code stringlengths 2.5k 150k | kind stringclasses 1 value |
|---|---|
<a href="https://colab.research.google.com/github/kpe/bert-for-tf2/blob/master/examples/movie_reviews_with_bert_for_tf2_on_gpu.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
This is a modification of https://github/google-research/bert/blob/master/predicting_movie_reviews_with_bert_on_tf_hub.ipynb using the Tensorflow 2.0 Keras implementation of BERT from [kpe/bert-for-tf2](https://github.com/kpe/bert-for-tf2) with the original [google-research/bert](https://github.com/google-research/bert) weights.
```
# Copyright 2019 Google Inc.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Predicting Movie Review Sentiment with [kpe/bert-for-tf2](https://github.com/kpe/bert-for-tf2)
First install some prerequisites:
```
!pip install tqdm >> /dev/null
import os
import math
import datetime
from tqdm import tqdm
import pandas as pd
import numpy as np
import tensorflow as tf
tf.__version__
if tf.__version__.startswith("1."):
tf.enable_eager_execution()
```
In addition to the standard libraries we imported above, we'll need to install the [bert-for-tf2](https://github.com/kpe/bert-for-tf2) python package, and do the imports required for loading the pre-trained weights and tokenizing the input text.
```
!pip install bert-for-tf2 >> /dev/null
import bert
from bert import BertModelLayer
from bert.loader import StockBertConfig, map_stock_config_to_params, load_stock_weights
from bert.tokenization.bert_tokenization import FullTokenizer
```
#Data
First, let's download the dataset, hosted by Stanford. The code below, which downloads, extracts, and imports the IMDB Large Movie Review Dataset, is borrowed from [this Tensorflow tutorial](https://www.tensorflow.org/hub/tutorials/text_classification_with_tf_hub).
```
from tensorflow import keras
import os
import re
# Load all files from a directory in a DataFrame.
def load_directory_data(directory):
data = {}
data["sentence"] = []
data["sentiment"] = []
for file_path in tqdm(os.listdir(directory), desc=os.path.basename(directory)):
with tf.io.gfile.GFile(os.path.join(directory, file_path), "r") as f:
data["sentence"].append(f.read())
data["sentiment"].append(re.match("\d+_(\d+)\.txt", file_path).group(1))
return pd.DataFrame.from_dict(data)
# Merge positive and negative examples, add a polarity column and shuffle.
def load_dataset(directory):
pos_df = load_directory_data(os.path.join(directory, "pos"))
neg_df = load_directory_data(os.path.join(directory, "neg"))
pos_df["polarity"] = 1
neg_df["polarity"] = 0
return pd.concat([pos_df, neg_df]).sample(frac=1).reset_index(drop=True)
# Download and process the dataset files.
def download_and_load_datasets(force_download=False):
dataset = tf.keras.utils.get_file(
fname="aclImdb.tar.gz",
origin="http://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz",
extract=True)
train_df = load_dataset(os.path.join(os.path.dirname(dataset),
"aclImdb", "train"))
test_df = load_dataset(os.path.join(os.path.dirname(dataset),
"aclImdb", "test"))
return train_df, test_df
```
Let's use the `MovieReviewData` class below, to prepare/encode
the data for feeding into our BERT model, by:
- tokenizing the text
- trim or pad it to a `max_seq_len` length
- append the special tokens `[CLS]` and `[SEP]`
- convert the string tokens to numerical `ID`s using the original model's token encoding from `vocab.txt`
```
import bert
from bert import BertModelLayer
from bert.loader import StockBertConfig, map_stock_config_to_params, load_stock_weights
from bert.tokenization import FullTokenizer
class MovieReviewData:
DATA_COLUMN = "sentence"
LABEL_COLUMN = "polarity"
def __init__(self, tokenizer: FullTokenizer, sample_size=None, max_seq_len=1024):
self.tokenizer = tokenizer
self.sample_size = sample_size
self.max_seq_len = 0
train, test = download_and_load_datasets()
train, test = map(lambda df: df.reindex(df[MovieReviewData.DATA_COLUMN].str.len().sort_values().index),
[train, test])
if sample_size is not None:
assert sample_size % 128 == 0
train, test = train.head(sample_size), test.head(sample_size)
# train, test = map(lambda df: df.sample(sample_size), [train, test])
((self.train_x, self.train_y),
(self.test_x, self.test_y)) = map(self._prepare, [train, test])
print("max seq_len", self.max_seq_len)
self.max_seq_len = min(self.max_seq_len, max_seq_len)
((self.train_x, self.train_x_token_types),
(self.test_x, self.test_x_token_types)) = map(self._pad,
[self.train_x, self.test_x])
def _prepare(self, df):
x, y = [], []
with tqdm(total=df.shape[0], unit_scale=True) as pbar:
for ndx, row in df.iterrows():
text, label = row[MovieReviewData.DATA_COLUMN], row[MovieReviewData.LABEL_COLUMN]
tokens = self.tokenizer.tokenize(text)
tokens = ["[CLS]"] + tokens + ["[SEP]"]
token_ids = self.tokenizer.convert_tokens_to_ids(tokens)
self.max_seq_len = max(self.max_seq_len, len(token_ids))
x.append(token_ids)
y.append(int(label))
pbar.update()
return np.array(x), np.array(y)
def _pad(self, ids):
x, t = [], []
token_type_ids = [0] * self.max_seq_len
for input_ids in ids:
input_ids = input_ids[:min(len(input_ids), self.max_seq_len - 2)]
input_ids = input_ids + [0] * (self.max_seq_len - len(input_ids))
x.append(np.array(input_ids))
t.append(token_type_ids)
return np.array(x), np.array(t)
```
## A tweak
Because of a `tf.train.load_checkpoint` limitation requiring list permissions on the google storage bucket, we need to copy the pre-trained BERT weights locally.
```
bert_ckpt_dir="gs://bert_models/2018_10_18/uncased_L-12_H-768_A-12/"
bert_ckpt_file = bert_ckpt_dir + "bert_model.ckpt"
bert_config_file = bert_ckpt_dir + "bert_config.json"
%%time
bert_model_dir="2018_10_18"
bert_model_name="uncased_L-12_H-768_A-12"
!mkdir -p .model .model/$bert_model_name
for fname in ["bert_config.json", "vocab.txt", "bert_model.ckpt.meta", "bert_model.ckpt.index", "bert_model.ckpt.data-00000-of-00001"]:
cmd = f"gsutil cp gs://bert_models/{bert_model_dir}/{bert_model_name}/{fname} .model/{bert_model_name}"
!$cmd
!ls -la .model .model/$bert_model_name
bert_ckpt_dir = os.path.join(".model/",bert_model_name)
bert_ckpt_file = os.path.join(bert_ckpt_dir, "bert_model.ckpt")
bert_config_file = os.path.join(bert_ckpt_dir, "bert_config.json")
```
# Preparing the Data
Now let's fetch and prepare the data by taking the first `max_seq_len` tokenens after tokenizing with the BERT tokenizer, und use `sample_size` examples for both training and testing.
To keep training fast, we'll take a sample of about 2500 train and test examples, respectively, and use the first 128 tokens only (transformers memory and computation requirements scale quadraticly with the sequence length - so with a TPU you might use `max_seq_len=512`, but on a GPU this would be too slow, and you will have to use a very small `batch_size`s to fit the model into the GPU memory).
```
%%time
tokenizer = FullTokenizer(vocab_file=os.path.join(bert_ckpt_dir, "vocab.txt"))
data = MovieReviewData(tokenizer,
sample_size=10*128*2,#5000,
max_seq_len=128)
print(" train_x", data.train_x.shape)
print("train_x_token_types", data.train_x_token_types.shape)
print(" train_y", data.train_y.shape)
print(" test_x", data.test_x.shape)
print(" max_seq_len", data.max_seq_len)
```
## Adapter BERT
If we decide to use [adapter-BERT](https://arxiv.org/abs/1902.00751) we need some helpers for freezing the original BERT layers.
```
def flatten_layers(root_layer):
if isinstance(root_layer, keras.layers.Layer):
yield root_layer
for layer in root_layer._layers:
for sub_layer in flatten_layers(layer):
yield sub_layer
def freeze_bert_layers(l_bert):
"""
Freezes all but LayerNorm and adapter layers - see arXiv:1902.00751.
"""
for layer in flatten_layers(l_bert):
if layer.name in ["LayerNorm", "adapter-down", "adapter-up"]:
layer.trainable = True
elif len(layer._layers) == 0:
layer.trainable = False
l_bert.embeddings_layer.trainable = False
def create_learning_rate_scheduler(max_learn_rate=5e-5,
end_learn_rate=1e-7,
warmup_epoch_count=10,
total_epoch_count=90):
def lr_scheduler(epoch):
if epoch < warmup_epoch_count:
res = (max_learn_rate/warmup_epoch_count) * (epoch + 1)
else:
res = max_learn_rate*math.exp(math.log(end_learn_rate/max_learn_rate)*(epoch-warmup_epoch_count+1)/(total_epoch_count-warmup_epoch_count+1))
return float(res)
learning_rate_scheduler = tf.keras.callbacks.LearningRateScheduler(lr_scheduler, verbose=1)
return learning_rate_scheduler
```
#Creating a model
Now let's create a classification model using [adapter-BERT](https//arxiv.org/abs/1902.00751), which is clever way of reducing the trainable parameter count, by freezing the original BERT weights, and adapting them with two FFN bottlenecks (i.e. `adapter_size` bellow) in every BERT layer.
**N.B.** The commented out code below show how to feed a `token_type_ids`/`segment_ids` sequence (which is not needed in our case).
```
def create_model(max_seq_len, adapter_size=64):
"""Creates a classification model."""
#adapter_size = 64 # see - arXiv:1902.00751
# create the bert layer
with tf.io.gfile.GFile(bert_config_file, "r") as reader:
bc = StockBertConfig.from_json_string(reader.read())
bert_params = map_stock_config_to_params(bc)
bert_params.adapter_size = adapter_size
bert = BertModelLayer.from_params(bert_params, name="bert")
input_ids = keras.layers.Input(shape=(max_seq_len,), dtype='int32', name="input_ids")
# token_type_ids = keras.layers.Input(shape=(max_seq_len,), dtype='int32', name="token_type_ids")
# output = bert([input_ids, token_type_ids])
output = bert(input_ids)
print("bert shape", output.shape)
cls_out = keras.layers.Lambda(lambda seq: seq[:, 0, :])(output)
cls_out = keras.layers.Dropout(0.5)(cls_out)
logits = keras.layers.Dense(units=768, activation="tanh")(cls_out)
logits = keras.layers.Dropout(0.5)(logits)
logits = keras.layers.Dense(units=2, activation="softmax")(logits)
# model = keras.Model(inputs=[input_ids, token_type_ids], outputs=logits)
# model.build(input_shape=[(None, max_seq_len), (None, max_seq_len)])
model = keras.Model(inputs=input_ids, outputs=logits)
model.build(input_shape=(None, max_seq_len))
# load the pre-trained model weights
load_stock_weights(bert, bert_ckpt_file)
# freeze weights if adapter-BERT is used
if adapter_size is not None:
freeze_bert_layers(bert)
model.compile(optimizer=keras.optimizers.Adam(),
loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=[keras.metrics.SparseCategoricalAccuracy(name="acc")])
model.summary()
return model
adapter_size = None # use None to fine-tune all of BERT
model = create_model(data.max_seq_len, adapter_size=adapter_size)
%%time
log_dir = ".log/movie_reviews/" + datetime.datetime.now().strftime("%Y%m%d-%H%M%s")
tensorboard_callback = keras.callbacks.TensorBoard(log_dir=log_dir)
total_epoch_count = 50
# model.fit(x=(data.train_x, data.train_x_token_types), y=data.train_y,
model.fit(x=data.train_x, y=data.train_y,
validation_split=0.1,
batch_size=48,
shuffle=True,
epochs=total_epoch_count,
callbacks=[create_learning_rate_scheduler(max_learn_rate=1e-5,
end_learn_rate=1e-7,
warmup_epoch_count=20,
total_epoch_count=total_epoch_count),
keras.callbacks.EarlyStopping(patience=20, restore_best_weights=True),
tensorboard_callback])
model.save_weights('./movie_reviews.h5', overwrite=True)
%%time
_, train_acc = model.evaluate(data.train_x, data.train_y)
_, test_acc = model.evaluate(data.test_x, data.test_y)
print("train acc", train_acc)
print(" test acc", test_acc)
```
# Evaluation
To evaluate the trained model, let's load the saved weights in a new model instance, and evaluate.
