code stringlengths 2.5k 150k | kind stringclasses 1 value |
|---|---|
```
import exmp
import qiime2
import tempfile
import os.path
import pandas as pd
from qiime2.plugins.feature_table.methods import filter_samples
from qiime2.plugins.taxa.methods import collapse
```
# EXMP 1
```
taxonomy = exmp.load_taxonomy()
sample_metadata = exmp.load_sample_metadata()
data_dir = exmp.cm_path
rarefied_table = qiime2.Artifact.load(os.path.join(data_dir, "rarefied_table.qza"))
uu_dm = qiime2.Artifact.load(os.path.join(data_dir, "unweighted_unifrac_distance_matrix.qza"))
wu_dm = qiime2.Artifact.load(os.path.join(data_dir, "weighted_unifrac_distance_matrix.qza"))
faith_pd = qiime2.Artifact.load(os.path.join(data_dir, "faith_pd_vector.qza"))
shannon = qiime2.Artifact.load(os.path.join(data_dir, "shannon_vector.qza"))
evenness = qiime2.Artifact.load(os.path.join(data_dir, "evenness_vector.qza"))
with tempfile.TemporaryDirectory() as output_dir:
_, _, _, sample_metadata = exmp.ols_and_anova('RER_change', 'exmp1', '1.0',
output_dir, 'week',
sample_metadata, uu_dm, wu_dm,
faith_pd, shannon, evenness)
rarefied_table = filter_samples(table=rarefied_table, metadata=sample_metadata).filtered_table
taxa_table = collapse(table=rarefied_table, taxonomy=taxonomy, level=6).collapsed_table.view(pd.DataFrame)
sample_metadata = sample_metadata.to_dataframe()
sorted_wu_pc3_correlations = pd.DataFrame(taxa_table.corrwith(sample_metadata['Weighted_UniFrac_PC3'], method='spearman').sort_values(), columns=['Spearman rho'])
sorted_wu_pc3_correlations['25th percentile rarefied count'] = taxa_table[sorted_wu_pc3_correlations.index].quantile(0.25)
sorted_wu_pc3_correlations['Median rarefied count'] = taxa_table[sorted_wu_pc3_correlations.index].quantile(0.50)
sorted_wu_pc3_correlations['75th percentile rarefied count'] = taxa_table[sorted_wu_pc3_correlations.index].quantile(0.75)
```
The data are most easily interpreted if the ordination axes are positively correlated with the RER change. Since the direction of the PCs are arbitrary, I generally just run this a few times till I get a positive correlation.
```
sample_metadata['Weighted_UniFrac_PC3'].corr(sample_metadata['RER_change'])
output_dir = os.path.join(exmp.cm_path, 'ols-and-anova', 'exmp1-RER_change-week1.0')
sorted_wu_pc3_correlations.to_csv(open(os.path.join(output_dir, 'wu-pcoa3-genus-correlations.csv'), 'w'))
```
# EXMP 2
```
taxonomy = exmp.load_taxonomy()
sample_metadata = exmp.load_sample_metadata()
data_dir = exmp.cm_path
rarefied_table = qiime2.Artifact.load(os.path.join(data_dir, "rarefied_table.qza"))
uu_dm = qiime2.Artifact.load(os.path.join(data_dir, "unweighted_unifrac_distance_matrix.qza"))
wu_dm = qiime2.Artifact.load(os.path.join(data_dir, "weighted_unifrac_distance_matrix.qza"))
faith_pd = qiime2.Artifact.load(os.path.join(data_dir, "faith_pd_vector.qza"))
shannon = qiime2.Artifact.load(os.path.join(data_dir, "shannon_vector.qza"))
evenness = qiime2.Artifact.load(os.path.join(data_dir, "evenness_vector.qza"))
with tempfile.TemporaryDirectory() as output_dir:
_, _, _, sample_metadata = exmp.ols_and_anova('three_rep_max_squat_change', 'exmp2', '1.0',
output_dir, 'week',
sample_metadata, uu_dm, wu_dm,
faith_pd, shannon, evenness)
rarefied_table = filter_samples(table=rarefied_table, metadata=sample_metadata).filtered_table
taxa_table = collapse(table=rarefied_table, taxonomy=taxonomy, level=6).collapsed_table.view(pd.DataFrame)
sample_metadata = sample_metadata.to_dataframe()
sorted_wu_pc2_correlations = pd.DataFrame(taxa_table.corrwith(sample_metadata['Weighted_UniFrac_PC2'], method='spearman').sort_values(), columns=['Spearman rho'])
sorted_wu_pc2_correlations['25th percentile rarefied count'] = taxa_table[sorted_wu_pc2_correlations.index].quantile(0.25)
sorted_wu_pc2_correlations['Median rarefied count'] = taxa_table[sorted_wu_pc2_correlations.index].quantile(0.50)
sorted_wu_pc2_correlations['75th percentile rarefied count'] = taxa_table[sorted_wu_pc2_correlations.index].quantile(0.75)
sorted_wu_pc3_correlations = pd.DataFrame(taxa_table.corrwith(sample_metadata['Weighted_UniFrac_PC3'], method='spearman').sort_values(), columns=['Spearman rho'])
sorted_wu_pc3_correlations['25th percentile rarefied count'] = taxa_table[sorted_wu_pc3_correlations.index].quantile(0.25)
sorted_wu_pc3_correlations['Median rarefied count'] = taxa_table[sorted_wu_pc3_correlations.index].quantile(0.50)
sorted_wu_pc3_correlations['75th percentile rarefied count'] = taxa_table[sorted_wu_pc3_correlations.index].quantile(0.75)
sample_metadata['Weighted_UniFrac_PC2'].corr(sample_metadata['three_rep_max_squat_change'])
sample_metadata['Weighted_UniFrac_PC3'].corr(sample_metadata['three_rep_max_squat_change'])
output_dir = os.path.join(exmp.cm_path, 'ols-and-anova', 'exmp2-three_rep_max_squat_change-week1.0')
sorted_wu_pc2_correlations.to_csv(open(os.path.join(output_dir, 'wu-pcoa2-genus-correlations.csv'), 'w'))
sorted_wu_pc3_correlations.to_csv(open(os.path.join(output_dir, 'wu-pcoa3-genus-correlations.csv'), 'w'))
```
| github_jupyter |
```
import numpy
import urllib
import scipy.optimize
import random
from math import *
def parseData(fname):
for l in urllib.urlopen(fname):
yield eval(l)
print "Reading data..."
data = list(parseData("file:beer_50000.json"))
print "done"
def feature(datum):
text = datum['review/text'].lower().replace(',',' ').replace('?',' ')\
.replace('!',' ').replace(':',' ').replace('"',' ').replace('.',' ')\
.replace('(',' ').replace(')',' ').split()
num_lactic = 0
num_tart = 0
num_sour = 0
num_citric = 0
num_sweet = 0
num_acid = 0
num_hop = 0
num_fruit = 0
num_salt = 0
num_spicy = 0
for word in text:
if word == 'lactic': num_lactic += 1
if word == 'tart': num_tart += 1
if word == 'sour': num_sour += 1
if word == 'citric': num_citric += 1
if word == 'sweet': num_sweet += 1
if word == 'acid': num_acid += 1
if word == 'hop': num_hop += 1
if word == 'fruit': num_fruit += 1
if word == 'salt': num_salt += 1
if word == 'spicy': num_spicy += 1
feat = [1, num_lactic, num_tart, num_sour, \
num_citric, num_sweet, num_acid, num_hop, \
num_fruit, num_salt, num_spicy]
return feat
X = [feature(d) for d in data]
y = [d['beer/ABV'] >= 6.5 for d in data]
def inner(x,y):
return sum([x[i]*y[i] for i in range(len(x))])
def sigmoid(x):
res = 1.0 / (1 + exp(-x))
return res
length = int(len(data)/3)
X_train = X[:length]
y_train = y[:length]
X_validation = X[length:2*length]
y_validation = y[length:2*length]
X_test = X[2*length:]
y_test = y[2*length:]
# Count for number of total data, y=0 and y=1
num_total = len(y_train)
num_y0 = y_train.count(0)
num_y1 = y_train.count(1)
# NEGATIVE Log-likelihood
def f(theta, X, y, lam):
loglikelihood = 0
for i in range(len(X)):
logit = inner(X[i], theta)
if y[i]:
loglikelihood -= log(1 + exp(-logit)) * num_total / (2 * num_y1)
if not y[i]:
loglikelihood -= (log(1 + exp(-logit)) + logit ) * num_total / (2 * num_y0)
for k in range(len(theta)):
loglikelihood -= lam * theta[k]*theta[k]
# for debugging
# print("ll =" + str(loglikelihood))
return -loglikelihood
# NEGATIVE Derivative of log-likelihood
def fprime(theta, X, y, lam):
dl = [0]*len(theta)
for i in range(len(X)):
logit = inner(X[i], theta)
for k in range(len(theta)):
if y[i]:
dl[k] += X[i][k] * (1 - sigmoid(logit)) * num_total / (2 * num_y1)
if not y[i]:
dl[k] -= X[i][k] * (1 - sigmoid(logit)) * num_total / (2 * num_y0)
for k in range(len(theta)):
dl[k] -= lam*2*theta[k]
return numpy.array([-x for x in dl])
def train(lam):
theta,_,_ = scipy.optimize.fmin_l_bfgs_b(f, [0]*len(X[0]), fprime, pgtol = 10, args = (X_train, y_train, lam))
return theta
lam = 1.0
theta = train(lam)
print theta
X_data = [X_train, X_validation, X_test]
y_data = [y_train, y_validation, y_test]
symbol = ['train', 'valid', 'test']
print 'λ\tDataset\t\tTruePositive\tFalsePositive\tTrueNegative\tFalseNegative\tAccuracy\tBER'
for i in range(3):
def TP(theta):
scores = [inner(theta,x) for x in X_data[i]]
predictions = [s > 0 for s in scores]
correct = [((a==1) and (b==1)) for (a,b) in zip(predictions,y_data[i])]
tp = sum(correct) * 1.0
return tp
def TN(theta):
scores = [inner(theta,x) for x in X_data[i]]
predictions = [s > 0 for s in scores]
correct = [((a==0) and (b==0)) for (a,b) in zip(predictions,y_data[i])]
tn = sum(correct) * 1.0
return tn
def FP(theta):
scores = [inner(theta,x) for x in X_data[i]]
predictions = [s > 0 for s in scores]
correct = [((a==1) and (b==0)) for (a,b) in zip(predictions,y_data[i])]
fp = sum(correct) * 1.0
return fp
def FN(theta):
scores = [inner(theta,x) for x in X_data[i]]
predictions = [s > 0 for s in scores]
correct = [((a==0) and (b==1)) for (a,b) in zip(predictions,y_data[i])]
fn = sum(correct) * 1.0
return fn
tp = TP(theta)
fp = FP(theta)
tn = TN(theta)
fn = FN(theta)
TPR = tp / (tp + fn)
TNR = tn / (tn + fp)
BER = 1 - 0.5 * (TPR + TNR)
accuracy = (tp+tn)/(tp+tn+fp+fn)
print str(lam)+'\t'+symbol[i]+'\t\t'+str(tp)+'\t\t'+str(fp)+'\t\t'+str(tn)+'\t\t'+str(fn)+'\t\t'+str(accuracy)+'\t'+str(BER)
# Original Algorithm
# NEGATIVE Log-likelihood
def f(theta, X, y, lam):
loglikelihood = 0
for i in range(len(X)):
logit = inner(X[i], theta)
loglikelihood -= log(1 + exp(-logit))
if not y[i]:
loglikelihood -= logit
for k in range(len(theta)):
loglikelihood -= lam * theta[k]*theta[k]
# for debugging
# print("ll =" + str(loglikelihood))
return -loglikelihood
# NEGATIVE Derivative of log-likelihood
def fprime(theta, X, y, lam):
dl = [0]*len(theta)
for i in range(len(X)):
logit = inner(X[i], theta)
for k in range(len(theta)):
dl[k] += X[i][k] * (1 - sigmoid(logit))
if not y[i]:
dl[k] -= X[i][k]
for k in range(len(theta)):
dl[k] -= lam*2*theta[k]
return numpy.array([-x for x in dl])
def train(lam):
theta,_,_ = scipy.optimize.fmin_l_bfgs_b(f, [0]*len(X[0]), fprime, pgtol = 10, args = (X_train, y_train, lam))
return theta
lam = 1.0
theta = train(lam)
X_data = [X_train, X_validation, X_test]
y_data = [y_train, y_validation, y_test]
symbol = ['train', 'valid', 'test']
print 'λ\tDataset\t\tTruePositive\tFalsePositive\tTrueNegative\tFalseNegative\tAccuracy\tBER'
for i in range(3):
def TP(theta):
scores = [inner(theta,x) for x in X_data[i]]
predictions = [s > 0 for s in scores]
correct = [((a==1) and (b==1)) for (a,b) in zip(predictions,y_data[i])]
tp = sum(correct) * 1.0
return tp
def TN(theta):
scores = [inner(theta,x) for x in X_data[i]]
predictions = [s > 0 for s in scores]
correct = [((a==0) and (b==0)) for (a,b) in zip(predictions,y_data[i])]
tn = sum(correct) * 1.0
return tn
def FP(theta):
scores = [inner(theta,x) for x in X_data[i]]
predictions = [s > 0 for s in scores]
correct = [((a==1) and (b==0)) for (a,b) in zip(predictions,y_data[i])]
fp = sum(correct) * 1.0
return fp
def FN(theta):
scores = [inner(theta,x) for x in X_data[i]]
predictions = [s > 0 for s in scores]
correct = [((a==0) and (b==1)) for (a,b) in zip(predictions,y_data[i])]
fn = sum(correct) * 1.0
return fn
tp = TP(theta)
fp = FP(theta)
tn = TN(theta)
fn = FN(theta)
TPR = tp / (tp + fn)
TNR = tn / (tn + fp)
BER = 1 - 0.5 * (TPR + TNR)
accuracy = (tp+tn)/(tp+tn+fp+fn)
print str(lam)+'\t'+symbol[i]+'\t\t'+str(tp)+'\t\t'+str(fp)+'\t\t'+str(tn)+'\t\t'+str(fn)+'\t\t'+str(accuracy)+'\t'+str(BER)
```
| github_jupyter |
# Strategies
High-performance solvers, such as Z3, contain many tightly integrated, handcrafted heuristic combinations of algorithmic proof methods. While these heuristic combinations tend to be highly tuned for known classes of problems, they may easily perform very badly on new classes of problems. This issue is becoming increasingly pressing as solvers begin to gain the attention of practitioners in diverse areas of science and engineering. In many cases, changes to the solver heuristics can make a tremendous difference.
More information on Z3 is available from https://github.com/z3prover/z3.git
## Introduction
Z3 implements a methodology for orchestrating reasoning engines where "big" symbolic reasoning steps are represented as functions known as tactics, and tactics are composed using combinators known as tacticals. Tactics process sets of formulas called Goals.
When a tactic is applied to some goal G, four different outcomes are possible. The tactic succeeds in showing G to be satisfiable (i.e., feasible); succeeds in showing G to be unsatisfiable (i.e., infeasible); produces a sequence of subgoals; or fails. When reducing a goal G to a sequence of subgoals G1, ..., Gn, we face the problem of model conversion. A model converter construct a model for G using a model for some subgoal Gi.
In the following example, we create a goal g consisting of three formulas, and a tactic t composed of two built-in tactics: simplify and solve-eqs. The tactic simplify apply transformations equivalent to the ones found in the command simplify. The tactic solver-eqs eliminate variables using Gaussian elimination. Actually, solve-eqs is not restricted only to linear arithmetic. It can also eliminate arbitrary variables. Then, combinator Then applies simplify to the input goal and solve-eqs to each subgoal produced by simplify. In this example, only one subgoal is produced.
```
!pip install "z3-solver"
from z3 import *
x, y = Reals('x y')
g = Goal()
g.add(x > 0, y > 0, x == y + 2)
print(g)
t1 = Tactic('simplify')
t2 = Tactic('solve-eqs')
t = Then(t1, t2)
print(t(g))
```
In the example above, variable x is eliminated, and is not present the resultant goal.
In Z3, we say a clause is any constraint of the form Or(f_1, ..., f_n). The tactic split-clause will select a clause Or(f_1, ..., f_n) in the input goal, and split it n subgoals. One for each subformula f_i.
```
x, y = Reals('x y')
g = Goal()
g.add(Or(x < 0, x > 0), x == y + 1, y < 0)
t = Tactic('split-clause')
r = t(g)
for g in r:
print(g)
```
Tactics
Z3 comes equipped with many built-in tactics. The command describe_tactics() provides a short description of all built-in tactics.
describe_tactics()
Z3Py comes equipped with the following tactic combinators (aka tacticals):
* Then(t, s) applies t to the input goal and s to every subgoal produced by t.
* OrElse(t, s) first applies t to the given goal, if it fails then returns the result of s applied to the given goal.
* Repeat(t) Keep applying the given tactic until no subgoal is modified by it.
* Repeat(t, n) Keep applying the given tactic until no subgoal is modified by it, or the number of iterations is greater than n.
* TryFor(t, ms) Apply tactic t to the input goal, if it does not return in ms milliseconds, it fails.
* With(t, params) Apply the given tactic using the given parameters.
The following example demonstrate how to use these combinators.
```
x, y, z = Reals('x y z')
g = Goal()
g.add(Or(x == 0, x == 1),
Or(y == 0, y == 1),
Or(z == 0, z == 1),
x + y + z > 2)
# Split all clauses"
split_all = Repeat(OrElse(Tactic('split-clause'),
Tactic('skip')))
print(split_all(g))
split_at_most_2 = Repeat(OrElse(Tactic('split-clause'),
Tactic('skip')),
1)
print(split_at_most_2(g))
# Split all clauses and solve equations
split_solve = Then(Repeat(OrElse(Tactic('split-clause'),
Tactic('skip'))),
Tactic('solve-eqs'))
print(split_solve(g))
```
In the tactic split_solver, the tactic solve-eqs discharges all but one goal. Note that, this tactic generates one goal: the empty goal which is trivially satisfiable (i.e., feasible)
The list of subgoals can be easily traversed using the Python for statement.
```
x, y, z = Reals('x y z')
g = Goal()
g.add(Or(x == 0, x == 1),
Or(y == 0, y == 1),
Or(z == 0, z == 1),
x + y + z > 2)
# Split all clauses"
split_all = Repeat(OrElse(Tactic('split-clause'),
Tactic('skip')))
for s in split_all(g):
print(s)
```
A tactic can be converted into a solver object using the method solver(). If the tactic produces the empty goal, then the associated solver returns sat. If the tactic produces a single goal containing False, then the solver returns unsat. Otherwise, it returns unknown.
```
bv_solver = Then('simplify',
'solve-eqs',
'bit-blast',
'sat').solver()
x, y = BitVecs('x y', 16)
solve_using(bv_solver, x | y == 13, x > y)
```
In the example above, the tactic bv_solver implements a basic bit-vector solver using equation solving, bit-blasting, and a propositional SAT solver. Note that, the command Tactic is suppressed. All Z3Py combinators automatically invoke Tactic command if the argument is a string. Finally, the command solve_using is a variant of the solve command where the first argument specifies the solver to be used.
In the following example, we use the solver API directly instead of the command solve_using. We use the combinator With to configure our little solver. We also include the tactic aig which tries to compress Boolean formulas using And-Inverted Graphs.
```
bv_solver = Then(With('simplify', mul2concat=True),
'solve-eqs',
'bit-blast',
'aig',
'sat').solver()
x, y = BitVecs('x y', 16)
bv_solver.add(x*32 + y == 13, x & y < 10, y > -100)
print(bv_solver.check())
m = bv_solver.model()
print(m)
print(x*32 + y, "==", m.evaluate(x*32 + y))
print(x & y, "==", m.evaluate(x & y))
```
The tactic smt wraps the main solver in Z3 as a tactic.
```
x, y = Ints('x y')
s = Tactic('smt').solver()
s.add(x > y + 1)
print(s.check())
print(s.model())
```
Now, we show how to implement a solver for integer arithmetic using SAT. The solver is complete only for problems where every variable has a lower and upper bound.
```
s = Then(With('simplify', arith_lhs=True, som=True),
'normalize-bounds', 'lia2pb', 'pb2bv',
'bit-blast', 'sat').solver()
x, y, z = Ints('x y z')
solve_using(s,
x > 0, x < 10,
y > 0, y < 10,
z > 0, z < 10,
3*y + 2*x == z)
# It fails on the next example (it is unbounded)
s.reset()
solve_using(s, 3*y + 2*x == z)
```
Tactics can be combined with solvers. For example, we can apply a tactic to a goal, produced a set of subgoals, then select one of the subgoals and solve it using a solver. The next example demonstrates how to do that, and how to use model converters to convert a model for a subgoal into a model for the original goal.
```
t = Then('simplify',
'normalize-bounds',
'solve-eqs')
x, y, z = Ints('x y z')
g = Goal()
g.add(x > 10, y == x + 3, z > y)
r = t(g)
# r contains only one subgoal
print(r)
s = Solver()
s.add(r[0])
print(s.check())
# Model for the subgoal
print(s.model())
# Model for the original goal
print(r[0].convert_model(s.model()))
```
## Probes
Probes (aka formula measures) are evaluated over goals. Boolean expressions over them can be built using relational operators and Boolean connectives. The tactic FailIf(cond) fails if the given goal does not satisfy the condition cond. Many numeric and Boolean measures are available in Z3Py. The command describe_probes() provides the list of all built-in probes.
```
describe_probes()
```
In the following example, we build a simple tactic using FailIf. It also shows that a probe can be applied directly to a goal.
```
x, y, z = Reals('x y z')
g = Goal()
g.add(x + y + z > 0)
p = Probe('num-consts')
print("num-consts:", p(g))
t = FailIf(p > 2)
try:
t(g)
except Z3Exception:
print("tactic failed")
print("trying again...")
g = Goal()
g.add(x + y > 0)
print(t(g))
```
Z3Py also provides the combinator (tactical) If(p, t1, t2) which is a shorthand for:
OrElse(Then(FailIf(Not(p)), t1), t2)
The combinator When(p, t) is a shorthand for:
If(p, t, 'skip')
The tactic skip just returns the input goal. The following example demonstrates how to use the If combinator.
```
x, y, z = Reals('x y z')
g = Goal()
g.add(x**2 - y**2 >= 0)
p = Probe('num-consts')
t = If(p > 2, 'simplify', 'factor')
print(t(g))
g = Goal()
g.add(x + x + y + z >= 0, x**2 - y**2 >= 0)
print(t(g))
```
| github_jupyter |
### Specify a text string to examine with NEMO
```
# specify query string
payload = 'The World Health Organization on Sunday reported the largest single-day increase in coronavirus cases by its count, at more than 183,000 new cases in the latest 24 hours. The UN health agency said Brazil led the way with 54,771 cases tallied and the U.S. next at 36,617. Over 15,400 came in in India.'
payload = 'is strongly affected by large ground-water withdrawals at or near Tupelo, Aberdeen, and West Point.'
# payload = 'Overall design: Teliospores of pathogenic races T-1, T-5 and T-16 of T. caries provided by a collection in Aberdeen, ID, USA'
payload = 'The results provide evidence of substantial population structure in C. posadasii and demonstrate presence of distinct geographic clades in Central and Southern Arizona as well as dispersed populations in Texas, Mexico and South and Central America'
payload = 'Most frequent numerical abnormalities in B-NHL were gains of chromosomes 3 and 18, although gains of chromosome 3 were less prominent in FL.'
```
### Load functions
```
# import credentials file
import yaml
with open("config.yml", 'r') as ymlfile:
cfg = yaml.safe_load(ymlfile)
# general way to extract values for a given key. Returns an array. Used to parse Nemo response and extract wikipedia id
# from https://hackersandslackers.com/extract-data-from-complex-json-python/
def extract_values(obj, key):
"""Pull all values of specified key from nested JSON."""
arr = []
def extract(obj, arr, key):
"""Recursively search for values of key in JSON tree."""
if isinstance(obj, dict):
for k, v in obj.items():
if isinstance(v, (dict, list)):
extract(v, arr, key)
elif k == key:
arr.append(v)
elif isinstance(obj, list):
for item in obj:
extract(item, arr, key)
return arr
results = extract(obj, arr, key)
return results
# getting wikipedia ID
# see he API at https://www.mediawiki.org/wiki/API:Query#Example_5:_Batchcomplete
# also, https://stackoverflow.com/questions/37024807/how-to-get-wikidata-id-for-an-wikipedia-article-by-api
def get_WPID (name):
import json
url = 'https://en.wikipedia.org/w/api.php?action=query&prop=pageprops&ppprop=wikibase_item&redirects=1&format=json&titles=' +name
r=requests.get(url).json()
return extract_values(r,'wikibase_item')
```
### Send a request to NEMO, and get a response
```
# make a service request
import requests
# payloadutf = payload.encode('utf-8')
url = "https://nemoservice.azurewebsites.net/nemo?appid=" + cfg['api_creds']['nmo1']
newHeaders = {'Content-type': 'application/json', 'Accept': 'text/plain'}
response = requests.post(url,
data='"{' + payload + '}"',
headers=newHeaders)
# display the results as string (remove json braces)
a = response.content.decode()
resp_full = a[a.find('{')+1 : a.find('}')]
resp_full
```
### Parse the response and load all found elements into a dataframe
```
# create a dataframe with entities, remove duplicates, then add wikipedia/wikidata concept IDs
import pandas as pd
import re
import xml.etree.ElementTree as ET
df = pd.DataFrame(columns=["Type","Ref","EntityType","Name","Form","WP","Value","Alt","WP_ID"])
# note that the last column is to be populated later, via Wikipedia API
# all previous columns are from Nemo: based on "e" (entity) and "d" (data) elements. "c" (concept) to be explored
# get starting and ending positions of xml fragments in the Nemo output
pattern_start = "<(e|d|c)\s"
iter = re.finditer(pattern_start,resp_full)
indices1 = [m.start(0) for m in iter]
pattern_end = "</(e|d|c)>"
iter = re.finditer(pattern_end,resp_full)
indices2 = [m.start(0) for m in iter]
# iterate over xml fragments returned by Nemo, extracting attributes from each and adding to dataframe
for i, entity in enumerate(indices1):
a = resp_full[indices1[i] : indices2[i]+4]
root = ET.fromstring(a)
tag = root.tag
attributes = root.attrib
df = df.append({"Type":root.tag,
"Ref":attributes.get('ref'),
"EntityType":attributes.get('type'),
"Name":attributes.get('name'),
"Form":attributes.get('form'),
"WP":attributes.get('wp'),
"Value":attributes.get('value'),
"Alt":attributes.get('alt')},
ignore_index=True)
```
E stands for entity;
the attribute ref gives you the title of the corresponding Wikipedia page when the attribute wp has the value “y”;
the attribute type gives you the type of entity for known entities; the types of interest for you are G, which is geo-political entity, L – geographic form/location (such as a mountain), and F, which is facility (such as an airport).
D stands for datafield, which comprises dates, NUMEX, email addresses and URLs, tracking numbers, and so on.
C stands for concept; these appear in Wikipedia and are deemed as relevant for the input text, but they do not get disambiguated
```
# remove duplicate records from the df
df = df.drop_duplicates(keep='first')
# for each found entity, add wikidata unique identifiers to the dataframe
for index, row in df.iterrows():
if (row['WP']=='y'):
row['WP_ID'] = get_WPID(row['Name'])[0]
df
```
| github_jupyter |
# Audiobooks business case
## Preprocessing exercise
It makes sense to shuffle the indices prior to balancing the dataset.
Using the code from the lesson (below), shuffle the indices and then balance the dataset.
At the end of the course, you will have an exercise to create the same machine learning algorithm, with preprocessing done in this way.
Note: This is more of a programming exercise rather than a machine learning one. Being able to complete it successfully will ensure you understand the preprocessing.
Good luck!
**Solution:**
Scroll down to the 'Exercise Solution' section
### Extract the data from the csv
```
import numpy as np
# We will use the sklearn preprocessing library, as it will be easier to standardize the data.
from sklearn import preprocessing
# Load the data
raw_csv_data = np.loadtxt('Audiobooks_data.csv',delimiter=',')
# The inputs are all columns in the csv, except for the first one [:,0]
# (which is just the arbitrary customer IDs that bear no useful information),
# and the last one [:,-1] (which is our targets)
unscaled_inputs_all = raw_csv_data[:,1:-1]
# The targets are in the last column. That's how datasets are conventionally organized.
targets_all = raw_csv_data[:,-1]
```
### EXERCISE SOLUTION
We shuffle the indices before balancing (to remove any day effects, etc.)
However, we still have to shuffle them AFTER we balance the dataset as otherwise, all targets that are 1s will be contained in the train_targets.
This code is suboptimal, but is the easiest way to complete the exercise. Still, as we do the preprocessing only once, speed in not something we are aiming for.
We record the variables in themselves, so we don't amend the code that follows.
```
# When the data was collected it was actually arranged by date
# Shuffle the indices of the data, so the data is not arranged in any way when we feed it.
# Since we will be batching, we want the data to be as randomly spread out as possible
shuffled_indices = np.arange(unscaled_inputs_all.shape[0])
np.random.shuffle(shuffled_indices)
# Use the shuffled indices to shuffle the inputs and targets.
unscaled_inputs_all = unscaled_inputs_all[shuffled_indices]
targets_all = targets_all[shuffled_indices]
```
### Balance the dataset
```
# Count how many targets are 1 (meaning that the customer did convert)
num_one_targets = int(np.sum(targets_all))
# Set a counter for targets that are 0 (meaning that the customer did not convert)
zero_targets_counter = 0
# We want to create a "balanced" dataset, so we will have to remove some input/target pairs.
# Declare a variable that will do that:
indices_to_remove = []
# Count the number of targets that are 0.
# Once there are as many 0s as 1s, mark entries where the target is 0.
for i in range(targets_all.shape[0]):
if targets_all[i] == 0:
zero_targets_counter += 1
if zero_targets_counter > num_one_targets:
indices_to_remove.append(i)
# Create two new variables, one that will contain the inputs, and one that will contain the targets.
# We delete all indices that we marked "to remove" in the loop above.
unscaled_inputs_equal_priors = np.delete(unscaled_inputs_all, indices_to_remove, axis=0)
targets_equal_priors = np.delete(targets_all, indices_to_remove, axis=0)
```
### Standardize the inputs
```
# That's the only place we use sklearn functionality. We will take advantage of its preprocessing capabilities
# It's a simple line of code, which standardizes the inputs, as we explained in one of the lectures.
# At the end of the business case, you can try to run the algorithm WITHOUT this line of code.
# The result will be interesting.
scaled_inputs = preprocessing.scale(unscaled_inputs_equal_priors)
```
### Shuffle the data
```
# When the data was collected it was actually arranged by date
# Shuffle the indices of the data, so the data is not arranged in any way when we feed it.
# Since we will be batching, we want the data to be as randomly spread out as possible
shuffled_indices = np.arange(scaled_inputs.shape[0])
np.random.shuffle(shuffled_indices)
# Use the shuffled indices to shuffle the inputs and targets.
shuffled_inputs = scaled_inputs[shuffled_indices]
shuffled_targets = targets_equal_priors[shuffled_indices]
```
### Split the dataset into train, validation, and test
```
# Count the total number of samples
samples_count = shuffled_inputs.shape[0]
# Count the samples in each subset, assuming we want 80-10-10 distribution of training, validation, and test.
# Naturally, the numbers are integers.
train_samples_count = int(0.8 * samples_count)
validation_samples_count = int(0.1 * samples_count)
# The 'test' dataset contains all remaining data.
test_samples_count = samples_count - train_samples_count - validation_samples_count
# Create variables that record the inputs and targets for training
# In our shuffled dataset, they are the first "train_samples_count" observations
train_inputs = shuffled_inputs[:train_samples_count]
train_targets = shuffled_targets[:train_samples_count]
# Create variables that record the inputs and targets for validation.
# They are the next "validation_samples_count" observations, folllowing the "train_samples_count" we already assigned
validation_inputs = shuffled_inputs[train_samples_count:train_samples_count+validation_samples_count]
validation_targets = shuffled_targets[train_samples_count:train_samples_count+validation_samples_count]
# Create variables that record the inputs and targets for test.
# They are everything that is remaining.
test_inputs = shuffled_inputs[train_samples_count+validation_samples_count:]
test_targets = shuffled_targets[train_samples_count+validation_samples_count:]
# We balanced our dataset to be 50-50 (for targets 0 and 1), but the training, validation, and test were
# taken from a shuffled dataset. Check if they are balanced, too. Note that each time you rerun this code,
# you will get different values, as each time they are shuffled randomly.
# Normally you preprocess ONCE, so you need not rerun this code once it is done.
# If you rerun this whole sheet, the npzs will be overwritten with your newly preprocessed data.
# Print the number of targets that are 1s, the total number of samples, and the proportion for training, validation, and test.
print(np.sum(train_targets), train_samples_count, np.sum(train_targets) / train_samples_count)
print(np.sum(validation_targets), validation_samples_count, np.sum(validation_targets) / validation_samples_count)
print(np.sum(test_targets), test_samples_count, np.sum(test_targets) / test_samples_count)
```
### Save the three datasets in *.npz
```
# Save the three datasets in *.npz.
# In the next lesson, you will see that it is extremely valuable to name them in such a coherent way!
np.savez('Audiobooks_data_train', inputs=train_inputs, targets=train_targets)
np.savez('Audiobooks_data_validation', inputs=validation_inputs, targets=validation_targets)
np.savez('Audiobooks_data_test', inputs=test_inputs, targets=test_targets)
```
| github_jupyter |
This notebook will cover the assumed knowledge of pandas.
Here's a few questions to check if you already know the material in this notebook.
1. Does a NumPy array have a single dtype or multiple dtypes?
2. Why is broadcasting useful?
3. How do you slice a DataFrame by row label?
4. How do you select a column of a DataFrame?
5. Is the Index a column in the DataFrame?
If you feel pretty comfortable with those, go ahead and skip this notebook.
[Answers](#Answers) are at the end. We'll meet up at the next notebook.
# Aside: IPython Notebook
- two modes command and edit
- command -> edit: `Enter`
- edit -> command: `Esc`
- `h` : Keyboard Shortcuts: (from command mode)
- `j` / `k` : navigate cells
- `shift+Enter` executes a cell
Outline:
- [NumPy Foundation](#NumPy-Foundation)
- [Pandas](#Pandas)
- [Data Structures](#Data-Structures)
## Numpy Foundation
pandas is built atop NumPy, historically and in the actual library.
It's helpful to have a good understanding of some NumPyisms. [Speak the vernacular](https://www.youtube.com/watch?v=u2yvNw49AX4).
### ndarray
The core of numpy is the `ndarray`, N-dimensional array. These are homogenously-typed, fixed-length data containers.
NumPy also provides many convenient and fast methods implemented on the `ndarray`.
```
from __future__ import print_function
import numpy as np
import pandas as pd
x = np.array([1, 2, 3])
x
x.dtype
y = np.array([[True, False], [False, True]])
y
y.shape
```
### dtypes
Unlike python lists, NumPy arrays care about the type of data stored within.
The full list of NumPy dtypes can be found in the [NumPy documentation](http://docs.scipy.org/doc/numpy/user/basics.types.html).
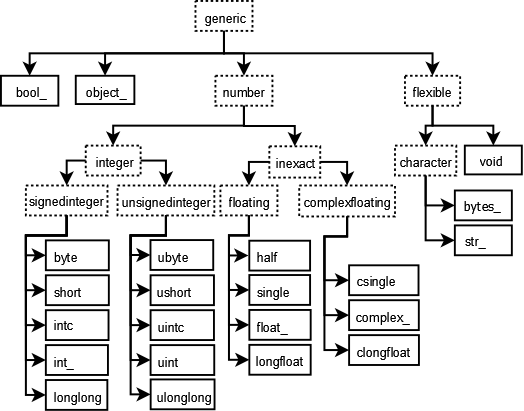
We sacrifice the convinience of mixing bools and ints and floats within an array for much better performance.
However, an unexpected `dtype` change will probably bite you at some point in the future.
The two biggest things to remember are
- Missing values (NaN) cast integer or boolean arrays to floats
- the object dtype is the fallback
You'll want to avoid object dtypes. It's typically slow.
### Broadcasting
It's super cool and super useful. The one-line explanation is that when doing elementwise operations, things expand to the "correct" shape.
```
# add a scalar to a 1-d array
x = np.arange(5)
print('x: ', x)
print('x+1:', x + 1, end='\n\n')
y = np.random.uniform(size=(2, 5))
print('y: ', y, sep='\n')
print('y+1:', y + 1, sep='\n')
```
Since `x` is shaped `(5,)` and `y` is shaped `(2,5)` we can do operations between them.
```
x * y
```
Without broadcasting we'd have to manually reshape our arrays, which quickly gets annoying.
```
x.reshape(1, 5).repeat(2, axis=0) * y
```
# Pandas
We'll breeze through the basics here, and get onto some interesting applications in a bit. I want to provide the *barest* of intuition so things stick down the road.
## Why pandas
NumPy is great. But it lacks a few things that are conducive to doing statisitcal analysis. By building on top of NumPy, pandas provides
- labeled arrays
- heterogenous data types within a table
- better missing data handling
- convenient methods
- more data types (Categorical, Datetime)
## Data Structures
This is the typical starting point for any intro to pandas.
We'll follow suit.
### The DataFrame
Here we have the workhorse data structure for pandas.
It's an in-memory table holding your data, and provides a few conviniences over lists of lists or NumPy arrays.
```
import numpy as np
import pandas as pd
# Many ways to construct a DataFrame
# We pass a dict of {column name: column values}
np.random.seed(42)
df = pd.DataFrame({'A': [1, 2, 3], 'B': [True, True, False],
'C': np.random.randn(3)},
index=['a', 'b', 'c']) # also this weird index thing
df
from IPython.display import Image
Image('dataframe.png')
```
### Selecting
Our first improvement over numpy arrays is labeled indexing. We can select subsets by column, row, or both. Column selection uses the regular python `__getitem__` machinery. Pass in a single column label `'A'` or a list of labels `['A', 'C']` to select subsets of the original `DataFrame`.
```
# Single column, reduces to a Series
df['A']
cols = ['A', 'C']
df[cols]
```
For row-wise selection, use the special `.loc` accessor.
```
df.loc[['a', 'b']]
```
When your index labels are ordered, you can use ranges to select rows or columns.
```
df.loc['a':'b']
```
Notice that the slice is *inclusive* on both sides, unlike your typical slicing of a list. Sometimes, you'd rather slice by *position* instead of label. `.iloc` has you covered:
```
df.iloc[0:2]
```
This follows the usual python slicing rules: closed on the left, open on the right.
As I mentioned, you can slice both rows and columns. Use `.loc` for label or `.iloc` for position indexing.
```
df.loc['a', 'B']
```
Pandas, like NumPy, will reduce dimensions when possible. Select a single column and you get back `Series` (see below). Select a single row and single column, you get a scalar.
You can get pretty fancy:
```
df.loc['a':'b', ['A', 'C']]
```
#### Summary
- Use `[]` for selecting columns
- Use `.loc[row_lables, column_labels]` for label-based indexing
- Use `.iloc[row_positions, column_positions]` for positional index
I've left out boolean and hierarchical indexing, which we'll see later.
## Series
You've already seen some `Series` up above. It's the 1-dimensional analog of the DataFrame. Each column in a `DataFrame` is in some sense a `Series`. You can select a `Series` from a DataFrame in a few ways:
```
# __getitem__ like before
df['A']
# .loc, like before
df.loc[:, 'A']
# using `.` attribute lookup
df.A
```
You'll have to be careful with the last one. It won't work if you're column name isn't a valid python identifier (say it has a space) or if it conflicts with one of the (many) methods on `DataFrame`. The `.` accessor is extremely convient for interactive use though.
You should never *assign* a column with `.` e.g. don't do
```python
# bad
df.A = [1, 2, 3]
```
It's unclear whether your attaching the list `[1, 2, 3]` as an attirbute of `df`, or whether you want it as a column. It's better to just say
```python
df['A'] = [1, 2, 3]
# or
df.loc[:, 'A'] = [1, 2, 3]
```
`Series` share many of the same methods as `DataFrame`s.
## Index
`Index`es are something of a peculiarity to pandas.
First off, they are not the kind of indexes you'll find in SQL, which are used to help the engine speed up certain queries.
In pandas, `Index`es are about lables. This helps with selection (like we did above) and automatic alignment when performing operations between two `DataFrame`s or `Series`.
R does have row labels, but they're nowhere near as powerful (or complicated) as in pandas. You can access the index of a `DataFrame` or `Series` with the `.index` attribute.
```
df.index
```
There are special kinds of `Index`es that you'll come across. Some of these are
- `MultiIndex` for multidimensional (Hierarchical) labels
- `DatetimeIndex` for datetimes
- `Float64Index` for floats
- `CategoricalIndex` for, you guessed it, `Categorical`s
We'll talk *a lot* more about indexes. They're a complex topic and can introduce headaches.
<blockquote class="twitter-tweet" lang="en"><p lang="en" dir="ltr"><a href="https://twitter.com/gjreda">@gjreda</a> <a href="https://twitter.com/treycausey">@treycausey</a> in some cases row indexes are the best thing since sliced bread, in others they simply get in the way. Hard problem</p>— Wes McKinney (@wesmckinn) <a href="https://twitter.com/wesmckinn/status/547177248768659457">December 22, 2014</a></blockquote>
Pandas, for better or for worse, does usually provide ways around row indexes being obstacles. The problem is knowing *when* they are just getting in the way, which mostly comes by experience. Sorry.
# Answers
1. Does a NumPy array have a single dtype or multiple dtypes?
- NumPy arrays are homogenous: they only have a single dtype (unlike DataFrames).
You can have an array that holds mixed types, e.g. `np.array(['a', 1])`, but the
dtype of that array is `object`, which you probably want to avoid.
2. Why is broadcasting useful?
- It lets you perform operations between arrays that are compatable, but not nescessarily identical,
in shape. This makes your code cleaner.
3. How do you slice a DataFrame by row label?
- Use `.loc[label]`. For position based use `.iloc[integer]`.
4. How do you select a column of a DataFrame?
- Standard `__getitem__`: `df[column_name]`
5. Is the Index a column in the DataFrame?
- No. It isn't included in any operations (`mean`, etc). It can be inserted as a regular
column with `df.reset_index()`.
| github_jupyter |
```
import sympy
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import cm
from sympy import sin, cos, pi, Function
from sympy import Symbol, symbols, Matrix, Transpose, init_session, Array, tensorproduct
from sympy.physics.vector import ReferenceFrame, outer, dynamicsymbols, Point
```
## Definindo as funções para h e psi
```
#Defining h
def h(t, L, w, e_mais, e_cruzado,A):
h_mais = A*cos(w*t-w*L)
h_cruzado = A*sin(w*t-w*L)
return h_mais*e_mais + h_cruzado*e_cruzado
```
\begin{equation}
h = h_+ + h_\times
\end{equation}
```
#função PSI(t)
def PSIj(j, k, L, N, A, w, T, ep, ec):
H = h(T, L[j-1], w, ep, ec,A)
phij = N[j-1].dot(H.dot(N[j-1]))/2
return phij/(1-(k.dot(N[j-1]))**2) #expandir aqui
```
\begin{equation}
\Psi (t) = \frac{n^i h_{ij} n^j}{2(1 - (\hat{k}\cdot \hat{n})^2)}
\end{equation}
## Símbolos
```
phi, theta, t, w, L, A , psi, sigma= symbols('ϕ θ t ω L A ψ σ')
```
## Sistemas de coordenadas e vetores usando o sympy
```
DetFrame = ReferenceFrame("Det")
WaveFrame = ReferenceFrame("Wave")
WaveFrame.orient(DetFrame, "body", (phi, theta, psi), 'zxz')
vx = WaveFrame.x
vy = WaveFrame.y
vz = WaveFrame.z
dbaseii = outer(vx, vx)
dbaseij = outer(vx, vy)
dbaseik = outer(vx, vz)
dbaseji = outer(vy, vx)
dbasejj = outer(vy, vy)
dbasejk = outer(vy, vz)
dbaseki = outer(vz, vx)
dbasekj = outer(vz, vy)
dbasekk = outer(vz, vz)
e_plus = dbaseii - dbasejj
e_cross = dbaseij + dbaseji
#n no referencial do detector
n2 = cos(sigma)*DetFrame.x + sin(sigma)*DetFrame.y
n3 = cos(sigma)*DetFrame.x - sin(sigma)*DetFrame.y
k = WaveFrame.z
```
## Defining posições dos satélites
```
O = Point('O') #origin
O.set_vel(DetFrame, 0)
#seting p1, p2, p3
p1 = Point(r'P_1')
p2 = Point(r'P_2')
p3 = Point(r'P_3')
#r1, r2, r3, gamma1, gamma2, gamma3 = symbols(r'r_1 r_2 r_3 \gamma_1 \gamma_2 \gamma_3') #dist from org & phase angle
l = Symbol('l')
p1.set_pos(O, l*cos(0 )*DetFrame.x + l*sin(0 )*DetFrame.y + 0*DetFrame.z)
p2.set_pos(O, l*cos(2*pi/3)*DetFrame.x + l*sin(2*pi/3)*DetFrame.y + 0*DetFrame.z)
p3.set_pos(O, l*cos(4*pi/3)*DetFrame.x + l*sin(4*pi/3)*DetFrame.y + 0*DetFrame.z)
P1 = p1.pos_from(O)
P2 = p2.pos_from(O)
P3 = p3.pos_from(O)
P = [P1, P2, P3]
#setting n's, according to KTV notation
n1 = p2.pos_from(p3)
n2 = p3.pos_from(p1)
n3 = p1.pos_from(p2)
L1 = n1.magnitude()
L2 = n2.magnitude()
L3 = n3.magnitude()
N = [n1, n2, n3]
L = [L1, L2, L3]
```
## Início do cálculo do interferômetro
```
PARAMETERS = (k,L,N,P,A,w,t, e_plus, e_cross)
def delay(func, D):
return func.subs(w*t, w*t - L[D-1])
def ygw(i,j,k,L,N,P,A,w,T, ep, ec):
m = abs(6-i-j)-1
return (1+ k.dot(N[m]))*\
(PSIj(m, k, L, N, A, w, T + k.dot(P[i-1]) - L[m], ep, ec)\
- PSIj(m,k, L, N, A, w, T + k.dot(P[j-1]), ep, ec)) # # T + k.dot(P[i]) - L[m]) , T + k.dot(P[j]))
def ygwD(i,j,k,L,N,P,A,w,T, ep, ec, D): #Ygw com delay
#delay = L[D]
return delay(ygw(i,j,k,L,N,P,A,w,T, ep, ec), D)
def yij(i,j, parms = PARAMETERS):
k,L,N,P,A,w,T, ep, ec = parms
return ygw(i,j,k,L,N,P,A,w,T, ep, ec)
def yijD(i,j,D):
return delay(yij(i,j),D)
def yijDD(i,j,D, E):
return delay(delay(yij(i,j),D),E)
f = A*cos(w*t)
f
delay(f, 2)
X = (yij(3,1) + yijD(1,3,2))\
+ delay(delay((yij(2,1) + yijD(1,2,3)),2),2)\
- (yij(2,1) + yijD(1,2,3))\
- delay(delay(yij(3,1)+yijD(1,3,2),2),2)\
- delay(delay(delay(delay(\
(yij(3,1) + yijD(1,3,2))\
+ delay(delay((yij(2,1) + yijD(1,2,3)),2),2)\
- (yij(2,1) + yijD(1,2,3))\
- delay(delay(yij(3,1)+yijD(1,3,2),2),2)\
,2),2),3),3)
#X = sympy.trigsimp(X)
y1 = yijD(3,1,2) - yij(2,3)
#calculando M
X=sympy.trigsimp(y1)
X=sympy.expand(X)
X
#M=sympy.trigsimp(M)
F_mais=X.coeff(cos(w*t))
F_cruzado=X.coeff(sin(w*t))
F_cruzado
f_mais = sympy.lambdify([ phi, theta, w, A, l], F_mais)
f_cruzado = sympy.lambdify([phi, theta, w, A, l], F_cruzado)
M_eval = sympy.lambdify([phi, theta, w, A, l], M)
f_mais
#defining parameters
phi_value, theta_value = np.mgrid[-np.pi:np.pi:100j, 0:np.pi:100j]
arm=5e9/3e8 #segundos
f=10**-3 #Hz
freq=2*np.pi*f
a=1
#atribuindos os valores acima nas funções
# [phi , theta , w , A, r1, r2, r3, gamma1, gamma2, gamma3]
f_mais_data = f_mais((phi_value), (theta_value), freq, a, arm)
f_cruzado_data = f_cruzado((phi_value),(theta_value), freq, a, arm)
f_mais_data
#plot phi, theta e F
fig = plt.figure()
ax = fig.gca(projection='3d')
ax.plot_surface(phi_value, theta_value,(f_mais_data),color='b')
#ax.plot_surface(phi_value, theta_value,(f_cruzado_data),color='g')
#ax.plot_surface(phi_value, theta_value,(f_cruzado_data-f_mais_data),color='g')
ax.set_xlabel('phi')
ax.set_ylabel('theta')
ax.set_zlabel('F+')
plt.show()
#plot x,y,z
fig = plt.figure()
ax = fig.gca(projection='3d')
x_mais=(f_mais_data)*np.sin(theta_value)*np.sin(phi_value)
y_mais=-(f_mais_data)*np.sin(theta_value)*np.cos(phi_value)
z_mais=(f_mais_data)*np.cos(theta_value)
x_cruzado=(f_cruzado_data)*np.sin(theta_value)*np.sin(phi_value)
y_cruzado=-(f_cruzado_data)*np.sin(theta_value)*np.cos(phi_value)
z_cruzado=(f_cruzado_data)*np.cos(theta_value)
ax.plot_surface(x_mais,y_mais,z_mais,color='b')
#ax.plot_surface(x_cruzado,y_cruzado,z_cruzado,color='g')
#ax.plot_surface((x_cruzado-x_mais),(y_cruzado-y_mais),(z_cruzado-z_mais),color='g', label = 'F_cruzado')
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_zlabel('z')
plt.show()
```
| github_jupyter |
## LND REST API example
### senario
* multi-hop payment (Alice -> Bob -> Charlie)
### setup
```
# load libraries
import json
from threading import Thread
from base64 import b64decode
from time import sleep
from client import LndClient, BtcClient
from util import p, dump, generate_blocks
import requests, codecs, json
from time import sleep
from configs import *
bitcoin = BtcClient(RPC_USER, RPC_PASS, BITCOIN_IP, RPC_PORT)
# initialize mainchain
bitcoin.generate(101)
p('block height = {}'.format(bitcoin.getblockcount()))
# node
alice = LndClient("alice", LND_IP, LND_REST_PORT)
bob = LndClient("bob", LND_IP_BOB, LND_REST_PORT)
charlie = LndClient("charlie", LND_IP_CHARLIE, LND_REST_PORT)
```
### 1. fund Alice, Bob
* Alice: 0.09 btc
* Charlie: 0.08 btc
```
p('[wallet balance before funding]')
p(" alice = {}".format(alice.get('/balance/blockchain')))
p(" bob = {}".format( bob.get('/balance/blockchain')))
addr_a = alice.get('/newaddress', {'type': 'np2wkh'})['address']
addr_b = bob.get('/newaddress', {'type': 'np2wkh'})['address']
bitcoin.sendtoaddress(addr_a, 0.09)
bitcoin.sendtoaddress(addr_b, 0.08)
bitcoin.generate(6)
p('[wallet balance after funding]')
p(" alice = {}".format(alice.get('/balance/blockchain')))
p(" bob = {}".format( bob.get('/balance/blockchain')))
```
### 2. connect nodes
* Alice -> Bob
* Bob -> Charlie
```
# Alice -> Bob
pubkey_b = bob.get('/getinfo')['identity_pubkey']
host_b = f'{LND_IP_BOB}:9735'
alice.post('/peers', { 'addr': { 'pubkey': pubkey_b, 'host': host_b }, 'perm': True })
# Bob -> Charlie
pubkey_c = charlie.get('/getinfo')['identity_pubkey']
host_c = f'{LND_IP_CHARLIE}:9735'
bob.post('/peers', { 'addr': { 'pubkey': pubkey_c, 'host': host_c }, 'perm': True })
p('[identity pubkey]')
p(" bob = {}".format(pubkey_b))
p(" charlie = {}".format(pubkey_c))
p('[peer]')
p(' alice <-> ', end=''); dump( alice.get('/peers'))
p(' bob <-> ', end=''); dump( bob.get('/peers'))
p(' charlie <-> ', end=''); dump(charlie.get('/peers'))
```
### 3. open the channel
```
# Alice to Bob
point_a = alice.post('/channels', {
"node_pubkey_string": pubkey_b, "local_funding_amount": "7000000", "push_sat": "0"
})
# Bob to Charlie
point_b = bob.post('/channels', {
"node_pubkey_string": pubkey_c, "local_funding_amount": "6000000", "push_sat": "0"
})
# open the channel
sleep(2)
bitcoin.generate(6)
# check the channel state
p('[channels: alice]'); dump(alice.get('/channels'))
p('[channels: bob]' ); dump( bob.get('/channels'))
p('[channel: Alice <-> Bob]')
funding_tx_id_a = b64decode(point_a['funding_txid_bytes'].encode())[::-1].hex()
output_index_a = point_a['output_index']
p(' funding tx txid = {}'.format(funding_tx_id_a))
p(' funding tx vout n = {}'.format(output_index_a))
p('[channel: Bob <-> Charlie]')
funding_tx_id_b = b64decode(point_b['funding_txid_bytes'].encode())[::-1].hex()
output_index_b = point_b['output_index']
p(' funding tx txid = {}'.format(funding_tx_id_b))
p(' funding tx vout n = {}'.format(output_index_b))
```
### 4. create invoice
* Charlie charges Alice 40,000 satoshi
```
# add a invoice to the invoice database
invoice = charlie.post('/invoices', { "value": "40000" })
# check the invoice
invoice_info = charlie.get('/invoice/' + b64decode(invoice['r_hash'].encode()).hex())
p('[invoice]'); dump(invoice_info)
# check the payment request
payreq = charlie.get('/payreq/' + invoice['payment_request'])
p('[payment request]'); dump(payreq)
```
### 5. send the payment
* Alice pays 40,000 satoshi to Charlie
* If you have the error "unable to find a path to destination", please wait a little while and then try again.
```
# check the channel balance
p('[channel balance before paying]')
p(' alice = {}'.format( alice.get('/balance/channels')))
p(' charlie = {}'.format(charlie.get('/balance/channels')))
# send the payment
payment = alice.post('/channels/transactions', { 'payment_request': invoice['payment_request'] })
p('[payment]'); dump(payment)
# check the payment
# p('[payment]'); dump(alice.get('/payments'))
# wait
sleep(2)
# check the channel balance
p('[channel balance after paying]')
p(' alice = {}'.format( alice.get('/balance/channels')))
p(' charlie = {}'.format(charlie.get('/balance/channels')))
```
### 6. close the channel
```
# check the balance
p('[channel balance]')
p(' alice = {}'.format( alice.get('/balance/channels')))
p(' bob = {}'.format( bob.get('/balance/channels')))
p(' charlie = {}'.format(charlie.get('/balance/channels')))
p('[wallet balance]')
p(' alice = ', end=''); dump( alice.get('/balance/blockchain')['confirmed_balance'])
p(' bob = ', end=''); dump( bob.get('/balance/blockchain')['confirmed_balance'])
p(' charlie = ', end=''); dump(charlie.get('/balance/blockchain')['confirmed_balance'])
p('[channel: Alice <-> Bob]')
# mine mainchain 1 block after 3 sec
Thread(target=generate_blocks, args=(bitcoin, 1, 3)).start()
# check the channel state before closing
p(' number of channels : {}'.format(len(alice.get('/channels')['channels'])))
# close the channel
res = alice.delete('/channels/' + funding_tx_id_a + '/' + str(output_index_a)).text.split("\n")[1]
closing_txid_a = json.loads(res)['result']['chan_close']['closing_txid']
p(' closing_txid = {}'.format(closing_txid_a))
sleep(5)
# check the channel state after closing
p(' number of channels : {}'.format(len(alice.get('/channels')['channels'])))
p('[channel: Bob <-> Charlie]')
# mine mainchain 1 block after 3 sec
Thread(target=generate_blocks, args=(bitcoin, 1, 3)).start()
# close the channel
res = bob.delete('/channels/' + funding_tx_id_b + '/' + str(output_index_b)).text.split("\n")[1]
closing_txid_b = json.loads(res)['result']['chan_close']['closing_txid']
p(' closing_txid = {}'.format(closing_txid_b))
sleep(5)
# check the balance
p('[channel balance]')
p(' alice = {}'.format( alice.get('/balance/channels')))
p(' bob = {}'.format( bob.get('/balance/channels')))
p(' charlie = {}'.format(charlie.get('/balance/channels')))
p('[wallet balance]')
p(' alice = ', end=''); dump( alice.get('/balance/blockchain')['confirmed_balance'])
p(' bob = ', end=''); dump( bob.get('/balance/blockchain')['confirmed_balance'])
p(' charlie = ', end=''); dump(charlie.get('/balance/blockchain')['confirmed_balance'])
```
| github_jupyter |
# Table Visualization
This section demonstrates visualization of tabular data using the [Styler][styler]
class. For information on visualization with charting please see [Chart Visualization][viz]. This document is written as a Jupyter Notebook, and can be viewed or downloaded [here][download].
[styler]: ../reference/api/pandas.io.formats.style.Styler.rst
[viz]: visualization.rst
[download]: https://nbviewer.ipython.org/github/pandas-dev/pandas/blob/master/doc/source/user_guide/style.ipynb
## Styler Object and HTML
Styling should be performed after the data in a DataFrame has been processed. The [Styler][styler] creates an HTML `<table>` and leverages CSS styling language to manipulate many parameters including colors, fonts, borders, background, etc. See [here][w3schools] for more information on styling HTML tables. This allows a lot of flexibility out of the box, and even enables web developers to integrate DataFrames into their exiting user interface designs.
The `DataFrame.style` attribute is a property that returns a [Styler][styler] object. It has a `_repr_html_` method defined on it so they are rendered automatically in Jupyter Notebook.
[styler]: ../reference/api/pandas.io.formats.style.Styler.rst
[w3schools]: https://www.w3schools.com/html/html_tables.asp
```
import matplotlib.pyplot
# We have this here to trigger matplotlib's font cache stuff.
# This cell is hidden from the output
import pandas as pd
import numpy as np
df = pd.DataFrame([[38.0, 2.0, 18.0, 22.0, 21, np.nan],[19, 439, 6, 452, 226,232]],
index=pd.Index(['Tumour (Positive)', 'Non-Tumour (Negative)'], name='Actual Label:'),
columns=pd.MultiIndex.from_product([['Decision Tree', 'Regression', 'Random'],['Tumour', 'Non-Tumour']], names=['Model:', 'Predicted:']))
df.style
```
The above output looks very similar to the standard DataFrame HTML representation. But the HTML here has already attached some CSS classes to each cell, even if we haven't yet created any styles. We can view these by calling the [.render()][render] method, which returns the raw HTML as string, which is useful for further processing or adding to a file - read on in [More about CSS and HTML](#More-About-CSS-and-HTML). Below we will show how we can use these to format the DataFrame to be more communicative. For example how we can build `s`:
[render]: ../reference/api/pandas.io.formats.style.Styler.render.rst
```
# Hidden cell to just create the below example: code is covered throughout the guide.
s = df.style\
.hide_columns([('Random', 'Tumour'), ('Random', 'Non-Tumour')])\
.format('{:.0f}')\
.set_table_styles([{
'selector': '',
'props': 'border-collapse: separate;'
},{
'selector': 'caption',
'props': 'caption-side: bottom; font-size:1.3em;'
},{
'selector': '.index_name',
'props': 'font-style: italic; color: darkgrey; font-weight:normal;'
},{
'selector': 'th:not(.index_name)',
'props': 'background-color: #000066; color: white;'
},{
'selector': 'th.col_heading',
'props': 'text-align: center;'
},{
'selector': 'th.col_heading.level0',
'props': 'font-size: 1.5em;'
},{
'selector': 'th.col2',
'props': 'border-left: 1px solid white;'
},{
'selector': '.col2',
'props': 'border-left: 1px solid #000066;'
},{
'selector': 'td',
'props': 'text-align: center; font-weight:bold;'
},{
'selector': '.true',
'props': 'background-color: #e6ffe6;'
},{
'selector': '.false',
'props': 'background-color: #ffe6e6;'
},{
'selector': '.border-red',
'props': 'border: 2px dashed red;'
},{
'selector': '.border-green',
'props': 'border: 2px dashed green;'
},{
'selector': 'td:hover',
'props': 'background-color: #ffffb3;'
}])\
.set_td_classes(pd.DataFrame([['true border-green', 'false', 'true', 'false border-red', '', ''],
['false', 'true', 'false', 'true', '', '']],
index=df.index, columns=df.columns))\
.set_caption("Confusion matrix for multiple cancer prediction models.")\
.set_tooltips(pd.DataFrame([['This model has a very strong true positive rate', '', '', "This model's total number of false negatives is too high", '', ''],
['', '', '', '', '', '']],
index=df.index, columns=df.columns),
css_class='pd-tt', props=
'visibility: hidden; position: absolute; z-index: 1; border: 1px solid #000066;'
'background-color: white; color: #000066; font-size: 0.8em;'
'transform: translate(0px, -24px); padding: 0.6em; border-radius: 0.5em;')
s
```
## Formatting the Display
### Formatting Values
Before adding styles it is useful to show that the [Styler][styler] can distinguish the *display* value from the *actual* value. To control the display value, the text is printed in each cell, and we can use the [.format()][formatfunc] method to manipulate this according to a [format spec string][format] or a callable that takes a single value and returns a string. It is possible to define this for the whole table or for individual columns.
Additionally, the format function has a **precision** argument to specifically help formatting floats, as well as **decimal** and **thousands** separators to support other locales, an **na_rep** argument to display missing data, and an **escape** argument to help displaying safe-HTML or safe-LaTeX. The default formatter is configured to adopt pandas' regular `display.precision` option, controllable using `with pd.option_context('display.precision', 2):`
Here is an example of using the multiple options to control the formatting generally and with specific column formatters.
[styler]: ../reference/api/pandas.io.formats.style.Styler.rst
[format]: https://docs.python.org/3/library/string.html#format-specification-mini-language
[formatfunc]: ../reference/api/pandas.io.formats.style.Styler.format.rst
```
df.style.format(precision=0, na_rep='MISSING', thousands=" ",
formatter={('Decision Tree', 'Tumour'): "{:.2f}",
('Regression', 'Non-Tumour'): lambda x: "$ {:,.1f}".format(x*-1e6)
})
```
### Hiding Data
The index and column headers can be completely hidden, as well subselecting rows or columns that one wishes to exclude. Both these options are performed using the same methods.
The index can be hidden from rendering by calling [.hide_index()][hideidx] without any arguments, which might be useful if your index is integer based. Similarly column headers can be hidden by calling [.hide_columns()][hidecols] without any arguments.
Specific rows or columns can be hidden from rendering by calling the same [.hide_index()][hideidx] or [.hide_columns()][hidecols] methods and passing in a row/column label, a list-like or a slice of row/column labels to for the ``subset`` argument.
Hiding does not change the integer arrangement of CSS classes, e.g. hiding the first two columns of a DataFrame means the column class indexing will start at `col2`, since `col0` and `col1` are simply ignored.
We can update our `Styler` object to hide some data and format the values.
[hideidx]: ../reference/api/pandas.io.formats.style.Styler.hide_index.rst
[hidecols]: ../reference/api/pandas.io.formats.style.Styler.hide_columns.rst
```
s = df.style.format('{:.0f}').hide_columns([('Random', 'Tumour'), ('Random', 'Non-Tumour')])
s
# Hidden cell to avoid CSS clashes and latter code upcoding previous formatting
s.set_uuid('after_hide')
```
## Methods to Add Styles
There are **3 primary methods of adding custom CSS styles** to [Styler][styler]:
- Using [.set_table_styles()][table] to control broader areas of the table with specified internal CSS. Although table styles allow the flexibility to add CSS selectors and properties controlling all individual parts of the table, they are unwieldy for individual cell specifications. Also, note that table styles cannot be exported to Excel.
- Using [.set_td_classes()][td_class] to directly link either external CSS classes to your data cells or link the internal CSS classes created by [.set_table_styles()][table]. See [here](#Setting-Classes-and-Linking-to-External-CSS). These cannot be used on column header rows or indexes, and also won't export to Excel.
- Using the [.apply()][apply] and [.applymap()][applymap] functions to add direct internal CSS to specific data cells. See [here](#Styler-Functions). These cannot be used on column header rows or indexes, but only these methods add styles that will export to Excel. These methods work in a similar way to [DataFrame.apply()][dfapply] and [DataFrame.applymap()][dfapplymap].
[table]: ../reference/api/pandas.io.formats.style.Styler.set_table_styles.rst
[styler]: ../reference/api/pandas.io.formats.style.Styler.rst
[td_class]: ../reference/api/pandas.io.formats.style.Styler.set_td_classes.rst
[apply]: ../reference/api/pandas.io.formats.style.Styler.apply.rst
[applymap]: ../reference/api/pandas.io.formats.style.Styler.applymap.rst
[dfapply]: ../reference/api/pandas.DataFrame.apply.rst
[dfapplymap]: ../reference/api/pandas.DataFrame.applymap.rst
## Table Styles
Table styles are flexible enough to control all individual parts of the table, including column headers and indexes.
However, they can be unwieldy to type for individual data cells or for any kind of conditional formatting, so we recommend that table styles are used for broad styling, such as entire rows or columns at a time.
Table styles are also used to control features which can apply to the whole table at once such as creating a generic hover functionality. The `:hover` pseudo-selector, as well as other pseudo-selectors, can only be used this way.
To replicate the normal format of CSS selectors and properties (attribute value pairs), e.g.
```
tr:hover {
background-color: #ffff99;
}
```
the necessary format to pass styles to [.set_table_styles()][table] is as a list of dicts, each with a CSS-selector tag and CSS-properties. Properties can either be a list of 2-tuples, or a regular CSS-string, for example:
[table]: ../reference/api/pandas.io.formats.style.Styler.set_table_styles.rst
```
cell_hover = { # for row hover use <tr> instead of <td>
'selector': 'td:hover',
'props': [('background-color', '#ffffb3')]
}
index_names = {
'selector': '.index_name',
'props': 'font-style: italic; color: darkgrey; font-weight:normal;'
}
headers = {
'selector': 'th:not(.index_name)',
'props': 'background-color: #000066; color: white;'
}
s.set_table_styles([cell_hover, index_names, headers])
# Hidden cell to avoid CSS clashes and latter code upcoding previous formatting
s.set_uuid('after_tab_styles1')
```
Next we just add a couple more styling artifacts targeting specific parts of the table. Be careful here, since we are *chaining methods* we need to explicitly instruct the method **not to** ``overwrite`` the existing styles.
```
s.set_table_styles([
{'selector': 'th.col_heading', 'props': 'text-align: center;'},
{'selector': 'th.col_heading.level0', 'props': 'font-size: 1.5em;'},
{'selector': 'td', 'props': 'text-align: center; font-weight: bold;'},
], overwrite=False)
# Hidden cell to avoid CSS clashes and latter code upcoding previous formatting
s.set_uuid('after_tab_styles2')
```
As a convenience method (*since version 1.2.0*) we can also pass a **dict** to [.set_table_styles()][table] which contains row or column keys. Behind the scenes Styler just indexes the keys and adds relevant `.col<m>` or `.row<n>` classes as necessary to the given CSS selectors.
[table]: ../reference/api/pandas.io.formats.style.Styler.set_table_styles.rst
```
s.set_table_styles({
('Regression', 'Tumour'): [{'selector': 'th', 'props': 'border-left: 1px solid white'},
{'selector': 'td', 'props': 'border-left: 1px solid #000066'}]
}, overwrite=False, axis=0)
# Hidden cell to avoid CSS clashes and latter code upcoding previous formatting
s.set_uuid('xyz01')
```
## Setting Classes and Linking to External CSS
If you have designed a website then it is likely you will already have an external CSS file that controls the styling of table and cell objects within it. You may want to use these native files rather than duplicate all the CSS in python (and duplicate any maintenance work).
### Table Attributes
It is very easy to add a `class` to the main `<table>` using [.set_table_attributes()][tableatt]. This method can also attach inline styles - read more in [CSS Hierarchies](#CSS-Hierarchies).
[tableatt]: ../reference/api/pandas.io.formats.style.Styler.set_table_attributes.rst
```
out = s.set_table_attributes('class="my-table-cls"').render()
print(out[out.find('<table'):][:109])
```
### Data Cell CSS Classes
*New in version 1.2.0*
The [.set_td_classes()][tdclass] method accepts a DataFrame with matching indices and columns to the underlying [Styler][styler]'s DataFrame. That DataFrame will contain strings as css-classes to add to individual data cells: the `<td>` elements of the `<table>`. Rather than use external CSS we will create our classes internally and add them to table style. We will save adding the borders until the [section on tooltips](#Tooltips).
[tdclass]: ../reference/api/pandas.io.formats.style.Styler.set_td_classes.rst
[styler]: ../reference/api/pandas.io.formats.style.Styler.rst
```
s.set_table_styles([ # create internal CSS classes
{'selector': '.true', 'props': 'background-color: #e6ffe6;'},
{'selector': '.false', 'props': 'background-color: #ffe6e6;'},
], overwrite=False)
cell_color = pd.DataFrame([['true ', 'false ', 'true ', 'false '],
['false ', 'true ', 'false ', 'true ']],
index=df.index,
columns=df.columns[:4])
s.set_td_classes(cell_color)
# Hidden cell to avoid CSS clashes and latter code upcoding previous formatting
s.set_uuid('after_classes')
```
## Styler Functions
We use the following methods to pass your style functions. Both of those methods take a function (and some other keyword arguments) and apply it to the DataFrame in a certain way, rendering CSS styles.
- [.applymap()][applymap] (elementwise): accepts a function that takes a single value and returns a string with the CSS attribute-value pair.
- [.apply()][apply] (column-/row-/table-wise): accepts a function that takes a Series or DataFrame and returns a Series, DataFrame, or numpy array with an identical shape where each element is a string with a CSS attribute-value pair. This method passes each column or row of your DataFrame one-at-a-time or the entire table at once, depending on the `axis` keyword argument. For columnwise use `axis=0`, rowwise use `axis=1`, and for the entire table at once use `axis=None`.
This method is powerful for applying multiple, complex logic to data cells. We create a new DataFrame to demonstrate this.
[apply]: ../reference/api/pandas.io.formats.style.Styler.apply.rst
[applymap]: ../reference/api/pandas.io.formats.style.Styler.applymap.rst
```
np.random.seed(0)
df2 = pd.DataFrame(np.random.randn(10,4), columns=['A','B','C','D'])
df2.style
```
For example we can build a function that colors text if it is negative, and chain this with a function that partially fades cells of negligible value. Since this looks at each element in turn we use ``applymap``.
```
def style_negative(v, props=''):
return props if v < 0 else None
s2 = df2.style.applymap(style_negative, props='color:red;')\
.applymap(lambda v: 'opacity: 20%;' if (v < 0.3) and (v > -0.3) else None)
s2
# Hidden cell to avoid CSS clashes and latter code upcoding previous formatting
s2.set_uuid('after_applymap')
```
We can also build a function that highlights the maximum value across rows, cols, and the DataFrame all at once. In this case we use ``apply``. Below we highlight the maximum in a column.
```
def highlight_max(s, props=''):
return np.where(s == np.nanmax(s.values), props, '')
s2.apply(highlight_max, props='color:white;background-color:darkblue', axis=0)
# Hidden cell to avoid CSS clashes and latter code upcoding previous formatting
s2.set_uuid('after_apply')
```
We can use the same function across the different axes, highlighting here the DataFrame maximum in purple, and row maximums in pink.
```
s2.apply(highlight_max, props='color:white;background-color:pink;', axis=1)\
.apply(highlight_max, props='color:white;background-color:purple', axis=None)
```
This last example shows how some styles have been overwritten by others. In general the most recent style applied is active but you can read more in the [section on CSS hierarchies](#CSS-Hierarchies). You can also apply these styles to more granular parts of the DataFrame - read more in section on [subset slicing](#Finer-Control-with-Slicing).
It is possible to replicate some of this functionality using just classes but it can be more cumbersome. See [item 3) of Optimization](#Optimization)
<div class="alert alert-info">
*Debugging Tip*: If you're having trouble writing your style function, try just passing it into ``DataFrame.apply``. Internally, ``Styler.apply`` uses ``DataFrame.apply`` so the result should be the same, and with ``DataFrame.apply`` you will be able to inspect the CSS string output of your intended function in each cell.
</div>
## Tooltips and Captions
Table captions can be added with the [.set_caption()][caption] method. You can use table styles to control the CSS relevant to the caption.
[caption]: ../reference/api/pandas.io.formats.style.Styler.set_caption.rst
```
s.set_caption("Confusion matrix for multiple cancer prediction models.")\
.set_table_styles([{
'selector': 'caption',
'props': 'caption-side: bottom; font-size:1.25em;'
}], overwrite=False)
# Hidden cell to avoid CSS clashes and latter code upcoding previous formatting
s.set_uuid('after_caption')
```
Adding tooltips (*since version 1.3.0*) can be done using the [.set_tooltips()][tooltips] method in the same way you can add CSS classes to data cells by providing a string based DataFrame with intersecting indices and columns. You don't have to specify a `css_class` name or any css `props` for the tooltips, since there are standard defaults, but the option is there if you want more visual control.
[tooltips]: ../reference/api/pandas.io.formats.style.Styler.set_tooltips.rst
```
tt = pd.DataFrame([['This model has a very strong true positive rate',
"This model's total number of false negatives is too high"]],
index=['Tumour (Positive)'], columns=df.columns[[0,3]])
s.set_tooltips(tt, props='visibility: hidden; position: absolute; z-index: 1; border: 1px solid #000066;'
'background-color: white; color: #000066; font-size: 0.8em;'
'transform: translate(0px, -24px); padding: 0.6em; border-radius: 0.5em;')
# Hidden cell to avoid CSS clashes and latter code upcoding previous formatting
s.set_uuid('after_tooltips')
```
The only thing left to do for our table is to add the highlighting borders to draw the audience attention to the tooltips. We will create internal CSS classes as before using table styles. **Setting classes always overwrites** so we need to make sure we add the previous classes.
```
s.set_table_styles([ # create internal CSS classes
{'selector': '.border-red', 'props': 'border: 2px dashed red;'},
{'selector': '.border-green', 'props': 'border: 2px dashed green;'},
], overwrite=False)
cell_border = pd.DataFrame([['border-green ', ' ', ' ', 'border-red '],
[' ', ' ', ' ', ' ']],
index=df.index,
columns=df.columns[:4])
s.set_td_classes(cell_color + cell_border)
# Hidden cell to avoid CSS clashes and latter code upcoding previous formatting
s.set_uuid('after_borders')
```
## Finer Control with Slicing
The examples we have shown so far for the `Styler.apply` and `Styler.applymap` functions have not demonstrated the use of the ``subset`` argument. This is a useful argument which permits a lot of flexibility: it allows you to apply styles to specific rows or columns, without having to code that logic into your `style` function.
The value passed to `subset` behaves similar to slicing a DataFrame;
- A scalar is treated as a column label
- A list (or Series or NumPy array) is treated as multiple column labels
- A tuple is treated as `(row_indexer, column_indexer)`
Consider using `pd.IndexSlice` to construct the tuple for the last one. We will create a MultiIndexed DataFrame to demonstrate the functionality.
```
df3 = pd.DataFrame(np.random.randn(4,4),
pd.MultiIndex.from_product([['A', 'B'], ['r1', 'r2']]),
columns=['c1','c2','c3','c4'])
df3
```
We will use subset to highlight the maximum in the third and fourth columns with red text. We will highlight the subset sliced region in yellow.
```
slice_ = ['c3', 'c4']
df3.style.apply(highlight_max, props='color:red;', axis=0, subset=slice_)\
.set_properties(**{'background-color': '#ffffb3'}, subset=slice_)
```
If combined with the ``IndexSlice`` as suggested then it can index across both dimensions with greater flexibility.
```
idx = pd.IndexSlice
slice_ = idx[idx[:,'r1'], idx['c2':'c4']]
df3.style.apply(highlight_max, props='color:red;', axis=0, subset=slice_)\
.set_properties(**{'background-color': '#ffffb3'}, subset=slice_)
```
This also provides the flexibility to sub select rows when used with the `axis=1`.
```
slice_ = idx[idx[:,'r2'], :]
df3.style.apply(highlight_max, props='color:red;', axis=1, subset=slice_)\
.set_properties(**{'background-color': '#ffffb3'}, subset=slice_)
```
There is also scope to provide **conditional filtering**.
Suppose we want to highlight the maximum across columns 2 and 4 only in the case that the sum of columns 1 and 3 is less than -2.0 *(essentially excluding rows* `(:,'r2')`*)*.
```
slice_ = idx[idx[(df3['c1'] + df3['c3']) < -2.0], ['c2', 'c4']]
df3.style.apply(highlight_max, props='color:red;', axis=1, subset=slice_)\
.set_properties(**{'background-color': '#ffffb3'}, subset=slice_)
```
Only label-based slicing is supported right now, not positional, and not callables.
If your style function uses a `subset` or `axis` keyword argument, consider wrapping your function in a `functools.partial`, partialing out that keyword.
```python
my_func2 = functools.partial(my_func, subset=42)
```
## Optimization
Generally, for smaller tables and most cases, the rendered HTML does not need to be optimized, and we don't really recommend it. There are two cases where it is worth considering:
- If you are rendering and styling a very large HTML table, certain browsers have performance issues.
- If you are using ``Styler`` to dynamically create part of online user interfaces and want to improve network performance.
Here we recommend the following steps to implement:
### 1. Remove UUID and cell_ids
Ignore the `uuid` and set `cell_ids` to `False`. This will prevent unnecessary HTML.
<div class="alert alert-warning">
<font color=red>This is sub-optimal:</font>
</div>
```
df4 = pd.DataFrame([[1,2],[3,4]])
s4 = df4.style
```
<div class="alert alert-info">
<font color=green>This is better:</font>
</div>
```
from pandas.io.formats.style import Styler
s4 = Styler(df4, uuid_len=0, cell_ids=False)
```
### 2. Use table styles
Use table styles where possible (e.g. for all cells or rows or columns at a time) since the CSS is nearly always more efficient than other formats.
<div class="alert alert-warning">
<font color=red>This is sub-optimal:</font>
</div>
```
props = 'font-family: "Times New Roman", Times, serif; color: #e83e8c; font-size:1.3em;'
df4.style.applymap(lambda x: props, subset=[1])
```
<div class="alert alert-info">
<font color=green>This is better:</font>
</div>
```
df4.style.set_table_styles([{'selector': 'td.col1', 'props': props}])
```
### 3. Set classes instead of using Styler functions
For large DataFrames where the same style is applied to many cells it can be more efficient to declare the styles as classes and then apply those classes to data cells, rather than directly applying styles to cells. It is, however, probably still easier to use the Styler function api when you are not concerned about optimization.
<div class="alert alert-warning">
<font color=red>This is sub-optimal:</font>
</div>
```
df2.style.apply(highlight_max, props='color:white;background-color:darkblue;', axis=0)\
.apply(highlight_max, props='color:white;background-color:pink;', axis=1)\
.apply(highlight_max, props='color:white;background-color:purple', axis=None)
```
<div class="alert alert-info">
<font color=green>This is better:</font>
</div>
```
build = lambda x: pd.DataFrame(x, index=df2.index, columns=df2.columns)
cls1 = build(df2.apply(highlight_max, props='cls-1 ', axis=0))
cls2 = build(df2.apply(highlight_max, props='cls-2 ', axis=1, result_type='expand').values)
cls3 = build(highlight_max(df2, props='cls-3 '))
df2.style.set_table_styles([
{'selector': '.cls-1', 'props': 'color:white;background-color:darkblue;'},
{'selector': '.cls-2', 'props': 'color:white;background-color:pink;'},
{'selector': '.cls-3', 'props': 'color:white;background-color:purple;'}
]).set_td_classes(cls1 + cls2 + cls3)
```
### 4. Don't use tooltips
Tooltips require `cell_ids` to work and they generate extra HTML elements for *every* data cell.
### 5. If every byte counts use string replacement
You can remove unnecessary HTML, or shorten the default class names with string replace functions.
```
html = Styler(df4, uuid_len=0, cell_ids=False)\
.set_table_styles([{'selector': 'td', 'props': props},
{'selector': '.col1', 'props': 'color:green;'},
{'selector': '.level0', 'props': 'color:blue;'}])\
.render()\
.replace('blank', '')\
.replace('data', '')\
.replace('level0', 'l0')\
.replace('col_heading', '')\
.replace('row_heading', '')
import re
html = re.sub(r'col[0-9]+', lambda x: x.group().replace('col', 'c'), html)
html = re.sub(r'row[0-9]+', lambda x: x.group().replace('row', 'r'), html)
print(html)
from IPython.display import HTML
HTML(html)
```
## Builtin Styles
Some styling functions are common enough that we've "built them in" to the `Styler`, so you don't have to write them and apply them yourself. The current list of such functions is:
- [.highlight_null][nullfunc]: for use with identifying missing data.
- [.highlight_min][minfunc] and [.highlight_max][maxfunc]: for use with identifying extremeties in data.
- [.highlight_between][betweenfunc] and [.highlight_quantile][quantilefunc]: for use with identifying classes within data.
- [.background_gradient][bgfunc]: a flexible method for highlighting cells based or their, or other, values on a numeric scale.
- [.text_gradient][textfunc]: similar method for highlighting text based on their, or other, values on a numeric scale.
- [.bar][barfunc]: to display mini-charts within cell backgrounds.
The individual documentation on each function often gives more examples of their arguments.
[nullfunc]: ../reference/api/pandas.io.formats.style.Styler.highlight_null.rst
[minfunc]: ../reference/api/pandas.io.formats.style.Styler.highlight_min.rst
[maxfunc]: ../reference/api/pandas.io.formats.style.Styler.highlight_max.rst
[betweenfunc]: ../reference/api/pandas.io.formats.style.Styler.highlight_between.rst
[quantilefunc]: ../reference/api/pandas.io.formats.style.Styler.highlight_quantile.rst
[bgfunc]: ../reference/api/pandas.io.formats.style.Styler.background_gradient.rst
[textfunc]: ../reference/api/pandas.io.formats.style.Styler.text_gradient.rst
[barfunc]: ../reference/api/pandas.io.formats.style.Styler.bar.rst
### Highlight Null
```
df2.iloc[0,2] = np.nan
df2.iloc[4,3] = np.nan
df2.loc[:4].style.highlight_null(null_color='yellow')
```
### Highlight Min or Max
```
df2.loc[:4].style.highlight_max(axis=1, props='color:white; font-weight:bold; background-color:darkblue;')
```
### Highlight Between
This method accepts ranges as float, or NumPy arrays or Series provided the indexes match.
```
left = pd.Series([1.0, 0.0, 1.0], index=["A", "B", "D"])
df2.loc[:4].style.highlight_between(left=left, right=1.5, axis=1, props='color:white; background-color:purple;')
```
### Highlight Quantile
Useful for detecting the highest or lowest percentile values
```
df2.loc[:4].style.highlight_quantile(q_left=0.85, axis=None, color='yellow')
```
### Background Gradient and Text Gradient
You can create "heatmaps" with the `background_gradient` and `text_gradient` methods. These require matplotlib, and we'll use [Seaborn](https://stanford.edu/~mwaskom/software/seaborn/) to get a nice colormap.
```
import seaborn as sns
cm = sns.light_palette("green", as_cmap=True)
df2.style.background_gradient(cmap=cm)
df2.style.text_gradient(cmap=cm)
```
[.background_gradient][bgfunc] and [.text_gradient][textfunc] have a number of keyword arguments to customise the gradients and colors. See the documentation.
[bgfunc]: ../reference/api/pandas.io.formats.style.Styler.background_gradient.rst
[textfunc]: ../reference/api/pandas.io.formats.style.Styler.text_gradient.rst
### Set properties
Use `Styler.set_properties` when the style doesn't actually depend on the values. This is just a simple wrapper for `.applymap` where the function returns the same properties for all cells.
```
df2.loc[:4].style.set_properties(**{'background-color': 'black',
'color': 'lawngreen',
'border-color': 'white'})
```
### Bar charts
You can include "bar charts" in your DataFrame.
```
df2.style.bar(subset=['A', 'B'], color='#d65f5f')
```
Additional keyword arguments give more control on centering and positioning, and you can pass a list of `[color_negative, color_positive]` to highlight lower and higher values.
Here's how you can change the above with the new `align` option, combined with setting `vmin` and `vmax` limits, the `width` of the figure, and underlying css `props` of cells, leaving space to display the text and the bars:
```
df2.style.bar(align=0, vmin=-2.5, vmax=2.5, color=['#d65f5f', '#5fba7d'], height=50,
width=60, props="width: 120px; border-right: 1px solid black;").format('{:.3f}', na_rep="")
```
The following example aims to give a highlight of the behavior of the new align options:
```
# Hide the construction of the display chart from the user
import pandas as pd
from IPython.display import HTML
# Test series
test1 = pd.Series([-100,-60,-30,-20], name='All Negative')
test2 = pd.Series([-10,-5,0,90], name='Both Pos and Neg')
test3 = pd.Series([10,20,50,100], name='All Positive')
test4 = pd.Series([100, 103, 101, 102], name='Large Positive')
head = """
<table>
<thead>
<th>Align</th>
<th>All Negative</th>
<th>Both Neg and Pos</th>
<th>All Positive</th>
<th>Large Positive</th>
</thead>
</tbody>
"""
aligns = ['left', 'right', 'zero', 'mid', 'mean', 99]
for align in aligns:
row = "<tr><th>{}</th>".format(align)
for series in [test1,test2,test3, test4]:
s = series.copy()
s.name=''
row += "<td>{}</td>".format(s.to_frame().style.hide_index().bar(align=align,
color=['#d65f5f', '#5fba7d'],
width=100).render()) #testn['width']
row += '</tr>'
head += row
head+= """
</tbody>
</table>"""
HTML(head)
```
## Sharing styles
Say you have a lovely style built up for a DataFrame, and now you want to apply the same style to a second DataFrame. Export the style with `df1.style.export`, and import it on the second DataFrame with `df1.style.set`
```
style1 = df2.style.applymap(style_negative, props='color:red;')\
.applymap(lambda v: 'opacity: 20%;' if (v < 0.3) and (v > -0.3) else None)
style2 = df3.style
style2.use(style1.export())
style2
```
Notice that you're able to share the styles even though they're data aware. The styles are re-evaluated on the new DataFrame they've been `use`d upon.
## Limitations
- DataFrame only `(use Series.to_frame().style)`
- The index and columns must be unique
- No large repr, and construction performance isn't great; although we have some [HTML optimizations](#Optimization)
- You can only style the *values*, not the index or columns (except with `table_styles` above)
- You can only apply styles, you can't insert new HTML entities
Some of these might be addressed in the future.
## Other Fun and Useful Stuff
Here are a few interesting examples.
### Widgets
`Styler` interacts pretty well with widgets. If you're viewing this online instead of running the notebook yourself, you're missing out on interactively adjusting the color palette.
```
from ipywidgets import widgets
@widgets.interact
def f(h_neg=(0, 359, 1), h_pos=(0, 359), s=(0., 99.9), l=(0., 99.9)):
return df2.style.background_gradient(
cmap=sns.palettes.diverging_palette(h_neg=h_neg, h_pos=h_pos, s=s, l=l,
as_cmap=True)
)
```
### Magnify
```
def magnify():
return [dict(selector="th",
props=[("font-size", "4pt")]),
dict(selector="td",
props=[('padding', "0em 0em")]),
dict(selector="th:hover",
props=[("font-size", "12pt")]),
dict(selector="tr:hover td:hover",
props=[('max-width', '200px'),
('font-size', '12pt')])
]
np.random.seed(25)
cmap = cmap=sns.diverging_palette(5, 250, as_cmap=True)
bigdf = pd.DataFrame(np.random.randn(20, 25)).cumsum()
bigdf.style.background_gradient(cmap, axis=1)\
.set_properties(**{'max-width': '80px', 'font-size': '1pt'})\
.set_caption("Hover to magnify")\
.format(precision=2)\
.set_table_styles(magnify())
```
### Sticky Headers
If you display a large matrix or DataFrame in a notebook, but you want to always see the column and row headers you can use the [.set_sticky][sticky] method which manipulates the table styles CSS.
[sticky]: ../reference/api/pandas.io.formats.style.Styler.set_sticky.rst
```
bigdf = pd.DataFrame(np.random.randn(16, 100))
bigdf.style.set_sticky(axis="index")
```
It is also possible to stick MultiIndexes and even only specific levels.
```
bigdf.index = pd.MultiIndex.from_product([["A","B"],[0,1],[0,1,2,3]])
bigdf.style.set_sticky(axis="index", pixel_size=18, levels=[1,2])
```
### HTML Escaping
Suppose you have to display HTML within HTML, that can be a bit of pain when the renderer can't distinguish. You can use the `escape` formatting option to handle this, and even use it within a formatter that contains HTML itself.
```
df4 = pd.DataFrame([['<div></div>', '"&other"', '<span></span>']])
df4.style
df4.style.format(escape="html")
df4.style.format('<a href="https://pandas.pydata.org" target="_blank">{}</a>', escape="html")
```
## Export to Excel
Some support (*since version 0.20.0*) is available for exporting styled `DataFrames` to Excel worksheets using the `OpenPyXL` or `XlsxWriter` engines. CSS2.2 properties handled include:
- `background-color`
- `color`
- `font-family`
- `font-style`
- `font-weight`
- `text-align`
- `text-decoration`
- `vertical-align`
- `white-space: nowrap`
- Currently broken: `border-style`, `border-width`, `border-color` and their {`top`, `right`, `bottom`, `left` variants}
- Only CSS2 named colors and hex colors of the form `#rgb` or `#rrggbb` are currently supported.
- The following pseudo CSS properties are also available to set excel specific style properties:
- `number-format`
Table level styles, and data cell CSS-classes are not included in the export to Excel: individual cells must have their properties mapped by the `Styler.apply` and/or `Styler.applymap` methods.
```
df2.style.\
applymap(style_negative, props='color:red;').\
highlight_max(axis=0).\
to_excel('styled.xlsx', engine='openpyxl')
```
A screenshot of the output:

## Export to LaTeX
There is support (*since version 1.3.0*) to export `Styler` to LaTeX. The documentation for the [.to_latex][latex] method gives further detail and numerous examples.
[latex]: ../reference/api/pandas.io.formats.style.Styler.to_latex.rst
## More About CSS and HTML
Cascading Style Sheet (CSS) language, which is designed to influence how a browser renders HTML elements, has its own peculiarities. It never reports errors: it just silently ignores them and doesn't render your objects how you intend so can sometimes be frustrating. Here is a very brief primer on how ``Styler`` creates HTML and interacts with CSS, with advice on common pitfalls to avoid.
### CSS Classes and Ids
The precise structure of the CSS `class` attached to each cell is as follows.
- Cells with Index and Column names include `index_name` and `level<k>` where `k` is its level in a MultiIndex
- Index label cells include
+ `row_heading`
+ `level<k>` where `k` is the level in a MultiIndex
+ `row<m>` where `m` is the numeric position of the row
- Column label cells include
+ `col_heading`
+ `level<k>` where `k` is the level in a MultiIndex
+ `col<n>` where `n` is the numeric position of the column
- Data cells include
+ `data`
+ `row<m>`, where `m` is the numeric position of the cell.
+ `col<n>`, where `n` is the numeric position of the cell.
- Blank cells include `blank`
The structure of the `id` is `T_uuid_level<k>_row<m>_col<n>` where `level<k>` is used only on headings, and headings will only have either `row<m>` or `col<n>` whichever is needed. By default we've also prepended each row/column identifier with a UUID unique to each DataFrame so that the style from one doesn't collide with the styling from another within the same notebook or page. You can read more about the use of UUIDs in [Optimization](#Optimization).
We can see example of the HTML by calling the [.render()][render] method.
[render]: ../reference/api/pandas.io.formats.style.Styler.render.rst
```
print(pd.DataFrame([[1,2],[3,4]], index=['i1', 'i2'], columns=['c1', 'c2']).style.render())
```
### CSS Hierarchies
The examples have shown that when CSS styles overlap, the one that comes last in the HTML render, takes precedence. So the following yield different results:
```
df4 = pd.DataFrame([['text']])
df4.style.applymap(lambda x: 'color:green;')\
.applymap(lambda x: 'color:red;')
df4.style.applymap(lambda x: 'color:red;')\
.applymap(lambda x: 'color:green;')
```
This is only true for CSS rules that are equivalent in hierarchy, or importance. You can read more about [CSS specificity here](https://www.w3schools.com/css/css_specificity.asp) but for our purposes it suffices to summarize the key points:
A CSS importance score for each HTML element is derived by starting at zero and adding:
- 1000 for an inline style attribute
- 100 for each ID
- 10 for each attribute, class or pseudo-class
- 1 for each element name or pseudo-element
Let's use this to describe the action of the following configurations
```
df4.style.set_uuid('a_')\
.set_table_styles([{'selector': 'td', 'props': 'color:red;'}])\
.applymap(lambda x: 'color:green;')
```
This text is red because the generated selector `#T_a_ td` is worth 101 (ID plus element), whereas `#T_a_row0_col0` is only worth 100 (ID), so is considered inferior even though in the HTML it comes after the previous.
```
df4.style.set_uuid('b_')\
.set_table_styles([{'selector': 'td', 'props': 'color:red;'},
{'selector': '.cls-1', 'props': 'color:blue;'}])\
.applymap(lambda x: 'color:green;')\
.set_td_classes(pd.DataFrame([['cls-1']]))
```
In the above case the text is blue because the selector `#T_b_ .cls-1` is worth 110 (ID plus class), which takes precendence.
```
df4.style.set_uuid('c_')\
.set_table_styles([{'selector': 'td', 'props': 'color:red;'},
{'selector': '.cls-1', 'props': 'color:blue;'},
{'selector': 'td.data', 'props': 'color:yellow;'}])\
.applymap(lambda x: 'color:green;')\
.set_td_classes(pd.DataFrame([['cls-1']]))
```
Now we have created another table style this time the selector `T_c_ td.data` (ID plus element plus class) gets bumped up to 111.
If your style fails to be applied, and its really frustrating, try the `!important` trump card.
```
df4.style.set_uuid('d_')\
.set_table_styles([{'selector': 'td', 'props': 'color:red;'},
{'selector': '.cls-1', 'props': 'color:blue;'},
{'selector': 'td.data', 'props': 'color:yellow;'}])\
.applymap(lambda x: 'color:green !important;')\
.set_td_classes(pd.DataFrame([['cls-1']]))
```
Finally got that green text after all!
## Extensibility
The core of pandas is, and will remain, its "high-performance, easy-to-use data structures".
With that in mind, we hope that `DataFrame.style` accomplishes two goals
- Provide an API that is pleasing to use interactively and is "good enough" for many tasks
- Provide the foundations for dedicated libraries to build on
If you build a great library on top of this, let us know and we'll [link](https://pandas.pydata.org/pandas-docs/stable/ecosystem.html) to it.
### Subclassing
If the default template doesn't quite suit your needs, you can subclass Styler and extend or override the template.
We'll show an example of extending the default template to insert a custom header before each table.
```
from jinja2 import Environment, ChoiceLoader, FileSystemLoader
from IPython.display import HTML
from pandas.io.formats.style import Styler
```
We'll use the following template:
```
with open("templates/myhtml.tpl") as f:
print(f.read())
```
Now that we've created a template, we need to set up a subclass of ``Styler`` that
knows about it.
```
class MyStyler(Styler):
env = Environment(
loader=ChoiceLoader([
FileSystemLoader("templates"), # contains ours
Styler.loader, # the default
])
)
template_html_table = env.get_template("myhtml.tpl")
```
Notice that we include the original loader in our environment's loader.
That's because we extend the original template, so the Jinja environment needs
to be able to find it.
Now we can use that custom styler. It's `__init__` takes a DataFrame.
```
MyStyler(df3)
```
Our custom template accepts a `table_title` keyword. We can provide the value in the `.render` method.
```
HTML(MyStyler(df3).render(table_title="Extending Example"))
```
For convenience, we provide the `Styler.from_custom_template` method that does the same as the custom subclass.
```
EasyStyler = Styler.from_custom_template("templates", "myhtml.tpl")
HTML(EasyStyler(df3).render(table_title="Another Title"))
```
#### Template Structure
Here's the template structure for the both the style generation template and the table generation template:
Style template:
```
with open("templates/html_style_structure.html") as f:
style_structure = f.read()
HTML(style_structure)
```
Table template:
```
with open("templates/html_table_structure.html") as f:
table_structure = f.read()
HTML(table_structure)
```
See the template in the [GitHub repo](https://github.com/pandas-dev/pandas) for more details.
```
# # Hack to get the same style in the notebook as the
# # main site. This is hidden in the docs.
# from IPython.display import HTML
# with open("themes/nature_with_gtoc/static/nature.css_t") as f:
# css = f.read()
# HTML('<style>{}</style>'.format(css))
```
| github_jupyter |
```
import numpy as np
import scipy.sparse as sparse
import scipy.fftpack as fft
import matplotlib.pyplot as plt
%matplotlib notebook
shape = (5, 5)
nx, ny = shape
charges = np.zeros(shape)
charges[:] = 1.0 / (nx * ny)
charges[nx // 2, ny // 2] = 1.0 / (nx * ny) - 1.0
print(charges[:3, :3])
charges = charges.flatten()
#Build Laplacian
ex = np.append(np.ones(nx - 2), [2, 2])
ey = np.append(np.ones(ny - 2), [2, 2])
Dxx = sparse.spdiags([ex, -2 * np.ones(nx), ex[::-1]], [-1, 0, 1], nx, nx)
Dyy = sparse.spdiags([ey, -2 * np.ones(ny), ey[::-1]], [-1, 0, 1], ny, ny)
L = sparse.kronsum(Dxx, Dyy).todense()
print(Dxx.shape, Dyy.shape, L.shape)
fig, axes = plt.subplots(1, 3)
axes[0].imshow(Dxx.todense())
axes[1].imshow(Dyy.todense())
axes[2].imshow(L)
###############
#Fourier method
rhofft = np.zeros(shape, dtype = float)
for i in range(shape[0]):
rhofft[i,:] = fft.dct(charges.reshape(shape)[i,:], type = 1) / (shape[1] - 1.0)
for j in range(shape[1]):
rhofft[:,j] = fft.dct(rhofft[:,j], type = 1) / (shape[0] - 1.0)
for i in range(shape[0]):
for j in range(shape[1]):
factor = 2.0 * (np.cos((np.pi * i) / (shape[0] - 1)) + np.cos((np.pi * j) / (shape[1] - 1)) - 2.0)
if factor != 0.0:
rhofft[i, j] /= factor
else:
rhofft[i, j] = 0.0
potential = np.zeros(shape, dtype = float)
for i in range(shape[0]):
potential[i,:] = 0.5 * fft.dct(rhofft[i,:], type = 1)
for j in range(shape[1]):
potential[:,j] = 0.5 * fft.dct(potential[:,j], type = 1)
################
charges = charges.reshape(shape)
charges_hat = (L @ potential.flatten()).reshape(shape)
print(np.sum(np.abs(charges_hat - charges)))
# print(potential)
fig, axes = plt.subplots(2, 2)
axr = axes.ravel()
axim = axr[0].imshow(charges_hat)
fig.colorbar(axim, ax=axr[0])
axim = axr[1].imshow(charges)
fig.colorbar(axim, ax=axr[1])
axim = axr[2].imshow(charges - charges_hat)
fig.colorbar(axim, ax=axr[2])
axim = axr[3].imshow(potential)
fig.colorbar(axim, ax=axr[3])
charges[:4, :4]
```
# Diagonalizing operators:
```
import scipy.linalg as LA
# Example from Strang, 1999
A0 = LA.circulant([2,-1,0,-1])
print(A0)
# LA.LU
Lam, V = LA.eig(A0)
print(Lam)
print(V)
print(V[:, 0])
LA.norm(V, axis=1)
LA.norm(V[:, 0])
1/np.sqrt(2)
```
# Periodic A: Diagonlized by the DFT
The columns of the N = 4 DFT matrix are eigenvectors of the $A_0$ matrix.
Using $\omega = e^{i 2 \pi / n}$,
```
N = A0.shape[0]
omega = np.exp(1j*2*np.pi / N)
print(omega)
```
$v_k = \omega^{jk}, j \in {0,1,\ldots,N-1}$
```
Vs = []
for j in range(N):
v = omega ** (j * np.arange(0, N))
Vs.append(v)
print(f"{j = }, {np.around(v, 2)}")
```
To see eigenvalues, divide the product $Av$ by $v$:
```
for j in range(N):
lam = np.around((A0 @ Vs[j]) / Vs[j], 2)
print(f"{j=}, {lam}")
print(V[:, 1])
LA.eig?
import numpy.linalg as NLA
L2, V2 = NLA.eig(A0.astype(complex))
np.around(V2, 2)
```
| github_jupyter |
# Testing performance of different 2D Feature detectors in OpenCV
Imports...
```
import cv2
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sn
import time
sn.set()
# Utilities
r2b = lambda x: cv2.cvtColor(x, cv2.COLOR_BGR2RGB)
r2ba = lambda x: cv2.cvtColor(x, cv2.COLOR_BGRA2RGBA)
```
## Create an artificial split image
Read image
```
im = cv2.imread('full.jpg')
```
Find an overlapping horizontal (width) split at: 5/8 and 3/8.
E.g. the left image will end at 5/8 the width, and the right image will start at 3/8 the width.
This was the images will overlap in 2/8 or 1/4 of the width.
```
im.shape[1] * (5./8.), im.shape[1] * (3./8.)
```
Using the above information we split the image horizontally.
```
im_left = im[:,:1617]
im_right = im[:,970:]
plt.subplot(131),plt.imshow(r2b(im)),plt.axis('off'),plt.title('Original')
plt.subplot(132),plt.imshow(r2b(im_left)),plt.axis('off'),plt.title('Left')
plt.subplot(133),plt.imshow(r2b(im_right)),plt.axis('off'),plt.title('Right');
h_orig,w_orig = im.shape[:2]
h,w = im_right.shape[:2]
```
## A feature-type-oblivious test
To measure the success of features we will need to change the feature types while keeping the same API.
Luckily this is easily possible in OpenCV.
The following function will provide us with a `cv2.Feature2D` feature detector for each algorithm as well as a `cv2.DetectorMatcher` to match the features.
```
# adapted from: https://github.com/opencv/opencv/blob/master/samples/python/find_obj.py
def init_feature(name):
chunks = name.split('-')
if chunks[0] == 'sift':
detector = cv2.xfeatures2d.SIFT_create(2500)
norm = cv2.NORM_L2
elif chunks[0] == 'surf':
detector = cv2.xfeatures2d.SURF_create(6500)
norm = cv2.NORM_L2
elif chunks[0] == 'orb':
detector = cv2.ORB_create(2500)
norm = cv2.NORM_HAMMING
elif chunks[0] == 'akaze':
detector = cv2.AKAZE_create(threshold=0.0065)
norm = cv2.NORM_HAMMING
elif chunks[0] == 'brisk':
detector = cv2.BRISK_create(100)
norm = cv2.NORM_HAMMING
else:
return None, None
if 'flann' in chunks:
FLANN_INDEX_KDTREE = 1
FLANN_INDEX_LSH = 6
if norm == cv2.NORM_L2:
flann_params = dict(algorithm = FLANN_INDEX_KDTREE, trees = 5)
else:
flann_params= dict(algorithm = FLANN_INDEX_LSH,
table_number = 6, # 12
key_size = 12, # 20
multi_probe_level = 1) #2
matcher = cv2.FlannBasedMatcher(flann_params, {}) # bug : need to pass empty dict (#1329)
else:
matcher = cv2.BFMatcher(norm)
return detector, matcher
# A utility function to align feature points using the given matching, also apply the 2-NN ratio test
def filter_matches(kp1, kp2, matches, ratio = 0.75):
mkp = [(kp1[m[0].queryIdx], kp2[m[0].trainIdx]) for m in matches if len(m) == 2 and m[0].distance < m[1].distance * ratio]
mkp1,mkp2 = zip(*mkp)
p1 = np.float32([kp.pt for kp in mkp1])
p2 = np.float32([kp.pt for kp in mkp2])
return p1, p2, mkp
```
Visualize some features
```
out = im_left.copy()
# get keypoints
detector, matcher = init_feature('surf-flann')
im_kpts, im_desc = detector.detectAndCompute(im_left, None)
cv2.drawKeypoints(out, im_kpts, out, flags=cv2.DRAW_MATCHES_FLAGS_DRAW_RICH_KEYPOINTS+cv2.DRAW_MATCHES_FLAGS_DRAW_OVER_OUTIMG)
plt.figure(figsize=(6,6))
plt.imshow(r2b(out)), plt.grid(False);
```
### Pre-cache the rotated images
In order to save time in the main loop, we will create the rotated images
```
num_stops = 19
w_r = int(np.sqrt(w*w+h*h)) # the diagonal of the image can serve as the width of the new images
h_r = int(np.sqrt(w*w+h*h)) # since a 45deg rotation will have the diagonal as width.
s_y,s_x = int(h_r/2-h/2),int(w_r/2-w/2) # offset for central-rotation
ims, masks = [],[]
for i,a in enumerate(np.linspace(-90,90,num_stops)):
im_right_ext = np.ones((h_r,w_r,3), np.uint8) * 255 # white border...
im_right_ext[s_y:s_y+h, s_x:s_x+w] = im_right # offset the image to the center
im_right_mask = np.zeros((h_r,w_r,3), np.uint8) # a mask is also needed
im_right_mask[s_y:s_y+h, s_x:s_x+w] = 255
M = cv2.getRotationMatrix2D((w_r/2.0+0.5,h_r/2.0+0.5), a, 1.0) # rotate about the center
ims += [cv2.warpAffine(im_right_ext, M, (w_r,h_r), borderValue=(255,255,255))]
masks += [cv2.warpAffine(im_right_mask, M, (w_r,h_r))] # also rotate the mask...
cv2.imwrite('rotations/%03d.jpg'%i, ims[-1])
```
The major test loop follows.
We will go through all feature types, extract the features for the left image and then for each of the rotated images. Then we match and align.
```
results = {}
for feature_name in ['akaze','surf','sift','orb','brisk']:
# extract features for left image
detector, matcher = init_feature(feature_name + '-flann')
im_kpts, im_desc = detector.detectAndCompute(im_left, None)
matcher.add([im_desc]) # cache the left features
results[feature_name] = {'MSE': [], 'time': []}
# loop the right image rotations
for i,a in enumerate(np.linspace(-90,90,num_stops)):
t = time.process_time()
# extract features
kpts,desc = detector.detectAndCompute(ims[i],masks[i][:,:,0])
# match with the left image
raw_matches = matcher.knnMatch(desc,k=2) #2
p1, p2, _ = filter_matches(kpts, im_kpts, raw_matches) # align and filter
results[feature_name]['time'].append(time.process_time() - t) # record execution time
if len(p1) < 4:
print('%d matches found, not enough for transform estimation' % len(p1))
continue
# recover the transformation (rotation + translation)
H, status = cv2.estimateAffine2D(p1, p2, method=cv2.RANSAC, ransacReprojThreshold=5.0)
p1, p2 = p1[status], p2[status]
warped = cv2.warpAffine(ims[i], H, (w_orig,h_orig)) # undo th rotation on the right image
warped_mask = cv2.warpAffine(masks[i], H, (w_orig,h_orig)) / 255.0
# stitch the images using the mask
stitched_out = np.zeros((h_orig,w_orig,3),np.uint8)
stitched_out[:,:w-1] = im_left
stitched_out = np.uint8(stitched_out * (1 - warped_mask)) + np.uint8(warped * warped_mask)
# calculate the metrics
MSE = np.mean((stitched_out-im)**2)
results[feature_name]['MSE'].append(MSE)
print('%s, angle= %d, %d / %d inliers/matched, MSE = %.3f' % (feature_name,
int(a),
np.sum(status),
len(status),
MSE))
from scipy.interpolate import make_interp_spline, BSpline
```
## Charting the results
```
xold = np.linspace(-90,90,num_stops)
for f in results:
plt.plot(xold,results[f]['MSE'],label=f)
plt.legend(loc='upper center',ncol=3,fancybox=True, shadow=True)
plt.ylim(0,65),plt.xticks(np.linspace(-90,90,13).astype(np.int))
plt.ylabel('MSE'),plt.xlabel('Rotation Angle')
plt.savefig('rotation_perf.svg')
times = [np.mean(results[r]['time']) for r in results]
mses = [np.mean(results[r]['MSE']) for r in results]
fig = plt.figure()
ax = fig.add_subplot(111) # Create matplotlib axes
ax2 = ax.twinx() # Create another axes that shares the same x-axis as ax.
indices = np.arange(len(results))
width = 0.35
ax.bar(indices - width/2, times, width = width, color='b', label='Mean Time')
ax.set_label('Maan Time')
ax.set_ylabel('Mean Time (seconds)')
ax.set_xlabel('Feature Type')
ax.set_ylim(0,7.7)
ax.axes.set_xticklabels([0]+list(results.keys()))
ax2.bar(indices + width/2, mses, width = width, color='r', label='Mean MSE')
ax2.set_ylabel('Mean MSE')
ax2.grid(None)
ax2.set_ylim(0,49)
fig.legend(loc='upper center', ncol=2,fancybox=True, shadow=True, bbox_to_anchor=(0.5,0.9))
plt.savefig('time_vs_mse.svg')
```
At this point we can decide on the best feature type for the job.
In this case it seems AKAZE is the best performer.
| github_jupyter |
# Operadores Booleanos
Un booleano es $1$ o $0$, verdadero o falso. Con operadores como **and**, **or** o **not**, se pueden realizar consultas nuevas combinando estos operadores.
```
# Define Variables
my_house = 120.0 # mts^2
your_apartment = 72.0 # mts^2
# my_house is bigger than 100 and smaller than 120?
print(my_house>100 and my_house<120)
# my_house smaller than 72 or bigger than 110?
print(my_house<72 or my_house>110)
# Double my_house smaller than triple your_apartment?
print(2*my_house < 3*your_apartment)
# Using not
print(not my_house > 100)
```
## Ejercicio 1
Para el siguiente codigo de python:
```python
x = 9
y = 10
not(not(x < 4) and not(y > 15 or y > 10))
````
Cuál sería el resultado si ejecutamos este código?
a. True
b. False
c. Correr este comando genera un error
# Estructuras de Control
## IF
Condición es de tipo booleano, y cuando ésta se cumple se ejecuta el bloque.
```python
if condicion1:
bloque1
elif condicion2:
bloque2
else:
bloque3
```
### Ejercicio 2
Teniendo este código en Python:
```python
area = 10.0
if(area < 9) :
print("small")
elif(area < 12) :
print("medium")
else :
print("large")
````
Cuál sera la salida si se ejecuta?
a. small
b. medium
c. large
d. Error de sintaxis
```
# Define variables
room = "bed"
size = 180.0
# f-elif-else construct for room
if room == "bed" :
print("looking around in the bedroom.")
elif room == "kit" :
print("looking around in the kitchen.")
else :
print("looking around elsewhere.")
# if statement for area
if size > 180 :
print("big place!")
elif size > 100 :
print("medium size, nice!")
else:
print("pretty small.")
```
### Ejercicio 3
Dado un número entero $n$, realice las siguientes acciones condicionales:
1. Si es impar imprima 'Extraño'
2. Si es par y esta en el rango inclusive de 2 a 5, imprima 'No Extraño'
3. Si es par y esta en el rango inclusive de 6 a 20, imprima 'Extraño'
4. Si es par y es más grande que 20, imprima 'No Extraño'
***Ejemplos:***
$$n = 3$$
$n$ es impar y los números impares son de tipo 'Extraño', así que debe de imprimir 'Extraño'.
$$n = 24$$
$n$ es par y es mayor que 20, los números pares mayores que 20 son de tipo ' No Extraño', así que debe de imprimir 'No Extraño'.
```
# Your code
```
## WHILE
El ciclo while se ejecuta mientras la condición sea cierta, si la condición es falsa al inicio, el bloque no se ejecuta y se pasan a ejecutar las sentencias que le siguen. En Python, cualquier valor entero distinto de cero es verdadero y 0 es falso. La condición también puede ser una lista o cualquier secuencia, siendo la secuencia vacía falsa. El cuerpo del bucle debe estar identado, ya que de este modo Python agrupa las sentencias.
```python
while condicion:
bloque
```
### Ejercicio 4
Cuántos impresiones hará este ciclo de while?
```python
x = 1
while x < 4 :
print(x)
x = x + 1
```
a. 0
b. 1
c. 2
d. 3
```
while True:
print("Ciclo Infinito")
# Initialize offset
offset = 8
# Code the while loop
while offset != 0:
print("correcting...")
offset = offset - 1 # You can use too: offset -= 1
print(offset)
# Iterate Lists
lista = ['a','e','i','o','u']
while lista:
print(lista.pop(0))
z = 3
while z != 1 : # any boolean condition could be placed here. Also, notice the : at the end!
print( f"z = {z}" )
if z % 2 == 0 : # if z is even, divide by 2
z = z // 2
else :
z = 3 * z + 1 # if z is odd, multiply by 3 add 1
```
## FOR
Ejecuta un bloque $n$ cantidad de veces, inicia desde $0$ hasta $n-1$. Python permite iterar no solo sobre progresiones aritméticas, sino que recorre los elementos de una secuencia, ya sea lista o cadena, en el orden en que estos aparecen en la misma.
```python
for idx in range(0,n):
bloque
for elem in lista:
bloque
```
El uso del ciclo para iterar la lista solo da acceso a todos los elementos de la lista, si también se desea acceder a la información del índice se puede usar el método `enumerate()`
```python
for indice,elemento in enumerate(lista):
print("Indice" + str(indice) + ":" + str(elemento)
````
### FOR sobre Listas
```
# areas list
areas = [11.25, 18.0, 20.0, 10.75, 9.50]
# for loop using enumerate()
for idx,a in enumerate(areas) :
print("room" + str(idx) + ":" + str(a))
# house list of lists
house = [["hallway", 11.25],
["kitchen", 18.0],
["living room", 20.0],
["bedroom", 10.75],
["bathroom", 9.50]]
# For structure
for elem in house:
print("the "+str(elem[0]) + " is "+ str(elem[1]) + "sqm")
```
### FOR sobre Diccionarios
```
# Definition of dictionary
europe = {'spain':'madrid', 'france':'paris', 'germany':'berlin',
'norway':'oslo', 'italy':'rome', 'poland':'warsaw', 'austria':'vienna' }
# Iterate over europe
for key in europe:
print("the capital of "+key+ " is "+europe[key])
# Iterate over europe.items()
for key, value in europe.items() :
print("the capital of "+key+ " is "+value)
# Iterate over europe.keys()
for key in europe.keys():
print("the capital of "+key+ " is "+europe[key])
# Iterate over europe.values()
for value in europe.values() :
print("the capitals is "+value)
```
### Ejercicio 5
Escriba una función que genere un diccionario donde las claves son los numeros del 1 al 15 (ambos incluidos) y los valores son el cuadrado de las claves.
Ejemplo:
```python
{
1 : 1,
2 : 4,
3 : 9,
...
}
```
```
# Your code
```
## Control de ciclos
### Break
Esta función se puede usar en ciclos FOR y WHILE y simplemente termina el ciclo actual y continua con la ejecución de la siguiente instrucción.
```
# Break in FOR
for char in "Python":
if char == "h":
break
print ("char : " + char)
# Break in FOR
for i in range(1, 12) :
print( f"i = {i}" )
if i == 7 :
print( "Found 7, everybody's favorite number! No need to keep working!")
break
# Break in WHILE
value = 10
while value > 0:
value = value -1
if value == 5:
break
print ("value : " + str(value))
print ("End Script")
```
### Continue
Esta función regresa al comienzo del ciclo, ignorando las siguientes lineas de código del ciclo actual e inicia la siguiente iteración. Se puede usar en ciclos FOR y WHILE.
```
# Continue in FOR
for char in "Python":
if char == "h":
continue
print ("char : " + char)
# Continue in FOR
for i in range(1, 12):
if i % 5 == 0 : # if i is a multiple of 5
print( "I don't like multiples of 5, skipping this one")
continue # this jumps directly to the beginning of the loop, skipping the rest of _this_ iteration!
print( f"i = {i}")
# Continue in WHILE
value = 10
while value > 0:
value = value -1
if value == 5:
continue
print ("value : " + str(value))
print ("End Script")
```
### Pass
Es una operación no sucede nada cuando se ejecuta. Se utiliza cuando se requiere por sintaxis una declaración pero no se quiere ejecutar ningún código. También se usa como un código temporal, hasta que se escriba el código final.
```
# Pass in FOR
for char in "Python":
if char == "h":
pass
print ("char : " + char)
# Pass in WHILE
value = 10
while value > 0:
value = value -1
if value == 5:
pass
print ("value : " + str(value))
print ("End Script")
```
# Errores y Excepciones
Hay dos tipos de errores diferentes: *errores de sintaxis* y *excepciones*.
## Errores de sintaxis
En los errores de sintaxis, el interprete repite la línea culpable del error.
```python
while True print('Hola mundo')
```
```python
File "<stdin>", line 1
while True print('Hola mundo')
^
SyntaxError: invalid syntax
```
## Excepciones
Sin importar que la declaración este sintácticamente correcta, puede generar un error cuando se intenta ejecutarla. Los errores detectados durante la ejecución se llaman excepciones pueden ser manejados por los programas en Python.
La declaración `try` nos permite manejar las excepciones y trabaja de la siguiente manera:
1. Se ejecuta el *bloque try* (el código entre la declaración `try`y `except`)
2. Si no ocurre una excepción el `boque except` se omite y termina la ejecución de la declaración `try`.
3. Si ocurre una excepción durante la ejecución del *bloque try*, el resto del bloque se omite. Si el tipo de excepción coincide con la excepción nombreada luevo de la palabra reservada `except`, se ejecuta el *bloque except*, y la ejecución continúa luego de la declaración `try`.
4. Si ocurre una excepción que no coincida con la excepción nombrada en el `except`, esta es una excepción no manejada y la ejecución se frena con un mensaje de error indicando lo que sucedió, por ejemplo: **ZeroDivisionError**, **NameError**, **TypeError** y **ValueError**.
```
# ZeroDivisionError
10 * (1/0)
# NameError
4 + spam*3
# TypeError
'2' + 2
#ValueError
float("juan")
# try-except structure with particular type of exception defined
my_string = "asd"
try :
my_string_as_float = 2 / 0 #float( my_string )
print( "the entered value could be converted directly to a number")
except ValueError as err :
# capture this particular type of exception and bind it to the name 'err'
print( "an exception was thrown: defaulting to special float value = nan")
a_as_float = float( "nan")
print(err)
#print( f"a_as_float = {a_as_float}")
# try-except structure with general type of exception
my_string = 12
try :
my_string_as_float = float( my_string )
print( "the entered value could be converted directly to a number")
except:
# capture this particular type of exception and bind it to the name 'err'
print( "an exception was thrown: defaulting to special float value = nan")
a_as_float = float( "nan")
print( f"a_as_float = {a_as_float}")
# try-except structure with general type of exception and print error
my_string = "asdad"
try :
my_string_as_float = float( my_string )
print( "the entered value could be converted directly to a number")
except Exception as inst:
# capture this particular type of exception and bind it to the name 'err'
print( f"an exception was thrown: {inst}")
a_as_float = float( "nan")
print( f"a_as_float = {a_as_float}")
# try-except structure with general type of exception
my_string = "asdad"
try :
my_string_as_float = float( my_string )
print( "the entered value could be converted directly to a number")
except:
# capture this particular type of exception and bind it to the name 'err'
print( "an exception was thrown: defaulting to special float value = nan")
a_as_float = float( "nan")
print( f"a_as_float = {a_as_float}")
```
La declaración `try` tiene otra clausula opcional `finally` que intenta definir acciones de limpieza que deben ser ejecutadas bajo ciertas circustancias. La cláusula `finally` siempre es ejecutada antes de salir de la declaración `try`, ya sea que una excepción haya ocurrido o no.
```
# structure try-except-finally
try:
#result = x / 0
result = 4/2
except ZeroDivisionError:
print("¡división por cero!")
else:
print("el resultado es", result)
finally:
print("ejecutando la clausula finally")
```
# Clases
Las clases proveen una forma de empaquetar datos y funcionalidad juntos. Al crear una nueva clase, se crea un nuevo *tipo* de objeto, permitiendo crear nuevas *instancias* de este tipo. Cada instancia de clase puede tener atributos adjuntos para mantener su estado. Las instancias de clase también pueden tener métodos (definidos por su clase) para modificar su estado.
La variable *self* es una instancia de la clase y no es ua palabra reservada de Python, cualquier etiqueta utilizada como primer parámetro tendría el mismo valor (no se aconseja usar otro valor por convención). Cada vez que declaremos un método en Python, vamos a tener uqe agregarle la variable *self* para que cuando sea invocado el método, Python pase el objeto instanciado y opere con los valores actuales de esa instancia.
```
# Definimos la clase con nombre Mascota
class Mascota:
# Atributos
especie = 'Gato'
_oculto = 2
def __init__(self,nombre,especie=especie):
"""
El método __init__ es una función para inicializar la clase, si no se especifica la especie toma el valor por defecto definido en el atributo.
"""
self.nombre = nombre
self.especie = especie
self._private=None
def __str__(self):
"""
El método __str__ es una función que proporciona una cadena informal de la representación del objeto.
"""
return "%s es un %s" % (self.nombre, self.especie)
def darNombre(self):
"""
Retorna el nombre de la mascota.
"""
return self.nombre
def darEspecie(self):
"""
Retorna la especie de la mascota.
"""
return self.especie
Teo = Mascota("Teo","Perro")
print(Teo)
Teo.especie='Gato'
print(Teo)
?Teo.darEspecie
```
# Funciones lambda
Lambdas es la forma de Python de definir funciones anónimas muy simples cuyo valor de retorno es el resultado de evaluar una sola expresión.
```python
lambda argumentos: resultado
```
Una de las razones por las que desees utilizar este tipo de funciones es por rapidez y en ciertos casos claridad del código.
```
# Define function
def function_sum(x,y):
return x + y
function_sum(2,3)
# Define lambda
lambda_function = lambda x,y: x + y
lambda_function(2,3)
```
## Usando Lambda con Sorted
```
data_recs = [
[ "Maria", 7.0, 8.0, 3.0 ],
[ "Juan" , 5.0, 2.0, 2.0 ],
[ "Mateo", 1.0, 4.0, 4.0 ],
]
```
Se quiere ordenar estos registros por el valor del primer componente numérico (índice 1 en la lista)
```
data_recs[:][1][1]
sorted(data_recs)
records_sorted = sorted(data_recs, key = lambda rec : rec[1])
records_sorted
```
## Ejercicio 6
Modifique el anterior ejemplo y ordene los registros en order descendiente por el ultimo record. Ayuda ( reverse = True )
```
help(sorted)
# Your Code
```
# Función filter()
Es una función que permite filtrar, a partir de una lista o iterador y una función condicional, es capaz de devolver una nueva colección con los elementos filtrados que cumplan la condición. En este tipo de función las funciones `lambda` son muy utiles.
La función `filter(funcion,iterable)` construye una lista con aquellos elementos en los cuales `funcion(iterable[i])` retorna `True`.
