(x, [-1, 64, 64, 1])
x_imgs = tf.map_fn(
lambda img: tf.image.per_image_standardization(img), imgs)
# 1. Layer - Convulation
h_conv1 = tf.nn.relu(conv2d(x_imgs, W_conv1_) + b_conv1_, name='h_conv1_')
# 2. Layer - Max Pool
h_pool1 = max_pool_2x2(h_conv1, name='h_pool1_')
# 3. Layer - Convulation
h_conv2 = tf.nn.relu(conv2d(h_pool1, W_conv2_) + b_conv2_, name='h_conv2_')
# Reshape filters into flat arraty
h_pool2 = max_pool_2x2(h_conv2, name='h_pool2_')
h_conv3 = tf.nn.relu(conv2d(h_pool2, W_conv3_) + b_conv3_, name='h_conv3_')
# Reshape filters into flat arraty
h_pool3 = max_pool_2x2(h_conv3, name='h_pool3_')
h_flat = tf.reshape(h_pool3, [-1, 8*8*20])
# 6. Dropout
h_flat_drop = tf.nn.dropout(h_flat, keep_prob)
# 7. Output layer
out = tf.matmul(h_flat_drop, W_fc1_) + b_fc1_
if clas:
return tf.reshape(out, [b_size, seq_size, -1])
else:
return tf.reshape(out, [b_size, -1])
# Helper without embedding, can add param: 'next_inputs_fn'
helper_infer = seq2seq.InferenceHelper(
sample_fn=(lambda x: x),
sample_shape=[letter_size],
sample_dtype=tf.float32,
start_inputs=tf.cast(tf.fill([batch_size, letter_size], LETTER_PAD), tf.float32), # PAD <- EOS, need flaot32
end_fn=(lambda sample_ids:
# tf.greater(sample_ids[:, 0], 0)))
tf.equal(tf.argmax(CNN_2(sample_ids, False), axis=-1, output_type=tf.int32), 1)))
decoder_infer = seq2seq.BasicDecoder(
decoder_cell, helper_infer, decoder_initial_state,
output_layer=projection_layer)
# Dynamic decoding
outputs_infer, final_context_state, final_seq_lengths = seq2seq.dynamic_decode(
decoder_infer,
impute_finished=True,
maximum_iterations=output_length)
letter_prediction_infer = tf.identity(outputs_infer.rnn_output, # sample_id
name='letter_prediction_infer')
```
## RNN
```
word_inputs = tf.cond(is_training,
lambda: letter_targets,
lambda: letter_prediction_infer)
word_inputs_length_ = tf.cond(is_training,
lambda: letter_targets_length,
lambda: final_seq_lengths)
word_inputs_length = word_inputs_length_ # tf.subtract(word_inputs_length_, 1)
# Input images CNN
word_outputs = CNN_2(word_inputs)
# Word RNN
# cell_RNN = create_cell(wordRNN_units,
# wordRNN_layers,
# wordRNN_residual_layers,
# is_dropout=True,
# keep_prob=keep_prob)
# word_outputs, _ = tf.nn.dynamic_rnn(
# cell = cell_RNN,
# inputs = word_inputs,
# sequence_length = word_inputs_length,
# dtype = tf.float32)
# word_logits = tf.layers.dense(
# inputs=word_outputs,
# units=char_size,
# name='pred')
word_logits = word_outputs
word_prediction = tf.argmax(
word_logits, axis=-1, output_type=tf.int32, # word_logits = tf.layers.dense(
name='word_prediction')
```
## Optimizer
#### Weights + Paddings
```
# Pad test accuracy
letter_test_targets = tf.pad(
letter_targets,
[[0, 0],
[0, output_length - tf.reduce_max(letter_targets_length)],
[0, 0]],
constant_values=LETTER_PAD,
mode='CONSTANT')
# Pad prediction to match lengths
letter_pred_infer_pad = tf.pad(
letter_prediction_infer,
[[0, 0],
[0, output_length - tf.reduce_max(word_inputs_length_)],
[0, 0]],
constant_values=LETTER_PAD,
mode='CONSTANT')
word_pad_lenght = tf.maximum(
tf.reduce_max(word_inputs_length_),
tf.reduce_max(word_targets_length))
word_logits_pad = tf.pad(
word_logits,
[[0, 0],
[0, word_pad_lenght - tf.reduce_max(word_inputs_length_)],
[0, 0]],
constant_values=PAD,
mode='CONSTANT')
word_pred_pad = tf.pad(
word_prediction,
[[0, 0],
[0, word_pad_lenght - tf.reduce_max(word_inputs_length_)]],
constant_values=PAD,
mode='CONSTANT')
word_targets_pad = tf.pad(
word_targets,
[[0, 0],
[0, word_pad_lenght - tf.reduce_max(word_targets_length)]],
constant_values=PAD,
mode='CONSTANT')
# Weights
letter_loss_weights = tf.sequence_mask(
letter_train_length,
tf.reduce_max(letter_train_length),
dtype=tf.float32)
letter_test_weights = tf.sequence_mask(
letter_train_length,
output_length,
dtype=tf.float32)
word_loss_weights = tf.sequence_mask(
word_targets_length, # word_inputs_length, try max(targets, inputs)
word_pad_lenght,
dtype=tf.float32)
word_acc_weights = tf.sequence_mask(
tf.subtract(final_seq_lengths, 1), # word_inputs_length, try max(targets, inputs)
word_pad_lenght,
dtype=tf.float32)
## Loss
letter_loss = tf.losses.mean_squared_error(
predictions=letter_logits_train,
labels=letter_targets,
weights=tf.stack([letter_loss_weights for i in range(letter_size)], axis=-1))
word_seq_loss = seq2seq.sequence_loss(
logits=word_logits_pad,
targets=word_targets_pad,
weights=word_loss_weights,
name='word_loss')
regularization = tf.get_collection(tf.GraphKeys.REGULARIZATION_LOSSES)
word_loss = word_seq_loss + sum(regularization)
loss = tf.cond(is_words,
lambda: word_loss,
lambda: letter_loss)
# learning_rate_ = learning_rate
learning_rate_ = tf.cond(tf.logical_and(is_words, tf.logical_not(is_training)),
lambda: learning_rate, # * 0.1,
lambda: learning_rate)
## Calculate and clip gradients
params = tf.trainable_variables()
gradients = tf.gradients(loss, params)
clipped_gradients, _ = tf.clip_by_global_norm(
gradients, max_gradient_norm)
### Optimization
optimizer = tf.train.AdamOptimizer(learning_rate_)
train_step = optimizer.apply_gradients(
zip(clipped_gradients, params),
name='train_step')
### Evaluate model
correct_prediction = tf.equal(
word_pred_pad,
word_targets_pad)
## Advanced accuracy only the elements of seq including EOS symbol
# accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
accuracy = (tf.reduce_sum(tf.cast(correct_prediction, tf.float32) * word_acc_weights) \
/ tf.reduce_sum(word_acc_weights))
sess.run(tf.global_variables_initializer())
fd = test_iterator.next_feed(3)
pre = word_logits.eval(fd)
print(pre.shape)
```
## Training
```
sess.run(tf.global_variables_initializer())
saver = tf.train.Saver()
# Creat plot for live stats ploting
trainPlot = TrainingPlot(TRAIN_STEPS, TEST_ITER, LOSS_ITER)
try:
for i_batch in range(TRAIN_STEPS):
# Three steps (can overflow): 1. train letters, 2. train words on known letter 3. combine
is_words_, is_train_ = (True, False) # (True, True) - without partial sep letters
if i_batch < 1000 or i_batch % 2 == 0: # 2000 # 5000
is_words_, is_train_ = (False, True)
# elif i_batch < 3000: # 12000
# is_words_, is_train_ = (True, True)
elif i_batch % 4 == 3:
is_words_, is_train_ = (True, False)
fd = train_iterator.next_feed(BATCH_SIZE, words=is_words_, train=is_train_)
if is_train_:
train_step.run(train_letters_iterator.next_feed(BATCH_SIZE, words=is_words_, train=is_train_))
else:
train_step.run(fd)
if i_batch % LOSS_ITER == 0:
# Plotting loss
tmpLoss = loss.eval(fd)
trainPlot.updateCost(tmpLoss, i_batch // LOSS_ITER)
if i_batch % TEST_ITER == 0:
# Plotting accuracy
fd_test = test_iterator.next_feed(BATCH_SIZE)
fd = train_iterator.next_feed(BATCH_SIZE, words=True, train=False)
accTest = accuracy.eval(fd_test)
accTrain = accuracy.eval(fd)
trainPlot.updateAcc(accTest, accTrain, i_batch // TEST_ITER)
if i_batch % SAVE_ITER == 0:
saver.save(sess, save_location)
if i_batch % EPOCH == 0:
fd_test = test_iterator.next_feed(BATCH_SIZE)
print('batch %r - loss: %r' % (i_batch, sess.run(loss, fd_test)))
predict_, target_ = sess.run([word_prediction, word_targets], fd_test)
for i, (inp, pred) in enumerate(zip(target_, predict_)):
print(' expected > {}'.format(inp))
print(' predicted > {}'.format(pred))
if i >= 1:
break
print()
except KeyboardInterrupt:
print('Training interrupted, model saved.')
saver.save(sess, save_location)
%matplotlib inline
def evalLetters(feed):
predict_, target_, predict_lengths_, target_lengths_ = sess.run(
[letter_prediction_infer,
letter_targets,
final_seq_lengths,
letter_targets_length],
feed)
for i, (inp, pred) in enumerate(zip(target_, predict_)):
print("Expected images:", target_lengths_[i])
for x in range(len(inp)):
implt(inp[x].reshape((64, 64)), 'gray')
print("Predicted images:", predict_lengths_[i])
for x in range(len(pred)):
implt(pred[x].reshape((64, 64)), 'gray')
if i >= 0:
break
fd_test = test_iterator.next_feed(BATCH_SIZE)
fd = train_iterator.next_feed(BATCH_SIZE, words=False, train=False)
evalLetters(fd_test)
evalLetters(fd)
```
| github_jupyter |
# Visualizing CNN Layers
---
In this notebook, we load a trained CNN (from a solution to FashionMNIST) and implement several feature visualization techniques to see what features this network has learned to extract.
### Load the [data](http://pytorch.org/docs/master/torchvision/datasets.html)
In this cell, we load in just the **test** dataset from the FashionMNIST class.
```
# our basic libraries
import torch
import torchvision
# data loading and transforming
from torchvision.datasets import FashionMNIST
from torch.utils.data import DataLoader
from torchvision import transforms
# The output of torchvision datasets are PILImage images of range [0, 1].
# We transform them to Tensors for input into a CNN
## Define a transform to read the data in as a tensor
data_transform = transforms.ToTensor()
test_data = FashionMNIST(root='./data', train=False,
download=True, transform=data_transform)
# Print out some stats about the test data
print('Test data, number of images: ', len(test_data))
# prepare data loaders, set the batch_size
## TODO: you can try changing the batch_size to be larger or smaller
## when you get to training your network, see how batch_size affects the loss
batch_size = 20
test_loader = DataLoader(test_data, batch_size=batch_size, shuffle=True)
# specify the image classes
classes = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
```
### Visualize some test data
This cell iterates over the training dataset, loading a random batch of image/label data, using `dataiter.next()`. It then plots the batch of images and labels in a `2 x batch_size/2` grid.
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
# obtain one batch of training images
dataiter = iter(test_loader)
images, labels = dataiter.next()
images = images.numpy()
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(batch_size):
ax = fig.add_subplot(2, batch_size/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
ax.set_title(classes[labels[idx]])
```
### Define the network architecture
The various layers that make up any neural network are documented, [here](http://pytorch.org/docs/master/nn.html). For a convolutional neural network, we'll use a simple series of layers:
* Convolutional layers
* Maxpooling layers
* Fully-connected (linear) layers
```
import torch.nn as nn
import torch.nn.functional as F
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# 1 input image channel (grayscale), 10 output channels/feature maps
# 3x3 square convolution kernel
## output size = (W-F)/S +1 = (28-3)/1 +1 = 26
# the output Tensor for one image, will have the dimensions: (10, 26, 26)
# after one pool layer, this becomes (10, 13, 13)
self.conv1 = nn.Conv2d(1, 10, 3)
# maxpool layer
# pool with kernel_size=2, stride=2
self.pool = nn.MaxPool2d(2, 2)
# second conv layer: 10 inputs, 20 outputs, 3x3 conv
## output size = (W-F)/S +1 = (13-3)/1 +1 = 11
# the output tensor will have dimensions: (20, 11, 11)
# after another pool layer this becomes (20, 5, 5); 5.5 is rounded down
self.conv2 = nn.Conv2d(10, 20, 3)
# 20 outputs * the 5*5 filtered/pooled map size
self.fc1 = nn.Linear(20*5*5, 50)
# dropout with p=0.4
self.fc1_drop = nn.Dropout(p=0.4)
# finally, create 10 output channels (for the 10 classes)
self.fc2 = nn.Linear(50, 10)
# define the feedforward behavior
def forward(self, x):
# two conv/relu + pool layers
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
# prep for linear layer
# this line of code is the equivalent of Flatten in Keras
x = x.view(x.size(0), -1)
# two linear layers with dropout in between
x = F.relu(self.fc1(x))
x = self.fc1_drop(x)
x = self.fc2(x)
# final output
return x
```
### Load in our trained net
This notebook needs to know the network architecture, as defined above, and once it knows what the "Net" class looks like, we can instantiate a model and load in an already trained network.
The architecture above is taken from the example solution code, which was trained and saved in the directory `saved_models/`.
```
# instantiate your Net
net = Net()
# load the net parameters by name
net.load_state_dict(torch.load('saved_models/fashion_net_ex.pt'))
print(net)
```
## Feature Visualization
Sometimes, neural networks are thought of as a black box, given some input, they learn to produce some output. CNN's are actually learning to recognize a variety of spatial patterns and you can visualize what each convolutional layer has been trained to recognize by looking at the weights that make up each convolutional kernel and applying those one at a time to a sample image. These techniques are called feature visualization and they are useful for understanding the inner workings of a CNN.
In the cell below, you'll see how to extract and visualize the filter weights for all of the filters in the first convolutional layer.
Note the patterns of light and dark pixels and see if you can tell what a particular filter is detecting. For example, the filter pictured in the example below has dark pixels on either side and light pixels in the middle column, and so it may be detecting vertical edges.
<img src='images/edge_filter_ex.png' width= 30% height=30%/>
```
# Get the weights in the first conv layer
weights = net.conv1.weight.data
w = weights.numpy()
# for 10 filters
fig=plt.figure(figsize=(20, 8))
columns = 5
rows = 2
for i in range(0, columns*rows):
fig.add_subplot(rows, columns, i+1)
plt.imshow(w[i][0], cmap='gray')
print('First convolutional layer')
plt.show()
weights = net.conv2.weight.data
w = weights.numpy()
```
### Activation Maps
Next, you'll see how to use OpenCV's `filter2D` function to apply these filters to a sample test image and produce a series of **activation maps** as a result. We'll do this for the first and second convolutional layers and these activation maps whould really give you a sense for what features each filter learns to extract.
```
# obtain one batch of testing images
dataiter = iter(test_loader)
images, labels = dataiter.next()
images = images.numpy()
# select an image by index
idx = 10
img = np.squeeze(images[idx])
# Use OpenCV's filter2D function
# apply a specific set of filter weights (like the one's displayed above) to the test image
import cv2
plt.imshow(img, cmap='gray')
weights = net.conv1.weight.data
w = weights.numpy()
# 1. first conv layer
# for 10 filters
fig=plt.figure(figsize=(30, 10))
columns = 5*2
rows = 2
for i in range(0, columns*rows):
fig.add_subplot(rows, columns, i+1)
if ((i%2)==0):
plt.imshow(w[int(i/2)][0], cmap='gray')
else:
c = cv2.filter2D(img, -1, w[int((i-1)/2)][0])
plt.imshow(c, cmap='gray')
plt.show()
# Same process but for the second conv layer (20, 3x3 filters):
plt.imshow(img, cmap='gray')
# second conv layer, conv2
weights = net.conv2.weight.data
w = weights.numpy()
# 1. first conv layer
# for 20 filters
fig=plt.figure(figsize=(30, 10))
columns = 5*2
rows = 2*2
for i in range(0, columns*rows):
fig.add_subplot(rows, columns, i+1)
if ((i%2)==0):
plt.imshow(w[int(i/2)][0], cmap='gray')
else:
c = cv2.filter2D(img, -1, w[int((i-1)/2)][0])
plt.imshow(c, cmap='gray')
plt.show()
```
### Question: Choose a filter from one of your trained convolutional layers; looking at these activations, what purpose do you think it plays? What kind of feature do you think it detects?
**Answer**: In the first convolutional layer (conv1), the sixth filter, pictured as the first filter in the second row of the conv1 filters, appears to detect vertical/slightly-left leaning edges. It has a positively weighted left-most column and seems to detect the vertical edges of sleeves in a pullover.
| github_jupyter |
```
import arviz as az
import pymc as pm
# import arviz.labels as azl
import matplotlib.pyplot as plt
import seaborn as sns
from pathlib import Path
import pickle
import matplotlib as mpl
import warnings
import numpy as np
warnings.simplefilter(action='ignore', category=FutureWarning)
import pandas as pd
```
## Reestimate model because of pickle problems
```
def gamma(alpha, beta):
def g(x):
return pm.Gamma(x, alpha=alpha, beta=beta)
return g
def hcauchy(beta):
def g(x):
return pm.HalfCauchy(x, beta=beta)
return g
def fit_gp(y, X, l_prior, eta_prior, sigma_prior, kernel_type='M52', bayes_kws=dict(draws=1000, tune=1000, chains=2, cores=1), prop_Xu=None):
"""
function to return a pymc3 model
y : dependent variable
X : independent variables
prop_Xu : number of inducing varibles to use. If None, use full marginal likelihood. If not none, use FTIC.
bayes_kw : kws for pm.sample
X, y are dataframes. We'll use the column names.
"""
kernel_type = kernel_type.lower()
with pm.Model() as model:
# Covert arrays
X_a = X.values
y_a = y.values.flatten()
X_cols = list(X.columns)
# Kernels
# 3 way interaction
eta = eta_prior('eta')
cov = eta**2
for i in range(X_a.shape[1]):
var_lab = 'l_'+X_cols[i]
if kernel_type=='rbf':
cov = cov*pm.gp.cov.ExpQuad(X_a.shape[1], ls=l_prior(var_lab), active_dims=[i])
if kernel_type=='exponential':
cov = cov*pm.gp.cov.Exponential(X_a.shape[1], ls=l_prior(var_lab), active_dims=[i])
if kernel_type=='m52':
cov = cov*pm.gp.cov.Matern52(X_a.shape[1], ls=l_prior(var_lab), active_dims=[i])
if kernel_type=='m32':
cov = cov*pm.gp.cov.Matern32(X_a.shape[1], ls=l_prior(var_lab), active_dims=[i])
# Covariance model
cov_tot = cov
# Noise model
sigma_n =sigma_prior('sigma_n')
# Model
if not (prop_Xu is None):
# Inducing variables
num_Xu = int(X_a.shape[0]*prop_Xu)
Xu = pm.gp.util.kmeans_inducing_points(num_Xu, X_a)
gp = pm.gp.MarginalSparse(cov_func=cov_tot, approx="FITC")
y_ = gp.marginal_likelihood('y_', X=X_a, y=y_a, Xu=Xu, noise=sigma_n)
else:
gp = pm.gp.Marginal(cov_func=cov_tot)
y_ = gp.marginal_likelihood('y_', X=X_a, y=y_a, noise=sigma_n)
if not (bayes_kws is None):
trace = pm.sample(**bayes_kws)
result = trace
else:
mp = pm.find_MAP(progressbar=False)
result = mp
return gp, result, model
def bootstrap(n_samples,y, X, seed=42, **kwargs):
rng = np.random.default_rng(seed)
all_params = []
all_preds = []
i = 0
while i < n_samples:
print(' ', i, end=', ')
ix = np.array(y.index)
bs_ix = rng.choice(ix, size=len(ix), replace=True)
new_y, new_X = y.loc[bs_ix, :], X.loc[bs_ix, :]
try:
gp, mp, model = fit_gp(new_y, new_X, **kwargs)
all_params.append(mp)
with model:
y_pred, var = gp.predict(new_X.values, point=mp, diag=True)
all_preds.append((new_y, y_pred, var))
i += 1
except:
print('error')
print()
return all_params, all_preds
root_path = '1fme/sensitivity'
experiments = [['dihedrals', None, 'exponential'], ['distances', 'linear', 'exponential'], ['distances', 'logistic', 'exponential']]
models = []
l_prior = gamma(2, 0.5)
eta_prior = hcauchy(2)
sigma_prior = hcauchy(2)
params = []
preds = []
n_boot = 250
for i, (feat, trans, kernel) in enumerate(experiments):
print(feat, trans, kernel)
kwargs = dict(l_prior=l_prior, eta_prior=eta_prior, sigma_prior=sigma_prior, # Priors
kernel_type=kernel, # Kernel
prop_Xu=None, # proportion of data points which are inducing variables.
bayes_kws=None)
results_path = Path(root_path).joinpath(f"{feat}_{trans}_{kernel}_mml.pkl")
results = pickle.load(results_path.open('rb'))
data_s = results['data']
y, X = data_s.iloc[:, [0]], data_s.iloc[:, 1:]
param, pred = bootstrap(n_boot, y, X, **kwargs)
params.append(param)
preds.append(pred)
param_df = []
keep_params = ['l_dim', 'l_lag', 'l_states', 'feature', 'l_cent', 'l_steep']
for i, experiment in enumerate(experiments):
bs_params = params[i]
lab = experiment[0]
if experiment[1] is not None:
lab += f"-{experiment[1]}"
df = pd.concat([pd.DataFrame(x, index=[j]) for j, x in enumerate(bs_params)])
df['feature'] = lab
param_df.append(df)
param_df = pd.concat(param_df)
param_df = param_df.loc[:, keep_params]
param_df_m = param_df.melt(id_vars=['feature'], var_name='hyperparameter')
param_df_m['R'] = 1.0/param_df_m['value']
var_name_map={"l_states": r"Num. states",
"l_dim": r"Num. dims",
"l_lag": r"$\tau_{tICA}$",
"l_cent": r"Centre",
"l_steep": r"Steepness"}
param_df_m['hyperparameter'] = param_df_m['hyperparameter'].apply(lambda x: var_name_map[x])
```
Sensitivity
```
with sns.plotting_context('paper', font_scale=1.5):
fig, ax = plt.subplots(1)
sns.boxplot(y='feature', x='R', hue='hyperparameter', data=param_df_m,ax=ax,
whis=(0, 100), orient='h', linewidth=0.5)
h, l = ax.get_legend_handles_labels()
ax.legend(h, l, loc='upper left', bbox_to_anchor=(1, 1))
ax.set_xlim(0.2, 10)
ax.set_xscale('log')
plt.savefig(Path(root_path).parent.joinpath('sensitivity.pdf'), bbox_inches='tight')
```
| github_jupyter |
# 目次
1. 機械学習の例
- ここではシンプルなLinear Regressionを例に出します
2. アニーリングに移行してみる
- 重みをバイナリ変数で扱う方法を紹介します
# 機械学習の例 | Linear Regression
Linear Regression(線型回帰)は確率を回帰する機械学習モデルです。$x$が入力、$Y$を出力とし、パラメータ$\begin{pmatrix}\beta_0 \\ \vdots \\ \beta_k \end{pmatrix}$を学習させます。
線型回帰のモデルは以下のような式で定義されます。
$$
Y = \beta_0 x_0 + \cdots + \beta_k x_k = \sum_{i=0}^{k} \beta_{i} x_i
$$
```
import numpy as np
import pandas as pd
from sklearn.linear_model import LinearRegression
from sklearn.metrics import roc_auc_score, accuracy_score, mean_squared_error
```
そしたらLinear Regressionを使って学習を行ってみようと思います。
`calc_y`関数で定義されるように、各特徴量に対して1か0の重みを掛け、それらの和をtargetとして与えました。
回帰係数が[1, 0, 1, 1]になったら完璧ですね。
```
def calc_y(x):
output = x[0] * 1 + x[1] * 0 + x[2] * 1 + x[3] * 1
# output = 1 if res >= 2
return output
nrows = 100
nfea = 4
# nrows x nfeaの行列を作る
df = np.random.randint(2, size=(nrows, nfea)).astype(bool)
target = []
for i in range(nrows):
t = int(calc_y(df[i, :]))
target.append(t)
df = pd.DataFrame(df).astype(int)
df["target"] = target
df.head()
```
`sklearn`の`LinearRegression`モジュールで学習を行います。特にパラメータの指定は行いません。
```
lr = LinearRegression()
lr.fit(df.iloc[:, 0:4], df.iloc[:, 4])
```
回帰係数は`calc_y`関数で定義した通りと、意図した結果が得られました。
```
# 回帰係数の確認
lr.coef_
```
# アニーリングに移行してみる
どうやってやるか、端的に説明すると「重みをバイナリ変数にし、二乗誤差をエネルギーとして定式化する」になります。
## 重みをバイナリ変数にする
バイナリ変数は0-1の二値をとる変数です。
$$
\hat{y} = \beta_0 x_0 + \cdots + \beta_k x_k
$$
↑これが ↓こう ($q$は$\{0, 1\}$を取るバイナリ変数です)
$$
\hat{y} = q_0 x_0 + \cdots + q_k x_k
$$
線型回帰の重みは任意の実数がとれますが、バイナリ変数を使うことで二値しかとれないことがわかります。
### 二値じゃ全然役に立たないのでは?
ここで、バイナリ変数にすることで生じる課題を解決する方法を紹介します。しかし、これからの解説では上に書いた0と1の二値の重みを使用するので、興味がある人が読んでください
#### 逆量子化 | より多くの整数値をとる
例えば下のような式で$q_0$を定義したらどうでしょうか。$q_0$は0から10の11個の整数値をとることができます。
$$
q_0 = \sum_{i=0}^9 q_{0, i}
$$
実数の値を離散値に丸めることを離散値といいますが、これは離散値をたくさん用意することで実数の値に近づける"逆量子化"を行っています。ちなみに量子化という言葉は学術的に使用されているワードですが、逆量子化は私が命名した言葉なので、この分野における正しいワードかはわかりません。
#### スケーリング | 整数値だけでなく、小数も扱えるように
逆量子化では整数値しかとれませんでしたが、各バイナリ変数に対して小数の重み付けをしてあげることで"全体として"小数の数を扱うことができます。
ex. 1
逆量子化の例に出した0 ~ 10の重みの範囲を0 ~ 1.0にする
$$
q_0 = \sum_{i=0}^9 0.1 q_{0, i}
$$
ex. 2
重み$\begin{pmatrix}w_0 \\ \vdots \\ w_k \end{pmatrix}$を使う。一般化させただけ。
$$
q_0 = \sum_{i=0}^{k} w_i q_{0, i}
$$
## 二乗誤差で定式化する
誤差関数を二乗誤差としたらアニーリングにおけるエネルギーと誤差関数を同一視することができます。
ターゲット$y$に対して予測値を$\hat{y}$としたとき、エネルギー(誤差関数)$H$は以下のように与えられます。
$$
H = (y - \hat{y}) ^2
$$
$\hat{y}$は`重みをバイナリ変数にする`チャプターで定義したので、それを代入します。
$$
H = (y - (q_0 x_0 + \cdots + q_k x_k))^2 = (y - \sum_0^k q_i x_i)^2
$$
## 実装してみる
先程、様々な小数の値をとることのできる逆量子化を紹介しましたが、今回の重みは0か1をとる1つのバイナリ変数のみで実装します。
```
from amplify import IsingPoly, IsingMatrix, gen_symbols, sum_poly, BinaryPoly
from amplify import Solver, decode_solution, BinarySymbolGenerator
from amplify.client import FixstarsClient
import json
```
自分はローカルのファイルからトークンを読み込んでいますが、動かすときには`token読み込み`を消して以下ようにベタ書きしてください。
余談ですが、ローカルから読み込むようにすればトークンの流出が防げたり、トークンを更新してもコードを変える必要がなくなったりします。
```python
client.token = "token"
```
```
client = FixstarsClient()
"""token読み込み はじまり"""
token_path = "/home/yuma/.amplify/token.json"
with open(token_path) as f:
client.token = json.load(f)["AMPLIFY_TOKEN"]
"""token読み込み おわり"""
client.parameters.timeout = 100
solver = Solver(client)
```
学習させる4つのバイナリ変数(0か1をとるイジング変数)を用意します。
```
gen = BinarySymbolGenerator()
q = gen.array(nfea)
q
```
エネルギーの定式化を行います。各行のデータに対して二乗誤差を計算し、エネルギーに足していきます。
```
f = 0
for i in range(nrows):
# amplifyのドキュメントでは標準のsum関数は推奨されていません。
# イテレートする回数が多い時はamplifyのsum_poly関数を使いましょう。
y_hat = sum(
q[j] * df.iloc[i, j]
for j in range(nfea)
)
# 二乗誤差
f += (df.iloc[i, 4] - y_hat) ** 2
if i == 0:
print(f"{'dfの1行目':-^30}")
print(df.iloc[[0], :])
print(f"{'y_hat, y':-^30}")
print(f"{y_hat}, {df.iloc[0, 4]}")
print(f"{'エネルギー':-^30}")
print(f"{f}")
```
定式化は完了したので、Amplify AEに計算式を投げて解きます。
```
result = solver.solve(f)
binary_weight = q.decode(result[0].values)
binary_weight
```
上の解のエネルギー(誤差関数)はいくらでしょうか! 0です!! 最強のAIができました。
これはyを定義した`calc_y`の定義式、Linear Regressionの解とも一致しています。
```
result[0].energy
```
以上を通してアニーリングを用いて機械学習を行うことができます。
今回は限定的な線型回帰でしたが、この考え方を使うことで幅広い機械学習モデルを実装することが可能になります!
| github_jupyter |
# The TxTl Toolbox in BioCRNpyler
### A recreation of the original MATLAB TxTl Toolbox, as seen in [Singhal et al. 2020](https://www.biorxiv.org/content/10.1101/2020.08.05.237990v1)
This tutorial shows how to use the EnergyTxTlExtract Mixture with a parameter file derived from the paper above. This Mixture is a simplification of the models used in the original toolbox. Notable changes include:
1. Using only a single nucleotide species NTPs (instead of GTP, ATP, UTP, and CTP)
2. A slightly different NTP regeneration Mechanism which explicitly incorporates the amount of fuel, 3PGA, put into the extract and metabolic leak of the extract.
3. Degredation of RNA bound to ribosomes (which releases the ribosome).
4. A modification of the Energy consumption reactions for Transcription and Translation so that there is only a single binding reaction.
## The CRN displayed below shows the energy utilization process model
```
from biocrnpyler import *
#A = DNAassembly("A", promoter = "P", rbs = "rbs")
E = EnergyTxTlExtract(parameter_file = "txtl_toolbox_parameters.txt")
CRN = E.compile_crn()
print(CRN.pretty_print())
try:
import numpy as np
maxtime = 30000
timepoints = np.arange(0, maxtime, 100)
R = CRN.simulate_with_bioscrape_via_sbml(timepoints)
if R is not None:
%matplotlib inline
import pylab as plt
plt.plot(timepoints, R[str(E.ntps.get_species())], label = E.ntps.get_species())
plt.plot(timepoints, R[str(E.amino_acids.get_species())], label = E.amino_acids.get_species())
plt.plot(timepoints, R[str(E.fuel.get_species())], label = E.fuel.get_species())
plt.xticks(np.arange(0, maxtime, 3600), [str(i) for i in range(0, int(np.ceil(maxtime/3600)))])
plt.legend()
except ModuleNotFoundError:
print('please install the plotting libraries: pip install biocrnpyler[all]')
```
## Adding a DNA assembly
This will produce protein expression, but for a limited time. The
```
A = DNAassembly("A", promoter = "P", rbs = "rbs", initial_concentration = 1*10**-6)
E = EnergyTxTlExtract(components = [A], parameter_file = "txtl_toolbox_parameters.txt")
CRN = E.compile_crn()
print(CRN.pretty_print())
try:
maxtime = 30000
timepoints = np.arange(0, maxtime, 100)
R = CRN.simulate_with_bioscrape_via_sbml(timepoints)
if R is not None:
%matplotlib inline
plt.subplot(121)
plt.plot(timepoints, R[str(E.ntps.get_species())], label = E.ntps.get_species())
plt.plot(timepoints, R[str(E.amino_acids.get_species())], label = E.amino_acids.get_species())
plt.plot(timepoints, R[str(E.fuel.get_species())], label = E.fuel.get_species())
plt.xticks(np.arange(0, maxtime, 3600), [str(i) for i in range(0, int(np.ceil(maxtime/3600)))])
plt.legend()
plt.subplot(122)
plt.plot(timepoints, R[str(A.transcript)], label = A.transcript)
plt.plot(timepoints, R[str(A.protein)], label = A.protein)
plt.xticks(np.arange(0, maxtime, 3600), [str(i) for i in range(0, int(np.ceil(maxtime/3600)))])
plt.legend()
except ModuleNotFoundError:
print('please install the plotting libraries: pip install biocrnpyler[all]')
```
| github_jupyter |
# Excercises Electric Machinery Fundamentals
## Chapter 2
## Problem 2-4
```
%pylab inline
```
### Description
The secondary winding of a real transformer has a terminal voltage of
* $v_S(t) = \sqrt{2}\cdot 200 \sin(2\pi\cdot 60\,Hz\cdot t)\,V = 282.8 \sin(377t)\,V$.
The turns ratio of the transformer is
* $100:200 \rightarrow a = 0.5$ .
If the secondary current of the transformer is
* $i_S(t) = \sqrt{2}\cdot 5 \sin(377t - 36.87°)\, A$.
1. What is the primary current of this transformer?
2. What are its voltage regulation and efficiency?
The impedances of this transformer referred to the primary side are:
$$R_{eq} = 0.20\,\Omega \qquad R_{C} = 300\,\Omega$$
$$X_{eq} = 0.80\,\Omega \qquad X_{M} = 100\,\Omega$$
```
VS_m = sqrt(2) * 200 # [V]
VS_angle = 0 # [rad]|[deg]
IS_m = sqrt(2)*5 # [A]
IS_deg = -36.87 # angle of IS in [deg]
IS_rad = IS_deg/180*pi # angle of IS in [rad]
a = 0.5
Req = 0.20 # [Ohm]
Rc = 300.0 # [Ohm]
Xeq = 0.80 # [Ohm]
Xm = 100.0 # [Ohm]
```
### SOLUTION
The equivalent circuit of this transformer is shown below. (Since no particular equivalent circuit was specified, we are using the approximate equivalent circuit referred to the primary side.)
<img src="figs/FigC_2-18a.jpg" width="70%">
The secondary voltage and current are:
```
VS = VS_m / sqrt(2) * (cos(VS_angle) + sin(VS_angle)*1j) # [V]
IS = IS_m / sqrt(2) * (cos(IS_rad) + sin(IS_rad)*1j) # [A]
print('VS = {:>5.1f} V ∠{:>3.0f}°'.format(
abs(VS), VS_angle))
print('IS = {:>5.1f} A ∠{:>3.2f}°'.format(
abs(IS), (IS_deg)))
```
The secondary voltage referred to the primary side is:
$$\vec{V}'_S = a\vec{V}_S$$
```
VSp = a * VS
print('VSp = {:.1f} V ∠{:.0f}°'.format(
abs(VSp), VS_angle)) # the turns ratio has no effect on the angle
```
The secondary current referred to the primary side is:
$$\vec{I}'_S = \frac{\vec{I}_S}{a}$$
```
ISp = IS / a
print('ISp = {:.1f} A ∠{:.2f}°'.format(
abs(ISp), IS_deg)) # the turns ratio has no effect on the angle
```
The primary circuit voltage is given by:
$$\vec{V}_P = \vec{V}'_S + \vec{I}'_S(R_{eq} + jX_{eq})$$
```
VP = VSp + ISp * (Req + Xeq*1j)
VP_angle = arctan(VP.imag/VP.real)
print('VP = {:.1f} V ∠{:.1f}°'.format(
abs(VP), degrees(VP_angle)))
```
The excitation current of this transformer is:
$$\vec{I}_{EX} = \vec{I}_C + \vec{I}_M$$
```
IC = VP / Rc
IM = VP / (Xm*1j)
Iex = IC + IM
Iex_angle = arctan(Iex.imag/Iex.real)
print('Iex = {:.2f} A ∠{:.1f}°'.format(
abs(Iex), Iex_angle/pi*180)) # angle in [deg]
```
**Therefore, the total primary current of this transformer is:**
$$\vec{I}_P = \vec{I}'_S + \vec{I}_{EX}$$
```
IP = ISp + Iex
IP_angle = arctan(IP.imag/IP.real)
print('VS = {:.1f} V ∠{:.1f}°'.format(
abs(IP), IP_angle/pi*180)) # angle in [deg]
print('===================')
```
**The voltage regulation of the transformer at this load is:**
$$VR = \frac{V_P - aV_S}{aV_S} \cdot 100\%$$
```
VR = (abs(VP) - a*abs(VS))/(a*abs(VS)) * 100
print('VR = {:.1f}%'.format(VR))
print('=========')
```
The input power to this transformer is:
$$P_{IN} = V_PI_P\cos\theta$$
```
Pin = abs(VP) * abs(IP) * cos(VP_angle - IP_angle)
print('Pin = {:.0f} W'.format(Pin))
```
The output power from this transformer is:
$$P_{OUT} = V_SI_S\cos\theta$$
```
Pout = abs(VS) * abs(IS) * cos(VS_angle - IS_rad)
print('Pout = {:.0f} W'.format(Pout))
```
**Therefore, the transformer's efficiency is:**
$$\eta = \frac{P_{OUT}}{P_{IN}} \cdot 100\,\%$$
```
eta = Pout / Pin * 100
print('η = {:.1f} %'.format(eta))
print('==========')
```
| github_jupyter |
# Lesson 6: Practice with Pandas and Bokeh
## Practice 1: Axes with logarithmic scale
Sometimes you need to plot your data with a logarithmic scale. As an example, let's consider the classic genetic switch engineered by Jim Collins and coworkers ([Gardner, et al., *Nature*, **403**, 339, 2000](http://www.nature.com/nature/journal/v403/n6767/full/403339a0.html)). This genetic switch was incorporated into *E. coli* and is inducible by adjusting the concentration of the lactose analog IPTG. The readout is the fluorescence intensity of GFP.
Let's load in some data that have the IPTG concentrations and GFP fluorescence intensity. The data are in the file `~/git/data/collins_switch.csv`. Let's look at it.
```
!cat Dataset/collins_switch.csv
```
It has two rows of non-data. Then, Column 1 is the IPTG concentration, column 2 is the normalized GFP expression level, and the last column is the standard error of the mean normalized GFP intensity. This gives the error bars, which we will look at in the next exercise. For now, we will just plot IPTG versus normalized GFP intensity.
In looking at the data set, note that there are two entries for [IPTG] = 0.04 mM. At this concentration, the switch happens, and there are two populations of cells, one with high expression of GFP and one with low. The two data points represent these two populations of cells.
Now, let's make a plot of IPTG versus GFP.
>1. Load in the data set using Pandas. Make sure you use the `comment` kwarg of pd.read_csv() properly.
2. Make a plot of normalized GFP intensity (y-axis) versus IPTG concentration (x-axis).
Now that you have done that, there are some problems with the plot. It is really hard to see the data points with low concentrations of IPTG. In fact, looking at the data set, the concentration of IPTG varies over four orders of magnitude. When you have data like this, it is wise to plot them on a logarithmic scale. You can specify the x-axis as logarithmic when you instantiate a figure with `bokeh.plotting.figure()` by using the `x_axis_type='log'` kwarg. (The obvious analogous kwarg applied for the y-axis.) For this data set, it is definitely best to have the x-axis on a logarithmic scale. Remake the plot you just did with the x-axis logarithmically scaled.
When you make the x-axis logarithmically scaled, you will notice the Bokeh's formatting for the tick labels is pretty awful. Fixing this is a surprisingly difficult problem, and many plotting packages do not make pretty superscripts.
<br />
## Practice 2: Plots with error bars
The data set also contains the standard error of the mean, or SEM. The SEM is often displayed on plots as error bars. Now construct the plot with error bars.
>1. Add columns `error_low` and `error_high` to the `DataFrame` containing the Collins data. These will set the bottoms and tops of the error bars. You should base the values in these columns on the standard error of the mean (`sem`). Assuming a Gaussian model, the 95% confidence interval is ±1.96 times the s.e.m.
2. Make a plot with the measured expression levels and the error bars. *Hint*: Check out the [Bokeh docs](https://bokeh.pydata.org/en/latest/docs/user_guide/plotting.html) and think about what kind of glyph works best for error bars.
## Computing environment
```
%load_ext watermark
%watermark -v -p jupyterlab
```
| github_jupyter |
# Deploy a People Counter App at the Edge
| Details | |
|-----------------------|---------------|
| Programming Language: | Python 3.5 or 3.6 |

## What it Does
The people counter application will demonstrate how to create a smart video IoT solution using Intel® hardware and software tools. The app will detect people in a designated area, providing the number of people in the frame, average duration of people in frame, and total count.
## How it Works
The counter will use the Inference Engine included in the Intel® Distribution of OpenVINO™ Toolkit. The model used should be able to identify people in a video frame. The app should count the number of people in the current frame, the duration that a person is in the frame (time elapsed between entering and exiting a frame) and the total count of people. It then sends the data to a local web server using the Paho MQTT Python package.
You will choose a model to use and convert it with the Model Optimizer.

## Requirements
### Hardware
* 6th to 10th generation Intel® Core™ processor with Iris® Pro graphics or Intel® HD Graphics.
* OR use of Intel® Neural Compute Stick 2 (NCS2)
* OR Udacity classroom workspace for the related course
### Software
* Intel® Distribution of OpenVINO™ toolkit 2019 R3 release
* Node v6.17.1
* Npm v3.10.10
* CMake
* MQTT Mosca server
<span style="color:blue">
<strong>Table of Contents</strong>
<li> Page 1: Overview and Requirements
<li> Page 2: Setup Instructions
<li> Page 3: What Model to Use
<li> Page 4: Running Your Code
<li> Page 5: Tips for Running Locally
<li> Page 6: Helper Page for Button
</span>
<!--
%%ulab_page_divider
--><hr/>
## Setup
**<span style="color:red">
You do not need to install the Toolkit or Node in the workspace as they come pre-installed; you can skip to "Install npm".
</span>**
### Install Intel® Distribution of OpenVINO™ toolkit
Utilize the classroom workspace, or refer to the relevant instructions for your operating system for this step.
- [Linux/Ubuntu](./linux-setup.md)
- [Mac](./mac-setup.md)
- [Windows](./windows-setup.md)
### Install Nodejs and its dependencies
Utilize the classroom workspace, or refer to the relevant instructions for your operating system for this step.
- [Linux/Ubuntu](./linux-setup.md)
- [Mac](./mac-setup.md)
- [Windows](./windows-setup.md)
### Install npm
There are three components that need to be running in separate terminals for this application to work:
- MQTT Mosca server
- Node.js* Web server
- FFmpeg server
From the main directory:
* For MQTT/Mosca server:
```
cd webservice/server
npm install
```
* For Web server:
```
cd ../ui
npm install
```
**Note:** If any configuration errors occur in mosca server or Web server while using **npm install**, use the below commands:
```
sudo npm install npm -g
rm -rf node_modules
npm cache clean
npm config set registry "http://registry.npmjs.org"
npm install
```
<span style="color:blue">
<strong>Table of Contents</strong>
<li> Page 1: Overview and Requirements
<li> Page 2: Setup Instructions
<li> Page 3: What Model to Use
<li> Page 4: Running Your Code
<li> Page 5: Tips for Running Locally
<li> Page 6: Helper Page for Button
</span>
<!--
%%ulab_page_divider
--><hr/>
## What model to use
It is up to you to decide on what model to use for the application. You need to find a model not already converted to Intermediate Representation format (i.e. not one of the Intel® Pre-Trained Models), convert it, and utilize the converted model in your application.
Note that you may need to do additional processing of the output to handle incorrect detections, such as adjusting confidence threshold or accounting for 1-2 frames where the model fails to see a person already counted and would otherwise double count.
**If you are otherwise unable to find a suitable model after attempting and successfully converting at least three other models**, you can document in your write-up what the models were, how you converted them, and why they failed, and then utilize any of the Intel® Pre-Trained Models that may perform better.
<span style="color:blue">
<strong>Table of Contents</strong>
<li> Page 1: Overview and Requirements
<li> Page 2: Setup Instructions
<li> Page 3: What Model to Use
<li> Page 4: Running Your Code
<li> Page 5: Tips for Running Locally
<li> Page 6: Helper Page for Button
</span>
<!--
%%ulab_page_divider
--><hr/>
## Run the application
You can use the button to source the environment in the initial terminal, although any new terminals using the toolkit will need the command as detailed in Step 4 below.
<button id="ulab-button-3be1e29c" class="ulab-btn--primary"></button>
From the main directory:
### Step 1 - Start the Mosca server
```
cd webservice/server/node-server
node ./server.js
```
You should see the following message, if successful:
```
Mosca server started.
```
### Step 2 - Start the GUI
Open new terminal and run below commands.
```
cd webservice/ui
npm run dev
```
You should see the following message in the terminal.
```
webpack: Compiled successfully
```
### Step 3 - FFmpeg Server
Open new terminal and run the below commands.
```
sudo ffserver -f ./ffmpeg/server.conf
```
### Step 4 - Run the code
Open a new terminal to run the code.
#### Setup the environment
You must configure the environment to use the Intel® Distribution of OpenVINO™ toolkit one time per session by running the following command:
```
source /opt/intel/openvino/bin/setupvars.sh -pyver 3.5
```
You should also be able to run the application with Python 3.6, although newer versions of Python will not work with the app.
#### Running on the CPU
When running Intel® Distribution of OpenVINO™ toolkit Python applications on the CPU, the CPU extension library is required. This can be found at:
```
/opt/intel/openvino/deployment_tools/inference_engine/lib/intel64/
```
*Depending on whether you are using Linux or Mac, the filename will be either `libcpu_extension_sse4.so` or `libcpu_extension.dylib`, respectively.* (The Linux filename may be different if you are using a AVX architecture)
Though by default application runs on CPU, this can also be explicitly specified by ```-d CPU``` command-line argument:
```
python main.py -i resources/Pedestrian_Detect_2_1_1.mp4 -m your-model.xml -l /opt/intel/openvino/deployment_tools/inference_engine/lib/intel64/libcpu_extension_sse4.so -d CPU -pt 0.6 | ffmpeg -v warning -f rawvideo -pixel_format bgr24 -video_size 768x432 -framerate 24 -i - http://0.0.0.0:3004/fac.ffm
```
If you are in the classroom workspace, use the “Open App” button to view the output.
<button id="ulab-button-c7ebc45c" class="ulab-btn--primary"></button>
If working locally, to see the output on a web based interface, open the link [http://0.0.0.0:3004](http://0.0.0.0:3004/) in a browser.
#### Running on the Intel® Neural Compute Stick
**<span style="color:red">Not available in Udacity workspace.</span>**
To run on the Intel® Neural Compute Stick, use the ```-d MYRIAD``` command-line argument:
```
python3.5 main.py -d MYRIAD -i resources/Pedestrian_Detect_2_1_1.mp4 -m your-model.xml -pt 0.6 | ffmpeg -v warning -f rawvideo -pixel_format bgr24 -video_size 768x432 -framerate 24 -i - http://0.0.0.0:3004/fac.ffm
```
To see the output on a web based interface, open the link [http://0.0.0.0:3004](http://0.0.0.0:3004/) in a browser.
**Note:** The Intel® Neural Compute Stick can only run FP16 models at this time. The model that is passed to the application, through the `-m <path_to_model>` command-line argument, must be of data type FP16.
#### Using a camera stream instead of a video file
To get the input video from the camera, use the `-i CAM` command-line argument. Specify the resolution of the camera using the `-video_size` command line argument.
For example:
```
python main.py -i CAM -m your-model.xml -l /opt/intel/openvino/deployment_tools/inference_engine/lib/intel64/libcpu_extension_sse4.so -d CPU -pt 0.6 | ffmpeg -v warning -f rawvideo -pixel_format bgr24 -video_size 768x432 -framerate 24 -i - http://0.0.0.0:3004/fac.ffm
```
To see the output on a web based interface, open the link [http://0.0.0.0:3004](http://0.0.0.0:3004/) in a browser.
**Note:**
User has to give `-video_size` command line argument according to the input as it is used to specify the resolution of the video or image file.
<span style="color:blue">
<strong>Table of Contents</strong>
<li> Page 1: Overview and Requirements
<li> Page 2: Setup Instructions
<li> Page 3: What Model to Use
<li> Page 4: Running Your Code
<li> Page 5: Tips for Running Locally
<li> Page 6: Helper Page for Button
</span>
<!--
%%ulab_page_divider
--><hr/>
## A Note on Running Locally
The servers herein are configured to utilize the Udacity classroom workspace. As such,
to run on your local machine, you will need to change the below file:
```
webservice/ui/src/constants/constants.js
```
The `CAMERA_FEED_SERVER` and `MQTT_SERVER` both use the workspace configuration.
You can change each of these as follows:
```
CAMERA_FEED_SERVER: "http://localhost:3004"
...
MQTT_SERVER: "ws://localhost:3002"
```
<span style="color:blue">
<strong>Table of Contents</strong>
<li> Page 1: Overview and Requirements
<li> Page 2: Setup Instructions
<li> Page 3: What Model to Use
<li> Page 4: Running Your Code
<li> Page 5: Tips for Running Locally
<li> Page 6: Helper Page for Buttons
</span>
<!--
%%ulab_page_divider
--><hr/>
## Helper Page: Source Environment & Open App
As a helper to consolidate location of the buttons, find below buttons to source the environment and open the app in the preview window (assuming your app is running).
<button id="ulab-button-5e98247e" class="ulab-btn--primary"></button>
The `SOURCE ENV` button only works on the initial terminal; you can also source the environment with the below command:
```
source /opt/intel/openvino/bin/setupvars.sh -pyver 3.5
```
<button id="ulab-button-2174bf4d" class="ulab-btn--primary"></button>
| github_jupyter |
Deep Learning
=============
Assignment 1
------------
The objective of this assignment is to learn about simple data curation practices, and familiarize you with some of the data we'll be reusing later.
This notebook uses the [notMNIST](http://yaroslavvb.blogspot.com/2011/09/notmnist-dataset.html) dataset to be used with python experiments. This dataset is designed to look like the classic [MNIST](http://yann.lecun.com/exdb/mnist/) dataset, while looking a little more like real data: it's a harder task, and the data is a lot less 'clean' than MNIST.
```
# These are all the modules we'll be using later. Make sure you can import them
# before proceeding further.
from __future__ import print_function
import imageio
import matplotlib.pyplot as plt
import numpy as np
import os
import sys
import tarfile
from IPython.display import display, Image
from sklearn.linear_model import LogisticRegression
from six.moves.urllib.request import urlretrieve
from six.moves import cPickle as pickle
# Config the matplotlib backend as plotting inline in IPython
%matplotlib inline
```
First, we'll download the dataset to our local machine. The data consists of characters rendered in a variety of fonts on a 28x28 image. The labels are limited to 'A' through 'J' (10 classes). The training set has about 500k and the testset 19000 labeled examples. Given these sizes, it should be possible to train models quickly on any machine.
```
url = 'https://commondatastorage.googleapis.com/books1000/'
last_percent_reported = None
data_root = '.' # Change me to store data elsewhere
def download_progress_hook(count, blockSize, totalSize):
"""A hook to report the progress of a download. This is mostly intended for users with
slow internet connections. Reports every 5% change in download progress.
"""
global last_percent_reported
percent = int(count * blockSize * 100 / totalSize)
if last_percent_reported != percent:
if percent % 5 == 0:
sys.stdout.write("%s%%" % percent)
sys.stdout.flush()
else:
sys.stdout.write(".")
sys.stdout.flush()
last_percent_reported = percent
def maybe_download(filename, expected_bytes, force=False):
"""Download a file if not present, and make sure it's the right size."""
dest_filename = os.path.join(data_root, filename)
if force or not os.path.exists(dest_filename):
print('Attempting to download:', filename)
filename, _ = urlretrieve(url + filename, dest_filename, reporthook=download_progress_hook)
print('\nDownload Complete!')
statinfo = os.stat(dest_filename)
if statinfo.st_size == expected_bytes:
print('Found and verified', dest_filename)
else:
raise Exception(
'Failed to verify ' + dest_filename + '. Can you get to it with a browser?')
return dest_filename
train_filename = maybe_download('notMNIST_large.tar.gz', 247336696)
test_filename = maybe_download('notMNIST_small.tar.gz', 8458043)
```
Extract the dataset from the compressed .tar.gz file.
This should give you a set of directories, labeled A through J.
```
num_classes = 10
np.random.seed(133)
def maybe_extract(filename, force=False):
root = os.path.splitext(os.path.splitext(filename)[0])[0] # remove .tar.gz
if os.path.isdir(root) and not force:
# You may override by setting force=True.
print('%s already present - Skipping extraction of %s.' % (root, filename))
else:
print('Extracting data for %s. This may take a while. Please wait.' % root)
tar = tarfile.open(filename)
sys.stdout.flush()
tar.extractall(data_root)
tar.close()
data_folders = [
os.path.join(root, d) for d in sorted(os.listdir(root))
if os.path.isdir(os.path.join(root, d))]
if len(data_folders) != num_classes:
raise Exception(
'Expected %d folders, one per class. Found %d instead.' % (
num_classes, len(data_folders)))
print(data_folders)
return data_folders
train_folders = maybe_extract(train_filename)
test_folders = maybe_extract(test_filename)
```
---
Problem 1
---------
Let's take a peek at some of the data to make sure it looks sensible. Each exemplar should be an image of a character A through J rendered in a different font. Display a sample of the images that we just downloaded. Hint: you can use the package IPython.display.
---
Now let's load the data in a more manageable format. Since, depending on your computer setup you might not be able to fit it all in memory, we'll load each class into a separate dataset, store them on disk and curate them independently. Later we'll merge them into a single dataset of manageable size.
We'll convert the entire dataset into a 3D array (image index, x, y) of floating point values, normalized to have approximately zero mean and standard deviation ~0.5 to make training easier down the road.
A few images might not be readable, we'll just skip them.
```
image_size = 28 # Pixel width and height.
pixel_depth = 255.0 # Number of levels per pixel.
def load_letter(folder, min_num_images):
"""Load the data for a single letter label."""
image_files = os.listdir(folder)
dataset = np.ndarray(shape=(len(image_files), image_size, image_size),
dtype=np.float32)
print(folder)
num_images = 0
for image in image_files:
image_file = os.path.join(folder, image)
try:
image_data = (imageio.imread(image_file).astype(float) -
pixel_depth / 2) / pixel_depth
if image_data.shape != (image_size, image_size):
raise Exception('Unexpected image shape: %s' % str(image_data.shape))
dataset[num_images, :, :] = image_data
num_images = num_images + 1
except (IOError, ValueError) as e:
print('Could not read:', image_file, ':', e, '- it\'s ok, skipping.')
dataset = dataset[0:num_images, :, :]
if num_images < min_num_images:
raise Exception('Many fewer images than expected: %d < %d' %
(num_images, min_num_images))
print('Full dataset tensor:', dataset.shape)
print('Mean:', np.mean(dataset))
print('Standard deviation:', np.std(dataset))
return dataset
def maybe_pickle(data_folders, min_num_images_per_class, force=False):
dataset_names = []
for folder in data_folders:
set_filename = folder + '.pickle'
dataset_names.append(set_filename)
if os.path.exists(set_filename) and not force:
# You may override by setting force=True.
print('%s already present - Skipping pickling.' % set_filename)
else:
print('Pickling %s.' % set_filename)
dataset = load_letter(folder, min_num_images_per_class)
try:
with open(set_filename, 'wb') as f:
pickle.dump(dataset, f, pickle.HIGHEST_PROTOCOL)
except Exception as e:
print('Unable to save data to', set_filename, ':', e)
return dataset_names
train_datasets = maybe_pickle(train_folders, 45000)
test_datasets = maybe_pickle(test_folders, 1800)
```
---
Problem 2
---------
Let's verify that the data still looks good. Displaying a sample of the labels and images from the ndarray. Hint: you can use matplotlib.pyplot.
---
---
Problem 3
---------
Another check: we expect the data to be balanced across classes. Verify that.
---
Merge and prune the training data as needed. Depending on your computer setup, you might not be able to fit it all in memory, and you can tune `train_size` as needed. The labels will be stored into a separate array of integers 0 through 9.
Also create a validation dataset for hyperparameter tuning.
```
def make_arrays(nb_rows, img_size):
if nb_rows:
dataset = np.ndarray((nb_rows, img_size, img_size), dtype=np.float32)
labels = np.ndarray(nb_rows, dtype=np.int32)
else:
dataset, labels = None, None
return dataset, labels
def merge_datasets(pickle_files, train_size, valid_size=0):
num_classes = len(pickle_files)
valid_dataset, valid_labels = make_arrays(valid_size, image_size)
train_dataset, train_labels = make_arrays(train_size, image_size)
vsize_per_class = valid_size // num_classes
tsize_per_class = train_size // num_classes
start_v, start_t = 0, 0
end_v, end_t = vsize_per_class, tsize_per_class
end_l = vsize_per_class+tsize_per_class
for label, pickle_file in enumerate(pickle_files):
try:
with open(pickle_file, 'rb') as f:
letter_set = pickle.load(f)
# let's shuffle the letters to have random validation and training set
np.random.shuffle(letter_set)
if valid_dataset is not None:
valid_letter = letter_set[:vsize_per_class, :, :]
valid_dataset[start_v:end_v, :, :] = valid_letter
valid_labels[start_v:end_v] = label
start_v += vsize_per_class
end_v += vsize_per_class
train_letter = letter_set[vsize_per_class:end_l, :, :]
train_dataset[start_t:end_t, :, :] = train_letter
train_labels[start_t:end_t] = label
start_t += tsize_per_class
end_t += tsize_per_class
except Exception as e:
print('Unable to process data from', pickle_file, ':', e)
raise
return valid_dataset, valid_labels, train_dataset, train_labels
train_size = 200000
valid_size = 10000
test_size = 10000
valid_dataset, valid_labels, train_dataset, train_labels = merge_datasets(
train_datasets, train_size, valid_size)
_, _, test_dataset, test_labels = merge_datasets(test_datasets, test_size)
print('Training:', train_dataset.shape, train_labels.shape)
print('Validation:', valid_dataset.shape, valid_labels.shape)
print('Testing:', test_dataset.shape, test_labels.shape)
```
Next, we'll randomize the data. It's important to have the labels well shuffled for the training and test distributions to match.
```
def randomize(dataset, labels):
permutation = np.random.permutation(labels.shape[0])
shuffled_dataset = dataset[permutation,:,:]
shuffled_labels = labels[permutation]
return shuffled_dataset, shuffled_labels
train_dataset, train_labels = randomize(train_dataset, train_labels)
test_dataset, test_labels = randomize(test_dataset, test_labels)
valid_dataset, valid_labels = randomize(valid_dataset, valid_labels)
```
---
Problem 4
---------
Convince yourself that the data is still good after shuffling!
---
Finally, let's save the data for later reuse:
```
pickle_file = os.path.join(data_root, 'notMNIST.pickle')
try:
f = open(pickle_file, 'wb')
save = {
'train_dataset': train_dataset,
'train_labels': train_labels,
'valid_dataset': valid_dataset,
'valid_labels': valid_labels,
'test_dataset': test_dataset,
'test_labels': test_labels,
}
pickle.dump(save, f, pickle.HIGHEST_PROTOCOL)
f.close()
except Exception as e:
print('Unable to save data to', pickle_file, ':', e)
raise
statinfo = os.stat(pickle_file)
print('Compressed pickle size:', statinfo.st_size)
```
---
Problem 5
---------
By construction, this dataset might contain a lot of overlapping samples, including training data that's also contained in the validation and test set! Overlap between training and test can skew the results if you expect to use your model in an environment where there is never an overlap, but are actually ok if you expect to see training samples recur when you use it.
Measure how much overlap there is between training, validation and test samples.
Optional questions:
- What about near duplicates between datasets? (images that are almost identical)
- Create a sanitized validation and test set, and compare your accuracy on those in subsequent assignments.
---
---
Problem 6
---------
Let's get an idea of what an off-the-shelf classifier can give you on this data. It's always good to check that there is something to learn, and that it's a problem that is not so trivial that a canned solution solves it.
Train a simple model on this data using 50, 100, 1000 and 5000 training samples. Hint: you can use the LogisticRegression model from sklearn.linear_model.
Optional question: train an off-the-shelf model on all the data!
---
| github_jupyter |
# DAT210x - Programming with Python for DS
## Module4- Lab5
```
import pandas as pd
from scipy import misc
from mpl_toolkits.mplot3d import Axes3D
import matplotlib
import matplotlib.pyplot as plt
%matplotlib inline
# Look pretty...
# matplotlib.style.use('ggplot')
plt.style.use('ggplot')
```
Create a regular Python list (not NDArray) and name it `samples`:
```
samples = []
color = []
```
Code up a for-loop that iterates over the images in the `Datasets/ALOI/32/` folder. Look in the folder first, so you know how the files are organized, and what file number they start from and end at.
Load each `.png` file individually in your for-loop using the instructions provided in the Feature Representation reading. Once loaded, flatten the image into a single-dimensional NDArray and append it to your `samples` list.
**Optional**: You can resample the image down by a factor of two if you have a slower computer. You can also scale the image from `0-255` to `0.0-1.0` if you'd like--doing so shouldn't have any effect on the algorithm's results.
```
for i in range(0,355,5):
img = misc.imread('32_r%s.png' % i)
samples.append((img[::2,::2]/255.0).reshape(-1))
color.append('b')
```
Convert `samples` to a DataFrame named `df`:
```
df = pd.DataFrame(samples)
```
Import any necessary libraries to perform Isomap here, reduce `df` down to three components and using `K=6` for your neighborhood size:
```
from sklearn import manifold
iso = manifold.Isomap(n_neighbors=2, n_components=3)
iso.fit(df)
isot = iso.transform(df)
dft = pd.DataFrame(isot)
```
Create a 2D Scatter plot to graph your manifold. You can use either `'o'` or `'.'` as your marker. Graph the first two isomap components:
```
dft.plot.scatter(0, 1, marker='o')
```
Chart a 3D Scatter plot to graph your manifold. You can use either `'o'` or `'.'` as your marker:
```
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.set_xlabel('principle')
ax.set_ylabel('second')
ax.set_zlabel('third')
ax. scatter(dft.iloc[:,0],dft.iloc[:,1],dft.iloc[:,2], marker='o')
```
Answer the first three lab questions!
Create another for loop. This time it should iterate over all the images in the `Datasets/ALOI/32_i` directory. Just like last time, load up each image, process them the way you did previously, and append them into your existing `samples` list:
```
for i in range(110,220,10):
img = misc.imread('32_i%s.png' % i)
samples.append((img[::2,::2]/255.0).reshape(-1))
color.append('r')
```
Convert `samples` to a DataFrame named `df`:
```
df2 = pd.DataFrame(samples)
```
Import any necessary libraries to perform Isomap here, reduce `df` down to three components and using `K=6` for your neighborhood size:
```
iso = manifold.Isomap(n_neighbors=6, n_components=3)
iso.fit(df2)
isot = iso.transform(df2)
dft = pd.DataFrame(isot)
```
Create a 2D Scatter plot to graph your manifold. You can use either `'o'` or `'.'` as your marker. Graph the first two isomap components:
```
dft.plot.scatter(0,1,c=color,marker='o')
```
Chart a 3D Scatter plot to graph your manifold. You can use either `'o'` or `'.'` as your marker:
```
fig = plt.figure()
ax = fig.add_subplot('111',projection='3d')
ax.set_xlabel('primary')
ax.set_ylabel('secondary')
ax.set_zlabel('tirshiary')
ax.scatter(dft.iloc[:,0],dft.iloc[:,1],dft.iloc[:,2], c=color, marker='o')
```
| github_jupyter |
# LAB 03: Basic Feature Engineering in Keras
**Learning Objectives**
1. Create an input pipeline using tf.data
2. Engineer features to create categorical, crossed, and numerical feature columns
## Introduction
In this lab, we utilize feature engineering to improve the prediction of housing prices using a Keras Sequential Model.
Each learning objective will correspond to a __#TODO__ in the notebook where you will complete the notebook cell's code before running. Refer to the [solution](https://github.com/GoogleCloudPlatform/training-data-analyst/blob/master/courses/machine_learning/deepdive2/feature_engineering/solutions/3_keras_basic_feat_eng.ipynb) for reference.
Start by importing the necessary libraries for this lab.
```
# Install Sklearn
!python3 -m pip install --user sklearn
# Ensure the right version of Tensorflow is installed.
!pip freeze | grep tensorflow==2.1 || pip install tensorflow==2.1
```
**Note:** After executing the above cell you will see the output
`tensorflow==2.1.0` that is the installed version of tensorflow.
```
import os
import tensorflow.keras
import matplotlib.pyplot as plt
import pandas as pd
import tensorflow as tf
from tensorflow import feature_column as fc
from tensorflow.keras import layers
from sklearn.model_selection import train_test_split
from keras.utils import plot_model
print("TensorFlow version: ",tf.version.VERSION)
```
Many of the Google Machine Learning Courses Programming Exercises use the [California Housing Dataset](https://developers.google.com/machine-learning/crash-course/california-housing-data-description
), which contains data drawn from the 1990 U.S. Census. Our lab dataset has been pre-processed so that there are no missing values.
First, let's download the raw .csv data by copying the data from a cloud storage bucket.
```
if not os.path.isdir("../data"):
os.makedirs("../data")
!gsutil cp gs://cloud-training-demos/feat_eng/housing/housing_pre-proc.csv ../data
!ls -l ../data/
```
Now, let's read in the dataset just copied from the cloud storage bucket and create a Pandas dataframe.
```
housing_df = pd.read_csv('../data/housing_pre-proc.csv', error_bad_lines=False)
housing_df.head()
```
We can use .describe() to see some summary statistics for the numeric fields in our dataframe. Note, for example, the count row and corresponding columns. The count shows 20433.000000 for all feature columns. Thus, there are no missing values.
```
housing_df.describe()
```
#### Split the dataset for ML
The dataset we loaded was a single CSV file. We will split this into train, validation, and test sets.
```
train, test = train_test_split(housing_df, test_size=0.2)
train, val = train_test_split(train, test_size=0.2)
print(len(train), 'train examples')
print(len(val), 'validation examples')
print(len(test), 'test examples')
```
Now, we need to output the split files. We will specifically need the test.csv later for testing. You should see the files appear in the home directory.
```
train.to_csv('../data/housing-train.csv', encoding='utf-8', index=False)
val.to_csv('../data/housing-val.csv', encoding='utf-8', index=False)
test.to_csv('../data/housing-test.csv', encoding='utf-8', index=False)
!head ../data/housing*.csv
```
## Lab Task 1: Create an input pipeline using tf.data
Next, we will wrap the dataframes with [tf.data](https://www.tensorflow.org/guide/datasets). This will enable us to use feature columns as a bridge to map from the columns in the Pandas dataframe to features used to train the model.
Here, we create an input pipeline using tf.data. This function is missing two lines. Correct and run the cell.
```
# A utility method to create a tf.data dataset from a Pandas Dataframe
def df_to_dataset(dataframe, shuffle=True, batch_size=32):
dataframe = dataframe.copy()
# TODO 1a -- Your code here
if shuffle:
ds = ds.shuffle(buffer_size=len(dataframe))
ds = ds.batch(batch_size)
return ds
```
Next we initialize the training and validation datasets.
```
batch_size = 32
train_ds = df_to_dataset(train)
val_ds = df_to_dataset(val, shuffle=False, batch_size=batch_size)
```
Now that we have created the input pipeline, let's call it to see the format of the data it returns. We have used a small batch size to keep the output readable.
```
# TODO 1b -- Your code here
```
We can see that the dataset returns a dictionary of column names (from the dataframe) that map to column values from rows in the dataframe.
#### Numeric columns
The output of a feature column becomes the input to the model. A numeric is the simplest type of column. It is used to represent real valued features. When using this column, your model will receive the column value from the dataframe unchanged.
In the California housing prices dataset, most columns from the dataframe are numeric. Let' create a variable called **numeric_cols** to hold only the numerical feature columns.
```
# TODO 1c -- Your code here
```
#### Scaler function
It is very important for numerical variables to get scaled before they are "fed" into the neural network. Here we use min-max scaling. Here we are creating a function named 'get_scal' which takes a list of numerical features and returns a 'minmax' function, which will be used in tf.feature_column.numeric_column() as normalizer_fn in parameters. 'Minmax' function itself takes a 'numerical' number from a particular feature and return scaled value of that number.
Next, we scale the numerical feature columns that we assigned to the variable "numeric cols".
```
# Scalar def get_scal(feature):
# TODO 1d -- Your code here
# TODO 1e -- Your code here
```
Next, we should validate the total number of feature columns. Compare this number to the number of numeric features you input earlier.
```
print('Total number of feature coLumns: ', len(feature_columns))
```
### Using the Keras Sequential Model
Next, we will run this cell to compile and fit the Keras Sequential model.
```
# Model create
feature_layer = tf.keras.layers.DenseFeatures(feature_columns, dtype='float64')
model = tf.keras.Sequential([
feature_layer,
layers.Dense(12, input_dim=8, activation='relu'),
layers.Dense(8, activation='relu'),
layers.Dense(1, activation='linear', name='median_house_value')
])
# Model compile
model.compile(optimizer='adam',
loss='mse',
metrics=['mse'])
# Model Fit
history = model.fit(train_ds,
validation_data=val_ds,
epochs=32)
```
Next we show loss as Mean Square Error (MSE). Remember that MSE is the most commonly used regression loss function. MSE is the sum of squared distances between our target variable (e.g. housing median age) and predicted values.
```
loss, mse = model.evaluate(train_ds)
print("Mean Squared Error", mse)
```
#### Visualize the model loss curve
Next, we will use matplotlib to draw the model's loss curves for training and validation. A line plot is also created showing the mean squared error loss over the training epochs for both the train (blue) and test (orange) sets.
```
def plot_curves(history, metrics):
nrows = 1
ncols = 2
fig = plt.figure(figsize=(10, 5))
for idx, key in enumerate(metrics):
ax = fig.add_subplot(nrows, ncols, idx+1)
plt.plot(history.history[key])
plt.plot(history.history['val_{}'.format(key)])
plt.title('model {}'.format(key))
plt.ylabel(key)
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='upper left');
plot_curves(history, ['loss', 'mse'])
```
### Load test data
Next, we read in the test.csv file and validate that there are no null values.
Again, we can use .describe() to see some summary statistics for the numeric fields in our dataframe. The count shows 4087.000000 for all feature columns. Thus, there are no missing values.
```
test_data = pd.read_csv('../data/housing-test.csv')
test_data.describe()
```
Now that we have created an input pipeline using tf.data and compiled a Keras Sequential Model, we now create the input function for the test data and to initialize the test_predict variable.
```
# TODO 1f -- Your code here
test_predict = test_input_fn(dict(test_data))
```
#### Prediction: Linear Regression
Before we begin to feature engineer our feature columns, we should predict the median house value. By predicting the median house value now, we can then compare it with the median house value after feature engineeing.
To predict with Keras, you simply call [model.predict()](https://keras.io/models/model/#predict) and pass in the housing features you want to predict the median_house_value for. Note: We are predicting the model locally.
```
predicted_median_house_value = model.predict(test_predict)
```
Next, we run two predictions in separate cells - one where ocean_proximity=INLAND and one where ocean_proximity= NEAR OCEAN.
```
# Ocean_proximity is INLAND
model.predict({
'longitude': tf.convert_to_tensor([-121.86]),
'latitude': tf.convert_to_tensor([39.78]),
'housing_median_age': tf.convert_to_tensor([12.0]),
'total_rooms': tf.convert_to_tensor([7653.0]),
'total_bedrooms': tf.convert_to_tensor([1578.0]),
'population': tf.convert_to_tensor([3628.0]),
'households': tf.convert_to_tensor([1494.0]),
'median_income': tf.convert_to_tensor([3.0905]),
'ocean_proximity': tf.convert_to_tensor(['INLAND'])
}, steps=1)
# Ocean_proximity is NEAR OCEAN
model.predict({
'longitude': tf.convert_to_tensor([-122.43]),
'latitude': tf.convert_to_tensor([37.63]),
'housing_median_age': tf.convert_to_tensor([34.0]),
'total_rooms': tf.convert_to_tensor([4135.0]),
'total_bedrooms': tf.convert_to_tensor([687.0]),
'population': tf.convert_to_tensor([2154.0]),
'households': tf.convert_to_tensor([742.0]),
'median_income': tf.convert_to_tensor([4.9732]),
'ocean_proximity': tf.convert_to_tensor(['NEAR OCEAN'])
}, steps=1)
```
The arrays returns a predicted value. What do these numbers mean? Let's compare this value to the test set.
Go to the test.csv you read in a few cells up. Locate the first line and find the median_house_value - which should be 249,000 dollars near the ocean. What value did your model predicted for the median_house_value? Was it a solid model performance? Let's see if we can improve this a bit with feature engineering!
## Lab Task 2: Engineer features to create categorical and numerical features
Now we create a cell that indicates which features will be used in the model.
Note: Be sure to bucketize 'housing_median_age' and ensure that 'ocean_proximity' is one-hot encoded. And, don't forget your numeric values!
```
# TODO 2a -- Your code here
```
Next, we scale the numerical, bucktized, and categorical feature columns that we assigned to the variables in the precding cell.
```
# Scalar def get_scal(feature):
def get_scal(feature):
def minmax(x):
mini = train[feature].min()
maxi = train[feature].max()
return (x - mini)/(maxi-mini)
return(minmax)
# All numerical features - scaling
feature_columns = []
for header in numeric_cols:
scal_input_fn = get_scal(header)
feature_columns.append(fc.numeric_column(header,
normalizer_fn=scal_input_fn))
```
### Categorical Feature
In this dataset, 'ocean_proximity' is represented as a string. We cannot feed strings directly to a model. Instead, we must first map them to numeric values. The categorical vocabulary columns provide a way to represent strings as a one-hot vector.
Next, we create a categorical feature using 'ocean_proximity'.
```
# TODO 2b -- Your code here
```
### Bucketized Feature
Often, you don't want to feed a number directly into the model, but instead split its value into different categories based on numerical ranges. Consider our raw data that represents a homes' age. Instead of representing the house age as a numeric column, we could split the home age into several buckets using a [bucketized column](https://www.tensorflow.org/api_docs/python/tf/feature_column/bucketized_column). Notice the one-hot values below describe which age range each row matches.
Next we create a bucketized column using 'housing_median_age'
```
# TODO 2c -- Your code here
```
### Feature Cross
Combining features into a single feature, better known as [feature crosses](https://developers.google.com/machine-learning/glossary/#feature_cross), enables a model to learn separate weights for each combination of features.
Next, we create a feature cross of 'housing_median_age' and 'ocean_proximity'.
```
# TODO 2d -- Your code here
```
Next, we should validate the total number of feature columns. Compare this number to the number of numeric features you input earlier.
```
print('Total number of feature coumns: ', len(feature_columns))
```
Next, we will run this cell to compile and fit the Keras Sequential model. This is the same model we ran earlier.
```
# Model create
feature_layer = tf.keras.layers.DenseFeatures(feature_columns,
dtype='float64')
model = tf.keras.Sequential([
feature_layer,
layers.Dense(12, input_dim=8, activation='relu'),
layers.Dense(8, activation='relu'),
layers.Dense(1, activation='linear', name='median_house_value')
])
# Model compile
model.compile(optimizer='adam',
loss='mse',
metrics=['mse'])
# Model Fit
history = model.fit(train_ds,
validation_data=val_ds,
epochs=32)
```
Next, we show loss and mean squared error then plot the model.
```
loss, mse = model.evaluate(train_ds)
print("Mean Squared Error", mse)
plot_curves(history, ['loss', 'mse'])
```
Next we create a prediction model. Note: You may use the same values from the previous prediciton.
```
# TODO 2e -- Your code here
```
### Analysis
The array returns a predicted value. Compare this value to the test set you ran earlier. Your predicted value may be a bit better.
Now that you have your "feature engineering template" setup, you can experiment by creating additional features. For example, you can create derived features, such as households per population, and see how they impact the model. You can also experiment with replacing the features you used to create the feature cross.
Copyright 2020 Google Inc.
Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| github_jupyter |
```
# default_exp models.XCMPlus
```
# XCM (An Explainable Convolutional Neural Network for Multivariate Time Series Classification)
> This is an unofficial PyTorch implementation by Ignacio Oguiza of - oguiza@gmail.com based on Temporal Convolutional Network (Bai, 2018).
**References:**
* Fauvel, K., Lin, T., Masson, V., Fromont, É., & Termier, A. (2020). XCM: An Explainable Convolutional Neural Network for Multivariate Time Series Classification. arXiv preprint arXiv:2009.04796.