```
%%time
model = create_model(data.max_seq_len, adapter_size=None)
model.load_weights("movie_reviews.h5")
_, train_acc = model.evaluate(data.train_x, data.train_y)
_, test_acc = model.evaluate(data.test_x, data.test_y)
print("train acc", train_acc)
print(" test acc", test_acc)
```
# Prediction
For prediction, we need to prepare the input text the same way as we did for training - tokenize, adding the special `[CLS]` and `[SEP]` token at begin and end of the token sequence, and pad to match the model input shape.
```
pred_sentences = [
"That movie was absolutely awful",
"The acting was a bit lacking",
"The film was creative and surprising",
"Absolutely fantastic!"
]
tokenizer = FullTokenizer(vocab_file=os.path.join(bert_ckpt_dir, "vocab.txt"))
pred_tokens = map(tokenizer.tokenize, pred_sentences)
pred_tokens = map(lambda tok: ["[CLS]"] + tok + ["[SEP]"], pred_tokens)
pred_token_ids = list(map(tokenizer.convert_tokens_to_ids, pred_tokens))
pred_token_ids = map(lambda tids: tids +[0]*(data.max_seq_len-len(tids)),pred_token_ids)
pred_token_ids = np.array(list(pred_token_ids))
print('pred_token_ids', pred_token_ids.shape)
res = model.predict(pred_token_ids).argmax(axis=-1)
for text, sentiment in zip(pred_sentences, res):
print(" text:", text)
print(" res:", ["negative","positive"][sentiment])
```
| github_jupyter |
# Torch Connector and Hybrid QNNs
This tutorial introduces Qiskit's `TorchConnector` class, and demonstrates how the `TorchConnector` allows for a natural integration of any `NeuralNetwork` from Qiskit Machine Learning into a PyTorch workflow. `TorchConnector` takes a Qiskit `NeuralNetwork` and makes it available as a PyTorch `Module`. The resulting module can be seamlessly incorporated into PyTorch classical architectures and trained jointly without additional considerations, enabling the development and testing of novel **hybrid quantum-classical** machine learning architectures.
## Content:
[Part 1: Simple Classification & Regression](#Part-1:-Simple-Classification-&-Regression)
The first part of this tutorial shows how quantum neural networks can be trained using PyTorch's automatic differentiation engine (`torch.autograd`, [link](https://pytorch.org/tutorials/beginner/blitz/autograd_tutorial.html)) for simple classification and regression tasks.
1. [Classification](#1.-Classification)
1. Classification with PyTorch and `OpflowQNN`
2. Classification with PyTorch and `CircuitQNN`
2. [Regression](#2.-Regression)
1. Regression with PyTorch and `OpflowQNN`
[Part 2: MNIST Classification, Hybrid QNNs](#Part-2:-MNIST-Classification,-Hybrid-QNNs)
The second part of this tutorial illustrates how to embed a (Quantum) `NeuralNetwork` into a target PyTorch workflow (in this case, a typical CNN architecture) to classify MNIST data in a hybrid quantum-classical manner.
***
```
# Necessary imports
import numpy as np
import matplotlib.pyplot as plt
from torch import Tensor
from torch.nn import Linear, CrossEntropyLoss, MSELoss
from torch.optim import LBFGS
from qiskit import Aer, QuantumCircuit
from qiskit.utils import QuantumInstance, algorithm_globals
from qiskit.opflow import AerPauliExpectation
from qiskit.circuit import Parameter
from qiskit.circuit.library import RealAmplitudes, ZZFeatureMap
from qiskit_machine_learning.neural_networks import CircuitQNN, TwoLayerQNN
from qiskit_machine_learning.connectors import TorchConnector
# Set seed for random generators
algorithm_globals.random_seed = 42
# declare quantum instance
qi = QuantumInstance(Aer.get_backend('aer_simulator_statevector'))
```
## Part 1: Simple Classification & Regression
### 1. Classification
First, we show how `TorchConnector` allows to train a Quantum `NeuralNetwork` to solve a classification tasks using PyTorch's automatic differentiation engine. In order to illustrate this, we will perform **binary classification** on a randomly generated dataset.
```
# Generate random dataset
# Select dataset dimension (num_inputs) and size (num_samples)
num_inputs = 2
num_samples = 20
# Generate random input coordinates (X) and binary labels (y)
X = 2*algorithm_globals.random.random([num_samples, num_inputs]) - 1
y01 = 1*(np.sum(X, axis=1) >= 0) # in { 0, 1}, y01 will be used for CircuitQNN example
y = 2*y01-1 # in {-1, +1}, y will be used for OplowQNN example
# Convert to torch Tensors
X_ = Tensor(X)
y01_ = Tensor(y01).reshape(len(y)).long()
y_ = Tensor(y).reshape(len(y), 1)
# Plot dataset
for x, y_target in zip(X, y):
if y_target == 1:
plt.plot(x[0], x[1], 'bo')
else:
plt.plot(x[0], x[1], 'go')
plt.plot([-1, 1], [1, -1], '--', color='black')
plt.show()
```
#### A. Classification with PyTorch and `OpflowQNN`
Linking an `OpflowQNN` to PyTorch is relatively straightforward. Here we illustrate this using the `TwoLayerQNN`, a sub-case of `OpflowQNN` introduced in previous tutorials.
```
# Set up QNN
# Note: we are not providing them explicitly in this examples,
# but TwoLayerQNN requires a feature_map and ansatz to work.
# By default, these parameters are set to ZZFeatureMap
# and RealAmplitudes (respectively).
qnn1 = TwoLayerQNN(num_qubits=num_inputs, quantum_instance=qi)
print(qnn1.operator)
# Set up PyTorch module
# Note: If we don't explicitly declare the initial weights
# they are chosen uniformly at random from [-1, 1].
initial_weights = 0.1*(2*algorithm_globals.random.random(qnn1.num_weights) - 1)
model1 = TorchConnector(qnn1, initial_weights=initial_weights)
print("Initial weights: ", initial_weights)
# Test with a single input
model1(X_[0, :])
```
##### Optimizer
The choice of optimizer for training any machine learning model can be crucial in determining the success of our training's outcome. When using `TorchConnector`, we get access to all of the optimizer algorithms defined in the [`torch.optim`] package ([link](https://pytorch.org/docs/stable/optim.html)). Some of the most famous algorithms used in popular machine learning architectures include *Adam*, *SGD*, or *Adagrad*. However, for this tutorial we will be using the L-BFGS algorithm (`torch.optim.LBFGS`), one of the most well know second-order optimization algorithms for numerical optimization.
##### Loss Function
As for the loss function, we can also take advantage of PyTorch's pre-defined modules from `torch.nn`, such as the [Cross-Entropy](https://pytorch.org/docs/stable/generated/torch.nn.CrossEntropyLoss.html) or [Mean Squared Error](https://pytorch.org/docs/stable/generated/torch.nn.MSELoss.html) losses.
**💡 Clarification :**
In classical machine learning, the general rule of thumb is to apply a Cross-Entropy loss to classification tasks, and MSE loss to regression tasks. However, this recommendation is given under the assumption that the output of the classification network is a class probability value in the [0,1] range (usually this is achieved through a Softmax layer). Because the following example for `TwoLayerQNN` does not include such layer, and we don't apply any mapping to the output (the following section shows an example of application of parity mapping with `CircuitQNNs`), the QNN's output can take any value in the range [-1,1]. In case you were wondering, this is the reason why this particular example uses MSELoss for classification despite it not being the norm (but we encourage you to experiment with different loss functions and see how they can impact training results).
```
# Define optimizer and loss
optimizer = LBFGS(model1.parameters())
f_loss = MSELoss(reduction='sum')
# Start training
model1.train() # set model to training mode
# Note from (https://pytorch.org/docs/stable/optim.html):
# Some optimization algorithms such as LBFGS need to
# reevaluate the function multiple times, so you have to
# pass in a closure that allows them to recompute your model.
# The closure should clear the gradients, compute the loss,
# and return it.
def closure():
optimizer.zero_grad() # Initialize/clear gradients
loss = f_loss(model1(X_), y_) # Evaluate loss function
loss.backward() # Backward pass
print(loss.item()) # Print loss
return loss
# Run optimizer step4
optimizer.step(closure)
# Evaluate model and compute accuracy
y_predict = []
for x, y_target in zip(X, y):
output = model1(Tensor(x))
y_predict += [np.sign(output.detach().numpy())[0]]
print('Accuracy:', sum(y_predict == y)/len(y))
# Plot results
# red == wrongly classified
for x, y_target, y_p in zip(X, y, y_predict):
if y_target == 1:
plt.plot(x[0], x[1], 'bo')
else:
plt.plot(x[0], x[1], 'go')
if y_target != y_p:
plt.scatter(x[0], x[1], s=200, facecolors='none', edgecolors='r', linewidths=2)
plt.plot([-1, 1], [1, -1], '--', color='black')
plt.show()
```
The red circles indicate wrongly classified data points.
#### B. Classification with PyTorch and `CircuitQNN`
Linking an `CircuitQNN` to PyTorch requires a bit more attention than `OpflowQNN`. Without the correct setup, backpropagation is not possible.
In particular, we must make sure that we are returning a dense array of probabilities in the network's forward pass (`sparse=False`). This parameter is set up to `False` by default, so we just have to make sure that it has not been changed.
**⚠️ Attention:**
If we define a custom interpret function ( in the example: `parity`), we must remember to explicitly provide the desired output shape ( in the example: `2`). For more info on the initial parameter setup for `CircuitQNN`, please check out the [official qiskit documentation](https://qiskit.org/documentation/machine-learning/stubs/qiskit_machine_learning.neural_networks.CircuitQNN.html).
```
# Define feature map and ansatz
feature_map = ZZFeatureMap(num_inputs)
ansatz = RealAmplitudes(num_inputs, entanglement='linear', reps=1)
# Define quantum circuit of num_qubits = input dim
# Append feature map and ansatz
qc = QuantumCircuit(num_inputs)
qc.append(feature_map, range(num_inputs))
qc.append(ansatz, range(num_inputs))
# Define CircuitQNN and initial setup
parity = lambda x: '{:b}'.format(x).count('1') % 2 # optional interpret function
output_shape = 2 # parity = 0, 1
qnn2 = CircuitQNN(qc, input_params=feature_map.parameters, weight_params=ansatz.parameters,
interpret=parity, output_shape=output_shape, quantum_instance=qi)
# Set up PyTorch module
# Reminder: If we don't explicitly declare the initial weights
# they are chosen uniformly at random from [-1, 1].
initial_weights = 0.1*(2*algorithm_globals.random.random(qnn2.num_weights) - 1)
print("Initial weights: ", initial_weights)
model2 = TorchConnector(qnn2, initial_weights)
```
For a reminder on optimizer and loss function choices, you can go back to [this section](#Optimizer).