```
a = [0, 1, -1, -2, 3, -4, 5, 6, 7]
resultado = filter(lambda x: x > 0, a)
for val in resultado:
print(val)
list(resultado)
def multiple(numero): # Primero declaramos una función condicional
if numero % 5 == 0: # Comprobamos si un numero es múltiple de cinco
return True # Sólo devolvemos True si lo es
numeros = [2, 5, 10, 23, 50, 33]
list(filter(multiple, numeros))
list( filter(lambda numero: numero%5 == 0, numeros) )
```
## Filtrando objetos
```
# Definimos una clase persona
class Persona:
def __init__(self, nombre, edad):
self.nombre = nombre
self.edad = edad
def __str__(self):
return "{} de {} años".format(self.nombre, self.edad)
# Se crea una lista con 4 personas
personas = [
Persona("Juan", 35),
Persona("Marta", 16),
Persona("Manuel", 78),
Persona("Eduardo", 12)
]
# Visualizamos la lista
for p in personas:
print(p)
# Se desea filtrar los menores de 18
menores = filter(lambda persona: persona.edad < 18, personas)
# Visualizamos los objetos de la lista filtrada
for menor in menores:
print(menor)
```
# Función map()
Funciona similar a `filter()`, con la diferencia que en lugar de aplicar una condición a un elemento de una lista o secuencia, aplica una función sobre todos los elementos y como resultado se devuelve un iterable de tipo map.
```
numeros = [2, 5, 10, 23, 50, 33]
list( map(lambda x: x*2, numeros) )
```
La función `map()` se utiliza mucho junto a expresiones lambda ya que permite ahorrar esfuerzo creando ciclos FOR. Adicional se puede utilizar sobre más de un iterable con la condición que tengan la misma longitud.
```
a = [1, 2, 3, 4, 5]
b = [6, 7, 8, 9, 10]
list( map(lambda x,y : x*y, a,b) )
c = [11, 12, 13, 14, 15]
list( map(lambda x,y,z : x*y*z, a,b,c) )
```
## Mapeando objetos
```
personas = map(lambda p: Persona(p.nombre, p.edad+1), personas)
for persona in personas:
print(persona)
```
# List comprehensions
Es una funcionalidad que nos permite crear listas en una misma linea de codigo.
## Ciclos y List Comprenhensions
Supongamos que nos dan:
* Una lista de entrada input_arr = [elem1, elem2, elem3, ...]
* Una función fun
Queremos producir el array output_arr = [fun (elem1), fun (elem2), fun (elem3)], es decir, con aplicar fun a cada elemento de la matriz y devolver los resultados.
Aquí es cómo la mayoría de los programadores abordarían este problema:
```
input_arr = ["maria", "ana", "sara"]
capitalize = lambda a_str : a_str[0].upper() + a_str[1:]
# NON-Idiomatic Python code follows:
output_arr = []
for elem in input_arr :
output_arr.append( capitalize( elem ) )
output_arr
```
Eso es aproximadamente 3 líneas. No está mal, pero podría ser mucho mejor.
La solución idiomática de Python se llama **List (or for) comprehension**.
```python
[funcion(x) for item in list1]
```
```
output_arr2 = [ capitalize(elem) for elem in input_arr ]
print(output_arr2)
```
## Condicionales en List Comprehensions
En este caso se desea realizar un condicional para definir el valor, se tienen dos estructuras, si la condición solo tiene IF, o si se tiene la estructura IF-ELSE
```python
[result1 if conditional else result2 for item in list1 ]
[result for item in list1 if conditional]
```
```
squares_cubes = [n**2 if n%2 == 0 else n**3 for n in range(1,16)]
print(squares_cubes)
evens = [n for n in range(1,21) if n%2 == 0]
print(evens)
```
## List Comprehensions para ciclos anidados
También puede usarse para realizar operaciones en ciclos anidados como recorrer una lista de listas.
```python
[ <the_expression> for <element_a> in <iterable_a> (optional if <condition_a>)
for <element_b> in <iterable_b> (optional if <condition_b>)
for <element_c> in <iterable_c> (optional if <condition_c>)
... and so on ...]
```
```
matrix = [
[1, 2, 3, 4],
[5, 6, 7, 8],
[9, 10, 11, 12],
]
flatten = [n for row in matrix if sum(row)>11 for n in row if n%2==0]
print(flatten)
```
## List Comprehensions anidados
```
matrix = [
[1, 2, 3, 4],
[5, 6, 7, 8],
[9, 10, 11, 12],
]
transpose = [[row[n] for row in matrix] for n in range(4)]
print(transpose)
```
## Ejercicio 7
Defina una función avg1 que tome una lista que conste del nombre de una persona (str) y luego solo números y calcula y devuelve el promedio de esos números.
Por ejemplo:
```python
avg1 (['Mateo', 5.0, 2.0, 2.0])
```
debe devolver 3, es el resultado de realizar la siguiente operación: `((5 + 2 + 2) / 3))`
```
def avg1(lst):
# Your code
result = 0
return result
print(avg1(["mateo",5,2,2]))
```
## Ejercicio 8
Use list-comprehension para aplicar la función avg1 definida anteriormente a cada registro en data_recs.
```python
data_recs = [
[ "Juan" , 5.0, 2.0, 2.0 ],
[ "Maria", 7.0, 8.0, 3.0 ],
[ "Mateo", 1.0, 4.0, 4.0 ],
]
````
```
# Your Code
```
# Módulos y Paquetes
Los módulos son ficheros que contienen definiciones que se pueden importar en otros scripts para reutilizar funcionalidades. Se pueden organizar en paquetes y subpaquetes, adicional se importarian sólo cuando es necesario.
Nosotros podemos crear nuestros propios módulos o paquetes, o utilizar ya algunos creados por otras personas y que estan disponibles para su uso.
Para instalar un paquete en Python se realiza de la siguiente manera:
```python
pip install pandas
````
Si se desea desinstalar:
```python
pip uninstall pandas
````
```
!pip install pandas
!ls
!mkdir temp
```
## Módulos estándar
Estos son algunos módulos esenciales de Python:
###**collections**
Añade funcionalidades especificas a las listas.
```
from collections import Counter
l = [1,2,3,4,1,2,3,1,2,1]
Counter(l)
animales = "gato perro canario perro canario perro"
c = Counter(animales.split())
print(c)
```
###**datetime**
Utilizado para manejar las fechas y las horas.
```
from datetime import datetime
dt = datetime.now() # Fecha y hora actual
print(dt)
print(dt.year) # año
print(dt.month) # mes
print(dt.day) # día
print(dt.hour) # hora
print(dt.minute) # minutos
print(dt.second) # segundos
print(dt.microsecond) # microsegundos
print("{}:{}:{}".format(dt.hour, dt.minute, dt.second))
print("{}/{}/{}".format(dt.day, dt.month, dt.year))
from datetime import datetime, timedelta
dt = datetime.now()
print(dt.strftime("%A %d de %b del %y - %H:%M"))
# Generamos 14 días con 4 horas y 1000 segundos de tiempo
t = timedelta(days=14, hours=4, seconds=1000)
# Lo operamos con el datetime de la fecha y hora actual
dentro_de_dos_semanas = dt + t
print(dentro_de_dos_semanas.strftime("%A %d de %B del %Y - %H:%M"))
hace_dos_semanas = dt - t
print(hace_dos_semanas.strftime("%A %d de %B del %Y - %H:%M"))
```
###**math**
Incluye funciones matematicas.
```
import math
print(math.floor(3.99)) # Redondeo a la baja (suelo)
print(math.ceil(3.01)) # Redondeo al alta (techo)
print(math.pi) # Constante pi
print(math.e) # Constante e
```
###**random**
Sirve para generar contenidos aleatorios, escoger aleatoriamente valores y este tipo de cosas que hacen que un programa tenga comportamientos al azar.
```
import random
# Flotante aleatorio >= 0 y < 1.0
print(random.random())
# Flotante aleatorio >= 1 y <10.0
print(random.uniform(1,10))
# Entero aleatorio de 0 a 9, 10 excluído
print(random.randrange(10))
# Entero aleatorio de 0 a 100
print(random.randrange(0,101))
# Entero aleatorio de 0 a 100 cada 2 números, múltiples de 2
print(random.randrange(0,101,2))
# Entero aleatorio de 0 a 100 cada 5 números, múltiples de 5
print(random.randrange(0,101,5))
help(random)
# Letra aleatoria
print(random.choice('Hola mundo'))
# Elemento aleatorio
print(random.choice([1,2,3,4,5]))
# Dos elementos aleatorios
print(random.sample([1,2,3,4,5], 2))
# Barajar una lista, queda guardado
lista = [1,2,3,4,5]
random.shuffle(lista)
print(lista)
```
###**re**
Es el módulo de las expresiones regulares (Regular Expressions). Utilizando una sintaxis especial, sirven para hacer comprobaciones y búsquedas. Son especialmente útiles trabajando con cadenas de caracteres, pero incluye muchos métodos alternativos.
```
import re
patron1 = '[^A-Za-z0-9 ]+'
patron2 = '[^A-Za-z ]+'
cadena = 'Juan4 $Perez5%'
print(re.sub(patron1, '_', cadena))
print(re.sub(patron2, '_', cadena))
```
###**os**
Funcionalidades dependientes del sistema Operativo.
```
import os
# Conocer el Directorio Actual
print(os.getcwd())
# Saber si el directorio Existe
print(os.path.exists('/content'))
```
###**sys**
Nos permite conseguir información del entorno del sistema.
```
import sys
#Retorna el número de versión de Python
print(sys.version)
#Retorna la plataforma sobre la cuál se está ejecutando el intérprete
print(sys.platform)
```
###**csv**
El módulo CSV tiene varias funciones y clases disponibles para leer y escribir CSVs.
#### Integrando Google Drive con Colab
```
# Solo si se ejecuta desde Google Drive
from google.colab import drive
drive.mount('/content/gdrive')
```
#### Leyendo un archivo CSV
El archivo que se va a cargar fue tomado de los datos abiertos de Accidentes de tránsito registrados por la Secretaría de Movilidad de la Alcaldía de Medellin, desde el año 2014 al año en curso. [Fuente](http://medata.gov.co/dataset/accidentalidad)
```
from csv import reader
opened_file = open('accidentalidad.csv') # Ruta del archivo si falla podria ser necesario adicionar encoding="utf-8"
read_file = reader(opened_file,delimiter=';')
read_file
# Visualizando la información
apps_data = list(read_file)
apps_data[:2]
# Funcion que genera diccionario de frecuencias
def freq_table(data_set, index):
frequency_table = {}
for row in data_set[1:]:
value = row[index]
if value in frequency_table:
frequency_table[value] += 1
else:
frequency_table[value] = 1
return frequency_table
# Clases de Accidentes
freq_table(apps_data,5)
# Localizacion de Accidentes
freq_table(apps_data,9)
# Gravedad de los Accidentes
freq_table(apps_data,17)
```
| github_jupyter |
# Logistic regression model
## Setup
```
!git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
%cd -q /pyprobml/scripts
!pip install -q optax
!pip install -q blackjax
!pip install -q sgmcmcjax
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import itertools
import warnings
from functools import partial
import jax
import jax.numpy as jnp
from jax.random import uniform, normal, bernoulli, split
from jax import jit, grad, value_and_grad, vmap
from jax.experimental import optimizers
from jax.scipy.special import logsumexp
from blackjax import nuts, stan_warmup
import optax
import sgmcmc_subspace_lib as sub
from sgmcmc_utils import build_optax_optimizer, build_nuts_sampler
from sgmcmcjax.samplers import *
```
## Generate Data
This part is based on https://github.com/jeremiecoullon/SGMCMCJax/blob/master/docs/nbs/models/logistic_regression.py
```
#ignore by GPU/TPU message (generated by jax module)
warnings.filterwarnings("ignore", message='No GPU/TPU found, falling back to CPU.')
# Sample initial beta values from random normal
def init_params(rng_key, d):
return normal(rng_key, (d,))
def gen_cov_mat(key, d, rho):
Sigma0 = np.diag(np.ones(d))
for i in range(1,d):
for j in range(0, i):
Sigma0[i,j] = (uniform(key)*2*rho - rho)**(i-j)
Sigma0[j,i] = Sigma0[i,j]
return jnp.array(Sigma0)
def logistic(theta, x):
return 1/(1+jnp.exp(-jnp.dot(theta, x)))
def gen_data(key, dim, N):
"""
Generate data with dimension `dim` and `N` data points
Parameters
----------
key: uint32
random key
dim: int
dimension of data
N: int
Size of dataset
Returns
-------
theta_true: ndarray
Theta array used to generate data
X: ndarray
Input data, shape=(N,dim)
y: ndarray
Output data: 0 or 1s. shape=(N,)
"""
key, theta_key, cov_key, x_key = split(key, 4)
rho = 0.4
print(f"Generating data, with N={N} and dim={dim}")
theta_true = normal(theta_key, shape=(dim, ))*jnp.sqrt(10)
covX = gen_cov_mat(cov_key, dim, rho)
X = jnp.dot(normal(x_key, shape=(N,dim)), jnp.linalg.cholesky(covX))
p_array = batch_logistic(theta_true, X)
keys = split(key, N)
y = batch_benoulli(keys, p_array).astype(jnp.int32)
return theta_true, X, y
@jit
def predict(params, inputs):
return batch_logistic(params, inputs) > 0.5
@jit
def accuracy(params, batch):
inputs, targets = batch
predicted_class = predict(params, inputs)
return jnp.mean(predicted_class == targets)
@jit
def loglikelihood(theta, x_val, y_val):
return -logsumexp(jnp.array([0., (1.-2.*y_val)*jnp.dot(theta, x_val)]))
@jit
def logprior(theta):
return -(0.5/10)*jnp.dot(theta,theta)
batch_logistic = jit(vmap(logistic, in_axes=(None, 0)))
batch_benoulli = vmap(bernoulli, in_axes=(0, 0))
batch_loglikelihood = vmap(loglikelihood, in_axes=(None, 0, 0))
dim = 10 # Choose a dimension for the parameters (10, 50,100 in the paper)
subspace_dim = 2 # Choose a dimension for the subspace parameters
ndata = 10000 # Number of data points
nwarmup = 1000 # Number of iterations during warmup phase
nsamples = 10000 # Number of SGMCMC iterations
nsamplesCV = nsamples // 2
key = jax.random.PRNGKey(42)
theta_true, X, y = gen_data(key, dim, ndata)
batch_size = int(0.01*X.shape[0])
data = (X, y)
init_key, key = split(key)
theta_init = init_params(init_key, dim)
```
## SGD
```
niters = 5000
learning_rate = 6e-5
opt = optax.sgd(learning_rate=learning_rate)
optimizer = build_optax_optimizer(opt, loglikelihood, logprior,
data, batch_size, pbar=False)
opt_key, key = split(key)
sgd_params, logpost_array = optimizer(opt_key, niters, theta_init)
train_accuracy = accuracy(sgd_params, data)
print("Training set accuracy {}".format(train_accuracy))
plt.plot(-logpost_array, color='tab:orange')
plt.show()
```
### Subspace Model
```
sub_opt_key, key = split(key)
sgd_sub_params, _, opt_log_post_trace, _ = sub.subspace_optimizer(
sub_opt_key, loglikelihood, logprior, theta_init, data, batch_size,
subspace_dim, nwarmup, nsamples, opt, pbar=False)
train_accuracy = accuracy(sgd_sub_params, data)
print("Training set accuracy {}".format(train_accuracy))
plt.plot(-opt_log_post_trace, color='tab:pink')
plt.show()
```
## NUTS
```
nuts_sampler = build_nuts_sampler(nwarmup, loglikelihood, logprior,
data, batch_size=ndata, pbar=False)
nuts_key, key = split(key)
nuts_params = nuts_sampler(nuts_key, nsamples//10, theta_init)
train_accuracy = accuracy(jnp.mean(nuts_params, axis=0), data)
print("Training set accuracy {}".format(train_accuracy))
```
### Subspace Model
```
build_nuts_sampler_partial = partial(build_nuts_sampler, nwarmup=nwarmup)
nuts_key, key = split(key)
nuts_sub_params = sub.subspace_sampler(nuts_key, loglikelihood, logprior,
theta_init, build_nuts_sampler_partial,
data, batch_size, subspace_dim, nsamples,
nsteps_full=0, nsteps_sub=0, use_cv=False, opt=None, pbar=False)
```
## SGLD
```
dt = 1e-5
# Run sampler
sgld_sampler = build_sgld_sampler(dt, loglikelihood, logprior, data, batch_size)
sgld_key, key = split(key)
sgld_output = sgld_sampler(sgld_key, nsamples, theta_init)
```
### Subspace Sampler
```
build_sgld_sampler_partial = partial(build_sgld_sampler, dt=dt)
sgld_key, key = split(key)
sgld_sub_output = sub.subspace_sampler(sgld_key, loglikelihood, logprior,
theta_init, build_sgld_sampler_partial,
data, batch_size, subspace_dim, nsamples,
nsteps_full=0, nsteps_sub=0, use_cv=False,
opt=None, pbar=False)
```
## SGLDCV
```
dt = 1e-5
sgldCV_sampler = build_sgldCV_sampler(dt, loglikelihood, logprior,
data, batch_size, sgd_params)
sgldCV_key, key = split(key)
sgldCV_output = sgldCV_sampler(sgldCV_key, nsamplesCV, sgd_params)
```
### Subspace Sampler
```
build_sgldCV_sampler_partial = partial(build_sgldCV_sampler, dt=dt)
sgldCV_key, key = split(key)
sgldCV_sub_output = sub.subspace_sampler(sgldCV_key, loglikelihood, logprior,
theta_init, build_sgldCV_sampler_partial,
data, batch_size, subspace_dim, nsamples,
nsteps_full=niters//2, nsteps_sub=niters//2,
use_cv=True, opt=opt, pbar=False)
```
## SGHMC
```
L = 5
dt = 1e-6
sghmc_sampler = build_sghmc_sampler(dt, L, loglikelihood,
logprior, data, batch_size)
sghmc_key, key = split(key)
sghmc_output = sghmc_sampler(sghmc_key, nsamples, theta_init)
```
### Subspace Sampler
```
build_sghmc_sampler_partial = partial(build_sghmc_sampler, dt=dt, L=L)
sghmc_key, key = split(key)
sghmc_sub_output = sub.subspace_sampler(sghmc_key, loglikelihood, logprior,
theta_init, build_sghmc_sampler_partial,
data, batch_size, subspace_dim,
nsamples, nsteps_full=0, nsteps_sub=0,
use_cv=False, opt=None, pbar=False)
```
## SGHMCCV
```
dt = 1e-7 # step size parameter
L = 5
sghmcCV_sampler = build_sghmcCV_sampler(dt, L, loglikelihood, logprior, data, batch_size, sgd_params)
sghmcCV_key, key = split(key)
sghmcCV_output = sghmcCV_sampler(sghmcCV_key, nsamplesCV, sgd_params)
```
### Subspace Sampler
```
build_sghmcCV_sampler_partial = partial(build_sghmcCV_sampler, dt=dt, L=L)
sghmcCV_key, key = split(key)
sghmcCV_sub_output = sub.subspace_sampler(sghmcCV_key, loglikelihood, logprior,
theta_init, build_sghmcCV_sampler_partial,
data, batch_size, subspace_dim, nsamples,
nsteps_full=niters//2, nsteps_sub=niters//2,
use_cv=True, opt=opt, pbar=False)
```
## SGNHT
```
dt = 1e-6 # step size parameter
a = 0.02
sgnht_sampler = build_sgnht_sampler(dt, loglikelihood, logprior,
data, batch_size, a=a)
sgnht_key, key = split(key)
sgnht_output = sgnht_sampler(sgnht_key, nsamples, theta_init)
```
### Subspace Sampler
```
build_sgnht_sampler_partial = partial(build_sgnht_sampler, dt=dt, a=a)
sgnht_key, key = split(key)
sgnht_sub_output = sub.subspace_sampler(sgnht_key, loglikelihood, logprior,
theta_init, build_sgnht_sampler_partial,
data, batch_size, subspace_dim, nsamples,
nsteps_full=0, nsteps_sub=0, use_cv=False,
opt=opt, pbar=False)
```
## SGHNTCV
```
dt = 1e-6 # step size parameter
a = 0.02
sgnhtCV_sampler = build_sgnhtCV_sampler(dt, loglikelihood, logprior,
data, batch_size, sgd_params, a=a)
sgnhtCV_key, key = split(key)
sgnhtCV_output = sgnhtCV_sampler(sgnhtCV_key, nsamplesCV, sgd_params)
```
### Subspace Sampler
```
build_sgnhtCV_sampler_partial = partial(build_sgnhtCV_sampler, dt=dt, a=a)
sgnhtCV_key, key = split(key)
sgnhtCV_sub_output = sub.subspace_sampler(sgnhtCV_key, loglikelihood, logprior,
theta_init, build_sgnhtCV_sampler_partial,
data, batch_size, subspace_dim, nsamples,
nsteps_full=niters//2, nsteps_sub=niters//2,
use_cv=True, opt=opt, pbar=False)
```
## ULA - SGLD with the full dataset
```
dt = 4e-5 # step size parameter
ula_sampler = build_sgld_sampler(dt, loglikelihood, logprior,
data, batch_size=ndata)
ula_key, key = split(key)
ula_output = ula_sampler(ula_key, nsamples, theta_init)
```
### Subspace Sampler
```
build_sgld_sampler_partial = partial(build_sgld_sampler, dt=dt)
ula_key, key = split(key)
ula_sub_output = sub.subspace_sampler(ula_key, loglikelihood, logprior,
theta_init, build_sgld_sampler_partial,
data, ndata, subspace_dim, nsamples,
nsteps_full=0, nsteps_sub=0, use_cv=False,
opt=None, pbar=False)
```
## Trace plots
```
def trace_plot(outs):
nrows, ncols = 2, 4
fig, axes = plt.subplots(nrows=nrows, ncols=ncols, figsize=(16, 12))
for ax, (title, out) in zip(axes.flatten(), outs.items()):
ax.plot(out)
ax.set_title(title)
ax.set_xlabel("Iteration")
ax.grid(color='white', linestyle='-', linewidth=2)
ax.set_axisbelow(True)
ax.set_facecolor("#EAEBF0")
plt.tight_layout()
plt.savefig("traceplot.pdf", dpi=300)
plt.show()
outs = {"STAN": nuts_params, "SGLD": sgld_output, "SGLDCV": sgldCV_output, "ULA": ula_output, "SGHMC": sghmc_output, "SGHMCCV":sghmcCV_output, "SGNHT": sgnht_output, "SGNHTCV": sgnhtCV_output}
trace_plot(outs)
subspace_outs = {"STAN": nuts_sub_params, "SGLD": sgld_sub_output, "SGLDCV": sgldCV_sub_output, "ULA": ula_sub_output, "SGHMC": sghmc_sub_output, "SGHMCCV":sghmcCV_sub_output, "SGNHT": sgnht_sub_output, "SGNHTCV": sgnhtCV_sub_output}
trace_plot(subspace_outs)
```
| github_jupyter |
# Finding locations to establish temporary emergency facilities
Run this notebook to create a Decision Optimization model with Decision Optimization for Watson Studio and deploy the model using Watson Machine Learning.
The deployed model can later be accessed using the [Watson Machine Learning client library](https://wml-api-pyclient-dev-v4.mybluemix.net/) to find optimal location based on given constraints.
The model created here is a basic Decision Optimization model. The main purpose is to demonstrate creating a model and deploying using Watson Machine Learning. This model can and should be improved upon to include better constraints that can provide a more optimal solution.
## Steps
**Build and deploy model**
1. [Provision a Watson Machine Learning service](#provision-a-watson-machine-learning-service)
1. [Set up the Watson Machine Learning client library](#set-up-the-watson-machine-learning-client-library)
1. [Build the Decision Optimization model](#build-the-decision-optimization-model)
1. [Deploy the Decision Optimization model](#deploy-the-decision-optimization-model)
**Test the deployed model**
1. [Generate an API Key from the HERE Developer Portal](#generate-an-api-key-from-the-here-developer-portal)
1. [Query HERE API for Places](#query-here-api-for-places)
1. [Create and monitor a job to test the deployed model](#create-and-monitor-a-job-to-test-the-deployed-model)
1. [Extract and display solution](#extract-and-display-solution)
<br>
### Provision a Watson Machine Learning service
- If you do not have an IBM Cloud account, [register for a free trial account](https://cloud.ibm.com/registration).
- Log into [IBM Cloud](https://cloud.ibm.com/login)
- Create a [create a Watson Machine Learning instance](https://cloud.ibm.com/catalog/services/machine-learning)
<br>
### Set up the Watson Machine Learning client library
Install the [Watson Machine Learning client library](https://wml-api-pyclient-dev-v4.mybluemix.net/). This notebook uses the preview Python client based on v4 of Watson Machine Learning APIs.
> **Important** Do not load both (V3 and V4) WML API client libraries into a notebook.
```
# Uninstall the Watson Machine Learning client Python client based on v3 APIs
!pip uninstall watson-machine-learning-client -y
# Install the WML client API v4
!pip install watson-machine-learning-client-V4
```
<br>
#### Create a client instance
Use your [Watson Machine Learning service credentials](https://dataplatform.cloud.ibm.com/docs/content/wsj/analyze-data/ml-get-wml-credentials.html) and update the next cell.
```
# @hidden_cell
WML_API_KEY = '...'
WML_INSTANCE_ID = '...'
WML_URL = 'https://us-south.ml.cloud.ibm.com'
from watson_machine_learning_client import WatsonMachineLearningAPIClient
# Instantiate a client using credentials
wml_credentials = {
'apikey': WML_API_KEY,
'instance_id': WML_INSTANCE_ID,
'url': WML_URL
}
client = WatsonMachineLearningAPIClient(wml_credentials)
client.version
```
<br>
### Build the Decision Optimization model
- The Decision Optimization model will be saved to a `model.py` file in a subdirectory (i.e., `model/`) of the current working directory.
- The model will be placed in a tar archive and uploaded to Watson Machine Learning.
Set up variables for model and deployment
```
import os
model_dir = 'model'
model_file = 'model.py'
model_path = '{}/{}'.format(model_dir, model_file)
model_tar = 'model.tar.gz'
model_tar_path = '{}/{}'.format(os.getcwd(), model_tar)
model_name = 'DO_HERE_DEMO'
model_desc = 'Finding locations for short-term emergency facilities'
deployment_name = 'DO_HERE_DEMO Deployment'
deployment_desc = 'Deployment of DO_HERE_DEMO model'
print(model_path)
print(model_tar_path)
```
<br>
#### Create the model.py in a model subdirectory
Use the `mkdir` and `write_file` commands to create the subdirectory and write the model code to a file.
```
%mkdir $model_dir
%%writefile $model_path
from docplex.util.environment import get_environment
from os.path import splitext
import pandas
from six import iteritems
import json
def get_all_inputs():
'''Utility method to read a list of files and return a tuple with all
read data frames.
Returns:
a map { datasetname: data frame }
'''
result = {}
env = get_environment()
for iname in [f for f in os.listdir('.') if splitext(f)[1] == '.csv']:
with env.get_input_stream(iname) as in_stream:
df = pandas.read_csv(in_stream)
datasetname, _ = splitext(iname)
result[datasetname] = df
return result
def write_all_outputs(outputs):
'''Write all dataframes in ``outputs`` as .csv.
Args:
outputs: The map of outputs 'outputname' -> 'output df'
'''
for (name, df) in iteritems(outputs):
if isinstance(df, pandas.DataFrame):
csv_file = '%s.csv' % name
print(csv_file)
with get_environment().get_output_stream(csv_file) as fp:
if sys.version_info[0] < 3:
fp.write(df.to_csv(index=False, encoding='utf8'))
else:
fp.write(df.to_csv(index=False).encode(encoding='utf8'))
elif isinstance(df, str):
txt_file = '%s.txt' % name
with get_environment().get_output_stream(txt_file) as fp:
fp.write(df.encode(encoding='utf8'))
if len(outputs) == 0:
print('Warning: no outputs written')
%%writefile -a $model_path
from docplex.mp.model import Model
from statistics import mean
def get_distance(routes_df, start, destination):
s = getattr(start, 'geocode', start)
d = getattr(destination, 'geocode', destination)
row = routes_df.loc[
(routes_df['start'] == s) &
(routes_df['destination'] == d)
]
return row['distance'].values[0]
def build_and_solve(places_df, routes_df, number_sites=3):
print('Building and solving model')
mean_dist = mean(routes_df['distance'].unique())
p_only = places_df.loc[places_df['is_medical'] == False]
h_only = places_df.loc[places_df['is_medical'] == True]
places = list(p_only.itertuples(name='Place', index=False))
postal_codes = p_only['postal_code'].unique()
hospital_geocodes = h_only['geocode'].unique()
mdl = Model(name='temporary emergency sites')
## decision variables
places_vars = mdl.binary_var_dict(places, name='is_place')
postal_link_vars = mdl.binary_var_matrix(postal_codes, places, 'link')
hosp_link_vars = mdl.binary_var_matrix(hospital_geocodes, places, 'link')
## objective function
# minimize hospital distances
h_total_distance = mdl.sum(hosp_link_vars[h, p] * abs(mean_dist - get_distance(routes_df, h, p)) for h in hospital_geocodes for p in places)
mdl.minimize(h_total_distance)
## constraints
# match places with their correct postal_code
for p in places:
for c in postal_codes:
if p.postal_code != c:
mdl.add_constraint(postal_link_vars[c, p] == 0, 'ct_forbid_{0!s}_{1!s}'.format(c, p))
# # each postal_code should have one only place
# mdl.add_constraints(
# mdl.sum(postal_link_vars[c, p] for p in places) == 1 for c in postal_codes
# )
# # each postal_code must be associated with a place
# mdl.add_constraints(
# postal_link_vars[c, p] <= places_vars[p] for p in places for c in postal_codes
# )
# solve for 'number_sites' places
mdl.add_constraint(mdl.sum(places_vars[p] for p in places) == number_sites)
## model info
mdl.print_information()
stats = mdl.get_statistics()
## model solve
mdl.solve(log_output=True)
details = mdl.solve_details
status = '''
Model stats
number of variables: {}
number of constraints: {}
Model solve
time (s): {}
status: {}
'''.format(
stats.number_of_variables,
stats.number_of_constraints,
details.time,
details.status
)
possible_sites = [p for p in places if places_vars[p].solution_value == 1]
return possible_sites, status
%%writefile -a $model_path
import pandas
def run():
# Load CSV files into inputs dictionary
inputs = get_all_inputs()
places_df = inputs['places']
routes_df = inputs['routes']
site_suggestions, status = build_and_solve(places_df, routes_df)
solution_df = pandas.DataFrame(site_suggestions)
outputs = {
'solution': solution_df,
'status': status
}
# Generate output files
write_all_outputs(outputs)
run()
```
<br>
#### Create the model tar archive
Use the `tar` command to create a tar archive with the model file.
```
import tarfile
def reset(tarinfo):
tarinfo.uid = tarinfo.gid = 0
tarinfo.uname = tarinfo.gname = 'root'
return tarinfo
tar = tarfile.open(model_tar, 'w:gz')
tar.add(model_path, arcname=model_file, filter=reset)
tar.close()
```
<br>
### Deploy the Decision Optimization model
Store model in Watson Machine Learning with:
- the tar archive previously created,
- metadata including the model type and runtime
```
# All available meta data properties
client.repository.ModelMetaNames.show()
# All available runtimes
client.runtimes.list(pre_defined=True)
```
<br>
#### Upload the model to Watson Machine Learning
Configure the model metadata and set the model type (i.e., `do-docplex_12.9`) and runtime (i.e., `do_12.9`)
```
import os
model_metadata = {
client.repository.ModelMetaNames.NAME: model_name,
client.repository.ModelMetaNames.DESCRIPTION: model_desc,
client.repository.ModelMetaNames.TYPE: 'do-docplex_12.9',
client.repository.ModelMetaNames.RUNTIME_UID: 'do_12.9'
}
model_details = client.repository.store_model(model=model_tar_path, meta_props=model_metadata)
model_uid = client.repository.get_model_uid(model_details)
print('Model GUID: {}'.format(model_uid))
```
<br>
#### Create a deployment
Create a batch deployment for the model, providing deployment metadata and model UID.
```
deployment_metadata = {
client.deployments.ConfigurationMetaNames.NAME: deployment_name,
client.deployments.ConfigurationMetaNames.DESCRIPTION: deployment_desc,
client.deployments.ConfigurationMetaNames.BATCH: {},
client.deployments.ConfigurationMetaNames.COMPUTE: {'name': 'S', 'nodes': 1}
}
deployment_details = client.deployments.create(model_uid, meta_props=deployment_metadata)
deployment_uid = client.deployments.get_uid(deployment_details)
print('Deployment GUID: {}'.format(deployment_uid))
```
<br>
**Congratulations!** The model has been succesfully deployed. Please make a note of the deployment UID.
<br>
## Test the deployed model
### Generate an API Key from the HERE Developer Portal
To test your deployed model using actual data from HERE Location services, you'll need an API key.
Follow the instructions outlined in the [HERE Developer Portal](https://developer.here.com/sign-up) to [generate an API key](https://developer.here.com/documentation/authentication/dev_guide/topics/api-key-credentials.html).
Use your [HERE.com API key](https://developer.here.com/sign-up) and update the next cell.
```
# @hidden_cell
HERE_APIKEY = '...'
```
<br>
Set up helper functions to query HERE APIs
```
import re
import requests
geocode_endpoint = 'https://geocode.search.hereapi.com/v1/geocode?q={address}&apiKey={api_key}'
browse_endpoint = 'https://browse.search.hereapi.com/v1/browse?categories=%s&at=%s&apiKey=%s'
matrix_routing_endpoint = 'https://matrix.route.ls.hereapi.com/routing/7.2/calculatematrix.json?mode=%s&summaryAttributes=%s&apiKey=%s'
coordinates_regex = '^[-+]?([1-8]?\d(\.\d+)?|90(\.0+)?),\s*[-+]?(180(\.0+)?|((1[0-7]\d)|([1-9]?\d))(\.\d+)?)$'
def is_geocode (location):
geocode = None
if isinstance(location, str):
l = location.split(',')
if len(l) == 2:
geocode = '{},{}'.format(l[0].strip(), l[1].strip())
elif isinstance(location, list) and len(location) == 2:
geocode = ','.join(str(l) for l in location)
if geocode is not None and re.match(coordinates_regex, geocode):
return [float(l) for l in geocode.split(',')]
else:
return False
def get_geocode (address):
g = is_geocode(address)
if not g:
url = geocode_endpoint.format(address=address, api_key=HERE_APIKEY)
response = requests.get(url)
if response.ok:
jsonResponse = response.json()
position = jsonResponse['items'][0]['position']
g = [position['lat'], position['lng']]
else:
print(response.text)
return g
def get_browse_url (location, categories, limit=25):
categories = ','.join(c for c in categories)
geocode = get_geocode(location)
coordinates = ','.join(str(g) for g in geocode)
browse_url = browse_endpoint % (
categories,
coordinates,
HERE_APIKEY
)
if limit > 0:
browse_url = '{}&limit={}'.format(browse_url, limit)
return browse_url
def browse_places (location, categories=[], results_limit=100):
places_list = []
browse_url = get_browse_url(location, categories, limit=results_limit)
response = requests.get(browse_url)
if response.ok:
json_response = response.json()
places_list = json_response['items']
else:
print(response.text)
return places_list
def get_places_nearby (location, categories=[], results_limit=100, max_distance_km=50):
places_list = browse_places(location, categories=categories, results_limit=results_limit)
filtered_places = []
for p in places_list:
if p['distance'] <= max_distance_km * 1000:
filtered_places.append(Place(p))
return filtered_places
def get_hospitals_nearby (location, results_limit=100, max_distance_km=50):
h_cat = ['800-8000-0159']
hospitals_list = browse_places(location, categories=h_cat, results_limit=results_limit)
filtered_hospitals = []
for h in hospitals_list:
if h['distance'] <= max_distance_km * 1000:
filtered_hospitals.append(Place(h, is_medical=True))
return filtered_hospitals
def get_matrix_routing_url ():
route_mode = 'shortest;car;traffic:disabled;'
summary_attributes = 'routeId,distance'
matrix_routing_url = matrix_routing_endpoint % (
route_mode,
summary_attributes,
HERE_APIKEY
)
return matrix_routing_url
def get_route_summaries (current_geocode, places, hospitals):
# Request should not contain more than 15 start positions
num_starts = 15
postal_codes_set = set()
postal_codes_geocodes = []
places_waypoints = {}
for i, p in enumerate(places):
if p.postal_code:
postal_codes_set.add('{}:{}'.format(p.postal_code, p.country))
places_waypoints['destination{}'.format(i)] = p.geocode
for p in postal_codes_set:
geocode = get_geocode(p)
postal_codes_geocodes.append({
'postal_code': p.split(':')[0],
'geocode': ','.join(str(g) for g in geocode)
})
current = {
'geocode': ','.join(str(g) for g in current_geocode)
}
start_geocodes = [current] + postal_codes_geocodes + [h.to_dict() for h in hospitals]
start_coords = [
start_geocodes[i:i+num_starts]
for i in range(0, len(start_geocodes), num_starts)
]
route_summaries = []
matrix_routing_url = get_matrix_routing_url()
for sc in start_coords:
start_waypoints = {}
for i, s in enumerate(sc):
start_waypoints['start{}'.format(i)] = s['geocode']
coords = {**start_waypoints, **places_waypoints}
response = requests.post(matrix_routing_url, data = coords)
if not response.ok:
print(response.text)
else:
json_response = response.json()
for entry in json_response['response']['matrixEntry']:
start_geocode = start_waypoints['start{}'.format(entry['startIndex'])]
dest_geocode = places_waypoints[
'destination{}'.format(entry['destinationIndex'])
]
for s in sc:
if 'address' not in s and 'postal_code' in s and s['geocode'] == start_geocode:
route_summaries.append({
'start': s['postal_code'],
'destination': dest_geocode,
'distance': entry['summary']['distance'],
'route_id': entry['summary']['routeId']
})
break
route_summaries.append({
'start': start_geocode,
'destination': dest_geocode,
'distance': entry['summary']['distance'],
'route_id': entry['summary']['routeId']
})
return route_summaries
```
<br>
Define a Place class
```
class Place(object):
def __init__(self, p, is_medical=False):
self.id = p['id']
self.title = p['title']
self.address = p['address']['label'] if 'label' in p['address'] else p['address']
self.postal_code = p['address']['postalCode'] if 'postalCode' in p['address'] else p['postal_code']
self.distance = p['distance']
self.primary_category = p['categories'][0]['id'] if 'categories' in p else p['primary_category']
self.geocode = '{},{}'.format(p['position']['lat'], p['position']['lng']) if 'position' in p else p['geocode']
self.country = p['address']['countryCode'] if 'countryCode' in p['address'] else p['country']
self.is_medical = p['is_medical'] if 'is_medical' in p else is_medical
if isinstance(self.is_medical, str):
self.is_medical = self.is_medical.lower() in ['true', '1']
def to_dict(self):
location = self.geocode.split(',')
return({
'id': self.id,
'title': self.title,
'address': self.address,
'postal_code': self.postal_code,
'distance': self.distance,
'primary_category': self.primary_category,
'geocode': self.geocode,
'country': self.country,
'is_medical': self.is_medical
})
def __str__(self):
return self.address
```
<br>
### Query HERE API for Places
Use the HERE API to get a list of Places in the vicinity of an address
Example of `Place` entity returned by HERE API:
```json
{
'title': 'Duane Street Hotel',
'id': 'here:pds:place:840dr5re-fba2a2b91f944ee4a699eea7556896bd',
'resultType': 'place',
'address': {
'label': 'Duane Street Hotel, 130 Duane St, New York, NY 10013, United States',
'countryCode': 'USA',
'countryName': 'United States',
'state': 'New York',
'county': 'New York',
'city': 'New York',
'district': 'Tribeca',
'street': 'Duane St',
'postalCode': '10013',
'houseNumber': '130'
},
'position': { 'lat': 40.71599, 'lng': -74.00735 },
'access': [ { 'lat': 40.71608, 'lng': -74.00728 } ],
'distance': 161,
'categories': [
{ 'id': '100-1000-0000' },
{ 'id': '200-2000-0000' },
{ 'id': '500-5000-0000' },
{ 'id': '500-5000-0053' },
{ 'id': '500-5100-0000' },
{ 'id': '700-7400-0145' }
],
'foodTypes': [ { 'id': '101-000' } ],
'contacts': [ ],
'openingHours': [
{
'text': [ 'Mon-Sun: 00:00 - 24:00' ],
'isOpen': true,
'structured': [
{
'start': 'T000000',
'duration': 'PT24H00M',
'recurrence': 'FREQ:DAILY;BYDAY:MO,TU,WE,TH,FR,SA,SU'
}
]
}
]
}
```
```
address = 'New York, NY'
max_results = 20
# HERE Place Category System
# https://developer.here.com/documentation/geocoding-search-api/dev_guide/topics-places/places-category-system-full.html
places_categories = ['500-5000'] # Hotel-Motel
current_geocode = get_geocode(address)
places = get_places_nearby(
current_geocode,
categories=places_categories,
results_limit=max_results
)
hospitals = get_hospitals_nearby(
current_geocode,
results_limit=3
)
print('Places:')
for p in places:
print(p)
print('\nHospitals:')
for h in hospitals:
print(h)
```
<br>
### Create and monitor a job to test the deployed model
Create a payload containing places data received from HERE
```
import pandas as pd
places_df = pd.DataFrame.from_records([p.to_dict() for p in (places + hospitals)])
places_df.head()
route_summaries = get_route_summaries(current_geocode, places, hospitals)
routes_df = pd.DataFrame.from_records(route_summaries)
routes_df.drop_duplicates(keep='last', inplace=True)
routes_df.head()
solve_payload = {
client.deployments.DecisionOptimizationMetaNames.INPUT_DATA: [
{ 'id': 'places.csv', 'values' : places_df },
{ 'id': 'routes.csv', 'values' : routes_df }
],
client.deployments.DecisionOptimizationMetaNames.OUTPUT_DATA: [
{ 'id': '.*\.csv' },
{ 'id': '.*\.txt' }
]
}
```
<br>
Submit a new job with the payload and deployment.
Set the UID of the deployed model.
```
# deployment_uid = '...'
job_details = client.deployments.create_job(deployment_uid, solve_payload)
job_uid = client.deployments.get_job_uid(job_details)
print('Job UID: {}'.format(job_uid))
```
Display job status until it is completed.
The first job of a new deployment might take some time as a compute node must be started.
```
from time import sleep
while job_details['entity']['decision_optimization']['status']['state'] not in ['completed', 'failed', 'canceled']:
print(job_details['entity']['decision_optimization']['status']['state'] + '...')
sleep(3)
job_details=client.deployments.get_job_details(job_uid)
print(job_details['entity']['decision_optimization']['status']['state'])
# job_details
job_details['entity']['decision_optimization']['status']
```
<br>
### Extract and display solution
Display the output solution.
```
import base64
output_data = job_details['entity']['decision_optimization']['output_data']
solution = None
stats = None
for i, d in enumerate(output_data):
if d['id'] == 'solution.csv':
solution = pd.DataFrame(output_data[i]['values'],
columns = job_details['entity']['decision_optimization']['output_data'][0]['fields'])
else:
stats = base64.b64decode(output_data[i]['values'][0][0]).decode('utf-8')
print(stats)
solution.head()
```
<br>
Check out the online documentation at <a href="https://dataplatform.cloud.ibm.com/docs" target="_blank" rel="noopener noreferrer">https://dataplatform.cloud.ibm.com/docs</a> for more samples, tutorials and documentation.
<br>
## Helper functions
See `watson-machine-learning-client(V4)` Python library documentation for more info on the API:
https://wml-api-pyclient-dev-v4.mybluemix.net/
```
## List models
def list_models(wml_client):
wml_client.repository.list_models()
## List deployments
def list_deployments(wml_client):
wml_client.deployments.list()
## Delete a model
def delete_model(wml_client, model_uid):
wml_client.repository.delete(model_uid)
## Delete a deployment
def delete_deployment(wml_client, deployment_uid):
wml_client.deployments.delete(deployment_uid)
## Get details of all models
def details_all_models(wml_client):
return wml_client.repository.get_model_details()['resources']
## Get details of all deployments
def details_all_deployments(wml_client):
return wml_client.deployments.get_details()['resources']
# Find model using model name
def get_models_by_name(wml_client, model_name):
all_models = wml_client.repository.get_model_details()['resources']
models = [m for m in all_models if m['entity']['name'] == model_name]
return models
# Find deployment using deployment name
def get_deployments_by_name(wml_client, deployment_name):
all_deployments = wml_client.deployments.get_details()['resources']
deployments = [d for d in all_deployments if d['entity']['name'] == deployment_name][0]
return deployments
delete_deployment(client, deployment_uid)
delete_model(client, model_uid)
list_deployments(client)
list_models(client)
```
| github_jupyter |
```
%matplotlib inline
import os
import time
import pickle
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy import ndimage
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from IPython.display import clear_output
from datetime import datetime
from lib.utils import SamplePool, make_seed, make_circle_masks, get_rand_avail
from lib.utils import get_sobel, softmax
from lib.NCCAModel2 import NCCAModel2
with open('anchor_loc.pickle', 'rb') as handle:
anchor_loc = pickle.load(handle)
root = "_maps/"
full_size = (100,100)
map_size = (80,80)
color_map = [(0.5,0.5,0.5),
(0.5,1.0,0.5),
(1.0,1.0,0.5),
(1.0,0.7,0.2),
(1.0,0.5,0.5),
(1.0,0.5,1.0)]
################################################################
d_trains = []
d_tests = []
alive_maps = []
for d_i, obj_name in enumerate(list(anchor_loc.keys())[:10]):
filenames = []
common_index = {}
for filename in os.listdir(root):
if filename[:len(obj_name)]==obj_name:
filenames.append(root+filename)
for filename in filenames:
with open(filename, 'rb') as handle:
map_dict = pickle.load(handle)
for index in map_dict:
try:
tmp = int(map_dict[index]['status'])
if index in common_index:
common_index[index]+= 1
else:
common_index[index] = 1
except (TypeError, KeyError):
continue
common_index = [x for x in common_index.keys() if common_index[x]==len(filenames)]
d_train = np.zeros([64, full_size[0], full_size[1], 4])
d_test = np.zeros([len(filenames)-d_train.shape[0], full_size[0], full_size[1], d_train.shape[-1]])
for i,filename in enumerate(filenames[:d_train.shape[0]]):
with open(filename, 'rb') as handle:
map_dict = pickle.load(handle)
for index in common_index:
try:
status = min(int(map_dict[index]['status'])-1, 3)
d_train[i, index[0], index[1]] = np.zeros(d_train.shape[-1])
d_train[i, index[0], index[1], status] = 1
except (TypeError, KeyError):
continue
for i,filename in enumerate(filenames[d_train.shape[0]:]):
with open(filename, 'rb') as handle:
map_dict = pickle.load(handle)
for index in common_index:
try:
status = min(int(map_dict[index]['status'])-1, 3)
d_test[i, index[0], index[1]] = np.zeros(d_test.shape[-1])
d_test[i, index[0], index[1], status] = 1
except (TypeError, KeyError):
continue
alive_map = np.expand_dims(np.expand_dims(np.sum(d_train[0, ...], -1)>0.001, 0), -1)
cut_off = ((full_size[0]-map_size[0])//2, (full_size[1]-map_size[1])//2)
d_train = d_train[:, cut_off[0]:(cut_off[0]+map_size[0]),
cut_off[1]:(cut_off[1]+map_size[1]), :]
d_test = d_test[:, cut_off[0]:(cut_off[0]+map_size[0]),
cut_off[1]:(cut_off[1]+map_size[1]), :]
alive_map = alive_map[:, cut_off[0]:(cut_off[0]+map_size[0]),
cut_off[1]:(cut_off[1]+map_size[1]), :]
print(d_train.shape, d_test.shape, alive_map.shape)
d_trains.append(d_train)
d_tests.append(d_test)
alive_maps.append(alive_map)
DEVICE = torch.device("cuda:0")
model_path = "models/ncca_softmax_multi_traffic.pth"
CHANNEL_N = 16
ALPHA_CHANNEL = 4
lr = 8e-4
lr_gamma = 0.99997
betas = (0.8, 0.9)
n_epoch = 60000
BATCH_SIZE = 8
N_STEPS = 128
POOL_SIZE = 16
CELL_FIRE_RATE = 0.5
CALIBRATION = 1.0
eps = 1e-3
USE_PATTERN_POOL = 1
DAMAGE_N = 4
TRANS_N = 2
valid_masks = []
for alive_map in alive_maps:
valid_masks.append(alive_map.astype(bool))
valid_masks = np.concatenate(valid_masks, 0)
pools_list = []
for d_i, d_train in enumerate(d_trains):
pools = []
for _ in range(d_train.shape[0]):
init_coord = get_rand_avail(valid_masks[d_i:(d_i+1)])
seed = make_seed(map_size, CHANNEL_N, np.arange(CHANNEL_N-ALPHA_CHANNEL)+ALPHA_CHANNEL, init_coord)
pools.append(SamplePool(x=np.repeat(seed[None, ...], POOL_SIZE, 0)))
pools_list.append(pools)
my_model = NCCAModel2(CHANNEL_N, ALPHA_CHANNEL, CELL_FIRE_RATE, DEVICE).to(DEVICE)
# my_model.load_state_dict(torch.load(model_path))
optimizer = optim.Adam(my_model.parameters(), lr=lr, betas=betas)
scheduler = optim.lr_scheduler.ExponentialLR(optimizer, lr_gamma)
# torch.autograd.set_detect_anomaly(True)
loss_log = []
def plot_loss(loss_log):
plt.figure(figsize=(10, 4))
plt.title('Loss history (log10)')
plt.plot(np.log10(loss_log), '.', alpha=0.1)
plt.show()
return
def train(x, target, valid_mask_t, calibration_map, steps, optimizer, scheduler):
for _ in range(steps):
x = my_model(x, valid_mask_t, 1)
h = torch.softmax(x[..., :ALPHA_CHANNEL], -1)
t = target[..., :ALPHA_CHANNEL]
_delta = t*(h-1)
delta = _delta * calibration_map * CALIBRATION
y1 = x[..., :ALPHA_CHANNEL]-delta
alpha_h = x[..., ALPHA_CHANNEL:(ALPHA_CHANNEL+1)]
y2 = alpha_h - 2 * (alpha_h-valid_mask_t) * calibration_map * CALIBRATION
x = torch.cat((y1,y2,x[..., (ALPHA_CHANNEL+1):]), -1)
loss_kldiv, loss_alpha = loss_f(x, target)
loss_kldiv, loss_alpha = torch.mean(loss_kldiv), torch.mean(loss_alpha)
loss = loss_kldiv+loss_alpha
print(loss_kldiv.item(), loss_alpha.item())
optimizer.zero_grad()
loss.backward()
optimizer.step()
scheduler.step()
return x, loss
def loss_f(x, target, epsilon=1e-8):
h = torch.softmax(x[..., :ALPHA_CHANNEL], -1)
t = target[..., :ALPHA_CHANNEL]
loss_kldiv = torch.mean(torch.sum(h*torch.log(h/t+epsilon), -1), [-1,-2])
loss_alpha = torch.mean(torch.pow(x[..., ALPHA_CHANNEL]-target[..., ALPHA_CHANNEL], 2), [-1,-2])
return loss_kldiv, loss_alpha
starting_time = time.time()
for i_epoch in range(n_epoch+1):
targets = []
target_is = []
for _ in range(BATCH_SIZE):
d_i = np.random.randint(len(d_trains))
target_i = np.random.randint(d_trains[d_i].shape[0])
target_is.append((d_i, target_i))
target = np.concatenate((d_trains[d_i][target_i:target_i+1], valid_masks[d_i:(d_i+1)]), -1)
targets.append(target)
targets = np.concatenate(targets, 0).astype(np.float32)
targets[..., :-1] += eps
targets[..., :-1] /= np.sum(targets[..., :-1], axis=-1, keepdims=True)
_target = torch.from_numpy(targets).to(DEVICE)
calibration_map = make_circle_masks(_target.size(0), map_size[0], map_size[1], rmin=0.5, rmax=0.5)[..., None]
calibration_map = torch.from_numpy(calibration_map.astype(np.float32)).to(DEVICE)
if USE_PATTERN_POOL:
batches = []
batch_x = []
train_x = []
for target_index, (d_i, target_i) in enumerate(target_is):
batch = pools_list[d_i][target_i].sample(1)
batches.append(batch)
batch_x.append(batch.x)
if target_index<(len(target_is)-TRANS_N):
train_x.append(batch.x)
else:
new_target_i = np.random.randint(d_trains[d_i].shape[0])
batch = pools_list[d_i][new_target_i].sample(1)
train_x.append(batch.x)
x0 = np.concatenate(train_x, 0)
init_coord = get_rand_avail(valid_masks[:1])
seed = make_seed(map_size, CHANNEL_N, np.arange(CHANNEL_N-ALPHA_CHANNEL)+ALPHA_CHANNEL, init_coord)
x0[:1] = seed
if DAMAGE_N:
damage = 1.0-make_circle_masks(DAMAGE_N, map_size[0], map_size[1])[..., None]
x0[1:1+DAMAGE_N] *= damage
else:
x0 = np.repeat(seed[None, ...], BATCH_SIZE, 0)
x0 = torch.from_numpy(x0.astype(np.float32)).to(DEVICE)
valid_mask_t = valid_masks[[tmp[0] for tmp in target_is]]
valid_mask_t = torch.from_numpy(valid_mask_t.astype(np.float32)).to(DEVICE)
x, loss = train(x0, _target, valid_mask_t, calibration_map, N_STEPS, optimizer, scheduler)
if USE_PATTERN_POOL:
for batch_i, batch in enumerate(batches):
batch.x[:] = x.detach().cpu().numpy()[batch_i:(batch_i+1)]
batch.commit()
loss_log.append(loss.item())
if (i_epoch)%50 == 0:
clear_output()
x0 = x0.detach().cpu().numpy()
x0 = softmax(x0, -1)
hyp = x.detach().cpu().numpy()
hyp = softmax(hyp, -1)
cali_map_numpy = calibration_map.detach().cpu().numpy()
for i in range(targets.shape[0]):
plt.figure(figsize=(18,4))
for j in range(4):
plt.subplot(1,15,j+1)
rotated_img = ndimage.rotate(targets[i, ..., j], 90)
plt.imshow(rotated_img, cmap=plt.cm.gray, vmin=0, vmax=1)
plt.axis('off')
# all white
plt.subplot(1,15,5)
plt.imshow(np.ones(map_size), cmap='binary', vmin=1, vmax=1)
plt.axis('off')
for j in range(4):
plt.subplot(1,15,j+6)
rotated_img = ndimage.rotate(x0[i, ..., j], 90)
plt.imshow(rotated_img, cmap=plt.cm.gray, vmin=0, vmax=1)
plt.axis('off')
# all white
plt.subplot(1,15,10)
plt.imshow(np.ones(map_size), cmap='binary', vmin=1, vmax=1)
plt.axis('off')
# calibration_map
plt.subplot(1,15,11)
rotated_img = ndimage.rotate(cali_map_numpy[i, ..., 0], 90)
plt.imshow(rotated_img, cmap=plt.cm.gray, vmin=0, vmax=1)
plt.axis('off')
for j in range(4):
plt.subplot(1,15,j+12)
rotated_img = ndimage.rotate(hyp[i, ..., j], 90)
plt.imshow(rotated_img, cmap=plt.cm.gray, vmin=0, vmax=1)
plt.axis('off')
plt.show()
plot_loss(loss_log)
total_time_cost = np.round((time.time()-starting_time)/60, 4)
ave_time_cost = np.round((time.time()-starting_time)/60/(i_epoch+1), 4)
print(i_epoch, "loss =", loss.item(), "ave_log_loss", np.log(np.mean(loss_log[-100:]))/np.log(10))
print("Toal Time Cost:", total_time_cost, "min")
print("Ave Time Cost:", ave_time_cost, "min/epoch")
torch.save(my_model.state_dict(), model_path)
np.save("loss_logs/loss_log_train_2_hidden_16_pool", loss_log)
```
| github_jupyter |
```
import keras
keras.__version__
```
# Neural style transfer
This notebook contains the code samples found in Chapter 8, Section 3 of [Deep Learning with Python](https://www.manning.com/books/deep-learning-with-python?a_aid=keras&a_bid=76564dff). Note that the original text features far more content, in particular further explanations and figures: in this notebook, you will only find source code and related comments.
----
Besides Deep Dream, another major development in deep learning-driven image modification that happened in the summer of 2015 is neural
style transfer, introduced by Leon Gatys et al. The neural style transfer algorithm has undergone many refinements and spawned many
variations since its original introduction, including a viral smartphone app, called Prisma. For simplicity, this section focuses on the
formulation described in the original paper.
Neural style transfer consists in applying the "style" of a reference image to a target image, while conserving the "content" of the target
image:

What is meant by "style" is essentially textures, colors, and visual patterns in the image, at various spatial scales, while the "content"
is the higher-level macrostructure of the image. For instance, blue-and-yellow circular brush strokes are considered to be the "style" in
the above example using Starry Night by Van Gogh, while the buildings in the Tuebingen photograph are considered to be the "content".
The idea of style transfer, tightly related to that of texture generation, has had a long history in the image processing community prior
to the development of neural style transfer in 2015. However, as it turned out, the deep learning-based implementations of style transfer
offered results unparalleled by what could be previously achieved with classical computer vision techniques, and triggered an amazing
renaissance in creative applications of computer vision.
The key notion behind implementing style transfer is same idea that is central to all deep learning algorithms: we define a loss function
to specify what we want to achieve, and we minimize this loss. We know what we want to achieve: conserve the "content" of the original image,
while adopting the "style" of the reference image. If we were able to mathematically define content and style, then an appropriate loss
function to minimize would be the following:
```
loss = distance(style(reference_image) - style(generated_image)) +
distance(content(original_image) - content(generated_image))
```
Where `distance` is a norm function such as the L2 norm, `content` is a function that takes an image and computes a representation of its
"content", and `style` is a function that takes an image and computes a representation of its "style".
Minimizing this loss would cause `style(generated_image)` to be close to `style(reference_image)`, while `content(generated_image)` would
be close to `content(generated_image)`, thus achieving style transfer as we defined it.
A fundamental observation made by Gatys et al is that deep convolutional neural networks offer precisely a way to mathematically defined
the `style` and `content` functions. Let's see how.
## The content loss
As you already know, activations from earlier layers in a network contain _local_ information about the image, while activations from
higher layers contain increasingly _global_ and _abstract_ information. Formulated in a different way, the activations of the different
layers of a convnet provide a decomposition of the contents of an image over different spatial scales. Therefore we expect the "content" of
an image, which is more global and more abstract, to be captured by the representations of a top layer of a convnet.
A good candidate for a content loss would thus be to consider a pre-trained convnet, and define as our loss the L2 norm between the
activations of a top layer computed over the target image and the activations of the same layer computed over the generated image. This
would guarantee that, as seen from the top layer of the convnet, the generated image will "look similar" to the original target image.
Assuming that what the top layers of a convnet see is really the "content" of their input images, then this does work as a way to preserve
image content.
## The style loss
While the content loss only leverages a single higher-up layer, the style loss as defined in the Gatys et al. paper leverages multiple
layers of a convnet: we aim at capturing the appearance of the style reference image at all spatial scales extracted by the convnet, not
just any single scale.
For the style loss, the Gatys et al. paper leverages the "Gram matrix" of a layer's activations, i.e. the inner product between the feature maps
of a given layer. This inner product can be understood as representing a map of the correlations between the features of a layer. These
feature correlations capture the statistics of the patterns of a particular spatial scale, which empirically corresponds to the appearance
of the textures found at this scale.
Hence the style loss aims at preserving similar internal correlations within the activations of different layers, across the style
reference image and the generated image. In turn, this guarantees that the textures found at different spatial scales will look similar
across the style reference image and the generated image.
## In short
In short, we can use a pre-trained convnet to define a loss that will:
* Preserve content by maintaining similar high-level layer activations between the target content image and the generated image. The
convnet should "see" both the target image and the generated image as "containing the same things".
* Preserve style by maintaining similar _correlations_ within activations for both low-level layers and high-level layers. Indeed, feature
correlations capture _textures_: the generated and the style reference image should share the same textures at different spatial scales.
Now let's take a look at a Keras implementation of the original 2015 neural style transfer algorithm. As you will see, it shares a lot of
similarities with the Deep Dream implementation we developed in the previous section.
## Neural style transfer in Keras
Neural style transfer can be implemented using any pre-trained convnet. Here we will use the VGG19 network, used by Gatys et al in their paper.
VGG19 is a simple variant of the VGG16 network we introduced in Chapter 5, with three more convolutional layers.
This is our general process:
* Set up a network that will compute VGG19 layer activations for the style reference image, the target image, and the generated image at
the same time.
* Use the layer activations computed over these three images to define the loss function described above, which we will minimize in order
to achieve style transfer.
* Set up a gradient descent process to minimize this loss function.
Let's start by defining the paths to the two images we consider: the style reference image and the target image. To make sure that all
images processed share similar sizes (widely different sizes would make style transfer more difficult), we will later resize them all to a
shared height of 400px.
```
from keras.preprocessing.image import load_img, img_to_array
path = 'C:/Users/gaborstefanics/Pictures/Saved Pictures/'
# This is the path to the image you want to transform.
# target_image_path = '/home/ubuntu/data/portrait.png'
target_image_path = path + 'IMG_3077_2.jpg'
# This is the path to the style image.
style_reference_image_path = path + 'picasso02.jpg'
# Dimensions of the generated picture.
width, height = load_img(target_image_path).size
img_height = 400
img_width = int(width * img_height / height)
```
We will need some auxiliary functions for loading, pre-processing and post-processing the images that will go in and out of the VGG19
convnet:
```
import numpy as np
from keras.applications import vgg19
def preprocess_image(image_path):
img = load_img(image_path, target_size=(img_height, img_width))
img = img_to_array(img)
img = np.expand_dims(img, axis=0)
img = vgg19.preprocess_input(img)
return img
def deprocess_image(x):
# Remove zero-center by mean pixel
x[:, :, 0] += 103.939
x[:, :, 1] += 116.779
x[:, :, 2] += 123.68
# 'BGR'->'RGB'
x = x[:, :, ::-1]
x = np.clip(x, 0, 255).astype('uint8')
return x
```
Let's set up the VGG19 network. It takes as input a batch of three images: the style reference image, the target image, and a placeholder
that will contain the generated image. A placeholder is simply a symbolic tensor, the values of which are provided externally via Numpy
arrays. The style reference and target image are static, and thus defined using `K.constant`, while the values contained in the placeholder
of the generated image will change over time.
```
from keras import backend as K
target_image = K.constant(preprocess_image(target_image_path))
style_reference_image = K.constant(preprocess_image(style_reference_image_path))
# This placeholder will contain our generated image
combination_image = K.placeholder((1, img_height, img_width, 3))
# We combine the 3 images into a single batch
input_tensor = K.concatenate([target_image,
style_reference_image,
combination_image], axis=0)
# We build the VGG19 network with our batch of 3 images as input.
# The model will be loaded with pre-trained ImageNet weights.
model = vgg19.VGG19(input_tensor=input_tensor,
weights='imagenet',
include_top=False)
print('Model loaded.')
```
Let's define the content loss, meant to make sure that the top layer of the VGG19 convnet will have a similar view of the target image and
the generated image:
```
def content_loss(base, combination):
return K.sum(K.square(combination - base))
```
Now, here's the style loss. It leverages an auxiliary function to compute the Gram matrix of an input matrix, i.e. a map of the correlations
found in the original feature matrix.
```
def gram_matrix(x):
features = K.batch_flatten(K.permute_dimensions(x, (2, 0, 1)))
gram = K.dot(features, K.transpose(features))
return gram
def style_loss(style, combination):
S = gram_matrix(style)
C = gram_matrix(combination)
channels = 3
size = img_height * img_width
return K.sum(K.square(S - C)) / (4. * (channels ** 2) * (size ** 2))
```
To these two loss components, we add a third one, the "total variation loss". It is meant to encourage spatial continuity in the generated
image, thus avoiding overly pixelated results. You could interpret it as a regularization loss.
```
def total_variation_loss(x):
a = K.square(
x[:, :img_height - 1, :img_width - 1, :] - x[:, 1:, :img_width - 1, :])
b = K.square(
x[:, :img_height - 1, :img_width - 1, :] - x[:, :img_height - 1, 1:, :])
return K.sum(K.pow(a + b, 1.25))
```
The loss that we minimize is a weighted average of these three losses. To compute the content loss, we only leverage one top layer, the
`block5_conv2` layer, while for the style loss we use a list of layers than spans both low-level and high-level layers. We add the total
variation loss at the end.
Depending on the style reference image and content image you are using, you will likely want to tune the `content_weight` coefficient, the
contribution of the content loss to the total loss. A higher `content_weight` means that the target content will be more recognizable in
the generated image.
```
# Dict mapping layer names to activation tensors
outputs_dict = dict([(layer.name, layer.output) for layer in model.layers])
# Name of layer used for content loss
content_layer = 'block5_conv2'
# Name of layers used for style loss
style_layers = ['block1_conv1',
'block2_conv1',
'block3_conv1',
'block4_conv1',
'block5_conv1']
# Weights in the weighted average of the loss components
total_variation_weight = 1e-4
style_weight = 1.
content_weight = 0.025
# Define the loss by adding all components to a `loss` variable
loss = K.variable(0.)
layer_features = outputs_dict[content_layer]
target_image_features = layer_features[0, :, :, :]
combination_features = layer_features[2, :, :, :]
loss += content_weight * content_loss(target_image_features,
combination_features)
for layer_name in style_layers:
layer_features = outputs_dict[layer_name]
style_reference_features = layer_features[1, :, :, :]
combination_features = layer_features[2, :, :, :]
sl = style_loss(style_reference_features, combination_features)
loss += (style_weight / len(style_layers)) * sl
loss += total_variation_weight * total_variation_loss(combination_image)
```
Finally, we set up the gradient descent process. In the original Gatys et al. paper, optimization is performed using the L-BFGS algorithm,
so that is also what we will use here. This is a key difference from the Deep Dream example in the previous section. The L-BFGS algorithms
comes packaged with SciPy. However, there are two slight limitations with the SciPy implementation:
* It requires to be passed the value of the loss function and the value of the gradients as two separate functions.
* It can only be applied to flat vectors, whereas we have a 3D image array.
It would be very inefficient for us to compute the value of the loss function and the value of gradients independently, since it would lead
to a lot of redundant computation between the two. We would be almost twice slower than we could be by computing them jointly. To by-pass
this, we set up a Python class named `Evaluator` that will compute both loss value and gradients value at once, will return the loss value
when called the first time, and will cache the gradients for the next call.
```
# Get the gradients of the generated image wrt the loss
grads = K.gradients(loss, combination_image)[0]
# Function to fetch the values of the current loss and the current gradients
fetch_loss_and_grads = K.function([combination_image], [loss, grads])
class Evaluator(object):
def __init__(self):
self.loss_value = None
self.grads_values = None
def loss(self, x):
assert self.loss_value is None
x = x.reshape((1, img_height, img_width, 3))
outs = fetch_loss_and_grads([x])
loss_value = outs[0]
grad_values = outs[1].flatten().astype('float64')
self.loss_value = loss_value
self.grad_values = grad_values
return self.loss_value
def grads(self, x):
assert self.loss_value is not None
grad_values = np.copy(self.grad_values)
self.loss_value = None
self.grad_values = None
return grad_values
evaluator = Evaluator()
```
Finally, we can run the gradient ascent process using SciPy's L-BFGS algorithm, saving the current generated image at each iteration of the
algorithm (here, a single iteration represents 20 steps of gradient ascent):
```
from scipy.optimize import fmin_l_bfgs_b
from scipy.misc import imsave
import time
result_prefix = 'style_transfer_result'
iterations = 20
# Run scipy-based optimization (L-BFGS) over the pixels of the generated image
# so as to minimize the neural style loss.
# This is our initial state: the target image.
# Note that `scipy.optimize.fmin_l_bfgs_b` can only process flat vectors.
x = preprocess_image(target_image_path)
x = x.flatten()
for i in range(iterations):
print('Start of iteration', i)
start_time = time.time()
x, min_val, info = fmin_l_bfgs_b(evaluator.loss, x,
fprime=evaluator.grads, maxfun=20)
print('Current loss value:', min_val)
# Save current generated image
img = x.copy().reshape((img_height, img_width, 3))
img = deprocess_image(img)
fname = result_prefix + '_at_iteration_%d.png' % i
imsave(fname, img)
end_time = time.time()
print('Image saved as', fname)
print('Iteration %d completed in %ds' % (i, end_time - start_time))
```
Here's what we get:
```
from matplotlib import pyplot as plt
# Content image
plt.imshow(load_img(target_image_path, target_size=(img_height, img_width)))
plt.figure()
# Style image
plt.imshow(load_img(style_reference_image_path, target_size=(img_height, img_width)))
plt.figure()
# Generate image
plt.imshow(img)
plt.show()
```
Keep in mind that what this technique achieves is merely a form of image re-texturing, or texture transfer. It will work best with style
reference images that are strongly textured and highly self-similar, and with content targets that don't require high levels of details in
order to be recognizable. It would typically not be able to achieve fairly abstract feats such as "transferring the style of one portrait to
another". The algorithm is closer to classical signal processing than to AI, so don't expect it to work like magic!
Additionally, do note that running this style transfer algorithm is quite slow. However, the transformation operated by our setup is simple
enough that it can be learned by a small, fast feedforward convnet as well -- as long as you have appropriate training data available. Fast
style transfer can thus be achieved by first spending a lot of compute cycles to generate input-output training examples for a fixed style
reference image, using the above method, and then training a simple convnet to learn this style-specific transformation. Once that is done,
stylizing a given image is instantaneous: it's a just a forward pass of this small convnet.
## Take aways
* Style transfer consists in creating a new image that preserves the "contents" of a target image while also capturing the "style" of a
reference image.
* "Content" can be captured by the high-level activations of a convnet.
* "Style" can be captured by the internal correlations of the activations of different layers of a convnet.
* Hence deep learning allows style transfer to be formulated as an optimization process using a loss defined with a pre-trained convnet.
* Starting from this basic idea, many variants and refinements are possible!
| github_jupyter |
# Dummy Variables Exercise
In this exercise, you'll create dummy variables from the projects data set. The idea is to transform categorical data like this:
| Project ID | Project Category |
|------------|------------------|
| 0 | Energy |
| 1 | Transportation |
| 2 | Health |
| 3 | Employment |
into new features that look like this:
| Project ID | Energy | Transportation | Health | Employment |
|------------|--------|----------------|--------|------------|
| 0 | 1 | 0 | 0 | 0 |
| 1 | 0 | 1 | 0 | 0 |
| 2 | 0 | 0 | 1 | 0 |
| 3 | 0 | 0 | 0 | 1 |
(Note if you were going to use this data with a model influenced by multicollinearity, you would want to eliminate one of the columns to avoid redundant information.)
The reasoning behind these transformations is that machine learning algorithms read in numbers not text. Text needs to be converted into numbers. You could assign a number to each category like 1, 2, 3, and 4. But a categorical variable has no inherent order.
Pandas makes it very easy to create dummy variables with the [get_dummies](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.get_dummies.html) method. In this exercise, you'll create dummy variables from the World Bank projects data; however, there's a caveat. The World Bank data is not particularly clean, so you'll need to explore and wrangle the data first.
You'll focus on the text values in the sector variables.
Run the code cells below to read in the World Bank projects data set and then to filter out the data for text variables.
```
import pandas as pd
import numpy as np
# read in the projects data set and do basic wrangling
projects = pd.read_csv('../data/projects_data.csv', dtype=str)
projects.drop('Unnamed: 56', axis=1, inplace=True)
projects['totalamt'] = pd.to_numeric(projects['totalamt'].str.replace(',', ''))
projects['countryname'] = projects['countryname'].str.split(';', expand=True)[0]
projects['boardapprovaldate'] = pd.to_datetime(projects['boardapprovaldate'])
# keep the project name, lending, sector and theme data
sector = projects.copy()
sector = sector[['project_name', 'lendinginstr', 'sector1', 'sector2', 'sector3', 'sector4', 'sector5', 'sector',
'mjsector1', 'mjsector2', 'mjsector3', 'mjsector4', 'mjsector5',
'mjsector', 'theme1', 'theme2', 'theme3', 'theme4', 'theme5', 'theme ',
'goal', 'financier', 'mjtheme1name', 'mjtheme2name', 'mjtheme3name',
'mjtheme4name', 'mjtheme5name']]
```
Run the code cell below. This cell shows the percentage of each variable that is null. Notice the mjsector1 through mjsector5 variables are all null. The mjtheme1name through mjtheme5name are also all null as well as the theme variable.
Because these variables contain so many null values, they're probably not very useful.
```
# output percentage of values that are missing
#100 * sector.isnull().sum() / sector.shape[0]
sector.isna().mean()
```
The sector1 variable looks promising; it doesn't contain any null values at all. In the next cell, store the unique sector1 values in a list and output the results. Use the sort_values() and unique() methods.
```
# TODO: Create a list of the unique values in sector1. Use the sort_values() and unique() pandas methods.
# And then convert those results into a Python list
uniquesectors1 = list(sector.sector1.sort_values().unique())
uniquesectors1
# run this code cell to see the number of unique values
print('Number of unique values in sector1:', len(uniquesectors1))
```
3060 different categories is quite a lot! Remember that with dummy variables, if you have n categorical values, you need n - 1 new variables! That means 3059 extra columns!
There are a few issues with this 'sector1' variable. First, there are values labeled '!$!0'. These should be substituted with NaN.
Furthermore, each sector1 value ends with a ten or eleven character string like '!$!49!$!EP'. Some sectors show up twice in the list like:
'Other Industry; Trade and Services!$!70!$!YZ',
'Other Industry; Trade and Services!$!63!$!YZ',
But it seems like those are actually the same sector. You'll need to remove everything past the exclamation point.
Many values in the sector1 variable start with the term '(Historic)'. Try removing that phrase as well.
Fix these issues in the code cell below.
```
# TODO: In the sector1 variable, replace the string '!$!0' with nan
# Put the results back into the sector1 variable
# HINT: you can use the pandas replace() method and numpy.nan
sector.sector1.replace('!$!0', np.nan, inplace=True)
# TODO: In the sector1 variable, remove the last 10 or 11 characters from the sector1 variable.
# HINT: There is more than one way to do this. For example,
# you can use the replace method with a regex expression '!.+'
# That regex expression looks for a string with an exclamation
# point followed by one or more characters
sector.sector1.replace('\!\$.+', '', regex=True, inplace=True)
# TODO: Remove the string '(Historic)' from the sector1 variable
# HINT: You can use the replace method
sector.sector1.replace('\(Historic\)', '', regex=True, inplace=True)
print('Number of unique sectors after cleaning:', len(list(sector['sector1'].unique())))
print('Percentage of null values after cleaning:', 100 * sector['sector1'].isnull().sum() / sector['sector1'].shape[0])
```
Now there are 156 unique categorical values. That's better than 3060. If you were going to use this data with a supervised learning machine model, you could try converting these 156 values to dummy variables. You'd still have to train and test a model to see if those are good features.
But can you do anything else with the sector1 variable?
The percentage of null values for 'sector1' is now 3.49%. That turns out to be the same number as the null values for the 'sector' column. You can see this if you scroll back up to where the code calculated the percentage of null values for each variable.
Perhaps the 'sector1' and 'sector' variable have the same information. If you look at the 'sector' variable, however, it also needs cleaning. The values look like this:
'Urban Transport;Urban Transport;Public Administration - Transportation'
It turns out the 'sector' variable combines information from the 'sector1' through 'sector5' variables and the 'mjsector' variable. Run the code cell below to look at the sector variable.
```
sector['sector']
```
What else can you do? If you look at all of the diferent sector1 categories, it might be useful to combine a few of them together. For example, there are various categories with the term "Energy" in them. And then there are other categories that seem related to energy but don't have the word energy in them like "Thermal" and "Hydro". Some categories have the term "Renewable Energy", so perhaps you could make a separate "Renewable Energy" category.
Similarly, there are categories with the term "Transportation" in them, and then there are related categories like "Highways".
In the next cell, find all sector1 values with the term 'Energy' in them. For each of these rows, put the string 'energy' in a new column called 'sector1_aggregates'. Do the same for "Transportation".
```
sector.sector1.unique()
import re
# Create the sector1_aggregates variable
sector.loc[:,'sector1_aggregates'] = sector['sector1']
# TODO: The code above created a new variable called sector1_aggregates.
# Currently, sector1_aggregates has all of the same values as sector1
# For this task, find all the rows in sector1_aggregates with the term 'Energy' in them,
# For all of these rows, replace whatever is the value is with the term 'Energy'.
# The idea is to simplify the category names by combining various categories together.
# Then, do the same for the term 'Transportation
# HINT: You can use the contains() methods. See the documentation for how to ignore case using the re library
# HINT: You might get an error saying "cannot index with vector containing NA / NaN values."
# Try converting NaN values to something else like False or a string
# Fill nan with False
#sector.sector1_aggregates.fillna(False, inplace=True)
# Replace str containing 'Energy' with 'Energy'
sector.sector1_aggregates.replace('.*Energy.*', 'Energy', inplace=True, regex=True)
# Replace str containing 'Transportation' with 'Transportation'
sector.sector1_aggregates.replace('.*Transportation.*', 'Transportation', inplace=True, regex=True)
print('Number of unique sectors after cleaning:', len(list(sector['sector1_aggregates'].unique())))
```
The number of unique sectors continues to go down. Keep in mind that how much to consolidate will depend on your machine learning model performance and your hardware's ability to handle the extra features in memory. If your hardware's memory can handle 3060 new features and your machine learning algorithm performs better, then go for it!
There are still 638 entries with NaN values. How could you fill these in? You might try to determine an appropriate category from the 'project_name' or 'lendinginstr' variables. If you make dummy variables including NaN values, then you could consider a feature with all zeros to represent NaN. Or you could delete these records from the data set. Pandas will ignore NaN values by default. That means, for a given row, all dummy variables will have a value of 0 if the sector1 value was NaN.
Don't forget about the bigger context! This data is being prepared for a machine learning algorithm. Whatever techniques you use to engineer new features, you'll need to use those when running your model on new data. So if your new data does not contain a sector1 value, you'll have to run whatever feature engineering processes you did on your training set.
In this final set, use the pandas pd.get_dummies() method to create dummy variables. Then use the concat() method to concatenate the dummy variables to a dataframe that contains the project totalamt variable and the project year from the boardapprovaldate.
```
%%time
# TODO: Create dummy variables from the sector1_aggregates data. Put the results into a dataframe called dummies
# Hint: Use the get_dummies method
dummies = pd.get_dummies(pd.get_dummies(sector.sector1_aggregates))
# TODO: Create a new dataframe called df by
# filtering the projects data for the totalamt and
# the year from boardapprovaldate
projects['year'] = projects.boardapprovaldate.dt.year
df = projects.loc[:, ['totalamt', 'year']]
# TODO: Concatenate the results of dummies and projects
# into a single data frame
df_final = pd.concat([df, dummies], axis=1)
df_final.head()
```
# Conclusion
Pandas makes it relatively easy to create dummy variables; however, oftentimes you'll need to clean the data first.
| github_jupyter |
# Modificaciones de https://github.com/MIT-LCP/wfdb-python
# Demo Scripts for the wfdb-python package
Run this script from the base directory of the git repository to access the included demo files
```
import wfdb
import numpy as np
import os
from IPython.display import display
import matplotlib.pyplot as plt
import pandas as pd
url = "charisdb/"
senial = wfdb.rdsamp('charis2',pbdir = url)
wfdb.plotrec(senial, title = 'prueba')
#diplay(senial.__dict__)
!rsync physionet.org::charisdb
!rsync -a physionet.org::mirror-setup mirror-setup
!rsync physionet.org::charisdb /home/pic/Escritorio/wfdb-python-master/carpeta/
# See the help documentation for the read functions
help(wfdb.rdsamp)
#help(wfdb.srdsamp)
#help(wfdb.rdann)
```
## Accedemos a la ruta
```
# Demo 1 - Read a wfdb record using the 'rdsamp' function into a wfdb.Record object.
# Plot the signals, and show the data.
#record = wfdb.rdsamp('sampledata/t107l')
#wfdb.plotrec(record, title='Record a105l from Physionet Challenge 2015')
#display(record.__dict__)
# Can also read the same files hosted on Physiobank https://physionet.org/physiobank/database/
# in the challenge/2015/training/ database subdirectory. Full url = urecord2 = wfdb.rdsamp('charis1', pbdir = 'charisdb/')
wfdb.plotrec(record2, title='Record CHARIS1 from Physionet CharisDB')
display(record2.__dict__)
len(record2.p_signals[:])
signales = np.ndarray.flatten(record2.p_signals)
tamanio = len(signales)/3
reFactorizacion = signales.reshape(tamanio,3)
senialABP = []
senialECG = []
senialICP = []
for reFactor in reFactorizacion:
senialABP.append(reFactor[0])
senialECG.append(reFactor[1])
senialICP.append(reFactor[2])
senialABP = np.asarray(senialABP)
senialECG = np.asarray(senialECG)
senialICP = np.asarray(senialICP)
senialABP[0:100]
plt.plot(senialICP[0:1200])
plt.show()
senialICP[0:1200]
#Guardar datos en CSV
df = pd.DataFrame(reFactorizacion,columns = ['ABP', 'ECG', 'ICP'])
df.to_csv('charis.csv')
# Demo 2 - Read certain channels and sections of the WFDB record using the simplified 'srdsamp' function
# which returns a numpy array and a dictionary. Show the data.
signals, fields=wfdb.srdsamp('sampledata/s0010_re', channels=[14, 0, 5, 10], sampfrom=100, sampto=15000)
display(signals)
display(fields)
# Can also read the same files hosted on Physiobank
signals2, fields2=wfdb.srdsamp('s0010_re', channels=[14, 0, 5, 10], sampfrom=100, sampto=15000, pbdir = 'ptbdb/patient001/')
# Demo 3 - Read a WFDB header file only (without the signals)
record = wfdb.rdheader('sampledata/drive02')
display(record.__dict__)
# Can also read the same file hosted on Physiobank
record2 = wfdb.rdheader('drive02', pbdir = 'drivedb')
# Demo 4 - Read part of a WFDB annotation file into a wfdb.Annotation object, and plot the samples
annotation = wfdb.rdann('sampledata/100', 'atr', sampfrom = 100000, sampto = 110000)
annotation.fs = 360
wfdb.plotann(annotation, timeunits = 'minutes')
# Can also read the same file hosted on PhysioBank
annotation2 = wfdb.rdann('100', 'atr', sampfrom = 100000, sampto = 110000, pbdir = 'mitdb')
# Demo 5 - Read a WFDB record and annotation. Plot all channels, and the annotation on top of channel 0.
record = wfdb.rdsamp('sampledata/100', sampto = 15000)
annotation = wfdb.rdann('sampledata/100', 'atr', sampto = 15000)
wfdb.plotrec(record, annotation = annotation, title='Record 100 from MIT-BIH Arrhythmia Database', timeunits = 'seconds')
```
### Multisegment waveform examples
The following sections load and plots waveforms from the MIMIC matched waveform database. These waveforms have been matched to clinical data in the MIMIC Clinical database. The input records are multi-segment (made up of multiple individual WFDB records) and relatively long.
Note that these kinds of records contain segments in which certain channels are missing. <strong>matplotlib</strong> automatically zooms in on sections without Nans in individual channels but the entire durations of the signals input into <strong>plotrec</strong> are actually plotted.
```
# Demo 6 - Read the multi-segment record and plot waveforms from the MIMIC matched waveform database.
import wfdb
from IPython.display import display
record=wfdb.rdsamp('sampledata/multisegment/s25047/s25047-2704-05-04-10-44')
wfdb.plotrec(record, title='Record s25047-2704-05-04-10-44')
display(record.__dict__)
# Can also read the same files hosted on PhysioBank (takes long to stream the many large files)
#record2 = wfdb.rdsamp('s25047-2704-05-04-10-44', pbdir = 'mimic2wdb/matched/s25047')
# Demo 7 - Read the multi-segment record and plot waveforms from the MIMIC matched waveform database.
# Notice that some channels have no valid values to plot
record = wfdb.rdsamp('sampledata/multisegment/s00001/s00001-2896-10-10-00-31', sampfrom = 3000000, sampto = 4000000)
wfdb.plotrec(record, title='Record s00001/s00001-2896-10-10-00-31')
display(record.__dict__)
# Can also read the same files hosted on PhysioBank
record2 = wfdb.rdsamp('s00001-2896-10-10-00-31', sampfrom = 3000000, sampto = 4000000, pbdir = 'mimic2wdb/matched/s00001')
```
### Multiple sample/frame examples
Although there can only be one base sampling frequency per record, a single wfdb record can store multiple channels with different sampling frequencies, as long as their sampling frequencies can all be expressed by an integer multiple of a base value. This is done by using the `sampsperframe` attribute in each channel, which indicates the number of samples of each channel present in each frame.
ie: To capture three signals with `fs = 120, 240, and 360 Hz` in a single record, they can be combined into a record with `fs = 120` and `sampsperframe = [1, 2, 3]`.
#### Reading Options
This package allows signals in records with multiple samples/frame to be read in two ways:
1. smoothed - An uniform mxn numpy is returned as the d_signals or p_signals field. Channels with multiple samples/frame have their values averaged within each frame. This is like the behaviour of the `rdsamp` function of the original WFDB c package. Note that `wfdb.plotrec` only works if the record object has the `p_signals` field.
2. expanded - A list of 1d numpy arrays is returned as the e_d_signals or e_p_signals field. All samples for each channel are returned in its respective numpy array. The arrays may have different lengths depending on their `sampsperframe` values.
Set the `smoothframes` *(default=True)* option in `rdsamp` to return the desired signal type.
```
# Demo 8 - Read a wfdb record in which one channel has multiple samples/frame. Return a smoothed uniform array.
record = wfdb.rdsamp('sampledata/test01_00s_frame')
wfdb.plotrec(record)
# Demo 9 - Read a wfdb record in which one channel has multiple samples/frame. Return a list of all the expanded samples.
record = wfdb.rdsamp('sampledata/test01_00s_frame', smoothframes = False)
display(record.e_p_signals)
# Show that different channels have different lengths. Channel 1 has 2 samples/frame, hence has 2x as many samples.
print([len(s) for s in record.e_p_signals])
# wfdb.plotrec doesn't work because the Record object is missing its p_signals field.
```
## Writing Records and Annotations
```
# Demo 10 - Read a WFDB record's digital samples and create a copy via the wrsamp() instance method
# of the Record object.
# Read a record as a Record object.
record = wfdb.rdsamp('sampledata/100', physical = False)
record.recordname = '100x'
# Call the instance method of the object
record.wrsamp()
# The new file can be read
recordx = wfdb.rdsamp('100x')
# Demo 11 - Write a WFDB record without using a Record object via the gateway wrsamp function.
# This is the basic way to write physical signals to a WFDB file.
# Read part of a record from Physiobank
sig, fields = wfdb.srdsamp('a103l', sampfrom = 50000, channels = [0,1], pbdir = 'challenge/2015/training')
# Call the gateway wrsamp function, manually inserting fields as function input parameters
wfdb.wrsamp('ecgrecord', fs = 250, units = ['mV', 'mV'], signames = ['I', 'II'], p_signals = sig, fmt = ['16', '16'])
# The new file can be read
recordecg = wfdb.rdsamp('ecgrecord')
# Demo 12 - Write a WFDB record with multiple samples/frame in a channel
# Read a record as a Record object.
record = wfdb.rdsamp('sampledata/test01_00s_frame', physical = False, smoothframes=False)
record.recordname = 'test01_00s_framex'
# Call the instance method of the object with expanded=True to write the record using the e_d_signals field
record.wrsamp(expanded=True)
# The new file can be read
recordx = wfdb.rdsamp('test01_00s_framex')
# Demo 13 - Read a WFDB annotation file and create a copy via the wrann() instance method
# of the Annotation object
# Read an annotation from Physiobank
annotation = wfdb.rdann('sampledata/100', 'atr')
annotation.annotator = 'cpy'
# Call the instance method of the object
annotation.wrann()
# The new file can be read
ann100copy = wfdb.rdann('100', 'cpy')
# Demo 14 - Write a WFDB annotation file without using an Annotator object via the gateway wrann function.
# Read an annotation as an Annotation object
annotation = wfdb.rdann('b001', 'atr', pbdir='cebsdb')
# Call the gateway wrann function, manually inserting fields as function input parameters
wfdb.wrann('b001', 'cpy', annotation.annsamp, annotation.anntype)
# The new file can be read
annbcopy = wfdb.rdann('b001', 'cpy')
# Demo 15 - View what the 'anntype' symbols mean in the standard WFDB library
wfdb.showanncodes()
```
## Downloading Content from Physiobank
- The downloads are made via http
- See the above demos for examples on streaming WFDB files stored in Physiobank without downloading them to local disk
- Physionet has rsync modules for downloading entire databases for users who have access to rsync.
```
# Demo 16 - List the Physiobank Databases
dbs = wfdb.getdblist()
display(dbs)
# Demo 17 - Download all the WFDB records and annotations from a small Physiobank Database
# Make a temporary download directory in your current working directory
cwd = os.getcwd()
dldir = os.path.join(cwd, 'tmp_dl_dir')
# Make sure to use a new directory
while os.path.exists(dldir):
dldir = dldir+'1'
# Download all the WFDB content
wfdb.dldatabase('ahadb', dlbasedir = dldir)
# Display the downloaded content in the folder
display(os.listdir(dldir))
# Demo 18 - Download specified files from a Physiobank database
# The files to download
filelist = ['STAFF-Studies-bibliography-2016.pdf', 'data/001a.hea', 'data/001a.dat', 'data/001b.hea', 'data/001b.dat']
# Make a temporary download directory in your current working directory
cwd = os.getcwd()
dldir = os.path.join(cwd, 'tmp_dl_dir')
# Make sure to use a new directory
while os.path.exists(dldir):
dldir = dldir+'1'
# Download the listed files
wfdb.dldatabasefiles('staffiii', dldir, filelist)
# Display the downloaded content in the folder
display(os.listdir(dldir))
display(os.listdir(os.path.join(dldir, 'data')))
```
| github_jupyter |
# Basics of Data Visualisation
## Context
This example is based on sample dataset for a telo predicting customer behavior to retain them. The key target of the analysis is the churn feature - customers that left the service with the last month.
Source: https://www.kaggle.com/blastchar/telco-customer-churn/
## Dataset
Each row represents a customer, each column contains customer’s attributes described oin the column Metadata.
- Customer Account Information
- `CustomerID`: Unique ID for the customer *(unique String)*?
- `Churn`: Customers who left within the last month *(Yes or No)*?
- `Tenure`: How long they’ve been a customer *(In months, Integer)*?
- `Contract`: What type of contract do they have *(Month-to-month, One year, Two year)*?
- `PaymentMethod`: What payment method is used by the customer *(Electronic check, Mailed check, Bank transfer, or Credit card)*?
- `PaperlessBilling`: Whether the customer has subscribed to paperless billing *(Yes or No)*?
- `MonthlyCharges`: What was the monthly charges for the customer (Amount, Float)?
- `TotalCharges`: What were the total charges for the customer (Amount, Float)?
- Customer Demographics
- `Gender`: What is the gender of the customer *(Male or Female)*?
- `SeniorCitizen`: Whether the customer is a Senior Citizen or not *(0 or 1)*?
- `Partner`: Whether the customer has a partner *(Yes or No)*?
- `Dependents`: Whether the customer has any dependents *(Yes or No)*?
- Customer Signed Up Service Status
- `PhoneService`: Signed up for Phone Service *(Yes or No)*?
- `MultipleLines`: Signed up for Multiple Lines *(Yes or No or No Phone Service)*?
- `InternetService`: Signed up for Internet Service *(DSL or Fiber optic or No)*?
- `OnlineSecurity`: Signed up for Online Security *(Yes or No or No internet service)*?
- `OnlineBackup`: Signed up for Online Backup *(Yes or No or No internet service)*?
- `DeviceProtection`: Signed up for Device Protection plan *(Yes or No or No internet service)*?
- `TechSupport`: Signed up for Tech Support *(Yes or No or No internet service)*?
- `StreamingTV`: Signed up for Streaming TV *(Yes or No or No internet service)*?
- `StreamingMovies`: Signed up for Streaming Movies *(Yes or No or No internet service)*?
```
import numpy as np
import pandas as pd
import altair as alt
alt.data_transformers.disable_max_rows()
alt.data_transformers.enable('json')
df = pd.read_csv("data/churn.csv")
df.shape
df.head()
df.dtypes
```
## 1D Continuous Vis
Let us look at the `Tenure` variables
### Tenure
#### Plotting all the data
- We can plot all the points directly
- This usually results in overplotting, so we add opacity to handle this
```
alt.Chart(df).mark_tick().encode(
alt.X("Tenure:Q"),
alt.Opacity(value=0.01)
)
alt.Chart(df).mark_circle().encode(
alt.X("Tenure:Q"),
alt.Opacity(value=0.01)
)
# Adding random jitter to the chart
alt.Chart(df).mark_circle().encode(
alt.X("Tenure:Q"),
alt.Y("jitter:Q"),
alt.Size(value=50),
alt.Opacity(value=0.05)
).transform_calculate(
jitter=alt.expr.random()
).properties(height=100)
```
#### Binning the Data
```
alt.Chart(df).mark_bar().encode(
alt.X("Tenure:Q", bin=True),
alt.Y("count()")
)
alt.Chart(df).mark_bar().encode(
alt.X("Tenure:Q", bin=alt.BinParams(maxbins=20)),
alt.Y("count()")
)
alt.Chart(df).mark_area().encode(
alt.X("Tenure:Q", bin=True),
alt.Y("count()")
)
```
#### Binning using top-level transform
```
alt.Chart(df).mark_bar().encode(
alt.X("Binned_Tenure:O"),
alt.Y("count()")
).transform_bin(
'Binned_Tenure', field='Tenure'
)
```
## 1D Categorical
### Churn
```
alt.Chart(df).mark_point().encode(
alt.Y("Churn:N"),
)
alt.Chart(df).mark_point().encode(
alt.Y("Churn:N"),
alt.X('count()')
)
alt.Chart(df).mark_bar().encode(
alt.Y("Churn:N"),
alt.X('count()')
)
```
## 2D Continuous + Categorical
```
alt.Chart(df).mark_bar().encode(
alt.Y("Churn:N"),
alt.X('Tenure:Q', bin=True),
alt.Color("count()")
)
alt.Axis?
```
| github_jupyter |
# Data Analysis
This is the main notebook performing all feature engineering, model selection, training, evaluation etc.
The different steps are:
- Step1 - import dependencies
- Step2 - load payloads into memory
- Step3A - Feature engineering custom features
- Step3B - Feature engineering bag-of-words
- Step3C - Feature space visualization
- Step4 - Model selection
- (Step4B - Load pre-trained classifiers)
- Step5 - Visualization
- Step6 - Website integration extract
# Step1
import dependencies
```
%matplotlib inline
import pandas as pd
import numpy as np
import pickle
import matplotlib.pyplot as plt
import seaborn
import string
from IPython.display import display
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.model_selection import train_test_split
from sklearn.model_selection import RandomizedSearchCV
from sklearn.model_selection import learning_curve
from sklearn.decomposition import TruncatedSVD
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import chi2
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
from sklearn.svm import SVC
from sklearn.gaussian_process import GaussianProcessClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.naive_bayes import MultinomialNB
from sklearn.ensemble import AdaBoostClassifier
from sklearn.linear_model import SGDClassifier
from sklearn.neighbors import NearestNeighbors
from sklearn.neighbors.nearest_centroid import NearestCentroid
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
from sklearn.tree import DecisionTreeClassifier
import sklearn.gaussian_process.kernels as kernels
from sklearn.cross_validation import ShuffleSplit
from sklearn.cross_validation import KFold
from sklearn.pipeline import Pipeline
from sklearn.metrics import confusion_matrix
from sklearn.metrics import roc_curve
from sklearn.metrics import roc_auc_score
from scipy.stats import expon
```
# Step2
load the payloads into memory
```
payloads = pd.read_csv("data/payloads.csv",index_col='index')
display(payloads.head(30))
```
# Step3A - feature engineering custom features
We will create our own feature space with features that might be important for this task, this includes:
- length of payload
- number of non-printable characters in payload
- number of punctuation characters in payload
- the minimum byte value of payload
- the maximum byte value of payload
- the mean byte value of payload
- the standard deviation of payload byte values
- number of distinct bytes in payload
- number of SQL keywords in payload
- number of javascript keywords in payload
```
def plot_feature_distribution(features):
print('Properties of feature: ' + features.name)
print(features.describe())
f, ax = plt.subplots(1, figsize=(10, 6))
ax.hist(features, bins=features.max()-features.min()+1, normed=1)
ax.set_xlabel('value')
ax.set_ylabel('fraction')
plt.show()
def create_feature_length(payloads):
'''
Feature describing the lengh of the input
'''
payloads['length'] = [len(str(row)) for row in payloads['payload']]
return payloads
payloads = create_feature_length(payloads)
display(payloads.head())
plot_feature_distribution(payloads['length'])
def create_feature_non_printable_characters(payloads):
'''
Feature
Number of non printable characthers within payload
'''
payloads['non-printable'] = [ len([1 for letter in str(row) if letter not in string.printable]) for row in payloads['payload']]
return payloads
create_feature_non_printable_characters(payloads)
display(payloads.head())
plot_feature_distribution(payloads['non-printable'])
def create_feature_punctuation_characters(payloads):
'''
Feature
Number of punctuation characthers within payload
'''
payloads['punctuation'] = [ len([1 for letter in str(row) if letter in string.punctuation]) for row in payloads['payload']]
return payloads
create_feature_punctuation_characters(payloads)
display(payloads.head())
plot_feature_distribution(payloads['punctuation'])
def create_feature_min_byte_value(payloads):
'''
Feature
Minimum byte value in payload
'''
payloads['min-byte'] = [ min(bytearray(str(row), 'utf8')) for row in payloads['payload']]
return payloads
create_feature_min_byte_value(payloads)
display(payloads.head())
plot_feature_distribution(payloads['min-byte'])
def create_feature_max_byte_value(payloads):
'''
Feature
Maximum byte value in payload
'''
payloads['max-byte'] = [ max(bytearray(str(row), 'utf8')) for row in payloads['payload']]
return payloads
create_feature_max_byte_value(payloads)
display(payloads.head())
plot_feature_distribution(payloads['max-byte'])
def create_feature_mean_byte_value(payloads):
'''
Feature
Maximum byte value in payload
'''
payloads['mean-byte'] = [ np.mean(bytearray(str(row), 'utf8')) for row in payloads['payload']]
return payloads
create_feature_mean_byte_value(payloads)
display(payloads.head())
plot_feature_distribution(payloads['mean-byte'].astype(int))
def create_feature_std_byte_value(payloads):
'''
Feature
Standard deviation byte value in payload
'''
payloads['std-byte'] = [ np.std(bytearray(str(row), 'utf8')) for row in payloads['payload']]
return payloads
create_feature_std_byte_value(payloads)
display(payloads.head())
plot_feature_distribution(payloads['std-byte'].astype(int))
def create_feature_distinct_bytes(payloads):
'''
Feature
Number of distinct bytes in payload
'''
payloads['distinct-bytes'] = [ len(list(set(bytearray(str(row), 'utf8')))) for row in payloads['payload']]
return payloads
create_feature_distinct_bytes(payloads)
display(payloads.head())
plot_feature_distribution(payloads['distinct-bytes'])
sql_keywords = pd.read_csv('data/SQLKeywords.txt', index_col=False)
def create_feature_sql_keywords(payloads):
'''
Feature
Number of SQL keywords within payload
'''
payloads['sql-keywords'] = [ len([1 for keyword in sql_keywords['Keyword'] if str(keyword).lower() in str(row).lower()]) for row in payloads['payload']]
return payloads
create_feature_sql_keywords(payloads)
display(type(sql_keywords))
display(payloads.head())
plot_feature_distribution(payloads['sql-keywords'])
js_keywords = pd.read_csv('data/JavascriptKeywords.txt', index_col=False)
def create_feature_javascript_keywords(payloads):
'''
Feature
Number of Javascript keywords within payload
'''
payloads['js-keywords'] = [len([1 for keyword in js_keywords['Keyword'] if str(keyword).lower() in str(row).lower()]) for row in payloads['payload']]
return payloads
create_feature_javascript_keywords(payloads)
display(payloads.head())
plot_feature_distribution(payloads['js-keywords'])
```
define a function that makes a feature vector from the payload using the custom features
```
def create_features(payloads):
features = create_feature_length(payloads)
features = create_feature_non_printable_characters(features)
features = create_feature_punctuation_characters(features)
features = create_feature_max_byte_value(features)
features = create_feature_min_byte_value(features)
features = create_feature_mean_byte_value(features)
features = create_feature_std_byte_value(features)
features = create_feature_distinct_bytes(features)
features = create_feature_sql_keywords(features)
features = create_feature_javascript_keywords(features)
del features['payload']
return features
```
### Scoring custom features
Score the custom features using the SelectKBest function, then visualize the scores in a graph
to see which features are less significant
```
Y = payloads['is_malicious']
X = create_features(pd.DataFrame(payloads['payload'].copy()))
test = SelectKBest(score_func=chi2, k='all')
fit = test.fit(X, Y)
# summarize scores
print(fit.scores_)
features = fit.transform(X)
# summarize selected features
# summarize scores
np.set_printoptions(precision=2)
print(fit.scores_)
# Get the indices sorted by most important to least important
indices = np.argsort(fit.scores_)
# To get your top 10 feature names
featuress = []
for i in range(10):
featuress.append(X.columns[indices[i]])
display(featuress)
display([featuress[i] + ' ' + str(fit.scores_[i]) for i in indices[range(10)]])
plt.rcdefaults()
fig, ax = plt.subplots()
y_pos = np.arange(len(featuress))
performance = 3 + 10 * np.random.rand(len(featuress))
error = np.random.rand(len(featuress))
ax.barh(y_pos, fit.scores_[indices[range(10)]], align='center',
color='green', ecolor='black')
ax.set_yticks(y_pos)
ax.set_yticklabels(featuress)
ax.set_xscale('log')
#ax.invert_yaxis() # labels read top-to-bottom
ax.set_xlabel('Points')
ax.set_title('SelectKBest()')
plt.show()
```
# Step3B - Feature engineering using bag of words techniques.
Additional to our custom feature space, we will create 6 more feature spaces using bag-of-words techniques
The following vectorizers below is another way of creating features for text input.
We will test the performance of these techniques independently from our custom features in Step 3A.
We will create vectorizers of these combinations:
- 1-grams CountVectorizer
- 2-grams CountVectorizer
- 3-grams CountVectorizer
- 1-grams TfidfVectorizer
- 2-grams TfidfVectorizer
- 3-grams TfidfVectorizer
The type of N-gram function determines how the actual "words" should be created from the payload string
Each vectorizer is used later in Step4 in Pipeline objects before training
See report for further explanation
### 1-Grams features
create a Countvectorizer and TF-IDFvectorizer that uses 1-grams.
1-grams equals one feature for each letter/symbol recorded
```
def get1Grams(payload_obj):
'''Divides a string into 1-grams
Example: input - payload: "<script>"
output- ["<","s","c","r","i","p","t",">"]
'''
payload = str(payload_obj)
ngrams = []
for i in range(0,len(payload)-1):
ngrams.append(payload[i:i+1])
return ngrams
tfidf_vectorizer_1grams = TfidfVectorizer(tokenizer=get1Grams)
count_vectorizer_1grams = CountVectorizer(min_df=1, tokenizer=get1Grams)
```
### 2-Grams features
create a Countvectorizer and TF-IDFvectorizer that uses 2-grams.
```
def get2Grams(payload_obj):
'''Divides a string into 2-grams
Example: input - payload: "<script>"
output- ["<s","sc","cr","ri","ip","pt","t>"]
'''
payload = str(payload_obj)
ngrams = []
for i in range(0,len(payload)-2):
ngrams.append(payload[i:i+2])
return ngrams
tfidf_vectorizer_2grams = TfidfVectorizer(tokenizer=get2Grams)
count_vectorizer_2grams = CountVectorizer(min_df=1, tokenizer=get2Grams)
```
### 3-Grams features
Create a Countvectorizer and TF-IDFvectorizer that uses 3-grams
```
def get3Grams(payload_obj):
'''Divides a string into 3-grams
Example: input - payload: "<script>"
output- ["<sc","scr","cri","rip","ipt","pt>"]
'''
payload = str(payload_obj)
ngrams = []
for i in range(0,len(payload)-3):
ngrams.append(payload[i:i+3])
return ngrams
tfidf_vectorizer_3grams = TfidfVectorizer(tokenizer=get3Grams)
count_vectorizer_3grams = CountVectorizer(min_df=1, tokenizer=get3Grams)
```
## Step3C - Feature space visualization
After creating our different feature spaces to later train each classifier on,
we first examine them visually by projecting the feature spaces into two dimensions using Principle Component Analysis
Graphs are shown below displaying the data in 3 out of 7 of our feature spaces
```
def visualize_feature_space_by_projection(X,Y,title='PCA'):
'''Plot a two-dimensional projection of the dataset in the specified feature space
input: X - data
Y - labels
title - title of plot
'''
pca = TruncatedSVD(n_components=2)
X_r = pca.fit(X).transform(X)
# Percentage of variance explained for each components
print('explained variance ratio (first two components): %s'
% str(pca.explained_variance_ratio_))
plt.figure()
colors = ['blue', 'darkorange']
lw = 2
#Plot malicious and non-malicious separately with different colors
for color, i, y in zip(colors, [0, 1], Y):
plt.scatter(X_r[Y == i, 0], X_r[Y == i, 1], color=color, alpha=.3, lw=lw,
label=i)
plt.legend(loc='best', shadow=False, scatterpoints=1)
plt.title(title)
plt.show()
```
### 1-Grams CountVectorizer feature space visualization
```
X = count_vectorizer_1grams.fit_transform(payloads['payload'])
Y = payloads['is_malicious']
visualize_feature_space_by_projection(X,Y,title='PCA visualization of 1-grams CountVectorizer feature space')
```
### 3-Grams TFIDFVectorizer feature space visualization
```
X = tfidf_vectorizer_3grams.fit_transform(payloads['payload'])
Y = payloads['is_malicious']
visualize_feature_space_by_projection(X,Y,title='PCA visualization of 3-grams TFIDFVectorizer feature space')
```
### Custom feature space visualization
```
X = create_features(pd.DataFrame(payloads['payload'].copy()))
Y = payloads['is_malicious']
visualize_feature_space_by_projection(X,Y,title='PCA visualization of custom feature space')
```
# Step4 - Model selection and evaluation
First, we will automate hyperparameter tuning and out of sample testing using train_model below
```
def train_model(clf, param_grid, X, Y):
'''Trains and evaluates the model clf from input
The function selects the best model of clf by optimizing for the validation data,
then evaluates its performance using the out of sample test data.
input - clf: the model to train
param_grid: a dict of hyperparameters to use for optimization
X: features
Y: labels
output - the best estimator (trained model)
the confusion matrix from classifying the test data
'''
#First, partition into train and test data
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.2, random_state=42)
n_iter = 5
#If number of possible iterations are less than prefered number of iterations,
#set it to the number of possible iterations
#number of possible iterations are not less than prefered number of iterations if any argument is expon()
#because expon() is continous (writing 100 instead, could be any large number)
n_iter = min(n_iter,np.prod([
100 if type(xs) == type(expon())
else len(xs)
for xs in param_grid.values()
]))
#perform a grid search for the best parameters on the training data.
#Cross validation is made to select the parameters, so the training data is actually split into
#a new train data set and a validation data set, K number of times
cv = ShuffleSplit(n=len(X_train), n_iter=5, test_size=0.2, random_state=0) #DEBUG: n_iter=10
#cv = KFold(n=len(X), n_folds=10)
random_grid_search = RandomizedSearchCV(
clf,
param_distributions=param_grid,
cv=cv,
scoring='f1',
n_iter=n_iter, #DEBUG 1
random_state=5,
refit=True,
verbose=10
)
'''Randomized search used instead. We have limited computing power
grid_search = GridSearchCV(
clf,
param_grid=param_grid,
cv=cv,
scoring='f1', #accuracy/f1/f1_weighted all give same result?
verbose=10,
n_jobs=-1
)
grid_search.fit(X_train, Y_train)
'''
random_grid_search.fit(X_train, Y_train)
#Evaluate the best model on the test data
Y_test_predicted = random_grid_search.best_estimator_.predict(X_test)
Y_test_predicted_prob = random_grid_search.best_estimator_.predict_proba(X_test)[:, 1]
confusion = confusion_matrix(Y_test, Y_test_predicted)
TP = confusion[1, 1]
TN = confusion[0, 0]
FP = confusion[0, 1]
FN = confusion[1, 0]
#Calculate recall (sensitivity) from confusion matrix
sensitivity = TP / float(TP + FN)
#Calculate specificity from confusion matrix
specificity = TN / float(TN + FP)
#Calculate accuracy
accuracy = (confusion[0][0] + confusion[1][1]) / (confusion.sum().sum())
#Calculate axes of ROC curve
fpr, tpr, thresholds = roc_curve(Y_test, Y_test_predicted_prob)
#Area under the ROC curve
auc = roc_auc_score(Y_test, Y_test_predicted_prob)
return {
'conf_matrix':confusion,
'accuracy':accuracy,
'sensitivity':sensitivity,
'specificity':specificity,
'auc':auc,
'params':random_grid_search.best_params_,
'model':random_grid_search.best_estimator_,
'roc':{'fpr':fpr,'tpr':tpr,'thresholds':thresholds}
}
```
Then, we will use the train_model function to train, optimize and retrieve out of sample testing results from a range of classifiers.
Classifiers tested using our custom feature space:
- AdaBoost
- SGD classifier
- MultiLayerPerceptron classifier
- Logistic Regression
- Support Vector Machine
- Random forest
- Decision Tree
- Multinomial Naive Bayes
Classifiers tested using bag-of-words feature spaces:
- MultiLayerPerceptron classifier
- Logistic Regression
- Support Vector Machine
- Random forest
- Multinomial Naive Bayes
Some classifiers were unable to train using a bag-of-words feature space because they couldn't handle sparse graphs
All their best parameters with their performance is stored in a dataframe called classifier_results
Make dictionary of models with parameters to optimize using bag-of-words feature spaces
```
def create_classifier_inputs_using_vectorizers(vectorizer, subscript):
'''make pipelines of the specified vectorizer with the classifiers to train
input - vectorizer: the vectorizer to add to the pipelines
subscript: subscript name for the dictionary key
output - A dict of inputs to use for train_model(); a pipeline and a dict of params to optimize
'''
classifier_inputs = {}
classifier_inputs[subscript + ' MLPClassifier'] = {
'pipeline':Pipeline([('vect', vectorizer),('clf',MLPClassifier(
activation='relu',
solver='adam',
early_stopping=False,
verbose=True
))]),
'dict_params': {
'vect__min_df':[1,2,5,10,20,40],
'clf__hidden_layer_sizes':[(500,250,125,62)],
'clf__alpha':[0.0005,0.001,0.01,0.1,1],
'clf__learning_rate':['constant','invscaling'],
'clf__learning_rate_init':[0.001,0.01,0.1,1],
'clf__momentum':[0,0.9],
}
}
'''
classifier_inputs[subscript + ' MultinomialNB'] = {
'pipeline':Pipeline([('vect', vectorizer),('clf',MultinomialNB())]),
'dict_params': {
'vect__min_df':[1,2,5,10,20,40]
}
}
classifier_inputs[subscript + ' RandomForest'] = {
'pipeline':Pipeline([('vect', vectorizer),('clf',RandomForestClassifier(
max_depth=None,min_samples_split=2, random_state=0))]),
'dict_params': {
'vect__min_df':[1,2,5,10,20,40],
'clf__n_estimators':[10,20,40,60]
}
}
classifier_inputs[subscript + ' Logistic'] = {
'pipeline':Pipeline([('vect', vectorizer), ('clf',LogisticRegression())]),
'dict_params': {
'vect__min_df':[1,2,5,10,20,40],
'clf__C':[0.001, 0.01, 0.1, 1, 10, 100, 1000]
}
}
classifier_inputs[subscript + ' SVM'] = {
'pipeline':Pipeline([('vect', vectorizer), ('clf',SVC(probability=True))]),
'dict_params': {
'vect__min_df':[1,2,5,10,20,40],
'clf__C':[0.001, 0.01, 0.1, 1, 10, 100, 1000],
'clf__gamma':[0.001, 0.0001,'auto'],
'clf__kernel':['rbf']
}
}
'''
return classifier_inputs
```
Make dictionary of models with parameters to optimize using custom feature spaces
```
def create_classifier_inputs(subscript):
classifier_inputs = {}
'''classifier_inputs[subscript + ' GPC'] = {
'pipeline':GaussianProcessClassifier(),
'dict_params': {
'kernel':[
1.0*kernels.RBF(1.0),
1.0*kernels.Matern(),
1.0*kernels.RationalQuadratic(),
1.0*kernels.DotProduct()
]
}
}'''
classifier_inputs[subscript + ' AdaBoostClassifier'] = {
'pipeline':AdaBoostClassifier(n_estimators=100),
'dict_params': {
'n_estimators':[10,20,50, 100],
'learning_rate':[0.1, 0.5, 1.0, 2.0]
}
}
classifier_inputs[subscript + ' SGD'] = {
'pipeline':SGDClassifier(loss="log", penalty="l2"),
'dict_params': {
'learning_rate': ['optimal']
}
}
classifier_inputs[subscript + ' RandomForest'] = {
'pipeline':RandomForestClassifier(
max_depth=None,min_samples_split=2, random_state=0),
'dict_params': {
'n_estimators':[10,20,40,60]
}
}
classifier_inputs[subscript + ' DecisionTree'] = {
'pipeline': DecisionTreeClassifier(max_depth=5),
'dict_params': {
'min_samples_split': [2]
}
}
'''classifier_inputs[subscript + ' MLPClassifier'] = {
'pipeline':MLPClassifier(
activation='relu',
solver='adam',
early_stopping=False,
verbose=True
),
'dict_params': {
'hidden_layer_sizes':[(300, 200, 150, 150), (30, 30, 30), (150, 30, 30, 150),
(400, 250, 100, 100) , (150, 200, 300)],
'alpha':[0.0005,0.001,0.01,0.1,1],
'learning_rate':['constant','invscaling'],
'learning_rate_init':[0.0005,0.001,0.01,0.1,1],
'momentum':[0,0.9],
}
}'''
classifier_inputs[subscript + ' Logistic'] = {
'pipeline':LogisticRegression(),
'dict_params': {
'C': [0.001, 0.01, 0.1, 1, 10, 100, 1000]
}
}
classifier_inputs[subscript + ' MultinomialNB'] = {
'pipeline':MultinomialNB(),
'dict_params': {
'alpha': [1.0]
}
}
'''classifier_inputs[subscript + ' SVM'] = {
'pipeline':SVC(probability=True),
'dict_params': {
'C':[0.001, 0.01, 0.1, 1, 10, 100, 1000],
'gamma':[0.001, 0.0001,'auto'],
'kernel':['rbf']
}
}'''
return classifier_inputs
```
Create a new result table
```
classifier_results = pd.DataFrame(columns=['accuracy','sensitivity','specificity','auc','conf_matrix','params','model','roc'])#,index=classifier_inputs.keys())
```
Use the 6 different feature spaces generated from the vectorizers previously above,
and train every classifier in classifier_inputs in every feature space
### P.S! Don't try to run this, it will take several days to complete
### Instead skip to Step4B
```
classifier_inputs = {}
classifier_inputs.update(create_classifier_inputs_using_vectorizers(count_vectorizer_1grams,'count 1grams'))
classifier_inputs.update(create_classifier_inputs_using_vectorizers(count_vectorizer_2grams,'count 2grams'))
classifier_inputs.update(create_classifier_inputs_using_vectorizers(count_vectorizer_3grams,'count 3grams'))
classifier_inputs.update(create_classifier_inputs_using_vectorizers(tfidf_vectorizer_1grams,'tfidf 1grams'))
classifier_inputs.update(create_classifier_inputs_using_vectorizers(tfidf_vectorizer_2grams,'tfidf 2grams'))
classifier_inputs.update(create_classifier_inputs_using_vectorizers(tfidf_vectorizer_3grams,'tfidf 3grams'))
X = payloads['payload']
Y = payloads['is_malicious']
for classifier_name, inputs in classifier_inputs.items():
display(inputs['dict_params'])
if classifier_name in classifier_results.index.values.tolist():
print('Skipping ' + classifier_name + ', already trained')
else:
result_dict = train_model(inputs['pipeline'],inputs['dict_params'],X,Y)
classifier_results.loc[classifier_name] = result_dict
display(classifier_results)
display(pd.DataFrame(payloads['payload'].copy()))
```
Use our custom feature space,
and train every classifier in classifier_inputs_custom with
### P.S! Don't try to run this, it will take many hours to complete
### Instead skip to Step4B
```
classifier_inputs_custom = {}
#Get classifiers and parameters to optimize
classifier_inputs_custom.update(create_classifier_inputs('custom'))
#Extract payloads and labels
Y = payloads['is_malicious']
X = create_features(pd.DataFrame(payloads['payload'].copy()))
#Select the best features
X_new = SelectKBest(score_func=chi2, k=4).fit_transform(X,Y)
#Call train_model for every classifier and save results to classifier_results
for classifier_name, inputs in classifier_inputs_custom.items():
if classifier_name in classifier_results.index.values.tolist():
print('Skipping ' + classifier_name + ', already trained')
else:
result_dict = train_model(inputs['pipeline'],inputs['dict_params'],X,Y)
classifier_results.loc[classifier_name] = result_dict
display(classifier_results)
#pickle.dump( classifier_results, open( "data/trained_classifiers_custom_all_features.p", "wb" ) )
#Save classifiers in a pickle file to be able to re-use them without re-training
pickle.dump( classifier_results, open( "data/trained_classifiers.p", "wb" ) )
```
### Classifier results
```
#Display the results for the classifiers that were trained using our custom feature space
custom_features_classifiers = pickle.load( open("data/trained_classifier_custom_all_features.p", "rb"))
display(custom_features_classifiers)
#Display the results for the classifiers that were using bag of words feature spaces
classifier_results = pickle.load( open( "data/trained_classifiers.p", "rb" ) )
display(classifier_results)
#Combine the two tables into one table
classifier_results = classifier_results.append(custom_features_classifiers)
classifier_results = classifier_results.sort_values(['sensitivity','accuracy'], ascending=[False,False])
display(classifier_results)
```
### F1-score
Calculate F1-score of each classifier and add to classifiers table
(We didn't implement this in the train_model function as with the other performance metrics because we've already done a 82 hour training session before this and didn't want to re-run the entire training just to add F1-score from inside train_model)
```
def f1_score(conf_matrix):
precision = conf_matrix[0][0] / (conf_matrix[0][0] + conf_matrix[0][1] )
recall = conf_matrix[0][0] / (conf_matrix[0][0] + conf_matrix[1][0] )
return (2 * precision * recall) / (precision + recall)
#load classifier table if not yet loaded
classifier_results = pickle.load( open( "data/trained_classifiers.p", "rb" ) )
#Calculate F1-scores
classifier_results['F1-score'] = [ f1_score(conf_matrix) for conf_matrix in classifier_results['conf_matrix']]
#Re-arrange columns
classifier_results = classifier_results[['F1-score','accuracy','sensitivity','specificity','auc','conf_matrix','params','model','roc']]
#re-sort on F1-score
classifier_results = classifier_results.sort_values(['F1-score','accuracy'], ascending=[False,False])
display(classifier_results)
```
Final formating
Convert numeric columns to float
Round numeric columns to 4 decimals
```
classifier_results[['F1-score','accuracy','sensitivity','specificity','auc']] = classifier_results[['F1-score','accuracy','sensitivity','specificity','auc']].apply(pd.to_numeric)
classifier_results = classifier_results.round({'F1-score':4,'accuracy':4,'sensitivity':4,'specificity':4,'auc':4})
#classifier_results[['F1-score','accuracy','sensitivity','specificity','auc','conf_matrix','params']].to_csv('data/classifiers_result_table.csv')
display(classifier_results.dtypes)
```
### Export classifiers
First, export full list of trained classifiers for later use
Second, pick one classifier to save in a separate pickle, used later to implement in a dummy server
```
#save complete list of classifiers to 'trained_classifiers'
pickle.dump( classifier_results, open( "data/trained_classifiers.p", "wb" ) )
#In this case, we are going to implement tfidf 2grams RandomForest in our dummy server
classifier = (custom_features_classifiers['model'].iloc[0])
print(classifier)
#Save classifiers in a pickle file to be able to re-use them without re-training
pickle.dump( classifier, open( "data/tfidf_2grams_randomforest.p", "wb" ) )
```
## Step4B - load pre-trained classifiers
Instead of re-training all classifiers, load the classifiers from disk that we have already trained
```
classifier_results = pickle.load( open( "data/trained_classifiers.p", "rb" ) )
```
## Step5 - Visualization
In this section we will visualize:
- Histogram of classifier performances
- Learning curves
- ROC curves
### Performance histogram
First, make a histogram of classifier performance measured by F1-score.
Same classifier using different feature spaces are clustered together in the graph
Also, print the table of F1-scores and computes the averages along the x-axis and y-axis,
e.g. the average F1-score for each classifier, and the average F1-score for each feature space
```
def get_classifier_name(index):
'''
Returns the name of the classifier at the given index name
'''
return index.split()[len(index.split())-1]
#Group rows together using same classifier
grouped = classifier_results.groupby(get_classifier_name)
hist_df = pd.DataFrame(columns=['custom','count 1grams','count 2grams','count 3grams','tfidf 1grams','tfidf 2grams','tfidf 3grams'])
for classifier, indices in grouped.groups.items():
#Make a list of feature spaces
feature_spaces = indices.tolist()
feature_spaces = [feature_space.replace(classifier,'') for feature_space in feature_spaces]
feature_spaces = [feature_space.strip() for feature_space in feature_spaces]
#If no result exists, it will stay as 0
hist_df.loc[classifier] = {
'custom':0,
'count 1grams':0,
'count 2grams':0,
'count 3grams':0,
'tfidf 1grams':0,
'tfidf 2grams':0,
'tfidf 3grams':0
}
#Extract F1-score from classifier_results to corrensponding entry in hist_df
for fs in feature_spaces:
hist_df[fs].loc[classifier] = classifier_results['F1-score'].loc[fs + ' ' + classifier]
#Plot the bar plot
f, ax = plt.subplots()
ax.set_ylim([0.989,1])
hist_df.plot(kind='bar', figsize=(12,7), title='F1-score of all models grouped by classifiers', ax=ax, width=0.8)
#Make Avgerage F1-score row and cols for the table and print the table
hist_df_nonzero = hist_df.copy()
hist_df_nonzero[hist_df > 0] = True
hist_df['Avg Feature'] = (hist_df.sum(axis=1) / np.array(hist_df_nonzero.sum(axis=1)))
hist_df_nonzero = hist_df.copy()
hist_df_nonzero[hist_df > 0] = True
hist_df.loc['Avg Classifier'] = (hist_df.sum(axis=0) / np.array(hist_df_nonzero.sum(axis=0)))
hist_df = hist_df.round(4)
display(hist_df)
```
### Learning curves
Create learning curves for a sample of classifiers. This is to visualize how the dataset size impacts the performance
```
def plot_learning_curve(df_row,X,Y):
'''Plots the learning curve of a classifier with its parameters
input - df_row: row of classifier_result
X: payload data
Y: labels
'''
#The classifier to plot learning curve for
estimator = df_row['model']
title = 'Learning curves for classifier ' + df_row.name
train_sizes = np.linspace(0.1,1.0,5)
cv = ShuffleSplit(n=len(X), n_iter=3, test_size=0.2, random_state=0)
#plot settings
plt.figure()
plt.title(title)
plt.xlabel("Training examples")
plt.ylabel("Score")
print('learning curve in process...')
train_sizes, train_scores, test_scores = learning_curve(
estimator, X, Y, cv=cv, n_jobs=-1, train_sizes=train_sizes, verbose=0) #Change verbose=10 to print progress
print('Learning curve done!')