* Official XCM PyTorch implementation: not available as of Nov 27th, 2020
```
#export
from tsai.imports import *
from tsai.utils import *
from tsai.models.layers import *
from tsai.models.utils import *
from tsai.models.explainability import *
#export
# This is an unofficial PyTorch implementation by Ignacio Oguiza - oguiza@gmail.com based on:
# Fauvel, K., Lin, T., Masson, V., Fromont, É., & Termier, A. (2020). XCM: An Explainable Convolutional Neural Network for
# Multivariate Time Series Classification. arXiv preprint arXiv:2009.04796.
# Official XCM PyTorch implementation: not available as of Nov 27th, 2020
class XCMPlus(nn.Sequential):
def __init__(self, c_in:int, c_out:int, seq_len:Optional[int]=None, nf:int=128, window_perc:float=1., flatten:bool=False, custom_head:callable=None,
concat_pool:bool=False, fc_dropout:float=0., bn:bool=False, y_range:tuple=None, **kwargs):
window_size = int(round(seq_len * window_perc, 0))
backbone = _XCMPlus_Backbone(c_in, c_out, seq_len=seq_len, nf=nf, window_perc=window_perc)
self.head_nf = nf
self.c_out = c_out
self.seq_len = seq_len
if custom_head: head = custom_head(self.head_nf, c_out, seq_len, **kwargs)
else: head = self.create_head(self.head_nf, c_out, seq_len, flatten=flatten, concat_pool=concat_pool,
fc_dropout=fc_dropout, bn=bn, y_range=y_range)
super().__init__(OrderedDict([('backbone', backbone), ('head', head)]))
def create_head(self, nf, c_out, seq_len=None, flatten=False, concat_pool=False, fc_dropout=0., bn=False, y_range=None):
if flatten:
nf *= seq_len
layers = [Flatten()]
else:
if concat_pool: nf *= 2
layers = [GACP1d(1) if concat_pool else GAP1d(1)]
layers += [LinBnDrop(nf, c_out, bn=bn, p=fc_dropout)]
if y_range: layers += [SigmoidRange(*y_range)]
return nn.Sequential(*layers)
def show_gradcam(self, x, y=None, detach=True, cpu=True, apply_relu=True, cmap='inferno', figsize=None, **kwargs):
att_maps = get_attribution_map(self, [self.backbone.conv2dblock, self.backbone.conv1dblock], x, y=y, detach=detach, cpu=cpu, apply_relu=apply_relu)
att_maps[0] = (att_maps[0] - att_maps[0].min()) / (att_maps[0].max() - att_maps[0].min())
att_maps[1] = (att_maps[1] - att_maps[1].min()) / (att_maps[1].max() - att_maps[1].min())
figsize = ifnone(figsize, (10, 10))
fig = plt.figure(figsize=figsize, **kwargs)
ax = plt.axes()
plt.title('Observed variables')
im = ax.imshow(att_maps[0], cmap=cmap)
cax = fig.add_axes([ax.get_position().x1+0.01,ax.get_position().y0,0.02,ax.get_position().height])
plt.colorbar(im, cax=cax)
plt.show()
fig = plt.figure(figsize=figsize, **kwargs)
ax = plt.axes()
plt.title('Time')
im = ax.imshow(att_maps[1], cmap=cmap)
cax = fig.add_axes([ax.get_position().x1+0.01,ax.get_position().y0,0.02,ax.get_position().height])
plt.colorbar(im, cax=cax)
plt.show()
class _XCMPlus_Backbone(Module):
def __init__(self, c_in:int, c_out:int, seq_len:Optional[int]=None, nf:int=128, window_perc:float=1.):
window_size = int(round(seq_len * window_perc, 0))
self.conv2dblock = nn.Sequential(*[Unsqueeze(1), Conv2d(1, nf, kernel_size=(1, window_size), padding='same'), BatchNorm(nf), nn.ReLU()])
self.conv2d1x1block = nn.Sequential(*[nn.Conv2d(nf, 1, kernel_size=1), nn.ReLU(), Squeeze(1)])
self.conv1dblock = nn.Sequential(*[Conv1d(c_in, nf, kernel_size=window_size, padding='same'), BatchNorm(nf, ndim=1), nn.ReLU()])
self.conv1d1x1block = nn.Sequential(*[nn.Conv1d(nf, 1, kernel_size=1), nn.ReLU()])
self.concat = Concat()
self.conv1d = nn.Sequential(*[Conv1d(c_in + 1, nf, kernel_size=window_size, padding='same'), BatchNorm(nf, ndim=1), nn.ReLU()])
def forward(self, x):
x1 = self.conv2dblock(x)
x1 = self.conv2d1x1block(x1)
x2 = self.conv1dblock(x)
x2 = self.conv1d1x1block(x2)
out = self.concat((x2, x1))
out = self.conv1d(out)
return out
from tsai.data.all import *
from tsai.models.XCM import *
dsid = 'NATOPS'
X, y, splits = get_UCR_data(dsid, split_data=False)
tfms = [None, Categorize()]
dls = get_ts_dls(X, y, splits=splits, tfms=tfms)
model = XCMPlus(dls.vars, dls.c, dls.len)
learn = Learner(dls, model, metrics=accuracy)
xb, yb = dls.one_batch()
bs, c_in, seq_len = xb.shape
c_out = len(np.unique(yb))
model = XCMPlus(c_in, c_out, seq_len, fc_dropout=.5)
test_eq(model(xb).shape, (bs, c_out))
model = XCMPlus(c_in, c_out, seq_len, concat_pool=True)
test_eq(model(xb).shape, (bs, c_out))
model = XCMPlus(c_in, c_out, seq_len)
test_eq(model(xb).shape, (bs, c_out))
test_eq(count_parameters(XCMPlus(c_in, c_out, seq_len)), count_parameters(XCM(c_in, c_out, seq_len)))
model
model.show_gradcam(xb[0], yb[0])
bs = 16
n_vars = 3
seq_len = 12
c_out = 10
xb = torch.rand(bs, n_vars, seq_len)
new_head = partial(conv_lin_3d_head, d=(5, 2))
net = XCMPlus(n_vars, c_out, seq_len, custom_head=new_head)
print(net(xb).shape)
net.head
bs = 16
n_vars = 3
seq_len = 12
c_out = 2
xb = torch.rand(bs, n_vars, seq_len)
net = XCMPlus(n_vars, c_out, seq_len)
change_model_head(net, create_pool_plus_head, concat_pool=False)
print(net(xb).shape)
net.head
#hide
out = create_scripts(); beep(out)
```
| github_jupyter |
## Intro to deep learning for medical imaging by [MD.ai](https://www.md.ai)
## Lesson 3. RSNA Pneumonia Detection Challenge (Kaggel API)
The [Radiological Society of North America](http://www.rsna.org/) Pneumonia Detection Challenge: https://www.kaggle.com/c/rsna-pneumonia-detection-challenge
This notebook covers the basics of parsing the competition dataset, training using a detector basd on the [Mask-RCNN algorithm](https://arxiv.org/abs/1703.06870) for object detection and instance segmentation.
This notebook is developed by [MD.ai](https://www.md.ai), the platform for medical AI.
This notebook requires Google's [TensorFlow](https://www.tensorflow.org/) machine learning framework.
**Intro to deep learning for medical imaging lessons**
- Lesson 1. Classification of chest vs. adominal X-rays using TensorFlow/Keras [Github](https://github.com/mdai/ml-lessons/blob/master/lesson1-xray-images-classification.ipynb) [Annotator](https://public.md.ai/annotator/project/PVq9raBJ)
- Lesson 2. Lung X-Rays Semantic Segmentation using UNets. [Github](https://github.com/mdai/ml-lessons/blob/master/lesson2-lung-xrays-segmentation.ipynb)
[Annotator](https://public.md.ai/annotator/project/aGq4k6NW/workspace)
- Lesson 3. RSNA Pneumonia detection using Kaggle data format [Github](https://github.com/mdai/ml-lessons/blob/master/lesson3-rsna-pneumonia-detection-kaggle.ipynb) [Annotator](https://public.md.ai/annotator/project/LxR6zdR2/workspace)
- Lesson 3. RSNA Pneumonia detection using MD.ai python client library [Github](https://github.com/mdai/ml-lessons/blob/master/lesson3-rsna-pneumonia-detection-mdai-client-lib.ipynb) [Annotator](https://public.md.ai/annotator/project/LxR6zdR2/workspace)
*Copyright 2018 MD.ai, Inc.
Licensed under the Apache License, Version 2.0*
```
# install dependencies not included by Colab
# use pip3 to ensure compatibility w/ Google Deep Learning Images
!pip3 install -q pydicom
!pip3 install -q tqdm
!pip3 install -q imgaug
import os
import sys
import random
import math
import numpy as np
import cv2
import matplotlib.pyplot as plt
import json
import pydicom
from imgaug import augmenters as iaa
from tqdm import tqdm
import pandas as pd
import glob
```
### First: Install Kaggle API for download competition data.
```
# Install Kaggle API for download competition data
!pip3 install -q kaggle
```
### Next: You must accept the user agreement on the competition website! Then follow [instructions to obtain your Kaggle Credentials.](https://github.com/Kaggle/kaggle-api#api-credentials)
If you are unable to download the competition dataset, check to see if you have accepted the **user agreement** on the competition website.
```
# enter your Kaggle credentionals here
os.environ['KAGGLE_USERNAME']=""
os.environ['KAGGLE_KEY']=""
# Root directory of the project
ROOT_DIR = os.path.abspath('./lesson3-data')
# Directory to save logs and trained model
MODEL_DIR = os.path.join(ROOT_DIR, 'logs')
if not os.path.exists(ROOT_DIR):
os.makedirs(ROOT_DIR)
os.chdir(ROOT_DIR)
# If you are unable to download the competition dataset, check to see if you have
# accepted the user agreement on the competition website.
!kaggle competitions download -c rsna-pneumonia-detection-challenge
```
### Data is downloaded as zip files. Unzip the test and train datasets as well as the csv of annotations.
```
# unzipping takes a few minutes
!unzip -q -o stage_1_test_images.zip -d stage_1_test_images
!unzip -q -o stage_1_train_images.zip -d stage_1_train_images
!unzip -q -o stage_1_train_labels.csv.zip
```
### MD.ai Annotator
Additionally, If you are interested in augmenting the existing annotations, you can use the MD.ai annotator to view DICOM images, and create annotatios to be exported.
MD.ai annotator project URL for the Kaggle dataset: https://public.md.ai/annotator/project/LxR6zdR2/workspace
**Annotator features**
- The annotator can be used to view DICOM images and create image and exam level annotations.
- You can apply the annotator to filter by label, adjudicate annotations, and assign annotation tasks to your team.
- Notebooks can be built directly within the annotator for rapid model development.
- The data wrangling is abstracted away by the interface and by our MD.ai library.
- Simplifies image annotation in order to widen the participation in the futrue of medical image deep learning.
The annotator allows you to create initial annotations, build and run models, modify/finetune the annotations based on predicted values, and repeat.
The MD.ai python client library implements functions to easily download images and annotations and to prepare the datasets used to train the model for classification. See the following example notebook for parsing annotations and training using MD.ai annotator:
https://github.com/mdai/ml-lessons/blob/master/lesson3-rsna-pneumonia-detection-mdai-client-lib.ipynb
- MD.ai URL: https://www.md.ai
- MD.ai documentation URL: https://docs.md.ai/
### Install Matterport's Mask-RCNN model from github.
See the [Matterport's implementation of Mask-RCNN](https://github.com/matterport/Mask_RCNN).
```
os.chdir(ROOT_DIR)
!git clone https://github.com/matterport/Mask_RCNN.git
os.chdir('Mask_RCNN')
!python setup.py -q install
# Import Mask RCNN
sys.path.append(os.path.join(ROOT_DIR, 'Mask_RCNN')) # To find local version of the library
from mrcnn.config import Config
from mrcnn import utils
import mrcnn.model as modellib
from mrcnn import visualize
from mrcnn.model import log
train_dicom_dir = os.path.join(ROOT_DIR, 'stage_1_train_images')
test_dicom_dir = os.path.join(ROOT_DIR, 'stage_1_test_images')
```
### Some setup functions and classes for Mask-RCNN
- dicom_fps is a list of the dicom image path and filenames
- image_annotions is a dictionary of the annotations keyed by the filenames
- parsing the dataset returns a list of the image filenames and the annotations dictionary
```
def get_dicom_fps(dicom_dir):
dicom_fps = glob.glob(dicom_dir+'/'+'*.dcm')
return list(set(dicom_fps))
def parse_dataset(dicom_dir, anns):
image_fps = get_dicom_fps(dicom_dir)
image_annotations = {fp: [] for fp in image_fps}
for index, row in anns.iterrows():
fp = os.path.join(dicom_dir, row['patientId']+'.dcm')
image_annotations[fp].append(row)
return image_fps, image_annotations
# The following parameters have been selected to reduce running time for demonstration purposes
# These are not optimal
class DetectorConfig(Config):
"""Configuration for training pneumonia detection on the RSNA pneumonia dataset.
Overrides values in the base Config class.
"""
# Give the configuration a recognizable name
NAME = 'pneumonia'
# Train on 1 GPU and 8 images per GPU. We can put multiple images on each
# GPU because the images are small. Batch size is 8 (GPUs * images/GPU).
GPU_COUNT = 1
IMAGES_PER_GPU = 8
BACKBONE = 'resnet50'
NUM_CLASSES = 2 # background + 1 pneumonia classes
# Use small images for faster training. Set the limits of the small side
# the large side, and that determines the image shape.
IMAGE_MIN_DIM = 64
IMAGE_MAX_DIM = 64
RPN_ANCHOR_SCALES = (32, 64)
TRAIN_ROIS_PER_IMAGE = 16
MAX_GT_INSTANCES = 3
DETECTION_MAX_INSTANCES = 3
DETECTION_MIN_CONFIDENCE = 0.9
DETECTION_NMS_THRESHOLD = 0.1
RPN_TRAIN_ANCHORS_PER_IMAGE = 16
STEPS_PER_EPOCH = 100
TOP_DOWN_PYRAMID_SIZE = 32
STEPS_PER_EPOCH = 100
config = DetectorConfig()
config.display()
class DetectorDataset(utils.Dataset):
"""Dataset class for training pneumonia detection on the RSNA pneumonia dataset.
"""
def __init__(self, image_fps, image_annotations, orig_height, orig_width):
super().__init__(self)
# Add classes
self.add_class('pneumonia', 1, 'Lung Opacity')
# add images
for i, fp in enumerate(image_fps):
annotations = image_annotations[fp]
self.add_image('pneumonia', image_id=i, path=fp,
annotations=annotations, orig_height=orig_height, orig_width=orig_width)
def image_reference(self, image_id):
info = self.image_info[image_id]
return info['path']
def load_image(self, image_id):
info = self.image_info[image_id]
fp = info['path']
ds = pydicom.read_file(fp)
image = ds.pixel_array
# If grayscale. Convert to RGB for consistency.
if len(image.shape) != 3 or image.shape[2] != 3:
image = np.stack((image,) * 3, -1)
return image
def load_mask(self, image_id):
info = self.image_info[image_id]
annotations = info['annotations']
count = len(annotations)
if count == 0:
mask = np.zeros((info['orig_height'], info['orig_width'], 1), dtype=np.uint8)
class_ids = np.zeros((1,), dtype=np.int32)
else:
mask = np.zeros((info['orig_height'], info['orig_width'], count), dtype=np.uint8)
class_ids = np.zeros((count,), dtype=np.int32)
for i, a in enumerate(annotations):
if a['Target'] == 1:
x = int(a['x'])
y = int(a['y'])
w = int(a['width'])
h = int(a['height'])
mask_instance = mask[:, :, i].copy()
cv2.rectangle(mask_instance, (x, y), (x+w, y+h), 255, -1)
mask[:, :, i] = mask_instance
class_ids[i] = 1
return mask.astype(np.bool), class_ids.astype(np.int32)
```
### Examine the annotation data, parse the dataset, and view dicom fields
```
# training dataset
anns = pd.read_csv(os.path.join(ROOT_DIR, 'stage_1_train_labels.csv'))
anns.head(6)
image_fps, image_annotations = parse_dataset(train_dicom_dir, anns=anns)
ds = pydicom.read_file(image_fps[0]) # read dicom image from filepath
image = ds.pixel_array # get image array
# show dicom fields
ds
# Original DICOM image size: 1024 x 1024
ORIG_SIZE = 1024
```
### Split the data into training and validation datasets
**Note: We have only used only a portion of the images for demonstration purposes. See comments below.**
- To use all the images do: image_fps_list = list(image_fps)
- Or change the number of images from 100 to a custom number
```
######################################################################
# Modify this line to use more or fewer images for training/validation.
# To use all images, do: image_fps_list = list(image_fps)
image_fps_list = list(image_fps[:1000])
#####################################################################
# split dataset into training vs. validation dataset
# split ratio is set to 0.9 vs. 0.1 (train vs. validation, respectively)
sorted(image_fps_list)
random.seed(42)
random.shuffle(image_fps_list)
validation_split = 0.1
split_index = int((1 - validation_split) * len(image_fps_list))
image_fps_train = image_fps_list[:split_index]
image_fps_val = image_fps_list[split_index:]
print(len(image_fps_train), len(image_fps_val))
```
### Create and prepare the training dataset using the DetectorDataset class.
```
# prepare the training dataset
dataset_train = DetectorDataset(image_fps_train, image_annotations, ORIG_SIZE, ORIG_SIZE)
dataset_train.prepare()
```
### Let's look at a sample annotation. We see a bounding box with (x, y) of the the top left corner as well as the width and height.
```
# Show annotation(s) for a DICOM image
test_fp = random.choice(image_fps_train)
image_annotations[test_fp]
# prepare the validation dataset
dataset_val = DetectorDataset(image_fps_val, image_annotations, ORIG_SIZE, ORIG_SIZE)
dataset_val.prepare()
```
### Display a random image with bounding boxes
```
# Load and display random samples and their bounding boxes
# Suggestion: Run this a few times to see different examples.
image_id = random.choice(dataset_train.image_ids)
image_fp = dataset_train.image_reference(image_id)
image = dataset_train.load_image(image_id)
mask, class_ids = dataset_train.load_mask(image_id)
print(image.shape)
plt.figure(figsize=(10, 10))
plt.subplot(1, 2, 1)
plt.imshow(image[:, :, 0], cmap='gray')
plt.axis('off')
plt.subplot(1, 2, 2)
masked = np.zeros(image.shape[:2])
for i in range(mask.shape[2]):
masked += image[:, :, 0] * mask[:, :, i]
plt.imshow(masked, cmap='gray')
plt.axis('off')
print(image_fp)
print(class_ids)
model = modellib.MaskRCNN(mode='training', config=config, model_dir=MODEL_DIR)
```
### Image Augmentation. Try finetuning some variables to custom values
```
# Image augmentation
augmentation = iaa.SomeOf((0, 1), [
iaa.Fliplr(0.5),
iaa.Affine(
scale={"x": (0.8, 1.2), "y": (0.8, 1.2)},
translate_percent={"x": (-0.2, 0.2), "y": (-0.2, 0.2)},
rotate=(-25, 25),
shear=(-8, 8)
),
iaa.Multiply((0.9, 1.1))
])
```
### Now it's time to train the model. Note that training even a basic model can take a few hours.
Note: the following model is for demonstration purpose only. We have limited the training to one epoch, and have set nominal values for the Detector Configuration to reduce run-time.
- dataset_train and dataset_val are derived from DetectorDataset
- DetectorDataset loads images from image filenames and masks from the annotation data
- model is Mask-RCNN
```
NUM_EPOCHS = 1
# Train Mask-RCNN Model
import warnings
warnings.filterwarnings("ignore")
model.train(dataset_train, dataset_val,
learning_rate=config.LEARNING_RATE,
epochs=NUM_EPOCHS,
layers='all',
augmentation=augmentation)
# select trained model
dir_names = next(os.walk(model.model_dir))[1]
key = config.NAME.lower()
dir_names = filter(lambda f: f.startswith(key), dir_names)
dir_names = sorted(dir_names)
if not dir_names:
import errno
raise FileNotFoundError(
errno.ENOENT,
"Could not find model directory under {}".format(self.model_dir))
fps = []
# Pick last directory
for d in dir_names:
dir_name = os.path.join(model.model_dir, d)
# Find the last checkpoint
checkpoints = next(os.walk(dir_name))[2]
checkpoints = filter(lambda f: f.startswith("mask_rcnn"), checkpoints)
checkpoints = sorted(checkpoints)
if not checkpoints:
print('No weight files in {}'.format(dir_name))
else:
checkpoint = os.path.join(dir_name, checkpoints[-1])
fps.append(checkpoint)
model_path = sorted(fps)[-1]
print('Found model {}'.format(model_path))
class InferenceConfig(DetectorConfig):
GPU_COUNT = 1
IMAGES_PER_GPU = 1
inference_config = InferenceConfig()
# Recreate the model in inference mode
model = modellib.MaskRCNN(mode='inference',
config=inference_config,
model_dir=MODEL_DIR)
# Load trained weights (fill in path to trained weights here)
assert model_path != "", "Provide path to trained weights"
print("Loading weights from ", model_path)
model.load_weights(model_path, by_name=True)
# set color for class
def get_colors_for_class_ids(class_ids):
colors = []
for class_id in class_ids:
if class_id == 1:
colors.append((.941, .204, .204))
return colors
```
### How does the predicted box compared to the expected value? Let's use the validation dataset to check.
Note that we trained only one epoch for **demonstration purposes ONLY**. You might be able to improve performance running more epochs.
```
# Show few example of ground truth vs. predictions on the validation dataset
dataset = dataset_val
fig = plt.figure(figsize=(10, 30))
for i in range(4):
image_id = random.choice(dataset.image_ids)
original_image, image_meta, gt_class_id, gt_bbox, gt_mask =\
modellib.load_image_gt(dataset_val, inference_config,
image_id, use_mini_mask=False)
plt.subplot(6, 2, 2*i + 1)
visualize.display_instances(original_image, gt_bbox, gt_mask, gt_class_id,
dataset.class_names,
colors=get_colors_for_class_ids(gt_class_id), ax=fig.axes[-1])
plt.subplot(6, 2, 2*i + 2)
results = model.detect([original_image]) #, verbose=1)
r = results[0]
visualize.display_instances(original_image, r['rois'], r['masks'], r['class_ids'],
dataset.class_names, r['scores'],
colors=get_colors_for_class_ids(r['class_ids']), ax=fig.axes[-1])
# Get filenames of test dataset DICOM images
test_image_fps = get_dicom_fps(test_dicom_dir)
```
### Final steps - Create the submission file
```
# Make predictions on test images, write out sample submission
def predict(image_fps, filepath='sample_submission.csv', min_conf=0.98):
# assume square image
resize_factor = ORIG_SIZE / config.IMAGE_SHAPE[0]
with open(filepath, 'w') as file:
for image_id in tqdm(image_fps):
ds = pydicom.read_file(image_id)
image = ds.pixel_array
# If grayscale. Convert to RGB for consistency.
if len(image.shape) != 3 or image.shape[2] != 3:
image = np.stack((image,) * 3, -1)
patient_id = os.path.splitext(os.path.basename(image_id))[0]
results = model.detect([image])
r = results[0]
out_str = ""
out_str += patient_id
assert( len(r['rois']) == len(r['class_ids']) == len(r['scores']) )
if len(r['rois']) == 0:
pass
else:
num_instances = len(r['rois'])
out_str += ","
for i in range(num_instances):
if r['scores'][i] > min_conf:
out_str += ' '
out_str += str(round(r['scores'][i], 2))
out_str += ' '
# x1, y1, width, height
x1 = r['rois'][i][1]
y1 = r['rois'][i][0]
width = r['rois'][i][3] - x1
height = r['rois'][i][2] - y1
bboxes_str = "{} {} {} {}".format(x1*resize_factor, y1*resize_factor, \
width*resize_factor, height*resize_factor)
out_str += bboxes_str
file.write(out_str+"\n")
# predict only the first 50 entries
sample_submission_fp = 'sample_submission.csv'
predict(test_image_fps[:50], filepath=sample_submission_fp)
output = pd.read_csv(sample_submission_fp, names=['id', 'pred_string'])
output.head(50)
```
| github_jupyter |
## Linear least-squares and a bland dense network
We're going to use the MIT-BIH datasets to train and test a basic feedforward network and see how it does. We'll compare the results to a linear regression.
We'll use two different inputs: a mostly unprocessed version of the dataset, and a version in the frequency domain obtained by applying the FFT.
```
import datetime
import os
import logging
import numpy as np
import tensorflow as tf
import tools.plot as plot
import tools.train as train
import tools.models as models
## Read in data
files = ("../data/mitbih_train.csv", "../data/mitbih_test.csv")
inputs, labels, sparse_labels, df = train.preprocess(*files, fft=False)
inputs_fft = train.dataset_fft(inputs)
train.class_count(df)
```
Let's look at a few random samples of the training data:
```
plot.plot_ecg(files[0], 125, 1)
```
### Least-squares
Let's try least-squares regression with numpy.
```
lstsq_soln = np.linalg.lstsq(inputs["train"], labels["train"], rcond=None)
lstsq_soln_fft = np.linalg.lstsq(inputs_fft["train"], labels["train"], rcond=None)
print("Rank of training dataset:", lstsq_soln[2])
print("Rank of training dataset after (real) FFT:", lstsq_soln_fft[2])
```
Now let's see how accurate it is.
```
def lstsq_accuracy(inputs, labels, coeffs):
predict = {}
accuracy = {}
for key in inputs:
predict[key] = np.argmax(np.dot(inputs[key], coeffs), axis=1)
num_correct = np.sum(
labels[key][range(labels[key].shape[0]), predict[key]] == 1
)
accuracy[key] = num_correct / labels[key].shape[0]
print("Training accuracy:", accuracy["train"])
print("Test accuracy:", accuracy["test"])
return predict
print("Regular least-squares")
predict = lstsq_accuracy(inputs, labels, lstsq_soln[0])
plot.plot_cm(sparse_labels["test"], predict["test"], classes=np.arange(5), normalize=True)
print("After FFT")
predict_fft = lstsq_accuracy(inputs_fft, labels, lstsq_soln_fft[0])
plot.plot_cm(sparse_labels["test"], predict_fft["test"], classes=np.arange(5), normalize=True)
```
### Dense feed-forward network
Let's try an unregularized, bland feed-forward network with a couple of hidden layers.
```
# Tensorboard logging
rightnow = datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
nofftpath = os.path.join("..", "logs", rightnow, "nofft")
config = {
"optimizer": "Nadam",
"loss": "categorical_crossentropy",
"batch_size": 200,
"val_split": 0.05,
"epochs": 300,
"verbose": 0,
"patience": 20,
"logdir": nofftpath,
}
inputsize = inputs["train"].shape[1]
ncategories = labels["train"].shape[1]
hiddenlayers = [(100, "relu")]
# Suppress tensorflow warnings about internal deprecations
logger = tf.get_logger()
logger.setLevel(logging.ERROR)
print("Unprocessed data")
model = models.create_dense(inputsize, hiddenlayers, ncategories)
history = train.train_print(model, inputs, labels, config)
plot.plot_fit_history(history)
test_pred = np.argmax(model.predict(inputs["test"]), axis=1)
plot.plot_cm(
sparse_labels["test"],
test_pred,
classes=np.array(["N", "S", "V", "F", "Q"]),
normalize=True,
)
# Tensorboard logging
fftpath = os.path.join("..", "logs", rightnow, "fft")
config_fft = config
config_fft["logdir"] = fftpath
print("After FFT")
model_fft = models.create_dense(inputs_fft["train"].shape[1], hiddenlayers, ncategories)
history_fft = train.train_print(model_fft, inputs_fft, labels, config_fft)
plot.plot_fit_history(history_fft)
test_pred_fft = np.argmax(model_fft.predict(inputs_fft["test"]), axis=1)
plot.plot_cm(
sparse_labels["test"],
test_pred_fft,
classes=np.array(["N", "S", "V", "F", "Q"]),
normalize=True,
)
```
The results don't tend to be very consistent. The final test accuracy varies from run to run generally fairly significantly and it's not clear if the FFT "does" anything for the accuracy of the training.
| github_jupyter |
```
import requests
import altair as alt
import pandas as pd
import sys
import warnings
from collections import OrderedDict
from datetime import datetime, timezone
from io import StringIO
%load_ext watermark
%watermark -iv -v
```
### Load latest data
```
try:
url = 'https://raw.githubusercontent.com/owid/covid-19-data/master/public/data/vaccinations/vaccinations.csv'
r = requests.get(url, allow_redirects=True)
df = pd.read_csv(StringIO(r.content.decode("utf-8")))[['location', 'date', 'total_vaccinations_per_hundred']]
max_values = df.groupby('location').max('total_vaccinations_per_hundred').sort_values('total_vaccinations_per_hundred', ascending=False).reset_index(drop=False)
print(','.join(sorted(df.location.unique().tolist())))
display(df.sample(5))
except:
warnings.warn('Cannot read vaccination data to pandas dataframe!')
```
### Plot routine
```
def plot_chart(df, cmap):
return alt.Chart(df[(df.location.isin(cmap.keys())) & (~df.total_vaccinations_per_hundred.isna())]).mark_line(point=True, strokeWidth=2).encode(
x = alt.X('date:T', title='Date'),
y = alt.Y('total_vaccinations_per_hundred:Q', title='Vaccinated [%]'),
color = alt.Color('location:N', title='Country', scale=alt.Scale(
domain = list(cmap.keys()),
range = list(cmap.values())
)), tooltip = ['location', 'date:T', 'total_vaccinations_per_hundred']
).configure_point(
size=50
).properties(width=300*1.61, height=300, title=f'Vaccination status {datetime.now().astimezone(timezone.utc).strftime("%Y-%m-%d %H:%M")} (UTC)')
```
### Main countries
```
country_vs_color = OrderedDict({
'Israel': '#1A85FF',
'Bahrain': '#FEFE62',
'United Kingdom': '#40B0A6',
'United States': '#4B0092',
'Canada': '#D35FB7',
'China': '#DC3220',
'Russia': '#994F00',
'European Union': '#E66100',
'Japan': '#E1BE6A',
'Australia': '#000000',
'India': '#D35FB7'
})
cmap = OrderedDict({k:country_vs_color[k] for k in [x for x in max_values.location if x in country_vs_color.keys()]})
print('Main countries: ', ",".join(list(cmap.keys())))
chart = plot_chart(df, cmap)
chart.save('blah.png', log_path = 'out')
display(chart)
#chart.save('out/main-countries.png')
df[(df.location=='United Kingdom') & (~df.total_vaccinations_per_hundred.isna())]
```
### European countries
```
country_vs_color = OrderedDict({
'Austria': '#000000',
'Bulgaria': '#004949',
'Croatia': '#009292',
'Denmark': '#ff6db6',
'Estonia': '#ffb6db',
'France': '#490092',
'Germany': '#006ddb',
'Greece': '#b66dff',
'Hungary': '#6db6ff',
'Italy': '#b6dbff',
'Latvia': '#920000',
'Lithuania': '#924900',
'Luxembourg': '#db6d00',
'Poland': '#24ff24',
'Portugal': '#ffff6d',
'Romania': '#000000'
})
cmap = OrderedDict({k:country_vs_color[k] for k in [x for x in max_values.location if x in country_vs_color.keys()]})
print('European countries: ', ",".join(list(cmap.keys())))
chart = plot_chart(df, cmap)
display(chart)
#chart.save('out/european-countries.png')
```
| github_jupyter |
```
from pathlib import Path
import pandas as pd
import numpy as np
DATA_DIR = (Path.cwd() / ".." / "Data").resolve()
assert (DATA_DIR / "CH 2020-2021.csv").exists()
assert (DATA_DIR / "metervalues_pseudonymized_1_neu.csv").exists()
assert (DATA_DIR / "metervalues_pseudonymized_2_neu.csv").exists()
carbon = pd.DataFrame(pd.read_csv(DATA_DIR / "CH 2020-2021.csv"))
mvs_1 = pd.DataFrame(pd.read_csv(DATA_DIR / "metervalues_pseudonymized_1_neu.csv", delimiter=';', low_memory=False))
mvs_2 = pd.DataFrame(pd.read_csv(DATA_DIR / "metervalues_pseudonymized_2_neu.csv", delimiter=';', low_memory=False))
carbon.head()
mvs_1.head()
mvs_2.head()
mvs = pd.concat([mvs_1, mvs_2], ignore_index=True)
mvs.head()
df = mvs.astype({"Chargepoint": str, "connector": str})
df['unique_charge_point'] = df['Chargepoint'] + df['connector']
df.head()
df['unique_charge_point'].nunique()
df['increment'].dtype
df['increment'] = pd.to_numeric(df['increment'])
df[696713:696715]
df
df = df.drop(696713)
df['increment'] = pd.to_numeric(df['increment'])
df['increment'].dtype
df[696713:696715]
df['increment'] > 5500
l = df[df['increment'] > 5500]
l['charge_log_id'].nunique
l_list = l['charge_log_id'].tolist()
l_list
df_new = df[~df['charge_log_id'].isin(l_list)]
df_new.head()
df_new
df_new[df_new['increment'] > 5500]
df_new
df_new.loc[df_new['charge_log_id'] == '2188']
from datetime import datetime
datetime_object = datetime.strptime('01.01.2021 00:15', '%d.%m.%Y %H:%M')
datetime_object.timestamp()
df_new['n_timestamp'] = df_new.apply(lambda t: datetime.strptime(t['timestamp'], '%d.%m.%Y %H:%M').timestamp(), axis=1)
df_sorted = df_new.sort_values('n_timestamp')
df_sorted.head()
#Warning! Very slow (sorry but I wasn't bothered to make it fast)
charge_ids = list(set(df_sorted['charge_log_id']))
dataframes = {}
for charge_id in charge_ids:
df_charges = df_sorted.loc[df_sorted['charge_log_id'] == charge_id]
dataframes[charge_id] = df_charges
dataframes
times = []
for i in charge_ids:
times.append((dataframes[i]['n_timestamp'].max() - dataframes[i]['n_timestamp'].min())/ 3600)
import seaborn as sns
sns.histplot(times)
to_drop = {}
for (c_id, df) in dataframes.items():
powers = df['increment']
starting_valley = True
second_valley = False
for inc in powers:
if starting_valley and inc >= 25:
starting_valley = False
continue
if not starting_valley and inc == 0:
second_valley = True
continue
if second_valley and inc >= 25:
to_drop[c_id] = df
break
print(len(to_drop))
len(dataframes)
len([val for val in to_drop.values() if val['increment'].iloc[-1] >= 25])
df_sorted.loc[~df_sorted['charge_log_id'].isin(to_drop.keys())].to_csv(DATA_DIR / "seperated_meter_data.csv")
```
| github_jupyter |
### Data Visualization
#### `matplotlib` - from the documentation:
https://matplotlib.org/3.1.1/tutorials/introductory/pyplot.html
`matplotlib.pyplot` is a collection of command style functions that make matplotlib work like MATLAB. <br>
Each pyplot function makes some change to a figure: e.g., creates a figure, creates a plotting area in a figure, plots some lines in a plotting area, decorates the plot with labels, etc.
In `matplotlib.pyplot` various states are preserved across function calls, so that it keeps track of things like the current figure and plotting area, and the plotting functions are directed to the current axes.<br>
"axes" in most places in the documentation refers to the axes part of a figure and not the strict mathematical term for more than one axis).
```
%matplotlib inline
import matplotlib.pyplot as plt
```
Call signatures::
```
plot([x], y, [fmt], data=None, **kwargs)
plot([x], y, [fmt], [x2], y2, [fmt2], ..., **kwargs)
```
Quick plot
The main usage of `plt` is the `plot()` and `show()` functions
```
plt.plot()
plt.show()
```
List
```
plt.plot([8, 24, 27, 42])
plt.ylabel('numbers')
plt.show()
# Plot the two lists, add axes labels
x=[4,5,6,7]
y=[2,5,1,7]
```
`matplotlib` can use *format strings* to quickly declare the type of plots you want. Here are *some* of those formats:
|**Character**|**Description**|
|:-----------:|:--------------|
|'--'|Dashed line|
|':'|Dotted line|
|'o'|Circle marker|
|'^'|Upwards triangle marker|
|'b'|Blue|
|'c'|Cyan|
|'g'|Green|
```
plt.plot([3, 4, 9, 20], 'gs')
plt.axis([-1, 4, 0, 22])
plt.show()
plt.plot([3, 4, 9, 20], 'b^--', linewidth=2, markersize=12)
plt.show()
plt.plot([3, 4, 9, 20], color='blue', marker='^', linestyle='dashed', linewidth=2, markersize=12)
plt.show()
# Plot a list with 10 numbers with a magenta dotted line and circles for points.
import numpy as np
# evenly sampled time
time = np.arange(0, 7, 0.3)
# gene expression
ge = np.arange(1, 8, 0.3)
# red dashes, blue squares and green triangles
plt.plot(time, ge, 'r--', time, ge**2, 'bs', time, ge**3, 'g^')
plt.show()
```
linestyle or ls [ '-' | '--' | '-.' | ':' |
```
lines = plt.plot([1, 2, 3])
plt.setp(lines)
names = ['A', 'B', 'C', 'D']
values = [7, 20, 33, 44]
values1 = np.random.rand(100)
plt.figure(figsize=(9, 3))
plt.subplot(131)
plt.bar(names, values)
plt.subplot(132)
plt.scatter(names, values)
plt.subplot(133)
plt.hist(values1)
plt.suptitle('Categorical Plotting')
plt.show()
import pandas as pd
df_iris = pd.read_csv('https://raw.githubusercontent.com/uiuc-cse/data-fa14/gh-pages/data/iris.csv')
df_iris.head()
x1 = df_iris.petal_length
y1 = df_iris.petal_width
x2 = df_iris.sepal_length
y2 = df_iris.sepal_width
plt.plot(x1, y1, 'g^', x2, y2, 'bs')
plt.show()
```
#### Histogram
```
help(plt.hist)
n, bins, patches = plt.hist(df_iris.petal_length, bins=20,facecolor='#8303A2', alpha=0.8, rwidth=.8, align='mid')
# Add a title
plt.title('Iris dataset petal length')
# Add y axis label
plt.ylabel('number of plants')
```
#### Boxplot
```
help(plt.boxplot)
plt.boxplot(df_iris.petal_length)
# Add a title
plt.title('Iris dataset petal length')
# Add y axis label
plt.ylabel('petal length')
```
The biggest issue with `matplotlib` isn't its lack of power...it is that it is too much power. With great power, comes great responsibility. When you are quickly exploring data, you don't want to have to fiddle around with axis limits, colors, figure sizes, etc. Yes, you *can* make good figures with `matplotlib`, but you probably won't.
https://python-graph-gallery.com/matplotlib/
Pandas works off of `matplotlib` by default. You can easily start visualizing dataframs and series just by a simple command.
#### Using pandas `.plot()`
Pandas abstracts some of those initial issues with data visualization. However, it is still a `matplotlib` plot</br></br>
Every plot that is returned from `pandas` is subject to `matplotlib` modification.
```
df_iris.plot.box()
plt.show()
# Plot the histogram of the petal lengths
# Plot the histograms of all 4 numerical characteristics in a plot
df_iris.groupby("species")['petal_length'].mean().plot(kind='bar')
plt.show()
df_iris.plot(x='petal_length', y='petal_width', kind = "scatter")
plt.savefig('output.png')
plt.savefig('output.png')
```
https://github.com/pandas-dev/pandas/blob/v0.25.0/pandas/plotting/_core.py#L504-L1533
#### Multiple Plots
```
df_iris.petal_length.plot(kind='density')
df_iris.sepal_length.plot(kind='density')
df_iris.petal_width.plot(kind='density')
```
`matplotlib` allows users to define the regions of their plotting canvas. If the user intends to create a canvas with multiple plots, they would use the `subplot()` function. The `subplot` function sets the number of rows and columns the canvas will have **AND** sets the current index of where the next subplot will be rendered.
```
plt.figure(1)
# Plot all three columns from df in different subplots
# Rows first index (top-left)
plt.subplot(3, 1, 1)
df_iris.petal_length.plot(kind='density')
plt.subplot(3, 1, 2)
df_iris.sepal_length.plot(kind='density')
plt.subplot(3, 1, 3)
df_iris.petal_width.plot(kind='density')
# Some plot configuration
plt.subplots_adjust(top=.92, bottom=.08, left=.1, right=.95, hspace=.25, wspace=.35)
plt.show()
# Temporary styles
with plt.style.context(('ggplot')):
plt.figure(1)
# Plot all three columns from df in different subplots
# Rows first index (top-left)
plt.subplot(3, 1, 1)
df_iris.petal_length.plot(kind='density')
plt.subplot(3, 1, 2)
df_iris.sepal_length.plot(kind='density')
plt.subplot(3, 1, 3)
df_iris.petal_width.plot(kind='density')
# Some plot configuration
plt.subplots_adjust(top=.92, bottom=.08, left=.1, right=.95, hspace=.25, wspace=.35)
plt.show()
# Plot the histograms of the petal length and width and sepal length and width
# Display them on the columns of a figure with 2X2 subplots
# color them red, green, blue and yellow, respectivelly
```
### `seaborn` - dataset-oriented plotting
Seaborn is a library that specializes in making *prettier* `matplotlib` plots of statistical data. <br>
It is built on top of matplotlib and closely integrated with pandas data structures.
https://seaborn.pydata.org/introduction.html<br>
https://python-graph-gallery.com/seaborn/
```
import seaborn as sns
```
`seaborn` lets users *style* their plotting environment.
```
sns.set(style='whitegrid')
```
However, you can always use `matplotlib`'s `plt.style`
```
#dir(sns)
sns.scatterplot(x='petal_length',y='petal_width',data=df_iris)
plt.show()
sns.scatterplot(x='petal_length',y='petal_width', hue = "species",data=df_iris)
plt.show()
```
#### Violin plot
Fancier box plot that gets rid of the need for 'jitter' to show the inherent distribution of the data points
```
columns = ['petal_length', 'petal_width', 'sepal_length']
fig, axes = plt.subplots(figsize=(10, 10))
sns.violinplot(data=df_iris.loc[:,columns], ax=axes)
axes.set_ylabel('number')
axes.set_xlabel('columns', )
plt.show()
```
#### Distplot
```
sns.set(style='darkgrid', palette='muted')
# 1 row, 3 columns
f, axes = plt.subplots(4,1, figsize=(10,10), sharex=True)
sns.despine(left=True)
# Regular displot
sns.distplot(df_iris.petal_length, ax=axes[0])
# Change the color
sns.distplot(df_iris.petal_width, kde=False, ax=axes[1], color='orange')
# Show the Kernel density estimate
sns.distplot(df_iris.sepal_width, hist=False, kde_kws={'shade':True}, ax=axes[2], color='purple')
# Show the rug
sns.distplot(df_iris.sepal_length, hist=False, rug=True, ax=axes[3], color='green')
```
#### FacetGrid
```
sns.set()
columns = ['species', 'petal_length', 'petal_width']
facet_column = 'species'
g = sns.FacetGrid(df_iris.loc[:,columns], col=facet_column, hue=facet_column, col_wrap=5)
g.map(plt.scatter, 'petal_length', 'petal_width')
sns.relplot(x="petal_length", y="petal_width", col="species",
hue="species", style="species", size="species",
data=df_iris)
plt.show()
```
https://jakevdp.github.io/PythonDataScienceHandbook/04.14-visualization-with-seaborn.html
### `plotnine` - R ggplot2 in python
plotnine is an implementation of a grammar of graphics in Python, it is based on ggplot2. The grammar allows users to compose plots by explicitly mapping data to the visual objects that make up the plot.
Plotting with a grammar is powerful, it makes custom (and otherwise complex) plots are easy to think about and then create, while the simple plots remain simple.
```
!pip install plotnine
```
https://plotnine.readthedocs.io/en/stable/
```
from plotnine import *
ggplot(data=df_iris) + aes(x="petal_length", y = "petal_width") + geom_point()
# add transparency - to avoid over plotting
ggplot(data=df_iris) + aes(x="petal_length", y = "petal_width") + geom_point(alpha=0.7)
# change point size
ggplot(data=df_iris) + aes(x="petal_length", y = "petal_width") + geom_point(size = 0.7, alpha=0.7)
# more parameters
ggplot(data=df_iris) + aes(x="petal_length", y = "petal_width") + geom_point() + scale_x_log10() + xlab("Petal Length")
n = "3"
ft = "length and width"
title = 'species : %s, petal : %s' % (n,ft)
ggplot(data=df_iris) +aes(x='petal_length',y='petal_width',color="species") + geom_point(size=0.7,alpha=0.7) + facet_wrap('~species',nrow=3) + theme(figure_size=(9,5)) + ggtitle(title)
p = ggplot(data=df_iris) + aes(x='petal_length') + geom_histogram(binwidth=1,color='black',fill='grey')
p
ggsave(plot=p, filename='hist_plot_with_plotnine.png')
```
http://cmdlinetips.com/2018/05/plotnine-a-python-library-to-use-ggplot2-in-python/ <br>
https://www.rstudio.com/wp-content/uploads/2015/03/ggplot2-cheatsheet.pdf
<img src = "https://www.rstudio.com/wp-content/uploads/2015/03/ggplot2-cheatsheet.pdf" width = "1000"/>
Use ggplot to plot the sepal_length in boxplots separated by species, add new axes labels and make the y axis values log10.
* Write a function that takes as a parameter a line of the dataframe and if the species is
** setosa it returns the petal_length
** versicolor it returns the petal_width
** virginica it returns the sepal_length
Apply this function to every line in the dataset in a for loop and save the result in an array
Use ggplot to make a histogram of the values
| github_jupyter |
# NumPy Basics: Arrays and Vectorized Computation
```
import numpy as np
np.random.seed(12345)
import matplotlib.pyplot as plt
plt.rc('figure', figsize=(10, 6))
np.set_printoptions(precision=4, suppress=True)
import numpy as np
my_arr = np.arange(1000000)
my_list = list(range(1000000))
%time for _ in range(10): my_arr2 = my_arr * 2
%time for _ in range(10): my_list2 = [x * 2 for x in my_list]
```
## The NumPy ndarray: A Multidimensional Array Object
```
import numpy as np
# Generate some random data
data = np.random.randn(2, 3)
data
data * 10
data + data
data.shape
data.dtype
```
### Creating ndarrays
```
data1 = [6, 7.5, 8, 0, 1]
arr1 = np.array(data1)
arr1
data2 = [[1, 2, 3, 4], [5, 6, 7, 8]]
arr2 = np.array(data2)
arr2
arr2.ndim
arr2.shape
arr1.dtype
arr2.dtype
np.zeros(10)
np.zeros((3, 6))
np.empty((2, 3, 2))
np.arange(15)
```
### Data Types for ndarrays
```
arr1 = np.array([1, 2, 3], dtype=np.float64)
arr2 = np.array([1, 2, 3], dtype=np.int32)
arr1.dtype
arr2.dtype
arr = np.array([1, 2, 3, 4, 5])
arr.dtype
float_arr = arr.astype(np.float64)
float_arr.dtype
arr = np.array([3.7, -1.2, -2.6, 0.5, 12.9, 10.1])
arr
arr.astype(np.int32)
numeric_strings = np.array(['1.25', '-9.6', '42'], dtype=np.string_)
numeric_strings.astype(float)
int_array = np.arange(10)
calibers = np.array([.22, .270, .357, .380, .44, .50], dtype=np.float64)
int_array.astype(calibers.dtype)
empty_uint32 = np.empty(8, dtype='u4')
empty_uint32
```
### Arithmetic with NumPy Arrays
```
arr = np.array([[1., 2., 3.], [4., 5., 6.]])
arr
arr * arr
arr - arr
1 / arr
arr ** 0.5
arr2 = np.array([[0., 4., 1.], [7., 2., 12.]])
arr2
arr2 > arr
```
### Basic Indexing and Slicing
```
arr = np.arange(10)
arr
arr[5]
arr[5:8]
arr[5:8] = 12
arr
arr_slice = arr[5:8]
arr_slice
arr_slice[1] = 12345
arr
arr_slice[:] = 64
arr
arr2d = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
arr2d[2]
arr2d[0][2]
arr2d[0, 2]
arr3d = np.array([[[1, 2, 3], [4, 5, 6]], [[7, 8, 9], [10, 11, 12]]])
arr3d
arr3d[0]
old_values = arr3d[0].copy()
arr3d[0] = 42
arr3d
arr3d[0] = old_values
arr3d
arr3d[1, 0]
x = arr3d[1]
x
x[0]
```
#### Indexing with slices
```
arr
arr[1:6]
arr2d
arr2d[:2]
arr2d[:2, 1:]
arr2d[1, :2]
arr2d[:2, 2]
arr2d[:, :1]
arr2d[:2, 1:] = 0
arr2d
```
### Boolean Indexing
```
names = np.array(['Bob', 'Joe', 'Will', 'Bob', 'Will', 'Joe', 'Joe'])
data = np.random.randn(7, 4)
names
data
names == 'Bob'
data[names == 'Bob']
print(data[names == 'Bob', 2:])
print(data[names == 'Bob', 3])
names != 'Bob'
data[~(names == 'Bob')]
cond = names == 'Bob'
data[~cond]
mask = (names == 'Bob') | (names == 'Will')
print(mask)
data[mask]
data[data < 0] = 0
data
data[names != 'Joe'] = 7
data
```
### Fancy Indexing
```
arr = np.empty((8, 4))
for i in range(8):
arr[i] = i
arr
arr[[4, 3, 0, 6]]
arr[[-3, -5, -7]]
arr = np.arange(32).reshape((8, 4))
print(arr)
arr[[1, 5, 7, 2], [0, 3, 1, 2]]
arr[[1, 5, 7, 2]][:, [0, 3, 1, 2]]
```
### Transposing Arrays and Swapping Axes
```
arr = np.arange(15).reshape((3, 5))
print(arr)
arr.T
arr = np.random.randn(6, 3)
print(arr)
np.dot(arr.T, arr)
arr = np.arange(16).reshape((2, 2, 4))
print(arr)
arr.transpose((1, 0, 2))
arr
arr.swapaxes(1, 2)
```
## Universal Functions: Fast Element-Wise Array Functions
```
arr = np.arange(10)
print(arr)
print(np.sqrt(arr))
np.exp(arr)
x = np.random.randn(8)
y = np.random.randn(8)
x
y
np.maximum(x, y)
arr = np.random.randn(7) * 5
arr
remainder, whole_part = np.modf(arr)
remainder
whole_part
arr
np.sqrt(arr)
np.sqrt(arr, arr)
arr
```
## 4.3Array-Oriented Programming with Arrays
```
points = np.arange(-5, 5, 0.01) # 1000 equally spaced points
xs, ys = np.meshgrid(points, points)
ys
z = np.sqrt(xs ** 2 + ys ** 2)
z
import matplotlib.pyplot as plt
plt.imshow(z, cmap=plt.cm.gray); plt.colorbar()
plt.title("Image plot of $\sqrt{x^2 + y^2}$ for a grid of values")
plt.draw()
plt.close('all')
```
### Expressing Conditional Logic as Array Operations
```
import numpy as np
xarr = np.array([1.1, 1.2, 1.3, 1.4, 1.5])
yarr = np.array([2.1, 2.2, 2.3, 2.4, 2.5])
cond = np.array([True, False, True, True, False])
result = [(x if c else y)
for x, y, c in zip(xarr, yarr, cond)]
result
result = np.where(cond, xarr, yarr)
result
arr = np.random.randn(4, 4)
print(arr)
arr > 0
np.where(arr > 0, 2, -2)
np.where(arr > 0, 2, arr) # set only positive values to 2
```
### Mathematical and Statistical Methods
```
arr = np.random.randn(5, 4)
arr
arr.mean()
np.mean(arr)
arr.sum()
arr.mean(axis=1)
arr.sum(axis=0)
arr = np.array([0, 1, 2, 3, 4, 5, 6, 7])
arr.cumsum()
arr = np.array([[0, 1, 2], [3, 4, 5], [6, 7, 8]])
arr
arr.cumsum(axis=0)
arr.cumprod(axis=1)
```
### Methods for Boolean Arrays
```
arr = np.random.randn(100)
(arr > 0).sum() # Number of positive values
bools = np.array([False, False, True, False])
print(bools.any())
bools.all()
```
### Sorting
```
arr = np.random.randn(6)
arr
arr.sort()
arr
arr = np.random.randn(5, 3)
arr
arr.sort(1)
arr
large_arr = np.random.randn(1000)
large_arr.sort()
large_arr[int(0.05 * len(large_arr))] # 5% quantile
```
### Unique and Other Set Logic
```
names = np.array(['Bob', 'Joe', 'Will', 'Bob', 'Will', 'Joe', 'Joe'])
np.unique(names)
ints = np.array([3, 3, 3, 2, 2, 1, 1, 4, 4])
np.unique(ints)
sorted(set(names))
values = np.array([6, 0, 0, 3, 2, 5, 6])
np.in1d(values, [2, 3, 6])
```
## File Input and Output with Arrays
```
arr = np.arange(10)
np.save('some_array', arr)
np.load('some_array.npy')
np.savez('array_archive.npz', a=arr, b=arr)
arch = np.load('array_archive.npz')
arch['b']
np.savez_compressed('arrays_compressed.npz', a=arr, b=arr)
!rm some_array.npy
!rm array_archive.npz
!rm arrays_compressed.npz
```
## Linear Algebra
```
x = np.array([[1., 2., 3.], [4., 5., 6.]])
y = np.array([[6., 23.], [-1, 7], [8, 9]])
x
y
x.dot(y)
np.dot(x, y)
np.dot(x, np.ones(3))
x @ np.ones(3)
from numpy.linalg import inv, qr
X = np.random.randn(5, 5)
mat = X.T.dot(X)
inv(mat)
mat.dot(inv(mat))
q, r = qr(mat)
r
```
## Pseudorandom Number Generation
```
samples = np.random.normal(size=(4, 4))
samples
from random import normalvariate
N = 1000000
%timeit samples = [normalvariate(0, 1) for _ in range(N)]
%timeit np.random.normal(size=N)
np.random.seed(1234)
rng = np.random.RandomState(1234)
rng.randn(10)
```
## Example: Random Walks
```
import random
position = 0
walk = [position]
steps = 1000
for i in range(steps):
step = 1 if random.randint(0, 1) else -1
position += step
walk.append(position)
plt.figure()
plt.plot(walk[:100])
np.random.seed(12345)
nsteps = 1000
draws = np.random.randint(0, 2, size=nsteps)
steps = np.where(draws > 0, 1, -1)
walk = steps.cumsum()
walk.min()
walk.max()
(np.abs(walk) >= 10).argmax()
```
### Simulating Many Random Walks at Once
```
nwalks = 5000
nsteps = 1000
draws = np.random.randint(0, 2, size=(nwalks, nsteps)) # 0 or 1
steps = np.where(draws > 0, 1, -1)
walks = steps.cumsum(1)
walks
walks.max()
walks.min()
hits30 = (np.abs(walks) >= 30).any(1)
hits30
hits30.sum() # Number that hit 30 or -30
crossing_times = (np.abs(walks[hits30]) >= 30).argmax(1)
crossing_times.mean()
steps = np.random.normal(loc=0, scale=0.25,
size=(nwalks, nsteps))
```
## Conclusion
| github_jupyter |
<img src="images/qiskit-heading.gif" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="250 px" align="left">
## _*E91 quantum key distribution protocol*_
***
### Contributors
Andrey Kardashin
## *Introduction*
Suppose that Alice wants to send a message to Bob.
In order to protect the information in the message from the eavesdropper Eve, it must be encrypted.
Encryption is the process of encoding the *plaintext* into *ciphertext*.
The strength of encryption, that is, the property to resist decryption, is determined by its algorithm.
Any encryption algorithm is based on the use of a *key*.
In order to generate the ciphertext, the [one-time pad technique](https://en.wikipedia.org/wiki/One-time_pad) is usually used.
The idea of this technique is to apply the *exclusive or* (XOR) $\oplus$ operation to bits of the plaintext and bits of the key to obtain the ciphertext.
Thus, if $m=(m_1 \ldots m_n)$, $c=(c_1 \ldots c_n)$ and $k=(k_1 \ldots k_n)$ are binary strings of plaintext, ciphertext and key respectively, then the encryption is defined as $c_i=m_i \oplus k_i$, and decryption as $m_i=c_i \oplus k_i$.

The one-time pad method is proved to be be absolutely secure.
Thus, if Eve intercepted the ciphertext $c$, she will not get any information from the message $m$ until she has the key $k$.
The main problem of modern cryptographic systems is the distribution among the participants of the communication session of a secret key, possession of which should not be available to third parties.
The rapidly developing methods of quantum key distribution can solve this problem regardless of the capabilities of the eavesdropper.
In this tutorial, we show how Alice and Bob can generate a secret key using the *E91* quantum key distribution protocol.
## *Quantum entanglement*
The E91 protocol developed by Artur Ekert in 1991 is based on the use of entangled states and Bell's theorem (see [Entanglement Revisited](https://github.com/Qiskit/qiskit-tutorial/blob/master/reference/qis/entanglement_revisited.ipynb) QISKit tutorial).
It is known that two electrons *A* and *B* can be prepared in such a state that they can not be considered separately from each other.
One of these states is the singlet state
$$\lvert\psi_s\rangle =
\frac{1}{\sqrt{2}}(\lvert0\rangle_A\otimes\lvert1\rangle_B - \lvert1\rangle_A\otimes\lvert0\rangle_B) =
\frac{1}{\sqrt{2}}(\lvert01\rangle - \lvert10\rangle),$$
where the vectors $\lvert 0 \rangle$ and $\lvert 1 \rangle$ describe the states of each electron with the [spin](https://en.wikipedia.org/wiki/Spin_(physics%29) projection along the positive and negative direction of the *z* axis.
The observable of the projection of the spin onto the direction $\vec{n}=(n_x, n_y, n_z)$ is given by
$$\vec{n} \cdot \vec{\sigma} =
n_x X + n_y Y + n_z Z,$$
where $\vec{\sigma} = (X, Y, Z)$ and $X, Y, Z$ are the Pauli matrices.
For two qubits *A* and *B*, the observable $(\vec{a} \cdot \vec{\sigma})_A \otimes (\vec{b} \cdot \vec{\sigma})_B$ describes the joint measurement of the spin projections onto the directions $\vec{a}$ and $\vec{b}$.
It can be shown that the expectation value of this observable in the singlet state is
$$\langle (\vec{a} \cdot \vec{\sigma})_A \otimes (\vec{b} \cdot \vec{\sigma})_B \rangle_{\psi_s} =
-\vec{a} \cdot \vec{b}. \qquad\qquad (1)$$
Here we see an interesting fact: if Alice and Bob measure the spin projections of electrons A and B onto the same direction, they will obtain the opposite results.
Thus, if Alice got the result $\pm 1$, then Bob *definitely* will get the result $\mp 1$, i.e. the results will be perfectly anticorrelated.
## *CHSH inequality*
In the framework of classical physics, it is impossible to create a correlation inherent in the singlet state $\lvert\psi_s\rangle$.
Indeed, let us measure the observables $X$, $Z$ for qubit *A* and observables $W = \frac{1}{\sqrt{2}} (X + Z)$, $V = \frac{1}{\sqrt{2}} (-X + Z)$ for qubit *B*.
Performing joint measurements of these observables, the following expectation values can be obtained:
\begin{eqnarray*}
\langle X \otimes W \rangle_{\psi_s} &= -\frac{1}{\sqrt{2}}, \quad
\langle X \otimes V \rangle_{\psi_s} &= \frac{1}{\sqrt{2}}, \qquad\qquad (2) \\
\langle Z \otimes W \rangle_{\psi_s} &= -\frac{1}{\sqrt{2}}, \quad
\langle Z \otimes V \rangle_{\psi_s} &= -\frac{1}{\sqrt{2}}.
\end{eqnarray*}
Now we can costruct the *Clauser-Horne-Shimony-Holt (CHSH) correlation value*:
$$C =
\langle X\otimes W \rangle - \langle X \otimes V \rangle + \langle Z \otimes W \rangle + \langle Z \otimes V \rangle =
-2 \sqrt{2}. \qquad\qquad (3)$$
The [local hidden variable theory](https://en.wikipedia.org/wiki/Local_hidden_variable_theory) which was developed in particular to explain the quantum correlations gives that $\lvert C \rvert \leqslant 2$.
But [Bell's theorem](https://en.wikipedia.org/wiki/Bell's_theorem) states that "no physical theory of local hidden variables can ever reproduce all of the predictions of quantum mechanics."
Thus, the violation of the [CHSH inequality](https://en.wikipedia.org/wiki/Bell's_theorem#Bell_inequalities_are_violated_by_quantum_mechanical_predictions) (i.e. $C = -2 \sqrt{2}$ for the singlet state), which is a generalized form of Bell's inequality, can serve as an *indicator of quantum entanglement*.
This fact finds its application in the E91 protocol.
## *The protocol*
To implement the E91 quantum key distribution protocol, there must be a source of qubits prepared in the singlet state.
It does not matter to whom this source belongs: to Alice, to Bob, to some trusted third-party Charlie or even to Eve.
The steps of the E91 protocol are following.
1. Charlie, the owner of the singlet state preparation device, creates $N$ entangled states $\lvert\psi_s\rangle$ and sends qubits *A* to Alice and qubits *B* to Bob via the quantum channel.

2. Participants Alice and Bob generate strings $b=(b_1 \ldots b_N)$ and $b^{'}=(b_1^{'} \ldots b_N^{'})$, where $b_i, b^{'}_j = 1, 2, 3$.
Depending on the elements of these strings, Alice and Bob measure the spin projections of their qubits along the following directions:
\begin{align*}
b_i = 1: \quad \vec{a}_1 &= (1,0,0) \quad (X \text{ observable}) &
b_j^{'} = 1: \quad \vec{b}_1 &= \left(\frac{1}{\sqrt{2}},0,\frac{1}{\sqrt{2}}\right) \quad (W \text{ observable})
\\
b_i = 2: \quad \vec{a}_2 &= \left(\frac{1}{\sqrt{2}},0,\frac{1}{\sqrt{2}}\right) \quad (W \text{ observable}) &
b_j^{'} = 2: \quad \vec{b}_2 &= (0,0,1) \quad ( \text{Z observable})
\\
b_i = 3: \quad \vec{a}_3 &= (0,0,1) \quad (Z \text{ observable}) &
b_j^{'} = 3: \quad \vec{b}_3 &= \left(-\frac{1}{\sqrt{2}},0,\frac{1}{\sqrt{2}}\right) \quad (V \text{ observable})
\end{align*}
<img src="images/vectors.png" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="center">
We can describe this process as a measurement of the observables $(\vec{a}_i \cdot \vec{\sigma})_A \otimes (\vec{b}_j \cdot \vec{\sigma})_B$ for each singlet state created by Charlie.
3. Alice and Bob record the results of their measurements as elements of strings $a=(a_1 \ldots a_N)$ and $a^{'} =(a_1^{'} \ldots a_N^{'})$ respectively, where $a_i, a^{'}_j = \pm 1$.
4. Using the classical channel, participants compare their strings $b=(b_1 \ldots b_N)$ and $b^{'}=(b_1^{'} \ldots b_N^{'})$.
In other words, Alice and Bob tell each other which measurements they have performed during the step 2.
If Alice and Bob have measured the spin projections of the $m$-th entangled pair of qubits onto the same direction (i.e. $\vec{a}_2/\vec{b}_1$ or $\vec{a}_3/\vec{b}_2$ for Alice's and Bob's qubit respectively), then they are sure that they obtained opposite results, i.e. $a_m = - a_m^{'}$ (see Eq. (1)).
Thus, for the $l$-th bit of the key strings $k=(k_1 \ldots k_n),k^{'}=(k_1^{'} \ldots k_n^{'})$ Alice and Bob can write $k_l = a_m, k_l^{'} = -a_m^{'}$.

5. Using the results obtained after measuring the spin projections onto the $\vec{a}_1/\vec{b}_1$, $\vec{a}_1/\vec{b}_3$, $\vec{a}_3/\vec{b}_1$ and $\vec{a}_3/\vec{b}_3$ directions (observables $(2)$), Alice and Bob calculate the CHSH correlation value $(3)$.
If $C = -2\sqrt{2}$, then Alice and Bob can be sure that the states they had been receiving from Charlie were entangled indeed.
This fact tells the participants that there was no interference in the quantum channel.
## *Simulation*
In this section we simulate the E91 quantum key distribution protocol *without* the presence of an eavesdropper.
### *Step one: creating the singlets*
In the first step Alice and Bob receive their qubits of the singlet states $\lvert\psi_s\rangle$ created by Charlie.
For our simulation, we need registers with two quantum bits and four classical bits.
```
# Checking the version of PYTHON; we only support > 3.5
import sys
if sys.version_info < (3,5):
raise Exception('Please use Python version 3.5 or greater.')
# useful additional packages
import numpy as np
import random
import re # regular expressions module
# importing the QISKit
from qiskit import QuantumCircuit, QuantumProgram
#import Qconfig
# Quantum program setup
Q_program = QuantumProgram()
#Q_program.set_api(Qconfig.APItoken, Qconfig.config['url']) # set the APIToken and API url
# Creating registers
qr = Q_program.create_quantum_register("qr", 2)
cr = Q_program.create_classical_register("cr", 4)
```
Let us assume that qubits *qr\[0\]* and *qr\[1\]* belong to Alice and Bob respetively.
In classical bits *cr\[0\]* and *cr\[1\]* Alice and Bob store their measurement results, and classical bits *cr\[2\]* and *cr\[3\]* are used by Eve to store her measurement results of Alice's and Bob's qubits.
Now Charlie creates a singlet state:
```
singlet = Q_program.create_circuit('singlet', [qr], [cr])
singlet.x(qr[0])
singlet.x(qr[1])
singlet.h(qr[0])
singlet.cx(qr[0],qr[1])
```
Qubits *qr\[0\]* and *qr\[1\]* are now entangled.
After creating a singlet state, Charlie sends qubit *qr\[0\]* to Alice and qubit *qr\[1\]* to Bob.

### *Step two: measuring*
First let us prepare the measurements which will be used by Alice and Bob.
We define $A(\vec{a}_i) = \vec{a}_i \cdot \vec{\sigma}$ and $B(\vec{b}_j) = \vec{b}_j \cdot \vec{\sigma}$ as the spin projection observables used by Alice and Bob for their measurements.
To perform these measurements, the standard basis $Z$ must be rotated to the proper basis when it is needed (see [Superposition](https://quantumexperience.ng.bluemix.net/proxy/tutorial/full-user-guide/002-The_Weird_and_Wonderful_World_of_the_Qubit/020-Superposition.html) and [Entanglement and Bell Tests](https://quantumexperience.ng.bluemix.net/proxy/tutorial/full-user-guide/003-Multiple_Qubits_Gates_and_Entangled_States/050-Entanglement_and_Bell_Tests.html) user guides).
Here we define the notation of possible measurements of Alice and Bob:

Blocks on the left side can be considered as *detectors* used by the participants to measure $X, W, Z$ and $V$ observables.
Now we prepare the corresponding curcuits.
```
## Alice's measurement circuits
# measure the spin projection of Alice's qubit onto the a_1 direction (X basis)
measureA1 = Q_program.create_circuit('measureA1', [qr], [cr])
measureA1.h(qr[0])
measureA1.measure(qr[0],cr[0])
# measure the spin projection of Alice's qubit onto the a_2 direction (W basis)
measureA2 = Q_program.create_circuit('measureA2', [qr], [cr])
measureA2.s(qr[0])
measureA2.h(qr[0])
measureA2.t(qr[0])
measureA2.h(qr[0])
measureA2.measure(qr[0],cr[0])
# measure the spin projection of Alice's qubit onto the a_3 direction (standard Z basis)
measureA3 = Q_program.create_circuit('measureA3', [qr], [cr])
measureA3.measure(qr[0],cr[0])
## Bob's measurement circuits
# measure the spin projection of Bob's qubit onto the b_1 direction (W basis)
measureB1 = Q_program.create_circuit('measureB1', [qr], [cr])
measureB1.s(qr[1])
measureB1.h(qr[1])
measureB1.t(qr[1])
measureB1.h(qr[1])
measureB1.measure(qr[1],cr[1])
# measure the spin projection of Bob's qubit onto the b_2 direction (standard Z basis)
measureB2 = Q_program.create_circuit('measureB2', [qr], [cr])
measureB2.measure(qr[1],cr[1])
# measure the spin projection of Bob's qubit onto the b_3 direction (V basis)
measureB3 = Q_program.create_circuit('measureB3', [qr], [cr])
measureB3.s(qr[1])
measureB3.h(qr[1])
measureB3.tdg(qr[1])
measureB3.h(qr[1])
measureB3.measure(qr[1],cr[1])
## Lists of measurement circuits
aliceMeasurements = [measureA1, measureA2, measureA3]
bobMeasurements = [measureB1, measureB2, measureB3]
```
Supose Alice and Bob want to generate a secret key using $N$ singlet states prepared by Charlie.
```
# Define the number of singlets N
numberOfSinglets = 500
```
The participants must choose the directions onto which they will measure the spin projections of their qubits.
To do this, Alice and Bob create the strings $b$ and $b^{'}$ with randomly generated elements.
```
aliceMeasurementChoices = [random.randint(1, 3) for i in range(numberOfSinglets)] # string b of Alice
bobMeasurementChoices = [random.randint(1, 3) for i in range(numberOfSinglets)] # string b' of Bob
```
Now we combine Charlie's device and Alice's and Bob's detectors into one circuit (singlet + Alice's measurement + Bob's measurement).
```
circuits = [] # the list in which the created circuits will be stored
for i in range(numberOfSinglets):
# create the name of the i-th circuit depending on Alice's and Bob's measurement choices
circuitName = str(i) + ':A' + str(aliceMeasurementChoices[i]) + '_B' + str(bobMeasurementChoices[i])
# create the joint measurement circuit
# add Alice's and Bob's measurement circuits to the singlet state curcuit
Q_program.add_circuit(circuitName,
singlet + # singlet state circuit
aliceMeasurements[aliceMeasurementChoices[i]-1] + # measurement circuit of Alice
bobMeasurements[bobMeasurementChoices[i]-1] # measurement circuit of Bob
)
# add the created circuit to the circuits list
circuits.append(circuitName)
```
Let us look at the name of one of the prepared circuits.
```
print(circuits[0])
```
It tells us about the number of the singlet state received from Charlie, and the measurements applied by Alice and Bob.
In the *circuits* list we have stored $N$ (*numberOfSinglets*) circuits similar to those shown in the figure below.

The idea is to model every act of the creation of the singlet state, the distribution of its qubits among the participants and the measurement of the spin projection onto the chosen direction in the E91 protocol by executing each circuit from the *circuits* list with one shot.
### *Step three: recording the results*
First let us execute the circuits on the simulator.
```
result = Q_program.execute(circuits, backend='local_qasm_simulator', shots=1, max_credits=5, wait=10, timeout=240)
print(result)
```
Look at the output of the execution of the first circuit.
```
result.get_counts(circuits[0])
```
It consists of four digits.
Recall that Alice and Bob store the results of the measurement in classical bits *cr\[0\]* and *cr\[1\]* (two digits on the right).
Since we model the secret key generation process without the presence of an eavesdropper, the classical bits *cr\[2\]* and *cr\[3\]* are always 0.
Also note that the output is the Python dictionary, in which the keys are the obtained results, and the values are the counts.
Alice and Bob record the results of their measurements as bits of the strings $a$ and $a^{'}$.
To simulate this process we need to use regular expressions module *[re](https://docs.python.org/3/howto/regex.html#regex-howto)*.
First, we compile the search patterns.
```
abPatterns = [
re.compile('..00$'), # search for the '..00' output (Alice obtained -1 and Bob obtained -1)
re.compile('..01$'), # search for the '..01' output
re.compile('..10$'), # search for the '..10' output (Alice obtained -1 and Bob obtained 1)
re.compile('..11$') # search for the '..11' output
]
```
Using these patterns, we can find particular results in the outputs and fill strings the $a$ and $a^{'}$ with the results of Alice's and Bob's measurements.
```
aliceResults = [] # Alice's results (string a)
bobResults = [] # Bob's results (string a')
for i in range(numberOfSinglets):
res = list(result.get_counts(circuits[i]).keys())[0] # extract the key from the dict and transform it to str; execution result of the i-th circuit
if abPatterns[0].search(res): # check if the key is '..00' (if the measurement results are -1,-1)
aliceResults.append(-1) # Alice got the result -1
bobResults.append(-1) # Bob got the result -1
if abPatterns[1].search(res):
aliceResults.append(1)
bobResults.append(-1)
if abPatterns[2].search(res): # check if the key is '..10' (if the measurement results are -1,1)
aliceResults.append(-1) # Alice got the result -1
bobResults.append(1) # Bob got the result 1
if abPatterns[3].search(res):
aliceResults.append(1)
bobResults.append(1)
```
### *Step four: revealing the bases*
In the previos step we have stored the measurement results of Alice and Bob in the *aliceResults* and *bobResults* lists (strings $a$ and $a^{'}$).
Now the participants compare their strings $b$ and $b^{'}$ via the public classical channel.
If Alice and Bob have measured the spin projections of their qubits of the *i*-th singlet onto the same direction, then Alice records the result $a_i$ as the bit of the string $k$, and Bob records the result $-a_i$ as the bit of the string $k^{'}$ (see Eq. (1)).
```
aliceKey = [] # Alice's key string k
bobKey = [] # Bob's key string k'
# comparing the stings with measurement choices
for i in range(numberOfSinglets):
# if Alice and Bob have measured the spin projections onto the a_2/b_1 or a_3/b_2 directions
if (aliceMeasurementChoices[i] == 2 and bobMeasurementChoices[i] == 1) or (aliceMeasurementChoices[i] == 3 and bobMeasurementChoices[i] == 2):
aliceKey.append(aliceResults[i]) # record the i-th result obtained by Alice as the bit of the secret key k
bobKey.append(- bobResults[i]) # record the multiplied by -1 i-th result obtained Bob as the bit of the secret key k'
keyLength = len(aliceKey) # length of the secret key
```
The keys $k$ and $k'$ are now stored in the *aliceKey* and *bobKey* lists, respectively.
The remaining results which were not used to create the keys can now be revealed.
It is important for Alice and Bob to have the same keys, i.e. strings $k$ and $k^{'}$ must be equal.
Let us compare the bits of strings $k$ and $k^{'}$ and find out how many there are mismatches in the keys.
```
abKeyMismatches = 0 # number of mismatching bits in Alice's and Bob's keys
for j in range(keyLength):
if aliceKey[j] != bobKey[j]:
abKeyMismatches += 1
```
Note that since the strings $k$ and $k^{'}$ are secret, Alice and Bob have no information about mismatches in the bits of their keys.
To find out the number of errors, the participants can perform a random sampling test.
Alice randomly selects $\delta$ bits of her secret key and tells Bob which bits she selected.
Then Alice and Bob compare the values of these check bits.
For large enough $\delta$ the number of errors in the check bits will be close to the number of errors in the remaining bits.
### *Step five: CHSH correlation value test*
Alice and Bob want to be sure that there was no interference in the communication session.
To do that, they calculate the CHSH correlation value $(3)$ using the results obtained after the measurements of spin projections onto the $\vec{a}_1/\vec{b}_1$, $\vec{a}_1/\vec{b}_3$, $\vec{a}_3/\vec{b}_1$ and $\vec{a}_3/\vec{b}_3$ directions.
Recall that it is equivalent to the measurement of the observables $X \otimes W$, $X \otimes V$, $Z \otimes W$ and $Z \otimes V$ respectively.
According to the Born-von Neumann statistical postulate, the expectation value of the observable $E = \sum_j e_j \lvert e_j \rangle \langle e_j \rvert$ in the state $\lvert \psi \rangle$ is given by
$$\langle E \rangle_\psi =
\mathrm{Tr}\, \lvert\psi\rangle \langle\psi\rvert \, E = \\
\mathrm{Tr}\, \lvert\psi\rangle \langle\psi\rvert \sum_j e_j \lvert e_j \rangle \langle e_j \rvert =
\sum_j \langle\psi\rvert(e_j \lvert e_j \rangle \langle e_j \rvert) \lvert\psi\rangle =
\sum_j e_j \left|\langle\psi\lvert e_j \rangle \right|^2 = \\
\sum_j e_j \mathrm{P}_\psi (E \models e_j),$$
where $\lvert e_j \rangle$ is the eigenvector of $E$ with the corresponding eigenvalue $e_j$, and $\mathrm{P}_\psi (E \models e_j)$ is the probability of obtainig the result $e_j$ after measuring the observable $E$ in the state $\lvert \psi \rangle$.
A similar expression can be written for the joint measurement of the observables $A$ and $B$:
$$\langle A \otimes B \rangle_\psi =
\sum_{j,k} a_j b_k \mathrm{P}_\psi (A \models a_j, B \models b_k) =
\sum_{j,k} a_j b_k \mathrm{P}_\psi (a_j, b_k). \qquad\qquad (4)$$
Note that if $A$ and $B$ are the spin projection observables, then the corresponding eigenvalues are $a_j, b_k = \pm 1$.
Thus, for the observables $A(\vec{a}_i)$ and $B(\vec{b}_j)$ and singlet state $\lvert\psi\rangle_s$ we can rewrite $(4)$ as
$$\langle A(\vec{a}_i) \otimes B(\vec{b}_j) \rangle =
\mathrm{P}(-1,-1) - \mathrm{P}(1,-1) - \mathrm{P}(-1,1) + \mathrm{P}(1,1). \qquad\qquad (5)$$
In our experiments, the probabilities on the right side can be calculated as follows:
$$\mathrm{P}(a_j, b_k) = \frac{n_{a_j, b_k}(A \otimes B)}{N(A \otimes B)}, \qquad\qquad (6)$$
where the numerator is the number of results $a_j, b_k$ obtained after measuring the observable $A \otimes B$, and the denominator is the total number of measurements of the observable $A \otimes B$.
Since Alice and Bob revealed their strings $b$ and $b^{'}$, they know what measurements they performed and what results they have obtained.
With this data, participants calculate the expectation values $(2)$ using $(5)$ and $(6)$.
```
# function that calculates CHSH correlation value
def chsh_corr(result):
# lists with the counts of measurement results
# each element represents the number of (-1,-1), (-1,1), (1,-1) and (1,1) results respectively
countA1B1 = [0, 0, 0, 0] # XW observable
countA1B3 = [0, 0, 0, 0] # XV observable
countA3B1 = [0, 0, 0, 0] # ZW observable
countA3B3 = [0, 0, 0, 0] # ZV observable
for i in range(numberOfSinglets):
res = list(result.get_counts(circuits[i]).keys())[0]
# if the spins of the qubits of the i-th singlet were projected onto the a_1/b_1 directions
if (aliceMeasurementChoices[i] == 1 and bobMeasurementChoices[i] == 1):
for j in range(4):
if abPatterns[j].search(res):
countA1B1[j] += 1
if (aliceMeasurementChoices[i] == 1 and bobMeasurementChoices[i] == 3):
for j in range(4):
if abPatterns[j].search(res):
countA1B3[j] += 1
if (aliceMeasurementChoices[i] == 3 and bobMeasurementChoices[i] == 1):
for j in range(4):
if abPatterns[j].search(res):
countA3B1[j] += 1
# if the spins of the qubits of the i-th singlet were projected onto the a_3/b_3 directions
if (aliceMeasurementChoices[i] == 3 and bobMeasurementChoices[i] == 3):
for j in range(4):
if abPatterns[j].search(res):
countA3B3[j] += 1
# number of the results obtained from the measurements in a particular basis
total11 = sum(countA1B1)
total13 = sum(countA1B3)
total31 = sum(countA3B1)
total33 = sum(countA3B3)
# expectation values of XW, XV, ZW and ZV observables (2)
expect11 = (countA1B1[0] - countA1B1[1] - countA1B1[2] + countA1B1[3])/total11 # -1/sqrt(2)
expect13 = (countA1B3[0] - countA1B3[1] - countA1B3[2] + countA1B3[3])/total13 # 1/sqrt(2)
expect31 = (countA3B1[0] - countA3B1[1] - countA3B1[2] + countA3B1[3])/total31 # -1/sqrt(2)
expect33 = (countA3B3[0] - countA3B3[1] - countA3B3[2] + countA3B3[3])/total33 # -1/sqrt(2)
corr = expect11 - expect13 + expect31 + expect33 # calculate the CHSC correlation value (3)
return corr
```
### *Output*
Now let us print all the interesting values.
```
corr = chsh_corr(result) # CHSH correlation value
# CHSH inequality test
print('CHSH correlation value: ' + str(round(corr, 3)))
# Keys
print('Length of the key: ' + str(keyLength))
print('Number of mismatching bits: ' + str(abKeyMismatches) + '\n')
```
Finaly, Alice and Bob have the secret keys $k$ and $k^{'}$ (*aliceKey* and *bobKey*)!
Now they can use the one-time pad technique to encrypt and decrypt messages.
Since we simulate the E91 protocol without the presence of Eve, the CHSH correlation value should be close to $-2\sqrt{2} \approx -2.828$.
In addition, there should be no mismatching bits in the keys of Alice and Bob.
Note also that there are 9 possible combinations of measurements that can be performed by Alice and Bob, but only 2 of them give the results using which the secret keys can be created.
Thus, the ratio of the length of the keys to the number of singlets $N$ should be close to $2/9$.
## *Simulation of eavesdropping*
Suppose some third party wants to interfere in the communication session of Alice and Bob and obtain a secret key.
The eavesdropper can use the *intercept-resend* attacks: Eve intercepts one or both of the entangled qubits prepared by Charlie, measures the spin projections of these qubits, prepares new ones depending on the results obtained ($\lvert 01 \rangle$ or $\lvert 10 \rangle$) and sends them to Alice and Bob.
A schematic representation of this process is shown in the figure below.

Here $E(\vec{n}_A) = \vec{n}_A \cdot \vec{\sigma}$ and $E(\vec{n}_B) = \vec{n}_A \cdot \vec{\sigma}$ are the observables of the of the spin projections of Alice's and Bob's qubits onto the directions $\vec{n}_A$ and $\vec{n}_B$.
It would be wise for Eve to choose these directions to be $\vec{n}_A = \vec{a}_2,\vec{a}_3$ and $\vec{n}_B = \vec{b}_1,\vec{b}_2$ since the results obtained from other measurements can not be used to create a secret key.
Let us prepare the circuits for Eve's measurements.