```
# Define model, optimizer, and loss
optimizer = LBFGS(model2.parameters())
f_loss = CrossEntropyLoss() # Our output will be in the [0,1] range
# Start training
model2.train()
# Define LBFGS closure method (explained in previous section)
def closure():
optimizer.zero_grad(set_to_none=True) # Initialize gradient
loss = f_loss(model2(X_), y01_) # Calculate loss
loss.backward() # Backward pass
print(loss.item()) # Print loss
return loss
# Run optimizer (LBFGS requires closure)
optimizer.step(closure);
# Evaluate model and compute accuracy
y_predict = []
for x in X:
output = model2(Tensor(x))
y_predict += [np.argmax(output.detach().numpy())]
print('Accuracy:', sum(y_predict == y01)/len(y01))
# plot results
# red == wrongly classified
for x, y_target, y_ in zip(X, y01, y_predict):
if y_target == 1:
plt.plot(x[0], x[1], 'bo')
else:
plt.plot(x[0], x[1], 'go')
if y_target != y_:
plt.scatter(x[0], x[1], s=200, facecolors='none', edgecolors='r', linewidths=2)
plt.plot([-1, 1], [1, -1], '--', color='black')
plt.show()
```
The red circles indicate wrongly classified data points.
### 2. Regression
We use a model based on the `TwoLayerQNN` to also illustrate how to perform a regression task. The chosen dataset in this case is randomly generated following a sine wave.
```
# Generate random dataset
num_samples = 20
eps = 0.2
lb, ub = -np.pi, np.pi
f = lambda x: np.sin(x)
X = (ub - lb)*algorithm_globals.random.random([num_samples, 1]) + lb
y = f(X) + eps*(2*algorithm_globals.random.random([num_samples, 1])-1)
plt.plot(np.linspace(lb, ub), f(np.linspace(lb, ub)), 'r--')
plt.plot(X, y, 'bo')
plt.show()
```
#### A. Regression with PyTorch and `OpflowQNN`
The network definition and training loop will be analogous to those of the classification task using `TwoLayerQNN`. In this case, we define our own feature map and ansatz, instead of using the default values.
```
# Construct simple feature map
param_x = Parameter('x')
feature_map = QuantumCircuit(1, name='fm')
feature_map.ry(param_x, 0)
# Construct simple feature map
param_y = Parameter('y')
ansatz = QuantumCircuit(1, name='vf')
ansatz.ry(param_y, 0)
# Construct QNN
qnn3 = TwoLayerQNN(1, feature_map, ansatz, quantum_instance=qi)
print(qnn3.operator)
# Set up PyTorch module
# Reminder: If we don't explicitly declare the initial weights
# they are chosen uniformly at random from [-1, 1].
initial_weights = 0.1*(2*algorithm_globals.random.random(qnn3.num_weights) - 1)
model3 = TorchConnector(qnn3, initial_weights)
```
For a reminder on optimizer and loss function choices, you can go back to [this section](#Optimizer).
```
# Define optimizer and loss function
optimizer = LBFGS(model3.parameters())
f_loss = MSELoss(reduction='sum')
# Start training
model3.train() # set model to training mode
# Define objective function
def closure():
optimizer.zero_grad(set_to_none=True) # Initialize gradient
loss = f_loss(model3(Tensor(X)), Tensor(y)) # Compute batch loss
loss.backward() # Backward pass
print(loss.item()) # Print loss
return loss
# Run optimizer
optimizer.step(closure)
# Plot target function
plt.plot(np.linspace(lb, ub), f(np.linspace(lb, ub)), 'r--')
# Plot data
plt.plot(X, y, 'bo')
# Plot fitted line
y_ = []
for x in np.linspace(lb, ub):
output = model3(Tensor([x]))
y_ += [output.detach().numpy()[0]]
plt.plot(np.linspace(lb, ub), y_, 'g-')
plt.show()
```
***
## Part 2: MNIST Classification, Hybrid QNNs
In this second part, we show how to leverage a hybrid quantum-classical neural network using `TorchConnector`, to perform a more complex image classification task on the MNIST handwritten digits dataset.
For a more detailed (pre-`TorchConnector`) explanation on hybrid quantum-classical neural networks, you can check out the corresponding section in the [Qiskit Textbook](https://qiskit.org/textbook/ch-machine-learning/machine-learning-qiskit-pytorch.html).
```
# Additional torch-related imports
from torch import cat, no_grad, manual_seed
from torch.utils.data import DataLoader
from torchvision import datasets, transforms
import torch.optim as optim
from torch.nn import (Module, Conv2d, Linear, Dropout2d, NLLLoss,
MaxPool2d, Flatten, Sequential, ReLU)
import torch.nn.functional as F
```
### Step 1: Defining Data-loaders for train and test
We take advantage of the `torchvision` [API](https://pytorch.org/vision/stable/datasets.html) to directly load a subset of the [MNIST dataset](https://en.wikipedia.org/wiki/MNIST_database) and define torch `DataLoader`s ([link](https://pytorch.org/docs/stable/data.html)) for train and test.
```
# Train Dataset
# -------------
# Set train shuffle seed (for reproducibility)
manual_seed(42)
batch_size = 1
n_samples = 100 # We will concentrate on the first 100 samples
# Use pre-defined torchvision function to load MNIST train data
X_train = datasets.MNIST(root='./data', train=True, download=True,
transform=transforms.Compose([transforms.ToTensor()]))
# Filter out labels (originally 0-9), leaving only labels 0 and 1
idx = np.append(np.where(X_train.targets == 0)[0][:n_samples],
np.where(X_train.targets == 1)[0][:n_samples])
X_train.data = X_train.data[idx]
X_train.targets = X_train.targets[idx]
# Define torch dataloader with filtered data
train_loader = DataLoader(X_train, batch_size=batch_size, shuffle=True)
```
If we perform a quick visualization we can see that the train dataset consists of images of handwritten 0s and 1s.
```
n_samples_show = 6
data_iter = iter(train_loader)
fig, axes = plt.subplots(nrows=1, ncols=n_samples_show, figsize=(10, 3))
while n_samples_show > 0:
images, targets = data_iter.__next__()
axes[n_samples_show - 1].imshow(images[0, 0].numpy().squeeze(), cmap='gray')
axes[n_samples_show - 1].set_xticks([])
axes[n_samples_show - 1].set_yticks([])
axes[n_samples_show - 1].set_title("Labeled: {}".format(targets[0].item()))
n_samples_show -= 1
# Test Dataset
# -------------
# Set test shuffle seed (for reproducibility)
# manual_seed(5)
n_samples = 50
# Use pre-defined torchvision function to load MNIST test data
X_test = datasets.MNIST(root='./data', train=False, download=True,
transform=transforms.Compose([transforms.ToTensor()]))
# Filter out labels (originally 0-9), leaving only labels 0 and 1
idx = np.append(np.where(X_test.targets == 0)[0][:n_samples],
np.where(X_test.targets == 1)[0][:n_samples])
X_test.data = X_test.data[idx]
X_test.targets = X_test.targets[idx]
# Define torch dataloader with filtered data
test_loader = DataLoader(X_test, batch_size=batch_size, shuffle=True)
```
### Step 2: Defining the QNN and Hybrid Model
This second step shows the power of the `TorchConnector`. After defining our quantum neural network layer (in this case, a `TwoLayerQNN`), we can embed it into a layer in our torch `Module` by initializing a torch connector as `TorchConnector(qnn)`.
**⚠️ Attention:**
In order to have an adequate gradient backpropagation in hybrid models, we MUST set the initial parameter `input_gradients` to TRUE during the qnn initialization.
```
# Define QNN
feature_map = ZZFeatureMap(2)
ansatz = RealAmplitudes(2, reps=1)
# REMEMBER TO SET input_gradients=True FOR ENABLING HYBRID GRADIENT BACKPROP
qnn4 = TwoLayerQNN(2, feature_map, ansatz, input_gradients=True, exp_val=AerPauliExpectation(), quantum_instance=qi)
print(qnn4.operator)
# Define torch NN module
class Net(Module):
def __init__(self):
super().__init__()
self.conv1 = Conv2d(1, 2, kernel_size=5)
self.conv2 = Conv2d(2, 16, kernel_size=5)
self.dropout = Dropout2d()
self.fc1 = Linear(256, 64)
self.fc2 = Linear(64, 2) # 2-dimensional input to QNN
self.qnn = TorchConnector(qnn4) # Apply torch connector, weights chosen
# uniformly at random from interval [-1,1].
self.fc3 = Linear(1, 1) # 1-dimensional output from QNN
def forward(self, x):
x = F.relu(self.conv1(x))
x = F.max_pool2d(x, 2)
x = F.relu(self.conv2(x))
x = F.max_pool2d(x, 2)
x = self.dropout(x)
x = x.view(x.shape[0], -1)
x = F.relu(self.fc1(x))
x = self.fc2(x)
x = self.qnn(x) # apply QNN
x = self.fc3(x)
return cat((x, 1 - x), -1)
model4 = Net()
```
### Step 3: Training
```
# Define model, optimizer, and loss function
optimizer = optim.Adam(model4.parameters(), lr=0.001)
loss_func = NLLLoss()
# Start training
epochs = 10 # Set number of epochs
loss_list = [] # Store loss history
model4.train() # Set model to training mode
for epoch in range(epochs):
total_loss = []
for batch_idx, (data, target) in enumerate(train_loader):
optimizer.zero_grad(set_to_none=True) # Initialize gradient
output = model4(data) # Forward pass
loss = loss_func(output, target) # Calculate loss
loss.backward() # Backward pass
optimizer.step() # Optimize weights
total_loss.append(loss.item()) # Store loss
loss_list.append(sum(total_loss)/len(total_loss))
print('Training [{:.0f}%]\tLoss: {:.4f}'.format(
100. * (epoch + 1) / epochs, loss_list[-1]))
# Plot loss convergence
plt.plot(loss_list)
plt.title('Hybrid NN Training Convergence')
plt.xlabel('Training Iterations')
plt.ylabel('Neg. Log Likelihood Loss')
plt.show()
```
### Step 4: Evaluation
```
model4.eval() # set model to evaluation mode
with no_grad():
correct = 0
for batch_idx, (data, target) in enumerate(test_loader):
output = model4(data)
if len(output.shape) == 1:
output = output.reshape(1, *output.shape)
pred = output.argmax(dim=1, keepdim=True)
correct += pred.eq(target.view_as(pred)).sum().item()
loss = loss_func(output, target)
total_loss.append(loss.item())
print('Performance on test data:\n\tLoss: {:.4f}\n\tAccuracy: {:.1f}%'
.format(sum(total_loss) / len(total_loss),
correct / len(test_loader) / batch_size * 100)
)
# Plot predicted labels
n_samples_show = 6
count = 0
fig, axes = plt.subplots(nrows=1, ncols=n_samples_show, figsize=(10, 3))
model4.eval()
with no_grad():
for batch_idx, (data, target) in enumerate(test_loader):
if count == n_samples_show:
break
output = model4(data[0:1])
if len(output.shape) == 1:
output = output.reshape(1, *output.shape)
pred = output.argmax(dim=1, keepdim=True)
axes[count].imshow(data[0].numpy().squeeze(), cmap='gray')
axes[count].set_xticks([])
axes[count].set_yticks([])
axes[count].set_title('Predicted {}'.format(pred.item()))
count += 1
```
🎉🎉🎉🎉
**You are now able to experiment with your own hybrid datasets and architectures using Qiskit Machine Learning.**
**Good Luck!**
```
import qiskit.tools.jupyter
%qiskit_version_table
%qiskit_copyright
```
| github_jupyter |
# The soil production function
This lesson produced by Simon M Mudd and Fiona J Clubb. Last update (13/09/2021)
Back in the late 1800s, people (including G.K. Gilbert) were speculating about the rates at which soil was formed. This might depend on things like the number of burrowing animals, the rock type, the number of plants, and other factors.
The soil is "produced" from somewhere, and usually it is produced from some combination of conversion of rock to sediments, addition of organic matter, and deposition of dust. But we are going to focus on the conversion of rock material to sediment that can move.
Gilbert suggested that the rate soil was produced (from underlying rocks) depended on the thickness of the soil. We can make a prediction about the relationship between soil thickness and the rate soil is produced, and we call this the *soil production function*.
This function has proposed to have a few different forms, which we will explore below.