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
plt.grid()
plt.fill_between(train_sizes, train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std, alpha=0.1,
color="r")
plt.fill_between(train_sizes, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.1, color="g")
plt.plot(train_sizes, train_scores_mean, 'o-', color="r",
label="Training score")
plt.plot(train_sizes, test_scores_mean, 'o-', color="g",
label="Cross-validation score")
plt.legend(loc="best")
plt.show()
```
Three examples of learning curves from the trained classifiers.
All learning curves have upsloping cross-validation score at the end,
which means that adding more data would potentially increase the accuracy
```
#plot learning curve for tfidf 1grams RandomForest
X = payloads['payload']
Y = payloads['is_malicious']
plot_learning_curve(classifier_results.iloc[0],X,Y)
#plot learning curve for count 3grams MultinomialNB
X = payloads['payload']
Y = payloads['is_malicious']
plot_learning_curve(classifier_results.iloc[6],X,Y)
#plot learning curve for custom svm
X = create_features(pd.DataFrame(payloads['payload'].copy()))
Y = payloads['is_malicious']
plot_learning_curve(classifier_results.iloc[5],X,Y)
```
### ROC curves
Plot ROC curves for a range of classifiers to visualize the sensitivity/specificity trade-off and the AUC
```
def visualize_result(classifier_list):
'''Plot the ROC curve for a list of classifiers in the same graph
input - classifier_list: a subset of classifier_results
'''
f, (ax1, ax2) = plt.subplots(1,2)
f.set_figheight(6)
f.set_figwidth(15)
#Subplot 1, ROC curve
for classifier in classifier_list:
ax1.plot(classifier['roc']['fpr'], classifier['roc']['tpr'])
ax1.scatter(1-classifier['specificity'],classifier['sensitivity'], edgecolor='k')
ax1.set_xlim([0, 1])
ax1.set_ylim([0, 1.0])
ax1.set_title('ROC curve for top3 and bottom3 classifiers')
ax1.set_xlabel('False Positive Rate (1 - Specificity)')
ax1.set_ylabel('True Positive Rate (Sensitivity)')
ax1.grid(True)
#subplot 2, ROC curve zoomed
for classifier in classifier_list:
ax2.plot(classifier['roc']['fpr'], classifier['roc']['tpr'])
ax2.scatter(1-classifier['specificity'],classifier['sensitivity'], edgecolor='k')
ax2.set_xlim([0, 0.3])
ax2.set_ylim([0.85, 1.0])
ax2.set_title('ROC curve for top3 and bottom3 classifiers (Zoomed)')
ax2.set_xlabel('False Positive Rate (1 - Specificity)')
ax2.set_ylabel('True Positive Rate (Sensitivity)')
ax2.grid(True)
#Add further zoom
left, bottom, width, height = [0.7, 0.27, 0.15, 0.15]
ax3 = f.add_axes([left, bottom, width, height])
for classifier in classifier_list:
ax3.plot(classifier['roc']['fpr'], classifier['roc']['tpr'])
ax3.scatter(1-classifier['specificity'],classifier['sensitivity'], edgecolor='k')
ax3.set_xlim([0, 0.002])
ax3.set_ylim([0.983, 1.0])
ax3.set_title('Zoomed even further')
ax3.grid(True)
plt.show()
```
Plot ROC curves for the top3 classifiers and the bottom 3 classifiers, sorted by F1-score
Left: standard scale ROC curve
Right: zoomed in version of same graph, to easier see in the upper right corner
```
indices = [0,1,2, len(classifier_results)-1,len(classifier_results)-2,len(classifier_results)-3]
visualize_result([classifier_results.iloc[index] for index in indices])
```
## Step6 - Website integration extract
This is the code needed when implementing the saved classifier in tfidf_2grams_randomforest.p on a server
```
import pickle
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.ensemble import RandomForestClassifier
def get2Grams(payload_obj):
'''Divides a string into 2-grams
Example: input - payload: "<script>"
output- ["<s","sc","cr","ri","ip","pt","t>"]
'''
payload = str(payload_obj)
ngrams = []
for i in range(0,len(payload)-2):
ngrams.append(payload[i:i+2])
return ngrams
classifier = pickle.load( open("data/tfidf_2grams_randomforest.p", "rb"))
def injection_test(inputs):
variables = inputs.split('&')
values = [ variable.split('=')[1] for variable in variables]
print(values)
return 'MALICIOUS' if classifier.predict(values).sum() > 0 else 'NOT_MALICIOUS'
#test injection_test
display(injection_test("val1=%3Cscript%3Ekiddie"))
```
# (Step7)
we can display which types of queries the classifiers failed to classify. These are interesting to examine for further work on how to improve the classifiers and the quality of the data set
```
pipe = Pipeline([('vect', vectorizer), ('clf',LogisticRegression(C=10))])
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.2, random_state=42)
cv = ShuffleSplit(n=len(X_train), n_iter=1, test_size=0.2, random_state=0) #DEBUG: n_iter=10
random_grid_search = RandomizedSearchCV(
pipe,
param_distributions={
'clf__C':[10]
},
cv=cv,
scoring='roc_auc',
n_iter=1,
random_state=5,
refit=True
)
random_grid_search.fit(X_train, Y_train)
#Evaluate the best model on the test data
Y_test_predicted = random_grid_search.best_estimator_.predict(X_test)
#Payloads classified incorrectly
pd.options.display.max_colwidth = 200
print('False positives')
print(X_test[(Y_test == 0) & (Y_test_predicted == 1)])
print('False negatives')
print(X_test[(Y_test == 1) & (Y_test_predicted == 0)])
```
| github_jupyter |
Handling files belongs also to the basic skills in programming, that's why this chapter was added as a completion by me (Kinga Sipos).
<!--NAVIGATION-->
< [Strings and Regular Expressions](13-Strings-and-Regular-Expressions.ipynb) | [Contents](Index.ipynb) | [Modules and Packages](15-Modules-and-Packages.ipynb) >
# File Input and Output
## Filesystem operations
Filesystem operations can be carried out by executing a normal shell command preceded by exclamation mark:
```
!ls
```
Another alternative to operate files is to use the ``os`` module:
* ``os.getcwd()``- Returns the path to the current working directory.
* ``os.chdir(path)`` - Changes the current working directory to path.
* ``os.listdir(dir)`` - Returns the list of entries in directory dir (omitting ‘.’ and ‘..’)
* ``os.makedirs(path)`` - Creates a directory; nothing happens if the directory already exists. Creates all the intermediate-level directories needed to contain the leaf.
* ``os.rename(old,new)`` - Renames a file or directory from old to new.
Specific path related functions are methods of ``os.path``:
* ``os.path.exists(path)`` - Returns True if path exists.
* ``os.path.isdir(path)`` - Returns True if path is a directory.
* ``os.path.isfile(path)`` - Returns True if path is a regular file.
* ``os.path.basename(path)`` - Returns the base name (the part after the last ‘/’ character)
* ``os.path.dirname(path)`` - Returns the directory name (the part before the last / character).
* ``os.path.abspath(path)`` - Make path absolute (i.e., start with a /).
## Read from a file and write to a file
### Reading from and writing to textfiles
1. The **first line** of code for processing a text file usually looks like this:
`with open(filename, mode) as stream:` - which prepares the file for processing.
Mode is one of ``'r'``, ``'w'`` or ``'a'`` for reading, writing, appending. You can add a ‘+’ character to enable read+write (other effects being the same).
`stream = open(filename, mode)` is equivalent to the first line of the previous code, the difference is that `with` ensures that the file is closed after the desired operation is carried out, otherwise one should close the file explicitely by the `stream.close()` command.
2. a) If the chosen processing mode is read, the **second line** can be something like
`content = stream.read()` - which returns the whole content of a file as a multiline string
or
`content = stream.readlines()` - which returns the content of the file as a list of one line strings
or
`for line in stream:` - which reads the lines of file line by line.
2. b) If the chosen processing is write, the **second line** can be
`stream.write(text)`
### Exercise
As an exercise we will create a file with the haikus from the previous chapter.
```
mytext = """WORKSHOP HAIKU
translated by Éva Antal
Perhaps do not even touch it.
Just look at it, look at it,
until it becomes beautiful.
TEST QUESTION FOR EVERY DAY
translated by Éva Antal
Do you still see
what you look at, or you only
know: "there" "it" "is"?
FROM THE BEST OF INTENTIONS
translated by Gábor G. Gyukics and Michael Castro
fall asleep;
die the same way a child
bites into an apple.
MEETING
translated by Gábor G. Gyukics and Michael Castro
I plan it as a farewell
THE HAIKU
translated by Tamás Révbíró
in front of my feet
a bird sat, and then took flight.
Now I'm heavier.
AXIOM
translated by Tamás Révbíró
You should try and help
everything to be the way
it is anyway.
ECHO ON EPICTETUS
translated by Tamás Révbíró
Don't say, "I lost it",
about anything. Rather
say, "I gave it back".
AXIOM
translated by Tamás Révbíró
Parents and killers:
almost-innocent servants.
They just execute.
ZENsation
translated by Tamás Révbíró
Look, the snow gives body to the wind!
DISILLUSIONIST
translated by Tamás Révbíró
Why should I travel
when I can be a stranger
right here, standing still?"""
with open('Haikus.txt', 'w') as outstream:
outstream.write(mytext)
```
One can check whether the file is closed.
```
outstream.closed
```
Now let's read the first two lines from the created file.
```
with open('Haikus.txt', 'r') as instream:
print(instream.readline())
print(instream.readline())
```
This time let's read all the lines of the file into a list and print the first 6 lines.
```
with open('Haikus.txt', 'r') as instream:
textlines = instream.readlines()
for i in range(6):
print(textlines[i])
```
### Reading from and writing to Comma Separated Values files
Reading and writing can be performed in the same way as above.
For example one can create a CSV file by the following code:
```
import csv
with open('employee_file.csv', mode='w') as employee_file:
employee_writer = csv.writer(employee_file, delimiter=',', quotechar='"', quoting=csv.QUOTE_MINIMAL)
employee_writer.writerow(['John Smith', 'Accounting', 'November', '27'])
employee_writer.writerow(['Erica Meyers', 'IT', 'March', '31'])
```
or with the following code:
```
import csv
with open('employee_file2.csv', mode='w') as csv_file:
fieldnames = ['emp_name', 'dept', 'birth_month', 'age']
writer = csv.DictWriter(csv_file, fieldnames=fieldnames)
writer.writeheader()
writer.writerow({'emp_name': 'John Smith', 'dept': 'Accounting', 'birth_month': 'November', 'age': '27'})
writer.writerow({'emp_name': 'Erica Meyers', 'dept': 'IT', 'birth_month': 'March', 'age': 31})
```
Reading can be performed in the following way:
```
with open('employee_file2.csv') as csv_file:
csv_reader = csv.reader(csv_file, delimiter=',')
line_count = 0
for row in csv_reader:
if line_count == 0:
print(f'Column names are {", ".join(row)}')
line_count += 1
else:
print(f'\t{row[0]} works at {row[1]}, has birthday in {row[2]} and is {row[3]} years old.')
line_count += 1
print(f'Processed {line_count} lines.')
with open('employee_file2.csv', mode='r') as csv_file:
csv_reader = csv.DictReader(csv_file)
line_count = 0
for row in csv_reader:
if line_count == 0:
print(f'Column names are {", ".join(row)}')
line_count += 1
print(f'\t{row["emp_name"]} works in the {row["dept"]} department, and was born in {row["birth_month"]}.')
line_count += 1
print(f'Processed {line_count} lines.')
```
One can import a CSV file directly as dataframe.
```
import pandas as pd
df = pd.read_csv('employee_file2.csv')
print(df)
```
Experiment with possible methods and attributes of dataframes!
```
df.head()
df.describe()
df.dtypes
```
<!--NAVIGATION-->
< [Strings and Regular Expressions](13-Strings-and-Regular-Expressions.ipynb) | [Contents](Index.ipynb) | [Modules and Packages](15-Modules-and-Packages.ipynb) >
| github_jupyter |
# 📝 Exercise M5.01
In the previous notebook, we showed how a tree with a depth of 1 level was
working. The aim of this exercise is to repeat part of the previous
experiment for a depth with 2 levels to show how the process of partitioning
is repeated over time.
Before to start, we will:
* load the dataset;
* split the dataset into training and testing dataset;
* define the function to show the classification decision function.
```
import pandas as pd
penguins = pd.read_csv("../datasets/penguins_classification.csv")
culmen_columns = ["Culmen Length (mm)", "Culmen Depth (mm)"]
target_column = "Species"
```
<div class="admonition note alert alert-info">
<p class="first admonition-title" style="font-weight: bold;">Note</p>
<p class="last">If you want a deeper overview regarding this dataset, you can refer to the
Appendix - Datasets description section at the end of this MOOC.</p>
</div>
```
from sklearn.model_selection import train_test_split
data, target = penguins[culmen_columns], penguins[target_column]
data_train, data_test, target_train, target_test = train_test_split(
data, target, random_state=0
)
```
Create a decision tree classifier with a maximum depth of 2 levels and fit
the training data. Once this classifier trained, plot the data and the
decision boundary to see the benefit of increasing the depth. To plot the
decision boundary, you should import the class `DecisionBoundaryDisplay`
from the module `helpers.plotting` as shown in the previous course notebook.
```
# solution
from sklearn.tree import DecisionTreeClassifier
tree = DecisionTreeClassifier(max_depth=2)
tree.fit(data_train, target_train)
import matplotlib.pyplot as plt
import seaborn as sns
from helpers.plotting import DecisionBoundaryDisplay
palette = ["tab:red", "tab:blue", "black"]
DecisionBoundaryDisplay.from_estimator(
tree, data_train, response_method="predict", cmap="RdBu", alpha=0.5
)
ax = sns.scatterplot(data=penguins, x=culmen_columns[0], y=culmen_columns[1],
hue=target_column, palette=palette)
plt.legend(bbox_to_anchor=(1.05, 1), loc='upper left')
_ = plt.title("Decision boundary using a decision tree")
```
Did we make use of the feature "Culmen Length"?
Plot the tree using the function `sklearn.tree.plot_tree` to find out!
```
# solution
from sklearn.tree import plot_tree
_, ax = plt.subplots(figsize=(16, 12))
_ = plot_tree(tree, feature_names=culmen_columns,
class_names=tree.classes_, impurity=False, ax=ax)
```
The resulting tree has 7 nodes: 3 of them are "split nodes" and 4
are "leaf nodes" (or simply "leaves"), organized in 2 levels.
We see that the second tree level used the "Culmen Length" to make
two new decisions. Qualitatively, we saw that such a simple tree was enough
to classify the penguins' species.
Compute the accuracy of the decision tree on the testing data.
```
# solution
test_score = tree.fit(data_train, target_train).score(data_test, target_test)
print(f"Accuracy of the DecisionTreeClassifier: {test_score:.2f}")
```
At this stage, we have the intuition that a decision tree is built by
successively partitioning the feature space, considering one feature at a
time.
We predict an Adelie penguin if the feature value is below the threshold,
which is not surprising since this partition was almost pure. If the feature
value is above the threshold, we predict the Gentoo penguin, the class that
is most probable.
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Dynamic-Schedule" data-toc-modified-id="Dynamic-Schedule-1"><span class="toc-item-num">1 </span>Dynamic Schedule</a></span><ul class="toc-item"><li><span><a href="#Homogeneous-Exponential-Case" data-toc-modified-id="Homogeneous-Exponential-Case-1.1"><span class="toc-item-num">1.1 </span>Homogeneous Exponential Case</a></span></li><li><span><a href="#Heterogeneous-Exponential-Case" data-toc-modified-id="Heterogeneous-Exponential-Case-1.2"><span class="toc-item-num">1.2 </span>Heterogeneous Exponential Case</a></span></li><li><span><a href="#Phase-Type-Case" data-toc-modified-id="Phase-Type-Case-1.3"><span class="toc-item-num">1.3 </span>Phase-Type Case</a></span><ul class="toc-item"><li><span><a href="#Phase-Type-Fit" data-toc-modified-id="Phase-Type-Fit-1.3.1"><span class="toc-item-num">1.3.1 </span>Phase-Type Fit</a></span></li><li><span><a href="#Weighted-Erlang-Distribution" data-toc-modified-id="Weighted-Erlang-Distribution-1.3.2"><span class="toc-item-num">1.3.2 </span>Weighted Erlang Distribution</a></span></li><li><span><a href="#Hyperexponential-Distribution" data-toc-modified-id="Hyperexponential-Distribution-1.3.3"><span class="toc-item-num">1.3.3 </span>Hyperexponential Distribution</a></span></li></ul></li></ul></li></ul></div>
# Dynamic Schedule
_Roshan Mahes, Michel Mandjes, Marko Boon_
In this notebook we determine dynamic schedules that minimize the following cost function:
\begin{align*}
\omega \sum_{i=1}^{n}\mathbb{E}I_i + (1 - \omega)\sum_{i=1}^{n}\mathbb{E}W_i,\quad \omega\in(0,1),
\end{align*}
where $I_i$ and $W_i$ are the expected idle and waiting time associated to client $i$, respectively. We assume that the service tasks $B_1,\dots,B_n$ are independent and solve the problem assuming different types of distributions.
The following packages are required:
```
# math
import numpy as np
import scipy
import math
from scipy.stats import binom, erlang, poisson
from scipy.optimize import minimize
# web scraping
from urllib.request import urlopen
from bs4 import BeautifulSoup as soup
import pandas as pd
# plotting
import plotly.graph_objects as go
import plotly.express as px
from itertools import cycle
# caching
from functools import cache
```
## Homogeneous Exponential Case
In the first case, we assume $B_1,\dots,B_n \stackrel{i.i.d.}{\sim} B \stackrel{d}{=} \text{Exp}(\mu)$ for some $\mu > 0$. In our thesis, we have determined a recursive procedure. We state the result.
<div class="alert alert-warning">
<b>Corollary 2.5.</b>
For arrival time $t$ we have, with $X_t \sim \text{Pois}(\mu t)$ and $\ell = 2,\dots,k+1$,
\begin{align*}
p_{k1}(t) = \mathbb{P}(X_t\geq k),\quad
p_{k\ell}(t) = \mathbb{P}(X_t = k-\ell+1).
\end{align*}
</div>
<div class="alert alert-warning">
<b>Proposition 2.7.</b>
Let $X_t \sim \text{Pois}(\mu t)$. Then
\begin{align*}
f_k(t) &= t\mathbb{P}(X_t\geq k) - \frac{k}{\mu}\mathbb{P}(X_t\geq k+1), \\
g_k(t) &= \frac{k(k-1)}{2\mu}\mathbb{P}(X_t\geq k+1) + (k-1)t\mathbb{P}(X_t\leq k-1) - \frac{\mu t^2}{2}\mathbb{P}(X_t\leq k-2).
\end{align*}
</div>
<div class="alert alert-warning">
<b>Theorem 3.5.</b>
Let $p_{k\ell}(t)$, $f_k(t)$ and $g_k(t)$ be given by Corollary 2.5 and Proposition 2.7. The following recursion holds: for $i=1,\dots,n-1$ and $k=1,\dots,i$,
\[
C_i^{\star}(k) = \inf_{t\geq 0}\left(\omega f_k(t) + (1 - \omega)g_k(t) + \sum_{\ell=1}^{k+1}p_{k\ell}(t)C_{i+1}^{\star}(\ell)\right),
\]
whereas, for $k=1,\dots,n$,
\[
C_n^{\star}(k) = (1-\omega)g_{k}(\infty) = (1-\omega)\frac{k(k-1)}{2\mu}.
\]
</div>
We have implemented the formulas as follows.
```
def cost(t,i,k,mu,omega,n,C_matrix,use_h=True):
"""
Computes the cost of the (remaining) schedule
when t is the next interarrival time.
"""
Fk = [poisson.cdf(k,mu*t), poisson.cdf(k-2,mu*t), poisson.cdf(k-1,mu*t)]
f = (1 - Fk[-1]) * t - (1 - Fk[0]) * k / mu
if use_h:
g = (k - 1) / mu
else:
g = Fk[-1] * (k - 1) * t - Fk[-2] * mu * t**2 / 2 + (1 - Fk[0]) * k * (k - 1) / (2 * mu)
cost = omega * f + (1 - omega) * g
cost += (1 - Fk[-1]) * Cstar_homexp(i+1,1,mu,omega,n,C_matrix,use_h)
for l in range(2,k+2):
cost += poisson.pmf(k-l+1,mu*t) * Cstar_homexp(i+1,l,mu,omega,n,C_matrix,use_h)
return cost
def Cstar_homexp(i,k,mu=1,omega=1/2,n=15,C_matrix=None,use_h=True):
"""
Computes C*_i(k) in the homogeneous exponential case.
"""
if C_matrix[i-1][k-1] != None: # retrieve stored value
pass
elif i == n: # initial condition
if use_h:
C_matrix[i-1][k-1] = (1 - omega) * (k - 1) / mu
else:
C_matrix[i-1][k-1] = (1 - omega) * k * (k - 1) / (2 * mu)
else:
optimization = minimize(cost,0,args=(i,k,mu,omega,n,C_matrix,use_h),method='Nelder-Mead')
C_matrix[i-1][k-1] = optimization.fun
minima[i-1][k-1] = optimization.x[0]
return C_matrix[i-1][k-1]
```
Now we plot our dynamic schedule for $n = 15$ and $\omega = 0.5$:
```
omega = 0.5
n = 15
# compute schedule
C_matrix = [[None for k in range(n+1)] for i in range(n)]
minima = [[None for k in range(n+1)] for i in range(n)]
for i in range(1,n+1):
for k in range(1,i+1):
Cstar_homexp(i,k,mu=1,omega=omega,n=n,C_matrix=C_matrix,use_h=True)
# plot schedule
palette = cycle(px.colors.cyclical.mrybm[2:])
fig = go.Figure()
for k in range(1,n):
fig.add_trace(go.Scatter(x=np.arange(1,n+2), y=[minima[i][k-1] for i in range(n)],
name=k, marker_color=next(palette)))
fig.update_layout(
template='plotly_white',
title='$\\text{Dynamic Schedule}\ (n=' + f'{n},\ \omega={omega})$',
legend_title='$\\text{Clients in System}\ (k)$',
xaxis = {'title': '$\\text{Client Position}\ (i)$', 'range': [0.7, n - 0.7], 'dtick': 1},
yaxis = {'title': '$\\text{Interarrival Time}\ (\\tau_{i}(k))$', 'dtick': 1}
)
fig.show()
print(f'Cost: {C_matrix[0][0]}')
minima
```
## Heterogeneous Exponential Case
Now we consider the case that the service tasks $B_i$ are independent and _heterogeneous exponentially_ distributed, i.e. $B_i \sim \text{Exp}(\mu_i)$, $i=1,\dots,n$. For ease we assume that all $\mu_i$ are distinct, i.e., $\mu_i \neq \mu_j$ for $i,j = 1,\dots,n$, $i\neq j$, but the case that some of the $\mu_i$ coincide can be considered analogously. We obtain the following result.
<div class="alert alert-warning">
<b>Lemma 2.12.</b>
For $k=1,\dots,n$ and $\ell=0,\dots,n-k$, we can write the density $\varphi_{k\ell}$ as
\[
\varphi_{k\ell}(s) := \mathbb{P}\left(\sum_{j=k}^{k+\ell}B_j \in\mathrm{d}s\right)
= \sum_{j=k}^{k+\ell}c_{k\ell j}e^{-\mu_j s},\quad s \geq 0.
\]
The coefficients $c_{k\ell j}$ are given recursively through $c_{k0k} = \mu_k$ and
\[
c_{k,\ell+1,j} = c_{k\ell j}\frac{\mu_{k+\ell+1}}{\mu_{k+\ell+1} - \mu_j}\quad \text{for}\ j = k,\dots,k+\ell,\quad c_{k,\ell+1,k+\ell+1} = \sum_{j=k}^{k+\ell}c_{k\ell j}\frac{\mu_{k+\ell+1}}{\mu_j - \mu_{k+\ell+1}}.
\]
</div>
<div class="alert alert-warning">
<b>Proposition 2.16.</b>
For $i=1,\dots,n-1$, $k=1,\dots,i$, $\ell = 2,\dots,k+1$ and $t\geq 0$,
\[
p_{k1,i}(t) = 1 - \sum_{\ell=2}^{k+1}p_{k\ell,i}(t),\quad
p_{k\ell,i}(t) = \frac{\varphi_{i-k+1,k-\ell+1}(t)}{\mu_{i-\ell+2}}.
\]
</div>
<div class="alert alert-warning">
<b>Proposition 2.17.</b>
For $i=1,\dots,n-1$ and $k=1,\dots,i$,
\begin{align*}
f_{k,i}(t) = t - \sum_{j=i-k+1}^{i}\frac{c_{i-k+1,k-1,j}}{\mu_j}\psi_{j}(t),
\quad
g_{k,i}(t) = \sum_{\ell=0}^{k-1}(k-\ell-1)\sum_{j=i-k+1}^{i-k+\ell+1}\frac{c_{i-k+1,\ell,j}}{\mu_{i-k+\ell+1}}\psi_{j}(t),
\end{align*}
with $\psi_{j}(t) = (1 - e^{-\mu_j t})/\mu_j$.
</div>
<div class="alert alert-warning">
<b>Theorem 3.9.</b>
We can determine the $C^{\star}_i(k)$ recursively: for $i=1,\dots,n-1$ and $k=1,\dots,i$,
\[
C^{\star}_i(k) = \inf_{t\ge 0}\left(\omega f_{k,i}(t) + (1-\omega)g_{k,i}(t) + \sum_{\ell=1}^{k+1}p_{k\ell,i}(t)C^{\star}_{i+1}(\ell)\right),
\]
whereas, for $k=1,\dots,n$,
\[
C^{\star}_n(k) = (1 - \omega)g_{k,n}(\infty) = (1 - \omega)\sum_{\ell=0}^{k-1}(k-\ell-1)\frac{1}{\mu_{n-k+\ell+1}}.
\]
</div>
These formulas lead to the following implementation.
```
# helper functions
def c(k,l,j,mu):
"""Computes the weights c of phi recursively (Lemma 2.23)."""
# storage indices
k_, l_, j_ = k - 1, l, j - 1
if c_stored[k_][l_][j_] != None:
pass
elif k == j and not l:
c_stored[k_][l_][j_] = mu[k_]
elif l:
if j >= k and j < k + l:
c_stored[k_][l_][j_] = c(k,l-1,j,mu) * mu[k_+l_] / (mu[k_+l_] - mu[j-1])
elif k + l == j:
c_stored[k_][l_][j_] = sum([c(k,l-1,m,mu) * mu[j-1] / (mu[m-1] - mu[j-1])
for m in range(k,k+l)])
return c_stored[k_][l_][j_]
def phi(k,l,s,mu):
return sum([c(k,l,j,mu) * math.exp(-mu[j-1] * s) for j in range(k,k+l+1)])
def psi(j,t,mu):
return (1 - math.exp(-mu[j-1] * t)) / mu[j-1]
# transition probabilities
def trans_prob_het(t,i,k,mu):
"""Computes the transition probabilities (Prop. 2.25)."""
p = [phi(i-k+1,k-l+1,t,mu) / mu[i-l+1] for l in range(2,k+2)]
return [1 - sum(p)] + p
# cost function
def cost_het(t,i,k,mu,omega,n,C_matrix,use_h=True):
"""Computes the cost of the (remaining) schedule
when t is the next interarrival time."""
f = t - sum([c(i-k+1,k-1,j,mu) * psi(j,t,mu) / mu[j-1] for j in range(i-k+1,i+1)])
if use_h:
g = sum(1 / mu[i-k:i-1])
else:
g = 0
for l in range(k-1):
g += (k - l - 1) * sum([c(i-k+1,l,j,mu) * psi(j,t,mu) / mu[i-k+l] for j in range(i-k+1,i-k+l+2)])
p = trans_prob_het(t,i,k,mu)
cost = omega * f + (1 - omega) * g
cost += sum([Cstar_het(i+1,l,mu,omega,n,C_matrix,use_h) * p[l-1] for l in range(1,k+2)])
return cost
def Cstar_het(i,k,mu,omega,n,C_matrix,use_h=True):
"""Computes C*_i(k) in the heterogeneous exponential case."""
if C_matrix[i-1][k-1] != None: # retrieve stored value
pass
elif i == n: # initial condition
if use_h:
C_matrix[i-1][k-1] = (1 - omega) * sum(1 / mu[i-k:i-1])
else:
C_matrix[i-1][k-1] = (1 - omega) * sum([(k - l - 1) / mu[n-k+l] for l in range(k)])
else:
optimization = minimize(cost_het,0,args=(i,k,mu,omega,n,C_matrix,use_h))#,bounds=((0,500),))
C_matrix[i-1][k-1] = optimization.fun
minima[i-1][k-1] = optimization.x[0]
return C_matrix[i-1][k-1]
```
Again we can plot our dynamic schedule:
```
omega = 0.5
n = 11
mus = np.linspace(0.5,1.5,n)
# plot schedule
palette = cycle(px.colors.cyclical.mrybm[2:])
fig = go.Figure()
print(f'omega = {omega}\nmu = {mus}\n')
C_matrix = [[None for k in range(n)] for i in range(n)]
minima = [[None for k in range(n)] for i in range(n)]
c_stored = [[[None for j in range(n)] for l in range(n)] for k in range(n)]
# compute values
for i in range(1,n+1):
for k in range(1,i+1):
Cstar_het(i,k,mus,omega=omega,n=n,C_matrix=C_matrix,use_h=True)
# cost
print(f'Cost: {C_matrix[0][0]}')
for k in range(1,n):
fig.add_trace(go.Scatter(x=np.arange(1,n+2), y=[minima[i][k-1] for i in range(n)],
name=k, marker_color=next(palette)))
fig.update_layout(
template='plotly_white',
title='$\\text{Dynamic Schedule}\ (n=' + f'{n},\ \omega={omega})$',
legend_title='$\\text{Clients in System}\ (k)$',
xaxis = {'title': '$\\text{Client Position}\ (i)$', 'range': [0.7, n - 0.7], 'dtick': 1},
yaxis = {'title': '$\\text{Interarrival Time}\ (\\tau_{i}(k))$', 'dtick': 1},
width=800,
height=600
)
fig.show()
```
## Phase-Type Case
Our most general case consists of service time distributions constructed by convolutions and mixtures of exponential distributions, the so-called _phase-type distributions_.
### Phase-Type Fit
There are two special cases of phase-type distributions that are of particular interest: the weighted Erlang distribution and the hyperexponential distribution. The idea is to fit the first two moments of the real service-time distribution. The former distribution can be used to approximate any non-negative distribution with coefficient of variation below 1, whereas the latter can be used if this coefficient of variation is larger than 1. The parameters of the weighted Erlang and hyperexponential distribution are obtained with the following function.
```
def SCV_to_params(SCV, mean=1):
# weighted Erlang case
if SCV <= 1:
K = math.floor(1/SCV)
p = ((K + 1) * SCV - math.sqrt((K + 1) * (1 - K * SCV))) / (SCV + 1)
mu = (K + 1 - p) / mean
return K, p, mu
# hyperexponential case
else:
p = 0.5 * (1 + np.sqrt((SCV - 1) / (SCV + 1)))
mu = 1 / mean
mu1 = 2 * p * mu
mu2 = 2 * (1 - p) * mu
return p, mu1, mu2
```
In the following subsections we develop procedures for finding the optimal static schedule in the weighted Erlang case and the hyperexponential case, respectively.
### Weighted Erlang Distribution
In this case, we assume that the service time $B$ equals w.p. $p\in[0,1]$ an Erlang-distributed random variable with $K$ exponentially distributed phases, each of them having mean $\mu^{-1}$, and with probability $1-p$ an Erlang-distributed random variable with $K+1$ exponentially distributed phases, again with mean $\mu^{-1}$:
\begin{align*}
B \stackrel{\text{d}}{=} \sum_{i=1}^{K}X_i + X_{K+1}\mathbb{1}_{\{U > p\}},
\end{align*}
where $X_i \stackrel{iid}{\sim} \text{Exp}(\mu)$ and $U\sim\text{Unif}[0,1]$. The following recursion can be found in the thesis.
<div class="alert alert-warning">
<b>Theorem 3.16 (discrete version).</b>
For $i=1,\dots,n-1$, $k=1,\dots,i$, and $m\in\mathbb{N}_0$,
\[
\xi_i(k,m) = \inf_{t\in \mathbb{N}_0}\Bigg(\omega \bar{f}^{\circ}_{k,m\Delta}(t\Delta)
+ (1 - \omega)\bar{h}^{\circ}_{k,m\Delta} + \sum_{\ell=2}^{k}\sum_{j=0}^{t}\bar{q}_{k\ell,mj}(t)\xi_{i+1}(\ell,j)
+ P^{\downarrow}_{k,m\Delta}(t\Delta)\xi_{i+1}(1,0) + P^{\uparrow}_{k,m\Delta}(t\Delta)\xi_{i+1}(k+1,m+t) \Bigg),
\]
whereas, for $k=1,\dots,n$ and $m \in \mathbb{N}_0$,
\[
\xi_n(k,m) = (1 - \omega)\bar{h}^{\circ}_{k,m\Delta}.
\]
</div>
Below is our implementation.
```
### helper functions
@cache
def gamma(z, u):
gamma_circ = poisson.pmf(z-1, mu*u)
if z == K + 1:
gamma_circ *= (1 - p)
return gamma_circ / B_sf(u)
@cache
def B_sf(t):
"""The survival function P(B > t)."""
return poisson.cdf(K-1, mu*t) + (1 - p) * poisson.pmf(K, mu*t)
@cache
def P_k0(k, z, t):
"""Computes P(N_t- = 0 | N_0 = k, Z_0 = z)."""
if z <= K:
return sum([binom.pmf(m, k, 1-p) * erlang.cdf(t, k*K-z+1+m, scale=1/mu) for m in range(k+1)])
elif z == K + 1:
return sum([binom.pmf(m, k-1, 1-p) * erlang.cdf(t, (k-1)*K+1+m, scale=1/mu) for m in range(k)])
@cache
def psi(v, t, k, l):
"""
Computes P(t-v < Erl(k,mu) < t, Erl(k,mu) + Erl(l-k,mu) > t),
where Erl(k,mu) and Erl(l-k,mu) are independent.
"""
return sum([poisson.pmf(j, mu*t) * binom.sf(j-k, j, v/t) for j in range(k, l)])
@cache
def f(k, t):
return poisson.sf(k-1, mu*t) * t - poisson.sf(k, mu*t) * k / mu
@cache
def f_bar(k, z, t):
"""Computes the mean idle time given (N_0, Z_0) = (k,z)."""
if z <= K:
return sum([binom.pmf(m, k, 1 - p) * f(k*K+1-z+m, t) for m in range(k+1)])
elif z == K + 1:
return sum([binom.pmf(m, k-1, 1 - p) * f((k-1)*K+1+m, t) for m in range(k)])
@cache
def f_circ(k, u, t):
"""Computes the mean idle time given (N_0, B_0) = (k,u)."""
return sum([gamma(z, u) * f_bar(k, z, t) for z in range(1, K+2)])
@cache
def h_bar(k, z):
"""Computes the mean waiting time given (N_0, Z_0) = (k,z)."""
if k == 1:
return 0
elif z <= K:
return ((k - 1) * (K + 1 - p) + 1 - z) / mu
elif z == K + 1:
return ((k - 2) * (K + 1 - p) + 1) / mu
@cache
def h_circ(k, u):
"""Computes the mean waiting time given (N_0, B_0) = (k,u)."""
return sum([gamma(z, u) * h_bar(k, z) for z in range(1, K+2)])
### transition probabilities
# 1. No client has been served before time t.
@cache
def P_up(k, u, t):
"""Computes P(N_t- = k | N_0 = k, B_0 = u)."""
return B_sf(u+t) / B_sf(u)
# 2. All clients have been served before time t.
@cache
def P_down(k, u, t):
"""Computes P(N_t- = 0 | N_0 = k, B_0 = u)."""
return sum([gamma(z, u) * P_k0(k, z, t) for z in range(1, K+2)])
# 3. Some (but not all) clients have been served before time t.
@cache
def q(diff, z, v, t):
"""
Computes P(N_t = l, B_t < v | N_0 = k, Z_0 = z).
Note: diff = k-l.
"""
q = 0
if z <= K:
for m in range(diff+2):
I_klmz = (diff + 1) * K - z + m + 1
E = p * psi(v, t, I_klmz, I_klmz+K) + (1 - p) * psi(v, t, I_klmz, I_klmz+K+1)
q += binom.pmf(m, diff+1, 1-p) * E
elif z == K + 1:
for m in range(diff+1):
I_klm = diff * K + m + 1
E = p * psi(v, t, I_klm, I_klm+K) + (1 - p) * psi(v, t, I_klm, I_klm+K+1)
q += binom.pmf(m, diff, 1-p) * E
return q
@cache
def q_bar(diff, m, j, t):
"""
Approximates P(N_{t*Delta} = l, B_{t*Delta} in d(j*Delta) | N_0 = k, B_0 = m * Delta).
Note: diff = k-l.
"""
lower = min(max(0, (j - 0.5) * Delta), t*Delta)
upper = min(max(0, (j + 0.5) * Delta), t*Delta)
q_bar = sum([gamma(z, m*Delta) * (q(diff, z, upper, t*Delta) - q(diff, z, lower, t*Delta)) for z in range(1, K+2)])
return q_bar
### cost function
@cache
def cost_we(t, i, k, m):
"""Computes (approximately) the cost when
t/Delta is the next interarrival time."""
cost = omega * f_circ(k, m*Delta, t*Delta) + (1 - omega) * h_circ(k, m*Delta)
cost += P_down(k, m*Delta, t*Delta) * xi_we(i+1, 1, 0) + P_up(k, m*Delta, t*Delta) * xi_we(i+1, k+1, m+t) ####
# print('f_circ(k, m*Delta, t*Delta)', f_circ(k, m*Delta, t*Delta))
# print('h_circ(k, m*Delta)', h_circ(k, m*Delta))
# print('P_down(k, m*Delta, t*Delta)', P_down(k, m*Delta, t*Delta))
# print('xi_we(i+1, 1, 0)', xi_we(i+1, 1, 0))
# print('P_up(k, m*Delta, t*Delta', P_up(k, m*Delta, t*Delta))
# print('xi_we(i+1, k+1, m+t)', xi_we(i+1, k+1, m+t))
for l in range(2, k+1):
for j in range(t+1):
cost += q_bar(k-l, m, j, t) * xi_we(i+1, l, j)
return cost
k, u = 3, 4
h_circ(k, u)
i = 2
k = 1
m = 1
t = 9
# cost_we(t, i, k, m)
# for t in range(1,21):
# print(t, cost_we(t, i, k, m) - cost_we(t-1, i, k, m))
(1 - 0.5) * h_circ(2, 1)
xi_we(3,2,10) #### 0.4362059564857282
i = 3
k = 2
m = 1
t = 9
(1 - omega) * h_circ(k, (m+t)*Delta)
# def xi_we(i, k, m):
# """Implements the Weighted Erlang Case."""
# # truncate time in service m
# if m >= t_MAX:
# m_new = t_MAX-1
# else:
# m_new = m
# if xi_matrix[i-1][k-1][m]: # retrieve stored value
# pass
# elif i == n: # initial condition
# xi_matrix[i-1][k-1][m] = (1 - omega) * h_circ(k, m*Delta)
# else:
# # initial guess
# if m > 0 and minima[i-1][k-1][m-1]:
# t_guess = minima[i-1][k-1][m-1]
# else:
# t_guess = eval(old_minima[i-1][k-1])[m]
# cost_guess = cost_we(t_guess, i, k, m)
# t_new = t_guess
# # walk to the left
# while True:
# t_new -= 1
# cost_new = cost_we(t_new, i, k, m)
# if cost_new < cost_guess:
# t_guess = t_new
# cost_guess = cost_new
# elif cost_new > cost_guess:
# break
# # walk to the right
# while True:
# t_new += 1
# cost_new = cost_we(t_new, i, k, m)
# if cost_new < cost_guess:
# t_guess = t_new
# cost_guess = cost_new
# elif cost_new > cost_guess:
# break
# xi_matrix[i-1][k-1][m] = cost_guess
# minima[i-1][k-1][m] = t_guess
# print("end",i,k,m,t_guess,cost_guess)
# return xi_matrix[i-1][k-1][m]
def xi_we(i, k, m):
"""Implements the Weighted Erlang Case."""
if m <= t_MAX and xi_matrix[i-1][k-1][m]: # retrieve stored value
pass
elif i == n: # initial condition
if m <= t_MAX:
xi_matrix[i-1][k-1][m] = (1 - omega) * h_circ(k, m*Delta)
else:
return (1 - omega) * h_circ(k, m*Delta)
else:
if m <= t_MAX:
# initial guess
if m > 0 and minima[i-1][k-1][m-1]:
t_guess = minima[i-1][k-1][m-1]
else:
t_guess = eval(old_minima[i-1][k-1])[m]
else:
if minima[i-1][k-1][t_MAX]:
t_guess = minima[i-1][k-1][t_MAX]
else:
t_guess = old_minima[i-1][k-1][t_MAX]
cost_guess = cost_we(t_guess, i, k, m)
t_new = t_guess
# walk to the left
while True:
t_new -= 1
cost_new = cost_we(t_new, i, k, m)
if cost_new < cost_guess:
t_guess = t_new
cost_guess = cost_new
elif cost_new > cost_guess:
break
# walk to the right
while True:
t_new += 1
cost_new = cost_we(t_new, i, k, m)
if cost_new < cost_guess:
t_guess = t_new
cost_guess = cost_new
elif cost_new > cost_guess:
break
if m <= t_MAX:
xi_matrix[i-1][k-1][m] = cost_guess
minima[i-1][k-1][m] = t_guess
else:
return cost_guess
if m <= 2:
print("end",i,k,m,t_guess,cost_guess)
return xi_matrix[i-1][k-1][m]
SCV = 0.6
K, p, mu = SCV_to_params(SCV)
Delta = 0.01
# epsilon = 0.005
t_MAX = int(5/Delta) # int(5/Delta)
n = 5
omega = 0.5
import csv
C_matrix = [[None for k in range(n)] for i in range(n)]
minima = [[None for k in range(n-1)] for i in range(n-1)]
# compute values
for i in range(1,n+1):
for k in range(1,i+1):
Cstar_homexp(i,k,mu=1,omega=omega,n=n,C_matrix=C_matrix)
# # cost
print("\nCost:", C_matrix[0][0])
new_minima = [[[None for m in range(t_MAX+1)] for k in range(n-1)] for i in range(n-1)]
for i in range(n-1):
for k in range(i+1):
new_minima[i][k] = [int(round(minima[i][k],2) / Delta)] * t_MAX * 2
with open(f'SCV_1.00_omega_{omega}_minima.csv','w', newline='') as myfile:
out = csv.writer(myfile)
out.writerows(new_minima)
with open(f'SCV_1.00_omega_{omega:.1f}_minima.csv','r') as csvfile:
reader = csv.reader(csvfile)
old_minima = list(reader)
xi_matrix = [[[None for m in range(t_MAX+1)] for k in range(i+1)] for i in range(n)]
minima = [[[None for m in range(t_MAX+1)] for k in range(i+1)] for i in range(n)]
for i in np.arange(n,0,-1):
for k in range(1,i+1):
print("i =",i,"k =",k)
for m in range(t_MAX+1):
xi_we(i,k,m)
i, k, m = 5, 4, 2
print(xi_we(i,k,m))
print(minima[i-1][k-1][m])
```
We proceed by analyzing the second case, i.e., the hyperexponential case.
### Hyperexponential Distribution
In this case the service times $B_i$ are independent and distributed as $B$, where $B$ equals with probability $p\in [0,1]$ an exponentially distributed random variable with mean $\mu_1^{-1}$, and with probability $1-p$ an exponentially distributed random variable with mean $\mu_{2}^{-1}$. The following recursion can be derived from the thesis.
<div class="alert alert-warning">
<b>Theorem 3.19 (discrete version).</b>
For $i=1,\dots,n-1$, $k=1,\dots,i$, and $m\in\mathbb{N}_0$,
\[
\xi_i(k,m) = \inf_{t\in \mathbb{N}_0}\Bigg(\omega \bar{f}^{\circ}_{k,m\Delta}(t\Delta)
+ (1 - \omega)\bar{h}^{\circ}_{k,m\Delta} + \sum_{\ell=2}^{k}\sum_{j=0}^{t}\bar{q}_{k\ell,mj}(t)\xi_{i+1}(\ell,j)
+ P^{\downarrow}_{k,m\Delta}(t\Delta)\xi_{i+1}(1,0) + P^{\uparrow}_{k,m\Delta}(t\Delta)\xi_{i+1}(k+1,m+t) \Bigg),
\]
whereas, for $k=1,\dots,n$ and $m \in \mathbb{N}_0$,
\[
\xi_n(k,m) = (1 - \omega)\bar{h}^{\circ}_{k,m\Delta}.
\]
</div>
Below is our implementation.
```
### helper functions
# @cache
def gamma(z, u):
if z == 1:
return p * np.exp(-mu1 * u) / B_sf(u)
elif z == 2:
return (1 - p) * np.exp(-mu2 * u) / B_sf(u)
# @cache
def B_sf(t):
return p * np.exp(-mu1 * t) + (1 - p) * np.exp(-mu2 * t) ### gamma_circ
# @cache
def zeta(alpha, t, k):
if not k:
return (np.exp(alpha * t) - 1) / alpha
else:
return ((t ** k) * np.exp(alpha * t) - k * zeta(alpha, t, k-1)) / alpha
# @cache
def rho(t,m,k):
if not k:
return np.exp(-mu2 * t) * (mu1 ** m) / ((mu1 - mu2) ** (m + 1)) * erlang.cdf(t, m+1, scale=1/(mu1 - mu2))
elif not m:
return np.exp(-mu1 * t) * (mu2 ** k) / math.factorial(k) * zeta(mu1-mu2, t, k)
else:
return (mu1 * rho(t, m-1, k) - mu2 * rho(t, m, k-1)) / (mu1 - mu2)
# @cache
def Psi(t,m,k):
if not m:
return erlang.cdf(t, k, scale=1/mu2)
else:
return erlang.cdf(t, m, scale=1/mu1) - mu1 * sum([rho(t, m-1, i) for i in range(k)])
# @cache
def chi(v, t, z, k, l):
"""
Computes P(t-v < Erl(k,mu1) + Erl(l,mu2) < t, Erl(k,mu1) + Erl(l,mu2) + E(1,mu_z) > t),
where Erl(k,mu1) and Erl(l,mu2) are independent.
"""
if z == 1:
if not k and l:
return np.exp(-mu1 * t) * ((mu2) ** l) \
* (zeta(mu1-mu2, t, l-1) - zeta(mu1-mu2, t-v, l-1)) / math.factorial(l-1)
elif k and not l:
return poisson.pmf(k, mu1*t) * binom.sf(0, k, v/t)
else:
return mu2 * (rho(t, k, l-1) - np.exp(-mu1 * v) * rho(t-v, k, l-1))
elif z == 2:
if not k and l:
return poisson.pmf(l, mu2*t) * binom.sf(0, l, v/t)
elif k and not l:
return np.exp(-mu2 * t) * (erlang.cdf(t, k, scale=1/(mu1-mu2)) - erlang.cdf(t-v, k, scale=1/(mu1-mu2))) \
* (mu1 / (mu1 - mu2)) ** k
else:
return mu1 * (rho(t, k-1, l) - np.exp(-mu2 * v) * rho(t-v, k-1, l))
# @cache
def sigma(t, m, k):
if not k:
return t * erlang.cdf(t, m, scale=1/mu1) - (m / mu1) * erlang.cdf(t, m+1, scale=1/mu1)
elif not m:
return t * erlang.cdf(t, k, scale=1/mu2) - (k / mu2) * erlang.cdf(t, k+1, scale=1/mu2)
else:
return (t - k / mu2) * erlang.cdf(t, m, scale=1/mu1) - (m / mu1) * erlang.cdf(t, m+1, scale=1/mu1) \
+ (mu1 / mu2) * sum([(k - i) * rho(t, m-1, i) for i in range(k)])
# @cache
def f_bar(k, z, t):
"""Computes the mean idle time given (N_0, Z_0) = (k,z)."""
if z == 1:
return sum([binom.pmf(m, k-1, p) * sigma(t, m+1, k-1-m) for m in range(k)])
elif z == 2:
return sum([binom.pmf(m, k-1, p) * sigma(t, m, k-m) for m in range(k)])
# @cache
def h_bar(k, z):
"""Computes the mean waiting time given (N_0, Z_0) = (k,z)."""
if k == 1:
return 0
else:
if z == 1:
return (k-2) + (1/mu1)
elif z == 2:
return (k-2) + (1/mu2)
# @cache
def f_circ(k, u, t):
"""Computes the mean idle time given (N_0, B_0) = (k,u)."""
return gamma(1, u) * f_bar(k, 1, t) + gamma(2, u) * f_bar(k, 2, t)
# @cache
def h_circ(k, u):
"""Computes the mean waiting time given (N_0, B_0) = (k,u)."""
return gamma(1, u) * h_bar(k, 1) + gamma(2, u) * h_bar(k, 2)
### transition probabilities
# 1. No client has been served before time t.
# @cache
def P_up(k, u, t):
"""Computes P(N_t- = k | N_0 = k, B_0 = u)."""
return B_sf(u + t) / B_sf(u)
# 2. All clients have been served before time t.
# @cache
def P_down(k, u, t):
"""Computes P(N_t- = 0 | N_0 = k, B_0 = u)."""
return sum([binom.pmf(m, k-1, p) * (Psi(t, m+1, k-1-m) * gamma(1, u) \
+ Psi(t, m, k-m) * gamma(2, u)) for m in range(k)])
# 3. Some (but not all) clients have been served before time t.
# @cache
def q(diff, z, v, t):
"""
Computes P(N_t = l, B_t < v | N_0 = k, Z_0 = z).
Note: diff = k-l.
"""
if z == 1:
return sum([binom.pmf(m, diff, p) * (p * chi(v, t, 1, m+1, diff-m) \
+ (1 - p) * chi(v, t, 2, m+1, diff-m)) for m in range(diff+1)])
elif z == 2:
return sum([binom.pmf(m, diff, p) * (p * chi(v, t, 1, m, diff-m+1) \
+ (1 - p) * chi(v, t, 2, m, diff-m+1)) for m in range(diff+1)])
# @cache
def q_bar(diff, m, j, t):
"""
Approximates P(N_{t*Delta} = l, B_{t*Delta} in d(j*Delta) | N_0 = k, B_0 = m * Delta).
Note: diff = k-l.
"""
lower = min(max(0, (j - 0.5) * Delta), t*Delta)
upper = min(max(0, (j + 0.5) * Delta), t*Delta)
q1_low = q(diff, 1, lower, t*Delta)
q1_upp = q(diff, 1, upper, t*Delta)
q2_low = q(diff, 2, lower, t*Delta)
q2_upp = q(diff, 2, upper, t*Delta)
return gamma(1, m*Delta) * (q1_upp - q1_low) + gamma(2, m*Delta) * (q2_upp - q2_low)
### cost function
# @cache
def cost_he(t, i, k, m):
"""
Computes (approximately) the cost when
t/Delta is the next interarrival time.
"""
cost = omega * f_circ(k, m*Delta, t*Delta) + (1 - omega) * h_circ(k, m*Delta)
cost += P_down(k, m*Delta, t*Delta) * xi_he(i+1, 1, 0) + P_up(k, m*Delta, t*Delta) * xi_he(i+1, k+1, m+t)
for l in range(2, k+1):
for j in range(t+1):
cost_diff = q_bar(k-l, m, j, t) * xi_he(i+1, l, j)
# if cost_diff > 1e-10:
cost += cost_diff
return cost
# k = 2
# np.exp(-mu1 * t) * (mu2 ** k) / math.factorial(k) * zeta(mu1-mu2, t, k)
# (np.exp(-mu1 * t) * (mu2 ** k) / (mu2 - mu1) ** (k+1)) * \
# (1 - sum([np.exp((mu1 - mu2) * t) * ((((mu2 - mu1) * t) ** i) / math.factorial(i)) for i in range(k+1)]))
l = 2
# chi_1[0,l]
np.exp(-mu1 * t) * ((mu2) ** l) \
* (zeta(mu1-mu2, t, l-1) - zeta(mu1-mu2, t-v, l-1)) / math.factorial(l-1)
(np.exp(-mu1 * t) * ((mu2 / (mu2 - mu1)) ** l)) * \
(sum([np.exp(-(mu2-mu1)*(t-v)) * (((mu2 - mu1) * (t - v)) ** i) / math.factorial(i) for i in range(l)]) - \
sum([np.exp(-(mu2-mu1)*t) * (((mu2 - mu1) * t) ** i) / math.factorial(i) for i in range(l)]))
f_circ(k, m*Delta, t*Delta)
h_circ(k, m*Delta)
P_down(k, m*Delta, t*Delta)
xi_he(i+1, 1, 0)
P_up(k, m*Delta, t*Delta)
xi_he(i+1, k+1, m+t)
t = 2
i = 4
k = 2 ### k > 1
m = 0
cost_he(t,i,k,m)
v = 1.3
t = 2.8
z = 2
k = 4
l = 0
q(k-l,z,v,t) ### q hangt alleen af van k-l
q_bar(k-l, v, v, t)
np.exp(-mu2 * t) * ((mu1 ** k) / math.factorial(k-1)) * (zeta(mu2 - mu1, t, k-1) - zeta(mu2 - mu1, t-v, k-1))
SCV = 2
p, mu1, mu2 = SCV_to_params(SCV)
n = 5
v = 0.05
t = 0.10
print(chi(v,t,1,1,0)) ## 0.00776 (klopt)
print(chi(v,t,1,0,1)) ## 0.02081 (FOUT) bij mij 0????
print(chi(v,t,2,0,1)) ## 0.0021 (klopt)
print(chi(v,t,2,1,0)) ## 0.0077 (klopt)
mu2-mu1
l = 1
np.exp(-mu1 * t) * ((mu2 / (mu1 - mu2)) ** l) * \
(
sum([np.exp(-(mu1-mu2)*(t-v)) * (((mu2 - mu1) * (t - v)) ** i) / math.factorial(i) for i in range(l)])) - \
sum([np.exp(-(mu1-mu2)*t) * (((mu2 - mu1) * t) ** i) / math.factorial(i) for i in range(l)]
)
l = 1
np.exp(-mu1 * t) * ((mu2 / (mu2 - mu1)) ** l) * \
(1 - sum([np.exp(-(mu2-mu1)*t) * (((mu2 - mu1) * t) ** i) / math.factorial(i) for i in range(l)])) \
- np.exp(-mu1*(t-v)) * ((mu2 / (mu2 - mu1)) ** l) * \
(1 - sum([np.exp(-(mu2-mu1)*(t-v)) * (((mu2 - mu1) * (t - v)) ** i) / math.factorial(i) for i in range(l)]))
def xi_he(i, k, m):
"""Implements the Hyperexponential Case."""
# truncate time in service m
if m >= t_MAX:
m = t_MAX-1
if xi_matrix[i-1][k-1][m]: # retrieve stored value
pass
elif i == n: # initial condition
xi_matrix[i-1][k-1][m] = (1 - omega) * h_circ(k, m*Delta)
else:
# if m >= 2 and xi_matrix[i-1][k-1][m-1] and xi_matrix[i-1][k-1][m-2]:
# # fill all coming values with current cost & minimum
# if abs(xi_matrix[i-1][k-1][m-1] - xi_matrix[i-1][k-1][m-2]) < epsilon:
# xi_matrix[i-1][k-1][m:] = [xi_matrix[i-1][k-1][m-1]] * (t_MAX - (m - 1))
# minima[i-1][k-1][m:] = [minima[i-1][k-1][m-1]] * (t_MAX - (m - 1))
# print(i,k,m,"break")
# return xi_matrix[i-1][k-1][m]
# initial guess
if m > 0 and minima[i-1][k-1][m-1]:
t_guess = minima[i-1][k-1][m-1]
else:
t_guess = eval(old_minima[i-1][k-1])[m]
cost_guess = cost_he(t_guess, i, k, m)
t_new = t_guess
# walk to the left
while True:
t_new -= 1
cost_new = cost_he(t_new, i, k, m)
if cost_new < cost_guess:
t_guess = t_new
cost_guess = cost_new
elif cost_new > cost_guess:
break
# walk to the right
while True:
t_new += 1
cost_new = cost_he(t_new, i, k, m)
if cost_new < cost_guess:
t_guess = t_new
cost_guess = cost_new
elif cost_new > cost_guess:
break
xi_matrix[i-1][k-1][m] = cost_guess
minima[i-1][k-1][m] = t_guess
if m <= 20:
print("end",i,k,m,t_guess,cost_guess)
return xi_matrix[i-1][k-1][m]
```
With this program, we can obtain dynamic schedules in the hyperexponential case:
```
SCV = 2.5
p, mu1, mu2 = SCV_to_params(SCV)
Delta = 0.01
epsilon = 0.005
t_MAX = int(5/Delta)
n = 5
omega = 0.5
import csv
C_matrix = [[None for k in range(n)] for i in range(n)]
minima = [[None for k in range(n-1)] for i in range(n-1)]
# compute values
for i in range(1,n+1):
for k in range(1,i+1):
Cstar_homexp(i,k,mu=1,omega=omega,n=n,C_matrix=C_matrix)
# # cost
print("\nCost:", C_matrix[0][0])
new_minima = [[[None for m in range(t_MAX)] for k in range(n-1)] for i in range(n-1)]
for i in range(n-1):
for k in range(i+1):
new_minima[i][k] = [int(round(minima[i][k],2) / Delta)] * t_MAX * 2
with open(f'SCV_1.00_omega_{omega}_minima.csv','w', newline='') as myfile:
out = csv.writer(myfile)
out.writerows(new_minima)
with open(f'SCV_1.00_omega_{omega:.1f}_minima.csv','r') as csvfile:
reader = csv.reader(csvfile)
old_minima = list(reader)
xi_matrix = [[[None for m in range(t_MAX)] for k in range(i+1)] for i in range(n)]
minima = [[[None for m in range(t_MAX)] for k in range(i+1)] for i in range(n)]
# i = 3
# k = 1
# # m = 0
# # for k in np.arange(1,5):
# for m in np.arange(3):
# print(i,k,m,xi_he(i,k,m))
xi_matrix = [[[None for m in range(t_MAX)] for k in range(i+1)] for i in range(n)]
minima = [[[None for m in range(t_MAX)] for k in range(i+1)] for i in range(n)]
for i in np.arange(n,0,-1):
for k in range(1,i+1):
print("i =",i,"k =",k)
for m in range(101):
xi_he(i,k,m)
xi_he(1,1,0)
print('Function Summary')
functions = ['gamma', 'B_sf', 'zeta', 'rho', 'Psi', 'chi', 'sigma', 'f_bar', 'h_bar',
'f_circ', 'h_circ', 'P_up', 'P_down', 'q', 'q_bar', 'cost_he']
for function in functions:
info = eval(function).cache_info()
print(f'{str(function):8s}: {info.hits:8d} hits\
{info.misses:8d} misses\
{info.hits/(info.hits + info.misses):.2%} gain')
```
| github_jupyter |
# Circuit Simulation
This tutorial demonstrates how to compute (simulate) the outcome probabilities of circuits in pyGSTi. There are currently two basic ways to to this - but constructing and simulating a `Circuit` object, or by constructing and propagating a state.
## Method 1: `Circuit` simulation
This is the primary way circuit simulation is done in pyGSTi. `Model` objects are statistical models that predict the outcome probabilities of events, and (at least for all current model types) "events" are circuits, described by `Circuit` objects. Thus, the three steps to simulating a circuit using this approach are:
1. create a `Model`
2. create a `Circuit`
3. call `model.probs(circuit)`
Building models and circuits (steps 1 and 2) are largely covered in other tutorials (see the [essential objects tutorial](../01-EssentialObjects.ipynb), [circuits tutorial](../objects/Circuit.ipynb), and [explicit-op model](../objects/ExplicitModel.ipynb) and [implicit-op model](../objects/ImplicitModel.ipynb) tutorials). This section focuses on step 3 and `Model` options which impact the way in which a model computes probabilities. This approach to circuit simulation is most convenient when you have a large number of circuits which are known (and fixed) beforehand.
Let's begin with a simple example, essentially the same as the one in the [using-essential-objects tutorial](../02-Using-Essential-Objects.ipynb):
```
import pygsti
mdl = pygsti.construction.build_explicit_model((0,1),
[(), ('Gxpi2',0), ('Gypi2',0), ('Gxpi2',1), ('Gypi2',1), ('Gcnot',0,1)],
["I(0,1)","X(pi/2,0)", "Y(pi/2,0)", "X(pi/2,1)", "Y(pi/2,1)", "CNOT(0,1)"])
c = pygsti.objects.Circuit([('Gxpi2',0),('Gcnot',0,1),('Gypi2',1)] , line_labels=[0,1])
print(c)
mdl.probs(c) # Compute the outcome probabilities of circuit `c`
```
This example builds an `ExplicitOpModel` (best for 1-2 qubits) on 2 qubits with $X(\pi/2)$ and $Y(\pi/2)$ rotation gates on each qubit and a CNOT gate between them. This model is able to simulate any circuit *layer* (a.k.a. "time-step" or "clock-cycle") that contains any *one* of these gates (this is what it means to be an explicit-op model: the operation for every simulate-able circuit layer must be explicitly supplied to the `Model`). For example, this model cannot simulate a circuit layer where two `Gxpi2` gates occur in parallel:
```
c2 = pygsti.objects.Circuit([ [('Gxpi2',0), ('Gxpi2',1)],('Gcnot',0,1) ] , line_labels=[0,1])
print(c2)
try:
mdl.probs(c2)
except KeyError as e:
print("KEY ERROR (can't simulate this layer): " + str(e))
```
As is detailed in the [implicit-op model tutorial](../objects/ImplicitModel.ipynb), an "implicit-operation" model *is* able to implicitly create layer operations from constituent gates, and thus perform the simulation of `c2`:
```
implicit_mdl = pygsti.construction.build_localnoise_model(2, ('Gxpi2', 'Gypi2', 'Gcnot'))
print(c2)
implicit_mdl.probs(c2)
```
## Method 2: state propagation
In this method of circuit simulation, a state object (a `SPAMVec` in pyGSTi) is propagated circuit-layer by circuit-layer. This method is convenient when a there are few (or just one!) circuit that involves substantial classical logic or needs to be probed at various points in time. It is slower to simulate circuits in this way, as it requires calls more calls between pyGSTi's Python and C routines than method 1 does.
The two cells below show how to perform the same two circuits above using the state-propagation method.
```
#Simulating circuit `c` above using `mdl`: [('Gxpi2',0),('Gcnot',0,1),('Gypi2',1)]
rho = mdl['rho0']
rho = mdl[('Gxpi2',0)].acton(rho)
rho = mdl[('Gcnot',0,1)].acton(rho)
rho = mdl[('Gypi2',1)].acton(rho)
probs = mdl['Mdefault'].acton(rho)
print(probs)
```
Note that, especially for implicit models, the interface is a bit clunky. <font style="color:red">Simulation by state propagation is a work in progress in pyGSTi, and users should expect that this interface may change (improve!) in the future</font>.
```
#Simulating circuit `c2` above using `implicit_mdl`: [ [('Gxpi2',0), ('Gxpi2',1)], ('Gcnot',0,1) ]
from pygsti.objects import Label as L
liz = implicit_mdl._layer_lizard()
rho = liz.get_prep( L('rho0') )
rho = liz.get_operation( L((('Gxpi2',0),('Gxpi2',1))) ).acton(rho)
rho = liz.get_operation( L('Gcnot',(0,1)) ).acton(rho)
probs = liz.povm_blks['layers']['Mdefault'].acton(rho)
print(probs)
```
## Method 3: hybrid
(an addition planned in future releases of pyGSTi)
## Forward-simulation types
PyGSTi refers to the process of computing circuit-outcome probabilities as *forward simulation*, and there are several methods of forward simulation currently available. The default method for 1- and 2-qubit models multiplies together dense process matrices, and is named `"matrix"` (because operations are *matrices*). The default method for 3+ qubit models performs sparse matrix-vector products, and is named `"map"` (because operations are abstract *maps*). A `Model` is constructed for a single type of forward simulation, and it stores this within its `.simtype` member. For more information on using different types of forward simulation see the [forward simulation types tutorial](algorithms/advanced/ForwardSimulationTypes.ipynb).
Here are some examples showing which method is being used and how to switch between them. Usually you don't need to worry about the forward-simulation type, but in the future pyGSTi may have more options for specialized purposes.
```
c3 = pygsti.objects.Circuit([('Gxpi2',0),('Gcnot',0,1)] , line_labels=[0,1])
explicit_mdl = pygsti.construction.build_explicit_model((0,1),
[(), ('Gxpi2',0), ('Gypi2',0), ('Gxpi2',1), ('Gypi2',1), ('Gcnot',0,1)],
["I(0,1)","X(pi/2,0)", "Y(pi/2,0)", "X(pi/2,1)", "Y(pi/2,1)", "CNOT(0,1)"])
print("2Q explicit_mdl will simulate probabilities using the '%s' forward-simulation method." % explicit_mdl.simtype)
explicit_mdl.probs(c3)
implicit_mdl = pygsti.construction.build_localnoise_model(3, ('Gxpi2', 'Gypi2', 'Gcnot'))
print("3Q implicit_mdl will simulate probabilities using the '%s' forward-simulation method." % implicit_mdl.simtype)
implicit_mdl.probs(c)
implicit_mdl.set_simtype('matrix')
print("3Q implicit_mdl will simulate probabilities using the '%s' forward-simulation method." % implicit_mdl.simtype)
implicit_mdl.probs(c)
```
| github_jupyter |
### Easy string manipulation
```
x = 'a string'
y = "a string"
if x == y:
print("they are the same")
fox = "tHe qUICk bROWn fOx."
```
To convert the entire string into upper-case or lower-case, you can use the ``upper()`` or ``lower()`` methods respectively:
```
fox.upper()
fox.lower()
```
A common formatting need is to capitalize just the first letter of each word, or perhaps the first letter of each sentence.
This can be done with the ``title()`` and ``capitalize()`` methods:
```
fox.title()
fox.capitalize()
```
The cases can be swapped using the ``swapcase()`` method:
```
fox.swapcase()
line = ' this is the content '
line.strip()
```
To remove just space to the right or left, use ``rstrip()`` or ``lstrip()`` respectively:
```
line.rstrip()
line.lstrip()
```
To remove characters other than spaces, you can pass the desired character to the ``strip()`` method:
```
num = "000000000000435"
num.strip('0')
line = 'the quick brown fox jumped over a lazy dog'
line.find('fox')
line.index('fox')
line[16:21]
```
The only difference between ``find()`` and ``index()`` is their behavior when the search string is not found; ``find()`` returns ``-1``, while ``index()`` raises a ``ValueError``:
```
line.find('bear')
line.index('bear')
line.partition('fox')
```
The ``rpartition()`` method is similar, but searches from the right of the string.
The ``split()`` method is perhaps more useful; it finds *all* instances of the split-point and returns the substrings in between.
The default is to split on any whitespace, returning a list of the individual words in a string:
```
line_list = line.split()
print(line_list)
print(line_list[1])
```
A related method is ``splitlines()``, which splits on newline characters.
Let's do this with a Haiku, popularly attributed to the 17th-century poet Matsuo Bashō:
```
haiku = """matsushima-ya
aah matsushima-ya
matsushima-ya"""
haiku.splitlines()
```
Note that if you would like to undo a ``split()``, you can use the ``join()`` method, which returns a string built from a splitpoint and an iterable:
```
'--'.join(['1', '2', '3'])
```
A common pattern is to use the special character ``"\n"`` (newline) to join together lines that have been previously split, and recover the input:
```
print("\n".join(['matsushima-ya', 'aah matsushima-ya', 'matsushima-ya']))
pi = 3.14159
str(pi)
print ("The value of pi is " + pi)
```
Pi is a float number so it must be transform to sting.
```
print( "The value of pi is " + str(pi))
```
A more flexible way to do this is to use *format strings*, which are strings with special markers (noted by curly braces) into which string-formatted values will be inserted.
Here is a basic example:
```
"The value of pi is {}".format(pi)
```
### Easy regex manipulation!
```
import re
line = 'the quick brown fox jumped over a lazy dog, fox'
```
With this, we can see that the ``regex.search()`` method operates a lot like ``str.index()`` or ``str.find()``:
```
line.index('fox')
regex = re.compile('fox')
match = regex.search(line)
match.start()
```
Similarly, the ``regex.sub()`` method operates much like ``str.replace()``:
```
line.replace('fox', 'BEAR')
regex.sub('BEAR', line)
```
The following is a table of the repetition markers available for use in regular expressions:
| Character | Description | Example |
|-----------|-------------|---------|
| ``?`` | Match zero or one repetitions of preceding | ``"ab?"`` matches ``"a"`` or ``"ab"`` |
| ``*`` | Match zero or more repetitions of preceding | ``"ab*"`` matches ``"a"``, ``"ab"``, ``"abb"``, ``"abbb"``... |
| ``+`` | Match one or more repetitions of preceding | ``"ab+"`` matches ``"ab"``, ``"abb"``, ``"abbb"``... but not ``"a"`` |
| ``.`` | Any character | ``.*`` matches everything |
| ``{n}`` | Match ``n`` repetitions of preeeding | ``"ab{2}"`` matches ``"abb"`` |
| ``{m,n}`` | Match between ``m`` and ``n`` repetitions of preceding | ``"ab{2,3}"`` matches ``"abb"`` or ``"abbb"`` |
```
bool(re.search(r'ab', "Boabab"))
bool(re.search(r'.*ma.*', "Ala ma kota"))
bool(re.search(r'.*(psa|kota).*', "Ala ma kota"))
bool(re.search(r'.*(psa|kota).*', "Ala ma psa"))
bool(re.search(r'.*(psa|kota).*', "Ala ma chomika"))
zdanie = "Ala ma kota."
wzor = r'.*' #pasuje do każdego zdania
zamiennik = r"Ala ma psa."
re.sub(wzor, zamiennik, zdanie)
wzor = r'(.*)kota.'
zamiennik = r"\1 psa."
re.sub(wzor, zamiennik, zdanie)
wzor = r'(.*)ma(.*)'
zamiennik = r"\1 posiada \2"
re.sub(wzor, zamiennik, zdanie)
```
| github_jupyter |
```
# fundamentals
import os, glob
import numpy as np
import pandas as pd
from calendar import monthrange, month_name
import scipy.stats as stats
import funcs as funcs
import datetime
import imp
# plotting libraries and setup
from matplotlib.colors import BoundaryNorm
import matplotlib.pyplot as plt
%matplotlib inline
plt.rc('font', family='serif')
plt.rc('font', size=12)
plt.rc('facecolor', )
# met mast functions and utilities
import met_funcs as MET
import vis as vis
import utils as utils
# paths (must mount volume smb://nrel.gov/shared/wind/WindWeb/MetData/135mData/)
towerID = 'M5'
metPathLoHz = '/Users/nhamilto/Documents/Wake_Dynamics/SiteChar/data/{}/txt/'.format(towerID)
figPath = '../../figs/{}'.format(towerID)
# time range
years = [ int(a) for a in np.arange(2012,2018,1) ] #
months = [ int(a) for a in np.arange(1,12.1,1) ]
# or just get all?
inputfiles = [] # list of files to be read into metdata object
filecount = 0
for year in years:
for month in months:
fName = glob.glob(os.path.join(metPathLoHz,'{0}_{1}.txt'.format(year,month_name[month])))
if len(fName)>0:
fName = fName[0]
inputfiles.append(fName)
print('{} files to be read into MetData'.format(len(inputfiles)))
## load data from list of input data files
metdat = MET.load_met_data(inputfiles, verbose=False)
## remove columns that are all nans
MET.drop_nan_cols(metdat)
## use qc columns to mask data (qc != 1 --> questionable data)
metdat = MET.qc_mask(metdat)
## flag data by stability class
stabconds, stabcat = MET.flag_stability(metdat)
## group columns based on category, assign units, labels, savenames
varcats, varunits, varlabels, varsave = MET.categorize_fields(metdat, keeplist=True)
## drop columns not in any of the categories, filter TI, temperature, stability parameters
MET.groom_data(metdat, varcats)
filtcols = [col for col in metdat.columns if 'air' not in col.lower() and 'humidity' not in col.lower()]
## Finally, reject outliers more than 5 standard deviations from the mean
for col in metdat.columns:
try:
metdat[col] = MET.reject_outliers(metdat[col], m=6)
except:
continue
catinfo = {}
catinfo['columns'] = varcats
catinfo['units'] = varunits
catinfo['labels'] = varlabels
catinfo['save'] = varsave
# M5 excluded angles,
# Obstruction, start ang. end ang.
# GE 46 146
# Alstom 123 203
# CART-3 172 213
# CART-2 177 212
# Siemens 165 210
# Gamesa 189 228
exclude_angles = [(46,146),(123,203),(172,213),(177,212),(165,210),(189,228)]
tempcol,_,_ = utils.get_vertical_locations(catinfo['columns']['air temperature'])
temperaturedata = metdat[tempcol]
presscol,_,_ = utils.get_vertical_locations(catinfo['columns']['air pressure'])
pressuredata = metdat[presscol]
tempcol[::2]
fig, ax = plt.subplots(2,1, figsize = (8,5), sharex=True)
colors = utils.get_colors(len(presscol), basecolor='blue')
pressuredata.plot.line(ax=ax.flatten()[0], color=colors, legend=False, alpha=0.75)
leg = ax.flatten()[0].legend(presscol, frameon=False, loc=6, bbox_to_anchor=(1,0.5))
ax.flatten()[0].set_ylabel(catinfo['labels']['air pressure'])
colors = utils.get_colors(len(tempcol[::2]), basecolor='red')
temperaturedata[tempcol[::2]].plot.line(ax=ax.flatten()[1], color=colors, legend=False, alpha=0.75)
leg = ax.flatten()[1].legend(tempcol[::2], frameon=False, loc=6, bbox_to_anchor=(1,0.5))
ax.flatten()[1].set_ylabel(catinfo['labels']['air temperature'])
fig.tight_layout()
fig.savefig(os.path.join(figPath,'M5_pressure_v_temperature_timeseries.png'), dpi=200, bbox_inches='tight')
```
# Cut by TI
```
turbclasses = np.linspace(0,50,6)
turbcol, probe_heights, _ = utils.get_vertical_locations(catinfo['columns']['ti'], 87)
metdat['turbclass'] = pd.cut(metdat[turbcol], turbclasses, include_lowest=False).astype(str)
metdat['turbclass'].value_counts()
temp = metdat.groupby('turbclass')
turbclasses = list(temp.groups.keys())[:-1]
lowTI = temp.get_group(turbclasses[0])
plotcats = ['air density',
'air pressure',
'air temperature',
'direction',
'relative humidity',
'speed',
'wind shear',
'wind veer']
lowtifigpath = '../../figs/lowTI'
try:
os.makedirs(lowtifigpath)
except:
pass
```
# Full data histograms
```
catinfo['labels']['direction']
nrelcolors = utils.get_nrelcolors()
for cat in ['direction']:#plotcats:
height = 87
if 'shear' in cat.lower():
height = 110
plotvar, probe_height, _ = utils.get_vertical_locations(catinfo['columns'][cat], height)
fulldat = metdat[plotvar].dropna().sort_values()
fulldat = MET.reject_outliers(fulldat,m=4)
fig, ax = plt.subplots(figsize=(5,3))
binwidth = np.round((fulldat.max()-fulldat.min())/35.0,decimals=3)
bins = np.arange(fulldat.min(), fulldat.max(), binwidth)
fulldat.plot.hist(bins = bins,
color=nrelcolors['blue'][0],
edgecolor='k',
alpha=1,
ax=ax,
weights=np.ones(len(fulldat))/len(fulldat)*100,
legend=False)
ax.set_xlabel(catinfo['labels'][cat])
ax.set_ylabel('Frequency [%]')
fig.savefig(os.path.join(figPath,'M5_{}_hist_comp_{}m.png'.format(catinfo['save'][cat],probe_height)),dpi=200,bbox_inches='tight')
plt.clf()
```
# Low TI histograms comparisons
```
nrelcolors = utils.get_nrelcolors()
for cat in ['direction']:#plotcats:
height = 87
if 'shear' in cat.lower():
height = 110
plotvar, _, _ = utils.get_vertical_locations(catinfo['columns'][cat], height)
fulldat = metdat[plotvar].dropna().sort_values()
fulldat = MET.reject_outliers(fulldat,m=4)
lowtidat = lowTI[plotvar].dropna().sort_values()
lowtidat = MET.reject_outliers(lowtidat,m=4)
result = pd.concat([fulldat, lowtidat], axis=1)
fig, ax = plt.subplots(figsize=(5,3))
binwidth = np.round((fulldat.max()-fulldat.min())/35.0,decimals=3)
bins = np.arange(fulldat.min(), fulldat.max(), binwidth)
fulldat.plot.hist(bins = bins,
color=nrelcolors['blue'][0],
edgecolor='k',
alpha=0.5,
ax=ax,
weights=np.ones(len(fulldat))/len(fulldat)*100,
legend=False)
lowtidat.plot.hist(bins = bins,
color=nrelcolors['red'][0],
edgecolor='k',
alpha=0.5,
ax=ax,
weights=np.ones(len(lowtidat))/len(lowtidat)*100,
legend=False)
ax.set_xlabel(catinfo['labels'][cat])
ax.set_ylabel('Frequency [%]')
leg = ax.legend(['Full Data', 'Low TI'], frameon=False)
fig.savefig(os.path.join(lowtifigpath,'LOWTI_{}_hist_comp.png'.format(catinfo['save'][cat])),dpi=200,bbox_inches='tight')
plt.clf()
turbcol,_,_ = utils.get_vertical_locations(catinfo['columns']['ti'], 87)
nrelcolors= utils.get_nrelcolors()
colors = utils.get_colors(5, basecolor='span')
# test = metdat.groupby([metdat.index.weekofyear,'turbclass'])
test = metdat.groupby([metdat.index.dayofyear,'turbclass'])
test2 = test[turbcol].count().unstack().drop('nan',axis=1).transpose()
test2 = test2/test2.sum()
test2 = test2.transpose()
test2.mean()*100
test2.std()*100
test3 = test2[test2.columns[-1::-1]]
colors = utils.get_colors(5, basecolor='span',reverse=True)
fig, ax = plt.subplots(figsize=(5,3))
for ii,turb in enumerate(turbclasses[-1::-1]):
data = test3[turb].dropna()
plt.hist(data, bins=np.arange(data.min(),data.max(),0.01),
color=colors[ii],
edgecolor='k',
alpha=0.9,
weights= np.ones(len(data))/len(data), density=False)
ax.set_xlabel('Daily Contribution [%]')
ax.set_ylabel('Frequency [%]')
leg = ax.legend(turbclasses[-1::-1], loc=6, bbox_to_anchor = (1,0.5), frameon=False)
leg.set_title(catinfo['labels']['ti'])
# fig.savefig(os.path.join(lowtifigpath,'TI_frequency_hist.png'),dpi=200,bbox_inches='tight')
test3 = test2[test2.columns[-1::-1]]
colors = utils.get_colors(5, basecolor='span',reverse=True)
fig, ax = plt.subplots(figsize=(5,3))
for ii,turb in enumerate(turbclasses[-1::-1]):
data = test3[turb]
ax.hist(data, bins=np.arange(data.min(),data.max(),0.02),
color=colors[ii],
edgecolor='k',
alpha=0.85,
weights= 100*np.ones(len(data))/len(data), density=False)
# data.plot.kde(color=colors[ii], ax=ax)
ax.set_xlim(0,0.65)
ax.set_xlabel('Daily Contribution [%]')
ax.set_ylabel('Frequency [%]')
leg = ax.legend(turbclasses[-1::-1], loc=6, bbox_to_anchor = (1,0.5), frameon=False)
leg.set_title(catinfo['labels']['ti'])
fig.savefig(os.path.join(lowtifigpath,'LOWTI_frequency_hist.png'),dpi=200,bbox_inches='tight')
test3 = test2[test2.columns[-1::-1]]
times = pd.to_datetime(test3.index, format='%j')
pd.DatetimeIndex(times, format='%m-%d')
import matplotlib.dates as mdates
test3 = test2[test2.columns[-1::-1]]
times = pd.to_datetime(test3.index, format='%j')
test3 = test3.set_index(times.format('%m'))
fig, ax = plt.subplots(figsize=(5,3))
ax = test3.plot(x=test3.index, color=colors, ax=ax)
leg = ax.legend(turbclasses[-1::-1], loc=6, bbox_to_anchor=(1,0.5), frameon=False)
leg.set_title(catinfo['labels']['ti'])
ax.set_ylabel('Daily Contribution [%]')
ax.set_xlabel('Day of Year')
ax.format_xdata = mdates.DateFormatter('%m')
# fig.savefig(os.path.join(lowtifigpath,'LOWTI_plot_by_day.png'),dpi=200,bbox_inches='tight')
```
## Low TI figs
```
categories = list(catinfo['columns'].keys())
for cat in ['speed']:#categories:
if 'stability flag' in cat.lower():
continue
# # savepath for new figs
# savecat = catinfo['save'][cat]
# catfigpath = os.makedirs(os.path.join(figPath,savecat), mode=0o777, exist_ok=True)
# catfigpath = os.path.join(figPath,savecat)
# Profiles
## cumulative profile
fig, ax = vis.cumulative_profile(lowTI, catinfo, cat)
# fig.savefig(os.path.join(catfigpath,'{}_{}_profile.png'.format(towerID, savecat)), dpi=200, bbox_inches='tight')
## monthly profile
fig, ax = vis.monthly_profile(lowTI, catinfo, cat)
# fig.savefig(os.path.join(catfigpath,'{}_{}_profile_monthly.png'.format(towerID, savecat)), dpi=200, bbox_inches='tight')
## stability profile
fig,ax = vis.stability_profile(lowTI, catinfo, cat)
# fig.savefig(os.path.join(catfigpath,'{}_{}_profile_stability.png'.format(towerID, savecat)), dpi=200, bbox_inches='tight')
## monthly stability profile
fig,ax = vis.monthly_stability_profiles(lowTI, catinfo, cat)
# fig.savefig(os.path.join(catfigpath,'{}_{}_profile_monthly_stability.png'.format(towerID, savecat)), dpi=200, bbox_inches='tight')
# Diurnal cycle
## cumulative hourly plot
fig,ax = vis.hourlyplot(lowTI, catinfo, cat)
# fig.savefig(os.path.join(catfigpath,'{}_{}_hourly.png'.format(towerID, savecat)), dpi=200, bbox_inches='tight')
## monthly hourly plot
fig,ax = vis.monthlyhourlyplot(lowTI, catinfo, cat)
# fig.savefig(os.path.join(catfigpath,'{}_{}_hourly_monthly.png'.format(towerID, savecat)), dpi=200, bbox_inches='tight')
plt.close('all')
temp = lowTI.copy()
temp = temp.groupby(temp.index.month)
fig,ax = vis.monthlyhourlyplot(lowTI,catinfo,'direction')
fig.savefig(os.path.join(lowtifigpath,'TI_hourly_monthly.png'),dpi=200,bbox_inches='tight')
fig, ax, leg = vis.monthly_rose_fig(lowTI,catinfo,'speed',vertloc=90, bins=[0,3,5,7,12], ylim=12)
fig.savefig(os.path.join(lowtifigpath,'TI_monthly_wind_rose.png'),dpi=200,bbox_inches='tight')
dircol, probe_heights, _ = utils.get_vertical_locations(catinfo['columns']['direction'], 87)
spdcol, probe_heights, _ = utils.get_vertical_locations(catinfo['columns']['speed'], 87)
colors = utils.get_colors(5,basecolor='span')
fig,ax = plt.subplots(figsize=(8,3))
for ii, tclass in enumerate(turbclasses):
test.get_group((1,tclass)).plot.scatter(dircol, spdcol, color=colors[ii], alpha = 0.35, edgecolor='k', ax=ax)
ax.legend(turbclasses)
dircol, probe_heights, _ = utils.get_vertical_locations(catinfo['columns']['direction'], 87)
fig, ax = plt.subplots(figsize=(8,3))
cat = 'gradient richardson'
stabcol, probe_heights, _ = utils.get_vertical_locations(catinfo['columns'][cat], 87)
test.get_group((1,turbclasses[0])).plot.scatter(dircol, stabcol, color=colors[0], alpha = 0.35, edgecolor='k', ax=ax)
ax.set_title(catinfo['labels'][cat])
fig, ax = plt.subplots(figsize=(8,3))
cat = 'stability parameter z/l'
stabcol, probe_heights, _ = utils.get_vertical_locations(catinfo['columns'][cat], 87)
test.get_group((1,turbclasses[0])).plot.scatter(dircol, stabcol, color=colors[0], alpha = 0.35, edgecolor='k', ax=ax)
ax.set_title(catinfo['labels'][cat])
fig, ax = plt.subplots(figsize=(8,3))
cat = 'monin-obukhov length'
stabcol, probe_heights, _ = utils.get_vertical_locations(catinfo['columns'][cat], 87)
test.get_group((1,turbclasses[0])).plot.scatter(dircol, stabcol, color=colors[0], alpha = 0.35, edgecolor='k', ax=ax)
ax.set_title(catinfo['labels'][cat])
```
# Weibull distribution
```
import windrose
import scipy.stats as stats
speedcols, _, _ = utils.get_vertical_locations(catinfo['columns']['speed'])
for plotvar in speedcols[0:1]:
fulldat = metdat[plotvar].dropna()
fulldat = MET.reject_outliers(fulldat,m=4)
lowtidat = lowTI[plotvar].dropna()
lowtidat = MET.reject_outliers(lowtidat,m=4)
binwidth = np.round((lowtidat.max()-lowtidat.min())/35.0,decimals=3)
bins = np.arange(lowtidat.min(), lowtidat.max(), binwidth)
nrecolors = utils.get_nrelcolors()
binwidth = np.round((lowtidat.max()-lowtidat.min())/35.0,decimals=3)
bins = np.arange(lowtidat.min(), lowtidat.max(), binwidth)
fig, ax = plt.subplots(figsize = (5,3))
lowtiparams = stats.exponweib.fit(lowtidat, fc=1)
ax.plot(bins, stats.exponweib.pdf(bins, *lowtiparams), color=nrecolors['red'][0])
# thing,stuff =
output = ax.hist(lowtidat,
bins = bins,
facecolor=nrecolors['red'][0],
edgecolor='k',
alpha=0.3,
normed=True)
fullparams = stats.exponweib.fit(fulldat, fc=1)
pdf = stats.exponweib.pdf(bins, *fullparams)
ax.plot(bins, pdf, color=nrecolors['blue'][0])
fullparams = stats.weibull_min.fit(fulldat)
pdf = stats.weibull_min.pdf(bins, *fullparams)
ax.plot(bins, pdf)
# thing,stuff =
output = ax.hist(fulldat,
bins = bins,
facecolor=nrecolors['blue'][0],
edgecolor='k',
alpha=0.3,
normed=True)
leg = fig.legend()
# fig.savefig(os.path.join(lowtifigpath,'TI_monthly_wind_rose.png'),dpi=200,bbox_inches='tight')
plotvar, _, _ = utils.get_vertical_locations(catinfo['columns']['speed'], height)
fulldat = metdat[plotvar].dropna().sort_values()
fulldat = MET.reject_outliers(fulldat,m=4)
binwidth = np.round((fulldat.max()-fulldat.min())/35.0,decimals=3)
bins = np.arange(fulldat.min(), fulldat.max(), binwidth)
fullparams = stats.rayleigh.fit(fulldat)
plt.plot(bins, stats.rayleigh.pdf(bins, *fullparams), color=nrecolors['blue'][0])
output = plt.hist(fulldat,
bins = bins,
facecolor=nrecolors['blue'][0],
edgecolor='k',
alpha=0.3,
normed=True)
fullparams = stats.weibull_min.fit(lowtidat, floc=1)
pdf = stats.weibull_min.pdf(bins, *fullparams)
plt.plot(bins, pdf, color=nrecolors['blue'][0])
# thing,stuff =
output = plt.hist(lowtidat,
bins = bins,
facecolor=nrecolors['blue'][0],
edgecolor='k',
alpha=0.3,
normed=True)
fullparams
```
| github_jupyter |
```
import os
import sys
from skimage import io, img_as_float, img_as_ubyte
from skimage.util import view_as_windows
import json
from multiprocessing import cpu_count, Pool
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%load_ext autotime
# VISUALIZE SOME SALIENCY MAPS
directory = '../Datasets/AWS-Colours-Quantitative/'
num_imgs = 0
for filename in os.listdir(directory):
path = directory+filename
num_an = filename.split('.')[0].split('_')[-1]
img = img_as_float(io.imread(path))
print(img.dtype, img.shape, img.max(), img.min())
plt.imshow(img, cmap='gray')
plt.title(num_an)
plt.show()
print(path)
num_imgs += 1
if num_imgs == 10: break
print('Total num images',num_imgs)
# DEFINITIONS
def append_df_to_excel(filename, df, sheet_name='Sheet1', startrow=None,
truncate_sheet=False,
**to_excel_kwargs):
"""
Append a DataFrame [df] to existing Excel file [filename]
into [sheet_name] Sheet.
If [filename] doesn't exist, then this function will create it.
Parameters:
filename : File path or existing ExcelWriter
(Example: '/path/to/file.xlsx')
df : dataframe to save to workbook
sheet_name : Name of sheet which will contain DataFrame.
(default: 'Sheet1')
startrow : upper left cell row to dump data frame.
Per default (startrow=None) calculate the last row
in the existing DF and write to the next row...
truncate_sheet : truncate (remove and recreate) [sheet_name]
before writing DataFrame to Excel file
to_excel_kwargs : arguments which will be passed to `DataFrame.to_excel()`
[can be dictionary]
Returns: None
"""
from openpyxl import load_workbook
import pandas as pd
# ignore [engine] parameter if it was passed
if 'engine' in to_excel_kwargs:
to_excel_kwargs.pop('engine')
writer = pd.ExcelWriter(filename, engine='openpyxl')
try:
# try to open an existing workbook
writer.book = load_workbook(filename)
# get the last row in the existing Excel sheet
# if it was not specified explicitly
if startrow is None and sheet_name in writer.book.sheetnames:
startrow = writer.book[sheet_name].max_row
# truncate sheet
if truncate_sheet and sheet_name in writer.book.sheetnames:
# index of [sheet_name] sheet
idx = writer.book.sheetnames.index(sheet_name)
# remove [sheet_name]
writer.book.remove(writer.book.worksheets[idx])
# create an empty sheet [sheet_name] using old index
writer.book.create_sheet(sheet_name, idx)
# copy existing sheets
writer.sheets = {ws.title:ws for ws in writer.book.worksheets}
except FileNotFoundError:
# file does not exist yet, we will create it
pass
if startrow is None:
startrow = 0
# write out the new sheet
df.to_excel(writer, sheet_name, startrow=startrow, **to_excel_kwargs)
# save the workbook
writer.save()
def json_to_dict(path):
with open(path) as json_file:
data = json.load(json_file)
return data
def compute_windows(img, window_size=128, window_stride=64):
windows = view_as_windows(img, window_size, step=window_stride)
num_windows = windows.shape[0]**2
return np.reshape(windows, newshape=(num_windows, windows.shape[-2], windows.shape[-1]) )
def compute_candidates(img, window_size=128, window_stride=64):
windows = compute_windows(img, window_size=window_size, window_stride=window_stride)
#print(windows.shape)
windows = np.array([np.sum(w) for w in windows])
return np.argsort(windows)[::-1], np.sort(windows)[::-1]
def compute_detections(indexs, values, outlier_threshold=-999999, top_anomalies=-1):
mean, std = np.mean(values), np.std(values)
threshold_index = np.where(values >= mean+outlier_threshold*std)[0]
if len(threshold_index) <= 0:
return [], []
#print(threshold_index)
threshold_index = threshold_index[-1]+1
#print(mean, std)
indexs, values = indexs[:threshold_index], values[:threshold_index]
if top_anomalies >= 0: return indexs[:top_anomalies], values[:top_anomalies]
return indexs, values
def check_detections(center, detected_indexs, window_size, num_windows, sliding_window_stride, center_tolerance=0):
assert center_tolerance >= 0,'Center tolerance must be >= 0'
if len(detected_indexs) <= 0: return 0
anomaly_detected = False
i = 0
if center_tolerance == 0:
while not anomaly_detected and i < len(detected_indexs):
index_candidate = detected_indexs[i]
start_row, start_col = int(index_candidate/num_windows)*sliding_window_stride, (index_candidate % num_windows)*sliding_window_stride
end_row, end_col = start_row+window_size, start_col+window_size
anomaly_detected = (start_row <= center[0] <= end_row) and (start_col <= center[1] <= end_col)
#print(center)
#print('Rows:',start_row, end_row)
#print('Cols:',start_col, end_col)
i += 1
else:
raise NotImplementedError
return int(anomaly_detected)
def evaluate_all_images(dataset_type, saliency_type, window_size = 128, sliding_window_stride = 64, top=1, outlier_threshold=-999999):
directory = '../Datasets/'+saliency_type+'-'+dataset_type+'-Quantitative/'
json_path = '../JSONs/Bark-'+dataset_type+'-Quantitative.json'
centers = json_to_dict(json_path)
num_windows = int( (image_size-window_size)/sliding_window_stride )+1
num_imgs = 0
positives = 0
for filename in os.listdir(directory):
path = directory+filename
num_an = filename.split('.')[0].split('_')[-1]
img = img_as_float(io.imread(path))
indexs, values = compute_candidates(img, window_size=window_size, window_stride=sliding_window_stride)
indexs, values = compute_detections(indexs, values, outlier_threshold=outlier_threshold, top_anomalies=top)
detected = check_detections(centers[str(num_an)], indexs, window_size, num_windows, sliding_window_stride)
positives += detected
num_imgs += 1
#break
return positives*100 / num_imgs
def compute_accuracy(threshold):
if threshold < 0: threshold_label = 'All'
else: threshold_label = (str(threshold))[:3]
accuracies = {}
for n in range(1, max_top):
accuracy = evaluate_all_images(dataset_type, saliency_type, window_size, sliding_window_stride, top=n, outlier_threshold=threshold)
accuracies[columns[n-1]] = accuracy
#print('Top',n,'accuracy:',accuracy)
if accuracy >= 100: break
return saliency_type+'-'+str(window_size)+'-'+str(sliding_window_stride)+'-'+threshold_label, pd.Series(accuracies)
image_size = 256
img_index = 474
window_size = 128
sliding_window_stride = 64
num_windows = int( (image_size-window_size)/sliding_window_stride )+1
json_path = '../JSONs/Bark-Colours-Quantitative.json'
centers = json_to_dict(json_path)
img = img_as_float(io.imread('../Datasets/AWS-Colours-Quantitative/an_474.jpg'))
indexs, values = compute_candidates(img, window_size=window_size, window_stride=sliding_window_stride)
indexs, values = compute_detections(indexs, values, outlier_threshold=-np.inf, top_anomalies=1)
print(indexs)
print(check_detections(centers[str(img_index)], indexs, window_size, num_windows, sliding_window_stride))
print( evaluate_all_images('Colours', 'AWS') )
#-----------------------------------#
dataset_type = 'Colours'
saliency_type = 'AWS'
image_size = 256
window_size = 128
sliding_window_stride = 64
#-----------------------------------#
num_windows = int( (image_size-window_size)/sliding_window_stride )+1
max_top = min(50, num_windows**2) + 1
columns = ['Top '+str(n) for n in range(1, 51)]
df = pd.DataFrame(columns=columns)
thresholds = np.insert(np.arange(0.0, 3.1, 0.1), 0, -np.inf)
print('Available cores:',cpu_count())
pool = Pool(4)
thresholds = pool.map(compute_accuracy, thresholds)
#accuracy_computer = functools.partial(compute_accuracy, anomalies_results=anomalies_results, max_top=max_top, centers=centers, columns=columns)
#thresholds = pool.map(accuracy_computer, thresholds)
pool.close()
pool.join()
for model_name, series in thresholds:
df.loc[model_name] = series
print(df.head(3))
# SAVE DATAFRAME TO DRIVE EXCEL
excel_path = '../SALIENCY-RESULTS/'+dataset_type+'_Accuracy_results.xlsx'
print(excel_path)
append_df_to_excel(excel_path, df)
image_size = 256
for dt_type in ['External']:
global dataset_type
dataset_type = dt_type
for s_type in ['AWS', 'WMAP']:
global saliency_type
saliency_type = s_type
for ws in [128, 64]:
global window_size
window_size = ws
for i in [2,4]:
global sliding_window_stride
sliding_window_stride = int(window_size/i)
num_windows = int( (image_size-window_size)/sliding_window_stride )+1
max_top = min(50, num_windows**2) + 1
columns = ['Top '+str(n) for n in range(1, 51)]
df = pd.DataFrame(columns=columns)
thresholds = np.insert(np.arange(0.0, 3.1, 0.1), 0, -np.inf)
print('Available cores:',cpu_count())
pool = Pool(4)
thresholds = pool.map(compute_accuracy, thresholds)
#accuracy_computer = functools.partial(compute_accuracy, anomalies_results=anomalies_results, max_top=max_top, centers=centers, columns=columns)
#thresholds = pool.map(accuracy_computer, thresholds)
pool.close()
pool.join()
for model_name, series in thresholds:
df.loc[model_name] = series
print(df.head(3))
# SAVE DATAFRAME TO DRIVE EXCEL
excel_path = '../SALIENCY-RESULTS/'+dataset_type+'_Rectified_Accuracy_results.xlsx'
print(excel_path)
append_df_to_excel(excel_path, df)
def evaluate_all_clear_images(dataset_type, saliency_type, window_size = 128, sliding_window_stride = 64, top=1, outlier_threshold=-999999):
directory = '../Datasets/'+saliency_type+'-'+dataset_type+'-Quantitative/'
json_path = '../JSONs/Bark-'+dataset_type+'-Quantitative.json'
centers = json_to_dict(json_path)
num_windows = int( (image_size-window_size)/sliding_window_stride )+1
num_imgs = 0
positives = 0
for filename in os.listdir(directory):
path = directory+filename
num_an = filename.split('.')[0].split('_')[-1]
img = img_as_float(io.imread(path))
indexs, values = compute_candidates(img, window_size=window_size, window_stride=sliding_window_stride)
indexs, values = compute_detections(indexs, values, outlier_threshold=outlier_threshold, top_anomalies=top)
detected = len(indexs) != 0
positives += detected
num_imgs += 1
#break
return positives*100 / num_imgs
def compute_clear_accuracy(threshold):
if threshold < 0: threshold_label = 'All'
else: threshold_label = (str(threshold))[:3]
accuracies = {}
for n in range(1, max_top):
accuracy = evaluate_all_clear_images(dataset_type, saliency_type, window_size, sliding_window_stride, top=n, outlier_threshold=threshold)
accuracies[columns[n-1]] = accuracy
#print('Top',n,'accuracy:',accuracy)
if accuracy >= 100: break
return saliency_type+'-'+str(window_size)+'-'+str(sliding_window_stride)+'-'+threshold_label, pd.Series(accuracies)
image_size = 256
for dt_type in ['Clear']:
global dataset_type
dataset_type = dt_type
for s_type in ['AWS', 'WMAP']:
global saliency_type
saliency_type = s_type
for ws in [128, 64]:
global window_size
window_size = ws
for i in [4,2]:
global sliding_window_stride
sliding_window_stride = int(window_size/i)
num_windows = int( (image_size-window_size)/sliding_window_stride )+1
max_top = min(50, num_windows**2) + 1
columns = ['Top '+str(n) for n in range(1, 51)]
df = pd.DataFrame(columns=columns)
thresholds = np.insert(np.arange(0.0, 3.1, 0.1), 0, -np.inf)
print('Available cores:',cpu_count())
pool = Pool(4)
thresholds = pool.map(compute_clear_accuracy, thresholds)
#accuracy_computer = functools.partial(compute_accuracy, anomalies_results=anomalies_results, max_top=max_top, centers=centers, columns=columns)
#thresholds = pool.map(accuracy_computer, thresholds)
pool.close()
pool.join()
for model_name, series in thresholds:
df.loc[model_name] = series
print(df.head(3))
# SAVE DATAFRAME TO DRIVE EXCEL
excel_path = '../SALIENCY-RESULTS/'+dataset_type+'_Rectified_Accuracy_results.xlsx'
print(excel_path)
append_df_to_excel(excel_path, df)
```
| github_jupyter |
# Create a circuit to generate any two-qubit quantum state in Qiskit
Build a general 2-qubit circuit that could output all Hilbert space of states by tuning its parameters.
```
from qiskit import *
import numpy as np
def state_maker(theta, ang0, ang1):
circ = QuantumCircuit(2,2)
circ.u3(theta, 0, 0, 0)
circ.cx(0, 1)
circ.u3(*ang1, 1)
circ.u3(*ang0, 0)
return circ
def get_ensemble(theta0, theta1, theta2, N=1024):
circuit = state_maker(theta0, [theta1,0,0], [theta2,0,0])
circuit.measure(0,0)
circuit.measure(1,1)
simulator = Aer.get_backend('qasm_simulator')
result = execute(circuit, backend = simulator, shots = N).result()
counts = result.get_counts()
return counts
from qiskit.tools.visualization import plot_histogram
#angi = [theta, phi, lam]
ang0 = [0,0,0]
ang1 = [0,0,0]
theta = 0
circ = state_maker(theta, ang0, ang1)
%matplotlib inline
#circ.draw(output='mpl')
circ.measure(0,0)
circ.measure(1,1)
circ.draw(output='mpl')
```
Example of the count result for some parameters $\theta_0$, $\theta_1$ and $\theta_2$.
```
from ttq.optimizer import optimize
pi = np.pi
_EXPECTED_VALUES = {
'00' : 0.5,
'01' : 0.2,
'10' : 0.2,
'11' : 0.1
}
_MAX_ERROR = 0.05
_N = 1024
_STEP = 0.1
_PARAMS = 3
theta0, theta1, theta2 = optimize(conf = {
'bound': [0, 2 * pi],
'expected_values': _EXPECTED_VALUES,
'max_error': _MAX_ERROR,
'max_iter': None,
'n_states': _N,
'step': _STEP,
'x0': [0] * _PARAMS
})
counts = get_ensemble(theta0, theta1, theta2)
print(counts)
plot_histogram(counts)
```
Example of the generated state result for some parameters $\theta_0$, $\theta_1$ and $\theta_2$.
```
circ = state_maker(theta0, [theta1, 0, 0], [theta2, 0, 0])
simulator = Aer.get_backend('statevector_simulator')
result = execute(circ, backend = simulator).result()
statevector = result.get_statevector()
print(statevector)
```
# For some $\theta$'s
Plot the probability of measuring each state for a given set of parameters.
```
ntheta = 100
N = 1024
theta = np.linspace(0, 2*np.pi, ntheta)
prob00, prob01, prob10, prob11 = [], [], [], []
for t in theta: # to check it we only change one parameter
counts = get_ensemble(t, t, t, N)
prob00.append(counts['00']/N if '00' in counts.keys() else 0)
prob01.append(counts['01']/N if '01' in counts.keys() else 0)
prob10.append(counts['10']/N if '10' in counts.keys() else 0)
prob11.append(counts['11']/N if '11' in counts.keys() else 0)
import matplotlib.pyplot as plt
plt.plot(theta, prob00, label='| 00 >')
plt.plot(theta, prob01, label='| 01 >')
plt.plot(theta, prob10, label='| 10 >')
plt.plot(theta, prob11, label='| 11 >')
plt.legend(loc = 'upper right')
plt.show()
```
# Measuring the 'entanglement'
We measure the realtion between the amplidudes of states $| 00 >$ and $| 01 >$ for different $\theta_0$'s.
```
import matplotlib.pyplot as plt
ang0 = [0,0,0]
ang1 = [0,0,0]
entang = []
e00 = []
e11 = []
thetas = np.linspace(0, 2*np.pi, 10)
thetas = thetas[2:-2]
for theta in thetas:
circ = state_maker(theta, ang0, ang1)
simulator = Aer.get_backend('statevector_simulator')
result = execute(circ, backend = simulator).result()
statevector = result.get_statevector()
print('theta = {:2.2f}pi '.format(theta/np.pi) )
print('state = ', statevector)
print()
entang.append(abs(statevector[0])/(abs(statevector[3])+.0001))
plt.plot(thetas, entang)
```
| github_jupyter |
# DiscRimNN
### By Willie Maddox
## Problem A
Create a model that reads a sequence of numbers and classifies the last number in the sequence based on the previous numbers in the sequence.
## Problem B
Create a model that reads a sequence of numbers, one number at a time, and classify that number based on all previously seen numbers.
## Build a signal generator
The signal generator builds waves using the standard form:
$$x(t) = h + A\sin\left(\frac{2\pi t}{T} + \phi\right)$$
where $h$ is the height (vertical offset), $A$ is the amplitude (vertical scale), $T$ is the period (horizontal scale), and $\phi$ is the phase (horizontal offset). An optional $\Delta$ value can also be assigned to each of these 4 variables. These variables give us fine grained control over how we construct our waves and the $\Delta$ values allow us to introduce randomness in the variables.
```
from discrimnn.signal import MixedSignal
```
# Trivial Case: Offset (-3, 0, 3)
<img src="out/offset_30/signals.png" width="1000">
<img src="out/offset_30/mixedsignal.png" width="1000">
<img src="out/offset_30/mixedsignal_with_truth.png" width="1000">
```
# start off with simplest case for proof of concept
time_coeffs = {'start': 0, 'stop': 75, 'n_timestamps': 301, 'n_timesteps': 10}
sig1_coeffs = {'name': 'A', 'offset': {'mean': -3}, 'color': '#ff0000'}
sig2_coeffs = {'name': 'B', 'offset': {'mean': 0}, 'color': '#00ff00'}
sig3_coeffs = {'name': 'C', 'offset': {'mean': 3}, 'color': '#0000ff'}
sig_coeffs = [sig1_coeffs, sig2_coeffs, sig3_coeffs]
msig_coeffs = {'phase': {'mean': 0, 'delta': np.pi}, 'period': {'mean': 25}}
msig = MixedSignal(time_coeffs, sig_coeffs, msig_coeffs, method='sliding')
```
<img src="out/offset_30/mixedsignal_table.png" width="300">
## Next define the RNN
```python
model = Sequential()
model.add(LSTM(32, batch_input_shape=(4, 10, 1), stateful=True, return_sequences=True))
model.add(LSTM(32, stateful=True))
model.add(Dense(n_signals, activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='rmsprop', metrics=['accuracy'])
model.summary()
```
<img src="out/offset_30/model_plot.png" width="400">
## Now train the model
```python
n_generations = 20
for i in range(n_generations):
X, y = msig.generate()
history = model.fit(X, y, epochs=1, batch_size=batch_size, verbose=1, shuffle=False)
model.reset_states()
```
## Results (training loss, accuracy, and test)
<img src="out/offset_30/loss_acc.png" width="1000">
<img src="out/offset_30/eval_pred.png" width="1000">
# Offset (-0.5, 0, 0.5)
<img src="out/offset_050/signals.png" width="1000">
<img src="out/offset_050/mixedsignal.png" width="1000">
<img src="out/offset_050/mixedsignal_with_truth.png" width="1000">
# Offset (-0.5, 0, 0.5) - Results
<img src="out/offset_050/loss_acc.png" width="1000">
<img src="out/offset_050/eval_pred.png" width="1000">
# Offset (-0.1, 0, 0.1)
<img src="out/offset_010/signals.png" width="1000">
<img src="out/offset_010/mixedsignal.png" width="1000">
<img src="out/offset_010/mixedsignal_with_truth.png" width="1000">
# Offset (-0.1, 0, 0.1) - Results
<img src="out/offset_010/loss_acc.png" width="1000">
<img src="out/offset_010/eval_pred.png" width="1000">
# Phase (0, $\pi$, 0)
<img src="out/phase_010/signals.png" width="1000">
<img src="out/phase_010/mixedsignal.png" width="1000">
<img src="out/phase_010/mixedsignal_with_truth.png" width="1000">
# Phase (0, $\pi$, 0) - Results
<img src="out/phase_010/loss_acc.png" width="1000">
<img src="out/phase_010/eval_pred.png" width="1000">
# Summary
1. Full control over dataset
2. Dataset is infinite
3. Model never sees the same sample twice.
4. Can study all three RNN base problems (classification, prediction, forcasting).
5. Great way to zero in on and study a particular aspect of RNN's
# Future work (TODO list)
- [x] create single signal generator class
- [x] create mixed signal generator class
- [ ] create signal noise functions (Gaussian, OU, etc.)
- [ ] create timestep noise functions
- [x] add legends to plots.
- [ ] during training, save outlier X, y train sets to file for further analysis.
- [x] save configuration of mixed signal properties as json for housekeeping.
- [ ] make plots of the mixed signal with colors mapped to hidden layers, lstm states, etc.
- [ ] unit tests for signal.py
- [ ] create startup.py to handle project directories and other goodies.
- [ ] fix savefig clipping the bottoms of our figures.
| github_jupyter |
# First fitting from amalgams
In this phase, we are not considering sequences, leave alone syntax trees, in prediction. Instead we are using the frequency of (shallow) occurence of names in types to predict the (shallow) occurence in definitions.
Here we consider the first two models. The second has some depth and shows overfitting.
## Structure of the models
Both the models have a similar structure.
* there is a common representation of the input data.
* a prediction is made from this of a component the output name distribution (we call this the _low rank prediction_).
* the other component is the input scaled, i.e., it is assumed that elements in the statement are in the proofs.
- this should be rectified, currently the scaling is uniform, depending on the amalgams. It should depend on the specific elements.
* the scaling is also determined from the representation (not too good as mentioned)
* the components are put together.
```
import amalgam_predict as pred
import keras
from matplotlib import pyplot as plt
```
We fit the first model.
* The fit is reasonable.
* More importantly, the validation data fits almost as well as the training data.
```
hist1 = pred.fit(1024, pred.model1)
plt.rcParams['figure.figsize'] = [20, 15]
plt.plot(hist1.history['kullback_leibler_divergence'])
plt.plot(hist1.history['val_kullback_leibler_divergence'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'val'], loc='upper left')
plt.show()
```
As we see, the final KL-divergence is `2.7425` for the training data, and `2.8078` for the validation data.
We now fit the second model. As mentioned, this fits much better, but that is clearly a case of overfitting.
```
hist2 = pred.fit(1024, pred.model2)
plt.plot(hist2.history['kullback_leibler_divergence'])
plt.plot(hist2.history['val_kullback_leibler_divergence'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'val'], loc='upper left')
plt.show()
```
We see the fit keeps improving (this was before the early stop), reaching `1.2060`, but the validation error flattens, ending at `2.4163`
To do:
* use better model for persistence
After adding the dropout layer, we get a similar validation fit without the overfitting.
```
hist3 = pred.fit(1024, pred.model3)
plt.plot(hist3.history['kullback_leibler_divergence'])
plt.plot(hist3.history['val_kullback_leibler_divergence'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'val'], loc='upper left')
plt.show()
```
The above shows really no overfitting. So one can try to improve the model.
### Better persistence model (TODO)
* Have a variable which is a row by which to scale each term by pointwise multiplication.
* Initialize from data.
* Multiply and then apply sigmoid to get a probability distribution on terms.
* Use this instead of the input when mixing in.
```
pred.data.keys()
pred.data['types']
import numpy as np
def count_matrix(pairs, dim):
vec = np.zeros((dim, ), np.float32)
for d in pairs:
name = d['name']
count = d['count']
vec[pred.indices[name]] = count
return vec
term_count = count_matrix(pred.data['terms'], pred.dim)
term_count
np.sum(term_count)
```
| github_jupyter |
```
#pip install covidcast
import covidcast
from datetime import date
import pandas as pd
import pathlib
covidcast.metadata()
```
## Data to Use
Covidcast:
* SafeGraph (social distancing metrics, weekly patterns) - https://cmu-delphi.github.io/delphi-epidata/api/covidcast-signals/safegraph.html - may want to look at data prior to pandemic as well since the data goes back all the way to January 1st, 2019.
* NCHS (CDC) mortality data - https://cmu-delphi.github.io/delphi-epidata/api/covidcast-signals/nchs-mortality.html - this data is different from the death data from USAFacts and JHU because in this data, deaths are reported by the date they occur, not the date they were reported. Also, see details regarding "Missingness" since some data was removed when death counts were 50% less of the expected number.
* USAFacts Cases and Deaths - https://cmu-delphi.github.io/delphi-epidata/api/covidcast-signals/usa-facts.html - I plan to use this priimarily for the case count.
IPUMS USA:
Will not use this since there's no 2020 data.
U.S. Bureau of Labor Statistics: https://www.bls.gov/
Use for employment and salary data.
```
data = covidcast.signal("usa-facts", "confirmed_cumulative_num", date(2020, 1, 25), date(2020, 12, 31))
data.head()
data.tail()
#nchs_new_death_data = covidcast.signal("nchs-mortality", "deaths_covid_incidence_num")
#covidcast.signal("jhu-csse", "")
usa_facts_data = covidcast.signal("usa-facts", "confirmed_cumulative_prop", date(2020, 1, 25), date(2020, 12, 31), geo_type = "state")
usa_facts_data.head()
usa_facts_data.tail()
kerstin_folder_path = pathlib.Path.cwd()
# This worked, but not like how I wanted. There's just too much extra stuff.
usa_facts_data.to_csv(kerstin_folder_path/"usafacts_cum_cases_pop_state.csv")
```
Notice the many rows in the below dataframe. Ignore the index because that only matches with the state name, so the index repeats.
```
#usafacts_cum_cases_pop_state = usa_facts_data[['geo_value','time_value','value']]
#usafacts_cum_cases_pop_state
usafacts_il_county_cum_num = covidcast.signal("usa-facts", "confirmed_cumulative_prop", date(2020, 1, 25), date(2020, 12, 31), geo_type = "county", geo_values = "il")
usafacts_il_county_cum_num
usafacts_county_cum_prop = covidcast.signal("usa-facts", "confirmed_cumulative_prop", date(2020, 1, 25), date(2020, 12, 31), geo_type = "county")
# As seen by the warnings, there is no data for Feb 13-19
usafacts_county_cum_prop
usafacts_il_county_cum_7day_prop = covidcast.signal("usa-facts", "confirmed_7day_cumulative_prop", date(2020, 1, 25), date(2020, 12, 31), geo_type = "county")
usafacts_il_county_cum_7day_prop
```
| github_jupyter |
### Deutsch Jozsa Algorithm!
We are given an oracle that implements either a constant function or a balanced function.
With just one query, we can find out which one it is.
---
Done as part of the NPTEL Course - Introduction to Quantum Computing: Quantum Algorithms and Qiskit
https://onlinecourses.nptel.ac.in/noc21_cs103/preview
```
# Importingstandard Qiskit libraries
from qiskit import QuantumCircuit, execute, Aer, IBMQ
from qiskit.compiler import transpile, assemble
from qiskit.tools.jupyter import *
from qiskit.visualization import plot_histogram
from ibm_quantum_widgets import *
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit
from numpy import pi
# Loading the IBM Q Account
provider = IBMQ.load_account()
print("Process Complete!")
"""
f is constant, f(x) = 0
There's only 1 way of making a constant function such that f(x) = 0
Important!
While giving an arbitrary input like 011, apply the NOT gate to to q1 and q2.
Then, before taking the final measurement, apply the NOT gate to q1 and q2 again for it again.
"""
qreg_q = QuantumRegister(4, 'q')
creg_c = ClassicalRegister(4, 'c')
circuit = QuantumCircuit(qreg_q, creg_c)
"""
Making the input |0001>
"""
circuit.x(qreg_q[3])
circuit.barrier(qreg_q[3], qreg_q[0], qreg_q[1], qreg_q[2])
"""
Step 1 - Hadamard Transform
Setting the state to |+++->
"""
circuit.h(qreg_q[0])
circuit.h(qreg_q[1])
circuit.h(qreg_q[2])
circuit.h(qreg_q[3])
circuit.barrier(qreg_q[3], qreg_q[0], qreg_q[1], qreg_q[2])
"""
Step 2 - Implementation of the Oracle
Constant function, gives 0
"""
None
circuit.barrier(qreg_q[3], qreg_q[0], qreg_q[1], qreg_q[2])
"""
Step 3 - Hadamard on Input Qubits
"""
circuit.h(qreg_q[0])
circuit.h(qreg_q[1])
circuit.h(qreg_q[2])
circuit.barrier()
"""
Step 4 - Measure the first n qubits
"""
circuit.measure(qreg_q[0], creg_c[0])
circuit.measure(qreg_q[1], creg_c[1])
circuit.measure(qreg_q[2], creg_c[2])
circuit.draw()
backend = Aer.get_backend('qasm_simulator')
job_simulator = execute(circuit, backend, shots = 1024)
results_simulator = job_simulator.result()
counts = results_simulator.get_counts(circuit)
print("Counts:", counts)
plot_histogram(counts)
"""
f is constant, f(x) = 1
There's only 1 way of making a constant function such that f(x) = 1
"""
qreg_q = QuantumRegister(4, 'q')
creg_c = ClassicalRegister(4, 'c')
circuit = QuantumCircuit(qreg_q, creg_c)
"""
Making the input |0001>
"""
circuit.x(qreg_q[3])
circuit.barrier()
"""
Step 1 - Hadamard Transform
Setting the state to |+++->
"""
circuit.h(qreg_q[0])
circuit.h(qreg_q[1])
circuit.h(qreg_q[2])
circuit.h(qreg_q[3])
circuit.barrier()
"""
Step 2 - Implementation of the Oracle
Constant function, gives 1
"""
circuit.x(qreg_q[3])
circuit.barrier()
"""
Step 3 - Hadamard on Input Qubits
"""
circuit.h(qreg_q[0])
circuit.h(qreg_q[1])
circuit.h(qreg_q[2])
circuit.barrier()
"""
Step 4 - Measure the first n qubits
"""
circuit.measure(qreg_q[0], creg_c[0])
circuit.measure(qreg_q[1], creg_c[1])
circuit.measure(qreg_q[2], creg_c[2])
circuit.draw()
backend = Aer.get_backend('qasm_simulator')
job_simulator = execute(circuit, backend, shots = 1024)
results_simulator = job_simulator.result()
counts = results_simulator.get_counts(circuit)
print("Counts:", counts)
plot_histogram(counts)
"""
f is balanced
There are many ways of implementing a balanced function.
One way is by using consecutive CNOTs
"""
qreg_q = QuantumRegister(4, 'q')
creg_c = ClassicalRegister(4, 'c')
circuit = QuantumCircuit(qreg_q, creg_c)
"""
Making the input |0001>
"""
circuit.x(qreg_q[3])
circuit.barrier()
"""
Step 1 - Hadamard Transform
Setting the state to |+++->
"""
circuit.h(qreg_q[0])
circuit.h(qreg_q[1])
circuit.h(qreg_q[2])
circuit.h(qreg_q[3])
circuit.barrier()
"""
Step 2 - Implementation of the Oracle
Balanced function with CNOT
"""
circuit.cx(qreg_q[0], qreg_q[3])
circuit.cx(qreg_q[1], qreg_q[3])
circuit.cx(qreg_q[2], qreg_q[3])
circuit.barrier()
"""
Step 3 - Hadamard on Input Qubits
"""
circuit.h(qreg_q[0])
circuit.h(qreg_q[1])
circuit.h(qreg_q[2])
circuit.barrier()
"""
Step 4 - Measure the first n qubits
"""
circuit.measure(qreg_q[0], creg_c[0])
circuit.measure(qreg_q[1], creg_c[1])
circuit.measure(qreg_q[2], creg_c[2])
circuit.draw()
backend = Aer.get_backend('qasm_simulator')
job_simulator = execute(circuit, backend, shots = 1024)
results_simulator = job_simulator.result()
counts = results_simulator.get_counts(circuit)
print("Counts:", counts)
plot_histogram(counts)
"""
f is balanced
There are many ways of implementing a balanced function.
One way is by using consecutive CNOTs.
We can use any number of NOTs as well in this oracle
"""
qreg_q = QuantumRegister(4, 'q')
creg_c = ClassicalRegister(4, 'c')
circuit = QuantumCircuit(qreg_q, creg_c)
"""
Making the input |0001>
"""
circuit.x(qreg_q[3])
circuit.barrier()
"""
Step 1 - Hadamard Transform
Setting the state to |+++->
"""
circuit.h(qreg_q[0])
circuit.h(qreg_q[1])
circuit.h(qreg_q[2])
circuit.h(qreg_q[3])
circuit.barrier()
"""
Step 2 - Implementation of the Oracle
Balanced function with CNOTs and NOTs
"""
circuit.x(qreg_q[0])
circuit.cx(qreg_q[0], qreg_q[3])
circuit.x(qreg_q[1])
circuit.cx(qreg_q[1], qreg_q[3])
circuit.x(qreg_q[2])
circuit.cx(qreg_q[2], qreg_q[3])
circuit.barrier()
"""
Step 3 - Hadamard on Input Qubits
"""
circuit.h(qreg_q[0])
circuit.h(qreg_q[1])
circuit.h(qreg_q[2])
circuit.barrier()
"""
Step 4 - Measure the first n qubits
"""
circuit.measure(qreg_q[0], creg_c[0])
circuit.measure(qreg_q[1], creg_c[1])
circuit.measure(qreg_q[2], creg_c[2])
circuit.draw()
backend = Aer.get_backend('qasm_simulator')
job_simulator = execute(circuit, backend, shots = 1024)
results_simulator = job_simulator.result()
counts = results_simulator.get_counts(circuit)
print("Counts:", counts)
plot_histogram(counts)
```
| github_jupyter |

# Magnetometer Calibration
Kevin Walchko, Phd
30 May 2020
---
To calibrate a magnetometer, you need to get readings from all directions in 3D space. Ideally, when you plot the readings out, you should get a perfect sphere centered at (0,0,0). However, due to misalignments, offset, etc ... you end up with ellipsoids centered at some biased location.
Here we are going to try and get enough readings to estimate these errors and properly calibrate the sensor. We will load in a pre-recorded data set, where the sensor was tumbled around and calibrate it.
## Errors

- **Soft iron errors:** caused by distortion of the Earth's magnetic field due to materials in the environment. Think of it like electricity - the magnetic field is looking for the easiest path to get to where it is going. Since magnetic fields can flow more easily through ferromagnetic materials than air, more of the field will flow through the ferromagnetic material than you would expect if it were just air. This distortion effect causes the magnetic field lines to be bent sometimes quite a bit. Note that unlike hard iron interference which is the result of materials which actually have a magnetic field of their own, soft iron interference is caused by non-magnetic materials distorting the Earth's magnetic field. This type of interference has a squishing effect on the magnetic data circle turning it into more of an ellipsoid shape. The distortion in this case depends on the direction that the compass is facing. Because of this, the distortion cannot be calibrated out with a simple offset
- **Hard iron errors:** caused by static magnetic fields associated with the enviornment. For example, this could include any minor (or major) magnetism in the metal chassis or frame of a vehicle, any actual magnets such as speakers, etc... This interference pattern is unique to the environment but is constant. If you have your compass in an enclosure that is held together with metal screws, these relatively small amounts of ferromagnetic material can cause issues. If we consider the magnetic data circle, hard iron interference has the effect of shifting the entire circle away from the origin by some amount. The amount is dependent on any number of different factors and can be very large.
## References
- Ozyagcilar, T. ["Calibrating an eCompass in the Presence of Hard and Soft-iron Interference."](AN4246.pdf) Freescale Semiconductor Ltd. 1992, pp. 1-17.