```
# measurement of the spin projection of Alice's qubit onto the a_2 direction (W basis)
measureEA2 = Q_program.create_circuit('measureEA2', [qr], [cr])
measureEA2.s(qr[0])
measureEA2.h(qr[0])
measureEA2.t(qr[0])
measureEA2.h(qr[0])
measureEA2.measure(qr[0],cr[2])
# measurement of the spin projection of Allice's qubit onto the a_3 direction (standard Z basis)
measureEA3 = Q_program.create_circuit('measureEA3', [qr], [cr])
measureEA3.measure(qr[0],cr[2])
# measurement of the spin projection of Bob's qubit onto the b_1 direction (W basis)
measureEB1 = Q_program.create_circuit('measureEB1', [qr], [cr])
measureEB1.s(qr[1])
measureEB1.h(qr[1])
measureEB1.t(qr[1])
measureEB1.h(qr[1])
measureEB1.measure(qr[1],cr[3])
# measurement of the spin projection of Bob's qubit onto the b_2 direction (standard Z measurement)
measureEB2 = Q_program.create_circuit('measureEB2', [qr], [cr])
measureEB2.measure(qr[1],cr[3])
# lists of measurement circuits
eveMeasurements = [measureEA2, measureEA3, measureEB1, measureEB2]
```
Like Alice and Bob, Eve must choose the directions onto which she will measure the spin projections of the qubits.
In our simulation, the eavesdropper randomly chooses one of the observables $W \otimes W$ or $Z \otimes Z$ to measure.
```
# list of Eve's measurement choices
# the first and the second elements of each row represent the measurement of Alice's and Bob's qubits by Eve respectively
eveMeasurementChoices = []
for j in range(numberOfSinglets):
if random.uniform(0, 1) <= 0.5: # in 50% of cases perform the WW measurement
eveMeasurementChoices.append([0, 2])
else: # in 50% of cases perform the ZZ measurement
eveMeasurementChoices.append([1, 3])
```
Like we did before, now we create the circuits with singlet states and detectors of Eve, Alice and Bob.
```
circuits = [] # the list in which the created circuits will be stored
for j in range(numberOfSinglets):
# create the name of the j-th circuit depending on Alice's, Bob's and Eve's choices of measurement
circuitName = str(j) + ':A' + str(aliceMeasurementChoices[j]) + '_B' + str(bobMeasurementChoices[j] + 2) + '_E' + str(eveMeasurementChoices[j][0]) + str(eveMeasurementChoices[j][1] - 1)
# create the joint measurement circuit
# add Alice's and Bob's measurement circuits to the singlet state curcuit
Q_program.add_circuit(circuitName,
singlet + # singlet state circuit
eveMeasurements[eveMeasurementChoices[j][0]-1] + # Eve's measurement circuit of Alice's qubit
eveMeasurements[eveMeasurementChoices[j][1]-1] + # Eve's measurement circuit of Bob's qubit
aliceMeasurements[aliceMeasurementChoices[j]-1] + # measurement circuit of Alice
bobMeasurements[bobMeasurementChoices[j]-1] # measurement circuit of Bob
)
# add the created circuit to the circuits list
circuits.append(circuitName)
```
Now we execute all the prepared circuits on the simulator.
```
result = Q_program.execute(circuits, backend='local_qasm_simulator', shots=1, max_credits=5, wait=10, timeout=240)
print(result)
```
Let us look at the name of the first circuit and the output after it is executed.
```
print(str(circuits[0]) + '\t' + str(result.get_counts(circuits[0])))
```
We can see onto which directions Eve, Alice and Bob measured the spin projections and the results obtained.
Recall that the bits *cr\[2\]* and *cr\[3\]* (two digits on the left) are used by Eve to store the results of her measurements.
To extract Eve's results from the outputs, we need to compile new search patterns.
```
ePatterns = [
re.compile('00..$'), # search for the '00..' result (Eve obtained the results -1 and -1 for Alice's and Bob's qubits)
re.compile('01..$'), # search for the '01..' result (Eve obtained the results 1 and -1 for Alice's and Bob's qubits)
re.compile('10..$'),
re.compile('11..$')
]
```
Now Eve, Alice and Bob record the results of their measurements.
```
aliceResults = [] # Alice's results (string a)
bobResults = [] # Bob's results (string a')
# list of Eve's measurement results
# the elements in the 1-st column are the results obtaned from the measurements of Alice's qubits
# the elements in the 2-nd column are the results obtaned from the measurements of Bob's qubits
eveResults = []
# recording the measurement results
for j in range(numberOfSinglets):
res = list(result.get_counts(circuits[j]).keys())[0] # extract a key from the dict and transform it to str
# Alice and Bob
if abPatterns[0].search(res): # check if the key is '..00' (if the measurement results are -1,-1)
aliceResults.append(-1) # Alice got the result -1
bobResults.append(-1) # Bob got the result -1
if abPatterns[1].search(res):
aliceResults.append(1)
bobResults.append(-1)
if abPatterns[2].search(res): # check if the key is '..10' (if the measurement results are -1,1)
aliceResults.append(-1) # Alice got the result -1
bobResults.append(1) # Bob got the result 1
if abPatterns[3].search(res):
aliceResults.append(1)
bobResults.append(1)
# Eve
if ePatterns[0].search(res): # check if the key is '00..'
eveResults.append([-1, -1]) # results of the measurement of Alice's and Bob's qubits are -1,-1
if ePatterns[1].search(res):
eveResults.append([1, -1])
if ePatterns[2].search(res):
eveResults.append([-1, 1])
if ePatterns[3].search(res):
eveResults.append([1, 1])
```
As before, Alice, Bob and Eve create the secret keys using the results obtained after measuring the observables $W \otimes W$ and $Z \otimes Z$.
```
aliceKey = [] # Alice's key string a
bobKey = [] # Bob's key string a'
eveKeys = [] # Eve's keys; the 1-st column is the key of Alice, and the 2-nd is the key of Bob
# comparing the strings with measurement choices (b and b')
for j in range(numberOfSinglets):
# if Alice and Bob measured the spin projections onto the a_2/b_1 or a_3/b_2 directions
if (aliceMeasurementChoices[j] == 2 and bobMeasurementChoices[j] == 1) or (aliceMeasurementChoices[j] == 3 and bobMeasurementChoices[j] == 2):
aliceKey.append(aliceResults[j]) # record the i-th result obtained by Alice as the bit of the secret key k
bobKey.append(-bobResults[j]) # record the multiplied by -1 i-th result obtained Bob as the bit of the secret key k'
eveKeys.append([eveResults[j][0], -eveResults[j][1]]) # record the i-th bits of the keys of Eve
keyLength = len(aliceKey) # length of the secret skey
```
To find out the number of mismatching bits in the keys of Alice, Bob and Eve we compare the lists *aliceKey*, *bobKey* and *eveKeys*.
```
abKeyMismatches = 0 # number of mismatching bits in the keys of Alice and Bob
eaKeyMismatches = 0 # number of mismatching bits in the keys of Eve and Alice
ebKeyMismatches = 0 # number of mismatching bits in the keys of Eve and Bob
for j in range(keyLength):
if aliceKey[j] != bobKey[j]:
abKeyMismatches += 1
if eveKeys[j][0] != aliceKey[j]:
eaKeyMismatches += 1
if eveKeys[j][1] != bobKey[j]:
ebKeyMismatches += 1
```
It is also good to know what percentage of the keys is known to Eve.
```
eaKnowledge = (keyLength - eaKeyMismatches)/keyLength # Eve's knowledge of Bob's key
ebKnowledge = (keyLength - ebKeyMismatches)/keyLength # Eve's knowledge of Alice's key
```
Using the *chsh_corr* function defined above we calculate the CSHS correlation value.
```
corr = chsh_corr(result)
```
And now we print all the results.
```
# CHSH inequality test
print('CHSH correlation value: ' + str(round(corr, 3)) + '\n')
# Keys
print('Length of the key: ' + str(keyLength))
print('Number of mismatching bits: ' + str(abKeyMismatches) + '\n')
print('Eve\'s knowledge of Alice\'s key: ' + str(round(eaKnowledge * 100, 2)) + ' %')
print('Eve\'s knowledge of Bob\'s key: ' + str(round(ebKnowledge * 100, 2)) + ' %')
```
Due to Eve's interference in the communication session, the CHSH correlation value is far away from $-2 \sqrt{2}$.
Alice and Bob see it and will not use the secret key to encrypt and decrypt any messages.
It has been shown by Ekert that for any eavesdropping strategy and for any directions $\vec{n}_A$, $\vec{n}_B$ onto which Eve measures the spin projections of Alice's and Bob's qubits the following inequality can be written:
$$ -\sqrt{2} \leqslant C \leqslant \sqrt{2},$$
where $C$ is CHSH correlation value (3).
The more Eve interferes in the communication session, the more she knows about the secret keys.
But at the same time, the deviation of the CHSH correlation value from $-2\sqrt{2}$ also increases.
We can see that there are the mismatches in the keys of Alice and Bob.
Where do they come from?
After Eve measures the qubits of the singlet state $\lvert \psi_s \rangle$, she randomly obtains the results $-1,1$ or $1,-1$ (see Eq. (1)).
Depending on the results obtained, the eavesdropper prepares the state $\lvert \varphi_1 \rangle = \lvert 01 \rangle$ or $\lvert \varphi_2 \rangle = \lvert 10 \rangle$ (in our simulation it is automatically provided by a measurement in the $Z$ basis) and sends its qubits to Alice and Bob.
When Alice and Bob measure the observable $W \otimes W$, they obtain any combination of results with probability $\mathrm{P}_{\varphi_{n}}(a_i, b_j)$.
To see this, one can compare the results of the execution of Quantum Scores of [$W_E \otimes W_E \vert W_A \otimes W_B$](https://quantumexperience.ng.bluemix.net/share/code/1c4d96685cb20c2b99e43f9999b28313/execution/917dca7c81dfda7886af97eef85d5946) and [$W_E \otimes W_E \vert Z_A \otimes Z_B$](https://quantumexperience.ng.bluemix.net/share/code/0d378f5f16990ab3e47546ae4b0c39d2/execution/e836b67e10e9d11d7828a67a834cf4fd) measurements (the subscripts denote who performs the measurement).
In order to correct the mismatches in the keys of Alice and Bob classical error reconciliation algorithms are used.
A very good description of the error correction methods can be found in [Quantum cryptography](https://arxiv.org/abs/quant-ph/0101098) by N. Gisin et al.
| github_jupyter |
# 2A.eco - Python et la logique SQL - correction
Correction d'exercices sur SQL.
```
from jyquickhelper import add_notebook_menu
add_notebook_menu()
```
SQL permet de créer des tables, de rechercher, d'ajouter, de modifier ou de supprimer des données dans les bases de données.
Un peu ce que vous ferez bientôt tous les jours. C’est un langage de management de données, pas de nettoyage, d’analyse ou de statistiques avancées.
Les instructions SQL s'écrivent d'une manière qui ressemble à celle de phrases ordinaires en anglais. Cette ressemblance voulue vise à faciliter l'apprentissage et la lecture. Il est néanmoins important de respecter un ordre pour les différentes instructions.
Dans ce TD, nous allons écrire des commandes en SQL via Python.
Pour plus de précisions sur SQL et les commandes qui existent, rendez-vous là [SQL, PRINCIPES DE BASE](http://www.xavierdupre.fr/app/ensae_teaching_cs/helpsphinx/ext2a/sql_doc.html).
## Se connecter à une base de données
A la différence des tables qu'on utilise habituellement, la base de données n'est pas visible directement en ouvrant Excel ou un éditeur de texte. Pour avoir une vue de ce que contient la base de données, il est nécessaire d'avoir un autre type de logiciel.
Pour le TD, nous vous recommandans d'installer SQLLiteSpy (disponible à cette adresse [SqliteSpy](http://www.yunqa.de/delphi/products/sqlitespy/index) ou [sqlite_bro](https://pypi.python.org/pypi/sqlite_bro) si vous voulez voir à quoi ressemble les données avant de les utiliser avec Python.
```
import sqlite3
# on va se connecter à une base de données SQL vide
# SQLite stocke la BDD dans un simple fichier
filepath = "./DataBase.db"
open(filepath, 'w').close() #crée un fichier vide
CreateDataBase = sqlite3.connect(filepath)
QueryCurs = CreateDataBase.cursor()
```
La méthode cursor() est un peu particulière :
Il s'agit d'une sorte de tampon mémoire intermédiaire, destiné à mémoriser temporairement les données en cours de traitement, ainsi que les opérations que vous effectuez sur elles, avant leur transfert définitif dans la base de données. Tant que la méthode .commit() n'aura pas été appelée, aucun ordre ne sera appliqué à la base de données.
--------------------
A présent que nous sommes connectés à la base de données, on va créer une table qui contient plusieurs variables de format différents
- ID sera la clé primaire de la base
- Nom, Rue, Ville, Pays seront du text
- Prix sera un réel
```
# On définit une fonction de création de table
def CreateTable(nom_bdd):
QueryCurs.execute('''CREATE TABLE IF NOT EXISTS ''' + nom_bdd + '''
(id INTEGER PRIMARY KEY, Name TEXT,City TEXT, Country TEXT, Price REAL)''')
# On définit une fonction qui permet d'ajouter des observations dans la table
def AddEntry(nom_bdd, Nom,Ville,Pays,Prix):
QueryCurs.execute('''INSERT INTO ''' + nom_bdd + '''
(Name,City,Country,Price) VALUES (?,?,?,?)''',(Nom,Ville,Pays,Prix))
def AddEntries(nom_bdd, data):
""" data : list with (Name,City,Country,Price) tuples to insert
"""
QueryCurs.executemany('''INSERT INTO ''' + nom_bdd + '''
(Name,City,Country,Price) VALUES (?,?,?,?)''',data)
### On va créer la table clients
CreateTable('Clients')
AddEntry('Clients','Toto','Munich','Germany',5.2)
AddEntries('Clients',
[('Bill','Berlin','Germany',2.3),
('Tom','Paris','France',7.8),
('Marvin','Miami','USA',15.2),
('Anna','Paris','USA',7.8)])
# on va "commit" c'est à dire qu'on va valider la transaction.
# > on va envoyer ses modifications locales vers le référentiel central - la base de données SQL
CreateDataBase.commit()
```
### Voir la table
Pour voir ce qu'il y a dans la table, on utilise un premier Select où on demande à voir toute la table
```
QueryCurs.execute('SELECT * FROM Clients')
Values = QueryCurs.fetchall()
print(Values)
```
### Passer en pandas
Rien de plus simple : plusieurs manières de faire
```
import pandas as pd
# méthode SQL Query
df1 = pd.read_sql_query('SELECT * FROM Clients', CreateDataBase)
print("En utilisant la méthode read_sql_query \n", df1.head(), "\n")
#méthode DataFrame en utilisant la liste issue de .fetchall()
df2 = pd.DataFrame(Values, columns=['ID','Name','City','Country','Price'])
print("En passant par une DataFrame \n", df2.head())
```
## Comparaison SQL et pandas
### SELECT
En SQL, la sélection se fait en utilisant des virgules ou * si on veut sélectionner toutes les colonnes
```
# en SQL
QueryCurs.execute('SELECT ID,City FROM Clients LIMIT 2')
Values = QueryCurs.fetchall()
print(Values)
```
En pandas, la sélection de colonnes se fait en donnant une liste
```
#sur la table
df2[['ID','City']].head(2)
```
### WHERE
En SQL, on utilise WHERE pour filtrer les tables selon certaines conditions
```
QueryCurs.execute('SELECT * FROM Clients WHERE City=="Paris"')
print(QueryCurs.fetchall())
```
Avec Pandas, on peut utiliser plusieurs manières de faire :
- avec un booléen
- en utilisant la méthode 'query'
```
df2[df2['City'] == "Paris"]
df2.query('City == "Paris"')
```
Pour mettre plusieurs conditions, on utilise :
- & en Python, AND en SQL
- | en python, OR en SQL
```
QueryCurs.execute('SELECT * FROM Clients WHERE City=="Paris" AND Country == "USA"')
print(QueryCurs.fetchall())
df2.query('City == "Paris" & Country == "USA"')
df2[(df2['City'] == "Paris") & (df2['Country'] == "USA")]
```
## GROUP BY
En pandas, l'opération GROUP BY de SQL s'effectue avec une méthode similaire : groupby()
groupby() sert à regrouper des observations en groupes selon les modalités de certaines variables en appliquant une fonction d'aggrégation sur d'autres variables.
```
QueryCurs.execute('SELECT Country, count(*) FROM Clients GROUP BY Country')
print(QueryCurs.fetchall())
```
Attention, en pandas, la fonction count() ne fait pas la même chose qu'en SQL. Count() s'applique à toutes les colonnes et compte toutes les observations non nulles.
```
df2.groupby('Country').count()
```
Pour réaliser la même chose qu'en SQL, il faut utiliser la méthode size()
```
df2.groupby('Country').size()
```
On peut aussi appliquer des fonctions plus sophistiquées lors d'un groupby
```
QueryCurs.execute('SELECT Country, AVG(Price), count(*) FROM Clients GROUP BY Country')
print(QueryCurs.fetchall())
```
Avec pandas, on peut appeler les fonctions classiques de numpy
```
import numpy as np
df2.groupby('Country').agg({'Price': np.mean, 'Country': np.size})
```
Ou utiliser des fonctions lambda
```
# par exemple calculer le prix moyen et le multiplier par 2
df2.groupby('Country')['Price'].apply(lambda x: 2*x.mean())
QueryCurs.execute('SELECT Country, 2*AVG(Price) FROM Clients GROUP BY Country').fetchall()
QueryCurs.execute('SELECT * FROM Clients WHERE Country == "Germany"')
print(QueryCurs.fetchall())
QueryCurs.execute('SELECT * FROM Clients WHERE City=="Berlin" AND Country == "Germany"')
print(QueryCurs.fetchall())
QueryCurs.execute('SELECT * FROM Clients WHERE Price BETWEEN 7 AND 20')
print(QueryCurs.fetchall())
```
## Enregistrer une table SQL sous un autre format
On utilise le package csv, l'option 'w' pour 'write'.
On crée l'objet "writer", qui vient du package csv.
Cet objet a deux méthodes :
- writerow pour les noms de colonnes : une liste
- writerows pour les lignes : un ensemble de liste
```
data = QueryCurs.execute('SELECT * FROM Clients')
import csv
with open('./output.csv', 'w') as file:
writer = csv.writer(file)
writer.writerow(['id','Name','City','Country','Price'])
writer.writerows(data)
```
On peut également passer par un DataFrame pandas et utiliser .to_csv()
```
QueryCurs.execute('''DROP TABLE Clients''')
#QueryCurs.close()
```
## Exercice
Dans cet exercice, nous allons manipuler les tables de la base de données World.
Avant tout, connectez vous à la base de donénes en utilisant sqlite3 et connect
Lien vers la base de données : [World.db3](https://github.com/sdpython/ensae_teaching_cs/raw/master/src/ensae_teaching_cs/data/data_sql/World.db3) ou
```
from ensae_teaching_cs.data import simple_database
name = simple_database()
```
```
#Se connecter à la base de données WORLD
CreateDataBase = sqlite3.connect("./World.db3")
QueryCurs = CreateDataBase.cursor()
```
Familiarisez vous avec la base de données : quelles sont les tables ? quelles sont les variables de ces tables ?
- utilisez la fonction PRAGMA pour obtenir des informations sur les tables
```
# pour obtenir la liste des tables dans la base de données
tables = QueryCurs.execute("SELECT name FROM sqlite_master WHERE type='table';").fetchall()
# on veut voir les colonnes de chaque table ainsi que la première ligne
for table in tables :
print("Table :", table[0])
schema = QueryCurs.execute("PRAGMA table_info({})".format(table[0])).fetchall()
print("Colonnes", ["{}".format(x[1]) for x in schema])
print("1ère ligne", QueryCurs.execute('SELECT * FROM {} LIMIT 1'.format(table[0])).fetchall(), "\n")
```
## Question 1
- Quels sont les 10 pays qui ont le plus de langues ?
- Quelle langue est présente dans le plus de pays ?
```
QueryCurs.execute("""SELECT CountryCode, COUNT(*) as NB
FROM CountryLanguage
GROUP BY CountryCode
ORDER BY NB DESC
LIMIT 10""").fetchall()
QueryCurs.execute('''SELECT Language, COUNT(*) as NB
FROM CountryLanguage
GROUP BY Language
ORDER BY -NB
LIMIT 1''').fetchall()
```
## Question 2
- Quelles sont les différentes formes de gouvernements dans les pays du monde ?
- Quels sont les 3 gouvernements où la population est la plus importante ?
```
QueryCurs.execute('''SELECT DISTINCT GovernmentForm FROM Country''').fetchall()
QueryCurs.execute('''SELECT GovernmentForm, SUM(Population) as Pop_Totale_Gouv
FROM Country
GROUP BY GovernmentForm
ORDER BY Pop_Totale_Gouv DESC
LIMIT 3
''').fetchall()
```
## Question 3
- Combien de pays ont Elisabeth II à la tête de leur gouvernement ?
- Quelle proporition des sujets de Sa Majesté ne parlent pas anglais ?
- 78 % ou 83% ?
```
QueryCurs.execute('''SELECT HeadOfState, Count(*)
FROM Country
WHERE HeadOfState = "Elisabeth II" ''').fetchall()
# la population totale
population_queen_elisabeth = QueryCurs.execute('''SELECT HeadOfState, SUM(Population)
FROM Country
WHERE HeadOfState = "Elisabeth II"''').fetchall()
# La part de la population parlant anglais
Part_parlant_anglais= QueryCurs.execute('''SELECT Language, SUM(Percentage*0.01*Population)
FROM
Country
LEFT JOIN
CountryLanguage
ON Country.Code = CountryLanguage.CountryCode
WHERE HeadOfState = "Elisabeth II"
AND Language = "English"
''').fetchall()
# La réponse est 78% d'après ces données
Part_parlant_anglais[0][1]/population_queen_elisabeth[0][1]
## on trouve 83% si on ne fait pas attention au fait que dans certaines zones, 0% de la population parle anglais
## La population totale n'est alors pas la bonne, comme dans cet exemple
QueryCurs.execute('''SELECT Language,
SUM(Population_pays*0.01*Percentage) as Part_parlant_anglais, SUM(Population_pays) as Population_totale
FROM (SELECT Language, Code, Percentage, SUM(Population) as Population_pays
FROM
Country
LEFT JOIN
CountryLanguage
ON Country.Code = CountryLanguage.CountryCode
WHERE HeadOfState = "Elisabeth II" AND Language == "English"
GROUP BY Code)''').fetchall()
```
Conclusion: il vaut mieux écrire deux requêtes simples et lisibles pour obtenir le bon résultat, plutôt qu'une requête qui fait tout en une seule passe mais dont on va devoir vérifier la correction longuement...
## Question 4 - passons à Pandas
Créer une DataFrame qui contient les informations suivantes par pays :
- le nom
- le code du pays
- le nombre de langues parlées
- le nombre de langues officielles
- la population
- le GNP
- l'espérance de vie
**Indice : utiliser la commande pd.read_sql_query**
Que dit la matrice de corrélation de ces variables ?
```
df = pd.read_sql_query('''SELECT Code, Name, Population, GNP , LifeExpectancy,
COUNT(*) as Nb_langues_parlees, SUM(IsOfficial) as Nb_langues_officielles
FROM Country
INNER JOIN CountryLanguage ON Country.Code = CountryLanguage.CountryCode
GROUP BY Country.Code''',
CreateDataBase)
df.head()
df.corr()
```
| github_jupyter |
```
# default_exp text.data.language_modeling
# default_cls_lvl 3
# all_slow
#hide
%reload_ext autoreload
%autoreload 2
%matplotlib inline
```
# text.data.language_modeling
> This module contains the bits required to use the fastai DataBlock API and/or mid-level data processing pipelines to organize your data for causal and masked language modeling tasks. This includes things like training BERT from scratch or fine-tuning a particular pre-trained LM on your own corpus.
```
# export
import os, random
from abc import ABC, abstractmethod
from enum import Enum
from datasets import Dataset
from fastcore.all import *
from fastai.imports import *
from fastai.losses import CrossEntropyLossFlat
from fastai.torch_core import *
from fastai.torch_imports import *
from transformers import (
AutoModelForCausalLM,
AutoModelForMaskedLM,
logging,
PretrainedConfig,
PreTrainedTokenizerBase,
PreTrainedModel,
BatchEncoding,
)
from blurr.text.data.core import TextInput, BatchTokenizeTransform, Preprocessor, first_blurr_tfm
from blurr.text.utils import get_hf_objects
logging.set_verbosity_error()
# hide_input
import pdb
from fastai.data.block import DataBlock, ColReader, ColSplitter
from fastai.data.core import DataLoader, DataLoaders, TfmdDL
from fastai.data.external import untar_data, URLs
from fastai.data.transforms import *
from fastcore.test import *
from nbdev.showdoc import show_doc
from blurr.utils import print_versions
from blurr.text.data.core import TextBlock
from blurr.text.utils import BlurrText
NLP = BlurrText()
os.environ["TOKENIZERS_PARALLELISM"] = "false"
print("What we're running with at the time this documentation was generated:")
print_versions("torch fastai transformers")
# hide
# cuda
torch.cuda.set_device(1)
print(f"Using GPU #{torch.cuda.current_device()}: {torch.cuda.get_device_name()}")
```
## Setup
For this example, we'll use the `WIKITEXT_TINY` dataset available from fastai. In addition to using the `Datasets` library from Hugging Face, fastai provides a lot of smaller datasets that are really useful when experimenting and/or in the early development of your training/validation/inference coding.
```
wiki_path = untar_data(URLs.WIKITEXT_TINY)
wiki_path.ls()
train_df = pd.read_csv(wiki_path / "train.csv", header=None)
valid_df = pd.read_csv(wiki_path / "test.csv", header=None)
print(len(train_df), len(valid_df))
train_df.head()
train_df["is_valid"] = False
valid_df["is_valid"] = True
df = pd.concat([train_df, valid_df])
df.head()
model_cls = AutoModelForCausalLM
pretrained_model_name = "gpt2"
hf_arch, hf_config, hf_tokenizer, hf_model = get_hf_objects(pretrained_model_name, model_cls=model_cls)
# some tokenizers like gpt and gpt2 do not have a pad token, so we add it here mainly for the purpose
# of setting the "labels" key appropriately (see below)
if hf_tokenizer.pad_token is None:
hf_tokenizer.pad_token = "[PAD]"
hf_tokenizer.pad_token, hf_tokenizer.pad_token_id
# special_tokens_dict = {'additional_special_tokens': ['[C1]']}
# num_added_toks = hf_tokenizer.add_special_tokens(special_tokens_dict)
# hf_model.resize_token_embeddings(len(hf_tokenizer))
```
## Preprocessing
Starting with version 2.0, `BLURR` provides a language preprocessing class that can be used to preprocess DataFrames or Hugging Face Datasets for both causal and masked language modeling tasks.
### `LMPreprocessor` -
```
# export
class LMPreprocessor(Preprocessor):
def __init__(
self,
# A Hugging Face tokenizer
hf_tokenizer: PreTrainedTokenizerBase,
# The number of examples to process at a time
batch_size: int = 1000,
# How big each chunk of text should be (default: hf_tokenizer.model_max_length)
chunk_size: Optional[int] = None,
# How to indicate the beginning on a new text example (default is hf_tokenizer.eos_token|sep_token
sep_token: Optional[str] = None,
# The attribute holding the text
text_attr: str = "text",
# The attribute that should be created if your are processing individual training and validation
# datasets into a single dataset, and will indicate to which each example is associated
is_valid_attr: Optional[str] = "is_valid",
# Tokenization kwargs that will be applied with calling the tokenizer
tok_kwargs: dict = {},
):
tok_kwargs = {**tok_kwargs, "truncation": False, "return_offsets_mapping": True}
super().__init__(hf_tokenizer, batch_size, text_attr, None, is_valid_attr, tok_kwargs)
self.chunk_size = chunk_size or hf_tokenizer.model_max_length
self.sep_token = sep_token or hf_tokenizer.eos_token or hf_tokenizer.sep_token
def process_df(self, training_df: pd.DataFrame, validation_df: Optional[pd.DataFrame] = None):
# process df in mini-batches
final_train_df = pd.DataFrame()
for g, batch_df in training_df.groupby(np.arange(len(training_df)) // self.batch_size):
final_train_df = final_train_df.append(self._process_df_batch(batch_df))
final_train_df.reset_index(drop=True, inplace=True)
final_val_df = pd.DataFrame() if validation_df is not None else None
if final_val_df is not None:
for g, batch_df in validation_df.groupby(np.arange(len(validation_df)) // self.batch_size):
final_val_df = final_val_df.append(self._process_df_batch(batch_df))
final_val_df.reset_index(drop=True, inplace=True)
final_df = super().process_df(final_train_df, final_val_df)
return final_df
def process_hf_dataset(self, training_ds: Dataset, validation_ds: Optional[Dataset] = None):
ds = super().process_hf_dataset(training_ds, validation_ds)
return Dataset.from_pandas(self.process_df(pd.DataFrame(ds)))
# ----- utility methods -----
def _process_df_batch(self, batch_df):
batch_df.reset_index(drop=True, inplace=True)
# concatenate our texts
concat_txts = {self.text_attr: f" {self.sep_token} ".join(batch_df[self.text_attr].values.tolist())}
inputs = self._tokenize_function(concat_txts)
# compute the length of our concatenated texts
n_total_toks = len(inputs["input_ids"])
# need to modify chunk_size to included the # of special tokens added
max_chunk_size = self.chunk_size - self.hf_tokenizer.num_special_tokens_to_add() - 1
# drop the last chunk of text if it is smaller than chunk size (see the HF course, section 7 on training MLMs)
total_length = (n_total_toks // max_chunk_size) * max_chunk_size
# break our concatenated into chunks of text of size max_chunk_size
examples = []
for i in range(0, total_length, max_chunk_size):
chunked_offsets = inputs["offset_mapping"][i : i + max_chunk_size]
chunked_text = concat_txts[self.text_attr][min(chunked_offsets)[0] : max(chunked_offsets)[1]]
examples.append(chunked_text)
return pd.DataFrame(examples, columns=[f"proc_{self.text_attr}"])
```
#### Using a `DataFrame`
```
preprocessor = LMPreprocessor(hf_tokenizer, chunk_size=128, text_attr=0)
proc_df = preprocessor.process_df(train_df, valid_df)
print(len(proc_df))
proc_df.head(2)
```
#### Using a Hugging Face `Dataset`
```
# TODO
```
## LM Strategies
### `LMType` -
```
# export
class LMType(Enum):
"""Use this enum to indicate what kind of language model you are training"""
CAUSAL = 1
MASKED = 2
```
### `BaseLMStrategy` and implementations -
```
# export
class BaseLMStrategy(ABC):
"""ABC for various language modeling strategies (e.g., causal, BertMLM, WholeWordMLM, etc...)"""
def __init__(self, hf_tokenizer, ignore_token_id=CrossEntropyLossFlat().ignore_index):
store_attr(["hf_tokenizer", "ignore_token_id"])
@abstractmethod
def build_inputs_targets(self, samples, include_labels: bool = True, inputs: Optional[BatchEncoding] = None):
pass
# utility methods
def _get_random_token_id(self, n):
return random.sample(list(self.hf_tokenizer.get_vocab().values()), n)
@classmethod
@abstractmethod
def get_lm_type(cls):
pass
```
Here we include a `BaseLMStrategy` abstract class and several different strategies for building your inputs and targets for causal and masked language modeling tasks. With CLMs, the objective is to simply predict the next token, but with MLMs, a variety of masking strategies may be used (e.g., mask random tokens, mask random words, mask spans, etc...). A `BertMLMStrategy` is introduced below that follows the "mask random tokens" strategy used in the BERT paper, but users can create their own `BaseLMStrategy` subclass to support any masking strategy they desire.
#### `CausalLMStrategy` -
```
# export
class CausalLMStrategy(BaseLMStrategy):
"""For next token prediction language modeling tasks, we want to use the `CausalLMStrategy` which makes the
necessary changes in your inputs/targets for causal LMs
"""
def build_inputs_targets(self, samples, include_labels: bool = True, inputs: Optional[BatchEncoding] = None):
updated_samples = []
for s in samples:
if include_labels:
s[0]["labels"] = s[0]["input_ids"].clone()
s[0]["labels"][s[0]["labels"] == self.hf_tokenizer.pad_token_id] = self.ignore_token_id
targ_ids = torch.cat([s[0]["input_ids"][1:], tensor([self.hf_tokenizer.eos_token_id])])
updated_samples.append((s[0], targ_ids))
return updated_samples
@classmethod
def get_lm_type(cls: LMType):
return LMType.CAUSAL
```
#### `BertMLMStrategy` -
```
# export
class BertMLMStrategy(BaseLMStrategy):
"""A masked language modeling strategy using the default BERT masking definition."""
def __init__(self, hf_tokenizer, ignore_token_id=CrossEntropyLossFlat().ignore_index):
super().__init__(hf_tokenizer, ignore_token_id)
vocab = hf_tokenizer.get_vocab()
self.dnm_tok_ids = [
vocab[tok] for tok in list(hf_tokenizer.special_tokens_map.values()) if vocab[tok] != hf_tokenizer.mask_token_id
]
def build_inputs_targets(self, samples, include_labels: bool = True, inputs: Optional[BatchEncoding] = None):
updated_samples = []
for s in samples:
# mask the input_ids
masked_input_ids = s[0]["input_ids"].clone()
# we want to mask 15% of the non-special tokens(e.g., special tokens inclue [CLS], [SEP], etc...)
idxs = torch.randperm(len(masked_input_ids))
total_masked_idxs = int(len(idxs) * 0.15)
# of the 15% for masking, replace 80% with [MASK] token, 10% with random token, and 10% with correct token
n_mask_idxs = int(total_masked_idxs * 0.8)
n_rnd_idxs = int(total_masked_idxs * 0.1)
# we only want non-special tokens
mask_idxs = [idx for idx in idxs if masked_input_ids[idx] not in self.dnm_tok_ids][:total_masked_idxs]
# replace 80% with [MASK]
if n_mask_idxs > 0 and len(mask_idxs) >= n_mask_idxs:
masked_input_ids[[mask_idxs[:n_mask_idxs]]] = self.hf_tokenizer.mask_token_id
# replace 10% with a random token
if n_rnd_idxs > 0 and len(mask_idxs) >= (n_mask_idxs + n_rnd_idxs):
rnd_tok_ids = self._get_random_token_id(n_rnd_idxs)
masked_input_ids[[mask_idxs[n_mask_idxs : (n_mask_idxs + n_rnd_idxs)]]] = tensor(rnd_tok_ids)
# ignore padding when calculating the loss
lbls = s[0]["input_ids"].clone()
lbls[[[idx for idx in idxs if idx not in mask_idxs]]] = self.ignore_token_id
# update the inputs to use our masked input_ids and labels; set targ_ids = labels (will use when
# we calculate the loss ourselves)
s[0]["input_ids"] = masked_input_ids
targ_ids = lbls
if include_labels:
s[0]["labels"] = targ_ids.clone()
updated_samples.append((s[0], targ_ids))
return updated_samples
@classmethod
def get_lm_type(cls: LMType):
return LMType.MASKED
```
Follows the masking strategy used in the [BERT paper](https://arxiv.org/abs/1810.04805) for random token masking
## Mid-level API
#### `CausalLMTextInput` and `MLMTextInput` -
```
# export
class CausalLMTextInput(TextInput):
pass
# export
class MLMTextInput(TextInput):
pass
```
Again, we define a custom classes for the `@typedispatch`ed methods to use so that we can override how both causal and masked language modeling inputs/targets are assembled, as well as, how the data is shown via methods like `show_batch` and `show_results`.
#### `LMBatchTokenizeTransform` -
```
# export
class LMBatchTokenizeTransform(BatchTokenizeTransform):
def __init__(
self,
# The abbreviation/name of your Hugging Face transformer architecture (e.b., bert, bart, etc..)
hf_arch: str,
# A specific configuration instance you want to use
hf_config: PretrainedConfig,
# A Hugging Face tokenizer
hf_tokenizer: PreTrainedTokenizerBase,
# A Hugging Face model
hf_model: PreTrainedModel,
# To control whether the "labels" are included in your inputs. If they are, the loss will be calculated in
# the model's forward function and you can simply use `PreCalculatedLoss` as your `Learner`'s loss function to use it
include_labels: bool = True,
# The token ID that should be ignored when calculating the loss
ignore_token_id: int = CrossEntropyLossFlat().ignore_index,
# The language modeling strategy (or objective)
lm_strategy_cls: BaseLMStrategy = CausalLMStrategy,
# To control the length of the padding/truncation. It can be an integer or None,
# in which case it will default to the maximum length the model can accept. If the model has no
# specific maximum input length, truncation/padding to max_length is deactivated.
# See [Everything you always wanted to know about padding and truncation](https://huggingface.co/transformers/preprocessing.html#everything-you-always-wanted-to-know-about-padding-and-truncation)
max_length: int = None,
# To control the `padding` applied to your `hf_tokenizer` during tokenization. If None, will default to
# `False` or `'do_not_pad'.
# See [Everything you always wanted to know about padding and truncation](https://huggingface.co/transformers/preprocessing.html#everything-you-always-wanted-to-know-about-padding-and-truncation)
padding: Union[bool, str] = True,
# To control `truncation` applied to your `hf_tokenizer` during tokenization. If None, will default to
# `False` or `do_not_truncate`.
# See [Everything you always wanted to know about padding and truncation](https://huggingface.co/transformers/preprocessing.html#everything-you-always-wanted-to-know-about-padding-and-truncation)
truncation: Union[bool, str] = True,
# The `is_split_into_words` argument applied to your `hf_tokenizer` during tokenization. Set this to `True`
# if your inputs are pre-tokenized (not numericalized)
is_split_into_words: bool = False,
# Any other keyword arguments you want included when using your `hf_tokenizer` to tokenize your inputs
tok_kwargs={},
# Any keyword arguments you want included when generated text
# See [How to generate text](https://huggingface.co/blog/how-to-generate)
text_gen_kwargs={},
# Keyword arguments to apply to `BatchTokenizeTransform`
**kwargs
):
super().__init__(
hf_arch,
hf_config,
hf_tokenizer,
hf_model,
include_labels=include_labels,
ignore_token_id=ignore_token_id,
max_length=max_length,
padding=padding,
truncation=truncation,
is_split_into_words=is_split_into_words,
tok_kwargs=tok_kwargs.copy(),
**kwargs
)
self.lm_strategy = lm_strategy_cls(hf_tokenizer=hf_tokenizer, ignore_token_id=ignore_token_id)
self.text_gen_kwargs, self.ignore_token_id = text_gen_kwargs, ignore_token_id
def encodes(self, samples, return_batch_encoding=False):
# because no target is specific in CLM, fastai will duplicate the inputs (which is just the raw text)
samples, inputs = super().encodes(samples, return_batch_encoding=True)
if len(samples[0]) == 1:
return samples
updated_samples = self.lm_strategy.build_inputs_targets(samples, self.include_labels, inputs)
if return_batch_encoding:
return updated_samples, inputs
return updated_samples
```
Our `LMBatchTokenizeTransform` allows us to update the input's `labels` and our targets appropriately given any language modeling task.
The `labels` argument allows you to forgo calculating the loss yourself by letting Hugging Face return it for you should you choose to do that. Padding tokens are set to -100 by default (e.g., `CrossEntropyLossFlat().ignore_index`) and prevent cross entropy loss from considering token prediction for tokens it should ... i.e., the padding tokens. For more information on the meaning of this argument, see the [Hugging Face glossary entry for "Labels"](https://huggingface.co/transformers/glossary.html#labels)
## Examples
### Using the mid-level API
#### Causal LM
##### Step 1: Get your Hugging Face objects.
```
model_cls = AutoModelForCausalLM
pretrained_model_name = "gpt2"
hf_arch, hf_config, hf_tokenizer, hf_model = get_hf_objects(pretrained_model_name, model_cls=model_cls)
# some tokenizers like gpt and gpt2 do not have a pad token, so we add it here mainly for the purpose
# of setting the "labels" key appropriately (see below)
if hf_tokenizer.pad_token is None:
hf_tokenizer.pad_token = "[PAD]"
```
##### Step 2: Preprocess data
```
preprocessor = LMPreprocessor(hf_tokenizer, chunk_size=128, text_attr=0)
proc_df = preprocessor.process_df(train_df, valid_df)
print(len(proc_df))
proc_df.head(2)
```
##### Step 3: Create your `DataBlock`
```
batch_tok_tfm = LMBatchTokenizeTransform(hf_arch, hf_config, hf_tokenizer, hf_model, lm_strategy_cls=CausalLMStrategy)
blocks = (TextBlock(batch_tokenize_tfm=batch_tok_tfm, input_return_type=CausalLMTextInput), noop)
dblock = DataBlock(blocks=blocks, get_x=ColReader("proc_0"), splitter=ColSplitter(col="is_valid"))
```
##### Step 4: Build your `DataLoaders`
```
dls = dblock.dataloaders(proc_df, bs=4)
b = dls.one_batch()
b[0]["input_ids"].shape, b[0]["labels"].shape, b[1].shape
explode_types(b)
# export
@typedispatch
def show_batch(
# This typedispatched `show_batch` will be called for `CausalLMTextInput` typed inputs
x: CausalLMTextInput,
# Your targets
y,
# Your raw inputs/targets
samples,
# Your `DataLoaders`. This is required so as to get at the Hugging Face objects for
# decoding them into something understandable
dataloaders,
# Your `show_batch` context
ctxs=None,
# The maximum number of items to show
max_n=6,
# Any truncation your want applied to your decoded inputs
trunc_at=None,
# Any other keyword arguments you want applied to `show_batch`
**kwargs
):
# grab our tokenizer and ignore token to decode
tfm = first_blurr_tfm(dataloaders)
hf_tokenizer = tfm.hf_tokenizer
ignore_token_id = tfm.ignore_token_id
res = L(
[
(
hf_tokenizer.decode(s[0], skip_special_tokens=False)[:trunc_at],
hf_tokenizer.decode(s[1][s[1] != ignore_token_id], skip_special_tokens=True)[:trunc_at],
)
for s in samples
]
)
display_df(pd.DataFrame(res, columns=["text", "target"])[:max_n])
return ctxs
dls.show_batch(dataloaders=dls, max_n=2, trunc_at=500)
```
#### Masked LM
##### Step 1: Get your Hugging Face objects.
```
model_cls = AutoModelForMaskedLM
pretrained_model_name = "bert-base-uncased"
hf_arch, hf_config, hf_tokenizer, hf_model = get_hf_objects(pretrained_model_name, model_cls=model_cls)
# some tokenizers like gpt and gpt2 do not have a pad token, so we add it here mainly for the purpose
# of setting the "labels" key appropriately (see below)
if hf_tokenizer.pad_token is None:
hf_tokenizer.pad_token = "[PAD]"
```
##### Step 2: Preprocess data
```
preprocessor = LMPreprocessor(hf_tokenizer, chunk_size=128, text_attr=0)
proc_df = preprocessor.process_df(train_df, valid_df)
print(len(proc_df))
proc_df.head(2)
```
##### Step 3: Create your `DataBlock`
```
batch_tok_tfm = LMBatchTokenizeTransform(hf_arch, hf_config, hf_tokenizer, hf_model, lm_strategy_cls=BertMLMStrategy)
blocks = (TextBlock(batch_tokenize_tfm=batch_tok_tfm, input_return_type=MLMTextInput), noop)
dblock = DataBlock(blocks=blocks, get_x=ColReader("proc_0"), splitter=ColSplitter(col="is_valid"))
```
##### Step 4: Build your `DataLoaders`
```
dls = dblock.dataloaders(proc_df, bs=4)
b = dls.one_batch()
b[0]["input_ids"].shape, b[0]["labels"].shape, b[1].shape
b[0]["input_ids"][0][:20], b[0]["labels"][0][:20], b[1][0][:20]
explode_types(b)
# export
@typedispatch
def show_batch(
# This typedispatched `show_batch` will be called for `MLMTextInput` typed inputs
x: MLMTextInput,
# Your targets
y,
# Your raw inputs/targets
samples,
# Your `DataLoaders`. This is required so as to get at the Hugging Face objects for
# decoding them into something understandable
dataloaders,
# Your `show_batch` context
ctxs=None,
# The maximum number of items to show
max_n=6,
# Any truncation your want applied to your decoded inputs
trunc_at=None,
# Any other keyword arguments you want applied to `show_batch`
**kwargs,
):
# grab our tokenizer and ignore token to decode
tfm = first_blurr_tfm(dataloaders)
hf_tokenizer = tfm.hf_tokenizer
ignore_token_id = tfm.ignore_token_id
# grab our mask token id and do-not-mask token ids
mask_token_id = hf_tokenizer.mask_token_id
vocab = hf_tokenizer.get_vocab()
dnm_tok_ids = [vocab[tok] for tok in list(hf_tokenizer.special_tokens_map.values()) if vocab[tok] != mask_token_id]
res = L()
for s in samples:
# exclue dnm tokens from input
inps = [
hf_tokenizer.decode(tok_id) if (tok_id == mask_token_id or s[1][idx] == ignore_token_id) else f"[{hf_tokenizer.decode(tok_id)}]"
for idx, tok_id in enumerate(s[0])
if (tok_id not in dnm_tok_ids)
]
# replaced masked tokens with "[{actual_token}]"
trgs = [
hf_tokenizer.decode(s[0][idx]) if (tok_id == ignore_token_id) else f"[{hf_tokenizer.decode(tok_id)}]"
for idx, tok_id in enumerate(s[1])
if (s[0][idx] not in dnm_tok_ids)
]
res.append((" ".join(inps[:trunc_at]).strip(), " ".join(trgs[:trunc_at]).strip()))
display_df(pd.DataFrame(res, columns=["text", "target"])[:max_n])
return ctxs
dls.show_batch(dataloaders=dls, max_n=2, trunc_at=250)
```
## Export -
```
# hide
from nbdev.export import notebook2script
notebook2script()
```
| github_jupyter |
<h1 align="center">NEURAL NETWORKS</h1>
Neural networks, also known as artificial neural networks (ANNs) or simulated neural networks (SNNs), are a subset of machine learning and are at the heart of deep learning algorithms. Their name and structure are inspired by the human brain, mimicking the way that biological neurons signal to one another.

Artificial neural networks (ANNs) are comprised of a node layers, containing an input layer, one or more hidden layers, and an output layer. Each node, or artificial neuron, connects to another and has an associated weight and threshold. If the output of any individual node is above the specified threshold value, that node is activated, sending data to the next layer of the network. Otherwise, no data is passed along to the next layer of the network.
Neural networks rely on training data to learn and improve their accuracy over time. However, once these learning algorithms are fine-tuned for accuracy, they are powerful tools in computer science and artificial intelligence, allowing us to classify and cluster data at a high velocity. Tasks in speech recognition or image recognition can take minutes versus hours when compared to the manual identification by human experts. One of the most well-known neural networks is Google’s search algorithm.
Source: https://www.ibm.com/cloud/learn/neural-networks
### Task
The objetive of this project is to analized the clasification using neural networks of coffee samples coming from Colombia, Guatamela y Taiwan. Sets from two countries (Colombia-Guatemala and Guatemala-Taiwan) with 2 parameters (altitude, and flavor) and 5 input parameters (Total_points, Body, Balance, Altitude, Flavor) are analyzed. Finally, the 3 countries will be analyzed together with the 5 parameters and the prediction and raining precisions, as well as the confusion matrix will be used as an indicator of the quality of the classification. The Sklearn library will be used for the creation and testing of the neural network (8,8,8).
### Cool coffee facts! Aftertaste
A coffee’s aftertaste is largely the product of all the other sensory elements you find in each sip. When you swallow, all these elements mingle to form the thing we call aftertaste. Acidity in the aftertaste can be crisp and bright, or mellow and clean. It can cause an aftertaste to linger for ages or disappear quickly. If the coffee is under extracted, you’re drinking espresso, or it just happens to be a really acidic bean, you may feel a light burn that travels down your throat.
Sweetness is a great thing to find in the aftertaste. A coffee that goes down with a pleasant sweetness is probably going to leave a hint of that sweetness there for you to enjoy for a while. Think of how your throat feels sweet and clean after a bite of fresh cake. Bitterness is rarely something you want to experience in your coffee’s aftertaste. More often than not, it’s harsh and biting. It can even create a slight scratchy feeling as you swallow. Generally, a highly bitter aftertaste is the result of over roasting or over extraction.
```
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.neural_network import MLPClassifier
from sklearn.neural_network import MLPRegressor
# Import necessary modules
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from math import sqrt
from sklearn.metrics import r2_score
from sklearn.metrics import confusion_matrix
from sklearn import metrics
from sklearn.metrics import classification_report
# Set theme for plotting
sns.set_theme()
# Import the data
cafe = pd.read_csv("coffee_data.csv")
cafe.rename(columns = {'Country.of.Origin':'Country', 'Total.Cup.Points':'Total_points'}, inplace = True)
# Plot the data
countries = ["Taiwan", "Guatemala","Colombia"]
colors = ["red", "magenta", "lightseagreen"]
fig, ax = plt.subplots(figsize = (10, 8))
for country, color in zip(countries, colors):
temp_df = cafe[cafe.Country == country]
ax.scatter(temp_df.Flavor,
temp_df.altitude_mean_meters,
c = color,
label = country,
)
ax.set_xlabel("Flavor", fontsize = 15)
ax.set_ylabel("Altitude [m]", fontsize = 15)
ax.legend()
plt.show()
```
As mentioned before, these three countries (Colombia, Guatemala, Taiwan) represent 3 types of high quality coffees with different environmental conditions, but with similar cup quality values. Classification is a complex task and we will analyze the results using an 8,8,8 layer neural network.
### Guatemala Colombia
#### Five elements
```
values=['Guatemala','Colombia']
cafe_filter= cafe[cafe.Country.isin(values)]
X1 = cafe_filter[['Total_points', 'Body', 'Balance', 'Flavor',
'altitude_mean_meters']]
y1 = cafe_filter['Country']
X1_train, X1_test, y1_train, y1_test = train_test_split(X1, y1, test_size=0.3,random_state=42)
mlp1 = MLPClassifier(hidden_layer_sizes=(8,8,8), activation='relu', solver='adam', max_iter=500)
mlp1.fit(X1_train,y1_train)
predict_train1 = mlp1.predict(X1_train)
predict_test1 = mlp1.predict(X1_test)
print ("Train - Accuracy :", metrics.accuracy_score(y1_train, predict_train1))
print ("Test - Accuracy :", metrics.accuracy_score(y1_test, predict_test1))
print(classification_report(y1_test,predict_test1))
fig = plt.figure(figsize = (10,8))
cm1 = confusion_matrix(y1_test, predict_test1)
sns.heatmap(cm1,annot=True,fmt='g')
plt.title('Confusion matrix for Neural Networks (Colombia-Guatemala) 5 elements',fontsize=14)
```
#### Guatemala-Colombia Using 2 elements
```
values=['Guatemala','Colombia']
cafe_filter= cafe[cafe.Country.isin(values)]
X2 = cafe_filter[['Flavor','altitude_mean_meters']]
y2 = cafe_filter['Country']
X2_train, X2_test, y2_train, y2_test = train_test_split(X2, y2, test_size=0.3,random_state=42)
mlp2 = MLPClassifier(hidden_layer_sizes=(8,8,8), activation='relu', solver='adam', max_iter=500)
mlp2.fit(X2_train,y2_train)
predict_train2 = mlp2.predict(X2_train)
predict_test2 = mlp2.predict(X2_test)
print ("Train - Accuracy :", metrics.accuracy_score(y2_train, predict_train2))
print ("Test - Accuracy :", metrics.accuracy_score(y2_test, predict_test2))
print(classification_report(y2_test,predict_test2))
fig = plt.figure(figsize = (10,8))
cm2 = confusion_matrix(y2_test, predict_test2)
sns.heatmap(cm2,annot=True,fmt='g')
plt.title('Confusion matrix for Neural Networks (Colombia-Guatemala) 2 elements',fontsize=14)
```
Several tests were performed in which the result of the accuracy in the classification varied, getting values from 20% to a maximum of 80% for the 5 elements, and from 10% to 70% for the classifier with 3 parameters. This fluctuation is due to the random selection of the training and test data values. If we use the maximum value obtained we have 80% for 5 elements and 70% for 2 elements using neural networks, although the average result could be lower.
### Guatemala- Taiwan
#### Five elements
```
values=['Guatemala','Taiwan']
cafe_filter= cafe[cafe.Country.isin(values)]
X3 = cafe_filter[['Total_points', 'Body', 'Balance', 'Flavor',
'altitude_mean_meters']]
y3 = cafe_filter['Country']
X3_train, X3_test, y3_train, y3_test = train_test_split(X3, y3, test_size=0.3,random_state=42)
mlp3 = MLPClassifier(hidden_layer_sizes=(8,8,8), activation='relu', solver='adam', max_iter=500)
mlp3.fit(X3_train,y3_train)
predict_train3 = mlp3.predict(X3_train)
predict_test3 = mlp3.predict(X3_test)
print ("Train - Accuracy :", metrics.accuracy_score(y3_train, predict_train3))
print ("Test - Accuracy :", metrics.accuracy_score(y3_test, predict_test3))
print(classification_report(y3_test,predict_test3))
fig = plt.figure(figsize = (10,8))
cm3 = confusion_matrix(y3_test, predict_test3)
sns.heatmap(cm3,annot=True,fmt='g')
plt.title('Confusion matrix for Neural Networks (Guatemala-Taiwan) 5 elements',fontsize=14)
```
#### Guatemala-Taiwan Using 2 elements
```
values=['Guatemala','Taiwan']
cafe_filter= cafe[cafe.Country.isin(values)]
X4 = cafe_filter[['Flavor', 'altitude_mean_meters']]
y4 = cafe_filter['Country']
X4_train, X4_test, y4_train, y4_test = train_test_split(X4, y4, test_size=0.2,random_state=42)
mlp4 = MLPClassifier(hidden_layer_sizes=(8,8,8), activation='relu', solver='adam', max_iter=500)
mlp4.fit(X4_train,y4_train)
predict_train4 = mlp4.predict(X4_train)
predict_test4 = mlp4.predict(X4_test)
print ("Train - Accuracy :", metrics.accuracy_score(y4_train, predict_train4))
print ("Test - Accuracy :", metrics.accuracy_score(y4_test, predict_test4))
print(classification_report(y4_test,predict_test4))
fig = plt.figure(figsize = (10,8))
cm4 = confusion_matrix(y4_test, predict_test4)
sns.heatmap(cm4,annot=True,fmt='g')
plt.title('Confusion matrix for Neural Networks (Guatemala-Taiwan) 2 elements',fontsize=14)
```
Several tests were performed for Guatemala-Taiwan, in which the result of the accuracy in the classification varied, getting values from 25% to a maximum of 95% for the 5 elements, and from 30% to 93% for the classifier with 3 parameters. This fluctuation is due to the random selection of the training and test data values. If we use the maximum value obtained we have 95% for 5 elements and 93% for 2 elements using neural networks, although the average result could be lower. This eaiser clasification task fits better for neural networks algorithm.
### Colombia-Guatemala-Taiwan
```
values=['Guatemala','Taiwan', 'Colombia']
cafe_filter= cafe[cafe.Country.isin(values)]
X5 = cafe_filter[['Total_points', 'Body', 'Balance', 'Flavor',
'altitude_mean_meters']]
y5 = cafe_filter['Country']
X5_train, X5_test, y5_train, y5_test = train_test_split(X5, y5, test_size=0.3,random_state=42)
mlp5 = MLPClassifier(hidden_layer_sizes=(8,8,8), activation='relu', solver='adam', max_iter=500)
mlp5.fit(X5_train,y5_train)
predict_train5 = mlp5.predict(X5_train)
predict_test5 = mlp5.predict(X5_test)
print ("Train - Accuracy :", metrics.accuracy_score(y5_train, predict_train5))
print ("Test - Accuracy :", metrics.accuracy_score(y5_test, predict_test5))
print(classification_report(y5_test,predict_test5))
fig = plt.figure(figsize = (10,8))
cm5 = confusion_matrix(y5_test, predict_test5)
sns.heatmap(cm5,annot=True,fmt='g')
plt.title('Confusion matrix for Neural Network, 5 elements, three countries',fontsize=14)
```
For this case, the 5 parameters clasifier was teste several time . The result of the accuracy in the classification varied, getting values from 15% to a maximum of 75%. This more complicated clasification task was harder for neural networks algorithm.
| github_jupyter |
```
import numpy as np
import pandas as pd
import scipy
import scipy.linalg
import matplotlib.pyplot as plt
import sklearn.metrics
import sklearn.neighbors
import time
import os
import torch
from torch import nn
import torch.nn.functional as F
from torch.utils.data import DataLoader, Dataset, TensorDataset
import ipdb
import bda_utils
bda_utils.setup_seed(10)
```
# 1. BDA Part
## 1.a. Define BDA methodology
```
def kernel(ker, X1, X2, gamma):
K = None
if not ker or ker == 'primal':
K = X1
elif ker == 'linear':
if X2 is not None:
K = sklearn.metrics.pairwise.linear_kernel(
np.asarray(X1).T, np.asarray(X2).T)
else:
K = sklearn.metrics.pairwise.linear_kernel(np.asarray(X1).T)
elif ker == 'rbf':
if X2 is not None:
K = sklearn.metrics.pairwise.rbf_kernel(
np.asarray(X1).T, np.asarray(X2).T, gamma)
else:
K = sklearn.metrics.pairwise.rbf_kernel(
np.asarray(X1).T, None, gamma)
return K
def proxy_a_distance(source_X, target_X):
"""
Compute the Proxy-A-Distance of a source/target representation
"""
nb_source = np.shape(source_X)[0]
nb_target = np.shape(target_X)[0]
train_X = np.vstack((source_X, target_X))
train_Y = np.hstack((np.zeros(nb_source, dtype=int),
np.ones(nb_target, dtype=int)))
clf = svm.LinearSVC(random_state=0)
clf.fit(train_X, train_Y)
y_pred = clf.predict(train_X)
error = metrics.mean_absolute_error(train_Y, y_pred)
dist = 2 * (1 - 2 * error)
return dist
def estimate_mu(_X1, _Y1, _X2, _Y2):
adist_m = proxy_a_distance(_X1, _X2)
C = len(np.unique(_Y1))
epsilon = 1e-3
list_adist_c = []
for i in range(1, C + 1):
ind_i, ind_j = np.where(_Y1 == i), np.where(_Y2 == i)
Xsi = _X1[ind_i[0], :]
Xtj = _X2[ind_j[0], :]
adist_i = proxy_a_distance(Xsi, Xtj)
list_adist_c.append(adist_i)
adist_c = sum(list_adist_c) / C
mu = adist_c / (adist_c + adist_m)
if mu > 1:
mu = 1
if mu < epsilon:
mu = 0
return mu
class BDA:
def __init__(self, kernel_type='primal', dim=30, lamb=1, mu=0.5, gamma=1, T=10, mode='BDA', estimate_mu=False):
'''
Init func
:param kernel_type: kernel, values: 'primal' | 'linear' | 'rbf'
:param dim: dimension after transfer
:param lamb: lambda value in equation
:param mu: mu. Default is -1, if not specificied, it calculates using A-distance
:param gamma: kernel bandwidth for rbf kernel
:param T: iteration number
:param mode: 'BDA' | 'WBDA'
:param estimate_mu: True | False, if you want to automatically estimate mu instead of manally set it
'''
self.kernel_type = kernel_type
self.dim = dim
self.lamb = lamb
self.mu = mu
self.gamma = gamma
self.T = T
self.mode = mode
self.estimate_mu = estimate_mu
def fit(self, Xs, Ys, Xt, Yt):
'''
Transform and Predict using 1NN as JDA paper did
:param Xs: ns * n_feature, source feature
:param Ys: ns * 1, source label
:param Xt: nt * n_feature, target feature
:param Yt: nt * 1, target label
:return: acc, y_pred, list_acc
'''
# ipdb.set_trace()
list_acc = []
X = np.hstack((Xs.T, Xt.T)) # X.shape: [n_feature, ns+nt]
X_mean = np.linalg.norm(X, axis=0) # why it's axis=0? the average of features
X_mean[X_mean==0] = 1
X /= X_mean
m, n = X.shape
ns, nt = len(Xs), len(Xt)
e = np.vstack((1 / ns * np.ones((ns, 1)), -1 / nt * np.ones((nt, 1))))
C = np.unique(Ys)
H = np.eye(n) - 1 / n * np.ones((n, n))
mu = self.mu
M = 0
Y_tar_pseudo = None
Xs_new = None
for t in range(self.T):
print('\tStarting iter %i'%t)
N = 0
M0 = e * e.T * len(C)
# ipdb.set_trace()
if Y_tar_pseudo is not None:
for i in range(len(C)):
e = np.zeros((n, 1))
Ns = len(Ys[np.where(Ys == C[i])])
Nt = len(Y_tar_pseudo[np.where(Y_tar_pseudo == C[i])])
if self.mode == 'WBDA':
Ps = Ns / len(Ys)
Pt = Nt / len(Y_tar_pseudo)
alpha = Pt / Ps
# mu = 1
else:
alpha = 1
tt = Ys == C[i]
e[np.where(tt == True)] = 1 / Ns
# ipdb.set_trace()
yy = Y_tar_pseudo == C[i]
ind = np.where(yy == True)
inds = [item + ns for item in ind]
try:
e[tuple(inds)] = -alpha / Nt
e[np.isinf(e)] = 0
except:
e[tuple(inds)] = 0 # ?
N = N + np.dot(e, e.T)
# ipdb.set_trace()
# In BDA, mu can be set or automatically estimated using A-distance
# In WBDA, we find that setting mu=1 is enough
if self.estimate_mu and self.mode == 'BDA':
if Xs_new is not None:
mu = estimate_mu(Xs_new, Ys, Xt_new, Y_tar_pseudo)
else:
mu = 0
# ipdb.set_trace()
M = (1 - mu) * M0 + mu * N
M /= np.linalg.norm(M, 'fro')
# ipdb.set_trace()
K = kernel(self.kernel_type, X, None, gamma=self.gamma)
n_eye = m if self.kernel_type == 'primal' else n
a, b = np.linalg.multi_dot([K, M, K.T]) + self.lamb * np.eye(n_eye), np.linalg.multi_dot([K, H, K.T])
w, V = scipy.linalg.eig(a, b)
ind = np.argsort(w)
A = V[:, ind[:self.dim]]
Z = np.dot(A.T, K)
Z_mean = np.linalg.norm(Z, axis=0) # why it's axis=0?
Z_mean[Z_mean==0] = 1
Z /= Z_mean
Xs_new, Xt_new = Z[:, :ns].T, Z[:, ns:].T
global device
model = sklearn.svm.SVC(kernel='linear').fit(Xs_new, Ys.ravel())
Y_tar_pseudo = model.predict(Xt_new)
# ipdb.set_trace()
acc = sklearn.metrics.mean_squared_error(Y_tar_pseudo, Yt) # Yt is already in classes
print(acc)
return Xs_new, Xt_new, A #, acc, Y_tar_pseudo, list_acc
```
## 1.b. Load Data
```
Xs, Xt = bda_utils.load_data(if_weekday=1, if_interdet=1)
Xs = Xs[:,8:9]
Xt = Xt[:,8:9]
Xs, Xs_min, Xs_max = bda_utils.normalize2D(Xs)
Xt, Xt_min, Xt_max = bda_utils.normalize2D(Xt)
for i in range(Xs.shape[1]):
plt.figure(figsize=[20,4])
plt.plot(Xs[:, i])
plt.plot(Xt[:, i])
```
## 1.d. Hyperparameters
```
label_seq_len = 7
# batch_size = full batch
seq_len = 12
reduced_dim = 4
inp_dim = min(Xs.shape[1], Xt.shape[1])
label_dim = min(Xs.shape[1], Xt.shape[1])
hid_dim = 12
layers = 1
lamb = 2
MU = 0.7
bda_dim = label_seq_len-4
kernel_type = 'linear'
hyper = {
'inp_dim':inp_dim,
'label_dim':label_dim,
'label_seq_len':label_seq_len,
'seq_len':seq_len,
'reduced_dim':reduced_dim,
'hid_dim':hid_dim,
'layers':layers,
'lamb':lamb,
'MU': MU,
'bda_dim':bda_dim,
'kernel_type':kernel_type}
hyper = pd.DataFrame(hyper, index=['Values'])
hyper
```
## 1.e. Apply BDA and get $Xs_{new}$, $Xt_{new}$
```
Xs = Xs[:96, :]
# [sample size, seq_len, inp_dim (dets)], [sample size, label_seq_len, inp_dim (dets)]
Xs_3d, Ys_3d = bda_utils.sliding_window(Xs, Xs, seq_len, label_seq_len)
Xt_3d, Yt_3d = bda_utils.sliding_window(Xt, Xt, seq_len, label_seq_len)
Ys_3d = Ys_3d[:, label_seq_len-1:, :]
Yt_3d = Yt_3d[:, label_seq_len-1:, :]
print(Xs_3d.shape)
print(Ys_3d.shape)
print(Xt_3d.shape)
print(Yt_3d.shape)
t_s = time.time()
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
Xs_train_3d = []
Ys_train_3d = []
Xt_valid_3d = []
Xt_train_3d = []
Yt_valid_3d = []
Yt_train_3d = []
for i in range(Xs_3d.shape[2]):
print('Starting det %i'%i)
bda = BDA(kernel_type='linear', dim=seq_len-reduced_dim, lamb=lamb, mu=MU, gamma=1, T=2) # T is iteration time
Xs_new, Xt_new, A = bda.fit(
Xs_3d[:, :, i], bda_utils.get_class(Ys_3d[:, :, i]), Xt_3d[:, :, i], bda_utils.get_class(Yt_3d[:, :, i])
) # input shape: ns, n_feature | ns, n_label_feature
# normalize
Xs_new, Xs_new_min, Xs_new_max = bda_utils.normalize2D(Xs_new)
Xt_new, Xt_new_min, Xt_new_max = bda_utils.normalize2D(Xt_new)
print(Xs_new.shape)
print(Xt_new.shape)
day_train_t = 1
Xs_train = Xs_new.copy()
Ys_train = Ys_3d[:, :, i]
Xt_valid = Xt_new.copy()[int(96*day_train_t):, :]
Xt_train = Xt_new.copy()[:int(96*day_train_t), :]
Yt_valid = Yt_3d[:, :, i].copy()[int(96*day_train_t):, :]
Yt_train = Yt_3d[:, :, i].copy()[:int(96*day_train_t), :]
print('Time spent:%.5f'%(time.time()-t_s))
print(Xs_train.shape)
print(Ys_train.shape)
print(Xt_valid.shape)
print(Xt_train.shape)
print(Yt_valid.shape)
print(Yt_train.shape)
train_x = np.vstack([Xs_train, Xt_train])
train_y = np.vstack([Ys_train, Yt_train])
```
# 2. Regression Part
```
from sklearn.ensemble import RandomForestRegressor
regr = RandomForestRegressor(max_depth=3, random_state=10)
regr.fit(train_x, train_y.flatten())
```
# 3. Evaluation
```
g_t = Yt_valid.flatten()
pred = regr.predict(Xt_valid)
plt.figure(figsize=[16,4])
plt.plot(g_t, label='label')
plt.plot(pred, label='predict')
plt.legend()
print(bda_utils.nrmse_loss_func(pred, g_t, 0))
print(bda_utils.mape_loss_func(pred, g_t, 0))
print(bda_utils.smape_loss_func(pred, g_t, 0))
print(bda_utils.mae_loss_func(pred, g_t, 0))
pred_base = pd.read_csv('./runs_base/base_data_plot/pred_base_RF.csv', header=None)
g_t_base = pd.read_csv('./runs_base/base_data_plot/g_t_base_RF.csv', header=None)
plt.rc('text', usetex=True)
plt.rcParams["font.family"] = "Times New Roman"
plt.figure(figsize=[20, 6], dpi=300)
diff = g_t_base.shape[0]-g_t.shape[0]
plt.plot(range(g_t.shape[0]), g_t_base[diff:]*(903-15)+15, 'b', label='Ground Truth')
plt.plot(range(g_t.shape[0]), pred_base[diff:]*(903-15)+15, 'g', label='Base Model (RF)')
# plt.figure()
# plt.plot(range(371), g_t_bda)
plt.plot(range(g_t.shape[0]), pred*(903-15)+15, 'r', label='BDA (RF)')
plt.legend(loc=1, fontsize=18)
plt.xlabel('Time [15 min]', fontsize=18)
plt.ylabel('Flow [veh/hr]', fontsize=18)
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.

# Automated Machine Learning
**Beer Production Forecasting**
## Contents
1. [Introduction](#Introduction)
1. [Setup](#Setup)
1. [Data](#Data)
1. [Train](#Train)
1. [Evaluate](#Evaluate)
## Introduction
This notebook demonstrates demand forecasting for Beer Production Dataset using AutoML.
AutoML highlights here include using Deep Learning forecasts, Arima, Prophet, Remote Execution and Remote Inferencing, and working with the `forecast` function. Please also look at the additional forecasting notebooks, which document lagging, rolling windows, forecast quantiles, other ways to use the forecast function, and forecaster deployment.
Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.
An Enterprise workspace is required for this notebook. To learn more about creating an Enterprise workspace or upgrading to an Enterprise workspace from the Azure portal, please visit our [Workspace page.](https://docs.microsoft.com/azure/machine-learning/service/concept-workspace#upgrade)
Notebook synopsis:
1. Creating an Experiment in an existing Workspace
2. Configuration and remote run of AutoML for a time-series model exploring Regression learners, Arima, Prophet and DNNs
4. Evaluating the fitted model using a rolling test
## Setup
```
import os
import azureml.core
import pandas as pd
import numpy as np
import logging
import warnings
from pandas.tseries.frequencies import to_offset
# Squash warning messages for cleaner output in the notebook
warnings.showwarning = lambda *args, **kwargs: None
from azureml.core.workspace import Workspace
from azureml.core.experiment import Experiment
from azureml.train.automl import AutoMLConfig
from matplotlib import pyplot as plt
from sklearn.metrics import mean_absolute_error, mean_squared_error
from azureml.train.estimator import Estimator
```
This sample notebook may use features that are not available in previous versions of the Azure ML SDK.
```
print("This notebook was created using version 1.16.0 of the Azure ML SDK")
print("You are currently using version", azureml.core.VERSION, "of the Azure ML SDK")
```
As part of the setup you have already created a <b>Workspace</b>. To run AutoML, you also need to create an <b>Experiment</b>. An Experiment corresponds to a prediction problem you are trying to solve, while a Run corresponds to a specific approach to the problem.
```
ws = Workspace.from_config()
# choose a name for the run history container in the workspace
experiment_name = 'beer-remote-cpu'
experiment = Experiment(ws, experiment_name)
output = {}
output['Subscription ID'] = ws.subscription_id
output['Workspace'] = ws.name
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Run History Name'] = experiment_name
pd.set_option('display.max_colwidth', -1)
outputDf = pd.DataFrame(data = output, index = [''])
outputDf.T
```
### Using AmlCompute
You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for your AutoML run. In this tutorial, you use `AmlCompute` as your training compute resource.
```
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# Choose a name for your CPU cluster
cpu_cluster_name = "beer-cluster"
# Verify that cluster does not exist already
try:
compute_target = ComputeTarget(workspace=ws, name=cpu_cluster_name)
print('Found existing cluster, use it.')
except ComputeTargetException:
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D2_V2',
max_nodes=4)
compute_target = ComputeTarget.create(ws, cpu_cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
```
## Data
Read Beer demand data from file, and preview data.
Let's set up what we know about the dataset.
**Target column** is what we want to forecast.
**Time column** is the time axis along which to predict.
**Time series identifier columns** are identified by values of the columns listed `time_series_id_column_names`, for example "store" and "item" if your data has multiple time series of sales, one series for each combination of store and item sold.
This dataset has only one time series. Please see the [orange juice notebook](https://github.com/Azure/MachineLearningNotebooks/tree/master/how-to-use-azureml/automated-machine-learning/forecasting-orange-juice-sales) for an example of a multi-time series dataset.
```
import pandas as pd
from pandas import DataFrame
from pandas import Grouper
from pandas import concat
from pandas.plotting import register_matplotlib_converters
register_matplotlib_converters()
plt.figure(figsize=(20, 10))
plt.tight_layout()
plt.subplot(2, 1, 1)
plt.title('Beer Production By Year')
df = pd.read_csv("Beer_no_valid_split_train.csv", parse_dates=True, index_col= 'DATE').drop(columns='grain')
test_df = pd.read_csv("Beer_no_valid_split_test.csv", parse_dates=True, index_col= 'DATE').drop(columns='grain')
plt.plot(df)
plt.subplot(2, 1, 2)
plt.title('Beer Production By Month')
groups = df.groupby(df.index.month)
months = concat([DataFrame(x[1].values) for x in groups], axis=1)
months = DataFrame(months)
months.columns = range(1,13)
months.boxplot()
plt.show()
target_column_name = 'BeerProduction'
time_column_name = 'DATE'
time_series_id_column_names = []
freq = 'M' #Monthly data
```
### Split Training data into Train and Validation set and Upload to Datastores
```
from helper import split_fraction_by_grain
from helper import split_full_for_forecasting
train, valid = split_full_for_forecasting(df, time_column_name)
train.to_csv("train.csv")
valid.to_csv("valid.csv")
test_df.to_csv("test.csv")
datastore = ws.get_default_datastore()
datastore.upload_files(files = ['./train.csv'], target_path = 'beer-dataset/tabular/', overwrite = True,show_progress = True)
datastore.upload_files(files = ['./valid.csv'], target_path = 'beer-dataset/tabular/', overwrite = True,show_progress = True)
datastore.upload_files(files = ['./test.csv'], target_path = 'beer-dataset/tabular/', overwrite = True,show_progress = True)
from azureml.core import Dataset
train_dataset = Dataset.Tabular.from_delimited_files(path = [(datastore, 'beer-dataset/tabular/train.csv')])
valid_dataset = Dataset.Tabular.from_delimited_files(path = [(datastore, 'beer-dataset/tabular/valid.csv')])
test_dataset = Dataset.Tabular.from_delimited_files(path = [(datastore, 'beer-dataset/tabular/test.csv')])
```
### Setting forecaster maximum horizon
The forecast horizon is the number of periods into the future that the model should predict. Here, we set the horizon to 12 periods (i.e. 12 months). Notice that this is much shorter than the number of months in the test set; we will need to use a rolling test to evaluate the performance on the whole test set. For more discussion of forecast horizons and guiding principles for setting them, please see the [energy demand notebook](https://github.com/Azure/MachineLearningNotebooks/tree/master/how-to-use-azureml/automated-machine-learning/forecasting-energy-demand).
```
forecast_horizon = 12
```
## Train
Instantiate a AutoMLConfig object. This defines the settings and data used to run the experiment.
|Property|Description|
|-|-|
|**task**|forecasting|
|**primary_metric**|This is the metric that you want to optimize.<br> Forecasting supports the following primary metrics <br><i>spearman_correlation</i><br><i>normalized_root_mean_squared_error</i><br><i>r2_score</i><br><i>normalized_mean_absolute_error</i>
|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|
|**training_data**|Input dataset, containing both features and label column.|
|**label_column_name**|The name of the label column.|
|**enable_dnn**|Enable Forecasting DNNs|
This step requires an Enterprise workspace to gain access to this feature. To learn more about creating an Enterprise workspace or upgrading to an Enterprise workspace from the Azure portal, please visit our [Workspace page.](https://docs.microsoft.com/azure/machine-learning/service/concept-workspace#upgrade).
```
from azureml.automl.core.forecasting_parameters import ForecastingParameters
forecasting_parameters = ForecastingParameters(
time_column_name=time_column_name, forecast_horizon=forecast_horizon
)
automl_config = AutoMLConfig(task='forecasting',
primary_metric='normalized_root_mean_squared_error',
experiment_timeout_hours = 1,
training_data=train_dataset,
label_column_name=target_column_name,
validation_data=valid_dataset,
verbosity=logging.INFO,
compute_target=compute_target,
max_concurrent_iterations=4,
max_cores_per_iteration=-1,
enable_dnn=True,
forecasting_parameters=forecasting_parameters)
```
We will now run the experiment, starting with 10 iterations of model search. The experiment can be continued for more iterations if more accurate results are required. Validation errors and current status will be shown when setting `show_output=True` and the execution will be synchronous.
```
remote_run = experiment.submit(automl_config, show_output= False)
remote_run
# If you need to retrieve a run that already started, use the following code
# from azureml.train.automl.run import AutoMLRun
# remote_run = AutoMLRun(experiment = experiment, run_id = '<replace with your run id>')
remote_run.wait_for_completion()
```
Displaying the run objects gives you links to the visual tools in the Azure Portal. Go try them!
### Retrieve the Best Model for Each Algorithm
Below we select the best pipeline from our iterations. The get_output method on automl_classifier returns the best run and the fitted model for the last fit invocation. There are overloads on get_output that allow you to retrieve the best run and fitted model for any logged metric or a particular iteration.
```
from helper import get_result_df
summary_df = get_result_df(remote_run)
summary_df
from azureml.core.run import Run
from azureml.widgets import RunDetails
forecast_model = 'TCNForecaster'
if not forecast_model in summary_df['run_id']:
forecast_model = 'ForecastTCN'
best_dnn_run_id = summary_df['run_id'][forecast_model]
best_dnn_run = Run(experiment, best_dnn_run_id)
best_dnn_run.parent
RunDetails(best_dnn_run.parent).show()
best_dnn_run
RunDetails(best_dnn_run).show()
```
## Evaluate on Test Data
We now use the best fitted model from the AutoML Run to make forecasts for the test set.
We always score on the original dataset whose schema matches the training set schema.
```
from azureml.core import Dataset
test_dataset = Dataset.Tabular.from_delimited_files(path = [(datastore, 'beer-dataset/tabular/test.csv')])
# preview the first 3 rows of the dataset
test_dataset.take(5).to_pandas_dataframe()
compute_target = ws.compute_targets['beer-cluster']
test_experiment = Experiment(ws, experiment_name + "_test")
import os
import shutil
script_folder = os.path.join(os.getcwd(), 'inference')
os.makedirs(script_folder, exist_ok=True)
shutil.copy('infer.py', script_folder)
from helper import run_inference
test_run = run_inference(test_experiment, compute_target, script_folder, best_dnn_run, test_dataset, valid_dataset, forecast_horizon,
target_column_name, time_column_name, freq)
RunDetails(test_run).show()
from helper import run_multiple_inferences
summary_df = run_multiple_inferences(summary_df, experiment, test_experiment, compute_target, script_folder, test_dataset,
valid_dataset, forecast_horizon, target_column_name, time_column_name, freq)
for run_name, run_summary in summary_df.iterrows():
print(run_name)
print(run_summary)
run_id = run_summary.run_id
test_run_id = run_summary.test_run_id
test_run = Run(test_experiment, test_run_id)
test_run.wait_for_completion()
test_score = test_run.get_metrics()[run_summary.primary_metric]
summary_df.loc[summary_df.run_id == run_id, 'Test Score'] = test_score
print("Test Score: ", test_score)
summary_df
```
| github_jupyter |
## Fern Fractals
```
import numpy as np
import matplotlib.pyplot as plt
```
The Barnsley Fern is a fractal that resembles the Black Spleenwort species of fern. It is constructed by plotting a sequence of points in the $(x,y)$ plane, starting at $(0,0)$, generated by following the affine transformations $f_1$, $f_2$, $f_2$, and $f_4$ where each transformation is applied to the previous point and chosen at random with probabilities $p_1 = 0.01$, $p_2 = 0.85$, $p_3 = 0.07$, and $p_4 = 0.07$
$$ f_1(x,y) =
\begin{bmatrix}
0 & 0 \\
0 & 0.16
\end{bmatrix}
\begin{bmatrix}
x \\
y
\end{bmatrix}
$$
$$ f_2(x,y) =
\begin{bmatrix}
0.85 & 0.04 \\
-0.04 & 0.85
\end{bmatrix}
\begin{bmatrix}
x \\
y
\end{bmatrix}
+
\begin{bmatrix}
0 \\
1.6
\end{bmatrix}
$$
$$ f_3(x,y) =
\begin{bmatrix}
0.2 & -.26 \\
0.23 & 0.22
\end{bmatrix}
\begin{bmatrix}
x \\
y
\end{bmatrix}
+
\begin{bmatrix}
0 \\
1.6
\end{bmatrix}
$$
$$ f_4(x,y) =
\begin{bmatrix}
-0.15 & 0.28 \\
0.26 & 0.24
\end{bmatrix}
\begin{bmatrix}
x \\
y
\end{bmatrix}
+
\begin{bmatrix}
0 \\
0.44
\end{bmatrix}
$$
### Part 0: Introduction to vector tranformations
The functions listed above are known as vector transformations. They are extremely similar to functions you're used to, but instead of operating on scalars, they operate on vectors. Here is an extremely brief introduction to what vector functions are and how we'll use them in this assignment:
The functions we're used to seeing in math take a number input and produce some other number.
$$ f(x) = y$$
Similarily, vector functions take a vector input and produce some other vector.
$$ f(\ \vec v\ ) = \vec u $$
While programming, you've probably already dealt with something extremely similar. For example this following function is a vector transformation. This function takes a vector as an argument ($\vec{v} = [x, y]$) and returns some other vector ($\vec u = [a,b]$).
def my_program(x, y):
a = 3 * x
b = 5 * y
return a, b
In the functions defined above, the input vector [x, y] is multiplied by a matrix and added to another vector. Here's an example of how matrix multiplication works.
$$
\begin{bmatrix}
a & b \\
c & d
\end{bmatrix}
\begin{bmatrix}
x \\
y
\end{bmatrix} =
\begin{bmatrix}
ax + by \\
cx + dy
\end{bmatrix}
$$
The programming function that correspond to this would look like:
def function(x, y):
new_x = a*x + b*y
new_y = c*x + d*y
return new_x, new_y
### Part 1: Defining the transformation functions
Let's start by creating functions for the transformations defined above.