## Exponential Soil Production
In lectures we identified that the rate of weathering on a hillslope could be described as an exponential function that depends on soil depth, with weathering rates declining as soil gets deeper (Heimsath et al., 1997):
$p = W_0 e^{-\frac{h}{\gamma}}$
where $W_0$ is the soil production rate with no soil, and $\gamma$ is a length scale that determines how quickly soil production falls off with depth.
Typical values for $W_0$ are in the range 0.01-1 mm/yr [(Perron, 2017)](http://www.annualreviews.org/doi/abs/10.1146/annurev-earth-060614-105405). Note that when you're doing numerical calculations you have to be consistent with units. We will always do calculations in length units of ***metres*** (m), time units of ***years*** (y) and mass units of ***kilograms*** (kg). However we might convert to other units for the purposes of plotting sensible numbers (e.g. Weathering rates in mm/y = m/y $\times$ 1000).
Let's take a look at what this function looks like by plotting it with python:
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
h_locs = np.arange(0,2,0.1)
```
We define the soil production function:
```
def soil_prod_function(h_locs, W_0 = 0.0001, gamma = 0.4):
P = np.multiply(W_0, np.exp( - np.divide(h_locs,gamma) ) )
return P
```
Now lets plot the function and see what it looks like.
In the code below there are two soil production functions that you can compare. For example if you make `W_0` twice as much as `W_02` that means the second soil production function produces soil twice as fast as the first when there is no soil.
```
plt.rcParams['figure.figsize'] = [10, 4]
plt.clf()
# TRY CHANGING THE FOUR PARAMETER BELOW
# These two are for the first soil production function
W_0 = 0.0001
gamma = 0.4
# These two are for the second soil production function
W_02 = 0.0002
gamma2 = 0.4
# This bit calculates the functions
P = soil_prod_function(h_locs, W_0 = W_0, gamma = gamma)
P2 = soil_prod_function(h_locs, W_0 = W_02, gamma = gamma2)
# The rest of this stuff makes the figure
f, ax = plt.subplots(1, 1)
ax.plot(h_locs, P*1000,label="P1")
ax.plot(h_locs, P2*1000,label="P2")
ax.set_xlabel("Soil thickness ($m$)")
ax.set_ylabel("Soil production (mm/yr)")
plt.title("Two soil production function. Try playing with the parameters!")
ax.legend()
plt.tight_layout()
```
## The peaked soil production function
We also discussed in the lecture an alternative way in which soil may be produced: where there are very slow rates of soil production where there is bare bedrock, then soil peaks at some intermediate thickness before decreasing exponentially with increasing soil thickness. This model dates back to Gilbert (1877), and makes intuitive sense: water is needed for weathering processes as we discussed today. If there is bare bedrock, water is quickly transported through overland flow and little weathering can take place. If there is too much soil, then it's unlikely to be fully saturated down to the bedrock--soil interface.
In this section, we will make some plots of a hypothetical peaked (or humped) soil production function.
We will use the theoretical formulation from [Cox (1980)](https://onlinelibrary.wiley.com/doi/abs/10.1002/esp.3760050305) to calculate the weathering rate for a range of soil depths. This is a bit more complicated than the exponential function and has a bigger range of parameters:
\begin{equation}
W = W_0 (\alpha e^{-kh}) + (1 - \alpha)f \\
f = \Bigg(1 + c\frac{h}{h_c} - \frac{h^2}{{h_c}^2}\Bigg)
\end{equation}
You should recognise some of these parameters from the exponential equation. The first part of the equation is the exponential function multiplied by a coefficient, $\alpha$. $W$ is still the weathering rate, $W_0$ is the inital rate of soil production where there is no soil, and $h$ is soil depth. There are two new parameters: $h_c$ is a critical soil depth (m), and $c$ is an empirical constant. Anhert (1977) suggests that $c$ might vary between 1.7 - 2.3, $h_c$ might vary between 0.6 - 1.5, and $\alpha$ between 0 - 1. If $\alpha = 1$, then the relationship is simply the exponential function.
```
# first, let's clear the original figure
plt.clf()
# make a new figure
fig, ax = plt.subplots()
k = 1
# define the critical depth for soil production
h_c = 0.5 #metres
# define the initial rate of soil production
W_0 = 0.0005 #m/year
# define the constant c
c = 2 #dimensionless
# define alpha
alpha = 0.2
# calculate the weathering rate for the range of soil depths, h
f = (1 + c*(h_locs/h_c) - h_locs**2/h_c**2)
W = W_0 * (alpha * np.exp(-k*h_locs) + (1 - alpha)*f)
# plot the new result with a blue dashed line
ax.plot(h_locs,W*1000.,'--', color='blue', label = 'Peaked function')
# add a legend
plt.legend(loc = 'upper right')
# set the y limit of the humped function to 0 (so we don't get negative weathering rates), and set the axis labels
ax.set_ylim(0,)
plt.xlabel("Soil Depth (m)")
plt.ylabel("Weathering Rate (mm/y)")
plt.title("The peaked soil production function")
plt.tight_layout()
```
Optional Exercise 1
---
1. Have a play around and try to change some of the parameters in the peaked function (simply modify in the code block above). How does this affect the curve?
2. Try to make a plot with the exponential and peaked functions on the same set of axes, so you can compare them (HINT - copy the line that creates the exponential soil production function into the code block above, and then give it a different colour).
---
---
## Optional Exercise 2
<p>Create a figure from the practical today that shows the difference between the peaked and exponential soil production functions for different initial soil production rates. You should write a figure caption that annotates what your soil production plot is showing. The caption should be a paragraph of text that describes each line, and the parameters that have been used to create them, and offers a brief explanation of how the parameters used influence rates of soil production. For an indication of the level of detail required, you could look at examples of captions to figures in journal articles, such as Figure 3 in
[Heimsath et al. (2012)](https://www.nature.com/ngeo/journal/v5/n3/pdf/ngeo1380.pdf). You can use any program you like, such as Microsoft Word, to create your figure.
**Note**: the exercises from the practicals in this module will not be marked, but they are all teaching you important skills that will be used in the summative assessment. I would therefore really encourage you to engage with them. I will go over the answers and discuss the exercises at the start of the next session.
For your independent project, you will be expected to present 5 figures with captions, so this is a good chance to practice how to write a good figure caption!
| github_jupyter |
# Chapter 1: Building Abstractions with Functions
## 1.1 Get started
> All computing begins with representing information, specifying logic to process it, and designing abstractions that manage the complexity of that logic.
**Statements and Expressions**
Computer programs consist of instruction to either
1. Compute some value
2. Carry out some action
**Functions**
Functions encapsulate logic that manipulates data.
**Objects**
An object seamlessly bundles together data and the logic that manipulates that data in a way that manages the complexity of both
**Interpreters**
A program that implements such a procedure, evaluating compound expressions, is called an interpreter.
> In the end, we will find that all of these core concepts are closely related: functions are objects, objects are functions, and interpreters are instances of both. However, developing a clear understanding of each of these concepts and their role in organizing code is critical to mastering the art of programming.
> computer = powerful + stupid
Learning to interpret errors and diagnose the cause of unexpected errors is called *debugging*.
Debugging principles
1. Test incrementally
2. Isolate errors
3. Check your assumptions
4. Consult others
5. Concentrate on what happens instead of what does not happen
```
from urllib.request import urlopen
shakespeare = urlopen('http://composingprograms.com/shakespeare.txt')
words = set(shakespeare.read().decode().split())
{w for w in words if len(w) == 6 and w[::-1] in words}
```
## 1.2 Elements of Programming
> Programs must be written for people to read, and only incidentally for machines to execute.
Pay particular attention to the means that the language provides for combining simple ideas to form more complex ideas. Every powerful language has three such mechanisms.
* **primitive expressions and statements**, which represent the simplest building blocks that the language provides,
* **means of combination**, by which compound elements are built from simpler ones and
* **means of abstraction**, by which compound element can be named and manipulated as units.
In programming, we deal with two kinds of elements: functions and data. Informally, data is stuff that we want to manipulate, and functions describe the rules for manipulating the data.
Use names to refer to computational objects. If a value has been given a name, we say that the name binds to the value.
**Pure functions** Functions have some input (their arguments) and return some output (the result of applying them). Pure functions can be composed more reliably into compound call expressions and tend to be simpler to test
**Non-pure functions** In addition to returning a value, applying a non-pure function can generate side effects, which make some change to the state of the interpreter or computer.
```
print(print(1), print(2))
```
## 1.3 Defining New Functions
Function: powerful abstraction technique
**Environment**
An environment in which an expression is evaluated consists of a sequence of frames, depicted as boxes. Each frame contains bindings, each of which associates a name with its corresponding value. There is a single global frame.
**Name Evaluation**
A name evaluates to the value bound to that name in the earliest frame of the current environment in which that name is found.
**Aspects of a functional abstraction**
To master the use of a functional abstraction, it is often useful to consider its three core attributes. The domain of a function is the set of arguments it can take. The range of a function is the set of values it can return. The intent of a function is the relationship it computes between inputs and output (as well as any side effects it might generate). Understanding functional abstractions via their domain, range, and intent is critical to using them correctly in a complex program.
```
from operator import truediv, floordiv
print(truediv(3, 4))
print(floordiv(1, 4))
```
## 1.4 Designing Functions
Functions are an essential ingredient of all programs, large and small, and serve as our primary medium to express computational processes in a programming language.
1. Each function should have exactly one job
2. Don't repeat yourself is a central tenet of software engineering
3. Functions should be defined generally
**Decomposing a complex task into concise functions is a skill that takes experience to master.**
```
def func(a, b):
"""
one line briefly describe its functionality
Args:
a (int): xxx
b (str): xxx
Returns:
a bool value
"""
pass
```
## 1.5 Control
Controls are statements that control the flow of a program's execution based on the results of logical comparisons.
Rather than being evaluated, statements are executed.
Testing a function is the **act** of verifying that the function's behavior matches expectations.
```
# testing
assert abs(1) == 1
assert(abs(2) == 2)
```
## 1.6 Higher-Order Functions
> One of the things we should demand from a powerful programming language is the ability to build abstractions by assigning names to common patterns and then to work in terms of the names directly.
1. naming and functions allow us to abstract away a vast amount of complexity
2. it is only by virtue of the fact that we have an extremely general evaluation procedure for the Python language that small components can be composed into complex processes.
This discipline of sharing names among nested definitions is called **lexical scoping**. Critically, the inner functions have access to the names in the environment where they are defined (not where they are called).
We require two extensions to our environment model to enable lexical scoping.
1. Each user-defined function has a parent environment: the environment in which it was defined.
2. When a user-defined function is called, its local frame extends its parent environment.
```
def improve(update, close, guess=1):
while not close(guess):
guess = update(guess)
return guess
def golden_update(guess):
return 1/guess + 1
def square_close_to_successor(guess):
return approx_eq(guess * guess, guess + 1)
def approx_eq(x, y, tolerance=1e-15):
return abs(x - y) < tolerance
print(improve(golden_update, square_close_to_successor))
```
| github_jupyter |
# Cleaning WaDEQA data for ArcGIS Web App development, not through Excel.
# Creating 3 Tables: Sites, Allocations, BeneficialUse
#### General Approach:
1) Read in csv sheet of data. Export fields into new database.
2) Remove bad rows and elements.
3) Fix date datatype.
4) Include ID int value for beneficial use.
5) Fix and remove misisng or nan values for allocatoin flow and volume.
6) Remove duplicates based on AllocationNativeID.
7) Create API link for each siteUUID.
8) Export completed df as processed csv.