- Teslabs: [Magnetometer Calibration](https://teslabs.com/articles/magnetometer-calibration/)
- ThePoorEngineer: [Calibrating the Magnetometer](https://www.thepoorengineer.com/en/calibrating-the-magnetometer/)
- Mathworks: [magcal](https://www.mathworks.com/help/fusion/ref/magcal.html#mw_34252c54-1f78-46b9-8c30-1a2b7351b0ce)
```
import numpy as np
np.set_printoptions(precision=3)
np.set_printoptions(suppress=True)
from scipy import linalg
import sys
from squaternion import Quaternion
import pandas as pd
%matplotlib inline
from matplotlib import pyplot as plt
# from math import sqrt, atan2, asin, pi
from math import radians as deg2rad
from math import degrees as rad2deg
from slurm import storage
from datetime import datetime
import os
import pickle
def loadPickle(filename):
with open(filename, 'rb') as fd:
d = pickle.load(fd)
return d
# let's load in some data and have a look at what we have
def bag_info(bag):
print('Bag keys:')
print('-'*50)
for k in bag.keys():
print(f' {k:>10}: {len(bag[k]):<7}')
# fname = "../../software/python/data.pickle"
fname = "../../software/python/dddd.pickle"
data = loadPickle(fname)
accel = []
gyro = []
mag = []
pres = []
temp = []
stamp = []
# bnoq = []
# bnoe = []
bno = {
"euler": {
"roll": [],
"pitch": [],
"yaw": [],
"time": []
},
"q": {
"w": [],
"x": [],
"y": [],
"z": [],
"time": []
}
}
tstart = data[0][-1]
for d in data:
a,g,m,p,t,q,e,dt = d
accel.append(a)
gyro.append(g)
mag.append(m)
pres.append(p)
temp.append(t)
bno["q"]["w"].append(q[0])
bno["q"]["x"].append(q[1])
bno["q"]["y"].append(q[2])
bno["q"]["z"].append(q[3])
bno["q"]["time"].append(dt - tstart)
bno["euler"]["roll"].append(e[0])
bno["euler"]["pitch"].append(e[1])
bno["euler"]["yaw"].append(e[2])
bno["euler"]["time"].append(dt - tstart)
stamp.append(dt)
accel = np.array(accel)
gyro = np.array(gyro)
uT = 50.8862
Bpp = np.array(mag)
print(f">> Mag data size: {Bpp.shape}")
def plotMagnetometer(data):
x = [v[0] for v in data]
rx = (max(x)-min(x))/2
cx = min(x)+rx
y = [v[1] for v in data]
ry = (max(y)-min(y))/2
cy = min(y)+ry
z = [v[2] for v in data]
rz = (max(z)-min(z))/2
cz = min(z)+rz
alpha = 0.1
u = np.linspace(0, 2 * np.pi, 100)
plt.plot(rx*np.cos(u)+cx, ry*np.sin(u)+cy,'-r',label='xy')
plt.plot(x,y,'.r',alpha=alpha)
plt.plot(rx*np.cos(u)+cx, rz*np.sin(u)+cz,'-g',label='xz')
plt.plot(x,z,'.g',alpha=alpha)
plt.plot(rz*np.cos(u)+cz, ry*np.sin(u)+cy,'-b',label='zy')
plt.plot(z,y, '.b',alpha=alpha)
plt.title(f"CM:({cx:.1f}, {cy:.1f}, {cz:.1f}) uT R:({rx:.1f}, {ry:.1f}, {rz:.1f}) uT")
plt.xlabel('$\mu$T')
plt.ylabel('$\mu$T')
plt.grid(True);
plt.axis('equal')
plt.legend();
def magcal(Bp, uT=None):
"""
Modelled after the matlab function: magcal(D) -> A, b, expmfs
inputs:
Bp: data points
uT: expected field strength for longitude/altitude. If None
is given, then automatically calculated and used
returns:
A: soft-iron 3x3 matrix of scaling
b: hard-iron offsets
expmfs: expected field strength"""
Y = np.array([v[0]**2+v[1]**2+v[2]**2 for v in Bp])
X = np.hstack((Bp,np.ones((Bp.shape[0],1))))
beta = np.linalg.inv(X.T.dot(X)).dot(X.T.dot(Y))
b=0.5*beta[:3]
# expected mag field strength
expmfs=np.sqrt(beta[3]+b[0]**2+b[1]**2+b[2]**2)
if uT is None:
uT = expmfs
x = [v[0] for v in Bp]
rx = (max(x)-min(x))/2
y = [v[1] for v in Bp]
ry = (max(y)-min(y))/2
z = [v[2] for v in Bp]
rz = (max(z)-min(z))/2
A = np.diag([uT/rx,uT/ry,uT/rz])
return A,b,expmfs
# Raw uncalibrated values - you can see the hard-iron offsets
# and the soft-iron ellipses
plotMagnetometer(Bpp)
# calibrated w/o expected field strength
A,vv,bb = magcal(Bpp)
print(f">> soft-iron correction:\n{A}")
print(f">> hard-iron offset: {vv}uT expmfs: {bb:.1f}uT")
plotMagnetometer((Bpp-vv).dot(A))
# calibrated with expected field strength - it only changes
# the radius of the circles
A,vv,bb = magcal(Bpp,uT)
print(f">> soft-iron correction:\n{A}")
print(f">> hard-iron offset: {vv}uT expmfs: {bb:.1f}uT")
plotMagnetometer((Bpp-vv).dot(A))
```
```
>> soft-iron correction:
[[0.983 0. 0. ]
[0. 0.947 0. ]
[0. 0. 0.941]]
>> hard-iron offset: [-20.438 34.429 -2.368]uT expmfs: 52.6uT
```
```
>> soft-iron correction:
[[0.951 0. 0. ]
[0. 0.916 0. ]
[0. 0. 0.91 ]]
>> hard-iron offset: [-20.438 34.429 -2.368]uT expmfs: 52.6uT
```
# Save Parameters
```
M = np.vstack((A,vv))
print(M)
params = {}
params["imu"] = "adafruit NXP"
params["timestamp"] = datetime.now()
params["mag"] = M.tolist()
params["shape"] = M.shape
storage.write("magnetometer-alt.yaml", params)
np.hstack((A,vv.reshape((3,1))))
rr = ["# hello",
{
"A": A.tolist(),
"b": vv.tolist()
}]
storage.write("temp.yaml", rr)
```
| github_jupyter |
# Developing an AI application
Going forward, AI algorithms will be incorporated into more and more everyday applications. For example, you might want to include an image classifier in a smart phone app. To do this, you'd use a deep learning model trained on hundreds of thousands of images as part of the overall application architecture. A large part of software development in the future will be using these types of models as common parts of applications.
In this project, you'll train an image classifier to recognize different species of flowers. You can imagine using something like this in a phone app that tells you the name of the flower your camera is looking at. In practice you'd train this classifier, then export it for use in your application. We'll be using [this dataset](http://www.robots.ox.ac.uk/~vgg/data/flowers/102/index.html) of 102 flower categories, you can see a few examples below.
<img src='assets/Flowers.png' width=500px>
The project is broken down into multiple steps:
* Load and preprocess the image dataset
* Train the image classifier on your dataset
* Use the trained classifier to predict image content
We'll lead you through each part which you'll implement in Python.
When you've completed this project, you'll have an application that can be trained on any set of labeled images. Here your network will be learning about flowers and end up as a command line application. But, what you do with your new skills depends on your imagination and effort in building a dataset. For example, imagine an app where you take a picture of a car, it tells you what the make and model is, then looks up information about it. Go build your own dataset and make something new.
First up is importing the packages you'll need. It's good practice to keep all the imports at the beginning of your code. As you work through this notebook and find you need to import a package, make sure to add the import up here.
```
# Imports here
import torch
import matplotlib
import matplotlib.pyplot as plt
import torch.nn.functional as F
import numpy as np
import time
from torch import nn
from torch import optim
from torchvision import datasets, transforms, models
from PIL import Image
```
## Load the data
Here you'll use `torchvision` to load the data ([documentation](http://pytorch.org/docs/0.3.0/torchvision/index.html)). The data should be included alongside this notebook, otherwise you can [download it here](https://s3.amazonaws.com/content.udacity-data.com/nd089/flower_data.tar.gz). The dataset is split into three parts, training, validation, and testing. For the training, you'll want to apply transformations such as random scaling, cropping, and flipping. This will help the network generalize leading to better performance. You'll also need to make sure the input data is resized to 224x224 pixels as required by the pre-trained networks.
The validation and testing sets are used to measure the model's performance on data it hasn't seen yet. For this you don't want any scaling or rotation transformations, but you'll need to resize then crop the images to the appropriate size.
The pre-trained networks you'll use were trained on the ImageNet dataset where each color channel was normalized separately. For all three sets you'll need to normalize the means and standard deviations of the images to what the network expects. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`, calculated from the ImageNet images. These values will shift each color channel to be centered at 0 and range from -1 to 1.
```
data_dir = 'flowers'
train_dir = data_dir + '/train'
valid_dir = data_dir + '/valid'
test_dir = data_dir + '/test'
# TODO: Define your transforms for the training, validation, and testing sets
train_transforms = transforms.Compose([transforms.RandomRotation(30),
transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
test_transforms = transforms.Compose([transforms.Resize(255),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
validation_transforms = transforms.Compose([transforms.Resize(255),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
# TODO: Load the datasets with ImageFolder
train_data = datasets.ImageFolder(train_dir, transform=train_transforms)
test_data = datasets.ImageFolder(test_dir, transform=test_transforms)
validation_data = datasets.ImageFolder(valid_dir, transform=validation_transforms)
# TODO: Using the image datasets and the trainforms, define the dataloaders
trainloader = torch.utils.data.DataLoader(train_data, batch_size=64, shuffle=True)
testloader = torch.utils.data.DataLoader(test_data, batch_size=64)
validloader = torch.utils.data.DataLoader(validation_data, batch_size=64)
```
### Label mapping
You'll also need to load in a mapping from category label to category name. You can find this in the file `cat_to_name.json`. It's a JSON object which you can read in with the [`json` module](https://docs.python.org/2/library/json.html). This will give you a dictionary mapping the integer encoded categories to the actual names of the flowers.
```
import json
with open('cat_to_name.json', 'r') as f:
cat_to_name = json.load(f)
```
# Building and training the classifier
Now that the data is ready, it's time to build and train the classifier. As usual, you should use one of the pretrained models from `torchvision.models` to get the image features. Build and train a new feed-forward classifier using those features.
We're going to leave this part up to you. Refer to [the rubric](https://review.udacity.com/#!/rubrics/1663/view) for guidance on successfully completing this section. Things you'll need to do:
* Load a [pre-trained network](http://pytorch.org/docs/master/torchvision/models.html) (If you need a starting point, the VGG networks work great and are straightforward to use)
* Define a new, untrained feed-forward network as a classifier, using ReLU activations and dropout
* Train the classifier layers using backpropagation using the pre-trained network to get the features
* Track the loss and accuracy on the validation set to determine the best hyperparameters
We've left a cell open for you below, but use as many as you need. Our advice is to break the problem up into smaller parts you can run separately. Check that each part is doing what you expect, then move on to the next. You'll likely find that as you work through each part, you'll need to go back and modify your previous code. This is totally normal!
When training make sure you're updating only the weights of the feed-forward network. You should be able to get the validation accuracy above 70% if you build everything right. Make sure to try different hyperparameters (learning rate, units in the classifier, epochs, etc) to find the best model. Save those hyperparameters to use as default values in the next part of the project.
One last important tip if you're using the workspace to run your code: To avoid having your workspace disconnect during the long-running tasks in this notebook, please read in the earlier page in this lesson called Intro to
GPU Workspaces about Keeping Your Session Active. You'll want to include code from the workspace_utils.py module.
**Note for Workspace users:** If your network is over 1 GB when saved as a checkpoint, there might be issues with saving backups in your workspace. Typically this happens with wide dense layers after the convolutional layers. If your saved checkpoint is larger than 1 GB (you can open a terminal and check with `ls -lh`), you should reduce the size of your hidden layers and train again.
```
# Load a pre-trained network
model = models.vgg16(pretrained=True)
print(model)
# Train the classifier layers using backpropagation using the pre-trained network to get the features
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# Freeze parameters so we don't backprop through them
for param in model.parameters():
param.requires_grad = False
model.classifier = nn.Sequential((nn.Dropout(0.5)),
(nn.Linear(25088, 120)),
(nn.ReLU()),
(nn.Linear(120, 90)),
(nn.ReLU()),
(nn.Linear(90,80)),
(nn.ReLU()),
(nn.Linear(80,102)),
(nn.LogSoftmax(dim=1)))
criterion = nn.NLLLoss()
# Only train the classifier parameters, feature parameters are frozen
optimizer = optim.Adam(model.classifier.parameters(), lr=0.001)
model.to(device);
#Track the loss and accuracy on the validation set to determine the best hyperparameters
epochs = 3
steps = 0
running_loss = 0
print_every = 5
for epoch in range(epochs):
for inputs, labels in trainloader:
steps += 1
# Move input and label tensors to the default device
inputs, labels = inputs.to(device), labels.to(device)
optimizer.zero_grad()
logps = model.forward(inputs)
loss = criterion(logps, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
if steps % print_every == 0:
test_loss = 0
accuracy = 0
model.eval()
with torch.no_grad():
for inputs, labels in validloader:
inputs, labels = inputs.to(device), labels.to(device)
logps = model.forward(inputs)
batch_loss = criterion(logps, labels)
test_loss += batch_loss.item()
# Calculate accuracy
ps = torch.exp(logps)
top_p, top_class = ps.topk(1, dim=1)
equals = top_class == labels.view(*top_class.shape)
accuracy += torch.mean(equals.type(torch.FloatTensor)).item()
print(f"Epoch {epoch+1}/{epochs}.. "
f"Train loss: {running_loss/print_every:.3f}.. "
f"Validation loss: {test_loss/len(testloader):.3f}.. "
f"Accuracy: {accuracy/len(testloader):.3f}")
running_loss = 0
model.train()
```
## Testing your network
It's good practice to test your trained network on test data, images the network has never seen either in training or validation. This will give you a good estimate for the model's performance on completely new images. Run the test images through the network and measure the accuracy, the same way you did validation. You should be able to reach around 70% accuracy on the test set if the model has been trained well.
```
# TODO: Do validation on the test set
def check_test_acc(testloader):
correct = 0
total = 0
with torch.no_grad():
for data in testloader:
images, labels = data
images, labels = images.to(device), labels.to(device)
outputs = model(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy on test images %d %%' % (100 * correct / total))
check_test_acc(testloader)
```
## Save the checkpoint
Now that your network is trained, save the model so you can load it later for making predictions. You probably want to save other things such as the mapping of classes to indices which you get from one of the image datasets: `image_datasets['train'].class_to_idx`. You can attach this to the model as an attribute which makes inference easier later on.
```model.class_to_idx = image_datasets['train'].class_to_idx```
Remember that you'll want to completely rebuild the model later so you can use it for inference. Make sure to include any information you need in the checkpoint. If you want to load the model and keep training, you'll want to save the number of epochs as well as the optimizer state, `optimizer.state_dict`. You'll likely want to use this trained model in the next part of the project, so best to save it now.
```
# TODO: Save the checkpoint
model.class_to_idx = train_data.class_to_idx
torch.save({
'epochs': epochs,
'model_state_dict': model.state_dict(),
'optimizer_state_dict': optimizer.state_dict(),
'class_to_idx':model.class_to_idx,
'arch': 'vgg16',
}, 'checkpoint.pt')
```
## Loading the checkpoint
At this point it's good to write a function that can load a checkpoint and rebuild the model. That way you can come back to this project and keep working on it without having to retrain the network.
```
# TODO: Write a function that loads a checkpoint and rebuilds the model
def load_model(path):
checkpoint = torch.load(path)
model.class_to_idx = checkpoint['class_to_idx']
model.load_state_dict(checkpoint['model_state_dict'])
epochs = checkpoint['epochs']
load_model('checkpoint.pt')
print(model)
```
# Inference for classification
Now you'll write a function to use a trained network for inference. That is, you'll pass an image into the network and predict the class of the flower in the image. Write a function called `predict` that takes an image and a model, then returns the top $K$ most likely classes along with the probabilities. It should look like
```python
probs, classes = predict(image_path, model)
print(probs)
print(classes)
> [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]
> ['70', '3', '45', '62', '55']
```
First you'll need to handle processing the input image such that it can be used in your network.
## Image Preprocessing
You'll want to use `PIL` to load the image ([documentation](https://pillow.readthedocs.io/en/latest/reference/Image.html)). It's best to write a function that preprocesses the image so it can be used as input for the model. This function should process the images in the same manner used for training.
First, resize the images where the shortest side is 256 pixels, keeping the aspect ratio. This can be done with the [`thumbnail`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) or [`resize`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) methods. Then you'll need to crop out the center 224x224 portion of the image.
Color channels of images are typically encoded as integers 0-255, but the model expected floats 0-1. You'll need to convert the values. It's easiest with a Numpy array, which you can get from a PIL image like so `np_image = np.array(pil_image)`.
As before, the network expects the images to be normalized in a specific way. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`. You'll want to subtract the means from each color channel, then divide by the standard deviation.
And finally, PyTorch expects the color channel to be the first dimension but it's the third dimension in the PIL image and Numpy array. You can reorder dimensions using [`ndarray.transpose`](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.ndarray.transpose.html). The color channel needs to be first and retain the order of the other two dimensions.
```
def process_image(image):
img_pil = Image.open(image)
width, height = img_pil.size
size = 256
if width > height:
ratio = width / height
new_height = size
new_width = int(size * ratio)
else:
ratio = height / width
new_width = size
new_height = int(size * ratio)
adjustments = transforms.Compose([
transforms.Resize((new_width, new_height)),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225])
])
img_tensor = adjustments(img_pil)
return img_tensor
# TODO: Process a PIL image for use in a PyTorch model
img = (data_dir + '/test' + '/1/' + 'image_06752.jpg')
img = process_image(img)
print(img.shape)
```
To check your work, the function below converts a PyTorch tensor and displays it in the notebook. If your `process_image` function works, running the output through this function should return the original image (except for the cropped out portions).
```
def imshow(image, ax=None, title=None):
"""Imshow for Tensor."""
if ax is None:
fig, ax = plt.subplots()
# PyTorch tensors assume the color channel is the first dimension
# but matplotlib assumes is the third dimension
image = image.numpy().transpose((1, 2, 0))
# Undo preprocessing
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
image = std * image + mean
# Image needs to be clipped between 0 and 1 or it looks like noise when displayed
image = np.clip(image, 0, 1)
ax.imshow(image)
return ax
imshow(process_image("flowers/test/1/image_06743.jpg"))
```
## Class Prediction
Once you can get images in the correct format, it's time to write a function for making predictions with your model. A common practice is to predict the top 5 or so (usually called top-$K$) most probable classes. You'll want to calculate the class probabilities then find the $K$ largest values.
To get the top $K$ largest values in a tensor use [`x.topk(k)`](http://pytorch.org/docs/master/torch.html#torch.topk). This method returns both the highest `k` probabilities and the indices of those probabilities corresponding to the classes. You need to convert from these indices to the actual class labels using `class_to_idx` which hopefully you added to the model or from an `ImageFolder` you used to load the data ([see here](#Save-the-checkpoint)). Make sure to invert the dictionary so you get a mapping from index to class as well.
Again, this method should take a path to an image and a model checkpoint, then return the probabilities and classes.
```python
probs, classes = predict(image_path, model)
print(probs)
print(classes)
> [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]
> ['70', '3', '45', '62', '55']
```
```
def predict(image_path, model, topk=5):
# TODO: Implement the code to predict the class from an image file
img_torch = process_image(image_path)
img_torch = img_torch.unsqueeze_(0)
img_torch = img_torch.float()
with torch.no_grad():
output = model.forward(img_torch.cuda())
probability = F.softmax(output.data,dim=1)
return probability.topk(topk)
img = (data_dir + '/test' + '/1/' + 'image_06743.jpg')
val1, val2 = predict(img, model)
print(val1)
print(val2)
```
## Sanity Checking
Now that you can use a trained model for predictions, check to make sure it makes sense. Even if the testing accuracy is high, it's always good to check that there aren't obvious bugs. Use `matplotlib` to plot the probabilities for the top 5 classes as a bar graph, along with the input image. It should look like this:
<img src='assets/inference_example.png' width=300px>
You can convert from the class integer encoding to actual flower names with the `cat_to_name.json` file (should have been loaded earlier in the notebook). To show a PyTorch tensor as an image, use the `imshow` function defined above.
```
# TODO: Display an image along with the top 5 classes
def check_sanity(path):
plt.rcParams["figure.figsize"] = (10,5)
plt.subplot(211)
index = 1
probabilities = predict(path, model)
image = process_image(path)
probabilities = probabilities
axs = imshow(image, ax = plt)
axs.axis('off')
axs.title(cat_to_name[str(index)])
axs.show()
a = np.array((probabilities[0][0]).cpu())
b = [cat_to_name[str(index + 1)] for index in np.array((probabilities[1][0]).cpu())]
N=float(len(b))
fig,ax = plt.subplots(figsize=(8,3))
width = 0.8
tickLocations = np.arange(N)
ax.bar(tickLocations, a, width, linewidth=4.0, align = 'center')
ax.set_xticks(ticks = tickLocations)
ax.set_xticklabels(b)
ax.set_xlim(min(tickLocations)-0.6,max(tickLocations)+0.6)
ax.set_yticks([0.2,0.4,0.6,0.8,1,1.2])
ax.set_ylim((0,1))
ax.yaxis.grid(True)
plt.show()
check_sanity(test_dir + '/1/image_06743.jpg')
```
| github_jupyter |
#### New to Plotly?
Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
<br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
<br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
### Imports
This tutorial imports [Plotly](https://plot.ly/python/getting-started/), [Numpy](http://www.numpy.org/), and [Pandas](https://plot.ly/pandas/intro-to-pandas-tutorial/).
```
import plotly.plotly as py
from plotly.tools import FigureFactory as FF
import numpy as np
import pandas as pd
```
#### Import Data
For this histogram example, we will import some real data.
```
import plotly.plotly as py
from plotly.tools import FigureFactory as FF
data = pd.read_csv('https://raw.githubusercontent.com/plotly/datasets/master/wind_speed_laurel_nebraska.csv')
df = data[0:10]
table = FF.create_table(df)
py.iplot(table, filename='wind-data-sample')
```
#### Histogram
Using `np.histogram()` we can compute histogram data from a data array. This function returns the values of the histogram (i.e. the number for each bin) and the bin endpoints as well, which denote the intervals for which the histogram values correspond to.
```
import plotly.plotly as py
import plotly.graph_objs as go
data_array = np.array((data['10 Min Std Dev']))
hist_data = np.histogram(data_array)
binsize = hist_data[1][1] - hist_data[1][0]
trace1 = go.Histogram(
x=data_array,
histnorm='count',
name='Histogram of Wind Speed',
autobinx=False,
xbins=dict(
start=hist_data[1][0],
end=hist_data[1][-1],
size=binsize
)
)
trace_data = [trace1]
layout = go.Layout(
bargroupgap=0.3
)
fig = go.Figure(data=trace_data, layout=layout)
py.iplot(fig)
hist_data
help(np.histogram)
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'Histogram.ipynb', 'numpy/histogram/', 'Histogram | plotly',
'A histogram is a chart which divides data into bins with a numeric range, and each bin gets a bar corresponding to the number of data points in that bin.',
title = 'Numpy Histogram | plotly',
name = 'Histogram',
has_thumbnail='true', thumbnail='thumbnail/histogram.jpg',
language='numpy', page_type='example_index',
display_as='numpy-statistics', order=2)
```
| github_jupyter |
# A simple example
To illustrate the application of quantile regression neural networks we consider a simple, 1-dimensional regression probelm with heteroscedastic uncertainty:
\begin{align}
y = \sin(x) + \cos(x) \cdot \mathcal{N}(0, 1)
\end{align}
The code below generates the synthetic training data consisting of $10^7$ randomly sampled $x$ values and corresponding $y$ values.
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import quantnn
import quantnn.plotting
quantnn.plotting.set_style()
from quantnn.examples import simple
x, y = simple.create_training_data(10_000_000)
simple.plot_histogram(x, y)
plt.savefig("simple_training_data.png")
```
## Quantile regression neural networks (QRNNs)
The task of predicting the conditional distribution $P(y|x)$ of $y$ given input $x$ can be solved using quantile regression. The ``QRNN`` class of the quantnn package provides an implementation of a Quantile Regression Neural Network (QRNN), i.e. it uses a neural network to learn to predict the quantiles of the posterior distribution $P(y|x)$.
### Model definition and training
The simplest way to train a quantile regression neural network is to use the ``quantnn.QRNN`` class, which provides a simple interface to define and and train QRNNs based on fully-connected, feed-forward neural networks.
To do this all we need to do is to choose which quantiles to predict as well as the basic parameters of the network architecture. More specifically, the following information must be provided to the constructor of the ``QRNN`` class:
1. The list of quantiles that we want to predict.
2. The number of input features (``input_dimensions``). For this simple example this just ``1``.
3. A tuple describing the model architecture ``(n_layers, n_neurons, activation_function)``
> **Note:** quantnn will automatically choose the backend between Keras and PyTorch depending on which one is available on your system. You can specifically choose one over the other using the ``quantnn.qrnn.set_backend`` function.
```
import quantnn as q
quantiles = np.linspace(0.01, 0.99, 99)
layers = 4
neurons = 128
activation = "relu"
model = (layers, neurons, activation)
qrnn = q.QRNN(quantiles, n_inputs=1, model=model)
```
Finally, we train the neural network on the training data. Here, we use only a subset of the data, in order to speed up the training process. We restart the training three times while lowering the learning rate, which leads to more accurate predicted quantiles.
```
training_data = (x[::5].reshape(-1, 1), y[::5].reshape(-1, 1))
results = qrnn.train(training_data=training_data,
n_epochs=15)
```
### Evaluation
To evaluate the QRNN we generate 1000 new $x$-value and use the ``predict`` method of the trained ``qrnn`` object to predict the quantiles of $y$.
```
n = 1_000
x_val = np.linspace(-np.pi, np.pi, n)
y_val = simple.create_validation_data(x_val)
y_pred = qrnn.predict(x_val.reshape(-1, 1))
y_mean = qrnn.posterior_mean(y_pred=y_pred)
```
We can now compare the quantiles predicted using the QRNN to the empirical quantiles from the training data. The QRNN manages fairly well to predict the true conditional cumulative distribution function (CDF). With additional optimization of the training process, the
```
from quantnn.plotting import plot_quantiles
from matplotlib.gridspec import GridSpec
gs = GridSpec(1, 3, width_ratios=[1.0, 1.0, 0.05])
f = plt.figure(figsize=(12, 5))
# Calculate empirical CDF
bins = np.linspace(-np.pi, np.pi, 201)
counts, _, _ = np.histogram2d(y, x, bins=bins)
norm = np.sum(counts * (bins[1:] - bins[:-1]), axis=0, keepdims=True)
counts /= norm
cdf = np.cumsum(counts * (bins[1:] - bins[:-1]), axis=0)
# Plot QRNN quantiles
ax = plt.subplot(gs[0, 0])
m = plot_quantiles(ax, x_val, y_pred[:, 1::6], qrnn.quantiles[1::6])
ax.set_xlim([-np.pi, np.pi])
ax.set_ylim([-2.5, 2.5])
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.set_title("(a) Predicted CDF", loc="left")
ax.grid(False)
# Plot true cdf
ax = plt.subplot(gs[0, 1])
ax.pcolormesh(bins, bins, cdf, cmap=m.cmap, norm=m.norm)
ax.set_xlim([-np.pi, np.pi])
ax.set_ylim([-2.5, 2.5])
ax.set_xlabel("x")
ax.set_title("(b) True CDF", loc="left")
ax = plt.subplot(gs[0, 2])
plt.colorbar(m, cax=ax, label="$P(Y \leq y | X=x)$")
plt.tight_layout()
```
## Density regression neural network
An alternative way to learn to predict the distribution $P(y|x)$ is to use a neural network to predict a gridded approximation of the probability density function, a technique we will, for simplicity, refer to as Density Regression Neural Network (DRNN).
To create a DRNN model, we need to choose a suitable binning for the output value. Here we simply use an equidistant grid covering the range $-3$ to $3$. Apart from having to provide the bins of the output PDF instead of the quantiles to predict, the definition, training and evaluation of the DRNN works in exactly the same way as for the QRNN.
```
from quantnn.drnn import DRNN
layers = 4
neurons = 128
activation = "relu"
model = (layers, neurons, activation)
bins = np.linspace(-3, 3, 101)
drnn = DRNN(bins, n_inputs=1, model=model)
training_data = (x[::5].reshape(-1, 1), y[::5].reshape(-1, 1))
results = drnn.train(training_data=training_data,
n_epochs=15)
y_pred = drnn.predict(x_val.reshape(-1, 1))
```
To evaluate the quality of the predicted PDF, we can now simply plot the predicted PDFs to the empirical PDFs of the training data. Similar as for the QRNN, the prediction match the reference values fairly well.
```
from quantnn.plotting import plot_quantiles
from matplotlib.colors import Normalize
from matplotlib.gridspec import GridSpec
gs = GridSpec(1, 3, width_ratios=[1.0, 1.0, 0.05])
f = plt.figure(figsize=(12, 5))
norm=Normalize(0.0, 0.5)
# Plot DRNN PDF
bin_centers = 0.5 * (drnn.bins[1:] + drnn.bins[:-1])
ax = plt.subplot(gs[0, 0])
m = ax.pcolormesh(x_val, bin_centers, y_pred.T, norm=norm)
ax.set_xlim([-np.pi, np.pi])
ax.set_ylim([-2.5, 2.5])
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.set_title("(a) Predicted PDF", loc="left")
ax.grid(False)
# Plot true cdf
bins_x = np.linspace(-np.pi, np.pi, 201)
counts, _, _ = np.histogram2d(y, x, bins=((bins, bins_x)))
counts /= np.sum(counts * (bins[1:] - bins[:-1])[:, np.newaxis], axis=0, keepdims=True)
ax = plt.subplot(gs[0, 1])
ax.pcolormesh(bins_x, bins, counts, norm=norm)
ax.set_xlim([-np.pi, np.pi])
ax.set_ylim([-2.5, 2.5])
ax.set_xlabel("x")
ax.set_title("(b) True PDF", loc="left")
ax = plt.subplot(gs[0, 2])
plt.colorbar(m, cax=ax, label="$p(y | x)$")
plt.tight_layout()
```
## Calculating posterior statistics
Both the QRNN and DRNN allow us to predict the posterior distribution of the quantity $y$ given input $x$. In most applications, however, we are not interested in the full distribution but rather specific statistics. The quantnn package provides functions to calculate a range of such statistics in the ``quantnn.quantiles`` module for QRNN predictions and in the ``quantnn.desntiy`` module for predictions produced by a DRNN.
```
y_pred_qrnn = qrnn.predict(x_val.reshape(-1, 1))
y_pred_drnn = drnn.predict(x_val.reshape(-1, 1))
```
### Mean and median
```
import quantnn.quantiles as qq
import quantnn.density as qd
qrnn_mean = qrnn.posterior_mean(y_pred=y_pred_qrnn)
drnn_mean = drnn.posterior_mean(y_pred=y_pred_drnn)
qrnn_median = y_pred_qrnn[:, 49]
drnn_median = drnn.posterior_quantiles(y_pred=y_pred_drnn, quantiles=[0.5])
gs = GridSpec(2, 2, height_ratios=[1.0, 0.1])
f, axs = plt.subplots(1, 2, figsize=(12, 5))
y_mean = np.sin(x_val)
ax = plt.subplot(gs[0, 0])
ax.plot(x_val, qrnn_mean)
ax.plot(x_val, drnn_mean)
ax.plot(x_val, y_mean, c="k", ls="--", label="Truth")
ax.set_title("(a) Mean", loc="left")
ax = plt.subplot(gs[0, 1])
handles = []
handles += ax.plot(x_val, qrnn_median)
handles += ax.plot(x_val, drnn_median)
handles += ax.plot(x_val, y_mean, c="k", ls="--", label="Truth")
ax.set_title("(b) Median", loc="left")
ax = plt.subplot(gs[1, :])
ax.set_axis_off()
ax.legend(handles=handles, labels=["QRNN", "DRNN", "truth"], ncol=3, loc="center")
```
### Quantiles
```
quantiles = np.linspace(0.1, 0.9, 9)
quantiles_qrnn = qrnn.posterior_quantiles(y_pred=y_pred_qrnn, quantiles=quantiles)
quantiles_drnn = drnn.posterior_quantiles(y_pred=y_pred_drnn, quantiles=quantiles)
gs = GridSpec(1, 3, width_ratios=[1.0, 1.0, 0.05])
f, axs = plt.subplots(1, 2, figsize=(13, 5))
y_mean = np.sin(x_val)
ax = plt.subplot(gs[0, 0])
m = plot_quantiles(ax, x_val, quantiles_qrnn, quantiles)
ax.set_title("(a) QRNN quantiles", loc="left")
ax.set_xlim([-np.pi, np.pi])
ax.set_ylim([-2, 2])
ax.set_xlabel("x")
ax.set_ylabel("y")
ax = plt.subplot(gs[0, 1])
plot_quantiles(ax, x_val, quantiles_drnn, quantiles)
ax.set_title("(b) DRNN quantiles", loc="left")
ax.set_xlim([-np.pi, np.pi])
ax.set_ylim([-2, 2])
ax.set_xlabel("x")
ax.set_ylabel("y")
ax = plt.subplot(gs[0, 2])
plt.colorbar(m, cax=ax, label="$P(Y \leq y | X=x)$")
```
## Binned PDF
```
bins = drnn.bins
centers = 0.5 * (bins[1:] + bins[:-1])
binned_pdf_qrnn = qq.pdf_binned(y_pred_qrnn.numpy(),
qrnn.quantiles,
bins)
gs = GridSpec(1, 3, width_ratios=[1.0, 1.0, 0.05])
f, axs = plt.subplots(1, 2, figsize=(13, 5))
norm = Normalize(0, 0.5)
ax = plt.subplot(gs[0, 0])
m = ax.pcolormesh(x_val, centers, binned_pdf_qrnn.T, norm=norm)
ax.set_title("(a) QRNN", loc="left")
ax.set_xlim([-np.pi, np.pi])
ax.set_ylim([-2, 2])
ax.set_xlabel("x")
ax.set_ylabel("y")
ax = plt.subplot(gs[0, 1])
m = ax.pcolormesh(x_val, centers, y_pred_drnn.T, norm=norm)
ax.set_title("(b) DRNN", loc="left")
ax.set_xlim([-np.pi, np.pi])
ax.set_ylim([-2, 2])
ax.set_xlabel("x")
ax = plt.subplot(gs[0, 2])
plt.colorbar(m, cax=ax, label="$p(y | x)$")
plt.tight_layout()
```
## Classification
Finally, we consider the case of trying to classfying whether the output $y$ corresponding to a given $x$ is larger than a certain threshold value. For this we can use the ``probability_larger_than`` functions of the ``quantnn.quantiles`` and ``quantnn.density`` modules. This function returns the probability that the output $y$ is larger than the given threshold.
```
t = 0.1
p_qrnn = qq.probability_larger_than(y_pred_qrnn.numpy(), qrnn.quantiles, t)
p_drnn = qd.probability_larger_than(y_pred_drnn.numpy(), drnn.bins, t)
# Calculate empirical probability
bins = np.linspace(-np.pi, np.pi, 201)
counts, _, _ = np.histogram2d(y, x, bins=bins)
centers = 0.5 * (bins[1:] + bins[:-1])
counts_larger_t = counts * (centers > t).reshape(-1, 1)
p = np.sum(counts_larger_t, axis=0) / np.sum(counts, axis=0)
f, ax = plt.subplots(1, 1)
ax.plot(x_val, p_qrnn)
ax.plot(x_val, p_drnn)
ax.plot(centers, p, c="k", ls="--")
ax.set_xlim([-np.pi, np.pi])
ax.set_ylim([0, 1])
ax.set_xlabel("x")
ax.set_ylabel("$P(Y > 0.1 | X = x)$")
```
| github_jupyter |
Toy Image
```
import tensorflow as tf
import numpy as np
import random
import matplotlib.pyplot as plt
tf.set_random_seed(777) # for reproducibility
sess = tf.InteractiveSession()
image = np.array([[[[1],[2],[3]],
[[4],[5],[6]],
[[7],[8],[9]]]], dtype=np.float32)
print("image.shape", image.shape)
plt.imshow(image.reshape(3,3), cmap='Greys')
```
Image: 1,3,3,1 image, Filter: 2,2,1,1, Stride: 1x1, Padding: VALID
```
weight = tf.constant([[[[1.]],[[1.]]],
[[[1.]],[[1.]]]])
print("weight.shape", weight.shape)
conv2d = tf.nn.conv2d(image, weight, strides=[1,1,1,1], padding='VALID')
conv2d_img = conv2d.eval()
print("conv2d_img.shape", conv2d_img.shape)
conv2d_img = np.swapaxes(conv2d_img, 0, 3)
for i, one_img in enumerate(conv2d_img):
print(one_img.reshape(2,2))
plt.subplot(1,2,i+1), plt.imshow(one_img.reshape(2,2), cmap="gray")
```
Image: 1,3,3,1 image, Filter: 2,2,1,1, Stride: 1x1, Padding: SAME
```
print("weight.shape", weight.shape)
conv2d = tf.nn.conv2d(image, weight, strides=[1,1,1,1], padding='SAME')
conv2d_img = conv2d.eval()
print("conv2d_img.shape", conv2d_img.shape)
conv2d_img = np.swapaxes(conv2d_img, 0, 3)
for i, one_img in enumerate(conv2d_img):
print(one_img.reshape(3,3))
plt.subplot(1,2,i+1), plt.imshow(one_img.reshape(3,3), cmap="gray")
```
3 filters (2,2,1,3)
```
weight = tf.constant([[[[1.,10.,-1.]],[[1.,10.,-1.]]],
[[[1.,10.,-1.]],[[1.,10.,-1.]]]])
print("weight.shape", weight.shape)
conv2d = tf.nn.conv2d(image, weight, strides=[1,1,1,1], padding='SAME')
conv2d_img = conv2d.eval()
print("conv2d_img.shape", conv2d_img.shape)
conv2d_img = np.swapaxes(conv2d_img, 0, 3)
for i, one_img in enumerate(conv2d_img):
print(one_img.reshape(3,3))
plt.subplot(1,3,i+1), plt.imshow(one_img.reshape(3,3), cmap="gray")
```
Max Pooling
```
image = np.array([[[[4],[3]],
[[2],[1]]]], dtype=np.float32)
pool = tf.nn.max_pool(image, ksize=[1,2,2,1],
strides=[1,1,1,1], padding='SAME')
print(image.shape, pool.shape)
print(pool.eval())
```
MNIST image loading
```
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True)
img = mnist.train.images[0].reshape(28,28)
plt.imshow(img, cmap='gray')
```
MNIST Convolution layer
```
img = img.reshape(-1, 28, 28, 1)
W1 = tf.Variable(tf.random_normal([3, 3, 1, 5], stddev=0.01))
conv2d = tf.nn.conv2d(img, W1, strides=[1,2,2,1], padding='SAME')
print(conv2d)
sess.run(tf.global_variables_initializer())
conv2d_img = conv2d.eval()
conv2d_img = np.swapaxes(conv2d_img, 0, 3)
for i, one_img in enumerate(conv2d_img):
plt.subplot(1,5,i+1), plt.imshow(one_img.reshape(14,14), cmap='gray')
```
| github_jupyter |
[[source]](../api/alibi.explainers.cem.rst)
# Contrastive Explanation Method
## Overview
The *Contrastive Explanation Method* (CEM) is based on the paper [Explanations based on the Missing: Towards Constrastive Explanations with Pertinent Negatives](https://arxiv.org/abs/1802.07623) and extends the [code](https://github.com/IBM/Contrastive-Explanation-Method) open sourced by the authors. CEM generates instance based local black box explanations for classification models in terms of Pertinent Positives (PP) and Pertinent Negatives (PN). For a PP, the method finds the features that should be minimally and sufficiently present (e.g. important pixels in an image) to predict the same class as on the original instance. PN's on the other hand identify what features should be minimally and necessarily absent from the instance to be explained in order to maintain the original prediction class. The aim of PN's is not to provide a full set of characteristics that should be absent in the explained instance, but to provide a minimal set that differentiates it from the closest different class. Intuitively, the Pertinent Positives could be compared to Anchors while Pertinent Negatives are similar to Counterfactuals. As the authors of the paper state, CEM can generate clear explanations of the form: "An input x is classified in class y because features $f_{i}$, ..., $f_{k}$ are present and because features $f_{m}$, ..., $f_{p}$ are absent." The current implementation is most suitable for images and tabular data without categorical features.
In order to create interpretable PP's and PN's, feature-wise perturbation needs to be done in a meaningful way. To keep the perturbations sparse and close to the original instance, the objective function contains an elastic net ($\beta$$L_{1}$ + $L_{2}$) regularizer. Optionally, an auto-encoder can be trained to reconstruct instances of the training set. We can then introduce the $L_{2}$ reconstruction error of the perturbed instance as an additional loss term in our objective function. As a result, the perturbed instance lies close to the training data manifold.
The ability to add or remove features to arrive at respectively PN's or PP's implies that there are feature values that contain no information with regards to the model's predictions. Consider for instance the MNIST image below where the pixels are scaled between 0 and 1. The pixels with values close to 1 define the number in the image while the background pixels have value 0. We assume that perturbations towards the background value 0 are equivalent to removing features, while perturbations towards 1 imply adding features.

It is intuitive to understand that adding features to get a PN means changing 0's into 1's until a different number is formed, in this case changing a 4 into a 9.

To find the PP, we do the opposite and change 1's from the original instance into 0's, the background value, and only keep a vague outline of the original 4.

It is however often not trivial to find these non-informative feature values and domain knowledge becomes very important.
For more details, we refer the reader to the original [paper](https://arxiv.org/abs/1802.07623).
## Usage
### Initialization
The optimizer is defined in TensorFlow (TF) internally. We first load our MNIST classifier and the (optional) auto-encoder. The example below uses Keras or TF models. This allows optimization of the objective function to run entirely with automatic differentiation because the TF graph has access to the underlying model architecture. For models built in different frameworks (e.g. scikit-learn), the gradients of part of the loss function with respect to the input features need to be evaluated numerically. We'll handle this case later.
```python
# define models
cnn = load_model('mnist_cnn.h5')
ae = load_model('mnist_ae.h5')
```
We can now initialize the CEM explainer:
```python
# initialize CEM explainer
shape = (1,) + x_train.shape[1:]
mode = 'PN'
cem = CEM(cnn, mode, shape, kappa=0., beta=.1,
feature_range=(x_train.min(), x_train.max()),
gamma=100, ae_model=ae, max_iterations=1000,
c_init=1., c_steps=10, learning_rate_init=1e-2,
clip=(-1000.,1000.), no_info_val=-1.)
```
Besides passing the the predictive and auto-encoder models, we set a number of **hyperparameters** ...
... **general**:
* `mode`: 'PN' or 'PP'.
* `shape`: shape of the instance to be explained, starting with batch dimension. Currently only single explanations are supported, so the batch dimension should be equal to 1.
* `feature_range`: global or feature-wise min and max values for the perturbed instance.
... related to the **optimizer**:
* `max_iterations`: number of loss optimization steps for each value of *c*; the multiplier of the first loss term.
* `learning_rate_init`: initial learning rate, follows polynomial decay.
* `clip`: min and max gradient values.
... related to the **non-informative value**:
* `no_info_val`: as explained in the previous section, it is important to define which feature values are considered background and not crucial for the class predictions. For MNIST images scaled between 0 and 1 or -0.5 and 0.5 as in the notebooks, pixel perturbations in the direction of the (low) background pixel value can be seen as removing features, moving towards the non-informative value. As a result, the `no_info_val` parameter is set at a low value like -1. `no_info_val` can be defined globally or feature-wise. For most applications, domain knowledge becomes very important here. If a representative sample of the training set is available, we can always (naively) infer a `no_info_val` by taking the feature-wise median or mean:
```python
cem.fit(x_train, no_info_type='median')
```
... related to the **objective function**:
* `c_init` and `c_steps`: the multiplier $c$ of the first loss term is updated for `c_steps` iterations, starting at `c_init`. The first loss term encourages the perturbed instance to be predicted as a different class for a PN and the same class for a PP. If we find a candidate PN or PP for the current value of $c$, we reduce the value of $c$ for the next optimization cycle to put more emphasis on the regularization terms and improve the solution. If we cannot find a solution, $c$ is increased to put more weight on the prediction class restrictions of the PN and PP before focusing on the regularization.
* `kappa`: the first term in the loss function is defined by a difference between the predicted probabilities for the perturbed instance of the original class and the max of the other classes. $\kappa \geq 0$ defines a cap for this difference, limiting its impact on the overall loss to be optimized. Similar to the original paper, we set $\kappa$ to 0. in the examples.
* `beta`: $\beta$ is the $L_{1}$ loss term multiplier. A higher value for $\beta$ means more weight on the sparsity restrictions of the perturbations. Similar to the paper, we set $\beta$ to 0.1 for the MNIST and Iris datasets.
* `gamma`: multiplier for the optional $L_{2}$ reconstruction error. A higher value for $\gamma$ means more emphasis on the reconstruction error penalty defined by the auto-encoder. Similar to the paper, we set $\gamma$ to 100 when we have an auto-encoder available.
While the paper's default values for the loss term coefficients worked well for the simple examples provided in the notebooks, it is recommended to test their robustness for your own applications.
### Explanation
We can finally explain the instance:
```python
explanation = cem.explain(X)
```
The ```explain``` method returns a dictionary with the following *key: value* pairs:
* *X*: original instance
* *X_pred*: predicted class of original instance
* *PN* or *PP*: Pertinent Negative or Pertinant Positive
* *PN_pred* or *PP_pred*: predicted class of PN or PP
* *grads_graph*: gradient values computed from the TF graph with respect to the input features at the PN or PP
* *grads_num*: numerical gradient values with respect to the input features at the PN or PP
### Numerical Gradients
So far, the whole optimization problem could be defined within the internal TF graph, making autodiff possible. It is however possible that we do not have access to the model architecture and weights, and are only provided with a ```predict``` function returning probabilities for each class. We initialize the CEM in the same way as before:
```python
# define model
lr = load_model('iris_lr.h5')
predict_fn = lambda x: lr.predict(x)
# initialize CEM explainer
shape = (1,) + x_train.shape[1:]
mode = 'PP'
cem = CEM(predict_fn, mode, shape, kappa=0., beta=.1,
feature_range=(x_train.min(), x_train.max()),
eps=(1e-2, 1e-2), update_num_grad=100)
```
In this case, we need to evaluate the gradients of the loss function with respect to the input features numerically:
\begin{equation*} \frac{\partial L}{\partial x} = \frac{\partial L}{\partial p} \frac{\partial p}{\partial x} \end{equation*}
where $L$ is the loss function, $p$ the predict function and $x$ the input features to optimize. There are now 2 additional hyperparameters to consider:
* `eps`: a tuple to define the perturbation size used to compute the numerical gradients. `eps[0]` and `eps[1]` are used respectively for $^{\delta L}/_{\delta p}$ and $^{\delta p}/_{\delta x}$. `eps[0]` and `eps[1]` can be a combination of float values or numpy arrays. For `eps[0]`, the array dimension should be *(1 x nb of prediction categories)* and for `eps[1]` it should be *(1 x nb of features)*. For the Iris dataset, `eps` could look as follows:
```python
eps0 = np.array([[1e-2, 1e-2, 1e-2]]) # 3 prediction categories, equivalent to 1e-2
eps1 = np.array([[1e-2, 1e-2, 1e-2, 1e-2]]) # 4 features, also equivalent to 1e-2
eps = (eps0, eps1)
```
- `update_num_grad`: for complex models with a high number of parameters and a high dimensional feature space (e.g. Inception on ImageNet), evaluating numerical gradients can be expensive as they involve prediction calls for each perturbed instance. The `update_num_grad` parameter allows you to set a batch size on which to evaluate the numerical gradients, reducing the number of prediction calls required.
## Examples
[Contrastive Explanations Method (CEM) applied to MNIST](../examples/cem_mnist.nblink)
[Contrastive Explanations Method (CEM) applied to Iris dataset](../examples/cem_iris.nblink)
| github_jupyter |
수많은 마라톤 선수들이 마라톤에 참여하였습니다. 단 한 명의 선수를 제외하고는 모든 선수가 마라톤을 완주하였습니다.
마라톤에 참여한 선수들의 이름이 담긴 배열 participant와 완주한 선수들의 이름이 담긴 배열 completion이 주어질 때, 완주하지 못한 선수의 이름을 return 하도록 solution 함수를 작성해주세요.
* 마라톤 경기에 참여한 선수의 수는 1명 이상 100,000명 이하입니다.
* completion의 길이는 participant의 길이보다 1 작습니다.
* 참가자의 이름은 1개 이상 20개 이하의 알파벳 소문자로 이루어져 있습니다.
* 참가자 중에는 동명이인이 있을 수 있습니다.
* https://programmers.co.kr/learn/courses/30/lessons/42576
### 00_1 dict() & polynomial rolling
* dict 구조
* polynomial rolling을 통해 해쉬 충돌을 방지한 구조
```
def solution(participant, completion):
#no refactoring
#completion = ["leo","kiki","eden"]
p = 31
m = 0xfffff
x = 0
hash_table = dict()
# polynomial rolling hash function.
for i in participant:
mod_value=0
x = 0
for j in i:
mod_value = mod_value + ord(j)*pow(p,x)
x+=1
#print("par_hash("+i+") : "+ str((mod_value % m)))
if i in hash_table:
completion.remove(i)
#print("remove : "+i)
hash_table[i] = (mod_value % m)
#hash for completion
for k in completion:
mod_value=0
x = 0
for l in k:
mod_value = mod_value + ord(l)*pow(p,x)
x+=1
#print("com_hash("+k+") : "+ str((mod_value % m)))
if k in hash_table:
#print("del :("+k+")")
del hash_table[k]
for key in hash_table:
#print(key)
return key
```
### 00_2. dict & polynomial rolling & decimal key
* dict 구조
* polynomial rolling을 통해 해쉬 충돌을 방지한 구조
* map key 값으로 숫자 content 로 문자
```
def solution(participant, completion):
#no refactoring
#completion = ["leo","kiki","eden"]
p = 31
m = 0xfffff
x = 0
hash_table = dict()
# polynomial rolling hash function.
for i in participant:
mod_value=0
x = 0
for j in i:
mod_value = mod_value + ord(j)*pow(p,x)
x+=1
mod_result = (mod_value % m)
if mod_result in hash_table:
completion.remove(i)
#print("remove : "+i)
hash_table[mod_result] = i
#hash for completion
for k in completion:
mod_value=0
x = 0
for l in k:
mod_value = mod_value + ord(l)*pow(p,x)
x+=1
mod_result = (mod_value % m)
#print("com_hash("+k+") : "+ str((mod_value % m)))
if mod_result in hash_table:
#print("del :("+k+")")
del hash_table[mod_result]
for val in hash_table.values():
return val
solution([],[])
```
### 00_3. LIST INDEX & 참가 +1, 완주 -1
* 10만개 리스트 인덱스 생성
* MOD 연산을 통해 해당 INDEX 값에 +1
* 값이 있을경우 +1 뺼 경우 -1
* +1인 인덱스값을 출력
<b>평가(실패)</b>
(57.34ms, 26.3MB)
dict 보다 시간, 공간 효율성이 떨어짐
```
def solution(participant, completion):
#example
participant = ["leo","kiki","kiki","eden"]
completion = ["leo","kiki",]
#constant
p = 31
m = 0xfffff
str_list = list([0 for i in range(m)])
#update participant on str_list
for i in participant:
mod_value=0; x=0
for j in i:
mod_value = mod_value + ord(j)*pow(p,x)
x+=1
if str_list[(mod_value % m)]!= 0:
completion.remove(i)
print("dup remove : "+i+"("+str(mod_value % m)+")")
else:
str_list[(mod_value % m)]=i
print("add :"+i+"("+str(mod_value % m)+")")
#update completion on str_list
for k in completion:
mod_value=0; x=0
for z in k:
mod_value = mod_value + ord(z)*pow(p,x)
x+=1
str_list[(mod_value % m)]=0
print("remove :",k+"("+str(mod_value % m)+")")
for y in str_list:
if y!=0:
print(y)
solution([],[])
```
### 00_3. hash() & dict()
* dict 구조
* 내장 hash 함수를 이용하여 사용
<b>결과</b>
또 실패(0.01ms, 10.2MB)
```
def solution(participant, completion):
#no refactoring
participant = ["leo","kiki","eden","eden"]
completion = ["leo","kiki","eden"]
p = 31
m = 0xfffff
x = 0
hash_table = dict()
# polynomial rolling hash function.
for i in participant:
if hash(i) in hash_table:
completion.remove(i)
#print("remove : "+i)
else:
hash_table[hash(i)] = i
#hash for completion
for k in completion:
if hash(k) in hash_table:
del hash_table[hash(k)]
for val in hash_table.values():
return val
solution([],[])
```
### 00_4 hash() & dict() & string key
* dict 구조
* 내장 hash 함수를 이용하여 사용
* 재시도(dict key값 string으로)
결과 또 실패(0.01ms, 10.2MB)
```
def solution(participant, completion):
#no refactoring
participant = ["leo","kiki","eden","eden"]
completion = ["leo","kiki","eden"]
p = 31
m = 0xfffff
x = 0
hash_table = dict()
# polynomial rolling hash function.
for i in participant:
if i in hash_table:
completion.remove(i)
#print("remove : "+i)
else:
hash_table[i] = hash(i)
#hash for completion
for k in completion:
if k in hash_table:
del hash_table[k]
for val in hash_table.values():
return val
solution([],[])
```
### 00_s_1 collections.Counter
| github_jupyter |
```
import pandas as pd
import numpy as np
#?pd.read_csv
```
# Read Daily Shareprices and Quarterlöy Income statements (Source SimFin)
```
# Import the main functionality from the SimFin Python API.
import simfin as sf
# Import names used for easy access to SimFin's data-columns.
from simfin.names import *
sf.set_data_dir('data/')
#sf.load_api_key(path='../../keys/simfin.key', default_key='free')
sf.set_api_key(api_key='free')
df = sf.load(dataset='income', variant='quarterly', market='us')
stock = sf.load(dataset='shareprices', variant='daily', market='us')
```
# Re-read Files, Parse dates and cleanup column names
```
df = pd.read_csv("data/us-income-quarterly.csv",sep=';',header=0, parse_dates=[5,6,7])
stock = pd.read_csv("data/us-shareprices-daily.csv",sep=';',header=0, parse_dates=[2])
for s in [' ','.',',','(',')']:
df.columns = df.columns.str.replace(s, '')
stock.columns = stock.columns.str.replace(s, '')
symbol = 'MSFT'
msft = stock[stock.Ticker == symbol].set_index('Date')
df1 = df[df.Ticker == symbol].set_index('PublishDate')
df1['PublishDate'] = df1.index
df1.index.name = 'Date'
ms = msft.join(df1,how='left',rsuffix='inc').fillna(method='ffill')
ms.shape
ms = ms[~ms.Tickerinc.isnull()]
ms = ms.dropna(axis=1)
```
# Create Target to Predict
Target: Adjusted Close of Tomorrow
```
data = ms
tgt = 'AdjClose'
data[f"{tgt}_s1"] = ms[tgt].shift(-1)
data[f"spread"] = data[f"{tgt}_s1"] - data[tgt]
data[f"target"] = data[f"spread"] > 0
# Create Date/Timebased Features
data['weekday'] = data.index.weekday.astype(float)
data['dayofyear'] = data.index.dayofyear.astype(float)
data['month'] = data.index.month.astype(float)
# Create Hist. targes/values
for var in [f"spread",f"target"]:
for lag in np.arange(1,10):
data[f"{var}_lag{lag}"] = data[var].shift(lag)
for var in [f"target"]:
for lag in np.arange(1,10):
data[f"{var}_lag{lag}"] = data[var].shift(lag).astype(bool)
data["Days_Since_Report"] = (data.index-data["PublishDate"]).apply(lambda x: x.days).astype(float)
data["Days_Since_Report"]
data = data.dropna()
data.to_parquet("data/msft.parq")
import matplotlib.pyplot as plt
# create figure and axis objects with subplots()
fig,ax = plt.subplots()
# twin object for two different y-axis on the sample plot
ax2=ax.twinx()
display(ms[['Revenue','AdjClose']].corr())
ms['Revenue'].plot(ax=ax,color='red')
ms['AdjClose'].plot(ax=ax2)
fig,ax = plt.subplots()
# twin object for two different y-axis on the sample plot
ax2=ax.twinx()
display(ms[['NetIncome','AdjClose']].corr())
ms['NetIncome'].plot(ax=ax,color='red')
ms['AdjClose'].plot(ax=ax2)
```
| github_jupyter |
# Compound Video Player Widget
Most everything in this notebook is a work in progress.
```
import os
import IPython
import ipywidgets
# import nutmeg
from jpy_video import Video, TimeCode, compound
# Display cells full width
txt = """
<style>
div#notebook-container { width: 95%; }
div#menubar-container { width: 65%; }
div#maintoolbar-container { width: 99%; }
</style>
"""
IPython.display.display(IPython.display.HTML(data=txt))
```
# Setup
```
f = '/home/pierre/Projects/GoProHelper/notebooks/data/GOPR8802.intra.mp4'
os.path.isfile(f)
fps = 59.9
wid = compound.VideoPlayer(f, 1/fps)
wid.display()
wid.wid_video.
wid.wid_timecode.layout.top
wid.wid_info
wid.wid_timecode._model_module_version
```
# Components
```
# HTML5 video widget
wid_video = Video(f)
wid_video.set_property('controls', False)
# Timecode
wid_timecode = TimeCode()
# Slider
wid_slider = ipywidgets.FloatSlider(step=1/fps, continuous_update=True, readout=False)
wid_slider.layout.width='500pt'
# wid_button = ipywidgets.Button(icon='play') # http://fontawesome.io/icon/pause/
# self.wid_slider = ipywidgets.FloatSlider(min=0, max=60, step=timebase,
# continuous_update=True, orientation='horizontal',
# readout=False,
# slider_color='blue')
# self.wid_slider.layout.width = '50%'
```
# Assemble
```
wid_controls = ipywidgets.HBox(children=[wid_timecode, wid_slider])
wid_outer = ipywidgets.VBox(children=[wid_video, wid_controls])
# Link widgets at front end
ipywidgets.jslink((wid_video, 'current_time'), (wid_slider, 'value'))
ipywidgets.jsdlink((wid_video, 'current_time'), (wid_timecode, 'timecode'))
```
# Event Handlers
```
# def handle_any(wid, **event):
# """Respond to any event type
# """
# update_timecode(wid_time, wid_video.properties.currentTime)
def handle_displayed(wid, **event):
"""Do stuff that can only be done after widget is displayed
"""
wid.set_property('controls', False)
def handle_loaded_metadata(wid, **event):
"""Function to be called when sufficient video metadata has been loaded at the frontend
"""
pass
# print(wid.properties)
def handle_duration_change(wid, **event):
"""Update anything that depends on video duration
"""
wid_slider.max = wid.properties.duration
wid_video.on_displayed(handle_displayed)
# wid_video.on_event(handle_any)
wid_video.on_event(handle_loaded_metadata, 'loadedmetadata')
wid_video.on_event(handle_duration_change, 'loadedmetadata')
wid_video.on_event(handle_duration_change, 'durationchange')
```
| github_jupyter |
# Parse BPA for input into pipeline
We want files for each strain for each omics data and merge the conditions.
```
import matplotlib
import pandas as pd
# pd.set_option('display.max_rows', None)
infile_path = "../multi_omics_master_heatmap_table.tsv"
data = pd.read_csv(infile_path, sep="\t")
# two different metabolomics and proteomics platforms used
exp_old = data["Type_of_Experiment"].unique().tolist()
exp_new = ["Proteomics", "Proteomics", "Metabolomics",
"Metabolomics", "Transcriptomics"]
exp_map = dict(zip(exp_old, exp_new))
data.replace({"Type_of_Experiment": exp_map}, inplace=True)
# take one strain as example first
strain = "03-311-0071"
data = data[data["Strain"] == strain]
sample_info = data[[
"replicate_name", "Treatment_Type", "Type_of_Experiment"
]].drop_duplicates()
sample_info.set_index("replicate_name", inplace=True)
meta = data[data["Type_of_Experiment"] == "Metabolomics"]
prot = data[data["Type_of_Experiment"] == "Proteomics"]
tran = data[data["Type_of_Experiment"] == "Transcriptomics"]
meta = meta[[
"entity_id", "replicate_name", "Log_Counts"
]].pivot_table(
index="replicate_name",
columns="entity_id",
values="Log_Counts",
)
prot = prot[[
"entity_id", "replicate_name", "Log_Counts"
]].pivot_table(
index="replicate_name",
columns="entity_id",
values="Log_Counts",
)
tran = tran[[
"entity_id", "replicate_name", "Log_Counts"
]].pivot_table(
index="replicate_name",
columns="entity_id",
values="Log_Counts",
)
# check for missing values in data
missing = [
meta.isnull().sum(axis=1).sum(),
prot.isnull().sum(axis=1).sum(),
tran.isnull().sum(axis=1).sum(),
]
def remap_samples(sample_info, omics_block, omics_name):
index = sample_info[sample_info["Type_of_Experiment"] == omics_name]["Treatment_Type"]
mapped = pd.DataFrame(index).merge(omics_block, left_index=True, right_index=True)
mapped = mapped.reset_index().set_index("Treatment_Type").drop("replicate_name", axis=1)
mapped.sort_values("Treatment_Type", inplace=True)
mapped.reset_index(inplace=True)
treat = pd.DataFrame(mapped.Treatment_Type)
count = pd.DataFrame(pd.Series(mapped.index, dtype=str))
treat_count = pd.merge(treat, count, left_index=True, right_index=True)
mapped["tmp"] = treat_count["Treatment_Type"] + "_" + treat_count[0]
mapped.drop("Treatment_Type", axis=1, inplace=True)
mapped.set_index("tmp", inplace=True)
mapped.index.name = None
mapped.to_csv(".".join([omics_name, "tsv"]) ,sep="\t")
return mapped
mapped_meta = remap_samples(sample_info, meta, "Metabolomics")
mapped_prot = remap_samples(sample_info, prot, "Proteomics")
mapped_tran = remap_samples(sample_info, tran, "Transcriptomics")
pheno = pd.DataFrame(mapped_meta.index)
pheno["Growth_Media"] = pheno[0]
pheno.set_index(0, inplace=True)
pheno.index.name = None
pheno = pd.DataFrame(pheno.Growth_Media.str.split("_", expand=True)[0])
pheno.columns = ["Growth_Media"]
pheno.to_csv("targets.tsv", sep="\t")
!md5 Metabolomics.tsv Proteomics.tsv Transcriptomics.tsv targets.tsv ../multi_omics_master_heatmap_table.tsv
```
| github_jupyter |
# Chapter 4
This is the fourth in a series of notebooks related to astronomy data.
As a running example, we are replicating parts of the analysis in a recent paper, "[Off the beaten path: Gaia reveals GD-1 stars outside of the main stream](https://arxiv.org/abs/1805.00425)" by Adrian M. Price-Whelan and Ana Bonaca.
In the first lesson, we wrote ADQL queries and used them to select and download data from the Gaia server.
In the second lesson, we write a query to select stars from the region of the sky where we expect GD-1 to be, and save the results in a FITS file.
In the third lesson, we read that data back and identified stars with the proper motion we expect for GD-1.
## Outline
Here are the steps in this lesson:
1. Using data from the previous lesson, we'll identify the values of proper motion for stars likely to be in GD-1.
2. Then we'll compose an ADQL query that selects stars based on proper motion, so we can download only the data we need.
3. We'll also see how to write the results to a CSV file.
That will make it possible to search a bigger region of the sky in a single query.
After completing this lesson, you should be able to
* Convert proper motion between frames.
* Write an ADQL query that selects based on proper motion.
## Installing libraries
If you are running this notebook on Colab, you can run the following cell to install Astroquery and the other libraries we'll use.
If you are running this notebook on your own computer, you might have to install these libraries yourself. See the instructions in the preface.
```
# If we're running on Colab, install libraries
import sys
IN_COLAB = 'google.colab' in sys.modules
if IN_COLAB:
!pip install astroquery astro-gala pyia python-wget
```
## Reload the data
The following cells download the data from the previous lesson, if necessary, and load it into a Pandas `DataFrame`.
```
import os
from wget import download
filename = 'gd1_dataframe.hdf5'
path = 'https://github.com/AllenDowney/AstronomicalData/raw/main/data/'
if not os.path.exists(filename):
print(download(path+filename))
import pandas as pd
df = pd.read_hdf(filename, 'df')
centerline = pd.read_hdf(filename, 'centerline')
selected = pd.read_hdf(filename, 'selected')
```
## Selection by proper motion
At this point we have downloaded data for a relatively large number of stars (more than 100,000) and selected a relatively small number (around 1000).
It would be more efficient to use ADQL to select only the stars we need. That would also make it possible to download data covering a larger region of the sky.
However, the selection we did was based on proper motion in the `GD1Koposov10` frame. In order to do the same selection in ADQL, we have to work with proper motions in ICRS.
As a reminder, here's the rectangle we selected based on proper motion in the `GD1Koposov10` frame.
```
pm1_min = -8.9
pm1_max = -6.9
pm2_min = -2.2
pm2_max = 1.0
import astropy.units as u
pm1_rect = [pm1_min, pm1_min, pm1_max, pm1_max, pm1_min] * u.mas/u.yr
pm2_rect = [pm2_min, pm2_max, pm2_max, pm2_min, pm2_min] * u.mas/u.yr
```
The following figure shows:
* Proper motion for the stars we selected along the center line of GD-1,
* The rectangle we selected, and
* The stars inside the rectangle highlighted in green.
```
import matplotlib.pyplot as plt
pm1 = centerline['pm_phi1']
pm2 = centerline['pm_phi2']
plt.plot(pm1, pm2, 'ko', markersize=0.3, alpha=0.3)
pm1 = selected['pm_phi1']
pm2 = selected['pm_phi2']
plt.plot(pm1, pm2, 'gx', markersize=0.3, alpha=0.3)
plt.plot(pm1_rect, pm2_rect, '-')
plt.xlabel('Proper motion phi1 (GD1 frame)')
plt.ylabel('Proper motion phi2 (GD1 frame)')
plt.xlim(-12, 8)
plt.ylim(-10, 10);
```
Now we'll make the same plot using proper motions in the ICRS frame, which are stored in columns `pmra` and `pmdec`.
```
pm1 = centerline['pmra']
pm2 = centerline['pmdec']
plt.plot(pm1, pm2, 'ko', markersize=0.3, alpha=0.3)
pm1 = selected['pmra']
pm2 = selected['pmdec']
plt.plot(pm1, pm2, 'gx', markersize=1, alpha=0.3)
plt.xlabel('Proper motion ra (ICRS frame)')
plt.ylabel('Proper motion dec (ICRS frame)')
plt.xlim([-10, 5])
plt.ylim([-20, 5]);
```
The proper motions of the selected stars are more spread out in this frame, which is why it was preferable to do the selection in the GD-1 frame.
But now we can define a polygon that encloses the proper motions of these stars in ICRS,
and use the polygon as a selection criterion in an ADQL query.
SciPy provides a function that computes the [convex hull](https://en.wikipedia.org/wiki/Convex_hull) of a set of points, which is the smallest convex polygon that contains all of the points.
To use it, I'll select columns `pmra` and `pmdec` and convert them to a NumPy array.
```
import numpy as np
points = selected[['pmra','pmdec']].to_numpy()
points.shape
```
NOTE: If you are using an older version of Pandas, you might not have `to_numpy()`; you can use `values` instead, like this:
```
points = selected[['pmra','pmdec']].values
```
We'll pass the points to `ConvexHull`, which returns an object that contains the results.
```
from scipy.spatial import ConvexHull
hull = ConvexHull(points)
hull
```
`hull.vertices` contains the indices of the points that fall on the perimeter of the hull.
```
hull.vertices
```
We can use them as an index into the original array to select the corresponding rows.
```
pm_vertices = points[hull.vertices]
pm_vertices
```
To plot the resulting polygon, we have to pull out the x and y coordinates.
```
pmra_poly, pmdec_poly = np.transpose(pm_vertices)
```
The following figure shows proper motion in ICRS again, along with the convex hull we just computed.
```
pm1 = centerline['pmra']
pm2 = centerline['pmdec']
plt.plot(pm1, pm2, 'ko', markersize=0.3, alpha=0.3)
pm1 = selected['pmra']
pm2 = selected['pmdec']
plt.plot(pm1, pm2, 'gx', markersize=0.3, alpha=0.3)
plt.plot(pmra_poly, pmdec_poly)
plt.xlabel('Proper motion phi1 (ICRS frame)')
plt.ylabel('Proper motion phi2 (ICRS frame)')
plt.xlim([-10, 5])
plt.ylim([-20, 5]);
```
To use `pm_vertices` as part of an ADQL query, we have to convert it to a string.
We'll use `flatten` to convert from a 2-D array to a 1-D array, and `str` to convert each element to a string.
```
t = [str(x) for x in pm_vertices.flatten()]
t
```
Now `t` is a list of strings; we can use `join` to make a single string with commas between the elements.
```
pm_point_list = ', '.join(t)
pm_point_list
```
## Selecting the region
Let's review how we got to this point.
1. We made an ADQL query to the Gaia server to get data for stars in the vicinity of GD-1.
2. We transformed to `GD1` coordinates so we could select stars along the centerline of GD-1.
3. We plotted the proper motion of the centerline stars to identify the bounds of the overdense region.
4. We made a mask that selects stars whose proper motion is in the overdense region.
The problem is that we downloaded data for more than 100,000 stars and selected only about 1000 of them.
It will be more efficient if we select on proper motion as part of the query. That will allow us to work with a larger region of the sky in a single query, and download less unneeded data.
This query will select on the following conditions:
* `parallax < 1`
* `bp_rp BETWEEN -0.75 AND 2`
* Coordinates within a rectangle in the GD-1 frame, transformed to ICRS.
* Proper motion with the polygon we just computed.
The first three conditions are the same as in the previous query. Only the last one is new.
Here's the rectangle in the GD-1 frame we'll select.
```
phi1_min = -70
phi1_max = -20
phi2_min = -5
phi2_max = 5
phi1_rect = [phi1_min, phi1_min, phi1_max, phi1_max] * u.deg
phi2_rect = [phi2_min, phi2_max, phi2_max, phi2_min] * u.deg
```
Here's how we transform it to ICRS, as we saw in the previous lesson.
```
import gala.coordinates as gc
import astropy.coordinates as coord
corners = gc.GD1Koposov10(phi1=phi1_rect, phi2=phi2_rect)
corners_icrs = corners.transform_to(coord.ICRS)
```
To use `corners_icrs` as part of an ADQL query, we have to convert it to a string. Here's how we do that, as we saw in the previous lesson.
```
point_base = "{point.ra.value}, {point.dec.value}"
t = [point_base.format(point=point)
for point in corners_icrs]
point_list = ', '.join(t)
point_list
```
Now we have everything we need to assemble the query.
## Assemble the query
Here's the base string we used for the query in the previous lesson.
```
query_base = """SELECT
{columns}
FROM gaiadr2.gaia_source
WHERE parallax < 1
AND bp_rp BETWEEN -0.75 AND 2
AND 1 = CONTAINS(POINT(ra, dec),
POLYGON({point_list}))
"""
```
**Exercise:** Modify `query_base` by adding a new clause to select stars whose coordinates of proper motion, `pmra` and `pmdec`, fall within the polygon defined by `pm_point_list`.
```
# Solution
query_base = """SELECT
{columns}
FROM gaiadr2.gaia_source
WHERE parallax < 1
AND bp_rp BETWEEN -0.75 AND 2
AND 1 = CONTAINS(POINT(ra, dec),
POLYGON({point_list}))
AND 1 = CONTAINS(POINT(pmra, pmdec),
POLYGON({pm_point_list}))
"""
```
Here again are the columns we want to select.
```
columns = 'source_id, ra, dec, pmra, pmdec, parallax, parallax_error, radial_velocity'
```
**Exercise:** Use `format` to format `query_base` and define `query`, filling in the values of `columns`, `point_list`, and `pm_point_list`.
```
# Solution
query = query_base.format(columns=columns,
point_list=point_list,
pm_point_list=pm_point_list)
print(query)
```
Here's how we run it.
```
from astroquery.gaia import Gaia
job = Gaia.launch_job_async(query)
print(job)
```
And get the results.
```
candidate_table = job.get_results()
len(candidate_table)
```
## Plotting one more time
Let's see what the results look like.
```
x = candidate_table['ra']
y = candidate_table['dec']
plt.plot(x, y, 'ko', markersize=0.3, alpha=0.3)
plt.xlabel('ra (degree ICRS)')
plt.ylabel('dec (degree ICRS)');
```
Here we can see why it was useful to transform these coordinates. In ICRS, it is more difficult to identity the stars near the centerline of GD-1.
So, before we move on to the next step, let's collect the code we used to transform the coordinates and make a Pandas `DataFrame`:
```
from pyia import GaiaData
def make_dataframe(table):
"""Transform coordinates from ICRS to GD-1 frame.
table: Astropy Table
returns: Pandas DataFrame
"""
gaia_data = GaiaData(table)
c_sky = gaia_data.get_skycoord(distance=8*u.kpc,
radial_velocity=0*u.km/u.s)
c_gd1 = gc.reflex_correct(
c_sky.transform_to(gc.GD1Koposov10))
df = table.to_pandas()
df['phi1'] = c_gd1.phi1
df['phi2'] = c_gd1.phi2
df['pm_phi1'] = c_gd1.pm_phi1_cosphi2
df['pm_phi2'] = c_gd1.pm_phi2
return df
```
Here's how we can use this function:
```
candidate_df = make_dataframe(candidate_table)
```
And let's see the results.
```
x = candidate_df['phi1']
y = candidate_df['phi2']
plt.plot(x, y, 'ko', markersize=0.5, alpha=0.5)
plt.xlabel('ra (degree GD1)')
plt.ylabel('dec (degree GD1)');
```
We're starting to see GD-1 more clearly.
We can compare this figure with one of these panels in Figure 1 from the original paper:
<img height="150" src="https://github.com/datacarpentry/astronomy-python/raw/gh-pages/fig/gd1-2.png">
<img height="150" src="https://github.com/datacarpentry/astronomy-python/raw/gh-pages/fig/gd1-4.png">
The top panel shows stars selected based on proper motion only, so it is comparable to our figure (although notice that it covers a wider region).
In the next lesson, we will use photometry data from Pan-STARRS to do a second round of filtering, and see if we can replicate the bottom panel.
We'll also learn how to add annotations like the ones in the figure from the paper, and customize the style of the figure to present the results clearly and compellingly.
## Saving the DataFrame
Let's save this `DataFrame` so we can pick up where we left off without running this query again.
```
filename = 'gd1_candidates.hdf5'
candidate_df.to_hdf(filename, 'candidate_df', mode='w')
```
We can use `ls` to confirm that the file exists and check the size:
```
!ls -lh gd1_candidates.hdf5
```
If you are using Windows, `ls` might not work; in that case, try:
```
!dir gd1_candidates.hdf5
```
## CSV
Pandas can write a variety of other formats, [which you can read about here](https://pandas.pydata.org/pandas-docs/stable/user_guide/io.html).
We won't cover all of them, but one other important one is [CSV](https://en.wikipedia.org/wiki/Comma-separated_values), which stands for "comma-separated values".
CSV is a plain-text format with minimal formatting requirements, so it can be read and written by pretty much any tool that works with data. In that sense, it is the "least common denominator" of data formats.
However, it has an important limitation: some information about the data gets lost in translation, notably the data types. If you read a CSV file from someone else, you might need some additional information to make sure you are getting it right.
Also, CSV files tend to be big, and slow to read and write.
With those caveats, here's how to write one:
```
candidate_df.to_csv('gd1_candidates.csv')
```
We can check the file size like this:
```
!ls -lh gd1_candidates.csv
```
The CSV file about 2 times bigger than the HDF5 file (so that's not that bad, really).
We can see the first few lines like this:
```
!head -3 gd1_candidates.csv
```
The CSV file contains the names of the columns, but not the data types.
We can read the CSV file back like this:
```
read_back_csv = pd.read_csv('gd1_candidates.csv')
```
Let's compare the first few rows of `candidate_df` and `read_back_csv`
```
candidate_df.head(3)
read_back_csv.head(3)
```
Notice that the index in `candidate_df` has become an unnamed column in `read_back_csv`. The Pandas functions for writing and reading CSV files provide options to avoid that problem, but this is an example of the kind of thing that can go wrong with CSV files.
## Summary
In the previous lesson we downloaded data for a large number of stars and then selected a small fraction of them based on proper motion.
In this lesson, we improved this process by writing a more complex query that uses the database to select stars based on proper motion. This process requires more computation on the Gaia server, but then we're able to either:
1. Search the same region and download less data, or
2. Search a larger region while still downloading a manageable amount of data.
In the next lesson, we'll learn about the databased `JOIN` operation and use it to download photometry data from Pan-STARRS.
## Best practices
* When possible, "move the computation to the data"; that is, do as much of the work as possible on the database server before downloading the data.
* For most applications, saving data in FITS or HDF5 is better than CSV. FITS and HDF5 are binary formats, so the files are usually smaller, and they store metadata, so you don't lose anything when you read the file back.
* On the other hand, CSV is a "least common denominator" format; that is, it can be read by practically any application that works with data.
| github_jupyter |
## Preprocessing
```
# Import our dependencies
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
import os
import pandas as pd
import tensorflow as tf
from keras.callbacks import Callback
from tensorflow.keras.callbacks import ModelCheckpoint
# Import and read the charity_data.csv.
import pandas as pd
application_df = pd.read_csv("../Resources/charity_data.csv")
application_df.head()
# Drop the non-beneficial ID columns, 'EIN' and 'NAME'.
application_df = application_df.drop(["EIN","NAME", "USE_CASE", "INCOME_AMT"], 1)
# Determine the number of unique values in each column.
application_df.nunique()
# Look at APPLICATION_TYPE value counts for binning
application_counts = application_df['APPLICATION_TYPE'].value_counts()
application_counts
# Choose a cutoff value and create a list of application types to be replaced
# use the variable name `application_types_to_replace`
application_types_to_replace = list(application_counts[application_counts < 1000].index)
# Replace in dataframe
for app in application_types_to_replace:
application_df['APPLICATION_TYPE'] = application_df['APPLICATION_TYPE'].replace(app,"Other")
# Check to make sure binning was successful
application_df['APPLICATION_TYPE'].value_counts()
# Look at CLASSIFICATION value counts for binning
class_counts = application_df['CLASSIFICATION'].value_counts()
class_counts
# You may find it helpful to look at CLASSIFICATION value counts >1
class_count_great_one = class_counts[class_counts > 1]
class_count_great_one
# Choose a cutoff value and create a list of classifications to be replaced
# use the variable name `classifications_to_replace`
classifications_to_replace = list(class_counts[class_counts < 1750].index)
# Replace in dataframe
for cls in classifications_to_replace:
application_df['CLASSIFICATION'] = application_df['CLASSIFICATION'].replace(cls,"Other")
# Check to make sure binning was successful
application_df['CLASSIFICATION'].value_counts()
# Convert categorical data to numeric with `pd.get_dummies`
application_df_dummies = pd.get_dummies(application_df)
application_df_dummies.head()
# Split our preprocessed data into our features and target arrays
X = application_df_dummies.drop(['IS_SUCCESSFUL'], axis = 'columns').values
y = application_df_dummies['IS_SUCCESSFUL'].values
# Split the preprocessed data into a training and testing dataset
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state = 42)
# Create a StandardScaler instances
scaler = StandardScaler()
# Fit the StandardScaler
X_scaler = scaler.fit(X_train)
# Scale the data
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
```
## Compile, Train and Evaluate the Model
```
# Define the model - deep neural net, i.e., the number of input features and hidden nodes for each layer.
num_input_layer = len(X_train[0])
hidden_node_layers1 = 90
hidden_node_layers2 = 50
hidden_node_layers3 = 1
nn = tf.keras.models.Sequential()
# First hidden layer
nn.add(tf.keras.layers.Dense(units = hidden_node_layers1, input_dim = num_input_layer, activation = 'relu'))
# Second hidden layer
nn.add(tf.keras.layers.Dense(units = hidden_node_layers2, activation = 'relu'))
# Output layer
nn.add(tf.keras.layers.Dense(units = hidden_node_layers3, activation = 'sigmoid'))
# Check the structure of the model
nn.summary()
# Checkpoint file creation
os.makedirs('checkpoints/', exist_ok = True)
checkpoint_dir = r'checkpoints/Checkpoints.{epoch:02d}.hdf5'
# Compile the model
nn.compile(loss = 'binary_crossentropy', optimizer = 'adam', metrics = ['accuracy'])
checkpoint = ModelCheckpoint(filepath = checkpoint_dir, monitor = 'accuracy', verbose = 1, save_weights_only = True,
mode = 'auto', period = 1)
# Train the model
fit_model = nn.fit(X_train_scaled, y_train, epochs = 750, callbacks = [checkpoint])
# Evaluate the model using the test data
model_loss, model_accuracy = nn.evaluate(X_test_scaled,y_test,verbose=2)
print(f"Loss: {model_loss}, Accuracy: {model_accuracy}")
# Export our model to HDF5 file
nn.save('AlphabetSoupCharity_Optimization3.h5')
```
| github_jupyter |
# Exploring Seattle Weather
**Learning Objective:** Apply data visualization practices to study the weather in Seattle.
In this notebook, we will create visualizations to explore weather data for Seattle, taken from NOAA. The dataset is a CSV file with columns for the temperature (in Celsius), precipitation (in centimeters), wind speed (in meter/second), and weather type. We have one row for each day from January 1st, 2012 to December 31st, 2015. This notebook is based on the Seattle weather example in the [Altair](https://altair-viz.github.io/tutorials/exploring-weather.html) and [Vega-Lite](https://vega.github.io/vega-lite/tutorials/explore.html) documentation.
## Imports
```
import altair as alt
alt.data_transformers.enable('json')
from vega_datasets import data
```
## Data
Load the Seattle weather data from Altair:
```
df = data.seattle_weather()
df.head()
```
## Explore
Let’s start by looking at the precipitation, using tick marks to see the distribution of precipitation values:
```
alt.Chart(df).mark_tick().encode(
x='precipitation'
)
```
It looks as though precipitation is skewed towards lower values; that is, when it rains in Seattle, it usually doesn’t rain very much. It is difficult to see patterns across continuous variables, and so to better see this, we can create a histogram of the precipitation data. For this we first discretize the precipitation values by adding a binning to x. Additionally, we set our encoding channel y with the special field `*` that is aggregated with `count`. The result is a histogram of precipitation values:
```
alt.Chart(df, width=400, height=200).mark_bar().encode(
alt.X('precipitation', bin=True),
alt.Y('count(*):Q')
)
```
Next, let’s look at how precipitation in Seattle changes throughout the year. Altair natively supports dates and discretization of dates when we set the type to temporal (shorthand `T`). For example, in the following plot, we compute the total precipitation for each month. To discretize the data into months, we set the keyword `timeUnit="month"`:
```
alt.Chart(df, width=400, height=200).mark_line().encode(
alt.X('date:T', timeUnit='month'),
alt.Y('average(precipitation)')
)
```
This chart shows that in Seattle the precipitation in the winter is, on average, much higher than summer (an unsurprising observation to those who live there!). By changing the mapping of encoding channels to data features, you can begin to explore the relationships within the data.
When looking at precipitation and temperature, we might want to aggregate by year and month (`yearmonth`) rather than just month. This allows us to see seasonal trends, with daily variation smoothed out. We might also wish to see the maximum and minimum temperature in each month:
```
alt.Chart(df, width=400, height=200).mark_line().encode(
alt.X('date:T', timeUnit='yearmonth'),
alt.Y('max(temp_max)'),
)
```
In this chart, it looks as though the maximum temperature is increasing from year to year over the course of this relatively short baseline. To look closer into this, let’s instead look at the mean of the maximum daily temperatures for each year:
```
alt.Chart(df, width=400, height=200).mark_line().encode(
alt.X('date:T', timeUnit='year'),
alt.Y('mean(temp_max)'),
)
```
And in fact, the chart indicates that yes, the annual average of the daily high temperatures increased over the course of these four years, a fact that you can confirm for minimum daily temperatures as well.
You might also wonder how the variability of the temperatures changes throughout the year. For this, we have to add a computation to derive a new field. You can do with with Pandas:
```
df['temp_range'] = df.temp_max - df.temp_min
```
Now, let's look at the temperature range over time:
```
alt.Chart(df, width=400, height=200).mark_line().encode(
alt.X('date:T', timeUnit='month'),
y='mean(temp_range):Q'
)
```
Next we will explore the weather field, which encodes a categorical variable describing the weather on a given day. We might wish to know how different kinds of weather (e.g. sunny days or rainy days) are distributed throughout the year. To answer this, we can discretize the date by month and then count the number of records on the y-Axis. We then break down the bars by the weather type by mapping this column to a color channel. When a bar chart has a field mapped to color, Altair will automatically stack the bars atop each other:
```
alt.Chart(df).mark_bar().encode(
x=alt.X('date:T', timeUnit='month'),
y='count()',
color='weather',
)
```
The default color palette’s semantics might not match our expectation. For example, we probably do not expect “sun” (sunny) to be purple. We can tune the chart by providing a color scale range that maps the values from the weather field to meaningful colors, using standard hex color codes:
```
scale = alt.Scale(domain=['sun', 'fog', 'drizzle', 'rain', 'snow'],
range=['#e7ba52', '#c7c7c7', '#aec7e8', '#1f77b4', '#9467bd'])
```
This scale can be passed to the color encoding to be applied to the plot style. In addition, we can customize the titles for the axis and legend to make the meaning of the plot more clear:
```
alt.Chart(df, width=400, height=200).mark_bar().encode(
x=alt.X('date:T', timeUnit='month', axis=alt.Axis(title='Month of the year')),
y='count():Q',
color=alt.Color('weather', legend=alt.Legend(title='Weather type'), scale=scale),
)
```
Combining the above ideas lets us create any number of flexible visualizations of this dataset. For example, here is a plot that uses the customizations we have developed above to explore the relationship between weather, precipitation, maximum temperature, and temperature range, configured to use a larger canvas:
```
alt.Chart(df).mark_point().encode(
alt.X('temp_max', axis=alt.Axis(title='Maximum Daily Temperature (C)')),
alt.Y('temp_range', axis=alt.Axis(title='Daily Temperature Range (C)')),
alt.Color('weather', scale=scale),
size='precipitation',
).interactive()
```
This gives us even more insight into the weather patterns in Seattle: rainy and foggy days tend to be cooler with a narrower range of temperatures, while warmer days tend to be dry and sunny, with a wider spread between low and high temperature.
| github_jupyter |
# Introduction
This notebook uses a Faster RCNN model trained with [this notebook](https://colab.research.google.com/drive/13KQoEpEG8vP76_OXHPJjgqI6lOxsaLxY#scrollTo=S557gOSdQgAi). A model of this type should therefore be downloaded to use this demo. A test model can be downloaded [here](https://drive.google.com/file/d/1eqj1dcnDcjguICUyLHNfflyZsmF3BNA9/view?usp=sharing)
To use this demo on the validation data set, following the structure of the training data, please structure the data as described in the training notebook linked above and run the data section of this notebook.
For a single image downloaded from e.g. Google, the data section (except the dictionaries to convert classes to labels and vice versa) can be skipped.
Below is code to mount your Google drive if running this code in colab.
```
# mount drive, this should be set to the folder where the data folder is stored
from google.colab import drive
drive.mount('/content/drive')
cd 'drive/My Drive/DTU/Deep learning/FoodRecognition' #must match tour own path to the directory where data is stored
```
# Load necetities
## Packages
```
import torchvision.datasets as datasets
import numpy as np
import torch
import torchvision.transforms as transforms
import os
from engine import train_one_epoch, evaluate
import utils
import transforms as T
import matplotlib.pyplot as plt
import matplotlib.patches as patches
from PIL import Image, ImageDraw
```
## Data
```
# dicts for converting classes to labels
classes = ['__background__', 'boiled peas', 'boiled potatoes', 'chopped lettuce', 'fried egg',
'glass of milk', 'glass of water', 'meatballs', 'plain rice', 'plain spaghetti',
'slice of bread']
cls_to_label_dict = {j:i for i,j in enumerate(classes)}
label_to_cls_dict = {i:j for i,j in enumerate(classes)}
"""
Load data into a dataset class that can be used to train our model.
"""
torch.manual_seed(1)
class foodDataset(torch.utils.data.Dataset):
"""
Class to store the food data
"""
def __init__(self, data_root:str, datatype:str = "train", transforms = None):
self.data_root = data_root
self.transforms = transforms
self.splitImgPath = data_root + "VOCdevkit/VOC2007/ImageSets/Main/" + datatype + ".txt"
with open(self.splitImgPath, "r") as splitIdx:
self.imgNames = splitIdx.readlines()
# self.dataset = datasets.VOCDetection(data_root, year='2007', image_set = datatype, transform=self.transforms)
self.dataset = datasets.VOCDetection(data_root, year='2007', image_set = datatype)
def __getitem__(self, idx):
img = self.dataset[idx][0]
obs = self.dataset[idx][1]["annotation"]["object"]
num_objs = len(obs)
image_id = torch.tensor([int(os.path.splitext(self.dataset[idx][1]["annotation"]["filename"])[0])])
boxes = []
labels = torch.ones((num_objs,), dtype=torch.int64)
for i in range(num_objs):
xmin = int(obs[i]["bndbox"]["xmin"])
xmax = int(obs[i]["bndbox"]["xmax"])
ymin = int(obs[i]["bndbox"]["ymin"])
ymax = int(obs[i]["bndbox"]["ymax"])
boxes.append([xmin, ymin, xmax, ymax])
cls = obs[i]["name"]
try:
labels[i] *= cls_to_label_dict[cls]
except KeyError:
raise KeyError(f"Image {image_id} had an unavailable label: {cls} ")
boxes = torch.as_tensor(boxes, dtype=torch.float32)
area = (boxes[:, 3] - boxes[:, 1]) * (boxes[:, 2] - boxes[:, 0])
# suppose all instances are not crowd
iscrowd = torch.zeros((num_objs,), dtype=torch.int64)
target = {}
target["boxes"] = boxes
target["labels"] = labels
target["area"] = area
target["image_id"] = image_id
target["iscrowd"] = iscrowd
return (img, target) if self.transforms is None else self.transforms(img, target)
def __len__(self):
return len(self.imgNames)
def get_transform(train):
transforms = []
# converts the image, a PIL image, into a PyTorch Tensor
transforms.append(T.ToTensor())
if train:
# during training, randomly flip the training images
# and ground-truth for data augmentation
transforms.append(T.RandomHorizontalFlip(0.5))
return T.Compose(transforms)
# use our dataset and defined transformations
root = "data/VOC/"
dataset_train = foodDataset(root, "train", get_transform(train=False))
dataset_test = foodDataset(root, "test", get_transform(train=False))
dataset_val = foodDataset(root, "val", get_transform(train=False))
# define training and validation data loaders
data_loader_train = torch.utils.data.DataLoader(
dataset_train, batch_size=1, shuffle=False, num_workers=4,
collate_fn=utils.collate_fn)
data_loader_test = torch.utils.data.DataLoader(
dataset_test, batch_size=1, shuffle=False, num_workers=4,
collate_fn=utils.collate_fn)
data_loader_val = torch.utils.data.DataLoader(
dataset_val, batch_size=1, shuffle=False, num_workers=4,
collate_fn=utils.collate_fn)
```
## Model
Please change path model, to match the path to where the model is stored
```
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
print(f"Device used is: {device}")
model = torch.load("models/opt_model/model_epoch39.pth")
# evaluate(model, data_loader_val, device=device)
```
# Prediction
## Plot functions
```
# Create figure, axes, and display the image
def show_img(img, target = None, prediction = None, score_threshold = 0.2, dist_threshold = 100,
pred_col = '#000000', trg_col = '#7f32a8', txt_col = 'white', savename = None):
"""
Shows a test image with the boundary boxes and their labels
-----------------------------------------------------------
img: tensor with img values
target: the target boundary box, typically obtained directly from dataset, if None the box will not be drawn
prediction: the prediction obtained when giving the img to the model, if None the box will not be drawn
score_threshold: a value set to sort out box predictions with low scores
dist_threshold: a value set to sort out overlapping boxes by looking at the abosolute value of the difference between the box and boxes already found
pred_col: color used to draw prediction box
trg_col: color used to draw target box
txt_col: color used for displaying the class
savename: name used to save image with boxes, if None the iamge will not be saved
"""
# Draw the bounding box: target
def draw_target(target):
for i in range(len(target["boxes"])):
bbox = target["boxes"][i]
left = bbox[0]
top = bbox[1]
width = bbox[2]-bbox[0]
height = bbox[3]-bbox[1]
rect = patches.Rectangle((left, top), width, height,linewidth=2,edgecolor=trg_col,facecolor='none')
ax.add_patch(rect)
def draw_prediction(prediction, score_thres = 0.2, dist_thres = 100):
found = []
boxes = prediction[0]["boxes"]
for i in range(len(boxes)):
if prediction[0]["scores"][i] < score_thres:
break
if i != 0:
dist = []
for j in range(len(found)):
dist.append((boxes[i]-found[j]).abs().sum())
if any(x < dist_thres for x in dist):
break
bbox = boxes[i]
left = bbox[0]
top = bbox[1]
width = bbox[2]-bbox[0]
height = bbox[3]-bbox[1]
rect = patches.Rectangle((left, top), width, height,linewidth=2,edgecolor=pred_col,facecolor='none')
label = "%s (%.3f)" % (label_to_cls_dict[prediction[0]["labels"][i].item()],prediction[0]["scores"][i])
# print(label)
plt.text(left, top-20, label, color=txt_col)
ax.add_patch(rect)
found.append(bbox)
im = Image.fromarray(img.mul(255).permute(1, 2, 0).byte().numpy())
fig,ax = plt.subplots(1, figsize = (10, 10))
ax.imshow(im)
if target is not None:
draw_target(target)
if prediction is not None:
draw_prediction(prediction, score_threshold, dist_threshold)
if savename != None:
plt.savefig(savename)
```
## Try single test image
### From validation set
```
# pick one image from the test set (876, 842, 270, 296 used in report)
# img, target = dataset_val[876]
img, target = dataset_val[np.random.randint(0, len(dataset_val))] #get random image from validation set
# put the model in evaluation mode
model.eval()
with torch.no_grad():
prediction = model([img.to(device)])
# print("-"*25+"Prediction" + "-"*25)
# display(prediction)
# print("-"*25+"Target" + "-"*25)
# display(target)
# Show the bounding box
show_img(img, None, prediction, score_threshold = 0.7, dist_threshold = 24, txt_col = "black", pred_col = "blue", savename = None)
plt.show()
```
### Image from external sources
```
# load image and transform
img = Image.open("potato.jpg") #change to match own image
trans = transforms.ToTensor()
img = trans(img)
# predict
model.eval()
with torch.no_grad():
prediction = model([img.to(device)])
show_img(img, None, prediction, score_threshold = 0.7, dist_threshold = 24, txt_col = "black", pred_col = "blue", savename = None)
```
| github_jupyter |
# Mentoria Evolution - Python para Data Science
https://minerandodados.com.br
* Para executar uma célula digite **Control + enter** ou clique em **Run**.
* As celulas para rodar script Python devem ser do tipo code.
* As celular aceitam comandos python e já executam um "print" automáticamente.
## Mentoria Evolution - Aula 2
### Estruturas de Dados - Listas ###
** Sintaxe: nome = [elementos] **
```
idades = [25,38,18,47]
type(idades)
endereco = ['Rua dos Fulanos, Belo Horizonte.',2500]
endereco
```
- Acessando elementos
```
rua = endereco[0]
rua
numero = endereco[1]
print(rua,numero)
```
- Atualizando elementos
```
endereco[1] = 2750
endereco
endereco[1] = 2750
numero = endereco[1]
print(rua,numero)
```
- Operações com Listas
```
nomes = ['Felipe','Joao','Maria']
nomes
```
- Contando elementos
```
len(nomes)
```
- Verificando elementos
```
'Felipe' in nomes
```
- Valores máximo e mínimo
```
max(nomes)
min(nomes)
```
- Concatenando listas
```
nomes
nomes + ['Jose','Carla']
nomes = nomes + ['Jose','Carla']
print(nomes)
```
- Adiciona novos elementos
```
nomes.append('Marcelo')
nomes
```
- Índice de um determinado elemento
```
nomes.index('Joao')
```
- Removendo um elemento
```
nomes.remove('Marcelo')
nomes
```
- Contando elementos
```
nomes.count('Joao')
```
- Ordenando elementos
```
nomes.sort(reverse=True)
print(nomes)
```
### Estruturas de Dados - Dicionários ###
- Objeto do tipo Chave e Valor
- Sintaxe: nome = {'chave':'valor'}
```
dic = {'nome':'Rodrigo'}
pessoas = {
'Felipe':30, 'Fulana':18, 'Maria':55, 'Jose':80,
'valores':[1,3.5,400,5,6],
'pesos':{'Felipe':68,'Fulana':55}
}
pessoas
```
- Acessando o valor a partir de uma chave
```
pessoas['Jose']
pessoas['pesos']
```
- Dicionario Aninhado
```
pessoas = {
'Felipe': {'Idade':30,'Cidade':'Belo Horizonte','Peso':65},
'valores':[1,3.5,400,5,6],
}
pessoas
pessoas['Felipe']['Cidade']
pessoas['Felipe']['Peso']
Cadastro_pessoas = {
'Clientes': {'Cliente_01': {'Nome':'Rodrigo',
'Idade':30,
'Cidade':'Belo Horizonte',
'Peso':65
},
'Cliente_02': {'Nome': 'Felipe',
'e-mail': 'felipe10@gmail.com'
},
},
'valores':[1,3.5,400,5,6]}
Cadastro_pessoas['Clientes']['Cliente_01']['Cidade']
Cadastro_pessoas['Clientes']['Cliente_02']
```
**Métodos**
- keys() – Retorna as chaves do dicionário
```
pessoas
pessoas.keys()
```
- values() - Retorna os valores do dicionário
```
pessoas.values()
```
- get() – Retorna o valor de uma determinada chave senão existir retorna o valor passado como parâmetro
```
pessoas.get('Juca','Não existe')
```
- setdefault() - Retorna o valor da chave caso ela exista, senão inseri a chave e o valor no dicionário
```
pessoas.setdefault('Felipe',40)
pessoas
pessoas.setdefault('Marcos',30)
```
- items() – Retorna itens e valores
```
pessoas.items()
```
- clear() – Limpa o objeto dicionário
```
pessoas.clear()
pessoas
```
### Estruturas Condicionais e Loops ###
**Controles de fluxo**
**Sintaxe**<br>
**if <condição>:**
>**instruções**
```
if 1 > 10:
print('O numero 10 é maior que 1')
```
**Sintaxe**<br>
**if <condição>:**
>**instruções**
**else:**
>**instruções**
```
x=1
y=10
if x > y:
print("O numero em X é maior que Y")
else:
print('O numero em Y é maior que X')
```
**Trabalhando com loops FOR**
** Sintaxe: for <variável> in <condição>: instruções **
- Loop em elementos de uma lista
```
for i in [1,2,3,4,5]:
print ("Valor: %s" %i)
print (i + 2)
```
- Loop em elementos de uma string
```
for c in "Python é uma linguagem de programação":
print (c)
```
**Loops FOR aninhados**
**Sintaxe**<br>
**for <variável> in <condição>:**
>**instruções**<br>
>**for <variável> in <condição>:**
>>**instruções**
```
for i in ['1a Fase','2a Fase','3a Fase']:
print (i)
for y in ['manha','tarde','noite']:
print (y)
print(' ')
```
**Loops com While**
**Sintaxe**<br>
**while <condição>: **
>**instruções**
```
i = 0
while i < 10:
print (i)
i = i + 1
```
**Loops while com instrução else**
**Sintaxe**<br>
**while <condição>:**
>**instruções**
**else:**
>**instruções**
```
i = 0
while i < 10:
print (i)
i = i + 1
else:
print("Numero é maior ou igual a 10")
```
- Ao concluir, salve seu notebook e envie suas respostas para **contato@minerandodados.com.br**
| github_jupyter |
## Exercise 3 - Quantum error correction
### Importing Packages
```
from qiskit import QuantumCircuit, QuantumRegister, ClassicalRegister, Aer, transpile
from qc_grader import grade_ex3
import qiskit.tools.jupyter
from qiskit.test.mock import FakeTokyo
```
#### --------------------------------------------------------------------------------------------------------------------
### 1. Circuit
In this example we'll use 5 qubits that we'll call code qubits. To keep track of them, we'll define a special quantum register.
```
code = QuantumRegister(5,'code')
```
We'll also have an additional four qubits we'll call syndrome qubits.
```
syn = QuantumRegister(4,'syn')
```
Similarly we define a register for the four output bits, used when measuring the syndrome qubits.
```
out = ClassicalRegister(4,'output')
```
We consider the qubits to be laid out as follows, with the code qubits forming the corners of four triangles, and the syndrome qubits living inside each triangle.
```
c0----------c1
| \ s0 / |
| \ / |
| s1 c2 s2 |
| / \ |
| / s3 \ |
c3----------c4
```
For each triangle we associate a stabilizer operation on its three qubits. For the qubits on the sides, the stabilizers are $ZZZ$. For the top and bottom ones, they are $XXX$.
The syndrome measurement circuit corresponds to a measurement of these observables. This is done in a similar way to surface code stabilizers (in fact, this code is a small version of a surface code).
```
qc_syn = QuantumCircuit(code,syn,out)
# Left ZZZ
qc_syn.cx(code[0],syn[1])
qc_syn.cx(code[2],syn[1])
qc_syn.cx(code[3],syn[1])
#qc_syn.barrier()
# Right ZZZ
qc_syn.swap(code[1],code[2])
qc_syn.cx(code[2],syn[2])
qc_syn.swap(code[1],code[2])
qc_syn.cx(code[2],syn[2])
qc_syn.cx(code[4],syn[2])
#qc_syn.barrier()
# Top XXX
qc_syn.h(syn[0])
qc_syn.cx(syn[0],code[0])
qc_syn.cx(syn[0],code[1])
qc_syn.cx(syn[0],code[2])
qc_syn.h(syn[0])
#qc_syn.barrier()
# Bottom XXX
qc_syn.h(syn[3])
qc_syn.cx(syn[3],code[2])
qc_syn.cx(syn[3],code[3])
qc_syn.cx(syn[3],code[4])
qc_syn.h(syn[3])
#qc_syn.barrier()
# Measure the auxilliary qubits
qc_syn.measure(syn,out)
qc_syn.draw('mpl')
qc_init = QuantumCircuit(code,syn,out)
qc_init.h(syn[0])
qc_init.cx(syn[0],code[0])
qc_init.cx(syn[0],code[1])
qc_init.cx(syn[0],code[2])
qc_init.cx(code[2],syn[0])
qc_init.h(syn[3])
qc_init.cx(syn[3],code[2])
qc_init.cx(syn[3],code[3])
qc_init.cx(syn[3],code[4])
qc_init.cx(code[4],syn[3])
#qc_init.barrier()
qc_init.draw('mpl')
```
The initialization circuit prepares an eigenstate of these observables, such that the output of the syndrome measurement will be `0000` with certainty.
```
qc = qc_init.compose(qc_syn)
display(qc.draw('mpl'))
job = Aer.get_backend('qasm_simulator').run(qc)
job.result().get_counts()
```
#### --------------------------------------------------------------------------------------------------------------------
### 2. Error Qubits
```
error_qubits = [0,4]
```
Here 0 and 4 refer to the positions of the qubits in the following list, and hence are qubits `code[0]` and `code[4]`.
```
qc.qubits
```
To check that the code does as we require, we can use the following function to create circuits for inserting artificial errors. Here the errors we want to add are listed in `errors` as a simple text string, such as `x0` for an `x` on `error_qubits[0]`.
```
def insert(errors,error_qubits,code,syn,out):
qc_insert = QuantumCircuit(code,syn,out)
if 'x0' in errors:
qc_insert.x(error_qubits[0])
if 'x1' in errors:
qc_insert.x(error_qubits[1])
if 'z0' in errors:
qc_insert.z(error_qubits[0])
if 'z1' in errors:
qc_insert.z(error_qubits[1])
return qc_insert
```
Rather than all 16 possibilities, let's just look at the four cases where a single error is inserted.
```
for error in ['x0','x1','z0','z1']:
qc = qc_init.compose(insert([error],error_qubits,code,syn,out)).compose(qc_syn)
job = Aer.get_backend('qasm_simulator').run(qc)
print('\nFor error '+error+':')
counts = job.result().get_counts()
for output in counts:
print('Output was',output,'for',counts[output],'shots.')
```
### 2. Backend
```
backend = FakeTokyo()
backend
```
As a simple idea of how our original circuit is laid out, let's see how many two-qubit gates it contains.
```
qc = qc_init.compose(qc_syn)
qc = transpile(qc, basis_gates=['u','cx'])
qc.num_nonlocal_gates()
qc1 = transpile(qc,backend,basis_gates=['u','cx'], optimization_level=3)
qc1.num_nonlocal_gates()
```
#### --------------------------------------------------------------------------------------------------------------------
### 3. Initial Layout
```
initial_layout = [0,2,6,10,12,1,5,7,11]
qc2 = transpile(qc,backend,initial_layout=initial_layout, basis_gates=['u','cx'], optimization_level=3)
qc2.num_nonlocal_gates()
```
#### --------------------------------------------------------------------------------------------------------------------
### 4. Grading
```
grade_ex3(qc_init,qc_syn,error_qubits,initial_layout)
```
#### --------------------------------------------------------------------------------------------------------------------
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import sys
from pathlib import Path
sys.path.append(str(Path().cwd().parent))
import matplotlib.pyplot as plt
import load_dataset
import plotting
dataset = load_dataset.Dataset('../data/dataset/')
```
## Разбор (не запускать ячейки)
### Выгрузим временной ряд индекса Доу-Джонса, один из ярких примеров рядов, который, будучи случайным блужданием, очень сложно предсказать лучше наивного метода.
```
ts = dataset['dow_jones_3.csv']
ts.plot()
len(ts)
```
### Разобьем ряд на трейн и тест и попробуем зафитить нашу "сложную" ML модель TimeSeriesPredictor с базой в виде линейной регрессии.
```
ts_train, ts_test = ts[:200], ts[200:]
from model import TimeSeriesPredictor
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.linear_model import LinearRegression
predictor = TimeSeriesPredictor(
num_lags=12,
granularity='P1D',
model=LinearRegression
)
predictor.fit(ts_train)
```
### Сделаем one step ahead предсказания на тесте, предсказание выглядит неплохо.
```
preds = predictor.predict_batch(ts_train, ts_test)
# кажется, получилось весьма неплохо
ts.plot()
preds.plot()
```
### Давайте посмотрим на абсолютные метрики.
```
from sklearn.metrics import mean_squared_error, mean_absolute_error
mean_squared_error(preds, ts_test)
mean_absolute_error(preds, ts_test)
# выдать какой-то вердикт по ним, впрочем, сложно
```
### Остатки тоже выглядят неплохо
```
(ts_test-preds).plot()
```
### Дикки-Фуллер говорит о стационарности остатков.
```
from statsmodels.tsa.stattools import adfuller
adfuller(ts_test-preds)[1]
```
### По всем признакам, мы получили неплохую модель. Пора выкатывать на прод!
### А теперь давайте сравним с наивным предсказанием.
```
from metrics import mase
mase(preds, ts_test, method='naive')
```
### Модель показывает себя на 10% хуже наивного предсказания!

## Задание
Реализуйте метрику MASE, которая записывается, как
`mase = MAE(y_pred, y_true) / MAE(y_reference, y_true)`, где
`y_pred` - one step ahead предсказания вашей модели на некой тестовой выборке
`y_true` - реальные значения тестовой выборки
`y_reference` - предсказания сравниваемой модели на той же тестовой выборке
* чтобы получить y_reference для наивного метода, достаточно только сдвинуть y_true (потерей одной точки в данном задании перенебрегаем
* чтобы получить y_reference для всех остальных методов, необходимо передать в функцию историю ряда до y_true.
* реализацию методов mean, median можете сделать как на всем ts, так и на произвольном скользящем окне по вашему усмотрению
```
def mase(y_pred, y_true, method='naive', ts=None):
"""
Реализуйте метрику mase для сравнения предсказаний y_pred вашего метода и y_method, являющегося
предсказанием метода, указанного в параметре method.
Формула для расчета mase = MAE(y_pred, y_true) / MAE(y_reference, y_true),
где MAE - mean absolute error - средняя абсолютная ошибка.
В качестве method реализуйте поддержку {'naive', 'mean', 'median'}.
Для наивного метода достаточно использовать только y_true.
Для методов средней и медианы понадобится использовать временной ряд ts.
"""
'<your code here>'
return
mase(preds, ts_test)
```
| github_jupyter |
# Step 4: Feature Engineering
Use the code below to run TensorFlow Transform on some example data using the schema from your pipeline. Start by importing and opening the metadata store.
```
from __future__ import print_function
import os
import tempfile
import pandas as pd
import tfx_utils
import tensorflow_transform as tft
from tensorflow_transform import beam as tft_beam
from tfx.utils import io_utils
from tensorflow_metadata.proto.v0 import schema_pb2
# For DatasetMetadata boilerplate
from tensorflow_transform.tf_metadata import dataset_metadata
from tensorflow_transform.tf_metadata import dataset_schema
from tensorflow_transform.tf_metadata import schema_utils
def _make_default_sqlite_uri(pipeline_name):
return os.path.join(os.environ['HOME'], 'airflow/tfx/metadata', pipeline_name, 'metadata.db')
def get_metadata_store(pipeline_name):
return tfx_utils.TFXReadonlyMetadataStore.from_sqlite_db(_make_default_sqlite_uri(pipeline_name))
pipeline_name = 'TFX_Example'
pipeline_db_path = _make_default_sqlite_uri(pipeline_name)
print('Pipeline DB:\n{}'.format(pipeline_db_path))
store = get_metadata_store(pipeline_name)
```
Get the schema URI from the metadata store
```
# Get the schema URI from the metadata store
schemas = store.get_artifacts_of_type_df(tfx_utils.TFXArtifactTypes.SCHEMA)
assert len(schemas.URI) == 1
schema_uri = schemas.URI.iloc[0] + 'schema.pbtxt'
print ('Schema URI:\n{}'.format(schema_uri))
```
Get the schema that was inferred by TensorFlow Data Validation
```
schema_proto = io_utils.parse_pbtxt_file(file_name=schema_uri, message=schema_pb2.Schema())
feature_spec, domains = schema_utils.schema_as_feature_spec(schema_proto)
legacy_metadata = dataset_metadata.DatasetMetadata(dataset_schema.from_feature_spec(feature_spec, domains))
```
Define features and create functions for TensorFlow Transform
```
# Need to re-import because currently taxi_utils.py imports as `transform` not `tft`
import tensorflow_transform as transform
import tensorflow as tf
# Categorical features are assumed to each have a maximum value in the dataset.
_MAX_CATEGORICAL_FEATURE_VALUES = [24, 31, 12]
_CATEGORICAL_FEATURE_KEYS = [
'trip_start_hour', 'trip_start_day', 'trip_start_month',
'pickup_census_tract', 'dropoff_census_tract', 'pickup_community_area',
'dropoff_community_area'
]
_DENSE_FLOAT_FEATURE_KEYS = ['trip_miles', 'fare', 'trip_seconds']
# Number of buckets used by tf.transform for encoding each feature.
_FEATURE_BUCKET_COUNT = 10
_BUCKET_FEATURE_KEYS = [
'pickup_latitude', 'pickup_longitude', 'dropoff_latitude',
'dropoff_longitude'
]
# Number of vocabulary terms used for encoding VOCAB_FEATURES by tf.transform
_VOCAB_SIZE = 1000
# Count of out-of-vocab buckets in which unrecognized VOCAB_FEATURES are hashed.
_OOV_SIZE = 10
_VOCAB_FEATURE_KEYS = [
'payment_type',
'company',
]
# Keys
_LABEL_KEY = 'tips'
_FARE_KEY = 'fare'
def _transformed_name(key):
return key + '_xf'
def _transformed_names(keys):
return [_transformed_name(key) for key in keys]
def _fill_in_missing(x):
"""Replace missing values in a SparseTensor.
Fills in missing values of `x` with '' or 0, and converts to a dense tensor.
Args:
x: A `SparseTensor` of rank 2. Its dense shape should have size at most 1
in the second dimension.
Returns:
A rank 1 tensor where missing values of `x` have been filled in.
"""
default_value = '' if x.dtype == tf.string else 0
return tf.squeeze(
tf.sparse_to_dense(x.indices, [x.dense_shape[0], 1], x.values,
default_value),
axis=1)
def preprocessing_fn(inputs):
"""tf.transform's callback function for preprocessing inputs.
Args:
inputs: map from feature keys to raw not-yet-transformed features.
Returns:
Map from string feature key to transformed feature operations.
"""
outputs = {}
for key in _DENSE_FLOAT_FEATURE_KEYS:
# Preserve this feature as a dense float, setting nan's to the mean.
outputs[_transformed_name(key)] = transform.scale_to_z_score(
_fill_in_missing(inputs[key]))
for key in _VOCAB_FEATURE_KEYS:
# Build a vocabulary for this feature.
outputs[_transformed_name(key)] = transform.compute_and_apply_vocabulary(
_fill_in_missing(inputs[key]),
top_k=_VOCAB_SIZE,
num_oov_buckets=_OOV_SIZE)
for key in _BUCKET_FEATURE_KEYS:
outputs[_transformed_name(key)] = transform.bucketize(
_fill_in_missing(inputs[key]), _FEATURE_BUCKET_COUNT)
for key in _CATEGORICAL_FEATURE_KEYS:
outputs[_transformed_name(key)] = _fill_in_missing(inputs[key])
# Was this passenger a big tipper?
taxi_fare = _fill_in_missing(inputs[_FARE_KEY])
tips = _fill_in_missing(inputs[_LABEL_KEY])
outputs[_transformed_name(_LABEL_KEY)] = tf.where(
tf.is_nan(taxi_fare),
tf.cast(tf.zeros_like(taxi_fare), tf.int64),
# Test if the tip was > 20% of the fare.
tf.cast(
tf.greater(tips, tf.multiply(taxi_fare, tf.constant(0.2))), tf.int64))
return outputs
```
Display the results of transforming some example data
```
from IPython.display import display
with tft_beam.Context(temp_dir=tempfile.mkdtemp()):
raw_examples = [
{
"fare": [100.0],
"trip_start_hour": [12],
"pickup_census_tract": ['abcd'],
"dropoff_census_tract": [12345.0], # No idea why this is a float
"company": ['taxi inc.'],
"trip_start_timestamp": [123456],
"pickup_longitude": [12.0],
"trip_start_month": [5],
"trip_miles": [8.0],
"dropoff_longitude": [12.05],
"dropoff_community_area": [123],
"pickup_community_area": [123],
"payment_type": ['visa'],
"trip_seconds": [600.0],
"trip_start_day": [12],
"tips": [10.0],
"pickup_latitude": [80.0],
"dropoff_latitude": [80.01],
}
]
(transformed_examples, transformed_metadata), transform_fn = (
(raw_examples, legacy_metadata)
| 'AnalyzeAndTransform' >> tft_beam.AnalyzeAndTransformDataset(
preprocessing_fn))
display(pd.DataFrame(transformed_examples))
```
| github_jupyter |
# Simulating Clifford randomized benchmarking using a generic noise model
This tutorial demonstrates shows how to simulate Clifford RB sequences using arbitrary $n$-qubit process matrices. In this example $n=2$.
```
import pygsti
import numpy as np
```
## Get some CRB circuits
First, we follow the [Clifford RB](../CliffordRB.ipynb) tutorial to generate a set of sequences. If you want to perform Direct RB instead, just replace this cell with the contents of the [Direct RB](../DirectRB.ipynb) tutorial up until the point where it creates `circuitlist`:
```
#Specify the device to be benchmarked - in this case 2 qubits
nQubits = 2
qubit_labels = [0,1]
gate_names = ['Gxpi2', 'Gypi2','Gcphase']
availability = {'Gcphase':[(0,1)]}
pspec = pygsti.obj.ProcessorSpec(nQubits, gate_names, availability=availability,
qubit_labels=qubit_labels)
#Specify RB parameters (k = number of repetitions at each length)
lengths = [0,1,2,4,8,16]
k = 10
subsetQs = [0,1]
randomizeout = False # ==> all circuits have the *same* ideal outcome (the all-zeros bitstring)
#Generate clifford RB circuits
exp_design = pygsti.protocols.CliffordRBDesign(pspec, lengths, k, qubit_labels=subsetQs, randomizeout=randomizeout)
#Collect all the circuits into one list:
circuitlist = exp_design.all_circuits_needing_data
```
## Create a model to simulate these circuits
Now we need to create a model that can simulate circuits like this. Two things to note:
1. RB circuits use our "multi-qubit" gate naming, so you have gates like `Gxpi2:0` and `Gcphase:0:1`.
2. RB circuits do gates in parallel (this only matters for >1 qubits), so you have layers like `[Gypi2:0Gypi2:1]`
In this example, we'll make a model with $n$-qubit process matrices, so this will be practically limited to small $n$. We construct a model based on our standard 2-qubit X, Y, and CPHASE model, since this
has all the appropriate gates. To get a model with the multi-qubit labels, we'll use a standard multi-qubit "model-pack", which packages a `Model` object with relevant meta information needed by other protocols (like GST). If you can't start with a standard model, then you'll need to create an `ExplicitOpModel` object of the appropriate dimension (see the [explicit models tutorial](../../objects/ExplicitModel.ipynb)) and assign to it gates with are, for instance `('Gxpi2',0)` rather than just `'Gxpi2'`.
Here we import the `smq2Q_XYCPHASE` model pack:
```
from pygsti.modelpacks import smq2Q_XYCPHASE
```
We'll depolarize the target model and set one of the process matrices to a custom value as a demonstration. Here is where you can set any 2-qubit process matrices you want to any of the gates:
```
myModel = smq2Q_XYCPHASE.target_model().depolarize(op_noise=0.01, spam_noise=0.01)
myModel[('Gx',0)] = np.kron(
np.array([[1, 0, 0, 0],
[0, 0.85, 0, 0],
[0, 0, 0, -0.85],
[0, 0, 0.85, 0]], 'd'),
np.array([[1, 0, 0, 0],
[0, 0.95, 0, 0],
[0, 0, 0.95, 0],
[0, 0, 0, 0.95]], 'd'))
#print(myModel[('Gx',0)])
myModel.operations.keys() #voila! you have gates like "Gx:0" rather than "Gxi"
```
Since, `ExplicitOpModel` objects (e.g., those in the model packs) don't know how to automatically simulate multiple gates in parallel (you'd need to add an operation for each layer explicitly), we'll just *serialize* the circuits so they don't contain any parallel gates. This addresses point 2) above. Then we can simulate our circuits using our `ExplicitOpModel`, creating a `DataSet`.
```
serial_circuits = [c.serialize() for c in circuitlist]
ds = pygsti.construction.generate_fake_data(myModel, serial_circuits, 100, seed=1234)
#See how the DataSet contains serialized circuits (just printing the first several layers for clarity)
print(ds.keys()[10][0:7])
print(circuitlist[10][0:5])
```
Next, we "un-serialize" the circuits in the resulting data-set (`ds`) using the `process_circuits` function. This is needed because the RB experiment design calls for the original (parallel-gate) circuits, not the serialized ones. The cell below updates the circuits for all the data we just simulated so the data counts are associated with the original circuits.
```
#map circuits in dataset back to non-serialized RB circuits that we expect to have data for:
unserialize_map = { serial_circuit: orig_circuit for (serial_circuit, orig_circuit) in zip(serial_circuits, circuitlist)}
ds = ds.copy_nonstatic()
ds.process_circuits(lambda c: unserialize_map[c])
ds.done_adding_data()
```
## Running RB on the simulated `DataSet`
To run an RB analysis, we just package up the experiment design and data set into a `ProtocolData` object and give this to a `RB` protocol's `run` method. This returns a `RandomizedBenchmarkingResults` object that can be used to plot the RB decay curve. (See the [RB analysis tutorial](../RBAnalysis.ipynb) for more details.)
```
data = pygsti.protocols.ProtocolData(exp_design, ds)
results = pygsti.protocols.RB().run(data)
%matplotlib inline
results.plot()
```
| github_jupyter |
# Introduction to programming 1
João Pedro Malhado and Clyde Fare, Imperial College London (contact: [chemistry-git@imperial.ac.uk](mailto:chemistry-git@imperial.ac.uk))
This notebook is licensed under a [Creative Commons Attribution 4.0 (CC-by) license](http://creativecommons.org/licenses/by/4.0/)
This is an interactive tutorial! As you go through it any time you see something that looks like this:
a = "Hello"
that's followed by an empty *code cell* (a light grey rectangle with a label like *"In[ ]"*), you should type the expression in the code cell, hit Shift+Return to *execute* it, and note the output.
No copying and pasting! You'll learn the concepts better if you type them out yourself.
Learning how to program is a skill that is developed by experimenting, trying ideas out, thinking about what works and what doesn't, and asking for help. You are strongly encouraged to open new cells in the notebook and trying things out. The computer will not complain!
## Overview
In this workshop we will distinguish different data types that can be manipulated in computer programs, and what kind of operations can be performed on each type. These will be the building blocks we will be using when constructing our programs. As these workshops develop we will see how to bring things together and construct functionality.
## Data types
At the centre of all computer programs is manipulating and performing logical operations on data and data structures. Data structures can be very complex and abstract, but we'll start with simple data types which are present in almost all programming languages. We will first identify what they are, and look at what simple operations can be performed on them.
### Different types of numbers
In scientific computing, the most important data types are numbers. In a similar way to mathematics where we can distinguish different types of numbers (integers, rationals, reals, complex, etc...), in computing we also have different types of numbers.
Is there a difference between 1 and 1.0?
In order to enquire the data type of a given object we can use a function (more on what a function is later on) called *type* that tells you what kind of thing -- what data type -- something is. We can check for ourselves that Python considers 1 and 1.0 to be different data types:
type(1)
type(1.0)
Python is identifying the two numbers above as an *integer* and a *[floating point number](https://en.wikipedia.org/wiki/Floating_point)*. This difference is related to the way computers represent numbers, and can be important in some operation on some programming languages. Fortunately in Python (version 3 or above) we will not have to worry too much about them.
What do you think the types of -1, 0.2, 9753 and 6.626e-34 are?
Check your guesses in the cell below.
The last number of the sequence above is the Plank constant to 4 significant figures: 6.626×10<sup>-34</sup>. Therefore we see that we can use the *e* to specify scientific notation.
What would be the result of
3e-3
If we change the sign
3e3
What is the type of *3e3*?
Is it the same as the type of 300? Test it out!
The last numerical type available in Python is that of complex numbers. Python uses the convention followed in engineering, where the imaginary constant is represented by the letter *j*.
type(1+2j)
What will the type of the difference of the complex number 1+2j and the pure imaginary number 2j be?
#### Operations with numbers
The simplest programming operations you can do with numbers are the same as the ones you would do in a simple calculator. In fact you can use the notebook as a sophisticated calculator. Let's try a few.
1+1
Try a couple of expressions in the cell below. Note that you can use brackets to specify the order of the operations.
(5+7)/2
Calculate the energy of a photon of violet light with a wavelength of 440 nm (you can remind yourself of the value of the [constants you need](http://physics.nist.gov/cgi-bin/cuu/Category?view=html&Universal.x=70&Universal.y=18)).
Two \* can be used for exponentiation. We can try it on a complex number.
(3-1j)**2
Or to do a fourth root
16**(1/4)
One operation that we learn in primary school, seldom used in our daily lives, but is somewhat surprisingly useful in programming is the modulo operation which calculates the remainder of the division of two numbers. This is done using the '%' symbol
3%2
The mathematical operations available in de bare Python language are relatively simple, and common functions like trigonometric functions, exponentials or logarithms are not available by default. These can be made available by loading special modules (to be discussed further later). By loading the [math module](https://docs.python.org/3/library/math.html), many more mathematical functions and constants are made available.
import math
math.sin(3*math.pi/2)
The math module is loaded when you use the pylab environment you may already be familiar with. However, we shall delay the discussion of these extensions until later, as we want to focus on the main characteristics of the programming language.