```
# Define f1
def f1(x, y):
# TO DO: fill me in
newx = 0
newy = .16 * y
return newx, newy
assert(abs (f1(1,1)[0] - (0)) < 0.0001)
assert(abs (f1(1,1)[1] - (0.16)) < 0.0001)
# TO DO: create an additional assert statement
assert(abs(f1(0,1)[0]) == 0)
# Define f2
def f2(x, y):
# TO DO: fill me in
newx = .85*x + .04*y
newy = -0.04*x + .85*y + 1.6
return newx, newy
assert(abs (f2(1,1)[0] - (0.89)) < 0.0001)
assert(abs (f2(1,1)[1] - (2.41)) < 0.0001)
# TO DO: create an additional assert statement
assert(f2(0,1)[0] == .04)
# Define f3
def f3(x, y):
# TO DO: fill me in
newx = .2*x - .26*y
newy = .23*x + .22*y + 1.6
return newx, newy
assert(abs (f3(1,1)[0] - (-0.06)) < 0.0001)
assert(abs (f3(1,1)[1] - (2.05)) < 0.0001)
# TO DO: create an additional assert statement
assert(f3(0,1)[0] == -.26)
# Define f4
def f4(x, y):
# TO DO: fill me in
newx = -.15*x + .28*y
newy = .26*x + .24*y + .44
return newx, newy
assert(abs (f4(1,1)[0] - (0.13)) < 0.0001)
assert(abs (f4(1,1)[1] - (0.94)) < 0.0001)
# TO DO: create an additional assert statement
assert(f4(0,1)[0] == .28)
```
### Part 2: Applying the transformations
Now that we have our transformations defined, let's apply them! Defining this function is not hard, but it's also not intuitive. Below I've outlined one way you can create your `generate_fractal` function.
To give you some intuition into what this function is doing, you can think of it as creating a "picture". We start with an `x` and `y` value and apply one of the transformations above. The tranformation gives us a new `x` and `y` value. We plot this `x` and `y` and then we repeat this process for some number of iterations (N).
Here's a walk through of one way you can implement the function below:
1. Initialize `x` and `y` to 0.
2. Create two arrays (I'll refer to them as `XA` and `YA`) both with length N. As you iterate and apply your transformation functions, you'll store your resulting `x` and `y` values in these arrays.
3. Create a list of the functions you defined above
4. Make a `for` loop that iterates N times over the following:
1. Randomly select one of your functions above using the probabilities listed in the introduction, I recommend `np.random.choice(...)`. Make sure your list of probabilities is in the same order as your list of functions.
2. Get your new `x`, `y` values from calling the function selected in the previous step with your existing `x`, `y` values
3. Set the `ith` index of `XA` and `YA` to the new `x` and `y` values respectively. (The `ith` index in this case is whatever number loop you are on)
5. Create a scatter plot of your X and Y value arrays
1. Use `plt.scatter(....)` and include the arguments `s = 1` and `marker = "o"` to get the cleanest image
2. Additionally, add color! To do this, create an additional array of length N (I'll refer to this as `color_array`). The numbers in this array are up to you, but should not be all zero).
3. In your `plt.scatter(...)` call, add arguments `c = color_array` and `cmap = "Greens"`. Once this is working for you, you're welcome to look up and use other colormaps.
<b> I recommend increasing your figure size so your plot is easier to see. Do this by adding `plt.figure(figsize=(10,10))` before you call `plt.scatter(...)`</b>
When your testing your function, use at least N = 5000, or it'll be extremely hard to tell from the plot if you've done it correctly.
```
def generate_fractal(N):
# TO DO: Fill me in!
XA = np.zeros(N)
YA = np.zeros(N)
x = 0
y = 0
func_arr = np.random.choice(np.array([f1, f2, f3, f4]),N,p=[.01, .85, .07, .07])
for i in np.arange(N):
func = func_arr[i]
XA[i] = x
YA[i] = y
x = func(XA[i],YA[i])[0]
y = func(XA[i],YA[i])[1]
color = np.arange(N)
plt.figure(figsize=(10,10))
plt.scatter(XA, YA, s=1, marker = "o", c= color, cmap = "Greens")
generate_fractal(1000)
```
### Part 3: Run your code with N = 100,000.
```
# Call your function with N = 100,000.
# NOTE: This should only take a few seconds to run.
# If your code takes longer, see if there's anything you can change that'll
# improve performance (is there anything in the loop that doesn't need to be?)
N = 100000
generate_fractal(N)
```
| github_jupyter |
```
!pip3 install torch==1.9.0+cu111 torchvision==0.10.0+cu111 torchaudio==0.9.0 -f https://download.pytorch.org/whl/torch_stable.html
!pip install matplotlib scikit-learn pytorch-lightning
!unzip ImageLibrary_6_11_19.zip
import os
from typing import (
List, Any, Callable,
Optional, Tuple, Union, Dict)
import numpy as np
from PIL import Image
import pytorch_lightning as pl
from pytorch_lightning.callbacks import EarlyStopping
from sklearn.metrics import precision_recall_fscore_support
from sklearn.model_selection import train_test_split
import torch
from torch import nn
import torchvision.models as models
from torchvision import transforms
import warnings
warnings.filterwarnings('ignore')
```
# Initial setup
```
np.random.seed(42)
data_path = './ImageLibrary_6_11_19'
training_fraction = 0.7
validation_fraction = 0.2
test_fraction = 0.1
```
# Main classes and helper functions
```
class ImageFilelistDataset(torch.utils.data.Dataset):
def __init__(self,
image_paths: List[str],
labels: List[int],
transform: Optional[Callable] = None,
test: bool = False
):
self.img_paths = image_paths
self.labels = labels
self.transform = transform or transforms.ToTensor()
self.test = test
def _loader(self, path: str) -> Any:
return Image.open(path).convert('RGB')
def __getitem__(self, index: int) -> Union[torch.Tensor, Tuple[torch.Tensor, int]]:
img_path = self.img_paths[index]
img_rgb = self._loader(img_path)
img = self.transform(img_rgb)
img_rgb.close()
if self.test:
return img
target = self.labels[index]
return img, target
def __len__(self) -> int:
return len(self.img_paths)
class ImageDataModule(pl.LightningDataModule):
def __init__(
self,
data_path: str,
batch_size: int = 32,
img_transforms: Optional[Callable] = None
):
super().__init__()
self.data_path = data_path
self.batch_size = batch_size
self.transforms = img_transforms
def _load_paths(self) -> Tuple[List[str], List[int]]:
"""
Finds paths of images of each class
and randomly selects the same number of images of each class.
Thus, the number of samples of each class is the same.
Returns a list with image paths and a list of class labels.
"""
class_dirs = os.listdir(self.data_path)
min_class_imgs_count = min([
len(os.listdir(os.path.join(data_path, class_dir)))
for class_dir in class_dirs
])
data_paths = []
classes = []
for i, class_dir in enumerate(class_dirs):
class_files = os.listdir(os.path.join(data_path, class_dir))
class_files_random = np.random.choice(
class_files,
size=min_class_imgs_count,
replace=False
)
data_paths.extend([
os.path.join(data_path, class_dir, class_file)
for class_file in class_files_random
])
classes.extend([i]*min_class_imgs_count)
return data_paths, classes
def _data_split(self, data_paths: List[str], classes: List[int]):
self.X_train, X_rem, self.y_train, y_rem = train_test_split(
data_paths, classes,
test_size=validation_fraction + test_fraction,
stratify=classes
)
self.X_val, self.X_test, self.y_val, self.y_test = train_test_split(
X_rem, y_rem,
test_size=test_fraction / (validation_fraction+test_fraction),
stratify=y_rem
)
def setup(self, stage: Optional[str] = None):
data_paths, classes = self._load_paths()
self._data_split(data_paths, classes)
def train_dataloader(self):
data = ImageFilelistDataset(
self.X_train, self.y_train, transform=self.transforms['train'])
return torch.utils.data.DataLoader(
dataset=data,
batch_size=self.batch_size,
shuffle=True,
num_workers=8
)
def val_dataloader(self):
data = ImageFilelistDataset(
self.X_val, self.y_val, transform=self.transforms['default'])
return torch.utils.data.DataLoader(
dataset=data,
batch_size=self.batch_size,
num_workers=8
)
def test_dataloader(self):
data = ImageFilelistDataset(
self.X_test, self.y_test, transform=self.transforms['default'])
return torch.utils.data.DataLoader(
dataset=data,
batch_size=self.batch_size,
num_workers=8
)
def predict_dataloader(self):
data = ImageFilelistDataset(
self.X_test, self.y_test, transform=self.transforms['default'], test=True)
return torch.utils.data.DataLoader(
dataset=data,
num_workers=8
)
class ResNet50(pl.LightningModule):
def __init__(self, num_target_classes: int):
super().__init__()
self.model = self._build_model(num_target_classes)
self.num_target_classes = num_target_classes
self.loss = nn.CrossEntropyLoss()
def _build_model(self, num_classes: int) -> models.resnet.ResNet:
model = models.resnet50(pretrained=True)
model.fc = torch.nn.Linear(model.fc.in_features, num_classes)
return model
def _log_metric(self, metric: str, value: float):
self.log(metric, value, prog_bar=True, on_epoch=True, on_step=False)
def forward(self, x):
return self.model(x)
def training_step(self, batch, batch_idx: int):
loss, acc = self._step(batch)
self._log_metric("train_loss", loss)
self._log_metric("train_acc", acc)
return loss
def validation_step(self, batch, batch_idx: int):
loss, acc = self._step(batch)
self._log_metric("val_loss", loss)
self._log_metric("val_acc", acc)
def test_step(self, batch, batch_idx: int):
loss, acc = self._step(batch)
self._log_metric("test_loss", loss)
self._log_metric("test_acc", acc)
def _step(self, batch):
x, y = batch
y_pred = self.forward(x)
acc = self.acc(y_pred, y)
loss = self.loss(y_pred, y)
return loss, acc
def acc(self, y_pred, y_target):
return (y_target == torch.argmax(y_pred, 1)).type(torch.FloatTensor).mean()
def configure_optimizers(self):
return torch.optim.Adam(self.parameters(), lr=1e-4)
def get_binary_metrics(
y_true: List[int],
y_pred: List[torch.Tensor],
target_class: int = 3) -> Dict[str, float]:
"""
Converts the results to a binary form according to `target_class`.
Calculates the precision, recall and F1 score.
"""
y_true = np.array(y_true)
y_true[y_true != target_class] = 0
y_true[y_true == target_class] = 1
y_pred = [v[0] for v in y_pred]
y_pred = torch.argmax(torch.stack(y_pred), dim=1)
y_pred[y_pred != target_class] = 0
y_pred[y_pred == target_class] = 1
precision, recall, f1, _ = precision_recall_fscore_support(
y_true, y_pred.cpu(), average='binary')
return {
"precision": precision,
"recall": recall,
"f1_score": f1
}
```
# Training
```
img_transforms = {
'train': transforms.Compose([
transforms.ToTensor(),
transforms.RandomAffine(
degrees=(-180, 180),
translate=(0.228, 0.228) # Translate in pixels is 175*0.228 = 39.9 px
),
transforms.RandomVerticalFlip(),
transforms.RandomHorizontalFlip(),
transforms.Resize((224, 224))
]),
'default': transforms.Compose([
transforms.ToTensor(),
transforms.Resize((224, 224))
])
}
data_module = ImageDataModule(data_path, img_transforms=img_transforms)
data_module.setup()
num_target_classes = len(os.listdir(data_path))
target_class = os.listdir(data_path).index('trophallaxis')
model = ResNet50(num_target_classes)
trainer = pl.Trainer(
gpus=1, max_epochs=20,
callbacks=[EarlyStopping('val_loss', patience=5)]
)
trainer.fit(model, data_module)
```
# Result metrics
## Train loss and accuracy
```
trainer.validate(model, data_module.train_dataloader())
```
## Validation loss and accuracy
```
trainer.validate(model, data_module.val_dataloader())
```
## Test loss and accuracy
```
trainer.validate(model, data_module.test_dataloader())
```
## Test binary classification metrics
```
y_pred = trainer.predict(model, data_module.predict_dataloader())
get_binary_metrics(data_module.y_test, y_pred)
```
# Save models
```
torch.save(model.model, 'resnet50.pt')
trainer.save_checkpoint('resnet50.ckpt')
```
# Check models loading
```
loaded_model = torch.load('resnet50.pt')
loaded_model.eval()
new_model = ResNet50(num_target_classes)
new_model.model = loaded_model
y_pred = trainer.predict(new_model, data_module.predict_dataloader())
get_binary_metrics(data_module.y_test, y_pred)
loaded_model = ResNet50.load_from_checkpoint('resnet50.ckpt', num_target_classes=num_target_classes)
y_pred = trainer.predict(loaded_model, data_module.predict_dataloader())
get_binary_metrics(data_module.y_test, y_pred)
```
# Binary metrics for the whole dataset
```
full_data = data_module.X_train + data_module.X_val + data_module.X_test
full_target = data_module.y_train + data_module.y_val + data_module.y_test
data = ImageFilelistDataset(full_data, full_target, transform=img_transforms['default'], test=True)
full_dataloader = torch.utils.data.DataLoader(
dataset=data,
num_workers=8
)
y_pred = trainer.predict(model, full_dataloader)
get_binary_metrics(full_target, y_pred)
```
| github_jupyter |
# LSTM PREDICCIÓN DE CONSUMOS
## Import de las librerías necesarias
```
import itertools
import pickle
import keras
import matplotlib.pyplot as plt
import numpy as np # linear algebra
import pandas as pd
import seaborn as sns
import sweetviz as sv
from keras.callbacks import EarlyStopping
from keras.layers import Dense
from keras.layers import Dropout
from keras.layers import LSTM
from keras.models import Sequential
from keras.utils import np_utils
from sklearn import metrics # for the check the error and accuracy of the model
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split # to split the data into two parts
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import StandardScaler # for normalization
```
## Análisis Exploratorio de Datos
```
#Lectura del archivo
df = pd.read_csv('household_power_consumption.txt', sep=';',
parse_dates={'Fecha' : ['Date', 'Time']},
infer_datetime_format=True,
low_memory=False,
na_values=['nan','?'],
index_col='Fecha')
#Tratamiento de NaN
df = df.interpolate()
df.isnull().sum()
#Eliminar voltaje y energia reativa
df = df.drop(['Voltage','Global_reactive_power'], axis=1)
#Cambio de unidades a KWh
df['Sub_metering_1'] = df['Sub_metering_1']/1000
df['Sub_metering_2'] = df['Sub_metering_2']/1000
df['Sub_metering_3'] = df['Sub_metering_3']/1000
#Cambio de potencia a energia (kWh)
df['Energia activa global'] = df['Global_active_power']/60
#df['Energia reactiva global'] = df['Global_reactive_power']/60
#Medidor 4
df['Sub_metering_4'] = df['Energia activa global']-(df['Sub_metering_1']+df['Sub_metering_2']+df['Sub_metering_3'])
#Creacion del nuevo dataframe
df_mean = df[['Global_intensity']].copy()
df_sum = df.drop(['Global_intensity', 'Global_active_power'], axis=1)
#Resampling del nuevo dataframe
df_mean = df_mean.resample('h').mean()
df_sum = df_sum.resample('h').sum()
df1 = df_mean.merge(df_sum, left_index=True, right_index=True)
#Mover energia activa a la ultima posicion
brb = df1.pop('Energia activa global') # remove column b and store it in brb
df1.insert(0, 'Energia activa global', brb) # en primera posicion
correlation_matrix = df1.corr()
plt.figure(figsize = (15,10))
plt.title('Matriz de correlación')
sns.heatmap(data=correlation_matrix, annot=True)
sns.set_theme(style='darkgrid')
sns.histplot(df1['Energia activa global'], stat='probability')
```
# Construcción del modelo
```
def series_to_supervised(data, n_in=1, n_out=1, dropnan=True):
n_vars = 1 if type(data) is list else data.shape[1]
dff = pd.DataFrame(data)
cols, names = list(), list()
# input sequence (t-n, ... t-1)
for i in range(n_in, 0, -1):
cols.append(dff.shift(i))
names += [('var%d(t-%d)' % (j+1, i)) for j in range(n_vars)]
# forecast sequence (t, t+1, ... t+n)
for i in range(0, n_out):
cols.append(dff.shift(-i))
if i == 0:
names += [('var%d(t)' % (j+1)) for j in range(n_vars)]
else:
names += [('var%d(t+%d)' % (j+1, i)) for j in range(n_vars)]
# put it all together
agg = pd.concat(cols, axis=1)
agg.columns = names
# drop rows with NaN values
if dropnan:
agg.dropna(inplace=True)
return agg
```
### Escalar los datos
```
n_hours = 24
n_features = df1.shape[1]
values = df1.values
scaler = MinMaxScaler()
scaler = scaler.fit(values)
scaled = scaler.transform(values)
#Save the scaler
pickle.dump(scaler, open('scaler.pkl','wb'))
supervised = series_to_supervised(scaled, n_hours, 1)
#Eliminamos variables que no vayan a predecirse
variables_eliminadas = supervised.columns[-(n_features-1):]
supervised.drop(variables_eliminadas, axis=1, inplace=True)
```
### Train test split
```
# Dividir en entrenamiento y test
values = supervised.values
n_train_time = int(supervised.shape[0]*0.75)
train = values[:n_train_time, :]
test = values[n_train_time:, :]
# Dividir en entradas y salidas
X_train, y_train = train[:, :-1], train[:, -1]
X_test, y_test = test[:, :-1], test[:, -1]
# Redimensionar el input a 3d [muestras, secuencias, variables]
X_train = X_train.reshape((X_train.shape[0], n_hours, n_features))
X_test = X_test.reshape((X_test.shape[0], n_hours, n_features))
print(X_train.shape, y_train.shape, X_test.shape, y_test.shape)
```
### Definir el modelo
```
early_stopping = EarlyStopping(
min_delta=0.0005, # minimium amount of change to count as an improvement
patience=10, # how many epochs to wait before stopping
restore_best_weights=True,
)
#Preparar la red LSTM
units = 100
#n_outputs = y_train.shape[1]
model = Sequential()
model.add(LSTM(units, input_shape=(X_train.shape[1], X_train.shape[2]), return_sequences=True))
model.add(Dropout(0.2))
model.add(LSTM(units, input_shape=(X_train.shape[1], X_train.shape[2]), return_sequences=False))
model.add(Dropout(0.2))
model.add(Dense(1))
model.compile(loss='mse', optimizer='adam')
```
### Adaptar el modelo a los datos
```
#Elegir hiperparametros
batch_size = 64
epochs = 50
# fit network
history = model.fit(X_train, y_train, epochs=epochs, batch_size=batch_size, validation_data=(X_test, y_test), verbose=1, shuffle=False, callbacks=[early_stopping])
# summarize history for loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper right')
plt.show()
```
### Resultados
```
# Hacer una predicción
yhat = model.predict(X_test)
X_test2 = X_test.reshape((X_test.shape[0], n_hours*n_features))
# Invertir el escalado de la predicción
inv_yhat = np.concatenate((yhat, X_test2[:, -(n_features-1):]), axis=1)
inv_yhat = scaler.inverse_transform(inv_yhat)
inv_yhat = inv_yhat[:,0]
# Invertir el escalado del real
test_y = y_test.reshape((len(y_test), 1))
inv_y = np.concatenate((test_y, X_test2[:, -(n_features-1):]), axis=1)
inv_y = scaler.inverse_transform(inv_y)
inv_y = inv_y[:,0]
# calcular RMSE
rmse = np.sqrt(mean_squared_error(inv_y, inv_yhat))
print('Test RMSE: %.3f' % rmse)
## time steps, every step is one hour (you can easily convert the time step to the actual time index)
## for a demonstration purpose, I only compare the predictions in 200 hours.
hours = 200
plt.figure(figsize = (16,7))
plt.title('Predicción vs real')
aa=[x for x in range(hours)]
plt.plot(aa, inv_y[:hours], marker='.', label="real")
plt.plot(aa, inv_yhat[:hours], 'r', label="prediction")
plt.ylabel('Energia activa global (kWh)', size=15)
plt.xlabel('Horas', size=15)
plt.legend(fontsize=15)
plt.show()
```
## Funciones añadidas
```
# Guardar el Modelo
#model.save('consumos_model.h5')
# Recrea exactamente el mismo modelo solo desde el archivo
model = keras.models.load_model('consumos_model.h5')
```
## Iterar para encontrar los mejores hiperparámetros
```
#Hiperparametros
n_units = [50,100,150,250]
n_capas = [1,2]
n_batch_size = [32, 64]
epochs = 50
i=0
best = 10000
errores = []
parametros = []
for capas in n_capas:
for batch_size in n_batch_size:
for units in n_units:
# Dividir en entrenamiento y test
values = supervised.values
n_train_time = int(supervised.shape[0]*0.75)
train = values[:n_train_time, :]
test = values[n_train_time:, :]
# Dividir en entradas y salidas
X_train, y_train = train[:, :-1], train[:, -1]
X_test, y_test = test[:, :-1], test[:, -1]
# Redimensionar el input a 3d [muestras, secuencias, variables]
X_train = X_train.reshape((X_train.shape[0], n_hours, n_features))
X_test = X_test.reshape((X_test.shape[0], n_hours, n_features))
#Configurar modelo
if(capas==1):
model = Sequential()
model.add(LSTM(units, input_shape=(X_train.shape[1], X_train.shape[2]), return_sequences=False))
model.add(Dropout(0.2))
model.add(Dense(1))
model.compile(loss='mse', optimizer='adam')
if(capas==2):
model = Sequential()
model.add(LSTM(units, input_shape=(X_train.shape[1], X_train.shape[2]), return_sequences=True))
model.add(Dropout(0.2))
model.add(LSTM(units, input_shape=(X_train.shape[1], X_train.shape[2]), return_sequences=False))
model.add(Dropout(0.2))
model.add(Dense(1))
model.compile(loss='mse', optimizer='adam')
# fit network
history = model.fit(X_train, y_train, epochs=epochs, batch_size=batch_size, validation_data=(X_test, y_test), verbose=1, shuffle=False, callbacks=[early_stopping])
# Hacer una predicción
yhat = model.predict(X_test)
X_test2 = X_test.reshape((X_test.shape[0], n_hours*n_features))
# Invertir el escalado de la predicción
inv_yhat = np.concatenate((yhat, X_test2[:, -(n_features-1):]), axis=1)
inv_yhat = scaler.inverse_transform(inv_yhat)
inv_yhat = inv_yhat[:,0]
# Invertir el escalado del real
test_y = y_test.reshape((len(y_test), 1))
inv_y = np.concatenate((test_y, X_test2[:, -(n_features-1):]), axis=1)
inv_y = scaler.inverse_transform(inv_y)
inv_y = inv_y[:,0]
# calcular RMSE
rmse = np.sqrt(mean_squared_error(inv_y, inv_yhat))
print('--------')
print('Iteración número ', i)
print('Parametros:')
print('Capas =', capas)
print('Units =', units)
print('Batch Size =', batch_size)
print('Test RMSE: %.3f' % rmse)
print('--------')
print()
current_parametros = {
'iteracion': i,
'units': units,
'batch_size': batch_size,
'capas': capas,
'error': rmse
}
parametros.append(current_parametros)
print('--------')
errores.append(rmse)
if(rmse < best):
best_model = model
best = rmse
best_parametros = current_parametros
i+=1
results=pd.DataFrame(parametros)
results.to_excel('errores.xlsx')
results.loc[results['error'].idxmin()]
# Guardar el Modelo
best_model.save('best_model.h5')
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# 初学者的 TensorFlow 2.0 教程
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/quickstart/beginner"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />在 TensorFlow.org 观看</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/quickstart/beginner.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />在 Google Colab 运行</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/quickstart/beginner.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />在 GitHub 查看源代码</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/zh-cn/tutorials/quickstart/beginner.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />下载笔记本</a>
</td>
</table>
Note: 我们的 TensorFlow 社区翻译了这些文档。因为社区翻译是尽力而为, 所以无法保证它们是最准确的,并且反映了最新的
[官方英文文档](https://www.tensorflow.org/?hl=en)。如果您有改进此翻译的建议, 请提交 pull request 到
[tensorflow/docs](https://github.com/tensorflow/docs) GitHub 仓库。要志愿地撰写或者审核译文,请加入
[docs-zh-cn@tensorflow.org Google Group](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-zh-cn)。
这是一个 [Google Colaboratory](https://colab.research.google.com/notebooks/welcome.ipynb) 笔记本文件。 Python程序可以直接在浏览器中运行,这是学习 Tensorflow 的绝佳方式。想要学习该教程,请点击此页面顶部的按钮,在Google Colab中运行笔记本。
1. 在 Colab中, 连接到Python运行环境: 在菜单条的右上方, 选择 *CONNECT*。
2. 运行所有的代码块: 选择 *Runtime* > *Run all*。
下载并安装 TensorFlow 2.0 测试版包。将 TensorFlow 载入你的程序:
```
from __future__ import absolute_import, division, print_function, unicode_literals
# 安装 TensorFlow
try:
# Colab only
%tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
```
载入并准备好 [MNIST 数据集](http://yann.lecun.com/exdb/mnist/)。将样本从整数转换为浮点数:
```
mnist = tf.keras.datasets.mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
```
将模型的各层堆叠起来,以搭建 `tf.keras.Sequential` 模型。为训练选择优化器和损失函数:
```
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
```
训练并验证模型:
```
model.fit(x_train, y_train, epochs=5)
model.evaluate(x_test, y_test, verbose=2)
```
现在,这个照片分类器的准确度已经达到 98%。想要了解更多,请阅读 [TensorFlow 教程](https://www.tensorflow.org/tutorials/)。
| github_jupyter |
# OpenVINO benchmarking with 2D U-Net
In this tutorial, we will use the Intel® Distribution of OpenVINO™ Toolkit to perform benchmarking
This tutorial assumes that you have already downloaded and installed [Intel® OpenVINO™](https://software.intel.com/en-us/openvino-toolkit/choose-download) on your computer.
In order to use Intel® OpenVINO™, we need to do a few steps:
1. Convert our Keras model to a Tensorflow model.
1. Freeze the Tensorflow saved format model
1. Use the OpenVINO Model Optimizer to convert the above freezed-model to the OpenVINO Intermediate Representation (IR) format
1. Benchmark using the OpenVINO benchmark tool: `/opt/intel/openvino/deployment_tools/tools/benchmark_tool/benchmark_app.py`
```
import keras
import os
import tensorflow as tf
import numpy as np
import keras as K
import shutil, sys
def dice_coef(y_true, y_pred, axis=(1, 2), smooth=1):
"""
Sorenson (Soft) Dice
\frac{ 2 \times \left | T \right | \cap \left | P \right |}{ \left | T \right | + \left | P \right | }
where T is ground truth mask and P is the prediction mask
"""
intersection = tf.reduce_sum(y_true * y_pred, axis=axis)
union = tf.reduce_sum(y_true + y_pred, axis=axis)
numerator = tf.constant(2.) * intersection + smooth
denominator = union + smooth
coef = numerator / denominator
return tf.reduce_mean(coef)
def soft_dice_coef(target, prediction, axis=(1, 2), smooth=0.01):
"""
Sorenson (Soft) Dice - Don't round the predictions
\frac{ 2 \times \left | T \right | \cap \left | P \right |}{ \left | T \right | + \left | P \right | }
where T is ground truth mask and P is the prediction mask
"""
intersection = tf.reduce_sum(target * prediction, axis=axis)
union = tf.reduce_sum(target + prediction, axis=axis)
numerator = tf.constant(2.) * intersection + smooth
denominator = union + smooth
coef = numerator / denominator
return tf.reduce_mean(coef)
def dice_coef_loss(target, prediction, axis=(1, 2), smooth=1.):
"""
Sorenson (Soft) Dice loss
Using -log(Dice) as the loss since it is better behaved.
Also, the log allows avoidance of the division which
can help prevent underflow when the numbers are very small.
"""
intersection = tf.reduce_sum(prediction * target, axis=axis)
p = tf.reduce_sum(prediction, axis=axis)
t = tf.reduce_sum(target, axis=axis)
numerator = tf.reduce_mean(intersection + smooth)
denominator = tf.reduce_mean(t + p + smooth)
dice_loss = -tf.log(2.*numerator) + tf.log(denominator)
return dice_loss
def combined_dice_ce_loss(y_true, y_pred, axis=(1, 2), smooth=1.,
weight=0.9):
"""
Combined Dice and Binary Cross Entropy Loss
"""
return weight*dice_coef_loss(y_true, y_pred, axis, smooth) + \
(1-weight)*K.losses.binary_crossentropy(y_true, y_pred)
inference_filename = "unet_decathlon_4_8814_128x128_randomcrop-any-input.h5"
model_filename = os.path.join("/home/ubuntu/models/unet", inference_filename)
# Load model
print("Loading Model... ")
model = K.models.load_model(model_filename, custom_objects={
"combined_dice_ce_loss": combined_dice_ce_loss,
"dice_coef_loss": dice_coef_loss,
"soft_dice_coef": soft_dice_coef,
"dice_coef": dice_coef})
print("Model loaded successfully from: " + model_filename)
sess = keras.backend.get_session()
sess.run(tf.global_variables_initializer())
import shutil, sys
output_directory = "/home/ubuntu/models/unet/output"
print("Freezing the graph.")
keras.backend.set_learning_phase(0)
signature = tf.saved_model.signature_def_utils.predict_signature_def(
inputs={'input': model.input}, outputs={'output': model.output})
#If directory exists, delete it and let builder rebuild the TF model.
if os.path.isdir(output_directory):
print (output_directory, "exists already. Deleting the folder")
shutil.rmtree(output_directory)
builder = tf.saved_model.builder.SavedModelBuilder(output_directory)
builder.add_meta_graph_and_variables(sess=sess,
tags=[tf.saved_model.tag_constants.SERVING],
signature_def_map={
tf.saved_model.signature_constants.DEFAULT_SERVING_SIGNATURE_DEF_KEY:signature
}, saver=tf.train.Saver())
builder.save()
print("TensorFlow protobuf version of model is saved in:", output_directory)
print("Model input name = ", model.input.op.name)
print("Model input shape = ", model.input.shape)
print("Model output name = ", model.output.op.name)
print("Model output shape = ", model.output.shape)
output_frozen_model_dir = "/home/ubuntu/models/unet/frozen_model"
output_frozen_graph = output_frozen_model_dir+'/saved_model_frozen.pb'
if not os.path.isdir(output_frozen_model_dir):
os.mkdir(output_frozen_model_dir)
else:
print('Directory', output_frozen_model_dir, 'already exists. Deleting it and re-creating it')
shutil.rmtree(output_frozen_model_dir)
os.mkdir(output_frozen_model_dir)
from tensorflow.python.tools.freeze_graph import freeze_graph
_ = freeze_graph(input_graph="",
input_saver="",
input_binary=False,
input_checkpoint="",
restore_op_name="save/restore_all",
filename_tensor_name="save/Const:0",
clear_devices=True,
initializer_nodes="",
input_saved_model_dir=output_directory,
output_node_names=model.output.op.name,
output_graph=output_frozen_graph)
print("TensorFlow Frozen model model is saved in:", output_frozen_graph)
output_frozen_model_dir = "/home/ubuntu/models/unet/frozen_model"
output_frozen_graph = output_frozen_model_dir+'/saved_model_frozen.pb'
if not os.path.exists(output_frozen_graph):
print(output_frozen_graph + ' doesn\'t exist. Please make sure you have a trained keras to TF frozen model')
!mo_tf.py \
--input_model '/home/ubuntu/models/unet/frozen_model/saved_model_frozen.pb' \
--input_shape=[1,160,160,4] \
--data_type FP32 \
--output_dir /home/ubuntu/models/unet/IR_models/FP32 \
--model_name saved_model
```
#### Run the following command in the terminal
```
mo_tf.py \
--input_model '/home/ubuntu/models/unet/frozen_model/saved_model_frozen.pb' \
--input_shape=[1,160,160,4] \
--data_type FP32 \
--output_dir /home/ubuntu/models/unet/IR_models/FP32 \
--model_name saved_model
```
#### Sample Output:
```
(tensorflow_p36) ubuntu@ip-172-31-46-30:~$ mo_tf.py \
> --input_model '/home/ubuntu/models/unet/frozen_model/saved_model_frozen.pb' \
> --input_shape=[1,160,160,4] \
> --data_type FP32 \
> --output_dir /home/ubuntu/models/unet/IR_models/FP32 \
> --model_name saved_model
Model Optimizer arguments:
Common parameters:
- Path to the Input Model: /home/ubuntu/models/unet/frozen_model/saved_model_frozen.pb
- Path for generated IR: /home/ubuntu/models/unet/IR_models/FP32
- IR output name: saved_model
- Log level: ERROR
- Batch: Not specified, inherited from the model
- Input layers: Not specified, inherited from the model
- Output layers: Not specified, inherited from the model
- Input shapes: [1,160,160,4]
- Mean values: Not specified
- Scale values: Not specified
- Scale factor: Not specified
- Precision of IR: FP32
- Enable fusing: True
- Enable grouped convolutions fusing: True
- Move mean values to preprocess section: False
- Reverse input channels: False
TensorFlow specific parameters:
- Input model in text protobuf format: False
- Path to model dump for TensorBoard: None
- List of shared libraries with TensorFlow custom layers implementation: None
- Update the configuration file with input/output node names: None
- Use configuration file used to generate the model with Object Detection API: None
- Operations to offload: None
- Patterns to offload: None
- Use the config file: None
Model Optimizer version: 2020.1.0-61-gd349c3ba4a
[ SUCCESS ] Generated IR version 10 model.
[ SUCCESS ] XML file: /home/ubuntu/models/unet/IR_models/FP32/saved_model.xml
[ SUCCESS ] BIN file: /home/ubuntu/models/unet/IR_models/FP32/saved_model.bin
[ SUCCESS ] Total execution time: 6.41 seconds.
[ SUCCESS ] Memory consumed: 443 MB.
```
## Benchmark
Benchmark using the following command:
```
python3 /opt/intel/openvino/deployment_tools/tools/benchmark_tool/benchmark_app.py \
-m /home/ubuntu/models/unet/IR_models/FP32/saved_model.xml \
-nireq 1 -nstreams 1
```
#### Sample Output
```
[Step 1/11] Parsing and validating input arguments
[Step 2/11] Loading Inference Engine
[ INFO ] InferenceEngine:
API version............. 2.1.37988
[ INFO ] Device info
CPU
MKLDNNPlugin............ version 2.1
Build................... 37988
[Step 3/11] Reading the Intermediate Representation network
[Step 4/11] Resizing network to match image sizes and given batch
[ INFO ] Network batch size: 1, precision: MIXED
[Step 5/11] Configuring input of the model
[Step 6/11] Setting device configuration
[Step 7/11] Loading the model to the device
[Step 8/11] Setting optimal runtime parameters
[Step 9/11] Creating infer requests and filling input blobs with images
[ INFO ] Network input 'MRImages' precision FP32, dimensions (NCHW): 1 4 160 160
[ WARNING ] No input files were given: all inputs will be filled with random values!
[ INFO ] Infer Request 0 filling
[ INFO ] Fill input 'MRImages' with random values (some binary data is expected)
[Step 10/11] Measuring performance (Start inference asyncronously, 1 inference requests using 1 streams for CPU, limits: 60000 ms duration)
[Step 11/11] Dumping statistics report
Count: 11079 iterations
Duration: 60014.36 ms
Latency: 5.11 ms
Throughput: 184.61 FPS
```
| github_jupyter |
## Agent-Based Model for Wealth
Agent-Based model of sacrifices/taxes based on the affine wealth model from Bruce M. Boghosian
"Is Inequality Inevitable?", Bruce M. Boghosian, Scientific American, October 30, 2019
"Follow the money", Brian Hayes, American Scientist, Vol. 90, 2002, 400-405
```
import pandas as pd
import numpy as np
import random
import matplotlib as mpl
import matplotlib.pyplot as plt
from matplotlib import cm
```
### The agents
```
class Agent:
def __init__(self, money = 10.0, stake = 0.10):
self.wealth = money
self.stake = stake
def tax_wealth(self, rate):
taxes = self.wealth * rate
self.subtract_wealth(taxes)
return taxes
def subtract_wealth(self, amount):
self.wealth = self.wealth - amount
def add_wealth(self, amount):
self.wealth = self.wealth + amount
def interact(self, opponent):
min_wealth = min(self.wealth, opponent.wealth)
stake = min_wealth * self.stake
flip = random.randint(0, 1)
if (flip == 0):
self.add_wealth(stake)
opponent.subtract_wealth(stake)
else:
self.subtract_wealth(stake)
opponent.add_wealth(stake)
a = Agent()
b = Agent()
a.interact(b)
a.interact(b)
a.interact(b)
a.interact(b)
a.interact(b)
print("Agent A wealth", a.wealth)
print("Agent B wealth", b.wealth)
```
### The agent based model
```
class AgentBasedModel:
def __init__(self, n, rate = 0):
self.apply_taxes = rate > 0
self.tax_rate = rate
self.agents = n
self.model = []
self.data = []
for x in range(self.agents): self.model.append(Agent())
def random_agent(self):
i = random.randrange(0, self.agents)
return self.model[i]
def play(self, timestep):
agent1 = self.random_agent()
agent2 = agent1
while (agent1 == agent2):
agent2 = self.random_agent()
agent1.interact(agent2)
if self.apply_taxes:
taxed_agent = self.random_agent()
taxes = taxed_agent.tax_wealth(self.tax_rate) / self.agents
for x in range(self.agents):
agent = self.random_agent()
if agent != taxed_agent: agent.add_wealth(taxes)
def wealth_distribution(self):
return [agent.wealth for agent in self.model]
def wealth_grouped(self, groups = 25):
distribution = self.wealth_distribution()
df = pd.DataFrame(distribution, columns=['Wealth'])
bins = np.arange(0, groups, 1)
return list(df.groupby(pd.cut(df.Wealth, bins)).size().reset_index()[0])
def run(self, timesteps):
for t in range(timesteps):
self.play(t)
iterations = 50000
x_1 = AgentBasedModel(250, 0)
x_1.run(iterations)
x_2 = AgentBasedModel(250, 0)
x_2.run(iterations)
x_3 = AgentBasedModel(250, 0)
x_3.run(iterations)
x_4 = AgentBasedModel(250, 0)
x_4.run(iterations)
x_5 = AgentBasedModel(250, 0)
x_5.run(iterations)
dict = {'w1': x_1.wealth_grouped() , 'w2': x_2.wealth_grouped(), 'w3': x_3.wealth_grouped(), 'w4': x_4.wealth_grouped(), 'w5': x_5.wealth_grouped()}
df_0 = pd.DataFrame(dict)
df_0['Wealth'] = df_0.mean(axis=1)
df_0.head()
x_1 = AgentBasedModel(250, 0.01)
x_1.run(iterations)
x_2 = AgentBasedModel(250, 0.01)
x_2.run(iterations)
x_3 = AgentBasedModel(250, 0.01)
x_3.run(iterations)
x_4 = AgentBasedModel(250, 0.01)
x_4.run(iterations)
x_5 = AgentBasedModel(250, 0.01)
x_5.run(iterations)
dict = {'w1': x_1.wealth_grouped(), 'w2': x_2.wealth_grouped(), 'w3': x_3.wealth_grouped(), 'w4': x_4.wealth_grouped(), 'w5': x_5.wealth_grouped()}
df_1 = pd.DataFrame(dict)
df_1['Wealth'] = df_1.mean(axis=1)
df_1.head()
x_1 = AgentBasedModel(250, 0.02)
x_1.run(iterations)
x_2 = AgentBasedModel(250, 0.02)
x_2.run(iterations)
x_3 = AgentBasedModel(250, 0.02)
x_3.run(iterations)
x_4 = AgentBasedModel(250, 0.02)
x_4.run(iterations)
x_5 = AgentBasedModel(250, 0.02)
x_5.run(iterations)
dict = {'w1': x_1.wealth_grouped(), 'w2': x_2.wealth_grouped(), 'w3': x_3.wealth_grouped(), 'w4': x_4.wealth_grouped(), 'w5': x_5.wealth_grouped()}
df_2 = pd.DataFrame(dict)
df_2['Wealth'] = df_2.mean(axis=1)
df_2.head()
x_1 = AgentBasedModel(250, 0.03)
x_1.run(iterations)
x_2 = AgentBasedModel(250, 0.03)
x_2.run(iterations)
x_3 = AgentBasedModel(250, 0.03)
x_3.run(iterations)
x_4 = AgentBasedModel(250, 0.03)
x_4.run(iterations)
x_5 = AgentBasedModel(250, 0.03)
x_5.run(iterations)
dict = {'w1': x_1.wealth_grouped(), 'w2': x_2.wealth_grouped(), 'w3': x_3.wealth_grouped(), 'w4': x_4.wealth_grouped(), 'w5': x_5.wealth_grouped()}
df_3 = pd.DataFrame(dict)
df_3['Wealth'] = df_3.mean(axis=1)
df_3.head()
x_1 = AgentBasedModel(250, 0.04)
x_1.run(iterations)
x_2 = AgentBasedModel(250, 0.04)
x_2.run(iterations)
x_3 = AgentBasedModel(250, 0.04)
x_3.run(iterations)
x_4 = AgentBasedModel(250, 0.04)
x_4.run(iterations)
x_5 = AgentBasedModel(250, 0.04)
x_5.run(iterations)
dict = {'w1': x_1.wealth_grouped(), 'w2': x_2.wealth_grouped(), 'w3': x_3.wealth_grouped(), 'w4': x_4.wealth_grouped(), 'w5': x_5.wealth_grouped()}
df_4 = pd.DataFrame(dict)
df_4['Wealth'] = df_4.mean(axis=1)
df_4.head()
x_1 = AgentBasedModel(250, 0.05)
x_1.run(iterations)
x_2 = AgentBasedModel(250, 0.05)
x_2.run(iterations)
x_3 = AgentBasedModel(250, 0.05)
x_3.run(iterations)
x_4 = AgentBasedModel(250, 0.05)
x_4.run(iterations)
x_5 = AgentBasedModel(250, 0.05)
x_5.run(iterations)
dict = {'w1': x_1.wealth_grouped(), 'w2': x_2.wealth_grouped(), 'w3': x_3.wealth_grouped(), 'w4': x_4.wealth_grouped(), 'w5': x_5.wealth_grouped()}
df_5 = pd.DataFrame(dict)
df_5['Wealth'] = df_5.mean(axis=1)
df_5.head()
x_1 = AgentBasedModel(250, 0.06)
x_1.run(iterations)
x_2 = AgentBasedModel(250, 0.06)
x_2.run(iterations)
x_3 = AgentBasedModel(250, 0.06)
x_3.run(iterations)
x_4 = AgentBasedModel(250, 0.06)
x_4.run(iterations)
x_5 = AgentBasedModel(250, 0.06)
x_5.run(iterations)
dict = {'w1': x_1.wealth_grouped(), 'w2': x_2.wealth_grouped(), 'w3': x_3.wealth_grouped(), 'w4': x_4.wealth_grouped(), 'w5': x_5.wealth_grouped()}
df_6 = pd.DataFrame(dict)
df_6['Wealth'] = df_6.mean(axis=1)
df_6.head()
x_1 = AgentBasedModel(250, 0.07)
x_1.run(iterations)
x_2 = AgentBasedModel(250, 0.07)
x_2.run(iterations)
x_3 = AgentBasedModel(250, 0.07)
x_3.run(iterations)
x_4 = AgentBasedModel(250, 0.07)
x_4.run(iterations)
x_5 = AgentBasedModel(250, 0.07)
x_5.run(iterations)
dict = {'w1': x_1.wealth_grouped(), 'w2': x_2.wealth_grouped(), 'w3': x_3.wealth_grouped(), 'w4': x_4.wealth_grouped(), 'w5': x_5.wealth_grouped()}
df_7 = pd.DataFrame(dict)
df_7['Wealth'] = df_7.mean(axis=1)
df_7.head()
x_1 = AgentBasedModel(250, 0.08)
x_1.run(iterations)
x_2 = AgentBasedModel(250, 0.08)
x_2.run(iterations)
x_3 = AgentBasedModel(250, 0.08)
x_3.run(iterations)
x_4 = AgentBasedModel(250, 0.08)
x_4.run(iterations)
x_5 = AgentBasedModel(250, 0.08)
x_5.run(iterations)
dict = {'w1': x_1.wealth_grouped(), 'w2': x_2.wealth_grouped(), 'w3': x_3.wealth_grouped(), 'w4': x_4.wealth_grouped(), 'w5': x_5.wealth_grouped()}
df_8 = pd.DataFrame(dict)
df_8['Wealth'] = df_8.mean(axis=1)
df_8.head()
```
### Wealth distribution
Wealth for the simple "yard sale model" without taxes follows a Pareto distribution
```
%matplotlib inline
from scipy.stats import pareto
t = np.arange(0.9, 40., 0.5)
R = pareto.pdf(x=t, b=0.05, loc=0, scale=0.1) * 1000
p2 = plt.bar(range(24), df_0.Wealth, label="Wealth")
p1 = plt.plot(t, R, label="Pareto Distribution", color="black")
plt.ylabel('Probability')
plt.xlabel('Wealth')
plt.title('No sacrifices or taxes')
plt.legend()
plt.show()
mpl.use('svg')
p2 = plt.bar(range(24), df_0.Wealth, label="Wealth")
p1 = plt.plot(t, R, label="Pareto Distribution", color="black")
plt.ylabel('Probability')
plt.xlabel('Wealth')
plt.title('No sacrifices or taxes')
plt.legend()
plt.savefig("WealthDistributionNoTaxes.svg", format="svg")
%matplotlib inline
fig = plt.figure(figsize=(11, 8))
ax = fig.add_subplot(111, projection='3d')
series = 5
n = 24
bins = np.arange(0,n+1,1)
d = list(df_0.Wealth) + list(df_2.Wealth) + list(df_4.Wealth) + list(df_6.Wealth) + list(df_8.Wealth)
_x = np.arange(n)
_y = np.arange(0, (series*2), 2)
_xx, _yy = np.meshgrid(_x, _y)
x, y = _xx.ravel(), _yy.ravel()
bottom = np.zeros_like(d)
width = 0.4
depth = 0.1
ax.bar3d(x, y, bottom, width, depth, d, shade=True)
ax.set_title('Distribution of Wealth\n')
ax.set_ylabel('Sacrifices in %')
ax.set_xlabel('Wealth')
ax.set_zlabel('Probability')
plt.show()
mpl.use('svg')
fig = plt.figure(figsize=(11, 8))
ax = fig.add_subplot(111, projection='3d')
series = 5
n = 24
bins = np.arange(0,n+1,1)
d = list(df_0.Wealth) + list(df_2.Wealth) + list(df_4.Wealth) + list(df_6.Wealth) + list(df_8.Wealth)
_x = np.arange(n)
_y = np.arange(0, (series*2), 2)
_xx, _yy = np.meshgrid(_x, _y)
x, y = _xx.ravel(), _yy.ravel()
bottom = np.zeros_like(d)
width = 0.4
depth = 0.1
ax.bar3d(x, y, bottom, width, depth, d, shade=True)
ax.set_title('Distribution of Wealth\n')
ax.set_ylabel('Sacrifices in %')
ax.set_xlabel('Wealth')
ax.set_zlabel('Probability')
fig.set_tight_layout(True)
plt.savefig("WealthDistributionTaxes3D.svg", format="svg")
%matplotlib inline
series = 8
bins = np.arange(0,25.0,1)
n = len(bins) - 1
d = list(df_1.Wealth) + list(df_2.Wealth) + list(df_3.Wealth) + list(df_4.Wealth) + list(df_5.Wealth) + list(df_6.Wealth) + list(df_7.Wealth) + list(df_8.Wealth)
_x = np.arange(n)
_y = np.arange(1, series+1, 1)
_xx, _yy = np.meshgrid(_x, _y)
x, y = _xx.ravel(), _yy.ravel()
fig = plt.figure(figsize=(12, 8))
ax = plt.axes(projection='3d')
data = np.array(d).reshape((series, n))
ax.plot_surface(_xx, _yy, data, cmap=cm.coolwarm)
# ax.plot_wireframe(_xx, _yy, data)
ax.set_title('Distribution of Wealth')
ax.set_ylabel('Sacrifices in %')
ax.set_xlabel('Wealth')
ax.set_zlabel('Probability')
mpl.use('svg')
fig = plt.figure(figsize=(12, 8))
ax = plt.axes(projection='3d')
data = np.array(d).reshape((series, n))
ax.plot_surface(_xx, _yy, data, cmap=cm.coolwarm)
ax.set_title('Distribution of Wealth')
ax.set_ylabel('Sacrifices in %')
ax.set_xlabel('Wealth')
ax.set_zlabel('Probability')
fig.set_tight_layout(True)
plt.savefig("WealthDistributionTaxes3DSurface.svg", format="svg")
%matplotlib inline
fig, axs = plt.subplots(4, 1, figsize=(10,10), sharex=True, sharey=True)
axs[0].bar(range(24), df_2.Wealth)
axs[0].axvline(x=float(5.0), color="Black")
axs[0].set_title('2% Sacrifices')
axs[1].bar(range(24), df_4.Wealth)
axs[1].axvline(x=float(6.0), color="Black")
axs[1].set_title('4% Sacrifices')
axs[2].bar(range(24), df_6.Wealth)
axs[2].axvline(x=float(7.0), color="Black")
axs[2].set_title('6% Sacrifices')
axs[3].bar(range(24), df_8.Wealth)
axs[3].axvline(x=float(8.0), color="Black")
axs[3].set_title('8% Sacrifices')
for ax in axs.flat:
ax.set(xlabel='Wealth', ylabel='Probability')
# Hide x labels and tick labels for top plots and y ticks for right plots.
for ax in axs.flat:
ax.label_outer()
plt.show()
mpl.use('svg')
fig, axs = plt.subplots(4, 1, figsize=(10,10), sharex=True, sharey=True)
axs[0].bar(range(24), df_2.Wealth)
axs[0].axvline(x=float(5.0), color="Black")
axs[0].set_title('2% Sacrifices')
axs[1].bar(range(24), df_4.Wealth)
axs[1].axvline(x=float(6.0), color="Black")
axs[1].set_title('4% Sacrifices')
axs[2].bar(range(24), df_6.Wealth)
axs[2].axvline(x=float(7.0), color="Black")
axs[2].set_title('6% Sacrifices')
axs[3].bar(range(24), df_8.Wealth)
axs[3].axvline(x=float(8.0), color="Black")
axs[3].set_title('8% Sacrifices')
fig.set_tight_layout(True)
for ax in axs.flat:
ax.set(xlabel='Wealth', ylabel='Probability')
# Hide x labels and tick labels for top plots and y ticks for right plots.
for ax in axs.flat:
ax.label_outer()
plt.savefig("WealthDistributionTaxes.svg", format="svg")
```
| github_jupyter |
# Convolutional Neural Networks: Step by Step
Welcome to Course 4's first assignment! In this assignment, you will implement convolutional (CONV) and pooling (POOL) layers in numpy, including both forward propagation and (optionally) backward propagation.
**Notation**:
- Superscript $[l]$ denotes an object of the $l^{th}$ layer.
- Example: $a^{[4]}$ is the $4^{th}$ layer activation. $W^{[5]}$ and $b^{[5]}$ are the $5^{th}$ layer parameters.
- Superscript $(i)$ denotes an object from the $i^{th}$ example.
- Example: $x^{(i)}$ is the $i^{th}$ training example input.
- Lowerscript $i$ denotes the $i^{th}$ entry of a vector.
- Example: $a^{[l]}_i$ denotes the $i^{th}$ entry of the activations in layer $l$, assuming this is a fully connected (FC) layer.
- $n_H$, $n_W$ and $n_C$ denote respectively the height, width and number of channels of a given layer. If you want to reference a specific layer $l$, you can also write $n_H^{[l]}$, $n_W^{[l]}$, $n_C^{[l]}$.
- $n_{H_{prev}}$, $n_{W_{prev}}$ and $n_{C_{prev}}$ denote respectively the height, width and number of channels of the previous layer. If referencing a specific layer $l$, this could also be denoted $n_H^{[l-1]}$, $n_W^{[l-1]}$, $n_C^{[l-1]}$.
We assume that you are already familiar with `numpy` and/or have completed the previous courses of the specialization. Let's get started!
## 1 - Packages
Let's first import all the packages that you will need during this assignment.
- [numpy](www.numpy.org) is the fundamental package for scientific computing with Python.
- [matplotlib](http://matplotlib.org) is a library to plot graphs in Python.
- np.random.seed(1) is used to keep all the random function calls consistent. It will help us grade your work.
```
import numpy as np
import h5py
import matplotlib.pyplot as plt
%matplotlib inline
plt.rcParams['figure.figsize'] = (5.0, 4.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
%load_ext autoreload
%autoreload 2
np.random.seed(1)
```
## 2 - Outline of the Assignment
You will be implementing the building blocks of a convolutional neural network! Each function you will implement will have detailed instructions that will walk you through the steps needed:
- Convolution functions, including:
- Zero Padding
- Convolve window
- Convolution forward
- Convolution backward (optional)
- Pooling functions, including:
- Pooling forward
- Create mask
- Distribute value
- Pooling backward (optional)
This notebook will ask you to implement these functions from scratch in `numpy`. In the next notebook, you will use the TensorFlow equivalents of these functions to build the following model:
<img src="images/model.png" style="width:800px;height:300px;">
**Note** that for every forward function, there is its corresponding backward equivalent. Hence, at every step of your forward module you will store some parameters in a cache. These parameters are used to compute gradients during backpropagation.
## 3 - Convolutional Neural Networks
Although programming frameworks make convolutions easy to use, they remain one of the hardest concepts to understand in Deep Learning. A convolution layer transforms an input volume into an output volume of different size, as shown below.
<img src="images/conv_nn.png" style="width:350px;height:200px;">
In this part, you will build every step of the convolution layer. You will first implement two helper functions: one for zero padding and the other for computing the convolution function itself.
### 3.1 - Zero-Padding
Zero-padding adds zeros around the border of an image:
<img src="images/PAD.png" style="width:600px;height:400px;">
<caption><center> <u> <font color='purple'> **Figure 1** </u><font color='purple'> : **Zero-Padding**<br> Image (3 channels, RGB) with a padding of 2. </center></caption>
The main benefits of padding are the following:
- It allows you to use a CONV layer without necessarily shrinking the height and width of the volumes. This is important for building deeper networks, since otherwise the height/width would shrink as you go to deeper layers. An important special case is the "same" convolution, in which the height/width is exactly preserved after one layer.
- It helps us keep more of the information at the border of an image. Without padding, very few values at the next layer would be affected by pixels as the edges of an image.
**Exercise**: Implement the following function, which pads all the images of a batch of examples X with zeros. [Use np.pad](https://docs.scipy.org/doc/numpy/reference/generated/numpy.pad.html). Note if you want to pad the array "a" of shape $(5,5,5,5,5)$ with `pad = 1` for the 2nd dimension, `pad = 3` for the 4th dimension and `pad = 0` for the rest, you would do:
```python
a = np.pad(a, ((0,0), (1,1), (0,0), (3,3), (0,0)), 'constant', constant_values = (..,..))
```
```
# GRADED FUNCTION: zero_pad
def zero_pad(X, pad):
"""
Pad with zeros all images of the dataset X. The padding is applied to the height and width of an image,
as illustrated in Figure 1.
Argument:
X -- python numpy array of shape (m, n_H, n_W, n_C) representing a batch of m images
pad -- integer, amount of padding around each image on vertical and horizontal dimensions
Returns:
X_pad -- padded image of shape (m, n_H + 2*pad, n_W + 2*pad, n_C)
"""
### START CODE HERE ### (≈ 1 line)
X_pad = np.pad(X, ((0, 0), (pad, pad), (pad, pad), (0, 0)), 'constant', constant_values=0)
### END CODE HERE ###
return X_pad
np.random.seed(1)
x = np.random.randn(4, 3, 3, 2)
x_pad = zero_pad(x, 2)
print ("x.shape =", x.shape)
print ("x_pad.shape =", x_pad.shape)
print ("x[1,1] =", x[1,1])
print ("x_pad[1,1] =", x_pad[1,1])
fig, axarr = plt.subplots(1, 2)
axarr[0].set_title('x')
axarr[0].imshow(x[0,:,:,0])
axarr[1].set_title('x_pad')
axarr[1].imshow(x_pad[0,:,:,0])
```
**Expected Output**:
<table>
<tr>
<td>
**x.shape**:
</td>
<td>
(4, 3, 3, 2)
</td>
</tr>
<tr>
<td>
**x_pad.shape**:
</td>
<td>
(4, 7, 7, 2)
</td>
</tr>
<tr>
<td>
**x[1,1]**:
</td>
<td>
[[ 0.90085595 -0.68372786]
[-0.12289023 -0.93576943]
[-0.26788808 0.53035547]]
</td>
</tr>
<tr>
<td>
**x_pad[1,1]**:
</td>
<td>
[[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]]
</td>
</tr>
</table>
### 3.2 - Single step of convolution
In this part, implement a single step of convolution, in which you apply the filter to a single position of the input. This will be used to build a convolutional unit, which:
- Takes an input volume
- Applies a filter at every position of the input
- Outputs another volume (usually of different size)
<img src="images/Convolution_schematic.gif" style="width:500px;height:300px;">
<caption><center> <u> <font color='purple'> **Figure 2** </u><font color='purple'> : **Convolution operation**<br> with a filter of 2x2 and a stride of 1 (stride = amount you move the window each time you slide) </center></caption>
In a computer vision application, each value in the matrix on the left corresponds to a single pixel value, and we convolve a 3x3 filter with the image by multiplying its values element-wise with the original matrix, then summing them up and adding a bias. In this first step of the exercise, you will implement a single step of convolution, corresponding to applying a filter to just one of the positions to get a single real-valued output.
Later in this notebook, you'll apply this function to multiple positions of the input to implement the full convolutional operation.
**Exercise**: Implement conv_single_step(). [Hint](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.sum.html).
```
# GRADED FUNCTION: conv_single_step
def conv_single_step(a_slice_prev, W, b):
"""
Apply one filter defined by parameters W on a single slice (a_slice_prev) of the output activation
of the previous layer.
Arguments:
a_slice_prev -- slice of input data of shape (f, f, n_C_prev)
W -- Weight parameters contained in a window - matrix of shape (f, f, n_C_prev)
b -- Bias parameters contained in a window - matrix of shape (1, 1, 1)
Returns:
Z -- a scalar value, result of convolving the sliding window (W, b) on a slice x of the input data
"""
### START CODE HERE ### (≈ 2 lines of code)
# Element-wise product between a_slice and W. Do not add the bias yet.
s = np.multiply(a_slice_prev, W)
# Sum over all entries of the volume s.
Z = np.sum(s)
# Add bias b to Z. Cast b to a float() so that Z results in a scalar value.
Z = Z + float(b)
### END CODE HERE ###
return Z
np.random.seed(1)
a_slice_prev = np.random.randn(4, 4, 3)
W = np.random.randn(4, 4, 3)
b = np.random.randn(1, 1, 1)
Z = conv_single_step(a_slice_prev, W, b)
print("Z =", Z)
```
**Expected Output**:
<table>
<tr>
<td>
**Z**
</td>
<td>
-6.99908945068
</td>
</tr>
</table>
### 3.3 - Convolutional Neural Networks - Forward pass
In the forward pass, you will take many filters and convolve them on the input. Each 'convolution' gives you a 2D matrix output. You will then stack these outputs to get a 3D volume:
<center>
<video width="620" height="440" src="images/conv_kiank.mp4" type="video/mp4" controls>
</video>
</center>
**Exercise**: Implement the function below to convolve the filters W on an input activation A_prev. This function takes as input A_prev, the activations output by the previous layer (for a batch of m inputs), F filters/weights denoted by W, and a bias vector denoted by b, where each filter has its own (single) bias. Finally you also have access to the hyperparameters dictionary which contains the stride and the padding.
**Hint**:
1. To select a 2x2 slice at the upper left corner of a matrix "a_prev" (shape (5,5,3)), you would do:
```python
a_slice_prev = a_prev[0:2,0:2,:]
```
This will be useful when you will define `a_slice_prev` below, using the `start/end` indexes you will define.
2. To define a_slice you will need to first define its corners `vert_start`, `vert_end`, `horiz_start` and `horiz_end`. This figure may be helpful for you to find how each of the corner can be defined using h, w, f and s in the code below.
<img src="images/vert_horiz_kiank.png" style="width:400px;height:300px;">
<caption><center> <u> <font color='purple'> **Figure 3** </u><font color='purple'> : **Definition of a slice using vertical and horizontal start/end (with a 2x2 filter)** <br> This figure shows only a single channel. </center></caption>
**Reminder**:
The formulas relating the output shape of the convolution to the input shape is:
$$ n_H = \lfloor \frac{n_{H_{prev}} - f + 2 \times pad}{stride} \rfloor +1 $$
$$ n_W = \lfloor \frac{n_{W_{prev}} - f + 2 \times pad}{stride} \rfloor +1 $$
$$ n_C = \text{number of filters used in the convolution}$$
For this exercise, we won't worry about vectorization, and will just implement everything with for-loops.
```
# GRADED FUNCTION: conv_forward
def conv_forward(A_prev, W, b, hparameters):
"""
Implements the forward propagation for a convolution function
Arguments:
A_prev -- output activations of the previous layer, numpy array of shape (m, n_H_prev, n_W_prev, n_C_prev)
W -- Weights, numpy array of shape (f, f, n_C_prev, n_C)
b -- Biases, numpy array of shape (1, 1, 1, n_C)
hparameters -- python dictionary containing "stride" and "pad"
Returns:
Z -- conv output, numpy array of shape (m, n_H, n_W, n_C)
cache -- cache of values needed for the conv_backward() function
"""
### START CODE HERE ###
# Retrieve dimensions from A_prev's shape (≈1 line)
(m, n_H_prev, n_W_prev, n_C_prev) = A_prev.shape
# Retrieve dimensions from W's shape (≈1 line)
(f, f, n_C_prev, n_C) = W.shape
# Retrieve information from "hparameters" (≈2 lines)
stride = hparameters["stride"]
pad = hparameters["pad"]
# Compute the dimensions of the CONV output volume using the formula given above. Hint: use int() to floor. (≈2 lines)
n_H = int((n_H_prev - f + 2*pad)/stride) + 1
n_W = int((n_H_prev - f + 2*pad)/stride) + 1
# Initialize the output volume Z with zeros. (≈1 line)
Z = np.zeros((m, n_H, n_W, n_C))
# Create A_prev_pad by padding A_prev
A_prev_pad = zero_pad(A_prev, pad)
for i in range(m): # loop over the batch of training examples
a_prev_pad = A_prev_pad[i] # Select ith training example's padded activation
for h in range(n_H): # loop over vertical axis of the output volume
for w in range(n_W): # loop over horizontal axis of the output volume
for c in range(n_C): # loop over channels (= #filters) of the output volume
# Find the corners of the current "slice" (≈4 lines)
vert_start = h*stride
vert_end = vert_start + f
horiz_start = w * stride
horiz_end = horiz_start + f
# Use the corners to define the (3D) slice of a_prev_pad (See Hint above the cell). (≈1 line)
a_slice_prev = a_prev_pad[vert_start:vert_end, horiz_start:horiz_end, :]
# Convolve the (3D) slice with the correct filter W and bias b, to get back one output neuron. (≈1 line)
Z[i, h, w, c] = conv_single_step(a_slice_prev, W[...,c], b[...,c])
### END CODE HERE ###
# Making sure your output shape is correct
assert(Z.shape == (m, n_H, n_W, n_C))
# Save information in "cache" for the backprop
cache = (A_prev, W, b, hparameters)
return Z, cache
np.random.seed(1)
A_prev = np.random.randn(10,4,4,3)
W = np.random.randn(2,2,3,8)
b = np.random.randn(1,1,1,8)
hparameters = {"pad" : 2,
"stride": 2}
Z, cache_conv = conv_forward(A_prev, W, b, hparameters)
print("Z's mean =", np.mean(Z))
print("Z[3,2,1] =", Z[3,2,1])
print("cache_conv[0][1][2][3] =", cache_conv[0][1][2][3])
```
**Expected Output**:
<table>
<tr>
<td>
**Z's mean**
</td>
<td>
0.0489952035289
</td>
</tr>
<tr>
<td>
**Z[3,2,1]**
</td>
<td>
[-0.61490741 -6.7439236 -2.55153897 1.75698377 3.56208902 0.53036437
5.18531798 8.75898442]
</td>
</tr>
<tr>
<td>
**cache_conv[0][1][2][3]**
</td>
<td>
[-0.20075807 0.18656139 0.41005165]
</td>
</tr>
</table>
Finally, CONV layer should also contain an activation, in which case we would add the following line of code:
```python
# Convolve the window to get back one output neuron
Z[i, h, w, c] = ...
# Apply activation
A[i, h, w, c] = activation(Z[i, h, w, c])
```
You don't need to do it here.
## 4 - Pooling layer
The pooling (POOL) layer reduces the height and width of the input. It helps reduce computation, as well as helps make feature detectors more invariant to its position in the input. The two types of pooling layers are:
- Max-pooling layer: slides an ($f, f$) window over the input and stores the max value of the window in the output.
- Average-pooling layer: slides an ($f, f$) window over the input and stores the average value of the window in the output.
<table>
<td>
<img src="images/max_pool1.png" style="width:500px;height:300px;">
<td>
<td>
<img src="images/a_pool.png" style="width:500px;height:300px;">
<td>
</table>
These pooling layers have no parameters for backpropagation to train. However, they have hyperparameters such as the window size $f$. This specifies the height and width of the fxf window you would compute a max or average over.
### 4.1 - Forward Pooling
Now, you are going to implement MAX-POOL and AVG-POOL, in the same function.
**Exercise**: Implement the forward pass of the pooling layer. Follow the hints in the comments below.
**Reminder**:
As there's no padding, the formulas binding the output shape of the pooling to the input shape is:
$$ n_H = \lfloor \frac{n_{H_{prev}} - f}{stride} \rfloor +1 $$
$$ n_W = \lfloor \frac{n_{W_{prev}} - f}{stride} \rfloor +1 $$
$$ n_C = n_{C_{prev}}$$
```
# GRADED FUNCTION: pool_forward
def pool_forward(A_prev, hparameters, mode = "max"):
"""
Implements the forward pass of the pooling layer
Arguments:
A_prev -- Input data, numpy array of shape (m, n_H_prev, n_W_prev, n_C_prev)
hparameters -- python dictionary containing "f" and "stride"
mode -- the pooling mode you would like to use, defined as a string ("max" or "average")
Returns:
A -- output of the pool layer, a numpy array of shape (m, n_H, n_W, n_C)
cache -- cache used in the backward pass of the pooling layer, contains the input and hparameters
"""
# Retrieve dimensions from the input shape
(m, n_H_prev, n_W_prev, n_C_prev) = A_prev.shape
# Retrieve hyperparameters from "hparameters"
f = hparameters["f"]
stride = hparameters["stride"]
# Define the dimensions of the output
n_H = int(1 + (n_H_prev - f) / stride)
n_W = int(1 + (n_W_prev - f) / stride)
n_C = n_C_prev
# Initialize output matrix A
A = np.zeros((m, n_H, n_W, n_C))
### START CODE HERE ###
for i in range(m): # loop over the training examples
for h in range(n_H): # loop on the vertical axis of the output volume
for w in range(n_W): # loop on the horizontal axis of the output volume
for c in range (n_C): # loop over the channels of the output volume
# Find the corners of the current "slice" (≈4 lines)
vert_start = h*stride
vert_end = vert_start + f
horiz_start = w * stride
horiz_end = horiz_start + f
# Use the corners to define the current slice on the ith training example of A_prev, channel c. (≈1 line)
a_prev_slice = A_prev[i, vert_start:vert_end, horiz_start:horiz_end, c]
# Compute the pooling operation on the slice. Use an if statment to differentiate the modes. Use np.max/np.mean.
if mode == "max":
A[i, h, w, c] = np.max(a_prev_slice)
elif mode == "average":
A[i, h, w, c] = np.mean(a_prev_slice)
### END CODE HERE ###
# Store the input and hparameters in "cache" for pool_backward()
cache = (A_prev, hparameters)
# Making sure your output shape is correct
assert(A.shape == (m, n_H, n_W, n_C))
return A, cache
np.random.seed(1)
A_prev = np.random.randn(2, 4, 4, 3)
hparameters = {"stride" : 2, "f": 3}
A, cache = pool_forward(A_prev, hparameters)
print("mode = max")
print("A =", A)
print()
A, cache = pool_forward(A_prev, hparameters, mode = "average")
print("mode = average")
print("A =", A)
```
**Expected Output:**
<table>
<tr>
<td>
A =
</td>
<td>
[[[[ 1.74481176 0.86540763 1.13376944]]]
[[[ 1.13162939 1.51981682 2.18557541]]]]
</td>
</tr>
<tr>
<td>
A =
</td>
<td>
[[[[ 0.02105773 -0.20328806 -0.40389855]]]
[[[-0.22154621 0.51716526 0.48155844]]]]
</td>
</tr>
</table>
Congratulations! You have now implemented the forward passes of all the layers of a convolutional network.
The remainer of this notebook is optional, and will not be graded.
## 5 - Backpropagation in convolutional neural networks (OPTIONAL / UNGRADED)
In modern deep learning frameworks, you only have to implement the forward pass, and the framework takes care of the backward pass, so most deep learning engineers don't need to bother with the details of the backward pass. The backward pass for convolutional networks is complicated. If you wish however, you can work through this optional portion of the notebook to get a sense of what backprop in a convolutional network looks like.
When in an earlier course you implemented a simple (fully connected) neural network, you used backpropagation to compute the derivatives with respect to the cost to update the parameters. Similarly, in convolutional neural networks you can to calculate the derivatives with respect to the cost in order to update the parameters. The backprop equations are not trivial and we did not derive them in lecture, but we briefly presented them below.
### 5.1 - Convolutional layer backward pass
Let's start by implementing the backward pass for a CONV layer.
#### 5.1.1 - Computing dA:
This is the formula for computing $dA$ with respect to the cost for a certain filter $W_c$ and a given training example:
$$ dA += \sum _{h=0} ^{n_H} \sum_{w=0} ^{n_W} W_c \times dZ_{hw} \tag{1}$$
Where $W_c$ is a filter and $dZ_{hw}$ is a scalar corresponding to the gradient of the cost with respect to the output of the conv layer Z at the hth row and wth column (corresponding to the dot product taken at the ith stride left and jth stride down). Note that at each time, we multiply the the same filter $W_c$ by a different dZ when updating dA. We do so mainly because when computing the forward propagation, each filter is dotted and summed by a different a_slice. Therefore when computing the backprop for dA, we are just adding the gradients of all the a_slices.
In code, inside the appropriate for-loops, this formula translates into:
```python
da_prev_pad[vert_start:vert_end, horiz_start:horiz_end, :] += W[:,:,:,c] * dZ[i, h, w, c]
```
#### 5.1.2 - Computing dW:
This is the formula for computing $dW_c$ ($dW_c$ is the derivative of one filter) with respect to the loss:
$$ dW_c += \sum _{h=0} ^{n_H} \sum_{w=0} ^ {n_W} a_{slice} \times dZ_{hw} \tag{2}$$
Where $a_{slice}$ corresponds to the slice which was used to generate the acitivation $Z_{ij}$. Hence, this ends up giving us the gradient for $W$ with respect to that slice. Since it is the same $W$, we will just add up all such gradients to get $dW$.
In code, inside the appropriate for-loops, this formula translates into:
```python
dW[:,:,:,c] += a_slice * dZ[i, h, w, c]
```
#### 5.1.3 - Computing db:
This is the formula for computing $db$ with respect to the cost for a certain filter $W_c$:
$$ db = \sum_h \sum_w dZ_{hw} \tag{3}$$
As you have previously seen in basic neural networks, db is computed by summing $dZ$. In this case, you are just summing over all the gradients of the conv output (Z) with respect to the cost.
In code, inside the appropriate for-loops, this formula translates into:
```python
db[:,:,:,c] += dZ[i, h, w, c]
```
**Exercise**: Implement the `conv_backward` function below. You should sum over all the training examples, filters, heights, and widths. You should then compute the derivatives using formulas 1, 2 and 3 above.
```
def conv_backward(dZ, cache):
"""
Implement the backward propagation for a convolution function
Arguments:
dZ -- gradient of the cost with respect to the output of the conv layer (Z), numpy array of shape (m, n_H, n_W, n_C)
cache -- cache of values needed for the conv_backward(), output of conv_forward()
Returns:
dA_prev -- gradient of the cost with respect to the input of the conv layer (A_prev),
numpy array of shape (m, n_H_prev, n_W_prev, n_C_prev)
dW -- gradient of the cost with respect to the weights of the conv layer (W)
numpy array of shape (f, f, n_C_prev, n_C)
db -- gradient of the cost with respect to the biases of the conv layer (b)
numpy array of shape (1, 1, 1, n_C)
"""
### START CODE HERE ###
# Retrieve information from "cache"
(A_prev, W, b, hparameters) = None
# Retrieve dimensions from A_prev's shape
(m, n_H_prev, n_W_prev, n_C_prev) = None
# Retrieve dimensions from W's shape
(f, f, n_C_prev, n_C) = None
# Retrieve information from "hparameters"
stride = None
pad = None
# Retrieve dimensions from dZ's shape
(m, n_H, n_W, n_C) = None
# Initialize dA_prev, dW, db with the correct shapes
dA_prev = None
dW = None
db = None
# Pad A_prev and dA_prev
A_prev_pad = None
dA_prev_pad = None
for i in range(None): # loop over the training examples
# select ith training example from A_prev_pad and dA_prev_pad
a_prev_pad = None
da_prev_pad = None
for h in range(None): # loop over vertical axis of the output volume
for w in range(None): # loop over horizontal axis of the output volume
for c in range(None): # loop over the channels of the output volume
# Find the corners of the current "slice"
vert_start = None
vert_end = None
horiz_start = None
horiz_end = None
# Use the corners to define the slice from a_prev_pad
a_slice = None
# Update gradients for the window and the filter's parameters using the code formulas given above
da_prev_pad[vert_start:vert_end, horiz_start:horiz_end, :] += None
dW[:,:,:,c] += None
db[:,:,:,c] += None
# Set the ith training example's dA_prev to the unpaded da_prev_pad (Hint: use X[pad:-pad, pad:-pad, :])
dA_prev[i, :, :, :] = None
### END CODE HERE ###
# Making sure your output shape is correct
assert(dA_prev.shape == (m, n_H_prev, n_W_prev, n_C_prev))
return dA_prev, dW, db
np.random.seed(1)
dA, dW, db = conv_backward(Z, cache_conv)
print("dA_mean =", np.mean(dA))
print("dW_mean =", np.mean(dW))
print("db_mean =", np.mean(db))
```
** Expected Output: **
<table>
<tr>
<td>
**dA_mean**
</td>
<td>
1.45243777754
</td>
</tr>
<tr>
<td>
**dW_mean**
</td>
<td>
1.72699145831
</td>
</tr>
<tr>
<td>
**db_mean**
</td>
<td>
7.83923256462
</td>
</tr>
</table>
## 5.2 Pooling layer - backward pass
Next, let's implement the backward pass for the pooling layer, starting with the MAX-POOL layer. Even though a pooling layer has no parameters for backprop to update, you still need to backpropagation the gradient through the pooling layer in order to compute gradients for layers that came before the pooling layer.
### 5.2.1 Max pooling - backward pass
Before jumping into the backpropagation of the pooling layer, you are going to build a helper function called `create_mask_from_window()` which does the following:
$$ X = \begin{bmatrix}
1 && 3 \\
4 && 2
\end{bmatrix} \quad \rightarrow \quad M =\begin{bmatrix}
0 && 0 \\
1 && 0
\end{bmatrix}\tag{4}$$
As you can see, this function creates a "mask" matrix which keeps track of where the maximum of the matrix is. True (1) indicates the position of the maximum in X, the other entries are False (0). You'll see later that the backward pass for average pooling will be similar to this but using a different mask.
**Exercise**: Implement `create_mask_from_window()`. This function will be helpful for pooling backward.
Hints:
- [np.max()]() may be helpful. It computes the maximum of an array.
- If you have a matrix X and a scalar x: `A = (X == x)` will return a matrix A of the same size as X such that:
```
A[i,j] = True if X[i,j] = x
A[i,j] = False if X[i,j] != x
```
- Here, you don't need to consider cases where there are several maxima in a matrix.
```
def create_mask_from_window(x):
"""
Creates a mask from an input matrix x, to identify the max entry of x.
Arguments:
x -- Array of shape (f, f)
Returns:
mask -- Array of the same shape as window, contains a True at the position corresponding to the max entry of x.
"""
### START CODE HERE ### (≈1 line)
mask = None
### END CODE HERE ###
return mask
np.random.seed(1)
x = np.random.randn(2,3)
mask = create_mask_from_window(x)
print('x = ', x)
print("mask = ", mask)
```
**Expected Output:**
<table>
<tr>
<td>
**x =**
</td>
<td>
[[ 1.62434536 -0.61175641 -0.52817175] <br>
[-1.07296862 0.86540763 -2.3015387 ]]
</td>
</tr>
<tr>
<td>
**mask =**
</td>
<td>
[[ True False False] <br>
[False False False]]
</td>
</tr>
</table>
Why do we keep track of the position of the max? It's because this is the input value that ultimately influenced the output, and therefore the cost. Backprop is computing gradients with respect to the cost, so anything that influences the ultimate cost should have a non-zero gradient. So, backprop will "propagate" the gradient back to this particular input value that had influenced the cost.
### 5.2.2 - Average pooling - backward pass
In max pooling, for each input window, all the "influence" on the output came from a single input value--the max. In average pooling, every element of the input window has equal influence on the output. So to implement backprop, you will now implement a helper function that reflects this.