```
#Needed Libaraies
import os
import numpy as np
import pandas as pd
#Setting work directory, reading inputs, creating dataframe
workingDir = "C:/Users/rjame/Documents/WSWC Documents/Portal Creation Research"
os.chdir(workingDir)
# fileInput = "dontopen_WaDEQAQuery_SitesAll.csv"
fileInput = "dontopen_LiveWaDEQAQuery_SitesAll.csv"
df = pd.read_csv(fileInput)
df
df.dtypes
#removeing bad rows of df because they are null for some odd reason
df = df[(df.WaterSourceName != 'test')]
df = df[df['SiteUUID'].notna()]
df = df[df['AllocationPriorityDateID'].notna()]
df = df[df['BeneficialUseCV'].notna()]
df = df[df['Longitude'].notna()]
df = df[df['Latitude'].notna()]
df.reset_index()
#changing data type of 'Date' to datatype, then changing format of 'Date' to yyyy-mm-dd
df['Date'] = pd.to_datetime(df['Date'], errors = 'coerce')
df['Date'] = pd.to_datetime(df["Date"].dt.strftime('%m/%d/%Ym'))
WSWCBenUseDict = {
"Irrigation" : "Agricultural",
"Agriculture" : "Agricultural",
"Commercial" : "Commercial",
"Domestic" : "Domestic",
"72-12-1 domestic one household" : "Domestic",
"72-12-1 domestic and livestock watering" : "Domestic",
"72-12-1 multiple domestic households" : "Domestic",
"Subdivision" : "Domestic",
"Mobile home parks" : "Domestic",
"Fire" : "Fire",
"Fire Protection" : "Fire",
"Industrial" : "Industrial",
"Oil production" : "Industrial",
"Highway construction" : "Industrial",
"Cemetery" : "Industrial",
"Poultry and egg operation" : "Industrial",
"Dairy operation" : "Industrial",
"Construction" : "Industrial",
"Domestic construction" : "Industrial",
"Mining" : "Mining",
"Municipal" : "Municipal",
"Power" : "Power",
"Power Generation" : "Power",
"Recreation" : "Recreation",
"Snow Making" : "Snow Making",
"Stockwatering" : "StockWatering",
"Stock" : "StockWatering",
"Wildlife" : "Wildlife",
"Fish and game propogation" : "Wildlife",
"Fish And Wildlife" : "Wildlife"}
def assignWSWCBU(colrowValue):
if colrowValue == '' or pd.isnull(colrowValue):
return np.nan
else:
String1 = colrowValue # remove whitespace chars
try:
outList = WSWCBenUseDict[String1]
except:
outList = "Other"
return outList
df['WSWCBeneficialUse'] = df.apply(lambda row: assignWSWCBU(row['BeneficialUseCV']), axis=1)
WSWCBenUseColorDict = {
"Agricultural " : "Yellow",
"Commercial " : "DarkGreen",
"Domestic " : "Blue",
"Fire " : "Crimson",
"Industrial " : "BlueViolet",
"Mining " : "Gold",
"Municipal " : "Black",
"Power " : "DarkOrange",
"Recreation " : "Fuchsia",
"Snow Making " : "MintCream",
"StockWatering " : "DarkTurquoise",
"Wildlife " : "PaleGreen",
"Other " : "SlateGray"}
def assignWSWCBUColor(colrowValue):
if colrowValue == '' or pd.isnull(colrowValue):
return "SlateGray"
else:
String1 = colrowValue # remove whitespace chars
try:
outList = WSWCBenUseColorDict[String1]
except:
outList = "SlateGray"
return outList
df['WBenUseColor'] = df.apply(lambda row: assignWSWCBUColor(row['WSWCBeneficialUse']), axis=1)
#Removing NaN, and missing (999) values from AllocaitonAmount, AllocaitonMaxium, IrrigatedAcerage.
df['AllocationAmount'] = df['AllocationAmount'].fillna(0)
df['AllocationAmount'] = df['AllocationAmount'].replace(999, 0)
df['AllocationMaximum'] = df['AllocationMaximum'].fillna(0)
df['AllocationMaximum'] = df['AllocationMaximum'].replace(999, 0)
df['IrrigatedAcreage'] = df['IrrigatedAcreage'].fillna(0)
df['IrrigatedAcreage'] = df['IrrigatedAcreage'].replace(999, 0)
#Sort by SiteUUID and BenUse (this way Agriculture should show up first in the below list)
df = df.sort_values(by=['SiteUUID', 'WSWCBeneficialUse'], ascending=True).reset_index()
## Create Sites Dataframe - with attached Allo and Benuse info ##
########################################
dfSite = pd.DataFrame()
#The Columns
dfSite['SiteUUID'] = df['SiteUUID'].astype(str)
dfSite['Lat'] = df['Latitude'].astype(float)
dfSite['Long'] = df['Longitude'].astype(float)
dfSite['SN_ID'] = df['AllocationNativeID'].astype(str)
dfSite['State'] = df['State'].astype(str)
dfSite['AA_CFS'] = df['AllocationAmount'].astype(float)
dfSite['AA_AF'] = df['IrrigatedAcreage'].astype(float)
dfSite['AAM_AF'] = df['AllocationMaximum'].astype(float)
dfSite['PD'] = df['Date']
dfSite['WSN'] = df['WaterSourceName'].astype(str)
dfSite['WBenUse'] = df['WSWCBeneficialUse'].astype(str)
dfSite['WBenUseColor'] = df['WBenUseColor'].astype(str)
#Groupby based on SiteUUID
dfNewSite = dfSite.groupby('SiteUUID', sort=True).agg(lambda x: ','.join([str(elem) for elem in (list(set(x)))])).reset_index()
dfNewSite
#Drop Duplicates - SiteUUID
dfNewSite = dfNewSite.drop_duplicates(['SiteUUID'], keep="first").reset_index(drop=True)
dfNewSite
## Create Allocations Dataframe ##
########################################
#AllocationAmountID, SiteUUID, State, AllocationNativeID, AllocationAmount
#, IrrigatedAcreage, AllocationMaximum, Date, WaterSourceName
#dfSite.SiuteUUId -to- dfAllo.SiuteUUId
dfAllo = pd.DataFrame()
#The Columns
dfAllo['SN_ID'] = df['AllocationNativeID'].astype(str)
dfAllo['SiteUUID'] = df['SiteUUID'].astype(str)
dfAllo['State'] = df['State'].astype(str)
dfAllo['AA_CFS'] = df['AllocationAmount'].astype(float)
dfAllo['AA_AF'] = df['IrrigatedAcreage'].astype(float)
dfAllo['AAM_AF'] = df['AllocationMaximum'].astype(float)
dfAllo['PD'] = df['Date']
dfAllo['WSN'] = df['WaterSourceName'].astype(str)
#Drop Duplicates - State Allocation Native ID & SiteUUID
dfAllo = dfAllo.drop_duplicates(['SN_ID', "SiteUUID"], keep="first")
#Sort & Reset Index
dfAllo = dfAllo.sort_values('SN_ID', ascending=True).reset_index(drop=True)
dfAllo
## Create Beneficial Use Dataframe ##
########################################
#AllocationAmountID, BeneficialUseCV,
#dfAllo.WSWC AllocationAmountID -to- dfBen.WSWC AllocationAmountID
dfBen = pd.DataFrame()
#The Columns
dfBen['SN_ID'] = df['AllocationNativeID'].astype(str)
dfBen['WBenUse'] = df['WSWCBeneficialUse'].astype(str)
#Drop Duplicates - State Allocation Native ID & WSWC Identified Beneficial Use
dfBen = dfBen.drop_duplicates(['SN_ID', 'WBenUse'], keep="first")
#Sort & Reset Index
dfBen = dfBen.sort_values('SN_ID', ascending=True).reset_index(drop=True)
dfBen
#Exporting Outbound DataFrame to working csv file.
dfNewSite.to_csv('P_dfSiteWithAll.csv', index=False) # The output
dfAllo.to_csv('P_dfAllo.csv', index=False) # The output
dfBen.to_csv('P_dfBen.csv', index=False) # The output
```
# Double Check. Sites that have multiple Allo and BenUse
```
# dfchecksite = pd.DataFrame() # The output dataframe for CSV.
# dfchecksite = df.groupby('SiteUUID', sort=False).agg(lambda x: ','.join([str(elem) for elem in (list(set(x)))])).reset_index()
# # dfchecksite.to_csv('P_dfchecksite.csv', index=False) # The output
# dfchecksite.to_csv('P_Livedfchecksite.csv', index=False) # The output
```
# Old Code to hold temporary hold on to
```
## Create Sites Dataframe ##
# ########################################
# #SiteUUID, Latitude, Longitude
# dfSite = pd.DataFrame()
# #The Columns
# dfSite['SiteUUID'] = df['SiteUUID'].astype(str)
# dfSite['Lat'] = df['Latitude'].astype(float)
# dfSite['Long'] = df['Longitude'].astype(float)
# #Drop Duplicates - SiteUUID
# dfSite = dfSite.drop_duplicates(['SiteUUID'], keep="first")
# #Sort & Reset Index
# dfSite = dfSite.sort_values('SiteUUID', ascending=True).reset_index(drop=True)
# dfSite
```
| github_jupyter |
# Image classification with modern MLP models
**Author:** [Khalid Salama](https://www.linkedin.com/in/khalid-salama-24403144/)<br>
**Date created:** 2021/05/30<br>
**Last modified:** 2021/05/30<br>
**Description:** Implementing the MLP-Mixer, FNet, and gMLP models for CIFAR-100 image classification.
## Introduction
This example implements three modern attention-free, multi-layer perceptron (MLP) based models for image
classification, demonstrated on the CIFAR-100 dataset:
1. The [MLP-Mixer](https://arxiv.org/abs/2105.01601) model, by Ilya Tolstikhin et al., based on two types of MLPs.
3. The [FNet](https://arxiv.org/abs/2105.03824) model, by James Lee-Thorp et al., based on unparameterized
Fourier Transform.
2. The [gMLP](https://arxiv.org/abs/2105.08050) model, by Hanxiao Liu et al., based on MLP with gating.
The purpose of the example is not to compare between these models, as they might perform differently on
different datasets with well-tuned hyperparameters. Rather, it is to show simple implementations of their
main building blocks.
This example requires TensorFlow 2.4 or higher, as well as
[TensorFlow Addons](https://www.tensorflow.org/addons/overview),
which can be installed using the following command:
```shell
pip install -U tensorflow-addons
```
## Setup
```
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
import tensorflow_addons as tfa
```
## Prepare the data
```
num_classes = 100
input_shape = (32, 32, 3)
(x_train, y_train), (x_test, y_test) = keras.datasets.cifar100.load_data()
print(f"x_train shape: {x_train.shape} - y_train shape: {y_train.shape}")
print(f"x_test shape: {x_test.shape} - y_test shape: {y_test.shape}")
```
## Configure the hyperparameters
```
weight_decay = 0.0001
batch_size = 128
num_epochs = 50
dropout_rate = 0.2
image_size = 64 # We'll resize input images to this size.
patch_size = 8 # Size of the patches to be extracted from the input images.
num_patches = (image_size // patch_size) ** 2 # Size of the data array.
embedding_dim = 256 # Number of hidden units.
num_blocks = 4 # Number of blocks.
print(f"Image size: {image_size} X {image_size} = {image_size ** 2}")
print(f"Patch size: {patch_size} X {patch_size} = {patch_size ** 2} ")
print(f"Patches per image: {num_patches}")
print(f"Elements per patch (3 channels): {(patch_size ** 2) * 3}")
```
## Build a classification model
We implement a method that builds a classifier given the processing blocks.
```
def build_classifier(blocks, positional_encoding=False):
inputs = layers.Input(shape=input_shape)
# Augment data.
augmented = data_augmentation(inputs)
# Create patches.
patches = Patches(patch_size, num_patches)(augmented)
# Encode patches to generate a [batch_size, num_patches, embedding_dim] tensor.
x = layers.Dense(units=embedding_dim)(patches)
if positional_encoding:
positions = tf.range(start=0, limit=num_patches, delta=1)
position_embedding = layers.Embedding(
input_dim=num_patches, output_dim=embedding_dim
)(positions)
x = x + position_embedding
# Process x using the module blocks.
x = blocks(x)
# Apply global average pooling to generate a [batch_size, embedding_dim] representation tensor.
representation = layers.GlobalAveragePooling1D()(x)
# Apply dropout.
representation = layers.Dropout(rate=dropout_rate)(representation)
# Compute logits outputs.
logits = layers.Dense(num_classes)(representation)
# Create the Keras model.
return keras.Model(inputs=inputs, outputs=logits)
```
## Define an experiment
We implement a utility function to compile, train, and evaluate a given model.