### Strings of text
Another very important data type we are interested in manipulating is text strings. This is evident when dealing with a text editor, a spell checker, or even how you are typing into this notebook, but even in simple computational tasks text strings will appear often.
Text strings appear in quotations. If you type a simple
Hello
below, Python will not understand what you mean and output an error message
If you type it in quotes
'Hello'
it will tolerate it much better
And if you ask what type this quoted thing is, Python will inform you that it is a *string*
type('Hello')
Note that the difference in type is related with the use of quotes and not the use of letters
type(1.2)
type('1.2')
```
type("We can even make 1 single string with spaces, punctuation and numbers such as 0.00729")
```
Note that in the string above we used double quotes instead of single quotes. Both are indeed equivalent in Python, but we must be consistent.
In the English language, because of the use of the apostrophe, single quotes can be problematic
'I wouldn't want to contradict you'
In the example above, Python would think the string finishes half way through the second word, and it will not understand what the rest of the command means. The use of double quotes helps in this case.
"I wouldn't want to contradict you"
Or to use triple quotes ('''), which have the advantage of being used for long strings that do not fit in one line. You could put a full novel in triple quotes.
```
'''This is a multiline string.
With some horrible characters that would normally create complications: '{}"/
As you can see, it extends over more than one line'''
```
#### Operations with strings
We know what to expect from the operation *1+1* and we tested this operation above. We have also seen that *1* and *"1"* are essentially different objects, for a computer in general, and Python in this case. So let us see what the following operation will result into
"1"+"1"
Any ideas on what happened, or is the numeracy of the computer is broken beyond repair? It is certainly helpful to query what is the type of the result of the operation we just did.
type("1"+"1")
Are things a bit more clear? What was achieved with the operation + acting on two strings ("1" and "1") was a string concatenation, i.e. we joined the two strings together. Let's try to do that again
"Hello "+"mate!"
We note here that there is no difference in the symbol we use for summation +, and the symbol we use for string concatenation +. Yet the operations are essentially different, because the objects we are operating on are of different types. Summing (in a mathematical sense) two strings is an ill defined operation, and concatenating two numbers together would not be a very useful thing to do.
Similarly, it is not defined the operation minus "-" between two strings
"Remove what?" - "what?"
Let's now try to outsmart the computer and "add" a string and a number
1+"Hello"
Let us look at this error message (a traceback) in some detail.
A traceback gives details on what was happening when Python encountered an Exception or Error -- something it doesn't know how to handle.
There are many kinds of Python errors, with descriptive names to help us understand what went wrong. In this case we are getting a TypeError: we tried to do some operation on a data type that isn't supported for that data type.
Python gives us a helpful error message as part of the TypeError:
unsupported operand type(s) for +: 'int' and 'str'
In order to render the operation meaningful we would need to convert the number into a string. This can be done using the *str* function
str(1)
We can confirm that the type of the previous operation is a string. We can now meaningfully concatenate the two objects
str(1)+"Hello"
It is important to understand what is going on. We are transforming the number 1 into the string "1" using the function *str*, and concatenating the resulting string "1" with the second string "Hello" via the operation + between two strings.
We can also be interested in the reverse operation, given a string we may want to use it as a number in order to perform some numerical operation. This can be done with the function *float*, that converts a string into a floating point number.
37+float("2.998e8")
The ability to convert between data types is very useful and often occurs in solving practical problems.
A second operation \*, this time involving an integer number and a string.
20*"pancake"
Before you try the command out, can you see that it would be strange if the result was a number?
There you have, 20 pancakes for you.
Note that the \* operation in this case is between an integer and a string. You can see that
"20"*"pancake"
is not as good
Using a floating point (real) number will also not work
1.5*"pancake"
Using the two operations with strings above, + and \*, we can start to be creative: produce the string "Hehehehehehehehehehehello" using strings with maximum 3 characters as building blocks
There are many operations that can be done with strings, and we will be looking at them as we go along, but one that often comes in handy is the *len* which gives the total number of characters (including spaces) in a string.
We can use it to determine the number of characters in the long string above
len(insert your previous expression here)
Write down the expression that would give you the difference in string length (character number) between the string above and a normal "Hello".
### Booleans
If we want to introduce any logic in a program, or instruct the computer on how to perform instructions according to specific conditions, we need to make use of a less obvious but very important data type: the logical statements of *True* and *False*, called booleans
type(True)
Note that they are capitalized
type(true)
And are essentially different from a string with the word "True"
type("True")
Booleans do not often show up explicitly written in programs. However they are present all the time as a result of some operation. For example a comparison of two numbers.
-2 <= 1
In the expression above, the computer is testing whether -2 is smaller or equal to 1, and we should be happy to see that it got the order of the numbers right. The examples we use here are obvious, and we use them to illustrate how booleans work. In practice we will be dealing with more realistic cases.
We can also test if two numbers are equal
5 == 5
Note the two = signs above. This is not a typo, == is used when we want to test for equality. We will see further down that one = sign has a different meaning.
Similarly, we can test if two numbers are different
5 != 5
Above we are testing if 5 is different from 5. Since they are equal, the result of the test is False.
We can also compare two strings
"introvert"=="shy"
We could also compare a string to a number, and we can expect these two things to be always different.
"1.0"==1.0
A common operation could be the comparison of the length of two strings
len("introvert") > len("shy")
Let us complicate things slightly. We can use the operation *not* to obtain the complementary of a boolean.
not True
If something is not True, then it is False! Note that what we wrote is not a string. It is the operation *not* on the boolean *True*.
It should be easy to understand what the result of not False should be (create a new cell and test it, if it is not clear).
Used on in these simple cases, the operation *not* does not seem very useful, but it can be used to negate increasingly more complex constructions
not (3 > 6/2)
Can you see why we obtained this result?
We can check the result of multiple logical operations together, using the logical operators *and* and *or*.
If two expressions are joined by an *and*, they both have to be True for the overall expression to be True.
If two expressions are joined by an *or*, as long as at least one is True, the overall expression is True.
Let us give it a try
1 > 0 and 1 < 2
Although the use of brackets is not necessary, we can think of these compound conditional expressions as written as (1 > 0) and (1 < 2). Since each of the two expressions is True, the overall result is True.
We can construct more complex logical statements
(1 > 0 or 1 < 2) and 1 > 10
Would the result be different if we change the position of the parenthesis?
1 > 0 or (1 < 2 and 1 > 10)
## Variables
We have now covered the three basic data types present in almost all programming languages: numbers (of several kinds), strings and booleans; as well as some simple operations specific to each type.
Another language element common to all programming languages are variables. Variables allow us to store the result of specific operations for later use, making it easier to write programs. If, for example, we were writing a program to perform quantum mechanical calculations, it would be very tedious and error prone to explicitly write the Plank constant each time it is used. Instead we can store the value of the Plank constant into a variable.
h=6.626e-34
We have just defined a variable "called" h, with the value 6.626e-34. Note that we used a single = sign in this process. One = is an assignment of the variable on the left hand side, the value on the right hand side; two == is a operation testing whether the left hand side is equal to the right hand side, and yields a boolean.
When using a notebook interface, assigning a variable does not yield an output (see above). The output of a code cell is in general the output of the operation in that cell, so any cell that ends with a variable assignment will yield no output. This does not mean the operations in the cell have not been performed. Indeed we can see that the value of the Plank constant has been stored in variable h, by just executing the cell with the variable name
h
We can now use h in doing operations instead of typing the value explicitly, and we can quickly see the advantage of this if we want to calculate the energy of a red photon with 400×10<sup>12</sup> Hz frequency
the energy of a yellow photon of 515×10<sup>12</sup> Hz
and the energy of a blue photon of 640×10<sup>12</sup> Hz
Defining a variable in programming is similar to defining a variable in mathematics, with some slight differences. We can define variables as numbers, but also as strings, booleans, or more complex structures.
w="fantastic"
w
type(w)
type(h)
In mathematics we define a variable and don't necessarily think of the specific values this variable takes. When we define a mathematical function $f(x)=x^2$, we often think of $x$ as a continuous "thing", taking all real values at the "same time". *In programming, variables always have one and only one well defined value at each point of the program.*
The point just made about variable values at each point in the program illustrates well the notion that a program is a sequence of instructions. Let us define the following variables
a=3
b=4
a=b
b="done"
Since in the above cell all instructions are variable assignments we don't see any output. But what is the value of each variable now? You can test them below. Any surprises?
In the examples above we have always used single letter names for variables. We are not restricted to this however, and although variable names can not have spaces or other special characters, it makes life easier for yourself and whoever reads your program, to use more explicit variable names.
magic_number = 1/137
amountOfFlour = 0.75
my_name = "Genghis"
Again we see no output, but we are happy there are no error messages.
In order to check the values of the variables all at once we can use the function *print*, which simply prints its content on the screen.
print(magic_number)
print(amountOfFlour)
print(my_name)
Note that unlike other cells, the code above does not have a "Out \[\_\]:" tag associated with it on the left of the cell. This is because the *print* function is not returning a result. The only thing that it does is printing to the screen, but otherwise has no other effect on the program.
The *print* function is surprisingly useful for a command that does so little. It is used to check values of variables without logically changing the code and thus is useful when checking for errors and debugging code. Never fear to use it!
```
a = "| (_| -()- -()- -()- -()- | -()- -()- -()- -()- ||\n"
b = "|_\_|_/___|__|__|__|___|__|___|__|___________________________||\n"
c = "|________________________________|__|__()_|__()_|__()__|_____||\n"
d = " ___|)_______________________________________________________\n"
e = "|_/(|,\____/_|___/_|____/_|______|___________________________||\n"
f = "|___/____________________________|___________________________||\n"
g = "| | | () | () | () | | ||\n"
h = "|__\___|.________________________|___\_|___\_|___\_|___|_____||\n"
i = "|__/|_______/|____/|_____/|______|___________________________||\n"
j = "|_____/__________________________|____\|____\|____\|_________||\n"
k = "|____/___________________________|___________________________||\n"
l = "|__/___\_._______________________|__|__|__|__|__|__|___|_____||\n"
print(d + f + i + e + b + g + a + c + l + h + j + k)
```
## Making decisions: if statements
Life is made of decisions, and any minimally versatile code will need to execute different instructions depending on some criteria. An example from daily life would be: if it is hot, open the window; if it is not hot, don't! All programming languages provide mechanism to conditionally execute a piece of code. In python it takes the form:
if 6 > 5:
print("Six is greater than five!")
That was our first multi-line piece of code, and we need to pay attention to the indentation on the second line (the notebook tries to take care of this for you).
Make sure to hit space four times before typing the second line; Python needs them to understand what we want it to do.
So what is going on here? When Python encounters the *if* keyword, it evaluates the expression following the keyword and before the colon. This expression must always evaluate to a boolean type, i.e. evaluate to True or evaluate to False.
If that expression is True, Python executes the code in the indented code block under the if line.
If that expression is False, Python skips over the code block.
a="same"
if 0 > 1:
a="changed"
a
By using one *if* statement in the examples above, we have seen that how to execute code if the expression in the if statement is True, and if it is False no code is executed. A "true choice" will be executing one piece of code if the expression is True, and another piece of code if it is False. This is done using the *else* statement.
sister_age = 15
brother_age = 12
if sister_age > brother_age:
verdict="sister is older"
else:
verdict="brother is older"
verdict
An *else* is not stand alone and is always associated with a particular *if*.
If we have more than two cases, we can use the *elif* keyword to check more cases. We can have as many *elif* cases as we want; Python will go down the code checking each *elif* until it finds a True condition or reaches the default *else* block.
sister_age = 15
brother_age = 12
if sister_age > brother_age:
verdict="sister is older"
elif sister_age == brother_age:
verdict="sister and brother are the same age"
else:
verdict="brother is older"
verdict
You don't have to have an else block, if you don't need it. That just means there isn't default code to execute when none of the if or elif conditions are True:
colour = "orange"
season = ""
if colour == "green" or colour == "red":
season = "Christmas colour!"
elif colour == "black" or colour == "orange":
season = "Halloween colour!"
elif colour == "pink":
season = "Valentine's Day colour!"
season
Now go back to the cell above and change the first line to read
colour = 'purple'
Execute the cell again. Notice that, this time, nothing is printed.
Let us use an *if* statement to conditionally define the value of a variable. First define a variable *x* to have a real number value of your choice.
Then build an *if* statement to define a variable *y* that should be the [absolute value](http://mathworld.wolfram.com/AbsoluteValue.html) of *x*. In this exercise we are aiming at a general solution, i.e. one that would work for any value of *x*. Change the value of *x*, take positive and negative values, to check your solution works.
## Summary
We covered the simplest data types found in most programming languages:
* Numbers, further divided into integers, floating point numbers (floats) and complex numbers.
* Strings, which are sequences of characters.
* Booleans, True and False logical variables.
Each of these data types has specific operations associated with them, and some operations which involve quantities of different types.
Variables provide a way to call objects by a name. A variable can be of any type.
The *if* statement provides a way to branch code execution: "if *something* do this, otherwise do that". It is a fundamental programming construct, and is present virtually in every program.
## Exercises
### Menu specials
Given a string *vegetable* and a string *entree*, create string "Today our specials are: <vegetable> and <entree>".
Try it on the following pairs:
vegetable = 'asparagus' ; entree = 'pasta primavera'
vegetable = 'artichoke' ; entree = 'steak frites'
vegetable = 'kale' ; entree = 'fondue'
### Too long for twitter?
Twitter messages can not be more than 140 characters. Given a string *tweet*, print to screen the sentence "Not for twits" if the string is longer that 140 characters, and the word "Soundbite" if the string is less than or equal to 140 characters.
Try it on:
tweet='The Analytical Engine weaves algebraic patterns, just as the Jacquard loom weaves flowers and leaves. -- Ada Lovelace, the first programmer'
tweet='Four score and seven years ago our fathers brought forth on this continent a new nation, conceived in liberty, and dedicated to the proposition that all men are created equal.'
### Greatest out of 3
Define 3 variables *x*, *y* and *z* as three numbers of your choice. Write a piece of code that outputs the greatest of the three values. Change the values of *x*, *y* and *z* to check if your solution works.
# Part 2
## Grouping data together
In the first part of this session we have looked at three basic data types used in programming: numbers, strings and booleans. All programs can in principle be written using just these data types, but to solve many problems it is often useful to use more sophisticated data types, provided by the programming language (or some extension).
A useful and versatile data structure provided by Python are *lists*, and we will be making use of these during this course.
*Lists* are similar to *arrays*, which you have seen before, but they are not the same, and we will highlight some differences below.
### Lists
Lists provide a way of keeping and treating data together. We can create a list by putting its elements in a comma separated list enclosed in square brackets.
days_of_the_week = ["Sunday", "Monday", "Tuesday", "Wednesday", "Thursday", "Friday", "Saturday"]
We can see that we have created an object with a different type
type(days_of_the_week)
We can access the members of a list using the *index* of that item:
days_of_the_week[2]
The index is an integer which specifies the element we want. A little confusingly Python uses 0 as the index of the first element of a list. Thus, in this example, the 0 element is "Sunday", 1 is "Monday", and so on.
If we want to count back from the last element of a list we use a negative index. An index of -1 corresponds to the last element of the list, whilst an index of -2 corresponds to the second to last element, -3 the third last, etc. etc..
days_of_the_week[-1]
If we try and provide an index that goes beyond the last element of a list Python will throw an IndexError (this is a common error in programming)
days_of_the_week[8]
As an exercise, retrieve "Thursday" from the list days_of_the_week.
Besides selecting individual elements, we can also select ranges within the list using two integer numbers separated by a colon ':'. The sublist starts with the element defined by the starting index and include all elements up to **but not including** the element defined by the ending index. This process is called *slicing*
working_days=days_of_the_week[1:6]
working_days
This behaviour of indexes can seem strange at first. It is perhaps helpful to think of the indexes when slicing as pointing *between* list elements, to the comma.
["Sunday", "Monday", "Tuesday", "Wednesday", "Thursday", "Friday", "Saturday"]
| | | | | | | |
0 1 2 3 4 5 6 7
When slicing between 1 and 6, we are including the elements between the commas so numbered. This is a picture that can help you visualize what is going on and it should not be taken too literally.
If we omit the ending index, the sublist will include all elements until the end of the list
catholic_working_days=days_of_the_week[1:]
catholic_working_days
Conversely, if we omit the starting index, the elements will be taken from the beginning of the list.
Define a variable jewish_working_days, including all the elements of days_of_the_week, except "Saturday".
Besides extracting elements of the list, indexes can also be used to modify parts of the list
days_of_the_week[1]="Bank holiday"
days_of_the_week
We can also specify a range in doing this
days_of_the_week[1:3]=["Hard work","Hard work"]
days_of_the_week
We may specify discontinuous ranges on the list, by indicating a third number after another colon. This is a step index, specifying the increment to the list index.
sport_days=days_of_the_week[1:6:2]
sport_days
This slicing mechanism can also be used to insert elements in the list
days_of_the_week[5:5]=["Extra day"]
days_of_the_week
Once more we can think of the slicing indexes as pointing between list elements. In the "space between" list elements, we are assigning a new element, an insertion into the list.
Lists are very general and its elements do not have to have the same data type. For example, we can have both strings and number is the same list
["Today", 7, 99.3, "", 5**2]
List elements can even be other lists. Tables of data usually take this form
outer_list=[["dozens","units"],[0.5,6],[1,12],[2,24]]
Just as we can access the members of a list with an index:
outer_list[2]
So too we can access the members of an inner list by an extra index (do not confuse this notation with the list range we used above)
outer_list[2][1]
The indices are operating sequentially. We can use round brackets to make this clearer.
(outer_list[2])[1]
The first index, is selecting from outer_list the element index 2, which is the list [1,12]. The second index is selecting from this list the elements index 1.
If the above looks a bit confusing here's an alternative way of getting the same answer that shows what's happening when this double index is being used:
inner_list = outer_list[2]
inner_element = inner_list[1]
inner_element
#### Operations on lists
There are many operations involving lists provided by the Python language. We shall not try to be exhaustive, we will start with a few simple operations that will allow us to do most of the work, and introduce useful features as we go along.
First we will note a certain analogy between lists and strings: lists are an ordered collection of elements, while strings are ordered collections of characters. Indeed we can use indexing in strings in a similar way we used them with lists
analogy="A string is an ordered collection of characters"
analogy[0]
analogy[15:33]
We could thus expect that some of the operations on strings could have a similar behaviour on lists. We can thus use the function *len* to determine the length of a list
len(days_of_the_week)
Try to predict the result of the following command
len(days_of_the_week[1:5])
We can use the opertor '+' to perform list concatenation, and can be used to build bigger lists
[1,2,3]+[4,5,6]
Build the list \[1,2,3,"something",10\] by combining 3 different lists.
The operator \* with an integer yields the repetition of the list elements
5*[0,1]
3*[[1,2]]
### Lists vs arrays
As mentioned earlier, there are many similarities between *lists* and *arrays*, but there are also very important differences, some of which may already be apparent from what we have covered so far.
Arrays are not a primitive type in Python, but is a data type made available by the NumPy package which is automatically loaded with the Pylab environment. So in order to use arrays we need to load this package in one form or the other
from numpy import array
Arrays can be formed from suitable lists by using the function *array()*
simple_list=[1,2,3]
simple_array=array(simple_list)
simple_array
Any array can be converted back to a list using the function *list()*
simple_list2=list(simple_array)
simple_list2==simple_list
While any array can be made a list, not all lists can be made arrays. While lists can be collections of any type of data, arrays must be **regular collections of numbers only**, i.e. if we wish to convert a list of lists of length 3, all elements must have length 3
complete_list=[[1,2,3],[4,5,6],[7,8,9],[10,11,12]]
complete_array=array(complete_list)
complete_array
The fact that arrays are in general regular, allows for a more sophisticated indexing.
<img src="numpy_indexing.png" />
For example we can select columns of the array (elements on the same position of every sub-array). Before the coma we specify row range, and after the comma the column
complete_array[:,1]
It is also possible to define some sort of mitigated arrays from incomplete lists, or lists of mixed types which are not numbers
incomplete_list=[[1,2,3],[4,5],[7,8,9],[10,11,12]]
incomplete_array=array(incomplete_list)
incomplete_array
mixed_list=[1/137,3,"word",False]
mixed_array=array(mixed_list)
mixed_array
Note that is both cases we obtain an odd object type: "dtype=object" or dtype='&lr;U21'. These type of arrays are in general much less useful, and in particular they don't allow for convenient [row,column] indexing
incomplete_array[:,1]
Operations on lists and arrays also behave differently. We have seen that operations with list have some resemblance to operations on strings. For example the '+' operator in lists gives concatenation
[1,2,3]+[2,2,2]
Arrays on the other hand behave like vectors. Adding two arrays gives a vector sum, i.e. summing elements in the same position
array([1,2,3])+array([2,2,2])
Also, the '\*' operation in lists gives element repetition
3*[1,2,3]
in arrays however it corresponds to vector multiplication by a scalar, each element of the array is multiplied
3*array([1,2,3])
Arrays can be extremely useful to work with numerical data. We shall make use of them at later stages of this course, but for the most part we will be making use of the more versatile lists.
### Extra: Variations of lists
Besides lists (and arrays), Python possesses other similar data structures with somewhat different properties that make them more suited for some applications. We shall not be making use of these during the course as lists are more general, but you may find them when looking at code written by others.
Below we cover only the simplest properties of these other data structures, but you can [read more](http://docs.python.org/3/tutorial/datastructures.html#tuples-and-sequences) about them elsewhere.
#### Tuples
Tuples are indicated by round brackets
tup=(1,2,3,4,"last")
We can access its elements like lists
tup[1:3]
But we cannot change its content
tup[0]=20
#### Sets
Sets are indicated by a sequence inside curly brackets (or the *set()* function). While lists can have many repeated elements, sets eliminate the redundancy and only keep one element of each
set([1,1,1,2,2,2])
If we make a set from a string, the string is "destroyed" as we obtain a set with all characters used in the string, but lose the order that potentially gave meaning to the string
set("The quick brown fox jumps over the lazy dog!")
#### Dictionaries
While list are accessed by a position index, dictionaries are accessed by a label
atomic_number={"He":2,"Ne":10,"Ar":18,"Kr":36,"Xe":54,"Rn":86}
atomic_number["Ar"]
## Summary
List provide a way to group together and manipulate data. Imagine we were dealing with a dataset with the maximum temperature of every day in the year. It would be unpractical if we had to assign each value to a different variable.
Lists can contain data of any type at the same time. We can also have lists of lists.
We learned how to access different elements inside a list.
Lists are related to the data structure *array*. There are however important differences to be aware of and operations carried on each have a different behaviour.
In the next workshop we will be looking at loop, which allow for manipulating lists in a more versatile way.
| github_jupyter |
```
#!/usr/bin/env python3
# -*- coding: utf-8 -*-
"""
Created on Mon Sep 24 11:59:40 2018
@author: jack.lingheng.meng
"""
import os
os.system('module load nixpkgs/16.09 gcc/5.4.0 cuda/8.0.44 cudnn/7.0 opencv/3.3.0 boost/1.65.1 openblas/0.2.20 hdf5/1.8.18 leveldb/1.18 mkl-dnn/0.14 python/3.5.2')
os.system('cd ~')
os.system('source openposeEnv_Python3/bin/activate')
os.system('export LD_LIBRARY_PATH=$LD_LIBRARY_PATH:$HOME/openpose_python_lib/lib:$HOME/openpose_python_lib/python/openpose:$HOME/caffe/build/lib:/cvmfs/soft.computecanada.ca/easybuild/software/2017/avx2/Compiler/gcc5.4/boost/1.65.1/lib')
# From Python
# It requires OpenCV installed for Python
import sys
import csv
import cv2
import os
from sys import platform
import argparse
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import pandas as pd
import numpy as np
import math
from scipy.stats import mode
import pdb
from IPython.core.debugger import Tracer
# Remember to add your installation path here
# Option b
# If you run `make install` (default path is `/usr/local/python` for Ubuntu), you can also access the OpenPose/python module from there. This will install OpenPose and the python library at your desired installation path. Ensure that this is in your python path in order to use it.
sys.path.insert(0,r'/home/lingheng/openpose_python_lib/python/openpose')
# Parameters for OpenPose. Take a look at C++ OpenPose example for meaning of components. Ensure all below are filled
try:
from openpose import *
except:
raise Exception('Error: OpenPose library could not be found. Did you enable `BUILD_PYTHON` in CMake and have this Python script in the right folder?')
params = dict()
params["logging_level"] = 3
params["output_resolution"] = "-1x-1"
params["net_resolution"] = "-1x368" # if crop video, this should be changged and must be mutplies of 16.
params["model_pose"] = "BODY_25"
params["alpha_pose"] = 0.6
params["scale_gap"] = 0.3
params["scale_number"] = 1
params["render_threshold"] = 0.05
# If GPU version is built, and multiple GPUs are available, set the ID here
params["num_gpu_start"] = 0
params["disable_blending"] = False
# Ensure you point to the correct path where models are located
params["default_model_folder"] = "/home/lingheng/openpose/models/"
# Construct OpenPose object allocates GPU memory
openpose = OpenPose(params)
def subplot_estimated_occupancy(occupancy_whole, occupancy_core, occupancy_margin, fig_filename):
"""
Plot and save estimated occupancy in Three Interest Area.
Args:
occupancy_whole (pd.DataFrame): occupancy in Whole Interest Area
occupancy_core (pd.DataFrame): occupancy in Core Interest Area
occupancy_margin (pd.DataFrame): occupancy in Margin Interest Area
fig_filename (string): filename of the saved figure
"""
ymin = 0
ymax = 20
ystep = 4
lw=1.5
plt.figure()
# Whole Interest Area
plt.subplot(3, 1, 1)
plt.plot(occupancy_whole['Time']/1000, occupancy_whole['Occupancy'], 'b-', lw, alpha=0.6)
plt.xlabel('time/second')
plt.ylabel('# of visitors')
plt.ylim(ymin, ymax)
plt.yticks(np.arange(ymin,ymax,ystep))
plt.title('Estimated # of visitors in Whole Interest Area')
plt.grid(True, linestyle=':')
# Core Interest Area
plt.subplot(3, 1, 2)
plt.plot(occupancy_core['Time']/1000, occupancy_core['Occupancy'], 'r-', lw, alpha=0.6)
plt.xlabel('time/second')
plt.ylabel('# of visitors')
plt.ylim(ymin, ymax)
plt.yticks(np.arange(ymin,ymax,ystep))
plt.title('Estimated # of visitors in Core Interest Area')
plt.grid(True, linestyle=':')
# Margin Interest Area
plt.subplot(3, 1, 3)
plt.plot(occupancy_margin['Time']/1000, occupancy_margin['Occupancy'], 'g-', lw, alpha=0.6)
plt.xlabel('time/second')
plt.ylabel('# of visitors')
plt.ylim(ymin, ymax)
plt.yticks(np.arange(ymin,ymax,ystep))
plt.title('Estimated # of visitors in Margin Interest Area')
plt.grid(True, linestyle=':')
plt.tight_layout()
#plt.show()
plt.savefig(fig_filename, dpi = 300)
def plot_estimated_occupancy(occupancy_whole, occupancy_core, occupancy_margin, fig_filename):
ymin=0
ymax=20
ystep=4
plt.figure()
# Whole Interest Area
plt.plot(occupancy_whole['Time']/1000, occupancy_whole['Occupancy'], 'r-', lw=1.5, alpha=0.6)
# Core Interest Area
plt.plot(occupancy_core['Time']/1000, occupancy_core['Occupancy'], 'g-', lw=1.5, alpha=0.6)
# Margin Interest Area
plt.plot(occupancy_margin['Time']/1000, occupancy_margin['Occupancy'], 'b-', lw=1.5, alpha=0.6)
plt.legend(('Whole Interest Area','Core Interest Area','Margin Interest Area'))
plt.xlabel('time/second')
plt.ylabel('# of visitors')
plt.ylim(ymin, ymax, ystep)
plt.title('Estimated # of visitors in Three Interest Areas')
plt.grid(True, linestyle=':')
plt.tight_layout()
plt.show()
plt.savefig(fig_filename, dpi = 300)
def moving_smoothing(values, window_size, smooth_type='mode', stride = 1):
"""
Smoothen estimated occupancy.
Args:
values (pandas.DataFrame):
values['Time']: time in millisecond
values['Occupancy']: estimated # of visitors
window_size(int): the size of sliding window
smooth_type (string):
1. 'mode'
2. 'mean'
3. 'min'
4. 'median'
stride (int): the stride between two consecutive windows
Returns:
smooth_time (np.array): smooth time i.e. the max time in each window
smooth_occupancy (np.array): smooth occupancy i.e. the mode occupancy in each window
"""
group_time = []
group_occupancy = []
for i in range(0, math.ceil((len(values['Time'])-window_size+1)/stride)):
group_time.append(values['Time'][i:i+window_size])
group_occupancy.append(values['Occupancy'][i:i+window_size])
smooth_time = []
smooth_occupancy = []
for i in range(len(group_time)):
smooth_time.append(min(group_time[i])) # max time in the group
if smooth_type == 'mode':
smooth_occupancy.append(mode(group_occupancy[i])[0][0]) # mode occupancy in the group
elif smooth_type == 'mean':
smooth_occupancy.append(np.round(np.mean(group_occupancy[i])))
elif smooth_type == 'min':
smooth_occupancy.append(np.round(np.min(group_occupancy[i])))
elif smooth_type == 'median':
smooth_occupancy.append(np.round(np.median(group_occupancy[i])))
else:
print('Please choose a proper smooth_type.')
smooth_values = pd.DataFrame(data={'Time': np.array(smooth_time),
'Occupancy': np.array(smooth_occupancy,dtype=int)})
return smooth_values#np.array(smooth_time), np.array(smooth_occupancy)
def interpret_senario(occupancy_whole, occupancy_core, occupancy_margin, senarios_truth_table):
"""
Args:
occupancy_whole (pd.DataFrame): estimation of coccupancy in whole intrest area
occupancy_core (pd.DataFrame): estimation of coccupancy in core intrest area
occupancy_margin (pd.DataFrame): estimation of coccupancy in margin intrest area
senarios_truth_table (pandas.DataFrame): senarios truth table which has information on
how to interpret senario.
Returns:
senario_sequence (np.array): sequnce of interpreted senario discription according to "Senario Truth Value Table"
event_sequence (np.array): sequence of interpreted senario code according to "Senario Truth Value Table"
Note: Different from "Senario Truth Value Table", in this sequence we convert all impossible cases into 0 rather than their original senario code.
event_time (np.array): the time of each event in millisecond.
"""
senario_sequence = []
event_sequence = []
event_time = []
for i in range(len(occupancy_whole['Occupancy'])-1):
change_x = occupancy_core['Occupancy'][i+1] - occupancy_core['Occupancy'][i]
change_y = occupancy_margin['Occupancy'][i+1] - occupancy_margin['Occupancy'][i]
change_z = occupancy_whole['Occupancy'][i+1] - occupancy_whole['Occupancy'][i]
# code:
# 0: hold
# 1: increase
# 2: decrease
if change_x == 0:
x = 0
elif change_x > 0:
x = 1
elif change_x < 0:
x = 2
if change_y == 0:
y = 0
elif change_y > 0:
y = 1
elif change_y < 0:
y = 2
if change_z == 0:
z = 0
elif change_z > 0:
z = 1
elif change_z < 0:
z = 2
# convert ternary to decimal
senario_index = z + y*3 + x*3^2
senario_sequence.append(senarios_truth_table['Explanation'][senario_index])
if senarios_truth_table['Truth value'][senario_index] == 0:
# convert all impossible cases into 0
event_sequence.append(0)
#event_sequence.append(senario_index)
else:
event_sequence.append(senario_index)
event_time.append(occupancy_whole['Time'][i])
return np.array(senario_sequence), np.array(event_sequence), np.array(event_time)
def plot_detected_interesting_event(senario_sequence, event_sequence, event_time, fig_filename):
ymin = 0
ymax = 26.0005
ystep = 1
plt.figure(figsize=(10, 6))
plt.scatter(event_time/1000, event_sequence)
plt.xlabel('time/second')
plt.ylabel('Event Description')
plt.ylim(ymin, ymax)
plt.yticks(np.arange(ymin,ymax,ystep), senarios_truth_table['Explanation'],
rotation=45, fontsize = 6)
ax2 = plt.twinx()
plt.ylabel('Event Code')
plt.yticks(np.arange(ymin,ymax,ystep), np.arange(ymin,ymax,ystep))
plt.title('Detected Interesting Events')
plt.grid(True, linestyle=':')
plt.tight_layout()
plt.savefig(fig_filename, dpi = 300)
def tag_interesting_event_description_on_video(video_filename,
smooth_type, window_size, stride,
senario_sequence, event_sequence, event_time):
"""
Args:
video_filename (string): filename of video
smooth_type (string): smooth type (hyper-parameter of smooth method)
window_size (int): size of smooth window (hyper-parameter of smooth method)
stride (int): stride size (hyper-parameter of smooth method)
senario_sequence (np.array): sequnce of interpreted senario discription according to "Senario Truth Value Table"
event_sequence (np.array): sequence of interpreted senario code according to "Senario Truth Value Table"
Note: Different from "Senario Truth Value Table", in this sequence we convert all impossible cases into 0 rather than their original senario code.
event_time (np.array): the time of each event in millisecond.
"""
camera = cv2.VideoCapture(video_filename)
(grabbed, frame) = camera.read()
fheight, fwidth, channels= frame.shape
fourcc = cv2.VideoWriter_fourcc(*'XVID')
out_tagged_camera_frame = cv2.VideoWriter(video_filename.split('.avi')[0]+'_tagged_smooth_type_{}_window_size_{}_stride_{}.avi'.format(smooth_type,window_size,stride),fourcc, camera.get(cv2.CAP_PROP_FPS), (fwidth,fheight))
# loop over the frames of the video
total_frame_number = camera.get(cv2.CAP_PROP_FRAME_COUNT)
max_line_character_num = 60 # 60 characters each line
detected_event_time = 0
detected_event_senario = ''
line_num = 1
for frame_count in range(len(event_time)):
if frame_count % 200 == 0:
print('Processing frame: {}'.format(frame_count))
(grabbed, frame) = camera.read()
if grabbed == True:
cv2.putText(frame, "smooth_type: {}, window_size: {}, stride: {}.".format(smooth_type,window_size,stride), (10, 120), cv2.FONT_HERSHEY_SIMPLEX, 1, (255, 0, 0), 2)
time = camera.get(cv2.CAP_PROP_POS_MSEC) #Current position of the video file in milliseconds.
event_index = frame_count
if event_sequence[event_index] != 0: # 0 means 'impossible event'
detected_event_time = time
detected_event_senario = senario_sequence[event_index]
cv2.putText(frame, "Detect Interesting Event at: {}s.".format(int(detected_event_time/1000)), (10, 150), cv2.FONT_HERSHEY_SIMPLEX, 1, (255, 0, 0), 2)
line_num = np.ceil(len(detected_event_senario)/max_line_character_num)
for i in range(int(line_num)):
if i < line_num:
cv2.putText(frame, "{}".format(detected_event_senario[i*max_line_character_num:(i+1)*max_line_character_num]), (10, 180+30*(i)), cv2.FONT_HERSHEY_SIMPLEX, 1, (255, 0, 0), 2)
else:
cv2.putText(frame, "{}".format(detected_event_senario[i*max_line_character_num:end]), (10, 180+30*(i)), cv2.FONT_HERSHEY_SIMPLEX, 1, (255, 0, 0), 2)
else: # repeat text from last detected event
cv2.putText(frame, "Detect Interesting Event at:{}s".format(int(detected_event_time/1000)), (10, 150), cv2.FONT_HERSHEY_SIMPLEX, 1, (255, 0, 0), 2)
for i in range(int(line_num)):
if i < line_num:
cv2.putText(frame, "{}".format(detected_event_senario[i*max_line_character_num:(i+1)*max_line_character_num]), (10, 180+30*(i)), cv2.FONT_HERSHEY_SIMPLEX, 1, (255, 0, 0), 2)
else:
cv2.putText(frame, "{}".format(detected_event_senario[i*max_line_character_num:end]), (10, 180+30*(i)), cv2.FONT_HERSHEY_SIMPLEX, 1, (255, 0, 0), 2)
# save processed videos
out_tagged_camera_frame.write(frame)
else:
# Pass this frame if cannot grab an image.
print('Frame: {}, grabbed={} and frame={}'.format(frame_count, grabbed, frame))
if __name__ == "__main__":
# construct the argument parser and parse the arguments
ap = argparse.ArgumentParser()
ap.add_argument("-v", "--video", default='/home/lingheng/project/lingheng/ROM_raw_videos/Camera1_test.mp4', help="path to the video file")
ap.add_argument("-o", "--output_directory", default='/home/lingheng/project/lingheng/ROM_processed_videos', help="directory to save processed video")
args = vars(ap.parse_args())
if args.get("video", None) is None:
raise Error("No input video!!")
# otherwise, we are reading from a video file
else:
camera = cv2.VideoCapture(args["video"])
########################################################################
# Estimate Occupancy #
########################################################################
# frames per second (fps) in the raw video
fps = camera.get(cv2.CAP_PROP_FPS)
frame_count = 1
print("Raw frames per second: {0}".format(fps))
# prepare to save video
(grabbed, frame) = camera.read()
## downsample frame
#downsample_rate = 0.5
#frame = cv2.resize(frame,None,fx=downsample_rate, fy=downsample_rate, interpolation = cv2.INTER_LINEAR)
# crop frame
original_h, original_w, channels= frame.shape
top_edge = int(original_h*(1/10))
down_edge = int(original_h*1)
left_edge = int(original_w*(1/5))
right_edge = int(original_w*(4/5))
frame_cropped = frame[top_edge:down_edge,left_edge:right_edge,:].copy() # must use copy(), otherwise slice only return address i.e. not hard copy
cropped_h, cropped_w, channels = frame_cropped.shape
fwidth = cropped_w
fheight = cropped_h
print("Frame width:{}, Frame height:{}.".format(cropped_w , cropped_h))
# Define the polygon of Core Interest Area
point_1 = [int(0.17 * cropped_w), int(0.20 * cropped_h)]
point_2 = [int(0.17 * cropped_w), int(0.62 * cropped_h)]
point_3 = [int(0.44 * cropped_w), int(0.82 * cropped_h)]
point_4 = [int(0.61 * cropped_w), int(0.72 * cropped_h)]
point_5 = [int(0.61 * cropped_w), int(0.20 * cropped_h)]
core_interest_area_polygon = np.array([point_1,point_2,point_3,point_4,point_5])
# get output video file name
file_path = args["video"].split('/')
file_name, _= file_path[-1].split('.')
fourcc = cv2.VideoWriter_fourcc(*'XVID')
output_video_filename = os.path.join(args['output_directory'],'{}_processed.avi'.format(file_name))
out_camera_frame_whole = cv2.VideoWriter(output_video_filename,fourcc, fps, (fwidth,fheight))
# get output estimated occupancy file name
out_occupancy_whole = os.path.join(args['output_directory'],'{}_processed_occupancy_whole.csv'.format(file_name))
out_occupancy_core = os.path.join(args['output_directory'],'{}_processed_occupancy_core.csv'.format(file_name))
out_occupancy_margin = os.path.join(args['output_directory'],'{}_processed_occupancy_margin.csv'.format(file_name))
with open(out_occupancy_whole, 'a') as csv_datafile:
fieldnames = ['Time', 'Occupancy']
writer = csv.DictWriter(csv_datafile, fieldnames = fieldnames)
writer.writeheader()
with open(out_occupancy_core, 'a') as csv_datafile:
fieldnames = ['Time', 'Occupancy']
writer = csv.DictWriter(csv_datafile, fieldnames = fieldnames)
writer.writeheader()
with open(out_occupancy_margin, 'a') as csv_datafile:
fieldnames = ['Time', 'Occupancy']
writer = csv.DictWriter(csv_datafile, fieldnames = fieldnames)
writer.writeheader()
# loop over the frames of the video
total_frame_number = camera.get(cv2.CAP_PROP_FRAME_COUNT)
for frame_count in range(int(total_frame_number)):
if frame_count % 200 == 0:
print('Processing frame: {}'.format(frame_count))
(grabbed, frame) = camera.read()
if grabbed == True:
time = camera.get(cv2.CAP_PROP_POS_MSEC) #Current position of the video file in milliseconds.
## downsample frame
#frame = cv2.resize(frame,None,fx=downsample_rate, fy=downsample_rate, interpolation = cv2.INTER_LINEAR)
# crop frame
frame_cropped = frame[top_edge:down_edge,left_edge:right_edge,:].copy() # must use copy()
# 1. Whole Interest Area
# Output keypoints and the image with the human skeleton blended on it
# (num_people, 25_keypoints, x_y_confidence) = keypoints_whole_interest_area.shape
keypoints_whole_interest_area, output_image_whole_interest_area = openpose.forward(frame_cropped, True)
# 2. Core Interest Area
core_interest_area_mask = np.zeros(frame_cropped.shape[:2], np.uint8)
cv2.drawContours(core_interest_area_mask, [core_interest_area_polygon], -1, (255, 255, 255), -1, cv2.LINE_AA)
core_interest_area = cv2.bitwise_and(output_image_whole_interest_area, frame_cropped, mask=core_interest_area_mask)
# 3. Margin Interest Area
margin_interest_area = cv2.bitwise_xor(output_image_whole_interest_area, core_interest_area)
# TODO: infer occupancy from "keypoints_whole_interest_area"
# draw the text and timestamp on the frame
occupancy_whole = keypoints_whole_interest_area.shape[0]
occupancy_core = 0
occupancy_margin = 0
for people in keypoints_whole_interest_area:
# Sort all keypoints and pick up the one with the highest confidence
# Meaning of keypoints (https://github.com/CMU-Perceptual-Computing-Lab/openpose/blob/master/doc/output.md)
ordered_keypoints = people[people[:,2].argsort(),:] # increasing order
x, y = ordered_keypoints[-1][:2]
#pdb.set_trace()
# Choose the one with higher confidence to calculatate occupancy and location
if cv2.pointPolygonTest(core_interest_area_polygon, (x, y), False) == 1:
occupancy_core += 1
else:
occupancy_margin += 1
cv2.drawContours(output_image_whole_interest_area, [core_interest_area_polygon], -1, (255, 255, 0), 2, cv2.LINE_AA)
cv2.putText(output_image_whole_interest_area, "Whole Occupancy: {}, Core Occupancy: {}, Margin Occupancy: {}".format(occupancy_whole, occupancy_core, occupancy_margin), (10, 80), cv2.FONT_HERSHEY_SIMPLEX, 1, (255, 255, 255), 2)
cv2.putText(core_interest_area, "Core Occupancy: {}".format(occupancy_core), (10, 50), cv2.FONT_HERSHEY_SIMPLEX, 1, (255, 255, 255), 2)
cv2.putText(margin_interest_area, "Margin Occupancy: {}".format(occupancy_margin), (10, 50), cv2.FONT_HERSHEY_SIMPLEX, 1, (255, 255, 255), 2)
# save estimated occupancy data
fieldnames = ['Time', 'Occupancy']
with open(out_occupancy_whole, 'a') as csv_datafile:
writer = csv.DictWriter(csv_datafile, fieldnames = fieldnames)
writer.writerow({'Time':time, 'Occupancy': occupancy_whole})
with open(out_occupancy_core, 'a') as csv_datafile:
writer = csv.DictWriter(csv_datafile, fieldnames = fieldnames)
writer.writerow({'Time':time, 'Occupancy': occupancy_core})
with open(out_occupancy_margin, 'a') as csv_datafile:
writer = csv.DictWriter(csv_datafile, fieldnames = fieldnames)
writer.writerow({'Time':time, 'Occupancy': occupancy_margin})
# save processed videos
out_camera_frame_whole.write(output_image_whole_interest_area)
else:
# Pass this frame if cannot grab an image.
print('Frame: {}, grabbed={} and frame={}'.format(frame_count, grabbed, frame))
########################################################################
# Smoothen Estimated Occupancy, then detect interesting event #
########################################################################
# read estimated occupancy in Three Interest Areas
occupancy_whole = pd.read_csv(out_occupancy_whole)
occupancy_core = pd.read_csv(out_occupancy_core)
occupancy_margin = pd.read_csv(out_occupancy_margin)
# save plot of estimated occupancy in Three Interest Areas
fig_filename = 'Subplot_Estimated_Occupancy.png'
subplot_estimated_occupancy(occupancy_whole, occupancy_core, occupancy_margin, fig_filename)
fig_filename = 'Plot_Estimated_Occupancy.png'
plot_estimated_occupancy(occupancy_whole, occupancy_core, occupancy_margin, fig_filename)
# smoothen
window_size = 25
smooth_type='mean'
stride = 1
smooth_occupancy_whole = moving_smoothing(occupancy_whole, window_size, smooth_type)
smooth_occupancy_core = moving_smoothing(occupancy_core, window_size, smooth_type)
smooth_occupancy_margin = moving_smoothing(occupancy_margin, window_size, smooth_type)
fig_filename = 'Subplot_Smooth_Estimated_Occupancy.png'
subplot_estimated_occupancy(smooth_occupancy_whole,smooth_occupancy_core,smooth_occupancy_margin, fig_filename)
fig_filename = 'Plot_Smooth_Estimated_Occupancy.png'
plot_estimated_occupancy(smooth_occupancy_whole,smooth_occupancy_core,smooth_occupancy_margin, fig_filename)
# load Senario Truth Table
senarios_truth_table = pd.read_csv('analize_visitor_in_and_out_senario_truth_table.csv')
# Interpret
senario_sequence, event_sequence, event_time = interpret_senario(smooth_occupancy_core,
smooth_occupancy_margin,
smooth_occupancy_whole,
senarios_truth_table)
# Plot interesting events
fig_filename = 'Plot_Interesting_Event_smooth_type_{}_window_size_{}_stride{}'.format(smooth_type, window_size, stride)
plot_detected_interesting_event(senario_sequence, event_sequence, event_time, fig_filename)
# Tag
tag_interesting_event_description_on_video(output_video_filename,
smooth_type, window_size, stride,
senario_sequence, event_sequence, event_time)
```
| github_jupyter |
# Quantile Regression
This is the first notebook in a series on performing (deep) Bayesian quantile regression.
In this notebook we look into quantile regression and write two classes to perform quantile regression quickly. These classes are based on linear programming. In order to know how to set it up, we first need to understand the mathematical derivation to get to the optimal solution. As a consequence, we first will derive how to rewrite the quantile regression problem into a linear programming problem.
After all this is done, we will perform quantile regression for [homoscedastic](https://en.wikipedia.org/wiki/Heteroscedasticity) as well as [heteroscedastic](https://en.wikipedia.org/wiki/Heteroscedasticity) data.
First we make some necessary imports and build a `generate_data` function such that we can simply call it later on.
```
import numpy as np
import matplotlib.pyplot as plt
from scipy import optimize
plt.style.use('seaborn')
SMALL_SIZE = 12
MEDIUM_SIZE = 13
BIGGER_SIZE = 16
plt.rc('font', size=SMALL_SIZE) # controls default text sizes
plt.rc('axes', titlesize=BIGGER_SIZE) # fontsize of the axes title
plt.rc('axes', labelsize=MEDIUM_SIZE) # fontsize of the x and y labels
plt.rc('xtick', labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc('ytick', labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc('legend', fontsize=SMALL_SIZE) # legend fontsize
plt.rc('figure', titlesize=BIGGER_SIZE) # fontsize of the figure title
%matplotlib inline
def generate_data(n_samples=100, x_min=0, x_max=3, intercept=1, slope=2, base_std=1.5, homoscedastic=True, extra_hetero_std=0.333, seed=42):
np.random.seed(seed)
x = np.linspace(x_min, x_max, n_samples)
y_true = intercept + slope * x
if homoscedastic:
y = y_true + np.random.normal(loc=0, scale=base_std, size=n_samples)
else:
y = y_true + np.random.normal(loc=0, scale=base_std, size=n_samples)*x + np.random.normal(loc=0, scale=extra_hetero_std, size=n_samples)
return x, y, y_true
```
## Rewriting the Quantile Regression into a linear programming problem
It is possible to rewrite the quantile regression loss function in a way that can easily be optimized. We will first derive this mathematically before coding everything. In case you are not necessarily interested in the mathematical details, you can just read the *summary* section to see what the solution is that we will code out.
Before diving into the mathematics below, it should be clear that any vector (e.g. $x$) is assumed to be a column vector. If it is a row vector it will be indicated by the transposed sign (e.g. $x^T$). All matrices will be indicated by a capital letter (e.g. $X$).
### Quantile Regression
The basic way that quantile regression works is that we set up a linear regression to estimate the true $y \in \mathbb{R}^n$ by:
$$ \hat{y} = X\theta$$
Where $X \in \mathbb{R}^{n\times p}$ is the data matrix, $\hat{y} \in \mathbb{R}^{n}$ the estimated output, and $\theta \in \mathbb{R}^p$ the parameters of the linear regression. The error term is defined as $\epsilon = y - \hat{y}$. The Quantile Loss function is then defined by:
$$ \mathcal{L}_Q(y, \hat{y}) = \sum_{i=1}^{n} \rho_{\tau} (\underbrace{y_i - x_i^T \theta}_{\epsilon_i}) $$
The function $\rho_\tau (r) = r \cdot (\tau - I[r < 0])$, with $I[r<0]$ being the [indicator function](https://en.wikipedia.org/wiki/Indicator_function) (I will not use the regular subscript notation, because I will need that later on for the identity matrix). Writing it out makes the loss function of the form
$$ \mathcal{L}_Q(y, \hat{y}) = \sum_{i=1}^{n} \tau |\epsilon_i| I[\epsilon_i \geq 0 ] + (1-\tau) |\epsilon_i| I[\epsilon_i < 0] $$
The loss function is also sometimes written in either of the following forms;
$$\begin{array}{rcl}
\mathcal{L}_Q(y, \hat{y}) &=& \tau \max(y-\hat{y},0) + (1-\tau) \left[- \min(y-\hat{y}, 0)\right] \\
\mathcal{L}_Q(y, \hat{y}) &=& \begin{cases}\tau (y-\hat{y}), & y-\hat{y} \leq 0 \\ (1-\tau) (y-\hat{y}), & y-\hat{y} < 0. \end{cases}
\end{array}
$$
They are all equivalent, but the method of using the $\rho_\tau(r)$ will make the derivation for linear programming easier to follow, so we will stick to that formulation for now. We find the optimum parameters $\theta$ by getting the arguments of minimizing the loss function.
$$ \hat{\theta} = \arg \min_{\theta} \mathcal{L}_Q(y, \hat{y}) = \arg \min_{\theta} \sum_{i=1}^{n} \tau |\epsilon_i| I[\epsilon_i \geq 0 ] + (1-\tau) |\epsilon_i| I[\epsilon_i < 0]. $$
In visual terms this is indicated by the fact that everything above all the data points above the regression line are weighted by $\tau$ in the loss function and everything underneath the regression line is weighted by $1-\tau$. This can be seen in the figure below.
<center>
<img src='img/quant_lossfunc.png' width="90%"></img>
</center>
### Linear Programming
There is a couple of ways to write an optimization problem. In terms of linear programming is this from time to time done by writing it as a [maximization problem](https://en.wikipedia.org/wiki/Linear_programming). It should be clear that it can quite quickly be rewritten as a minimization problem, because $\max_x f(x) = \min_x -f(x)$ (the same should be done with the boundary conditions). As a result, we will use the form
$$\begin{array}{crcl}
\min_z & c^T z, \\
\text{subject to} & Az & = & b \\
&z & \geq & 0&
\end{array}$$
This means we need to arrive somehow to this form. We are looking for a way to rewrite our quantile regression problem defined above into the matrix $A$ and the vectors $b, c, z$. Where I think it should be clear that the minimization should be equal for both terms.
Since we have the condition that $z \geq 0$, and we will see that $\epsilon$ will end up in the $z$ vector, we will need a way to make sure that $\epsilon_i \geq 0, ~ \forall i$. A classic way of doing this is by decomposing $\epsilon_i \in \mathbb{R}$ is by splitting it up in an absolute manner $\epsilon_i = u_i - v_i$ where $u_i \in \mathbb{R}_+$ as well as $v_i \in \mathbb{R}_+$. If we remember that $\epsilon_i$ is the error of the estimate, a natural way to decompose this is by saying:
$$ \epsilon_i = u_i - v_i, ~ \text{for} ~ \begin{cases} u_i = \max(0, y-\hat{y}) \\ v_i = - \min(0, y-\hat{y}) \end{cases} $$
This allows us to rewrite the minimization problem as
$$ \min_\theta \sum_{i=1}^n \tau u_i + (1-\tau) v_i = \tau \underbrace{\begin{bmatrix}1 & 1 & \ldots & 1 \end{bmatrix}}_{\mathbf{1}_n^T} \begin{bmatrix} u_1 \\ u_2 \\ \vdots \\ u_n \end{bmatrix} + (1-\tau) \begin{bmatrix}1 & 1 & \ldots & 1 \end{bmatrix} \begin{bmatrix} v_1 \\ v_2 \\ \vdots \\ v_n \end{bmatrix}
= \tau \mathbf{1}_n^T u + (1-\tau) \mathbf{1}_n^T v$$
Where the residuals $\epsilon_i$ should satisfy the constraints $y_i - x_i^T \theta = \epsilon_i$. This is how we earlier defined the estimation error, but now it will appear in the constraints. As a result the total optimization problem can be defined as
$$\begin{array}{crcl}
\min_{\theta, u, v} & \tau \mathbf{1}_n^T u + (1-\tau) \mathbf{1}_n^T v, \\
\text{s.t.} & y_i & = & x_i \theta + u_i - v_i, ~ i=1,\ldots,n
\end{array}$$
where $\theta \in \mathbb{R}^p,~ u \in \mathbb{R}_+^n,~ v \in \mathbb{R}_+^n$. It is now clear that $theta, u, v$ are all elements of $z$. As we have seen earlier, we have to invoke $z \geq 0$. We know the $u$ and $v$ satisfy this condition, but $\theta$ does not. To make sure this is the case, we decompose $\theta$ in the same way as before;
$$\theta = \theta^+ - \theta^-,~ \begin{cases} \theta^+ = \max(0, \theta) \\ \theta^- = - \min(0, \theta) \end{cases}$$.
This allows us to rewrite the original equation $y = X\theta + \epsilon$ as $y=X(\theta^+ - \theta^-) + I_n u - I_n v$ (with $I_n$ being the $n\times n$ identity matrix). In order to rewrite this as the linear programming problem, we define $b$ and rewrite it as
$$\begin{array}{rcl}
b:=y & = & X(\theta^+ - \theta^-) + I_n u + I_n v \\
&=& X \theta^+ - X \theta^- + I_n u - I_n v \\
\underbrace{y}_{b} &=& \underbrace{\begin{bmatrix} X & -X & I_n & -I_n \end{bmatrix}}_{A} \underbrace{ \begin{bmatrix}\theta^+ \\ \theta^- \\ u \\ v \end{bmatrix}}_{z}
\end{array}$$
As a result, we know have the constraint $Az=b$ and we satisfy $z \geq 0$. The last step is to find vector $c$. We know we should end up with $\min_z \tau \mathbf{1}_n^T u + (1-\tau) \mathbf{1}_n^T v$, but $z$ has the $theta^+, \theta^-$ terms which do not occur in the minimization problem. The easy way to make sure this works out all right is if we obtain
$$c^Tz = \mathbf{0}_p^T \theta^+ - \mathbf{0}_p^T \theta^- + \tau \mathbf{1}_n^T u + (1-\tau) \mathbf{1}_n^T v,$$
where $\mathbf{0}_p^T = \begin{bmatrix} 0 & 0 & \ldots & 0 \end{bmatrix}$ with $\mathbf{0}_p \in \mathbb{R}^p$. This means we can define $c$ to be
$$c := \begin{bmatrix} \mathbf{0}_p \\ \mathbf{0}_p \\ \tau \mathbf{1}_n \\ (1-\tau) \mathbf{1}_n \end{bmatrix} = \begin{bmatrix} \mathbf{0}_{2p} \\ \tau \mathbf{1}_n \\ (1-\tau) \mathbf{1}_n \end{bmatrix}. $$
This can now all be solved by the classic methods of solving linear programming problems (for example [the simplex algorithm](https://en.wikipedia.org/wiki/Simplex_algorithm) or [the interior point method](https://en.wikipedia.org/wiki/Interior-point_method)).
### Summary
To summarize quickly; we have written the quantile regression problem into a linear programming problem. The quantile regression problem was defined by estimating $y$ by the linear regression model $\hat{y} = X\theta$ subject to minimizing the loss function $\sum_{i=1}^{n} \tau |\epsilon_i| I[\epsilon_i \geq 0 ] + (1-\tau) |\epsilon_i| I[\epsilon_i < 0]$.
It has been shown that this can be written in the form
$$\begin{array}{crcl}
\min_z & c^T z, \\
\text{subject to} & Az & = & b \\
&z & \geq & 0&
\end{array}$$
with
$$ \begin{array}{rclrcl}
A & = & \begin{bmatrix} X & -X & I_n & -I_n \end{bmatrix}, & b & = & \begin{bmatrix} y_1 \\ y_2 \\ \vdots \\ y_n \end{bmatrix} \\
c^T & = & \begin{bmatrix} \mathbf{0}_p ^T &\tau \mathbf{1}_n^T & (1-\tau) \mathbf{1}_n^T \end{bmatrix} ,& z & = & \begin{bmatrix}\theta^+ \\ \theta^- \\ u \\ v \end{bmatrix}
\end{array}$$
If we carefully look at the shapes of all the elements, we find that $A \in \mathbb{R}^{n\times(2p + 2n)}$, $b \in \mathbb{R}^n$, $c \in \mathbb{R}^{2p + 2n}$ and $z \in \mathbb{R}^{2p + 2n}$. The convex optimization problem can then quickly be solved by a classic convex optimization method.
## Coding a `QuantileRegressor`
Now that we know what vectors and matrices we need, we can make a `scikit-learn` based regressor to learn the parameters by calling a `fit` function and then predict an output using a `predict` function. We will do this for a single quantile and subsequently make a `MultiQuantileRegressor` which can do it for a range of quantiles, because often multiple quantiles are desired.
### Single regressor
For the single regressor the only input when setting it up is going to be what specified quantile we need. This is the $\tau$ (or `tau`) parameter. The rest will be calculated when calling the `fit` function. This requires an input/data matrix $X$ and an output matrix $y$. In general this will be the data coming from a train/test split. It is important that $y$ gets transformed to a column vector and $X$ gets cast in the proper form. In order to incorporate an intercept in the parameters, we need to add a column $\mathbf{1}_n$ before the other columns of $X$.
All that rests then is building matrix $A$ and the vectors $b$ and $c$. These are then passed to `scipy`'s `linprog` method (which is in the `optimize` sub-module). The output from this linear programming is of a type called `scipy.optimize.optimize.OptimizeResult`. It has some specific attributes, among which a `success` attribute which indicates if the optimization was succesful.
In case it was succesful, the output is the $z$ vector. The first $p$ elements is $\theta^+$ and the second $p$ elements are $\theta^-$ elements (all non-negative). We just need to combine them to obtain $\theta = \theta^+ - \theta^-$. The array $\theta$ then contains all the necessary parameters to make predictions.
The `predict` function then just takes an array (vector or matrix) and makes prediction(s) for all the supplied data points.
```
class QuantileRegressor():
def __init__(self, tau=0.5):
self.tau = tau
def fit(self, X, y):
y = y.reshape(-1,1)
if len(X.shape) == 1: #check if it is a 1d dataset and then recast it into a column vector
X = X.reshape(-1,1)
intercept = np.ones([X.shape[0], 1])
X = np.hstack((intercept, X))
n = X.shape[0]
p = X.shape[1]
A = np.hstack((X, -X, np.eye(n), -np.eye(n)))
b = y.copy()
c = np.vstack((
np.zeros([p, 1]),
np.zeros([p, 1]),
self.tau * np.ones([n, 1]),
(1-self.tau) * np.ones([n, 1])
))
res = optimize.linprog(c, A_eq=A, b_eq=b, bounds=(0, None), method='interior-point')
if res.success:
theta_pos = res.x[0:p]
theta_neg = res.x[p:2*p]
self.theta = theta_pos - theta_neg
self.intercept = self.theta[0]
self.coef_ = self.theta[1:]
else:
raise ArithmeticError("The Linear Programming optimization was unsucessful.")
def predict(self, X):
if X.shape[0] == len(self.coef_): #check if this is a single sample
X = X.reshape(1,-1)
elif len(X.shape) == 1: #check if it is a 1d dataset and then recast it into a column vector
X = X.reshape(-1,1)
intercept = np.ones([X.shape[0], 1])
X = np.hstack((intercept, X))
y = np.dot(X, self.theta)
return y
```
### Setting up a regressor for multiple quantiles
As mentioned earlier, it can easily be the case that we would like to perform multiple quantile regressions in a row for a range of quantiles. For this, we have made the `MultiQuantileRegressor`. It takes as an input either a list of $\tau$ values as input (`l_tau`), or just an integer `n_tau` ($n_\tau$) of the amount of $\tau$ values desired. This is then uniformly spaced over the (open) interval $\langle 0,1 \rangle$.
The `fit` function works in the same way as of the `QuantileRegressor`, but first sets up the arrays which stay fixed ($A$ and $b$) and then for every $\tau$ calls the function `_fit_single` where it sets up the array $c$ and performs the linear programming optimization. All the different parameters are then appended into one big $\theta$ matrix where $\theta \in \mathbb{R}^{p\times n_{\tau}}$ corresponding to the $\tau$ values which can be found in `l_tau`.
```
class MultiQuantileRegressor():
def __init__(self, n_tau=None, l_tau=None):
assert (l_tau != None) | (n_tau != None), "Please provide either `l_tau` or `n_tau`"
if n_tau != None:
self.l_tau = np.linspace(0, 1, n_tau+2)[1:-1]
self.n_tau = n_tau
else:
self.l_tau = np.array(l_tau)
self.n_tau = len(l_tau)
def fit(self, X, y):
if len(y.shape) == 1:
y = y.reshape(-1,1)
if len(X.shape) == 1:
X = X.reshape(-1,1)
intercept = np.ones([X.shape[0], 1])
X = np.hstack((intercept, X))
n = X.shape[0]
p = X.shape[1]
A = np.hstack((X, -X, np.eye(n), -np.eye(n)))
b = y.copy()
self.theta = np.empty((self.n_tau, p))
for i, tau in enumerate(self.l_tau):
self.theta[i,:] = self._fit_single(A=A, b=b, n=n, p=p, tau=tau)
self.n_coef = p-1
self.intercept = self.theta[:,0]
self.coef_ = self.theta[:,1:]
def _fit_single(self, A, b, n, p, tau):
c = np.vstack((
np.zeros([2*p, 1]),
tau * np.ones([n, 1]),
(1-tau) * np.ones([n, 1])
))
res = optimize.linprog(c, A_eq=A, b_eq=b, bounds=(0, None), method='interior-point')
if res.success:
theta_pos = res.x[0:p]
theta_neg = res.x[p:2*p]
theta = theta_pos - theta_neg
return theta
else:
raise ArithmeticError(f"The Linear Programming optimization was unsucessful for tau = {tau}.")
def predict(self, X):
if X.shape[0] == self.n_coef:
X = X.reshape(1,-1)
elif len(X.shape) == 1:
X = X.reshape(-1,1)
intercept = np.ones([X.shape[0], 1])
X = np.hstack((intercept, X))
y = np.dot(X, self.theta.T)
return y
```
## Fitting the `QuantRegressor` and `MultiQuantRegressor`
Now we are going fit the regressors on homoscedastic data first and subsequently on heteroscedastic data. For this we have made the `generate_data` function at the start of this notebook, where you can indicate homoscedasticity or not and can set the standard deviation if desired.
### Homoscedastic data
The homoscedastic data is generated by:
$$ y = 1 + 2x + \eta, ~~~ \eta \sim \mathcal{N}\left(0, \sigma^2\right), ~~~ \text{with } \sigma = \frac{3}{2}$$
```
x, y, y_true = generate_data(n_samples=250)
```
Let's see what the data looks like. There is also an `y_true` in the data which shows the original linear line where the noise was added on top of.
```
plt.figure(figsize=(10,5))
plt.scatter(x, y, marker='x', label='Sampled data', color='tab:gray', alpha=0.6)
plt.plot(x, y_true, label='true regression line', lw=2, color='tab:blue')
plt.xlabel('x'); plt.ylabel('y'); plt.title('Generated data and underlying model'); plt.legend()
plt.show()
```
Now we set up the `QuantileRegressor` and `MultiQuantileRegressor`, for a `tau=0.33` and `n_tau=5` respectively. We then simply call the `fit` functions and `predict` on the entire domain to make it available for the plot.
```
tau_s = 0.33
n_tau = 5
reg_single = QuantileRegressor(tau=tau_s)
reg_multi = MultiQuantileRegressor(n_tau=n_tau)
reg_single.fit(x, y)
reg_multi.fit(x, y)
x_fit = np.linspace(0, 3, 100).reshape(-1,1)
y_singlefit = reg_single.predict(x_fit)
y_multifit = reg_multi.predict(x_fit)
fig, ax = plt.subplots(1, 2, figsize=(22,5))
ax[0].scatter(x, y, marker='x', label='Sampled train data', color='tab:gray', alpha=0.25)
ax[0].plot(x, y_true, label='True regression line', lw=2, ls='--', color='tab:blue', alpha=0.75)
ax[0].plot(x_fit, y_singlefit, label=f'Regression regression line for tau = {round(tau_s, 2)}', lw=2, ls='-', color='tab:green')
ax[0].set_xlabel('x')
ax[0].set_ylabel('y')
ax[0].set_title('Single Quantile Regression')
ax[0].legend()
ax[1].set_title('Multiple Quantile Regression')
ax[1].scatter(x, y, marker='x', label='Sampled train data', color='tab:gray', alpha=0.25)
ax[1].plot(x, y_true, label='True regression line', lw=2, ls='--', color='tab:blue', alpha=0.75)
for i, tau in enumerate(reg_multi.l_tau):
ax[1].plot(x_fit, y_multifit[:,i], label=f'Regression line for tau={round(tau, 2)}', lw=2, color='tab:green', alpha=(i+2*n_tau)/(3.5*n_tau))
ax[1].set_xlabel('x')
ax[1].set_ylabel('y')
ax[1].legend()
plt.show()
```
### Heteroscedastic data
The heteroscedastic data is generated by:
$$ y = 1 + 10x + \eta_1 x + \eta_2, ~~~ \begin{cases} \eta_1 \sim \mathcal{N}\left(0, \sigma_1^2\right), & \text{with } \sigma_1 = 6 \\ \eta_2 \sim \mathcal{N}\left(0, \sigma_2^2\right), & \text{with } \sigma_2 = \frac{1}{3} \end{cases} $$
```
slope = 10
base_std = 6
x2, y2, y2_true = generate_data(n_samples=200, slope=slope, base_std=base_std, homoscedastic=False)
```
Let's see what the data looks like. There is also an `y_true` in the data which shows the original linear line where the noise was added on top of.
```
plt.figure(figsize=(10,5))
plt.scatter(x2, y2, marker='x', label='Sampled data', color='tab:gray', alpha=0.55)
plt.plot(x2, y2_true, label='True regression line', lw=2, color='tab:blue')
plt.plot(x2, y2_true+base_std*x2, c='tab:green', ls='--', label='True regression with heteroscedastic variance')
plt.plot(x2, y2_true-base_std*x2, c='tab:green', ls='--')
plt.xlabel('x'); plt.ylabel('y'); plt.title('Generated data and underlying model'); plt.legend()
plt.show()
tau2_s = 0.33
n_tau2 = 5
reg_single2 = QuantileRegressor(tau=tau_s)
reg_multi2 = MultiQuantileRegressor(n_tau=n_tau)
reg_single2.fit(x2, y2)
reg_multi2.fit(x2, y2)
x2_fit = np.linspace(0, 3, 100).reshape(-1,1)
y2_singlefit = reg_single2.predict(x2_fit)
y2_multifit = reg_multi2.predict(x2_fit)
```
Now we set up the `QuantileRegressor` and `MultiQuantileRegressor`, for a `tau=0.33` and `n_tau=5` respectively. We then simply call the `fit` functions and `predict` on the entire domain to make it available for the plot.
```
fig, ax = plt.subplots(1, 2, figsize=(22,5))
ax[0].scatter(x2, y2, marker='x', label='Sampled train data', color='tab:grey', alpha=0.25)
ax[0].plot(x2, y2_true, label='True regression line', lw=2, ls='--', color='tab:blue', alpha=0.75)
ax[0].plot(x2_fit, y2_singlefit, label=f'Regression regression line for tau = {round(tau2_s, 2)}', lw=2, ls='-', color='tab:green')
ax[0].set_xlabel('x')
ax[0].set_ylabel('y')
ax[0].set_title('Single Quantile Regression')
ax[0].legend()
ax[1].set_title('Multiple Quantile Regression')
ax[1].scatter(x2, y2, marker='x', label='Sampled train data', color='tab:grey', alpha=0.25)
ax[1].plot(x2, y2_true, label='True regression line', lw=2, ls='--', color='tab:blue', alpha=0.75)
for i, tau in enumerate(reg_multi2.l_tau):
ax[1].plot(x2_fit, y2_multifit[:,i], label=f'Regression line for tau={round(tau, 2)}', lw=2, color='tab:green', alpha=(i+2*n_tau2)/(3.5*n_tau2))
ax[1].set_xlabel('x')
ax[1].set_ylabel('y')
ax[1].legend()
plt.show()
```
| github_jupyter |
```
# HIDDEN
from datascience import *
%matplotlib inline
path_data = '../../../data/'
import matplotlib.pyplot as plots
plots.style.use('fivethirtyeight')
import pylab as pl
import numpy as np
```
### Properties of the Mean ###
In this course, we have used the words "average" and "mean" interchangeably, and will continue to do so. The definition of the mean will be familiar to you from your high school days or even earlier.
**Definition.** The *average* or *mean* of a collection of numbers is the sum of all the elements of the collection, divided by the number of elements in the collection.
The methods `np.average` and `np.mean` return the mean of an array.
```
not_symmetric = make_array(2, 3, 3, 9)
np.average(not_symmetric)
np.mean(not_symmetric)
```
### Basic Properties ###
The definition and the example above point to some properties of the mean.
- It need not be an element of the collection.
- It need not be an integer even if all the elements of the collection are integers.
- It is somewhere between the smallest and largest values in the collection.
- It need not be halfway between the two extremes; it is not in general true that half the elements in a collection are above the mean.
- If the collection consists of values of a variable measured in specified units, then the mean has the same units too.
We will now study some other properties that are helpful in understanding the mean and its relation to other statistics.
### The Mean is a "Smoother" ##
You can think of taking the mean as an "equalizing" or "smoothing" operation. For example, imagine the entries in `not_symmetric` above as the dollars in the pockets of four different people. To get the mean, you first put all of the money into one big pot and then divide it evenly among the four people. They had started out with different amounts of money in their pockets (\$2, \$3, \$3, and \$9), but now each person has \$4.25, the mean amount.
### Proportions are Means ###
If a collection consists only of ones and zeroes, then the sum of the collection is the number of ones in it, and the mean of the collection is the proportion of ones.
```
zero_one = make_array(1, 1, 1, 0)
sum(zero_one)
np.mean(zero_one)
```
You can replace 1 by the Boolean `True` and 0 by `False`:
```
np.mean(make_array(True, True, True, False))
```
Because proportions are a special case of means, results about random sample means apply to random sample proportions as well.
### The Mean and the Histogram ###
The mean of the collection {2, 3, 3, 9} is 4.25, which is not the "halfway point" of the data. So then what does the mean measure?
To see this, notice that the mean can be calculated in different ways.
\begin{align*}
\mbox{mean} ~ &=~ 4.25 \\ \\
&=~ \frac{2 + 3 + 3 + 9}{4} \\ \\
&=~ 2 \cdot \frac{1}{4} ~~ + ~~ 3 \cdot \frac{1}{4} ~~ + ~~ 3 \cdot \frac{1}{4} ~~ + ~~ 9 \cdot \frac{1}{4} \\ \\
&=~ 2 \cdot \frac{1}{4} ~~ + ~~ 3 \cdot \frac{2}{4} ~~ + ~~ 9 \cdot \frac{1}{4} \\ \\
&=~ 2 \cdot 0.25 ~~ + ~~ 3 \cdot 0.5 ~~ + ~~ 9 \cdot 0.25
\end{align*}
The last expression is an example of a general fact: when we calculate the mean, each distinct value in the collection is *weighted* by the proportion of times it appears in the collection.
This has an important consequence. The mean of a collection depends only on the distinct values and their proportions, not on the number of elements in the collection. In other words, the mean of a collection depends only on the distribution of values in the collection.
Therefore, **if two collections have the same distribution, then they have the same mean.**
For example, here is another collection that has the same distribution as `not_symmetric` and hence the same mean.
```
not_symmetric
same_distribution = make_array(2, 2, 3, 3, 3, 3, 9, 9)
np.mean(same_distribution)
```
The mean is a physical attribute of the histogram of the distribution. Here is the histogram of the distribution of `not_symmetric` or equivalently the distribution of `same_distribution`.
```
# HIDDEN
t = Table().with_columns(
'Value', make_array(2, 3, 9),
'Proportion', make_array(0.25, 0.5, 0.25)
)
t.hist(counts='Value', bins=np.arange(1.5, 9.6, 1))
```
Imagine the histogram as a figure made out of cardboard attached to a wire that runs along the horizontal axis, and imagine the bars as weights attached at the values 2, 3, and 9. Suppose you try to balance this figure on a point on the wire. If the point is near 2, the figure will tip over to the right. If the point is near 9, the figure will tip over to the left. Somewhere in between is the point where the figure will balance; that point is the 4.25, the mean.
**The mean is the center of gravity or balance point of the histogram.**
To understand why that is, it helps to know some physics. The center of gravity is calculated exactly as we calculated the mean, by using the distinct values weighted by their proportions.
Because the mean is a balance point, it is sometimes displayed as a *fulcrum* or triangle at the base of the histogram.
```
# HIDDEN
mean = sum(t.column('Value')*t.column('Proportion'))
t.hist(counts='Value', bins=np.arange(1.5, 9.6, 1))
plots.scatter(mean, -0.009, marker='^', color='darkblue', s=60)
plots.plot([1.5, 9.5], [0, 0], color='grey')
plots.ylim(-0.05, 0.5);
```
### The Mean and the Median ###
If a student's score on a test is below average, does that imply that the student is in the bottom half of the class on that test?
Happily for the student, the answer is, "Not necessarily." The reason has to do with the relation between the average, which is the balance point of the histogram, and the median, which is the "half-way point" of the data.
The relationship is easy to see in a simple example. Here is a histogram of the collection {2, 3, 3, 4} which is in the array `symmetric`. The distribution is symmetric about 3. The mean and the median are both equal to 3.
```
symmetric = make_array(2, 3, 3, 4)
# HIDDEN
t2 = Table().with_columns(
'Value', make_array(2, 3, 4, 9),
'symmetric', make_array(0.25, 0.5, 0.25, 0)
)
mean1 = sum(t2.column('Value')*t2.column('symmetric'))
t2.hist(counts='Value', bins=np.arange(1.5, 4.6, 1))
plots.scatter(mean1, -0.009, marker='^', color='darkblue', s=60)
plots.xlim(1, 10)
plots.ylim(-0.05, 0.5);
np.mean(symmetric)
percentile(50, symmetric)
```
In general, **for symmetric distributions, the mean and the median are equal.**
What if the distribution is not symmetric? Let's compare `symmetric` and `not_symmetric`.
```
# HIDDEN
t2 = t2.with_column(
'not_symmetric', make_array(0.25, 0.5, 0, 0.25)
)
mean2 = sum(t2.column('Value')*t2.column('not_symmetric'))
t2.hist(counts='Value', bins=np.arange(1.5, 9.6, 1))
plots.scatter(mean1, -0.009, marker='^', color='darkblue', s=60)
plots.scatter(mean2, -0.009, marker='^', color='gold', s=60)
plots.ylim(-0.05, 0.5);
```
The blue histogram represents the original `symmetric` distribution. The gold histogram of `not_symmetric` starts out the same as the blue at the left end, but its rightmost bar has slid over to the value 9. The brown part is where the two histograms overlap.
The median and mean of the blue distribution are both equal to 3. The median of the gold distribution is also equal to 3, though the right half is distributed differently from the left.
But the mean of the gold distribution is not 3: the gold histogram would not balance at 3. The balance point has shifted to the right, to 4.25.
In the gold distribution, 3 out of 4 entries (75%) are below average. The student with a below average score can therefore take heart. He or she might be in the majority of the class.
In general, **if the histogram has a tail on one side (the formal term is "skewed"), then the mean is pulled away from the median in the direction of the tail.**
### Example ###
The table `sf2015` contains salary and benefits data for San Francisco City employees in 2015. As before, we will restrict our analysis to those who had the equivalent of at least half-time employment for the year.
```
sf2015 = Table.read_table(path_data + 'san_francisco_2015.csv').where('Salaries', are.above(10000))
```
As we saw earlier, the highest compensation was above \$600,000 but the vast majority of employees had compensations below \$300,000.
```
sf2015.select('Total Compensation').hist(bins = np.arange(10000, 700000, 25000))
```
This histogram is skewed to the right; it has a right-hand tail.
The mean gets pulled away from the median in the direction of the tail. So we expect the mean compensation to be larger than the median, and that is indeed the case.
```
compensation = sf2015.column('Total Compensation')
percentile(50, compensation)
np.mean(compensation)
```
Distributions of incomes of large populations tend to be right skewed. When the bulk of a population has middle to low incomes, but a very small proportion has very high incomes, the histogram has a long, thin tail to the right.
The mean income is affected by this tail: the farther the tail stretches to the right, the larger the mean becomes. But the median is not affected by values at the extremes of the distribution. That is why economists often summarize income distributions by the median instead of the mean.
| github_jupyter |
```
%pylab inline
from __future__ import division
from __future__ import print_function
import pandas as pd
import seaborn as sb
from collections import Counter
```
## Malware Classification Through Dynamically Mined Traces
### 1. The Dataset
The dataset used in this notebook can be freely downloaded from the [csmining website](http://www.csmining.org/index.php/malicious-software-datasets-.html), where there's also an easy explanation on the nature of the dataset and its strenghts/weaknesses.
For a quick recap: the dataset is a made of traces of API calls from 387 windows programs, some of which are malware.
The malware programs are labelled as 1, whereas the 'goodware' programs have a 0 label. Here's a line:
1,LoadLibraryW HeapAlloc HeapAlloc HeapFree HeapAlloc HeapFree HeapFree NtOpenKey LoadLibraryW GetProcAddress GetProcAddress [...]
Let's start exploring the dataset.
```
dataset = pd.read_csv('CSDMC_API_Train.csv')
dataset.columns = ['target','trace']
print(dataset.keys())
print(dataset.columns)
```
Each trace is simply a string, representing a list of API calls separated with a space and residing all in the first column.
So the first thing to do is to split it into an actual python list, creating a list of traces that will each be represented by a tuple containing the trace itself and its classification into 'malware' or 'goodware'.
```
traces = []
for i in dataset.index:
traces.append((dataset.iloc[i]['trace'].strip().split(' '), dataset.iloc[i]['target']))
print ('A trace: ' , type(traces[0][0]))
print ('A label: ', type(traces[8][1]))
```
To gain some additional knowledge on the dataset we could check its bias, or how well are the samples distributed between malware and goodware.
Let's count how many ones and zeroes there are in the target column.
```
c_target = Counter([x[1] for x in traces])
print(c_target)
```
It seems like the dataset is pretty biased towards malware, as there are few samples of benign applications being run.
It's almost the polar opposite of what would happen in a randomly sampled dataset from real world applications, as malware is usually a one digit percentage of the set of every application being released.
But let's not despair, this will actually make learning easier. It might hurt in the generalization tho.
Here's a graph showing the obvious bias:
```
plt.figure(figsize=(6,8))
plt.xticks(np.arange(2) + 1.4, ['Goodware', 'Malware'])
plt.title('Dataset Bias')
plt.ylabel('Number of Program Traces')
plt.xlabel('Classification')
plt.bar([1,2], c_target.values())
```
### 2. Initial Feature Mining
Now it's time to mine for features, as the dataset itself doesn't really lend itself to an easy classification with a Machine Learning algorithm.
Not out of the box at least.
The raw traces present some peculiar challenges for a smooth classification:
1. They are composed of strings
2. They have various length (makes it hard to fit them in a matrix with fixed width)
3. They present a lot of repeated data points
We need numerical features, and possibly a way to organize everything.
The first idea is to count how many times a given API call is contained in each trace, this should yield positive results during learning if there's any correlation at all with the quantity of calls made to a specific API during a program run and a malicious behaviour.
```
counted_traces = []
for trace, target in traces:
counted_traces.append((Counter(trace), target))
```
Just to get an immediate feedback let's print a list of the first 20 traces, and look at the 3 most used API calls in each trace.
The diagnosis is printed at the end to give some perspective.
```
diagnosis = ''
for i in range(20):
if counted_traces[i][1] == 0:
diagnosis = 'OK'
else:
diagnosis = 'MALWARE'
trace_sample = counted_traces[i][0].most_common(3)
print(i, ')', trace_sample, diagnosis)
```
We can obtain some good information and maybe some ideas from this alone:
1. The only two good samples have the shortest program run and the longest one, this might not be relevant in general but it's worth investigating
2. The most popular API calls are roughly the same for each program run, so maybe they won't be incredibly useful for classification
Also, this might be the shortest program run ever (my guess is it crashed soon after loading):
```
counted_traces[11][0]
```
Maybe then it's possible we'll need the length of each trace and the number of times an API has been called during a program run, and that's all information we can freely gather from the data we have assembled so far.
But the *absolute* number of API calls in a program trace isn't a very useful feature, as it mostly depends on the length of each trace, so we'll normalize it by searching for the **frequency** of each API call in a program run.
And since we will have the frequencies associated to each API call, maybe we can see if the frequency of the most used API call is useful for classification.
Since it's now time to gather more than one feature and it's better to keep everything tidy, let's generate a list of dictionaries that will contain the following fields:
**'ID'** : index of the trace, given by the enumerate() method
**'Counter'** : Counter containing the API calls and how many times they have been called
**'Freq'** : frequency at which a certain API call has been used in a program trace
**'Length'** : Length of the trace
**'MostFreqCall'** : The most common API call and its frequency
**'Target'** : 1 or 0, depending on the maliciousness of the sample
To be honest I'm just glad I could use the name 'dict_traces'.
```
dict_traces = [] #a list of dicts
for i, t in enumerate(counted_traces):
trace, target = t
max_freq = 0
most_common = ()
length = len(traces[i][0])
freq_dict = {}
for key in trace:
freq = trace[key] / length
freq_dict[key] = freq
if freq > max_freq:
max_freq = freq
most_common = (key, freq)
d = {'ID' : i,
'Counter' : trace,
'Freq' : freq_dict,
'Length' : length,
'MostFreqCall' : most_common,
'Target' : target}
dict_traces.append(d)
print(dict_traces[0].keys())
print(dict_traces[0]['MostFreqCall'])
```
What is the most frequent "most frequent call"?
Since the most popular API calls will inevitably be used by every program run, be it malicious or not, maybe we can avoid them.
```
most_freq_call_list = []
for d_t in dict_traces:
call, freq = d_t['MostFreqCall']
most_freq_call_list.append(call)
c = Counter(most_freq_call_list)
print('Maybe we can avoid these: ', c.most_common(3))
```
Here's a graph showing the N most frequent "most frequent call".
As we can see the first 4 are pretty noticeable, then they drop fast:
```
N = 12
plt.figure(figsize=(12,8))
plt.title('Most frequent "most frequent call"')
plt.ylabel('Frequency')
y = [x[1]/len(dict_traces) for x in c.most_common(N)]
plt.bar(np.arange(N) + 0.2, y)
plt.xticks(np.arange(N) + 0.6, [x[0] for x in c.most_common(N)], rotation=60)
plt.ylim((0, 0.5))
```
A further trasformation in our data is needed before we start learning, let's separate the target from the data points.
This will be useful to render the code more readable, and to have another quick glimpse into how biased the dataset is.
```
target = []
for d in dict_traces:
target.append(d['Target'])
print(target)
```
As we can see from the density of the ones, our algorithm would do pretty well if it just guessed 'malware' all the time:
```
p_malware = c_target[1] / len(target)
print('Accuracy if I always guess "malware" = ', p_malware)
print('False positives: ', 1 - p_malware)
```
Of course false negatives will be exactly 0% in this particular instance so, generally speaking, this wouldn't be a bad result.
But that wouldn't be a very realistic scenario in a learned classifier, and even then that would mean that it actually learnt something from the dataset (the target's distribution), although it shoudln't be useful at all for generalizing.
Let's see how a really dumb classifier would fare, by just guessing 'malware' and 'goodware' with 50% chance (this time accounting for both false positives and false negatives):
```
p_chance_mal = (p_malware * 0.5)
p_chance_good = (c_target[0] / len(target)) * 0.5
print ('''Probability of getting it right by guessing with 50%%:
- False Positive Ratio: %f
- False Negative Ratio: %f
''' % (1 - p_chance_mal, 1 - p_chance_good))
```
Now these are horrible ratios, let's hope we can do better than this.
## 3. Learning
It's time to finally try and learn something.
Throughout the rest of the notebook we'll use various classifiers and functions from the [scikit-learn](http://scikit-learn.org/) library.
First off we'll need a classifier, and since I'm a fan of ensemble learning we'll start with a Random Forest classifier initialized with 20 estimators and a random state set to 42.
The random state is very important, as it will help with the reproducibility of this study.
```
from sklearn.ensemble import RandomForestClassifier
model = RandomForestClassifier(n_estimators=20, random_state=42)
```
Now we can't start learning right away, as our dataset should be first divided into a 'train' set and a 'test' set, to ensure a bit of generalization.
We could do this manually, by randomly selecting a chunk of the dataset (usually 75%) for the training part and leaving the rest for testing, but we could still generate a lucky division and receive optimistic results that won't generalize well in the real world.
Thankfully scikit-learn has provided a neat implementation of the KFold algorithm that will allow us to generate how many folds we need.
```
from sklearn.cross_validation import KFold
kf = KFold(len(target), n_folds=3, random_state=42)
```
Another little adjustment is needed before using scikit-learn's algorithms, as they expect the data to be indexed vertically, but thankfully again, numpy has the solution.
We're going to create a numpy array with each trace length and reshape it accordingly.
Let'stry and learn only from the length of the traces:
```
scores = []
for train, test in kf:
train_data = np.array([dict_traces[i]['Length'] for i in train]).reshape(len(train), 1)
test_data = np.array([dict_traces[i]['Length'] for i in test]).reshape(len(test), 1)
model.fit(train_data, [target[i] for i in train])
scores.append(model.score(test_data, [target[i] for i in test]))
print(scores)
print(np.array(scores).mean())
```
We have chosen to learn from 3 folds and already our classifier seems to produce good results even with one feature.
This might be because it's a pretty small dataset and it's kinda biased.
Since we're going to try and learn from different features and maybe different classifiers, it's best to keep track of the scores in a global way, just to visualize the improvements over time (or lack thereof).
```
global_scores = {}
global_scores['Length'] = scores
```
Another feature we mined is the most frequent API call, let's see how well it does by itself.
Since classifiers work mainly in dimensional data, we need a way to encode the API call into an integer, maybe by using a dictionary.
This is a very rudementary but effective way:
```
most_freq_list = [x['MostFreqCall'][0] for x in dict_traces]
most_freq_counter = Counter(most_freq_list)
most_freq_dict = {}
index = 0
for call in most_freq_counter.keys():
most_freq_dict[call] = index
index += 1
print(most_freq_dict)
```
The learning process is basically the same as before, so maybe it's time to encode it in a function.
```
model = RandomForestClassifier(n_estimators=20, random_state=42)
scores = []
for train, test in kf:
train_data = np.array([most_freq_dict[dict_traces[i]['MostFreqCall'][0]] for i in train]).reshape(len(train), 1)
test_data = np.array([most_freq_dict[dict_traces[i]['MostFreqCall'][0]] for i in test]).reshape(len(test), 1)
model.fit(train_data, [target[i] for i in train])
scores.append(model.score(test_data, [target[i] for i in test]))
global_scores['MostFreqCall'] = scores
print(scores)
print(np.array(scores).mean())
```
Wait.
```
print(p_malware)
print(np.array(scores).mean())
```
This is probably just a coincidence, but the accuracy of our classifier is exactly the same as the hypothetical classifier that always guesses 'malware'.
And it's a weirdly good result, since the most frequent api call might be correlated with the classification but this is too much. Again, it's probably due to the size of the dataset or its skewness.
I think we can safely assume this is the lowest score we can get with any classifier.
To improve on this, lets try to learn from the 2 features we just mined.
```
model = RandomForestClassifier(n_estimators=20, random_state=42)
scores = []
for train, test in kf:
most_freq_train_data = np.array([most_freq_dict[dict_traces[i]['MostFreqCall'][0]] for i in train]).reshape(len(train), 1)
length_train_data = np.array([dict_traces[i]['Length'] for i in train]).reshape(len(train), 1)
train_data = np.append(most_freq_train_data, length_train_data,1)
most_freq_test_data = np.array([most_freq_dict[dict_traces[i]['MostFreqCall'][0]] for i in test]).reshape(len(test), 1)
length_test_data = np.array([dict_traces[i]['Length'] for i in test]).reshape(len(test), 1)
test_data = np.append(most_freq_test_data, length_test_data,1)
model.fit(train_data, [target[i] for i in train])
scores.append(model.score(test_data, [target[i] for i in test]))
global_scores['Length + MostFreqCall'] = scores
print(scores)
print(np.array(scores).mean())
```
This is a good improvement, even a 5% increase at this stage can be beneficial, we'll see if this is the right direction.
## 4. Reorganizing Features
One of the best aspects of the scikit-learn library is their readily-available datasets, which are either already present in the library's path, or provide a simple function that will download them.
Since we're mining features from our dataset, we could use the same structure as shown below:
```
from sklearn.datasets import load_iris
iris = load_iris()
print(iris.keys())
print(iris['feature_names'])
```
This is good practice, in case we later want to release this to the public or even in case someone wants to expand on this.
Starting with the easy things, a description and the target:
```
m_descr = """
Malware Traces Dataset
Notes:
---------
Dataset characteristics:
:Number of Instances: 387 (319 malware and 68 goodware)
:Number of Attributes: 2
:Attribute Information:
- trace length
- most frequent API call (encoded with an integer)
- class:
- Malware
- Goodware
"""
malware_dataset = {
'target_names' : ['Goodware', 'Malware'],
'DESCR' : m_descr,
'target' : np.array(target)
}
```
Now for the hard part: data and feature_names.
We'll need to unify the 2 features we have used until now:
```
m_most_freq_data = np.array([most_freq_dict[dict_traces[i]['MostFreqCall'][0]] for i in range(len(dict_traces))]).reshape(len(dict_traces), 1)
m_length_data = np.array([dict_traces[i]['Length'] for i in range(len(dict_traces))]).reshape(len(dict_traces), 1)
m_data = np.append(m_most_freq_data, m_length_data,1)
malware_dataset['data'] = m_data
malware_dataset['feature_names'] = ['trace length', 'most frequent call']
```
## 5. Reorganizing Learning
Since now our dataset is clean and organized, we can streamline the learning process aswell.
The function learn() will take in input a classifier, the features and the target points, while returning the scores as we have used until now (raw scores and their mean).
I also added an option to plot the confusion matrix for the learn classifier and the possibility to save the scores into the global_scores variable we have initialized a while ago.
```
from sklearn.metrics import confusion_matrix
def learn(model, data, target, descr=None, n_folds=3, plot=False):
'''
"descr" is an optional parameter that will save the results in global_scores for later visualization
"n_folds" is there just in case I want to change it
'''
kf = KFold(data.shape[0], n_folds=n_folds, random_state=42)
scores = []
best_score = 0
best_split = ()
for train, test in kf:
#this is easier to read
model.fit(data[train], target[train])
m_score = model.score(data[test], target[test])
scores.append(m_score)
if plot and m_score > best_score:
best_score = m_score
best_split = (train, test)
#this plots a simple confusion matrix
if plot:
train, test = best_split
model.fit(data[train], target[train])
cm = confusion_matrix(target[test], model.predict(data[test]))
plt.imshow(cm, interpolation='nearest', cmap=plt.cm.Blues)
plt.title('Confusion Matrix')
plt.xticks(np.arange(len(malware_dataset['target_names'])), malware_dataset['target_names'])
plt.yticks(np.arange(len(malware_dataset['target_names'])), malware_dataset['target_names'], rotation=90)
plt.ylabel('Actual Label')
plt.xlabel('Predicted Label')
plt.tight_layout()
if descr != None:
global_scores[descr] = scores
return (scores, np.array(scores).mean())
```
Let's try it out:
```
model = RandomForestClassifier(n_estimators=20, random_state = 42)
data = malware_dataset['data']
target = malware_dataset['target']
print('''Our algorithm has:
- scores: %s
- mean: %f
''' % learn(model, data, target, plot=True))
```
This is exactly the same result as before but this was expected, as that's why we initialized the same random state.
Now it's time to mine for more features, since the most frequent call kinda improved our classification when paired with the length of the trace, maybe the 2nd and 3rd most frequent calls will add to it?
```
print(dict_traces[0].keys())
print(dict_traces[0]['Counter'].most_common(4)[0])
m_second_most_freq = []
for trace in dict_traces:
m_second_most_freq.append(trace['Counter'].most_common(2)[1])
m_s_counter = Counter([x[0] for x in m_second_most_freq])
```
This is the same process we used to encode numerically the API calls in the first 'most frequent' feature, it's very rough but it gets the job done.
It could also be methodologically wrong, as we're using different encodings for some of the same API calls.
```
m_s_dict = {}
index = 0
for item in m_s_counter.keys():
m_s_dict[item] = index
index += 1
print(m_s_dict)
m_s_list = [m_s_dict[x[0]] for x in m_second_most_freq]
m_s_data = np.array(m_s_list).reshape(len(m_s_list), 1)
```
Let's add it to the existing feature set.
```
malware_dataset['data'] = np.append(malware_dataset['data'], m_s_data, 1)
malware_dataset['feature_names'].append('second most frequent call')
print(malware_dataset['feature_names'])
```
Has it already impacted the classification?
```
model = RandomForestClassifier(n_estimators=20, random_state=42)
data = malware_dataset['data']
target = malware_dataset['target']
descr = 'Length + MostFreqCall + SecondMostFreqCall'
print('''Our algorithm has:
- scores: %s
- mean: %f
''' % learn(model, data, target, descr, plot=True))
```
The overall improvement is negligible, but looking at the folds it seems like two of them were more susceptible to the new feature and the middle one didn't really take notice.
So it could be a step in the right direction.
Maybe it would help to visualize our improvements over time:
```
def plot_improvement():
plt.figure(figsize=(13,5))
t_counter = Counter(target)
#assuming an algorithm that always guesses "Malware" as the baseline
best_guess = t_counter[1] / len(target)
plt.plot(np.arange(len(global_scores.keys()) + 1), [best_guess] + [np.array(x).mean() for x in global_scores.values()])
plt.xticks(np.arange(len(global_scores.keys())) + 1, global_scores.keys(), rotation=60)
plt.xlim((0, len(global_scores.keys()) + 0.2))
plt.ylabel('Accuracy')
print('If we just guess "malware" we get an accuracy of: ', best_guess)
print('Our best classificator has an accuracy of: ', np.array([np.array(x).mean() for x in global_scores.values()]).max())
plot_improvement()
```
The improvement on the second most frequent call is clearly negligible, so we'll stop investigating in that direction.
But it seems apparent that the frequency of the api calls has to be correlated in some way with the malicious behaviors of the samples so we might aswell try this new approach.
The idea is simple, there are 10 most frequent apis throughout the dataset, and each trace presents them with a certain frequency:
```
m_10_most_common = []
for trace in dict_traces:
freq_list = []
for t, f in most_freq_counter.most_common(10):
freq_list.append(trace['Counter'][t] / trace['Length'])
m_10_most_common.append(freq_list)
print(m_10_most_common[2])
m_data_10 = np.array(m_10_most_common)
```
Let's update our feature set and the feature names:
```
malware_dataset['data'] = np.append(malware_dataset['data'], m_data_10, 1)
print(malware_dataset['data'].shape)
for i in range(10):
malware_dataset['feature_names'].append(str(i + 1) + ' API call')
```
Now we can try to learn from all the features just mined at once:
```
model = RandomForestClassifier(n_estimators=20, random_state=42)
data = malware_dataset['data']
target = malware_dataset['target']
descr = '10 MostFreqPerc'
print('''Our algorithm has:
- scores: %s
- mean: %f
''' % learn(model, data, target, descr, plot=True))
plot_improvement()
```
Now, this is a very good improvement, and it seems like every fold is responding in the same way so it's not dependent on the random selection of the training set.
But it's interesting to wonder if the newly mined features are just improving on the old ones or if they can be used on their own without any detraction from the classificiation.
So, what is the accuracy of our classifier if we only learn from the 10 new features?
```
model = RandomForestClassifier(n_estimators=20, random_state=42)
data = malware_dataset['data'][:, 2:]
target = malware_dataset['target']
print('''Our algorithm has:
- scores: %s
- mean: %f
''' % learn(model, data, target))
```
Not bad, as expected.
As a side note, I stopped plotting the confusion matrix as only false negatives were present.
## 6. Trying Different Classifiers
We could go on and mine for features for a while, but an algorithm wich can discern between malware and goodware with ~95% accuracy is already a pretty good result for such a short study.
Also, there's another direction where we could improve, and that's by trying out new models.
- ***AdaBoost***
Until now we used Random Forest, which is just an ensemble classifier that uses Decision Trees as base classifier, but the scikit-learn library also provides us with an implementation of AdaBoost, an ensemble classifier that seems to do just the same thing (its default base classifier is a Decision Tree).
So it might be interesting to see if we can get the same results.
*Note: on the surface Ada Boost and Random Forest seem to be fairly similar, as they both combine the results of many underlying 'weaker' classifiers and construct a stronger learner, but they differ a lot in their core.
Random Forest is a bagging algorithm and Ada Boost is a boosting algorithm.*
```
from sklearn.ensemble import AdaBoostClassifier
from sklearn.tree import DecisionTreeClassifier
base_class = DecisionTreeClassifier(random_state=42)
model = AdaBoostClassifier(n_estimators=120,learning_rate=1.2, random_state=42)
print('''Our algorithm has:
- scores: %s
- mean: %f
''' % learn(model, data, target))
```
The result is pretty similar, we can play a bit with the estimators and the learning rate but we won't get much better results than this.
Also, if we put back the first 2 features it actually becomes worse.
As a quick side note, we started with ensemble classifiers, but what about linear classifiers?
Well, there's a reason to ignore them:
```
from sklearn.linear_model import Perceptron
model = Perceptron(random_state=42)
data = malware_dataset['data'][:, 2:] #if we try to learn from the first 2 features, the perceptron will take a dive
print('''Our algorithm has:
- scores: %s
- mean: %f
''' % learn(model, data, target))
```
It would be pretty weird if the feature set we just constructed had any linear way to separate it into classes.
So any linear model is out of the question, but there are weaker models than Ada Boost to try against our feature set:
- ***Decision Tree Classifier***
This is the base classifier for both the Random Forest algorithm and for Ada Boost (at least in scikit-learn's implementation).
It's basically an algorithm that tries to learn a Decision Tree to classify the problem at hand, using several heuristics. The learned Decision Tree isn't guaranteed to be the optimal one, as that would entail solving an NP-complete problem and breaking everything.
```
model = DecisionTreeClassifier(random_state=42)
print('''Our algorithm has:
- scores: %s
- mean: %f
''' % learn(model, data, target))
```
- ***Naive Bayes***
Since ours is a simple classification job with only 2 classes, we might aswell try the most used classifier out there, Naive Bayes.
Now, this might not be a really good idea, since Naive Bayes assumes each feature to be independent from the others (and it's not really our case), but it's worth a try since it usually works anyway.
We'll try 3 different implementation of Naive Bayes, with different assumed probability distributions.
```
from sklearn.naive_bayes import MultinomialNB, GaussianNB, BernoulliNB
models = [MultinomialNB(), GaussianNB(), BernoulliNB()]
#try different first and last_index (0, 2)
first_index = 0
last_index = 4
data = malware_dataset['data'][:, first_index : last_index]
for model in models:
print('''Our algorithm has:
- scores: %s
- mean: %f
''' % learn(model, data, target))
```
- ***Support Vector Machines***
SVMs technically are non-probabilistic binary linear classifiers, but with the kernel trick they can easily perform non-linear classifications.
There are lots of parameters for SVMs (gamma, tolerance, penalty [...]) and of course the various kernels, so we'll see a handy way to automate the choice of these parameters with Grid Search.
```
from sklearn.svm import SVC
kernels = ['linear', 'poly', 'rbf', 'sigmoid']
model = SVC(kernel='rbf')
print('''Our algorithm has:
- scores: %s
- mean: %f
''' % learn(model, data, target))
```
Instead of manually trying out new models and parameters, we can automate everything using the handy GridSearch:
```
from sklearn.grid_search import GridSearchCV
param_grid = {'C': [0.001, 0.01, 0.1, 1, 10],
'gamma': [0.001, 0.01, 0.1, 1],
'kernel' : ['rbf', 'sigmoid'], #poly and linear hang up the whole notebook, beware
'degree' : [3, 4, 5]}
KF = KFold(len(target), n_folds=2, random_state=42)
grid = GridSearchCV(SVC(), param_grid=param_grid, cv=kf)#, verbose=3) #uncomment to see lots of prints
first_index = 0
last_index = 10
data = malware_dataset['data'][:, first_index : last_index]
grid.fit(data, target)
print(grid.best_score_)
print(grid.best_params_)
```
## 8. Future Improvements
This notebook might be updated or divided into more notebooks (it is pretty long), anyway there's lots of directions to take from here.
a. Ulterior Feature Mining
I doubt that we've found the best features for this classification job.
b. Dimensionality Reduction
Once we've mined for more features we can try to reduce the dimensionality of the problem using:
- Isomap
- TSNE (this works very well apparently)
c. Biclustering
| github_jupyter |
```
def fun_add(x,y):
return x+y
def fun_cal(a,b,c):
return fun_add(a,b)*c
fun_cal(12,13.98,-4)
fun_cal(12,43,4/3)
(lambda a,b,c:(lambda x,y:x+y)(a,b)*c)(12,-3,4/3)
(lambda a,b,c:(lambda x,y:x+y)(a,b)*c)(12,-3.89,4/3)
fun=lambda x,y:x+y
(lambda x,y:x+y)(-3/4,4.56)
(lambda a,b,c:(lambda x,y:x+y)(a,b)*c)(12,-4,4.789)
(lambda x,y,z:(lambda x,y:x+y)(x,y)*z)(12,-4,4.789)
def factorial(x):
if x>=0:
if x==0 or x==1:
return 1
else:
return x*factorial(x-1)
else:
print("Enter a non negative value!!!")
factorial(5)
factorial(-18)
factorial(7/9)
def factorial(x):
if x>=0:
if x==0 or x==1:
return 1
else:
return x*factorial(x+1)
else:
print("Enter a non negative value!!!")
factorial(14)
def fun_add(x):
if x>=1 and isinstance(x,int):
if x==1:
return 1
else:
return x+fun_add(x-1)
else:
print("Enter a non negative Integer value!!!")
fun_add(10)
def fun_add(x):
if x>=1 and isinstance(x,int):
if x==1:
return 1
else:
return x-fun_add(x-1)
else:
print("Enter a non negative Integer value!!!")
fun_add(15)
fun_add(12)
fun_add(-11)
fun_add(3/4)
fun_add(-3/4)
def fun(a):
if n<15:
print(n)
else:
print(n)
fun(a/15)
fun(10000)
def fun(a):
if a<15:
print(a)
else:
print(a)
fun(a/15)
fun(10000)
def fun(a):
if a<15:
print(a)
else:
print(a)
fun(a/15)
print(a)
fun(10000)
def fun(n):
return 2*n
fun(10)
def fun(*n):
return 2*n
fun(2)
fun(2,4)
def fun(*n):
return n
fun(2,4,5,6)
def fun(*n):
return n
T=fun(1,34,45,23)
T
def result(*mark):
return mark
result(13,34,56)
def result(*mark):
return["Pass" if m>=50 else "Fail" for m in mark]
result(25,34,56,67,68,75,85,95)
type(25)
type(result)
def fun(**n):
return n
fun(a=34,b=67,c=-67,d=4/3)
type(fun)
e=fun(a=34,b=67,c=-67,d=4/3)
e.items()
e.values()
e.keys()
e.extend()
e.pop()
a="Boy"
r="Cat"
s="Cat is catching the mouse"
type(a)
type(r)
type(s)
x="Kane is a nice boy"
len(x)
len(" ")
len(" ")
type(" ")
print(" ")
print("K")
x.pop()
x.append()
x.add()
x="Kane is a nice boy"
y="Sam is a good boy"
print("Today is a "+"beautiful day")
print("Today is a ","beautiful day")
x+y
m=x+y
m
x*3
x+3
y*3
n="Today is a sunny day"
m
m[0]
m[6]
m[5]
m[:2]
m[1:3]
m[-1]
m[-4]
m[6]
not "t" in n
s="Dog"
k="-67"
f="5/6"
str(f)
int(k)
v="8.976"
float(v)
int(float(v))
name=input("Enter your name:")
age=input("Enter your age:")
msg=name+" Is a good boy"+" and he is "+age+" years old"
msg
name="Harry"
age=29
msg="Hello (name), are you (age) years old?"
msg
name=input("Enter your name:")
age=21
msg=name+" Is a good boy"+" and he is "+str(age)+" years old"
msg
name="Harry"
age=29
msg="Hello (Harry), are you (29) years old?" .format(name,age)
msg
12+34
a=86
ord("A")
ord("B")
ord("&")
ord("*")
ord("^")
ord("%")
ord("#")
ord("@")
ord("!")
chr(33)
y="My name is kane"
y.replace("s","p")
y.add("name","names")
y.app("s","p")
y.replace("s"+"p")
y.upper()
y.lower()
y.swapcase()
y.title()
y.count("Name")
y.count("name")
y.find("name")
x="Happy birth day"
x.startswith("day")
x.endswith("Happy")
x.isalnum()
x.isalpha()
x.isdigit()
x.islower()
x.isupper()
msg="Today,is,a,rainy,day"
msg.split("a")
msg.split(",")
",".join(msg)
"****".join(msg)
msg="Today is a rainy day.But it may be sun shining in the evening"
msg
msg="Today is a rainy day.\b But it may be sun shining in the evening"
print(msg)
msg="Today is a rainy day.\n But it may be sun shining in the evening"
print(msg)
msg="Today is a rainy day.\t But it may be sun shining in the evening"
print(msg)
msg="Today is a rainy day.\\ But it may be sun shining in the evening"
print(msg)
```
| github_jupyter |
# Aqua Circuit Interoperability Update
_Donny Greenberg, Julien Gacon, Ali Javadi, Steve Wood, 24-Mar-19_
## Basic Vision & End Goal
* Make Aqua use circuits as a first-class currency, and feel more like an algorithms library _next to_ Terra, as users expect, rather than an independent library on top of it
* No more `construct_circuit()` wrappers in Aqua
* Promote Aqua’s best circuity features into Terra to be broadly useful
## Proposal - Three steps
1. Circuit as a First-class Citizen in Aqua
1. Aqua algorithms accept circuits directly, no more circuit wrappers
1. Circuit Library with Enhanced QuantumCircuit Families and Convenient Prebuilts
1. Destination for most of Aqua's enhanced circuit wrappers
1. Critically, allows for lazily constructed circuit placeholders.
1. Usability Improvements to Promote up to QuantumCircuit
1. Make circuit construction in Terra more powerful with features in Aqua users like
## 1. Circuit as a First-class Citizen in Aqua
* Anywhere previously calling `construct_circuit` now accepts circuits as-is, no questions asked
* Typehints ask for a circuit, and are indifferent whether a circuit is from the circuit library (below)
* Fully backwards compatible with Aqua's `construct_circuit`-based objects as long as we like
* Maybe warnings where behavior is strange, e.g. no parameters in VQE ansatz
### Demo - VQC with newly built circuits
Below, we demonstrate the execution of the Variational Quantum Classifier using no special circuit construction objects.
```
from qiskit import QuantumCircuit
from qiskit.circuit import ParameterVector
from qiskit.aqua.algorithms import VQC
from qiskit.aqua.components.optimizers import SLSQP
import numpy as np
import itertools
# Learning the one-hot encoding
train_feats = np.eye(3).tolist()
train_labels = [1,2,3]
train = dict(zip(train_labels, train_feats))
print(train)
feat_params = ParameterVector('ɸ', length=len(train_feats[0]))
feat_map = QuantumCircuit(3)
depth = 3
for _ in range(depth):
feat_map.h(qubit=range(3))
[feat_map.rz(phi=p, qubit=i) for i, p in enumerate(feat_params)]
[feat_map.crz(theta=p1*p2, control_qubit=q1, target_qubit=q2)
for ((q1, p1), (q2,p2)) in itertools.combinations(enumerate(feat_params), 2)]
feat_map.barrier()
feat_map.draw(output='mpl')
# Note: I need to calculate this number
classifier_params = ParameterVector('θ', length=19)
classifier = QuantumCircuit(3)
depth = 3
cp_iter = iter(classifier_params)
next(cp_iter)
for _ in range(depth):
[classifier.ry(theta=next(cp_iter), qubit=j) for j in classifier.qubits]
[classifier.crx(theta=next(cp_iter), control_qubit=q1, target_qubit=q2)
for (q1, q2) in itertools.combinations(classifier.qubits, 2)]
classifier.barrier()
classifier.draw(output='mpl')
vqc = VQC(optimizer=SLSQP(),
data_circuit=feat_map,
classifier_circuit=classifier,
training_dataset=train,
test_dataset=train)
vqc.run()
```
## 2. Circuit Library with Enhanced Flexibility Circuits and Convenient Prebuilts
_Proposal: Move Aqua's circuit-constructor objects - e.g. Ansatze, QFTs, Arithmetic - into a broadly useful circuit-library as flexible QuantumCircuit objects with enhanced features._
#### New Concepts in the Circuit Library
* Circuit Blueprints: Enhanced QuantumCircuit objects which are lazily populated and constructed, but print and interact as bona-fide circuits.
* Not a new class, simply subclasses of QuantumCircuit which match the QuantumCircuit interface
* Users generally shouldn't notice the difference, unless digging into circuit guts in debugging
* Properties such as `.data`, `.parameters`, etc. which require real circuits, trigger construction and caching of constructed circuit
* Meta-parameters, such as ansatz depth or connectivity, are mutable and edited lightly due to lazy construction. Setters trigger cached circuit wipe
* Circuit Families
* Collections of circuit blueprints or prebuilt circuits with extensions or use-case specific features - e.g. `PermutationCircuit`s can include properties which `ArithmeticCircuit`s do not.
* Allow for more aggressive convenience functionality for specific use cases e.g. Ansatz automatically allocating parameters during construction.
#### Options for Circuit Library Placement
1. Inside Terra, with integration tests
1. Pros - Consistent with user expectations that Terra contains circuit building blocks
1. Cons - Unlike other areas, Terra has many directories in the base qiskit level, include `qiskit/circuit/`. The library could not clearly be a `circuits` directory alongside Terra, but would likely be hidden inside `qiskit/circuit/library/`. May complicate Aqua development, requiring frequent multi-repo PRs and exactly synced releases.
1. Inside Aqua, with Qiskit-wide utility, no Aqua concepts
1. Pros - Can be placed in a `qiskit/circuit_library` directory alongside `qiskit/aqua`, giving clear delineation as an important library of circuits.
1. Cons - Users may not expect to find this in Aqua, and distinction between "complicated Terra gate" (e.g. MCT) and "simple library circuit" would make keeping these so far apart strange.
1. In its own repo
1. Pros - Clear importance and delineation.
1. Cons - Another repo.
#### Options for Circuit Family Organization
The circuit library is still a work in progress. In each of the below options, we can make all of the circuits importable from the base init, allowing us to iterate on the directory organization without breaking changes to circuit usage. This way, we can ensure that the circuits are in the correct location in Qiskit for Aqua to begin using them, rather than wait for the library to be complete, and then execute a breaking change to merge in Aqua's circuits.
1. **Organize By Circuit Purposes**
1. Data Preparation Circuits
1. Data Feature Maps
1. Probability Distributions
1. NLocal Circuits (name pending - needs to reflect purpose and availability of Optimization-specific features and properties, such as optimization_bounds and auto-parameterization).
1. TwoLocalCircuit
1. NLocalCircuit
1. Ry, RyRz, SwapRz
1. Arithmetic Circuits
1. Adders
1. Reciprocals
1. MCTs
1. Hamming weight
1. Basis Change Circuits
1. QFTs
1. QFT Placeholder/Base
1. QFT circuits
1. QWT Circuits
1. DCT Circuits
1. Pauli Basis Change
1. Oracle Circuits
1. Truth table
1. Logical expression
1. Phase oracle
1. Permutation oracle
1. Function Testing Circuits
1. Fourier Checking
1. Hidden shift with Bent Boolean functions
1. Ideal Hidden Linear Function circuits
1. Benchmarking
1. Near-Clifford / graph states
1. Random uniform
1. Quantum Volume
1. CNOT Dihedral (from Ignis)
1. **Organize By Circuit Form Factors** - Organization followed by internal quantum-circuits repo. Methodology is organization by the subcircuits and organization of the circuits themselves.
1. Random uniform
1. NLocal Circuits
1. NLocal, TwoLocal, Ry, RyRz, SwapRz
1. Linear rotation
1. Near-Clifford / graph states
1. Pauli Basis Change
1. Quantum volume
1. Quantum Fourier transform
1. Ideal HLF circuits
1. Hamming weight
1. Hidden shift with bent Boolean functions
1. Multiply-controlled NOT gate
1. IQP circuits
1. Fourier checking
1. Unresolved - It's unclear into which families the following circuits fall in the above grouping:
1. Artimetic
1. QFT vs. CNOT based adders
1. Cutoff vs. long-division based reciprocals
1. Oracle circuits
1. Data Feature Maps
1. Broader than forrelation by accepting different paulis to evolve, rather than just Z^n, but also each only half of a forrelation circuit
1. Can be other classes of hard circuits, not only forrelation-based
1. **Some Purpose, Some Complexity Families** - Allow both circuit purpose families and circuit form-factor families, allowing for custom enhancements or functionality in either family type. Circuits can act as placeholders (e.g. permutation) to be filled in by a choice of several synthesis implementations later. Circuits can also import circuits from other families so both practical and theoretical expectations are met without code duplication.
1. Data Preparation Circuits
1. Data Feature Maps
1. Probability Distributions
1. Arithmetic Circuits
1. Basis Change Circuits
1. Quantum Fourier Transform Circuits
1. Oracle Circuits
1. N Local Circuits
1. NLocal, TwoLocal, Ry, RyRz, SwapRz
1. Near-Clifford / Graph State Circuits
1. Quantum Volume Circuits
1. Ideal Hidden Linear Function Circuits
1. Hamming Weight Circuits
1. Hidden Shift with Bent Boolean Function Circuits
1. Multiply-controlled NOT Gate Circuits
1. IQP Circuits
1. Fourier Checking Circuits
1. **Two Subdirectories** - One corresponding to circuit purpose families, one corresponding to circuit complexity families. All circuits can be imported from `qiskit.circuit.library` so organization is aesthetic.
1. Circuit Purpose Families
1. Data Preparation Circuits
1. Data feature maps
1. probability_distributions
1. NLocal Circuits
1. TwoLocalCircuit, NLocalCircuit, Ry, RyRz, SwapRz
1. Arithmetic Circuits
1. Basis Change Circuits
1. QFT, QWT, DCT
1. Pauli Basis Change
1. Oracle Circuits - Truth table, Logical expression, Phase oracle, Permutation oracle
1. Circuit Complexity Families
1. Random uniform
1. Hardware efficient
1. Near-Clifford / graph states
1. Quantum volume
1. Quantum Fourier transform
1. Ideal HLF circuits
1. Hamming weight
1. Hidden shift with bent Boolean functions
1. Multiply-controlled NOT gate
1. IQP circuits
1. Fourier checking
_Additional Proposal: HardwareEfficient base class - abstract `target_backend` setter (QFTs, adders, NLocal, etc. can choose how to interpret what to do with backend information and provide specially tailored circuits for hardware which are not available by transpilation alone.)_
##### Organization Recommendation: Option 1 or 3
Option 1 or 3 introduces the most broadly useful organization to both the algorithms and complexity theory users, attempting to align with both of their expectations for circuit placement. They provide the greatest chance of mutual exclusivity in families, allowing families to be grouped in whichever way most naturally delineates them from the other circuit families based on industry conventions (rather than attempt at some form of objective delineation, which is unlikely to be future-proof). Minimizing ambiguities is a worthwhile long-term investment when this library could reasonably grow to hundreds of classes.
We recommend beginning with Option 1 _quietly_ to be able to migrate the circuits out of Aqua, and reorganize within the library as it is formally built out and new circuit families are added. This is functionally identical to Option 3, as the form-factor circuit families are not yet ready to be pushed to the library, and will need to be merged as a separate step. This will also allow for more time and deliberation about this complex organization question. Allowing all circuits to be imported from the base circuit library directory prevents the impending reorganizations from introducing breaking changes.
### Demo 2 - Powerful New QuantumCircuit Objects
```
from chemistry.code.molecule import Molecule
from qiskit.chemistry.components.initial_states import HartreeFock
from qiskit.aqua.algorithms import VQE
# from qiskit.circuit_.library import RyRz
from qiskit import BasicAer
qasm = BasicAer.get_backend('qasm_simulator')
hh = Molecule(geometry=[['H', [0., 0., 1.]],
['H', [0., 0.45, 1.]],
])
hamiltonian = hh.get_qubitop_hamiltonian()
molecular_wf = HartreeFock(hh) + RyRz(depth=3, entanglement='linear')
gse = VQE(wf_ansatz=molecular_wf, optimizer=SLSQP, backend=qasm).compute_minimum_eigenvalue(hamiltonian)
print(gse)
molecular_qf.target_backend = IBMQ.get_backend('ibmq_valencia')
```
## 3. QuantumCircuit Usability Improvements
Aqua's circuit_constructors have accumulated many powerful features not present in QuantumCircuit. No changes are strictly needed to QuantumCircuit to support the above proposals, but we can promote some of these improvements up to QuantumCircuit base to make these features broadly available.
* Suggested for immediate promotion:
* Mutable qubit number (delete qubits, or extend circuit) for anonymous register circuits
* `.parameters` returns a list instead of a set, as parameter randomization is inconvenient
* Further opportunities for radical circuit control
* Lazy parameterization - When no parameter is specified in a parameterized standard gate, create a new one for the user. We can do this lazily, and only create the full list when `.parameters` is called, in which case the list is "locked in."
* Lazy broadcasting - Similar to the NLocal Circuits. Allow the user to specify groups of qubits to which to apply a gate in circuit construction, but only construct these duplicates when the circuit data is actually needed. Allow users to manipulate these gate applications.