For example if we did average pooling in the forward pass using a 2x2 filter, then the mask you'll use for the backward pass will look like:
$$ dZ = 1 \quad \rightarrow \quad dZ =\begin{bmatrix}
1/4 && 1/4 \\
1/4 && 1/4
\end{bmatrix}\tag{5}$$
This implies that each position in the $dZ$ matrix contributes equally to output because in the forward pass, we took an average.
**Exercise**: Implement the function below to equally distribute a value dz through a matrix of dimension shape. [Hint](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.ones.html)
```
def distribute_value(dz, shape):
"""
Distributes the input value in the matrix of dimension shape
Arguments:
dz -- input scalar
shape -- the shape (n_H, n_W) of the output matrix for which we want to distribute the value of dz
Returns:
a -- Array of size (n_H, n_W) for which we distributed the value of dz
"""
### START CODE HERE ###
# Retrieve dimensions from shape (≈1 line)
(n_H, n_W) = None
# Compute the value to distribute on the matrix (≈1 line)
average = None
# Create a matrix where every entry is the "average" value (≈1 line)
a = None
### END CODE HERE ###
return a
a = distribute_value(2, (2,2))
print('distributed value =', a)
```
**Expected Output**:
<table>
<tr>
<td>
distributed_value =
</td>
<td>
[[ 0.5 0.5]
<br\>
[ 0.5 0.5]]
</td>
</tr>
</table>
### 5.2.3 Putting it together: Pooling backward
You now have everything you need to compute backward propagation on a pooling layer.
**Exercise**: Implement the `pool_backward` function in both modes (`"max"` and `"average"`). You will once again use 4 for-loops (iterating over training examples, height, width, and channels). You should use an `if/elif` statement to see if the mode is equal to `'max'` or `'average'`. If it is equal to 'average' you should use the `distribute_value()` function you implemented above to create a matrix of the same shape as `a_slice`. Otherwise, the mode is equal to '`max`', and you will create a mask with `create_mask_from_window()` and multiply it by the corresponding value of dZ.
```
def pool_backward(dA, cache, mode = "max"):
"""
Implements the backward pass of the pooling layer
Arguments:
dA -- gradient of cost with respect to the output of the pooling layer, same shape as A
cache -- cache output from the forward pass of the pooling layer, contains the layer's input and hparameters
mode -- the pooling mode you would like to use, defined as a string ("max" or "average")
Returns:
dA_prev -- gradient of cost with respect to the input of the pooling layer, same shape as A_prev
"""
### START CODE HERE ###
# Retrieve information from cache (≈1 line)
(A_prev, hparameters) = None
# Retrieve hyperparameters from "hparameters" (≈2 lines)
stride = None
f = None
# Retrieve dimensions from A_prev's shape and dA's shape (≈2 lines)
m, n_H_prev, n_W_prev, n_C_prev = None
m, n_H, n_W, n_C = None
# Initialize dA_prev with zeros (≈1 line)
dA_prev = None
for i in range(None): # loop over the training examples
# select training example from A_prev (≈1 line)
a_prev = None
for h in range(None): # loop on the vertical axis
for w in range(None): # loop on the horizontal axis
for c in range(None): # loop over the channels (depth)
# Find the corners of the current "slice" (≈4 lines)
vert_start = None
vert_end = None
horiz_start = None
horiz_end = None
# Compute the backward propagation in both modes.
if mode == "max":
# Use the corners and "c" to define the current slice from a_prev (≈1 line)
a_prev_slice = None
# Create the mask from a_prev_slice (≈1 line)
mask = None
# Set dA_prev to be dA_prev + (the mask multiplied by the correct entry of dA) (≈1 line)
dA_prev[i, vert_start: vert_end, horiz_start: horiz_end, c] += None
elif mode == "average":
# Get the value a from dA (≈1 line)
da = None
# Define the shape of the filter as fxf (≈1 line)
shape = None
# Distribute it to get the correct slice of dA_prev. i.e. Add the distributed value of da. (≈1 line)
dA_prev[i, vert_start: vert_end, horiz_start: horiz_end, c] += None
### END CODE ###
# Making sure your output shape is correct
assert(dA_prev.shape == A_prev.shape)
return dA_prev
np.random.seed(1)
A_prev = np.random.randn(5, 5, 3, 2)
hparameters = {"stride" : 1, "f": 2}
A, cache = pool_forward(A_prev, hparameters)
dA = np.random.randn(5, 4, 2, 2)
dA_prev = pool_backward(dA, cache, mode = "max")
print("mode = max")
print('mean of dA = ', np.mean(dA))
print('dA_prev[1,1] = ', dA_prev[1,1])
print()
dA_prev = pool_backward(dA, cache, mode = "average")
print("mode = average")
print('mean of dA = ', np.mean(dA))
print('dA_prev[1,1] = ', dA_prev[1,1])
```
**Expected Output**:
mode = max:
<table>
<tr>
<td>
**mean of dA =**
</td>
<td>
0.145713902729
</td>
</tr>
<tr>
<td>
**dA_prev[1,1] =**
</td>
<td>
[[ 0. 0. ] <br>
[ 5.05844394 -1.68282702] <br>
[ 0. 0. ]]
</td>
</tr>
</table>
mode = average
<table>
<tr>
<td>
**mean of dA =**
</td>
<td>
0.145713902729
</td>
</tr>
<tr>
<td>
**dA_prev[1,1] =**
</td>
<td>
[[ 0.08485462 0.2787552 ] <br>
[ 1.26461098 -0.25749373] <br>
[ 1.17975636 -0.53624893]]
</td>
</tr>
</table>
### Congratulations !
Congratulation on completing this assignment. You now understand how convolutional neural networks work. You have implemented all the building blocks of a neural network. In the next assignment you will implement a ConvNet using TensorFlow.
| github_jupyter |
```
import random
import copy
def generateKromosom(uk_krom):
krom = []
for i in range(uk_krom):
krom.append(random.randint(0,9))
return krom
def generatePopulasi(uk_pop):
pop = []
for i in range(uk_pop):
pop.append(generateKromosom(8))
return pop
def decodeKromosom(krom):
x1_val = -3 + (6 / (9 * (10**-1 + 10**-2 + 10**-3 + 10**-4))) * ((krom[0]*10**-1)+(krom[1]*10**-2)+(krom[2]*10**-3)+(krom[3]*10**-4))
x2_val = -2 + (4 / (9 * (10**-1 + 10**-2 + 10**-3 + 10**-4))) * ((krom[4]*10**-1)+(krom[5]*10**-2)+(krom[6]*10**-3)+(krom[7]*10**-4))
return [x1_val, x2_val]
def hitungFitness(krom):
k = decodeKromosom(krom)
fit_val = -((((4 - (2.1 * k[0]**2) + (k[0]**4 / 3)) * k[0]**2) + (k[0]*k[1]) + ((-4 + (4*k[1]**2))*k[1]**2)))
return fit_val
def hitungFitnessAll(pop, uk_pop):
fit_all = []
for i in range(uk_pop):
fit_all.append(hitungFitness(pop[i]))
return fit_all
def tournamentSelection(pop, uk_tour, uk_pop):
best_krom = []
for i in range(1, uk_tour):
krom = pop[random.randint(0,uk_pop-1)]
if (best_krom == [] or hitungFitness(krom) > hitungFitness(best_krom)):
best_krom = krom
return best_krom
def crossover(par1, par2, pc):
r = random.random()
if (r < pc):
point = random.randint(0,7)
for i in range(point):
par1[i], par2[i] = par2[i], par1[i]
return par1, par2
def mutasi(par1, par2, pm):
r = random.random()
if (r < pm):
par1[random.randint(0,7)] = random.randint(0,9)
par2[random.randint(0,7)] = random.randint(0,9)
return par1, par2
def getElitisme(fit_all):
return fit_all.index(max(fit_all))
```
### Hyperparameter
```
ukpop = 50
uktour = 5
generasi = 100
pc = 0.76
pm = 0.31
populasi = generatePopulasi(ukpop)
for i in range(generasi):
fitness = hitungFitnessAll(populasi, ukpop)
new_populasi = []
best = getElitisme(fitness)
new_populasi.append(populasi[best])
new_populasi.append(populasi[best])
i = 0
while (i < ukpop-2):
parent1 = tournamentSelection(populasi, uktour, ukpop)
parent2 = tournamentSelection(populasi, uktour, ukpop)
while (parent1 == parent2):
parent2 = tournamentSelection(populasi, uktour, ukpop)
par1 = copy.deepcopy(parent1)
par2 = copy.deepcopy(parent2)
child = crossover(par1, par2, pc)
child = mutasi(child[0], child[1], pm)
new_populasi += child
i += 2
populasi = new_populasi
fitness = hitungFitnessAll(populasi, ukpop)
result = getElitisme(fitness)
print(' HASIL MINIMASI FUNGSI')
print()
print('Kromosom terbaik:', populasi[result])
print('Fitness terbaik :', hitungFitness(populasi[result]))
print('Hasil decode :', decodeKromosom(populasi[result]))
# ukpop = 50
# uktour = 5
# generasi = 100
# pc = 0.76
# pm = 0.31
```
| github_jupyter |
<a href="https://colab.research.google.com/github/KordingLab/ENGR344/blob/master/tutorials/W4D1_How_do_we_know_how_certain_we_should_be/TA/W4D1_Tutorial4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Tutorial 4: Model Selection: Bias-variance trade-off
**Module 4: How do we know how certain we should be?**
**Originally By Neuromatch Academy**
**Content creators**: Pierre-Étienne Fiquet, Anqi Wu, Alex Hyafil with help from Byron Galbraith
**Content reviewers**: Lina Teichmann, Madineh Sarvestani, Patrick Mineault, Ella Batty, Michael Waskom
**Content Modifiers**: Konrad Kording, Ilenna Jones
**Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
<p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
```
# @title Due Dates Calendar
from ipywidgets import widgets
from IPython.display import display, IFrame, YouTubeVideo
out1 = widgets.Output()
with out1:
calendar = IFrame(src="https://calendar.google.com/calendar/embed?src=356b9d2nspjttvgbb3tvgk2f58%40group.calendar.google.com&ctz=America%2FNew_York", width=600, height=480)
display(calendar)
out = widgets.Tab([out1])
out.set_title(0, 'Calendar')
display(out)
```
---
# Tutorial Objectives
*Estimated timing of tutorial: 25 minutes*
This is Tutorial 5 of a series on fitting models to data. We start with simple linear regression, using least squares optimization (Tutorial 1) and Maximum Likelihood Estimation (Tutorial 2). We will use bootstrapping to build confidence intervals around the inferred linear model parameters (Tutorial 3). We'll finish our exploration of regression models by generalizing to multiple linear regression and polynomial regression (Tutorial 4). We end by learning how to choose between these various models. We discuss the bias-variance trade-off (Tutorial 5) and Cross Validation for model selection (Tutorial 6).
In this tutorial, we will learn about the bias-variance tradeoff and see it in action using polynomial regression models.
Tutorial objectives:
* Understand difference between test and train data
* Compare train and test error for models of varying complexity
* Understand how bias-variance tradeoff relates to what model we choose
```
# @title Tutorial slides
# @markdown These are the slides for the videos in all tutorials today
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/2mkq4/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
# @title Video 1: Bias Variance Tradeoff
from ipywidgets import widgets
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="O0KEY0xpzMk", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1])
out.set_title(0, 'Youtube')
display(out)
```
---
# Setup
```
# Imports
import numpy as np
import matplotlib.pyplot as plt
#@title Figure Settings
%config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
# @title Plotting Functions
def plot_MSE_poly_fits(mse_train, mse_test, max_order):
"""
Plot the MSE values for various orders of polynomial fits on the same bar
graph
Args:
mse_train (ndarray): an array of MSE values for each order of polynomial fit
over the training data
mse_test (ndarray): an array of MSE values for each order of polynomial fit
over the test data
max_order (scalar): max order of polynomial fit
"""
fig, ax = plt.subplots()
width = .35
ax.bar(np.arange(max_order + 1) - width / 2, mse_train, width, label="train MSE")
ax.bar(np.arange(max_order + 1) + width / 2, mse_test , width, label="test MSE")
ax.legend()
ax.set(xlabel='Polynomial order', ylabel='MSE', title ='Comparing polynomial fits');
# @title Helper functions
def ordinary_least_squares(x, y):
"""Ordinary least squares estimator for linear regression.
Args:
x (ndarray): design matrix of shape (n_samples, n_regressors)
y (ndarray): vector of measurements of shape (n_samples)
Returns:
ndarray: estimated parameter values of shape (n_regressors)
"""
return np.linalg.inv(x.T @ x) @ x.T @ y
def make_design_matrix(x, order):
"""Create the design matrix of inputs for use in polynomial regression
Args:
x (ndarray): input vector of shape (n_samples)
order (scalar): polynomial regression order
Returns:
ndarray: design matrix for polynomial regression of shape (samples, order+1)
"""
# Broadcast to shape (n x 1) so dimensions work
if x.ndim == 1:
x = x[:, None]
#if x has more than one feature, we don't want multiple columns of ones so we assign
# x^0 here
design_matrix = np.ones((x.shape[0],1))
# Loop through rest of degrees and stack columns
for degree in range(1, order+1):
design_matrix = np.hstack((design_matrix, x**degree))
return design_matrix
def solve_poly_reg(x, y, max_order):
"""Fit a polynomial regression model for each order 0 through max_order.
Args:
x (ndarray): input vector of shape (n_samples)
y (ndarray): vector of measurements of shape (n_samples)
max_order (scalar): max order for polynomial fits
Returns:
dict: fitted weights for each polynomial model (dict key is order)
"""
# Create a dictionary with polynomial order as keys, and np array of theta
# (weights) as the values
theta_hats = {}
# Loop over polynomial orders from 0 through max_order
for order in range(max_order+1):
X = make_design_matrix(x, order)
this_theta = ordinary_least_squares(X, y)
theta_hats[order] = this_theta
return theta_hats
```
---
# Section 1: Train vs test data
*Estimated timing to here from start of tutorial: 8 min*
The data used for the fitting procedure for a given model is the **training data**. In tutorial 4, we computed MSE on the training data of our polynomial regression models and compared training MSE across models. An additional important type of data is **test data**. This is held-out data that is not used (in any way) during the fitting procedure. When fitting models, we often want to consider both the train error (the quality of prediction on the training data) and the test error (the quality of prediction on the test data) as we will see in the next section.
We will generate some noisy data for use in this tutorial using a similar process as in Tutorial 4. However, now we will also generate test data. We want to see how our model generalizes beyond the range of values seen in the training phase. To accomplish this, we will generate x from a wider range of values ([-3, 3]). We then plot the train and test data together.
```
# @markdown Execute this cell to simulate both training and test data
### Generate training data
np.random.seed(0)
n_train_samples = 50
x_train = np.random.uniform(-2, 2.5, n_train_samples) # sample from a uniform distribution over [-2, 2.5)
noise = np.random.randn(n_train_samples) # sample from a standard normal distribution
y_train = x_train**2 - x_train - 2 + noise
### Generate testing data
n_test_samples = 20
x_test = np.random.uniform(-3, 3, n_test_samples) # sample from a uniform distribution over [-2, 2.5)
noise = np.random.randn(n_test_samples) # sample from a standard normal distribution
y_test = x_test**2 - x_test - 2 + noise
## Plot both train and test data
fig, ax = plt.subplots()
plt.title('Training & Test Data')
plt.plot(x_train, y_train, '.', markersize=15, label='Training')
plt.plot(x_test, y_test, 'g+', markersize=15, label='Test')
plt.legend()
plt.xlabel('x')
plt.ylabel('y');
```
---
# Section 2: Bias-variance tradeoff
*Estimated timing to here from start of tutorial: 10 min*
<details>
<summary> <font color='blue'>Click here for text recap of video </font></summary>
Finding a good model can be difficult. One of the most important concepts to keep in mind when modeling is the **bias-variance tradeoff**.
**Bias** is the difference between the prediction of the model and the corresponding true output variables you are trying to predict. Models with high bias will not fit the training data well since the predictions are quite different from the true data. These high bias models are overly simplified - they do not have enough parameters and complexity to accurately capture the patterns in the data and are thus **underfitting**.
**Variance** refers to the variability of model predictions for a given input. Essentially, do the model predictions change a lot with changes in the exact training data used? Models with high variance are highly dependent on the exact training data used - they will not generalize well to test data. These high variance models are **overfitting** to the data.
In essence:
* High bias, low variance models have high train and test error.
* Low bias, high variance models have low train error, high test error
* Low bias, low variance models have low train and test error
As we can see from this list, we ideally want low bias and low variance models! These goals can be in conflict though - models with enough complexity to have low bias also tend to overfit and depend on the training data more. We need to decide on the correct tradeoff.
In this section, we will see the bias-variance tradeoff in action with polynomial regression models of different orders.
</details>
Graphical illustration of bias and variance.
(Source: http://scott.fortmann-roe.com/docs/BiasVariance.html)
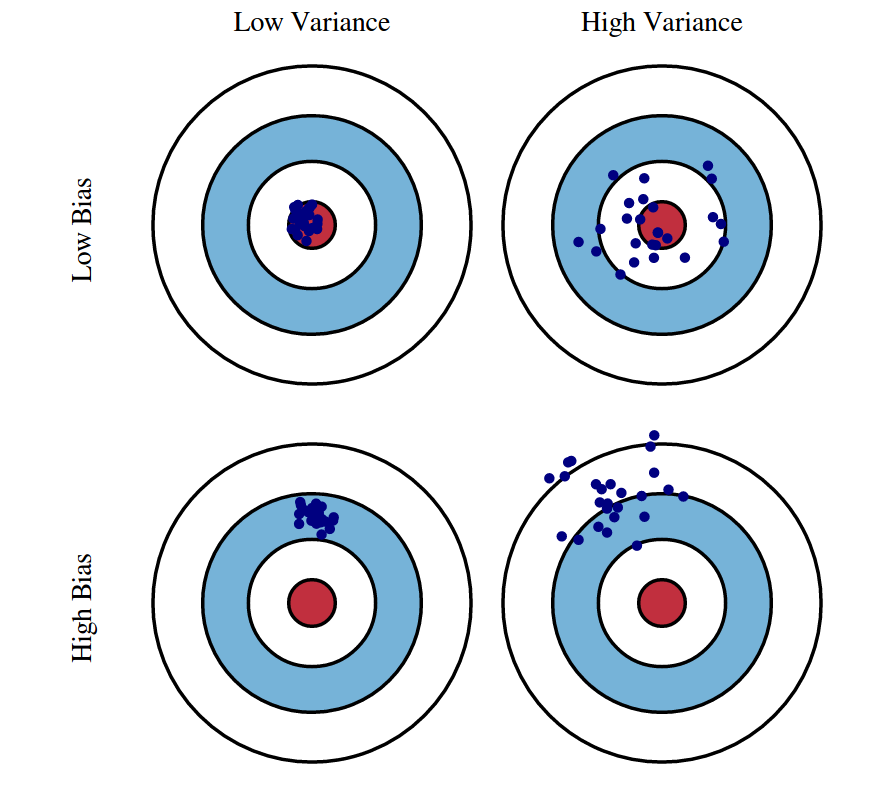
We will first fit polynomial regression models of orders 0-5 on our simulated training data just as we did in Tutorial 4.
```
# @markdown Execute this cell to estimate theta_hats
max_order = 5
theta_hats = solve_poly_reg(x_train, y_train, max_order)
```
## Coding Exercise 2: Compute and compare train vs test error
We will use MSE as our error metric again. Compute MSE on training data ($x_{train},y_{train}$) and test data ($x_{test}, y_{test}$) for each polynomial regression model (orders 0-5). Since you already developed code in T4 Exercise 4 for making design matrices and evaluating fit polynomials, we have ported that here into the functions `make_design_matrix` and `evaluate_poly_reg` for your use.
*Please think about after completing exercise before reading the following text! Do you think the order 0 model has high or low bias? High or low variance? How about the order 5 model?*
```
# @markdown Execute this cell for function `evalute_poly_reg`
def evaluate_poly_reg(x, y, theta_hats, max_order):
""" Evaluates MSE of polynomial regression models on data
Args:
x (ndarray): input vector of shape (n_samples)
y (ndarray): vector of measurements of shape (n_samples)
theta_hats (dict): fitted weights for each polynomial model (dict key is order)
max_order (scalar): max order of polynomial fit
Returns
(ndarray): mean squared error for each order, shape (max_order)
"""
mse = np.zeros((max_order + 1))
for order in range(0, max_order + 1):
X_design = make_design_matrix(x, order)
y_hat = np.dot(X_design, theta_hats[order])
residuals = y - y_hat
mse[order] = np.mean(residuals ** 2)
return mse
def compute_mse(x_train, x_test, y_train, y_test, theta_hats, max_order):
"""Compute MSE on training data and test data.
Args:
x_train(ndarray): training data input vector of shape (n_samples)
x_test(ndarray): test data input vector of shape (n_samples)
y_train(ndarray): training vector of measurements of shape (n_samples)
y_test(ndarray): test vector of measurements of shape (n_samples)
theta_hats(dict): fitted weights for each polynomial model (dict key is order)
max_order (scalar): max order of polynomial fit
Returns:
ndarray, ndarray: MSE error on training data and test data for each order
"""
#######################################################
## TODO for students: calculate mse error for both sets
## Hint: look back at tutorial 5 where we calculated MSE
# Fill out function and remove
raise NotImplementedError("Student excercise: calculate mse for train and test set")
#######################################################
mse_train = ...
mse_test = ...
return mse_train, mse_test
# Compute train and test MSE
mse_train, mse_test = compute_mse(x_train, x_test, y_train, y_test, theta_hats, max_order)
# Visualize
plot_MSE_poly_fits(mse_train, mse_test, max_order)
# to_remove solution
def compute_mse(x_train, x_test, y_train, y_test, theta_hats, max_order):
"""Compute MSE on training data and test data.
Args:
x_train(ndarray): training data input vector of shape (n_samples)
x_test(ndarray): test vector of shape (n_samples)
y_train(ndarray): training vector of measurements of shape (n_samples)
y_test(ndarray): test vector of measurements of shape (n_samples)
theta_hats(dict): fitted weights for each polynomial model (dict key is order)
max_order (scalar): max order of polynomial fit
Returns:
ndarray, ndarray: MSE error on training data and test data for each order
"""
mse_train = evaluate_poly_reg(x_train, y_train, theta_hats, max_order)
mse_test = evaluate_poly_reg(x_test, y_test, theta_hats, max_order)
return mse_train, mse_test
# Compute train and test MSE
mse_train, mse_test = compute_mse(x_train, x_test, y_train, y_test, theta_hats, max_order)
# Visualize
with plt.xkcd():
plot_MSE_poly_fits(mse_train, mse_test, max_order)
```
As we can see from the plot above, more complex models (higher order polynomials) have lower MSE for training data. The overly simplified models (orders 0 and 1) have high MSE on the training data. As we add complexity to the model, we go from high bias to low bias.
The MSE on test data follows a different pattern. The best test MSE is for an order 2 model - this makes sense as the data was generated with an order 2 model. Both simpler models and more complex models have higher test MSE.
So to recap:
Order 0 model: High bias, low variance
Order 5 model: Low bias, high variance
Order 2 model: Just right, low bias, low variance
---
# Summary
*Estimated timing of tutorial: 25 minutes*
- Training data is the data used for fitting, test data is held-out data.
- We need to strike the right balance between bias and variance. Ideally we want to find a model with optimal model complexity that has both low bias and low variance
- Too complex models have low bias and high variance.
- Too simple models have high bias and low variance.
**Note**
- Bias and variance are very important concepts in modern machine learning, but it has recently been observed that they do not necessarily trade off (see for example the phenomenon and theory of "double descent")
**Further readings:**
- [The elements of statistical learning](https://web.stanford.edu/~hastie/ElemStatLearn/) by Hastie, Tibshirani and Friedman
---
# Bonus
## Bonus Exercise
Prove the bias-variance decomposition for MSE
\begin{align}
\mathbb{E}_{x}\left[\left(y-\hat{y}(x ; \theta)\right)^{2}\right]=\left(\operatorname{Bias}_{x}[\hat{y}(x ; \theta)]\right)^{2}+\operatorname{Var}_{x}[\hat{y}(x ; \theta)]+\sigma^{2}
\end{align}
where
\begin{align}
\operatorname{Bias}_{x}[\hat{y}(x ; \theta)]=\mathbb{E}_{x}[\hat{y}(x ; \theta)]-y
\end{align}
and
\begin{align}
\operatorname{Var}_{x}[\hat{y}(x ; \theta)]=\mathbb{E}_{x}\left[\hat{y}(x ; \theta)^{2}\right]-\mathrm{E}_{x}[\hat{y}(x ; \theta)]^{2}
\end{align}
Hint: use
\begin{align}
\operatorname{Var}[X]=\mathbb{E}\left[X^{2}\right]-(\mathrm{E}[X])^{2}
\end{align}
| github_jupyter |
```
!pip install pyyaml==5.1
# This is the current pytorch version on Colab. Uncomment this if Colab changes its pytorch version
# !pip install torch==1.9.0+cu102 torchvision==0.10.0+cu102 -f https://download.pytorch.org/whl/torch_stable.html
# Install detectron2 that matches the above pytorch version
# See https://detectron2.readthedocs.io/tutorials/install.html for instructions
!pip install detectron2 -f https://dl.fbaipublicfiles.com/detectron2/wheels/cu102/torch1.9/index.html
# exit(0) # After installation, you need to "restart runtime" in Colab. This line can also restart runtime
# check pytorch installation:
import torch, torchvision
print(torch.__version__, torch.cuda.is_available())
assert torch.__version__.startswith("1.9") # please manually install torch 1.9 if Colab changes its default version
torchvision.__version__
# Some basic setup:
# Setup detectron2 logger
import detectron2
from detectron2.utils.logger import setup_logger
setup_logger()
# import some common libraries
import numpy as np
import os, json, cv2, random
from google.colab.patches import cv2_imshow
# import some common detectron2 utilities
from detectron2 import model_zoo
from detectron2.engine import DefaultPredictor
from detectron2.config import get_cfg
from detectron2.utils.visualizer import Visualizer
from detectron2.data import MetadataCatalog, DatasetCatalog
!pip install labelme2coco
!ls
import labelme2coco
folder = "./images"
save_json = "data.json"
labelme2coco.convert("./images/", "./images/data.json")
!git clone https://github.com/hpanwar08/detectron2
!wget https://www.dropbox.com/sh/1098ym6vhad4zi6/AAD8Y-SVN6EbfAWEDYuZHG8xa/model_final_trimmed.pth
%cd "/content/detectron2"
!python "/content/detectron2/tools/visualize_json_results.py" --input "aegon.json" --output dir/ --dataset "aegon"
from detectron2.data import MetadataCatalog
MetadataCatalog.get("dla_train").thing_classes = ['text', 'title', 'list', 'table', 'figure']
from detectron2.data.datasets import register_coco_instances
register_coco_instances("dla_train", {}, "/content/detectron2/aegon.json", "/content/detectron2/aegon")
!ls
from detectron2.engine import DefaultTrainer
cfg = get_cfg()
cfg.merge_from_file(model_zoo.get_config_file("COCO-InstanceSegmentation/mask_rcnn_R_101_FPN_3x.yaml"))
cfg.DATASETS.TRAIN = ("dla_train",)
cfg.DATASETS.TEST = ()
cfg.DATALOADER.NUM_WORKERS = 2
cfg.MODEL.WEIGHTS = "model_final_trimmed.pth" # Let training initialize from model zoo
#cfg.MODEL.WEIGHTS = model_zoo.get_checkpoint_url("detectron2/configs/DLA_mask_rcnn_R_101_FPN_3x.yaml") # Let training initialize from model zoo
cfg.SOLVER.IMS_PER_BATCH = 2
cfg.SOLVER.BASE_LR = 0.00025 # pick a good LR
cfg.SOLVER.MAX_ITER = 300 # 300 iterations seems good enough for this toy dataset; you will need to train longer for a practical dataset
cfg.SOLVER.STEPS = [] # do not decay learning rate
cfg.MODEL.ROI_HEADS.BATCH_SIZE_PER_IMAGE = 128 # faster, and good enough for this toy dataset (default: 512)
cfg.MODEL.ROI_HEADS.NUM_CLASSES = 1 # only has one class (ballon). (see https://detectron2.readthedocs.io/tutorials/datasets.html#update-the-config-for-new-datasets)
# NOTE: this config means the number of classes, but a few popular unofficial tutorials incorrect uses num_classes+1 here.
os.makedirs(cfg.OUTPUT_DIR, exist_ok=True)
trainer = DefaultTrainer(cfg)
trainer.resume_or_load(resume=False)
trainer.train()
!python "tools/train_net_dla.py"
!ls
import json
with open("/content/detectron2/aegon.json") as filejson:
data = json.load(filejson)
for idx, seg in enumerate(data["annotations"]):
data["annotations"][idx]['segmentation'] = []
with open("/content/detectron2/aegon.json") as filejson:
data = json.load(filejson)
jsonString = json.dumps(data)
jsonFile = open(cfg, "w")
jsonFile.write(jsonString)
jsonFile.close()
!python "detectron2/demo/demo.py" --config-file "detectron2/configs/DLA_mask_rcnn_X_101_32x8d_FPN_3x.yaml" --input "detectron2/2.png" --output "detectron2/output" --confidence-threshold 0.5 --opts MODEL.WEIGHTS "detectron2/model_final.pth"
!python "demo/demo.py" --config-file "configs/DLA_mask_rcnn_X_101_32x8d_FPN_3x.yaml" --input "210118-45-Aviva-Non-Indexation-2IncomeProtection-2Applicants-Deferred-8-2.png" --output "output" --confidence-threshold 0.5 --opts MODEL.WEIGHTS "output/20210825T0631/model_final.pth" MODEL.DEVICE "cpu"
%cd detectron2
!pwd
!ls
cp /content/model_final.pth /content/detectron2/model_final.pth
!zip -r /content/detectron2.zip /content/detectron2
```
| github_jupyter |
<img src="https://www.epfl.ch/about/overview/wp-content/uploads/2020/07/logo-epfl-1024x576.png" style="padding-right:10px;width:140px;float:left"></td>
<h2 style="white-space: nowrap">Image Processing Laboratory Notebooks</h2>
<hr style="clear:both">
<p style="font-size:0.85em; margin:2px; text-align:justify">
This Juypter notebook is part of a series of computer laboratories which are designed
to teach image-processing programming; they are running on the EPFL's Noto server. They are the practical complement of the theoretical lectures of the EPFL's Master course <b>Image Processing II</b>
(<a href="https://moodle.epfl.ch/course/view.php?id=463">MICRO-512</a>) taught by Dr. D. Sage, Dr. M. Liebling, Prof. M. Unser and Prof. D. Van de Ville.
</p>
<p style="font-size:0.85em; margin:2px; text-align:justify">
The project is funded by the Center for Digital Education and the School of Engineering. It is owned by the <a href="http://bigwww.epfl.ch/">Biomedical Imaging Group</a>.
The distribution or the reproduction of the notebook is strictly prohibited without the written consent of the authors. © EPFL 2022.
</p>
<p style="font-size:0.85em; margin:0px"><b>Authors</b>:
<a href="mailto:pol.delaguilapla@epfl.ch">Pol del Aguila Pla</a>,
<a href="mailto:kay.lachler@epfl.ch">Kay Lächler</a>,
<a href="mailto:alejandro.nogueronaramburu@epfl.ch">Alejandro Noguerón Arámburu</a>,
<a href="mailto:daniel.sage@epfl.ch">Daniel Sage</a>, and
<a href="mailto:kamil.seghrouchni@epfl.ch">Kamil Seghrouchni</a>.
</p>
<hr style="clear:both">
<h1>Lab 4.1: Orientation warm-up</h1>
<div style="background-color:#F0F0F0;padding:4px">
<p style="margin:4px;"><b>Released</b>: Thursday March 3, 2022</p>
<p style="margin:4px;"><b>Submission</b>: <span style="color:red">Friday March 11, 2022</span> (before 11:59PM) on <a href="https://moodle.epfl.ch/course/view.php?id=463">Moodle</a></p>
<p style="margin:4px;"><b>Grade weigth</b>: Lab 4 (18 points), 7.5 % of the overall grade</p>
<p style="margin:4px;"><b>Remote help</b>: Monday 7 March, 2022 on Zoom (12h-13h, see Moodle for link) and Thursday 10 March on campus</p>
<p style="margin:4px;"><b>Related lectures</b>: Chapter 6</p>
</div>
### Student Name: Guanqun LIU
### SCIPER: 334988
Double-click on this cell and fill your name and SCIPER number. Then, run the cell below to verify your identity in Noto and set the seed for random results.
```
import getpass
# This line recovers your camipro number to mark the images with your ID
uid = int(getpass.getuser().split('-')[2]) if len(getpass.getuser().split('-')) > 2 else ord(getpass.getuser()[0])
print(f'SCIPER: {uid}')
```
## <a name="imports_"></a> Imports
In the next cell we import Python libraries we will use throughout the lab, as well as the `ImageViewer` class, created specifically for this course, which provides interactive image visualization based on the `ipywidgets` library. We will import:
* [`matplotlib.pyplot`](https://matplotlib.org/3.2.2/api/_as_gen/matplotlib.pyplot.html), to display images,
* [`ipywidgets`](https://ipywidgets.readthedocs.io/en/latest/), to make the image display interactive,
* [`numpy`](https://numpy.org/doc/stable/reference/index.html), for mathematical operations on arrays,
* [`cv2`](https://docs.opencv.org/2.4/index.html), for image processing tasks.
We will then load the `ImageViewer` class (see the documentation [here](https://github.com/Biomedical-Imaging-Group/interactive-kit/wiki/Image-Viewer) or run the Python command `help(viewer)` after loading the class).
Finally, we load the images you will use in the exercise to test your functions.
```
# Configure plotting as dynamic
%matplotlib widget
# Import standard required packages for this exercise
import warnings
import matplotlib.pyplot as plt
import ipywidgets as widgets
import numpy as np
import cv2 as cv
import skimage
from skimage import feature
from interactive_kit import imviewer as viewer
# Load images to be used in this exercise
corner = cv.imread('images/corner.tif', cv.IMREAD_UNCHANGED)
dendrochronology = cv.imread('images/dendrochronolgy.tif', cv.IMREAD_UNCHANGED)
fingerprint = cv.imread('images/fingerprint.tif', cv.IMREAD_UNCHANGED)
harris_corner = cv.imread('images/harris-corner.tif', cv.IMREAD_UNCHANGED)
wave_ramp = cv.imread('images/wave-ramp.tif', cv.IMREAD_UNCHANGED)
```
# Orientation laboratory (13 points)
In this lab we will implement the computation of the structure tensor presented in Chapter 6.2, which can be used to perform directional image analysis.
The block-diagram of the complete system is shown in the following flowchart, where $f(x,y)$ is the graylevel input image.
<img src="images/block-diagram.png" alt="Drawing" style="width: 800px;"/>
Successively, you will implement the functions
* `structure_tensor` to generate the structure tensor matrix,
* `orientation_features`, which implements the whole chain of calculations to generate the features needed for directional analysis, and
* `colorize_features` to display the calculated features as a color image.
Once you ensured that the functions are correct, you will use them in two applications that rely on directional image analysis,
* a method to select specific orientations, and
* a keypoint detector (Harris corner detector).
Finally, we will analyze the `structure_tensor` function a bit more closely and see if we can improve the obtained results.
<div class = 'alert alert-info'>
<b>Note:</b> This part of the lab will be carried out completely in Python.
</div>
## <a id="ToC_2_Orientation"></a>Table of contents
1. [Structure tensor matrix](#1.-Structure-tensor-matrix-(2-points)) **(2 points)**
2. [Orientation features](#2.-Orientation-features)
1. [Feature calculation](#2.A.-Feature-calculation-(4-points)) **(4 points)**
2. [Feature visualization](#2.B.-Feature-visualization-(2-points)) **(2 points)**
3. [Application](#3.-Application)
1. [Selection of specific orientations](#3.A.-Selection-of-specific-orientations-(2-points)) **(2 points)**
2. [Harris corner detector](#3.B.-Harris-corner-detector-(2-points)) **(2 points)**
3. [*Advanced:* Isotropic filtering in the Fourier space](#3.C.-Advanced:-Isotropic-filtering-in-the-Fourier-space-(1-point)) **(1 point)**
### Visualize images
First of all, run the cell below to get familiar with the images we will be using. Remember you can use `Next` and `Prev` to cycle through the images.
```
# Declare image_list for ImageViewer
img_list = [corner, dendrochronology, fingerprint, harris_corner, wave_ramp]
imgs_viewer = viewer(img_list, widgets=True)
```
# 1. Structure tensor matrix (2 points)
[Back to table of contents](#ToC_2_Orientation)
To calculate the elements $J_{xx}$, $J_{xy}$ and $J_{yy}$ of the structure tensor, we only need two computational blocks: a Gaussian filter and a gradient filter. For the gradient, we will use the **OpenCV Sobel filter** [`cv.Sobel(src, ddepth, dx, dy, ksize)`](https://docs.opencv.org/3.4/d4/d86/group__imgproc__filter.html#gacea54f142e81b6758cb6f375ce782c8d). Click on the link to see its documentation and choose the right input parameters to apply a **first-order** Sobel filter of **size $3\times3$**.
For the smoothing filter we will use the standard Gaussian filter provided by OpenCV [`cv.GaussianBlur(src, ksize, sigmaX)`](https://docs.opencv.org/master/d4/d86/group__imgproc__filter.html#gaabe8c836e97159a9193fb0b11ac52cf1) with default boundary conditions (`BORDER_DEFAULT` or `BORDER_REFLECT_101`). If you are unsure about its input parameters, go check the documentation.
**For 2 points**, complete the function `structure_tensor` in the cell below. It may be useful to revisit the [figure](#Orientation-laboratory-(13-points)) at the start of this notebook before starting.
<div class="alert alert-info">
<b>Hints:</b>
<ul><li>Take a moment to go thoroughly through the documentation! It will make your life much easier.
</li><li><b><code>cv.Sobel</code>:</b> set <code>ddepth=cv.CV_64F</code> since the gradient can be a negative floating point number.
</li><li><b><code>cv.GaussianBlur</code>:</b> set <code>ksize=(0,0)</code> to let the function choose the filter size automatically from <code>sigmaX</code>.
</li><li><b>Multiplication:</b> Note that, contrary to Matlab and other languages, the Python operator <code>*</code> applied to a NumPy Array performs <i>element-wise</i> multiplication.
</li><li><b>Exponent:</b> Note that Python's exponent operator is <code>**</code>.
</li><li>To ensure that you define the right parameter in a function call always specify the name of the parameter. For example: <code>cv.Sobel(src=img, ddepth=cv.CV_64F, dx=1, dy=1, ksize=5)</code> is much easier to understand and debug than <code>cv.Sobel(img, cv.CV_64F, 1, 1, 5)</code>.</ul>
</div>
```
# Function that calculates the elements Jxx, Jxy and Jyy of the structure tensor matrix
def structure_tensor(img, sigma):
# Initialize output vatiables
Jxx = np.zeros(img.shape)
Jxy = np.zeros(img.shape)
Jyy = np.zeros(img.shape)
# YOUR CODE HERE
sobel_x = cv.Sobel(src=img, ddepth=cv.CV_64F, dx=1, dy=0, ksize=3)
sobel_y = cv.Sobel(src=img, ddepth=cv.CV_64F, dx=0, dy=1, ksize=3)
grad_x = sobel_x ** 2
grad_y = sobel_y ** 2
grad_xy = sobel_x * sobel_y
Jxx = cv.GaussianBlur(src=grad_x, ksize=(0,0), sigmaX=sigma, borderType=cv.BORDER_DEFAULT)
Jxy = cv.GaussianBlur(src=grad_xy, ksize=(0,0), sigmaX=sigma, borderType=cv.BORDER_DEFAULT)
Jyy = cv.GaussianBlur(src=grad_y, ksize=(0,0), sigmaX=sigma, borderType=cv.BORDER_DEFAULT)
return Jxx, Jxy, Jyy
```
Let's perform a quick sanity check on a simple $11 \times 11$ **impulse image** using `sigma=1`.
<div class="alert alert-info">
**Note:** You can modify the input image and the sigma value in the cell below to observe the different results.
</div>
```
# Define impulse image
size = 11
test_img = np.zeros((size,size))
test_img[size//2, size//2] = 1
# Run function and display the result
Jxx, Jxy, Jyy = structure_tensor(test_img, sigma = 1)
plt.close('all')
view = viewer([test_img, Jxx, Jyy, Jxy], subplots=(2,2))
```
Specifically, we will first check that $J_{xx}$ and $J_{yy}$ are non-negative and identical to each other when rotated by $90^{\circ}$, which should be the case for this impulse image. Then, we will also check that $J_{xy}$ contains both negative and non-negative numbers, and that all elements that are either in the fifth row or the fifth column of $J_{xy}$ are zero.
Because the structure tensor is a crucial part of this lab, we will also perform more sophisticated sanity checks comparing your results to our pre-computed correct results.
```
# Basic sanity checks
# Re-run example in case you've played around with the previous cell
test_img = np.zeros((11,11)); test_img[5, 5] = 1
Jxx, Jxy, Jyy = structure_tensor(test_img, sigma = 1)
# No negative values in Jxx and Jyy
if not (np.all(Jxx >= 0) and np.all(Jyy >= 0)):
print('WARNING!\nJxx and Jyy should not contain any negative values.\n')
if not np.allclose(np.rot90(Jxx), Jyy):
print('WARNING!\nJxx should be the same as Jyy but rotated 90 degrees.\n')
# Jxy both positive and negative
if not (np.any(Jxy > 0) and np.any(Jxy < 0)):
print('WARNING!\nJxy should contain both negative and positive values.\n')
# Fifth row/col zeros
if not (np.all(abs(Jxy[5, :]) < 1e-5) and np.all(abs(Jxy[:, 5]) < 1e-5)):
print('WARNING!\nThe fifth row/column of Jxy should only have zeros.\n')
# Comparison to pre-computed correct results
# Boundaries should be close to zero
error_check = [False, False, False]
mask = np.ones(test_img.shape, dtype=bool); mask[2:9, 2:9] = False
if not np.all(np.abs(Jxx[mask]) < 0.01):
print('WARNING!\nJxx is not yet correct, values outside the range x=[2,8], y=[2,8] should be close to 0.\n')
error_check[0] = True
if not np.all(np.abs(Jyy[mask]) < 0.01):
print('WARNING!\nJyy is not yet correct, values outside the range x=[2,8], y=[2,8] should be close to 0.\n')
error_check[1] = True
if not np.all(np.abs(Jxy[mask]) < 0.01):
print('WARNING!\nJxy is not yet correct, values outside the range x=[2,8], y=[2,8] should be close to 0.\n')
error_check[2] = True
# Correct outputs
Jxx_corr = np.array([[0.004, 0.018, 0.033, 0.035, 0.033, 0.018, 0.004],
[0.025, 0.114, 0.209, 0.224, 0.209, 0.114, 0.025],
[0.077, 0.35, 0.644, 0.688, 0.644, 0.35, 0.077],
[0.113, 0.512, 0.942, 1.006, 0.942, 0.512, 0.113],
[0.077, 0.35, 0.644, 0.688, 0.644, 0.35, 0.077],
[0.025, 0.114, 0.209, 0.224, 0.209, 0.114, 0.025],
[0.004, 0.018, 0.033, 0.035, 0.033, 0.018, 0.004]])
Jyy_corr = Jxx_corr.T
Jxy_corr = np.array([[ 0.003, 0.013, 0.019, 0. , -0.019, -0.013, -0.003],
[ 0.013, 0.056, 0.082, 0. , -0.082, -0.056, -0.013],
[ 0.019, 0.082, 0.119, 0. , -0.119, -0.082, -0.019],
[ 0. , 0. , 0. , 0. , 0. , 0. , 0. ],
[-0.019, -0.082, -0.119, 0. , 0.119, 0.082, 0.019],
[-0.013, -0.056, -0.082, 0. , 0.082, 0.056, 0.013],
[-0.003, -0.013, -0.019, 0. , 0.019, 0.013, 0.003]])
# Exact values in the center
if not np.allclose(Jxx[2:9, 2:9], Jxx_corr, atol=1e-3):
print('WARNING!\nThe non-zero values of Jxx inside the range x=[2,8], y=[2,8] are not yet correct.')
error_check[0] = True
if not np.allclose(Jyy[2:9, 2:9], Jyy_corr, atol=1e-3):
print('WARNING!\nThe non-zero values of Jyy inside the range x=[2,8], y=[2,8] are not yet correct.')
error_check[1] = True
if not np.allclose(Jxy[2:9, 2:9], Jxy_corr, atol=1e-3):
print('WARNING!\nThe non-zero values of Jxy inside the range x=[2,8], y=[2,8] are not yet correct.')
error_check[2] = True
# In the presence of errors, show differences to debug your code
if np.any(error_check):
print('\nLook at the output below to compare your solution to the correct one:\n')
Jxx_vis = np.zeros(test_img.shape); Jyy_vis = np.zeros(test_img.shape); Jxy_vis = np.zeros(test_img.shape)
Jxx_vis[mask == False] = Jxx_corr.flatten(); Jyy_vis[mask == False] = Jyy_corr.flatten(); Jxy_vis[mask == False] = Jxy_corr.flatten()
corr_imgs = [Jxx_vis, Jyy_vis, Jxy_vis]; imgs = [Jxx, Jyy, Jxy]; names = ['Jxx', 'Jyy', 'Jxy']
img_list = []; title_list = []; err_count = 0
for i, c in enumerate(error_check):
if c:
img_list.append(imgs[i]); img_list.append(corr_imgs[i])
title_list.append(names[i]); title_list.append(names[i] + ' correct')
err_count += 1
plt.close('all'); view = viewer(img_list, title=title_list, subplots=(err_count,2))
else:
print('Congratulations! Your structure_tensor passed the sanity check.\nRemember that this is not a guarantee that everything is correct.')
```
Now you can also apply `structure_tensor` to the various images we have imported before (corner, dendrochronology, fingerprint, harris_corner, or wave_ramp) and see what the elements of the structure tensor matrix look like. Run the cell below and change the variable `image` to different images. You can also change the `sigma` value and observe the effect this has on the result.
```
# you can change the image to any of the ones we imported: corner, dendrochronology, fingerprint, harris_corner, or wave_ramp
image = wave_ramp
Jxx, Jxy, Jyy = structure_tensor(image, sigma = 2)
plt.close('all')
view = viewer([image, Jxx, Jxy, Jyy], subplots=(2,2))
```
# 2. Orientation features
[Back to table of contents](#ToC_2_Orientation)
Now that we have the structure tensor, we can calculate interesting features from it that help us understand and visualize the orientation of an image. An easy way to think about the structure tensor is that, for each pixel location `[m,n]`, we can calculate a matrix $\mathbf{J}$ made out of the values of $J_{xx}$, $J_{yy}$, and $J_{xy}$ at that same pixel, i.e.,
$$
\mathbf{J}[m,n] = \left[ \begin{array}{cc} J_{xx}[m,n] & J_{xy}[m,n] \\ J_{xy}[m,n] & J_{yy}[m,n]\end{array} \right]\,.
$$
## 2.A. Feature calculation (4 points)
[Back to table of contents](#ToC_2_Orientation)
In the table below, you can see the four features that we are going to compute from this matrix $\mathbf{J}$, where we drop the pixel indeces `[m,n]` for simplicity. There, $\det(\mathbf{J})$ stands for the determinant of the matrix, and $\operatorname{tr}(\mathbf{J})$ for the trace.
| Feature | Relation to the structure tensor matrix $\mathbf{J}$ |
| :-: | :-: |
| Orientation | $$\Theta = \frac{1}{2}\arctan\left(\frac{2J_{xy}}{J_{yy}-J_{xx}}\right)$$ |
| Gradient Energy | $$E = J_{yy}+J_{xx}$$ |
| Coherence | $$C = \frac{\sqrt{(J_{yy}-J_{xx})^2+4J_{xy}^2}}{J_{yy}+J_{xx}}$$ |
| Harris Index | $$H = \det(\mathbf{J}) - \kappa \operatorname{tr}(\mathbf{J})^2\mbox{, with }\kappa = 0.05$$ |
In the cell below, write the function `orientation_features()` which implements the whole chain of processing of the algorithm described in the [flowchart](#Orientation-laboratory-(13-points)) at the start of this lab. Use the `structure_tensor` function you prepared in the previous section to get the structure tensor and use the relations shown in the table above to calculate the specified features (**1 point each**).
<div class="alert alert-info">
**Note:** For $\arctan\left(\frac{x1}{x2}\right)$ in $[-\pi, \pi]$ use `np.arctan2(x1, x2)`.
</div>
<div class="alert alert-warning">
**Beware:** At each pixel where $J_{xx}[m,n]+J_{yy}[m,n] < 0.01$, set the **coherence to 0** to avoid a division by zero.
</div>
```
# Function that calculates the orientation features of an image for a given sigma
def orientation_features(img, sigma):
orientation = np.zeros(img.shape)
energy = np.zeros(img.shape)
coherence = np.zeros(img.shape)
harris = np.zeros(img.shape)
# YOUR CODE HERE
# Calculate Jxx, Jxy, and Jyy
Jxx, Jxy, Jyy = structure_tensor(img, sigma)
J = np.array([[Jxx, Jxy], [Jxy, Jyy]])
# Calculate orientation features
orientation = np.arctan2(2 * Jxy, Jyy - Jxx) / 2
energy = Jyy + Jxx
for x in range(img.shape[0]):
for y in range(img.shape[1]):
if energy[x,y] < 0.01:
coherence[x,y] = 0
else:
coherence[x,y] = np.sqrt((Jyy[x,y] - Jxx[x,y]) ** 2 + (Jxy[x,y] ** 2) * 4) / energy[x,y]
harris = (Jxx * Jyy - Jxy ** 2) - 0.05 * (np.trace(J) ** 2)
return orientation, energy, coherence, harris
# Change the input image and sigma, to see the result on different images: corner, dendrochronology, fingerprint, harris_corner, or wave_ramp
input_img = wave_ramp
sigma = 2
image_list = [input_img] + list(orientation_features(input_img, sigma=sigma))
# Display the output features
title_list = ['Input image', 'Orientation', 'Energy', 'coherence', 'Harris Index']
plt.close('all')
view = viewer(image_list, title = title_list, colorbar=True, widgets=True)
```
As a sanity check, you can run the next four cells to check that each of the output features is in the correct range when applying the function to the `wave_ramp` image using `sigma = 2`. This does not mean that everything is correct but it can help to detect basic calculation errors.
```
# Sanity checks
features = orientation_features(wave_ramp, sigma=2)
# Orientation
if not abs(features[0].min() + np.pi/2) < 0.01 and abs(features[0].max() - np.pi/2) < 0.01:
print(f'WARNING!\nThe orientation should be in [-pi/2, pi/2], not in [{features[0].min():.3f}, {features[0].max():.3f}]')
else:
print('The orientation is in the correct range.')
# Sanity checks
features = orientation_features(wave_ramp, sigma=2)
# Energy
if not abs(features[1].min() - 17) < 1 and abs(features[1].max() - 29) < 1:
print(f'WARNING!\nFor this image, the energy should be in the range [~17, ~29], but it is in [{features[1].min():.3f}, {features[1].max():.3f}]')
else:
print('The energy is in the correct range for this image.')
# Sanity checks
features = orientation_features(wave_ramp, sigma=2)
# coherence
if not abs(features[2].min()) < 0.01 and abs(1 - features[2].max()) < 0.01:
print(f'WARNING\nThe coherence should be in the range [0, 1], not in [{features[2].min():.3f}, {features[2].max():.3f}]')
else:
print('The coherence is in the correct range.')
# Sanity checks
features = orientation_features(wave_ramp, sigma=2)
# Harris index
if not abs(features[3].min() + 41.5) < 1 and abs(features[3].max() - 111.3) < 1:
print(f'WARNING!\nFor this image, the Harris index should be in the range [~-41.5, ~111.3], not in [{features[3].min():.3f}, {features[3].max():.3f}]')
else:
print('The Harris index is in the correct range.')
```
## 2.B. Feature visualization (2 points)
[Back to table of contents](#ToC_2_Orientation)
Until now, we used grayscale (2D) images to visualize the orientation features. This only allows for single features to be displayed at once. It is often convenient to be able to visualize several features at once and integrate them into the original image to make the visual analysis more intuitive. One way to achieve this is to use the **hue, saturation, value (HSV)** color representation, which is sometimes also called HSB (for brightness), and is depicted in the figure below. Using this method, we can assign the orientation (a $2\pi$ periodic value) to the hue (which also lives in a circular scale), the coherence (a value between $0$ and $1$) to the saturation, and the brightness of the original image to the value, which will allow us to see the objects in the image. This will result on much clearer representations of the orientation features.
<img src="images/hsv_color_representation.png" alt="Drawing" style="width: 400px;"/>
In the cell below, **for 2 points**, implement the function `colorize_features` that takes as input parameters the *orientation*, the *coherence*, the *input image*, and a *mode* (of two possible modes, see table below). The function should
1. create an HSV image `hsv_image` from the orientation features, and
2. convert this HSV image to an RGB image using `rgb_img = cv.cvtColor(hsv_img, cv.COLOR_HSV2RGB)` in order to display it with the `viewer`.
Below, both `hsv_img` and `rgb_img` are 3D arrays with dimensions `(height, width, channels)` where the $i$-th channel can be accessed as `img[:, :, i]` and it contains the information relative to the letter in position $i$ of the name of the respective color representation (HSV or RGB: $i=0$, hue or red channel, $i=1$, saturation or green channel, $i=2$, value or blue channel).
The function will have two modes. Mode $0$ simply maps the orientation to the hue channel, while mode $1$ overlays the orientation and coherence features on top of the input image. See the table below for the specification of the two modes.
| Mode | H channel, range `[0, 180]` | S channel, range `[0, 255]` | V channel, range `[0, 255]` |
| :-: | :-: | :-: | :-: |
| 0: Orientation only | orientation | constant image of value `255` | constant image of value `255` |
| 1: Features on image | orientation | coherence | input image |
<div class="alert alert-warning">
**Beware:** Think carefully about how each channel should (or should not) be shifted and/or normalized. Any two pixels with the same brightness, orientation and coherence features in different images should look the same! Note that the orientation as you computed it above is in the range $[-\pi/2,\pi/2]$, and the coherence is in the range $[0,1]$. Assume the input image is in the range $[0,255]$.
</div>
<div class="alert alert-warning">
**Beware:** In order for the `cvtColor` function to work, it's important that the images are of type `uint8`. Please simply modify the predefined variable `hsv_img` to create the HSV image, and store the RGB image in the predefined variable `rgb_img`.
</div>
<div class="alert alert-info">
**Note:** If you are curious about how the conversion from HSV to RGB works, you can check out the formula [here](https://en.wikipedia.org/wiki/HSL_and_HSV#HSV_to_RGB).
</div>
```
# Function that returns a colorized rgb image depending on the orientation features
def colorize_features(orientation, coherence, img, mode):
# Fill hsv_img[:,:,0] to set the hue, hsv_img[:,:,1] to set the saturation, and hsv_img[:,:,2] to set the value
hsv_img = np.zeros((img.shape[0], img.shape[1], 3), dtype = np.uint8)
rgb_img = np.zeros((img.shape[0], img.shape[1], 3), dtype = np.uint8)
# YOUR CODE HERE
# Orientation Only
if mode == 0:
hsv_img[:, :, 0] = 90 + orientation * (180 / np.pi)
hsv_img[:, :, 1] = 255
hsv_img[:, :, 2] = 255
# Features On Image
elif mode == 1:
hsv_img[:, :, 0] = 90 + orientation * (180 / np.pi)
hsv_img[:, :, 1] = coherence * 255
hsv_img[:, :, 2] = img
# If not any mode, raise ValueError
else:
raise ValueError("Only mode = 0 and mode = 1 are acceptable values.")
print(hsv_img)
rgb_img = cv.cvtColor(hsv_img, cv.COLOR_HSV2RGB)
print(rgb_img)
return rgb_img
```
Now run the next two cells for a quick test on your function. As usual, remember that these tests are not definitive and that they do not guarantee the full points.
```
# Sanity check for mode 0. First we define a few arrays for which we know how the output will look like.
orientation = np.array([[-np.pi/2, -np.pi/4, 0, np.pi/4, np.pi/2]])
coherence = np.array([[0, 65./255, 130./255, 195./255, 255./255]])
img = np.array([[255, 195, 130, 65, 0]])
colorized_img = colorize_features(orientation, coherence, img, mode=0)
check_img = np.array([[[255, 0, 0], [127, 255, 0], [0, 255, 255],[128, 0, 255], [255, 0, 0]]], dtype=np.uint8)
if not np.allclose(colorized_img, check_img):
print('WARNING!\nYour colorization function is not yet correct for mode 0. Check the comparison below:')
plt.close('all')
view = viewer([check_img, colorized_img], title=['Expected output', 'Your output'], subplots=(1,2))
else:
print('Well done, your colorization function passed the sanity check for mode 0.')
# Sanity check for mode 1
orientation = np.array([[-np.pi/2, -np.pi/4, 0, np.pi/4, np.pi/2]])
coherence = np.array([[0, 65./255, 130./255, 195./255, 255./255]])
img = np.array([[255, 195, 130, 65, 0]])
colorized_img = colorize_features(orientation, coherence, img, mode=1)
check_img = np.array([[[255, 255, 255], [170, 195, 145], [64, 130, 130],[40, 15, 65], [0, 0, 0]]], dtype=np.uint8)
if not np.allclose(colorized_img, check_img):
print('WARNING!\nYour colorization function is not yet correct for mode 1. Check the comparison image below:')
plt.close('all')
view = viewer([check_img, colorized_img], title=['Expected output', 'Your output'], subplots=(1,2))
else:
print('Well done, your colorization function passed the sanity check for mode 1.')
```
Run the cell below to visualize the effect of the `colorize_features()` function on different images.
<div class="alert alert-info">
**Note:** Click on `Extra Widgets` to change the mode and sigma values and apply the colorization by clicking on `Apply Colorization`. Cycle through the different images by clicking on `Next` and `Prev`.
</div>
```
# Define control widgets for "Extra Widgets"
mode_dropdown = widgets.Dropdown(options=['0: Orientation only', '1: Features on image'],value='1: Features on image',description='Mode:',disabled=False)
mode_dictionary = {'0: Orientation only':0, '1: Features on image':1}
sigma_slider = widgets.IntSlider(value = 3, min = 1, max = 15, step = 1, description = r'$\sigma$')
button = widgets.Button(description = 'Apply Colorization')
def colorization_callback(img):
mode = mode_dictionary[mode_dropdown.value]
sigma = sigma_slider.value
# Get the features using the function from part 2.1
features = orientation_features(img, sigma=sigma)
# Create the colorized image
output = colorize_features(features[0], features[2], img, mode=mode)
return output
plt.close('all')
image_list = [255*(wave_ramp-wave_ramp.min())/(wave_ramp.max()-wave_ramp.min()), dendrochronology, fingerprint]
title = ["wave_ramp", "dendrochronology", "fingerprint"]
new_widgets = [mode_dropdown, sigma_slider, button]
view = viewer(image_list, new_widgets=new_widgets, callbacks=[colorization_callback], widgets=True, title=title)
```
# 3. Application
[Back to table of contents](#ToC_2_Orientation)
In this section you will implement some applications that rely on the functions you implemented in the previous sections, in order to show you what they could be used for in real-life scenarios.
## 3.A. Selection of specific orientations (2 points)
[Back to table of contents](#ToC_2_Orientation)
We propose to develop a function that only selects areas of the image with a specific orientation. In particular, the algorithm has to preserve pixels which have the following structure-tensor features,
- $E > T E_{max}$, where $E_{max}$ is the maximum energy in the image and $T\in[0,1]$ is the relative threshold,
- $C > 0.5$,
- $\theta_{min} \leq \theta(x, y) \leq \theta_{max}$, with $\theta$ in radians.
In the next cell, **for 2 points**, implement the function `select_direction(img, sigma, T, theta_min, theta_max)` that takes as input parameters:
* `img`: The input image
* `sigma`: $\sigma$ to be used in `orientation_features`
* `T`: The relative energy threshold
* `theta_min`: The minimum angle $\theta_{min}$
* `theta_max`: The maximum angle $\theta_{max}$
and that returns:
* `output`: Output image keeping the pixels with the given features, with all other pixels set to the minimum value of the image (which is not always 0).
Use the function `orientation_features` you implemented in [Part 2.A.](#2.A.-Feature-calculation-(4-points)) to get the features needed.
<div class="alert alert-warning">
**Note:** Since the angle $\theta$ is periodic, it is not always clear which angle is larger. You should account for the fact that $\theta_{\mathrm{min}}$ can be larger than $\theta_{\mathrm{max}}$: If $\theta_{\mathrm{min}} \leq \theta_{\mathrm{max}}$ then return the values inside the range [$\theta_{\mathrm{min}}$, $\theta_{\mathrm{max}}$], otherwise return the values that are outside this range, i.e., $[-\pi/2,\pi/2]\setminus (\theta_{\mathrm{max}}, \theta_{\mathrm{min}})$ (see [here](https://en.wikipedia.org/wiki/Complement_(set_theory)#Relative_complement)). For example: if $\theta_{\mathrm{min}} = \frac{\pi}{3}$ and $\theta_{\mathrm{max}} = -\frac{\pi}{3}$ the function should keep all orientation in the ranges $[\frac{\pi}{3}, \frac{\pi}{2}]$ and $[-\frac{\pi}{2}, -\frac{\pi}{3}]$ but discard all orientations in the range $(-\frac{\pi}{3}, \frac{\pi}{3})$. Drawing this example will certainly help you to understand this better.
</div>
<div class="alert alert-info">
**Hint:** You can use [`np.logical_and(condition_1, condition_2)`](https://numpy.org/doc/stable/reference/generated/numpy.logical_and.html) to get the boolean array for which both `condition_1` and `condition_2` are true.
</div>
```
# Function that extracts a range of orientations given by theta_min and theta_max
def select_direction(img, sigma, T, theta_min, theta_max):
# Check input params
assert 0 < T < 1, 'The threshold should be between 0 and 1'
assert -np.pi/2 <= theta_min <= np.pi/2, 'theta_min should be in [-pi/2, pi/2]'
assert -np.pi/2 <= theta_max <= np.pi/2, 'theta_max should be in [-pi/2, pi/2]'
output = img.copy()
# YOUR CODE HERE
# get orientation features
orientation, energy, coherence, _ = orientation_features(img, sigma)
logical_M = None
# normal case: theta_min <= theta_max
if theta_min <= theta_max:
logical_M = np.logical_and(((orientation >= theta_min) & (orientation <= theta_max)),
np.logical_and(energy > T * np.max(energy), coherence > 0.5))
# special case: theta_min > theta_max
else:
logical_M = np.logical_and((((orientation >= -np.pi/2) & (orientation <= theta_max)) | ((orientation >= theta_min) & (orientation <= np.pi/2))),
np.logical_and(energy > T * np.max(energy), coherence > 0.5))
output[logical_M == False] = np.min(img)
return output
```
The next cell will evaluate your function on a test image that consists of 4 lines at the angles $0$, $\frac{\pi}{4}$, $-\frac{\pi}{4}$ and $\frac{\pi}{2}$. The function will be called on this test image with the ranges $[-\frac{\pi}{6}, \frac{\pi}{6}]$, $[\frac{\pi}{6}, \frac{\pi}{3}]$, $[-\frac{\pi}{3}, -\frac{\pi}{6}]$, and $[\frac{\pi}{3}, -\frac{\pi}{3}]$, which should each exctract only one of the lines. Run the cell below to apply this sanity check.
```
# Create test image consisting of 4 lines at 0, pi/2, pi/4 and -pi/4
n = 51; r = n//2
line0 = np.zeros((n,n)); line0[r, :r-6] = 1; line0[r, r+7:] = 1
line90 = np.zeros((n,n)); line90[:r-6, r] = 1; line90[r+7:, r] = 1
line45 = np.zeros((n,n)); line45[range(n-1, r+6, -1), range(r-6)] = 1
line45[range(r-7, -1, -1), range(r+7, n)] = 1
lineM45 = np.zeros((n,n)); lineM45[range(r-6), range(r-6)] = 1
lineM45[range(r+7, n), range(r+7, n)] = 1
test_img = line0 + line90 + line45 + lineM45
plt.close('all')
view = viewer(test_img)
# Test the 4 lines
lines = [line0, line45, lineM45, line90]
ranges = [[-np.pi/6, np.pi/6], [np.pi/6, np.pi/3], [-np.pi/3, -np.pi/6], [np.pi/3, -np.pi/3]]
names = ['horizontal', 'diagonal ascending', 'diagonal descending', 'vertical']
check = True
for i, ran in enumerate(ranges):
test_dir = select_direction(test_img, T=0.1, sigma=2, theta_min=ran[0], theta_max=ran[1])
if not np.allclose(test_dir, lines[i]):
check = False
print(f'WARNING!\nOnly the {names[i]} line should be visible at theta_min={ran[0]:.3f}, theta_max={ran[1]:.3f}!\n')
view = viewer([test_dir, lines[i]], title=[f'Your output for min={ran[0]:.3f}, max={ran[1]:.3f}', 'Expected output'], subplots=(1,2))
if check:
print('Well done, your function passed the sanity check!')
```
Run the next cell to apply your function to the images we have been working with and play around with the different parameters by clicking on the button *Extra Widgets*. You can cycle through the different images by clicking on *Next* and *Prev*. Note that for some images the threshold $T$ needs to be set very low in order to extract any orientation.
```
# Define Sliders and button
min_slider = widgets.FloatSlider(value = -np.pi/2, min = -np.pi/2, max = np.pi/2, step=0.01, description = r'$\theta_\mathrm{min}$')
max_slider = widgets.FloatSlider(value = np.pi/2, min = -np.pi/2, max = np.pi/2, step=0.01, description = r'$\theta_\mathrm{max}$')
T_slider = widgets.FloatSlider(value = 0.5, min = 0.05, max = 0.95, step=0.05, description = r'$T$')
sigma_slider = widgets.IntSlider(value = 3, min = 1, max = 15, step = 1, description = r'$\sigma$')
button = widgets.Button(description = 'Extract Orientation')
# Define Callback function
def orientation_callback(img):
theta_min = min_slider.value
theta_max = max_slider.value
T = T_slider.value
sigma = sigma_slider.value
# Create the colorized image
output = select_direction(img, sigma, T, theta_min, theta_max)
return output
# Declare viewer parameters and start the visualization
plt.close('all')
image_list = [wave_ramp, dendrochronology, fingerprint]
new_widgets = [sigma_slider, T_slider, min_slider, max_slider, button]
view = viewer(image_list, new_widgets = new_widgets, callbacks = [orientation_callback], widgets = True)
```
## 3.B. Harris corner detector (2 points)
[Back to table of contents](#ToC_2_Orientation)
As you may remember from the lecture, the Harris index can be interpreted as the probability of having a corner at the corresponding location. This means we can implement a very basic corner detector by extracting the local maxima of the Harris index image and using them as the locations for our corners.
**For 1 point**, complete the function `detect_corners()` in the cell below. This function takes as parameters
* `img`: An image where to detect corners, in the form of a 2D NumPy array,
* `L`: the size (in integer number of pixels) of the square observation region around each pixel (that defines what a _local maxima_ is), and
* `T`: a relative threshold in the range $[0,1]$, where only local maxima that are above $T$ times the image maximum are kept.
The function returns:
* `output`: A list of coordinates where a local maximum has been identified, as specified by the `peak_local_max` function (see the hint below).
<div class = 'alert alert-info'>
**Hints:**
* To get the Harris index image, use the function `orientation_features` that you implemented in [Part 2.A](#2.-Orientation-features) **with `sigma=1`**.
* To extract the local maxima, use the function [`peak_local_max`](https://scikit-image.org/docs/dev/api/skimage.feature.html#skimage.feature.peak_local_max) provided by `skimage.feature`. It should be called as `feature.peak_local_max(input_img, min_distance, threshold_rel)`.
* The parameter `threshold_rel` (in the function `peak_local_max`) is given by the input parameter `T`.
* The parameter `min_distance` (in the function `peak_local_max`) is the minimum distance separating two local maxima in an $L\times L$ region. It is given by the relation $L = 2\;*$ `min_distance` $+ 1$.
</div>
```
# Function that detects corners using the Harris index method
def detect_corners(img, L, T):
# YOUR CODE HERE
_, _, _, harris = orientation_features(img, sigma=1)
output = feature.peak_local_max(harris, min_distance=int((L-1)/2), threshold_rel=T)
return output
```
Run the cell below define the function `show_red_crosses`, which produces a nice RGB image to visualize the results of the Harris corner detector.
```
# Function to prepare a nice visualization for Harris corners' detections
def show_red_crosses(img, corners):
# Create binary mask
peak_mask = np.zeros_like(img)
peak_mask[tuple(corners.T)] = True
# Create RGB image with red crosses in the corners
corners = cv.dilate(peak_mask, cv.getStructuringElement(cv.MORPH_CROSS, (5,5)))
output = cv.cvtColor(img, cv.COLOR_GRAY2RGB)
output[corners>0] = [255,0,0]
return output
```
Run the next cell to test your `detect_corners` function on a test image that contains 12 corners. Your function should be able to detect them all correctly.
```
# Create test image
n = 51; r = n // 2
test_img = np.zeros((n,n), dtype = np.uint8)
test_img[r-15:r+16,r-5:r+6] = 255
test_img[r-5:r+6,r-15:r+16] = 255
# Run corner detection
detected_corners = detect_corners(test_img, 3, 0.5)
# Visualize result
plt.close('all')
view = viewer(show_red_crosses(test_img,detected_corners))
# Check that the number of detected corners is 12
if not len(detected_corners) == 12:
print(f'WARNING!\nSorry but your function detected {len(detected_corners)} corners instead of 12. Check your code and try again!')
else:
print("Well done, your function detected 12 corners in the image! However, it's your job to verify the correct location by looking at the picture.")
```
Now, run the cell below to create an extra widget in the viewer and experiments with `L` and `T` to answer the upcoming MCQ.
```
qr_code = cv.imread('images/qr_code.png', cv.IMREAD_GRAYSCALE)
# Define Sliders and button
T_slider = widgets.FloatSlider(value = 0.5, min = 0.05, max = 0.95, step=0.05, description = r'$T$')
L_slider = widgets.IntSlider(value = 3, min = 1, max = 31, step = 2, description = r'$L$')
button = widgets.Button(description = 'Detect Corners')
# Define Callback function
def corner_callback(img):
T = T_slider.value
L = L_slider.value
return show_red_crosses(img,detect_corners(img, L=L, T=T))
# Start the visualization
plt.close('all')
view = viewer([harris_corner, corner, qr_code], new_widgets=[T_slider, L_slider, button],
callbacks=[corner_callback], widgets=True)
```
### Multiple Choice Question
* Q1: In general, a higher $L$ leads to
1. more corners, because a higher area is covered,
2. less corners, because values need to be a local maximum in a larger area,
3. less corners, because less areas of the image are observed, or
4. more corners, because more areas of the image are observed.
* Q2: In general, selecting a higher $T$ means that a corner has to be ... to be detected.
1. sharper, so that it's more defined,
2. rounder, so that it's less defined, or
3. more diffuse, so that it covers more area.
In the next cell, modify the variables `answer_one` and `answer_two` to reflect your answers. The following two cells are for you to check that your answer is in the valid range.
```
# Modify these variables
answer_one = 2
answer_two = 1
# YOUR CODE HERE
# Sanity check
if not answer_one in [1, 2, 3, 4]:
print('WARNING!\nChoose one of 1, 2, 3 or 4.')
# Sanity check
if not answer_two in [1, 2, 3]:
print('WARNING!\nChoose one of 1, 2 or 3.')
```
## 3.C. *Advanced:* Isotropic filtering in the Fourier space (1 point)
[Back to table of contents](#ToC_2_Orientation)
In the first exercise you were told to use the Sobel filter to compute the gradient for the structure tensor. As you have seen, this worked fine for everything we did until now. However, if we inspect the resulting orientation image more carefully, we can see some strange behavior. To illustrate this, let us first create a perfect test image, that contains (inside a certain radius of the image) exactly the same amount of pixels for every possible orientation.
Run the next cell to create this test image and define two useful functions that we will need later.
```
# Function that creates the orientation test image. This image is perfectly isotropic
def create_test_img(ny, nx):
# Minimum and maximum radial frequencies
fmin = 0.02; fmax = 8 * fmin;
# Center
hx = nx / 2; hy = ny / 2;
n = min(nx, ny);
def structure_func(j, i):
# Radius
r = np.sqrt((i - hx)**2 + (j - hy)**2);
# Radial envelope
u = 1.0 / (1.0 + np.exp((r - n * 0.45) / 2.0));
# Radial frequency profile
f = fmin + r * (fmax - fmin) / n;
# Radial modulating function
v = np.sin(np.pi * 2 * f * r);
return (1.0 + v * u) * 128
return np.fromfunction(structure_func, shape=(ny, nx)).astype(np.uint8)
# Calculates the Fourier Transform
def get_FT(img):
return np.fft.fftshift(np.fft.fft2(img))
# Calculates the inverse Fourier Transform
def get_iFT(img):
return np.fft.ifft2(np.fft.ifftshift(img)).real
test_img = create_test_img(2048, 2048)
plt.close('all')
view = viewer(test_img)
```
Now, in theory, if we extract the orientations of a circular cut-out of this test image using the `orientation_features` function that you coded in [Part 2.A.]((#2.A.-Feature-calculation-(4-points))) and plot the distribution of these orientations, we should get a completely flat line. Let's see how it actually looks: run the cell below to display the orientation image and its distribution inside a circular cut-out.
```
# Get the old orientation
orientation = orientation_features(test_img, sigma=5)[0]
# Generate the disk-shaped mask to only analize the colors inside the correct radius
mask = np.fromfunction(lambda i, j: np.sqrt((i-orientation.shape[0]//2)**2 + (j-orientation.shape[1]//2)**2),
shape=orientation.shape)
r = np.min(mask.shape)//2 - np.min(mask.shape)//20
# Generate histograms using the mask
hist, edg = np.histogram(orientation[mask < r], bins = 1000, range = (orientation.min(), orientation.max()))
cent = (edg[0:-1] + edg[1:])/2
# Display the masked orientation
orientation[mask >= r] = -np.pi/2
plt.close('all')
view = viewer([orientation], title=['Orientation of the test image'], cmap='hsv', colorbar=True)
# Display the angle distribution
f = plt.figure(); plt.title('Orientation distribution'); plt.xlabel('Angle [rad]'); plt.ylabel('# pixels')
plt.plot(cent, hist); plt.grid();
plt.xticks(ticks=[-np.pi/2,-np.pi/4,0,np.pi/4,np.pi/2],labels=[r'$-\pi/2$',r'$-\pi/4$',r'$0$',r'$\pi/4$',r'$\pi/2$']); plt.show()
```
In the first image, you will probably not see any problem since it's very hard to see it by just looking at the orientation image. However, in the distribution plot you can clearly see that some orientations are preferred over others. Specifically, ignoring the very sharp peaks at $0$, $\pm\frac{\pi}{4}$ and $\pm\frac{\pi}{2}$ (an artefact of having a limited number of pixels), there are more pixels with an orientation close to $0$ or $\pm\frac{\pi}{2}$ and less pixels close to $\pm\frac{\pi}{4}$.
Now why could that be? If we look more closely at the Sobel filter masks $h_x$ and $h_y$
$$h_x =
\begin{bmatrix}
1 & 0 & -1 \\
2 & 0 & -2 \\
1 & 0 & -1
\end{bmatrix}
,\;\;\;\;
h_y =
\begin{bmatrix}
1 & 2 & 1 \\
0 & 0 & 0 \\
-1 & -2 & -1
\end{bmatrix}
$$
we can see that this filter is not isotropic, meaning it doesn't treat all directions equally. In this case, as we saw in the orientation distribution above, the horizontal and vertical edges are favored, resulting in a biased orientation distribution.
In order to have a non-biased orientation detector, we need to re-implement the `structure_tensor` function using an isotropic gradient filter. There are several to choose from but in this exercise we will calculate the gradient using the Fourier property
$$\frac{\partial f(x,y)}{\partial x} \xrightarrow{\mathcal{F}} j\omega_x\operatorname{F}(\omega_x, \omega_y), \;\;\;
\frac{\partial f(x,y)}{\partial y} \xrightarrow{\mathcal{F}} j\omega_y\operatorname{F}(\omega_x, \omega_y)\,.$$
As you can see we only need to take the Fourier transform of our image and multiply it with either $j\omega_x$ or $j\omega_y$. Then we can take the inverse Fourier transform and we will have our derivatives in the $x$ and $y$ directions.
**For 1 point**, implement the functions `get_w` and `structure_tensor_improved` in the cells below.
The first step for you is to implement the function `get_w` that simply returns a vector $\omega$ consisting of $m$ equidistant points going from $-\frac{\pi}{2}$ to $\frac{\pi}{2}$.
This is a very simple function that will be used in `structure_tensor_improved` to generate the complex-valued vectors $j\omega_x$ and $j\omega_y$.
<div class="alert alert-info">
**Hint:** Use the function [`np.linspace`](https://numpy.org/doc/stable/reference/generated/numpy.linspace.html).
</div>
```
# Calculates omega (w) of length m
def get_w(m):
w = np.zeros(m)
# YOUR CODE HERE
w = np.linspace(-np.pi/2, np.pi/2, num=m, endpoint=True)
return w
# Visualize omega
plt.close('all')
plt.figure("Frequency vector")
plt.plot(get_w(100))
plt.ylabel(r'$\omega$'); plt.xlabel("Vector index"); plt.grid()
plt.yticks(ticks=[-np.pi/2,-np.pi/4,0,np.pi/4,np.pi/2],labels=[r'$-\pi/2$',r'$-\pi/4$',r'$0$',r'$\pi/4$',r'$\pi/2$']);
plt.show()
```
Run the next cell for a quick test on $\omega$.
```
# Perform test on a vector of 10 elements. We will check length, range, and a direct comparison with the solution
w_out = get_w(10)
check = True
if not len(w_out) == 10:
check = False
print(f'WARNING!\nThe length of the output vector should be equal to the input parameter m. Expected length=10, your length={len(w_out)}.')
if not np.min(w_out) == -np.pi/2 and np.max(w_out) == np.pi/2:
check = False
print(f'WARNING!\nOmega should go from -1.57079633 to 1.57079633 not from {np.min(w_out):.8f} to {np.max(w_out):.8f}!')
if not np.allclose(w_out, [-1.57079633, -1.22173048, -0.87266463, -0.52359878, -0.17453293,
0.17453293, 0.52359878, 0.87266463, 1.22173048, 1.57079633]):
check = False
print('WARNING!\nYour omega vector is not yet correct!')
if check:
print('Well done, the function passed the sanity check.')
```
In the next cell, complete the function `structure_tensor_improved` by implementing the Fourier domain gradient filters as explained above. The code that generates the arrays $j\omega_x$ and $j\omega_y$ is already provided. It uses your function `get_w` to fill both the rows of the imaginary part of `jw_x` and the columns of the imaginary part of `jw_y` with the vector $\omega$.
<div class="alert alert-info">
**Hint:** You can use the functions `get_FT(img)` to get the complex Fourier transform of `img` and `get_iFT(img_FT)` to get the inverse Fourier transform of `img_FT`. Those functions also take care of performing the correct shifting.
</div>
<div class="alert alert-info">
**Hint:** Implementing these filters might be easier than you think! Or exactly as easy as you thought, but did not know why. Have a look at the NumPy broadcasting rules [here](https://numpy.org/doc/stable/user/basics.broadcasting.html).