```
def run_experiment(model):
# Create Adam optimizer with weight decay.
optimizer = tfa.optimizers.AdamW(
learning_rate=learning_rate, weight_decay=weight_decay,
)
# Compile the model.
model.compile(
optimizer=optimizer,
loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=[
keras.metrics.SparseCategoricalAccuracy(name="acc"),
keras.metrics.SparseTopKCategoricalAccuracy(5, name="top5-acc"),
],
)
# Create a learning rate scheduler callback.
reduce_lr = keras.callbacks.ReduceLROnPlateau(
monitor="val_loss", factor=0.5, patience=5
)
# Create an early stopping callback.
early_stopping = tf.keras.callbacks.EarlyStopping(
monitor="val_loss", patience=10, restore_best_weights=True
)
# Fit the model.
history = model.fit(
x=x_train,
y=y_train,
batch_size=batch_size,
epochs=num_epochs,
validation_split=0.1,
callbacks=[early_stopping, reduce_lr],
)
_, accuracy, top_5_accuracy = model.evaluate(x_test, y_test)
print(f"Test accuracy: {round(accuracy * 100, 2)}%")
print(f"Test top 5 accuracy: {round(top_5_accuracy * 100, 2)}%")
# Return history to plot learning curves.
return history
```
## Use data augmentation
```
data_augmentation = keras.Sequential(
[
layers.experimental.preprocessing.Normalization(),
layers.experimental.preprocessing.Resizing(image_size, image_size),
layers.experimental.preprocessing.RandomFlip("horizontal"),
layers.experimental.preprocessing.RandomZoom(
height_factor=0.2, width_factor=0.2
),
],
name="data_augmentation",
)
# Compute the mean and the variance of the training data for normalization.
data_augmentation.layers[0].adapt(x_train)
```
## Implement patch extraction as a layer
```
class Patches(layers.Layer):
def __init__(self, patch_size, num_patches):
super(Patches, self).__init__()
self.patch_size = patch_size
self.num_patches = num_patches
def call(self, images):
batch_size = tf.shape(images)[0]
patches = tf.image.extract_patches(
images=images,
sizes=[1, self.patch_size, self.patch_size, 1],
strides=[1, self.patch_size, self.patch_size, 1],
rates=[1, 1, 1, 1],
padding="VALID",
)
patch_dims = patches.shape[-1]
patches = tf.reshape(patches, [batch_size, self.num_patches, patch_dims])
return patches
```
## The MLP-Mixer model
The MLP-Mixer is an architecture based exclusively on
multi-layer perceptrons (MLPs), that contains two types of MLP layers:
1. One applied independently to image patches, which mixes the per-location features.
2. The other applied across patches (along channels), which mixes spatial information.
This is similar to a [depthwise separable convolution based model](https://arxiv.org/pdf/1610.02357.pdf)
such as the Xception model, but with two chained dense transforms, no max pooling, and layer normalization
instead of batch normalization.
### Implement the MLP-Mixer module
```
class MLPMixerLayer(layers.Layer):
def __init__(self, num_patches, hidden_units, dropout_rate, *args, **kwargs):
super(MLPMixerLayer, self).__init__(*args, **kwargs)
self.mlp1 = keras.Sequential(
[
layers.Dense(units=num_patches),
tfa.layers.GELU(),
layers.Dense(units=num_patches),
layers.Dropout(rate=dropout_rate),
]
)
self.mlp2 = keras.Sequential(
[
layers.Dense(units=num_patches),
tfa.layers.GELU(),
layers.Dense(units=embedding_dim),
layers.Dropout(rate=dropout_rate),
]
)
self.normalize = layers.LayerNormalization(epsilon=1e-6)
def call(self, inputs):
# Apply layer normalization.
x = self.normalize(inputs)
# Transpose inputs from [num_batches, num_patches, hidden_units] to [num_batches, hidden_units, num_patches].
x_channels = tf.linalg.matrix_transpose(x)
# Apply mlp1 on each channel independently.
mlp1_outputs = self.mlp1(x_channels)
# Transpose mlp1_outputs from [num_batches, hidden_dim, num_patches] to [num_batches, num_patches, hidden_units].
mlp1_outputs = tf.linalg.matrix_transpose(mlp1_outputs)
# Add skip connection.
x = mlp1_outputs + inputs
# Apply layer normalization.
x_patches = self.normalize(x)
# Apply mlp2 on each patch independtenly.
mlp2_outputs = self.mlp2(x_patches)
# Add skip connection.
x = x + mlp2_outputs
return x
```
### Build, train, and evaluate the MLP-Mixer model
Note that training the model with the current settings on a V100 GPUs
takes around 8 seconds per epoch.
```
mlpmixer_blocks = keras.Sequential(
[MLPMixerLayer(num_patches, embedding_dim, dropout_rate) for _ in range(num_blocks)]
)
learning_rate = 0.005
mlpmixer_classifier = build_classifier(mlpmixer_blocks)
history = run_experiment(mlpmixer_classifier)
```
The MLP-Mixer model tends to have much less number of parameters compared
to convolutional and transformer-based models, which leads to less training and
serving computational cost.
As mentioned in the [MLP-Mixer](https://arxiv.org/abs/2105.01601) paper,
when pre-trained on large datasets, or with modern regularization schemes,
the MLP-Mixer attains competitive scores to state-of-the-art models.
You can obtain better results by increasing the embedding dimensions,
increasing, increasing the number of mixer blocks, and training the model for longer.
You may also try to increase the size of the input images and use different patch sizes.
## The FNet model
The FNet uses a similar block to the Transformer block. However, FNet replaces the self-attention layer
in the Transformer block with a parameter-free 2D Fourier transformation layer:
1. One 1D Fourier Transform is applied along the patches.
2. One 1D Fourier Transform is applied along the channels.
### Implement the FNet module
```
class FNetLayer(layers.Layer):
def __init__(self, num_patches, embedding_dim, dropout_rate, *args, **kwargs):
super(FNetLayer, self).__init__(*args, **kwargs)
self.ffn = keras.Sequential(
[
layers.Dense(units=embedding_dim),
tfa.layers.GELU(),
layers.Dropout(rate=dropout_rate),
layers.Dense(units=embedding_dim),
]
)
self.normalize1 = layers.LayerNormalization(epsilon=1e-6)
self.normalize2 = layers.LayerNormalization(epsilon=1e-6)
def call(self, inputs):
# Apply fourier transformations.
x = tf.cast(
tf.signal.fft2d(tf.cast(inputs, dtype=tf.dtypes.complex64)),
dtype=tf.dtypes.float32,
)
# Add skip connection.
x = x + inputs
# Apply layer normalization.
x = self.normalize1(x)
# Apply Feedfowrad network.
x_ffn = self.ffn(x)
# Add skip connection.
x = x + x_ffn
# Apply layer normalization.
return self.normalize2(x)
```
### Build, train, and evaluate the FNet model
Note that training the model with the current settings on a V100 GPUs
takes around 8 seconds per epoch.
```
fnet_blocks = keras.Sequential(
[FNetLayer(num_patches, embedding_dim, dropout_rate) for _ in range(num_blocks)]
)
learning_rate = 0.001
fnet_classifier = build_classifier(fnet_blocks, positional_encoding=True)
history = run_experiment(fnet_classifier)
```
As shown in the [FNet](https://arxiv.org/abs/2105.03824) paper,
better results can be achieved by increasing the embedding dimensions,
increasing, increasing the number of FNet blocks, and training the model for longer.
You may also try to increase the size of the input images and use different patch sizes.
The FNet scales very efficiently to long inputs, runs much faster faster than attention-based
Transformer models, and produces competitive accuracy results.
## The gMLP model
The gMLP is a MLP architecture that features a Spatial Gating Unit (SGU).
The SGU enables cross-patch interactions across the spatial (channel) dimension, by:
1. Transforming the input spatially by applying linear projection across patches (along channels).
2. Applying element-wise multiplication of the input and its spatial transformation.
### Implement the gMLP module
```
class gMLPLayer(layers.Layer):
def __init__(self, num_patches, embedding_dim, dropout_rate, *args, **kwargs):
super(gMLPLayer, self).__init__(*args, **kwargs)
self.channel_projection1 = keras.Sequential(
[
layers.Dense(units=embedding_dim * 2),
tfa.layers.GELU(),
layers.Dropout(rate=dropout_rate),
]
)
self.channel_projection2 = layers.Dense(units=embedding_dim)
self.spatial_projection = layers.Dense(
units=num_patches, bias_initializer="Ones"
)
self.normalize1 = layers.LayerNormalization(epsilon=1e-6)
self.normalize2 = layers.LayerNormalization(epsilon=1e-6)
def spatial_gating_unit(self, x):
# Split x along the channel dimensions.
# Tensors u and v will in th shape of [batch_size, num_patchs, embedding_dim].
u, v = tf.split(x, num_or_size_splits=2, axis=2)
# Apply layer normalization.
v = self.normalize2(v)
# Apply spatial projection.
v_channels = tf.linalg.matrix_transpose(v)
v_projected = self.spatial_projection(v_channels)
v_projected = tf.linalg.matrix_transpose(v_projected)
# Apply element-wise multiplication.
return u * v_projected
def call(self, inputs):
# Apply layer normalization.
x = self.normalize1(inputs)
# Apply the first channel projection. x_projected shape: [batch_size, num_patches, embedding_dim * 2].
x_projected = self.channel_projection1(x)
# Apply the spatial gating unit. x_spatial shape: [batch_size, num_patches, embedding_dim].
x_spatial = self.spatial_gating_unit(x_projected)
# Apply the second channel projection. x_projected shape: [batch_size, num_patches, embedding_dim].
x_projected = self.channel_projection2(x_spatial)
# Add skip connection.
return x + x_projected
```
### Build, train, and evaluate the gMLP model
Note that training the model with the current settings on a V100 GPUs
takes around 9 seconds per epoch.
```
gmlp_blocks = keras.Sequential(
[gMLPLayer(num_patches, embedding_dim, dropout_rate) for _ in range(num_blocks)]
)
learning_rate = 0.003
gmlp_classifier = build_classifier(gmlp_blocks)
history = run_experiment(gmlp_classifier)
```
As shown in the [gMLP](https://arxiv.org/abs/2105.08050) paper,
better results can be achieved by increasing the embedding dimensions,
increasing, increasing the number of gMLP blocks, and training the model for longer.
You may also try to increase the size of the input images and use different patch sizes.