* What we’d need to do to implement these two
Demo 4 - Interface demo of further opportunities
```
# Working notes - Captures new features but doesn't highlight them exactly
my_c = QuantumCircuit(4)
my_c.h(qubits='all')
my_c.cu3(qubits='full', theta=.5) # other two parameters are set to be parameterized under the hood
my_c.h(qubits='all')
my_c.rz(qubits='all', phi=Parameter('theta')) # Sets the same parameter for all of them
```
| github_jupyter |
```
from PIL import Image
from IPython.display import display
import random
import json
# Each image is made up a series of traits
# The weightings for each trait drive the rarity and add up to 100%
background = ["Blue", "Orange", "Purple", "Red", "Yellow"]
background_weights = [30, 40, 15, 5, 10]
circle = ["Blue", "Green", "Orange", "Red", "Yellow"]
circle_weights = [30, 40, 15, 5, 10]
square = ["Blue", "Green", "Orange", "Red", "Yellow"]
square_weights = [30, 40, 15, 5, 10]
# Dictionary variable for each trait.
# Eech trait corresponds to its file name
background_files = {
"Blue": "blue",
"Orange": "orange",
"Purple": "purple",
"Red": "red",
"Yellow": "yellow",
}
circle_files = {
"Blue": "blue-circle",
"Green": "green-circle",
"Orange": "orange-circle",
"Red": "red-circle",
"Yellow": "yellow-circle"
}
square_files = {
"Blue": "blue-square",
"Green": "green-square",
"Orange": "orange-square",
"Red": "red-square",
"Yellow": "yellow-square"
}
## Generate Traits
TOTAL_IMAGES = 30 # Number of random unique images we want to generate
all_images = []
# A recursive function to generate unique image combinations
def create_new_image():
new_image = {} #
# For each trait category, select a random trait based on the weightings
new_image ["Background"] = random.choices(background, background_weights)[0]
new_image ["Circle"] = random.choices(circle, circle_weights)[0]
new_image ["Square"] = random.choices(square, square_weights)[0]
if new_image in all_images:
return create_new_image()
else:
return new_image
# Generate the unique combinations based on trait weightings
for i in range(TOTAL_IMAGES):
new_trait_image = create_new_image()
all_images.append(new_trait_image)
# Returns true if all images are unique
def all_images_unique(all_images):
seen = list()
return not any(i in seen or seen.append(i) for i in all_images)
print("Are all images unique?", all_images_unique(all_images))
# Add token Id to each image
i = 0
for item in all_images:
item["tokenId"] = i
i = i + 1
print(all_images)
# Get Trait Counts
background_count = {}
for item in background:
background_count[item] = 0
circle_count = {}
for item in circle:
circle_count[item] = 0
square_count = {}
for item in square:
square_count[item] = 0
for image in all_images:
background_count[image["Background"]] += 1
circle_count[image["Circle"]] += 1
square_count[image["Square"]] += 1
print(background_count)
print(circle_count)
print(square_count)
#### Generate Metadata for all Traits
METADATA_FILE_NAME = './metadata/all-traits.json';
with open(METADATA_FILE_NAME, 'w') as outfile:
json.dump(all_images, outfile, indent=4)
#### Generate Images
for item in all_images:
im1 = Image.open(f'./trait-layers/backgrounds/{background_files[item["Background"]]}.jpg').convert('RGBA')
im2 = Image.open(f'./trait-layers/circles/{circle_files[item["Circle"]]}.png').convert('RGBA')
im3 = Image.open(f'./trait-layers/squares/{square_files[item["Square"]]}.png').convert('RGBA')
#Create each composite
com1 = Image.alpha_composite(im1, im2)
com2 = Image.alpha_composite(com1, im3)
#Convert to RGB
rgb_im = com2.convert('RGB')
file_name = str(item["tokenId"]) + ".png"
rgb_im.save("./images/" + file_name)
#### Generate Metadata for each Image
f = open('./metadata/all-traits.json',)
data = json.load(f)
IMAGES_BASE_URI = "ADD_IMAGES_BASE_URI_HERE"
PROJECT_NAME = "ADD_PROJECT_NAME_HERE"
def getAttribute(key, value):
return {
"trait_type": key,
"value": value
}
for i in data:
token_id = i['tokenId']
token = {
"image": IMAGES_BASE_URI + str(token_id) + '.png',
"tokenId": token_id,
"name": PROJECT_NAME + ' ' + str(token_id),
"attributes": []
}
token["attributes"].append(getAttribute("Background", i["Background"]))
token["attributes"].append(getAttribute("Circle", i["Circle"]))
token["attributes"].append(getAttribute("Square", i["Square"]))
with open('./metadata/' + str(token_id), 'w') as outfile:
json.dump(token, outfile, indent=4)
f.close()
```
| github_jupyter |
```
#import os
import re
import pandas as pd
import os
import openpyxl
# pd.set_option('display.max_rows', None)
import numpy as np
import nltk
from matplotlib import pyplot as plt
from konlpy.tag import Okt
from wordcloud import WordCloud
from gensim.corpora.dictionary import Dictionary
from gensim.models.ldamulticore import LdaMulticore
from gensim.models.coherencemodel import CoherenceModel
# import warnings
# warnings.filterwarnings('ignore', category = DeprecationWarning) # pyLDAvis.display 설치 후 발생
#import platform
# import csv
# import time
# import pyLDAvis.gensim
list_file = os.listdir("C:/Users/User/anaconda3/envs/NLP37/news/game/BigKinds/")
# list_file
index = 0
df_original_BigKinds = pd.DataFrame(columns = ['뉴스 식별자', '일자', '언론사', '기고자', '제목', '통합분류1', '통합분류2', '사건/사고 분류1', '사건/사고 분류2', '사건/사고 분류3', '인물', '위치', '기관', '키워드', '특성추출(가중치순 상위 50개)', '본문', 'URL', '분석제외 여부'])
for file in list_file:
index += 1
try:
df_original_BigKinds_temp = pd.read_excel("C:/Users/User/anaconda3/envs/NLP37/news/game/BigKinds/"+ file, engine='openpyxl')
df_original_BigKinds = pd.concat([df_original_BigKinds, df_original_BigKinds_temp])
except Exception as e:
print(e)
df_BigKinds = df_original_BigKinds[['일자', '키워드', '특성추출(가중치순 상위 50개)']]
df_BigKinds.rename(columns = {'키워드':'단어', '특성추출(가중치순 상위 50개)':'키워드'}, inplace = True)
df_BigKinds.to_csv('C:/Users/User/anaconda3/envs/NLP37/news/game/BigKinds_game_keyword.txt', sep = '\t', header=True, index = False)
df_BigKinds = pd.read_csv('C:/Users/User/anaconda3/envs/NLP37/news/game/BigKinds_game_keyword.txt', sep = '\t')
df_BigKinds
# month column 생성
df_BigKinds['연도'] = df_BigKinds['일자'].astype(str).str.slice(start=0, stop=4)
df_BigKinds['월'] = df_BigKinds['일자'].astype(str).str.slice(start=4, stop=6)
df_BigKinds
df_BigKinds.groupby(['연도','월']).count()
df_BigKinds['키워드'] = df_BigKinds['키워드'].str.split(',')
df_BigKinds['단어'] = df_BigKinds['단어'].str.split(',')
df_BigKinds
list_of_lists_keyword = df_BigKinds['키워드'].tolist()
list_of_lists_noun = df_BigKinds['단어'].tolist()
# 불용어 삭제
# f = open('C:/Users/User/anaconda3/envs/NLP37/news/TF/venture_stopword_list.txt', 'r')
# lines = f.readlines()
# list_stopword = []
# for line in lines:
# line = line.replace('\n', '')
# list_stopword.append(line)
# f.close()
# def cleaning(list_of_lists, list_stopword):
# list_of_lists_left = []
# for list_noun in list_of_lists:
# list_left = [noun for noun in list_noun if noun not in list_stopword]
# list_of_lists_left.append(list_left)
# return list_of_lists_left
# list_of_lists_keyword = cleaning(list_of_lists_keyword, list_stopword)
# list_of_lists_noun = cleaning(list_of_lists_noun, list_stopword)
# list of lists 병합
list_keyword = []
rows = len(list_of_lists_keyword)
i = 0
while i < rows:
list_keyword += list_of_lists_keyword[i]
i += 1
list_noun = []
rows = len(list_of_lists_noun)
i = 0
while i < rows:
list_noun += list_of_lists_noun[i]
i += 1
# wordcount_BigKinds = nltk.Text(list_nouns)
wordcount_BigKinds_game_keyword = nltk.Text(list_keyword)
print(wordcount_BigKinds_game_keyword.vocab().most_common(20))
wordcount_BigKinds_game_noun = nltk.Text(list_noun)
print(wordcount_BigKinds_game_noun.vocab().most_common(20))
most_common_words = wordcount_BigKinds_game_keyword.vocab().most_common(20)
# for window : font_path='c:/Windows/Fonts/malgun.ttf'
# for mac : font_path='/Library/Fonts/AppleGothic.ttf'
wordcloud = WordCloud(font_path='c:/Windows/Fonts/malgun.ttf',
relative_scaling = 0.2,
background_color = 'white',
width = 3000, height = 3000,
max_words = 100).generate_from_frequencies(dict(most_common_words))
plt.figure(figsize=(12, 8))
plt.imshow(wordcloud)
plt.axis('off')
plt.show()
wordcloud.to_file('C:/Users/User/anaconda3/envs/NLP37/news/game/BigKinds_game_keyword_wordcloud.png')
most_common_words = wordcount_BigKinds_game_noun.vocab().most_common(20)
# for window : font_path='c:/Windows/Fonts/malgun.ttf'
# for mac : font_path='/Library/Fonts/AppleGothic.ttf'
wordcloud = WordCloud(font_path='c:/Windows/Fonts/malgun.ttf',
relative_scaling = 0.2,
background_color = 'white',
width = 3000, height = 3000,
max_words = 100).generate_from_frequencies(dict(most_common_words))
plt.figure(figsize=(12, 8))
plt.imshow(wordcloud)
plt.axis('off')
plt.show()
wordcloud.to_file('C:/Users/User/anaconda3/envs/NLP37/news/game/BigKinds_game_noun_wordcloud.png')
keyword_dictionary = Dictionary(list_of_lists_keyword)
keyword_corpus = [keyword_dictionary.doc2bow(text) for text in list_of_lists_keyword]
# 출처: 서대호(2019), 잡아라! 텍스트마이닝 with 파이썬
print(keyword_dictionary)
keyword_coherences=[]
keyword_perplexities=[]
for i in range(1, 21):
num_topics = i
# tic = time.time()
lda_model = LdaMulticore(corpus = keyword_corpus, id2word = keyword_dictionary, num_topics = num_topics, passes = 1)
# print('number of topics',p,time.time() - tic)
coherence_model = CoherenceModel(model = lda_model, corpus = keyword_corpus, coherence = 'u_mass')
keyword_coherence = coherence_model.get_coherence()
print("coherence:", keyword_coherence)
keyword_coherences.append(keyword_coherence)
print("perplexity:", lda_model.log_perplexity(keyword_corpus), "\n")
keyword_perplexities.append(lda_model.log_perplexity(keyword_corpus))
with open('C:/Users/User/anaconda3/envs/NLP37/news/game/game_keyword_coherences.txt', 'w') as f:
for coherence_value in keyword_coherences:
f.write(str(coherence_value) +'\n')
with open('C:/Users/User/anaconda3/envs/NLP37/news/game/game_keyword_perplexities.txt', 'w') as f:
for perplexity_score in keyword_perplexities:
f.write(str(perplexity_score) +'\n')
keyword_coherences = []
with open('C:/Users/User/anaconda3/envs/NLP37/news/game/game_keyword_coherences.txt', 'r') as f:
keyword_coherences = [value.rstrip() for value in f.readlines()]
keyword_coherences = [float(i) for i in keyword_coherences]
keyword_perplexities = []
with open('C:/Users/User/anaconda3/envs/NLP37/news/game/game_keyword_perplexities.txt', 'r') as f:
keyword_perplexities = [value.rstrip() for value in f.readlines()]
keyword_perplexities = [float(i) for i in keyword_perplexities]
x = np.arange(1,21)
plt.plot(x, keyword_coherences)
plt.xlabel("number of topics")
plt.ylabel("coherence score")
plt.savefig("C:/Users/User/anaconda3/envs/NLP37/news/game/keyword_game_coherence.png")
plt.show()
x = np.arange(1,21)
plt.plot(x, keyword_perplexities)
plt.xlabel("number of topics")
plt.ylabel("perplexity score")
plt.savefig("C:/Users/User/anaconda3/envs/NLP37/news/game/keyword_game_perplexity.png")
plt.show()
keyword_lda_model = LdaMulticore(corpus = keyword_corpus, num_topics = 9, id2word = keyword_dictionary, passes = 1)
topics = keyword_lda_model.print_topics()
for topic in topics:
print(topic)
topic_words = {};
for i in range(9):
words = keyword_lda_model.show_topic(i, topn = 10)
topic_words['Topic # ' + '{:02d}'.format(i+1)] = [i[0] for i in words]
keyword_topic_table = pd.DataFrame(topic_words)
keyword_topic_table
keyword_topic_table.to_csv('C:/Users/User/anaconda3/envs/NLP37/news/game/BigKinds_keyword_topic_modeling.csv', header=True, index = False, encoding='cp949')
noun_dictionary = Dictionary(list_of_lists_noun)
noun_corpus = [noun_dictionary.doc2bow(text) for text in list_of_lists_noun]
# 출처: 서대호(2019), 잡아라! 텍스트마이닝 with 파이썬
print(noun_dictionary)
noun_coherences=[]
noun_perplexities=[]
for i in range(1, 21):
num_topics = i
# tic = time.time()
lda_model = LdaMulticore(corpus=noun_corpus, id2word = noun_dictionary, num_topics = num_topics, passes = 1)
# print('number of topics',p,time.time() - tic)
coherence_model = CoherenceModel(model = lda_model, corpus = noun_corpus, coherence = 'u_mass')
noun_coherence = coherence_model.get_coherence()
print("coherence:", noun_coherence)
noun_coherences.append(noun_coherence)
print("perplexity:", lda_model.log_perplexity(noun_corpus), "\n")
noun_perplexities.append(lda_model.log_perplexity(noun_corpus))
with open('C:/Users/User/anaconda3/envs/NLP37/news/game/game_noun_coherences.txt', 'w') as f:
for coherence_value in noun_coherences:
f.write(str(coherence_value) +'\n')
with open('C:/Users/User/anaconda3/envs/NLP37/news/game/game_noun_perplexities.txt', 'w') as f:
for perplexity_score in noun_perplexities:
f.write(str(perplexity_score) +'\n')
noun_coherences = []
with open('C:/Users/User/anaconda3/envs/NLP37/news/game/game_noun_coherences.txt', 'r') as f:
noun_coherences = [value.rstrip() for value in f.readlines()]
noun_coherences = [float(i) for i in noun_coherences]
noun_perplexities = []
with open('C:/Users/User/anaconda3/envs/NLP37/news/game/game_noun_perplexities.txt', 'r') as f:
noun_perplexities = [value.rstrip() for value in f.readlines()]
noun_perplexities = [float(i) for i in noun_perplexities]
x = np.arange(1,21)
plt.plot(x, noun_coherences)
plt.xlabel("number of topics")
plt.ylabel("coherence score")
plt.savefig("C:/Users/User/anaconda3/envs/NLP37/news/game/noun_game_coherence.png")
plt.show()
x = np.arange(1,21)
plt.plot(x, noun_perplexities)
plt.xlabel("number of topics")
plt.ylabel("perplexity score")
plt.savefig("C:/Users/User/anaconda3/envs/NLP37/news/game/noun_game_perplexity.png")
plt.show()
noun_lda_model = LdaMulticore(corpus =noun_corpus, num_topics = 5, id2word = noun_dictionary, passes = 1)
topics = noun_lda_model.print_topics()
for topic in topics:
print(topic)
# lda_model_5.show_topic(0, topn = 10)
topic_words = {};
for i in range(5):
words = noun_lda_model.show_topic(i, topn = 10)
topic_words['Topic # ' + '{:02d}'.format(i+1)] = [i[0] for i in words]
noun_topic_table = pd.DataFrame(topic_words)
noun_topic_table
noun_topic_table.to_csv('C:/Users/User/anaconda3/envs/NLP37/news/game/BigKinds_noun_topic_modeling.csv', header=True, index = False, encoding='cp949')
```
lda_display = pyLDAvis.gensim.prepare(lda_model_10, corpus, dictionary, sort_topics = False)
pyLDAvis.display(lda_display)
| github_jupyter |
```
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
movies = pd.read_csv('tmdb_5000_movies.csv')
credits = pd.read_csv('tmdb_5000_credits.csv')
movies.head()
credits.head()
movies = movies.merge(credits,on='title')
movies.head(1)
movies = movies[['movie_id','title','overview','genres','keywords','cast','crew']]
import ast
credits.head(1)['cast'].values
movies.head(1)
movies=movies.merge(credits,on='title')
movies.head(1)
credits.head(1)
movies=movies.merge(credits,on='title')
movies.head(1)
movies = movies[['movie_id','title','overview','genres','keywords','cast','crew']]
movies.isnull().sum()
movies.dropna(inplace=True)
movies.isnull().sum()
movies.duplicated().sum()
movies.iloc[0].genres
def convert(obj):
L = []
for i in ast.literal_eval(obj):
L.append(i['name'])
return L
movies['genres'] = movies['genres'].apply(convert)
movies.head()
movies['keywords'] = movies['keywords'].apply(convert)
movies.head()
def convert3(obj):
L = []
counter = 0
for i in ast.literal_eval(obj):
if counter!=3:
L.append(i['name'])
counter+=1
else:
break
return L
movies['cast'] = movies['cast'].apply(convert3)
movies.head()
def fetch_director(obj):
L = []
for i in ast.literal_eval(obj):
if i['job'] == 'Director':
L.append(i['name'])
break
return L
movies['crew'] = movies['crew'].apply(fetch_director)
movies.head()
movies['overview'][10]
movies['overview'] = movies['overview'].apply(lambda x:x.split())
def collapse(L):
L1 = []
for i in L:
L1.append(i.replace(" ",""))
return L1
movies['cast'] = movies['cast'].apply(collapse)
movies['crew'] = movies['crew'].apply(collapse)
movies['genres'] = movies['genres'].apply(collapse)
movies['keywords'] = movies['keywords'].apply(collapse)
movies.head()
movies['tags'] = movies['overview'] + movies['genres'] + movies['keywords'] + movies['cast'] + movies['crew']
movies.head()
new = movies.drop(columns=['overview','genres','keywords','cast','crew'])
new.head()
new['tags'] = new['tags'].apply(lambda x: " ".join(x))
new.head()
new['tags'] = new['tags'].apply(lambda x:x.lower())
import nltk
from nltk.stem.porter import PorterStemmer
ps=PorterStemmer()
def stem(text):
y=[]
for i in text.split():
y.append(ps.stem(i))
return " ".join(y)
new['tags']=new['tags'].apply(stem)
from sklearn.feature_extraction.text import CountVectorizer
cv = CountVectorizer(max_features=5000,stop_words='english')
vector = cv.fit_transform(new['tags']).toarray()
new.head()
from sklearn.metrics.pairwise import cosine_similarity
similarity = cosine_similarity(vector)
similarity[1]
def recommend(movie):
index = new[new['title'] == movie].index[0]
distances = sorted(list(enumerate(similarity[index])),reverse=True,key = lambda x: x[1])
for i in distances[1:6]:
print(new.iloc[i[0]].title)
recommend('Gandhi')
import pickle
pickle.dump(new,open('movie_list.pkl','wb'))
pickle.dump(similarity,open('similarity.pkl','wb'))
pickle.dump(new.to_dict(),open('movie_dict.pkl','wb'))
```
| github_jupyter |
# TensorFlow Visual Recognition Sample Application Part 1
## Define the model metadata
```
import tensorflow as tf
import requests
models = {
"mobilenet": {
"base_url":"https://github.com/DTAIEB/Thoughtful-Data-Science/raw/master/chapter%206/Visual%20Recognition/mobilenet_v1_0.50_224",
"model_file_url": "frozen_graph.pb",
"label_file": "labels.txt",
"output_layer": "MobilenetV1/Predictions/Softmax"
}
}
# helper method for reading attributes from the model metadata
def get_model_attribute(model, key, default_value = None):
if key not in model:
if default_value is None:
raise Exception("Require model attribute {} not found".format(key))
return default_value
return model[key]
```
## Helper methods for loading the graph and labels for a given model
```
# Helper method for resolving url relative to the selected model
def get_url(model, path):
return model["base_url"] + "/" + path
# Download the serialized model and create a TensorFlow graph
def load_graph(model):
graph = tf.Graph()
graph_def = tf.GraphDef()
graph_def.ParseFromString(
requests.get( get_url( model, model["model_file_url"] ) ).content
)
with graph.as_default():
tf.import_graph_def(graph_def)
return graph
# Load the labels
def load_labels(model, as_json = False):
labels = [line.rstrip() \
for line in requests.get( get_url( model, model["label_file"] ) ).text.split("\n") \
if line != ""]
if as_json:
return [{"index": item.split(":")[0], "label" : item.split(":")[1]} for item in labels]
return labels
```
## Use BeautifulSoup to scrape the images from a given url
```
from bs4 import BeautifulSoup as BS
import re
# return an array of all the images scraped from an html page
def get_image_urls(url):
# Instantiate a BeautifulSoup parser
soup = BS(requests.get(url).text, "html.parser")
# Local helper method for extracting url
def extract_url(val):
m = re.match(r"url\((.*)\)", val)
val = m.group(1) if m is not None else val
return "http:" + val if val.startswith("//") else val
# List comprehension that look for <img> elements and backgroud-image styles
return [extract_url(imgtag['src']) for imgtag in soup.find_all('img')] + [ \
extract_url(val.strip()) for key,val in \
[tuple(selector.split(":")) for elt in soup.select("[style]") \
for selector in elt["style"].strip(" ;").split(";")] \
if key.strip().lower()=='background-image' \
]
```
## Helper method for downloading an image into a temp file
```
import tempfile
def download_image(url):
response = requests.get(url, stream=True)
if response.status_code == 200:
with tempfile.NamedTemporaryFile(delete=False) as f:
for chunk in response.iter_content(2048):
f.write(chunk)
return f.name
else:
raise Exception("Unable to download image: {}".format(response.status_code))
```
## Decode an image into a tensor
```
# decode a given image into a tensor
def read_tensor_from_image_file(model, file_name):
file_reader = tf.read_file(file_name, "file_reader")
if file_name.endswith(".png"):
image_reader = tf.image.decode_png(file_reader, channels = 3,name='png_reader')
elif file_name.endswith(".gif"):
image_reader = tf.squeeze(tf.image.decode_gif(file_reader,name='gif_reader'))
elif file_name.endswith(".bmp"):
image_reader = tf.image.decode_bmp(file_reader, name='bmp_reader')
else:
image_reader = tf.image.decode_jpeg(file_reader, channels = 3, name='jpeg_reader')
float_caster = tf.cast(image_reader, tf.float32)
dims_expander = tf.expand_dims(float_caster, 0);
# Read some info from the model metadata, providing default values
input_height = get_model_attribute(model, "input_height", 224)
input_width = get_model_attribute(model, "input_width", 224)
input_mean = get_model_attribute(model, "input_mean", 0)
input_std = get_model_attribute(model, "input_std", 255)
resized = tf.image.resize_bilinear(dims_expander, [input_height, input_width])
normalized = tf.divide(tf.subtract(resized, [input_mean]), [input_std])
sess = tf.Session()
result = sess.run(normalized)
return result
```
## Score_image method that run the model and return the top 5 candidate answers
```
import numpy as np
# classify an image given its url
def score_image(graph, model, url):
# Get the input and output layer from the model
input_layer = get_model_attribute(model, "input_layer", "input")
output_layer = get_model_attribute(model, "output_layer")
# Download the image and build a tensor from its data
t = read_tensor_from_image_file(model, download_image(url))
# Retrieve the tensors corresponding to the input and output layers
input_tensor = graph.get_tensor_by_name("import/" + input_layer + ":0");
output_tensor = graph.get_tensor_by_name("import/" + output_layer + ":0");
with tf.Session(graph=graph) as sess:
# Execute the output, overriding the input tensor with the one corresponding
# to the image in the feed_dict argument
results = sess.run(output_tensor, {input_tensor: t})
results = np.squeeze(results)
# select the top 5 candidate and match them to the labels
top_k = results.argsort()[-5:][::-1]
labels = load_labels(model)
return [(labels[i].split(":")[1], results[i]) for i in top_k]
```
## Test the model using a Flickr page
```
model = models['mobilenet']
graph = load_graph(model)
image_urls = get_image_urls("https://www.flickr.com/search/?text=cats")
for url in image_urls:
results = score_image(graph, model, url)
print("Results for {}: \n\t{}".format(url, results))
```
| github_jupyter |
# k-NN movie reccomendation
| User\Film | Movie A | Movie B | Movie C | ... | Movie # |
|---------------------------------------------------------|
| **User A**| 3 | 4 | 0 | ... | 5 |
| **User B**| 0 | 3 | 2 | ... | 0 |
| **User C**| 4 | 1 | 3 | ... | 4 |
| **User D**| 5 | 3 | 2 | ... | 3 |
| ... | ... | ... | ... | ... | ... |
| **User #**| 2 | 1 | 1 | ... | 4 |
Task: For a new user find k similar users based on movie rating and recommend few new, previously unseen, movies to the new user. Use mean rating of k users to find which one to recommend. Use cosine similarity as distance function. User didnt't see a movie if he didn't rate the movie.
```
# Import necessary libraries
import tensorflow as tf
import numpy as np
# Define paramaters
set_size = 1000 # Number of users in dataset
n_features = 300 # Number of movies in dataset
K = 3 # Number of similary users
n_movies = 6 # Number of movies to reccomend
# Generate dummy data
data = np.array(np.random.randint(0, 6, size=(set_size, n_features)), dtype=np.float32)
new_user = np.array(np.random.randint(0, 6, size=(1, n_features)), dtype=np.float32)
# Find the number of movies that user did not rate
not_rated = np.count_nonzero(new_user == 0)
# Case in which the new user rated all movies in our dataset
if not_rated == 0:
print('Regenerate new user')
# Case in which we try to recommend more movies than user didn't see
if not_rated < n_movies:
print('Regenerate new user')
# Print few examples
# print(data[:3])
# print(new_user)
# Input train vector
X1 = tf.placeholder(dtype=tf.float32, shape=[None, n_features], name="X1")
# Input test vector
X2 = tf.placeholder(dtype=tf.float32, shape=[1, n_features], name="X2")
# Implement finding the k nearest users
```
# Locally weighted regression (LOWESS)
```
# Import necessary libraries
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# Load data as numpy array
x, y = np.loadtxt('../../data/02_LinearRegression/polynomial.csv', delimiter=',', unpack=True)
m = x.shape[0]
x = (x - np.mean(x, axis=0)) / np.std(x, axis=0)
y = (y - np.mean(y)) / np.std(y)
# Graphical preview
%matplotlib inline
fig, ax = plt.subplots()
ax.set_xlabel('X Labe')
ax.set_ylabel('Y Label')
ax.scatter(x, y, edgecolors='k', label='Data')
ax.grid(True, color='gray', linestyle='dashed')
X = tf.placeholder(tf.float32, name='X')
Y = tf.placeholder(tf.float32, name='Y')
w = tf.Variable(0.0, name='weights')
b = tf.Variable(0.0, name='bias')
# TODO: create model, cost function and optimization
with tf.Session() as sess:
# Initialize the necessary variables, in this case, w and b
sess.run(tf.global_variables_initializer())
# TODO: Implement optimization
# Output the values of w and b
w1, b1 = sess.run([w, b])
print(sess.run(t_w, feed_dict={X: 1.4}))
print('W: %f, b: %f' % (w1, b1))
print('Cost: %f' % sess.run(cost, feed_dict={X: x, Y: y}))
# Append hypothesis that we found on the plot
x1 = np.linspace(-1.0, 0.0, 50)
ax.plot(x1, x1 * w1 + b1, color='r', label='Predicted')
ax.plot(x1, np.exp(-(x1 - point_x) ** 2 / (2 * 0.15 ** 2)), color='g', label='Weight function')
ax.legend()
fig
```
| github_jupyter |
## Quantum Fourier Transform
```
import numpy as np
from numpy import pi
from qiskit import QuantumCircuit, transpile, assemble, Aer, IBMQ
from qiskit.providers.ibmq import least_busy
from qiskit.tools.monitor import job_monitor
from qiskit.visualization import plot_histogram, plot_bloch_multivector
# doing it for a 3 qubit case
qc = QuantumCircuit(3)
qc.h(2)
qc.draw('mpl')
# we want to turn this to extra quarter if qubit 1 is in |1>
# apply the CROT from qubit 1 to to qubit 2
qc.cp(pi/2,1,2)
qc.draw('mpl')
# we want an another eighsths turn if the least significant bit
# 0 has the value |1>
# apply CROT from qubit 2 to qubit 1
qc.cp(pi/4,0,2)
qc.draw('mpl')
# doing the same for the rest two qubits
qc.h(1)
qc.cp(pi/2,0,1)
qc.h(0)
qc.draw('mpl')
# and then swap the 0 and 2 qubit to complete the QFT
qc.swap(0,2)
qc.draw('mpl')
```
This is one way to create the QFT circuit, but we can also make a function to make that.
```
def qft_rotations(circuit,n):
if n == 0:
return circuit
n -= 1
circuit.h(0)
for qubit in range(n):
circuit.cp(pi/2**(n-qubit), qubit,n)
# so
qc = QuantumCircuit(4)
qft_rotations(qc,4)
qc.draw('mpl')
# how scaling works
from qiskit_textbook.widgets import scalable_circuit
scalable_circuit(qft_rotations)
# we can modify the prev function
def qft_rotations(circuit,n):
if n == 0:
return circuit
n -= 1
circuit.h(n)
for qubit in range(n):
circuit.cp(pi/2**(n-qubit), qubit,n)
qft_rotations(circuit,n)
qc = QuantumCircuit(4)
qft_rotations(qc,4)
qc.draw('mpl')
scalable_circuit(qft_rotations)
# now adding the swap gates
def swap_registeres(circuit, n):
for qubit in range(n//2):
circuit.swap(qubit, n-qubit-1)
return circuit
def qft(circuit,n):
qft_rotations(circuit,n)
swap_registeres(circuit,n)
return circuit
qc = QuantumCircuit(8)
qft(qc,8)
qc.draw('mpl')
scalable_circuit(qft)
```
## How the Circuit Works?
```
bin(7)
# encode this
qc = QuantumCircuit(3)
for i in range(3):
qc.x(i)
qc.draw('mpl')
# display in the aer simulator
sim = Aer.get_backend("aer_simulator")
qc_init = qc.copy()
qc_init.save_statevector()
statevector = sim.run(qc_init).result().get_statevector()
plot_bloch_multivector(statevector)
# now call the qft function
qft(qc,3)
qc.draw('mpl')
qc.save_statevector()
statevector = sim.run(qc).result().get_statevector()
plot_bloch_multivector(statevector)
```
### Running it on Real Quantum Device
```
def inverse_qft(circuit,n):
qft_circ = qft(QuantumCircuit(n), n)
invqft_circuit = qft_circ.inverse()
# add it to first n qubits
circuit.append(invqft_circuit, circuit.qubits[:n])
return circuit.decompose()
# now do it fo the 7
nqubits = 3
number = 7
qc = QuantumCircuit(nqubits)
for qubit in range(nqubits):
qc.h(qubit)
qc.p(number*pi/4,0)
qc.p(number*pi/2,1)
qc.p(number*pi,2)
qc.draw('mpl')
qc_init = qc.copy()
qc_init.save_statevector()
sim = Aer.get_backend("aer_simulator")
statevector = sim.run(qc_init).result().get_statevector()
plot_bloch_multivector(statevector)
# now the inverse QFT
qc = inverse_qft(qc, nqubits)
qc.measure_all()
qc.draw('mpl')
# Load our saved IBMQ accounts and get the least busy backend device with less than or equal to nqubits
IBMQ.load_account()
provider = IBMQ.get_provider(hub='ibm-q')
backend = least_busy(provider.backends(filters=lambda x: x.configuration().n_qubits >= nqubits
and not x.configuration().simulator
and x.status().operational==True))
print("least busy backend: ", backend)
shots = 2048
transpiled_qc = transpile(qc, backend, optimization_level=3)
job = backend.run(transpiled_qc, shots=shots)
job_monitor(job)
counts = job.result().get_counts()
plot_histogram(counts)
```
| github_jupyter |
# Expression trees in PyBaMM
The basic data structure that PyBaMM uses to express models is an expression tree. This data structure encodes a tree representation of a given equation. The expression tree is used to encode the equations of both the original symbolic model, and the discretised equations of that model. Once discretised, the model equations are then passed to the solver, which must then evaluate the discretised expression trees in order to perform the time-stepping.
The expression tree must therefore satisfy three requirements:
1. To encode the model equations, it must be able to encode an arbitrary equation, including unary and binary operators such as `*`, `-`, spatial gradients or divergence, symbolic parameters, scalar, matrices and vectors.
2. To perform the time-stepping, it must be able to be evaluated, given the current state vector $\mathbf{y}$ and the current time $t$
3. For solvers that require it, its gradient with respect to a given variable must be able to be evaluated (once again given $\mathbf{y}$ and $t$)
As an initial example, the code below shows how to construct an expression tree of the equation $2y(1 - y) + t$. We use the `pybamm.StateVector` to represent $\mathbf{y}$, which in this case will be a vector of size 1. The time variable $t$ is already provided by PyBaMM and is of class `pybamm.Time`.
```
%pip install pybamm -q # install PyBaMM if it is not installed
import pybamm
import numpy as np
y = pybamm.StateVector(slice(0,1))
t = pybamm.t
equation = 2*y * (1 - y) + t
equation.visualise('expression_tree1.png')
```

Once the equation is constructed, we can evaluate it at a given $t=1$ and $\mathbf{y}=\begin{pmatrix} 2 \end{pmatrix}$.
```
equation.evaluate(1, np.array([2]))
```
We can also calculate the expression tree representing the gradient of the equation with respect to $t$,
```
diff_wrt_equation = equation.diff(t)
diff_wrt_equation.visualise('expression_tree2.png')
```

...and evaluate this expression,
```
diff_wrt_equation.evaluate(t=1, y=np.array([2]), y_dot=np.array([2]))
```
## The PyBaMM Pipeline
Proposing, parameter setting and discretising a model in PyBaMM is a pipeline process, consisting of the following steps:
1. The model is proposed, consisting of equations representing the right-hand-side of an ordinary differential equation (ODE), and/or algebraic equations for a differential algebraic equation (DAE), and also associated boundary condition equations
2. The parameters present in the model are replaced by actual scalar values from a parameter file, using the [`pybamm.ParamterValues`](https://pybamm.readthedocs.io/en/latest/source/parameters/parameter_values.html) class
3. The equations in the model are discretised onto a mesh, any spatial gradients are replaced with linear algebra expressions and the variables of the model are replaced with state vector slices. This is done using the [`pybamm.Discretisation`](https://pybamm.readthedocs.io/en/latest/source/spatial_methods/discretisation.html) class.
## Stage 1 - Symbolic Expression Trees
At each stage, the expression tree consists of certain types of nodes. In the first stage, the model is first proposed using [`pybamm.Parameter`](https://pybamm.readthedocs.io/en/latest/source/expression_tree/parameter.html), [`pybamm.Variable`](https://pybamm.readthedocs.io/en/latest/source/expression_tree/variable.html), and other [unary](https://pybamm.readthedocs.io/en/latest/source/expression_tree/unary_operator.html) and [binary](https://pybamm.readthedocs.io/en/latest/source/expression_tree/binary_operator.html) operators (which also includes spatial operators such as [`pybamm.Gradient`](https://pybamm.readthedocs.io/en/latest/source/expression_tree/unary_operator.html#pybamm.Gradient) and [`pybamm.Divergence`](https://pybamm.readthedocs.io/en/latest/source/expression_tree/unary_operator.html#pybamm.Divergence)). For example, the right hand side of the equation
$$\frac{d c}{dt} = D \nabla \cdot \nabla c$$
can be constructed as an expression tree like so:
```
D = pybamm.Parameter('D')
c = pybamm.Variable('c', domain=['negative electrode'])
dcdt = D * pybamm.div(pybamm.grad(c))
dcdt.visualise('expression_tree3.png')
```

## Stage 2 - Setting parameters
In the second stage, the `pybamm.ParameterValues` class is used to replace all the parameter nodes with scalar values, according to an input parameter file. For example, we'll use a this class to set $D = 2$
```
parameter_values = pybamm.ParameterValues({'D': 2})
dcdt = parameter_values.process_symbol(dcdt)
dcdt.visualise('expression_tree4.png')
```

## Stage 3 - Linear Algebra Expression Trees
The third and final stage uses the `pybamm.Discretisation` class to discretise the spatial gradients and variables over a given mesh. After this stage the expression tree will encode a linear algebra expression that can be evaluated given the state vector $\mathbf{y}$ and $t$.
**Note:** for demonstration purposes, we use a dummy discretisation below. For a more complete description of the `pybamm.Discretisation` class, see the example notebook [here](https://github.com/pybamm-team/PyBaMM/blob/develop/examples/notebooks/spatial_methods/finite-volumes.ipynb).
```
from tests import get_discretisation_for_testing
disc = get_discretisation_for_testing()
disc.y_slices = {c.id: [slice(0, 40)]}
dcdt = disc.process_symbol(dcdt)
dcdt.visualise('expression_tree5.png')
```

After the third stage, our expression tree is now able to be evaluated by one of the solver classes. Note that we have used a single equation above to illustrate the different types of expression trees in PyBaMM, but any given models will consist of many RHS or algebraic equations, along with boundary conditions. See [here](https://github.com/pybamm-team/PyBaMM/tree/develop/examples/notebooks/Creating%20Models) for more details of PyBaMM models.
## References
The relevant papers for this notebook are:
```
pybamm.print_citations()
```
| github_jupyter |
# Kinematic chain in a plane (2D)
> Marcos Duarte, Renato Naville Watanabe
> [Laboratory of Biomechanics and Motor Control](http://pesquisa.ufabc.edu.br/bmclab)
> Federal University of ABC, Brazil
<h1>Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Properties-of-kinematic-chains" data-toc-modified-id="Properties-of-kinematic-chains-1"><span class="toc-item-num">1 </span>Properties of kinematic chains</a></span></li><li><span><a href="#The-kinematics-of-one-link-system" data-toc-modified-id="The-kinematics-of-one-link-system-2"><span class="toc-item-num">2 </span>The kinematics of one-link system</a></span><ul class="toc-item"><li><span><a href="#Forward-and-inverse-kinematics" data-toc-modified-id="Forward-and-inverse-kinematics-2.1"><span class="toc-item-num">2.1 </span>Forward and inverse kinematics</a></span></li><li><span><a href="#Matrix-representation-of-the-kinematics" data-toc-modified-id="Matrix-representation-of-the-kinematics-2.2"><span class="toc-item-num">2.2 </span>Matrix representation of the kinematics </a></span></li></ul></li><li><span><a href="#Differential-kinematics" data-toc-modified-id="Differential-kinematics-3"><span class="toc-item-num">3 </span>Differential kinematics</a></span><ul class="toc-item"><li><span><a href="#Linear-velocity-of-the-endpoint" data-toc-modified-id="Linear-velocity-of-the-endpoint-3.1"><span class="toc-item-num">3.1 </span>Linear velocity of the endpoint</a></span></li><li><span><a href="#Linear-acceleration-of-the-endpoint" data-toc-modified-id="Linear-acceleration-of-the-endpoint-3.2"><span class="toc-item-num">3.2 </span>Linear acceleration of the endpoint</a></span><ul class="toc-item"><li><span><a href="#Tangential-acceleration" data-toc-modified-id="Tangential-acceleration-3.2.1"><span class="toc-item-num">3.2.1 </span>Tangential acceleration</a></span></li><li><span><a href="#Centripetal-acceleration" data-toc-modified-id="Centripetal-acceleration-3.2.2"><span class="toc-item-num">3.2.2 </span>Centripetal acceleration</a></span></li></ul></li><li><span><a href="#Jacobian-matrix" data-toc-modified-id="Jacobian-matrix-3.3"><span class="toc-item-num">3.3 </span>Jacobian matrix</a></span></li><li><span><a href="#Derivative-of-a-vector-valued-function-using-the-Jacobian-matrix" data-toc-modified-id="Derivative-of-a-vector-valued-function-using-the-Jacobian-matrix-3.4"><span class="toc-item-num">3.4 </span>Derivative of a vector-valued function using the Jacobian matrix</a></span></li><li><span><a href="#Jacobian-matrix-in-the-context-of-kinematic-chains" data-toc-modified-id="Jacobian-matrix-in-the-context-of-kinematic-chains-3.5"><span class="toc-item-num">3.5 </span>Jacobian matrix in the context of kinematic chains</a></span><ul class="toc-item"><li><span><a href="#Jacobian-matrix-of-one-link-chain" data-toc-modified-id="Jacobian-matrix-of-one-link-chain-3.5.1"><span class="toc-item-num">3.5.1 </span>Jacobian matrix of one-link chain</a></span></li></ul></li></ul></li><li><span><a href="#The-kinematics-of-a-two-link-chain" data-toc-modified-id="The-kinematics-of-a-two-link-chain-4"><span class="toc-item-num">4 </span>The kinematics of a two-link chain</a></span><ul class="toc-item"><li><span><a href="#Joint-and-segment-angles" data-toc-modified-id="Joint-and-segment-angles-4.1"><span class="toc-item-num">4.1 </span>Joint and segment angles</a></span></li><li><span><a href="#Inverse-kinematics" data-toc-modified-id="Inverse-kinematics-4.2"><span class="toc-item-num">4.2 </span>Inverse kinematics </a></span></li></ul></li><li><span><a href="#Differential--kinematics" data-toc-modified-id="Differential--kinematics-5"><span class="toc-item-num">5 </span>Differential kinematics</a></span></li><li><span><a href="#Further-reading" data-toc-modified-id="Further-reading-6"><span class="toc-item-num">6 </span>Further reading</a></span></li><li><span><a href="#Video-lectures-on-the-Internet" data-toc-modified-id="Video-lectures-on-the-Internet-7"><span class="toc-item-num">7 </span>Video lectures on the Internet</a></span></li><li><span><a href="#Problems" data-toc-modified-id="Problems-8"><span class="toc-item-num">8 </span>Problems</a></span></li><li><span><a href="#References" data-toc-modified-id="References-9"><span class="toc-item-num">9 </span>References</a></span></li></ul></div>
Kinematic chain refers to an assembly of rigid bodies (links) connected by joints that is the mathematical model for a mechanical system which in turn can represent a biological system such as the human arm ([Wikipedia](http://en.wikipedia.org/wiki/Kinematic_chain)). The term chain refers to the fact that the links are constrained by their connections (typically, by a hinge joint which is also called pin joint or revolute joint) to other links. As consequence of this constraint, a kinematic chain in a plane is an example of circular motion of a rigid object.
Chapter 16 of Ruina and Rudra's book is a good formal introduction on the topic of circular motion of a rigid object. However, in this notebook we will not employ the mathematical formalism introduced in that chapter - the concept of a rotating reference frame and the related rotation matrix - we cover these subjects in the notebooks [Time-varying frame of reference](http://nbviewer.jupyter.org/github/BMClab/BMC/blob/master/notebooks/Time-varying%20frames.ipynb) and [Rigid-body transformations (2D)](https://nbviewer.jupyter.org/github/BMClab/BMC/blob/master/notebooks/Transformation2D.ipynb). Now, we will describe the kinematics of a chain in a Cartesian coordinate system using trigonometry and calculus. This approach is simpler and more intuitive but it gets too complicated for a kinematic chain with many links or in the 3D space. For such more complicated problems, it would be recommended using rigid transformations (see for example, Siciliano et al. (2009)).
We will deduce the kinematic properties of kinematic chains algebraically using [Sympy](http://sympy.org/), a Python library for symbolic mathematics. And in Sympy we could have used the [mechanics module](http://docs.sympy.org/latest/modules/physics/mechanics/index.html), a specific module for creation of symbolic equations of motion for multibody systems, but let's deduce most of the stuff by ourselves to understand the details.
## Properties of kinematic chains
For a kinematic chain, the base is the extremity (origin) of a kinematic chain which is typically considered attached to the ground, body or fixed. The endpoint is the other extremity (end) of a kinematic chain and typically can move. In robotics, the term end-effector is used and usually refers to a last link (rigid body) in this chain.
In topological terms, a kinematic chain is termed open when there is only one sequence of links connecting the two ends of the chain. Otherwise it's termed closed and in this case a sequence of links forms a loop. A kinematic chain can be classified as serial or parallel or a mixed of both. In a serial chain the links are connected in a serial order. A serial chain is an open chain, otherwise it is a parallel chain or a branched chain (e.g., hand and fingers).
Although the definition above is clear and classic in mechanics, it is unfortunately not the definition used by health professionals (clinicians and athletic trainers) when describing human movement. They refer to human joints and segments as a closed or open kinematic (or kinetic) chain simply if the distal segment (typically the foot or hand) is fixed (closed chain) or not (open chain).
In this text we will be consistent with mechanics, but keep in mind this difference when interacting with clinicians and athletic trainers.
Another important term to characterize a kinematic chain is <a href="https://en.wikipedia.org/wiki/Degrees_of_freedom_(mechanics)">degree of freedom (DOF)</a>. In mechanics, the degree of freedom of a mechanical system is the number of independent parameters that define its configuration or that determine the state of a physical system. A particle in the 3D space has three DOFs because we need three coordinates to specify its position. A rigid body in the 3D space has six DOFs because we need three coordinates of one point at the body to specify its position and three angles to to specify its orientation in order to completely define the configuration of the rigid body. For a link attached to a fixed body by a hinge joint in a plane, all we need to define the configuration of the link is one angle and then this link has only one DOF. A kinematic chain with two links in a plane has two DOFs, and so on.
The mobility of a kinematic chain is its total number of degrees of freedom. The redundancy of a kinematic chain is its mobility minus the number of degrees of freedom of the endpoint.
## The kinematics of one-link system
First, let's study the case of a system composed by one planar hinge joint and one link, which technically it's not a chain but it will be useful to review (or introduce) key concepts.
<br>
<figure><img src="./../images/onelink.gif" width=350 alt="onelink"/><figcaption><center><i>Figure. One link attached to a fixed body by a hinge joint in a plane.</i></center></figcaption> </figure>
First, let's import the necessary libraries from Python and its ecosystem:
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
sns.set_context("notebook", font_scale=1.2, rc={"lines.linewidth": 2,
"lines.markersize": 10})
from IPython.display import display, Math
from sympy import Symbol, symbols, Function
from sympy import Matrix, simplify, lambdify, expand, latex
from sympy import diff, cos, sin, sqrt, acos, atan2, atan, Abs
from sympy.vector import CoordSys3D
from sympy.physics.mechanics import dynamicsymbols, mlatex, init_vprinting
init_vprinting()
import sys
sys.path.insert(1, r'./../functions') # add to pythonpath
```
We need to define a Cartesian coordinate system and the symbolic variables, $t$, $\ell$, $\theta$ (and make $\theta$ a function of time):
```
G = CoordSys3D('')
t = Symbol('t')
l = Symbol('ell', real=True, positive=True)
# type \theta and press tab for the Greek letter θ
θ = dynamicsymbols('theta', real=True) # or Function('theta')(t)
```
Using trigonometry, the endpoint position in terms of the joint angle and link length is:
```
r_p = l*cos(θ)*G.i + l*sin(θ)*G.j + 0*G.k
r_p
```
With the components:
```
r_p.components
```
### Forward and inverse kinematics
Computing the configuration of a link or a chain (including the endpoint location) from the joint parameters (joint angles and link lengths) as we have done is called [forward or direct kinematics](https://en.wikipedia.org/wiki/Forward_kinematics).
If the linear coordinates of the endpoint position are known (for example, if they are measured with a motion capture system) and one wants to obtain the joint angle(s), this process is known as [inverse kinematics](https://en.wikipedia.org/wiki/Inverse_kinematics). For the one-link system above:
<span class="notranslate">
$$ \theta = arctan\left(\frac{y_P}{x_P}\right) $$
</span>
### Matrix representation of the kinematics
The mathematical manipulation will be easier if we use the matrix formalism (and let's drop the explicit dependence on <span class="notranslate">$t$</span>):
```
r = Matrix((r_p.dot(G.i), r_p.dot(G.j)))
r
```
Using the matrix formalism will simplify things, but we will loose some of the Sympy methods for vectors (for instance, the variable `r_p` has a method `magnitude` and the variable `r` does not.
If you prefer, you can keep the pure vector representation and just switch to matrix representation when displaying a variable:
```
r_p.to_matrix(G)
```
The third element of the matrix above refers to the <span class="notranslate"> $\hat{\mathbf{k}}$</span> component which is zero for the present case (planar movement).
## Differential kinematics
Differential kinematics gives the relationship between the joint velocities and the corresponding endpoint linear velocity. This mapping is described by a matrix, termed [Jacobian matrix](http://en.wikipedia.org/wiki/Jacobian_matrix_and_determinant), which depends on the kinematic chain configuration and it is of great use in the study of kinematic chains.
First, let's deduce the endpoint velocity without using the Jacobian and then we will see how to calculate the endpoint velocity using the Jacobian matrix.
The velocity of the endpoint can be obtained by the first-order derivative of the position vector. The derivative of a vector is obtained by differentiating each vector component:
<span class="notranslate">
$$
\frac{\mathrm{d}\overrightarrow{\mathbf{r}}}{\mathrm{d}t} =
\large
\begin{bmatrix}
\frac{\mathrm{d}x_P}{\mathrm{d}t} \\
\frac{\mathrm{d}y_P}{\mathrm{d}t} \\
\end{bmatrix}
$$
</span>
Note that the derivative is with respect to time but <span class="notranslate">$x_P$</span> and <span class="notranslate">$y_P$</span> depend explicitly on <span class="notranslate">$\theta$</span> and it's <span class="notranslate">$\theta$</span> that depends on <span class="notranslate">$t$ ($x_P$</span> and <span class="notranslate">$y_P$</span> depend implicitly on <span class="notranslate">$t$</span>). To calculate this type of derivative we will use the [chain rule](http://en.wikipedia.org/wiki/Chain_rule).
<br />
<div style="background-color:#FBFBEF;border:1px solid black;padding:10px;">
<b><a href="http://en.wikipedia.org/wiki/Chain_rule">Chain rule</a></b>
<br />
For variable <span class="notranslate">$f$</span> which is function of variable <span class="notranslate">$g$</span> which in turn is function of variable <span class="notranslate">$t$, $f(g(t))$</span> or <span class="notranslate">$(f\circ g)(t)$</span>, the derivative of <span class="notranslate">$f$</span> with respect to <span class="notranslate">$t$</span> is (using <a href="http://en.wikipedia.org/wiki/Notation_for_differentiation">Lagrange's notation</a>):
<br />
<span class="notranslate">
$$(f\circ g)^{'}(t) = f'(g(t)) \cdot g'(t)$$
</span>
Or using what is known as <a href="http://en.wikipedia.org/wiki/Notation_for_differentiation">Leibniz's notation</a>:
<br />
<span class="notranslate">
$$\frac{\mathrm{d}f}{\mathrm{d}t} = \frac{\mathrm{d}f}{\mathrm{d}g} \cdot \frac{\mathrm{d}g}{\mathrm{d}t}$$
</span>
If <span class="notranslate">$f$</span> is function of two other variables which both are function of <span class="notranslate"> $t$, $ f(x(t),y(t))$</span>, the chain rule for this case is:
<br />
<span class="notranslate">
$$\frac{\mathrm{d}f}{\mathrm{d}t} = \frac{\partial f}{\partial x} \cdot \frac{\mathrm{d}x}{\mathrm{d}t} + \frac{\partial f}{\partial y} \cdot \frac{\mathrm{d}y}{\mathrm{d}t}$$
</span>
Where <span class="notranslate">$df/dt$</span> represents the <a href="http://en.wikipedia.org/wiki/Total_derivative">total derivative</a> and <span class="notranslate">$\partial f / \partial x$</span> represents the <a href="http://en.wikipedia.org/wiki/Partial_derivative">partial derivative</a> of a function.
<br />
<b><a href="http://en.wikipedia.org/wiki/Product_rule">Product rule</a></b>
<br />
The derivative of the product of two functions is:
<br />
<span class="notranslate">
$$ (f \cdot g)' = f' \cdot g + f \cdot g' $$
</span>
</div>
### Linear velocity of the endpoint
For the planar one-link case, the linear velocity of the endpoint is:
```
v = r.diff(t)
v
```
Where we used the [Newton's notation](http://en.wikipedia.org/wiki/Notation_for_differentiation) for differentiation.
Note that <span class="notranslate">$\dot{\theta}$</span> represents the unknown angular velocity of the joint; this is why the derivative of <span class="notranslate">$\theta$</span> is not explicitly solved.
The magnitude or [Euclidian norm](http://en.wikipedia.org/wiki/Vector_norm) of the vector <span class="notranslate">$\overrightarrow{\mathbf{v}}$</span> is:
<span class="notranslate">
$$ ||\overrightarrow{\mathbf{v}}||=\sqrt{v_x^2+v_y^2} $$
</span>
```
simplify(sqrt(v[0]**2 + v[1]**2))
```
Which is <span class="notranslate">$\ell\dot{\theta}$</span>.<br>
We could have used the function `norm` of Sympy, but the output does not simplify nicely:
```
simplify(v.norm())
```
The direction of <span class="notranslate">$\overrightarrow{\mathbf{v}}$</span> is tangent to the circular trajectory of the endpoint as can be seen in the figure below where its components are also shown.
<figure><img src="./../images/onelink_vel.gif" width=350 alt="onelinkVel"/><figcaption><center><i>Figure. Endpoint velocity of one link attached to a fixed body by a hinge joint in a plane.</i></center></figcaption></figure>
### Linear acceleration of the endpoint
The acceleration of the endpoint position can be given by the second-order derivative of the position or by the first-order derivative of the velocity.
Using the chain and product rules for differentiation, the linear acceleration of the endpoint is:
```
acc = v.diff(t, 1)
acc
```
Examining the terms of the expression for the linear acceleration, we see there are two types of them: the term (in each direction) proportional to the angular acceleration <span class="notranslate">$\ddot{\theta}$</span> and other term proportional to the square of the angular velocity <span class="notranslate">$\dot{\theta}^{2}$</span>.
#### Tangential acceleration
The term proportional to angular acceleration, <span class="notranslate">$a_t$</span>, is always tangent to the trajectory of the endpoint (see figure below) and it's magnitude or Euclidean norm is:
```
A = θ.diff(t, 2)
simplify(sqrt(expand(acc[0]).coeff(A)**2 + expand(acc[1]).coeff(A)**2))*A
```
#### Centripetal acceleration
The term proportional to angular velocity, <span class="notranslate">$a_c$</span>, always points to the joint, the center of the circular motion (see figure below), because of that this term is termed [centripetal acceleration](http://en.wikipedia.org/wiki/Centripetal_acceleration#Tangential_and_centripetal_acceleration). Its magnitude is:
```
A = θ.diff(t)**2
simplify(sqrt(expand(acc[0]).coeff(A)**2+expand(acc[1]).coeff(A)**2))*A
```
This means that there will be a linear acceleration even if the angular acceleration is zero because although the magnitude of the linear velocity is constant in this case, its direction varies (due to the centripetal acceleration).
<br>
<figure><img src="./../images/onelink_acc.gif" width=350 alt="onelinkAcc"/><figcaption><center><i>Figure. Endpoint tangential and centripetal acceleration terms of one link attached to a fixed body by a hinge joint in a plane.</i></center></figcaption> </figure>
Let's plot some simulated data to have an idea of the one-link kinematics.
Consider <span class="notranslate"> $\ell=1\:m,\theta_i=0^o,\theta_f=90^o $</span>, and <span class="notranslate"> $1\:s$</span> of movement duration, and that it is a [minimum-jerk movement](https://nbviewer.jupyter.org/github/BMClab/bmc/blob/master/notebooks/MinimumJerkHypothesis.ipynb).
```
θ_i, θ_f, d = 0, np.pi/2, 1
ts = np.arange(0.01, 1.01, .01)
mjt = θ_i + (θ_f - θ_i)*(10*(t/d)**3 - 15*(t/d)**4 + 6*(t/d)**5)
ang = lambdify(t, mjt, 'numpy'); ang = ang(ts)
vang = lambdify(t, mjt.diff(t,1), 'numpy'); vang = vang(ts)
aang = lambdify(t, mjt.diff(t,2), 'numpy'); aang = aang(ts)
jang = lambdify(t, mjt.diff(t,3), 'numpy'); jang = jang(ts)
b, c, d, e = symbols('b c d e')
dicti = {l:1, θ:b, θ.diff(t, 1):c, θ.diff(t, 2):d, θ.diff(t, 3):e}
r2 = r.subs(dicti);
rxfu = lambdify(b, r2[0], modules = 'numpy')
ryfu = lambdify(b, r2[1], modules = 'numpy')
v2 = v.subs(dicti);
vxfu = lambdify((b, c), v2[0], modules = 'numpy')
vyfu = lambdify((b, c), v2[1], modules = 'numpy')
acc2 = acc.subs(dicti);
axfu = lambdify((b, c, d), acc2[0], modules = 'numpy')
ayfu = lambdify((b, c, d), acc2[1], modules = 'numpy')
jerk = r.diff(t,3)
jerk2 = jerk.subs(dicti);
jxfu = lambdify((b, c, d, e), jerk2[0], modules = 'numpy')
jyfu = lambdify((b, c, d, e), jerk2[1], modules = 'numpy')
fig, hax = plt.subplots(2, 4, sharex = True, figsize=(14, 7))
hax[0, 0].plot(ts, ang*180/np.pi, linewidth=3)
hax[0, 0].set_title('Angular displacement [ $^o$]');
hax[0, 0].set_ylabel('Joint')
hax[0, 1].plot(ts, vang*180/np.pi, linewidth=3)
hax[0, 1].set_title('Angular velocity [ $^o/s$]');
hax[0, 2].plot(ts, aang*180/np.pi, linewidth=3)
hax[0, 2].set_title('Angular acceleration [ $^o/s^2$]');
hax[0, 3].plot(ts, jang*180/np.pi, linewidth=3)
hax[0, 3].set_title('Angular jerk [ $^o/s^3$]');
hax[1, 0].plot(ts, rxfu(ang), 'r', linewidth=3, label = 'x')
hax[1, 0].plot(ts, ryfu(ang), 'k', linewidth=3, label = 'y')
hax[1, 0].set_title('Linear displacement [$m$]');
hax[1, 0].legend(loc='best').get_frame().set_alpha(0.8)
hax[1, 0].set_ylabel('Endpoint')
hax[1, 1].plot(ts,vxfu(ang, vang), 'r', linewidth=3)
hax[1, 1].plot(ts,vyfu(ang, vang), 'k', linewidth=3)
hax[1, 1].set_title('Linear velocity [$m/s$]');
hax[1, 2].plot(ts,axfu(ang, vang, aang), 'r', linewidth=3)
hax[1, 2].plot(ts,ayfu(ang, vang, aang), 'k', linewidth=3)
hax[1, 2].set_title('Linear acceleration [$m/s^2$]');
hax[1, 3].plot(ts, jxfu(ang, vang, aang, jang), 'r', linewidth=3)
hax[1, 3].plot(ts, jyfu(ang, vang, aang, jang), 'k', linewidth=3)
hax[1, 3].set_title('Linear jerk [$m/s^3$]');
fig.suptitle('Minimum jerk trajectory kinematics of one-link system', fontsize=20);
for i, hax2 in enumerate(hax.flat):
hax2.locator_params(nbins=5)
hax2.grid(True)
if i > 3:
hax2.set_xlabel('Time [s]');
plt.subplots_adjust(hspace=0.2, wspace=.3) #plt.tight_layout()
```
### Jacobian matrix
<br>
<div style="background-color:#FBFBEF;border:1px solid black;padding:10px;">
The <b><a href="https://en.wikipedia.org/wiki/Jacobian_matrix_and_determinant">Jacobian matrix</a></b> is the matrix of all first-order partial derivatives of a vector-valued function <span class="notranslate">$F$</span>:
<br>
<span class="notranslate">
$$
F(q_1,...q_n) = \begin{bmatrix}F_{1}(q_1,...q_n)\\
\vdots\\
F_{m}(q_1,...q_n)\\
\end{bmatrix}
$$
</span>
In a general form, the Jacobian matrix of the function <span class="notranslate">$F$</span> is:
<br>
<span class="notranslate">
$$
\mathbf{J}=
\large
\begin{bmatrix}
\frac{\partial F_{1}}{\partial q_{1}} & ... & \frac{\partial F_{1}}{\partial q_{n}} \\
\vdots & \ddots & \vdots \\
\frac{\partial F_{m}}{\partial q_{1}} & ... & \frac{\partial F_{m}}{\partial q_{n}} \\
\end{bmatrix}
$$
</span>
</div>
### Derivative of a vector-valued function using the Jacobian matrix
<br>
<div style="background-color:#FBFBEF;border:1px solid black;padding:10px;">
The time-derivative of a vector-valued function <span class="notranslate">$F$</span> can be computed using the Jacobian matrix:
<br>
<span class="notranslate">
$$
\frac{dF}{dt} = \mathbf{J} \cdot \begin{bmatrix}\frac{d q_1}{dt}\\
\vdots\\
\frac{d q_n}{dt}\\
\end{bmatrix}
$$
</span>
</div>
### Jacobian matrix in the context of kinematic chains
In the context of kinematic chains, the Jacobian is a matrix of all first-order partial derivatives of the linear position vector of the endpoint with respect to the angular position vector. The Jacobian matrix for a kinematic chain relates differential changes in the joint angle vector with the resulting differential changes in the linear position vector of the endpoint.
For a kinematic chain, the function <span class="notranslate">$F_{i}$</span> is the expression of the endpoint position in <span class="notranslate">$m$</span> coordinates and the variable <span class="notranslate">$q_{i}$</span> is the angle of each <span class="notranslate">$n$</span> joints.
#### Jacobian matrix of one-link chain
For the planar one-link case, the Jacobian matrix of the position vector of the endpoint <span class="notranslate">$r_P$</span> with respect to the angular position vector <span class="notranslate">$q_1=\theta$</span> is:
<br>
<span class="notranslate">
\begin{equation}
\mathbf{J}=
\large
\begin{bmatrix}
\frac{\partial x_P}{\partial \theta} \\
\frac{\partial y_P}{\partial \theta} \\
\end{bmatrix}
\end{equation}
</span>
Which evaluates to:
```
J = r.diff(θ)
J
```
And Sympy has a function to calculate the Jacobian:
```
J = r.jacobian([θ])
J
```
We can recalculate the kinematic expressions using the Jacobian matrix, which can be useful for simplifying the deduction.
The linear velocity of the end-effector is given by the product between the Jacobian of the kinematic link and the angular velocity:
<br>
<span class="notranslate">
$$ \overrightarrow{\mathbf{v}} = \mathbf{J} \cdot \dot{\theta}$$
</span>
Where:
```
ω = θ.diff(t)
ω
```
The angular velocity is also a vector; it's direction is perpendicular to the plane of rotation and using the [right-hand rule](http://en.wikipedia.org/wiki/Right-hand_rule) this direction is the same as of the versor <span class="notranslate">$\hat{\mathbf{k}}$</span> coming out of the screen (paper).
Then:
```
velJ = J*ω
velJ
```
And the linear acceleration of the endpoint is given by the derivative of this product:
<span class="notranslate">
$$ \overrightarrow{\mathbf{a}} = \dot{\mathbf{J}} \cdot \overrightarrow{\mathbf{\omega}} + \mathbf{J} \cdot \dot{\overrightarrow{\mathbf{\omega}}} $$
</span>
Let's calculate this derivative:
```
accJ = J.diff(t)*ω + J*ω.diff(t)
accJ
```
These two expressions derived with the Jacobian are the same as the direct derivatives of the equation for the endpoint position.
## The kinematics of a two-link chain
We now will look at the case of a planar kinematic chain with two links, as shown below. The deduction will be similar to the case with one link we just saw.
<br>
<figure><img src="./../images/twolinks.gif" width=400 alt="twolinks"/><figcaption><center><i>Figure. Kinematics of a two-link chain with hinge joints in a plane.</i></center></figcaption> </figure>
We need to define a Cartesian coordinate system and the symbolic variables <span class="notranslate">$t,\:\ell_1,\:\ell_2,\:\theta_1,\:\theta_2$</span> (and make <span class="notranslate">$\theta_1$</span> and <span class="notranslate">$\theta_2$</span> function of time):
```
G = CoordSys3D('')
t = Symbol('t')
l1, l2 = symbols('ell_1 ell_2', positive=True)
θ1, θ2 = dynamicsymbols('theta1 theta2')
```
The position of the endpoint in terms of the joint angles and link lengths is:
```
r2_p = (l1*cos(θ1) + l2*cos(θ1 + θ2))*G.i + (l1*sin(θ1) + l2*sin(θ1 + θ2))*G.j
r2_p
```
With the components:
```
r2_p.components
```
And in matrix form:
```
r2 = Matrix((r2_p.dot(G.i), r2_p.dot(G.j)))
r2
```
### Joint and segment angles
Note that <span class="notranslate"> $\theta_2$</span> is a joint angle (referred as measured in the joint space); the angle of the segment 2 with respect to the horizontal is <span class="notranslate"> $\theta_1+\theta_2$</span> and is referred as an angle in the segmental space.
Joint and segment angles are also referred as relative and absolute angles, respectively.
### Inverse kinematics
Using the [cosine rule](http://en.wikipedia.org/wiki/Law_of_cosines), in terms of the endpoint position, the angle <span class="notranslate">$\theta_2$</span> is:
<br>
<span class="notranslate">
\begin{equation}
x_P^2 + y_P^2 = \ell_1^2+\ell_2^2 - 2\ell_1 \ell_2 cos(\pi-\theta_2)
\end{equation}
</span>
<span class="notranslate">
\begin{equation}
\theta_2 = \arccos\left(\frac{x_P^2 + y_P^2 - \ell_1^2 - \ell_2^2}{2\ell_1 \ell_2}\;\;\right)
\end{equation}
</span>
To find the angle <span class="notranslate">$\theta_1$</span>, if we now look at the triangle in red in the figure below, its angle <span class="notranslate">$\phi$</span> is:
<br>
<span class="notranslate">
\begin{equation}
\phi = \arctan\left(\frac{\ell_2 \sin(\theta_2)}{\ell_1 + \ell_2 \cos(\theta_2)}\right)
\end{equation}
</span>
And the angle of its hypotenuse with the horizontal is:
<br>
<span class="notranslate">
\begin{equation}
\theta_1 + \phi = \arctan\left(\frac{y_P}{x_P}\right)
\end{equation}
</span>
Then, the angle <span class="notranslate">$\theta_1$</span> is:
<br>
<span class="notranslate">
\begin{equation}
\theta_1 = \arctan\left(\frac{y_P}{x_P}\right) - \arctan\left(\frac{\ell_2 \sin(\theta_2)}{\ell_1+\ell_2 \cos(\theta_2)}\right)
\end{equation}
</span>
Note that there are two possible sets of <span class="notranslate">$(\theta_1, \theta_2)$</span> angles for the same <span class="notranslate">$(x_P, y_P)$</span> coordinate that satisfy the equations above. The figure below shows in orange another possible configuration of the kinematic chain with the same endpoint coordinate. The other solution is <span class="notranslate">$\theta_2'=2\pi - \theta_2$</span>, but <span class="notranslate">$\sin(\theta_2')=-sin(\theta_{2})$</span> and then the <span class="notranslate">$arctan()$</span> term in the last equation becomes negative.
Even for a simple two-link chain we already have a problem of redundancy, there is more than one joint configuration for the same endpoint position; this will be much more problematic for chains with more links (more degrees of freedom).
<br>
<figure><img src="./../images/twolinks_ik.gif" width=350 alt="twolinks_ik"/><figcaption><center><i>Figure. Indetermination in the inverse kinematics approach to determine one of the joint angles for a two-link chain with hinge joints in a plane.</i></center></figcaption> </figure>
## Differential kinematics
The linear velocity of the endpoint is:
```
vel2 = r2.diff(t)
vel2
```
The linear velocity of the endpoint is the sum of the velocities at each joint, i.e., it is the velocity of the endpoint in relation to joint 2, for instance, <span class="notranslate">
$\ell_2cos(\theta_1 + \theta_2)\dot{\theta}_1$</span>, plus the velocity of joint 2 in relation to joint 1, for instance, <span class="notranslate">$\ell_1\dot{\theta}_1 cos(\theta_1)$</span>, and this last term we already saw for the one-link example. In classical mechanics this is known as [relative velocity](http://en.wikipedia.org/wiki/Relative_velocity), an example of [Galilean transformation](http://en.wikipedia.org/wiki/Galilean_transformation).
The linear acceleration of the endpoint is:
```
acc2 = r2.diff(t, 2)
acc2
```
We can separate the equation above for the linear acceleration in three types of terms: proportional to <span class="notranslate">$\ddot{\theta}$</span> and to <span class="notranslate">$\dot{\theta}^2$</span>, as we already saw for the one-link case, and a new term, proportional to <span class="notranslate">$\dot{\theta}_1\dot{\theta}_2$</span>:
```
acc2 = acc2.expand()
A = θ1.diff(t, 2)
B = θ2.diff(t, 2)
tg = A*Matrix((acc2[0].coeff(A),acc2[1].coeff(A)))+B*Matrix((acc2[0].coeff(B),acc2[1].coeff(B)))
A = θ1.diff(t)**2
B = θ2.diff(t)**2
ct = A*Matrix((acc2[0].coeff(A),acc2[1].coeff(A)))+B*Matrix((acc2[0].coeff(B),acc2[1].coeff(B)))
A = θ1.diff(t)*θ2.diff(t)
co = A*Matrix((acc2[0].coeff(A),acc2[1].coeff(A)))
display(Math(mlatex(r'Tangential:\:') + mlatex(tg)))
display(Math(mlatex(r'Centripetal:') + mlatex(ct)))
display(Math(mlatex(r'Coriolis:\;\;\;\;\:') + mlatex(co)))
```
This new term is called the [Coriolis acceleration](http://en.wikipedia.org/wiki/Coriolis_effect); it is 'felt' by the endpoint when its distance to the instantaneous center of rotation varies, due to the links' constraints, and as consequence the endpoint motion is deflected (its direction is perpendicular to the relative linear velocity of the endpoint with respect to the linear velocity at the second joint, <span class="notranslate">$\mathbf{v} - \mathbf{v}_{joint2}$</span>.
Let's now deduce the Jacobian for this planar two-link chain:
<br>
<span class="notranslate">
$$
\mathbf{J} =
\large
\begin{bmatrix}
\frac{\partial x_P}{\partial \theta_{1}} & \frac{\partial x_P}{\partial \theta_{2}} \\
\frac{\partial y_P}{\partial \theta_{1}} & \frac{\partial y_P}{\partial \theta_{2}} \\
\end{bmatrix}
$$
</span>
We could manually run:
```python
J = Matrix([[r2[0].diff(theta1), r2[0].diff(theta2)], [r2[1].diff(theta1), r2[1].diff(theta2)]])
```
But it's shorter with the Jacobian function from Sympy:
```
J2 = r2.jacobian([θ1, θ2])
J2
```
Using the Jacobian, the linear velocity of the endpoint is:
<br>
<span class="notranslate">
$$ \mathbf{v_J} = \mathbf{J} \cdot \begin{bmatrix}\dot{\theta_1}\\
\dot{\theta_2}\\
\end{bmatrix} $$
</span>
Where:
```
ω2 = Matrix((θ1, θ2)).diff(t)
ω2
```
Then:
```
vel2J = J2*ω2
vel2J
```
This expression derived with the Jacobian is the same as the first-order derivative of the equation for the endpoint position. We can show this equality by comparing the two expressions with Sympy:
```
vel2.expand() == vel2J.expand()
```
Once again, the linear acceleration of the endpoint is given by the derivative of the product between the Jacobian and the angular velocity:
<br>
<span class="notranslate">
\begin{equation}
\mathbf{a} = \dot{\mathbf{J}} \cdot \mathbf{\omega} + \mathbf{J} \cdot \dot{\mathbf{\omega}}
\end{equation}
</span>
Let's calculate this derivative:
```
acc2J = J2.diff(t)*ω2 + J2*ω2.diff(t)
acc2J
```
Once again, the expression above is the same as the second-order derivative of the equation for the endpoint position:
```
acc2.expand() == acc2J.expand()
```
Let's plot some simulated data to have an idea of the two-link kinematics.
Consider 1 s of movement duration, <span class="notranslate">$\ell_1=\ell_2=0.5m, \theta_1(0)=\theta_2(0)=0$</span>, <span class="notranslate">$\theta_1(1)=\theta_2(1)=90^o$</span>, and that the endpoint trajectory is a [minimum-jerk movement](https://nbviewer.jupyter.org/github/BMClab/bmc/blob/master/notebooks/MinimumJerkHypothesis.ipynb).
First, the simulated trajectories:
```
t, p0, pf, d = symbols('t p0 pf d')
rx = dynamicsymbols('rx', real=True) # or Function('rx')(t)
ry = dynamicsymbols('ry', real=True) # or Function('ry')(t)
# minimum jerk kinematics
mjt = p0 + (pf - p0)*(10*(t/d)**3 - 15*(t/d)**4 + 6*(t/d)**5)
rfu = lambdify((t, p0, pf, d), mjt, 'numpy')
vfu = lambdify((t, p0, pf, d), diff(mjt, t, 1), 'numpy')
afu = lambdify((t, p0, pf, d), diff(mjt, t, 2), 'numpy')
jfu = lambdify((t, p0, pf, d), diff(mjt, t, 3), 'numpy')
# values
d, L1, L2 = 1, .5, .5
#initial values:
p0, pf = [-0.5, 0.5], [0, .5*np.sqrt(2)]
ts = np.arange(0.01, 1.01, .01)
# endpoint kinematics
x = rfu(ts, p0[0], pf[0], d)
y = rfu(ts, p0[1], pf[1], d)
vx = vfu(ts, p0[0], pf[0], d)
vy = vfu(ts, p0[1], pf[1], d)
ax = afu(ts, p0[0], pf[0], d)
ay = afu(ts, p0[1], pf[1], d)
jx = jfu(ts, p0[0], pf[0], d)
jy = jfu(ts, p0[1], pf[1], d)
# inverse kinematics
ang2b = np.arccos((x**2 + y**2 - L1**2 - L2**2)/(2*L1*L2))
ang1b = np.arctan2(y, x) - (np.arctan2(L2*np.sin(ang2b), (L1+L2*np.cos(ang2b))))
ang2 = acos((rx**2 + ry**2 - l1**2 - l2**2)/(2*l1*l2))
ang2fu = lambdify((rx ,ry, l1, l2), ang2, 'numpy');
ang2 = ang2fu(x, y, L1, L2)
ang1 = atan2(ry, rx) - (atan(l2*sin(acos((rx**2 + ry**2 - l1**2 - l2**2)/(2*l1*l2)))/ \
(l1+l2*cos(acos((rx**2 + ry**2 - l1**2 - l2**2)/(2*l1*l2))))))
ang1fu = lambdify((rx, ry, l1, l2), ang1, 'numpy');
ang1 = ang1fu(x, y, L1, L2)
ang2b = acos((rx**2 + ry**2 - l1**2 - l2**2)/(2*l1*l2))
ang1b = atan2(ry, rx) - (atan(l2*sin(acos((rx**2 + ry**2 - l1**2 - l2**2)/(2*l1*l2)))/ \
(l1 + l2*cos(acos((rx**2 + ry**2-l1**2 - l2**2)/(2*l1*l2))))))
X, Y, Xd, Yd, Xdd, Ydd, Xddd, Yddd = symbols('X Y Xd Yd Xdd Ydd Xddd Yddd')
dicti = {rx:X, ry:Y, rx.diff(t, 1):Xd, ry.diff(t, 1):Yd, \
rx.diff(t, 2):Xdd, ry.diff(t, 2):Ydd, rx.diff(t, 3):Xddd, ry.diff(t, 3):Yddd, l1:L1, l2:L2}
vang1 = diff(ang1b, t, 1)
vang1 = vang1.subs(dicti)
vang1fu = lambdify((X, Y, Xd, Yd, l1, l2), vang1, 'numpy')
vang1 = vang1fu(x, y, vx, vy, L1, L2)
vang2 = diff(ang2b, t, 1)
vang2 = vang2.subs(dicti)
vang2fu = lambdify((X, Y, Xd, Yd, l1, l2), vang2, 'numpy')
vang2 = vang2fu(x, y, vx, vy, L1, L2)
aang1 = diff(ang1b, t, 2)
aang1 = aang1.subs(dicti)
aang1fu = lambdify((X, Y, Xd, Yd, Xdd, Ydd, l1, l2), aang1, 'numpy')
aang1 = aang1fu(x, y, vx, vy, ax, ay, L1, L2)
aang2 = diff(ang2b, t, 2)
aang2 = aang2.subs(dicti)
aang2fu = lambdify((X, Y, Xd, Yd, Xdd, Ydd, l1, l2), aang2, 'numpy')
aang2 = aang2fu(x, y, vx, vy, ax, ay, L1, L2)
jang1 = diff(ang1b, t, 3)
jang1 = jang1.subs(dicti)
jang1fu = lambdify((X, Y, Xd, Yd, Xdd, Ydd, Xddd, Yddd, l1, l2), jang1, 'numpy')
jang1 = jang1fu(x, y, vx, vy, ax, ay, jx, jy, L1, L2)
jang2 = diff(ang2b, t, 3)
jang2 = jang2.subs(dicti)
jang2fu = lambdify((X, Y, Xd, Yd, Xdd, Ydd, Xddd, Yddd, l1, l2), jang2, 'numpy')
jang2 = jang2fu(x, y, vx, vy, ax, ay, jx, jy, L1, L2)
```
And the plots for the trajectories:
```
fig, hax = plt.subplots(2, 4, sharex = True, figsize=(14, 7))
hax[0, 0].plot(ts, x, 'r', linewidth=3, label = 'x')
hax[0, 0].plot(ts, y, 'k', linewidth=3, label = 'y')
hax[0, 0].set_title('Linear displacement [$m$]')
hax[0, 0].legend(loc='best').get_frame().set_alpha(0.8)
hax[0, 0].set_ylabel('Endpoint')
hax[0, 1].plot(ts, vx, 'r', linewidth=3)
hax[0, 1].plot(ts, vy, 'k', linewidth=3)
hax[0, 1].set_title('Linear velocity [$m/s$]')
hax[0, 2].plot(ts, ax, 'r', linewidth=3)
hax[0, 2].plot(ts, ay, 'k', linewidth=3)
hax[0, 2].set_title('Linear acceleration [$m/s^2$]')
hax[0, 3].plot(ts, jx, 'r', linewidth=3)
hax[0, 3].plot(ts, jy, 'k', linewidth=3)
hax[0, 3].set_title('Linear jerk [$m/s^3$]')
hax[1, 0].plot(ts, ang1*180/np.pi, 'b', linewidth=3, label = 'Ang1')
hax[1, 0].plot(ts, ang2*180/np.pi, 'g', linewidth=3, label = 'Ang2')
hax[1, 0].set_title('Angular displacement [ $^o$]')
hax[1, 0].legend(loc='best').get_frame().set_alpha(0.8)
hax[1, 0].set_ylabel('Joint')
hax[1, 1].plot(ts, vang1*180/np.pi, 'b', linewidth=3)
hax[1, 1].plot(ts, vang2*180/np.pi, 'g', linewidth=3)
hax[1, 1].set_title('Angular velocity [ $^o/s$]')
hax[1, 2].plot(ts, aang1*180/np.pi, 'b', linewidth=3)
hax[1, 2].plot(ts, aang2*180/np.pi, 'g', linewidth=3)
hax[1, 2].set_title('Angular acceleration [ $^o/s^2$]')
hax[1, 3].plot(ts, jang1*180/np.pi, 'b', linewidth=3)
hax[1, 3].plot(ts, jang2*180/np.pi, 'g', linewidth=3)
hax[1, 3].set_title('Angular jerk [ $^o/s^3$]')
tit = fig.suptitle('Minimum jerk trajectory kinematics of a two-link chain', fontsize=20)
for i, hax2 in enumerate(hax.flat):
hax2.locator_params(nbins=5)
hax2.grid(True)
if i > 3:
hax2.set_xlabel('Time [$s$]')
plt.subplots_adjust(hspace=0.15, wspace=0.25) #plt.tight_layout()
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(14, 4))
ax1.plot(x, y, 'r', linewidth=3)
ax1.set_xlabel('Displacement in x [$m$]')
ax1.set_ylabel('Displacement in y [$m$]')
ax1.set_title('Endpoint space', fontsize=14)
ax1.axis('equal')
ax1.grid(True)
ax2.plot(ang1*180/np.pi, ang2*180/np.pi, 'b', linewidth=3)
ax2.set_xlabel('Displacement in joint 1 [ $^o$]')
ax2.set_ylabel('Displacement in joint 2 [ $^o$]')
ax2.set_title('Joint sapace', fontsize=14)
ax2.axis('equal')
ax2.grid(True)
```
## Further reading
- Read pages 477-494 of the 10th chapter of the [Ruina and Rudra's book](http://ruina.tam.cornell.edu/Book/index.html) for a review of differential equations and kinematics.
## Video lectures on the Internet
- Khan Academy: [Differential Calculus Review](https://khanacademy.org/math/differential-calculus)
- Khan Academy: [Chain Rule Review](https://khanacademy.org/math/differential-calculus/dc-chain)
- [Multivariate Calculus – Jacobian applied](https://www.youtube.com/watch?v=57q-2YxIZss)
## Problems
1. For the numerical example of the two-link chain plotted above, calculate and plot the values for the each type of acceleration (tangential, centripetal and Coriolis). See solution below.
2. For the two-link chain, calculate and interpret the Jacobian and the expressions for the position, velocity, and acceleration of the endpoint for the following cases:
a) When the first joint (the joint at the base) is fixed at <span class="notranslate"> $0^o$</span>.
b) When the second joint is fixed at <span class="notranslate">$0^o$</span>.
3. For the two-link chain, a special case of movement occurs when the endpoint moves along a line passing through the first joint (the joint at the base). A system with this behavior is known as a polar manipulator (Mussa-Ivaldi, 1986). For simplicity, consider that the lengths of the two links are equal to $\ell$. In this case, the two joint angles are related by: <span class="notranslate">$2\theta_1+\theta_2=\pi$</span>.
a) Calculate the Jacobian for this polar manipulator and compare it with the Jacobian for the standard two-link chain. Note the difference between the off-diagonal terms.
b) Calculate the expressions for the endpoint position, velocity, and acceleration.
c) For the endpoint acceleration of the polar manipulator, identify the tangential, centrifugal, and Coriolis components and compare them with the expressions for the standard two-link chain.
4. Deduce the equations for the kinematics of a two-link pendulum with the angles in relation to the vertical.
5. Deduce the equations for the kinematics of a two-link system using segment angles and compare with the deduction employing joint angles.
6. Calculate the Jacobian matrix for the following function:
<span class="notranslate">
\begin{equation}
f(x, y) = \begin{bmatrix}
x^2 y \\
5 x + \sin y \end{bmatrix}
\end{equation}
</span>
#### Calculation of each type of acceleration of the endpoint for the numerical example of the two-link system
```
# tangential acceleration
A1, A2, A1d, A2d, A1dd, A2dd = symbols('A1 A2 A1d A2d A1dd A2dd')
dicti = {θ1:A1, θ2:A2, θ1.diff(t, 1):A1d, θ2.diff(t,1):A2d, \
θ1.diff(t, 2):A1dd, θ2.diff(t, 2):A2dd, l1:L1, l2:L2}
tg2 = tg.subs(dicti)
tg2fu = lambdify((A1, A2, A1dd, A2dd), tg2, 'numpy');
tg2n = tg2fu(ang1, ang2, aang1, aang2)
tg2n = tg2n.reshape((2, 100)).T
# centripetal acceleration
ct2 = ct.subs(dicti)
ct2fu = lambdify((A1, A2, A1d, A2d), ct2, 'numpy');
ct2n = ct2fu(ang1, ang2, vang1, vang2)
ct2n = ct2n.reshape((2, 100)).T
# coriolis acceleration
co2 = co.subs(dicti)
co2fu = lambdify((A1, A2, A1d, A2d), co2, 'numpy');
co2n = co2fu(ang1, ang2, vang1, vang2)
co2n = co2n.reshape((2, 100)).T
# total acceleration (it has to be the same calculated before)
acc_tot = tg2n + ct2n + co2n
```
#### And the corresponding plots
```
fig, hax = plt.subplots(1, 3, sharex = True, sharey = True, figsize=(12, 5))
hax[0].plot(ts, acc_tot[:, 0], color=(1, 0, 0, .3), linewidth=5, label = 'x total')
hax[0].plot(ts, acc_tot[:, 1], color=(0, 0, 0, .3), linewidth=5, label = 'y total')
hax[0].plot(ts, tg2n[:, 0], 'r', linewidth=2, label = 'x')
hax[0].plot(ts, tg2n[:, 1], 'k', linewidth=2, label = 'y')
hax[0].set_title('Tangential')
hax[0].set_ylabel('Endpoint acceleration [$m/s^2$]')
hax[0].set_xlabel('Time [$s$]')
hax[1].plot(ts, acc_tot[:, 0], color=(1,0,0,.3), linewidth=5, label = 'x total')
hax[1].plot(ts, acc_tot[:, 1], color=(0,0,0,.3), linewidth=5, label = 'y total')
hax[1].plot(ts, ct2n[:, 0], 'r', linewidth=2, label = 'x')
hax[1].plot(ts, ct2n[:, 1], 'k', linewidth=2, label = 'y')
hax[1].set_title('Centripetal')
hax[1].set_xlabel('Time [$s$]')
hax[1].legend(loc='best').get_frame().set_alpha(0.8)
hax[2].plot(ts, acc_tot[:, 0], color=(1,0,0,.3), linewidth=5, label = 'x total')
hax[2].plot(ts, acc_tot[:, 1], color=(0,0,0,.3), linewidth=5, label = 'y total')
hax[2].plot(ts, co2n[:, 0], 'r', linewidth=2, label = 'x')
hax[2].plot(ts, co2n[:, 1], 'k', linewidth=2, label = 'y')
hax[2].set_title('Coriolis')
hax[2].set_xlabel('Time [$s$]')
tit = fig.suptitle('Acceleration terms for the minimum jerk trajectory of a two-link chain',
fontsize=16)
for hax2 in hax:
hax2.locator_params(nbins=5)
hax2.grid(True)
# plt.subplots_adjust(hspace=0.15, wspace=0.25) #plt.tight_layout()
```
## References
- Mussa-Ivaldi FA (1986) Compliance. In: Morasso P Tagliasco V (eds), [Human Movement Understanding: from computational geometry to artificial Intelligence](http://books.google.com.br/books?id=ZlZyLKNoAtEC). North-Holland, Amsterdam.
- Ruina A, Rudra P (2019) [Introduction to Statics and Dynamics](http://ruina.tam.cornell.edu/Book/index.html). Oxford University Press.
- Siciliano B et al. (2009) [Robotics - Modelling, Planning and Control](http://books.google.com.br/books/about/Robotics.html?hl=pt-BR&id=jPCAFmE-logC). Springer-Verlag London.
- Zatsiorsky VM (1998) [Kinematics of Human Motions](http://books.google.com.br/books/about/Kinematics_of_Human_Motion.html?id=Pql_xXdbrMcC&redir_esc=y). Champaign, Human Kinetics.
| github_jupyter |
# 응용통계학 (11주차) 5월 12일
> GLM, 일반화선형모형
- toc:true
- branch: master
- badges: true
- comments: false
- author: 최서연
- categories: [Applied Statistics, GLM, 일반화 선형 모형]
```
#hide
options(jupyter.plot_scale=4)
options(repr.plot.width=8,repr.plot.height=6,repr.plot.res=300)
#options(jupyter.rich_display=FALSE)
#options(max.print=1000)
```
다중공선성이 존재하는 상황을 가정하고
다중공선성을 어느 정도 제거한 모형 (M1)과 다중공선성이 내재되어 있는 모형 (M2) 을 고려하여
두 모형의 예측력을 모의실험을 통해 비교하여라,
단, 실험은 여러 번 반복하여 평균적인 결과를 report하되 설명변수의 개수는 3개 이상으로 설정하여라.
이미 존재하는 문서들을 참고하거나 재현해도 무방함.
(첨부된 문서 참고)
# 일반화선형모형 : Generalized linear model I
이 강의노트는 `Extending the linear model with R` (2016), 2nd edition, Julian J. Faraway, Chapman and Hall. 을 바탕으로 작성되었습니다.
## 1. Binary response
### 1.1 Heart Disease Example
샌프란시스코에 사는 39세에서 59세 사이의 건강한 성인남성 3154명에 대하여 8년 반 동안 관찰하여 심장관상동맥질환이 나타났는지 여부를 관찰하였다.
여기서 target 변수 `chd` ; coronary heat disease developed 는 factor변수로 취급할 수 있고 `no` `yes` 두 수준을 가진다. 다른 요인들이 질환여부와 어떻게 연관되는지를 알고자 한다면 target 변수가 연속이 아닌 이진(binary)형이므로 지금까지 다루었던 전형적인 선형모형은 적합하지 않다.
- no: 0, yes: 1
$$chd∼height+cigarette$$
- 담배는 하루에 몇 개피 피웠는지
- $y = \beta_0 + \beta_1 h + \beta_2 c + \epsilon$
- $\beta_0, \beta_1, \beta_2 \in R$
몇 가지 그림을 통해 자료를 살펴보자.
- 연속형에서는 산점도, 이진형에서는 박스플랏?!
```
data(wcgs, package="faraway")
summary(wcgs[,c("chd","height","cigs")])
plot(height ~ chd, wcgs)
```
- 키가 작으면 질병이 걸리는 건가? 명확하진 않다.
```
wcgs$y <- ifelse(wcgs$chd == "no",0,1)
plot(jitter(y,0.1) ~ jitter(height), wcgs, xlab="Height", ylab="Heart Disease", pch=".")
library(ggplot2)
ggplot(wcgs, aes(x=height, color=chd)) + geom_histogram(position="dodge", binwidth=1)
```
- no가 뚜렷하게 많이 분포해있는 모습
```
ggplot(wcgs, aes(x=cigs, color=chd)) + geom_histogram(position="dodge", binwidth=5, aes(y=..density..))
```
- yes가 높은 경향
```
ggplot(wcgs, aes(x=height,y=cigs))+geom_point(alpha=0.2, position=position_jitter())+facet_grid(~ chd)
```
키 혹은 흡연량이 질환여부와 연관이 있어 보이는가? 만약 그렇다면 어떻게 모형화 할 수 있겠는가?
### 1.2 Conditional mean regression
선형모형은 반응변수 $Y$를 설명변수 $x_1,…,x_p$의 선형겹합 $+ noise$ 로 표현한 것이다. 사실 `선형`이라는 의미는 오차항을 제외한 부분을 선형으로 모형화한다는 의미로 볼 수 있다.
- 설명변수를 linear로 모델링
$$E(Y|x_1,…,x_p)=β_0+β_1x_1+…+β_px_p$$
- $Y = f(X_1, \dots, X_p) + \epsilon$
- $E(\epsilon|x_1,\dots,x_p) = 0$
- $Y = 0 or 1$
- $E(Y) = p \times 1 + (1-p) \times 0 = p = P_r(Y=1)$
즉, 선형모형은 설명변수가 주어졌을 때 반응변수의 조건부 기대값(평균)을 설병변수들의 선형결합으로 모형화하고 있는 것이다. 반응변수 $Y$가 $Binary$인 경우 조건부 기대값은 조건부 확률과 같으므로,
$$E(Y|x_1,…,x_p)=P(Y=1|x_1,…,x_p)=β_0+β_1x_1+…+β_px_p$$
- R(실수 전체 가능)
과 같은 모형화가 가능하다.
하지만 위와 같은 모형은 모수추정에 따라 주어지는 조건부 확률의 추정치가 $proper range( [0,1] )$를 보장해주지 못한다. 이는 자료의 범위를 벗어나는 부분에서 특히 발생할 수 있는 문제이다.
- 음수예측이나 드문드문 있는 데이터로 불안정한 예측
이를 해결할 수 있는 간단한 해결책 중 하나는 $proper range$를 보장 할 수 있는 변환함수 $g$를 고려하는 것이다.
$$g(P(Y=1|x_1,…,x_p))=β_0+β_1x_1+…+β_px_p$$
- GLM
- $X^{\top} \beta$
반응변수가 $Binary$인 경우 이런 변환 함수는 다음과 같은 정의역과 공역을 가지는 것으로 선택한다.
$$g:(0,1)→R$$
변환함수는 다양하게 선택될 수 있으며, 반응변수의 특성에 따라 다른 변환함수를 고려하는 것이 자연스러울 것이다.
### 1.3 로지스틱 회귀 : Logistic regression
로지스틱 회귀모형은 $Binary response$인 경우 가장 흔하게 사용되는 모형이다.
$Y_i∈ \{ 0 ,1 \} , i=1,…,n$이고 $P(Y_i=1)=p_i∈(0,1)$이라 하자. 그리고 다음과 같은 모형을 생각하자.
$$η_i=g(p_i)=β_0+β_1x_{i1}+⋯+β_qx_{qi}$$
= $X_i^{\top} \beta$
여기서 $η_i$를 $linear predictor$라 한다. 이 모형에서는 설명변수의 선형결합이 $g$를 통해서 $p_i$와 연결된다. 이 때 변환함수 $g$를 연결함수($link function$)이라 하고 이런 형태의 모형화를 통칭하여 일반화선형모형 `Generalized linear model (GLM)`이라 부른다. 사실 앞서 배운 전통적인 선형모형은 $g(x)=x$인 GLM의 특수한 형태로 볼 수 있다.
앞서 언급했듯이 $g$에 대한 선택은 정답이 있는 것은 아니나 계산상의 이유, 추정량의 성질과 관련된 이유, 모형의 해석과 관련된 이유, 데이터와의 적합성과 관련된 이유 등으로 주로 쓰여지는 함수들이 존재한다. $Binary response$의 경우 가장 일반적으로 쓰이는 연결함수는 $logit$ 연결함수이며 다음과 같이 정의된다.
- link function 사용하기 때문에 추정량 성질과 관련될 수 있지
- 어떤 연결함수를 사용하는지에 따라 모형이 달라지낟.
- 더 좋은 적합도를 가진 link function을 찾아야 함.
$$g(t)=logit(t)=log\frac{t}{1−t}$$
- $h: R \to (0,1)$
- $lim_{t \to 0} g(t) = log 0 = - \infty$
- $lim_{t \to 1} g(t) = log \infty = \infty$
이 함수는 다음과 같이 정의되는 $logistic$ 함수의 역함수로 알려져 있다.
$$g^{−1}(t)=h(t)=\frac{1}{1+exp(−t)}=\frac{exp(t)}{1+exp(t)}$$
$logit$ 연결함수를 고려하는 $GLM$을 로지스틱 모형이라 부르고 다음과 같이 표현된다.
$$log\frac{p_i}{1−p_i}=β_0+β_1x_{i1}+⋯+β_qx_{qi}$$
- $p_i$ 자체의 움직임이 아니라 연결함수도 함께 고려해야한다. 달라질 수 있으니까
- $x$기 증가하면 $p$도 증가하고(positive), $x$가 감소하면 $p$도 감소할걸(negative)?
위에서 양변이 모두 $R$의 $range$를 가짐을 알 수 있다.
- $\hat{p}_i = P(Y_i=1|x_{i1}, \dots, x_{qi} ) = \frac{exp(\hat{η_i})}{1+exp(\hat{η_i})}$
- $1-\hat{p}_i = \frac{1}{1+exp(\hat{η_i})}$
```
library(faraway)
curve(ilogit(x),-6,6, xlab=expression(eta), ylab="p")
```
- linear로 하면 (0,1) 범위 벗어날 수 있다.
$logit$ 함수의 역함수 즉, $logistic$ 함수는 $S$자 형태의 곡선모양을 띠며 중심부분에서는 직선과 유사한 형태임을 알 수 있다.
모형이 정의되었으면 다음으로 모수의 추정량을 제시하여야 한다. 선형모형에서 사용하였던 최소제곱법의 원리를 차용할 수 있을까?
일반적으로 $GLM$에서는 최소제곱의 원리보다는 반응변수의 분포특성을 이용할 수 있는 최대가능도추정법을 사용한다.
$$l(β)=∑_{i=1}^{n}[y_iη_i−log(1+exp^{η_i})]$$
- $P(y_i = 1| x_{i1},\dots,x_{iq}) = p_i)$
- $y_{i|x_{i1},\dots,x_{iq}} \sim Bernoulli(p_i)$
- 확률질량함수$f(y_i) = p_i^{y_i}(1-p_i)^{1-y_i}, y_i = 0,1$
- $L = \Pi^{n}_{i=1} f(y_i) = \Pi^{n}_{i=1} p_{i}^{y_i}(1-p_i)^{1-y_i}$
- $l = \log L = \sum^{n}_{i=1} (y_i \log p_i + (1-y_i) \log(1-p_i))$
- $= \sum^{n}_{i=1}(y_i \log p_i - y_i \log (1-p_i) + \log (1-p_i))$
- $= \sum^{n}_{i=1} (y_i(log\frac{p_i}{1-p_i}) + \log (1-p_i))$
- $\log \frac{p_i}{1-p_i} = \beta_0 + \beta_1 x_{i1} + \dots + \beta_q x_{qi} = η_i = X_{i}^{\top}\beta$
- $\_i = \frac{exp(\hat{η_i})}{a+exp(\hat{η_i})}$
- $\sum^{n}_{i=1} [y_i η_i - \log (1+exp(\hat{η_i}))]$
- $argmax_{\beta = \beta_0,\dots, \beta_q} l(\beta) = MLE$
- $y_i = β_0+β_1x_{i1}+⋯+β_qx_{qi} + \epsilon$
- $argmin_{\beta_0,\dots,\beta_q}\sum(y_i - \hat{y})^2$
- 최소제곱법 사용?
- $y_i$는 0 또는 1의 값이 나오고
- $\hat{y}$는 0에서 1사이의 값이 나온다.
- 어떻게 계산해? $y_i$를 실수화시키자
위 로그가능도함수를 β에 대해서 최대화시키면 그 값이 추정량이 된다.
- 밑에서 family에 반응변수의 특성을 입력해주자
- continuous? binomial?
```
lmod <- glm(chd ~ height + cigs, family = binomial, wcgs)
summary(lmod)
```
- $g(\hat{p}_i) = log \frac{\hat{p}_i}{1-\hat{p}_i} = -4.50 + 0.025 h + 0.023 c$
- 키가 커지면, 담배를 많이 피면 질환 결릴 확률이 증가하는가?
- 명확히 말하면 $log \frac{\hat{p}_i}{1-\hat{p}_i}$이 증가하지
- monotone increasing function

- 즉, h,c가 증가하면 $\hat{p}_i$가 증가한다고 해석가능, 즉 방향으로 해석가능
null deviance에서 residual deviance로 32.3 정도 감소했다.
- null deviance는 parameter 다 뺀 가장 작은 deviance라고 생각
- residual deviance = $D_M = -2(\log L_M - \log L_s)$
```r
Null deviance: 1781.2 on 3153 degrees of freedom
Residual deviance: 1749.0 on 3151 degrees of freedom
```
```
(beta <- coef(lmod))
plot(jitter(y,0.1) ~ jitter(height), wcgs, xlab="Height", ylab="Heart Disease",pch=".")
curve(ilogit(beta[1] + beta[2]*x + beta[3]*0),add=TRUE) # predicted curve with non-smokers
curve(ilogit(beta[1] + beta[2]*x + beta[3]*20),add=TRUE,lty=2) # predicted curve with smokers
```
- 흡연을 안하는 집단과
- $\hat{\beta}_0 + \hat{\beta}_1 \times h + \hat{\beta}_2 \times 0$
- 흡연을 한 갑하는 집단
- $\hat{\beta}_0 + \hat{\beta}_1 \times h + \hat{\beta}_2 \times 20$
- 흡연량에 따라 어떻게 변할까?
- $\hat{\beta})_1 >0$
```
plot(jitter(y,0.1) ~ jitter(cigs), wcgs, xlab="Cigarette Use", ylab="Heart Disease",pch=".")
curve(ilogit(beta[1] + beta[2]*60 + beta[3]*x),add=TRUE) # predicted curve with short men
curve(ilogit(beta[1] + beta[2]*78 + beta[3]*x),add=TRUE,lty=2) # predicted curve with tall men
```
- 키 작은 집단과
- $\hat{\beta}_0 + \hat{\beta}_1 \times 60 + \hat{\beta}_2 \times c$
- 키 큰 집단
- $\hat{\beta}_0 + \hat{\beta}_1 \times 78 + \hat{\beta}_2 \times c$
- 키에 따라 어떻게 변할까?
계수에 대한 해석은 일반적인 선형모형과 유사하다. 하지만, Y와 직접 연결하여 해석할 수 없고 연결함수와 함께 해석하여함 함을 유의하여라.
- 자세한 해석을 위해
연관성의 방향은 연결함수와 상관없이 할 수 있다.(why?)
### 1.4 오즈 : Interpreting Odds
로지스틱 모형은 의학분야에서 특히 널리 활용된다. 그 이유 중 하나는 오즈값 혹은 오즈비에 대한 해석이 모형으로부터 direct하게 가능하기 때문이다. 오즈값은 chance에 대한 확률의 대체재로 볼 수 있다.
$$Odd=\frac{p}{1−p} ∈(0,∞)$$
- ex) $p=0.8, 1-p=0.2$
- $\frac{p}{1-p} = \frac{0.8}{0.2} = 4$
- 이길 확률이 질 확률의 4배
- ex) $p=0.75, 1-p=0.25$
- $\frac{p}{1-p} = \frac{0.75}{0.25} = 3$
- 이길 확률이 질 확률의 3배
만약 설명변수가 두 개라면 log-odds는 다음과 같이 표현된다.
$$log(\frac{p}{1−p})=β_0+β_1x_1+β_2x_2$$
때로는 서로 다른 개체들의 오즈값을 비교하고자 한다. 만약, $x_1$이 1단위만큼 변한다면 혹은 $x_1$이 0과 1의 값을 가지는 factor라면 두 경우의 오즈값의 변화는 $exp(β_1)$이 된다.
||$x_1$|$x_2$|
|---|---|---|
|A|65|10|
|B|65|11|
$\log \frac{p_A}{1-p_A} = \beta_0 + \beta_1 10 + \beta_2 65$
$\log \frac{p_B}{1-p_B} = \beta_0 + \beta_1 11 + \beta_2 65$
$\to \log \frac{p_B}{1-p_B} - \log \frac{p_A}{1-p_A} = \beta_1$
$\frac{\frac{p_B}{1-p_B}}{\frac{p_A}{1-p_A}} = exp(\beta_1)$
- $x$가 1 증가했을때 오즈값의 변화
```
exp(beta)
exp(beta[3]*20)
```
이를 일반화하여 오즈비(odds ratio)로 쓰기도 한다.
$$Odds ratio (OR)=\frac{p_1}{1−p_1} \big/ \frac{p_2}{1−p_2}, \begin{cases} 1 & 1\approx2\\ >1 &1>2\\ <1 &1<2\end{cases}$$
오즈비는 다음과 같이 정의되는 상대위험도(relative risk)와는 약간 다르다.
$$Relative risk (RR)=\frac{p_1}{p_2}, \begin{cases} 1 & 1\approx2\\ >1 &1>2\\ <1 &1<2\end{cases}$$
직관적으로는 상대위험도가 이해하기 더 쉽다. 하지만, 이 값은 어떤 실험환경에서는 계산이 불가능하다.
- 코호트 연구
- 돈, 시간이 많이 든다.
- 특히 희귀질병은 샘플이 많이 필요하다.
- 그룹을 나누어 위험도 측정
- Odds ratio 계산 가능하다.
- Rerative Risk 계산 가능하다.
- 후향적 연구
- 나중에 물어보는 연구
- Rerative Ratio 구하기 힘들다. 제어가 힘들기 때문
- Odds ratio 계산은 가능
||$x_1$|$x_2$|
|---|---|---|
|A|68|20|
|B|68|0|
```
c(ilogit(sum(beta*c(1,68,20))),ilogit(sum(beta*c(1,68,0))))
```
- 1보다 작네?
```
ilogit(sum(beta*c(1,68,20)))/ilogit(sum(beta*c(1,68,0)))
```
- 질병에 걸릴 확률이 담배를20개피 폈을때가 안 폈을때보다 1.5배 정도 높다.
- ilogit 은 logit의 역함수
여기서는 상대위험도와 오즈비의 값이 굉장히 비슷하다. 희박하게 발생하는 사건의 경우 위 두 값은 보통 크게 차이나지 않으나 그렇지 않으면 매우 달라진다.
- 만약, $p_1. p_2$가 작다면? $p_1,p_2 \approx 0$?
$\frac{\frac{p_1}{1-p_1}}{\frac{p_2}{1-p_2}} \approx \frac{p_1}{p_2} \neq \frac{p_1}{p_2}$
- 너무 작으면 값이 비슷하게 나온다.
- 커지면 비슷하지 않겠지만, 적어도 양/음수는 같다.
```
p1 = 0.05;p2 = 0.03
p2/p1; (p2/(1-p2))/(p1/(1-p1))
```
- 비슷하네
```
p1 = 0.5;p2 = 0.3
p2/p1; (p2/(1-p2))/(p1/(1-p1))
```
- 점점 차이나기 시작
```
p1 = 0.95;p2 = 0.97
p2/p1; (p2/(1-p2))/(p1/(1-p1))
```
- 거의 안 비슷하다고 봐야지.
if $exp(\beta_1)>1$?
- Odds ratio vs Rerative risk
- $\frac{p_B}{1-p_B} > \frac{p_A}{1-p_A} \therefore p_B > p_A , \beta > 0$
if $exp(\beta_1)<1$?
- Odds ratio vs Rerative risk
- $\frac{p_B}{1-p_B} < \frac{p_A}{1-p_A} \therefore p_B < p_A , \beta < 0$
Odds ratio 는 Rerative Risk 보다 증폭되는 경향이 있다.
$\frac{p_A}{1-p_A}, p_A = \frac{1}{3}, p_B=\frac{1}{4}, p_A > p_B$
- A가 더 위험하다. 즉, 질병발생 확률이 더 크다.
$\frac{\frac{p_A}{1-p_A}}{\frac{p_B}{1-p_B}} = \frac{\frac{\frac{1}{3}}{\frac{2}{3}}}{\frac{\frac{1}{4}}{\frac{3}{4}}} = \frac{\frac{1}{2}}{\frac{1}{3}} = \frac{3}{2} = 1.5$
- 즉, A의 Odds와 B의 Odds 사이의 ratio
- A의 Odds가 b의 Odds 보다 1.5만큼 크다,
$\frac{p_A}{p_B} = \frac{\frac{1}{3}}{\frac{1}{4}} = \frac{4}{3} = 1.333$
- 원래 1.3배인데 Odds ratio는 1.5?
- 증폭되었네?
if $p_A = p_B$?
- Odds ratio = 1
- Reratve Risk = 1
### 1.5 Inference
추정 이후에는 적절한 통계적 추론 과정이 필요할 수 있다. 이를 위해서는 잔차제곱합을 일반화한 개념이 필요하다. 선형모형에서 잔차제곱합은 다음과 같이 정의되었다.
$$∑^{n}_{i=1}(y_i−\hat{y_i})^2$$
- $s(\beta) \sum^{n}_{i=1}(y_i - X_{i}^{\top} \beta )^2$
- $s(\hat{\beta} = \sum^{n}_{i=1}(y - \hat{y})^2 = \sum^{n}_{i=1} (y-X\beta)^2 = SSE$
- $-2[l_{\mathcal{M}} - l_s] \ge 0$, 단 $l_{\mathcal{M}}\le l_s$
이는 선형모형이 자료를 얼마나 잘 적합하는지를 평가하는 하나의 측도이다. 이를 일반화하여 Deviance라는 개념을 소개한다. 특정 모형$(M)$에 대한 deviance는 다음과 같이 정의된다.
- deviance를 가장 많이 사용하고, deviance가 작을수록 포화모형이다.
- $SSE_s = 0, SSE_\mathcal{M} = \sum^{n}_{i=1} (y_i - \hat{y})^2$
$$D_M=−2 \log \big( \frac{L_{\mathcal{M}}}{L_S} \big)$$
- $-2(\log l_{\mathcal{M}} - \log l_s) \ge 0$
- $L(\hat{\beta}) = l_{\mathcal{M}}$
- $exp(\beta) = l_{\mathcal{M}}$
여기서 $L_{\mathcal{M}}$은 **최대화된 가능도 값**을 의미한다. $L_S$란 **포화모형(Saturated model)에서의 최대화된 가능도 값**을 의미한다. 포화모형이란 자료 하나당 모수를 하나씩 대응시켜 perfect fit을 가져오는 모형을 의미한다. 위 deviance값이 작을 수록 모형의 적합도가 높은 것으로 평가한다. 적절한 가정 하에서 deviance값은 적절한 자유도를 가지는 **카이제곱 분포**로 근사할 수 있는데 이를 이용하여 모형에 대한 유의성 검정을 수행할 수 있게 된다.
- $D_{\mathcal{M}} \sim \chi^2_{(K)}$
참고로 deviance는 오차항이 정규분포인 선형모형에서는 잔차제곱합의 정의와 일치하게 됨이 알려져 있다.
R 결과창에서 `Null deviance` 는 설명변수를 하나도 포함시키지 않은 모형에서의 deviance를 의미하고 `Residual deviance`가 현재 적합된 모형의 deviance를 의미한다. 만약 이 둘 사이에 큰 차이가 있다면 현재 모형이 유의성이 있다는 의미이다 (why?).
```
1-pchisq(32.2,2) # differnece between deviances, difference between models
```
- nulldeviance 17812 - deviance 1749 = 32.2
- 유의미한 변화일까?
- $H_0 : \beta_1 \in \beta_2 = 0$ vs $H_1 : not H_0$
모형은 유의수준 0.05에서 유의하다. 개별변수의 유의성은 anova함수를 이용하여 검정할 수 있다.
```
lmodc <- glm(chd ~ cigs, family = binomial, wcgs)
anova(lmodc,lmod, test="Chi")
```
- p값은 0.3374로 유의미한 변화(D감소)를 가져오지 않았다.
- 자유도도 1로, 파라메터 차이도 1이다.
- $H_0: \beta_\eta$ vs $H_1 : \beta_\eta \ne 0$
```
confint(lmod) # confidence intervals
```
### 1.6 Goodness of fit
- logistic의 필수 step은 아니다.
- 가장 잘 적합하는 $\hat{\beta}$ 잘 찾았나?
```
library(glmtoolbox)
hltest(lmod)
```
- 위 결과 보는데 obserded와 expected 값의 차이가 너무 커서 확인 후 알려주신대~
- p값도 좋지 않았다.(낮았다.)
The logistic model fits data well? : Hosmer-Lemeshow test
https://en.wikipedia.org/wiki/Hosmer%E2%80%93Lemeshow_test
### 1.7 Other link functions
로지스틱 모형이 가장 폭넓게 쓰이지만 다른 연결함수들을 고려할 수도 있다. 단, 연결함수는 다음의 성질을 만족해야 할 것이다.
- 증가함수
- 적절한 range를 보장
- $g(0,1) \to \cal{R}$ 실수 전체를 아우르면 좋겠다.
대체연결함수로 사용할 수 있는 것들에는 다음이 있다.
- Probit : $g=Φ^{−1}(p) , Φ : N(0,1)$의 CDF.
- CDF $Φ: \cal{R} \to (0,1)$
- $Φ^{-1} : (0,1) \to \cal{R}$
- Complementary log-log : $g=log(−log(1−p))$.
- $g(0,1) \to \cal{R}$
- Cauchit : $g=\tan(π(p−0.5))$
```
g = seq(-5,5,0.01)
plot(g,ilogit(g),type="l")
lines(g,pnorm(g),col=2)
lines(g,1-exp(-exp(g)),col=3)
lines(g,atan(g)/pi + 0.5,col=4)
abline(v=0,h=0.5,col=2,lty="dashed")
```
- 어떤 link function을 사용해야 하는가? 답은 없다.
- 검은색 선: logit link
- 점(0,0.5)에 대해 대칭
- 빨간색 선: Probit : $Φ^{-1}(p_i) = Φ(X_i \beta)$
- 점(0,0.5)에 대해 대칭
- 파란색 선: Cauchit
- 점(0,0.5)에 대해 대칭
- 연두색 선: Complementary log-log : $g=log(−log(1−p))$
- 빨리 증가 -> 천천히 증가
- 연두색 선만 대칭이 아닌 것을 확인 할 수 있다.
- 약물 반응 등
어떤 연결함수를 사용할지는 몇 가지 요소에 따라 결정할 수 있지만 정해진 답은 없다. 로지스틱 모형이 가장 널리 쓰이기는 하지만 상황에 따라 다른 모형을 더 자연스럽게 받아들이는 분야도 존재하며 조금 더 확장된 모형을 사용하는 경우에는 특정 연결함수를 사용하는 것이 필요할 수 있다.
한 가지 기억할 것은 cloglog 연결함수는 대칭성질을 가지지 않는다는 점이다.
```
lmod_l <- glm(chd ~ cigs + height, family = binomial, wcgs)
lmod_c <- glm(chd ~ cigs + height, family = binomial(link = cloglog), wcgs)
lmod_p <- glm(chd ~ cigs + height, family = binomial(link = probit), wcgs)
summary(lmod_l)
```
- Complementary log-log
```
summary(lmod_c)
```
- prob
```
summary(lmod_p)
```
- estimate값이 달라졌다?
- 하지만 실제로 probability를 다르게 주진 않았다.
- 적합도가 `1748.7`로 가장 좋다고 볼 수 있지만. 소수점 차이 밖에 나지 않았자.
- 적합도 좋은 것을 찾는게 좋으나, 큰 차이게 없다면 logistic을 쓰자!
| github_jupyter |
```
import os
import csv
import sys
import re
from surprise import Dataset
from surprise import Reader
from collections import defaultdict
import numpy as np
import pandas as pd
df = pd.read_csv('ml-latest-small/movies.csv')
class MovieLens:
movie_id_to_name = {}
name_to_movie_id = {}
ratings_path = 'ml-latest-small/ratings.csv'
moviespath = 'ml-latest-small/movies.csv'
def load_movie_data(self):
rating_data = Dataset.load_from_file(self.ratings_path)
df_movie = pd.read_csv(self.moviespath)
df_movie['movieId'].head(5)
self.movie_id_to_name = dict(zip(df_movie['movieId'], df_movie['title']))
self.name_to_movie_id = dict(zip(df_movie['title'], df_movie['movieId']))
return rating_data
def get_user_rating(self,user):
df_rating = pd.read_csv(self.ratings_path)
user_rating = list(zip(df_rating.query('userId=='+user)['movieId'],df_rating.query('userId=='+user)['rating']))
return user_rating
def get_popular_ranks(self):
df_popular = pd.read_csv(self.ratings_path)
y = df_popular.groupby('movieId')['movieId'].value_counts()
movieId = [x for x,y in list(y.keys())]
frequency = list(y.values)
ratings = dict(zip(movieId,frequency))
return ratings
def get_genres(self):
genres = defaultdict(list)
genre_ids = {}
max_genre_id = 0
df_movie = pd.read_csv(self.moviespath)
movie_id_list = df_movie['movieId']
movie_title_list = df_movie['title']
movie_genres_str = df_movie['genres']
for movie_id,genre in zip(movie_id_list,movie_genres_str):
genre_list = genre.split('|')
genre_id_list = []
for genre in genre_list:
if genre in genre_ids:
genre_id = genre_ids[genre]
else:
genre_id = max_genre_id
genre_ids[genre] = genre_id
max_genre_id = max_genre_id + 1
genre_id_list.append(genre_id)
genres[movie_id] = genre_id_list
for (movie_id,genre_id_list) in genres.items():
bitfield = [0]*max_genre_id
for genre_id in genre_id_list:
bitfield[genre_id] = 1
genres[movie_id] = bitfield
return genres
def get_years(self):
p = re.compile(r"(?:\((\d{4})\))?\s*$")
years = defaultdict(int)
df_movie = pd.read_csv(self.moviespath)
movie_title_list = df_movie['title']
movie_id_list = df_movie['movieId']
for title,movie_id in zip(movie_title_list,movie_id_list):
temp = p.search(title)
year = temp.group(1)
if year:
years[movie_id] = int(year)
return years
def get_mise_en_scene(self):
mes = defaultdict(list)
df_movie = pd.read_csv(self.moviespath)
movie_id_list = df_movie['ML_Id']
avg_shot_length_list = df_movie['f1']
mean_color_variance_list = df_movie['f2']
std_dev_color_variance_list = df_movie['f3']
mean_motion_list = df_movie['f4']
std_dev_motion_list = df_movie['f5']
mean_lighting_key_list = df_movie['f6']
num_shots_list = df_movie['f7']
for a,b,c,d,e,f,g,h in zip(movie_id_list,avg_shot_length_list,mean_color_variance_list,std_dev_color_variance_list,mean_motion_list,std_dev_motion_list,mean_lighting_key_list,num_shots_list):
mes[a] = [b,c,d,e,f,g,h]
return mes
def get_movie_name(self,movie_id):
if movie_id in self.movie_id_name:
return self.movie_id_name[movie_id]
else:
return ""
def get_movie_id(self,movie_name):
if movie_name in self.name_to_movie_id:
return self.name_to_movie_id[movie_name]
else:
return 0
from surprise import AlgoBase
from surprise import PredictionImpossible
# from MovieLens import MovieLens
import math
import numpy as np
import heapq
class ContentKNNAlgorithm(AlgoBase):
def __init__(self,k=40,sim_options={}):
AlgoBase.__init__(self)
self.k = k
def fit(self,trainset):
AlgoBase.fit(self,trainset)
ml = MovieLens()
genres = ml.get_genres()
years = ml.get_years()
mes = ml.get_mise_en_scene()
```
| github_jupyter |
### YAML Format
YAML, like JSON, is another data serialization standard. It is actually easier to read than JSON, and although it has been around for a long time (since 2001), it has gained a lot of popularity, especially in the Dev Ops world for configuration files (Docker, Kubernetes, etc).
Like JSON it is able to represent simple data types (strings, numbers, boolean, etc) as well as collections and associative arrays (dictionaries).
YAML focuses on human readability, and is a little more complex to parse.
Here is a sample YAML file:
```
title: Parrot Sketch
year: 1989
actors:
- first_name: John
last_name: Cleese
dob: 1939-10-27
- first_name: Michael
last_name: Palin
dob: 1943-05-05
```
As you can see this is much easier to read than JSON or XML.
To parse YAML into a Python dictionary would take a fair amount of work - especially since YAML is quite flexible.
Fortunately, we can use the 3rd party library, `pyyaml` to do this for us.
Again, I'm only going to show you a tiny bit of this library, and you can read more about it here:
https://pyyaml.org/wiki/PyYAMLDocumentation
(It's definitely less of a learning curve than Marshmallow!!)
#### Caution
When you load a yaml file using pyyaml, be careful - like pickling it can actually call out to Python functions - so do not load untrusted YAML files using `pyyaml`!
```
import yaml
data = '''
---
title: Parrot Sketch
year: 1989
actors:
- first_name: John
last_name: Cleese
dob: 1939-10-27
- first_name: Michael
last_name: Palin
dob: 1943-05-05
'''
d = yaml.load(data)
type(d)
from pprint import pprint
pprint(d)
```
You'll notice that unlike the built-in JSON parser, PyYAML was able to automatically deduce the `date` type in our YAML, as well of course as strings and integers.
Of course, serialization works the same way:
```
d = {'a': 100, 'b': False, 'c': 10.5, 'd': [1, 2, 3]}
print(yaml.dump(d))
```
You'll notice in the above example that the list was represented using `[1, 2, 3]` - this is valid YAML as well, and is equivalent to this notation:
```
d:
- 1
- 2
- 3
```
If you prefer this block style, you can force it this way:
```
print(yaml.dump(d, default_flow_style=False))
```
What's interesting about PyYAML is that it can also automatically serialize and deserialize complex objects:
```
class Person:
def __init__(self, name, dob):
self.name = name
self.dob = dob
def __repr__(self):
return f'Person(name={self.name}, dob={self.dob})'
from datetime import date
p1 = Person('John Cleese', date(1939, 10, 27))
p2 = Person('Michael Palin', date(1934, 5, 5))
print(yaml.dump({'john': p1, 'michael': p2}))
```
Notice that weird looking syntax? It's actually useful when we deserialize the YAML string - of course it means we must have a `Person` class defined with the appropriate init method.
```
yaml_data = '''
john: !!python/object:__main__.Person
dob: 1939-10-27
name: John Cleese
michael: !!python/object:__main__.Person
dob: 1934-05-05
name: Michael Palin
'''
d = yaml.load(yaml_data)
d
```
As you can see, `john` and `michael` were deserialized into `Person` type objects.
This is why you have to be quite careful with the source of any YAML you deserialize.
Here's an evil example:
```
yaml_data = '''
exec_paths:
!!python/object/apply:os.get_exec_path []
exec_command:
!!python/object/apply:subprocess.check_output [['ls', '/']]
'''
yaml.load(yaml_data)
```
So, be very careful with `load`. In general it is safer practice to use the `safe_load` method instead, but you will lose the ability to deserialize into custom Python objects, unless you override that behavior. You can always use Marshmallow to do that secondary step in a safer way.
```
yaml.safe_load(yaml_data)
```
To override and allow certain Python objects to be deserialized in `safe_load` we can proceed this way.
Firstly we are going to simplify the object tag notation by customizing it in our `Person` class, and we are also going to make our object as safe to be deserialized. Our `Person` class will now have to inherit from the `yaml.YAMLObject`:
```
from yaml import YAMLObject, SafeLoader
class Person(YAMLObject):
yaml_tag = '!Person'
def __init__(self, name, age):
self.name = name
self.age = age
def __repr__(self):
return f'Person(name={self.name}, age={self.age})'
```
First let's see how objects are now serialized:
```
yaml.dump(dict(john=Person('John Cleese', 79),
michael=Person('Michael Palin', 74)))
```
As you can see we have a slightly cleaner syntax.
Now let's try to load the serialized version:
```
yaml_data = '''
john: !Person
name: John Cleese
age: 79
michael: !Person
name: Michael Palin
age: 74
'''
yaml.load(yaml_data)
```
And `safe_load`:
```
yaml.safe_load(yaml_data)
```
So now let's mark our `Person` object as safe:
```
class Person(YAMLObject):
yaml_tag = '!Person'
yaml_loader = SafeLoader
def __init__(self, name, age):
self.name = name
self.age = age
def __repr__(self):
return f'Person(name={self.name}, age={self.age})'
yaml.safe_load(yaml_data)
```
And as you can see, the deserializtion now works for the `Person` class.
There's a lot more this library can do, so look at the reference if you want to use YAML.
Also, as I mentionmed before, you can combine this with `Marshmallow` for example to get to a full marshalling solution to complex (custom) Python types.
| github_jupyter |
# Introduction
Do higher film budgets lead to more box office revenue? Let's find out if there's a relationship using the movie budgets and financial performance data that I've scraped from [the-numbers.com](https://www.the-numbers.com/movie/budgets) on **May 1st, 2018**.
<img src=https://i.imgur.com/kq7hrEh.png>
# Import Statements
```
import pandas as pd
import matplotlib.pyplot as plt
```
# Notebook Presentation
```
pd.options.display.float_format = '{:,.2f}'.format
from pandas.plotting import register_matplotlib_converters
register_matplotlib_converters()
```
# Read the Data
```
data = pd.read_csv('cost_revenue_dirty.csv')
```
# Explore and Clean the Data
**Challenge**: Answer these questions about the dataset:
1. How many rows and columns does the dataset contain?
2. Are there any NaN values present?
3. Are there any duplicate rows?
4. What are the data types of the columns?
```
```
### Data Type Conversions
**Challenge**: Convert the `USD_Production_Budget`, `USD_Worldwide_Gross`, and `USD_Domestic_Gross` columns to a numeric format by removing `$` signs and `,`.
<br>
<br>
Note that *domestic* in this context refers to the United States.
```
```
**Challenge**: Convert the `Release_Date` column to a Pandas Datetime type.
```
```
### Descriptive Statistics
**Challenge**:
1. What is the average production budget of the films in the data set?
2. What is the average worldwide gross revenue of films?
3. What were the minimums for worldwide and domestic revenue?
4. Are the bottom 25% of films actually profitable or do they lose money?
5. What are the highest production budget and highest worldwide gross revenue of any film?
6. How much revenue did the lowest and highest budget films make?
```
```
# Investigating the Zero Revenue Films
**Challenge** How many films grossed $0 domestically (i.e., in the United States)? What were the highest budget films that grossed nothing?
```
```
**Challenge**: How many films grossed $0 worldwide? What are the highest budget films that had no revenue internationally?
```
```
### Filtering on Multiple Conditions
```
```
**Challenge**: Use the [`.query()` function](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.query.html) to accomplish the same thing. Create a subset for international releases that had some worldwide gross revenue, but made zero revenue in the United States.
Hint: This time you'll have to use the `and` keyword.
```
```
### Unreleased Films
**Challenge**:
* Identify which films were not released yet as of the time of data collection (May 1st, 2018).
* How many films are included in the dataset that have not yet had a chance to be screened in the box office?
* Create another DataFrame called data_clean that does not include these films.
```
# Date of Data Collection
scrape_date = pd.Timestamp('2018-5-1')
```
### Films that Lost Money
**Challenge**:
What is the percentage of films where the production costs exceeded the worldwide gross revenue?
```
```
# Seaborn for Data Viz: Bubble Charts
```
```
### Plotting Movie Releases over Time
**Challenge**: Try to create the following Bubble Chart:
<img src=https://i.imgur.com/8fUn9T6.png>
```
```
# Converting Years to Decades Trick
**Challenge**: Create a column in `data_clean` that has the decade of the release.
<img src=https://i.imgur.com/0VEfagw.png width=650>
Here's how:
1. Create a [`DatetimeIndex` object](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DatetimeIndex.html) from the Release_Date column.
2. Grab all the years from the `DatetimeIndex` object using the `.year` property.
<img src=https://i.imgur.com/5m06Ach.png width=650>
3. Use floor division `//` to convert the year data to the decades of the films.
4. Add the decades as a `Decade` column to the `data_clean` DataFrame.
```
```
### Separate the "old" (before 1969) and "New" (1970s onwards) Films
**Challenge**: Create two new DataFrames: `old_films` and `new_films`
* `old_films` should include all the films before 1969 (up to and including 1969)
* `new_films` should include all the films from 1970 onwards
* How many films were released prior to 1970?
* What was the most expensive film made prior to 1970?
```
```
# Seaborn Regression Plots
```
```
**Challenge**: Use Seaborn's `.regplot()` to show the scatter plot and linear regression line against the `new_films`.
<br>
<br>
Style the chart
* Put the chart on a `'darkgrid'`.
* Set limits on the axes so that they don't show negative values.
* Label the axes on the plot "Revenue in \$ billions" and "Budget in \$ millions".
* Provide HEX colour codes for the plot and the regression line. Make the dots dark blue (#2f4b7c) and the line orange (#ff7c43).
Interpret the chart
* Do our data points for the new films align better or worse with the linear regression than for our older films?
* Roughly how much would a film with a budget of $150 million make according to the regression line?
```
```
# Run Your Own Regression with scikit-learn
$$ REV \hat ENUE = \theta _0 + \theta _1 BUDGET$$
```
```
**Challenge**: Run a linear regression for the `old_films`. Calculate the intercept, slope and r-squared. How much of the variance in movie revenue does the linear model explain in this case?