</div>
```
# Function that calculates the elements Jxx, Jxy and Jyy of the structure tensor matrix
def structure_tensor_improved(img, sigma):
Jxx = None
Jxy = None
Jyy = None
# Generate the complex-numbered filters jw_x and jw_y
jw_x = 1j*get_w(img.shape[1])
jw_y = 1j*get_w(img.shape[0]).reshape((img.shape[0], 1))
# Gradient calculation using jw_x and jw_y in the Fourier space
# YOUR CODE HERE
grad_x = get_iFT(jw_x * get_FT(img))
grad_y = get_iFT(jw_y * get_FT(img))
# Calculate fxx, fxy, fyy and then Jxx, Jxy and Jyy from the gradients
# in exactly the same way as in the original structure tensor function (Part 1)
# YOUR CODE HERE
fxx = grad_x ** 2
fxy = grad_x * grad_y
fyy = grad_y ** 2
Jxx = cv.GaussianBlur(src=fxx, ksize=(0,0), sigmaX=sigma, borderType=cv.BORDER_DEFAULT)
Jxy = cv.GaussianBlur(src=fxy, ksize=(0,0), sigmaX=sigma, borderType=cv.BORDER_DEFAULT)
Jyy = cv.GaussianBlur(src=fyy, ksize=(0,0), sigmaX=sigma, borderType=cv.BORDER_DEFAULT)
return Jxx, Jxy, Jyy
```
In this part you will need to do the sanity check of the `structure_tensor_improved` function by eye. The cell below will run the function `orientation_features` twice on the `test_img`, once with the old `structure_tensor` and once with the improved version.
You will be able to see the results in an *IPLabViwer*, where you will see the orientation feature for each version of the `structure_tensor` function, as well as the difference between both results. Once you are sure on the correctness of `structure_tensor_improved`, reflect on the difference and its origins. Moreover, you will see a plot of both orientation distributions.
The displayed orientation feature is for you to see that you did not mess up the function, and you will most likely not be able to see much difference between the two versions. However, in the orientation distribution plot you should be able to clearly distinguish the Fourier version from the Sobel one. **The Fourier version should be completely flat (with exception of a few sharp spikes)** compared to the Sobel version that looks more like a sinusoidal wave.
```
# Get the old orientation
orientation = orientation_features(test_img, sigma=5)[0]
# Replace the structure tensor function by the improved one
old_ST = structure_tensor
structure_tensor = structure_tensor_improved
# Get the improved orientation
try:
orientation_improved = orientation_features(test_img, sigma=5)[0]
except:
# Handle errors in the code
structure_tensor = old_ST
raise
# Restore the old structure tensor function
structure_tensor = old_ST
# Generate the disk-shaped mask to only analize the colors inside the correct radius
mask = np.fromfunction(lambda i, j: np.sqrt((i-orientation.shape[0]//2)**2 + (j-orientation.shape[1]//2)**2),
shape=orientation.shape)
r = np.min(mask.shape)//2-np.min(mask.shape)//20
# Generate histograms using the mask
hist, edg = np.histogram(orientation[mask < r], bins = 1000, range = (orientation.min(), orientation.max()))
hist_improved, edg_improved = np.histogram(orientation_improved[mask < r], bins = 1000,
range = (orientation_improved.min(), orientation_improved.max()))
cent = (edg[0:-1] + edg[1:])/2;
cent_improved = (edg_improved[0:-1] + edg_improved[1:])/2;
# Display the two masked orieantations
orientation[mask >= r] = -np.pi/2
orientation_improved[mask >= r] = -np.pi/2
plt.close('all')
image_list = [orientation, orientation_improved]
title_list = ['Sobel orientation', 'Fourier orientation']
view = viewer(image_list, title=title_list, cmap='hsv', colorbar=True, widgets=True)
# Display the angle distribution
plt.figure()
plt.title('Orientation distribution'); plt.xlabel('Angle [rad]'); plt.ylabel('# pixels')
plt.xticks(ticks=[-np.pi/2,-np.pi/4,0,np.pi/4,np.pi/2],labels=[r'$-\pi/2$',r'$-\pi/4$',r'$0$',r'$\pi/4$',r'$\pi/2$']);
plt.plot(cent, hist)
plt.plot(cent_improved, hist_improved)
plt.legend(['Sobel orientation', 'Fourier orientation']); plt.grid(); plt.show()
```
<div class="alert alert-success">
<b>Congratulations on finishing the Orientation lab!!</b>
</div>
Make sure to save your notebook (you might want to keep a copy on your personal computer) and upload it to <a href="https://moodle.epfl.ch/course/view.php?id=463">Moodle</a>, **in a zip file with the other notebook of this lab.**
* Keep the name of the notebook as: *2_Orientation.ipynb*,
* Name the `zip` file: *Orientation_lab.zip*.
| github_jupyter |
# SageMaker endpoint
To deploy the model you previously trained, you need to create a Sagemaker Endpoint. This is a hosted prediction service that you can use to perform inference.
## Finding the model
This notebook uses a stored model if it exists. If you recently ran a training example that use the `%store%` magic, it will be restored in the next cell.
Otherwise, you can pass the URI to the model file (a .tar.gz file) in the `model_data` variable.
You can find your model files through the [SageMaker console](https://console.aws.amazon.com/sagemaker/home) by choosing **Training > Training jobs** in the left navigation pane. Find your recent training job, choose it, and then look for the `s3://` link in the **Output** pane. Uncomment the model_data line in the next cell that manually sets the model's URI.
```
# Retrieve a saved model from a previous notebook run's stored variable
%store -r model_data
# If no model was found, set it manually here.
# model_data = 's3://sagemaker-us-west-2-XXX/pytorch-smdataparallel-mnist-2020-10-16-17-15-16-419/output/model.tar.gz'
print("Using this model: {}".format(model_data))
```
## Create a model object
You define the model object by using SageMaker SDK's `PyTorchModel` and pass in the model from the `estimator` and the `entry_point`. The endpoint's entry point for inference is defined by `model_fn` as seen in the following code block that prints out `inference.py`. The function loads the model and sets it to use a GPU, if available.
```
!pygmentize code/inference.py
import sagemaker
role = sagemaker.get_execution_role()
from sagemaker.pytorch import PyTorchModel
model = PyTorchModel(model_data=model_data, source_dir='code',
entry_point='inference.py', role=role, framework_version='1.6.0', py_version='py3')
```
### Deploy the model on an endpoint
You create a `predictor` by using the `model.deploy` function. You can optionally change both the instance count and instance type.
```
predictor = model.deploy(initial_instance_count=1, instance_type='ml.m4.xlarge')
```
## Test the model
You can test the depolyed model using samples from the test set.
```
# Download the test set
from torchvision import datasets, transforms
from torch.utils.data import DataLoader
# TODO: can be removed after upgrade to torchvision==0.9.1
# see github.com/pytorch/vision/issues/1938 and github.com/pytorch/vision/issues/3549
datasets.MNIST.urls = [
'https://ossci-datasets.s3.amazonaws.com/mnist/train-images-idx3-ubyte.gz',
'https://ossci-datasets.s3.amazonaws.com/mnist/train-labels-idx1-ubyte.gz',
'https://ossci-datasets.s3.amazonaws.com/mnist/t10k-images-idx3-ubyte.gz',
'https://ossci-datasets.s3.amazonaws.com/mnist/t10k-labels-idx1-ubyte.gz'
]
test_set = datasets.MNIST('data', download=True, train=False,
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
]))
# Randomly sample 16 images from the test set
test_loader = DataLoader(test_set, shuffle=True, batch_size=16)
test_images, _ = iter(test_loader).next()
# inspect the images
import torchvision
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
def imshow(img):
img = img.numpy()
img = np.transpose(img, (1, 2, 0))
plt.imshow(img)
return
# unnormalize the test images for displaying
unnorm_images = (test_images * 0.3081) + 0.1307
print("Sampled test images: ")
imshow(torchvision.utils.make_grid(unnorm_images))
# Send the sampled images to endpoint for inference
outputs = predictor.predict(test_images.numpy())
predicted = np.argmax(outputs, axis=1)
print("Predictions: ")
print(predicted.tolist())
```
## Cleanup
If you don't intend on trying out inference or to do anything else with the endpoint, you should delete it.
```
predictor.delete_endpoint()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/PepeScott/daa_2021_1/blob/master/Tarea11.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
class NodoArbol:
def __init__(self,dato,hijo_izq=None,hijo_der=None):
self.dato=dato
self.left=hijo_izq
self.right=hijo_der
class BinarySearchTree:
def __init__(self):
self.__root=None
def insert(self,value):
if self.__root==None:
self.__root=NodoArbol(value,None,None)
else:
self.__insert_nodo__(self.__root,value)
def __insert_nodo__(self,nodo,value):
if nodo.dato == value:
pass
elif value < nodo.dato:
if nodo.left == None:
nodo.left=NodoArbol(value,None,None)
else:
self.__insert_nodo__(nodo.left,value)
else:
if nodo.right==None:
nodo.right=NodoArbol(value,None,None)
else:
self.__insert_nodo__(nodo.right,value)
def buscar(self,value):
if self.__root == None:
return None
else:
return self.__busca_nodo(self.__root, value)
def __busca_nodo(self,nodo,value):
if nodo == None:
return None
elif nodo.dato == value:
return nodo.dato
elif value < nodo.dato:
return self.__busca_nodo(nodo.left,value)
else:
return self.__busca_nodo(nodo.right,value)
def transversal(self,format="inorden"):
if format == "inorden":
self.__recorrido_in(self.__root)
elif format == "preorden":
self.__recorrido_pre(self.__root)
elif format == "posorden":
self.__recorrido_pos(self.__root)
else:
print("Formato de recorrido no valido")
def __recorrido_pre(self,nodo):
if nodo != None:
print(nodo.dato,end=",")
self.__recorrido_pre(nodo.left)
self.__recorrido_pre(nodo.right)
def __recorrido_in(self,nodo):
if nodo != None:
self.__recorrido_in(nodo.left)
print(nodo.dato,end=",")
self.__recorrido_in(nodo.right)
def __recorrido_pos(self,nodo):
if nodo != None:
self.__recorrido_pos(nodo.left)
self.__recorrido_pos(nodo.right)
print(nodo.dato,end=",")
def borrar(self, value):
if self.__root == None:
return None
else:
return self.__recorrido_borrar(self.__root,value)
def __recorrido_borrar(self,nodo,value):
if nodo.dato== None:
return None
if nodo.dato == value:
if nodo.left != None and nodo.right != None:
value=self.__remplazo_nodo_dos_hojas(nodo,nodo.right)
nodo.dato=value.dato
elif value < nodo.dato:
return self.__recorrido_borrar(nodo.left,value)
else:
return self.__recorrido_borrar(nodo.right,value)
def __remplazo_nodo_dos_hojas(self,nodor,nodo):
if nodo == None:
return None
elif nodo.left != None:
return self.__remplazo_nodo_dos_hojas(nodor,nodo.left)
else:
nodor.dato=nodo.dato
nodo.dato=None
return nodor
bst=BinarySearchTree()
bst.insert(50)
bst.insert(40)
bst.insert(20)
bst.insert(45)
bst.insert(80)
bst.insert(60)
bst.insert(90)
bst.insert(85)
bst.insert(100)
bst.insert(95)
bst.borrar(90)
print("recorrido pre:")
bst.transversal(format="preorden")
print("\nrecorrido in:")
bst.transversal(format="inorden")
```
| github_jupyter |
# Developing an AI application
Going forward, AI algorithms will be incorporated into more and more everyday applications. For example, you might want to include an image classifier in a smart phone app. To do this, you'd use a deep learning model trained on hundreds of thousands of images as part of the overall application architecture. A large part of software development in the future will be using these types of models as common parts of applications.
In this project, you'll train an image classifier to recognize different species of flowers. You can imagine using something like this in a phone app that tells you the name of the flower your camera is looking at. In practice you'd train this classifier, then export it for use in your application. We'll be using [this dataset](http://www.robots.ox.ac.uk/~vgg/data/flowers/102/index.html) of 102 flower categories, you can see a few examples below.
<img src='assets/Flowers.png' width=500px>
The project is broken down into multiple steps:
* Load and preprocess the image dataset
* Train the image classifier on your dataset
* Use the trained classifier to predict image content
We'll lead you through each part which you'll implement in Python.
When you've completed this project, you'll have an application that can be trained on any set of labeled images. Here your network will be learning about flowers and end up as a command line application. But, what you do with your new skills depends on your imagination and effort in building a dataset. For example, imagine an app where you take a picture of a car, it tells you what the make and model is, then looks up information about it. Go build your own dataset and make something new.
First up is importing the packages you'll need. It's good practice to keep all the imports at the beginning of your code. As you work through this notebook and find you need to import a package, make sure to add the import up here.
```
# Imports here
```
## Load the data
Here you'll use `torchvision` to load the data ([documentation](http://pytorch.org/docs/0.3.0/torchvision/index.html)). The data should be included alongside this notebook, otherwise you can [download it here](https://s3.amazonaws.com/content.udacity-data.com/nd089/flower_data.tar.gz). The dataset is split into three parts, training, validation, and testing. For the training, you'll want to apply transformations such as random scaling, cropping, and flipping. This will help the network generalize leading to better performance. You'll also need to make sure the input data is resized to 224x224 pixels as required by the pre-trained networks.
The validation and testing sets are used to measure the model's performance on data it hasn't seen yet. For this you don't want any scaling or rotation transformations, but you'll need to resize then crop the images to the appropriate size.
The pre-trained networks you'll use were trained on the ImageNet dataset where each color channel was normalized separately. For all three sets you'll need to normalize the means and standard deviations of the images to what the network expects. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`, calculated from the ImageNet images. These values will shift each color channel to be centered at 0 and range from -1 to 1.
```
data_dir = 'flowers'
train_dir = data_dir + '/train'
valid_dir = data_dir + '/valid'
test_dir = data_dir + '/test'
# TODO: Define your transforms for the training, validation, and testing sets
data_transforms =
# TODO: Load the datasets with ImageFolder
image_datasets =
# TODO: Using the image datasets and the trainforms, define the dataloaders
dataloaders =
```
### Label mapping
You'll also need to load in a mapping from category label to category name. You can find this in the file `cat_to_name.json`. It's a JSON object which you can read in with the [`json` module](https://docs.python.org/2/library/json.html). This will give you a dictionary mapping the integer encoded categories to the actual names of the flowers.
```
import json
with open('cat_to_name.json', 'r') as f:
cat_to_name = json.load(f)
```
# Building and training the classifier
Now that the data is ready, it's time to build and train the classifier. As usual, you should use one of the pretrained models from `torchvision.models` to get the image features. Build and train a new feed-forward classifier using those features.
We're going to leave this part up to you. If you want to talk through it with someone, chat with your fellow students! You can also ask questions on the forums or join the instructors in office hours.
Refer to [the rubric](https://review.udacity.com/#!/rubrics/1663/view) for guidance on successfully completing this section. Things you'll need to do:
* Load a [pre-trained network](http://pytorch.org/docs/master/torchvision/models.html) (If you need a starting point, the VGG networks work great and are straightforward to use)
* Define a new, untrained feed-forward network as a classifier, using ReLU activations and dropout
* Train the classifier layers using backpropagation using the pre-trained network to get the features
* Track the loss and accuracy on the validation set to determine the best hyperparameters
We've left a cell open for you below, but use as many as you need. Our advice is to break the problem up into smaller parts you can run separately. Check that each part is doing what you expect, then move on to the next. You'll likely find that as you work through each part, you'll need to go back and modify your previous code. This is totally normal!
When training make sure you're updating only the weights of the feed-forward network. You should be able to get the validation accuracy above 70% if you build everything right. Make sure to try different hyperparameters (learning rate, units in the classifier, epochs, etc) to find the best model. Save those hyperparameters to use as default values in the next part of the project.
```
# TODO: Build and train your network
```
## Testing your network
It's good practice to test your trained network on test data, images the network has never seen either in training or validation. This will give you a good estimate for the model's performance on completely new images. Run the test images through the network and measure the accuracy, the same way you did validation. You should be able to reach around 70% accuracy on the test set if the model has been trained well.
```
# TODO: Do validation on the test set
```
## Save the checkpoint
Now that your network is trained, save the model so you can load it later for making predictions. You probably want to save other things such as the mapping of classes to indices which you get from one of the image datasets: `image_datasets['train'].class_to_idx`. You can attach this to the model as an attribute which makes inference easier later on.
```model.class_to_idx = image_datasets['train'].class_to_idx```
Remember that you'll want to completely rebuild the model later so you can use it for inference. Make sure to include any information you need in the checkpoint. If you want to load the model and keep training, you'll want to save the number of epochs as well as the optimizer state, `optimizer.state_dict`. You'll likely want to use this trained model in the next part of the project, so best to save it now.
```
# TODO: Save the checkpoint
```
## Loading the checkpoint
At this point it's good to write a function that can load a checkpoint and rebuild the model. That way you can come back to this project and keep working on it without having to retrain the network.
```
# TODO: Write a function that loads a checkpoint and rebuilds the model
```
# Inference for classification
Now you'll write a function to use a trained network for inference. That is, you'll pass an image into the network and predict the class of the flower in the image. Write a function called `predict` that takes an image and a model, then returns the top $K$ most likely classes along with the probabilities. It should look like
```python
probs, classes = predict(image_path, model)
print(probs)
print(classes)
> [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]
> ['70', '3', '45', '62', '55']
```
First you'll need to handle processing the input image such that it can be used in your network.
## Image Preprocessing
You'll want to use `PIL` to load the image ([documentation](https://pillow.readthedocs.io/en/latest/reference/Image.html)). It's best to write a function that preprocesses the image so it can be used as input for the model. This function should process the images in the same manner used for training.
First, resize the images where the shortest side is 256 pixels, keeping the aspect ratio. This can be done with the [`thumbnail`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) or [`resize`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) methods. Then you'll need to crop out the center 224x224 portion of the image.
Color channels of images are typically encoded as integers 0-255, but the model expected floats 0-1. You'll need to convert the values. It's easiest with a Numpy array, which you can get from a PIL image like so `np_image = np.array(pil_image)`.
As before, the network expects the images to be normalized in a specific way. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`. You'll want to subtract the means from each color channel, then divide by the standard deviation.
And finally, PyTorch expects the color channel to be the first dimension but it's the third dimension in the PIL image and Numpy array. You can reorder dimensions using [`ndarray.transpose`](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.ndarray.transpose.html). The color channel needs to be first and retain the order of the other two dimensions.
```
def process_image(image):
''' Scales, crops, and normalizes a PIL image for a PyTorch model,
returns an Numpy array
'''
# TODO: Process a PIL image for use in a PyTorch model
```
To check your work, the function below converts a PyTorch tensor and displays it in the notebook. If your `process_image` function works, running the output through this function should return the original image (except for the cropped out portions).
```
def imshow(image, ax=None, title=None):
"""Imshow for Tensor."""
if ax is None:
fig, ax = plt.subplots()
# PyTorch tensors assume the color channel is the first dimension
# but matplotlib assumes is the third dimension
image = image.numpy().transpose((1, 2, 0))
# Undo preprocessing
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
image = std * image + mean
# Image needs to be clipped between 0 and 1 or it looks like noise when displayed
image = np.clip(image, 0, 1)
ax.imshow(image)
return ax
```
## Class Prediction
Once you can get images in the correct format, it's time to write a function for making predictions with your model. A common practice is to predict the top 5 or so (usually called top-$K$) most probable classes. You'll want to calculate the class probabilities then find the $K$ largest values.
To get the top $K$ largest values in a tensor use [`x.topk(k)`](http://pytorch.org/docs/master/torch.html#torch.topk). This method returns both the highest `k` probabilities and the indices of those probabilities corresponding to the classes. You need to convert from these indices to the actual class labels using `class_to_idx` which hopefully you added to the model or from an `ImageFolder` you used to load the data ([see here](#Save-the-checkpoint)). Make sure to invert the dictionary so you get a mapping from index to class as well.
Again, this method should take a path to an image and a model checkpoint, then return the probabilities and classes.
```python
probs, classes = predict(image_path, model)
print(probs)
print(classes)
> [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]
> ['70', '3', '45', '62', '55']
```
```
def predict(image_path, model, topk=5):
''' Predict the class (or classes) of an image using a trained deep learning model.
'''
# TODO: Implement the code to predict the class from an image file
```
## Sanity Checking
Now that you can use a trained model for predictions, check to make sure it makes sense. Even if the testing accuracy is high, it's always good to check that there aren't obvious bugs. Use `matplotlib` to plot the probabilities for the top 5 classes as a bar graph, along with the input image. It should look like this:
<img src='assets/inference_example.png' width=300px>
You can convert from the class integer encoding to actual flower names with the `cat_to_name.json` file (should have been loaded earlier in the notebook). To show a PyTorch tensor as an image, use the `imshow` function defined above.
```
# TODO: Display an image along with the top 5 classes
```
| github_jupyter |
```
%matplotlib inline
%load_ext autoreload
%autoreload 2
from __future__ import division
from __future__ import print_function
from __future__ import absolute_import
from __future__ import unicode_literals
import os
import numpy as np
import pandas as pd
from sklearn import linear_model, preprocessing, cluster
import matplotlib.pyplot as plt
import seaborn as sns
import scipy.linalg as slin
import scipy.sparse.linalg as sparselin
import scipy.sparse as sparse
import IPython
import copy
import tensorflow as tf
from tensorflow.contrib.learn.python.learn.datasets import base
from influence.inceptionModel import BinaryInceptionModel
from influence.binaryLogisticRegressionWithLBFGS import BinaryLogisticRegressionWithLBFGS
import influence.experiments as experiments
from influence.image_utils import plot_flat_bwimage, plot_flat_bwgrad, plot_flat_colorimage, plot_flat_colorgrad
from influence.dataset import DataSet
from influence.dataset_poisoning import generate_inception_features
from load_animals import load_animals, load_dogfish_with_koda
sns.set(color_codes=True)
```
# Attacking individual test images
```
num_classes = 2
num_train_ex_per_class = 900
num_test_ex_per_class = 300
dataset_name = 'dogfish_%s_%s' % (num_train_ex_per_class, num_test_ex_per_class)
image_data_sets = load_animals(
num_train_ex_per_class=num_train_ex_per_class,
num_test_ex_per_class=num_test_ex_per_class,
classes=['dog', 'fish'])
train_f = np.load('output/%s_inception_features_new_train.npz' % dataset_name)
train = DataSet(train_f['inception_features_val'], train_f['labels'])
test_f = np.load('output/%s_inception_features_new_test.npz' % dataset_name)
test = DataSet(test_f['inception_features_val'], test_f['labels'])
validation = None
data_sets = base.Datasets(train=train, validation=validation, test=test)
Y_train = image_data_sets.train.labels
Y_test = image_data_sets.test.labels
input_dim = 2048
weight_decay = 0.001
batch_size = 30
initial_learning_rate = 0.001
keep_probs = None
decay_epochs = [1000, 10000]
max_lbfgs_iter = 1000
num_classes = 2
tf.reset_default_graph()
model = BinaryLogisticRegressionWithLBFGS(
input_dim=input_dim,
weight_decay=weight_decay,
max_lbfgs_iter=max_lbfgs_iter,
num_classes=num_classes,
batch_size=batch_size,
data_sets=data_sets,
initial_learning_rate=initial_learning_rate,
keep_probs=keep_probs,
decay_epochs=decay_epochs,
mini_batch=False,
train_dir='output_ipynb',
log_dir='log',
model_name='%s_inception_onlytop' % dataset_name)
model.train()
weights = model.sess.run(model.weights)
orig_Y_train_pred = model.sess.run(model.preds, feed_dict=model.all_train_feed_dict)
orig_Y_pred = model.sess.run(model.preds, feed_dict=model.all_test_feed_dict)
num_train_attacks_needed = np.empty(len(Y_test))
num_train_attacks_needed[:] = -1
mask_orig_correct = np.zeros(len(Y_test), dtype=bool)
step_size = 0.02
weight_decay = 0.001
max_deviation = 0.5
model_name = '%s_inception_wd-%s' % (dataset_name, weight_decay)
for test_idx in range(len(Y_test)):
if orig_Y_pred[test_idx, int(Y_test[test_idx])] >= 0.5:
mask_orig_correct[test_idx] = True
else:
mask_orig_correct[test_idx] = False
filenames = [filename for filename in os.listdir('./output') if (
(('%s_attack_normal_loss_testidx-%s_trainidx-' % (model_name, test_idx)) in filename) and
(filename.endswith('stepsize-%s_proj_final.npz' % step_size)))]
assert len(filenames) <= 1
if len(filenames) == 1:
attack_f = np.load(os.path.join('output', filenames[0]))
indices_to_poison = attack_f['indices_to_poison']
num_train_attacks_needed[test_idx] = len(indices_to_poison)
poisoned_X_train_image = attack_f['poisoned_X_train_image']
for counter, idx_to_poison in enumerate(indices_to_poison):
image_diff = np.max(np.abs(image_data_sets.train.x[idx_to_poison, :] - poisoned_X_train_image[counter, :]) * 255 / 2)
assert image_diff < max_deviation + 1e-5
assert np.all(poisoned_X_train_image >= -1)
assert np.all(poisoned_X_train_image <= 1)
print('Number of test predictions flipped as the number of training images attacked increases:')
pd.Series(num_train_attacks_needed[mask_orig_correct]).value_counts()
```
| github_jupyter |
```
# Copyright 2021 NVIDIA Corporation. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
<img src="http://developer.download.nvidia.com/compute/machine-learning/frameworks/nvidia_logo.png" style="width: 90px; float: right;">
# Getting Started MovieLens: Training with TensorFlow
## Overview
We observed that TensorFlow training pipelines can be slow as the dataloader is a bottleneck. The native dataloader in TensorFlow randomly sample each item from the dataset, which is very slow. The window dataloader in TensorFlow is not much faster. In our experiments, we are able to speed-up existing TensorFlow pipelines by 9x using a highly optimized dataloader.<br><br>
Applying deep learning models to recommendation systems faces unique challenges in comparison to other domains, such as computer vision and natural language processing. The datasets and common model architectures have unique characteristics, which require custom solutions. Recommendation system datasets have terabytes in size with billion examples but each example is represented by only a few bytes. For example, the [Criteo CTR dataset](https://ailab.criteo.com/download-criteo-1tb-click-logs-dataset/), the largest publicly available dataset, is 1.3TB with 4 billion examples. The model architectures have normally large embedding tables for the users and items, which do not fit on a single GPU. You can read more in our [blogpost](https://medium.com/nvidia-merlin/why-isnt-your-recommender-system-training-faster-on-gpu-and-what-can-you-do-about-it-6cb44a711ad4).
### Learning objectives
This notebook explains, how to use the NVTabular dataloader to accelerate TensorFlow training.
1. Use **NVTabular dataloader** with TensorFlow Keras model
2. Leverage **multi-hot encoded input features**
### MovieLens25M
The [MovieLens25M](https://grouplens.org/datasets/movielens/25m/) is a popular dataset for recommender systems and is used in academic publications. The dataset contains 25M movie ratings for 62,000 movies given by 162,000 users. Many projects use only the user/item/rating information of MovieLens, but the original dataset provides metadata for the movies, as well. For example, which genres a movie has. Although we may not improve state-of-the-art results with our neural network architecture, the purpose of this notebook is to explain how to integrate multi-hot categorical features into a neural network.
## NVTabular dataloader for TensorFlow
We’ve identified that the dataloader is one bottleneck in deep learning recommender systems when training pipelines with TensorFlow. The dataloader cannot prepare the next batch fast enough and therefore, the GPU is not fully utilized.
We developed a highly customized tabular dataloader for accelerating existing pipelines in TensorFlow. In our experiments, we see a speed-up by 9x of the same training workflow with NVTabular dataloader. NVTabular dataloader’s features are:
- removing bottleneck of item-by-item dataloading
- enabling larger than memory dataset by streaming from disk
- reading data directly into GPU memory and remove CPU-GPU communication
- preparing batch asynchronously in GPU to avoid CPU-GPU communication
- supporting commonly used .parquet format
- easy integration into existing TensorFlow pipelines by using similar API - works with tf.keras models
More information in our [blogpost](https://medium.com/nvidia-merlin/training-deep-learning-based-recommender-systems-9x-faster-with-tensorflow-cc5a2572ea49).
```
# External dependencies
import os
import glob
import nvtabular as nvt
```
We define our base input directory, containing the data.
```
INPUT_DATA_DIR = os.environ.get(
"INPUT_DATA_DIR", os.path.expanduser("~/nvt-examples/movielens/data/")
)
# path to save the models
MODEL_BASE_DIR = os.environ.get("MODEL_BASE_DIR", os.path.expanduser("~/nvt-examples/"))
```
### Defining Hyperparameters
First, we define the data schema and differentiate between single-hot and multi-hot categorical features. Note, that we do not have any numerical input features.
```
BATCH_SIZE = 1024 * 32 # Batch Size
CATEGORICAL_COLUMNS = ["movieId", "userId"] # Single-hot
CATEGORICAL_MH_COLUMNS = ["genres"] # Multi-hot
NUMERIC_COLUMNS = []
# Output from ETL-with-NVTabular
TRAIN_PATHS = sorted(glob.glob(os.path.join(INPUT_DATA_DIR, "train", "*.parquet")))
VALID_PATHS = sorted(glob.glob(os.path.join(INPUT_DATA_DIR, "valid", "*.parquet")))
```
In the previous notebook, we used NVTabular for ETL and stored the workflow to disk. We can load the NVTabular workflow to extract important metadata for our training pipeline.
```
workflow = nvt.Workflow.load(os.path.join(INPUT_DATA_DIR, "workflow"))
```
The embedding table shows the cardinality of each categorical variable along with its associated embedding size. Each entry is of the form `(cardinality, embedding_size)`.
```
EMBEDDING_TABLE_SHAPES, MH_EMBEDDING_TABLE_SHAPES = nvt.ops.get_embedding_sizes(workflow)
EMBEDDING_TABLE_SHAPES.update(MH_EMBEDDING_TABLE_SHAPES)
EMBEDDING_TABLE_SHAPES
```
### Initializing NVTabular Dataloader for Tensorflow
We import TensorFlow and some NVTabular TF extensions, such as custom TensorFlow layers supporting multi-hot and the NVTabular TensorFlow data loader.
```
import os
import tensorflow as tf
# we can control how much memory to give tensorflow with this environment variable
# IMPORTANT: make sure you do this before you initialize TF's runtime, otherwise
# TF will have claimed all free GPU memory
os.environ["TF_MEMORY_ALLOCATION"] = "0.7" # fraction of free memory
from nvtabular.loader.tensorflow import KerasSequenceLoader, KerasSequenceValidater
from nvtabular.framework_utils.tensorflow import layers
```
First, we take a look on our data loader and how the data is represented as tensors. The NVTabular data loader are initialized as usually and we specify both single-hot and multi-hot categorical features as cat_names. The data loader will automatically recognize the single/multi-hot columns and represent them accordingly.
```
train_dataset_tf = KerasSequenceLoader(
TRAIN_PATHS, # you could also use a glob pattern
batch_size=BATCH_SIZE,
label_names=["rating"],
cat_names=CATEGORICAL_COLUMNS + CATEGORICAL_MH_COLUMNS,
cont_names=NUMERIC_COLUMNS,
engine="parquet",
shuffle=True,
buffer_size=0.06, # how many batches to load at once
parts_per_chunk=1,
)
valid_dataset_tf = KerasSequenceLoader(
VALID_PATHS, # you could also use a glob pattern
batch_size=BATCH_SIZE,
label_names=["rating"],
cat_names=CATEGORICAL_COLUMNS + CATEGORICAL_MH_COLUMNS,
cont_names=NUMERIC_COLUMNS,
engine="parquet",
shuffle=False,
buffer_size=0.06,
parts_per_chunk=1,
)
```
Let's generate a batch and take a look on the input features.<br><br>
We can see, that the single-hot categorical features (`userId` and `movieId`) have a shape of `(32768, 1)`, which is the batchsize (as usually).<br><br>
For the multi-hot categorical feature `genres`, we receive two Tensors `genres__values` and `genres__nnzs`.<br><br>
`genres__values` are the actual data, containing the genre IDs. Note that the Tensor has more values than the batch_size. The reason is, that one datapoint in the batch can contain more than one genre (multi-hot).<br>
`genres__nnzs` are a supporting Tensor, describing how many genres are associated with each datapoint in the batch.<br><br>
For example,
- if the first value in `genres__nnzs` is `5`, then the first 5 values in `genres__values` are associated with the first datapoint in the batch (movieId/userId).<br>
- if the second value in `genres__nnzs` is `2`, then the 6th and the 7th values in `genres__values` are associated with the second datapoint in the batch (continuing after the previous value stopped).<br>
- if the third value in `genres_nnzs` is `1`, then the 8th value in `genres__values` are associated with the third datapoint in the batch.
- and so on
```
batch = next(iter(train_dataset_tf))
batch[0]
```
We can see that the sum of `genres__nnzs` is equal to the shape of `genres__values`.
```
tf.reduce_sum(batch[0]["genres__nnzs"])
```
As each datapoint can have a different number of genres, it is more efficient to represent the genres as two flat tensors: One with the actual values (`genres__values`) and one with the length for each datapoint (`genres__nnzs`).
```
del batch
```
### Defining Neural Network Architecture
We will define a common neural network architecture for tabular data.
* Single-hot categorical features are fed into an Embedding Layer
* Each value of a multi-hot categorical features is fed into an Embedding Layer and the multiple Embedding outputs are combined via averaging
* The output of the Embedding Layers are concatenated
* The concatenated layers are fed through multiple feed-forward layers (Dense Layers with ReLU activations)
* The final output is a single number with sigmoid activation function
First, we will define some dictonary/lists for our network architecture.
```
inputs = {} # tf.keras.Input placeholders for each feature to be used
emb_layers = [] # output of all embedding layers, which will be concatenated
```
We create `tf.keras.Input` tensors for all 4 input features.
```
for col in CATEGORICAL_COLUMNS:
inputs[col] = tf.keras.Input(name=col, dtype=tf.int32, shape=(1,))
# Note that we need two input tensors for multi-hot categorical features
for col in CATEGORICAL_MH_COLUMNS:
inputs[col + "__values"] = tf.keras.Input(name=f"{col}__values", dtype=tf.int64, shape=(1,))
inputs[col + "__nnzs"] = tf.keras.Input(name=f"{col}__nnzs", dtype=tf.int64, shape=(1,))
```
Next, we initialize Embedding Layers with `tf.feature_column.embedding_column`.
```
for col in CATEGORICAL_COLUMNS + CATEGORICAL_MH_COLUMNS:
emb_layers.append(
tf.feature_column.embedding_column(
tf.feature_column.categorical_column_with_identity(
col, EMBEDDING_TABLE_SHAPES[col][0]
), # Input dimension (vocab size)
EMBEDDING_TABLE_SHAPES[col][1], # Embedding output dimension
)
)
emb_layers
```
NVTabular implemented a custom TensorFlow layer `layers.DenseFeatures`, which takes as an input the different `tf.Keras.Input` and pre-initialized `tf.feature_column` and automatically concatenate them into a flat tensor. In the case of multi-hot categorical features, `DenseFeatures` organizes the inputs `__values` and `__nnzs` to define a `RaggedTensor` and combine them. `DenseFeatures` can handle numeric inputs, as well, but MovieLens does not provide numerical input features.
```
emb_layer = layers.DenseFeatures(emb_layers)
x_emb_output = emb_layer(inputs)
x_emb_output
```
We can see that the output shape of the concatenated layer is equal to the sum of the individual Embedding output dimensions (1040 = 16+512+512).
```
EMBEDDING_TABLE_SHAPES
```
We add multiple Dense Layers. Finally, we initialize the `tf.keras.Model` and add the optimizer.
```
x = tf.keras.layers.Dense(128, activation="relu")(x_emb_output)
x = tf.keras.layers.Dense(128, activation="relu")(x)
x = tf.keras.layers.Dense(128, activation="relu")(x)
x = tf.keras.layers.Dense(1, activation="sigmoid", name="output")(x)
model = tf.keras.Model(inputs=inputs, outputs=x)
model.compile("sgd", "binary_crossentropy")
# You need to install the dependencies
tf.keras.utils.plot_model(model)
```
### Training the deep learning model
We can train our model with `model.fit`. We need to use a Callback to add the validation dataloader.
```
validation_callback = KerasSequenceValidater(valid_dataset_tf)
history = model.fit(train_dataset_tf, callbacks=[validation_callback], epochs=1)
MODEL_NAME_TF = os.environ.get("MODEL_NAME_TF", "movielens_tf")
MODEL_PATH_TEMP_TF = os.path.join(MODEL_BASE_DIR, MODEL_NAME_TF, "1/model.savedmodel")
model.save(MODEL_PATH_TEMP_TF)
```
Before moving to the next notebook, `04a-Triton-Inference-with-TF.ipynb`, we need to generate the Triton Inference Server configurations and save the models in the correct format. We just saved TensorFlow model to disk, and in the previous notebook `02-ETL-with-NVTabular`, we saved the NVTabular workflow. Let's load the workflow.
The TensorFlow input layers expect the input datatype to be int32. Therefore, we need to change the output datatypes to int32 for our NVTabular workflow.
```
workflow = nvt.Workflow.load(os.path.join(INPUT_DATA_DIR, "workflow"))
workflow.output_dtypes["userId"] = "int32"
workflow.output_dtypes["movieId"] = "int32"
MODEL_NAME_ENSEMBLE = os.environ.get("MODEL_NAME_ENSEMBLE", "movielens")
# model path to save the models
MODEL_PATH = os.environ.get("MODEL_PATH", os.path.join(MODEL_BASE_DIR, "models"))
```
NVTabular provides a function to save the NVTabular workflow, TensorFlow model and Triton Inference Server (IS) config files via `export_tensorflow_ensemble`. We provide the model, workflow, a model name for ensemble model, path and output column.
```
# Creates an ensemble triton server model, where
# model: The tensorflow model that should be served
# workflow: The nvtabular workflow used in preprocessing
# name: The base name of the various triton models
from nvtabular.inference.triton import export_tensorflow_ensemble
export_tensorflow_ensemble(model, workflow, MODEL_NAME_ENSEMBLE, MODEL_PATH, ["rating"])
```
Now, we can move to the next notebook, [04-Triton-Inference-with-TF.ipynb](https://github.com/NVIDIA/NVTabular/blob/main/examples/getting-started-movielens/04-Triton-Inference-with-TF.ipynb), to send inference request to the Triton IS.
| github_jupyter |
# Low-level functions
**This notebook is already a bit outdated. Please rather refer to the rest of the documentation in case of doubt**
First load the necessary libraries:
```
import miditapyr as mt
import mido
import pandas
import altair as alt
from rpy2.robjects import pandas2ri, r
pandas2ri.activate()
```
In order to run R, (we'll also use the R packages [pyramidi](https://github.com/urswilke/pyramidi) and the [tidyverse](https://www.tidyverse.org/)) in this jupyter notebook (via the cell magic **%%R**), the python package rpy2 needs to be used:
```
%load_ext rpy2.ipython
%%R
library(tidyverse)
library(pyramidi)
```
## Load midi file
There is a small test [midi file included in this package](https://github.com/urswilke/miditapyr/raw/master/notebooks/test_midi_file.mid):
```
mt.get_test_midi_file()
```
This file comes shipped when you install the package. You can also get the same result by specifying `as_string=True`
```
mid_file_str = mt.get_test_midi_file(as_string=True)
mid_file_str
```
and then loading it with `mido`
```
mido_mid_file = mido.MidiFile(mid_file_str)
```
Here you can listen to the midi file converted to mp3:
```
import IPython
IPython.display.Audio(url = "https://raw.githubusercontent.com/urswilke/miditapyr/master/docs/source/notebooks/test_midi_file.mp3")
```
## Midi to dataframe
Now the midi data can be loaded in a dataframe `dfc` and an integer `ticks_per_beat`:
```
dfc = mt.frame_midi(mido_mid_file)
ticks_per_beat = mido_mid_file.ticks_per_beat
dfc
ticks_per_beat
```
This dataframe `dfc` consists of 3 columns:
Column name | Meaning
----------- | -------
**i_track** | the track number
**meta** | whether the event in 'msg' is a [mido meta event](https://mido.readthedocs.io/en/latest/midi_files.html#meta-messages)
**msg** | the (meta) event information read by [mido.MidiFile()](https://mido.readthedocs.io/en/latest/midi_files.html) in a list of dictionaries
## One column per keyword
The information in the midi events in the `msg` column consists of keyword - value pairs. You can transform this information to a wide format with `mt.unnest_midi()`. Now every keyword gets its own column:
```
df = mt.unnest_midi(dfc)
df
```
## Add more time information
Now we are ready to run R code. We'll import the objects `df` and `ticks_per_beat` to R. The function `tab_measures()` [translates the time information](https://urswilke.github.io/pyramidi/reference/tab_measures.html) in the midi file (measured in ticks between events) to absolute times and also translates it to seconds (`t`), measures (`m`) and beats (`b`):
```
%%R -i df -i ticks_per_beat
dfm <- tab_measures(df, ticks_per_beat)
head(dfm)
```
Now we'll import this dataframe `dfm` back to python:
```
dfm = r['dfm']
dfm
```
As there are conversion problems of some columns we'll transform `meta` back to a boolean:
```
dfm['meta'] = dfm['meta'].astype(bool)
dfm
```
and split the dataframe in two by whether events are meta or not:
```
df_meta, df_notes = mt.split_df(dfm)
```
## Wide format
Now we'll import these back to R and [transform the data to a wide format](https://urswilke.github.io/pyramidi/reference/widen_events.html) with `widen_events()`:
```
%%R -i df_notes -i df_meta -i df
df_not_notes <- df_notes %>%
filter(!str_detect(.data$type, "^note_o[nf]f?$"))
df_notes_wide <- df_notes %>%
filter(str_detect(.data$type, "^note_o[nf]f?$")) %>%
widen_events() %>%
left_join(midi_defs)
```
## Plot in R
Now we can visualize the note events as piano roll plots for all tracks in the midi file:
```
%%R
p1 <- df_notes_wide %>%
ggplot() +
geom_segment(
aes(
x = m_note_on,
y = note_name,
xend = m_note_off,
yend = note_name,
color = velocity_note_on
)
) +
# each midi track is printed into its own facet:
facet_wrap( ~ i_track,
ncol = 1,
scales = "free_y")
p1
```
## Plot in python
We can also visualize a piano roll of the midi data in `df_notes_wide` with altair. We'll drop the name column as there were some more conversion problems:
```
df_notes_wide = r['df_notes_wide']
df_notes_wide.drop('name', axis=1)
# due to some more rpy2 conversion problems we'll only use a subset of the necessary columns:
df_subset = df_notes_wide[['m_note_on', 'm_note_off', 'velocity_note_on', 'note', 'i_track']]
p = alt.Chart(df_subset).mark_bar().encode(
x='m_note_on:T',
x2='m_note_off:T',
y='note:N',
color='velocity_note_on:Q',
tooltip=['m_note_on', 'm_note_off', 'note']
).properties(
width=200,
height=200
).facet(
facet='i_track:O',
columns=1
).resolve_scale(
y='independent'
)
p
# For an interactive version run:
# p.interactive()
```
## Back to long
Now we'll go back to R and convert the dataframe back to a long format:
```
%%R
df_notes_out <- df_notes_wide %>%
select(
c("i_track", "channel", "note", "i_note"),
matches("_note_o[nf]f?$")
) %>%
pivot_longer(
matches("_note_o[nf]f?$"),
names_to = c(".value", "type"),
names_pattern = "(.+?)_(.*)"
) %>%
mutate(meta = FALSE)
df_notes_out <-
df_notes_out %>%
full_join(df_meta) %>%
full_join(df_not_notes) %>%
arrange(i_track, ticks) %>%
group_by(i_track) %>%
mutate(time = ticks - lag(ticks) %>% {.[1] = 0; .}) %>%
ungroup()
df2 <-
df_notes_out %>%
select(names(df)) %>%
# mutate_if(is_numeric, as.integer) %>%
mutate_if(is.numeric, ~ifelse(is.na(.), NaN, .))
```
## Back to midi
```
df2 = r['df2']
df2['meta'] = df2['meta'].astype(bool)
df2.drop('name', axis=1, inplace = True)
df2
dfc2 = mt.nest_midi(df2, repair_reticulate_conversion = True)
dfc2
```
We can save the midi data back to a file:
```
mt.write_midi(dfc2,
ticks_per_beat,
"test.mid")
```
| github_jupyter |
# My Project
In addition to being a place to experiment, this project has been structured to build and serve your model in a Flask application. The purpose is to allow data science exploration to easily transition into deployed services and applications on the OpenShift platform. After saving this project to git, it can be built on the OpenShift platform to serve models.
Your dependencies will live in `requirements.txt` and your prediction function will live in `prediction.py`. As a Python based s2i application, this project can be configured and built upon to fit your needs.
### Project Organization
```
.
├── README.md
├── LICENSE
├── requirements.txt <- Used to install packages for s2i application
├── 0_start_here.ipynb <- Instructional notebook
├── 1_run_flask.ipynb <- Notebook for running flask locally to test
├── 2_test_flask.ipynb <- Notebook for testing flask requests
├── .gitignore <- standard python gitignore
├── .s2i <- hidden folder for advanced s2i configuration
│ └── environment <- s2i environment settings
├── gunicorn_config.py <- configuration for gunicorn when run in OpenShift
├── prediction.py <- the predict function called from Flask
└── wsgi.py <- basic Flask application
```
### Basic Flow
1. Install and manage dependencies in `requirements.txt`.
1. Experiment as usual.
1. Extract your prediction into the `prediction.py` file.
1. Update any dependencies.
1. Run and test your application locally.
1. Save to git.
For a complete overview, please read the [README.md](./README.md)
## Install Dependencies
```
import sys
!{sys.executable} -m pip install -r requirements.txt
```
## Experiment
Experiment with data and create your prediction function. Create any serialized models needed.
```
def predict(args_dict):
return {'prediction': 'not implemented'}
predict({'keys': 'values'})
```
## Create a Predict Function
Extract the prediction logic into a standalone python file, `prediction.py` in a `predict` function. Also, make sure `requirements.txt` is updated with any additional packages you've used and need for prediction.
```
def predict(args_dict):
return {'prediction': 'not implemented'}
```
## Test Predict Function
```
from prediction import predict
predict({'keys': 'values'})
```
### Run Flask
Run flask in a separate notebook ([1_run_flask.ipynb](./1_run_flask.ipynb)) to create a local service to try it out. You must run the application in a separate notebook since it will use the kernel until stopped.
```
!FLASK_ENV=development FLASK_APP=wsgi.py flask run
```
### Test the Flask Endpoint
Test your new service endpoint in this notebook or from a separate notebook ([2_test_flask.ipynb](./2_test_flask.ipynb)) to try it out. You can
```
!curl -X POST -H "Content-Type: application/json" --data '{"data": "hello world"}' http://localhost:5000/predictions
import requests
import json
response = requests.post('http://127.0.0.1:5000/predictions', '{"hello":"world"}')
response.json()
```
### Save Your Project to Git (and Build)
Now that you've created and tested your prediction and service endpoint, push the code up to git. This can be built as an s2i application on OpenShift.
| github_jupyter |
# What's in this exercise
Basics of how to work with Azure Cosmos DB - Cassandra API from Databricks <B>in batch</B>.<BR>
Section 07: Aggregation operations<BR>
**NOTE:**<br>
1) Server-side(Cassandra) filtering of non-partition key columns is not yet supported.<BR>
2) Server-side(Cassandra) aggregation operations are not yet supported yet<BR>
The samples below perform the same on the Spark-side<br>
### Prerequisites
The Datastax connector for Cassandra requires the Azure Comsos DB Cassandra API connection details to be initialized as part of the spark context. When you launch a Jupyter notebook, the spark session and context are already initialized and it is not advisable to stop and reinitialize the Spark context unless with every configuration set as part of the HDInsight default Jupyter notebook start-up. One workaround is to add the Cassandra instance details to Ambari, Spark2 service configuration directly. This is a one-time activity that requires a Spark2 service restart.<BR>
1. Go to Ambari, Spark2 service and click on configs
2. Then go to custom spark2-defaults and add a new property with the following, and restart Spark2 service:
spark.cassandra.connection.host=YOUR_COSMOSDB_ACCOUNT_NAME.cassandra.cosmosdb.azure.com<br>
spark.cassandra.connection.port=10350<br>
spark.cassandra.connection.ssl.enabled=true<br>
spark.cassandra.auth.username=YOUR_COSMOSDB_ACCOUNT_NAME<br>
spark.cassandra.auth.password=YOUR_COSMOSDB_KEY<br>
---------
## 1.0. Cassandra API connection
### 1.0.1. Configure dependencies
```
%%configure -f
{ "conf": {"spark.jars.packages": "com.datastax.spark:spark-cassandra-connector_2.11:2.3.0,com.microsoft.azure.cosmosdb:azure-cosmos-cassandra-spark-helper:1.0.0" }}
```
### 1.0.2. Cassandra API configuration
```
import org.apache.spark.rdd.RDD
import org.apache.spark.{SparkConf, SparkContext}
import spark.implicits._
import org.apache.spark.sql.functions._
import org.apache.spark.sql.Column
import org.apache.spark.sql.types.{StructType, StructField, StringType, IntegerType,LongType,FloatType,DoubleType, TimestampType}
import org.apache.spark.sql.cassandra._
//datastax Spark connector
import com.datastax.spark.connector._
import com.datastax.spark.connector.cql.CassandraConnector
import com.datastax.driver.core.{ConsistencyLevel, DataType}
import com.datastax.spark.connector.writer.WriteConf
//Azure Cosmos DB library for multiple retry
import com.microsoft.azure.cosmosdb.cassandra
// Specify connection factory for Cassandra
spark.conf.set("spark.cassandra.connection.factory", "com.microsoft.azure.cosmosdb.cassandra.CosmosDbConnectionFactory")
// Parallelism and throughput configs
spark.conf.set("spark.cassandra.output.batch.size.rows", "1")
spark.conf.set("spark.cassandra.connection.connections_per_executor_max", "10")
spark.conf.set("spark.cassandra.output.concurrent.writes", "100")
spark.conf.set("spark.cassandra.concurrent.reads", "512")
spark.conf.set("spark.cassandra.output.batch.grouping.buffer.size", "1000")
spark.conf.set("spark.cassandra.connection.keep_alive_ms", "60000000") //Increase this number as needed
spark.conf.set("spark.cassandra.output.ignoreNulls","true")
```
## 2.0. Data generator
```
//Delete data from prior runs
val cdbConnector = CassandraConnector(sc)
cdbConnector.withSessionDo(session => session.execute("delete from books_ks.books where book_id in ('b00300','b00001','b00023','b00501','b09999','b01001','b00999','b03999','b02999','b000009');"))
//Generate a few rows
val booksDF = Seq(
("b00001", "Arthur Conan Doyle", "A study in scarlet", 1887,11.33),
("b00023", "Arthur Conan Doyle", "A sign of four", 1890,22.45),
("b01001", "Arthur Conan Doyle", "The adventures of Sherlock Holmes", 1892,19.83),
("b00501", "Arthur Conan Doyle", "The memoirs of Sherlock Holmes", 1893,14.22),
("b00300", "Arthur Conan Doyle", "The hounds of Baskerville", 1901,12.25)
).toDF("book_id", "book_author", "book_name", "book_pub_year","book_price")
//Persist
booksDF.write.mode("append").format("org.apache.spark.sql.cassandra").options(Map( "table" -> "books", "keyspace" -> "books_ks", "output.consistency.level" -> "ALL", "ttl" -> "10000000")).save()
```
---
## 3.0. Count
### 3.0.1. RDD API
```
sc.cassandraTable("books_ks", "books").count
//count on server side - NOT SUPPORTED YET
//sc.cassandraTable("books_ks", "books").cassandraCount
```
### 3.0.2. Dataframe API
Count does not work currently for the dataframe API.<BR>
While we are pending release of count support, the sample below shows how we can execute counts currently using dataframe caching as a workaround-<br>
**Options for storage level**<br>
https://spark.apache.org/docs/2.2.0/rdd-programming-guide.html#which-storage-level-to-choose<br>
(1) MEMORY_ONLY:
Store RDD as deserialized Java objects in the JVM. If the RDD does not fit in memory, some partitions will not be cached and will be recomputed on the fly each time they're needed. This is the default level.<br>
(2) MEMORY_AND_DISK: <br>
Store RDD as deserialized Java objects in the JVM. If the RDD does not fit in memory, store the partitions that don't fit on disk, and read them from there when they're needed.<br>
(3) MEMORY_ONLY_SER: Java/Scala<br>
Store RDD as serialized Java objects (one byte array per partition). This is generally more space-efficient than deserialized objects, especially when using a fast serializer, but more CPU-intensive to read.<br>
(4) MEMORY_AND_DISK_SER: Java/Scala<br>
Similar to MEMORY_ONLY_SER, but spill partitions that don't fit in memory to disk instead of recomputing them on the fly each time they're needed.<br>
(5) DISK_ONLY: <br>
Store the RDD partitions only on disk.<br>
(6) MEMORY_ONLY_2, MEMORY_AND_DISK_2, etc. <br>
Same as the levels above, but replicate each partition on two cluster nodes.<br>
(7) OFF_HEAP (experimental):<br>
Similar to MEMORY_ONLY_SER, but store the data in off-heap memory. This requires off-heap memory to be enabled.<br>
```
//Workaround
import org.apache.spark.storage.StorageLevel
//Read from source
val readBooksDF = spark.read.cassandraFormat("books", "books_ks", "").load()
//Explain plan
readBooksDF.explain
//Materialize the dataframe
readBooksDF.persist(StorageLevel.MEMORY_ONLY)
//Subsequent execution against this DF hits the cache
readBooksDF.count
//Persist as temporary view
readBooksDF.createOrReplaceTempView("books_vw")
%%sql
--select * from books_vw
--select count(*) from books_vw where book_pub_year > 1900
--select count(book_id) from books_vw
select book_author, count(*) as count from books_vw group by book_author
```
## 4.0. Average
### 4.0.1. RDD API
```
sc.cassandraTable("books_ks", "books").select("book_price").as((c: Double) => c).mean
```
### 4.0.2. Dataframe API
```
spark.read.cassandraFormat("books", "books_ks", "").load().select("book_price").agg(avg("book_price")).show
```
### 4.0.3. SQL
```
%%sql
select min(book_price) from books_vw
```
## 5.0. Min
### 5.0.1. RDD API
```
sc.cassandraTable("books_ks", "books").select("book_price").as((c: Float) => c).min
```
### 5.0.2. Dataframe API
```
spark.read.cassandraFormat("books", "books_ks", "").load().select("book_id","book_price").agg(min("book_price")).show
```
### 5.0.3. SQL
```
%%sql
select min(book_price) from books_vw
```
## 6.0. Max
### 6.0.1. RDD API
```
sc.cassandraTable("books_ks", "books").select("book_price").as((c: Float) => c).max
```
### 6.0.2. Dataframe API
```
spark.read.cassandraFormat("books", "books_ks", "").load().select("book_price").agg(max("book_price")).show
```
### 6.0.3. SQL
```
%%sql
select max(book_price) from books_vw
```
## 7.0. Sum
### 7.0.1. RDD API
```
sc.cassandraTable("books_ks", "books").select("book_price").as((c: Float) => c).sum
```
### 7.0.2. Dataframe API
```
spark.read.cassandraFormat("books", "books_ks", "").load().select("book_price").agg(sum("book_price")).show
```
### 7.0.3. SQL
```
%%sql
select sum(book_price) from books_vw
```
## 8.0. Top or comparable
### 8.0.1. RDD API
```
val readCalcTopRDD = sc.cassandraTable("books_ks", "books").select("book_name","book_price").sortBy(_.getFloat(1), false)
readCalcTopRDD.zipWithIndex.filter(_._2 < 3).collect.foreach(println)
//delivers the first top n items without collecting the rdd to the driver.
```
### 8.0.2. Dataframe API
```
import org.apache.spark.sql.functions._
val readBooksDF = spark.read.format("org.apache.spark.sql.cassandra").options(Map( "table" -> "books", "keyspace" -> "books_ks")).load.select("book_name","book_price").orderBy(desc("book_price")).limit(3)
//Explain plan
readBooksDF.explain
//Top 3
readBooksDF.show
```
### 8.0.3. SQL
```
%%sql
select book_name,book_price from books_vw order by book_price desc limit 3
```
| github_jupyter |
```
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns #Control figure
import numpy as np
import os
from datetime import date
matplotlib.style.use('ggplot')
%matplotlib inline
from sodapy import Socrata
#MyAppToken = ''
#client = Socrata("data.cityofnewyork.us", MyAppToken)
#results = client.get("dsg6-ifza", limit=2000)
#df = pd.DataFrame.from_records(results)
cwd = os.getcwd() #to get current working directory
#print(cwd)
df = pd.read_csv('DOHMH_Childcare_Center_Inspections.csv', encoding = "L1")
df.info()
def clean_string(astr):
return astr.lower().replace('.', '') \
.replace(',', '') \
.replace(';', '') \
.replace(':', '') \
.replace('á', 'a') \
.replace('é', 'e') \
.replace('í', 'i') \
.replace('ó', 'o') \
.replace('ú', 'u') \
.replace(' ', '_') \
.replace('ñ', 'ni')
def clean_columns(df):
for series in df:
df.rename(columns={series:clean_string(series)}, inplace=True)
def execute(raw_dataset_path, clean_dataset_path):
print("\t-> Leyendo datos crudos a un DataFrame")
df = pd.read_csv('DOHMH_Childcare_Center_Inspections.csv')
print("\t-> Limpando columnas")
clean_columns(df)
df.info()
print("\t-> Reemplazando espacios en blanco")
for col in df.select_dtypes('object'):
df[col] = df[col].replace('\s+', ' ', regex=True)
print("\t-> Limpiando valores")
for col in df.select_dtypes('object'):
df[col] = df[col].str.strip()
df[col] = df[col].str.lower()
df[col] = df[col].str.replace('á', 'a')
df[col] = df[col].str.replace('é', 'e')
df[col] = df[col].str.replace('í', 'i')
df[col] = df[col].str.replace('ó', 'o')
df[col] = df[col].str.replace('ú', 'u')
df[col] = df[col].str.replace(' ', '_')
print("\t-> Cambiando NA por np.nan")
for col in df.select_dtypes('object'):
df.loc[df[col] == 'na', col] = np.nan
df.to_csv('df.csv', index=False)
print("\t-> Cuántos valores NaN tiene la base")
df.isnull().sum()
print("\t-> Eliminar duplicados")
df.duplicated().sum()
df = df.drop_duplicates()
df.shape
df.info()
```
### TABLA 3
```
tabla_3 = df.iloc[:, 0:28] #Seleccionamos sólo las columnas desde center_name hasta avg_critical_violation_rate
tabla_3.info()
tabla_3 = tabla_3.drop_duplicates()
tabla_3.shape
```
3.1
* Conservar únicamente las variables estáticas que se utilizaron en el modelo: daycareid, borough,maximum_capacity, program_type, facility_type, violation_rate_percent, total_educational_workers, public_health_hazard_violation_rate, critical_violation_rate.
```
dummies = ["program_type", "facility_type", "borough"]
df_1 = pd.get_dummies(tabla_3[dummies])
tabla_3 = tabla_3.join(df_1)
tabla_3 = tabla_3.drop(['program_type', 'facility_type', 'borough'], axis = 1)
tabla_3.info()
```
### TABLA 4
Conservar únicamente las variables que aportaban información sobre las inspecciones de la Tabla 2 (con la excepeción de borough): daycareid, inspection_date, inspection_summary, violation_category y borough.
```
df.info()
tabla_4 = df.iloc[:, [4,12,28,30,33]]
print("\t-> Reagrupar en tres variables Inspection Summary Result: reason, result_1 y result_2")
tabla_4['inspection_summary_result'] = tabla_4['inspection_summary_result'].astype('str')
df_3 = pd.DataFrame(tabla_4.inspection_summary_result.str.split('_-_',1).tolist(), columns= ['reason', 'result'])
df_3['result'] = df_3['result'].astype('str')
df_4 = pd.DataFrame(df_3.result.str.split(';_',1).tolist(), columns = ['result_1', 'result_2'])
df_3 = df_3.drop(df_3.columns[[1]], axis=1)
df_4 = df_4.join(df_3)
tabla_4 = tabla_4.join(df_4)
tabla_4 = tabla_4.drop(['inspection_summary_result'], axis = 1) #Eliminar inspection_summary_result
print("\t-> A la variable reason la hacemos dummy, es decir, initial annual inspection es 1 y en otro caso es cero")
tabla_4.reason.value_counts(dropna=False)
tabla_4['initial_annual_inspection'] = tabla_4.reason.apply(lambda x: 1 if x == "initial_annual_inspection" else 0)
tabla_4.initial_annual_inspection.value_counts(dropna=False)
tabla_4 = tabla_4.drop(['reason'], axis=1) #Eliminamos la variable reason
print("\t-> Creamos dummies a las variables result_1 y result_2")
dummies = ["result_1", "result_2"]
df_2 = pd.get_dummies(tabla_4[dummies])
tabla_4 = tabla_4.join(df_2)
tabla_4 = tabla_4.drop(['result_1', 'result_2'], axis = 1) #Eliminamos variables que no necesitamos
print("\t-> Creamos variables de año, mes y día a partir de Inspection date")
tabla_4['inspection_date'] = pd.to_datetime(tabla_4.inspection_date, format = '%m/%d/%Y')
tabla_4['inspection_year'] = tabla_4['inspection_date'].dt.year
tabla_4['inspection_month_name'] = tabla_4['inspection_date'].dt.month_name()
tabla_4['inspection_day_name'] = tabla_4['inspection_date'].dt.day_name()
print("\t-> Eliminamos días festivos, sábado y domingo ")
tabla_4 = tabla_4.drop(tabla_4.loc[tabla_4['inspection_day_name']== 'Saturday'].index)
tabla_4 = tabla_4.drop(tabla_4.loc[tabla_4['inspection_day_name']== 'Sunday'].index)
print("\t-> Poner como primer columna center_id e inspection_date")
tabla_4.rename(columns={'day_care_id':'center_id'}, inplace=True)
def order(frame,var):
varlist =[w for w in frame.columns if w not in var]
frame = frame[var+varlist]
return frame
tabla_4 = order(tabla_4,['center_id', 'inspection_date'])
print("\t-> Ordenamos la base por year, month y day en forma descendente")
tabla_4.sort_values(['inspection_date'], ascending=[False], inplace=True)
print("\t-> Creamos dummy = 1 si existió violación")
tabla_4.violation_category.value_counts(dropna=False)
tabla_4['violation'] = tabla_4['violation_category'].apply(lambda x: not pd.isnull(x))
tabla_4['violation'] = tabla_4['violation'].apply(lambda x: 1 if x == True else 0)
tabla_4.violation.value_counts(dropna=False)
print("\t-> Creamos dummy = 1 si existió violación y es un problema de salud pública")
tabla_4['public_hazard'] = tabla_4['violation_category'].apply(lambda x: 1 if x == 'public_health_hazard' else 0)
tabla_4.public_hazard.value_counts(dropna=False)
print("\t-> Creamos la variable violaciones_hist_salud_publica: Número de violaciones de salud pública históricas (2016-2019) por centro")
tabla_4['violaciones_hist_salud_publica'] = tabla_4.public_hazard[(tabla_4.inspection_year != 2020)]
df_4 = tabla_4.groupby('center_id').violaciones_hist_salud_publica.sum().reset_index()
tabla_4 = pd.merge(left=tabla_4,right=df_4, how='left', left_on='center_id', right_on='center_id')
tabla_4 = tabla_4.drop(['violaciones_hist_salud_publica_x'], axis=1) #Eliminamos la variable repetida
tabla_4.rename(columns={'violaciones_hist_salud_publica_y':'violaciones_hist_salud_publica'}, inplace=True)
print("\t-> Creamos la variable violaciones_2019_salud_publica: Número de violaciones de salud pública en el 2019 por centro")
tabla_4['violaciones_2019_salud_publica'] = tabla_4.public_hazard[(tabla_4.inspection_year == 2019)]
df_5 = tabla_4.groupby('center_id').violaciones_2019_salud_publica.sum().reset_index()
tabla_4 = pd.merge(left=tabla_4,right=df_5, how='left', left_on='center_id', right_on='center_id')
tabla_4 = tabla_4.drop(['violaciones_2019_salud_publica_x'], axis=1) #Eliminamos la variable repetida
tabla_4.rename(columns={'violaciones_2019_salud_publica_y':'violaciones_2019_salud_publica'}, inplace=True)
print("\t-> Creamos la variable violaciones_hist_criticas: Número de violaciones críticas históricas anteriores (2016-2019) por centro")
tabla_4['violation_critical'] = tabla_4['violation_category'].apply(lambda x: 1 if x == 'critical' else 0)
tabla_4['violaciones_hist_criticas'] = tabla_4.violation_critical[(tabla_4.inspection_year != 2020)]
df_6 = tabla_4.groupby('center_id').violaciones_hist_criticas.sum().reset_index()
tabla_4 = pd.merge(left=tabla_4,right=df_6, how='left', left_on='center_id', right_on='center_id')
tabla_4 = tabla_4.drop(['violaciones_hist_criticas_x'], axis=1) #Eliminamos la variable repetida
tabla_4.rename(columns={'violaciones_hist_criticas_y':'violaciones_hist_criticas'}, inplace=True)
print("\t-> Creamos la variable violaciones_2019_criticas: Número de violaciones críticas en el 2019 por centro")
tabla_4['violaciones_2019_criticas'] = tabla_4.violation_critical[(tabla_4.inspection_year == 2019)]
df_7 = tabla_4.groupby('center_id').violaciones_2019_criticas.sum().reset_index()
tabla_4 = pd.merge(left=tabla_4,right=df_7, how='left', left_on='center_id', right_on='center_id')
tabla_4 = tabla_4.drop(['violaciones_2019_criticas_x'], axis=1) #Eliminamos la variable repetida
tabla_4.rename(columns={'violaciones_2019_criticas_y':'violaciones_2019_criticas'}, inplace=True)
print("\t-> Creamos la variable ratio_violaciones_hist: Número de inspecciones en total de primera vez que resultaron en violación crítica o de salud pública/ número de inspecciones de primera vez por centro")
df_8 = tabla_4.loc[tabla_4['inspection_year'] != 2020]
df_9 = df_8[df_8.violation_category.isin(['critical', 'public_health_hazard']) & df_8['initial_annual_inspection']==1]
df_10 = df_9.groupby('center_id').initial_annual_inspection.sum().reset_index()
df_11 = tabla_4.groupby('center_id').initial_annual_inspection.sum().reset_index()
df_12 = pd.merge(left=df_11,right=df_10, how='left', left_on='center_id', right_on='center_id')
df_12['ratio_violaciones_hist'] = df_12['initial_annual_inspection_y'] / df_12['initial_annual_inspection_x']
tabla_4 = pd.merge(left=tabla_4,right=df_12, how='left', left_on='center_id', right_on='center_id')
tabla_4 = tabla_4.drop(['initial_annual_inspection_x', 'initial_annual_inspection_y'], axis=1) #Eliminamos variables que no necesitamos
print("\t-> Creamos la variable ratio_violaciones_2019: Número de inspecciones en total de primera vez que resultaron en violación crítica o de salud pública en el 2019 / número de inspecciones de primera vez por centro")
df_13 = tabla_4.loc[tabla_4['inspection_year'] == 2019]
df_14 = df_13[df_13.violation_category.isin(['critical', 'public_health_hazard']) & df_13['initial_annual_inspection']==1]
df_15 = df_14.groupby('center_id').initial_annual_inspection.sum().reset_index()
df_16 = pd.merge(left=df_11,right=df_15, how='left', left_on='center_id', right_on='center_id')
df_16['ratio_violaciones_2019'] = df_16['initial_annual_inspection_y'] / df_16['initial_annual_inspection_x']
tabla_4 = pd.merge(left=tabla_4,right=df_16, how='left', left_on='center_id', right_on='center_id')
tabla_4 = tabla_4.drop(['initial_annual_inspection_x','initial_annual_inspection_y'], axis=1) #Eliminamos variables que no necesitamos
print("\t-> Creamos la variable prom_violaciones_hist_borough: Promedio de violaciones históricas por distrito")
df_17 = tabla_4.loc[tabla_4['inspection_year'] != 2020]
df_18 = df_17.groupby('borough').violation.mean().reset_index()
tabla_4 = pd.merge(left=tabla_4,right=df_18, how='left', left_on='borough', right_on='borough')
tabla_4.rename(columns={'violation_y':'prom_violaciones_hist_borough'}, inplace=True)
tabla_4.rename(columns={'violation_x':'violation'}, inplace=True)
print("\t-> Creamos la variable prom_violaciones_2019_borough: Promedio de violaciones en el 2019 por distrito")
df_19 = tabla_4.loc[tabla_4['inspection_year'] == 2019]
df_20 = df_19.groupby('borough').violation.mean().reset_index()
tabla_4 = pd.merge(left=tabla_4,right=df_20, how='left', left_on='borough', right_on='borough')
tabla_4.rename(columns={'violation_y':'prom_violaciones_2019_borough'}, inplace=True)
tabla_4.rename(columns={'violation_x':'violation'}, inplace=True)
print("\t-> Creamos la variable ratio_violaciones_hist_sp: Número de violaciones de salud pública de primera vez por centro históricas (2017-2019)/ número de violaciones de primera vez de todo tipo por centro históricas (2017-2019) ")
df_21 = tabla_4.loc[tabla_4['inspection_year'] != 2020]
df_22 = df_21.loc[df_21['initial_annual_inspection'] == 1]
df_23 = df_22.groupby('center_id').public_hazard.sum().reset_index()
df_24 = df_22.groupby('center_id').violation.sum().reset_index()
df_25 = pd.merge(left=df_23,right=df_24, how='left', left_on='center_id', right_on='center_id')
df_25['ratio_violaciones_hist_sp'] = df_25['public_hazard'] / df_25['violation']
tabla_4 = pd.merge(left=tabla_4,right=df_25, how='left', left_on='center_id', right_on='center_id')
tabla_4 = tabla_4.drop(['public_hazard_y','violation_y'], axis=1) #Eliminamos variables que no necesitamos
tabla_4.rename(columns={'violation_x':'violation'}, inplace=True)
tabla_4.rename(columns={'public_hazard_x':'public_hazard'}, inplace=True)
print("\t-> Creamos la variable ratio_violaciones_2019_sp: Número de violaciones de salud pública de primera vez por centro en el 2019 / número de violaciones de primera vez de todo tipo por centro en el 2019 ")
df_26 = tabla_4.loc[tabla_4['inspection_year'] == 2019]
df_27 = df_26.loc[df_26['initial_annual_inspection'] == 1]
df_28 = df_27.groupby('center_id').public_hazard.sum().reset_index()
df_29 = df_27.groupby('center_id').violation.sum().reset_index()
df_30 = pd.merge(left=df_28,right=df_29, how='left', left_on='center_id', right_on='center_id')
df_30['ratio_violaciones_2019_sp'] = df_30['public_hazard'] / df_30['violation']
tabla_4 = pd.merge(left=tabla_4,right=df_30, how='left', left_on='center_id', right_on='center_id')
tabla_4 = tabla_4.drop(['public_hazard_y','violation_y'], axis=1) #Eliminamos variables que no necesitamos
tabla_4.rename(columns={'violation_x':'violation'}, inplace=True)
tabla_4.rename(columns={'public_hazard_x':'public_hazard'}, inplace=True)
print("\t-> Creamos la variable ratio_violaciones_hist_criticas: Número de violaciones críticas de primera vez por centro históricas (2017-2019)/ número de violaciones de primera vez de todo tipo por centro históricas (2017-2019)")
df_31 = tabla_4.loc[tabla_4['inspection_year'] != 2020]
df_32 = df_31.loc[df_31['initial_annual_inspection'] == 1]
df_33 = df_32.groupby('center_id').violation_critical.sum().reset_index()
df_34 = df_32.groupby('center_id').violation.sum().reset_index()
df_35 = pd.merge(left=df_33,right=df_34, how='left', left_on='center_id', right_on='center_id')
df_35['ratio_violaciones_hist_criticas'] = df_35['violation_critical'] / df_35['violation']
tabla_4 = pd.merge(left=tabla_4,right=df_35, how='left', left_on='center_id', right_on='center_id')
tabla_4 = tabla_4.drop(['violation_critical_y','violation_y'], axis=1) #Eliminamos variables que no necesitamos
tabla_4.rename(columns={'violation_x':'violation'}, inplace=True)
tabla_4.rename(columns={'violation_critical_x':'violation_critical'}, inplace=True)
print("\t-> Creamos la variable ratio_violaciones_2019_criticas: Número de violaciones críticas de primera vez por centro en el 2019/ número de violaciones de primera vez de todo tipo por centro en el 2019")
df_36 = tabla_4.loc[tabla_4['inspection_year'] == 2019]
df_37 = df_36.loc[df_36['initial_annual_inspection'] == 1]
df_38 = df_37.groupby('center_id').violation_critical.sum().reset_index()
df_39 = df_37.groupby('center_id').violation.sum().reset_index()
df_40 = pd.merge(left=df_38,right=df_39, how='left', left_on='center_id', right_on='center_id')
df_40['ratio_violaciones_2019_criticas'] = df_40['violation_critical'] / df_40['violation']
tabla_4 = pd.merge(left=tabla_4,right=df_40, how='left', left_on='center_id', right_on='center_id')
tabla_4 = tabla_4.drop(['violation_critical_y','violation_y'], axis=1) #Eliminamos variables que no necesitamos
tabla_4.rename(columns={'violation_x':'violation'}, inplace=True)
tabla_4.rename(columns={'violation_critical_x':'violation_critical'}, inplace=True)
tabla_4.info()
tabla_5 = tabla_4.join(tabla_3, lsuffix='_caller', rsuffix='_other')
tabla_5.info()
tabla_5 = tabla_5.set_index(['center_id', 'inspection_date'])
tabla_5.info()
tabla_5 = tabla_5.drop(tabla_5.columns[[0,1,14,15,31,32,33,34,35,36,37,38,39,40,42,43,44,45,46,47,49,51,53,55]], axis=1) #Eliminamos variables que no necesitamos
tabla_5.info()
tabla_5 = tabla_5.fillna(0)
```
Modelo Random Forest
- Para el entrenamiento se usaron todos los datos del 2017-2019 y para validación los datos correspondientes a lo que va del año 2020.
- Mediante una gráfica de barras se verifica si la muestra esta balanceada o no y se observa que no está balanceada.
```
sns.countplot(x='public_hazard', data=tabla_5, palette="Set3")
```
- Así que se utiliza _over-sampling_ para balancear la muestra.
```
count_class_0, count_class_1 = tabla_5.public_hazard.value_counts()
df_class_0 = tabla_5[tabla_5['public_hazard'] == 0]
df_class_1 = tabla_5[tabla_5['public_hazard'] == 1]
count_class_0
count_class_1
df_class_0_over = df_class_0.sample(count_class_1, replace=True)
df_test_over = pd.concat([df_class_1, df_class_0_over], axis=0)
print('Random over-sampling:')
print(df_test_over.public_hazard.value_counts())
df_test_over.public_hazard.value_counts().plot(kind='bar', title='Count (public_hazard)');
df_train = df_test_over.loc[df_test_over['inspection_year'] != 2020]
df_test = df_test_over.loc[df_test_over['inspection_year'] == 2020]
Y_train = df_train[['public_hazard']]
Y_test = df_test[['public_hazard']]
X_train = df_train[[i for i in df_train.keys() if i not in Y_train]]
X_test = df_test[[i for i in df_test.keys() if i not in Y_test]]
import sklearn as sk
from sklearn import preprocessing
from sklearn.ensemble import RandomForestClassifier
from sklearn import metrics
from sklearn.metrics import mean_squared_error
np.random.seed(0)
rforest = RandomForestClassifier(n_estimators=600, class_weight="balanced", max_depth=8, criterion='gini')
rforest.fit(X_train,Y_train.values.ravel())
Y_pred = rforest.predict(X_test)
print("Accuracy:",metrics.accuracy_score(Y_test, Y_pred))
print("Precision:",metrics.precision_score(Y_test, Y_pred, average='macro'))
print("Recall:",metrics.recall_score(Y_test, Y_pred, average='macro'))
rforest_matrix=metrics.confusion_matrix(Y_test,Y_pred)
pd.DataFrame(rforest_matrix)
class_names=[0,1]
fig, ax = plt.subplots()
tick_marks = np.arange(len(class_names))
plt.xticks(tick_marks, class_names)
plt.yticks(tick_marks, class_names)
sns.heatmap(pd.DataFrame(rforest_matrix), annot=True, cmap="YlGnBu" ,fmt='g')
ax.xaxis.set_label_position("top")
plt.tight_layout()
plt.title('Confusion matrix', y=1.1)
plt.ylabel('Actual label')
plt.xlabel('Predicted label')
feature_importance_frame = pd.DataFrame()
feature_importance_frame['features'] = list(X_train.keys())
feature_importance_frame['importance'] = list(rforest.feature_importances_)
feature_importance_frame = feature_importance_frame.sort_values(
'importance', ascending=False)
feature_importance_frame
```
Modelo XGBoost
```
import xgboost as xgb
from xgboost import XGBClassifier
from sklearn.metrics import accuracy_score
from collections import Counter
from sklearn.datasets import make_classification
import multiprocessing
xg_clas = xgb.XGBClassifier(n_estimators=500, max_depth=3, learning_rate=0.01, subsample=1, objective='binary:logistic', booster='gbtree', n_jobs=1, nthread=multiprocessing.cpu_count())
xg_clas.fit(X_train, Y_train)
Y_p = xg_clas.predict(X_test)
cnf_matrix = metrics.confusion_matrix(Y_test, Y_p)
pd.DataFrame(cnf_matrix)
class_names=[0,1]
fig, ax = plt.subplots()
tick_marks = np.arange(len(class_names))
plt.xticks(tick_marks, class_names)
plt.yticks(tick_marks, class_names)
sns.heatmap(pd.DataFrame(cnf_matrix), annot=True, cmap="YlGnBu" ,fmt='g')
ax.xaxis.set_label_position("top")
plt.tight_layout()
plt.title('Confusion matrix', y=1.1)
plt.ylabel('Actual label')
plt.xlabel('Predicted label')
feature_importance_frame = pd.DataFrame()
feature_importance_frame['features'] = list(X_train.keys())
feature_importance_frame['importance'] = list(xg_clas.feature_importances_)
feature_importance_frame = feature_importance_frame.sort_values(
'importance', ascending=False)
feature_importance_frame
```
| github_jupyter |
```
import tensorflow as tf
import keras
import numpy as np
import pandas as pd
nRowsRead = 194354 # specify 'None' if want to read whole file
# deliveries.csv has 150460 rows in reality, but we are only loading/previewing the first 1000 rows
df = pd.read_csv('all_matches.csv', delimiter=',', nrows = nRowsRead)
df.dataframeName = 'deliveries.csv'
nRow, nCol = df.shape
print(f'There are {nRow} rows and {nCol} columns')
df.head(10)
from mpl_toolkits.mplot3d import Axes3D
from sklearn.preprocessing import StandardScaler
import matplotlib.pyplot as plt # plotting
import os # accessing directory structure
df.info()
df.isnull().sum()
s = pd.value_counts(df['wides'])
s1 = pd.Series({'nunique': len(s), 'unique values': s.index.tolist()})
s.append(s1)
s = pd.value_counts(df['noballs'])
s1 = pd.Series({'nunique': len(s), 'unique values': s.index.tolist()})
s.append(s1)
s = pd.value_counts(df['byes'])
s1 = pd.Series({'nunique': len(s), 'unique values': s.index.tolist()})
s.append(s1)
s = pd.value_counts(df['legbyes'])
s1 = pd.Series({'nunique': len(s), 'unique values': s.index.tolist()})
s.append(s1)
s = pd.value_counts(df['penalty'])
s1 = pd.Series({'nunique': len(s), 'unique values': s.index.tolist()})
s.append(s1)
s = pd.value_counts(df['wicket_type'])
s1 = pd.Series({'nunique': len(s), 'unique values': s.index.tolist()})
s.append(s1)
#player_dismissed
s = pd.value_counts(df['player_dismissed'])
s1 = pd.Series({'nunique': len(s), 'unique values': s.index.tolist()})
s.append(s1)
#other_wicket_type
s = pd.value_counts(df['other_wicket_type'])
s1 = pd.Series({'nunique': len(s), 'unique values': s.index.tolist()})
s.append(s1)
#other_player_dismissed
s = pd.value_counts(df['other_player_dismissed'])
s1 = pd.Series({'nunique': len(s), 'unique values': s.index.tolist()})
s.append(s1)
import seaborn as sns
plt.rcParams['figure.figsize']=70,54
sns.countplot(x=df['striker'], hue='extras', data=df, linewidth=7, palette = "Set2")
plt.ylabel('Number of matches')
plt.xticks(rotation=90)
plt.xlabel('striker')
plt.title('striker V/s extras',size=50, fontweight="bold")
sns.countplot(x=df['season'], hue='extras', data=df, linewidth=7, palette = "Set2")
plt.ylabel('Number of matches')
plt.xticks(rotation=90)
plt.xlabel('year')
plt.title('Season V/s extras',size=50, fontweight="bold")
sns.countplot(x=df['season'], hue='wides', data=df, linewidth=7, palette = "Set2")
plt.ylabel('Number of matches')
plt.xticks(rotation=90)
plt.xlabel('year')
plt.title('Season V/s Wides',size=50, fontweight="bold")
sns.countplot(x=df['season'], hue='legbyes', data=df, linewidth=7, palette = "Set2")
plt.ylabel('Number of matches')
plt.xticks(rotation=90)
plt.xlabel('year')
plt.title('Season V/s legbyes',size=50, fontweight="bold")
sns.countplot(x=df['season'], hue='byes', data=df, linewidth=7, palette = "Set2")
plt.ylabel('Number of matches')
plt.xticks(rotation=90)
plt.xlabel('year')
plt.title('Season V/s byes',size=50, fontweight="bold")
```
| github_jupyter |
```
import glob
import numpy as np
from keras.layers import LSTM,Dense,Embedding,Dropout
from keras.models import Sequential
from keras.utils import np_utils
from music21 import converter,instrument,note,chord
from pickle import dump #used to save file
def get_notes():
#function created to get notes from the files dataset downloaded
notes=[]
for file in glob.glob("Piano-midi.de/train/*.mid"):
midi=converter.parse(file)
#Using that stream object we get a list of all the notes and chords in the file.
print(file)
notes_to_pass=None
parts=instrument.partitionByInstrument(midi)
if parts:#file has instrument part
notes_to_pass=parts.parts[0].recurse()
else:#file has notes in flat format
notes_to_pass=midi.flat.notes
for elements in notes_to_pass:
if isinstance(elements,note.Note):
notes.append(str(elements.pitch))
elif isinstance(elements,chord.Chord):
notes.append('.'.join(str(n) for n in elements.normalOrder))
# saving the notes we have created
with open("notes","wb") as filepath:
dump(notes,filepath)
return notes
```
# what i have done above
We start by loading each file into a Music21 stream object using the converter.parse(file) function. Using that stream object we get a list of all the notes and chords in the file. We append the pitch of every note object using its string notation since the most significant parts of the note can be recreated using the string notation of the pitch. And we append every chord by encoding the id of every note in the chord together into a single string, with each note being separated by a dot. These encodings allows us to easily decode the output generated by the network into the correct notes and chords.