Note that, the paper used advanced regularization strategies, such as MixUp and CutMix,
as well as AutoAugment.
| github_jupyter |
```
#%%appyter init
import os, sys; sys.path.insert(0, os.path.realpath('..'))
from appyter import magic
magic.init(lambda _=globals: _())
%matplotlib inline
# Imports
## Data processing
import pandas as pd
import numpy as np
import scipy as sp
## Machine Learning
import sklearn as sk
from sklearn import (
calibration,
decomposition,
ensemble,
feature_selection,
linear_model,
manifold,
metrics,
model_selection,
multioutput,
pipeline,
preprocessing,
svm,
tree,
feature_extraction,
neural_network,
)
from split import StratifiedGroupKFold, RepeatedStratifiedGroupKFold
import umap
## Plotting
from matplotlib import pyplot as plt
import seaborn as sns
import plotly.express as px
import plotly.graph_objects as go
## Drugmonizome API
from drugmonizome import Drugmonizome
## SEP-L1000 data retrieval
from sepl1000 import SEPL1000
## L1000FWD queries
import querysepl1000fwd
## Match drug name inputs using PubChem API
from DrugNameConverter import DrugNameConverter
# Utility
import os
import re
import json
from functools import reduce
from IPython.display import display, HTML
from tqdm import tqdm
# Interactive tables
from itables import show
# Plotly fix
import plotly.io as pio
pio.renderers.default = 'notebook'
rng = 2020
np.random.seed(rng)
# Notebook display util functions (adapted from Nicole Moiseyev's Patient Cohorts RNA-Seq Viewer appyter)
def make_clickable(link):
return f'<a target="_blank" href="{link}">{link}</a>'
table_number = 0
figure_number = 0
def figure_header(label,title):
global table_number
global figure_number
if label == 'Table':
table_number += 1
label = f'Table {table_number}'
elif label == 'Figure':
figure_number += 1
label = f'Figure {figure_number}'
display(HTML(f"<div style='font-size:2rem; padding:1rem 0;'><b>{label}</b>: {title}</div>"))
def figure_legend(label,title,content=''):
global table_number
global figure_number
if label == 'Table':
label = f'Table {table_number}'
elif label == 'Figure':
label = f'Figure {figure_number}'
display(HTML(f'<style>div.caption {{text-align: center;}}</style><div class=caption><b>{label}</b>: <i>{title}</i>. {content} </div>'))
```
## Select Input Datasets and Target Classes
Selected drug set libraries and phenotypic datasets are downloaded and joined on the compound InChI Key to produce a large input feature matrix. A machine learning model will be trained to predict the specified target labels from these features. This is a binary classification task that can be used to predict compounds that are likely to be associated with the target class.
```
%%appyter hide
{% do SectionField(
title='Input Dataset Selection',
subtitle='Select the input datasets to use for learning and classification. \
A model will be trained to predict the target labels from the selected features. \
If no datasets are selected, default features will be used.',
name='ATTRIBUTES',
img='attributes.png',
) %}
{% set sepl1000datasets = MultiCheckboxField(
name='sepl1000datasets',
label='Transcriptomic and Imaging Datasets after Perturbation<br>(from the SEP-L1000 project)',
description='These input datasets were used previously for side effect prediction (https://maayanlab.net/SEP-L1000/).',
choices=[
'LINCS Gene Expression Signatures',
'GO Transformed Signatures (PAEA)',
'MLPCN Cell Morphological Profiling',
'MACCS Chemical Fingerprint',
],
default=[],
section='ATTRIBUTES'
) %}
{% set exprdatasets = MultiCheckboxField(
name='exprdatasets',
label='L1000FWD<br>(drug set libraries from Drugmonizome)',
description='Top up and down-regulated genes after perturbation, along with enriched pathways.',
choices=[
'L1000FWD Downregulated GO Biological Processes',
'L1000FWD Downregulated GO Cellular Components',
'L1000FWD Downregulated GO Molecular Function',
'L1000FWD Downregulated KEGG Pathways',
'L1000FWD Downregulated Signatures',
'L1000FWD Predicted Side Effects',
'L1000FWD Upregulated GO Biological Process',
'L1000FWD Upregulated GO Cellular Components',
'L1000FWD Upregulated GO Molecular Function',
'L1000FWD Upregulated KEGG Pathways',
'L1000FWD Upregulated Signatures',
],
default=[],
section='ATTRIBUTES'
) %}
{% set targetdatasets = MultiCheckboxField(
name='targetdatasets',
label='Drug Targets and Associated Genes<br>(drug set libraries from Drugmonizome)',
choices=[
'Downregulated CREEDS Signatures',
'Upregulated CREEDS Signatures',
'DrugCentral Targets',
'DrugRepurposingHub Drug Targets',
'Drugbank Small Molecule Carriers',
'Drugbank Small Molecule Enzymes',
'Drugbank Small Molecule Targets',
'Drugbank Small Molecule Transporters',
'Geneshot Associated Genes',
'Geneshot Predicted AutoRIF Genes',
'Geneshot Predicted Coexpression Genes',
'Geneshot Predicted Enrichr Genes',
'Geneshot Predicted GeneRIF Genes',
'Geneshot Predicted Tagger Genes',
'KinomeScan Kinases',
'PharmGKB Single Nucleotide Polymorphisms',
'STITCH Targets',
],
default=[],
section='ATTRIBUTES'
) %}
{% set indicationdatasets = MultiCheckboxField(
name='indicationdatasets',
label='Indications, Modes of Action, and Side Effects<br>(drug set libraries from Drugmonizome)',
choices=[
'ATC Codes Drugsetlibrary',
'DrugRepurposingHub Mechanisms of Action',
'PharmGKB OFFSIDES Side Effects',
'SIDER Indications',
'SIDER Side Effects',
],
default=[],
section='ATTRIBUTES'
) %}
{% set structuraldatasets = MultiCheckboxField(
name='structuraldatasets',
label='Structural Features<br>(drug set libraries from Drugmonizome)',
choices=[
'RDKIT MACCS Chemical Fingerprints'
],
default=[],
section='ATTRIBUTES'
) %}
{% set keepmissing = BoolField(
name='keepmissing',
label='Keep drugs with missing data when joining datasets',
description='Keep drugs that appear in some datasets and not in others. \
Missing data is filled in with zeros. Otherwise, only drugs \
that are present in all datasets are preserved.',
default=False,
section='ATTRIBUTES',
) %}
{% set tfidf = BoolField(
name='tfidf',
label='Apply tf–idf normalization to binary inputs',
description='For binary drug-attribute associations in the input matrix, \
apply tf-idf transformation to normalize data.',
default=True,
section='ATTRIBUTES',
) %}
{% set attribute_datasets = exprdatasets.value +
targetdatasets.value +
indicationdatasets.value +
structuraldatasets.value %}
%%appyter markdown
To construct the input matrix, we download drug set libraries and phenotypic datasets and join them on the InChI Key.
{% if keepmissing.value %} Drugs that appear in some datasets and not in others are retained, and missing data is filled in with zeros.
{% else %} Only drugs that are present in all datasets are retained.
{% endif %}
%%appyter hide
{% do SectionField(
title='Target Label Selection',
subtitle='Upload a list of compounds or select an attribute from Drugmonizome to be assigned a positive class label for binary classification.',
name='TARGET',
img='target.png',
) %}
{% set target_field = TabField(
name='target_field',
label='Target Selection',
default='Attribute',
description='Select input method',
choices={
'List': [
ChoiceField(
name='drugformat',
label='Drug Identifier Format',
description='Compounds can be specified by either drug name or InChI Key.',
default='InChI Key',
choices=[
'Drug Name',
'InChI Key'
],
section='TARGET'
),
FileField(
name='drughitlist',
label='Upload List of Compounds',
description='Upload a list of compounds to be assigned positive class labels for binary classification. \
Compounds should be in a text file, specified by either drug name or InChI Key and separated by newlines.',
default='COVID19ScreenHitsInChIKeys.txt',
examples={
'COVID19ScreenHits.txt': 'https://appyters.maayanlab.cloud/storage/Drugmonizome_ML/COVID19ScreenHits.txt',
'COVID19ScreenHitsInChIKeys.txt': 'https://appyters.maayanlab.cloud/storage/Drugmonizome_ML/COVID19ScreenHitsInChIKeys.txt',
},
section='TARGET'
),
],
'Attribute': [
AutocompleteField(
name='target_attribute',
description='Enter a small molecule attribute from one of the Drugmonizome datasets that should be predicted.',
file_path="https://appyters.maayanlab.cloud/storage/Drugmonizome_ML/drugmonizome_terms.json",
label='Attribute',
hint='Enter Drugmonizome term...',
default='cyclooxygenase inhibitor (from DrugRepurposingHub Mechanisms of Action)',
constraint='(^(.+) \\(from (.+)\\)$|^$)',
)],
},
section='TARGET',
) %}
{% set includestereo = BoolField(
name='includestereo',
label='Include stereoisomers',
description='If true, compounds are matched to entries in the datasets by the first 14 characters of their InChI Keys, \
so stereoisomers of the compounds in the input list or with a particular attritube are also counted as hits. \
Note that different resources record different details for charge and stereochemistry, \
causing some compounds to have different full-length InChI Keys in different datasets. \
Selecting this option may allow such drugs to be better matched to entries in the datasets.',
default=True,
section='TARGET',
) %}
{% set target_name, target_dataset = '', '' %}
{% if target_field.raw_value == 'Attribute' %}
{% set target_name, target_dataset = target_field.value[0].value|re_match('^(.+) \\(from (.+)\\)$') %}
{% endif %}
%%appyter code_exec
{% if sepl1000datasets.value == [] and attribute_datasets == [] %}
# No datasets selected, so use default dataset
sepl1000datasets = ['LINCS Gene Expression Signatures']
{% else %}
# Use the selected SEP-L1000 datasets
sepl1000datasets = {{ sepl1000datasets }}
{% endif %}
{% if sepl1000datasets.value != [] or attribute_datasets == [] %}
name_to_file = {
'LINCS Gene Expression Signatures': 'LINCS_Gene_Experssion_signatures_CD.csv.gz',
'GO Transformed Signatures (PAEA)': 'GO_transformed_signatures_PAEA.csv.gz',
'MLPCN Cell Morphological Profiling': 'MLPCN_morplological_profiles.csv.gz',
'MACCS Chemical Fingerprint': 'MACCS_bitmatrix.csv.gz',
}
df_sepl1000_list = list(SEPL1000.download_df(list(name_to_file[dataset] for dataset in sepl1000datasets),
index_col=0))
dataset_sizes = list(zip(sepl1000datasets, [dataset.shape[1] for dataset in df_sepl1000_list]))
# Assemble all SEP-L1000 datasets
if len(df_sepl1000_list) > 1:
# Obtain merged dataframe with omics and target data
df_sepl1000 = reduce(
lambda a, b: pd.merge( # Merge two dataframes item by item
a, # left
b, # right
# Items with the same left and right index are merged
left_index=True,
right_index=True,
{% if keepmissing.value %}
how='outer', # Keep mis-matched indices
{% else %}
how='inner', # Keep only matched indices
{% endif %}
),
df_sepl1000_list,
)
else:
df_sepl1000 = df_sepl1000_list[0]
del(df_sepl1000_list)
# Mean-fill infinite and missing values
df_sepl1000 = df_sepl1000.replace([np.inf, -np.inf], np.nan)
df_sepl1000 = df_sepl1000.fillna(np.mean(df_sepl1000))
{% else %}
dataset_sizes = []
{% endif %}
%%appyter code_exec
# Use the selected attribute datasets
attribute_datasets = {{ attribute_datasets }}
{% if attribute_datasets == [] %}
X = df_sepl1000
{% else %}
df_attributes = list(Drugmonizome.download_df(
[dataset
for dataset in attribute_datasets]
))
dataset_sizes += list(zip(attribute_datasets, [dataset.shape[1] for dataset in df_attributes]))
# Assemble all attribute datasets
if len(df_attributes) > 1:
# Obtain merged dataframe with omics and target data
df = reduce(
lambda a, b: pd.merge( # Merge two dataframes item by item
a, # left
b, # right
# Items with the same left and right index are merged
left_index=True,
right_index=True,
{% if keepmissing.value %}
how='outer', # Keep mis-matched indices
{% else %}
how='inner', # Keep only matched indices
{% endif %}
),
df_attributes,
)
else:
df = df_attributes[0]
del(df_attributes)
df = df.fillna(0)
X = df.applymap(lambda f: 1 if f!=0 else 0)
{% if tfidf.value %}
# Apply tf-idf normalization
transformer = feature_extraction.text.TfidfTransformer()
X_tfidf = transformer.fit_transform(X).toarray()
X = pd.DataFrame(X_tfidf, columns=X.columns, index=X.index)
{% if sepl1000datasets.value != [] %}
{% if keepmissing.value %}
X = pd.merge(df_sepl1000, X, left_index=True, right_index=True, how='outer') # Keep mis-matched indices
{% else %}
X = pd.merge(df_sepl1000, X, left_index=True, right_index=True) # Keep only matched indices
{% endif %}
{% endif %}
{% endif %}
{% endif %}
# View input data
figure_header('Table', 'Input data')
display(X.head())
figure_legend('Table', 'Input data',
f'The input data contain {X.shape[0]} compounds and {X.shape[1]} features per compound, \
taken from the following datasets: {", ".join(sepl1000datasets + attribute_datasets)}.')