```
```
# Use Your Model to Make a Prediction
We just estimated the slope and intercept! Remember that our Linear Model has the following form:
$$ REV \hat ENUE = \theta _0 + \theta _1 BUDGET$$
**Challenge**: How much global revenue does our model estimate for a film with a budget of $350 million?
```
```
| github_jupyter |
```
import os
import matplotlib.pyplot as plt
import glob
from PIL import Image
import numpy as np
from sklearn.utils import shuffle
from tensorflow.python import keras
from tensorflow.python.keras import Sequential
from tensorflow.python.keras.layers import Dense, InputLayer, Conv2D, MaxPool2D, Flatten, BatchNormalization
from keras.preprocessing.image import ImageDataGenerator
from sklearn.model_selection import train_test_split
import keras.backend as K
from keras.callbacks import History
loss_ges = np.array([])
val_loss_ges = np.array([])
%matplotlib inline
np.set_printoptions(precision=4)
np.set_printoptions(suppress=True)
Input_dir='data_resize_all'
files = glob.glob(Input_dir + '/*.*')
x_data = []
y_data = []
for aktfile in files:
test_image = Image.open(aktfile)
test_image = np.array(test_image, dtype="float32")
test_image = np.reshape(test_image, (32,32,3))
base = os.path.basename(aktfile)
zahl = (int(base[8:10])) / 100
x_data.append(test_image)
zw = np.array([zahl])
y_data.append(zw)
x_data = np.array(x_data)
y_data = np.array(y_data)
print(x_data.shape)
print(y_data.shape)
x_data, y_data = shuffle(x_data, y_data)
X_train, X_test, y_train, y_test = train_test_split(x_data, y_data, test_size=0.2)
model = Sequential()
model.add(BatchNormalization(input_shape=(32,32,3)))
model.add(Conv2D(8, (5, 5), input_shape=(32,32,3), padding='same', activation="relu"))
model.add(MaxPool2D(pool_size=(4,4)))
model.add(Conv2D(4, (5, 5), padding='same'))
model.add(MaxPool2D(pool_size=(4,4)))
model.add(Conv2D(4, (3, 3), padding='same'))
model.add(MaxPool2D(pool_size=(2,2)))
model.add(Flatten())
model.add(Dense(16,activation="relu"))
model.add(Dense(2,activation="linear"))
model.add(Dense(1))
model.summary()
def Periodic_Loss(y_true, y_pred):
dif1 = K.abs(y_pred - y_true)
dif2 = K.abs(dif1 + K.constant(1))
# dif3 = K.abs(dif1 + K.constant(-1))
dif = K.minimum(dif1, dif2)
# dif = K.minimum(dif, dif3)
ret = K.mean(K.square(dif), axis=-1)
return ret
model.compile(loss=Periodic_Loss, optimizer=keras.optimizers.Adadelta(), metrics = ["accuracy"])
#model.compile(loss=keras.losses.mean_squared_error, optimizer=keras.optimizers.Adadelta(), metrics = ["accuracy"])
Batch_Size = 8
Epoch_Anz = 20
Shift_Range = 0
Brightness_Range = 0.3
datagen = ImageDataGenerator(width_shift_range=[-Shift_Range,Shift_Range], height_shift_range=[-Shift_Range,Shift_Range],brightness_range=[1-Brightness_Range,1+Brightness_Range])
train_iterator = datagen.flow(X_train, y_train, batch_size=Batch_Size)
validation_iterator = datagen.flow(X_test, y_test, batch_size=Batch_Size)
history = model.fit_generator(train_iterator, validation_data = validation_iterator, epochs = Epoch_Anz)
loss_ges = np.append(loss_ges, history.history['loss'])
val_loss_ges = np.append(val_loss_ges, history.history['val_loss'])
plt.semilogy(history.history['loss'])
plt.semilogy(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train','eval'], loc='upper left')
plt.show()
Input_dir='data_resize_all'
files = glob.glob(Input_dir + '/*.*')
res = []
for aktfile in files:
base = os.path.basename(aktfile)
zahl = (int(base[8:10])) / 100
test_image = Image.open(aktfile)
test_image = np.array(test_image, dtype="float32")
img = np.reshape(test_image,[1,32,32,3])
classes = model.predict(img)
zw1 = zahl
zw2 = round(classes[0][0], 2)
zw3 = round(classes[0][0] - zahl, 2)
zw4a = abs(zw3)
zw4b = abs(zw3+1)
zw4c = abs(zw3-1)
zw4 = zw3
if zw4b < zw4a:
zw4 = zw3+1
zw4a = zw4b
if zw4c < zw4a:
zw4 = zw3-1
res.append(np.array([zw1, zw2, zw3, zw4]))
# print(base, ', ', zw1, ', ', round(zw2, 2), ', ', round(zw3, 2), ', ', round(zw4, 2))
res = np.asarray(res)
statistic = np.array([np.mean(res[:,3]), np.std(res[:,3]), np.min(res[:,3]), np.max(res[:,3])])
print(statistic)
res_step_1 = res
plt.plot(res[:,0])
plt.plot(res[:,1])
plt.title('Result')
plt.ylabel('Analog Value')
plt.xlabel('#Picture')
plt.legend(['real','model'], loc='upper left')
plt.show()
plt.plot(res[:,3])
plt.title('Deviation')
plt.ylabel('Deviation from expected value')
plt.xlabel('#Picture')
plt.legend(['model'], loc='upper left')
plt.show()
Batch_Size = 8
Epoch_Anz = 40
Shift_Range = 1
Brightness_Range = 0.3
datagen = ImageDataGenerator(width_shift_range=[-Shift_Range,Shift_Range], height_shift_range=[-Shift_Range,Shift_Range],brightness_range=[1-Brightness_Range,1+Brightness_Range])
train_iterator = datagen.flow(X_train, y_train, batch_size=Batch_Size)
validation_iterator = datagen.flow(X_test, y_test, batch_size=Batch_Size)
history = model.fit_generator(train_iterator, validation_data = validation_iterator, epochs = Epoch_Anz)
loss_ges = np.append(loss_ges, history.history['loss'])
val_loss_ges = np.append(val_loss_ges, history.history['val_loss'])
plt.semilogy(history.history['loss'])
plt.semilogy(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train','eval'], loc='upper left')
plt.show()
Input_dir='data_resize_all'
files = glob.glob(Input_dir + '/*.*')
res = []
for aktfile in files:
base = os.path.basename(aktfile)
zahl = (int(base[8:10])) / 100
test_image = Image.open(aktfile)
test_image = np.array(test_image, dtype="float32")
img = np.reshape(test_image,[1,32,32,3])
classes = model.predict(img)
zw1 = zahl
zw2 = round(classes[0][0], 2)
zw3 = round(classes[0][0] - zahl, 2)
zw4a = abs(zw3)
zw4b = abs(zw3+1)
zw4c = abs(zw3-1)
zw4 = zw3
if zw4b < zw4a:
zw4 = zw3+1
zw4a = zw4b
if zw4c < zw4a:
zw4 = zw3-1
res.append(np.array([zw1, zw2, zw3, zw4]))
# print(base, ', ', zw1, ', ', round(zw2, 2), ', ', round(zw3, 2), ', ', round(zw4, 2))
res = np.asarray(res)
statistic = np.array([np.mean(res[:,3]), np.std(res[:,3]), np.min(res[:,3]), np.max(res[:,3])])
print(statistic)
res_step_1 = res
plt.plot(res[:,0])
plt.plot(res[:,1])
plt.title('Result')
plt.ylabel('Analog Value')
plt.xlabel('#Picture')
plt.legend(['real','model'], loc='upper left')
plt.show()
plt.plot(res[:,3])
plt.title('Deviation')
plt.ylabel('Deviation from expected value')
plt.xlabel('#Picture')
plt.legend(['model'], loc='upper left')
plt.ylim(-0.3, 0.3)
plt.show()
model.save("test.h5")
plt.semilogy(loss_ges)
plt.semilogy(val_loss_ges)
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train','eval'], loc='upper left')
plt.ylim(1E-1, 5E-5)
plt.show()
```
| github_jupyter |
# Training Neural Networks
The network we built in the previous part isn't so smart, it doesn't know anything about our handwritten digits. Neural networks with non-linear activations work like universal function approximators. There is some function that maps your input to the output. For example, images of handwritten digits to class probabilities. The power of neural networks is that we can train them to approximate this function, and basically any function given enough data and compute time.
<img src="https://github.com/Eylen/deep-learning-v2-pytorch/blob/master/intro-to-pytorch/assets/function_approx.png?raw=1" width=500px>
At first the network is naive, it doesn't know the function mapping the inputs to the outputs. We train the network by showing it examples of real data, then adjusting the network parameters such that it approximates this function.
To find these parameters, we need to know how poorly the network is predicting the real outputs. For this we calculate a **loss function** (also called the cost), a measure of our prediction error. For example, the mean squared loss is often used in regression and binary classification problems
$$
\large \ell = \frac{1}{2n}\sum_i^n{\left(y_i - \hat{y}_i\right)^2}
$$
where $n$ is the number of training examples, $y_i$ are the true labels, and $\hat{y}_i$ are the predicted labels.
By minimizing this loss with respect to the network parameters, we can find configurations where the loss is at a minimum and the network is able to predict the correct labels with high accuracy. We find this minimum using a process called **gradient descent**. The gradient is the slope of the loss function and points in the direction of fastest change. To get to the minimum in the least amount of time, we then want to follow the gradient (downwards). You can think of this like descending a mountain by following the steepest slope to the base.
<img src='https://github.com/Eylen/deep-learning-v2-pytorch/blob/master/intro-to-pytorch/assets/gradient_descent.png?raw=1' width=350px>
## Backpropagation
For single layer networks, gradient descent is straightforward to implement. However, it's more complicated for deeper, multilayer neural networks like the one we've built. Complicated enough that it took about 30 years before researchers figured out how to train multilayer networks.
Training multilayer networks is done through **backpropagation** which is really just an application of the chain rule from calculus. It's easiest to understand if we convert a two layer network into a graph representation.
<img src='https://github.com/Eylen/deep-learning-v2-pytorch/blob/master/intro-to-pytorch/assets/backprop_diagram.png?raw=1' width=550px>
In the forward pass through the network, our data and operations go from bottom to top here. We pass the input $x$ through a linear transformation $L_1$ with weights $W_1$ and biases $b_1$. The output then goes through the sigmoid operation $S$ and another linear transformation $L_2$. Finally we calculate the loss $\ell$. We use the loss as a measure of how bad the network's predictions are. The goal then is to adjust the weights and biases to minimize the loss.
To train the weights with gradient descent, we propagate the gradient of the loss backwards through the network. Each operation has some gradient between the inputs and outputs. As we send the gradients backwards, we multiply the incoming gradient with the gradient for the operation. Mathematically, this is really just calculating the gradient of the loss with respect to the weights using the chain rule.
$$
\large \frac{\partial \ell}{\partial W_1} = \frac{\partial L_1}{\partial W_1} \frac{\partial S}{\partial L_1} \frac{\partial L_2}{\partial S} \frac{\partial \ell}{\partial L_2}
$$
**Note:** I'm glossing over a few details here that require some knowledge of vector calculus, but they aren't necessary to understand what's going on.
We update our weights using this gradient with some learning rate $\alpha$.
$$
\large W^\prime_1 = W_1 - \alpha \frac{\partial \ell}{\partial W_1}
$$
The learning rate $\alpha$ is set such that the weight update steps are small enough that the iterative method settles in a minimum.
## Losses in PyTorch
Let's start by seeing how we calculate the loss with PyTorch. Through the `nn` module, PyTorch provides losses such as the cross-entropy loss (`nn.CrossEntropyLoss`). You'll usually see the loss assigned to `criterion`. As noted in the last part, with a classification problem such as MNIST, we're using the softmax function to predict class probabilities. With a softmax output, you want to use cross-entropy as the loss. To actually calculate the loss, you first define the criterion then pass in the output of your network and the correct labels.
Something really important to note here. Looking at [the documentation for `nn.CrossEntropyLoss`](https://pytorch.org/docs/stable/nn.html#torch.nn.CrossEntropyLoss),
> This criterion combines `nn.LogSoftmax()` and `nn.NLLLoss()` in one single class.
>
> The input is expected to contain scores for each class.
This means we need to pass in the raw output of our network into the loss, not the output of the softmax function. This raw output is usually called the *logits* or *scores*. We use the logits because softmax gives you probabilities which will often be very close to zero or one but floating-point numbers can't accurately represent values near zero or one ([read more here](https://docs.python.org/3/tutorial/floatingpoint.html)). It's usually best to avoid doing calculations with probabilities, typically we use log-probabilities.
```
import torch
from torch import nn
import torch.nn.functional as F
from torchvision import datasets, transforms
# Define a transform to normalize the data
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,)),
])
# Download and load the training data
trainset = datasets.MNIST('~/.pytorch/MNIST_data/', download=True, train=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
```
### Note
If you haven't seen `nn.Sequential` yet, please finish the end of the Part 2 notebook.
```
# Build a feed-forward network
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10))
# Define the loss
criterion = nn.CrossEntropyLoss()
# Get our data
images, labels = next(iter(trainloader))
# Flatten images
images = images.view(images.shape[0], -1)
# Forward pass, get our logits
logits = model(images)
# Calculate the loss with the logits and the labels
loss = criterion(logits, labels)
print(loss)
```
In my experience it's more convenient to build the model with a log-softmax output using `nn.LogSoftmax` or `F.log_softmax` ([documentation](https://pytorch.org/docs/stable/nn.html#torch.nn.LogSoftmax)). Then you can get the actual probabilities by taking the exponential `torch.exp(output)`. With a log-softmax output, you want to use the negative log likelihood loss, `nn.NLLLoss` ([documentation](https://pytorch.org/docs/stable/nn.html#torch.nn.NLLLoss)).
>**Exercise:** Build a model that returns the log-softmax as the output and calculate the loss using the negative log likelihood loss. Note that for `nn.LogSoftmax` and `F.log_softmax` you'll need to set the `dim` keyword argument appropriately. `dim=0` calculates softmax across the rows, so each column sums to 1, while `dim=1` calculates across the columns so each row sums to 1. Think about what you want the output to be and choose `dim` appropriately.
```
# TODO: Build a feed-forward network
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
# TODO: Define the loss
criterion = nn.NLLLoss()
### Run this to check your work
# Get our data
images, labels = next(iter(trainloader))
# Flatten images
images = images.view(images.shape[0], -1)
# Forward pass, get our logits
logits = model(images)
# Calculate the loss with the logits and the labels
loss = criterion(logits, labels)
print(loss)
```
## Autograd
Now that we know how to calculate a loss, how do we use it to perform backpropagation? Torch provides a module, `autograd`, for automatically calculating the gradients of tensors. We can use it to calculate the gradients of all our parameters with respect to the loss. Autograd works by keeping track of operations performed on tensors, then going backwards through those operations, calculating gradients along the way. To make sure PyTorch keeps track of operations on a tensor and calculates the gradients, you need to set `requires_grad = True` on a tensor. You can do this at creation with the `requires_grad` keyword, or at any time with `x.requires_grad_(True)`.
You can turn off gradients for a block of code with the `torch.no_grad()` content:
```python
x = torch.zeros(1, requires_grad=True)
>>> with torch.no_grad():
... y = x * 2
>>> y.requires_grad
False
```
Also, you can turn on or off gradients altogether with `torch.set_grad_enabled(True|False)`.
The gradients are computed with respect to some variable `z` with `z.backward()`. This does a backward pass through the operations that created `z`.
```
x = torch.randn(2,2, requires_grad=True)
print(x)
y = x**2
print(y)
```
Below we can see the operation that created `y`, a power operation `PowBackward0`.
```
## grad_fn shows the function that generated this variable
print(y.grad_fn)
```
The autograd module keeps track of these operations and knows how to calculate the gradient for each one. In this way, it's able to calculate the gradients for a chain of operations, with respect to any one tensor. Let's reduce the tensor `y` to a scalar value, the mean.
```
z = y.mean()
print(z)
```
You can check the gradients for `x` and `y` but they are empty currently.
```
print(x.grad)
```
To calculate the gradients, you need to run the `.backward` method on a Variable, `z` for example. This will calculate the gradient for `z` with respect to `x`
$$
\frac{\partial z}{\partial x} = \frac{\partial}{\partial x}\left[\frac{1}{n}\sum_i^n x_i^2\right] = \frac{x}{2}
$$
```
z.backward()
print(x.grad)
print(x/2)
```
These gradients calculations are particularly useful for neural networks. For training we need the gradients of the cost with respect to the weights. With PyTorch, we run data forward through the network to calculate the loss, then, go backwards to calculate the gradients with respect to the loss. Once we have the gradients we can make a gradient descent step.
## Loss and Autograd together
When we create a network with PyTorch, all of the parameters are initialized with `requires_grad = True`. This means that when we calculate the loss and call `loss.backward()`, the gradients for the parameters are calculated. These gradients are used to update the weights with gradient descent. Below you can see an example of calculating the gradients using a backwards pass.
```
# Build a feed-forward network
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
images, labels = next(iter(trainloader))
images = images.view(images.shape[0], -1)
logits = model(images)
loss = criterion(logits, labels)
print('Before backward pass: \n', model[0].weight.grad)
loss.backward()
print('After backward pass: \n', model[0].weight.grad)
```
## Training the network!
There's one last piece we need to start training, an optimizer that we'll use to update the weights with the gradients. We get these from PyTorch's [`optim` package](https://pytorch.org/docs/stable/optim.html). For example we can use stochastic gradient descent with `optim.SGD`. You can see how to define an optimizer below.
```
from torch import optim
# Optimizers require the parameters to optimize and a learning rate
optimizer = optim.SGD(model.parameters(), lr=0.01)
```
Now we know how to use all the individual parts so it's time to see how they work together. Let's consider just one learning step before looping through all the data. The general process with PyTorch:
* Make a forward pass through the network
* Use the network output to calculate the loss
* Perform a backward pass through the network with `loss.backward()` to calculate the gradients
* Take a step with the optimizer to update the weights
Below I'll go through one training step and print out the weights and gradients so you can see how it changes. Note that I have a line of code `optimizer.zero_grad()`. When you do multiple backwards passes with the same parameters, the gradients are accumulated. This means that you need to zero the gradients on each training pass or you'll retain gradients from previous training batches.
```
print('Initial weights - ', model[0].weight)
images, labels = next(iter(trainloader))
images.resize_(64, 784)
# Clear the gradients, do this because gradients are accumulated
optimizer.zero_grad()
# Forward pass, then backward pass, then update weights
output = model(images)
loss = criterion(output, labels)
loss.backward()
print('Gradient -', model[0].weight.grad)
# Take an update step and few the new weights
optimizer.step()
print('Updated weights - ', model[0].weight)
```
### Training for real
Now we'll put this algorithm into a loop so we can go through all the images. Some nomenclature, one pass through the entire dataset is called an *epoch*. So here we're going to loop through `trainloader` to get our training batches. For each batch, we'll doing a training pass where we calculate the loss, do a backwards pass, and update the weights.
>**Exercise:** Implement the training pass for our network. If you implemented it correctly, you should see the training loss drop with each epoch.
```
## Your solution here
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
optimizer = optim.SGD(model.parameters(), lr=0.003)
def train_step(optimizer, model, criterion, images, labels):
optimizer.zero_grad()
output = model(images)
loss = criterion(output, labels)
loss.backward()
optimizer.step()
return loss
epochs = 5
for e in range(epochs):
running_loss = 0
for images, labels in trainloader:
# Flatten MNIST images into a 784 long vector
images = images.view(images.shape[0], -1)
# TODO: Training pass
loss = train_step(optimizer, model, criterion, images, labels)
running_loss += loss.item()
else:
print(f"Training loss: {running_loss/len(trainloader)}")
```
With the network trained, we can check out it's predictions.
```
%matplotlib inline
import helper
images, labels = next(iter(trainloader))
img = images[0].view(1, 784)
# Turn off gradients to speed up this part
with torch.no_grad():
logps = model(img)
# Output of the network are log-probabilities, need to take exponential for probabilities
ps = torch.exp(logps)
helper.view_classify(img.view(1, 28, 28), ps)
```
Now our network is brilliant. It can accurately predict the digits in our images. Next up you'll write the code for training a neural network on a more complex dataset.
| github_jupyter |
```
import json
import pandas as pd
import numpy as np
import re
from sqlalchemy import create_engine
import psycopg2
from config import db_password
import time
# 1. Add the clean movie function that takes in the argument, "movie".
def clean_movie(movie):
movie = dict(movie) #create a non-destructive copy
alt_titles = {}
# combine alternate titles into one list
for key in ['Also known as','Arabic','Cantonese','Chinese','French',
'Hangul','Hebrew','Hepburn','Japanese','Literally',
'Mandarin','McCune-Reischauer','Original title','Polish',
'Revised Romanization','Romanized','Russian',
'Simplified','Traditional','Yiddish']:
if key in movie:
alt_titles[key] = movie[key]
movie.pop(key)
if len(alt_titles) > 0:
movie['alt_titles'] = alt_titles
# merge column names
def change_column_name(old_name, new_name):
if old_name in movie:
movie[new_name] = movie.pop(old_name)
change_column_name('Adaptation by', 'Writer(s)')
change_column_name('Country of origin', 'Country')
change_column_name('Directed by', 'Director')
change_column_name('Distributed by', 'Distributor')
change_column_name('Edited by', 'Editor(s)')
change_column_name('Length', 'Running time')
change_column_name('Original release', 'Release date')
change_column_name('Music by', 'Composer(s)')
change_column_name('Produced by', 'Producer(s)')
change_column_name('Producer', 'Producer(s)')
change_column_name('Productioncompanies ', 'Production company(s)')
change_column_name('Productioncompany ', 'Production company(s)')
change_column_name('Released', 'Release Date')
change_column_name('Release Date', 'Release date')
change_column_name('Screen story by', 'Writer(s)')
change_column_name('Screenplay by', 'Writer(s)')
change_column_name('Story by', 'Writer(s)')
change_column_name('Theme music composer', 'Composer(s)')
change_column_name('Written by', 'Writer(s)')
return movie
# 2 Add the function that takes in three arguments;
# Wikipedia data, Kaggle metadata, and MovieLens rating data (from Kaggle)
def extract_transform_load():
# Read in the kaggle metadata and MovieLens ratings CSV files as Pandas DataFrames.
kaggle_metadata = pd.read_csv(kaggle_file, low_memory=False)
ratings = pd.read_csv(ratings_file)
# Open and read the Wikipedia data JSON file.
with open(wiki_file, mode='r') as file:
wiki_movies_raw = json.load(file)
# 3. Write a list comprehension to filter out TV shows.
wiki_movies = [movie for movie in wiki_movies_raw
if ('Director' in movie or 'Directed by' in movie)
and 'imdb_link' in movie
and 'No. of episodes' not in movie]
#wiki_movies=[for movie in wiki_movies_raw if ("Director" in movie or "Directed by" in movie) and "imdb_link" in movie]
#and "No._of_episodes" not in movie]
# 4. Write a list comprehension to iterate through the cleaned wiki movies list
# and call the clean_movie function on each movie.
clean_movies=[clean_movie(movie) for movie in wiki_movies]
# 5. Read in the cleaned movies list from Step 4 as a DataFrame.
wiki_movies_df = pd.DataFrame(clean_movies)
# 6. Write a try-except block to catch errors while extracting the IMDb ID using a regular expression string and
# dropping any imdb_id duplicates. If there is an error, capture and print the exception.
try:
wiki_movies_df['imdb_id'] = wiki_movies_df['imdb_link'].str.extract(r'(tt\d{7})')
wiki_movies_df.drop_duplicates(subset='imdb_id', inplace=True)
except Exception as e:
print(e)
# 7. Write a list comprehension to keep the columns that don't have null values from the wiki_movies_df DataFrame.
wiki_columns_to_keep = [column for column in wiki_movies_df.columns if wiki_movies_df[column]\
.isnull().sum() < len(wiki_movies_df) * 0.9]
wiki_movies_df = wiki_movies_df[wiki_columns_to_keep]
# 8. Create a variable that will hold the non-null values from the “Box office” column.
box_office = wiki_movies_df['Box office'].dropna()
# 9. Convert the box office data created in Step 8 to string values using the lambda and join functions.
box_office = box_office.apply(lambda x: ' '.join(x) if type(x) == list else x)
# 10. Write a regular expression to match the six elements of "form_one" of the box office data.
form_one = r'\$\s*\d+\.?\d*\s*[mb]illi?on'
# 11. Write a regular expression to match the three elements of "form_two" of the box office data.
form_two = r'\$\d{1,3}(?:,\d{3})+'
# 12. Add the parse_dollars function.
def parse_dollars(s):
# if s is not a string, return NaN
if type(s) != str:
return np.nan
# if input is of the form $###.# million
if re.match(r'\$\s*\d+\.?\d*\s*milli?on', s, flags=re.IGNORECASE):
# remove dollar sign and " million"
s = re.sub('\$|\s|[a-zA-Z]','', s)
# convert to float and multiply by a million
value = float(s) * 10**6
# return value
return value
# if input is of the form $###.# billion
elif re.match(r'\$\s*\d+\.?\d*\s*billi?on', s, flags=re.IGNORECASE):
# remove dollar sign and " billion"
s = re.sub('\$|\s|[a-zA-Z]','', s)
# convert to float and multiply by a billion
value = float(s) * 10**9
# return value
return value
# if input is of the form $###,###,###
elif re.match(r'\$\s*\d{1,3}(?:[,\.]\d{3})+(?!\s[mb]illion)', s, flags=re.IGNORECASE):
# remove dollar sign and commas
s = re.sub('\$|,','', s)
# convert to float
value = float(s)
# return value
return value
# otherwise, return NaN
else:
return np.nan
# 13. Clean the box office column in the wiki_movies_df DataFrame.
box_office = box_office.str.replace(r'\$.*[-—–](?![a-z])', '$', regex=True)
box_office.str.extract(f'({form_one}|{form_two})')
wiki_movies_df['box_office'] = box_office.str.extract(f'({form_one}|{form_two})',
flags=re.IGNORECASE)[0].apply(parse_dollars)
wiki_movies_df.drop('Box office', axis=1, inplace=True)
# 14. Clean the budget column in the wiki_movies_df DataFrame.
budget = wiki_movies_df['Budget'].dropna()
budget = budget.map(lambda x: ' '.join(x) if type(x) == list else x)
budget = budget.str.replace(r'\$.*[-—–](?![a-z])', '$', regex=True)
wiki_movies_df['budget'] = budget.str.extract(f'({form_one}|{form_two})',
flags=re.IGNORECASE)[0].apply(parse_dollars)
# 15. Clean the release date column in the wiki_movies_df DataFrame.
release_date = wiki_movies_df['Release date'].dropna().apply(lambda x: ' '.join(x) if type(x) == list else x)
date_form_one = r'(?:January|February|March|April|May|June|July|August|September|October|November|December)\s[123]?\d,\s\d{4}'
date_form_two = r'\d{4}.[01]\d.[0123]\d'
date_form_three = r'(?:January|February|March|April|May|June|July|August|September|October|November|December)\s\d{4}'
date_form_four = r'\d{4}'
release_date.str.extract(f'({date_form_one}|{date_form_two}|{date_form_three}|{date_form_four})', flags=re.IGNORECASE)
wiki_movies_df['release_date'] = pd.to_datetime(release_date.str.extract(f'({date_form_one}|{date_form_two}|{date_form_three}|{date_form_four})')[0], infer_datetime_format=True)
# 16. Clean the running time column in the wiki_movies_df DataFrame.
running_time = wiki_movies_df['Running time'].dropna().apply(lambda x: ' '.join(x) if type(x) == list else x)
running_time.str.contains(r'^\d*\s*minutes$', flags=re.IGNORECASE, na=False).sum()
running_time.str.contains(r'^\d*\s*m', flags=re.IGNORECASE, na=False).sum()
running_time_extract = running_time.str.extract(r'(\d+)\s*ho?u?r?s?\s*(\d*)|(\d+)\s*m')
running_time_extract = running_time_extract.apply(lambda col: pd.to_numeric(col, errors='coerce')).fillna(0)
# Return three variables. The first is the wiki_movies_df DataFrame
return wiki_movies_df, kaggle_metadata, ratings
# 17. Create the path to your file directory and variables for the three files.
file_dir = 'C:/Users/erict/OneDrive/Desktop/Bootcamp Files/Movies ETL'
# The Wikipedia data
wiki_file = f'{file_dir}/wikipedia-movies.json'
# The Kaggle metadata
kaggle_file = f'{file_dir}/movies_metadata.csv'
# The MovieLens rating data.
ratings_file = f'{file_dir}/ratings.csv'
# 18. Set the three variables equal to the function created in D1.
wiki_file, kaggle_file, ratings_file = extract_transform_load()
# 19. Set the wiki_movies_df equal to the wiki_file variable.
wiki_movies_df = wiki_file
# 20. Check that the wiki_movies_df DataFrame looks like this.
wiki_movies_df.head()
# 21. Check that wiki_movies_df DataFrame columns are correct.
wiki_movies_df.columns.to_list()
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
from matplotlib import pyplot as plt
import koala.phase_space as ps
from koala.example_graphs import *
import koala.plotting as pl
from koala.graph_color import color_lattice
from matplotlib import cm
from koala.flux_finder import fluxes_from_bonds,fluxes_to_labels
from numpy import linalg as la
from koala.hamiltonian import generate_majorana_hamiltonian
```
## Here we look at a bunch of finite size effects and try figure out where they come from
### 1: Do system-wide loops actually change the energy?
```
# first we test a few bulk, non-phase spacey system
# parameters
J_vals = np.array([1,1,1])
system_size = 4
# define the lattices
h_comb = generate_honeycomb(system_size)
hex_sq = generate_hex_square_oct(system_size)
lattices = [
h_comb,
h_comb,
hex_sq,
hex_sq,
]
comb_color = color_lattice(h_comb)
hex_sq_color = color_lattice(hex_sq)
colorings = [
comb_color,
comb_color,
hex_sq_color,
hex_sq_color
]
# pick the spins to flip
flip_values = [
[],
[12,15,18,21],
[],
[132,133,134,135],
]
# set ujk
ujk_values = [np.full(lat.n_edges , 1) for lat in lattices]
for a in range(len(ujk_values)):
ujk_values[a][flip_values[a]] = -1
# find energies
energies = []
for n, lattice in enumerate(lattices):
H = generate_majorana_hamiltonian(lattice,colorings[n], ujk_values[n],J_vals)
energy_vals = la.eigvalsh(H)
ground_energy = np.sum(energy_vals*(energy_vals<0))
normalisation = len(energy_vals)/2
energies.append(ground_energy/normalisation)
# plot lattices
fig, ax = plt.subplots(2,2,figsize = (10,10))
ax = ax.flatten()
for n, lattice in enumerate(lattices):
pl.plot_edges(lattice, ax = ax[n],directions = ujk_values[n], labels = ujk_values[n], color_scheme = ['k', 'k', 'blue'] )
# pl.plot_vertex_indices(lattice, ax= ax[n])
# pl.plot_edge_indices(lattice, ax= ax[n])
fluxes = fluxes_from_bonds(lattice,ujk_values[n])
pl.plot_plaquettes(lattice, fluxes_to_labels(fluxes), color_scheme=['white','lightyellow'], ax = ax[n])
ax[n].set_title(f'Energy_per_state = {energies[n]:.4f}')
```
So it looks like adding a system-wide loop does affect things! However now let us use the phase space tricks to sample over the whole brillouin zone!
```
# phase space bastardry begins:
# use the lattices from before but now as unit cells:
k_sampling_number = 50
k_values = np.arange(k_sampling_number)*2*np.pi/k_sampling_number
KX,KY = np.meshgrid(k_values,k_values)
kx = KX.flatten()
ky = KY.flatten()
n_k_states = len(kx)
# find the energy by sampling over the wider k space
k_hamiltonians = [ps.k_hamiltonian_generator(lattices[n], colorings[n], ujk_values[n], J_vals) for n in range(len(lattices))]
energies = []
for H in k_hamiltonians:
energy = 0
for k in zip(kx,ky):
es = la.eigvalsh(H(k))
gs_energy = np.sum(es*(es<0))
normalisation = len(es)/2
energy += gs_energy/normalisation
energies.append(energy/n_k_states)
print(energies)
# plot lattices
fig, ax = plt.subplots(2,2,figsize = (10,10))
ax = ax.flatten()
for n, lattice in enumerate(lattices):
pl.plot_edges(lattice, ax = ax[n],directions = ujk_values[n], labels = ujk_values[n], color_scheme = ['k', 'k', 'blue'] )
# pl.plot_vertex_indices(lattice, ax= ax[n])
# pl.plot_edge_indices(lattice, ax= ax[n])
fluxes = fluxes_from_bonds(lattice,ujk_values[n])
pl.plot_plaquettes(lattice, fluxes_to_labels(fluxes), color_scheme=['white','lightyellow'], ax = ax[n])
ax[n].set_title(f'Energy_per_state = {energies[n]:.4f}')
```
You can see here that, once we average over more points in the brillouin zone by twisting the boundaries, the effect of the system wide spin flips vanishes. The explanation is that adding a system wide spin flip does not actually change the physics - it just adds a -1 to a line crossing the system - equivalent to adding an e^(i pi) phase to the boundary. This shifts the brillouin zone over and the effect is that we end up sampling 5 different points from a slightly different part of the phase space, getting a slightly different value of the energy!
### We can show this by looking at the gap size over phase space:
```
k_vals = np.concatenate([KX[:,:,np.newaxis],KY[:,:,np.newaxis]],axis=2)
gaps = []
for H in k_hamiltonians:
def find_gap(k):
h = H(k)
vals = la.eigvalsh(h)
return np.min(np.abs(vals))
gaps.append(np.apply_along_axis(find_gap,2,k_vals))
fig, axes = plt.subplots(2,2,figsize = (10,10))
for n, ax in enumerate(axes.flatten()):
cax = ax.matshow(gaps[n], cmap = 'gnuplot2_r',vmin = 0)
plt.colorbar(cax,ax=ax)
ax.set_title(f'min gap: {np.min(gaps[n]):.3e}')
```
| github_jupyter |
```
import os
import sys
import importlib
import copy
from collections import defaultdict
sys.path.insert(0, '/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc')
from tools_pattern import get_eucledean_dist
# script_n = os.path.basename(__file__).split('.')[0]
script_n = 'distribution_123share_bouton_210516'
import my_plot
importlib.reload(my_plot)
from my_plot import MyPlotData, my_box_plot
def to_ng_coord(coord):
return (
int(coord[0]/4),
int(coord[1]/4),
int(coord[2]/40),
)
import compress_pickle
# input_graph = compress_pickle.load('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/'\
# 'mf_grc_model/input_graph_201114_restricted_z.gz')
fname = ('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/'\
# 'mf_grc_model/input_graph_210407_xlim_90_140_zlim_0.0_44.0.gz')
'mf_grc_model/input_graph_210407_all.gz')
input_graph = compress_pickle.load(fname)
# z_min = 19800
# z_max = 29800
z_min = 19800
z_max = 29800
# GrCs are fully reconstructed and proofread from 90k to 150k
x_min = 105*1000*4
x_max = 135*1000*4
# radius = 200
n_randoms = 5
replication_hist2 = defaultdict(int)
grc_ids = set()
mf_ids = set()
replicated_2shares = defaultdict(int)
def get_prob(in_graph, unique_count=False, count_within_box=True, return_counted=False):
n_common_pairs = 0
processed = set()
total_n_pairs = 0
hist = defaultdict(int)
n = 0
counted_grcs = 0
for grc_i_id in in_graph.grcs:
n += 1
grc_i = in_graph.grcs[grc_i_id]
x, y, z = grc_i.soma_loc
if count_within_box:
if x < x_min or x > x_max:
continue
if z < z_min or z > z_max:
continue
counted_grcs += 1
grc_ids.add(grc_i_id)
rosettes_i = set([mf[1] for mf in grc_i.edges])
for r in rosettes_i:
mf_ids.add(r)
for grc_j_id in in_graph.grcs:
if grc_i_id == grc_j_id:
continue
if unique_count and (grc_i_id, grc_j_id) in processed:
continue
processed.add((grc_i_id, grc_j_id))
processed.add((grc_j_id, grc_i_id))
grc_j = in_graph.grcs[grc_j_id]
x, y, z = grc_j.soma_loc
# if count_within_box:
# if x < x_min or x > x_max:
# continue
# if z < z_min or z > z_max:
# continue
common_rosettes = set([mf[1] for mf in grc_j.edges])
common_rosettes = common_rosettes & rosettes_i
hist[len(common_rosettes)] += 1
if len(common_rosettes) == 2:
replication_hist2[grc_i_id] += 1
common_rosettes = tuple(sorted(list(common_rosettes)))
replicated_2shares[common_rosettes] += 1
for k in hist:
# fix 0 datapoint plots
if hist[k] == 0:
hist[k] = 1
if return_counted:
return hist, counted_grcs
else:
return hist
input_observed = copy.deepcopy(input_graph)
hist_data, n_grcs = get_prob(input_observed, count_within_box=True, return_counted=True)
print(n_grcs)
print(hist_data)
print(len(mf_ids))
# n_grcs = len(input_graph.grcs)
replication_hist2_list = []
for grc in grc_ids:
if grc in replication_hist2:
replication_hist2_list.append((grc, replication_hist2[grc]))
else:
replication_hist2_list.append((grc, 0))
replication_hist2_list_sorted = sorted(replication_hist2_list, key=lambda x: x[1])
mpd = MyPlotData()
mpd_count = MyPlotData()
i = 0
for grc_id, count in replication_hist2_list_sorted:
mpd_count.add_data_point(
count=count,
grc_id=grc_id,
i=i,
model='Observed',
)
i += 1
importlib.reload(my_plot); my_plot.my_relplot(
mpd_count,
x='i',
y='count',
# kind='hist',
context='paper',
linewidth=2.5,
# kde=True,
# stat='density',
ylim=[0, 18],
height=4,
aspect=2,
y_axis_label='# of 2-share GrC partners',
x_axis_label='Sorted GrCs',
save_filename=f'{script_n}_line.svg',
show=True,
)
# mpd_count_random = compress_pickle.load('distribution_123share_210512_random.gz')
# mpd_count_random = compress_pickle.load('distribution_123share_210512_random_test.gz')
# mpd_count_random2 = compress_pickle.load('distribution_123share_210512_random_test.gz')
mpd_all = MyPlotData()
mpd_all.append(mpd_count)
# mpd_all.append(compress_pickle.load(
# 'distribution_123share_210512_random_22000_5000.gz').add_key_value('model', '22000_5000'))
mpd_all.append(compress_pickle.load(
'sharedist_bouton_210514_random_test_420000_540000_19800_29800.gz').add_key_value('model', 'sphere_23k'))
importlib.reload(my_plot); my_plot.my_relplot(
mpd_all,
x='i',
y='count',
hue='model',
context='paper',
linewidth=2.5,
# kde=True,
# stat='density',
height=4,
aspect=2,
y_axis_label='# of 2-share GrC partners',
x_axis_label='Sorted GrCs',
save_filename=f'{script_n}_line.svg',
show=True,
)
# mpd_count_random = compress_pickle.load('distribution_123share_210512_random.gz')
# mpd_count_random = compress_pickle.load('distribution_123share_210512_random_test.gz')
# mpd_count_random2 = compress_pickle.load('distribution_123share_210512_random_test.gz')
mpd_all = MyPlotData()
mpd_all.append(mpd_count)
# mpd_all.append(compress_pickle.load(
# 'distribution_123share_210512_random_22000_5000.gz').add_key_value('model', '22000_5000'))
mpd_all.append(compress_pickle.load(
'sharedist_bouton_210514_random_sphere_23000_X_420000_540000_Z_19800_29800.gz').add_key_value(
'model', 'sphere_23'))
mpd_all.append(compress_pickle.load(
'sharedist_bouton_210514_random_sphere_15000_X_420000_540000_Z_19800_29800.gz').add_key_value(
'model', 'sphere_15'))
mpd_all.append(compress_pickle.load(
'sharedist_bouton_210514_random_sphere_30000_X_420000_540000_Z_19800_29800.gz').add_key_value(
'model', 'sphere_30'))
importlib.reload(my_plot); my_plot.my_relplot(
mpd_all,
x='i',
y='count',
hue='model',
context='paper',
linewidth=2.5,
# kde=True,
# stat='density',
height=4,
aspect=2,
y_axis_label='# of 2-share GrC partners',
x_axis_label='Sorted GrCs',
save_filename=f'{script_n}_line.svg',
show=True,
)
# mpd_count_random = compress_pickle.load('distribution_123share_210512_random.gz')
# mpd_count_random = compress_pickle.load('distribution_123share_210512_random_test.gz')
# mpd_count_random2 = compress_pickle.load('distribution_123share_210512_random_test.gz')
mpd_all = MyPlotData()
mpd_all.append(mpd_count)
# mpd_all.append(compress_pickle.load('distribution_123share_210512_random_22000_2500.gz'))
# mpd_all.append(compress_pickle.load(
# 'distribution_123share_210512_random_22000_5000.gz').add_key_value('model', '22000_5000'))
mpd_all.append(compress_pickle.load(
'distribution_123share_bouton_210516_random_circle_X_420000_540000_Z_19800_29800.gz').add_key_value(
'model', 'circle'))
# mpd_all.append(compress_pickle.load(
# 'distribution_123share_210512_random_22000_2500.gz').add_key_value('model', '22000_2500'))
# mpd_all.append(compress_pickle.load(
# 'distribution_123share_210512_random_edges.gz').add_key_value('model', 'edges'))
mpd_all.append(compress_pickle.load(
'distribution_123share_bouton_210516_random_edge_X_420000_540000_Z_19800_29800.gz').add_key_value(
'model', 'edge'))
mpd_all.append(compress_pickle.load(
'distribution_123share_bouton_210516_random_circle_dist_X_420000_540000_Z_19800_29800.gz').add_key_value(
'model', 'circle_dist'))
importlib.reload(my_plot); my_plot.my_relplot(
mpd_all,
x='i',
y='count',
hue='model',
context='paper',
linewidth=2.5,
# kde=True,
# stat='density',
height=4,
aspect=2,
y_axis_label='# of 2-share GrC partners',
x_axis_label='Sorted GrCs',
save_filename=f'{script_n}_line.svg',
show=True,
)
mpd_all = MyPlotData()
mpd_all.append(mpd_count)
mpd_all.append(compress_pickle.load(
'distribution_123share_210512_random_edges.gz').add_key_value('model', 'edges'))
mpd_all.append(compress_pickle.load(
'distribution_123share_210512_random_edges_10000.gz').add_key_value('model', 'edges_10k'))
importlib.reload(my_plot); my_plot.my_relplot(
mpd_all,
x='i',
y='count',
hue='model',
context='paper',
linewidth=2.5,
# kde=True,
# stat='density',
height=4,
aspect=2,
y_axis_label='# of 2-share GrC partners',
x_axis_label='Sorted GrCs',
save_filename=f'{script_n}_line.svg',
show=True,
)
for grc, count in replication_hist2_list_sorted:
print((grc, count))
print([k[0] for k in input_graph.grcs[grc].edges])
replicated_2shares_list = [(k, v) for k, v in replicated_2shares.items()]
replicated_2shares_list.sort(key=lambda x: x[1])
for i in replicated_2shares_list:
print(i)
import tools_mf_graph
importlib.reload(tools_mf_graph)
from tools_mf_graph import GCLGraph
fname = ('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/'\
'mf_grc_model/input_graph_210407_xlim_90_140_zlim_0.0_44.0.gz')
input_graph = compress_pickle.load(fname)
input_graph.avg_edge_count_by_depth = None
n_randoms = 5
hist_random_21 = []
for n in range(n_randoms):
# input_graph.randomize_graph(random_model=True)
input_graph.randomize_graph_by_grc2(
single_connection_per_pair=True,
constant_grc_degree='depth',
constant_dendrite_length=21000,
# always_pick_closest_rosette=True,
)
hist_random = get_prob(input_graph)
hist_random_21.append(hist_random)
n_randoms = 5
hist_random_17 = []
for n in range(n_randoms):
# input_graph.randomize_graph(random_model=True)
input_graph.randomize_graph_by_grc2(
single_connection_per_pair=True,
constant_grc_degree='depth',
constant_dendrite_length=21000,
# always_pick_closest_rosette=True,
)
hist_random = get_prob(input_graph)
hist_random_17.append(hist_random)
import tools_mf_graph
importlib.reload(tools_mf_graph)
from tools_mf_graph import GCLGraph
fname = ('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/'\
'mf_grc_model/input_graph_210407_xlim_90_140_zlim_0.0_44.0.gz')
input_graph = compress_pickle.load(fname)
input_graph.avg_edge_count_by_depth = None
input_graph.avg_dendrite_len = None
n_randoms = 5
hist_random_avg = []
for n in range(n_randoms):
# input_graph.randomize_graph(random_model=True)
input_graph.randomize_graph_by_grc2(
single_connection_per_pair=True,
constant_grc_degree='depth',
constant_dendrite_length=True,
# always_pick_closest_rosette=True,
)
hist = get_prob(input_graph)
hist_random_avg.append(hist)
n_randoms = 5
hist_shuffle = []
for n in range(n_randoms):
input_graph.randomize_graph_by_grc(
mf_dist_margin=4000,
single_connection_per_pair=True,
)
hist = get_prob(input_graph)
hist_shuffle.append(hist)
global_random_n_grcs, hist_global_random = compress_pickle.load('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/mf_grc_analysis/share_distribution/gen_global_random_7k_204k_data_2000.gz')
import compress_pickle
compress_pickle.dump((
hist_data,
hist_random,
global_random_n_grcs,
hist_global_random,
), f"{script_n}_data.gz")
# normalize
# total_n_pairs = hist_data[0] + hist_data[1] + hist_data[2] + hist_data[3]
# global_random_n_grcs = 204000
mpd_data = MyPlotData()
for n_share in [1, 2, 3]:
if n_share in hist_data:
mpd_data.add_data_point(
n_share=n_share,
count=hist_data[n_share]/n_grcs,
type='Observation',
)
for hist_random in hist_random_avg:
if n_share in hist_random:
mpd_data.add_data_point(
n_share=n_share,
count=hist_random[n_share]/n_grcs,
type='Local Random',
)
if n_share in hist_global_random:
mpd_data.add_data_point(
n_share=n_share,
# need to divide by 2 because we're sampling only 2/200 grcs
# (or multiply others by 2)
# count=hist_global_random[n_share]/global_random_n_grcs/2,
count=hist_global_random[n_share]/global_random_n_grcs,
type='Global Random',
)
def custom_legend_fn(plt):
# plt.legend(bbox_to_anchor=(1.025, .8), loc='upper left', borderaxespad=0.)
plt.legend(loc='upper right', frameon=False, markerscale=2, prop={'size': 8})
# importlib.reload(my_plot); my_plot.my_relplot(
# mpd_data,
# x='n_share',
# y='count',
# hue='type',
# # hue_order=['Data', 'Random Model'],
# kind='scatter',
# context='paper',
# # ylim=[.005, 50],
# ylim=[.005, 2000],
# xlim=[.7, 3.3],
# log_scale_y=True,
# s=150,
# # xticklabels=['', 1, '', 2, '', 3, ''],
# xticks=[1, 2, 3],
# height=4,
# aspect=1.1,
# custom_legend_fn=custom_legend_fn,
# y_axis_label='GrC pairs / # GrCs',
# x_axis_label='Shared Inputs',
# # save_filename=f'{script_n}.pdf',
# save_filename=f'{script_n}.svg',
# show=True,
# )
importlib.reload(my_plot); my_plot.my_relplot(
mpd_data,
x='n_share',
y='count',
hue='type',
# hue_order=['Data', 'Random Model'],
kind='line',
err_style="bars",
ci=68,
markers=True,
dashes=False,
# s=150,
context='paper',
# ylim=[.005, 50],
ylim=[.005, 2000],
xlim=[.9, 3.1],
log_scale_y=True,
linewidth=2.5,
# s=150,
# xticklabels=['', 1, '', 2, '', 3, ''],
xticks=[1, 2, 3],
# height=4,
width=3,
aspect=1.4,
font_scale=1,
custom_legend_fn=custom_legend_fn,
y_axis_label='GrC pairs / # GrCs',
x_axis_label='Shared MF inputs',
# save_filename=f'{script_n}.pdf',
save_filename=f'{script_n}_line.svg',
show=True,
)
n_grcs=142
mpd_data = MyPlotData()
for n_share in [1, 2, 3]:
if n_share in hist_data:
mpd_data.add_data_point(
n_share=n_share,
count=hist_data[n_share]/n_grcs,
type='Observation',
)
# for hist_random in hist_random_17:
# if n_share in hist_random:
# mpd_data.add_data_point(
# n_share=n_share,
# count=hist_random[n_share]/n_grcs,
# type='Local Random',
# )
for hist_random in hist_random_avg:
if n_share in hist_random:
mpd_data.add_data_point(
n_share=n_share,
count=hist_random[n_share]/n_grcs,
type='Local Random',
)
# for hist_random in hist_shuffle:
# if n_share in hist_random:
# mpd_data.add_data_point(
# n_share=n_share,
# count=hist_random[n_share]/n_grcs,
# type='Shuffle',
# )
palette = {
'Observation': 'black',
'Local Random': '#00a79d',
}
def custom_legend_fn(plt):
# plt.legend(bbox_to_anchor=(1.025, .8), loc='upper left', borderaxespad=0.)
plt.legend(loc='upper right', frameon=False, markerscale=2, prop={'size': 11})
importlib.reload(my_plot); my_plot.my_relplot(
mpd_data,
x='n_share',
y='count',
hue='type',
# hue_order=['Data', 'Random Model'],
kind='line',
err_style="bars",
# ci=68,
markers=True,
dashes=False,
# s=150,
context='paper',
# ylim=[.005, 50],
# ylim=[None, 100],
# ylim=[None, 20],
xlim=[.7, 3.3],
linewidth=1.5,
palette=palette,
# log_scale_y=True,
# s=150,
# xticklabels=['', 1, '', 2, '', 3, ''],
xticks=[1, 2, 3],
# width=3.1,
# height=3,
width=3,
aspect=1.4,
font_scale=1.2,
# aspect=1.1,
custom_legend_fn=custom_legend_fn,
# y_axis_label='GrC pairs / # GrCs',
y_axis_label='Avg. pairs per GrC',
x_axis_label='Shared MF inputs',
# save_filename=f'{script_n}.pdf',
save_filename=f'{script_n}_line_linear.svg',
show=True,
)
n_grcs=142
mpd_data = MyPlotData()
for n_share in [1, 2, 3]:
if n_share in hist_data:
mpd_data.add_data_point(
n_share=n_share,
count=hist_data[n_share]/n_grcs,
type='Observation',
)
# for hist_random in hist_random_17:
# if n_share in hist_random:
# mpd_data.add_data_point(
# n_share=n_share,
# count=hist_random[n_share]/n_grcs,
# type='Local Random',
# )
for hist_random in hist_random_avg:
if n_share in hist_random:
mpd_data.add_data_point(
n_share=n_share,
count=hist_random[n_share]/n_grcs,
type='Local Random',
)
# for hist_random in hist_shuffle:
# if n_share in hist_random:
# mpd_data.add_data_point(
# n_share=n_share,
# count=hist_random[n_share]/n_grcs,
# type='Shuffle',
# )
if n_share in hist_global_random:
mpd_data.add_data_point(
n_share=n_share,
# need to divide by 2 because we're sampling only 2/200 grcs
# (or multiply others by 2)
# count=hist_global_random[n_share]/global_random_n_grcs/2,
count=hist_global_random[n_share]/global_random_n_grcs,
type='Global Random',
)
palette = {
'Observation': 'black',
'Local Random': '#00a79d',
'Global Random': '#92278f',
}
def custom_legend_fn(plt):
# plt.legend(bbox_to_anchor=(1.025, .8), loc='upper left', borderaxespad=0.)
plt.legend(loc='upper right', frameon=False, markerscale=2, prop={'size': 11})
importlib.reload(my_plot); my_plot.my_relplot(
mpd_data,
x='n_share',
y='count',
hue='type',
# hue_order=['Data', 'Random Model'],
kind='line',
err_style="bars",
# ci=68,
markers=True,
dashes=False,
# s=150,
context='paper',
# ylim=[.005, 50],
# ylim=[None, 100],
# ylim=[None, 20],
xlim=[.7, 3.3],
linewidth=1.5,
palette=palette,
log_scale_y=True,
# s=150,
# xticklabels=['', 1, '', 2, '', 3, ''],
xticks=[1, 2, 3],
# width=3.1,
# height=3,
width=3,
aspect=1.4,
font_scale=1.2,
# aspect=1.1,
# custom_legend_fn=custom_legend_fn,
# y_axis_label='GrC pairs / # GrCs',
y_axis_label='Avg. pairs per GrC',
x_axis_label='Shared MF inputs',
# save_filename=f'{script_n}.pdf',
save_filename=f'{script_n}_line_log.svg',
show=True,
)
n_grcs=142
print(hist_data)
print(hist_random_17[0])
print(hist_data[2]/n_grcs)
print(hist_random_17[2][2]/n_grcs)
print(hist_random_avg[2][2]/n_grcs)
print(hist_shuffle[2][2]/n_grcs)
print()
print(hist_data[1]/n_grcs)
print(hist_random_17[2][1]/n_grcs)
print(hist_random_avg[2][1]/n_grcs)
print(hist_shuffle[2][1]/n_grcs)
```
| github_jupyter |
# 2D Advection-Diffusion equation
in this notebook we provide a simple example of the DeepMoD algorithm and apply it on the 2D advection-diffusion equation.
```
# General imports
import numpy as np
import torch
# DeepMoD functions
from deepymod import DeepMoD
from deepymod.model.func_approx import NN
from deepymod.model.library import Library2D_third
from deepymod.model.constraint import LeastSquares
from deepymod.model.sparse_estimators import Threshold,PDEFIND
from deepymod.training import train
from deepymod.training.sparsity_scheduler import TrainTestPeriodic
from scipy.io import loadmat
# Settings for reproducibility
np.random.seed(1)
torch.manual_seed(1)
if torch.cuda.is_available():
device = 'cuda'
else:
device = 'cpu'
```
## Prepare the data
Next, we prepare the dataset.
```
data = loadmat('Diffusion_2D_space41.mat')
data = np.real(data['Expression1']).reshape((41,41,41,4))[:,:,:,3]
x_dim, y_dim, t_dim = data.shape
time_range = [1,2,4,6,8,10,12,14]
for i in time_range:
# Downsample data and prepare data without noise:
down_data= np.take(np.take(np.take(data,np.arange(0,x_dim,3),axis=0),np.arange(0,y_dim,3),axis=1),np.arange(0,t_dim,i),axis=2)
print("Dowmsampled shape:",down_data.shape, "Total number of data points:", np.product(down_data.shape))
index = len(np.arange(0,t_dim,i))
width, width_2, steps = down_data.shape
x_arr, y_arr, t_arr = np.linspace(0,1,width), np.linspace(0,1,width_2), np.linspace(0,1,steps)
x_grid, y_grid, t_grid = np.meshgrid(x_arr, y_arr, t_arr, indexing='ij')
X, y = np.transpose((t_grid.flatten(), x_grid.flatten(), y_grid.flatten())), np.float32(down_data.reshape((down_data.size, 1)))
# Add noise
noise_level = 0.20
y_noisy = y + noise_level * np.std(y) * np.random.randn(y.size, 1)
# Randomize data
idx = np.random.permutation(y.shape[0])
X_train = torch.tensor(X[idx, :], dtype=torch.float32, requires_grad=True).to(device)
y_train = torch.tensor(y_noisy[idx, :], dtype=torch.float32).to(device)
# Configure DeepMoD
network = NN(3, [40, 40, 40, 40], 1)
library = Library2D_third(poly_order=0)
estimator = Threshold(0.05)
sparsity_scheduler = TrainTestPeriodic(periodicity=50, patience=200, delta=1e-5)
constraint = LeastSquares()
model = DeepMoD(network, library, estimator, constraint).to(device)
optimizer = torch.optim.Adam(model.parameters(), betas=(0.99, 0.99), amsgrad=True, lr=2e-3)
logdir='final_runs/20_noise_x14/'+str(index)+'/'
train(model, X_train, y_train, optimizer,sparsity_scheduler, log_dir=logdir, split=0.8, max_iterations=50000, delta=1e-6, patience=200)
```
| github_jupyter |
```
#!/usr/bin/python
# interpolate scalar gradient onto nedelec space
import petsc4py
import sys
petsc4py.init(sys.argv)
from petsc4py import PETSc
from dolfin import *
# from MatrixOperations import *
import numpy as np
import PETScIO as IO
import common
import scipy
import scipy.io
import time
import scipy.sparse as sp
import BiLinear as forms
import IterOperations as Iter
import MatrixOperations as MO
import CheckPetsc4py as CP
import Solver as S
import MHDmatrixPrecondSetup as PrecondSetup
import NSprecondSetup
import MHDprec as MHDpreconditioner
import gc
import MHDmulti
import MHDmatrixSetup as MHDsetup
import HartmanChannel
import ExactSol
# import matplotlib.pyplot as plt
#@profile
m = 2
def PETScToScipy(A):
data = A.getValuesCSR()
sparseSubMat = sp.csr_matrix(data[::-1], shape=A.size)
return sparseSubMat
def savePETScMat(A, name1, name2):
A_ = PETScToScipy(A)
scipy.io.savemat(name1, mdict={name2: A_})
set_log_active(False)
errL2u = np.zeros((m-1, 1))
errH1u = np.zeros((m-1, 1))
errL2p = np.zeros((m-1, 1))
errL2b = np.zeros((m-1, 1))
errCurlb = np.zeros((m-1, 1))
errL2r = np.zeros((m-1, 1))
errH1r = np.zeros((m-1, 1))
l2uorder = np.zeros((m-1, 1))
H1uorder = np.zeros((m-1, 1))
l2porder = np.zeros((m-1, 1))
l2border = np.zeros((m-1, 1))
Curlborder = np.zeros((m-1, 1))
l2rorder = np.zeros((m-1, 1))
H1rorder = np.zeros((m-1, 1))
NN = np.zeros((m-1, 1))
DoF = np.zeros((m-1, 1))
Velocitydim = np.zeros((m-1, 1))
Magneticdim = np.zeros((m-1, 1))
Pressuredim = np.zeros((m-1, 1))
Lagrangedim = np.zeros((m-1, 1))
Wdim = np.zeros((m-1, 1))
iterations = np.zeros((m-1, 1))
SolTime = np.zeros((m-1, 1))
udiv = np.zeros((m-1, 1))
MU = np.zeros((m-1, 1))
level = np.zeros((m-1, 1))
NSave = np.zeros((m-1, 1))
Mave = np.zeros((m-1, 1))
TotalTime = np.zeros((m-1, 1))
DimSave = np.zeros((m-1, 4))
dim = 2
ShowResultPlots = 'yes'
MU[0] = 1e0
for xx in xrange(1, m):
print xx
level[xx-1] = xx + 0
nn = 2**(level[xx-1])
# Create mesh and define function space
nn = int(nn)
NN[xx-1] = nn/2
L = 10.
y0 = 2.
z0 = 1.
# mesh, boundaries, domains = HartmanChannel.Domain(nn)
mesh = UnitSquareMesh(nn, nn)
parameters['form_compiler']['quadrature_degree'] = -1
order = 2
parameters['reorder_dofs_serial'] = False
Velocity = VectorElement("CG", mesh.ufl_cell(), order)
Pressure = FiniteElement("CG", mesh.ufl_cell(), order-1)
Magnetic = FiniteElement("N1curl", mesh.ufl_cell(), order-1)
Lagrange = FiniteElement("CG", mesh.ufl_cell(), order-1)
VelocityF = VectorFunctionSpace(mesh, "CG", order)
PressureF = FunctionSpace(mesh, "CG", order-1)
MagneticF = FunctionSpace(mesh, "N1curl", order-1)
LagrangeF = FunctionSpace(mesh, "CG", order-1)
W = FunctionSpace(mesh, MixedElement(
[Velocity, Pressure, Magnetic, Lagrange]))
Velocitydim[xx-1] = W.sub(0).dim()
Pressuredim[xx-1] = W.sub(1).dim()
Magneticdim[xx-1] = W.sub(2).dim()
Lagrangedim[xx-1] = W.sub(3).dim()
Wdim[xx-1] = W.dim()
print "\n\nW: ", Wdim[xx-1], "Velocity: ", Velocitydim[xx-1], "Pressure: ", Pressuredim[xx-1], "Magnetic: ", Magneticdim[xx-1], "Lagrange: ", Lagrangedim[xx-1], "\n\n"
dim = [W.sub(0).dim(), W.sub(1).dim(), W.sub(2).dim(), W.sub(3).dim()]
def boundary(x, on_boundary):
return on_boundary
FSpaces = [VelocityF, PressureF, MagneticF, LagrangeF]
DimSave[xx-1, :] = np.array(dim)
kappa = 1.0
Mu_m = 10.0
MU = 1.0
N = FacetNormal(mesh)
IterType = 'Full'
params = [kappa, Mu_m, MU]
n = FacetNormal(mesh)
u0, p0, b0, r0, Laplacian, Advection, gradPres, CurlCurl, gradR, NS_Couple, M_Couple = ExactSol.MHD2D(
4, 1)
MO.PrintStr("Seting up initial guess matricies", 2, "=", "\n\n", "\n")
BCtime = time.time()
BC = MHDsetup.BoundaryIndices(mesh)
MO.StrTimePrint("BC index function, time: ", time.time()-BCtime)
Hiptmairtol = 1e-6
HiptmairMatrices = PrecondSetup.MagneticSetup(
mesh, Magnetic, Lagrange, b0, r0, Hiptmairtol, params)
MO.PrintStr("Setting up MHD initial guess", 5, "+", "\n\n", "\n\n")
F_NS = -MU*Laplacian + Advection + gradPres - kappa*NS_Couple
if kappa == 0.0:
F_M = Mu_m*CurlCurl + gradR - kappa*M_Couple
else:
F_M = Mu_m*kappa*CurlCurl + gradR - kappa*M_Couple
du = TrialFunction(W)
(v, q, c, s) = TestFunctions(W)
u, p, b, r = split(du)
U = Function(W)
U.vector()[:] = 1.
u_k, p_k, b_k, r_k = split(U)
if kappa == 0.0:
m11 = params[1]*inner(curl(b), curl(c))*dx
else:
m11 = params[1]*params[0]*inner(curl(b), curl(c))*dx
m21 = inner(c, grad(r))*dx
m12 = inner(b, grad(s))*dx
a11 = params[2]*inner(grad(v), grad(u))*dx + inner((grad(u)*u_k), v)*dx + (
1./2)*div(u_k)*inner(u, v)*dx - (1./2)*inner(u_k, n)*inner(u, v)*ds
a12 = -div(v)*p*dx
a21 = -div(u)*q*dx
CoupleT = params[0]*(v[0]*b_k[1]-v[1]*b_k[0])*curl(b)*dx
Couple = -params[0]*(u[0]*b_k[1]-u[1]*b_k[0])*curl(c)*dx
Ftilde = inner((grad(u_k)*u), v)*dx + (1./2)*div(u) * \
inner(u_k, v)*dx - (1./2)*inner(u, n)*inner(u_k, v)*ds
Mtilde = -params[0]*(u_k[0]*b[1]-u_k[1]*b[0])*curl(c)*dx
Ctilde = params[0]*(v[0]*b[1]-v[1]*b[0])*curl(b_k)*dx
a = m11 + m12 + m21 + a11 + a21 + a12 + \
Couple + CoupleT + Ftilde + Mtilde + Ctilde
aa = m11 + m12 + m21 + a11 + a21 + a12 + Couple + CoupleT
if kappa == 0.0:
m11 = params[1]*inner(curl(b_k), curl(c))*dx
else:
m11 = params[1]*params[0]*inner(curl(b_k), curl(c))*dx
m21 = inner(c, grad(r_k))*dx
m12 = inner(b_k, grad(s))*dx
a11 = params[2]*inner(grad(v), grad(u_k))*dx + inner((grad(u_k)*u_k), v)*dx + (
1./2)*div(u_k)*inner(u_k, v)*dx - (1./2)*inner(u_k, n)*inner(u_k, v)*ds
a12 = -div(v)*p_k*dx
a21 = -div(u_k)*q*dx
CoupleT = params[0]*(v[0]*b_k[1]-v[1]*b_k[0])*curl(b_k)*dx
Couple = -params[0]*(u_k[0]*b_k[1]-u_k[1]*b_k[0])*curl(c)*dx
Lns = inner(v, F_NS)*dx
Lmaxwell = inner(c, F_M)*dx
L = Lns + Lmaxwell - (m11 + m12 + m21 + a11 + a21 + a12 + Couple + CoupleT)
J = derivative(L, U)
A, b = assemble_system(a, L)
A, b = CP.Assemble(A, b)
J = assemble(J)
J = CP.Assemble(J)
savePETScMat(J, "J", "J")
savePETScMat(A, "A", "A")
# print J
# J = assemble(J)
# J = CP.Assemble(J)
# x = Iter.u_prev(u_k, p_k, b_k, r_k)
# KSPlinearfluids, MatrixLinearFluids = PrecondSetup.FluidLinearSetup(
# PressureF, MU, mesh)
# kspFp, Fp = PrecondSetup.FluidNonLinearSetup(PressureF, MU, u_k, mesh)
# F = Lns + Lmaxwell - aa
# Hiptmairtol = 1e-4
# HiptmairMatrices = PrecondSetup.MagneticSetup(
# mesh, Magnetic, Lagrange, b0, r0, Hiptmairtol, params)
# IS = MO.IndexSet(W, 'Blocks')
# ones = Function(PressureF)
# ones.vector()[:] = (0*ones.vector().array()+1)
# eps = 1.0 # error measure ||u-u_k||
# tol = 1.0E-4 # tolerance
# iter = 0 # iteration counter
# maxiter = 1 # max no of iterations allowed
# SolutionTime = 0
# outer = 0
# # parameters['linear_algebra_backend'] = 'uBLAS'
# u_is = PETSc.IS().createGeneral(W.sub(0).dofmap().dofs())
# p_is = PETSc.IS().createGeneral(W.sub(1).dofmap().dofs())
# b_is = PETSc.IS().createGeneral(W.sub(2).dofmap().dofs())
# r_is = PETSc.IS().createGeneral(W.sub(3).dofmap().dofs())
# NS_is = PETSc.IS().createGeneral(range(VelocityF.dim()+PressureF.dim()))
# M_is = PETSc.IS().createGeneral(range(VelocityF.dim()+PressureF.dim(), W.dim()))
# bcu = DirichletBC(W.sub(0), Expression(("0.0", "0.0"), degree=4), boundary)
# bcb = DirichletBC(W.sub(2), Expression(("0.0", "0.0"), degree=4), boundary)
# bcr = DirichletBC(W.sub(3), Expression(("0.0"), degree=4), boundary)
# bcs = [bcu, bcb, bcr]
# U = Function(W) # the most recently computed solution
# F = action(F, U)
# # print assemble(dolfin.Jacobian(F))
# # OuterTol = 1e-5
# # InnerTol = 1e-5
# # NSits = 0
# # Mits = 0
# # TotalStart = time.time()
# # SolutionTime = 0
# # errors = np.array([])
# # bcu1 = DirichletBC(VelocityF, Expression(
# # ("0.0", "0.0"), degree=4), boundary)
# # U = x
# # while eps > tol and iter < maxiter:
# # iter += 1
# # MO.PrintStr("Iter "+str(iter), 7, "=", "\n\n", "\n\n")
# # A, b = assemble_system(aa, L)
# # A, b = CP.Assemble(A, b)
# # savePETScMat(J, "J", "J")
# # savePETScMat(A, "A", "A")
# # ss
J = assemble(J)
J = CP.Assemble(J)
savePETScMat(J, "J", "J")
savePETScMat(A, "A", "A")
print problem.jacobian_form()
solve(problem)
form = problem.jacobian_form()
Fw = action(F,U);
assemble(Fw)
problem.has_jacobian()
```
| github_jupyter |
#### 1 - 3 summarized below:
#### Lineraly Seperable Experiment
- **Training data:** X training points were randomly generated (values bounded between -100 and 100). Y training labels were generated by applying a randomly generated target function to the X training points.
- **Test data:** X test points were randomly generated (values bounded between -100 and 100). Y test labels were generated by applying the same target function to the X test points.
#### Non-lineraly Separable Experiment
- **Training data:** X training points were randomly generated (values bounded between -100 and 100). Y training labels randomly generated (-1 and 1). Then, the randomly generated target function was applied with a probaility of .75 to create 'somewhat' lineraly separable data.
- **Test data:** X test points were randomly generated (values bounded between -100 and 100). Y test labels randomly generated (-1 and 1). Then, the randomly generated target function was applied with a probaility of .75 to create 'somewhat' lineraly separable data.
**4.** The initial choice of the weights is random.
#### Answers to questions 5 - 8 can be seen in the statistics (and graphs) on pages 3-4.
#### Variation Results
1. The weights that give the lowest in-sample error rate is best.
2. The step size correlates with the amount the vector changes. i.e., A larger step size makes the vector adjustment larger.
3. It is best to consider training points that reduce the error rate the most first.
```
%matplotlib inline
import numpy as np
import random
from perceptron_learning import Perceptron
from perceptron_learning import two_d_vector as tdv
def main():
bound = 100 # the value that the x and y values are bounded by
num_pts = 80
num_train_pts = 50
perceptron = Perceptron(alpha=0.005)
target_fn = np.random.uniform(-10, 10, 3)
x = get_random_x(num_pts, bound)
x_train, x_test = x[:num_train_pts, :], x[num_train_pts:, :]
y_test = np.sign(np.dot(x_test, target_fn))
print('---------- Linearly Separable Data ----------')
perceptron.fit(x_train, target_fn=target_fn)
predictions = perceptron.predict(x_test)
print('{:28s}: y = {:.2f}x + {:.2f}'.format('Target Function',
tdv.get_slope(target_fn),
tdv.get_y_intercept(target_fn)))
print_error(predictions, y_test)
print()
y = get_y(x[:, 1:], target_fn)
y_train, y_test = y[:num_train_pts], y[num_train_pts:]
print('-------- Non-Linearly Separable Data --------')
perceptron.fit(x_train, y_train=y_train)
predictions = perceptron.predict(x_test)
print_error(predictions, y_test)
perceptron.visualize_training()
def print_error(predictions, y_test):
error = np.sum(np.not_equal(predictions, y_test)) / y_test.shape[0]
print('{0:28s}: {1:.2f}%'.format('Out of Sample (Test) Error', error * 100))
def get_y(training_pts, w_target):
# Have y be somewhat linearly separable
y = np.random.choice([-1, 1], training_pts.shape[0])
for i, pt in enumerate(training_pts):
pct_chance = .75
pt_above_line = tdv.pt_above_line(pt, w_target)
if pt_above_line and random.random() < pct_chance:
y[i] = 1
if not pt_above_line and random.random() < pct_chance:
y[i] = -1
return y
def get_random_x(num_points, bound):
pts = get_random_pts(num_points, bound)
x = np.insert(pts, 0, 1, axis=1) # Let x0 equal 1
return x
def get_random_pts(num_points, bound):
return np.random.randint(-bound, bound, size=(num_points, 2))
if __name__ == '__main__':
main()
"""
two_d_vector.py
Functions that operate on 2d vectors.
w0 (or x0) is a bias "dummy" weight,
so even though the vector is 3 dimensional,
we call it a 2 dimensional vector.
"""
import numpy as np
from random import uniform
def get_perpendicular_vector(w):
# Two lines are perpendicular if: m1 * m2 = -1.
# The two slopes must be negative reciprocals of each other.
m1 = get_slope(w)
m2 = -1 / m1
# m2 = - w[1] / w[2]
random_num = uniform(0, 10)
return np.array([uniform(0, 10), -1 * m2 * random_num, random_num])
def get_line(w, x_bound):
x_range = np.array(range(-x_bound, x_bound))
# Formula for line is: w1x1 + w2x2 + w0 = 0
# we let x2 = y, and x1 = x, then solve for y = mx + b
slope = get_slope(w)
y_intercept = get_y_intercept(w)
y_line = (slope * x_range) + y_intercept
return x_range, y_line
def pt_above_line(pt, w):
return pt[1] > get_slope(w) * pt[0] + get_y_intercept(w)
def get_y_intercept(w):
return - w[0] / w[2]
def get_slope(w):
return - w[1] / w[2]
"""
DataVisualizer.py
"""
import numpy as np
import matplotlib.pyplot as plt
from . import two_d_vector as tdv
class DataVisualizer:
def __init__(self, title, subtitle, x_bound, y_bound):
plt.style.use('seaborn-whitegrid')
self.fig, self.ax = plt.subplots()
self.title = title
self.subtitle = subtitle
self.x_bound = x_bound
self.y_bound = y_bound
def setup_axes(self):
self.ax.cla()
self.fig.canvas.set_window_title(self.subtitle)
self.fig.suptitle(self.title, fontsize=18)
self.ax.set_title(self.subtitle, fontsize=14)
self.ax.set_xlim(-self.x_bound, self.x_bound)
self.ax.set_ylim(-self.y_bound, self.y_bound)
@staticmethod
def red_pts_above_line(pts, w_target, true_classes):
pt_above_line = tdv.pt_above_line(pts[0, :], w_target)
pt_is_positive_class = true_classes[0] > 0
if pt_above_line and pt_is_positive_class:
# positive pt above line
return True
if not pt_above_line and not pt_is_positive_class:
# negative pt below line
return True
return False
def plot_hypothesis(self, pts, true_classes, w_hypothesis, w_target=None):
self.setup_axes()
self.ax.scatter(x=pts[:, 0], y=pts[:, 1], marker='x',
color=['r' if sign >= 0 else 'b' for sign in true_classes])
if w_target is not None:
x, y = tdv.get_line(w_target, self.x_bound)
self.ax.plot(x, y, label='target', color='m')
x, y = tdv.get_line(w_hypothesis, self.x_bound)
self.ax.plot(x, y, label='hypothesis', color='g')
if w_target is not None:
if self.red_pts_above_line(pts, w_target, true_classes):
self.ax.fill_between(x, y, np.full((1,), self.y_bound), color=(1, 0, 0, 0.15))
self.ax.fill_between(x, y, np.full((1,), -self.y_bound), color=(0, 0, 1, 0.15))
else:
self.ax.fill_between(x, y, np.full((1,), self.y_bound), color=(0, 0, 1, 0.15))
self.ax.fill_between(x, y, np.full((1,), -self.y_bound), color=(1, 0, 0, 0.15))
self.ax.legend(facecolor='w', fancybox=True, frameon=True, edgecolor='black', borderpad=1)
# plt.pause(0.01)
@staticmethod
def visualize():
plt.show()
"""
Logger.py
"""
class Logger:
def __init__(self):
self.num_iterations = 0
self.num_vector_updates = 0
def print_statistics(self):
print('{:28s}: {:}'.format('Number of iterations', self.num_iterations))
print('{:28s}: {:}'.format('Number of vector updates', self.num_vector_updates))
"""
Perceptron.py
"""
import numpy as np
from . import two_d_vector as tdv
from . import DataVisualizer, Logger
class Perceptron:
"""Uses 'pocket' algorithm to keep best hypothesis in it's 'pocket'"""
def __init__(self, alpha):
self.alpha = alpha
self.best_hypothesis = np.random.uniform(-10, 10, 3)
self.lowest_error = float('inf')
self.logger = Logger()
self.dv = None
def fit(self, x_train, y_train=None, target_fn=None):
"""Fits the model to the training data (class labels) or target function.
:param x_train: the training data
:param y_train: will be passed in in the non-linearly separable case
:param target_fn: will be passed in in the linearly separable case
:return: None
"""
self.best_hypothesis = np.random.uniform(-10, 10, 3)
self.lowest_error = float('inf')
self.logger = Logger()
self.dv = get_data_visualizer(target_fn, x_train)
if target_fn is not None:
y_train = np.sign(np.dot(x_train, target_fn))
self.best_hypothesis = tdv.get_perpendicular_vector(target_fn)
pts = x_train[:, 1:]
hypothesis = self.best_hypothesis
misclassified_pts = predict_and_evaluate(hypothesis, x_train, y_train)
while self.logger.num_vector_updates < 100000 and np.sum(misclassified_pts) > 0:
for i, misclassified_pt in enumerate(np.nditer(misclassified_pts)):
if misclassified_pt:
# update rule: w(t + 1) = w(t) + y(t) * x(t) * alpha
hypothesis += y_train[i] * x_train[i] * self.alpha
these_misclassified_pts = predict_and_evaluate(hypothesis, x_train, y_train)
this_error = calculate_error(np.sum(these_misclassified_pts), x_train.shape[0])
if this_error < self.lowest_error:
self.best_hypothesis = hypothesis
self.lowest_error = this_error
self.logger.num_vector_updates += 1
misclassified_pts = predict_and_evaluate(hypothesis, x_train, y_train)
self.logger.num_iterations += 1
self.dv.plot_hypothesis(pts, y_train, self.best_hypothesis, target_fn)
self.print_fit_statistics()
def print_fit_statistics(self):
self.logger.print_statistics()
print('{:28s}: y = {:.2f}x + {:.2f}'.format('Hypothesis',
tdv.get_slope(self.best_hypothesis),
tdv.get_y_intercept(self.best_hypothesis)))
print('{0:28s}: {1:.2f}%'.format('In Sample (Training) Error', self.lowest_error * 100))
def visualize_training(self):
self.dv.visualize()
def predict(self, x):
return predict(x, self.best_hypothesis)
def predict_and_evaluate(hypothesis, x_train, y_train):
pred_classes = predict(hypothesis, x_train)
misclassified_pts = np.not_equal(pred_classes, y_train)
return misclassified_pts
def predict(x, hypothesis):
return np.sign(np.dot(x, hypothesis.T))
def calculate_error(num_misclassified_pts, num_pts):
return num_misclassified_pts / float(num_pts)
def get_data_visualizer(target_fn, x_train):
plot_title = 'Perceptron Learning'
if target_fn is not None:
plot_subtitle = 'Linearly Separable Training Data'
else:
plot_subtitle = 'Non-linearly Separable Training Data'
x_bound = np.max(np.absolute(x_train[:, 1]))
y_bound = np.max(np.absolute(x_train[:, 2]))
return DataVisualizer(plot_title, plot_subtitle, x_bound, y_bound)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/bezerraluis/Mentoria/blob/master/Exploratory_Data_Analysis_.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Hypotheses
* Some cities have higher properties prices;
* Having animals impact on the prices ;
* Houses with a higher condominium fee have a higher rent value;
* Furnished houses have a higher rent value;
* The number of parking spaces impact on rent value.
# Understanding the dataset
#### city: City where the property is located
#### area: Property area
#### rooms: Quantity of rooms
#### bathroom: Quantity of bathroom
#### parking spaces: Quantity of parking spaces
#### floor: Floor
#### animals: Accept animals?
#### furniture: Furniture?
#### hoa (R$): Homeowners association tax
#### rent amount (R$)sort: Rent amount
#### property tax: municipal property tax
#### fire insurance (R$): fire insurance value
#### total (R$): the sum of all values
# Libraries
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
```
# Opening and Viewing the data
```
df_houses = pd.read_csv('/content/drive/My Drive/Todos os arquivos do pc antigo/datasets Kaggle/houses_to_rent_v2.csv')
#df_houses = pd.read_csv('houses_to_rent_v2.csv')
df_houses.head()
df_houses.info()
def missing_values(data):
# Null total
missing_values = df_houses.isnull().sum()
# Order nulls
total = missing_values.sort_values(ascending=True)
# Percentage
percent = (missing_values / len(df_houses.index)*100).round(2).sort_values(ascending=True)
table_missing = pd.concat([total, percent], axis=1, keys=['Number of Nulls', 'Percentagem of Nulls'])
return table_missing.tail(10)
```
We do not have missing values on our data set
```
missing_values(df_houses)
```
# Data Cleaning
With describe we can see many featrues have values to be Varified like area Max value 1000 too distante from the quartiles and fire insurance with Max value 677. But we are goint to see each of them individualy.
```
df_houses.describe().T
```
### City
I always use the method unique() to find any misplaced values
The column city has no missing values, they are object and the values are all written correctly, it is ok to work with it
```
df_houses['city'].unique()
```
### Area
We do not have a good distribution the points above 10.000 are skewing our data.
```
sns.boxplot(df_houses['area'])
```
We can see that the Max value is 46.335 and de min 11 our std is 537 the data is too dispersed.
```
df_houses['area'].describe()
sns.distplot(df_houses['area'],bins= 20)
df_houses['area'].unique()
```
At first, I thought in remove areas below 20 m2. However, after some search, I found out many apartments between 11 and 20 m2
```
df_houses.query('area<20')
```
We have areas with 1.600 ,46.335 ,24.606 and son on , if we compare these values with the prices we can notice that a house in São Paulo with these size should be much higher than these values.
```
highier_1000 = df_houses.query('area >1000')
highier_1000.head(9)
```
We have 9 values that we are going to eliminate
```
highier_1000.shape
```
We are going to create a new data set with values from 1000 below
```
df_houses = df_houses.query('area<=1000')
df_houses.head()
```
Now we have 10.683 data points
```
df_houses.shape
```
Now we have a better distribution
```
sns.boxplot(df_houses['area'])
```
Before we had a std of 537 and now 129 our data is less disperse
```
df_houses['area'].describe()
```
We have a right skewed distribution because of the values above 200. However , we are going to keep them because them can show us some important insights
```
sns.distplot(df_houses['area'],bins= 15)
```
### Rooms
```
df_houses['rooms'].describe()
```
This features looks like ok , std 1.17 ,quartiles between 1 and 3 , just max values is a little high 13 let's see it in depth.
```
sns.boxplot(df_houses['rooms'])
df_houses['rooms'].unique()
```
I thought about changing rooms equal 1 because it is not commom here in country - side. However after some search I found out many One Room Apartments that is normal in Big cities.
```
check =df_houses.query('rooms==1')
check
```
Now let's see more than 6 rooms and compare with area to find any wrong values, it looks like okay it would be weird properties with 60 m2 and 7 rooms.
```
rooms = df_houses.query('rooms >6 & area<150')
rooms
```
### Bathroom
```
df_houses['bathroom'].unique()
```
I want to filter properties with 1 bathroom and area more than 180 m2 to find some relation , because I know that properties with more than 100 m2 usually have more than 1 bathroom , that's my parameter to find some wrong datapoints.
```
filter_180 = df_houses.query('bathroom==1 & area>180')
filter_180
```
I found some propertiers with 1 bathroom and more than 180 m2 , I 'm going to change this values with the mean.
```
```
Spliting bathroom equal 1 and area less and equal 100 because I do not want to change those values
```
test_menor100 =df_houses.query('bathroom==1 & area<=100')
test_menor100
```
The mean of the column Bathroom
```
media = df_houses['bathroom'].mean()
media
```
I m going to make a function to convert the datapoints equal 1 from the column bathroom
```
def medias (valor):
if valor == 1 :
return round(df_houses['bathroom'].mean())
else :
return valor
```
I atributed bathroom equal 1 and area highier than 150 to test maior
```
test_maior =df_houses.query('bathroom==1 & area>150')
test_maior.head(10)
```
Now I am going to change the values equal 1 using the method apply and the function medias.
```
test_maior['bathroom'] = test_maior['bathroom'].apply(medias)
test_maior.head(10)
print('test_menor',test_menor100.shape)
print('test_maior',test_maior.shape)
```
Now I am going to concat test menor and test maior and create a dataframe with all values of bathroom equal 1
```
df_houses_bath_1 = pd.concat([test_menor100,test_maior])
df_houses_bath_1.head()
df_houses_bath_1.shape
```
Now I need to select all data from bathroom different from 1 from our original dataframe and concat with df_houses_bath_1 to create a new dataframe adjusted.
```
df_houses_bath_diff_1 = df_houses.query('bathroom!=1')
df_houses_bath_diff_1.shape
```
We can see the new dataframe df_houses_2 where we do not have datapoints in bathroom equal 1 and area highier than 150 m2
```
df_houses_2 = pd.concat([df_houses_bath_1,df_houses_bath_diff_1])
df_houses_2.head(30)
df_houses_2.info()
```
### Parking Spaces
Our data is distributed between 1 and 2 , with max value 10 and std of 1,5
```
df_houses_2['parking spaces'].describe()
df_houses_2['parking spaces'].unique()
sns.boxplot(df_houses_2['parking spaces'])
```
### Floor
If we try to use describe we are goin to have a problem because we have the symbol " - " between our datapoints first we have to correct this.
```
df_houses_2['floor'].unique()
```
We changed the name of the column floor to floors because it was a reserved word on python.
```
df_houses_2 = df_houses.rename(columns={'floor':'floors'})
df_houses_2.head()
```
We are going to replace the synbol - by 0 because according to the author of the dataset this values are houses.
```
df_houses_2['floors'] = df_houses_2['floors'].replace('-','0')
df_houses_2['floors'].unique()
df_houses_2['floors'] = df_houses_2['floors'].astype(int)
df_houses_2.info()
```
We can see the Max value is 301 and the Minimum is 1 we problably have a problem , most of our values are distributed between 1 to 8.
```
df_houses_2['floors'].describe()
sns.boxplot(df_houses_2['floors'])
df_houses_2['floors'].unique()
df_houses_2.info()
```
After some search I found ou that the highiest building in Brazil has 81 floor , so this is going to be our parameter. We can see that we only have 1 value higher than 81 probably some typo.
```
df_81 = (df_houses_2['floors']> 81).sum()
df_81
df_houses_2['floors'].value_counts()
```
Now we are going to drop the value above 81
```
df_houses_2 = df_houses_2.query('floors<81')
df_houses_2['floors'].unique()
```
Now our distribution is better , most of our values are between 0 and 20. However , I am going to keep those values above because they can show us some important insight.
```
sns.boxplot(df_houses_2['floors'])
```
We have a right skewed distribution
```
sns.distplot(df_houses_2['floors'],bins=15)
```
### Animal
Now let's analyse the comlumn animal and furniture. It is a categorical value so we are going to look for some typos.
```
df_houses_2['animal'].unique()
df_houses_2['furniture'].unique()
```
### Other Columns
Let's analyze the last 4 columns
```
df_houses_2[['hoa (R$)', 'rent amount (R$)', 'property tax (R$)' ,'fire insurance (R$)']]
```
We have some suspect values on hoa(Homeowners association tax) and property tax(municipal property tax) let's see them individually
```
df_houses_2[['hoa (R$)', 'rent amount (R$)', 'property tax (R$)' ,'fire insurance (R$)']].describe().T
df_houses_2['hoa (R$)'].describe()
sns.boxplot(df_houses_2['hoa (R$)'])
df_houses_2['hoa (R$)'].value_counts()
df_houses_2['hoa (R$)'] = df_houses_2.rename({'hoa (R$)':'hoa'},axis=1, inplace=True)
df_houses_2 = df_houses_2.drop('hoa (R$)',axis=1)
df_houses_2.head()
```
We are no going to use values above 10000 because they are consider luxury condominium [luxury](https://www1.folha.uol.com.br/sobretudo/morar/2017/06/1891861-condominios-oferecem-servicos-exclusivos-e-chegam-a-custar-r-30-mil.shtml)
```
filter = df_houses_2.query('hoa <10000')
filter['hoa'].value_counts()
sns.boxplot(filter['hoa'])
```
We going to use the max value of 3000 , because after some search I found out the mean value for apartment in São Paulo is 1.718,07
[hoa ,apartments](https://imoveis.estadao.com.br/noticias/jardins-e-moema-sao-os-bairros-com-condominios-mais-caros-de-sao-paulo/) and houses 3,1 mil [Hoa,houses](https://exame.com/seu-dinheiro/quanto-custa-ter-uma-casa-de-condominio-em-12-capitais/)
```
cond = df_houses_2.query('hoa <=3000')
df_houses_2 = cond
sns.boxplot(df_houses_2['hoa'])
df_houses_2.info()
```
Property Tax
Our data is too disperse std 3458 and the max value 313700 is out of range.
```
df_houses_2['property tax (R$)'].describe()
```
We have a value next to 50000 and another next to 300000 biasing our data
```
sns.boxplot(df_houses_2['property tax (R$)'])
df_houses_2['property tax (R$)'] = df_houses_2.rename({'property tax (R$)':'property'},axis=1, inplace=True)
df_houses_2 = df_houses_2.drop('property tax (R$)',axis=1)
df_houses_2.head()
```
We are going to remove those values and use [Link](https://noticias.r7.com/sao-paulo/paulistano-paga-em-media-r-1398-de-iptu-residencial-diz-estudo-07112018)
```
filter2 = df_houses_2.query('property >=20000')
filter2.head()
```
Let's remove just the value 313700 because a property of 42 m2 should not have a property tax of this value.
```
filter2['property'].unique()
df_houses_2 = df_houses_2.query('property != 313700')
```
Just to confirm we do not have this value anymore on our dataset
```
df_houses_2.query('property >300000')
```
# Exploratory Data Analysis
## Answering the Hypotheses
* Some cities have higher properties prices;
* Having animals impact on the prices ;
* Houses with a higher condominium fee have a higher rent value;
* Furnished houses have a higher rent value;
* The number of parking spaces impact on rent value.
```
df_houses_2.head()
```
## Some cities have higher properties prices
Yes ,we can notice that São Paulo , Rio de janeiro e Belo Horizonte have the highest rent values.
```
df_houses_2.groupby('city')['total (R$)'].median().sort_values(ascending = False)
plt.figure(figsize=(10,4))
sns.barplot(x= df_houses_2['city'], y= df_houses_2['total (R$)'],ci= False,estimator= np.median,order=["São Paulo", "Rio de Janeiro","Belo Horizonte","Campinas","Porto Alegre"]);
```
## Having animals impact on the prices
Yes , properties that accept animals are about 500 reais more expensive.
```
city = df_houses_2.groupby('animal')['rent amount (R$)']
display(city.agg(['mean', 'median']))
```
Analysing by city we can notice that not all cities have highier prices for animal owners as Rio de Janeiro and Porto Alegre.
```
plt.figure(figsize= (15,5))
sns.barplot(x= df_houses_2['city'], y= df_houses_2['rent amount (R$)'],hue=df_houses_2['animal'],ci= False,estimator= np.median,order=["São Paulo", "Rio de Janeiro","Belo Horizonte","Campinas","Porto Alegre"]);
```
## Houses with a higher condominium fee have a higher rent value
```
df_houses_2.groupby('city')['hoa'].mean().sort_values(ascending = False)
plt.figure(figsize=(10,4))
sns.barplot(x='city',y = 'hoa',data= df_houses_2,ci=False,
order= ('Rio de Janeiro','São Paulo','Campinas','Belo Horizonte','Porto Alegre'));
df_houses_2.groupby('city')['rent amount (R$)'].mean().sort_values(ascending = False)
plt.figure(figsize=(10,4))
sns.barplot(x='city',y = 'rent amount (R$)',ci= False,data= df_houses_2,order= ('São Paulo','Belo Horizonte','Rio de Janeiro','Campinas','Porto Alegre'));
```
No ,There is no relation between comdominium fee and the value of rent, because the city with the top value of comdominum is Rio de Janeiro and the city with the highiest rent value is São Paulo and the Pearson correlation between them is weak 0.30.
```
corresp = df_houses_2[['rent amount (R$)','hoa']].corr()
corresp
```
## Furnished houses have a higher rent value;
```
df_houses_2.groupby('furniture')['rent amount (R$)'].mean().sort_values(ascending = False)
plt.figure(figsize=(15,6))
sns.catplot(x='city',y ='rent amount (R$)',col = 'furniture',kind= 'bar',ci= False,data= df_houses_2);
```
Yes , properties with furniture is more expensive to rent on average 1200 reais and the graphic shows that all cities have the same pattern.
```
```
## The number of parking spaces impact on rent value.
```
df_houses_2.groupby('parking spaces')['rent amount (R$)'].mean().sort_values(ascending = False)
```
Yes , from 0 to 7 the number of parking spaces impact on prices , just when we have 8 parking spaces the value is a little bit lower than 7 and 10 parking spaces is lower than 2 parking spaces.
```
plt.figure(figsize=(10,4))
sns.barplot(x= 'parking spaces', y= 'rent amount (R$)', data= df_houses_2, ci= False)
```
We have a correlation of 0.56 between the price and the number of vacancies confirming the idea that the number of vacancies is related to the value of the property.
```
corresp = df_houses_2[['rent amount (R$)','parking spaces']].corr()
corresp
```
## Correlations
Finally, we can see which variables have a greater correlation with the rent value
Area
Bathroom
Parking Space
Fire Insurance
```
plt.figure(figsize=[15,15])
sns.set(font_scale=1)
sns.heatmap(df_houses_2.corr(),center=0,annot=True)
```
# Conclusion
* ## Some cities have higher properties prices;
##### Yes ,We can notice that São Paulo , Rio de janeiro e Belo Horizonte have the highest rent values.
* ## Having animals impact on the prices ;
##### Yes , properties that accept animals are about 500 reais more expensive.
* ## Houses with a higher condominium fee have a higher rent value
##### No ,There is no relation between comdominium fee and the value of rent, because the city with the top value of comdominum is Rio de Janeiro and the city with the highiest rent value is São Paulo.
* ## Furnished houses have a higher rent value
##### Yes , properties with furniture is more expensive to rent on average 1200 reais and the graphic shows that all cities have the same pattern.
* ## The number of parking spaces impact on rent value.
##### Yes , from 0 to 7 the number of parking spaces impact on prices , just when we have 8 parking spaces the value is a little bit lower than 7 and 10 parking spaces is lower than 2 parking spaces.
```
```
| github_jupyter |
## Predicting Survival on the Titanic
### History
Perhaps one of the most infamous shipwrecks in history, the Titanic sank after colliding with an iceberg, killing 1502 out of 2224 people on board. Interestingly, by analysing the probability of survival based on few attributes like gender, age, and social status, we can make very accurate predictions on which passengers would survive. Some groups of people were more likely to survive than others, such as women, children, and the upper-class. Therefore, we can learn about the society priorities and privileges at the time.
### Assignment:
Build a Machine Learning Pipeline, to engineer the features in the data set and predict who is more likely to Survive the catastrophe.
Follow the Jupyter notebook below, and complete the missing bits of code, to achieve each one of the pipeline steps.
```
import re
# to handle datasets
import pandas as pd
import numpy as np
# for visualization
import matplotlib.pyplot as plt
# to divide train and test set
from sklearn.model_selection import train_test_split
# feature scaling
from sklearn.preprocessing import StandardScaler
# to build the models
from sklearn.linear_model import LogisticRegression
# to evaluate the models
from sklearn.metrics import accuracy_score, roc_auc_score
# to persist the model and the scaler
import joblib
# to visualise al the columns in the dataframe
pd.pandas.set_option('display.max_columns', None)
```
## Prepare the data set
```
# load the data - it is available open source and online
data = pd.read_csv('https://www.openml.org/data/get_csv/16826755/phpMYEkMl')
# display data
data.head()
len(data.columns)
# replace interrogation marks by NaN values
data = data.replace('?', np.nan)
# retain only the first cabin if more than
# 1 are available per passenger
def get_first_cabin(row):
try:
return row.split()[0]
except:
return np.nan
data['cabin'] = data['cabin'].apply(get_first_cabin)
# extracts the title (Mr, Ms, etc) from the name variable
def get_title(passenger):
line = passenger
if re.search('Mrs', line):
return 'Mrs'
elif re.search('Mr', line):
return 'Mr'
elif re.search('Miss', line):
return 'Miss'
elif re.search('Master', line):
return 'Master'
else:
return 'Other'
data['title'] = data['name'].apply(get_title)
# cast numerical variables as floats
data['fare'] = data['fare'].astype('float')
data['age'] = data['age'].astype('float')
# drop unnecessary variables
data.drop(labels=['name','ticket', 'boat', 'body','home.dest'], axis=1, inplace=True)
# display data
data.head()
# save the data set
data.to_csv('titanic.csv', index=False)
```
## Data Exploration
### Find numerical and categorical variables
```
target = 'survived'
vars_cat = [var for var in data.columns if data[var].dtype=='O']
vars_num = [var for var in data.columns if var not in vars_cat and var != target]
print('Number of numerical variables: {}'.format(len(vars_num)))
print('Number of categorical variables: {}'.format(len(vars_cat)))
```
### Find missing values in variables
```
vars_with_na = [var for var in data.columns if data[var].isnull().sum() > 0]
data[vars_with_na].isnull().mean().sort_values(ascending=False)
# first in numerical variables
num_na = [var for var in vars_num if var in vars_with_na]
num_na
# now in categorical variables
cat_na = [var for var in vars_cat if var in vars_with_na]
cat_na
```
### Determine cardinality of categorical variables
```
data[vars_cat].nunique()
```
### Determine the distribution of numerical variables
```
data[vars_num].hist(bins=30, figsize=(10, 10))
```
## Separate data into train and test
Use the code below for reproducibility. Don't change it.
```
X_train, X_test, y_train, y_test = train_test_split(
data.drop('survived', axis=1), # predictors
data['survived'], # target
test_size=0.2, # percentage of obs in test set
random_state=0) # seed to ensure reproducibility
X_train.shape, X_test.shape
```
## Feature Engineering
### Extract only the letter (and drop the number) from the variable Cabin
```
X_train.cabin = X_train.cabin.str[0]
X_test.cabin = X_test.cabin.str[0]
X_train['cabin'].unique()
```
### Fill in Missing data in numerical variables:
- Add a binary missing indicator
- Fill NA in original variable with the median
```
for var in ['age', 'fare']:
X_train[var+'_NA'] = np.where(X_train[var].isnull(), 1, 0)
X_test[var+'_NA'] = np.where(X_test[var].isnull(), 1, 0)
median_val = X_train[var].median()
X_train[var].fillna(median_val, inplace=True)
X_test[var].fillna(median_val, inplace=True)
X_train[['age', 'fare']].isnull().sum()
```
### Replace Missing data in categorical variables with the string **Missing**
```
X_train[cat_na] = X_train[cat_na].fillna('Missing')
X_test[cat_na] = X_test[cat_na].fillna('Missing')
X_train.isnull().sum()
X_test.isnull().sum()
```
### Remove rare labels in categorical variables
- remove labels present in less than 5 % of the passengers
```
def find_frequent_labels(df, var, rare_perc):
df = df.copy()
tmp = df.groupby(var)[var].count() / len(df)
return tmp[tmp > rare_perc].index
for var in vars_cat:
# find the frequent categories
frequent_ls = find_frequent_labels(X_train, var, 0.05)
# replace rare categories by the string "Rare"
X_train[var] = np.where(X_train[var].isin(
frequent_ls), X_train[var], 'Rare')
X_test[var] = np.where(X_test[var].isin(
frequent_ls), X_test[var], 'Rare')
X_train[vars_cat].nunique()
X_test[vars_cat].nunique()
```
### Perform one hot encoding of categorical variables into k-1 binary variables
- k-1, means that if the variable contains 9 different categories, we create 8 different binary variables
- Remember to drop the original categorical variable (the one with the strings) after the encoding
```
for var in vars_cat:
X_train = pd.concat([X_train,
pd.get_dummies(X_train[var], prefix=var, drop_first=True)
], axis=1)
X_test = pd.concat([X_test,
pd.get_dummies(X_test[var], prefix=var, drop_first=True)
], axis=1)
X_train.drop(labels=vars_cat, axis=1, inplace=True)
X_test.drop(labels=vars_cat, axis=1, inplace=True)
X_train.shape, X_test.shape
X_train.head()
X_test.head()
X_test['embarked_Rare'] = 0
variables = [c for c in X_train.columns]
variables
```
### Scale the variables
- Use the standard scaler from Scikit-learn
```
# create scaler
scaler = StandardScaler()
# fit the scaler to the train set
scaler.fit(X_train[variables])
# transform the train and test set
X_train = scaler.transform(X_train[variables])
X_test = scaler.transform(X_test[variables])
```
## Train the Logistic Regression model
- Set the regularization parameter to 0.0005
- Set the seed to 0
```
# set up the model
# remember to set the random_state / seed
model = LogisticRegression(C=0.0005, random_state=0)
# train the model
model.fit(X_train, y_train)
```
## Make predictions and evaluate model performance
Determine:
- roc-auc
- accuracy
**Important, remember that to determine the accuracy, you need the outcome 0, 1, referring to survived or not. But to determine the roc-auc you need the probability of survival.**
```
# make predictions for test set
class_ = model.predict(X_train)
pred = model.predict_proba(X_train)[:,1]
# determine mse and rmse
print('train roc-auc: {}'.format(roc_auc_score(y_train, pred)))
print('train accuracy: {}'.format(accuracy_score(y_train, class_)))
print()
# make predictions for test set
class_ = model.predict(X_test)
pred = model.predict_proba(X_test)[:,1]
# determine mse and rmse
print('test roc-auc: {}'.format(roc_auc_score(y_test, pred)))
print('test accuracy: {}'.format(accuracy_score(y_test, class_)))
print()
```
That's it! Well done
**Keep this code safe, as we will use this notebook later on, to build production code, in our next assignement!!**
| github_jupyter |
# Authorise Notebook server to access Earth Engine
This notebook is a reproduction of the workflow originally developed by **Datalab**, which describes how to setup a Google Datalab container in your local machine using Docker.
You can check out the full tutorial by going to this link: https://developers.google.com/earth-engine/python_install-datalab-local
```
# Code to check the IPython Widgets library.
try:
import ipywidgets
except ImportError:
print('The IPython Widgets library is not available on this server.\n'
'Please see https://github.com/jupyter-widgets/ipywidgets '
'for information on installing the library.')
raise
print('The IPython Widgets library (version {0}) is available on this server.'.format(
ipywidgets.__version__
))
```
Next, check if the Earth Engine API is available on the server.
```
# Code to check the Earth Engine API library.
try:
import ee
except ImportError:
print('The Earth Engine Python API library is not available on this server.\n'
'Please see https://developers.google.com/earth-engine/python_install '
'for information on installing the library.')
raise
print('The Earth Engine Python API (version {0}) is available on this server.'.format(
ee.__version__
))
```
Finally, check if the notebook server is authorized to access the Earth Engine backend servers.
```
# Code to check if authorized to access Earth Engine.
import io
import os
import urllib
from IPython import display
# Define layouts used by the form.
row_wide_layout = ipywidgets.Layout(flex_flow="row nowrap", align_items="center", width="100%")
column_wide_layout = ipywidgets.Layout(flex_flow="column nowrap", align_items="center", width="100%")
column_auto_layout = ipywidgets.Layout(flex_flow="column nowrap", align_items="center", width="auto")
form_definition = {'form': None}
response_box = ipywidgets.HTML('')
def isAuthorized():
try:
ee.Initialize()
test = ee.Image(0).getInfo()
except:
return False
return True
def ShowForm(auth_status_button, instructions):
"""Show a form to the user."""
form_definition['form'] = ipywidgets.VBox([
auth_status_button,
instructions,
ipywidgets.VBox([response_box], layout=row_wide_layout)
], layout=column_wide_layout)
display.display(form_definition.get('form'))
def ShowAuthorizedForm():
"""Show a form for a server that is currently authorized to access Earth Engine."""
def revoke_credentials(sender):
credentials = ee.oauth.get_credentials_path()
if os.path.exists(credentials):
os.remove(credentials)
response_box.value = ''
Init()
auth_status_button = ipywidgets.Button(
layout=column_wide_layout,
disabled=True,
description='The server is authorized to access Earth Engine',
button_style='success',
icon='check'
)
instructions = ipywidgets.Button(
layout = row_wide_layout,
description = 'Click here to revoke authorization',
disabled = False,
)
instructions.on_click(revoke_credentials)
ShowForm(auth_status_button, instructions)
def ShowUnauthorizedForm():
"""Show a form for a server that is not currently authorized to access Earth Engine."""
auth_status_button = ipywidgets.Button(
layout=column_wide_layout,
button_style='danger',
description='The server is not authorized to access Earth Engine',
disabled=True
)
auth_link = ipywidgets.HTML(
'<a href="{url}" target="auth">Open Authentication Tab</a><br/>'
.format(url=ee.oauth.get_authorization_url()
)
)
instructions = ipywidgets.VBox(
[
ipywidgets.HTML(
'Click on the link below to start the authentication and authorization process. '
'Once you have received an authorization code, use it to replace the '
'REPLACE_WITH_AUTH_CODE in the code cell below and run the cell.'
),
auth_link,
],
layout=column_auto_layout
)
ShowForm(auth_status_button, instructions)
def Init():
# If a form is currently displayed, close it.
if form_definition.get('form'):
form_definition['form'].close()
# Display the appropriate form according to whether the server is authorized.
if isAuthorized():
ShowAuthorizedForm()
else:
ShowUnauthorizedForm()
Init()
```
If the server **is authorized**, you do not need to run the next code cell.
If the server **is not authorized**:
1. Copy the authentication code generated in the previous step.
2. Replace the REPLACE_WITH_AUTH_CODE string in the cell below with the authentication code.
3. Run the code cell to save authentication credentials.
```
auth_code = 'REPLACE_WITH_AUTH_CODE'
response_box = ipywidgets.HTML('')
try:
token = ee.oauth.request_token(auth_code.strip())
ee.oauth.write_token(token)
if isAuthorized():
Init()
else:
response_box.value = '<font color="red">{0}</font>'.format(
'The account was authenticated, but does not have permission to access Earth Engine.'
)
except Exception as e:
response_box.value = '<font color="red">{0}</font>'.format(e)
response_box
# Code to display an Earth Engine generated image.
from IPython.display import Image
url = ee.Image("CGIAR/SRTM90_V4").getThumbUrl({'min':0, 'max':3000})
Image(url=url)
```
| github_jupyter |
### speaker data
```
import pandas as pd
data = pd.read_csv("speaker_data.csv")
data
```
### bio data
```
# delete / in date
data = data.replace(to_replace= r'/', value= '', regex=True)
data
biodata = pd.read_csv("biographydata.csv")
biodata
```
### alternative data(policy decision
```
changedata = pd.read_csv("alternativedata.csv")
changedata = change.replace(to_replace= r'/', value= '', regex=True)
changedata
data['start_date'] = data['start_date'].astype(str)
data["Doc"] = data[['start_date', 'Speaker']].agg('-'.join, axis=1)
data
df=data[['Doc','content']]
df['Doc'] = df['Doc'].astype('str')
df['content'] = df['content'].astype('str')
df
```
### document create
```
# https://stackoverflow.com/questions/38127209/how-to-use-groupby-to-concatenate-strings-in-python-pandas
document = df.groupby('Doc')['content'].apply(','.join)
document
document = document.reset_index()
document.columns = ['id','text']
document
document[['date','speaker']] = document['id'].str.split('-',expand=True)
document
document['fed'] = None
document
```
## Add Fed(YES or NO) to df
```
for index, row in document.iterrows():
for i, r in biodata.iterrows():
if row["speaker"]==r["name"]:
row["fed"]=r['d_regional_fed']
document
for index, row in document.iterrows():
if row["fed"] is None:
row["fed"]="NA"
document
document['change'] = None
document
```
## Add change decision(a,b,c,d,e) to df
```
for index, row in document.iterrows():
for i, r in changedata.iterrows():
if row["date"]==r["start_date"]:
row["change"]=r['change']
document
for index, row in document.iterrows():
if row["change"] is None:
row["change"]="NA"
document
# convert to json list
document.to_json('speaker_doc.jsonlist',orient='records', lines=True)
document.to_csv("speaker_doc.csv")
```
## min-count 70, convert fed, change to one-hot
```
import preprocess_data
script = 'preprocess_data.py'
args = '/Users/Zhe/PycharmProjects/Research-Project/speaker_doc.jsonlist /Users/Zhe/PycharmProjects/Research-Project/FOMC --min-doc-count 70 --label fed,change'
print("python", script, args)
preprocess_data.main(args.split())
# load the vocabualry
import json
import os
with open(os.path.join('FOMC', 'train.vocab.json')) as f:
vocab = json.load(f)
print("First few words in the vocbulary:")
print(vocab[:6] + ['...'])
# load a covariate file
import pandas as pd
print("Start of a covariate file (train.fed.csv):")
df1 = pd.read_csv(os.path.join('FOMC','train.fed.csv'), header=0, index_col=0)
print(df1.head())
import run_scholar
script = 'run_scholar.py'
args = 'FOMC/ -k 10 --epochs 50 --dev-folds 10 --seed 42'
print("python", script, args)
run_scholar.main(args.split())
```
# Covars-fed
```
script = 'run_scholar.py'
args = 'FOMC/ -k 10 --epochs 150 --dev-folds 10 --seed 42 --topic-covars fed'
print("python", script, args)
run_scholar.main(args.split())
import numpy as np
from run_scholar import print_top_words
# load the stored (K x V) topic matrix (stored in a compressed numpy format)
beta = np.load(os.path.join('output', 'beta.npz'))['beta']
print_top_words(beta, vocab, n_pos=7, n_neg=0);
topic_covars = np.load(os.path.join('output', 'beta_c.npz'))
weights = topic_covars['beta']
names = topic_covars['names']
print_top_words(weights, vocab, topic_names=names, n_pos=7, n_neg=0);
```
# Interaction-fed
```
interactions = np.load(os.path.join('output', 'beta_ci.npz'))
weights = interactions['beta']
names = topic_covars['names']
names = [str(k) + ':' + c for k in range(10) for c in names]
print_top_words(weights, vocab, topic_names=names, n_pos=8, n_neg=0);
```
# label-decision, covars-fed
```
script = 'run_scholar.py'
args = 'FOMC/ -k 10 --epochs 150 --dev-folds 10 --seed 42 --topic-covars change,fed'
print("python", script, args)
run_scholar.main(args.split())
script = 'run_scholar.py'
args = 'FOMC/ -k 12 --epochs 150 --dev-folds 10 --seed 42 --topic-covars fed --label change'
print("python", script, args)
run_scholar.main(args.split())
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Eager modunun ana hatlari
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/tr/r1/tutorials/eager/eager_basics.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/tr/r1/tutorials/eager/eager_basics.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
Bu kitapcikta TensorFlow kullanarak konuya giris yapacagiz. Asagidaki konulari isleyecegiz:
* Gerekli paketleri iceri aktarma
* Tensorlari olusturma ve kullanma
* GPU hizlandirmayi kullanmak
* Veri setleri
## TensorFlow'u iceri alalim
'tensorflow' modulunu iceri alalim ver eager modunu secelim.
Eager modu, TensorFlow'a detaylarini daha sonra aciklayacagimiz etkilesimli bir arayuz saglar.
```
from __future__ import absolute_import, division, print_function
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow.compat.v1 as tf
```
## Tensorlar
Tensor kisaca cok boyutlu bir dizidir. NumPy'deki 'ndarray' nesneleri gibi, `Tensor` nesnesinin de bir veri turu ve sekli vardir. Ayrica Tensorlar GPU gibi hizlandirilmis hafizada bulunabilirler. TensorFlow, Tensorlari olusturmak ve kullanmak icin zengin islemlere sahiptir ([tf.add](https://www.tensorflow.org/api_docs/python/tf/add), [tf.matmul](https://www.tensorflow.org/api_docs/python/tf/matmul), [tf.linalg.inv](https://www.tensorflow.org/api_docs/python/tf/linalg/inv) etc.). Bu islemler Python tiplerini otomatik olarak degistirirler. Ornegin:
```
print(tf.add(1, 2))
print(tf.add([1, 2], [3, 4]))
print(tf.square(5))
print(tf.reduce_sum([1, 2, 3]))
print(tf.encode_base64("hello world"))
# Islec asiri yuklenmesi de desteklenir
print(tf.square(2) + tf.square(3))
```
Her Tensor'un bir sekli ve veri turu vardir
```
x = tf.matmul([[1]], [[2, 3]])
print(x.shape)
print(x.dtype)
```
NumPy dizileri ve TensorFlow Tensorlari arasindaki en belirgin farklar sunlardir:
1. Tensorlar hizlandirilmis hafizalar tarafindan desteklenebilr (GPU, TPU gibi).
2. Tensorlar degistirilemez.
### NumPy Uyumlulugu
TensorFlow Tensorlari ile NumPy 'ndarray'leri arasindaki donusum cok basittir:
* TensorFlow islemleri otomatik olarak NumPy ndarray'lerini Tensorlara donusturur.
* NumPy islemleri de otomatik olarak Tensorlari NumPy ndarray'lerine donusturur.
'.numpy()' metodunu kullanarak Tensorlari belirgin sekilde NumPy ndarray'lerine donusturebilirsiniz.
Tensorlar ve 'ndarray'ler temelde mumkun oldugunca ayni sekilde tanimlandigi icin bu donusturmeler ucuzdur. Fakat, NumPy dizileri her zaman ana hafizada calisirken Tensorlar GPU hafizasini da kullanabildigi icin her zaman benzer sekilde tanimlanamazlar ve donusturme isleminde GPU'dan ana hafizaya kopyalama da bulunur.
```
import numpy as np
ndarray = np.ones([3, 3])
print("TensorFlow operations convert numpy arrays to Tensors automatically")
tensor = tf.multiply(ndarray, 42)
print(tensor)
print("And NumPy operations convert Tensors to numpy arrays automatically")
print(np.add(tensor, 1))
print("The .numpy() method explicitly converts a Tensor to a numpy array")
print(tensor.numpy())
```
## GPU hizlandirmasi
Hesaplamalar icin GPU kullanarak bircok TensorFlow islemleri hizlandirilabilir. TensorFlow bir islem icin, ek aciklamaya gerek duymadan, otomatik olarak GPU ya da CPU kullanimina karar verir (ve gerektiginde tensorlari GPU ve CPU hafizalarina kopyalar). Bir islem sonucu olusan tensorlar o islem hangi hafizada yurutulduyse o hafizaya kopyalanir. Ornegin:
```
x = tf.random_uniform([3, 3])
print("Is there a GPU available: "),
print(tf.test.is_gpu_available())
print("Is the Tensor on GPU #0: "),
print(x.device.endswith('GPU:0'))
```
### Aygit Isimleri
`Tensor.device` ozelligi tensorlarin bulundugu aygitin tam adini dizgi olarak temin eder. Bu dizgide bircok detay bulunmaktadir: programin calistigi anasistemin bulundugu agin taniticisi ve anasistemdeki aygit. Bunlar TensorFlow programlarinin dagitiminda gerekli olan bilgilerdir. Eger tensor sistemdeki 'N'inci GPU'ya yerlestirilmisse bu dizgi `GPU:<N>` ile biter.
### Belirtilmis Aygit Yerlestirilmesi
TensorFlow'da "yerlestirme" terimi islemlerin uygulama sirasinda sistemde tek tek nasil atandigi (yerlestirildigi) anlaminda kullanilmistir. Yukarida da bahsettigimiz gibi, eger ozellikle belirtilmemisse TensorFlow bir islemi nerde calistiracagina otomatik olarak karar verir ve gerekirse tensorlari oraya kopyalar. Fakat, TensorFlow islemleri 'tf.device' baglam yoneticisi kullanilarak belirli aygitlara yerlestirilebilir. Ornegin:
```
import time
def time_matmul(x):
start = time.time()
for loop in range(10):
tf.matmul(x, x)
result = time.time()-start
print("10 loops: {:0.2f}ms".format(1000*result))
# CPU ustunde zorla calistirma
print("On CPU:")
with tf.device("CPU:0"):
x = tf.random_uniform([1000, 1000])
assert x.device.endswith("CPU:0")
time_matmul(x)
# Eger mumkunse GPU ustunde zorla calistirma #0
if tf.test.is_gpu_available():
with tf.device("GPU:0"): # Or GPU:1 for the 2nd GPU, GPU:2 for the 3rd etc.
x = tf.random_uniform([1000, 1000])
assert x.device.endswith("GPU:0")
time_matmul(x)
```
## Veri setleri
Simdi modelimize veri akimini saglamak icin [`tf.data.Dataset` API](https://www.tensorflow.org/r1/guide/datasets)'sini nasil kullanacagimizi gorecegiz:
* `Dataset`i olusturalim.
* Eager modunda `Dataset`in yinelenmesi.
Modelimizin egitim ve degerlendirme dongulerine verilen kompleks girdi hatlarini 'Dataset' API'si ile basit ve tekrar kullanilabilir parcalardan olusturmanizi tavsiye ederiz.
'Dataset' nesnesi olusturma API'si eager modunda iken TensorFlow graph'taki ile aynidir, fakat veri setindeki elemanlarin yinelenmesi islemi biraz daha basittir.
'tf.data.Dataset' nesnesi ustunde direk olarak Python yinelemesi yapabildiginiz icin `tf.data.Iterator` nesnesi olusturmaniza gerek yoktur.
Sonuc olarak, eger eager modunu kullaniyorsaniz, [TensorFlow Rehberi](https://www.tensorflow.org/r1/guide/datasets)'nde anlatilan yineleme gereksizdir.
### `Dataset` kaynagi olusturalim
Buradaki fonksiyonlardan birini [`Dataset.from_tensors`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#from_tensors), [`Dataset.from_tensor_slices`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#from_tensor_slices) ya da kutuklerden okunan nesneleri [`TextLineDataset`](https://www.tensorflow.org/api_docs/python/tf/data/TextLineDataset) veya [`TFRecordDataset`](https://www.tensorflow.org/api_docs/python/tf/data/TFRecordDataset) kullanarak _source_ dataset olusturabiliriz. [TensorFlow Rehberi](https://www.tensorflow.org/r1/guide/datasets#reading_input_data)'nde daha detayli bilgi bulabilirsiniz.
```
ds_tensors = tf.data.Dataset.from_tensor_slices([1, 2, 3, 4, 5, 6])
# CSV kutugunu olusturalim
import tempfile
_, filename = tempfile.mkstemp()
with open(filename, 'w') as f:
f.write("""Line 1
Line 2
Line 3
""")
ds_file = tf.data.TextLineDataset(filename)
```
### Transformations (donusumler) uygulayalim
Veri seti kayitlarini donusturmek icin transformations (donusumler) fonksiyonlarini kullanabiliriz: ornegin [`map`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#map), [`batch`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#batch), [`shuffle`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#shuffle). `tf.data.Dataset` API dokumanlari hakkinda daha fazla bilgi icin [buraya bakiniz](https://www.tensorflow.org/api_docs/python/tf/data/Dataset).
```
ds_tensors = ds_tensors.map(tf.square).shuffle(2).batch(2)
ds_file = ds_file.batch(2)
```
### Yineleme
Eager modunda 'Dataset' nesneleri yinelemeleri destekler.
Eger TensorFlow 'graphs'taki 'Dataset' kullanimina asina iseniz, `Dataset.make_one_shot_iterator()` ya da `get_next()` kullanimina gerek olmadigina lutfen dikkat ediniz.
```
print('Elements of ds_tensors:')
for x in ds_tensors:
print(x)
print('\nElements in ds_file:')
for x in ds_file:
print(x)
```
| github_jupyter |
# Demo - Bayesian Neural Network Regression
```
import numpy as np
from sklearn import datasets
import pandas as pd
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
import torchbnn as bnn
from sklearn.preprocessing import StandardScaler, MinMaxScaler
from sklearn.metrics import r2_score, mean_squared_error
import os
import warnings
import numpy as np
import pandas as pd
from tqdm import tqdm
from transistor_parameter_functions import subthreshold_swing, threshold_voltage, mobility_degradation, R_square
from collections import OrderedDict
import matplotlib.pyplot as plt
%matplotlib inline
cuda = torch.device("cuda")
X_train = pd.read_csv("./Train/Design.csv").to_numpy()# .iloc[:1000, :]
scaler = MinMaxScaler()
scaler.fit(X_train)
X_train = scaler.fit_transform(X_train)
y_train = pd.read_csv("./Train/Character.csv", header=None).iloc[:,1].to_numpy()# .iloc[:1000, :]
X_test = pd.read_csv("./Test/Design.csv").to_numpy()# [VARIABLE]
X_test = scaler.fit_transform(X_test )
y_test = pd.read_csv("./Test/Character.csv", header=None).iloc[:,1].to_numpy()
(X_train, X_test, y_train, y_test) = tuple(map(lambda x: torch.tensor(x, device=cuda), (X_train, X_test, y_train, y_test)))
```
## 1. Generate Sample Data
```
(X_train.shape, X_test.shape, y_train.shape, y_test.shape)
#X_train = torch.unsqueeze(X_train, dim=1)
#y_train = torch.unsqueeze(y_train, dim=1)
X_train = X_train.float()
y_train = y_train.float()
(X_train.shape, X_test.shape, y_train.shape, y_test.shape)
```
## 2. Define Model
```
model = nn.Sequential(
bnn.BayesLinear(prior_mu=0, prior_sigma=0.1, in_features=4, out_features=100),
nn.ReLU(),
bnn.BayesLinear(prior_mu=0, prior_sigma=0.1, in_features=100, out_features=1),
)
model.to("cuda")
mse_loss = nn.MSELoss()
kl_loss = bnn.BKLLoss(reduction='mean', last_layer_only=False)
kl_weight = 0.01
optimizer = optim.Adam(model.parameters(), lr=0.01)
```
## 3. Train Model
```
kl_weight = 0.1
for step in range(3001):
pre = model(X_train)
mse = mse_loss(pre, y_train)
kl = kl_loss(model)
cost = mse + kl_weight*kl
optimizer.zero_grad()
cost.backward()
optimizer.step()
if step%300==0:
print(f'{step}- MSE : %2.2f, KL : %2.2f' % (mse.item(), kl.item()))
```
## 4. Test Model
```
x_test = torch.linspace(-2, 2, 500)
y_test = x_test.pow(3) - x_test.pow(2) + 3*torch.rand(x_test.size())
x_test = torch.unsqueeze(x_test, dim=1)
y_test = torch.unsqueeze(y_test, dim=1)
plt.xlabel(r'$x$')
plt.ylabel(r'$y$')
plt.scatter(x_test.data.numpy(), y_test.data.numpy(), color='k', s=2)
y_predict = model(x_test)
plt.plot(x_test.data.numpy(), y_predict.data.numpy(), 'r-', linewidth=5, label='First Prediction')
y_predict = model(x_test)
plt.plot(x_test.data.numpy(), y_predict.data.numpy(), 'b-', linewidth=5, label='Second Prediction')
y_predict = model(x_test)
plt.plot(x_test.data.numpy(), y_predict.data.numpy(), 'g-', linewidth=5, label='Third Prediction')
plt.legend()
plt.show()
```
| github_jupyter |
# Optimización de portafolios
## Markowitz's mean-variance
### Planteamiento del problema
Sea $\mathbf{w} = (w_1, \ldots, w_N)$ el vector (columna) de ponderaciones del portafolio, es decir, $w_i$ representa la proporción que se tiene del activo $i$.
Dado un rendimiento deseado, $\mu^{*} \in \mathbb{R}$, se busca obtener un vector $\mathbf{w}^{*} = (w_{1}^{*}, \ldots, w_{N}^{*})$ que sea solución del siguiente problema de optimización
$$
\min_{\mathbf{w}} \sigma_{p}^{2} = \min_{\mathbf{w}} \mathbf{w}^{T} \mathbf{\Sigma} \mathbf{w}
$$
sujeto a las siguientes restricciones lineales
$$
\sum_{i = 1}^{N} w_{i} = 1
$$
y
$$
\mu_{p} = \sum_{i = 1}^{N} w_{i} \widehat{\mu_{i}} = \mu^{*}
$$
en donde $\widehat{\mu_{i}}$ es el rendimiento promedio del activo $i$.
De acuerdo al problema anterior, buscamos una combinación de ponderaciones que minimice la varianza del portafolio (y por lo tanto la volatilidad $\sigma_{p}$), invirtiendo todo nuestro capital disponible (restricción lineal 1) y a la vez obteniendo un rendimiento esperado $\mu^{*}$ (restriccion lineal 2).
El objetivo de este proyecto es resolver este problema de optimización y graficar la **frontera eficiente**.
## Datos
Se utilizarán los datos de los tres tipos de cambio `USD_MXN.csv` `EUR_MXN.csv` `GBP_MXN.csv`que vimos en clase.
## Actividades a realizar
* A partir de los archivos `csv` señalados, se deben de obtener los rendimientos logarítmicos de los precios de cierre ajustado `Adj Close` de cada archivo. Considere lo siguiente:
* En estos archivos un valor no disponible se establece con el string `"null"`.
* No se tiene la misma cantidad de información en los tres archivos.
* El rendimiento logarítmico en el tiempo $t$ está dado por $\ln(P_{t} / P_{t - 1})$.
* No puede calcular ninguna cantidad con una hoja de cálculo (excel) todo debe ser con python.
* A partir de los rendimientos logarítmicos obtenga el vector (ndarray, renglón) $\mathbf{\widehat{\mu}} = (\widehat{\mu_1}, \ldots, \widehat{\mu_N})$ en donde cada $\widehat{\mu_i}$ representa el promedio de los rendimientos logarítmicos del activo $i$. También, obtenga la matriz de varianzas y covarianzas $\mathbf{\Sigma}$. Considere lo siguiente
* El vector $\mathbf{\widehat{\mu}}$ debe de tener rendimientos anualizados, es decir cada entrada $\mu_{i}$ debe de ir multiplicada por $252$.