# next
we have to create input sequences for the network and their respective outputs. The output for each input sequence will be the first note or chord that comes after the sequence of notes in the input sequence in our list of notes.
```
def create_seq(notes,n_vocab):
length=50
pitchname=sorted(set(pitch for pitch in notes))
#mapping each node to int
note_to_int=dict((note,number) for number,note in enumerate(pitchname))
input_seq=[]
output_seq=[]
for i in range(0,len(notes)-length,1):
in_seq=notes[i:i+length]
out_seq=notes[i+length]
input_seq.append([note_to_int[char] for char in in_seq])
output_seq.append(note_to_int[out_seq])
n_pattern=len(input_seq)
#input_seq=np.array(input_seq)
#output_seq=np.array(output_seq)
#reshaping the input to amke it compatible with lstm
input_seq=np.reshape(input_seq,(n_pattern,length,1))
#normalizing
input_seq=input_seq/float(n_vocab)
output_seq=np_utils.to_categorical(output_seq)
return (input_seq,output_seq)
notes=get_notes()
n_vocab=len(set(notes))
input_seq,output_seq=create_seq(notes,n_vocab)
#cretaing deep learning model
model=Sequential()
model.add(LSTM(256,input_shape=(input_seq.shape[1],input_seq.shape[2]),return_sequences=True))
model.add(Dropout(0.3))
model.add(LSTM(512,return_sequences=True))
model.add(Dropout(0.3))
model.add(LSTM(256))
model.add(Dense(256,activation="relu"))
model.add(Dropout(0.3))
model.add(Dense(n_vocab,activation="softmax"))
model.compile(loss="categorical_crossentropy",optimizer="rmsprop")
model.summary()
from keras.callbacks import ModelCheckpoint
filepath = "weights-improvement-{epoch:02d}-{loss:.4f}-bigger.hdf5"
checkpoint = ModelCheckpoint(filepath, monitor='loss',verbose=0,save_best_only=True,mode='min')
callbacks_list = [checkpoint]
model.fit(input_seq,output_seq, epochs=200, batch_size=64, callbacks=callbacks_list)
```
| github_jupyter |
# JAX As Accelerated NumPy
[](https://colab.research.google.com/github/google/jax/blob/main/docs/jax-101/01-jax-basics.ipynb)
*Authors: Rosalia Schneider & Vladimir Mikulik*
In this first section you will learn the very fundamentals of JAX.
## Getting started with JAX numpy
Fundamentally, JAX is a library that enables transformations of array-manipulating programs written with a NumPy-like API.
Over the course of this series of guides, we will unpack exactly what that means. For now, you can think of JAX as *differentiable NumPy that runs on accelerators*.
The code below shows how to import JAX and create a vector.
```
import jax
import jax.numpy as jnp
x = jnp.arange(10)
print(x)
```
So far, everything is just like NumPy. A big appeal of JAX is that you don't need to learn a new API. Many common NumPy programs would run just as well in JAX if you substitute `np` for `jnp`. However, there are some important differences which we touch on at the end of this section.
You can notice the first difference if you check the type of `x`. It is a variable of type `DeviceArray`, which is the way JAX represents arrays.
```
x
```
One useful feature of JAX is that the same code can be run on different backends -- CPU, GPU and TPU.
We will now perform a dot product to demonstrate that it can be done in different devices without changing the code. We use `%timeit` to check the performance.
(Technical detail: when a JAX function is called, the corresponding operation is dispatched to an accelerator to be computed asynchronously when possible. The returned array is therefore not necessarily 'filled in' as soon as the function returns. Thus, if we don't require the result immediately, the computation won't block Python execution. Therefore, unless we `block_until_ready`, we will only time the dispatch, not the actual computation. See [Asynchronous dispatch](https://jax.readthedocs.io/en/latest/async_dispatch.html#asynchronous-dispatch) in the JAX docs.)
```
long_vector = jnp.arange(int(1e7))
%timeit jnp.dot(long_vector, long_vector).block_until_ready()
```
**Tip**: Try running the code above twice, once without an accelerator, and once with a GPU runtime (while in Colab, click *Runtime* → *Change Runtime Type* and choose `GPU`). Notice how much faster it runs on a GPU.
## JAX first transformation: `grad`
A fundamental feature of JAX is that it allows you to transform functions.
One of the most commonly used transformations is `jax.grad`, which takes a numerical function written in Python and returns you a new Python function that computes the gradient of the original function.
To use it, let's first define a function that takes an array and returns the sum of squares.
```
def sum_of_squares(x):
return jnp.sum(x**2)
```
Applying `jax.grad` to `sum_of_squares` will return a different function, namely the gradient of `sum_of_squares` with respect to its first parameter `x`.
Then, you can use that function on an array to return the derivatives with respect to each element of the array.
```
sum_of_squares_dx = jax.grad(sum_of_squares)
x = jnp.asarray([1.0, 2.0, 3.0, 4.0])
print(sum_of_squares(x))
print(sum_of_squares_dx(x))
```
You can think of `jax.grad` by analogy to the $\nabla$ operator from vector calculus. Given a function $f(x)$, $\nabla f$ represents the function that computes $f$'s gradient, i.e.
$$
(\nabla f)(x)_i = \frac{\partial f}{\partial x_i}(x).
$$
Analogously, `jax.grad(f)` is the function that computes the gradient, so `jax.grad(f)(x)` is the gradient of `f` at `x`.
(Like $\nabla$, `jax.grad` will only work on functions with a scalar output -- it will raise an error otherwise.)
This makes the JAX API quite different from other autodiff libraries like Tensorflow and PyTorch, where to compute the gradient we use the loss tensor itself (e.g. by calling `loss.backward()`). The JAX API works directly with functions, staying closer to the underlying math. Once you become accustomed to this way of doing things, it feels natural: your loss function in code really is a function of parameters and data, and you find its gradient just like you would in the math.
This way of doing things makes it straightforward to control things like which variables to differentiate with respect to. By default, `jax.grad` will find the gradient with respect to the first argument. In the example below, the result of `sum_squared_error_dx` will be the gradient of `sum_squared_error` with respect to `x`.
```
def sum_squared_error(x, y):
return jnp.sum((x-y)**2)
sum_squared_error_dx = jax.grad(sum_squared_error)
y = jnp.asarray([1.1, 2.1, 3.1, 4.1])
print(sum_squared_error_dx(x, y))
```
To find the gradient with respect to a different argument (or several), you can set `argnums`:
```
jax.grad(sum_squared_error, argnums=(0, 1))(x, y) # Find gradient wrt both x & y
```
Does this mean that when doing machine learning, we need to write functions with gigantic argument lists, with an argument for each model parameter array? No. JAX comes equipped with machinery for bundling arrays together in data structures called 'pytrees', on which more in a [later guide](https://colab.research.google.com/github/google/jax/blob/main/docs/jax-101/05.1-pytrees.ipynb). So, most often, use of `jax.grad` looks like this:
```
def loss_fn(params, data):
...
grads = jax.grad(loss_fn)(params, data_batch)
```
where `params` is, for example, a nested dict of arrays, and the returned `grads` is another nested dict of arrays with the same structure.
## Value and Grad
Often, you need to find both the value and the gradient of a function, e.g. if you want to log the training loss. JAX has a handy sister transformation for efficiently doing that:
```
jax.value_and_grad(sum_squared_error)(x, y)
```
which returns a tuple of, you guessed it, (value, grad). To be precise, for any `f`,
```
jax.value_and_grad(f)(*xs) == (f(*xs), jax.grad(f)(*xs))
```
## Auxiliary data
In addition to wanting to log the value, we often want to report some intermediate results obtained in computing the loss function. But if we try doing that with regular `jax.grad`, we run into trouble:
```
def squared_error_with_aux(x, y):
return sum_squared_error(x, y), x-y
jax.grad(squared_error_with_aux)(x, y)
```
This is because `jax.grad` is only defined on scalar functions, and our new function returns a tuple. But we need to return a tuple to return our intermediate results! This is where `has_aux` comes in:
```
jax.grad(squared_error_with_aux, has_aux=True)(x, y)
```
`has_aux` signifies that the function returns a pair, `(out, aux)`. It makes `jax.grad` ignore `aux`, passing it through to the user, while differentiating the function as if only `out` was returned.
## Differences from NumPy
The `jax.numpy` API closely follows that of NumPy. However, there are some important differences. We cover many of these in future guides, but it's worth pointing some out now.
The most important difference, and in some sense the root of all the rest, is that JAX is designed to be _functional_, as in _functional programming_. The reason behind this is that the kinds of program transformations that JAX enables are much more feasible in functional-style programs.
An introduction to functional programming (FP) is out of scope of this guide. If you already are familiar with FP, you will find your FP intuition helpful while learning JAX. If not, don't worry! The important feature of functional programming to grok when working with JAX is very simple: don't write code with side-effects.
A side-effect is any effect of a function that doesn't appear in its output. One example is modifying an array in place:
```
import numpy as np
x = np.array([1, 2, 3])
def in_place_modify(x):
x[0] = 123
return None
in_place_modify(x)
x
```
The side-effectful function modifies its argument, but returns a completely unrelated value. The modification is a side-effect.
The code below will run in NumPy. However, JAX arrays won't allow themselves to be modified in-place:
```
in_place_modify(jnp.array(x)) # Raises error when we cast input to jnp.ndarray
```
Helpfully, the error points us to JAX's side-effect-free way of doing the same thing via the [`jax.numpy.ndarray.at`](https://jax.readthedocs.io/en/latest/_autosummary/jax.numpy.ndarray.at.html) index update operators (be careful [`jax.ops.index_*`](https://jax.readthedocs.io/en/latest/jax.ops.html#indexed-update-functions-deprecated) functions are deprecated). They are analogous to in-place modification by index, but create a new array with the corresponding modifications made:
```
def jax_in_place_modify(x):
return x.at[0].set(123)
y = jnp.array([1, 2, 3])
jax_in_place_modify(y)
```
Note that the old array was untouched, so there is no side-effect:
```
y
```
Side-effect-free code is sometimes called *functionally pure*, or just *pure*.
Isn't the pure version less efficient? Strictly, yes; we are creating a new array. However, as we will explain in the next guide, JAX computations are often compiled before being run using another program transformation, `jax.jit`. If we don't use the old array after modifying it 'in place' using indexed update operators, the compiler can recognise that it can in fact compile to an in-place modify, resulting in efficient code in the end.
Of course, it's possible to mix side-effectful Python code and functionally pure JAX code, and we will touch on this more later. As you get more familiar with JAX, you will learn how and when this can work. As a rule of thumb, however, any functions intended to be transformed by JAX should avoid side-effects, and the JAX primitives themselves will try to help you do that.
We will explain other places where the JAX idiosyncracies become relevant as they come up. There is even a section that focuses entirely on getting used to the functional programming style of handling state: [Part 7: Problem of State](https://colab.research.google.com/github/google/jax/blob/main/docs/jax-101/07-state.ipynb). However, if you're impatient, you can find a [summary of JAX's sharp edges](https://jax.readthedocs.io/en/latest/notebooks/Common_Gotchas_in_JAX.html) in the JAX docs.
## Your first JAX training loop
We still have much to learn about JAX, but you already know enough to understand how we can use JAX to build a simple training loop.
To keep things simple, we'll start with a linear regression.
Our data is sampled according to $y = w_{true} x + b_{true} + \epsilon$.
```
import numpy as np
import matplotlib.pyplot as plt
xs = np.random.normal(size=(100,))
noise = np.random.normal(scale=0.1, size=(100,))
ys = xs * 3 - 1 + noise
plt.scatter(xs, ys);
```
Therefore, our model is $\hat y(x; \theta) = wx + b$.
We will use a single array, `theta = [w, b]` to house both parameters:
```
def model(theta, x):
"""Computes wx + b on a batch of input x."""
w, b = theta
return w * x + b
```
The loss function is $J(x, y; \theta) = (\hat y - y)^2$.
```
def loss_fn(theta, x, y):
prediction = model(theta, x)
return jnp.mean((prediction-y)**2)
```
How do we optimize a loss function? Using gradient descent. At each update step, we will find the gradient of the loss w.r.t. the parameters, and take a small step in the direction of steepest descent:
$\theta_{new} = \theta - 0.1 (\nabla_\theta J) (x, y; \theta)$
```
def update(theta, x, y, lr=0.1):
return theta - lr * jax.grad(loss_fn)(theta, x, y)
```
In JAX, it's common to define an `update()` function that is called every step, taking the current parameters as input and returning the new parameters. This is a natural consequence of JAX's functional nature, and is explained in more detail in [The Problem of State](https://colab.research.google.com/github/google/jax/blob/main/docs/jax-101/07-state.ipynb).
This function can then be JIT-compiled in its entirety for maximum efficiency. The next guide will explain exactly how `jax.jit` works, but if you want to, you can try adding `@jax.jit` before the `update()` definition, and see how the training loop below runs much faster.
```
theta = jnp.array([1., 1.])
for _ in range(1000):
theta = update(theta, xs, ys)
plt.scatter(xs, ys)
plt.plot(xs, model(theta, xs))
w, b = theta
print(f"w: {w:<.2f}, b: {b:<.2f}")
```
As you will see going through these guides, this basic recipe underlies almost all training loops you'll see implemented in JAX. The main difference between this example and real training loops is the simplicity of our model: that allows us to use a single array to house all our parameters. We cover managing more parameters in the later [pytree guide](https://colab.research.google.com/github/google/jax/blob/main/docs/jax-101/05.1-pytrees.ipynb). Feel free to skip forward to that guide now to see how to manually define and train a simple MLP in JAX.
| github_jupyter |
<a href="https://colab.research.google.com/github/amanvishnani/CSCI-6505/blob/master/A4_E.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
**Group Members**
* Aman Vishani - B00840115
* Karthikk Tamil mani - B00838575
```
%matplotlib inline
# PyTorch packages
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
# torchvision for loading MNIST dataset
import torchvision
from torchvision import datasets, transforms
# For plotting
import matplotlib.pyplot as plt
from sklearn import decomposition
from tqdm.notebook import tqdm
from sklearn import datasets as skdatasets
import numpy as np
from torch.utils.data import DataLoader
mnist_transformer = transforms.ToTensor()
# Download the training data
train_data = datasets.MNIST('./mnist_data', download=True, train=True,
transform=mnist_transformer)
# Download the test data
test_data = datasets.MNIST('./mnist_data', download=True, train=False,
transform=mnist_transformer)
classes = [i for i in range(10)]
print("Training examples: ", len(train_data))
print("Test examples: ", len(test_data))
def create_dataloader(x, y, batch_sz=200, image=True):
if type(x) is np.ndarray:
tensor_x = torch.Tensor(x) # transform to torch tensor
else:
tensor_x = x
if type(y) is np.ndarray:
tensor_y = torch.Tensor(y)
else:
tensor_y = y
tensor_x = tensor_x.reshape(-1,1,28,28)
if image:
tensor_y = tensor_y.reshape(-1,1,28,28)
my_dataset = torch.utils.data.TensorDataset(tensor_x,tensor_y) # create your datset
my_dataloader = torch.utils.data.DataLoader(my_dataset, batch_size=batch_sz , num_workers=0) # create your dataloader
return my_dataloader
train_data_clip = np.clip(train_data.data.reshape(-1, 28*28), 0., 1.)
test_data_clip = np.clip(test_data.data.reshape(-1, 28*28), 0., 1.)
from sklearn.decomposition import PCA
pca = PCA(0.90)
# Fit and reconstruct
x_train = pca.fit_transform(train_data_clip)
x_train = pca.inverse_transform(x_train)
x_test = pca.inverse_transform(pca.transform(test_data_clip))
plt.imshow(x_train[0].reshape(28,28))
class ConvDenoiseAutoEncoder(nn.Module):
def __init__(self):
super(ConvDenoiseAutoEncoder, self).__init__()
#encoder layers
self.conv1 = nn.Conv2d(1, 64, 3, padding=1) # conv layer (depth from 1 --> 64), 3x3 kernels
self.conv2 = nn.Conv2d(64, 16, 3, padding=1) # conv layer (depth from 64 --> 16), 3x3 kernels
self.conv3 = nn.Conv2d(16, 8, 3, padding=1) # conv layer (depth from 16 --> 8), 3x3 kernels
self.pool = nn.MaxPool2d(2, 2)
#decoder layers
self.t_conv1 = nn.ConvTranspose2d(8, 8, 3, stride=2) # kernel_size=3 to get to a 7x7 image output
self.t_conv2 = nn.ConvTranspose2d(8, 16, 2, stride=2) # two more transpose layers with a kernel of 2
self.t_conv3 = nn.ConvTranspose2d(16, 64, 2, stride=2)
self.conv_out = nn.Conv2d(64, 1, 3, padding=1) # one, final, normal conv layer to decrease the depth
def encode(self, x):
x = F.relu(self.conv1(x))
x = self.pool(x)
x = F.relu(self.conv2(x))
x = self.pool(x)
x = F.relu(self.conv3(x))
x = self.pool(x)
return x
def decode(self, x):
x = F.relu(self.t_conv1(x))
x = F.relu(self.t_conv2(x))
x = F.relu(self.t_conv3(x))
x = torch.sigmoid(self.conv_out(x))
return x
def forward(self, x):
x = self.encode(x)
x = self.decode(x)
return x
def denoisingCNN(dataLoader, epochs=10):
losses = []
train_acc = 0
test_acc = 0
print("Initializing the network ...")
net = ConvDenoiseAutoEncoder()
criterion = nn.MSELoss()
optimizer = torch.optim.Adam(net.parameters(), lr=0.001)
print("Training the network ...")
losses = []
net.cuda()
for e in tqdm(range(epochs)):
train_loss = 0.0
for data in dataLoader:
inputs, labels = data
inputs = inputs.float().cuda()
labels = labels.float().cuda()
optimizer.zero_grad()
# print(inputs.min())
# print(inputs.max())
outputs = net(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
train_loss += loss.item()*inputs.size(0)
losses.append(train_loss/len(dataLoader))
train_loss = train_loss/len(dataLoader)
print('Epoch: {} \tTraining Loss: {:.6f}'.format(
e,
train_loss
))
plt.plot(losses)
plt.figure()
return net, outputs, labels, inputs
myDataLoader = create_dataloader(x_train, train_data_clip, batch_sz=20)
trained_Model, outputs, origInput , noisy_Input = denoisingCNN(myDataLoader, epochs=10)
with torch.no_grad():
test_inputs = torch.Tensor(x_test).view(-1, 1, 28, 28).cuda()
outputs = trained_Model(test_inputs)
num_examples = 4
plt.figure()
for i in range(1,num_examples+1):
plt.subplot(1, num_examples+1, i).imshow(test_inputs.cpu()[i].reshape(28,28))
plt.figure()
for i in range(1,num_examples+1):
plt.subplot(1, num_examples+1, i).imshow(outputs.cpu()[i].reshape(28,28))
```
# CNN Code
```
def eval_CNN(train_loader, test_loader, epochs, batch_size):
##### Prepare return values ##################################################
losses = []
train_acc = 0
test_acc = 0
##### Define the network #####################################################
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 6, 3) # 1 channel in, 6 filters out, 3x3 filters
self.pool = nn.MaxPool2d(2, 2) # 2x2 pooling, with a stride of 2 (move the window by 2 pixels)
self.conv2 = nn.Conv2d(6, 16, 3) # 6 filters in, 16 filters out, 3x3 filters
self.fc1 = nn.Linear(16 * 5 * 5, 120) # the 16 filtered images are reduced to 5x5 now, connect to 120 hidden units out
self.fc2 = nn.Linear(120, 84) # 120 hidden units in, 84 hidden units out
self.fc3 = nn.Linear(84, 10) # 84 hidden units in, 10 outputs units
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1, 16 * 5 * 5) # .view() is similar to .reshape(), so this flattens x into a vector
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = F.log_softmax(self.fc3(x), dim=1)
return x
##### Initialize the network and optimizer ###################################
print("Initializing the network ...")
net = Net()
criterion = nn.CrossEntropyLoss()
optimizer = optim.RMSprop(net.parameters(), lr=0.001) # lr = learning rate/step size
##### Training the network ###################################################
print("Training the network ...")
for e in tqdm(range(epochs)): # loop over the dataset multiple times
for i, data in enumerate(train_loader, 0):
inputs, labels = data
optimizer.zero_grad() # zero the parameter gradients
outputs = net(inputs) # forward pass
loss = criterion(outputs, labels) # compute loss
loss.backward() # backward pass
optimizer.step() # gradient descent update
losses.append(loss.item())
print("Epoch: ", e+1, "\t Loss:", loss.item())
##### Evaluating the network on training data ################################
print("Evaluating on training data ...")
correct = 0
total = 0
with torch.no_grad():
for data in train_loader:
# if True:
inputs, labels = data
# inputs = torch.autograd.Variable(torch.from_numpy(inputs))
# inputs = inputs.float()
outputs = net(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
train_acc = correct / total
##### Evaluating the network on training data ################################
print("Evaluating on test data ...")
correct = 0
total = 0
with torch.no_grad():
for data in test_loader:
# if True:
inputs, labels = data
# inputs = torch.autograd.Variable(torch.from_numpy(inputs))
# inputs = inputs.float()
outputs = net(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
test_acc = correct / total
return losses, train_acc, test_acc
```
# Create DataLoader
```
train_list = []
with torch.no_grad():
for data, _ in myDataLoader:
train_inputs = torch.Tensor(data).view(-1, 1, 28, 28).cuda()
train_denoise = trained_Model(train_inputs)
train_list.append(train_denoise.cpu())
train_x = torch.cat(train_list, 0)
test_inputs = torch.Tensor(x_test).view(-1, 1, 28, 28).cuda()
test_x = trained_Model(test_inputs).cpu()
print(train_x.shape)
print(test_x.shape)
```
# Train CNN on Denoised Data
```
trainLoader = create_dataloader(train_x, train_data.targets, batch_sz=100, image=False)
testLoader = create_dataloader(test_x, test_data.targets, batch_sz=100, image=False)
losses, train_acc, test_acc = eval_CNN(trainLoader, testLoader, 10, 100)
plt.plot(losses)
print("Train Acc", train_acc)
print("Train Acc", test_acc)
```
# Train on reconstructed PCA
```
x_train = train_data.data.numpy()
x_train = x_train.reshape(-1, 28*28)
x_test = test_data.data.numpy()
x_test = x_test.reshape(-1, 28*28)
def inverse_compressUsingPCA(varianceLevel):
pca = decomposition.PCA(varianceLevel)
X_pca_train = pca.fit_transform(x_train)
X_train_reconstruct = pca.inverse_transform(X_pca_train)
X_pca_test = pca.fit_transform(x_test)
X_test_reconstruct = pca.inverse_transform(X_pca_test)
return X_train_reconstruct, X_test_reconstruct
compressedTrainData, compressedTestData = inverse_compressUsingPCA(0.50)
compressedTrainData = compressedTrainData.reshape(-1, 1, 28, 28)
compressedTestData = compressedTestData.reshape(-1, 1, 28, 28)
train_loader = create_dataloader(compressedTrainData, train_data.targets, image=False)
test_loader = create_dataloader(compressedTestData, test_data.targets, image=False)
losses, train_acc, cnn_50_acc = eval_CNN(train_loader, test_loader, epochs=5, batch_size=100)
print("Training accuracy: ", train_acc)
print("Test accuracy: ", cnn_50_acc)
plt.plot(losses)
plt.title("Loss with respect to iteration in Minibatch GD")
plt.figure()
```
**Inference**
1. CNN with denoised outputs of the denoising auto-encoder performed better than the reconstructed images from PCA, around 10% improvements in the performance.
2. This was because the ouputs from the denoised auto-encoder has less noise when compared to noisy compression of the PCA reconstruction.
References:
* [Keras Denoising Autoencoder](https://keras.io/examples/mnist_denoising_autoencoder/)
* [Convolutional Denoising Autoencoder](https://medium.com/activating-robotic-minds/how-to-reduce-image-noises-by-autoencoder-65d5e6de543)
* [Denoising Autoencoder](https://github.com/udacity/deep-learning-v2-pytorch/blob/master/autoencoder/denoising-autoencoder/Denoising_Autoencoder_Solution.ipynb)
| github_jupyter |
# 2.4 ネットワークモデルの実装、2.5 順伝搬関数の実装
本ファイルでは、SSDのネットワークモデルと順伝搬forward関数を作成します。
# 2.4 学習目標
1. SSDのネットワークモデルを構築している4つのモジュールを把握する
2. SSDのネットワークモデルを作成できるようになる
3. SSDで使用する様々な大きさのデフォルトボックスの実装方法を理解する
# 2.5 学習目標
1. Non-Maximum Suppressionを理解する
2. SSDの推論時に使用するDetectクラスの順伝搬を理解する
3. SSDの順伝搬を実装できるようになる
# 事前準備
とくになし
```
# パッケージのimport
from math import sqrt
from itertools import product
import pandas as pd
import torch
from torch.autograd import Function
import torch.nn as nn
import torch.nn.functional as F
import torch.nn.init as init
```
# vggモジュールを実装
```
# 34層にわたる、vggモジュールを作成
def make_vgg():
layers = []
in_channels = 3 # 色チャネル数
# vggモジュールで使用する畳み込み層やマックスプーリングのチャネル数
cfg = [64, 64, 'M', 128, 128, 'M', 256, 256,
256, 'MC', 512, 512, 512, 'M', 512, 512, 512]
for v in cfg:
if v == 'M':
layers += [nn.MaxPool2d(kernel_size=2, stride=2)]
elif v == 'MC':
# ceilは出力サイズを、計算結果(float)に対して、切り上げで整数にするモード
# デフォルトでは出力サイズを計算結果(float)に対して、切り下げで整数にするfloorモード
layers += [nn.MaxPool2d(kernel_size=2, stride=2, ceil_mode=True)]
else:
conv2d = nn.Conv2d(in_channels, v, kernel_size=3, padding=1)
layers += [conv2d, nn.ReLU(inplace=True)]
in_channels = v
pool5 = nn.MaxPool2d(kernel_size=3, stride=1, padding=1)
conv6 = nn.Conv2d(512, 1024, kernel_size=3, padding=6, dilation=6)
conv7 = nn.Conv2d(1024, 1024, kernel_size=1)
layers += [pool5, conv6,
nn.ReLU(inplace=True), conv7, nn.ReLU(inplace=True)]
return nn.ModuleList(layers)
# 動作確認
vgg_test = make_vgg()
print(vgg_test)
```
# extrasモジュールを実装
```
# 8層にわたる、extrasモジュールを作成
def make_extras():
layers = []
in_channels = 1024 # vggモジュールから出力された、extraに入力される画像チャネル数
# extraモジュールの畳み込み層のチャネル数を設定するコンフィギュレーション
cfg = [256, 512, 128, 256, 128, 256, 128, 256]
layers += [nn.Conv2d(in_channels, cfg[0], kernel_size=(1))]
layers += [nn.Conv2d(cfg[0], cfg[1], kernel_size=(3), stride=2, padding=1)]
layers += [nn.Conv2d(cfg[1], cfg[2], kernel_size=(1))]
layers += [nn.Conv2d(cfg[2], cfg[3], kernel_size=(3), stride=2, padding=1)]
layers += [nn.Conv2d(cfg[3], cfg[4], kernel_size=(1))]
layers += [nn.Conv2d(cfg[4], cfg[5], kernel_size=(3))]
layers += [nn.Conv2d(cfg[5], cfg[6], kernel_size=(1))]
layers += [nn.Conv2d(cfg[6], cfg[7], kernel_size=(3))]
# 活性化関数のReLUは今回はSSDモデルの順伝搬のなかで用意することにし、
# extraモジュールでは用意していません
return nn.ModuleList(layers)
# 動作確認
extras_test = make_extras()
print(extras_test)
```
# locモジュールとconfモジュールを実装
```
# デフォルトボックスのオフセットを出力するloc_layers、
# デフォルトボックスに対する各クラスの信頼度confidenceを出力するconf_layersを作成
def make_loc_conf(num_classes=21, bbox_aspect_num=[4, 6, 6, 6, 4, 4]):
loc_layers = []
conf_layers = []
# VGGの22層目、conv4_3(source1)に対する畳み込み層
loc_layers += [nn.Conv2d(512, bbox_aspect_num[0]
* 4, kernel_size=3, padding=1)]
conf_layers += [nn.Conv2d(512, bbox_aspect_num[0]
* num_classes, kernel_size=3, padding=1)]
# VGGの最終層(source2)に対する畳み込み層
loc_layers += [nn.Conv2d(1024, bbox_aspect_num[1]
* 4, kernel_size=3, padding=1)]
conf_layers += [nn.Conv2d(1024, bbox_aspect_num[1]
* num_classes, kernel_size=3, padding=1)]
# extraの(source3)に対する畳み込み層
loc_layers += [nn.Conv2d(512, bbox_aspect_num[2]
* 4, kernel_size=3, padding=1)]
conf_layers += [nn.Conv2d(512, bbox_aspect_num[2]
* num_classes, kernel_size=3, padding=1)]
# extraの(source4)に対する畳み込み層
loc_layers += [nn.Conv2d(256, bbox_aspect_num[3]
* 4, kernel_size=3, padding=1)]
conf_layers += [nn.Conv2d(256, bbox_aspect_num[3]
* num_classes, kernel_size=3, padding=1)]
# extraの(source5)に対する畳み込み層
loc_layers += [nn.Conv2d(256, bbox_aspect_num[4]
* 4, kernel_size=3, padding=1)]
conf_layers += [nn.Conv2d(256, bbox_aspect_num[4]
* num_classes, kernel_size=3, padding=1)]
# extraの(source6)に対する畳み込み層
loc_layers += [nn.Conv2d(256, bbox_aspect_num[5]
* 4, kernel_size=3, padding=1)]
conf_layers += [nn.Conv2d(256, bbox_aspect_num[5]
* num_classes, kernel_size=3, padding=1)]
return nn.ModuleList(loc_layers), nn.ModuleList(conf_layers)
# 動作確認
loc_test, conf_test = make_loc_conf()
print(loc_test)
print(conf_test)
```
# L2Norm層を実装
```
# convC4_3からの出力をscale=20のL2Normで正規化する層
class L2Norm(nn.Module):
def __init__(self, input_channels=512, scale=20):
super(L2Norm, self).__init__() # 親クラスのコンストラクタ実行
self.weight = nn.Parameter(torch.Tensor(input_channels))
self.scale = scale # 係数weightの初期値として設定する値
self.reset_parameters() # パラメータの初期化
self.eps = 1e-10
def reset_parameters(self):
'''結合パラメータを大きさscaleの値にする初期化を実行'''
init.constant_(self.weight, self.scale) # weightの値がすべてscale(=20)になる
def forward(self, x):
'''38×38の特徴量に対して、512チャネルにわたって2乗和のルートを求めた
38×38個の値を使用し、各特徴量を正規化してから係数をかけ算する層'''
# 各チャネルにおける38×38個の特徴量のチャネル方向の2乗和を計算し、
# さらにルートを求め、割り算して正規化する
# normのテンソルサイズはtorch.Size([batch_num, 1, 38, 38])になります
norm = x.pow(2).sum(dim=1, keepdim=True).sqrt()+self.eps
x = torch.div(x, norm)
# 係数をかける。係数はチャネルごとに1つで、512個の係数を持つ
# self.weightのテンソルサイズはtorch.Size([512])なので
# torch.Size([batch_num, 512, 38, 38])まで変形します
weights = self.weight.unsqueeze(
0).unsqueeze(2).unsqueeze(3).expand_as(x)
out = weights * x
return out
```
# デフォルトボックスを実装
```
# デフォルトボックスを出力するクラス
class DBox(object):
def __init__(self, cfg):
super(DBox, self).__init__()
# 初期設定
self.image_size = cfg['input_size'] # 画像サイズの300
# [38, 19, …] 各sourceの特徴量マップのサイズ
self.feature_maps = cfg['feature_maps']
self.num_priors = len(cfg["feature_maps"]) # sourceの個数=6
self.steps = cfg['steps'] # [8, 16, …] DBoxのピクセルサイズ
self.min_sizes = cfg['min_sizes']
# [30, 60, …] 小さい正方形のDBoxのピクセルサイズ(正確には面積)
self.max_sizes = cfg['max_sizes']
# [60, 111, …] 大きい正方形のDBoxのピクセルサイズ(正確には面積)
self.aspect_ratios = cfg['aspect_ratios'] # 長方形のDBoxのアスペクト比
def make_dbox_list(self):
'''DBoxを作成する'''
mean = []
# 'feature_maps': [38, 19, 10, 5, 3, 1]
for k, f in enumerate(self.feature_maps):
for i, j in product(range(f), repeat=2): # fまでの数で2ペアの組み合わせを作る f_P_2 個
# 特徴量の画像サイズ
# 300 / 'steps': [8, 16, 32, 64, 100, 300],
f_k = self.image_size / self.steps[k]
# DBoxの中心座標 x,y ただし、0~1で規格化している
cx = (j + 0.5) / f_k
cy = (i + 0.5) / f_k
# アスペクト比1の小さいDBox [cx,cy, width, height]
# 'min_sizes': [30, 60, 111, 162, 213, 264]
s_k = self.min_sizes[k]/self.image_size
mean += [cx, cy, s_k, s_k]
# アスペクト比1の大きいDBox [cx,cy, width, height]
# 'max_sizes': [60, 111, 162, 213, 264, 315],
s_k_prime = sqrt(s_k * (self.max_sizes[k]/self.image_size))
mean += [cx, cy, s_k_prime, s_k_prime]
# その他のアスペクト比のdefBox [cx,cy, width, height]
for ar in self.aspect_ratios[k]:
mean += [cx, cy, s_k*sqrt(ar), s_k/sqrt(ar)]
mean += [cx, cy, s_k/sqrt(ar), s_k*sqrt(ar)]
# DBoxをテンソルに変換 torch.Size([8732, 4])
output = torch.Tensor(mean).view(-1, 4)
# DBoxが画像の外にはみ出るのを防ぐため、大きさを最小0、最大1にする
output.clamp_(max=1, min=0)
return output
# 動作の確認
# SSD300の設定
ssd_cfg = {
'num_classes': 21, # 背景クラスを含めた合計クラス数
'input_size': 300, # 画像の入力サイズ
'bbox_aspect_num': [4, 6, 6, 6, 4, 4], # 出力するDBoxのアスペクト比の種類
'feature_maps': [38, 19, 10, 5, 3, 1], # 各sourceの画像サイズ
'steps': [8, 16, 32, 64, 100, 300], # DBOXの大きさを決める
'min_sizes': [30, 60, 111, 162, 213, 264], # DBOXの大きさを決める
'max_sizes': [60, 111, 162, 213, 264, 315], # DBOXの大きさを決める
'aspect_ratios': [[2], [2, 3], [2, 3], [2, 3], [2], [2]],
}
# DBox作成
dbox = DBox(ssd_cfg)
dbox_list = dbox.make_dbox_list()
# DBoxの出力を確認する
pd.DataFrame(dbox_list.numpy())
```
# SSDクラスを実装
```
# SSDクラスを作成する
class SSD(nn.Module):
def __init__(self, phase, cfg):
super(SSD, self).__init__()
self.phase = phase # train or inferenceを指定
self.num_classes = cfg["num_classes"] # クラス数=21
# SSDのネットワークを作る
self.vgg = make_vgg()
self.extras = make_extras()
self.L2Norm = L2Norm()
self.loc, self.conf = make_loc_conf(
cfg["num_classes"], cfg["bbox_aspect_num"])
# DBox作成
dbox = DBox(cfg)
self.dbox_list = dbox.make_dbox_list()
# 推論時はクラス「Detect」を用意します
if phase == 'inference':
self.detect = Detect()
# 動作確認
ssd_test = SSD(phase="train", cfg=ssd_cfg)
print(ssd_test)
```
# ここから2.5節 順伝搬の実装です
# 関数decodeを実装する
```
# オフセット情報を使い、DBoxをBBoxに変換する関数
def decode(loc, dbox_list):
"""
オフセット情報を使い、DBoxをBBoxに変換する。
Parameters
----------
loc: [8732,4]
SSDモデルで推論するオフセット情報。
dbox_list: [8732,4]
DBoxの情報
Returns
-------
boxes : [xmin, ymin, xmax, ymax]
BBoxの情報
"""
# DBoxは[cx, cy, width, height]で格納されている
# locも[Δcx, Δcy, Δwidth, Δheight]で格納されている
# オフセット情報からBBoxを求める
boxes = torch.cat((
dbox_list[:, :2] + loc[:, :2] * 0.1 * dbox_list[:, 2:],
dbox_list[:, 2:] * torch.exp(loc[:, 2:] * 0.2)), dim=1)
# boxesのサイズはtorch.Size([8732, 4])となります
# BBoxの座標情報を[cx, cy, width, height]から[xmin, ymin, xmax, ymax] に
boxes[:, :2] -= boxes[:, 2:] / 2 # 座標(xmin,ymin)へ変換
boxes[:, 2:] += boxes[:, :2] # 座標(xmax,ymax)へ変換
return boxes
```
# Non-Maximum Suppressionを行う関数を実装する
```
# Non-Maximum Suppressionを行う関数
def nm_suppression(boxes, scores, overlap=0.45, top_k=200):
"""
Non-Maximum Suppressionを行う関数。
boxesのうち被り過ぎ(overlap以上)のBBoxを削除する。
Parameters
----------
boxes : [確信度閾値(0.01)を超えたBBox数,4]
BBox情報。
scores :[確信度閾値(0.01)を超えたBBox数]
confの情報
Returns
-------
keep : リスト
confの降順にnmsを通過したindexが格納
count:int
nmsを通過したBBoxの数
"""
# returnのひな形を作成
count = 0
keep = scores.new(scores.size(0)).zero_().long()
# keep:torch.Size([確信度閾値を超えたBBox数])、要素は全部0
# 各BBoxの面積areaを計算
x1 = boxes[:, 0]
y1 = boxes[:, 1]
x2 = boxes[:, 2]
y2 = boxes[:, 3]
area = torch.mul(x2 - x1, y2 - y1)
# boxesをコピーする。後で、BBoxの被り度合いIOUの計算に使用する際のひな形として用意
tmp_x1 = boxes.new()
tmp_y1 = boxes.new()
tmp_x2 = boxes.new()
tmp_y2 = boxes.new()
tmp_w = boxes.new()
tmp_h = boxes.new()
# socreを昇順に並び変える
v, idx = scores.sort(0)
# 上位top_k個(200個)のBBoxのindexを取り出す(200個存在しない場合もある)
idx = idx[-top_k:]
# idxの要素数が0でない限りループする
while idx.numel() > 0:
i = idx[-1] # 現在のconf最大のindexをiに
# keepの現在の最後にconf最大のindexを格納する
# このindexのBBoxと被りが大きいBBoxをこれから消去する
keep[count] = i
count += 1
# 最後のBBoxになった場合は、ループを抜ける
if idx.size(0) == 1:
break
# 現在のconf最大のindexをkeepに格納したので、idxをひとつ減らす
idx = idx[:-1]
# -------------------
# これからkeepに格納したBBoxと被りの大きいBBoxを抽出して除去する
# -------------------
# ひとつ減らしたidxまでのBBoxを、outに指定した変数として作成する
torch.index_select(x1, 0, idx, out=tmp_x1)
torch.index_select(y1, 0, idx, out=tmp_y1)
torch.index_select(x2, 0, idx, out=tmp_x2)
torch.index_select(y2, 0, idx, out=tmp_y2)
# すべてのBBoxに対して、現在のBBox=indexがiと被っている値までに設定(clamp)
tmp_x1 = torch.clamp(tmp_x1, min=x1[i])
tmp_y1 = torch.clamp(tmp_y1, min=y1[i])
tmp_x2 = torch.clamp(tmp_x2, max=x2[i])
tmp_y2 = torch.clamp(tmp_y2, max=y2[i])
# wとhのテンソルサイズをindexを1つ減らしたものにする
tmp_w.resize_as_(tmp_x2)
tmp_h.resize_as_(tmp_y2)
# clampした状態でのBBoxの幅と高さを求める
tmp_w = tmp_x2 - tmp_x1
tmp_h = tmp_y2 - tmp_y1
# 幅や高さが負になっているものは0にする
tmp_w = torch.clamp(tmp_w, min=0.0)
tmp_h = torch.clamp(tmp_h, min=0.0)
# clampされた状態での面積を求める
inter = tmp_w*tmp_h
# IoU = intersect部分 / (area(a) + area(b) - intersect部分)の計算
rem_areas = torch.index_select(area, 0, idx) # 各BBoxの元の面積
union = (rem_areas - inter) + area[i] # 2つのエリアの和(OR)の面積
IoU = inter/union
# IoUがoverlapより小さいidxのみを残す
idx = idx[IoU.le(overlap)] # leはLess than or Equal toの処理をする演算です
# IoUがoverlapより大きいidxは、最初に選んでkeepに格納したidxと同じ物体に対してBBoxを囲んでいるため消去
# whileのループが抜けたら終了
return keep, count
```
# Detectクラスを実装する
```
# SSDの推論時にconfとlocの出力から、被りを除去したBBoxを出力する
class Detect(Function):
def __init__(self, conf_thresh=0.01, top_k=200, nms_thresh=0.45):
self.softmax = nn.Softmax(dim=-1) # confをソフトマックス関数で正規化するために用意
self.conf_thresh = conf_thresh # confがconf_thresh=0.01より高いDBoxのみを扱う
self.top_k = top_k # nm_supressionでconfの高いtop_k個を計算に使用する, top_k = 200
self.nms_thresh = nms_thresh # nm_supressionでIOUがnms_thresh=0.45より大きいと、同一物体へのBBoxとみなす
def forward(self, loc_data, conf_data, dbox_list):
"""
順伝搬の計算を実行する。
Parameters
----------
loc_data: [batch_num,8732,4]
オフセット情報。
conf_data: [batch_num, 8732,num_classes]
検出の確信度。
dbox_list: [8732,4]
DBoxの情報
Returns
-------
output : torch.Size([batch_num, 21, 200, 5])
(batch_num、クラス、confのtop200、BBoxの情報)
"""
# 各サイズを取得
num_batch = loc_data.size(0) # ミニバッチのサイズ
num_dbox = loc_data.size(1) # DBoxの数 = 8732
num_classes = conf_data.size(2) # クラス数 = 21
# confはソフトマックスを適用して正規化する
conf_data = self.softmax(conf_data)
# 出力の型を作成する。テンソルサイズは[minibatch数, 21, 200, 5]
output = torch.zeros(num_batch, num_classes, self.top_k, 5)
# cof_dataを[batch_num,8732,num_classes]から[batch_num, num_classes,8732]に順番変更
conf_preds = conf_data.transpose(2, 1)
# ミニバッチごとのループ
for i in range(num_batch):
# 1. locとDBoxから修正したBBox [xmin, ymin, xmax, ymax] を求める
decoded_boxes = decode(loc_data[i], dbox_list)
# confのコピーを作成
conf_scores = conf_preds[i].clone()
# 画像クラスごとのループ(背景クラスのindexである0は計算せず、index=1から)
for cl in range(1, num_classes):
# 2.confの閾値を超えたBBoxを取り出す
# confの閾値を超えているかのマスクを作成し、
# 閾値を超えたconfのインデックスをc_maskとして取得
c_mask = conf_scores[cl].gt(self.conf_thresh)
# gtはGreater thanのこと。gtにより閾値を超えたものが1に、以下が0になる
# conf_scores:torch.Size([21, 8732])
# c_mask:torch.Size([8732])
# scoresはtorch.Size([閾値を超えたBBox数])
scores = conf_scores[cl][c_mask]
# 閾値を超えたconfがない場合、つまりscores=[]のときは、何もしない
if scores.nelement() == 0: # nelementで要素数の合計を求める
continue
# c_maskを、decoded_boxesに適用できるようにサイズを変更します
l_mask = c_mask.unsqueeze(1).expand_as(decoded_boxes)
# l_mask:torch.Size([8732, 4])
# l_maskをdecoded_boxesに適応します
boxes = decoded_boxes[l_mask].view(-1, 4)
# decoded_boxes[l_mask]で1次元になってしまうので、
# viewで(閾値を超えたBBox数, 4)サイズに変形しなおす
# 3. Non-Maximum Suppressionを実施し、被っているBBoxを取り除く
ids, count = nm_suppression(
boxes, scores, self.nms_thresh, self.top_k)
# ids:confの降順にNon-Maximum Suppressionを通過したindexが格納
# count:Non-Maximum Suppressionを通過したBBoxの数
# outputにNon-Maximum Suppressionを抜けた結果を格納
output[i, cl, :count] = torch.cat((scores[ids[:count]].unsqueeze(1),
boxes[ids[:count]]), 1)
return output # torch.Size([1, 21, 200, 5])
```
# SSDクラスを実装する
```
# SSDクラスを作成する
class SSD(nn.Module):
def __init__(self, phase, cfg):
super(SSD, self).__init__()
self.phase = phase # train or inferenceを指定
self.num_classes = cfg["num_classes"] # クラス数=21
# SSDのネットワークを作る
self.vgg = make_vgg()
self.extras = make_extras()
self.L2Norm = L2Norm()
self.loc, self.conf = make_loc_conf(
cfg["num_classes"], cfg["bbox_aspect_num"])
# DBox作成
dbox = DBox(cfg)
self.dbox_list = dbox.make_dbox_list()
# 推論時はクラス「Detect」を用意します
if phase == 'inference':
self.detect = Detect()
def forward(self, x):
sources = list() # locとconfへの入力source1~6を格納
loc = list() # locの出力を格納
conf = list() # confの出力を格納
# vggのconv4_3まで計算する
for k in range(23):
x = self.vgg[k](x)
# conv4_3の出力をL2Normに入力し、source1を作成、sourcesに追加
source1 = self.L2Norm(x)
sources.append(source1)
# vggを最後まで計算し、source2を作成、sourcesに追加
for k in range(23, len(self.vgg)):
x = self.vgg[k](x)
sources.append(x)
# extrasのconvとReLUを計算
# source3~6を、sourcesに追加
for k, v in enumerate(self.extras):
x = F.relu(v(x), inplace=True)
if k % 2 == 1: # conv→ReLU→cov→ReLUをしたらsourceに入れる
sources.append(x)
# source1~6に、それぞれ対応する畳み込みを1回ずつ適用する
# zipでforループの複数のリストの要素を取得
# source1~6まであるので、6回ループが回る
for (x, l, c) in zip(sources, self.loc, self.conf):
# Permuteは要素の順番を入れ替え
loc.append(l(x).permute(0, 2, 3, 1).contiguous())
conf.append(c(x).permute(0, 2, 3, 1).contiguous())
# l(x)とc(x)で畳み込みを実行
# l(x)とc(x)の出力サイズは[batch_num, 4*アスペクト比の種類数, featuremapの高さ, featuremap幅]
# sourceによって、アスペクト比の種類数が異なり、面倒なので順番入れ替えて整える
# permuteで要素の順番を入れ替え、
# [minibatch数, featuremap数, featuremap数,4*アスペクト比の種類数]へ
# (注釈)
# torch.contiguous()はメモリ上で要素を連続的に配置し直す命令です。
# あとでview関数を使用します。
# このviewを行うためには、対象の変数がメモリ上で連続配置されている必要があります。
# さらにlocとconfの形を変形
# locのサイズは、torch.Size([batch_num, 34928])
# confのサイズはtorch.Size([batch_num, 183372])になる
loc = torch.cat([o.view(o.size(0), -1) for o in loc], 1)
conf = torch.cat([o.view(o.size(0), -1) for o in conf], 1)
# さらにlocとconfの形を整える
# locのサイズは、torch.Size([batch_num, 8732, 4])
# confのサイズは、torch.Size([batch_num, 8732, 21])
loc = loc.view(loc.size(0), -1, 4)
conf = conf.view(conf.size(0), -1, self.num_classes)
# 最後に出力する
output = (loc, conf, self.dbox_list)
if self.phase == "inference": # 推論時
# クラス「Detect」のforwardを実行
# 返り値のサイズは torch.Size([batch_num, 21, 200, 5])
return self.detect(output[0], output[1], output[2])
else: # 学習時
return output
# 返り値は(loc, conf, dbox_list)のタプル
```
以上
| github_jupyter |
# Self-Driving Car Engineer Nanodegree
## Deep Learning
## Project: Build a Traffic Sign Recognition Classifier
In this notebook, a template is provided for you to implement your functionality in stages, which is required to successfully complete this project. If additional code is required that cannot be included in the notebook, be sure that the Python code is successfully imported and included in your submission if necessary.
> **Note**: Once you have completed all of the code implementations, you need to finalize your work by exporting the iPython Notebook as an HTML document. Before exporting the notebook to html, all of the code cells need to have been run so that reviewers can see the final implementation and output. You can then export the notebook by using the menu above and navigating to \n",
"**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
In addition to implementing code, there is a writeup to complete. The writeup should be completed in a separate file, which can be either a markdown file or a pdf document. There is a [write up template](https://github.com/udacity/CarND-Traffic-Sign-Classifier-Project/blob/master/writeup_template.md) that can be used to guide the writing process. Completing the code template and writeup template will cover all of the [rubric points](https://review.udacity.com/#!/rubrics/481/view) for this project.
The [rubric](https://review.udacity.com/#!/rubrics/481/view) contains "Stand Out Suggestions" for enhancing the project beyond the minimum requirements. The stand out suggestions are optional. If you decide to pursue the "stand out suggestions", you can include the code in this Ipython notebook and also discuss the results in the writeup file.
>**Note:** Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. In addition, Markdown cells can be edited by typically double-clicking the cell to enter edit mode.
---
## Step 0: Load The Data
First, I augment the training data set with the following operations: apply histogram equalization, apply CLAHE algorithm, rotation with random angle between -15 and 15 degrees, translation, shearing, sher followed by rotation, as well as rotation of images with increased contrast.
```
import pickle
import cv2 as cv
import numpy as np
import random
import math
# Augment the training data
def grayscale(images):
new_images = np.zeros((images.shape[0], images.shape[1], images.shape[2]) )
for i in range(images.shape[0]):
new_images[i] = cv.cvtColor(images[i], cv.COLOR_RGB2GRAY)
return new_images
def equalizeHist(images):
new_images = np.zeros((images.shape[0], images.shape[1], images.shape[2]) )
for i in range(images.shape[0]):
img = cv.cvtColor(images[i], cv.COLOR_RGB2GRAY)
new_images[i] = cv.equalizeHist(img)
return new_images
def clahe(images):
new_images = np.zeros((images.shape[0], images.shape[1], images.shape[2]) )
clahe = cv.createCLAHE(clipLimit=2.0, tileGridSize=(8,8))
for i in range(images.shape[0]):
img = cv.cvtColor(images[i], cv.COLOR_RGB2GRAY)
new_images[i] = clahe.apply(img)
return new_images
def rotation(images):
new_images = np.copy(images)
for i in range(images.shape[0]):
# generate random angle between -15 and 15 degrees
theta = random.uniform(-math.radians(15), math.radians(15))
rows,cols = images[i].shape
M = cv.getRotationMatrix2D((cols/2,rows/2),theta,1)
new_images[i] = cv.warpAffine(images[i],M,(cols,rows))
return new_images
def translation(images):
new_images = np.copy(images)
for i in range(images.shape[0]):
# generate random translation params
delta_x = random.randint(-2,2)
delta_y = random.randint(-2,2)
rows,cols = images[i].shape
M = np.float32([[1,0,delta_x],[0,1,delta_y]])
new_images[i] = cv.warpAffine(images[i],M,(cols,rows))
return new_images
def scale(images):
new_images = np.copy(images)
for i in range(images.shape[0]):
factor = random.uniform(0.9,1.1)
new_images[i] = cv.resize(images[i],None,fx=factor, fy=factor, interpolation = cv.INTER_CUBIC)
return new_images
def shear(images):
new_images = np.copy(images)
for i in range(images.shape[0]):
src = np.float32([[5,5],[5,20],[20,5]])
dst = np.float32([[5 + random.randint(-2,2), 5 + random.randint(-2,2)],
[5 + random.randint(-2,2), 20 + random.randint(-2,2)],
[20 + random.randint(-2,2), 5 + random.randint(-2,2)]])
rows,cols = images[i].shape
M = cv.getAffineTransform(src,dst)
new_images[i] = cv.warpAffine(images[i],M,(cols,rows))
return new_images
# load file
training_file = '/home/ans5k/work/CarND-Traffic-Sign-Classifier-Project/traffic-signs-data/train.p'
with open(training_file, mode='rb') as f:
train = pickle.load(f)
X_train, y_train = train['features'], train['labels']
# convert to grayscale
gray_images = grayscale(X_train)
all_images = np.copy(gray_images)
all_y = np.copy(y_train)
# improve contrast
all_images = np.concatenate((all_images, equalizeHist(X_train)))
all_y = np.concatenate((all_y, y_train))
# clahe
all_images = np.concatenate((all_images, clahe(X_train)))
all_y = np.concatenate((all_y, y_train))
# rotation
all_images = np.concatenate((all_images, rotation(gray_images)))
all_y = np.concatenate((all_y, y_train))
all_images = np.concatenate((all_images, rotation(gray_images)))
all_y = np.concatenate((all_y, y_train))
# translation
all_images = np.concatenate((all_images, translation(gray_images)))
all_y = np.concatenate((all_y, y_train))
all_images = np.concatenate((all_images, translation(gray_images)))
all_y = np.concatenate((all_y, y_train))
# shear
all_images = np.concatenate((all_images, shear(gray_images)))
all_y = np.concatenate((all_y, y_train))
all_images = np.concatenate((all_images, shear(gray_images)))
all_y = np.concatenate((all_y, y_train))
all_images = np.concatenate((all_images, rotation(shear(gray_images))))
all_y = np.concatenate((all_y, y_train))
all_images = np.concatenate((all_images, rotation(clahe(X_train))))
all_y = np.concatenate((all_y, y_train))
all_images = np.concatenate((all_images, rotation(equalizeHist(X_train))))
all_y = np.concatenate((all_y, y_train))
augmented_training_file = '/home/ans5k/work/CarND-Traffic-Sign-Classifier-Project/traffic-signs-data/augmented-train.p'
with open(augmented_training_file, mode = 'wb') as f:
pickle.dump({'features': all_images, 'labels': all_y}, f)
```
Convert to grayscale the validation and test images.
```
validation_file = 'traffic-signs-data/valid.p'
with open(validation_file, mode='rb') as f:
valid = pickle.load(f)
X_valid, y_valid = valid['features'], valid['labels']
with open('traffic-signs-data/gray-valid.p', mode = 'wb') as f:
pickle.dump({'features': grayscale(X_valid), 'labels': y_valid}, f)
testing_file = 'traffic-signs-data/test.p'
with open(testing_file, mode='rb') as f:
test = pickle.load(f)
X_test, y_test = test['features'], test['labels']
with open('traffic-signs-data/gray-test.p', mode = 'wb') as f:
pickle.dump({'features': grayscale(X_test), 'labels': y_test}, f)
```
Run the cell below to load the new pickle files with augmented and preprocessed data:
```
# Load pickled data
import numpy as np
import pickle
training_file = 'traffic-signs-data/augmented-train.p'
validation_file = 'traffic-signs-data/gray-valid.p'
testing_file = 'traffic-signs-data/gray-test.p'
with open(training_file, mode='rb') as f:
train = pickle.load(f)
with open(validation_file, mode='rb') as f:
valid = pickle.load(f)
with open(testing_file, mode='rb') as f:
test = pickle.load(f)
X_train, y_train = train['features'], train['labels']
X_valid, y_valid = valid['features'], valid['labels']
X_test, y_test = test['features'], test['labels']
X_train = np.reshape(X_train, (X_train.shape[0],X_train.shape[1],X_train.shape[2], 1))
X_valid = np.reshape(X_valid, (X_valid.shape[0],X_valid.shape[1],X_valid.shape[2], 1))
X_test = np.reshape(X_test, (X_test.shape[0],X_test.shape[1],X_test.shape[2], 1))
```
---
## Step 1: Dataset Summary & Exploration
The pickled data is a dictionary with 4 key/value pairs:
- `'features'` is a 4D array containing raw pixel data of the traffic sign images, (num examples, width, height, channels).
- `'labels'` is a 1D array containing the label/class id of the traffic sign. The file `signnames.csv` contains id -> name mappings for each id.
- `'sizes'` is a list containing tuples, (width, height) representing the original width and height the image.
- `'coords'` is a list containing tuples, (x1, y1, x2, y2) representing coordinates of a bounding box around the sign in the image. **THESE COORDINATES ASSUME THE ORIGINAL IMAGE. THE PICKLED DATA CONTAINS RESIZED VERSIONS (32 by 32) OF THESE IMAGES**
Complete the basic data summary below. Use python, numpy and/or pandas methods to calculate the data summary rather than hard coding the results. For example, the [pandas shape method](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.shape.html) might be useful for calculating some of the summary results.
### Provide a Basic Summary of the Data Set Using Python, Numpy and/or Pandas
```
### Replace each question mark with the appropriate value.
### Use python, pandas or numpy methods rather than hard coding the results
# TODO: Number of training examples
n_train = X_train.shape[0]
# TODO: Number of validation examples
n_validation = X_valid.shape[0]
# TODO: Number of testing examples.
n_test = X_test.shape[0]
# TODO: What's the shape of an traffic sign image?
image_shape = X_train.shape[1:]
# TODO: How many unique classes/labels there are in the dataset.
n_classes = max(y_train) + 1
print("Number of training examples =", n_train)
print("Number of validation examples =", n_validation)
print("Number of testing examples =", n_test)
print("Image data shape =", image_shape)
print("Number of classes =", n_classes)
```
### Include an exploratory visualization of the dataset
Visualize the German Traffic Signs Dataset using the pickled file(s). This is open ended, suggestions include: plotting traffic sign images, plotting the count of each sign, etc.
The [Matplotlib](http://matplotlib.org/) [examples](http://matplotlib.org/examples/index.html) and [gallery](http://matplotlib.org/gallery.html) pages are a great resource for doing visualizations in Python.
**NOTE:** It's recommended you start with something simple first. If you wish to do more, come back to it after you've completed the rest of the sections. It can be interesting to look at the distribution of classes in the training, validation and test set. Is the distribution the same? Are there more examples of some classes than others?
```
### Data exploration visualization code goes here.
### Feel free to use as many code cells as needed.
import matplotlib.pyplot as plt
# Visualizations will be shown in the notebook.
%matplotlib inline
import random
import numpy as np
import csv
label2Name = {}
with open('signnames.csv') as namesFile:
nameReader = csv.reader(namesFile)
for row in nameReader:
label2Name[int(row[0])] = row[1]
fig, axes = plt.subplots(5, 5, figsize=(10,10),
subplot_kw={'xticks': [], 'yticks': []})
indexes = list(random.randint(0, n_train-1) for r in range(25))
labelLimit = 25
for ax, index in zip(axes.flat, indexes):
ax.imshow(np.reshape(X_train[index],(32,32)), cmap='gray')
```
----
## Step 2: Design and Test a Model Architecture
Design and implement a deep learning model that learns to recognize traffic signs. Train and test your model on the [German Traffic Sign Dataset](http://benchmark.ini.rub.de/?section=gtsrb&subsection=dataset).
The LeNet-5 implementation shown in the [classroom](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) at the end of the CNN lesson is a solid starting point. You'll have to change the number of classes and possibly the preprocessing, but aside from that it's plug and play!
With the LeNet-5 solution from the lecture, you should expect a validation set accuracy of about 0.89. To meet specifications, the validation set accuracy will need to be at least 0.93. It is possible to get an even higher accuracy, but 0.93 is the minimum for a successful project submission.
There are various aspects to consider when thinking about this problem:
- Neural network architecture (is the network over or underfitting?)
- Play around preprocessing techniques (normalization, rgb to grayscale, etc)
- Number of examples per label (some have more than others).
- Generate fake data.
Here is an example of a [published baseline model on this problem](http://yann.lecun.com/exdb/publis/pdf/sermanet-ijcnn-11.pdf). It's not required to be familiar with the approach used in the paper but, it's good practice to try to read papers like these.
### Pre-process the Data Set (normalization, grayscale, etc.)
Minimally, the image data should be normalized so that the data has mean zero and equal variance. For image data, `(pixel - 128)/ 128` is a quick way to approximately normalize the data and can be used in this project.
Other pre-processing steps are optional. You can try different techniques to see if it improves performance.
Use the code cell (or multiple code cells, if necessary) to implement the first step of your project.
```
### Preprocess the data here. It is required to normalize the data. Other preprocessing steps could include
### converting to grayscale, etc.
### Feel free to use as many code cells as needed.
import numpy as np
import cv2
# data is already in grayscale, performing normalization
X_train = (X_train-128)/128
X_valid = (X_valid-128)/128
X_test = (X_test-128)/128
```
### Architecture
The first of the architecture I tried is based on the one presented in class. It has two convolution layers with depths 6 and 16 and max poooling with kernel size 2x2. I added dropout regularization with rate 0.8. As in the Sermanet and LeCun paper, I added the output of the first convolutional layer to the input of the first fully conected layer.
```
### Define your architecture here.
### Feel free to use as many code cells as needed.
import tensorflow as tf
import math
from tensorflow.contrib.layers import flatten
def LeNet(x, dropout):
n_classes = 43
# Arguments used for tf.truncated_normal, randomly defines variables for the weights and biases for each layer
mu = 0
sigma = 0.1
# Store layers weight & bias
wc1 = tf.Variable(tf.truncated_normal(shape=(5, 5, 1, 6), mean = mu, stddev = sigma))
wc2 = tf.Variable(tf.truncated_normal(shape=(5, 5, 6, 16), mean = mu, stddev = sigma))
wd1 = tf.Variable(tf.truncated_normal(shape=(5*5*16 + 14*14*6, 512), mean = mu, stddev = sigma))
wd2 = tf.Variable(tf.truncated_normal(shape=(512, 128), mean = mu, stddev = sigma))
wout = tf.Variable(tf.truncated_normal(shape=(128, n_classes), mean = mu, stddev = sigma))
# biases
bc1 = tf.Variable(tf.zeros(6))
bc2 = tf.Variable(tf.zeros(16))
bd1 = tf.Variable(tf.zeros(512))
bd2 = tf.Variable(tf.zeros(128))
bout = tf.Variable(tf.zeros(n_classes))
# Layer 1: Convolutional. Input = 32x32x1. Output = 28x28x6.
conv1 = tf.nn.conv2d(x, wc1, strides=[1, 1, 1, 1], padding='VALID')
conv1 = tf.nn.bias_add(conv1, bc1)
# Activation.
conv1 = tf.nn.relu(conv1)
# Pooling. Input = 28x28x6. Output = 14x14x6.
conv1 = tf.nn.max_pool(conv1, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
l1_fwd = tf.contrib.layers.flatten(conv1)
# Layer 2: Convolutional. Output = 10x10x16.
conv2 = tf.nn.conv2d(conv1, wc2, strides=[1, 1, 1, 1], padding='VALID')
conv2 = tf.nn.bias_add(conv2, bc2)
# Activation.
conv2 = tf.nn.relu(conv2)
# Pooling. Input = 10x10x16. Output = 5x5x16.
conv2 = tf.nn.max_pool(conv2, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
# Flatten. Output = 5*5*16 + 14*14*6.
conv2 = tf.concat( [tf.contrib.layers.flatten(conv2), l1_fwd], 1)
# Layer 3: Fully Connected. Input = 5*5*16 + 14*14*6. Output = 512.
fc1 = tf.add(tf.matmul(conv2, wd1), bd1)
# Activation.
dropout1 = dropout # Dropout, probability to keep units
fc1 = tf.nn.relu(fc1)
fc1 = tf.nn.dropout(fc1, dropout1)
# Layer 4: Fully Connected. Input = 512. Output = 128.
fc2 = tf.add(tf.matmul(fc1, wd2), bd2)
# TODO: Activation.
dropout2 = dropout # Dropout, probability to keep units
fc2 = tf.nn.relu(fc2)
fc2 = tf.nn.dropout(fc2, dropout2)
# TODO: Layer 5: Fully Connected. Input = 128. Output = n_classes
logits = tf.add(tf.matmul(fc2, wout), bout)
return logits
```
A second architecture is based on the Sermanet and LeCun paper. The depth of the first convolutional layer is 108 and the one of the second is 200.
```
import tensorflow as tf
import math
from tensorflow.contrib.layers import flatten
def LeNet2(x, dropout):
# Arguments used for tf.truncated_normal, randomly defines variables for the weights and biases for each layer
mu = 0
sigma = 0.1
n_classes = 43
#weights
wc1 = tf.Variable(tf.truncated_normal(shape=(5, 5, 1, 108), mean = mu, stddev = sigma))
wc2 = tf.Variable(tf.truncated_normal(shape=(5, 5, 108, 200), mean = mu, stddev = sigma))
wd1 = tf.Variable(tf.truncated_normal(shape=(5*5*200 + 14*14*108, 1024), mean = mu, stddev = sigma))
wd2 = tf.Variable(tf.truncated_normal(shape=(1024, 128), mean = mu, stddev = sigma))
wout =tf.Variable(tf.truncated_normal(shape=(128, n_classes), mean = mu, stddev = sigma))
#biases
bc1 = tf.Variable(tf.zeros(108))
bc2 = tf.Variable(tf.zeros(200))
bd1 = tf.Variable(tf.zeros(1024))
bd2 = tf.Variable(tf.zeros(128))
bout = tf.Variable(tf.zeros(n_classes))
# Layer 1: Convolutional. Input = 32x32x1. Output = 28x28x108.
conv1 = tf.nn.conv2d(x, wc1, strides=[1, 1, 1, 1], padding='VALID')
conv1 = tf.nn.bias_add(conv1, bc1)
# Activation.
conv1 = tf.nn.relu(conv1)
# Pooling. Input = 28x28x108. Output = 14x14x108.
conv1 = tf.nn.max_pool(conv1, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
# Layer 2: Convolutional. Output = 10x10x200.
conv2 = tf.nn.conv2d(conv1, wc2, strides=[1, 1, 1, 1], padding='VALID')
conv2 = tf.nn.bias_add(conv2, bc2)
# Activation.
conv2 = tf.nn.relu(conv2)
# Pooling. Input = 10x10x200. Output = 5x5x200.
conv2 = tf.nn.max_pool(conv2, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
# TODO: Flatten. Input = 5x5x200 and 14x14x108..
conv2 = tf.concat([tf.contrib.layers.flatten(conv2),tf.contrib.layers.flatten(conv1)], axis=1)
# TODO: Layer 3: Fully Connected. Input = . Output = 1024.
fc1 = tf.add(tf.matmul(conv2, wd1), bd1)
# Activation.
dropout1 = dropout # Dropout, probability to keep units
fc1 = tf.nn.relu(fc1)
fc1 = tf.nn.dropout(fc1, dropout1)
# Layer 4: Fully Connected. Input = 1024. Output = 128.
fc2 = tf.add(tf.matmul(fc1, wd2), bd2)
# TODO: Activation.
dropout2 = dropout # Dropout, probability to keep units
fc2 = tf.nn.relu(fc2)
fc2 = tf.nn.dropout(fc2, dropout2)
# TODO: Layer 5: Fully Connected. Input = 128. Output = n_classes
logits = tf.add(tf.matmul(fc2, wout), bout)
return logits
def evaluate(X_data, y_data):
num_examples = len(X_data)
total_accuracy = 0
sess = tf.get_default_session()
for offset in range(0, num_examples, BATCH_SIZE):
batch_x, batch_y = X_data[offset:offset+BATCH_SIZE], y_data[offset:offset+BATCH_SIZE]
accuracy = sess.run(accuracy_operation, feed_dict={x: batch_x, y: batch_y})
total_accuracy += (accuracy * len(batch_x))
return total_accuracy / num_examples
```
### Train, Validate and Test the Model
A validation set can be used to assess how well the model is performing. A low accuracy on the training and validation
sets imply underfitting. A high accuracy on the training set but low accuracy on the validation set implies overfitting.
```
### Train your model here.
### Calculate and report the accuracy on the training and validation set.
### Once a final model architecture is selected,
### the accuracy on the test set should be calculated and reported as well.
### Feel free to use as many code cells as needed.
# Shuffle the training data.
from sklearn.utils import shuffle
def run(logits, filename):
EPOCHS = 40
BATCH_SIZE = 256
x = tf.placeholder(tf.float32, (None, 32, 32, 1))
y = tf.placeholder(tf.int32, (None))
one_hot_y = tf.one_hot(y, n_classes)
rate = 0.001
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(labels=one_hot_y, logits=logits)
loss_operation = tf.reduce_mean(cross_entropy)
optimizer = tf.train.AdamOptimizer(learning_rate = rate)
training_operation = optimizer.minimize(loss_operation)
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(one_hot_y, 1))
accuracy_operation = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
saver = tf.train.Saver()
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
num_examples = len(X_train)
print("Training...")
print()
for i in range(EPOCHS):
X_train, y_train = shuffle(X_train, y_train)
for offset in range(0, num_examples, BATCH_SIZE):
end = offset + BATCH_SIZE
batch_x, batch_y = X_train[offset:end], y_train[offset:end]
sess.run(training_operation, feed_dict={x: batch_x, y: batch_y})
print("EPOCH {} ...".format(i+1))
train_accuracy = evaluate(X_train, y_train)
print("Train Accuracy = {:.3f}".format(train_accuracy))
validation_accuracy = evaluate(X_valid, y_valid)
print("Validation Accuracy = {:.3f}".format(validation_accuracy))
print()
test_accuracy = evaluate(X_test, y_test)
print("Test Accuracy = {:.3f}".format(test_accuracy))
saver.save(sess, filename)
print("Model saved")
run(LeNet(x, 0.5), './lenet')
```
Here is an excerpt of the output of the first model training:
.......
EPOCH 37 ...
Train Accuracy = 0.983
Validation Accuracy = 0.959
EPOCH 38 ...
Train Accuracy = 0.986
Validation Accuracy = 0.967
EPOCH 39 ...
Train Accuracy = 0.987
Validation Accuracy = 0.967
EPOCH 40 ...
Train Accuracy = 0.987
Validation Accuracy = 0.969
Test Accuracy = 0.957
```
run(LeNet2(x, 0.5), './lenet2')
```
Here is an excerpt of the output of the second model training:
.....
EPOCH 37 ...
Train Accuracy = 0.998
Validation Accuracy = 0.975
EPOCH 38 ...
Train Accuracy = 0.997
Validation Accuracy = 0.971
EPOCH 39 ...
Train Accuracy = 0.998
Validation Accuracy = 0.976
EPOCH 40 ...
Train Accuracy = 0.998
Validation Accuracy = 0.966
Test Accuracy = 0.965
---
## Step 3: Test a Model on New Images
To give yourself more insight into how your model is working, download at least five pictures of German traffic signs from the web and use your model to predict the traffic sign type.
You may find `signnames.csv` useful as it contains mappings from the class id (integer) to the actual sign name.
### Load and Output the Images
I used the images from another Udacity project: https://github.com/darienmt/CarND-TrafficSignClassifier-P2/tree/master/webimages.
```
### Load the images and plot them here.
### Feel free to use as many code cells as needed.
### Data exploration visualization code goes here.
### Feel free to use as many code cells as needed.
import matplotlib.pyplot as plt
# Visualizations will be shown in the notebook.
%matplotlib inline
import glob
names = glob.glob('test-images/*.jpg')
images = [ plt.imread('./' + name ) for name in names ]
fig, axes = plt.subplots(1, len(images), figsize=(10,10),
subplot_kw={'xticks': [], 'yticks': []})
indexes = range(5)
for ax, index in zip(axes.flat, indexes):
ax.imshow(images[index])
# convert to grayscale
X_data = grayscale(np.array(images))
fig, axes = plt.subplots(1, len(X_data), figsize=(10,10),
subplot_kw={'xticks': [], 'yticks': []})
indexes = range(5)
for ax, index in zip(axes.flat, indexes):
ax.imshow(X_data[index], cmap = 'gray')
X_data = np.reshape(X_data, (X_data.shape[0],X_data.shape[1],X_data.shape[2], 1))
X_data = (X_data-128)/128
```
### Predict the Sign Type for Each Image
```
EPOCHS = 40
BATCH_SIZE = 256
x = tf.placeholder(tf.float32, (None, 32, 32, 1))
y = tf.placeholder(tf.int32, (None))
one_hot_y = tf.one_hot(y, n_classes)
logits = LeNet2(x, 1.0)
rate = 0.001
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(labels=one_hot_y, logits=logits)
loss_operation = tf.reduce_mean(cross_entropy)
optimizer = tf.train.AdamOptimizer(learning_rate = rate)
training_operation = optimizer.minimize(loss_operation)
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(one_hot_y, 1))
accuracy_operation = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
### Run the predictions here and use the model to output the prediction for each image.
### Make sure to pre-process the images with the same pre-processing pipeline used earlier.
### Feel free to use as many code cells as needed.
# load saved model
import tensorflow as tf
import numpy as np
import csv
label2Name = {}
with open('signnames.csv') as namesFile:
nameReader = csv.reader(namesFile)
for row in nameReader:
label2Name[int(row[0])] = row[1]
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
saver2 = tf.train.import_meta_graph('./lenet2.meta')
saver2.restore(sess, './lenet2')
y = sess.run(logits, feed_dict={x: X_data})
labels = sess.run(tf.argmax(y,1))
for i in range(len(labels)):
print('Label for file', names[i], 'is', label2Name[labels[i]])
```
The accuracy of the first architecture is 80%. The output is:
Label for file ../test-images/road_work.jpg is General caution
Label for file ../test-images/60_kmh.jpg is Speed limit (60km/h)
Label for file ../test-images/left_turn.jpg is Turn left ahead
Label for file ../test-images/yield_sign.jpg is Yield
Label for file ../test-images/stop_sign.jpg is Stop
The accuracy of the second architecture is 80%. The output is:
Label for file ../test-images/road_work.jpg is General caution
Label for file ../test-images/60_kmh.jpg is Speed limit (60km/h)
Label for file ../test-images/left_turn.jpg is Turn left ahead
Label for file ../test-images/yield_sign.jpg is Yield
Label for file ../test-images/stop_sign.jpg is Stop
### Analyze Performance
```
### Calculate the accuracy for these 5 new images.
### For example, if the model predicted 1 out of 5 signs correctly, it's 20% accurate on these new images.
```
### Output Top 5 Softmax Probabilities For Each Image Found on the Web
For each of the new images, print out the model's softmax probabilities to show the **certainty** of the model's predictions (limit the output to the top 5 probabilities for each image). [`tf.nn.top_k`](https://www.tensorflow.org/versions/r0.12/api_docs/python/nn.html#top_k) could prove helpful here.
The example below demonstrates how tf.nn.top_k can be used to find the top k predictions for each image.
`tf.nn.top_k` will return the values and indices (class ids) of the top k predictions. So if k=3, for each sign, it'll return the 3 largest probabilities (out of a possible 43) and the correspoding class ids.
Take this numpy array as an example. The values in the array represent predictions. The array contains softmax probabilities for five candidate images with six possible classes. `tf.nn.top_k` is used to choose the three classes with the highest probability:
```
# (5, 6) array
a = np.array([[ 0.24879643, 0.07032244, 0.12641572, 0.34763842, 0.07893497,
0.12789202],
[ 0.28086119, 0.27569815, 0.08594638, 0.0178669 , 0.18063401,
0.15899337],
[ 0.26076848, 0.23664738, 0.08020603, 0.07001922, 0.1134371 ,
0.23892179],
[ 0.11943333, 0.29198961, 0.02605103, 0.26234032, 0.1351348 ,
0.16505091],
[ 0.09561176, 0.34396535, 0.0643941 , 0.16240774, 0.24206137,
0.09155967]])
```
Running it through `sess.run(tf.nn.top_k(tf.constant(a), k=3))` produces:
```
TopKV2(values=array([[ 0.34763842, 0.24879643, 0.12789202],
[ 0.28086119, 0.27569815, 0.18063401],
[ 0.26076848, 0.23892179, 0.23664738],
[ 0.29198961, 0.26234032, 0.16505091],
[ 0.34396535, 0.24206137, 0.16240774]]), indices=array([[3, 0, 5],
[0, 1, 4],
[0, 5, 1],
[1, 3, 5],
[1, 4, 3]], dtype=int32))
```
Looking just at the first row we get `[ 0.34763842, 0.24879643, 0.12789202]`, you can confirm these are the 3 largest probabilities in `a`. You'll also notice `[3, 0, 5]` are the corresponding indices.