```python
import numpy as np
import pandas as pd
help(np.cov) #CUIDADO CON EL PARÁMETRO rowvar
#Los Dataframe tienen el método cov implementado
help(pd.DataFrame.cov)
#Si X es un data frame con sólo columnas
#numéricas, entonces
X.cov() # regresa la matriz de varianzas y covarianzas
#Los objetos DataFrame y Series de pandas
#al igual que los arreglos de numpy
#tienen una serie de métodos como
X.mean(axis = 0 o 1
X.std(axis = 0 o 1)
```
* Programe una función para calcular la varianza del portafolio $\sigma_{p}^{2} = \mathbf{w}^{T} \mathbf{\Sigma} \mathbf{w}$, esta es la función objetivo de nuestro problema de optimización.
* Recuerde que con **numpy** puede realizar la multiplicación de matrices (ndarrays) con `np.matmul` o con el símbolo `@`.
* La varianza debe de regresarse anualizada, es decir, debe de calcular $ 252 * \mathbf{w}^{T} \mathbf{\Sigma} \mathbf{w}$.
* Resuelva el problema de optimización utilizando la función `minimize` del módulo `scipy.optimize` para distintos valores valores de $\mu^{*}$, estos rendimientos objetivo (anualizados) estarán dados por
```python
mu_0 = np.linspace(0.02, 0.11, 100)
```
* Grafique la frontera eficiente, el eje X será la volatilidad (anualizada) del portafolio, $\sigma_{p}$, la cual se obtiene al aplicar la raíz cuadrada del valor regresado por la función `minimize`. El eje Y será el rendimiento esperado $\mu^{*}$ asociado con esa volatilidad.
* Finalmente cree las siguientes tablas:
* Una tabla con las ponderaciones de cada activo, el rendimiento esperado y la desviación estándar asociada a este rendimiento
* Una tabla con las ponderaciones de cada activo, el rendimiento esperado y la desviación estándar asociada a este rendimiento, considerando únicamente los casos en los que cada ponderación se encuentre en el intervalo $[-1,1]$.
**DEBE DE CREAR SU CÓDIGO PENSANDO EN QUE SEA ROBUSTO, ES DECIR, PENSANDO EN QUE LA VIDA REAL NO VA A TRABAJAR CON SÓLO 3 ARCHIVOS. PIENSE EN COMO AUTOMATIZAR EL PROCESO DESCRITO Y EL ÚNICO CAMBIO NECESARIO SERÍA MODIFICAR LO DE LA VARIABLE `rutas`**
# FECHA DE ENTREGA
Ya que tengo que entregar las calificaciones el día 30 de junio, la entrega de este proyecto es **a más tardar** el día **sábado 27 de junio a las 11:59 horas**.
**ESTA VEZ NO HABRÁ PRORROGA**
```
import numpy as np
import pandas as pd
from scipy.optimize import minimize
from scipy.optimize import LinearConstraint
from scipy.optimize import Bounds
from numpy import linalg
import matplotlib.pyplot as plt
#NA aparece como null
#revise el método dropna() que contienen
#los dataframe y los objetos Series
rutas = ['../datos/USD_MXN.csv',
'../datos/EUR_MXN.csv',
'../datos/GBP_MXN.csv']
#NO EJECUTE ESTA CELDA SI NO QUIERE PERDER LA TABLA
#SUGERENCIA: Crear un data frame que combina las columnas Adj Close
#NO EJECUTE ESTA CELDA SI NO QUIERE PERDER LA TABLA
#Se calculan los rendimientos (logarítmicos)
def calcula_varianza_port(pesos, *args):
pass
#NO EJECUTE ESTA CELDA SI NO QUIERE PERDER LA TABLA
#FRONTERA EFICIENTE
#NO PUEDO ASUMIR QUE TODOS
#SABEN ÁLGEBRA LINEAL
#LA PARTE DE IMPLEMENTAR
#LAS RESTRICCIONES LINEALES
#SE LAS HICE
#restricción de suma de pesos
n_activos = #NÚMERO DE ACTIVOS
unos = np.ones(n_activos)
rest_suma = LinearConstraint(unos, lb = 1, ub = 1)
#rendimientos deseados
mu_0 = np.linspace(0.02, 0.11, 100)
#Vector de pesos iniciales
#Parámetro x0 de minimize, puede inicializar
#considerando la misma ponderáción
#para cada activo, es decir, 1 / n_activos
#pesos =
#Se resuelve el problema de
#optimización para cada
#rendimiento deseado
for i in range(len(mu_0)):
#restricción de rendimiento deseado
rest_rend = LinearConstraint(mu,
lb = mu_0[i],
ub = mu_0[i])
#agrupa restricciones
rest = [rest_suma, rest_rend]
#busca solución
#Especifique el resto de los parámetros
solucion = minimize(
method = 'SLSQP'
constraints = rest)
#Verifica si se encontró solución
if solucion.success:
#TABLA DE PONDERACIONES
#NO EJECUTE ESTA CELDA SI NO QUIERE PERDER LA GRÁFICA
#Basta con graficar la curva azul
#NO EJECUTE ESTA CELDA
#TABLA DE PONDERACIONES EN DONDE CADA w_i está en [-1,1] únicamente
#NO EJECUTE ESTA CELDA
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import sklearn
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import KFold
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.linear_model import Perceptron
from sklearn.naive_bayes import GaussianNB
from sklearn.linear_model import LogisticRegression
from sklearn.neural_network import MLPClassifier
from sklearn.svm import SVC
from matplotlib import pyplot as plt
# plt.style.use('dark_background')
data = pd.read_csv('dataset_comb.csv')
data.head()
X = np.array(data[['Area','MajorAxisLength', 'MinorAxisLength', 'Eccentricity', 'ConvexArea', 'EquivDiameter', 'Extent', 'Perimeter', 'Roundness', 'AspectRation']])
y = np.array(data['Class'])
def train_evaluate(model):
train_accs = []
test_accs = []
kf = KFold(n_splits=7, shuffle=True, random_state=2021)
for train_index, test_index in kf.split(X):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
scaler = MinMaxScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
model.fit(X_train, y_train)
train_accs.append(model.score(X_train, y_train))
test_accs.append(model.score(X_test, y_test))
return train_accs, test_accs
linear_disc = LinearDiscriminantAnalysis()
linear_disc_scores_tr, linear_disc_scores_test = train_evaluate(linear_disc)
print(f'Train accurracy: {np.mean(linear_disc_scores_tr)}; Test accuracy: {np.mean(linear_disc_scores_test)}')
perceptron = Perceptron()
perceptron_scores_tr, perceptron_scores_test = train_evaluate(perceptron)
print(f'Train accurracy: {np.mean(perceptron_scores_tr)}; Test accuracy: {np.mean(perceptron_scores_test)}')
nb = GaussianNB()
nb_scores_tr, nb_scores_test = train_evaluate(nb)
print(f'Train accurracy: {np.mean(nb_scores_tr)}; Test accuracy: {np.mean(nb_scores_test)}')
lr = LogisticRegression()
lr_scores_tr, lr_scores_test = train_evaluate(lr)
print(f'Train accurracy: {np.mean(lr_scores_tr)}; Test accuracy: {np.mean(lr_scores_test)}')
ann = MLPClassifier(alpha=0.02, max_iter=10000)
ann_scores_tr, ann_scores_test = train_evaluate(ann)
print(f'Train accurracy: {np.mean(ann_scores_tr)}; Test accuracy: {np.mean(ann_scores_test)}')
svc = SVC()
svc_scores_tr, svc_scores_test = train_evaluate(svc)
print(f'Train accurracy: {np.mean(svc_scores_tr)}; Test accuracy: {np.mean(svc_scores_test)}')
fig, axs = plt.subplots(2, 3, figsize=(20, 10))
fig.suptitle('Accuracy boxplots for all 6 methods')
axs[0, 0].boxplot(linear_disc_scores_test, flierprops=dict(markerfacecolor='r', marker='s'))
axs[0, 1].boxplot(perceptron_scores_test, flierprops=dict(markerfacecolor='r', marker='s'))
axs[0, 2].boxplot(nb_scores_test, flierprops=dict(markerfacecolor='r', marker='s'))
axs[1, 0].boxplot(lr_scores_test, flierprops=dict(markerfacecolor='r', marker='s'))
axs[1, 1].boxplot(ann_scores_test, flierprops=dict(markerfacecolor='r', marker='s'))
axs[1, 2].boxplot(svc_scores_test, flierprops=dict(markerfacecolor='r', marker='s'))
axs[0, 0].set_title(f'Linear Discriminant Analysis: {round(np.mean(linear_disc_scores_test), 6) * 100}')
axs[0, 1].set_title(f'Perceptron: {round(np.mean(perceptron_scores_test), 6) * 100}')
axs[0, 2].set_title(f'Naïve Bayes: {round(np.mean(nb_scores_test), 6) * 100}')
axs[1, 0].set_title(f'Logistic Regression: {round(np.mean(lr_scores_test), 6) * 100}')
axs[1, 1].set_title(f'Artificial Neural Networks: {round(np.mean(ann_scores_test), 6) * 100}')
axs[1, 2].set_title(f'Support Vector Classifier: {round(np.mean(svc_scores_test), 6) * 100}')
```
| github_jupyter |
```
import scipy as sp
import numpy as np
import time
try:
from localgraphclustering import *
except:
# when the package is not installed, import the local version instead.
# the notebook must be placed in the original "notebooks/" folder
sys.path.append("../")
from localgraphclustering import *
import time
import networkx as nx
import random
import statistics as stat_
```
## Load data
```
g = GraphLocal('../datasets/sfld_brown_et_al_amidohydrolases_protein_similarities_for_beh.graphml','graphml',' ')
```
## TEMP
```
G = nx.read_graphml('../datasets/sfld_brown_et_al_amidohydrolases_protein_similarities_for_beh.graphml')
# groups = np.loadtxt('./datasets/ppi_mips.class', dtype = 'float')
groups = np.loadtxt('../datasets/sfld_brown_et_al_amidohydrolases_protein_similarities_for_beh_ground_truth.csv', dtype = 'str')
groups_by_id = dict()
for node in groups:
groups_by_id[node[0]] = node[1]
ids_clusters = set()
for node in groups:
ids_clusters.add(node[1])
ids_clusters = list(ids_clusters)
ground_truth_clusters_by_id = dict()
for node in groups:
ground_truth_clusters_by_id[node[1]] = []
for node in groups:
ground_truth_clusters_by_id[node[1]].append(node[0])
ground_truth_clusters_by_number = dict()
for node in groups:
ground_truth_clusters_by_number[node[1]] = []
counter = 0
for node in G.node:
if node == '1.0':
counter += 1
continue
what_group = groups_by_id[node]
ground_truth_clusters_by_number[what_group].append(counter)
counter += 1
all_clusters = []
counter = 0
for cluster_id in ground_truth_clusters_by_number:
cluster = ground_truth_clusters_by_number[cluster_id]
if len(cluster) == 1 or len(cluster) == 0:
counter += 1
continue
cond = g.compute_conductance(cluster)
counter += 1
if cond <= 0.57 and len(cluster) >= 10:
print("Id: ", cluster_id)
print("Cluster: ", counter, " conductance: ", cond, "Size: ", len(cluster))
all_clusters.append(cluster)
```
## Collect data for l1-reg. PR (with rounding)
```
nodes = {}
external_best_cond_acl = {}
external_best_pre_cond_acl = {}
vol_best_cond_acl = {}
vol_best_pre_acl = {}
size_clust_best_cond_acl = {}
size_clust_best_pre_acl = {}
f1score_best_cond_acl = {}
f1score_best_pre_acl = {}
true_positives_best_cond_acl = {}
true_positives_best_pre_acl = {}
precision_best_cond_acl = {}
precision_best_pre_acl = {}
recall_best_cond_acl = {}
recall_best_pre_acl = {}
cuts_best_cond_acl = {}
cuts_best_pre_acl = {}
cuts_acl_ALL = {}
ct_outer = 0
number_experiments = 0
for rr in all_clusters:
how_many = int(len(rr))
print(how_many)
random.seed(4)
nodes[ct_outer] = np.random.choice(rr, how_many, replace=False)
eigv, lambda_val = fiedler_local(g, rr)
lambda_val = np.real(lambda_val)
step = (2*lambda_val - lambda_val/2)/4
a_list = np.arange(lambda_val/2,2*lambda_val,step)
ct = 0
start = time.time()
for node in nodes[ct_outer]:
ref_node = [node]
max_precision = -1
min_conduct = 100
ct_inner = 0
for a in a_list:
if ct_outer <= 1:
rho = 0.15/np.sum(g.d[rr])
else:
rho = 0.2/np.sum(g.d[rr])
output_pr_clustering = approximate_PageRank(g,ref_node,method = "l1reg-rand", epsilon=1.0e-2, rho=rho, alpha=a, cpp = True, normalize=True,normalized_objective=True)
number_experiments += 1
output_pr_sc = sweep_cut(g,output_pr_clustering,cpp=True)
S = output_pr_sc[0]
cuts_acl_ALL[ct_outer,node,ct_inner] = S
size_clust_acl_ = len(S)
cond_val_l1pr = g.compute_conductance(S)
vol_ = sum(g.d[S])
true_positives_acl_ = set(rr).intersection(S)
if len(true_positives_acl_) == 0:
true_positives_acl_ = set(ref_node)
vol_ = g.d[ref_node][0,0]
precision = sum(g.d[np.array(list(true_positives_acl_))])/vol_
recall = sum(g.d[np.array(list(true_positives_acl_))])/sum(g.d[rr])
f1_score_ = 2*(precision*recall)/(precision + recall)
if f1_score_ >= max_precision:
max_precision = f1_score_
external_best_pre_cond_acl[ct_outer,node] = cond_val_l1pr
vol_best_pre_acl[ct_outer,node] = vol_
size_clust_best_pre_acl[ct_outer,node] = size_clust_acl_
true_positives_best_pre_acl[ct_outer,node] = true_positives_acl_
precision_best_pre_acl[ct_outer,node] = precision
recall_best_pre_acl[ct_outer,node] = recall
f1score_best_pre_acl[ct_outer,node] = f1_score_
cuts_best_pre_acl[ct_outer,node] = S
if cond_val_l1pr <= min_conduct:
min_conduct = cond_val_l1pr
external_best_cond_acl[ct_outer,node] = cond_val_l1pr
vol_best_cond_acl[ct_outer,node] = vol_
size_clust_best_cond_acl[ct_outer,node] = size_clust_acl_
true_positives_best_cond_acl[ct_outer,node] = true_positives_acl_
precision_best_cond_acl[ct_outer,node] = precision
recall_best_cond_acl[ct_outer,node] = recall
f1score_best_cond_acl[ct_outer,node] = f1_score_
cuts_best_cond_acl[ct_outer,node] = S
print('outer:', ct_outer, 'number of node: ',node, ' completed: ', ct/how_many, ' degree: ', g.d[node])
print('conductance: ', external_best_cond_acl[ct_outer,node], 'f1score: ', f1score_best_cond_acl[ct_outer,node], 'precision: ', precision_best_cond_acl[ct_outer,node], 'recall: ', recall_best_cond_acl[ct_outer,node])
ct += 1
end = time.time()
print(" ")
print("Outer: ", ct_outer," Elapsed time l1-reg. with rounding: ", end - start)
print("Outer: ", ct_outer," Number of experiments: ", number_experiments)
print(" ")
ct_outer += 1
```
## Performance of l1-reg. PR (with rounding).
```
all_data = []
xlabels_ = []
print('Results for l1-reg with rounding')
sum_precision = 0
sum_recall = 0
sum_f1 = 0
sum_conductance = 0
info_ref_nodes = all_clusters
l_info_ref_nodes = len(info_ref_nodes)
for i in range(l_info_ref_nodes):
temp_pre = []
temp_rec = []
temp_f1 = []
temp_conductance = []
for j in all_clusters[i]:
temp_pre.append(precision_best_cond_acl[i,j])
temp_rec.append(recall_best_cond_acl[i,j])
temp_f1.append(f1score_best_cond_acl[i,j])
temp_conductance.append(external_best_cond_acl[i,j])
print('Feature:', i,'Precision', stat_.mean(temp_pre), 'Recall', stat_.mean(temp_rec), 'F1', stat_.mean(temp_f1), 'Cond.', stat_.mean(temp_conductance))
```
## Function for seed set expansion using BFS
```
import queue
def seed_grow_bfs_steps(g,seeds,steps,vol_target,target_cluster):
"""
grow the initial seed set through BFS until its size reaches
a given ratio of the total number of nodes.
"""
Q = queue.Queue()
visited = np.zeros(g._num_vertices)
visited[seeds] = 1
for s in seeds:
Q.put(s)
if isinstance(seeds,np.ndarray):
seeds = seeds.tolist()
else:
seeds = list(seeds)
for step in range(steps):
for k in range(Q.qsize()):
node = Q.get()
si,ei = g.adjacency_matrix.indptr[node],g.adjacency_matrix.indptr[node+1]
neighs = g.adjacency_matrix.indices[si:ei]
for i in range(len(neighs)):
if visited[neighs[i]] == 0:
visited[neighs[i]] = 1
seeds.append(neighs[i])
Q.put(neighs[i])
vol_seeds = np.sum(g.d[seeds])
vol_target_intersection_input = np.sum(g.d[list(set(target_cluster).intersection(set(seeds)))])
sigma = vol_target_intersection_input/vol_target
if sigma > 0.75 or vol_seeds > 0.25*g.vol_G:
break
vol_seeds = np.sum(g.d[seeds])
vol_target_intersection_input = np.sum(g.d[list(set(target_cluster).intersection(set(seeds)))])
sigma = vol_target_intersection_input/vol_target
if sigma > 0.75 or vol_seeds > 0.25*g.vol_G:
break
vol_seeds = np.sum(g.d[seeds])
vol_target_intersection_input = np.sum(g.d[list(set(target_cluster).intersection(set(seeds)))])
sigma = vol_target_intersection_input/vol_target
if sigma > 0.75 or vol_seeds > 0.25*g.vol_G:
break
return seeds
```
## Collect data for seed set expansion + FlowImprove, try a lot of parameters
```
nodes = {}
external_best_cond_flBFS = {}
external_best_pre_cond_flBFS = {}
vol_best_cond_flBFS = {}
vol_best_pre_flBFS = {}
size_clust_best_cond_flBFS = {}
size_clust_best_pre_flBFS = {}
f1score_best_cond_flBFS = {}
f1score_best_pre_flBFS = {}
true_positives_best_cond_flBFS = {}
true_positives_best_pre_flBFS = {}
precision_best_cond_flBFS = {}
precision_best_pre_flBFS = {}
recall_best_cond_flBFS = {}
recall_best_pre_flBFS = {}
cuts_best_cond_flBFS = {}
cuts_best_pre_flBFS = {}
cuts_flBFS_ALL = {}
ct_outer = 0
number_experiments = 0
for rr in all_clusters:
how_many = int(len(rr))
print(how_many)
random.seed(4)
nodes[ct_outer] = np.random.choice(rr, how_many, replace=False)
n_step = 24
vol_target = np.sum(g.d[rr])
ct = 0
start = time.time()
for node in nodes[ct_outer]:
ref_node = [node]
max_precision = -1
min_conduct = 100
seeds = seed_grow_bfs_steps(g,[node],g._num_vertices,vol_target,rr)
vol_input = np.sum(g.d[seeds])
vol_graph_minus_input = np.sum(g.d[list(set(range(g._num_vertices)) - set(seeds))])
vol_target_intersection_input = np.sum(g.d[list(set(rr).intersection(set(seeds)))])
gamma = vol_input/vol_graph_minus_input
sigma = max(vol_target_intersection_input/vol_target,gamma)
delta = min(max((1/3)*(1.0/(1.0/sigma - 1)) - gamma,0),1)
S = flow_clustering(g,seeds,method="sl",delta=delta)[0]
number_experiments += 1
cuts_flBFS_ALL[ct_outer,node] = S
size_clust_flBFS_ = len(S)
cond_val_l1pr = g.compute_conductance(S)
vol_ = sum(g.d[S])
true_positives_flBFS_ = set(rr).intersection(S)
if len(true_positives_flBFS_) == 0:
true_positives_flBFS_ = set(ref_node)
vol_ = g.d[ref_node][0]
precision = sum(g.d[np.array(list(true_positives_flBFS_))])/vol_
recall = sum(g.d[np.array(list(true_positives_flBFS_))])/sum(g.d[rr])
f1_score_ = 2*(precision*recall)/(precision + recall)
if f1_score_ >= max_precision:
max_precision = f1_score_
external_best_pre_cond_flBFS[ct_outer,node] = cond_val_l1pr
vol_best_pre_flBFS[ct_outer,node] = vol_
size_clust_best_pre_flBFS[ct_outer,node] = size_clust_flBFS_
true_positives_best_pre_flBFS[ct_outer,node] = true_positives_flBFS_
precision_best_pre_flBFS[ct_outer,node] = precision
recall_best_pre_flBFS[ct_outer,node] = recall
f1score_best_pre_flBFS[ct_outer,node] = f1_score_
cuts_best_pre_flBFS[ct_outer,node] = S
if cond_val_l1pr <= min_conduct:
min_conduct = cond_val_l1pr
external_best_cond_flBFS[ct_outer,node] = cond_val_l1pr
vol_best_cond_flBFS[ct_outer,node] = vol_
size_clust_best_cond_flBFS[ct_outer,node] = size_clust_flBFS_
true_positives_best_cond_flBFS[ct_outer,node] = true_positives_flBFS_
precision_best_cond_flBFS[ct_outer,node] = precision
recall_best_cond_flBFS[ct_outer,node] = recall
f1score_best_cond_flBFS[ct_outer,node] = f1_score_
cuts_best_cond_flBFS[ct_outer,node] = S
print('outer:', ct_outer, 'number of node: ',node, ' completed: ', ct/how_many, ' degree: ', g.d[node])
print('conductance: ', external_best_cond_flBFS[ct_outer,node], 'f1score: ', f1score_best_cond_flBFS[ct_outer,node], 'precision: ', precision_best_cond_flBFS[ct_outer,node], 'recall: ', recall_best_cond_flBFS[ct_outer,node])
ct += 1
end = time.time()
print(" ")
print("Outer: ", ct_outer," Elapsed time BFS+SL: ", end - start)
print("Outer: ", ct_outer," Number of experiments: ", number_experiments)
print(" ")
ct_outer += 1
```
## Performance of BFS+FlowImp.
```
all_data = []
xlabels_ = []
print('Results for BFS+SL')
sum_precision = 0
sum_recall = 0
sum_f1 = 0
sum_conductance = 0
info_ref_nodes = all_clusters
l_info_ref_nodes = len(info_ref_nodes)
for i in range(l_info_ref_nodes):
temp_pre = []
temp_rec = []
temp_f1 = []
temp_conductance = []
for j in all_clusters[i]:
temp_pre.append(precision_best_cond_flBFS[i,j])
temp_rec.append(recall_best_cond_flBFS[i,j])
temp_f1.append(f1score_best_cond_flBFS[i,j])
temp_conductance.append(external_best_cond_flBFS[i,j])
print('Feature:', i,'Precision', stat_.mean(temp_pre), 'Recall', stat_.mean(temp_rec), 'F1', stat_.mean(temp_f1), 'Cond.', stat_.mean(temp_conductance))
```
## Collect data for L1+SL
```
nodes = {}
external_best_cond_l1SL = {}
external_best_pre_cond_l1SL = {}
vol_best_cond_l1SL = {}
vol_best_pre_l1SL = {}
size_clust_best_cond_l1SL = {}
size_clust_best_pre_l1SL = {}
f1score_best_cond_l1SL = {}
f1score_best_pre_l1SL = {}
true_positives_best_cond_l1SL = {}
true_positives_best_pre_l1SL = {}
precision_best_cond_l1SL = {}
precision_best_pre_l1SL = {}
recall_best_cond_l1SL = {}
recall_best_pre_l1SL = {}
cuts_best_cond_l1SL = {}
cuts_best_pre_l1SL = {}
cuts_l1SL_ALL = {}
ct_outer = 0
number_experiments = 0
for rr in all_clusters:
how_many = int(len(rr))
print(how_many)
random.seed(4)
nodes[ct_outer] = np.random.choice(rr, how_many, replace=False)
eigv, lambda_val = fiedler_local(g, rr)
lambda_val = np.real(lambda_val)
step = (2*lambda_val - lambda_val/2)/4
a_list = np.arange(lambda_val/2,2*lambda_val,step)
vol_target = np.sum(g.d[rr])
ct = 0
start = time.time()
for node in nodes[ct_outer]:
ref_node = [node]
max_precision = -1
min_conduct = 100
ct_inner = 0
for a in a_list:
if ct_outer <= 1:
rho = 0.15/np.sum(g.d[rr])
else:
rho = 0.2/np.sum(g.d[rr])
output_pr_clustering = approximate_PageRank(g,ref_node,method = "l1reg-rand", epsilon=1.0e-2, rho=rho, alpha=a, cpp = True, normalize=True,normalized_objective=True)
number_experiments += 1
output_pr_sc = sweep_cut(g,output_pr_clustering,cpp=True)
S = output_pr_sc[0]
vol_input = np.sum(g.d[S])
vol_graph_minus_input = np.sum(g.d[list(set(range(g._num_vertices)) - set(S))])
vol_target_intersection_input = np.sum(g.d[list(set(rr).intersection(set(S)))])
gamma = vol_input/vol_graph_minus_input
sigma = max(vol_target_intersection_input/vol_target,gamma)
delta = min(max((1/3)*(1.0/(1.0/sigma - 1)) - gamma,0),1)
S = flow_clustering(g,S,method="sl",delta=delta)[0]
cuts_l1SL_ALL[ct_outer,node,ct_inner] = S
size_clust_l1SL_ = len(S)
cond_val_l1pr = g.compute_conductance(S)
vol_ = sum(g.d[S])
true_positives_l1SL_ = set(rr).intersection(S)
if len(true_positives_l1SL_) == 0:
true_positives_l1SL_ = set(ref_node)
vol_ = g.d[ref_node][0]
precision = sum(g.d[np.array(list(true_positives_l1SL_))])/vol_
recall = sum(g.d[np.array(list(true_positives_l1SL_))])/sum(g.d[rr])
f1_score_ = 2*(precision*recall)/(precision + recall)
if f1_score_ >= max_precision:
max_precision = f1_score_
external_best_pre_cond_l1SL[ct_outer,node] = cond_val_l1pr
vol_best_pre_l1SL[ct_outer,node] = vol_
size_clust_best_pre_l1SL[ct_outer,node] = size_clust_l1SL_
true_positives_best_pre_l1SL[ct_outer,node] = true_positives_l1SL_
precision_best_pre_l1SL[ct_outer,node] = precision
recall_best_pre_l1SL[ct_outer,node] = recall
f1score_best_pre_l1SL[ct_outer,node] = f1_score_
cuts_best_pre_l1SL[ct_outer,node] = S
if cond_val_l1pr <= min_conduct:
min_conduct = cond_val_l1pr
external_best_cond_l1SL[ct_outer,node] = cond_val_l1pr
vol_best_cond_l1SL[ct_outer,node] = vol_
size_clust_best_cond_l1SL[ct_outer,node] = size_clust_l1SL_
true_positives_best_cond_l1SL[ct_outer,node] = true_positives_l1SL_
precision_best_cond_l1SL[ct_outer,node] = precision
recall_best_cond_l1SL[ct_outer,node] = recall
f1score_best_cond_l1SL[ct_outer,node] = f1_score_
cuts_best_cond_l1SL[ct_outer,node] = S
print('outer:', ct_outer, 'number of node: ',node, ' completed: ', ct/how_many, ' degree: ', g.d[node])
print('conductance: ', external_best_cond_l1SL[ct_outer,node], 'f1score: ', f1score_best_cond_l1SL[ct_outer,node], 'precision: ', precision_best_cond_l1SL[ct_outer,node], 'recall: ', recall_best_cond_l1SL[ct_outer,node])
ct += 1
end = time.time()
print(" ")
print("Outer: ", ct_outer," Elapsed time L1+SL with rounding: ", end - start)
print("Outer: ", ct_outer," Number of experiments: ", number_experiments)
print(" ")
ct_outer += 1
```
## Performance of l1+SL
```
all_data = []
xlabels_ = []
print('Results for L1+SL')
sum_precision = 0
sum_recall = 0
sum_f1 = 0
sum_conductance = 0
info_ref_nodes = all_clusters
l_info_ref_nodes = len(info_ref_nodes)
for i in range(l_info_ref_nodes):
temp_pre = []
temp_rec = []
temp_f1 = []
temp_conductance = []
for j in all_clusters[i]:
temp_pre.append(precision_best_cond_l1SL[i,j])
temp_rec.append(recall_best_cond_l1SL[i,j])
temp_f1.append(f1score_best_cond_l1SL[i,j])
temp_conductance.append(external_best_cond_l1SL[i,j])
print('Feature:', i,'Precision', stat_.mean(temp_pre), 'Recall', stat_.mean(temp_rec), 'F1', stat_.mean(temp_f1), 'Cond.', stat_.mean(temp_conductance))
```
| github_jupyter |
```
!wget https://gitlab.com/federicozzo/electiveai/raw/master/Desktop/uni/elective_AI/electiveai/bdd100K_img.zip?inline=false
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow as tf
import numpy as np
import IPython.display as display
import cv2
import json
import os
from tqdm import tqdm
import matplotlib.pyplot as plt
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
drive.flush_and_unmount()
with open("/content/bdd100k/labels/bdd100k_labels_images_train.json", "r") as f:
train_images_label = json.load(f)
with open("/content/bdd100k/labels/bdd100k_labels_images_val.json", "r") as f:
test_images_label = json.load(f)
def _int64_feature(value):
return tf.train.Feature(int64_list=tf.train.Int64List(value=[value]))
def _bytes_feature(value):
return tf.train.Feature(bytes_list=tf.train.BytesList(value=[value]))
def load_image(addr):
# read an image and resize to (128, 128)
# cv2 load images as BGR, convert it to RGB
img = cv2.imread(addr)
img = cv2.resize(img, (128, 128), interpolation=cv2.INTER_CUBIC)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
img = img.astype(np.float32)
return img
def find_img_in_labels(name):
name = name.split("/")[-1]
for image in train_images_label:
if image['name'] == name:
return image
return None
def find_img_in_test_labels(name):
name = name.split("/")[-1]
for image in test_images_label:
if image['name'] == name:
return image
return None
for image in train_images_label:
if image['attributes']['timeofday'] not in ['daytime', 'night', 'dawn/dusk', 'undefined']:
print(image['name'], image['attributes']['timeofday'])
def writeTfRecord(input_path, output_path, setname):
# open the TFRecords file
writer = tf.python_io.TFRecordWriter(output_path)
images_filenames = [input_path+fn for fn in os.listdir(path=input_path)]
for image_fn in tqdm(images_filenames):
# Load the image
img = load_image(image_fn)
if setname == 'train':
attributes = find_img_in_labels(image_fn)
else:
attributes = find_img_in_test_labels(image_fn)
if attributes is None:
print(image_fn)
continue
label = attributes['attributes']['timeofday']
if label not in ['daytime', 'night']:
continue
# Create a feature
feature = {'label': _bytes_feature(tf.compat.as_bytes(label)),
'image': _bytes_feature(tf.compat.as_bytes(img.tostring()))}
# Create an example protocol buffer
example = tf.train.Example(features=tf.train.Features(feature=feature))
# Serialize to string and write on the file
writer.write(example.SerializeToString())
writer.close()
train_path = '/content/bdd100k/images/100k/train/'
test_path = '/content/bdd100k/images/100k/test/'
val_path = '/content/bdd100k/images/100k/val/'
train_out = 'train.tfrecords'
test_out = 'test.tfrecords'
writeTfRecord(train_path, train_out)
writeTfRecord(val_path, test_out, 'val')
```
# ***Read from TfRecords File***
```
sess = tf.InteractiveSession()
### IMPORTANT : First you have to unzip the tfRecord to import! ###
tfrecord_path = 'test.tfrecords'
dataset = tf.data.TFRecordDataset(tfrecord_path)
def decode(serialized_example):
"""
Parses an image and label from the given `serialized_example`.
It is used as a map function for `dataset.map`
"""
IMAGE_SHAPE = (128,128,3)
# 1. define a parser
features = tf.parse_single_example(
serialized_example,
# Defaults are not specified since both keys are required.
features={
'image': tf.FixedLenFeature([], tf.string),
'label': tf.FixedLenFeature([], tf.string),
})
# 2. Convert the data
image = tf.decode_raw(features['image'], tf.float32)
label = features['label']
# 3. reshape
image = tf.convert_to_tensor(tf.reshape(image, IMAGE_SHAPE))
return image, label
def normalize(image, label):
"""Convert `image` from [0, 255] -> [-0.5, 0.5] floats."""
image = tf.cast(image, tf.float32) * (1. / 255)
return image, label
# Parse the record into tensors with map.
# map takes a Python function and applies it to every sample.
dataset = dataset.map(decode)
dataset = dataset.map(normalize)
batch_size = 1000
dataset = dataset.batch(batch_size)
# Creating an iterator
iterator = dataset.make_one_shot_iterator()
image_batch, label_batch = iterator.get_next()
image_batch, label_batch = sess.run([image_batch, label_batch])
print(image_batch.shape)
print(label_batch.shape)
plt.imshow(image_batch[0])
```
| github_jupyter |
##### 1

##### 2

##### 3

##### 4

##### 5

##### 6

##### 7

##### 8

##### 9

##### 10

##### 11

##### 12

##### 13

##### 14

##### 15

##### 16

##### 17

##### 18

##### 19

##### 20

##### 21

##### 22

##### 23

##### 24

##### 25

##### 26

##### 27

##### 28

##### 29

##### 30

##### 31

##### 32

##### 33

##### 34

##### 35

##### 36

##### 37

| github_jupyter |
# Loading Image Data
So far we've been working with fairly artificial datasets that you wouldn't typically be using in real projects. Instead, you'll likely be dealing with full-sized images like you'd get from smart phone cameras. In this notebook, we'll look at how to load images and use them to train neural networks.
We'll be using a [dataset of cat and dog photos](https://www.kaggle.com/c/dogs-vs-cats) available from Kaggle. Here are a couple example images:
<img src='assets/dog_cat.png'>
We'll use this dataset to train a neural network that can differentiate between cats and dogs. These days it doesn't seem like a big accomplishment, but five years ago it was a serious challenge for computer vision systems.
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
import torch
from torchvision import datasets, transforms
import helper
```
The easiest way to load image data is with `datasets.ImageFolder` from `torchvision` ([documentation](http://pytorch.org/docs/master/torchvision/datasets.html#imagefolder)). In general you'll use `ImageFolder` like so:
```python
dataset = datasets.ImageFolder('path/to/data', transform=transforms)
```
where `'path/to/data'` is the file path to the data directory and `transforms` is a list of processing steps built with the [`transforms`](http://pytorch.org/docs/master/torchvision/transforms.html) module from `torchvision`. ImageFolder expects the files and directories to be constructed like so:
```
root/dog/xxx.png
root/dog/xxy.png
root/dog/xxz.png
root/cat/123.png
root/cat/nsdf3.png
root/cat/asd932_.png
```
where each class has it's own directory (`cat` and `dog`) for the images. The images are then labeled with the class taken from the directory name. So here, the image `123.png` would be loaded with the class label `cat`. You can download the dataset already structured like this [from here](https://s3.amazonaws.com/content.udacity-data.com/nd089/Cat_Dog_data.zip). I've also split it into a training set and test set.
### Transforms
When you load in the data with `ImageFolder`, you'll need to define some transforms. For example, the images are different sizes but we'll need them to all be the same size for training. You can either resize them with `transforms.Resize()` or crop with `transforms.CenterCrop()`, `transforms.RandomResizedCrop()`, etc. We'll also need to convert the images to PyTorch tensors with `transforms.ToTensor()`. Typically you'll combine these transforms into a pipeline with `transforms.Compose()`, which accepts a list of transforms and runs them in sequence. It looks something like this to scale, then crop, then convert to a tensor:
```python
transforms = transforms.Compose([transforms.Resize(255),
transforms.CenterCrop(224),
transforms.ToTensor()])
```
There are plenty of transforms available, I'll cover more in a bit and you can read through the [documentation](http://pytorch.org/docs/master/torchvision/transforms.html).
### Data Loaders
With the `ImageFolder` loaded, you have to pass it to a [`DataLoader`](http://pytorch.org/docs/master/data.html#torch.utils.data.DataLoader). The `DataLoader` takes a dataset (such as you would get from `ImageFolder`) and returns batches of images and the corresponding labels. You can set various parameters like the batch size and if the data is shuffled after each epoch.
```python
dataloader = torch.utils.data.DataLoader(dataset, batch_size=32, shuffle=True)
```
Here `dataloader` is a [generator](https://jeffknupp.com/blog/2013/04/07/improve-your-python-yield-and-generators-explained/). To get data out of it, you need to loop through it or convert it to an iterator and call `next()`.
```python
# Looping through it, get a batch on each loop
for images, labels in dataloader:
pass
# Get one batch
images, labels = next(iter(dataloader))
```
>**Exercise:** Load images from the `Cat_Dog_data/train` folder, define a few transforms, then build the dataloader.
```
data_dir = '/home/cs/Downloads/Cat_Dog_data/train'
# TODO: compose transforms here
data_transforms = transforms.Compose([transforms.Resize(255),
transforms.CenterCrop(224),
transforms.ToTensor()])
# TODO: create the ImageFolder
dataset = datasets.ImageFolder(root=data_dir, transform=data_transforms)
# TODO: use the ImageFolder dataset to create the DataLoader
dataloader = torch.utils.data.DataLoader(dataset, batch_size=32, shuffle=True)
# Run this to test your data loader
images, labels = next(iter(dataloader))
helper.imshow(images[0], normalize=False)
```
If you loaded the data correctly, you should see something like this (your image will be different):
<img src='assets/cat_cropped.png', width=244>
## Data Augmentation
A common strategy for training neural networks is to introduce randomness in the input data itself. For example, you can randomly rotate, mirror, scale, and/or crop your images during training. This will help your network generalize as it's seeing the same images but in different locations, with different sizes, in different orientations, etc.
To randomly rotate, scale and crop, then flip your images you would define your transforms like this:
```python
train_transforms = transforms.Compose([transforms.RandomRotation(30),
transforms.RandomResizedCrop(100),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.5, 0.5, 0.5],
[0.5, 0.5, 0.5])])
```
You'll also typically want to normalize images with `transforms.Normalize`. You pass in a list of means and list of standard deviations, then the color channels are normalized like so
```input[channel] = (input[channel] - mean[channel]) / std[channel]```
Subtracting `mean` centers the data around zero and dividing by `std` squishes the values to be between -1 and 1. Normalizing helps keep the network work weights near zero which in turn makes backpropagation more stable. Without normalization, networks will tend to fail to learn.
You can find a list of all [the available transforms here](http://pytorch.org/docs/0.3.0/torchvision/transforms.html). When you're testing however, you'll want to use images that aren't altered (except you'll need to normalize the same way). So, for validation/test images, you'll typically just resize and crop.
>**Exercise:** Define transforms for training data and testing data below.
```
data_dir = '/home/cs/Downloads/Cat_Dog_data'
# TODO: Define transforms for the training data and testing data
train_transforms = transforms.Compose([transforms.RandomRotation(30),
transforms.RandomResizedCrop(100),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.5, 0.5, 0.5],
[0.5, 0.5, 0.5])])
test_transforms = transforms.Compose([transforms.Resize(255),
transforms.CenterCrop(224),
transforms.ToTensor()])
# Pass transforms in here, then run the next cell to see how the transforms look
train_data = datasets.ImageFolder(data_dir + '/train', transform=train_transforms)
test_data = datasets.ImageFolder(data_dir + '/test', transform=test_transforms)
trainloader = torch.utils.data.DataLoader(train_data, batch_size=32)
testloader = torch.utils.data.DataLoader(test_data, batch_size=32)
# change this to the trainloader or testloader
data_iter = iter(trainloader)
images, labels = next(data_iter)
fig, axes = plt.subplots(figsize=(10,4), ncols=4)
for ii in range(4):
ax = axes[ii]
helper.imshow(images[ii], ax=ax)
```
Your transformed images should look something like this.
<center>Training examples:</center>
<img src='assets/train_examples.png' width=500px>
<center>Testing examples:</center>
<img src='assets/test_examples.png' width=500px>
At this point you should be able to load data for training and testing. Now, you should try building a network that can classify cats vs dogs. This is quite a bit more complicated than before with the MNIST and Fashion-MNIST datasets. To be honest, you probably won't get it to work with a fully-connected network, no matter how deep. These images have three color channels and at a higher resolution (so far you've seen 28x28 images which are tiny).
In the next part, I'll show you how to use a pre-trained network to build a model that can actually solve this problem.
```
# Optional TODO: Attempt to build a network to classify cats vs dogs from this dataset
```
| github_jupyter |
## 📍 The Data
This example considers a hierarchical dataset. The world is split by continents. Continents are split by country. Each country has a value (population size). Our goal is to represent each country as a circle, its size being proportional to its population.
Let's create such a dataset:
```
data = [{'id': 'World', 'datum': 6964195249, 'children' : [
{'id' : "North America", 'datum': 450448697,
'children' : [
{'id' : "United States", 'datum' : 308865000},
{'id' : "Mexico", 'datum' : 107550697},
{'id' : "Canada", 'datum' : 34033000}
]},
{'id' : "South America", 'datum' : 278095425,
'children' : [
{'id' : "Brazil", 'datum' : 192612000},
{'id' : "Colombia", 'datum' : 45349000},
{'id' : "Argentina", 'datum' : 40134425}
]},
{'id' : "Europe", 'datum' : 209246682,
'children' : [
{'id' : "Germany", 'datum' : 81757600},
{'id' : "France", 'datum' : 65447374},
{'id' : "United Kingdom", 'datum' : 62041708}
]},
{'id' : "Africa", 'datum' : 311929000,
'children' : [
{'id' : "Nigeria", 'datum' : 154729000},
{'id' : "Ethiopia", 'datum' : 79221000},
{'id' : "Egypt", 'datum' : 77979000}
]},
{'id' : "Asia", 'datum' : 2745929500,
'children' : [
{'id' : "China", 'datum' : 1336335000},
{'id' : "India", 'datum' : 1178225000},
{'id' : "Indonesia", 'datum' : 231369500}
]}
]}]
```
## 🙇♂️ Compute circle position
We need an algorythm that computes the position of each country and continent circles, together with their radius. Fortunately, the `circlize` library is here. It's `circlify()` function does exactly that 😍
```
# import the circlify library
import circlify
# Compute circle positions thanks to the circlify() function
circles = circlify.circlify(
data,
show_enclosure=False,
target_enclosure=circlify.Circle(x=0, y=0, r=1)
)
```
Have a look to the `circles` object, it provides exactly that 🎉.
## 🔨 Build the viz
Let's be honnest, that's quite a bit of code to get a decent graph 😞. The `circlize` library has a `bubble()` function that allows to do a simple circle pack with one line of code, but it does not allow to customize the chart.
So once more `matplotlib` is our best friend for the rendering part. Here I'm printing the layers from the bottom to the top of the figure: first the cirles for the highest level of hierarchy (continent), then circle and labels for countries, then continent labels.
```
# import libraries
import circlify
import matplotlib.pyplot as plt
# Create just a figure and only one subplot
fig, ax = plt.subplots(figsize=(14,14))
# Title
ax.set_title('Repartition of the world population')
# Remove axes
ax.axis('off')
# Find axis boundaries
lim = max(
max(
abs(circle.x) + circle.r,
abs(circle.y) + circle.r,
)
for circle in circles
)
plt.xlim(-lim, lim)
plt.ylim(-lim, lim)
# Print circle the highest level (continents):
for circle in circles:
if circle.level != 2:
continue
x, y, r = circle
ax.add_patch( plt.Circle((x, y), r, alpha=0.5, linewidth=2, color="lightblue"))
# Print circle and labels for the highest level:
for circle in circles:
if circle.level != 3:
continue
x, y, r = circle
label = circle.ex["id"]
ax.add_patch( plt.Circle((x, y), r, alpha=0.5, linewidth=2, color="#69b3a2"))
plt.annotate(label, (x,y ), ha='center', color="white")
# Print labels for the continents
for circle in circles:
if circle.level != 2:
continue
x, y, r = circle
label = circle.ex["id"]
plt.annotate(label, (x,y ) ,va='center', ha='center', bbox=dict(facecolor='white', edgecolor='black', boxstyle='round', pad=.5))
```
| github_jupyter |
# Proyecto
## Instrucciones
1.- Completa los datos personales (nombre y rol USM) de cada integrante en siguiente celda.
* __Nombre-Rol__:
*
*
*
*
*
2.- Debes _pushear_ este archivo con tus cambios a tu repositorio personal del curso, incluyendo datos, imágenes, scripts, etc.
3.- Se evaluará:
- Soluciones
- Código
- Que Binder esté bien configurado.
- Al presionar `Kernel -> Restart Kernel and Run All Cells` deben ejecutarse todas las celdas sin error.
## I.- Sistemas de recomendación
### Introducción
El rápido crecimiento de la recopilación de datos ha dado lugar a una nueva era de información. Los datos se están utilizando para crear sistemas más eficientes y aquí es donde entran en juego los sistemas de recomendación. Los sistemas de recomendación son un tipo de sistemas de filtrado de información, ya que mejoran la calidad de los resultados de búsqueda y proporcionan elementos que son más relevantes para el elemento de búsqueda o están relacionados con el historial de búsqueda del usuario.
Se utilizan para predecir la calificación o preferencia que un usuario le daría a un artículo. Casi todas las grandes empresas de tecnología los han aplicado de una forma u otra: Amazon lo usa para sugerir productos a los clientes, YouTube lo usa para decidir qué video reproducir a continuación en reproducción automática y Facebook lo usa para recomendar páginas que me gusten y personas a seguir. Además, empresas como Netflix y Spotify dependen en gran medida de la efectividad de sus motores de recomendación para sus negocios y éxitos.
### Objetivos
Poder realizar un proyecto de principio a fin ocupando todos los conocimientos aprendidos en clase. Para ello deben cumplir con los siguientes objetivos:
* **Desarrollo del problema**: Se les pide a partir de los datos, proponer al menos un tipo de sistemas de recomendación. Como todo buen proyecto de Machine Learning deben seguir el siguiente procedimiento:
* **Lectura de los datos**: Describir el o los conjunto de datos en estudio.
* **Procesamiento de los datos**: Procesar adecuadamente los datos en estudio. Para este caso ocuparan técnicas de [NLP](https://en.wikipedia.org/wiki/Natural_language_processing).
* **Metodología**: Describir adecuadamente el procedimiento ocupado en cada uno de los modelos ocupados.
* **Resultados**: Evaluar adecuadamente cada una de las métricas propuesta en este tipo de problemas.
* **Presentación**: La presentación será levemente distinta a las anteriores, puesto que deberán ocupar la herramienta de Jupyter llamada [RISE](https://en.wikipedia.org/wiki/Natural_language_processing). Esta presentación debe durar aproximadamente entre 15-30 minutos, y deberán mandar sus videos (por youtube, google drive, etc.)
### Evaluación
* **Códigos**: Los códigos deben estar correctamente documentados (ocupando las *buenas prácticas* de python aprendidas en este curso).
* **Explicación**: La explicación de la metodología empleada debe ser clara, precisa y concisa.
* **Apoyo Visual**: Se espera que tengan la mayor cantidad de gráficos y/o tablas que puedan resumir adecuadamente todo el proceso realizado.
### Esquema del proyecto
El proyecto tendrá la siguiente estructura de trabajo:
```
- project
|
|- data
|- tmdb_5000_credits.csv
|- tmdb_5000_movies.csv
|- graficos.py
|- lectura.py
|- modelos.py
|- preprocesamiento.py
|- presentacion.ipynb
|- project.ipynb
```
donde:
* `data`: carpeta con los datos del proyecto
* `graficos.py`: módulo de gráficos
* `lectura.py`: módulo de lectura de datos
* `modelos.py`: módulo de modelos de Machine Learning utilizados
* `preprocesamiento.py`: módulo de preprocesamiento de datos
* `presentacion.ipynb`: presentación del proyecto (formato *RISE*)
* `project.ipynb`: descripción del proyecto
### Apoyo
Para que la carga del proyecto sea lo más amena posible, se les deja las siguientes referencias:
* **Sistema de recomendación**: Pueden tomar como referencia el proyecto de Kaggle [Getting Started with a Movie Recommendation System](https://www.kaggle.com/ibtesama/getting-started-with-a-movie-recommendation-system/data?select=tmdb_5000_credits.csv).
* **RISE**: Les dejo un video del Profesor Sebastían Flores denomindo *Presentaciones y encuestas interactivas en jupyter notebooks y RISE* ([link](https://www.youtube.com/watch?v=ekyN9DDswBE&ab_channel=PyConColombia)). Este material les puede ayudar para comprender mejor este nuevo concepto.
| github_jupyter |
# Introduction
This tutorial illustrates how to use *ObjTables* to revision datasets, revision schemas, and migrate datasets between revisions of their schemas. This tutorial uses an address book of CEOs as an example.
# Define a schema for an address book
First, as described in [Tutorial 1](1.%20Building%20and%20visualizing%20schemas.ipynb), use *ObjTables* to define a schema for an address book.
```
import enum
import obj_tables
import types
class Address(obj_tables.Model):
street = obj_tables.StringAttribute(unique=True, primary=True, verbose_name='Street')
city = obj_tables.StringAttribute(verbose_name='City')
state = obj_tables.StringAttribute(verbose_name='State')
zip_code = obj_tables.StringAttribute(verbose_name='Zip code')
country = obj_tables.StringAttribute(verbose_name='Country')
class Meta(obj_tables.Model.Meta):
table_format = obj_tables.TableFormat.multiple_cells
attribute_order = ('street', 'city', 'state', 'zip_code', 'country',)
verbose_name = 'Address'
verbose_name_plural = 'Addresses'
class Company(obj_tables.Model):
name = obj_tables.StringAttribute(unique=True, primary=True, verbose_name='Name')
url = obj_tables.UrlAttribute(verbose_name='URL')
address = obj_tables.OneToOneAttribute(Address, related_name='company', verbose_name='Address')
class Meta(obj_tables.Model.Meta):
table_format = obj_tables.TableFormat.row
attribute_order = ('name', 'url', 'address',)
verbose_name = 'Company'
verbose_name_plural = 'Companies'
class PersonType(str, enum.Enum):
family = 'family'
friend = 'friend'
business = 'business'
class Person(obj_tables.Model):
name = obj_tables.StringAttribute(unique=True, primary=True, verbose_name='Name')
type = obj_tables.EnumAttribute(PersonType, verbose_name='Type')
company = obj_tables.ManyToOneAttribute(Company, related_name='employees', verbose_name='Company')
email_address = obj_tables.EmailAttribute(verbose_name='Email address')
phone_number = obj_tables.StringAttribute(verbose_name='Phone number')
address = obj_tables.ManyToOneAttribute(Address, related_name='people', verbose_name='Address')
class Meta(obj_tables.Model.Meta):
table_format = obj_tables.TableFormat.row
attribute_order = ('name', 'type', 'company', 'email_address', 'phone_number', 'address',)
verbose_name = 'Person'
verbose_name_plural = 'People'
class AddressBook(obj_tables.Model):
id = obj_tables.StringAttribute(unique=True, primary=True, verbose_name='Id')
companies = obj_tables.OneToManyAttribute(Company, related_name='address_book')
people = obj_tables.OneToManyAttribute(Person, related_name='address_book')
class Meta(obj_tables.Model.Meta):
table_format = obj_tables.TableFormat.column
attribute_order = ('id', 'companies', 'people')
verbose_name = 'Address book'
verbose_name_plural = 'Address books'
```
# Revision an address book of the CEOs of technology companies
In many domains such as exploratory areas of science, datasets must often be built iteratively over time. For example, we believe that whole-cell models will be built by iteratively modeling additional biochemical species, reactions, and pathways over time as more experimental data and knowledge is generated and additional collaborators contribute to a model. Consequently, it is often helpful to track the provenence of a dataset including when the dataset was first created; when each revision was made; which objects and relationships were added, removed, or changed with each revision and why; and who contributed each revision.
##### Revisioning datasets with Git
We recommend using [Git](https://git-scm.com/) to track the revision provenance of a dataset as follows:
1. Create a Git repository.
2. Optionally, host the repository on a publicly accessible server such as [GitHub](https://github.com).
3. Save each revision in CSV, TSV, MULTI.CSV, or MULTI.TSV format so that Git can difference and merge the dataset.
4. For each revision, create an instance of `DataRepoMetadata`, use the `obj_tables.utils.set_git_repo_metadata_from_path` method to record the revision of the dataset into this instance, and use `obj_tables.io` to save this instance of `DataRepoMetadata` into the dataset.
6. Commit the revision, noting the rationale for the revision in the Git commit message. We also recommend [configuring Git to track the author of each revision](https://git-scm.com/book/en/v2/Getting-Started-First-Time-Git-Setup).
7. Optionally, push the revision to the public server.
##### (Step 1) Create a Git repository to track the revisions of the address book
```
from obj_tables.utils import DataRepoMetadata, set_git_repo_metadata_from_path
from wc_utils.util.git import RepoMetadataCollectionType
import git
import os
import shutil
repo_path = 'Address book'
repo_url = 'https://github.com/KarrLab/obj_tables_revisioning_tutorial_repo.git'
# create repository
if os.path.isdir(repo_path):
shutil.rmtree(repo_path)
repo = git.Repo.clone_from(repo_url, repo_path)
```
##### (Steps 3, 4) Create an initial address book of the CEOs of several technology companies as of 2011, and save it to multiple CSV files along with metadata about the current revision of the address book
```
import obj_tables.io
# Steve Jobs of Apple
apple = Company(name='Apple',
url='https://www.apple.com/',
address=Address(street='10600 N Tantau Ave',
city='Cupertino',
state='CA',
zip_code='95014',
country='US'))
jobs = Person(name='Steve Jobs',
type=PersonType.business,
company=apple,
email_address='sjobs@apple.com',
phone_number='408-996-1010',
address=apple.address)
# Reed Hasting of Netflix
netflix = Company(name='Netflix',
url='https://www.netflix.com/',
address=Address(street='100 Winchester Cir',
city='Los Gatos',
state='CA',
zip_code='95032',
country='US'))
hastings = Person(name='Reed Hastings',
type=PersonType.business,
company=netflix,
email_address='reed.hastings@netflix.com',
phone_number='408-540-3700',
address=netflix.address)
# Eric Schmidt of Google
google = Company(name='Google',
url='https://www.google.com/',
address=Address(street='1600 Amphitheatre Pkwy',
city='Mountain View',
state='CA',
zip_code='94043',
country='US'))
schmidt = Person(name='Eric Schmidt',
type=PersonType.business,
company=google,
email_address='eschmidt@google.com',
phone_number='650-253-0000',
address=google.address)
# Mark Zuckerberg of Facebook
facebook = Company(name='Facebook',
url='https://www.facebook.com/',
address=Address(street='1 Hacker Way #15',
city='Menlo Park',
state='CA',
zip_code='94025',
country='US'))
zuckerberg = Person(name='Mark Zuckerberg',
type=PersonType.business,
company=facebook,
email_address='zuck@fb.com',
phone_number='650-543-4800',
address=facebook.address)
# Merge the companies and CEOs into a single address book
ceos = AddressBook(
id = 'tech',
companies = [apple, facebook, google, netflix],
people = [schmidt, zuckerberg, hastings, jobs],
)
# Get the current revision of the repository
revision = DataRepoMetadata()
set_git_repo_metadata_from_path(revision, RepoMetadataCollectionType.DATA_REPO, path=repo_path)
# Save the address book to multiple CSV files along with its revision metadata
address_book_filename = os.path.join(repo_path, 'ceos-*.csv')
obj_tables.io.Writer().run(address_book_filename, [revision, ceos],
models=[DataRepoMetadata, AddressBook, Company, Person])
import pandas
pandas.read_csv(os.path.join(repo_path, 'ceos-Data repo metadata.csv'), delimiter=',')
pandas.read_csv(os.path.join(repo_path, 'ceos-Address book.csv'), delimiter=',')
pandas.read_csv(os.path.join(repo_path, 'ceos-Companies.csv'), delimiter=',')
pandas.read_csv(os.path.join(repo_path, 'ceos-People.csv'), delimiter=',')
```
##### (Step 5) Commit the initial address book
```
repo.index.add([
'ceos-Data repo metadata.csv',
'ceos-Address book.csv',
'ceos-Companies.csv',
'ceos-People.csv',
])
repo.index.commit('Initial version of address book')
```
##### (Steps 3, 4) Revise the address book to reflect the current CEOs as of 2020
```
# Tim Cook is now the CEO of Apple
jobs.cut_relations()
cook = Person(name='Tim Cook',
type=PersonType.business,
company=apple,
email_address='tcook@apple.com',
phone_number='408-996-1010',
address=apple.address)
# Sundar Pichai is now the CEO of Google
ceos.people.remove(schmidt)
google.employees.remove(schmidt)
google.address.people.remove(schmidt)
pichai = Person(name='Sundar Pichai',
type=PersonType.business,
company=google,
email_address='sundar@google.com',
phone_number='650-253-0000',
address=google.address)
# Get the current revision of the repository
set_git_repo_metadata_from_path(revision, RepoMetadataCollectionType.DATA_REPO, path=repo_path)
# Save the address book to a MULTI.CSV file along with its revision metadata
obj_tables.io.Writer().run(address_book_filename, [revision, ceos],
models=[DataRepoMetadata, AddressBook, Company, Person])
pandas.read_csv(os.path.join(repo_path, 'ceos-Data repo metadata.csv'), delimiter=',')
pandas.read_csv(os.path.join(repo_path, 'ceos-People.csv'), delimiter=',')
```
##### (Step 5) Commit the revised address book
```
repo.index.add([
'ceos-Data repo metadata.csv',
'ceos-Address book.csv',
'ceos-Companies.csv',
'ceos-People.csv',
])
repo.index.commit('Initial version of address book')
```
# Revise the address book schema and migrate the address book to the revised schema
Please check back soon! In the meantime, please contact us at [info@karrlab.org](mailto:info@karrlab.org) with any questions.
| github_jupyter |
This course covers the key Python skills you’ll need so you can start using Python for data science.
We'll start with a brief overview of Python syntax, variable assignment, and arithmetic operators. If you have previous Python experience, you can [skip straight to the hands-on exercise](https://www.kaggle.com/kernels/fork/1275163).
# Hello, Python!
Python was named for the British comedy troupe [Monty Python](https://en.wikipedia.org/wiki/Monty_Python), so we'll make our first Python program a homage to their skit about Spam.
Just for fun, try reading over the code below and predicting what it's going to do when run. (If you have no idea, that's fine!)
Then click the "output" button to see the results of our program.
```
spam_amount = 0
print(spam_amount)
# Ordering Spam, egg, Spam, Spam, bacon and Spam (4 more servings of Spam)
spam_amount = spam_amount + 4
if spam_amount > 0:
print("But I don't want ANY spam!")
viking_song = "Spam " * spam_amount
print(viking_song)
```
There's a lot to unpack here! This silly program demonstrates many important aspects of what Python code looks like and how it works. Let's review the code from top to bottom.
```
spam_amount = 0
```
**Variable assignment:** Here we create a variable called `spam_amount` and assign it the value of 0 using `=`, which is called the assignment operator.
> **Aside**: If you've programmed in certain other languages (like Java or C++), you might be noticing some things Python *doesn't* require us to do here:
- we don't need to "declare" `spam_amount` before assigning to it
- we don't need to tell Python what type of value `spam_amount` is going to refer to. In fact, we can even go on to reassign `spam_amount` to refer to a different sort of thing like a string or a boolean.
```
print(spam_amount)
```
**Function calls:**. `print` is a Python function that displays the value passed to it on the screen. We call functions by putting parentheses after their name, and putting the inputs (or *arguments*) to the function in those parentheses.
```
# Ordering Spam, egg, Spam, Spam, bacon and Spam (4 more servings of Spam)
spam_amount = spam_amount + 4
```
The first line above is a **comment**. In Python, comments begin with the `#` symbol.
Next we see an example of reassignment. Reassigning the value of an existing variable looks just the same as creating a variable - it still uses the `=` assignment operator.
In this case, the value we're assigning to `spam_amount` involves some simple arithmetic on its previous value. When it encounters this line, Python evaluates the expression on the right-hand-side of the `=` (0 + 4 = 4), and then assigns that value to the variable on the left-hand-side.
We won't talk much about "conditionals" until later, but, even if you've never coded before, you can probably guess what this does. Python is prized for its readability and the simplicity.
Note how we indicated which code belongs to the `if`. `"But I don't want ANY spam!"` is only supposed to be printed if `spam_amount` is positive. But the later code (like `print(viking_song)`) should be executed no matter what. How do we (and Python) know that?
The colon (`:`) at the end of the `if` line indicates that a new **code block** is starting. Subsequent lines which are indented are part of that code block.
> **Aside**: If you've coded before, you might know that some other languages use `{`curly braces`}` to mark the beginning and end of code blocks. Python's use of meaningful whitespace can be surprising to programmers who are accustomed to other languages, but in practice it can lead to more consistent and readable code than languages that do not enforce indentation of code blocks.
The later lines dealing with `viking_song` are not indented with an extra 4 spaces, so they're not a part of the `if`'s code block. We'll see more examples of indented code blocks later when we define functions and using loops.
This code snippet is also our first sighting of a **string** in Python:
```python
"But I don't want ANY spam!"
```
Strings can be marked either by double or single quotation marks. (But because this particular string *contains* a single-quote character, we might confuse Python by trying to surround it with single-quotes, unless we're careful.)
```
if spam_amount > 0:
print("But I don't want ANY spam!")
viking_song = "Spam Spam Spam"
print(viking_song)
viking_song = "Spam " * spam_amount
print(viking_song)
```
The `*` operator can be used to multiply two numbers (`3 * 3` evaluates to 9), but we can also multiply a string by a number, to get a version that's been repeated that many times. Python offers a number of cheeky little time-saving tricks like this where operators like `*` and `+` have a different meaning depending on what kind of thing they're applied to. (The technical term for this is [operator overloading](https://en.wikipedia.org/wiki/Operator_overloading).)
## Numbers and arithmetic in Python
We've already seen an example of a variable containing a number above:
```
spam_amount = 0
```
"Number" is a fine informal name for the kind of thing, but if we wanted to be more technical, we could ask Python how it would describe the type of thing that `spam_amount` is:
```
type(spam_amount)
```
It's an `int` - short for integer. There's another sort of number we commonly encounter in Python:
```
type(19.95)
```
A `float` is a number with a decimal place - very useful for representing things like weights or proportions.
`type()` is the second built-in function we've seen (after `print()`), and it's another good one to remember. It's very useful to be able to ask Python "what kind of thing is this?".
A natural thing to want to do with numbers is perform arithmetic. We've seen the `+` operator for addition, and the `*` operator for multiplication. Python also has us covered for the rest of the basic buttons on your calculator:
| Operator | Name | Description |
|--------------|----------------|--------------------------------------------------------|
| ``a + b`` | Addition | Sum of ``a`` and ``b`` |
| ``a - b`` | Subtraction | Difference of ``a`` and ``b`` |
| ``a * b`` | Multiplication | Product of ``a`` and ``b`` |
| ``a / b`` | True division | Quotient of ``a`` and ``b`` |
| ``a // b`` | Floor division | Quotient of ``a`` and ``b``, removing fractional parts |
| ``a % b`` | Modulus | Integer remainder after division of ``a`` by ``b`` |
| ``a ** b`` | Exponentiation | ``a`` raised to the power of ``b`` |
| ``-a`` | Negation | The negative of ``a`` |
<span style="display:none"></span>
One interesting observation here is that, whereas your calculator probably just has one button for division, Python can do two kinds. "True division" is basically what your calculator does:
```
print(5 / 2)
print(6 / 2)
```
It always gives us a `float`.
The `//` operator gives us a result that's rounded down to the next integer.
```
print(5 // 2)
print(6 // 2)
```
Can you think of where this would be useful? You'll see an example soon in the coding challenges.
### Order of operations
The arithmetic we learned in primary school has conventions about the order in which operations are evaluated. Some remember these by a mnemonic such as **PEMDAS** - **P**arentheses, **E**xponents, **M**ultiplication/**D**ivision, **A**ddition/**S**ubtraction.
Python follows similar rules about which calculations to perform first. They're mostly pretty intuitive.
```
8 - 3 + 2
-3 + 4 * 2
```
Sometimes the default order of operations isn't what we want:
```
hat_height_cm = 25
my_height_cm = 190
# How tall am I, in meters, when wearing my hat?
total_height_meters = hat_height_cm + my_height_cm / 100
print("Height in meters =", total_height_meters, "?")
```
Parentheses are useful here. You can add them to force Python to evaluate sub-expressions in whatever order you want.
```
total_height_meters = (hat_height_cm + my_height_cm) / 100
print("Height in meters =", total_height_meters)
```
### Builtin functions for working with numbers
`min` and `max` return the minimum and maximum of their arguments, respectively...
```
print(min(1, 2, 3))
print(max(1, 2, 3))
```
`abs` returns the absolute value of an argument:
```
print(abs(32))
print(abs(-32))
```
In addition to being the names of Python's two main numerical types, `int` and `float` can also be called as functions which convert their arguments to the corresponding type:
```
print(float(10))
print(int(3.33))
# They can even be called on strings!
print(int('807') + 1)
```
# Your Turn
Now is your chance. Try your **[first Python programming exercise](https://www.kaggle.com/kernels/fork/1275163)**!
---
*Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum/161283) to chat with other Learners.*
| github_jupyter |
## Scalability Experiment (Section 5.3)
The experiment is designed to compare the execution time of different coarsening schemes over increasingly large graphs.
* For consistency, we use a regular graph of increasing size (vertices, edges) but always the same degree
* The reduction is fixed to 0.5. The execution time will only slightly increase for larger ratios (since the problem that has to be solved becomes easier at consecutive levels where the graph is smaller)
* If the execution time exceeds a budget (set to 100 sec), computation is skipped.
The code accompanies paper [Graph reduction with spectral and cut guarantees](http://www.jmlr.org/papers/volume20/18-680/18-680.pdf) by Andreas Loukas published at JMLR/2019 ([bibtex](http://www.jmlr.org/papers/v20/18-680.bib)).
This work was kindly supported by the Swiss National Science Foundation (grant number PZ00P2 179981).
15 March 2019
[Andreas Loukas](https://andreasloukas.blog)
[](https://zenodo.org/badge/latestdoi/175851068)
Released under the Apache license 2.0
```
!pip install networkx
%load_ext autoreload
%autoreload 2
%matplotlib inline
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:80% !important; }</style>"))
from graph_coarsening.coarsening_utils import *
import graph_coarsening.graph_lib as graph_lib
import graph_coarsening.graph_utils as graph_utils
import numpy as np
import scipy as sp
from scipy import io
from scipy.linalg import circulant
import time
import os
import matplotlib
import matplotlib.pylab as plt
import pygsp as gsp
from pygsp import graphs, filters
gsp.plotting.BACKEND = 'matplotlib'
# Experiment parameters
N_all = np.logspace(2, 6, 30, dtype=np.int)
methods = ['heavy_edge', 'variation_edges', 'variation_neighborhoods', 'algebraic_JC', 'affinity_GS', 'kron']
K_all = [10,80] #[10, 20, 40]
r = 0.5
budget = 100 # don't run anything that takes longer than this (in seconds)
n_iterations = 10
deg = 10
algorithm = 'greedy'
n_methods = len(methods)
# print(deg*N_all/2)
```
### The actual experiment code (this will take long)
If one needs to just see the results, skip running this part.
```
rerun_all = False
rewrite_results = False
if rerun_all:
timings = np.zeros((len(N_all), len(K_all), n_methods, n_iterations)) * np.NaN
skip = np.zeros(len(methods))
for NIdx, N in enumerate(N_all):
G = graph_lib.models(N, 'regular', k=deg)
for KIdx, K in enumerate(K_all):
for methodIdx,method in enumerate(methods):
if skip[methodIdx] == 1 :
timings[NIdx, KIdx, methodIdx] = np.NaN
print('skipping: {}, {}, {}'.format(N, method, K))
continue
timing = 0
for iteration in range(n_iterations):
if method == 'kron':
start = time.time()
_, tmp = kron_coarsening(G, r=r, m=None)
end = time.time()
if tmp == None:
print('kron failed... skipping')
continue
else:
start = time.time()
_, _, Call, _ = coarsen(G, K=K, r=r, max_levels=4, method=method, algorithm=algorithm)
end = time.time()
if len(Call) >= 4: print('warning: too many levels for {}, r:{}, K:{}'.format(method, r, K) )
timings[NIdx, KIdx, methodIdx, iteration] = end-start
timing = np.mean(timings[NIdx, KIdx, methodIdx, :])
skip[methodIdx] = 1 if (timing > budget) else 0
print('N = {}, done!'.format(N))
if sum(skip) == len(methods): break
if rewrite_results:
filepath = os.path.join('..', 'results', 'experiment_scalability.npz')
print('.. saving to "' + filepath + '"')
np.savez(filepath, methods=methods, K_all=K_all, N_all=N_all, timings=timings, deg=deg, budget=budget)
print('done!')
```
### Load results
```
filepath = os.path.join('..', 'results', 'experiment_scalability.npz')
data = np.load(filepath)
methods, K_all, N_all, timings, deg, budget = data['methods'], data['K_all'], data['N_all'], data['timings'], data['deg'], data['budget']
```
### Visualize them
The produced figures are used in the paper
```
matplotlib.rcParams.update({'font.size': 25})
from matplotlib import cm
colors = [ cm.ocean(x) for x in np.linspace(0, 0.95, len(methods)+1)]
colors[1] = [0.8,0,0]
colors[-2] = (np.array([127, 77, 34])/255).tolist()
size = 2.7*2.7;
print('The figures are drawn in the following in order:')
for KIdx, K in enumerate(K_all):
fig, axes = plt.subplots(1, 1, figsize=(1.618*size, size));
for methodIdx,method in reversed(list(enumerate(methods))):
lineWidth = 1.5; marker = 's'
method = method.replace('_', ' ')
if method == 'heavy edge':
method = 'heavy edge'
cIdx, line, marker = 0, ':', 's'
elif 'variation edges' in method:
method = 'local var. (edges)'
cIdx, line, marker, lineWidth = 2, '-', 'o', 1.5
elif (method == 'variation neighborhoods') or (method == 'variation neighborhood'):
method = 'local var. (neigh)'
cIdx, line, marker, lineWidth = 1, '-', 'o', 1.5
elif 'algebraic' in method:
method = 'algebraic dist.'
cIdx, line = 3, ':'
elif 'affinity' in method:
method = 'affinity'
cIdx, line = 4, ':'
elif method == 'kron':
method = 'kron'
cIdx, line, marker = 5, '--', 'x'
else:
continue
style = line + marker
color = colors[cIdx]
tmp = np.mean(timings[:,KIdx,methodIdx,:], 1)
tmp[tmp>budget] = np.NaN
axes.plot(N_all*deg/2, tmp, style, label='{}'.format(method), color=color, lineWidth=lineWidth, markersize=6)
axes.plot(np.array([10, N_all[-1]])*deg/2, [budget, budget], 'k:')
axes.set_xscale('log')
axes.set_yscale('log')
axes.set_xlabel('number of edges (M)')
axes.set_ylabel('execution time (sec)')
axes.set_ylim([0.02, budget+30])
axes.set_xlim([300, N_all[-1]])
legend0 = axes.legend(fontsize=22, loc='lower right', edgecolor=[1,1,1])
axes.text(500, 63, 'max execution time', fontsize=21)
axes.spines['right'].set_visible(False)
axes.spines['top'].set_visible(False)
fig.tight_layout()
print('* experiment_scalability_K='+ str(K))
# fig.savefig(os.path.join('..', 'results', 'experiment_scalability_K='+ str(K) +'.pdf'))
```
| github_jupyter |
# Fairness and Explainability with SageMaker Clarify
1. [Overview](#Overview)
1. [Prerequisites and Data](#Prerequisites-and-Data)
1. [Initialize SageMaker](#Initialize-SageMaker)
1. [Download data](#Download-data)
1. [Loading the data: Adult Dataset](#Loading-the-data:-Adult-Dataset)
1. [Data inspection](#Data-inspection)
1. [Data encoding and upload to S3](#Encode-and-Upload-the-Data)
1. [Train and Deploy XGBoost Model](#Train-XGBoost-Model)
1. [Train Model](#Train-Model)
1. [Deploy Model to Endpoint](#Deploy-Model)
1. [Amazon SageMaker Clarify](#Amazon-SageMaker-Clarify)
1. [Detecting Bias](#Detecting-Bias)
1. [Writing BiasConfig](#Writing-BiasConfig)
1. [Pre-training Bias](#Pre-training-Bias)
1. [Post-training Bias](#Post-training-Bias)
1. [Viewing the Bias Report](#Viewing-the-Bias-Report)
1. [Explaining Predictions](#Explaining-Predictions)
1. [Viewing the Explainability Report](#Viewing-the-Explainability-Report)
1. [Clean Up](#Clean-Up)
## Overview
Amazon SageMaker Clarify helps improve your machine learning models by detecting potential bias and helping explain how these models make predictions. The fairness and explainability functionality provided by SageMaker Clarify takes a step towards enabling AWS customers to build trustworthy and understandable machine learning models. The product comes with the tools to help you with the following tasks.
* Measure biases that can occur during each stage of the ML lifecycle (data collection, model training and tuning, and monitoring of ML models deployed for inference).
* Generate model governance reports targeting risk and compliance teams and external regulators.
* Provide explanations of the data, models, and monitoring used to assess predictions.
This sample notebook walks you through:
1. Key terms and concepts needed to understand SageMaker Clarify
1. Measuring the pre-training bias of a dataset and post-training bias of a model
1. Explaining the importance of the various input features on the model's decision
1. Accessing the reports through SageMaker Studio if you have an instance set up.
In doing so, the notebook will first train a [SageMaker XGBoost](https://docs.aws.amazon.com/sagemaker/latest/dg/xgboost.html) model using training dataset, then use SageMaker Clarify to analyze a testing dataset in CSV format. SageMaker Clarify also supports analyzing dataset in [SageMaker JSONLines dense format](https://docs.aws.amazon.com/sagemaker/latest/dg/cdf-inference.html#common-in-formats), which is illustrated in [another notebook](https://github.com/aws/amazon-sagemaker-examples/blob/master/sagemaker_processing/fairness_and_explainability/fairness_and_explainability_jsonlines_format.ipynb).
## Prerequisites and Data
### Initialize SageMaker
```
from sagemaker import Session
session = Session()
bucket = session.default_bucket()
prefix = "sagemaker/DEMO-sagemaker-clarify"
region = session.boto_region_name
# Define IAM role
from sagemaker import get_execution_role
import pandas as pd
import numpy as np
import os
import boto3
role = get_execution_role()
s3_client = boto3.client("s3")
```
### Download data
Data Source: [https://archive.ics.uci.edu/ml/machine-learning-databases/adult/](https://archive.ics.uci.edu/ml/machine-learning-databases/adult/)
Let's __download__ the data and save it in the local folder with the name adult.data and adult.test from UCI repository$^{[2]}$.
$^{[2]}$Dua Dheeru, and Efi Karra Taniskidou. "[UCI Machine Learning Repository](http://archive.ics.uci.edu/ml)". Irvine, CA: University of California, School of Information and Computer Science (2017).
```
adult_columns = [
"Age",
"Workclass",
"fnlwgt",
"Education",
"Education-Num",
"Marital Status",
"Occupation",
"Relationship",
"Ethnic group",
"Sex",
"Capital Gain",
"Capital Loss",
"Hours per week",
"Country",
"Target",
]
if not os.path.isfile("adult.data"):
s3_client.download_file(
"sagemaker-sample-files", "datasets/tabular/uci_adult/adult.data", "adult.data"
)
print("adult.data saved!")
else:
print("adult.data already on disk.")
if not os.path.isfile("adult.test"):
s3_client.download_file(
"sagemaker-sample-files", "datasets/tabular/uci_adult/adult.test", "adult.test"
)
print("adult.test saved!")
else:
print("adult.test already on disk.")
```
### Loading the data: Adult Dataset
From the UCI repository of machine learning datasets, this database contains 14 features concerning demographic characteristics of 45,222 rows (32,561 for training and 12,661 for testing). The task is to predict whether a person has a yearly income that is more or less than $50,000.
Here are the features and their possible values:
1. **Age**: continuous.
1. **Workclass**: Private, Self-emp-not-inc, Self-emp-inc, Federal-gov, Local-gov, State-gov, Without-pay, Never-worked.
1. **Fnlwgt**: continuous (the number of people the census takers believe that observation represents).
1. **Education**: Bachelors, Some-college, 11th, HS-grad, Prof-school, Assoc-acdm, Assoc-voc, 9th, 7th-8th, 12th, Masters, 1st-4th, 10th, Doctorate, 5th-6th, Preschool.
1. **Education-num**: continuous.
1. **Marital-status**: Married-civ-spouse, Divorced, Never-married, Separated, Widowed, Married-spouse-absent, Married-AF-spouse.
1. **Occupation**: Tech-support, Craft-repair, Other-service, Sales, Exec-managerial, Prof-specialty, Handlers-cleaners, Machine-op-inspct, Adm-clerical, Farming-fishing, Transport-moving, Priv-house-serv, Protective-serv, Armed-Forces.
1. **Relationship**: Wife, Own-child, Husband, Not-in-family, Other-relative, Unmarried.
1. **Ethnic group**: White, Asian-Pac-Islander, Amer-Indian-Eskimo, Other, Black.
1. **Sex**: Female, Male.
* **Note**: this data is extracted from the 1994 Census and enforces a binary option on Sex
1. **Capital-gain**: continuous.
1. **Capital-loss**: continuous.
1. **Hours-per-week**: continuous.
1. **Native-country**: United-States, Cambodia, England, Puerto-Rico, Canada, Germany, Outlying-US(Guam-USVI-etc), India, Japan, Greece, South, China, Cuba, Iran, Honduras, Philippines, Italy, Poland, Jamaica, Vietnam, Mexico, Portugal, Ireland, France, Dominican-Republic, Laos, Ecuador, Taiwan, Haiti, Columbia, Hungary, Guatemala, Nicaragua, Scotland, Thailand, Yugoslavia, El-Salvador, Trinadad&Tobago, Peru, Hong, Holand-Netherlands.
Next, we specify our binary prediction task:
15. **Target**: <=50,000, >$50,000.
```
training_data = pd.read_csv(
"adult.data", names=adult_columns, sep=r"\s*,\s*", engine="python", na_values="?"
).dropna()
testing_data = pd.read_csv(
"adult.test", names=adult_columns, sep=r"\s*,\s*", engine="python", na_values="?", skiprows=1
).dropna()
training_data.head()
```
### Data inspection
Plotting histograms for the distribution of the different features is a good way to visualize the data. Let's plot a few of the features that can be considered _sensitive_.
Let's take a look specifically at the Sex feature of a census respondent. In the first plot we see that there are fewer Female respondents as a whole but especially in the positive outcomes, where they form ~$\frac{1}{7}$th of respondents.
```
training_data["Sex"].value_counts().sort_values().plot(kind="bar", title="Counts of Sex", rot=0)
training_data["Sex"].where(training_data["Target"] == ">50K").value_counts().sort_values().plot(
kind="bar", title="Counts of Sex earning >$50K", rot=0
)
```
### Encode and Upload the Dataset
Here we encode the training and test data. Encoding input data is not necessary for SageMaker Clarify, but is necessary for the model.
```
from sklearn import preprocessing
def number_encode_features(df):
result = df.copy()
encoders = {}
for column in result.columns:
if result.dtypes[column] == np.object:
encoders[column] = preprocessing.LabelEncoder()
# print('Column:', column, result[column])
result[column] = encoders[column].fit_transform(result[column].fillna("None"))
return result, encoders
training_data = pd.concat([training_data["Target"], training_data.drop(["Target"], axis=1)], axis=1)
training_data, _ = number_encode_features(training_data)
training_data.to_csv("train_data.csv", index=False, header=False)
testing_data, _ = number_encode_features(testing_data)
test_features = testing_data.drop(["Target"], axis=1)
test_target = testing_data["Target"]
test_features.to_csv("test_features.csv", index=False, header=False)
```
A quick note about our encoding: the "Female" Sex value has been encoded as 0 and "Male" as 1.
```
training_data.head()
```
Lastly, let's upload the data to S3
```
from sagemaker.s3 import S3Uploader
from sagemaker.inputs import TrainingInput
train_uri = S3Uploader.upload("train_data.csv", "s3://{}/{}".format(bucket, prefix))
train_input = TrainingInput(train_uri, content_type="csv")
test_uri = S3Uploader.upload("test_features.csv", "s3://{}/{}".format(bucket, prefix))
```
### Train XGBoost Model
#### Train Model
Since our focus is on understanding how to use SageMaker Clarify, we keep it simple by using a standard XGBoost model.
```
from sagemaker.image_uris import retrieve
from sagemaker.estimator import Estimator
container = retrieve("xgboost", region, version="1.2-1")
xgb = Estimator(
container,
role,
instance_count=1,
instance_type="ml.m5.xlarge",
disable_profiler=True,
sagemaker_session=session,
)
xgb.set_hyperparameters(
max_depth=5,
eta=0.2,
gamma=4,
min_child_weight=6,
subsample=0.8,
objective="binary:logistic",
num_round=800,
)
xgb.fit({"train": train_input}, logs=False)
```
#### Deploy Model
Here we create the SageMaker model.
```
model_name = "DEMO-clarify-model"
model = xgb.create_model(name=model_name)
container_def = model.prepare_container_def()
session.create_model(model_name, role, container_def)
```
## Amazon SageMaker Clarify
Now that you have your model set up. Let's say hello to SageMaker Clarify!
```
from sagemaker import clarify
clarify_processor = clarify.SageMakerClarifyProcessor(
role=role, instance_count=1, instance_type="ml.m5.xlarge", sagemaker_session=session
)
```
### Detecting Bias
SageMaker Clarify helps you detect possible pre- and post-training biases using a variety of metrics.
#### Writing DataConfig and ModelConfig
A `DataConfig` object communicates some basic information about data I/O to SageMaker Clarify. We specify where to find the input dataset, where to store the output, the target column (`label`), the header names, and the dataset type.
```
bias_report_output_path = "s3://{}/{}/clarify-bias".format(bucket, prefix)
bias_data_config = clarify.DataConfig(
s3_data_input_path=train_uri,
s3_output_path=bias_report_output_path,
label="Target",
headers=training_data.columns.to_list(),
dataset_type="text/csv",
)
```
A `ModelConfig` object communicates information about your trained model. To avoid additional traffic to your production models, SageMaker Clarify sets up and tears down a dedicated endpoint when processing.
* `instance_type` and `instance_count` specify your preferred instance type and instance count used to run your model on during SageMaker Clarify's processing. The testing dataset is small so a single standard instance is good enough to run this example. If your have a large complex dataset, you may want to use a better instance type to speed up, or add more instances to enable Spark parallelization.
* `accept_type` denotes the endpoint response payload format, and `content_type` denotes the payload format of request to the endpoint.
```
model_config = clarify.ModelConfig(
model_name=model_name,
instance_type="ml.m5.xlarge",
instance_count=1,
accept_type="text/csv",
content_type="text/csv",
)
```
A `ModelPredictedLabelConfig` provides information on the format of your predictions. XGBoost model outputs probabilities of samples, so SageMaker Clarify invokes the endpoint then uses `probability_threshold` to convert the probability to binary labels for bias analysis. Prediction above the threshold is interpreted as label value `1` and below or equal as label value `0`.
```
predictions_config = clarify.ModelPredictedLabelConfig(probability_threshold=0.8)
```
#### Writing BiasConfig
SageMaker Clarify also needs information on what the sensitive columns (`facets`) are, what the sensitive features (`facet_values_or_threshold`) may be, and what the desirable outcomes are (`label_values_or_threshold`).
SageMaker Clarify can handle both categorical and continuous data for `facet_values_or_threshold` and for `label_values_or_threshold`. In this case we are using categorical data.
We specify this information in the `BiasConfig` API. Here that the positive outcome is earning >$50,000, Sex is a sensitive category, and Female respondents are the sensitive group. `group_name` is used to form subgroups for the measurement of Conditional Demographic Disparity in Labels (CDDL) and Conditional Demographic Disparity in Predicted Labels (CDDPL) with regards to Simpson’s paradox.
```
bias_config = clarify.BiasConfig(
label_values_or_threshold=[1], facet_name="Sex", facet_values_or_threshold=[0], group_name="Age"
)
```
#### Pre-training Bias
Bias can be present in your data before any model training occurs. Inspecting your data for bias before training begins can help detect any data collection gaps, inform your feature engineering, and help you understand what societal biases the data may reflect.
Computing pre-training bias metrics does not require a trained model.
#### Post-training Bias
Computing post-training bias metrics does require a trained model.
Unbiased training data (as determined by concepts of fairness measured by bias metric) may still result in biased model predictions after training. Whether this occurs depends on several factors including hyperparameter choices.
You can run these options separately with `run_pre_training_bias()` and `run_post_training_bias()` or at the same time with `run_bias()` as shown below.
```
clarify_processor.run_bias(
data_config=bias_data_config,
bias_config=bias_config,
model_config=model_config,
model_predicted_label_config=predictions_config,
pre_training_methods="all",
post_training_methods="all",
)
```
#### Viewing the Bias Report
In Studio, you can view the results under the experiments tab.
<img src="./recordings/bias_report.gif">
Each bias metric has detailed explanations with examples that you can explore.
<img src="./recordings/bias_detail.gif">
You could also summarize the results in a handy table!
<img src="./recordings/bias_report_chart.gif">
If you're not a Studio user yet, you can access the bias report in pdf, html and ipynb formats in the following S3 bucket:
```
bias_report_output_path
```
### Explaining Predictions
There are expanding business needs and legislative regulations that require explanations of _why_ a model made the decision it did. SageMaker Clarify uses SHAP to explain the contribution that each input feature makes to the final decision.
Kernel SHAP algorithm requires a baseline (also known as background dataset). Baseline dataset type shall be the same as `dataset_type` of `DataConfig`, and baseline samples shall only include features. By definition, `baseline` should either be a S3 URI to the baseline dataset file, or an in-place list of samples. In this case we chose the latter, and put the first sample of the test dataset to the list.
```
shap_config = clarify.SHAPConfig(
baseline=[test_features.iloc[0].values.tolist()],
num_samples=15,
agg_method="mean_abs",
save_local_shap_values=True,
)
explainability_output_path = "s3://{}/{}/clarify-explainability".format(bucket, prefix)
explainability_data_config = clarify.DataConfig(
s3_data_input_path=train_uri,
s3_output_path=explainability_output_path,
label="Target",
headers=training_data.columns.to_list(),
dataset_type="text/csv",
)
clarify_processor.run_explainability(
data_config=explainability_data_config,
model_config=model_config,
explainability_config=shap_config,
)
```
#### Viewing the Explainability Report
As with the bias report, you can view the explainability report in Studio under the experiments tab
<img src="./recordings/explainability_detail.gif">
The Model Insights tab contains direct links to the report and model insights.
If you're not a Studio user yet, as with the Bias Report, you can access this report at the following S3 bucket.
```
explainability_output_path
```
#### Analysis of local explanations
It is possible to visualize the the local explanations for single examples in your dataset. You can use the obtained results from running Kernel SHAP algorithm for global explanations.
You can simply load the local explanations stored in your output path, and visualize the explanation (i.e., the impact that the single features have on the prediction of your model) for any single example.
```
local_explanations_out = pd.read_csv(explainability_output_path + "/explanations_shap/out.csv")
feature_names = [str.replace(c, "_label0", "") for c in local_explanations_out.columns.to_series()]
local_explanations_out.columns = feature_names
selected_example = 111
print(
"Example number:",
selected_example,
"\nwith model prediction:",
sum(local_explanations_out.iloc[selected_example]) > 0,
)
print("\nFeature values -- Label", training_data.iloc[selected_example])
local_explanations_out.iloc[selected_example].plot(
kind="bar", title="Local explanation for the example number " + str(selected_example), rot=90
)
```
### Clean Up
Finally, don't forget to clean up the resources we set up and used for this demo!
```
session.delete_model(model_name)
```
| github_jupyter |
```
import pmdarima
import pytz
import pandas as pd
import sm as sm
from pandas import DataFrame
dateparse = lambda dates: pd.datetime.strptime(dates, '%Y-%m-%d')
df = pd.read_csv('../doge_v1.csv', parse_dates=['Date'], date_parser=dateparse)
df.set_index(["Date"], drop=False, inplace=True)
df.head()
print(pd.date_range(
start="2017-11-09", end="2022-02-04").difference(df.index))
import matplotlib.pyplot as plt
df['Close'].plot(figsize=(15, 7))
from statsmodels.tsa.seasonal import seasonal_decompose
from dateutil.parser import parse
# Multiplicative Decomposition
result_mul = seasonal_decompose(df['Close'], model='multiplicative', extrapolate_trend='freq')
# Additive Decomposition
result_add = seasonal_decompose(df['Close'], model='additive', extrapolate_trend='freq')
# Plot
plt.rcParams.update({'figure.figsize': (10,10)})
result_mul.plot().suptitle('Multiplicative Decompose', fontsize=22)
result_add.plot().suptitle('Additive Decompose', fontsize=22)
plt.show()
from python_scripts.time_series_utilities import check_stationarity, plot_pacf_acf
check_stationarity(df, 'Close')
from scipy import signal
detrended = signal.detrend(df['Close'].values)
plt.plot(detrended)
result_mul = seasonal_decompose(df['Close'], model='multiplicative', extrapolate_trend='freq')
detrended_mul = df['Close'].values - result_mul.trend
plt.plot(detrended_mul)
import numpy as np
df['transformed_Close'] = np.log(df['Close'])
df['transformed_Close'].plot(figsize=(15, 7))
check_stationarity(df, 'transformed_Close')
df['transformed_Close'] = df.transformed_Close.diff()
df_transformed = df.iloc[1:, :]
df['transformed_Close'].plot(figsize=(15, 7))
check_stationarity(df_transformed, 'transformed_Close')
plot_pacf_acf(df_transformed, 'transformed_Close')
import statsmodels.api as sm
sm.stats.acorr_ljungbox(df_transformed.transformed_Close, lags=[30], return_df=True)
from pmdarima import auto_arima
import urllib3
arima_model = pmdarima.auto_arima(df_transformed.transformed_Close,
start_p = 0,
start_q= 0,
d=None,
max_p=27,
max_q=27,
test='adf',
seasonal=False,
trace=True,
error_action='ignore',
stepwise=True)
sm.stats.acorr_ljungbox(arima_model.resid(), lags=[30], return_df=True)
arima_model = pmdarima.auto_arima(df_transformed.transformed_Close,
start_p = 3,
start_q= 3,
d=None,
max_p=27,
max_q=27,
test='adf',
seasonal=False,
trace=True,
error_action='ignore',
stepwise=True)
sm.stats.acorr_ljungbox(arima_model.resid(), lags=[30], return_df=True)
from python_scripts.sarima_helper import train_test_split_continual
train,test = train_test_split_continual(df, 0.1)
model = sm.tsa.arima.ARIMA(np.log(train.Close), order=(3,1,2))
fitted_model = model.fit()
forecast = fitted_model.get_forecast(len(test))
sum_frame = (forecast.summary_frame())
sum_frame['mean'].plot(figsize=(15, 7))
plt.figure(figsize=(10, 6))
plt.grid(True)
plt.xlabel('Dates')
plt.ylabel('Closing Prices')
plt.plot(test.Close, 'blue', label='Test data')
plt.plot(np.exp(sum_frame['mean']), 'green', label='Predicted')
plt.legend()
plt.show()
from python_scripts.time_series_utilities import test_arima_one_step
predictions = test_arima_one_step(train, test, 3,1,2)
from python_scripts.time_series_utilities import grid_search_hyperparams
p_s = range(0,10)
q_s = range(0,10)
d_s = [1]
grid_search_hyperparams(p_s, q_s, d_s, train, test)
test_arima_one_step(train, test, 1,1,0)
```
| github_jupyter |
```
import requests
import pandas as pd
from splinter import Browser
from bs4 import BeautifulSoup as bs
from webdriver_manager.chrome import ChromeDriverManager
import pymongo
# Setup splinter
executable_path = {'executable_path': ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
```
# NASA Mars News
```
### NASA Mars News
# URL of page to be scarped
#url= 'https://redplanetscience.com/'
# Retrieve page with the requests module
url = 'https://redplanetscience.com/'
browser.visit(url)
html = browser.html
soup_mars = bs(html, "html.parser")
#print(soup_mars.prettify())
# News from March 16, 2022
url = 'https://redplanetscience.com/'
browser.visit(url)
#Dalay for loading the page
browser.is_element_present_by_css('div.list_text', wait_time=1)
html = browser.html
soup_mars = bs(html, "html.parser")
print(soup_mars.prettify())
# Identify and return title
news_title = soup_mars.find('div', class_="content_title")
news_title
# Identify and return full news content
news_nasa = soup_mars.find('div', class_='list_text')
news_nasa
# Identify and return paragraph
news_p = soup_mars.find('div', class_="article_teaser_body").get_text
news_p
```
# JPL Mars Space Images - Featured Image
```
### JPL Mars Space Images - Featured Image
featured_image_url = 'https://spaceimages-mars.com'
browser.visit(featured_image_url)
print(f"Featured_image_url = {featured_image_url}")
response = requests.get(featured_image_url, stream=True)
with open('Images/Featuredimage.png', 'wb') as out_file:
shutil.copyfileobj(response.raw, out_file)
Image(url='Images/Featuredimage.png')
# Find and click the full image button
full_image = browser.find_by_tag('button')[1]
full_image.click()
html = browser.html
image_mars = bs(html, "html.parser")
print(image_mars.prettify())
# Identify and return image
img_url_rel = image_mars.find('img', class_='fancybox-image').get('src')
img_url_rel
imag_url = f'https://spaceimages-mars.com/{img_url_rel}'
imag_url
```
# Mars Facts
# MARS PLANET PROFILE
```
### Mars Facts
# MARS PLANET PROFILE
url_facts = 'https://galaxyfacts-mars.com'
facts = ['Equatorial Diameter:', 'Polar Diameter:', 'Mass:', 'Moons:', 'Orbit Distance:', 'Orbit Period:',\
'Surface Temperature:', 'First Record:', 'Recorded By:']
# Use Panda's `read_html` to parse the url
### BEGIN SOLUTION
tables = pd.read_html(url_facts)
tables
### END SOLUTION'
### BEGIN SOLUTION
df = tables[0]
df.columns = ['Mars - Earth Comparison', 'Mars', 'Earth']
df.head()
### END SOLUTION
df.rename(columns = {'Mars - Earth Comparison':'Facts'}, inplace = True)
print(df.columns)
# Drop Earth
df_d= df.drop(['Earth'], axis = 1)
## Delete row 1 Mars-Earth comparsion
mars_facts = df_d.iloc[1:, :]
mars_facts
html_table = mars_facts.to_html()
html_table
```
# Mars Hemispheres
```
### Mars Hemispheres
url_hemis = 'https://marshemispheres.com/'
# Retrieve page with the requests module
response = requests.get(url_hemis)
# Create BeautifulSoup object; parse with 'lxml'
soup = BeautifulSoup(response.text, 'html.parser')
hemisphere_image = soup.find_all('div', class_='collapsible results')
hemisphere_image
url_hemis = 'https://marshemispheres.com/'
browser.visit(url_hemis)
# Getting the titles and images. Append the dictionary with the image url string and the hemisphere title to a list.
hemisphere_image_urls = []
# List Hemisphere
links = browser.find_by_css('a.product-item img')
# List the links
for i in range(len(links)):
# Hemisphere info Dictionary
hemisphereInfo = {}
# Find elements on each loop to avoid a stale element exception
browser.find_by_css('a.product-item img')[i].click()
# Find the image
sample = browser.find_by_text('Sample').first
hemisphereInfo['img_url']=sample['href']
# Find the Hemisphere Title
hemisphereInfo['title'] = browser.find_by_css('h2.title').text
#Append into a list
hemisphere_image_urls.append(hemisphereInfo)
# Navigate backwards
browser.back()
# Appended list
hemisphere_image_urls
browser.quit()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from scipy.cluster.hierarchy import dendrogram, linkage
import numpy as np
import glob
from functools import partial
from scipy.signal import savgol_filter
from scipy.signal import argrelmax, find_peaks_cwt
from scipy.signal import find_peaks
pd.set_option('display.max_rows', 10000, 'display.max_columns', 10000)
import matplotlib
import matplotlib.pyplot as plt
matplotlib.use('Qt5Agg')
Mediana= pd.read_excel(r'/Users/thaismedeiros/Documentos/mesocosmos/espectros/Espectros_Ma_Mh_TF.xlsx', index_col=0, sheet_name="m_alcicornis"
).T
Mediana.head()
```
# fazendo as integrais de linha¶
```
df= pd.read_excel(r'/Users/thaismedeiros/Documentos/mesocosmos/espectros/Espectros_Ma_Mh_TF.xlsx', index_col=0, sheet_name="m_alcicornis"
).T
df.head()
#Remoção do continuo
def get_interpolated_value(vetor):
return np.trapz(vetor)
def get_continuum(vetor):
x= vetor.index[[0, -1]]
array= vetor.values
y = [array[0], array[-1]]
b, a= np.polyfit(x,y,1)
continuum=a+b*vetor.index
return continuum
def continuum_remove(v, lambda_min, lambda_max):
vetor = v.loc[slice(lambda_min,lambda_max)]
remover= get_continuum(vetor)
vetor2 = vetor.values-remover
r=get_interpolated_value(vetor2)
plt.axes()
plt.plot(np.arange(vetor.size),vetor.values, color='b', label='original_data')
plt.plot(np.arange(remover.size),remover, color='r', label='continuo_removido')
plt.legend()
return r
def get_profundidade_from_continuum(v, lambda_min, lambda_max):
vetor = v.loc[slice(lambda_min,lambda_max)]
remover= get_continuum(vetor)
vetor2 = vetor.values-remover
xmax = vetor2.max()
xmin = vetor2.min()
return xmax - xmin
def get_value_depth(v, lambda_min, lambda_max):
vetor = v.loc[slice(lambda_min,lambda_max)]
xmax = vetor.max()
xmin = vetor.min()
return xmax - xmin
def get_grau_funilamento(v, lambda_min, lambda_max):
altura= get_value_depth(v, lambda_min, lambda_max)
delta_lambda= (lambda_max - lambda_min)//2
v1= v.loc[slice(lambda_min,lambda_min+delta_lambda)]
v2= v.loc[slice(lambda_min+delta_lambda,lambda_max)]
esquerdo= v1.values[np.argmin(np.abs(altura-v1.values))]
direito= v2.values[np.argmin(np.abs(altura-v2.values))]
return np.abs(esquerdo-direito)
lista = []
lista_prof_continuum= []
lambdas = [(650, 750)]
nomes = []
for l in lambdas:
variable = df.apply(continuum_remove, lambda_min=l[0], lambda_max=l[1], axis=1)
nomes.append('Remocao_continuum_'+str(l[0])+'_'+str(l[1]))
lista.append(variable)
profundidade=df.apply(get_profundidade_from_continuum, lambda_min=l[0], lambda_max=l[1], axis=1)
lista_prof_continuum.append(profundidade)
df_temp1 = pd.DataFrame(data=lista, index=nomes).T
df_temp2 = pd.DataFrame(data=lista_prof_continuum, index=nomes).T
df_temp2.columns = ['prof_cont']
result_df= pd.concat([df_temp1,df_temp2], axis=1)
result_df
```
## Largura a meia altura
```
result_df['prof_vale_ref'] = df.apply(get_value_depth, lambda_min=l[0], lambda_max=l[1], axis=1)
result_df ['convectividade_funil'] = df.apply(get_grau_funilamento, lambda_min=l[0], lambda_max=l[1], axis=1)
result_df ['area'] = df.apply(lambda x:get_interpolated_value(x.loc[[l[0],l[1]]]), axis=1)
result_df
```
| github_jupyter |
This notebook was prepared by [Donne Martin](http://donnemartin.com). Source and license info is on [GitHub](https://github.com/donnemartin/data-science-ipython-notebooks).
# Kaggle Machine Learning Competition: Predicting Titanic Survivors
* Competition Site
* Description
* Evaluation
* Data Set
* Setup Imports and Variables
* Explore the Data
* Feature: Passenger Classes
* Feature: Sex
* Feature: Embarked
* Feature: Age
* Feature: Family Size
* Final Data Preparation for Machine Learning
* Data Wrangling Summary
* Random Forest: Training
* Random Forest: Predicting
* Random Forest: Prepare for Kaggle Submission
* Support Vector Machine: Training
* Support Vector Machine: Predicting
## Competition Site
Description, Evaluation, and Data Set taken from the [competition site](https://www.kaggle.com/c/titanic-gettingStarted).
## Description

The sinking of the RMS Titanic is one of the most infamous shipwrecks in history. On April 15, 1912, during her maiden voyage, the Titanic sank after colliding with an iceberg, killing 1502 out of 2224 passengers and crew. This sensational tragedy shocked the international community and led to better safety regulations for ships.
One of the reasons that the shipwreck led to such loss of life was that there were not enough lifeboats for the passengers and crew. Although there was some element of luck involved in surviving the sinking, some groups of people were more likely to survive than others, such as women, children, and the upper-class.
In this challenge, we ask you to complete the analysis of what sorts of people were likely to survive. In particular, we ask you to apply the tools of machine learning to predict which passengers survived the tragedy.
## Evaluation
The historical data has been split into two groups, a 'training set' and a 'test set'. For the training set, we provide the outcome ( 'ground truth' ) for each passenger. You will use this set to build your model to generate predictions for the test set.
For each passenger in the test set, you must predict whether or not they survived the sinking ( 0 for deceased, 1 for survived ). Your score is the percentage of passengers you correctly predict.
The Kaggle leaderboard has a public and private component. 50% of your predictions for the test set have been randomly assigned to the public leaderboard ( the same 50% for all users ). Your score on this public portion is what will appear on the leaderboard. At the end of the contest, we will reveal your score on the private 50% of the data, which will determine the final winner. This method prevents users from 'overfitting' to the leaderboard.
## Data Set
| File Name | Available Formats |
|------------------|-------------------|
| train | .csv (59.76 kb) |
| gendermodel | .csv (3.18 kb) |
| genderclassmodel | .csv (3.18 kb) |
| test | .csv (27.96 kb) |
| gendermodel | .py (3.58 kb) |
| genderclassmodel | .py (5.63 kb) |
| myfirstforest | .py (3.99 kb) |
<pre>
VARIABLE DESCRIPTIONS:
survival Survival
(0 = No; 1 = Yes)
pclass Passenger Class
(1 = 1st; 2 = 2nd; 3 = 3rd)
name Name
sex Sex
age Age
sibsp Number of Siblings/Spouses Aboard
parch Number of Parents/Children Aboard
ticket Ticket Number
fare Passenger Fare
cabin Cabin
embarked Port of Embarkation
(C = Cherbourg; Q = Queenstown; S = Southampton)
SPECIAL NOTES:
Pclass is a proxy for socio-economic status (SES)
1st ~ Upper; 2nd ~ Middle; 3rd ~ Lower
Age is in Years; Fractional if Age less than One (1)
If the Age is Estimated, it is in the form xx.5
With respect to the family relation variables (i.e. sibsp and parch)
some relations were ignored. The following are the definitions used
for sibsp and parch.
Sibling: Brother, Sister, Stepbrother, or Stepsister of Passenger Aboard Titanic
Spouse: Husband or Wife of Passenger Aboard Titanic (Mistresses and Fiances Ignored)
Parent: Mother or Father of Passenger Aboard Titanic
Child: Son, Daughter, Stepson, or Stepdaughter of Passenger Aboard Titanic
Other family relatives excluded from this study include cousins,
nephews/nieces, aunts/uncles, and in-laws. Some children travelled
only with a nanny, therefore parch=0 for them. As well, some
travelled with very close friends or neighbors in a village, however,
the definitions do not support such relations.
</pre>
## Setup Imports and Variables
```
import pandas as pd
import numpy as np
import pylab as plt
# Set the global default size of matplotlib figures
plt.rc('figure', figsize=(10, 5))
# Size of matplotlib figures that contain subplots
fizsize_with_subplots = (10, 10)
# Size of matplotlib histogram bins
bin_size = 10
```
## Explore the Data
Read the data:
```
df_train = pd.read_csv('../data/titanic/train.csv')
df_train.head()
df_train.tail()
```
View the data types of each column:
```
df_train.dtypes
```
Type 'object' is a string for pandas, which poses problems with machine learning algorithms. If we want to use these as features, we'll need to convert these to number representations.
Get some basic information on the DataFrame:
```
df_train.info()
```
Age, Cabin, and Embarked are missing values. Cabin has too many missing values, whereas we might be able to infer values for Age and Embarked.
Generate various descriptive statistics on the DataFrame:
```
df_train.describe()
```
Now that we have a general idea of the data set contents, we can dive deeper into each column. We'll be doing exploratory data analysis and cleaning data to setup 'features' we'll be using in our machine learning algorithms.
Plot a few features to get a better idea of each:
```
# Set up a grid of plots
fig = plt.figure(figsize=fizsize_with_subplots)
fig_dims = (3, 2)
# Plot death and survival counts
plt.subplot2grid(fig_dims, (0, 0))
df_train['Survived'].value_counts().plot(kind='bar',
title='Death and Survival Counts')
# Plot Pclass counts
plt.subplot2grid(fig_dims, (0, 1))
df_train['Pclass'].value_counts().plot(kind='bar',
title='Passenger Class Counts')
# Plot Sex counts
plt.subplot2grid(fig_dims, (1, 0))
df_train['Sex'].value_counts().plot(kind='bar',
title='Gender Counts')
plt.xticks(rotation=0)
# Plot Embarked counts
plt.subplot2grid(fig_dims, (1, 1))
df_train['Embarked'].value_counts().plot(kind='bar',
title='Ports of Embarkation Counts')
# Plot the Age histogram
plt.subplot2grid(fig_dims, (2, 0))
df_train['Age'].hist()
plt.title('Age Histogram')
```
Next we'll explore various features to view their impact on survival rates.
## Feature: Passenger Classes
From our exploratory data analysis in the previous section, we see there are three passenger classes: First, Second, and Third class. We'll determine which proportion of passengers survived based on their passenger class.
Generate a cross tab of Pclass and Survived:
```
pclass_xt = pd.crosstab(df_train['Pclass'], df_train['Survived'])
pclass_xt
```
Plot the cross tab:
```
# Normalize the cross tab to sum to 1:
pclass_xt_pct = pclass_xt.div(pclass_xt.sum(1).astype(float), axis=0)
pclass_xt_pct.plot(kind='bar',
stacked=True,
title='Survival Rate by Passenger Classes')
plt.xlabel('Passenger Class')
plt.ylabel('Survival Rate')
```
We can see that passenger class seems to have a significant impact on whether a passenger survived. Those in First Class the highest chance for survival.
## Feature: Sex
Gender might have also played a role in determining a passenger's survival rate. We'll need to map Sex from a string to a number to prepare it for machine learning algorithms.
Generate a mapping of Sex from a string to a number representation:
```
sexes = sorted(df_train['Sex'].unique())
genders_mapping = dict(zip(sexes, range(0, len(sexes) + 1)))
genders_mapping
```
Transform Sex from a string to a number representation:
```
df_train['Sex_Val'] = df_train['Sex'].map(genders_mapping).astype(int)
df_train.head()
```
Plot a normalized cross tab for Sex_Val and Survived:
```
sex_val_xt = pd.crosstab(df_train['Sex_Val'], df_train['Survived'])
sex_val_xt_pct = sex_val_xt.div(sex_val_xt.sum(1).astype(float), axis=0)
sex_val_xt_pct.plot(kind='bar', stacked=True, title='Survival Rate by Gender')
```
The majority of females survived, whereas the majority of males did not.
Next we'll determine whether we can gain any insights on survival rate by looking at both Sex and Pclass.
Count males and females in each Pclass:
```
# Get the unique values of Pclass:
passenger_classes = sorted(df_train['Pclass'].unique())
for p_class in passenger_classes:
print 'M: ', p_class, len(df_train[(df_train['Sex'] == 'male') &
(df_train['Pclass'] == p_class)])
print 'F: ', p_class, len(df_train[(df_train['Sex'] == 'female') &
(df_train['Pclass'] == p_class)])
```
Plot survival rate by Sex and Pclass:
```
# Plot survival rate by Sex
females_df = df_train[df_train['Sex'] == 'female']
females_xt = pd.crosstab(females_df['Pclass'], df_train['Survived'])
females_xt_pct = females_xt.div(females_xt.sum(1).astype(float), axis=0)
females_xt_pct.plot(kind='bar',
stacked=True,
title='Female Survival Rate by Passenger Class')
plt.xlabel('Passenger Class')
plt.ylabel('Survival Rate')
# Plot survival rate by Pclass
males_df = df_train[df_train['Sex'] == 'male']
males_xt = pd.crosstab(males_df['Pclass'], df_train['Survived'])
males_xt_pct = males_xt.div(males_xt.sum(1).astype(float), axis=0)
males_xt_pct.plot(kind='bar',
stacked=True,
title='Male Survival Rate by Passenger Class')
plt.xlabel('Passenger Class')
plt.ylabel('Survival Rate')
```
The vast majority of females in First and Second class survived. Males in First class had the highest chance for survival.
## Feature: Embarked
The Embarked column might be an important feature but it is missing a couple data points which might pose a problem for machine learning algorithms:
```
df_train[df_train['Embarked'].isnull()]
```
Prepare to map Embarked from a string to a number representation:
```
# Get the unique values of Embarked
embarked_locs = sorted(df_train['Embarked'].unique())
embarked_locs_mapping = dict(zip(embarked_locs,
range(0, len(embarked_locs) + 1)))
embarked_locs_mapping
```
Transform Embarked from a string to a number representation to prepare it for machine learning algorithms:
```
df_train['Embarked_Val'] = df_train['Embarked'] \
.map(embarked_locs_mapping) \
.astype(int)
df_train.head()
```
Plot the histogram for Embarked_Val:
```
df_train['Embarked_Val'].hist(bins=len(embarked_locs), range=(0, 3))
plt.title('Port of Embarkation Histogram')
plt.xlabel('Port of Embarkation')
plt.ylabel('Count')
plt.show()
```
Since the vast majority of passengers embarked in 'S': 3, we assign the missing values in Embarked to 'S':
```
if len(df_train[df_train['Embarked'].isnull()] > 0):
df_train.replace({'Embarked_Val' :
{ embarked_locs_mapping[nan] : embarked_locs_mapping['S']
}
},
inplace=True)
```
Verify we do not have any more NaNs for Embarked_Val:
```
embarked_locs = sorted(df_train['Embarked_Val'].unique())
embarked_locs
```
Plot a normalized cross tab for Embarked_Val and Survived:
```
embarked_val_xt = pd.crosstab(df_train['Embarked_Val'], df_train['Survived'])
embarked_val_xt_pct = \
embarked_val_xt.div(embarked_val_xt.sum(1).astype(float), axis=0)
embarked_val_xt_pct.plot(kind='bar', stacked=True)
plt.title('Survival Rate by Port of Embarkation')
plt.xlabel('Port of Embarkation')
plt.ylabel('Survival Rate')
```
It appears those that embarked in location 'C': 1 had the highest rate of survival. We'll dig in some more to see why this might be the case. Below we plot a graphs to determine gender and passenger class makeup for each port:
```
# Set up a grid of plots
fig = plt.figure(figsize=fizsize_with_subplots)
rows = 2
cols = 3
col_names = ('Sex_Val', 'Pclass')
for portIdx in embarked_locs:
for colIdx in range(0, len(col_names)):
plt.subplot2grid((rows, cols), (colIdx, portIdx - 1))
df_train[df_train['Embarked_Val'] == portIdx][col_names[colIdx]] \
.value_counts().plot(kind='bar')
```
Leaving Embarked as integers implies ordering in the values, which does not exist. Another way to represent Embarked without ordering is to create dummy variables:
```
df_train = pd.concat([df_train, pd.get_dummies(df_train['Embarked_Val'], prefix='Embarked_Val')], axis=1)
```
## Feature: Age
The Age column seems like an important feature--unfortunately it is missing many values. We'll need to fill in the missing values like we did with Embarked.
Filter to view missing Age values:
```
df_train[df_train['Age'].isnull()][['Sex', 'Pclass', 'Age']].head()
```
Determine the Age typical for each passenger class by Sex_Val. We'll use the median instead of the mean because the Age histogram seems to be right skewed.
```
# To keep Age in tact, make a copy of it called AgeFill
# that we will use to fill in the missing ages:
df_train['AgeFill'] = df_train['Age']
# Populate AgeFill
df_train['AgeFill'] = df_train['AgeFill'] \
.groupby([df_train['Sex_Val'], df_train['Pclass']]) \
.apply(lambda x: x.fillna(x.median()))
```
Ensure AgeFill does not contain any missing values:
```
len(df_train[df_train['AgeFill'].isnull()])
```
Plot a normalized cross tab for AgeFill and Survived:
```
# Set up a grid of plots
fig, axes = plt.subplots(2, 1, figsize=fizsize_with_subplots)
# Histogram of AgeFill segmented by Survived
df1 = df_train[df_train['Survived'] == 0]['Age']
df2 = df_train[df_train['Survived'] == 1]['Age']
max_age = max(df_train['AgeFill'])
axes[0].hist([df1, df2],
bins=max_age / bin_size,
range=(1, max_age),
stacked=True)
axes[0].legend(('Died', 'Survived'), loc='best')
axes[0].set_title('Survivors by Age Groups Histogram')
axes[0].set_xlabel('Age')
axes[0].set_ylabel('Count')
# Scatter plot Survived and AgeFill
axes[1].scatter(df_train['Survived'], df_train['AgeFill'])
axes[1].set_title('Survivors by Age Plot')
axes[1].set_xlabel('Survived')
axes[1].set_ylabel('Age')
```
Unfortunately, the graphs above do not seem to clearly show any insights. We'll keep digging further.
Plot AgeFill density by Pclass:
```
for pclass in passenger_classes:
df_train.AgeFill[df_train.Pclass == pclass].plot(kind='kde')
plt.title('Age Density Plot by Passenger Class')
plt.xlabel('Age')
plt.legend(('1st Class', '2nd Class', '3rd Class'), loc='best')
```
When looking at AgeFill density by Pclass, we see the first class passengers were generally older then second class passengers, which in turn were older than third class passengers. We've determined that first class passengers had a higher survival rate than second class passengers, which in turn had a higher survival rate than third class passengers.
```
# Set up a grid of plots
fig = plt.figure(figsize=fizsize_with_subplots)
fig_dims = (3, 1)
# Plot the AgeFill histogram for Survivors
plt.subplot2grid(fig_dims, (0, 0))
survived_df = df_train[df_train['Survived'] == 1]
survived_df['AgeFill'].hist(bins=max_age / bin_size, range=(1, max_age))
# Plot the AgeFill histogram for Females
plt.subplot2grid(fig_dims, (1, 0))
females_df = df_train[(df_train['Sex_Val'] == 0) & (df_train['Survived'] == 1)]
females_df['AgeFill'].hist(bins=max_age / bin_size, range=(1, max_age))
# Plot the AgeFill histogram for first class passengers
plt.subplot2grid(fig_dims, (2, 0))
class1_df = df_train[(df_train['Pclass'] == 1) & (df_train['Survived'] == 1)]
class1_df['AgeFill'].hist(bins=max_age / bin_size, range=(1, max_age))
```
In the first graph, we see that most survivors come from the 20's to 30's age ranges and might be explained by the following two graphs. The second graph shows most females are within their 20's. The third graph shows most first class passengers are within their 30's.
## Feature: Family Size
Feature enginering involves creating new features or modifying existing features which might be advantageous to a machine learning algorithm.
Define a new feature FamilySize that is the sum of Parch (number of parents or children on board) and SibSp (number of siblings or spouses):
```
df_train['FamilySize'] = df_train['SibSp'] + df_train['Parch']
df_train.head()
```
Plot a histogram of FamilySize:
```
df_train['FamilySize'].hist()
plt.title('Family Size Histogram')
```
Plot a histogram of AgeFill segmented by Survived:
```
# Get the unique values of Embarked and its maximum
family_sizes = sorted(df_train['FamilySize'].unique())
family_size_max = max(family_sizes)
df1 = df_train[df_train['Survived'] == 0]['FamilySize']
df2 = df_train[df_train['Survived'] == 1]['FamilySize']
plt.hist([df1, df2],
bins=family_size_max + 1,
range=(0, family_size_max),
stacked=True)
plt.legend(('Died', 'Survived'), loc='best')
plt.title('Survivors by Family Size')
```
Based on the histograms, it is not immediately obvious what impact FamilySize has on survival. The machine learning algorithms might benefit from this feature.
Additional features we might want to engineer might be related to the Name column, for example honorrary or pedestrian titles might give clues and better predictive power for a male's survival.
## Final Data Preparation for Machine Learning
Many machine learning algorithms do not work on strings and they usually require the data to be in an array, not a DataFrame.
Show only the columns of type 'object' (strings):
```
df_train.dtypes[df_train.dtypes.map(lambda x: x == 'object')]
```
Drop the columns we won't use:
```
df_train = df_train.drop(['Name', 'Sex', 'Ticket', 'Cabin', 'Embarked'],
axis=1)
```
Drop the following columns:
* The Age column since we will be using the AgeFill column instead.
* The SibSp and Parch columns since we will be using FamilySize instead.
* The PassengerId column since it won't be used as a feature.
* The Embarked_Val as we decided to use dummy variables instead.
```
df_train = df_train.drop(['Age', 'SibSp', 'Parch', 'PassengerId', 'Embarked_Val'], axis=1)
df_train.dtypes
```
Convert the DataFrame to a numpy array:
```
train_data = df_train.values
train_data
```
## Data Wrangling Summary
Below is a summary of the data wrangling we performed on our training data set. We encapsulate this in a function since we'll need to do the same operations to our test set later.
```
def clean_data(df, drop_passenger_id):
# Get the unique values of Sex
sexes = sorted(df['Sex'].unique())
# Generate a mapping of Sex from a string to a number representation
genders_mapping = dict(zip(sexes, range(0, len(sexes) + 1)))
# Transform Sex from a string to a number representation
df['Sex_Val'] = df['Sex'].map(genders_mapping).astype(int)
# Get the unique values of Embarked
embarked_locs = sorted(df['Embarked'].unique())
# Generate a mapping of Embarked from a string to a number representation
embarked_locs_mapping = dict(zip(embarked_locs,
range(0, len(embarked_locs) + 1)))
# Transform Embarked from a string to dummy variables
df = pd.concat([df, pd.get_dummies(df['Embarked'], prefix='Embarked_Val')], axis=1)
# Fill in missing values of Embarked
# Since the vast majority of passengers embarked in 'S': 3,
# we assign the missing values in Embarked to 'S':
if len(df[df['Embarked'].isnull()] > 0):
df.replace({'Embarked_Val' :
{ embarked_locs_mapping[nan] : embarked_locs_mapping['S']
}
},
inplace=True)
# Fill in missing values of Fare with the average Fare
if len(df[df['Fare'].isnull()] > 0):
avg_fare = df['Fare'].mean()
df.replace({ None: avg_fare }, inplace=True)
# To keep Age in tact, make a copy of it called AgeFill
# that we will use to fill in the missing ages:
df['AgeFill'] = df['Age']
# Determine the Age typical for each passenger class by Sex_Val.
# We'll use the median instead of the mean because the Age
# histogram seems to be right skewed.
df['AgeFill'] = df['AgeFill'] \
.groupby([df['Sex_Val'], df['Pclass']]) \
.apply(lambda x: x.fillna(x.median()))
# Define a new feature FamilySize that is the sum of
# Parch (number of parents or children on board) and
# SibSp (number of siblings or spouses):
df['FamilySize'] = df['SibSp'] + df['Parch']
# Drop the columns we won't use:
df = df.drop(['Name', 'Sex', 'Ticket', 'Cabin', 'Embarked'], axis=1)
# Drop the Age column since we will be using the AgeFill column instead.
# Drop the SibSp and Parch columns since we will be using FamilySize.
# Drop the PassengerId column since it won't be used as a feature.
df = df.drop(['Age', 'SibSp', 'Parch'], axis=1)
if drop_passenger_id:
df = df.drop(['PassengerId'], axis=1)
return df
```
## Random Forest: Training
Create the random forest object:
```
from sklearn.ensemble import RandomForestClassifier
clf = RandomForestClassifier(n_estimators=100)
```
Fit the training data and create the decision trees:
```
# Training data features, skip the first column 'Survived'
train_features = train_data[:, 1:]
# 'Survived' column values
train_target = train_data[:, 0]
# Fit the model to our training data
clf = clf.fit(train_features, train_target)
score = clf.score(train_features, train_target)
"Mean accuracy of Random Forest: {0}".format(score)
```
## Random Forest: Predicting
Read the test data:
```
df_test = pd.read_csv('../data/titanic/test.csv')
df_test.head()
```
Note the test data does not contain the column 'Survived', we'll use our trained model to predict these values.
```
# Data wrangle the test set and convert it to a numpy array
df_test = clean_data(df_test, drop_passenger_id=False)
test_data = df_test.values
```
Take the decision trees and run it on the test data:
```
# Get the test data features, skipping the first column 'PassengerId'
test_x = test_data[:, 1:]
# Predict the Survival values for the test data
test_y = clf.predict(test_x)
```
## Random Forest: Prepare for Kaggle Submission
Create a DataFrame by combining the index from the test data with the output of predictions, then write the results to the output:
```
df_test['Survived'] = test_y
df_test[['PassengerId', 'Survived']] \
.to_csv('../data/titanic/results-rf.csv', index=False)
```
## Evaluate Model Accuracy
Submitting to Kaggle will give you an accuracy score. It would be helpful to get an idea of accuracy without submitting to Kaggle.
We'll split our training data, 80% will go to "train" and 20% will go to "test":
```
from sklearn import metrics
from sklearn.cross_validation import train_test_split
# Split 80-20 train vs test data
train_x, test_x, train_y, test_y = train_test_split(train_features,
train_target,
test_size=0.20,
random_state=0)
print (train_features.shape, train_target.shape)
print (train_x.shape, train_y.shape)
print (test_x.shape, test_y.shape)
```
Use the new training data to fit the model, predict, and get the accuracy score:
```
clf = clf.fit(train_x, train_y)
predict_y = clf.predict(test_x)
from sklearn.metrics import accuracy_score
print ("Accuracy = %.2f" % (accuracy_score(test_y, predict_y)))
```
View the Confusion Matrix:
| | condition True | condition false|
|------|----------------|---------------|
|prediction true|True Positive|False positive|
|Prediction False|False Negative|True Negative|
```
from IPython.core.display import Image
Image(filename='../data/confusion_matrix.png', width=800)
```
Get the model score and confusion matrix:
```
model_score = clf.score(test_x, test_y)
print ("Model Score %.2f \n" % (model_score))
confusion_matrix = metrics.confusion_matrix(test_y, predict_y)
print ("Confusion Matrix ", confusion_matrix)
print (" Predicted")
print (" | 0 | 1 |")
print (" |-----|-----|")
print (" 0 | %3d | %3d |" % (confusion_matrix[0, 0],
confusion_matrix[0, 1]))
print ("Actual |-----|-----|")
print (" 1 | %3d | %3d |" % (confusion_matrix[1, 0],
confusion_matrix[1, 1]))
print (" |-----|-----|")
```
Display the classification report:
$$Precision = \frac{TP}{TP + FP}$$
$$Recall = \frac{TP}{TP + FN}$$
$$F1 = \frac{2TP}{2TP + FP + FN}$$
```
from sklearn.metrics import classification_report
print(classification_report(test_y,
predict_y,
target_names=['Not Survived', 'Survived']))
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.