```
### Print out the top five softmax probabilities for the predictions on the German traffic sign images found on the web.
### Feel free to use as many code cells as needed.
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
saver2 = tf.train.import_meta_graph('./lenet.meta')
saver2.restore(sess, './lenet')
y = sess.run(logits, feed_dict={x: X_data})
top_prob = sess.run(tf.nn.top_k(tf.nn.softmax(y),3))
for i in range(len(labels)):
print('For file', names[i], ' top 3 are:')
for j in range(3):
print('%2.4f' % (top_prob[0][i][j]), 'for', label2Name[top_prob[1][i][j]])
```
For file ../test-images/road_work.jpg top 3 are:
0.9994 for General caution
0.0004 for Traffic signals
0.0002 for Pedestrians
For file ../test-images/60_kmh.jpg top 3 are:
1.0000 for Speed limit (60km/h)
0.0000 for Speed limit (80km/h)
0.0000 for No passing for vehicles over 3.5 metric tons
For file ../test-images/left_turn.jpg top 3 are:
1.0000 for Turn left ahead
0.0000 for Keep right
0.0000 for No vehicles
For file ../test-images/yield_sign.jpg top 3 are:
1.0000 for Yield
0.0000 for Speed limit (20km/h)
0.0000 for Speed limit (30km/h)
For file ../test-images/stop_sign.jpg top 3 are:
1.0000 for Stop
0.0000 for Turn left ahead
0.0000 for Speed limit (60km/h)
```
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
saver2 = tf.train.import_meta_graph('./lenet2.meta')
saver2.restore(sess, './lenet2')
y = sess.run(logits, feed_dict={x: X_data})
top_prob = sess.run(tf.nn.top_k(tf.nn.softmax(y),3))
for i in range(len(labels)):
print('For file', names[i], ' top 3 are:')
for j in range(3):
print('%2.4f' % (top_prob[0][i][j]), 'for', label2Name[top_prob[1][i][j]])
```
For file ../test-images/road_work.jpg top 3 are:
1.0000 for General caution
0.0000 for Traffic signals
0.0000 for Wild animals crossing
For file ../test-images/60_kmh.jpg top 3 are:
1.0000 for Speed limit (60km/h)
0.0000 for Speed limit (20km/h)
0.0000 for Speed limit (30km/h)
For file ../test-images/left_turn.jpg top 3 are:
1.0000 for Turn left ahead
0.0000 for Keep right
0.0000 for Stop
For file ../test-images/yield_sign.jpg top 3 are:
1.0000 for Yield
0.0000 for Speed limit (20km/h)
0.0000 for Speed limit (30km/h)
For file ../test-images/stop_sign.jpg top 3 are:
1.0000 for Stop
0.0000 for Speed limit (60km/h)
0.0000 for Keep right
### Project Writeup
Once you have completed the code implementation, document your results in a project writeup using this [template](https://github.com/udacity/CarND-Traffic-Sign-Classifier-Project/blob/master/writeup_template.md) as a guide. The writeup can be in a markdown or pdf file.
> **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to \n",
"**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
---
## Step 4 (Optional): Visualize the Neural Network's State with Test Images
This Section is not required to complete but acts as an additional excersise for understaning the output of a neural network's weights. While neural networks can be a great learning device they are often referred to as a black box. We can understand what the weights of a neural network look like better by plotting their feature maps. After successfully training your neural network you can see what it's feature maps look like by plotting the output of the network's weight layers in response to a test stimuli image. From these plotted feature maps, it's possible to see what characteristics of an image the network finds interesting. For a sign, maybe the inner network feature maps react with high activation to the sign's boundary outline or to the contrast in the sign's painted symbol.
Provided for you below is the function code that allows you to get the visualization output of any tensorflow weight layer you want. The inputs to the function should be a stimuli image, one used during training or a new one you provided, and then the tensorflow variable name that represents the layer's state during the training process, for instance if you wanted to see what the [LeNet lab's](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) feature maps looked like for it's second convolutional layer you could enter conv2 as the tf_activation variable.
For an example of what feature map outputs look like, check out NVIDIA's results in their paper [End-to-End Deep Learning for Self-Driving Cars](https://devblogs.nvidia.com/parallelforall/deep-learning-self-driving-cars/) in the section Visualization of internal CNN State. NVIDIA was able to show that their network's inner weights had high activations to road boundary lines by comparing feature maps from an image with a clear path to one without. Try experimenting with a similar test to show that your trained network's weights are looking for interesting features, whether it's looking at differences in feature maps from images with or without a sign, or even what feature maps look like in a trained network vs a completely untrained one on the same sign image.
<figure>
<img src="visualize_cnn.png" width="380" alt="Combined Image" />
<figcaption>
<p></p>
<p style="text-align: center;"> Your output should look something like this (above)</p>
</figcaption>
</figure>
<p></p>
```
### Visualize your network's feature maps here.
### Feel free to use as many code cells as needed.
# image_input: the test image being fed into the network to produce the feature maps
# tf_activation: should be a tf variable name used during your training procedure that represents the calculated state of a specific weight layer
# activation_min/max: can be used to view the activation contrast in more detail, by default matplot sets min and max to the actual min and max values of the output
# plt_num: used to plot out multiple different weight feature map sets on the same block, just extend the plt number for each new feature map entry
def outputFeatureMap(image_input, tf_activation, activation_min=-1, activation_max=-1 ,plt_num=1):
# Here make sure to preprocess your image_input in a way your network expects
# with size, normalization, ect if needed
# image_input =
# Note: x should be the same name as your network's tensorflow data placeholder variable
# If you get an error tf_activation is not defined it may be having trouble accessing the variable from inside a function
activation = tf_activation.eval(session=sess,feed_dict={x : image_input})
featuremaps = activation.shape[3]
plt.figure(plt_num, figsize=(15,15))
for featuremap in range(featuremaps):
plt.subplot(6,8, featuremap+1) # sets the number of feature maps to show on each row and column
plt.title('FeatureMap ' + str(featuremap)) # displays the feature map number
if activation_min != -1 & activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin =activation_min, vmax=activation_max, cmap="gray")
elif activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmax=activation_max, cmap="gray")
elif activation_min !=-1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin=activation_min, cmap="gray")
else:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", cmap="gray")
```
| github_jupyter |
```
from src import pysetperm as psp
import numpy as np
import pandas as pd
n_perms = 30000
cores = 6
# +-2kb of gene definition: range_modification=2000
gene_def_plus=2000
# can set minimum size of the candidate gene set.
min_size=10
annotations = psp.AnnotationSet(annotation_file='data/genes.txt', range_modification=gene_def_plus)
function_sets = psp.FunctionSets(function_set_file='data/vip.txt', min_set_size=min_size, annotation_obj=annotations)
# specific inputs
e_candidates = psp.Variants(variant_file='data/eastern-3.5e-05-candidate.snps.bed.gz')
e_candidates.annotate_variants(annotation_obj=annotations)
e_background = psp.Variants(variant_file='data/pbsnj-bg.snps.bed.gz')
e_background.annotate_variants(annotation_obj=annotations)
# central can use eastern background.
c_candidates = psp.Variants(variant_file='data/central-4e-05-candidate.snps.bed.gz')
c_candidates.annotate_variants(annotation_obj=annotations)
i_candidates = psp.Variants(variant_file='data/ancestral-0.001-candidate.snps.bed.gz')
i_candidates.annotate_variants(annotation_obj=annotations)
i_background = psp.Variants(variant_file='data/ancestral-bg.bed.gz')
i_background.annotate_variants(annotation_obj=annotations)
i_background.variants
# test objects
e_test_obj = psp.TestObject(e_candidates,
e_background,
function_sets,
n_cores=cores)
c_test_obj = psp.TestObject(c_candidates,
e_background,
function_sets,
n_cores=cores)
i_test_obj = psp.TestObject(i_candidates,
i_background,
function_sets,
n_cores=cores)
e_permutations = psp.Permutation(e_test_obj, n_perms, cores)
c_permutations = psp.Permutation(c_test_obj, n_perms, cores)
i_permutations = psp.Permutation(i_test_obj, n_perms, cores)
# distributions across permutations
e_per_set = psp.SetPerPerm(e_permutations,
function_sets,
e_test_obj,
cores)
c_per_set = psp.SetPerPerm(c_permutations,
function_sets,
c_test_obj,
cores)
i_per_set = psp.SetPerPerm(i_permutations,
function_sets,
i_test_obj,
cores)
# results tables
def make_results_table(test_obj, function_set_obj, set_perm_obj):
out = function_set_obj.function_sets.groupby('Id', as_index=False).agg({'FunctionName': pd.Series.unique})
out = out[out['Id'].isin(function_set_obj.function_array2d_ids)]
out['n_candidates'] = test_obj.n_candidate_per_function
out['mean_n_resample'] = set_perm_obj.mean_per_set
out['emp_p_e'] = set_perm_obj.p_enrichment
out['emp_p_d'] = set_perm_obj.p_depletion
out['fdr_e'] = psp.fdr_from_p_matrix(set_perm_obj.set_n_per_perm, out['emp_p_e'], method='enrichment')
out['fdr_d'] = psp.fdr_from_p_matrix(set_perm_obj.set_n_per_perm, out['emp_p_d'], method='depletion')
out['BH_fdr_e'] = psp.p_adjust_bh(out['emp_p_e'])
out['BH_fdr_d'] = psp.p_adjust_bh(out['emp_p_d'])
out = out.sort_values('emp_p_e')
return out
e_results = make_results_table(e_test_obj, function_sets, e_per_set)
c_results = make_results_table(c_test_obj, function_sets, c_per_set)
i_results = make_results_table(i_test_obj, function_sets, i_per_set)
e_results.sort_values('fdr_e')
c_results.sort_values('fdr_e')
i_results.sort_values('fdr_e')
# join objects
# test objs
ce_test_obj = psp.TestObject.add_objects(c_test_obj,e_test_obj)
ci_test_obj = psp.TestObject.add_objects(c_test_obj,i_test_obj)
ei_test_obj = psp.TestObject.add_objects(e_test_obj,i_test_obj)
cei_test_obj = psp.TestObject.add_objects(ce_test_obj,i_test_obj)
# n per permuation objs
ce_per_set=psp.SetPerPerm.join_objects(c_per_set,e_per_set)
ci_per_set=psp.SetPerPerm.join_objects(c_per_set,i_per_set)
ei_per_set=psp.SetPerPerm.join_objects(e_per_set,i_per_set)
cei_per_set=psp.SetPerPerm.join_objects(ce_per_set,i_per_set)
# joint results
ce_results = make_results_table(ce_test_obj, function_sets, ce_per_set)
ci_results = make_results_table(ci_test_obj, function_sets, ci_per_set)
ei_results = make_results_table(ei_test_obj, function_sets, ei_per_set)
cei_results = make_results_table(cei_test_obj, function_sets, cei_per_set)
ce_results.sort_values('fdr_e')
ci_results.sort_values('fdr_e')
ei_results.sort_values('fdr_e')
cei_results.sort_values('fdr_e')
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Pointwise-Local-Reconstruction-Error" data-toc-modified-id="Pointwise-Local-Reconstruction-Error-1"><span class="toc-item-num">1 </span>Pointwise Local Reconstruction Error</a></span></li></ul></div>
Pointwise Local Reconstruction Error
====================================
Example for the usage of the `skcosmo.metrics.pointwise_local_reconstruction_error` as pointwise local reconstruction error (LFRE) on the degenerate CH4 manifold. We apply the local reconstruction measure on the degenerate CH4 manifold dataset. This dataset was specifically constructed to be representable by a 4-body features (bispectrum) but not by a 3-body features (power spectrum). In other words the dataset contains environments which are different, but have the same 3-body features. For more details about the dataset please refer to [Pozdnyakov 2020](https://doi.org/10.1103/PhysRevLett.125.166001) .
The skcosmo dataset already contains the 3 and 4-body features computed with [librascal](https://github.com/lab-cosmo/librascal) so we can load it and compare it with the LFRE.
```
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mpl
mpl.rc('font', size=20)
from skcosmo.datasets import load_degenerate_CH4_manifold
from skcosmo.metrics import pointwise_local_reconstruction_error
# load features
degenerate_manifold = load_degenerate_CH4_manifold()
power_spectrum_features = degenerate_manifold.data.SOAP_power_spectrum
bispectrum_features = degenerate_manifold.data.SOAP_bispectrum
print(degenerate_manifold.DESCR)
n_local_points = 20
print("Computing pointwise LFRE...")
# local reconstruction error of power spectrum features using bispectrum features
power_spectrum_to_bispectrum_pointwise_lfre = pointwise_local_reconstruction_error(
power_spectrum_features,
bispectrum_features,
n_local_points,
train_idx = np.arange(0, len(power_spectrum_features), 2),
test_idx = np.arange(0, len(power_spectrum_features)),
estimator=None,
n_jobs=4,
)
# local reconstruction error of bispectrum features using power spectrum features
bispectrum_to_power_spectrum_pointwise_lfre = pointwise_local_reconstruction_error(
bispectrum_features,
power_spectrum_features,
n_local_points,
train_idx = np.arange(0, len(power_spectrum_features), 2),
test_idx = np.arange(0, len(power_spectrum_features)),
estimator=None,
n_jobs=4,
)
print("Computing pointwise LFRE finished.")
print(
"LFRE(3-body, 4-body) = ",
np.linalg.norm(power_spectrum_to_bispectrum_pointwise_lfre)/np.sqrt(len(power_spectrum_to_bispectrum_pointwise_lfre))
)
print(
"LFRE(4-body, 3-body) = ",
np.linalg.norm(bispectrum_to_power_spectrum_pointwise_lfre)/np.sqrt(len(power_spectrum_to_bispectrum_pointwise_lfre))
)
fig, (ax34, ax43) = plt.subplots(
1, 2, constrained_layout=True, figsize=(16, 7.5), sharey="row", sharex=True
)
vmax = 0.5
X, Y = np.meshgrid(np.linspace(0.7, 0.9, 9), np.linspace(-0.1, 0.1, 9))
pcm = ax34.contourf(
X,
Y,
power_spectrum_to_bispectrum_pointwise_lfre[81:].reshape(9, 9).T,
vmin=0,
vmax=vmax,
)
ax43.contourf(
X,
Y,
bispectrum_to_power_spectrum_pointwise_lfre[81:].reshape(9, 9).T,
vmin=0,
vmax=vmax,
)
ax34.axhline(y=0, color="red", linewidth=5)
ax43.axhline(y=0, color="red", linewidth=5)
ax34.set_ylabel(r"v/$\pi$")
ax34.set_xlabel(r"u/$\pi$")
ax43.set_xlabel(r"u/$\pi$")
ax34.set_title(r"$X^-$ LFRE(3-body, 4-body)")
ax43.set_title(r"$X^-$ LFRE(4-body, 3-body)")
cbar = fig.colorbar(pcm, ax=[ax34, ax43], label="LFRE", location="bottom")
plt.show()
```
The environments span a manifold which is described by the coordinates $v/\pi$ and $u/\pi$ (please refer to [Pozdnyakov 2020](https://doi.org/10.1103/PhysRevLett.125.166001) for a concrete understanding of the manifold). The LFRE is presented for each environment in the manifold in the two contour plots. It can be seen that the reconstruction error of 4-body features using 3-body features (the left plot) is most significant along the degenerate line (the horizontal red line). This agrees with the fact that the 3-body features remain the same on the degenerate line and can therefore not reconstruct the 4-body features. On the other hand the 4-body features can perfectly reconstruct the 3-body features as seen in the right plot.
| github_jupyter |
```
"""
Code modified from PyTorch DCGAN examples: https://github.com/pytorch/examples/tree/master/dcgan
"""
from __future__ import print_function
import argparse
import os
import numpy as np
import random
import torch
import torch.nn as nn
import torch.nn.parallel
import torch.backends.cudnn as cudnn
import torch.optim as optim
import torch.utils.data
import torchvision.datasets as dset
import torchvision.transforms as transforms
import torchvision.utils as vutils
from torch.autograd import Variable
# custom weights initialization called on netG and netD
def weights_init(m):
classname = m.__class__.__name__
if classname.find('Conv') != -1:
m.weight.data.normal_(0.0, 0.02)
elif classname.find('BatchNorm') != -1:
m.weight.data.normal_(1.0, 0.02)
m.bias.data.fill_(0)
# compute the current classification accuracy
def compute_acc(preds, labels):
correct = 0
preds_ = preds.data.max(1)[1]
correct = preds_.eq(labels.data).cpu().sum()
acc = float(correct) / float(len(labels.data)) * 100.0
return acc
import torch.utils.data as data
from PIL import Image
import os
import os.path
IMG_EXTENSIONS = ['.jpg', '.jpeg', '.png', '.ppm', '.bmp', '.pgm']
def is_image_file(filename):
"""Checks if a file is an image.
Args:
filename (string): path to a file
Returns:
bool: True if the filename ends with a known image extension
"""
filename_lower = filename.lower()
return any(filename_lower.endswith(ext) for ext in IMG_EXTENSIONS)
def find_classes(dir, classes_idx=None):
classes = [d for d in os.listdir(dir) if os.path.isdir(os.path.join(dir, d))]
classes.sort()
if classes_idx is not None:
assert type(classes_idx) == tuple
start, end = classes_idx
classes = classes[start:end]
class_to_idx = {classes[i]: i for i in range(len(classes))}
return classes, class_to_idx
def make_dataset(dir, class_to_idx):
images = []
dir = os.path.expanduser(dir)
for target in sorted(os.listdir(dir)):
if target not in class_to_idx:
continue
d = os.path.join(dir, target)
if not os.path.isdir(d):
continue
for root, _, fnames in sorted(os.walk(d)):
for fname in sorted(fnames):
if is_image_file(fname):
path = os.path.join(root, fname)
item = (path, class_to_idx[target])
images.append(item)
return images
def pil_loader(path):
# open path as file to avoid ResourceWarning (https://github.com/python-pillow/Pillow/issues/835)
with open(path, 'rb') as f:
with Image.open(f) as img:
return img.convert('RGB')
def accimage_loader(path):
import accimage
try:
return accimage.Image(path)
except IOError:
# Potentially a decoding problem, fall back to PIL.Image
return pil_loader(path)
def default_loader(path):
from torchvision import get_image_backend
if get_image_backend() == 'accimage':
return accimage_loader(path)
else:
return pil_loader(path)
class ImageFolder(data.Dataset):
"""A generic data loader where the images are arranged in this way: ::
root/dog/xxx.png
root/dog/xxy.png
root/dog/xxz.png
root/cat/123.png
root/cat/nsdf3.png
root/cat/asd932_.png
Args:
root (string): Root directory path.
transform (callable, optional): A function/transform that takes in an PIL image
and returns a transformed version. E.g, ``transforms.RandomCrop``
target_transform (callable, optional): A function/transform that takes in the
target and transforms it.
loader (callable, optional): A function to load an image given its path.
Attributes:
classes (list): List of the class names.
class_to_idx (dict): Dict with items (class_name, class_index).
imgs (list): List of (image path, class_index) tuples
"""
def __init__(self, root, transform=None, target_transform=None,
loader=default_loader, classes_idx=None):
self.classes_idx = classes_idx
classes, class_to_idx = find_classes(root, self.classes_idx)
imgs = make_dataset(root, class_to_idx)
if len(imgs) == 0:
raise(RuntimeError("Found 0 images in subfolders of: " + root + "\n"
"Supported image extensions are: " + ",".join(IMG_EXTENSIONS)))
self.root = root
self.imgs = imgs
self.classes = classes
self.class_to_idx = class_to_idx
self.transform = transform
self.target_transform = target_transform
self.loader = loader
def __getitem__(self, index):
"""
Args:
index (int): Index
Returns:
tuple: (image, target) where target is class_index of the target class.
"""
path, target = self.imgs[index]
img = self.loader(path)
if self.transform is not None:
img = self.transform(img)
if self.target_transform is not None:
target = self.target_transform(target)
return img, target
def __len__(self):
return len(self.imgs)
class _netG(nn.Module):
def __init__(self, ngpu, nz):
super(_netG, self).__init__()
self.ngpu = ngpu
self.nz = nz
# first linear layer
self.fc1 = nn.Linear(110, 768)
# Transposed Convolution 2
self.tconv2 = nn.Sequential(
nn.ConvTranspose2d(768, 384, 5, 2, 0, bias=False),
nn.BatchNorm2d(384),
nn.ReLU(True),
)
# Transposed Convolution 3
self.tconv3 = nn.Sequential(
nn.ConvTranspose2d(384, 256, 5, 2, 0, bias=False),
nn.BatchNorm2d(256),
nn.ReLU(True),
)
# Transposed Convolution 4
self.tconv4 = nn.Sequential(
nn.ConvTranspose2d(256, 192, 5, 2, 0, bias=False),
nn.BatchNorm2d(192),
nn.ReLU(True),
)
# Transposed Convolution 5
self.tconv5 = nn.Sequential(
nn.ConvTranspose2d(192, 64, 5, 2, 0, bias=False),
nn.BatchNorm2d(64),
nn.ReLU(True),
)
# Transposed Convolution 5
self.tconv6 = nn.Sequential(
nn.ConvTranspose2d(64, 3, 8, 2, 0, bias=False),
nn.Tanh(),
)
def forward(self, input):
if isinstance(input.data, torch.cuda.FloatTensor) and self.ngpu > 1:
input = input.view(-1, self.nz)
fc1 = nn.parallel.data_parallel(self.fc1, input, range(self.ngpu))
fc1 = fc1.view(-1, 768, 1, 1)
tconv2 = nn.parallel.data_parallel(self.tconv2, fc1, range(self.ngpu))
tconv3 = nn.parallel.data_parallel(self.tconv3, tconv2, range(self.ngpu))
tconv4 = nn.parallel.data_parallel(self.tconv4, tconv3, range(self.ngpu))
tconv5 = nn.parallel.data_parallel(self.tconv5, tconv4, range(self.ngpu))
tconv5 = nn.parallel.data_parallel(self.tconv6, tconv5, range(self.ngpu))
output = tconv5
else:
input = input.view(-1, self.nz)
fc1 = self.fc1(input)
fc1 = fc1.view(-1, 768, 1, 1)
tconv2 = self.tconv2(fc1)
tconv3 = self.tconv3(tconv2)
tconv4 = self.tconv4(tconv3)
tconv5 = self.tconv5(tconv4)
tconv5 = self.tconv6(tconv5)
output = tconv5
return output
class _netD(nn.Module):
def __init__(self, ngpu, num_classes=10):
super(_netD, self).__init__()
self.ngpu = ngpu
# Convolution 1
self.conv1 = nn.Sequential(
nn.Conv2d(3, 16, 3, 2, 1, bias=False),
nn.LeakyReLU(0.2, inplace=True),
nn.Dropout(0.5, inplace=False),
)
# Convolution 2
self.conv2 = nn.Sequential(
nn.Conv2d(16, 32, 3, 1, 0, bias=False),
nn.BatchNorm2d(32),
nn.LeakyReLU(0.2, inplace=True),
nn.Dropout(0.5, inplace=False),
)
# Convolution 3
self.conv3 = nn.Sequential(
nn.Conv2d(32, 64, 3, 2, 1, bias=False),
nn.BatchNorm2d(64),
nn.LeakyReLU(0.2, inplace=True),
nn.Dropout(0.5, inplace=False),
)
# Convolution 4
self.conv4 = nn.Sequential(
nn.Conv2d(64, 128, 3, 1, 0, bias=False),
nn.BatchNorm2d(128),
nn.LeakyReLU(0.2, inplace=True),
nn.Dropout(0.5, inplace=False),
)
# Convolution 5
self.conv5 = nn.Sequential(
nn.Conv2d(128, 256, 3, 2, 1, bias=False),
nn.BatchNorm2d(256),
nn.LeakyReLU(0.2, inplace=True),
nn.Dropout(0.5, inplace=False),
)
# Convolution 6
self.conv6 = nn.Sequential(
nn.Conv2d(256, 512, 3, 1, 0, bias=False),
nn.BatchNorm2d(512),
nn.LeakyReLU(0.2, inplace=True),
nn.Dropout(0.5, inplace=False),
)
# discriminator fc
self.fc_dis = nn.Linear(13*13*512, 1)
# aux-classifier fc
self.fc_aux = nn.Linear(13*13*512, num_classes)
# softmax and sigmoid
self.softmax = nn.Softmax()
self.sigmoid = nn.Sigmoid()
def forward(self, input):
if isinstance(input.data, torch.cuda.FloatTensor) and self.ngpu > 1:
conv1 = nn.parallel.data_parallel(self.conv1, input, range(self.ngpu))
conv2 = nn.parallel.data_parallel(self.conv2, conv1, range(self.ngpu))
conv3 = nn.parallel.data_parallel(self.conv3, conv2, range(self.ngpu))
conv4 = nn.parallel.data_parallel(self.conv4, conv3, range(self.ngpu))
conv5 = nn.parallel.data_parallel(self.conv5, conv4, range(self.ngpu))
conv6 = nn.parallel.data_parallel(self.conv6, conv5, range(self.ngpu))
flat6 = conv6.view(-1, 13*13*512)
fc_dis = nn.parallel.data_parallel(self.fc_dis, flat6, range(self.ngpu))
fc_aux = nn.parallel.data_parallel(self.fc_aux, flat6, range(self.ngpu))
else:
conv1 = self.conv1(input)
conv2 = self.conv2(conv1)
conv3 = self.conv3(conv2)
conv4 = self.conv4(conv3)
conv5 = self.conv5(conv4)
conv6 = self.conv6(conv5)
flat6 = conv6.view(-1, 13*13*512)
fc_dis = self.fc_dis(flat6)
fc_aux = self.fc_aux(flat6)
classes = self.softmax(fc_aux)
realfake = self.sigmoid(fc_dis).view(-1, 1).squeeze(1)
return realfake, classes
class _netG_CIFAR10(nn.Module):
def __init__(self, ngpu, nz):
super(_netG_CIFAR10, self).__init__()
self.ngpu = ngpu
self.nz = nz
# first linear layer
self.fc1 = nn.Linear(110, 384)
# Transposed Convolution 2
self.tconv2 = nn.Sequential(
nn.ConvTranspose2d(384, 192, 4, 1, 0, bias=False),
nn.BatchNorm2d(192),
nn.ReLU(True),
)
# Transposed Convolution 3
self.tconv3 = nn.Sequential(
nn.ConvTranspose2d(192, 96, 4, 2, 1, bias=False),
nn.BatchNorm2d(96),
nn.ReLU(True),
)
# Transposed Convolution 4
self.tconv4 = nn.Sequential(
nn.ConvTranspose2d(96, 48, 4, 2, 1, bias=False),
nn.BatchNorm2d(48),
nn.ReLU(True),
)
# Transposed Convolution 4
self.tconv5 = nn.Sequential(
nn.ConvTranspose2d(48, 3, 4, 2, 1, bias=False),
nn.Tanh(),
)
def forward(self, input):
if isinstance(input.data, torch.cuda.FloatTensor) and self.ngpu > 1:
input = input.view(-1, self.nz)
fc1 = nn.parallel.data_parallel(self.fc1, input, range(self.ngpu))
fc1 = fc1.view(-1, 384, 1, 1)
tconv2 = nn.parallel.data_parallel(self.tconv2, fc1, range(self.ngpu))
tconv3 = nn.parallel.data_parallel(self.tconv3, tconv2, range(self.ngpu))
tconv4 = nn.parallel.data_parallel(self.tconv4, tconv3, range(self.ngpu))
tconv5 = nn.parallel.data_parallel(self.tconv5, tconv4, range(self.ngpu))
output = tconv5
else:
input = input.view(-1, self.nz)
fc1 = self.fc1(input)
fc1 = fc1.view(-1, 384, 1, 1)
tconv2 = self.tconv2(fc1)
tconv3 = self.tconv3(tconv2)
tconv4 = self.tconv4(tconv3)
tconv5 = self.tconv5(tconv4)
output = tconv5
return output
class _netD_CIFAR10(nn.Module):
def __init__(self, ngpu, num_classes=10):
super(_netD_CIFAR10, self).__init__()
self.ngpu = ngpu
# Convolution 1
self.conv1 = nn.Sequential(
nn.Conv2d(3, 16, 3, 2, 1, bias=False),
nn.LeakyReLU(0.2, inplace=True),
nn.Dropout(0.5, inplace=False),
)
# Convolution 2
self.conv2 = nn.Sequential(
nn.Conv2d(16, 32, 3, 1, 1, bias=False),
nn.BatchNorm2d(32),
nn.LeakyReLU(0.2, inplace=True),
nn.Dropout(0.5, inplace=False),
)
# Convolution 3
self.conv3 = nn.Sequential(
nn.Conv2d(32, 64, 3, 2, 1, bias=False),
nn.BatchNorm2d(64),
nn.LeakyReLU(0.2, inplace=True),
nn.Dropout(0.5, inplace=False),
)
# Convolution 4
self.conv4 = nn.Sequential(
nn.Conv2d(64, 128, 3, 1, 1, bias=False),
nn.BatchNorm2d(128),
nn.LeakyReLU(0.2, inplace=True),
nn.Dropout(0.5, inplace=False),
)
# Convolution 5
self.conv5 = nn.Sequential(
nn.Conv2d(128, 256, 3, 2, 1, bias=False),
nn.BatchNorm2d(256),
nn.LeakyReLU(0.2, inplace=True),
nn.Dropout(0.5, inplace=False),
)
# Convolution 6
self.conv6 = nn.Sequential(
nn.Conv2d(256, 512, 3, 1, 1, bias=False),
nn.BatchNorm2d(512),
nn.LeakyReLU(0.2, inplace=True),
nn.Dropout(0.5, inplace=False),
)
# discriminator fc
self.fc_dis = nn.Linear(4*4*512, 1)
# aux-classifier fc
self.fc_aux = nn.Linear(4*4*512, num_classes)
# softmax and sigmoid
self.softmax = nn.Softmax()
self.sigmoid = nn.Sigmoid()
def forward(self, input):
if isinstance(input.data, torch.cuda.FloatTensor) and self.ngpu > 1:
conv1 = nn.parallel.data_parallel(self.conv1, input, range(self.ngpu))
conv2 = nn.parallel.data_parallel(self.conv2, conv1, range(self.ngpu))
conv3 = nn.parallel.data_parallel(self.conv3, conv2, range(self.ngpu))
conv4 = nn.parallel.data_parallel(self.conv4, conv3, range(self.ngpu))
conv5 = nn.parallel.data_parallel(self.conv5, conv4, range(self.ngpu))
conv6 = nn.parallel.data_parallel(self.conv6, conv5, range(self.ngpu))
flat6 = conv6.view(-1, 4*4*512)
fc_dis = nn.parallel.data_parallel(self.fc_dis, flat6, range(self.ngpu))
fc_aux = nn.parallel.data_parallel(self.fc_aux, flat6, range(self.ngpu))
else:
conv1 = self.conv1(input)
conv2 = self.conv2(conv1)
conv3 = self.conv3(conv2)
conv4 = self.conv4(conv3)
conv5 = self.conv5(conv4)
conv6 = self.conv6(conv5)
flat6 = conv6.view(-1, 4*4*512)
fc_dis = self.fc_dis(flat6)
fc_aux = self.fc_aux(flat6)
classes = self.softmax(fc_aux)
realfake = self.sigmoid(fc_dis).view(-1, 1).squeeze(1)
return realfake, classes
dataset = 'cifar10'
dataroot = './root'
workers = 32
batchSize = 1
imageSize = 32
nz = 100
ngf = 64
ndf = 64
niter = 25
lr = 0.0002
beta1 = 0.5
cuda = True
ngpu = 1
netG = ''
netD = ''
outf = '.'
manualSeed = 1
num_classes = 2
gpu_id = 0
# specify the gpu id if using only 1 gpu
if ngpu == 1:
os.environ['CUDA_VISIBLE_DEVICES'] = str(gpu_id)
if manualSeed is None:
manualSeed = random.randint(1, 10000)
print("Random Seed: ", manualSeed)
random.seed(manualSeed)
torch.manual_seed(manualSeed)
if cuda:
torch.cuda.manual_seed_all(manualSeed)
dataset = dset.CIFAR10(
root=dataroot, download=True,
transform=transforms.Compose([
transforms.Scale(imageSize),
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5)),
]))
assert dataset
dataloader = torch.utils.data.DataLoader(dataset, batch_size=batchSize,
shuffle=True, num_workers=int(workers))
# some hyper parameters
ngpu = int(ngpu)
nz = int(nz)
ngf = int(ngf)
ndf = int(ndf)
num_classes = int(num_classes)
nc = 3
netG = _netG_CIFAR10(ngpu, nz)
netG.apply(weights_init)
netD = _netD_CIFAR10(ngpu, num_classes)
netD.apply(weights_init)
# loss functions
dis_criterion = nn.BCELoss()
aux_criterion = nn.NLLLoss()
# tensor placeholders
input = torch.FloatTensor(batchSize, 3, imageSize, imageSize)
noise = torch.FloatTensor(batchSize, nz, 1, 1)
eval_noise = torch.FloatTensor(batchSize, nz, 1, 1).normal_(0, 1)
dis_label = torch.FloatTensor(batchSize)
aux_label = torch.LongTensor(batchSize)
real_label = 1
fake_label = 0
# if using cuda
if cuda:
netD.cuda()
netG.cuda()
dis_criterion.cuda()
aux_criterion.cuda()
input, dis_label, aux_label = input.cuda(), dis_label.cuda(), aux_label.cuda()
noise, eval_noise = noise.cuda(), eval_noise.cuda()
# define variables
input = Variable(input)
noise = Variable(noise)
eval_noise = Variable(eval_noise)
dis_label = Variable(dis_label)
aux_label = Variable(aux_label)
# noise for evaluation
eval_noise_ = np.random.normal(0, 1, (batchSize, nz))
eval_label = np.random.randint(0, num_classes, batchSize)
eval_onehot = np.zeros((batchSize, num_classes))
eval_onehot[np.arange(batchSize), eval_label] = 1
eval_noise_[np.arange(batchSize), :num_classes] = eval_onehot[np.arange(batchSize)]
eval_noise_ = (torch.from_numpy(eval_noise_))
eval_noise.data.copy_(eval_noise_.view(batchSize, nz, 1, 1))
# setup optimizer
optimizerD = optim.Adam(netD.parameters(), lr=lr, betas=(beta1, 0.999))
optimizerG = optim.Adam(netG.parameters(), lr=lr, betas=(beta1, 0.999))
avg_loss_D = 0.0
avg_loss_G = 0.0
avg_loss_A = 0.0
for epoch in range(niter):
for i, data in enumerate(dataloader, 0):
############################
# (1) Update D network: maximize log(D(x)) + log(1 - D(G(z)))
###########################
# train with real
netD.zero_grad()
real_cpu, label = data
batch_size = real_cpu.size(0)
if cuda:
real_cpu = real_cpu.cuda()
input.data.resize_as_(real_cpu).copy_(real_cpu)
dis_label.data.resize_(batch_size).fill_(real_label)
aux_label.data.resize_(batch_size).copy_(label)
dis_output, aux_output = netD(input)
dis_errD_real = dis_criterion(dis_output, dis_label)
aux_errD_real = aux_criterion(aux_output, aux_label)
errD_real = dis_errD_real + aux_errD_real
errD_real.backward()
D_x = dis_output.data.mean()
# compute the current classification accuracy
accuracy = compute_acc(aux_output, aux_label)
# train with fake
noise.data.resize_(batch_size, nz, 1, 1).normal_(0, 1)
label = np.random.randint(0, num_classes, batch_size)
noise_ = np.random.normal(0, 1, (batch_size, nz))
class_onehot = np.zeros((batch_size, num_classes))
class_onehot[np.arange(batch_size), label] = 1
noise_[np.arange(batch_size), :num_classes] = class_onehot[np.arange(batch_size)]
noise_ = (torch.from_numpy(noise_))
noise.data.copy_(noise_.view(batch_size, nz, 1, 1))
aux_label.data.resize_(batch_size).copy_(torch.from_numpy(label))
fake = netG(noise)
dis_label.data.fill_(fake_label)
dis_output, aux_output = netD(fake.detach())
dis_errD_fake = dis_criterion(dis_output, dis_label)
aux_errD_fake = aux_criterion(aux_output, aux_label)
errD_fake = dis_errD_fake + aux_errD_fake
errD_fake.backward()
D_G_z1 = dis_output.data.mean()
errD = errD_real + errD_fake
optimizerD.step()
############################
# (2) Update G network: maximize log(D(G(z)))
###########################
netG.zero_grad()
dis_label.data.fill_(real_label) # fake labels are real for generator cost
dis_output, aux_output = netD(fake)
dis_errG = dis_criterion(dis_output, dis_label)
aux_errG = aux_criterion(aux_output, aux_label)
errG = dis_errG + aux_errG
errG.backward()
D_G_z2 = dis_output.data.mean()
optimizerG.step()
# compute the average loss
curr_iter = epoch * len(dataloader) + i
all_loss_G = avg_loss_G * curr_iter
all_loss_D = avg_loss_D * curr_iter
all_loss_A = avg_loss_A * curr_iter
all_loss_G += errG.data[0]
all_loss_D += errD.data[0]
all_loss_A += accuracy
avg_loss_G = all_loss_G / (curr_iter + 1)
avg_loss_D = all_loss_D / (curr_iter + 1)
avg_loss_A = all_loss_A / (curr_iter + 1)
print('[%d/%d][%d/%d] Loss_D: %.4f (%.4f) Loss_G: %.4f (%.4f) D(x): %.4f D(G(z)): %.4f / %.4f Acc: %.4f (%.4f)'
% (epoch, niter, i, len(dataloader),
errD.data[0], avg_loss_D, errG.data[0], avg_loss_G, D_x, D_G_z1, D_G_z2, accuracy, avg_loss_A))
if i % 100 == 0:
vutils.save_image(
real_cpu, '%s/real_samples.png' % outf)
print('Label for eval = {}'.format(eval_label))
fake = netG(eval_noise)
vutils.save_image(
fake.data,
'%s/fake_samples_epoch_%03d.png' % (outf, epoch)
)
# do checkpointing
torch.save(netG.state_dict(), '%s/netG_epoch_%d.pth' % (outf, epoch))
torch.save(netD.state_dict(), '%s/netD_epoch_%d.pth' % (outf, epoch))
```
| github_jupyter |
```
%%init_spark
launcher.jars = ["file:///opt/benchmark-tools/spark-sql-perf/target/scala-2.12/spark-sql-perf_2.12-0.5.1-SNAPSHOT.jar"]
launcher.conf.set("spark.sql.warehouse.dir", "hdfs:///user/livy")
!hadoop fs -mkdir /user/livy
val scaleFactor = "1" // data scale 1GB
val iterations = 1 // how many times to run the whole set of queries.
val format = "parquet" // support parquet or orc
val storage = "hdfs" // choose HDFS
val bucket_name = "/user/livy" // scala notebook only has the write permission of "hdfs:///user/livy" directory
val partitionTables = true // create partition tables
val query_filter = Seq() // Seq() == all queries
//val query_filter = Seq("q1-v2.4", "q2-v2.4") // run subset of queries
val randomizeQueries = false // run queries in a random order. Recommended for parallel runs.
// detailed results will be written as JSON to this location.
var resultLocation = s"${storage}://${bucket_name}/results/tpcds_${format}/${scaleFactor}/"
var databaseName = s"tpcds_${format}_scale_${scaleFactor}_db"
val use_arrow = false // when you want to use gazella_plugin to run TPC-DS, you need to set it true.
if (use_arrow){
val data_path= s"${storage}://${bucket_name}/datagen/tpcds_${format}/${scaleFactor}"
resultLocation = s"${storage}://${bucket_name}/results/tpcds_arrow/${scaleFactor}/"
databaseName = s"tpcds_arrow_scale_${scaleFactor}_db"
val tables = Seq("call_center", "catalog_page", "catalog_returns", "catalog_sales", "customer", "customer_address", "customer_demographics", "date_dim", "household_demographics", "income_band", "inventory", "item", "promotion", "reason", "ship_mode", "store", "store_returns", "store_sales", "time_dim", "warehouse", "web_page", "web_returns", "web_sales", "web_site")
if (spark.catalog.databaseExists(s"$databaseName")) {
println(s"$databaseName has exists!")
}else{
spark.sql(s"create database if not exists $databaseName").show
spark.sql(s"use $databaseName").show
for (table <- tables) {
if (spark.catalog.tableExists(s"$table")){
println(s"$table has exists!")
}else{
spark.catalog.createTable(s"$table", s"$data_path/$table", "arrow")
}
}
if (partitionTables) {
for (table <- tables) {
try{
spark.sql(s"ALTER TABLE $table RECOVER PARTITIONS").show
}catch{
case e: Exception => println(e)
}
}
}
}
}
val timeout = 60 // timeout in hours
// COMMAND ----------
// Spark configuration
spark.conf.set("spark.sql.broadcastTimeout", "10000") // good idea for Q14, Q88.
// ... + any other configuration tuning
// COMMAND ----------
sql(s"use $databaseName")
import com.databricks.spark.sql.perf.tpcds.TPCDS
val tpcds = new TPCDS (sqlContext = spark.sqlContext)
def queries = {
val filtered_queries = query_filter match {
case Seq() => tpcds.tpcds2_4Queries
case _ => tpcds.tpcds2_4Queries.filter(q => query_filter.contains(q.name))
}
if (randomizeQueries) scala.util.Random.shuffle(filtered_queries) else filtered_queries
}
val experiment = tpcds.runExperiment(
queries,
iterations = iterations,
resultLocation = resultLocation,
tags = Map("runtype" -> "benchmark", "database" -> databaseName, "scale_factor" -> scaleFactor))
println(experiment.toString)
experiment.waitForFinish(timeout*60*60)
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Slice-specified-nodes-in-dimspec" data-toc-modified-id="Slice-specified-nodes-in-dimspec-1"><span class="toc-item-num">1 </span>Slice specified nodes in dimspec</a></span></li><li><span><a href="#Test-parallelism" data-toc-modified-id="Test-parallelism-2"><span class="toc-item-num">2 </span>Test parallelism</a></span><ul class="toc-item"><li><ul class="toc-item"><li><span><a href="#Example-task" data-toc-modified-id="Example-task-2.0.1"><span class="toc-item-num">2.0.1 </span>Example task</a></span></li></ul></li><li><span><a href="#Serial-invocation" data-toc-modified-id="Serial-invocation-2.1"><span class="toc-item-num">2.1 </span>Serial invocation</a></span><ul class="toc-item"><li><span><a href="#Maybe-sqash-dimensions-to-fit-into-einsum?" data-toc-modified-id="Maybe-sqash-dimensions-to-fit-into-einsum?-2.1.1"><span class="toc-item-num">2.1.1 </span>Maybe sqash dimensions to fit into einsum?</a></span></li><li><span><a href="#Many-var-parallelisation" data-toc-modified-id="Many-var-parallelisation-2.1.2"><span class="toc-item-num">2.1.2 </span>Many var parallelisation</a></span></li></ul></li><li><span><a href="#Plot-parallelisation-theoretical-speedup" data-toc-modified-id="Plot-parallelisation-theoretical-speedup-2.2"><span class="toc-item-num">2.2 </span>Plot parallelisation theoretical speedup</a></span></li><li><span><a href="#Use-unix-tools" data-toc-modified-id="Use-unix-tools-2.3"><span class="toc-item-num">2.3 </span>Use unix tools</a></span><ul class="toc-item"><li><span><a href="#Threading" data-toc-modified-id="Threading-2.3.1"><span class="toc-item-num">2.3.1 </span>Threading</a></span></li><li><span><a href="#Multiprocessing" data-toc-modified-id="Multiprocessing-2.3.2"><span class="toc-item-num">2.3.2 </span>Multiprocessing</a></span></li></ul></li></ul></li></ul></div>
```
#import ray
import pyrofiler as pyrof
from pyrofiler.pyrofiler import Profiler
from pyrofiler import callbacks
import matplotlib.pyplot as plt
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import sys
from multiprocessing import Pool, Array
from multiprocessing.dummy import Pool as ThreadPool
import os
sns.set_style('whitegrid')
np.random.seed(42)
def work(arg):
i,x,y, par_vars, result_idx= arg
patch = sliced_contract(x, y, par_vars, i)
sl = target_slice(result_idx, par_vars, i)
pool = ThreadPool(processes=2**7)
```
# Slice specified nodes in dimspec
```
def _none_slice():
return slice(None)
def _get_idx(x, idxs, slice_idx, shapes=None):
if shapes is None:
shapes = [2]*len(idxs)
point = np.unravel_index(slice_idx, shapes)
get_point = {i:p for i,p in zip(idxs, point)}
if x in idxs:
p = get_point[x]
return slice(p,p+1)
else:
return _none_slice()
def _slices_for_idxs(idxs, *args, shapes=None, slice_idx=0):
"""Return array of slices along idxs"""
slices = []
for indexes in args:
_slice = [_get_idx(x, idxs, slice_idx, shapes) for x in indexes ]
slices.append(tuple(_slice))
return slices
def log_log_scale():
plt.yscale('log')
plt.xscale('log')
def minorticks():
plt.minorticks_on()
plt.grid(which='minor', alpha=0.5, linestyle='-', axis='both')
```
# Test parallelism
### Example task
```
def get_example_task(A=8, B=10, C=7, dim1=0):
shape1 = [2]*(A+B)
shape2 = [2]*(A+C)
for i in range(dim1):
shape1[-i] = 1
shape2[-i] = 1
T1 = np.random.randn(*shape1)
T2 = np.random.randn(*shape2)
common = list(range(A))
idxs1 = common + list(range(A, A+B))
idxs2 = common + list(range(A+B, A+B+C))
return (T1, idxs1), (T2, idxs2)
x, y = get_example_task(A=9)
x[1], y[1]
```
## Serial invocation
```
def contract(A, B):
a, idxa = A
b, idxb = B
contract_idx = set(idxa) & set(idxb)
result_idx = set(idxa + idxb)
print('contract result idx',result_idx)
C = np.einsum(a,idxa, b,idxb, result_idx)
return C
def sliced_contract(x, y, idxs, num):
slices = _slices_for_idxs(idxs, x[1], y[1], slice_idx=num)
a = x[0][slices[0]]
b = y[0][slices[1]]
with pyrof.timing(f'\tcontract sliced {num}'):
C = contract((a, x[1]), (b, y[1]))
return C
def target_slice(result_idx, idxs, num):
slices = _slices_for_idxs(idxs, result_idx, slice_idx=num)
return slices
with pyrof.timing('contract'):
C = contract(x, y)
```
### Maybe sqash dimensions to fit into einsum?
```
def __contract_bound(A, B):
a, idxa = A
b, idxb = B
contract_idx = set(idxa) & set(idxb)
def glue_first(shape):
sh = [shape[0] * shape[1]] + list(shape[2:])
return sh
result_idx = set(idxa + idxb)
_map_a = {k:v for k,v in zip(idxa, a.shape)}
_map_b = {k:v for k,v in zip(idxb, b.shape)}
_map = {**_map_a, **_map_b}
print(_map)
result_idx = sorted(tuple(_map.keys()))
target_shape = tuple([_map[i] for i in result_idx])
_dimlen = len(result_idx)
_maxdims = 22
print('dimlen',_dimlen)
new_a, new_b = a.shape, b.shape
if _dimlen>_maxdims:
_contr_dim = _dimlen - _maxdims
print(len(new_a), len(new_b))
for i in range(_contr_dim):
idxa = idxa[1:]
idxb = idxb[1:]
new_a = glue_first(new_a)
new_b = glue_first(new_b)
_map_a = {k:v for k,v in zip(idxa, a.shape)}
_map_b = {k:v for k,v in zip(idxb, b.shape)}
_map = {**_map_a, **_map_b}
print(_map)
result_idx = sorted(tuple(_map.keys()))
print(len(new_a), len(new_b))
a = a.reshape(new_a)
b = b.reshape(new_b)
print(a.shape, b.shape)
print(idxa, idxb)
print('btsh',result_idx, target_shape)
C = np.einsum(a,idxa, b,idxb, result_idx)
return C.reshape(*target_shape)
def __add_dims(x, dims, ofs):
arr, idxs = x
arr = arr.reshape(list(arr.shape) + [1]*dims)
md = max(idxs)
return arr, idxs + list(range(md+ofs, ofs+md+dims))
```
### Many var parallelisation
```
prof_seq = Profiler()
prof_seq.use_append()
contract_idx = set(x[1]) & set(y[1])
result_idx = set(x[1] + y[1])
for i in range(1):
_ = contract(x,y)
for rank in range(1,7):
with prof_seq.timing('Single thread'):
C = contract(x,y)
par_vars = list(range(rank))
target_shape = C.shape
with prof_seq.timing('One patch: total'):
i = 0
with prof_seq.timing('One patch: compute'):
patch = sliced_contract(x, y, par_vars, i)
C_par = np.empty(target_shape)
with prof_seq.timing('One patch: assign'):
_slice = target_slice(result_idx, par_vars, i)
C_par[_slice[0]] = patch
```
## Plot parallelisation theoretical speedup
```
prof_seq.data
threads = 2**np.arange(1,7)
C_size = sys.getsizeof(C)
for k in prof_seq.data:
plt.plot(threads, prof_seq.data[k], label=k)
plt.loglog(basex=2, basey=2)
from matplotlib.ticker import FormatStrFormatter
plt.title(f'Single node parallelization one batch test. Task size: {C_size:e}')
plt.xlabel('Thread count')
plt.ylabel('Time')
minorticks()
plt.legend()
plt.savefig('figures/node_par_seqtest.pdf')
plt.close()
```
## Use unix tools
### Threading
```
x,y = get_example_task(A=20, B=9, C=8, dim1=2)
contract_idx = set(x[1]) & set(y[1])
result_idx = set(x[1] + y[1])
prof_thread = Profiler()
prof_thread.use_append()
for i in range(1):
C = contract(x,y)
C_size = sys.getsizeof(C)
target_shape = C.shape
C = None
for rank in range(1,7):
if rank==1:
with prof_thread.timing('Single thread'):
C = contract(x,y)
C = None
with prof_thread.timing('Multithread: total'):
par_vars = list(range(rank))
threads = 2**len(par_vars)
os.global_C = np.empty(target_shape)
with prof_thread.timing('Multithread: work'):
_ = pool.map(work, ((i,x,y,par_vars,result_idx)for i in range(threads)))
#assert np.array_equal(C, os.global_C)
_data = prof_thread.data
print(_data)
_data_knl = {'Single thread': [1.3409993648529053, 1.3587844371795654, 1.3243846893310547, 1.336273193359375, 1.3332529067993164, 1.3412296772003174], 'Multithread: work': [0.7453043460845947, 0.5046432018280029, 0.39226293563842773, 0.40014123916625977, 0.5875647068023682, 1.0763416290283203], 'Multithread: total': [0.7459092140197754, 0.5054154396057129, 0.3927571773529053, 0.4007418155670166, 0.588019847869873, 1.0771734714508057]}
_data_biggest = {'Single thread': [27.42847204208374, 26.855594873428345, 26.628530979156494, 26.862286806106567, 26.71247911453247, 27.049968957901], 'Multithread: work': [14.236661434173584, 7.511402368545532, 4.950175762176514, 3.012814521789551, 2.351712703704834, 1.994131088256836], 'Multithread: total': [14.23719048500061, 7.512014150619507, 4.950707912445068, 3.0133090019226074, 2.3522441387176514, 1.9946098327636719]}
#_data = _data_biggest
threads = 2**np.arange(1,7)
for k in _data:
plt.plot(threads, _data[k], label=k)
plt.loglog(basex=2, basey=2)
plt.yscale('linear')
from matplotlib.ticker import FormatStrFormatter
plt.title(f'Single node parallelization test. Task size: {C_size:e}')
plt.xlabel('Thread count')
plt.ylabel('Time')
minorticks()
plt.legend()
plt.savefig('figures/node_par_threadtest_biggest.pdf')
#plt.rcParams.update({"xtick.bottom" : True, "ytick.left" : True})
sns.set_style('whitegrid')
#sns.set()
_data_block = {
'28':{'Single thread': [4.890172481536865], 'Multithread: work': [5.31355881690979, 2.839036464691162, 1.6587004661560059, 1.4607517719268799, 1.1708364486694336, 1.3796212673187256], 'Multithread: total': [5.31405234336853, 2.839534282684326, 1.659132957458496, 1.4612171649932861, 1.1718018054962158, 1.380187749862671]}
,'29': {'Single thread': [12.708141088485718], 'Multithread: work': [12.543375015258789, 6.445459604263306, 3.702291250228882, 2.225062131881714, 1.7111496925354004, 1.9049854278564453], 'Multithread: total': [12.543986320495605, 6.445924997329712, 3.7027952671051025, 2.2256860733032227, 1.7118234634399414, 1.905548095703125]}
, '30': {'Single thread': [26.65827775001526], 'Multithread: work': [26.532104015350342, 13.471351146697998, 7.361323356628418, 4.6045496463775635, 2.9114484786987305, 2.138317108154297], 'Multithread: total': [26.532758712768555, 13.471930980682373, 7.363482475280762, 4.605044364929199, 2.91215181350708, 2.1388139724731445]}
, '31': {'Single thread': [54.215914249420166], 'Multithread: work': [53.743674755096436, 27.541589498519897, 15.45585584640503, 8.812772750854492, 5.398884296417236, 4.5649192333221436], 'Multithread: total': [53.74607563018799, 27.542162895202637, 15.456344604492188, 8.814988851547241, 5.399648427963257, 4.5654377937316895]}
, '32': {'Single thread': [107.05718398094177], 'Multithread: work': [106.85966396331787, 55.66744685173035, 31.097278356552124, 18.133748292922974, 10.42065167427063, 9.078657865524292], 'Multithread: total': [106.86018991470337, 55.669677734375, 31.099481344223022, 18.13595175743103, 10.421445369720459, 9.080750703811646]}
}
threads = 2**np.arange(1,7)
fig, axs = plt.subplots(1,1, figsize=(6,6))
colors = (plt.cm.gnuplot2(x) for x in np.linspace(.8,.2,len(_data_block)))
for size, _data in _data_block.items():
singl = _data['Single thread']
total = _data['Multithread: total']
c = next(colors)
plt.plot(threads, total, '-D',color=c, label=f'Tensor size {2**(4+int(size))/1e9:.2f}Gb')
plt.plot(threads, singl*len(threads), '--', alpha=.3, color=c )
#from matplotlib.ticker import FormatStrFormatter
plt.loglog(basex=2, basey=2)
#plt.yscale('linear')
plt.grid()
#minorticks()
ax = plt.gca()
#ax.yaxis.set_minor_locator(plt.ticker.LogLocator(base=10.0, subs='all'))
#ax.yaxis.set_minor_formatter(plt.ticker.NullFormatter())
plt.title(f'Single node contraction parallelization for different sizes')
plt.xlabel('Thread count')
plt.ylabel('Time')
plt.grid(True,which="both")
handles, labels = plt.gca().get_legend_handles_labels()
plt.legend(handles[::-1], labels[::-1], loc='upper right')
plt.savefig('figures/node_par_threadtest_gener_jlse.pdf')
#plt.rcParams.update({"xtick.bottom" : True, "ytick.left" : True})
sns.set_style('whitegrid')
#sns.set()
_data_block = {
'27':{'Single thread': [4.890172481536865], 'Multithread: work': [5.31355881690979, 2.839036464691162, 1.6587004661560059, 1.4607517719268799, 1.1708364486694336, 1.3796212673187256], 'Multithread: total': [5.31405234336853, 2.839534282684326, 1.659132957458496, 1.4612171649932861, 1.1718018054962158, 1.380187749862671]}
,'30': {'Single thread': [37.403658866882324], 'Multithread: work': [39.51915979385376, 21.37852430343628, 11.835341453552246, 7.165068864822388, 4.922534942626953, 4.410918235778809], 'Multithread: total': [39.519590854644775, 21.378950595855713, 11.83582329750061, 7.1655051708221436, 4.923001050949097, 4.411387205123901
]}
}
threads = 2**np.arange(1,7)
fig, axs = plt.subplots(1,1, figsize=(6,6))
colors = (plt.cm.gnuplot2(x) for x in np.linspace(.8,.2,len(_data_block)))
for size, _data in _data_block.items():
singl = _data['Single thread']
total = _data['Multithread: total']
c = next(colors)
plt.plot(threads, total, '-D',color=c, label=f'Tensor size {2**(4+int(size))/1e9:.2f}Gb')
plt.plot(threads, singl*len(threads), '--', alpha=.3, color=c )
#from matplotlib.ticker import FormatStrFormatter
plt.loglog(basex=2, basey=2)
#plt.yscale('linear')
plt.grid()
#minorticks()
ax = plt.gca()
#ax.yaxis.set_minor_locator(plt.ticker.LogLocator(base=10.0, subs='all'))
#ax.yaxis.set_minor_formatter(plt.ticker.NullFormatter())
plt.title(f'Single node contraction parallelization for different sizes')
plt.xlabel('Thread count')
plt.ylabel('Time')
plt.grid(True,which="both")
handles, labels = plt.gca().get_legend_handles_labels()
plt.legend(handles[::-1], labels[::-1], loc='upper right')
plt.savefig('figures/node_par_threadtest_gener_theta.pdf')
```
### Multiprocessing
```
flat_size = len(C.flatten())
with pyrof.timing('init array'):
os.global_C = np.empty(target_shape)
#os.global_C = tonumpyarray(Array('d', flat_size))
#us.global_C = os.global_C.reshape(target_shape)
pool = Pool(processes=threads)
print('inited pool')
with pyrof.timing('parallel work'):
print('started work')
_ = pool.map(work, range(threads))
C_size = sys.getsizeof(os.global_C)
print(f'result size: {C_size:e}')
assert np.array_equal(C, os.global_C)
del os.global_C
```
| github_jupyter |
##### Copyright 2021 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# TensorFlow Lite Model Analyzer
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/lite/guide/model_analyzer"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/tensorflow/blob/master/tensorflow/lite/g3doc/guide/model_analyzer.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/tensorflow/blob/master/tensorflow/lite/g3doc/guide/model_analyzer.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/tensorflow/tensorflow/lite/g3doc/guide/model_analyzer.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
TensorFlow Lite Model Analyzer API helps you analyze models in TensorFlow Lite format by listing a model's structure.
## Model Analyzer API
The following API is available for the TensorFlow Lite Model Analyzer.
```
tf.lite.experimental.Analyzer.analyze(model_path=None,
model_content=None,
gpu_compatibility=False)
```
You can find the API details from https://www.tensorflow.org/api_docs/python/tf/lite/experimental/Analyzer or run `help(tf.lite.experimental.Analyzer.analyze)` from a Python terminal.
## Basic usage with simple Keras model
The following code shows basic usage of Model Analyzer. It shows contents of the converted Keras model in TFLite model content, formatted as a flatbuffer object.
```
import tensorflow as tf
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(128, 128)),
tf.keras.layers.Dense(256, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10)
])
fb_model = tf.lite.TFLiteConverter.from_keras_model(model).convert()
tf.lite.experimental.Analyzer.analyze(model_content=fb_model)
```
## Basic usage with MobileNetV3Large Keras model
This API works with large models such as MobileNetV3Large. Since the output is large, you might want to browse it with your favorite text editor.
```
model = tf.keras.applications.MobileNetV3Large()
fb_model = tf.lite.TFLiteConverter.from_keras_model(model).convert()
tf.lite.experimental.Analyzer.analyze(model_content=fb_model)
```
## Check GPU delegate compatibility
The ModelAnalyzer API provides a way to check the [GPU delegate](https://www.tensorflow.org/lite/performance/gpu) compatibility of the given model by providing `gpu_compatibility=True` option.
### Case 1: When model is incompatibile
The following code shows a way to use `gpu_compatibility=True` option for simple tf.function which uses `tf.slice` with a 2D tensor and `tf.cosh` which are not compatible with GPU delegate.
You will see `GPU COMPATIBILITY WARNING` per every node which has compatibility issue(s).
```
import tensorflow as tf
@tf.function(input_signature=[
tf.TensorSpec(shape=[4, 4], dtype=tf.float32)
])
def func(x):
return tf.cosh(x) + tf.slice(x, [1, 1], [1, 1])
converter = tf.lite.TFLiteConverter.from_concrete_functions(
[func.get_concrete_function()], func)
converter.target_spec.supported_ops = [
tf.lite.OpsSet.TFLITE_BUILTINS,
tf.lite.OpsSet.SELECT_TF_OPS,
]
fb_model = converter.convert()
tf.lite.experimental.Analyzer.analyze(model_content=fb_model, gpu_compatibility=True)
```
### Case 2: When model is compatibile
In this example, the given model is compatbile with GPU delegate.
**Note:** Even though the tool doesn't find any compatibility issue, it doesn't guarantee that your model works well with GPU delegate on every device. There could be some runtime incompatibililty happen such as missing `CL_DEVICE_IMAGE_SUPPORT` feature by target OpenGL backend.
```
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(128, 128)),
tf.keras.layers.Dense(256, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10)
])
fb_model = tf.lite.TFLiteConverter.from_keras_model(model).convert()
tf.lite.experimental.Analyzer.analyze(model_content=fb_model, gpu_compatibility=True)
```
| github_jupyter |
<a href="http://landlab.github.io"><img style="float: left" src="../../landlab_header.png"></a>
WARNING: This tutorial has not been updated to work with Landlab 2.0 and is thus not tested to verify that it will run.
### Tutorial For Cellular Automaton Vegetation Model Coupled With Ecohydrologic Model
<hr>
<small>For more Landlab tutorials, click here: <a href="https://landlab.readthedocs.io/en/v2_dev/user_guide/tutorials.html">https://landlab.readthedocs.io/en/v2_dev/user_guide/tutorials.html</a></small>
<hr>
This tutorial demonstrates implementation of the Cellular Automaton Tree-GRass-Shrub Simulator (CATGRaSS) [Zhou et al., 2013] on a flat domain. This model is built using components from the Landlab component library. CATGRaSS is spatially explicit model of plant coexistence. It simulates local ecohydrologic dynamics (soil moisture, transpiration, biomass) and spatial evolution of tree, grass, and shrub Plant Functional Types (PFT) driven by rainfall and solar radiation.
Each cell in the model grid can hold a single PFT or remain empty. Tree and shrub plants disperse seeds to their neighbors. Grass seeds are assumed to be available at each cell. Establishment of plants in empty cells is determined probabilistically based on water stress of each PFT. Plants with lower water stress have higher probability of establishment. Plant mortality is simulated probabilistically as a result of aging and drought stress. Fires and grazing will be added to this model soon.
This model (driver) contains:
- A local vegetation dynamics model that simulates storm and inter-storm water balance and ecohydrologic fluxes (ET, runoff), and plant biomass dynamics by coupling the following components:
- PrecipitationDistribution
- Radiation
- PotentialEvapotranspiration
- SoilMoisture
- Vegetation
- A spatially explicit probabilistic cellular automaton component that simulates plant competition by tracking establishment and mortality of plants based on soil moisture stress:
- VegCA
To run this Jupyter notebook, please make sure that the following files are in the same folder:
- cellular_automaton_vegetation_flat_domain.ipynb (this notebook)
- Inputs_Vegetation_CA.txt (Input parameters for the model)
- Ecohyd_functions_flat.py (Utility functions)
[Ref: Zhou, X, E. Istanbulluoglu, and E.R. Vivoni. "Modeling the ecohydrological role of aspect-controlled radiation on tree-grass-shrub coexistence in a semiarid climate." Water Resources Research 49.5 (2013): 2872-2895]
In this tutorial, we are going to work with a landscape in central New Mexico, USA, where aspect controls the organization of PFTs. The climate in this area is semi-arid with Mean Annual Precipitation (MAP) of 254 mm [Zhou et. al 2013].
We will do the following:
- Import a landscape
- Initialize the landscape with random distribution of PFTs
- Run the coupled Ecohydrology and cellular automata plant competition model for 50 years
- Visualize and examine outputs
#### Let us walk through the code:
Import the required libraries
```
from __future__ import print_function
%matplotlib inline
import time
import numpy as np
from landlab import RasterModelGrid as rmg
from landlab import load_params
from Ecohyd_functions_flat import (
Initialize_,
Empty_arrays,
Create_PET_lookup,
Save_,
Plot_,
)
```
Note: 'Ecohyd_functions_flat.py' is a utility script that contains 'functions', which instantiates components and manages inputs and outputs, and help keep this driver concise. Contents of 'Ecohyd_functions_flat.py' can be a part of this driver (current file), however left out to keep driver concise.
To minimize computation time, we will use two grids in this driver. One grid will represent a flat landscape or domain (i.e., landscape with same elevation), on which the cellular automata plant competition will be simulated at an yearly time step. Another grid, with enough cells to house one cell for each of the plant functional types (PFTs), will be used to simulate soil moisture decay and local vegetation dynamics, in between successive storms (i.e. time step = one storm). Cumulative water stress (stress experienced by plants due to lack of enough soil moisture) will be calculated over an year and mapped to the other grid.
- grid: This grid represents the actual landscape. Each cell can be occupied by a single PFT such as tree, shrub, grass, or can be empty (bare). Initial PFT distribution is randomnly generated from inputs of percentage of cells occupied by each PFT.
- grid1: This grid allows us to calculate PFT specific cumulative water stress (cumulated over each storm in the year) and mapped with 'grid'.
Note: In this tutorial, the physical ecohydrological components and cellular automata plant competition will be run on grids with different resolution. To use grids with same resolution, see the tutorial 'cellular_automaton_vegetation_DEM.ipynb'.
```
grid1 = rmg((100, 100), spacing=(5.0, 5.0))
grid = rmg((5, 4), spacing=(5.0, 5.0))
```
Include the input file that contains all input parameters needed for all components. This file can either be a python dictionary or a text file that can be converted into a python dictionary. If a text file is provided, it will be converted to a Python dictionary. Here we use an existing text file prepared for this exercise.
```
InputFile = "Inputs_Vegetation_CA_flat.txt"
data = load_params(InputFile) # Create dictionary that holds the inputs
```
Instantiate landlab components to simulate corresponding attributes. In this example, we shall demonstrate the use of seasonal rainfall and PFT-specific potential evapotranspiration. The instantiated objects are:
- PD_D: object for dry season rainfall,
- PD_W: object for wet season rainfall,
- Rad: Radiation object computes radiation factor defined as the ratio of total shortwave radiation incident on a sloped surface to total shortwave radiation incident on a flat surface. Note: in this example a flat domain is considered. Radiation factor returned will be a cellular field of ones. This component is included because potential evaporanspiration (PET) component receives an input of radiation factor as a field.
- PET_PFT: Plant specific PET objects. PET is upper boundary to ET. For long-term simulations PET is represented using a cosine function as a function of day of year. Parameters of this function were obtained from P-M model application at a weather station. PET is spatially distributed by using the radiation factor.
- SM: Soil Moisture object simulates depth-averaged soil moisture at each cell using inputs of potential evapotranspiration, live leaf area index and vegetation cover.
- VEG: Vegetation dynamics object simulates net primary productivity, biomass and leaf area index (LAI) at each cell based on inputs of root-zone average soil moisture.
- vegca: Cellular Automaton plant competition object is run once every year. This object is initialized with a random cellular field of PFT. Every year, this object updates the cellular field of PFT based on probabilistic establishment and mortality of PFT at each cell.
Note: Almost every component in landlab is coded as a 'class' (to harness the advantages of objective oriented programming). An 'object' is the instantiation of the 'class' (for more information, please refer any objective oriented programming book). A 'field' refers to a Landlab field (please refer to the [Landlab documentation](https://github.com/landlab/landlab/wiki/Grid#adding-data-to-a-landlab-grid-element-using-fields) to learn more about Landlab fields).
Now let's instantiate all Landlab components that we are going to use for this tutorial:
```
PD_D, PD_W, Rad, PET_Tree, PET_Shrub, PET_Grass, SM, VEG, vegca = Initialize_(
data, grid, grid1
)
```
Lets look at the initial organization of PFTs
```
import matplotlib.pyplot as plt
import matplotlib as mpl
cmap = mpl.colors.ListedColormap(["green", "red", "black", "white", "red", "black"])
bounds = [-0.5, 0.5, 1.5, 2.5, 3.5, 4.5, 5.5]
norm = mpl.colors.BoundaryNorm(bounds, cmap.N)
description = "green: grass; red: shrub; black: tree; white: bare"
plt.figure(101)
grid1.imshow(
"vegetation__plant_functional_type",
at="cell",
cmap=cmap,
grid_units=("m", "m"),
norm=norm,
limits=[0, 5],
allow_colorbar=False,
)
plt.figtext(0.2, 0.0, description, weight="bold", fontsize=10)
```
Specify an approximate number of years for the model to run. For this example, we will run the simulation for 600 years. It might take less than 2+ minutes to run.
```
n_years = 600 # Approx number of years for model to run
# Calculate approximate number of storms per year
fraction_wet = (data["doy__end_of_monsoon"] - data["doy__start_of_monsoon"]) / 365.0
fraction_dry = 1 - fraction_wet
no_of_storms_wet = (
8760 * (fraction_wet) / (data["mean_interstorm_wet"] + data["mean_storm_wet"])
)
no_of_storms_dry = (
8760 * (fraction_dry) / (data["mean_interstorm_dry"] + data["mean_storm_dry"])
)
n = int(n_years * (no_of_storms_wet + no_of_storms_dry))
```
Create empty arrays to store spatio-temporal data over multiple iterations. The captured data can be used for plotting model outputs.
```
P, Tb, Tr, Time, VegType, PET_, Rad_Factor, EP30, PET_threshold = Empty_arrays(
n, grid, grid1
)
```
To reduce computational overhead, we shall create a lookup array for plant-specific PET values for each day of the year.
```
Create_PET_lookup(Rad, PET_Tree, PET_Shrub, PET_Grass, PET_, Rad_Factor, EP30, grid)
```
Specify current_time (in years). current_time is the current time in the simulation.
```
# # Represent current time in years
current_time = 0 # Start from first day of Jan
# Keep track of run time for simulation - optional
Start_time = time.clock() # Recording time taken for simulation
# declaring few variables that will be used in the storm loop
time_check = 0.0 # Buffer to store current_time at previous storm
yrs = 0 # Keep track of number of years passed
WS = 0.0 # Buffer for Water Stress
Tg = 270 # Growing season in days
```
The loop below couples the components introduced above in a for loop until all "n" number of storms are generated. Time is advanced by the soil moisture object based on storm and interstorm durations that are estimated by the strom generator object. The ecohydrologic model is run each storm whereas cellular automaton vegetation component is run once every year.
Note: This loop might take less than 2 minutes (depending on your computer) to run for 600 year simulation. Ignore any warnings you might see.
```
# # Run storm Loop
for i in range(0, n):
# Update objects
# Calculate Day of Year (DOY)
Julian = int(np.floor((current_time - np.floor(current_time)) * 365.0))
# Generate seasonal storms
# for Dry season
if Julian < data["doy__start_of_monsoon"] or Julian > data["doy__end_of_monsoon"]:
PD_D.update()
P[i] = PD_D.storm_depth
Tr[i] = PD_D.storm_duration
Tb[i] = PD_D.interstorm_duration
# Wet Season - Jul to Sep - NA Monsoon
else:
PD_W.update()
P[i] = PD_W.storm_depth
Tr[i] = PD_W.storm_duration
Tb[i] = PD_W.interstorm_duration
# Spatially distribute PET and its 30-day-mean (analogous to degree day)
grid["cell"]["surface__potential_evapotranspiration_rate"] = PET_[Julian]
grid["cell"]["surface__potential_evapotranspiration_30day_mean"] = EP30[Julian]
# Assign spatial rainfall data
grid["cell"]["rainfall__daily_depth"] = P[i] * np.ones(grid.number_of_cells)
# Update soil moisture component
current_time = SM.update(current_time, Tr=Tr[i], Tb=Tb[i])
# Decide whether its growing season or not
if Julian != 364:
if EP30[Julian + 1, 0] > EP30[Julian, 0]:
PET_threshold = 1
# 1 corresponds to ETThresholdup (begin growing season)
else:
PET_threshold = 0
# 0 corresponds to ETThresholddown (end growing season)
# Update vegetation component
VEG.update(PETThreshold_switch=PET_threshold, Tb=Tb[i], Tr=Tr[i])
# Update yearly cumulative water stress data
WS += (grid["cell"]["vegetation__water_stress"]) * Tb[i] / 24.0
# Record time (optional)
Time[i] = current_time
# Update spatial PFTs with Cellular Automata rules
if (current_time - time_check) >= 1.0:
if yrs % 100 == 0:
print("Elapsed time = {time} years".format(time=yrs))
VegType[yrs] = grid1["cell"]["vegetation__plant_functional_type"]
WS_ = np.choose(VegType[yrs], WS)
grid1["cell"]["vegetation__cumulative_water_stress"] = WS_ / Tg
vegca.update()
time_check = current_time
WS = 0
yrs += 1
VegType[yrs] = grid1["cell"]["vegetation__plant_functional_type"]
```
Time_Consumed is an optional variable that gives information about computer running time
```
Final_time = time.clock()
Time_Consumed = (Final_time - Start_time) / 60.0 # in minutes
print("Time_consumed = {time} minutes".format(time=Time_Consumed))
```
Save the outputs using ``numpy.save()``. These files have '.nc' extension, which can be loaded using ``numpy.load()``.
```
# # Saving
sim = "Sim_26Jul16_"
# Save_(sim, Tb, Tr, P, VegType, yrs, Time_Consumed, Time)
```
Let's look at outputs.
Plots of the cellular field of PFT at specified year step can be found below where:
GRASS = green; SHRUB = red; TREE = black; BARE = white;
At the end, percentage cover of each PFT is plotted with respect to time.
```
Plot_(grid1, VegType, yrs, yr_step=100)
```
If you want to explore this model further, open 'Inputs_Vegetation_CA.txt' and change the input parameters (e.g., initial PFT distribution percentages, storm characteristics, etc..).
### Click here for more <a href="https://landlab.readthedocs.io/en/v2_dev/user_guide/tutorials.html">Landlab tutorials</a>
| github_jupyter |
# Batch job analysis: Overview
Running batch jobs is an essential operation on IBM Z or mainframe. Everyday, up to 60,000 batch jobs are run on a typical mainframe system, and that number tends to double on the first and last day of each calendar month, quarter, and year. Analysis of logs and other output from this large workload requires the analytics of a machine learning solution like IBM Watson Machine Learning for z/OS (WMLz).
You can use WMLz to analyze your batch processing and extract insights to enhance the following aspects of your batch operation:
* Seasonability of batch jobs and trends of workload change
* Impact of transactions and other business operations to batch job elapsed time
* Prediction of elapsed time of long-running jobs
* Identification of potentially abnormal job instances as well as transaction volumes.
The following code pattern is a sample project that uses WMLz 2.1.0.3 for batch job analysis. The hypothetical project supposes that the system administrators at BankABC use the Python notebook and modeler flow capabilities of WMLz to analyze batch jobs.
* BankABC runs a Master Batch Job(MBJ) at midnight everyday. MBJ consists of more than 10,000 jobs that cover various types of mission-critical transactions.
* BankABC wants to analyze MBJ elapsed time and to find out the pattern of changes in elapsed time, the most impactful factors responsible for the changes, and the correlation between elapsed time and daily transaction volume.
* BankABC also wants to know if MBJ elapsed time is predictable. Usually, MBJ runs for 2~5 hours at midnight. The ability to predict the time of job completion will allow the bank to arrange MBJ and maintenance jobs to avoid interrupting its normal business operation.
After completing the project, you will learn how to perform the following tasks:
* Extract batch job operation data from SMF Type 30 records
* Explore log data to extract insights about batch job elapsed time
* Use various algorithms to predict batch job elapsed time
* Identify potentially abnormal job instances and correlation with transaction volume
* Learn to use WMLz web UI where you can use Jupyter Notebooks to code in Python and SPSS Modeler Flow to explore data and train model in canvas
* Read z/OS native files, including SMF Type 30 records with Python notebook based on mainframe data service.
## Architecture
<!--add an image in this path-->
<img src="https://raw.github.ibm.com/zhuoling/BatchJobAnalytics/contentedit/Image/architecture.png?token=AAAKI7J2PCW34LLH3VETWYK6QRBDI">
## What is included ?
**The project includes the following folders and files:**
### Data
This folder contains the following CSV files to be used as input data:
* `df_smf.csv` contains sample output of `1_BatchJob_SMF30Extract.ipynb`. The output is the batch job run time metrics that will be the most important input data for analysis.<br>
SMF provides a common interface a z/OS operation log for extracting system operation measurements. SMF Type 30 includes records of batch job operations. <br>
* `MasterBatchJob.csv` contains sample data of MBJ lapsed time over the period of one year. MBJ elapsed time refers to the minutes between the start time of the first job and the end time of the last job in MBJ recorded in `df_smf.csv`.
* `TxnVolume.csv` contains information about transaction volumes of various business types over the period of one year.
* `calendar_join.csv` contains calendar data with "calenday" elements like weekday, day, month, etc.
* `widetable_MBJ.csv` contains a joined wide table of MBJ elapsed time, transaction volumes, and calendar data, ready for model training in `3_BatchJob_MBJ_Prediction.ipynb`.
### Notebook
This folder contains the following Python notebooks. You can open and run each of them in your Jupyter web environment.
* `0_readme.ipynb` provides an overview of this sample project.
* `1_BatchJob_SMF30Extract.ipynb` extracts batch job run time operation data from SMF Type 30 log files. See IBM Knowledge Center for information about SMF Type 30 records. Because SMF Type 30 log files are usually large in size, they are not included in the project package. Only a small sample output from this `df_smf.csv` is included for your information.
* `2_BatchJob_MBJ_DataExploration.ipynb` explores MBJ elapsed time, gains insights on trend in timeline, and examines correlation between daytime business volumes and periodicity in day, week, and month.
* `3_BatchJob_MBJ_Prediction.ipynb` applies different methods for predicting MBJ elapsed time that is based on historical data, calendar information, or business transaction volume data.<br>
Note: In WMLz, reading a local data set in a notebook cell requires an authentication token. A WMLz administrator sets the expiration date of the token. In case of token expiration and authentication failure, click the top right button in the UI to replace the expired token.
### Flow
This folder contains the following SPSS modeler flows. You can open them in canvas and use drop-and-draw to explore.
* `4_BatchJob_MBJ_TSPredict.str` applies the time series algorithm to predict MBJ elapsed time.
* `5_BatchJob_MBJ_AnomalyDetect.str` detects anomaly in elapsed time and business transaction volumes.
### ProjectZIP
This folder contains compressed files `BatchJobAnalytics.zip` and `BatchJobAnalytics.tar.gz`. The content of the two files is the same, with `.zip` to be used on Windows or Mac and `tar.gz` on Linux. You can import one of the files to add the sample project to your WMLz workspace. Adding the project will also add the same Python notebooks and data sets.
### View
This folder contains the following output files of the sample project:
* Four read-only html files that are the output from the Python notebooks.
* Four screenshots from modeler flows that are prediction results and records of detected abnormal job instances.
# Related links
<a href="https://developer.ibm.com/patterns/analyze-batch-job-with-watson-machine-learning-for-zos"> Overview: Analyze Batch Jobs on IBM mainframe with machine learning</a><p>
<a href="https://github.com/IBM/analyze-batch-job-z"> GitHub: Analyze Batch Jobs via Watson Machine Learning on z/OS </a><p>
## Backgroud context
<a href="https://www.ibm.com/us-en/marketplace/machine-learning-for-zos">IBM Watson Machine Learning for z/OS </a><p>
<a href="http://www.redbooks.ibm.com/abstracts/sg248421.html?Open">Turning Data into Insight with IBM Machine Learning for z/OS </a><p>
<a href="https://www.ibm.com/support/knowledgecenter/zosbasics/com.ibm.zos.zmainframe/zconc_batchproc.htm">Mainframes working after hours: Batch processing </a><p>
<a href="https://en.wikipedia.org/wiki/IBM_System_Management_Facilities">IBM System Management Facilities </a><p>
<a href="https://www.ibm.com/support/knowledgecenter/en/SSLTBW_2.3.0/com.ibm.zos.v2r3.ieag200/rec30.htm">IBM SMF Type 30 record </a><p>
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.