%%appyter markdown
{% if target_field.raw_value == 'List' %}
The target labels are produced from the uploaded list of hits: 1 if the drug is specified as a hit, 0 otherwise.
{% if target_field.value[0].value == 'Drug Name' %} Drug names are matched to InChI Keys from PubChem, L1000FWD, and the Drugmonizome metadata.
{% endif %}
{% endif %}
%%appyter code_exec
{% if target_field.raw_value == 'List' %}
{% if target_field.value[1].value == '' %}
# Using default list of hits from COVID-19 in vitro drug screens
hits_filename = '../../COVID19ScreenHits.txt'
{% else %}
# Using user-specified list of positive drug hits
hits_filename = {{target_field.value[1]}}
{% endif %}
{% if target_field.value[0].value == 'InChI Key' %}
# Read InChI Keys from file
with open(hits_filename, 'r') as hits_file:
drug_hits = set(drug.strip().upper() for drug in hits_file.read().strip().split('\n')
if len(drug.strip()) > 0)
{% elif target_field.value[0].value == 'Drug Name' %}
# Helper functions
def merge(A, B, f):
"""
Merges two dictionaries, where items from shared keys are merged using a custom function.
"""
merged = {k: A.get(k, B.get(k)) for k in A.keys() ^ B.keys()}
merged.update({k: f(A[k], B[k]) for k in A.keys() & B.keys()})
return merged
def save_items(out_file, items):
"""
Saves list of items as rows in a file.
"""
with open(out_file, 'w') as f:
for i in range(len(items)):
if i < len(items) - 1:
f.write(items[i] + '\n')
else:
f.write(items[i])
def save_gmt(out_file, keys_to_sets, sep='\t'):
"""
Saves dict with key-set pairs as gmt file format.
"""
lines = []
for key in sorted(keys_to_sets):
lines.append(key + sep*2 + sep.join(sorted(keys_to_sets[key])))
save_items(out_file, lines)
# Read drug names from file
with open(hits_filename, 'r') as hits_file:
drug_hits = set(drug.strip().lower() for drug in hits_file.read().strip().split('\n')
if len(drug.strip()) > 0)
# Query PubChem API to map drug names to InChI Keys
print('Querying PubChem API...')
drug_hits_inchi_pubchem = DrugNameConverter.batch_to_inchi_keys(drug_hits)
# Query Drugmonizome API to map drug names to InChI Keys
print('Querying Drugmonizome API...')
drug_hits_inchi_drugmonizome = Drugmonizome.map_names_to_inchi_keys(drug_hits)
# Query L1000FWD API to map drug names to InChI Keys
print('Querying L1000FWD API...')
drug_hits_inchi_l1000fwd = querysepl1000fwd.map_names_to_inchi_keys(drug_hits)
# Combine InChI Keys from all resources
drug_hits_inchi = merge(drug_hits_inchi_pubchem, drug_hits_inchi_drugmonizome, lambda s1, s2: s1 | s2)
drug_hits_inchi = merge(drug_hits_inchi, drug_hits_inchi_l1000fwd, lambda s1, s2: s1 | s2)
save_gmt('hits_drug_name_to_inchi_keys.gmt', drug_hits_inchi)
# Unmatched drug names
unmatched_drugs = set(drug for drug in drug_hits
if drug not in drug_hits_inchi or len(drug_hits_inchi[drug]) == 0)
print(f'Drugs without InChI Keys ({ len(unmatched_drugs) }/{ len(drug_hits) }):', unmatched_drugs)
# Set of InChI Keys for user-specified hits
drug_hits = set(key for drug in drug_hits_inchi
for key in drug_hits_inchi[drug])
save_items('hits_inchi_keys.txt', sorted(drug_hits))
{% endif %}
{% else %}
df_target = list(Drugmonizome.download_df(
['{{ target_dataset }}']
))
df = df_target[0]
df = df.fillna(0)
Y = df.applymap(lambda f: 1 if f!=0 else 0)
drug_hits = set(Y[Y['{{ target_name }}'] == 1].index)
# Helper function
def save_items(out_file, items):
"""
Saves list of items as rows in a file.
"""
with open(out_file, 'w') as f:
for i in range(len(items)):
if i < len(items) - 1:
f.write(items[i] + '\n')
else:
f.write(items[i])
save_items('hits_inchi_keys.txt', sorted(drug_hits))
{% endif %}
%%appyter markdown
{% if target_field.raw_value == 'List' %}
{% if target_field.value[0].value == 'Drug Name' %}
For the user-inputted drug names:
* Mapping of drug name to InChI Key: [hits_drug_name_to_inchi_keys.gmt](./hits_drug_name_to_inchi_keys.gmt)
* List of InChI Keys: [hits_inchi_keys.txt](./hits_inchi_keys.txt)
{% endif %}
{% endif %}
%%appyter markdown
{% if target_field.raw_value == 'List' %}
We produce a target array containing 1 if the compound is specified as a hit and 0 otherwise.
{% else %}
We produce a target array containing 1 if the compound is associated with the attribute _{{ target_name }}_ in the Drugmonizome resource _{{ target_dataset }}_ and 0 otherwise.
{% endif %}
%%appyter code_exec
{% if includestereo.value %}
# Match first 14 characters of InChI Keys (hash of InChI connectivity information)
drug_hits_inchi_main_layer = set(key[:14] for key in drug_hits)
y = np.array([drug[:14] in drug_hits_inchi_main_layer for drug in X.index]).astype(np.int8)
{% else %}
# Match full InChI Keys
y = np.array([drug in drug_hits for drug in X.index]).astype(np.int8)
{% endif %}
print('Number of hits matched in input: %d (%0.3f %%)' % (y.sum(), 100*y.sum()/len(y)))
# Output data shapes
print('Input shape:', X.shape)
print('Target shape:', y.shape)
```
## Dimensionality Reduction and Visualization
```
%%appyter hide
{% do SectionField(
title='Machine Learning Pipeline',
subtitle='Select from available machine learning algorithms, their unique settings, and methods to use to evaluate the classifier.',
name='SETTINGS',
img='settings.png',
) %}
{% set visualization_reduction = ChoiceField(
name='visualization_reduction',
label='Data Visualization Method',
description='Select a dimensionality reduction algorithm for data visualization.',
default='UMAP',
choices={
'UMAP': 'umap.UMAP(low_memory=True, random_state=rng)',
'NMF': 'sk.decomposition.NMF(n_components=2)',
'PCA': 'sk.decomposition.PCA(n_components=2)',
'TruncatedSVD': 'sk.decomposition.TruncatedSVD(n_components=2)',
'IncrementalPCA': 'sk.decomposition.IncrementalPCA(n_components=2)',
'ICA': 'sk.decomposition.FastICA(n_components=2)',
'SparsePCA': 'sk.decomposition.SparsePCA(n_components=2)',
},
section='SETTINGS'
) %}
%%appyter markdown
We reduce the dimensionality of our omics feature space for visualization with {{ visualization_reduction.raw_value }}.
%%appyter code_exec
clf_dimensionality_reduction = {{ visualization_reduction }}
X_reduced = clf_dimensionality_reduction.fit_transform(X.values)
{% if visualization_reduction.raw_value == 'PCA' %}
print('Explained variance:', np.sum(clf_dimensionality_reduction.explained_variance_))
{% endif %}
X_reduced_df = pd.DataFrame(X_reduced, columns=['Component 1', 'Component 2'])
X_reduced_df['Drug Name'] = querysepl1000fwd.get_drug_names(X.index)
X_reduced_df['InChI Key'] = X.index
X_reduced_df['Label'] = y
X_reduced_df['marker symbol'] = ['x' if label else 'circle' for label in X_reduced_df['Label']]
X_reduced_df['text'] = ['<br>'.join(['Drug Name: ' + str(name),
'InChI Key: ' + str(inchi),
'Label: ' + str(label)])
for name, inchi, label in zip(X_reduced_df['Drug Name'],
X_reduced_df['InChI Key'],
X_reduced_df['Label'])]
%%appyter code_exec
fig = go.Figure()
for label in set(X_reduced_df['Label']):
X_plot = X_reduced_df[X_reduced_df['Label'] == label].sort_values('Label')
fig.add_trace(go.Scatter(mode='markers',
x=X_plot['Component 1'], y=X_plot['Component 2'],
text=X_plot['text'],
name=label,
marker=dict(
color=['#0d0887', '#f0f921'][label%2],
size=8,
symbol=X_plot['marker symbol'],
line_width=1,
line_color='white'
)))
fig.update_layout(height=600, width=800,
xaxis_title='Component 1',
yaxis_title='Component 2',
title_text='Known Labels ({{ visualization_reduction.raw_value }})',
legend_title_text='Target Label',
template='simple_white')
figure_header('Figure', 'Input feature space with {{ visualization_reduction.raw_value }} dimensionality reduction')
fig.show()
figure_legend('Figure', 'Input feature space with {{ visualization_reduction.raw_value }} dimensionality reduction',
f'Each point represents one of {X.shape[0]} compounds, with {X.shape[1]} features per compound, \
taken from the following datasets: {", ".join(sepl1000datasets + attribute_datasets)}. \
Compounds with known positive labels are marked by X\'s.')
```
## Machine Learning
We train and evaluate a machine learning model across multiple cross-validation splits by randomly dividing the input dataset into training and validation sets. For each round of cross-validation, a model is trained on the training set and is then used to make predictions for the compounds in the validation set. Each compound appears in at least one validation set, so the validation set predictions are used to assess model performance based on existing labels and to suggest novel predictions.
```
%%appyter hide
{% set dimensionality_reduction = ChoiceField(
name='dimensionality_reduction',
label='Dimensionality Reduction Algorithm',
description='Optionally select a dimensionality reduction algorithm as a data preprocessing step in the ML pipeline.',
default='None',
choices={
'None': 'None',
'PCA': 'sk.decomposition.PCA(n_components=64)',
'TruncatedSVD': 'sk.decomposition.TruncatedSVD(n_components=64)',
'IncrementalPCA': 'sk.decomposition.IncrementalPCA(n_components=64)',
'ICA': 'sk.decomposition.FastICA(n_components=64)',
'SparsePCA': 'sk.decomposition.SparsePCA(n_components=64)',
},
section='SETTINGS'
) %}
{% set feature_selection = ChoiceField(
name='feature_selection',
label='Machine Learning Feature Selection',
description='Optionally select a feature selection algorithm to include in the ML pipeline. \
If RecursiveSelectionFromExtraTrees is chosen, additional information can be obtained \
on the relative importance of different features based on which features are eliminated.',
default='None',
choices={
'None': 'None',
'SelectFromLinearSVC': 'sk.feature_selection.SelectFromModel(sk.svm.LinearSVC(loss="squared_hinge", penalty="l1", dual=False, class_weight="balanced"))',
'SelectFromExtraTrees': 'sk.feature_selection.SelectFromModel(sk.ensemble.ExtraTreesClassifier(class_weight="balanced"))',
'RecursiveSelectionFromExtraTrees': 'sk.feature_selection.RFE(sk.ensemble.ExtraTreesClassifier(class_weight="balanced"), n_features_to_select=256, step=0.1)',
'SelectKBest': 'sk.feature_selection.SelectKBest("f_classif")',
'SelectKBestChi2': 'sk.feature_selection.SelectKBest("chi2")',
'SelectKBestMultiInfo': 'sk.feature_selection.SelectKBest("mutual_info_classif")',
},
section='SETTINGS'
) %}
{% set algorithm = TabField(
name='algorithm',
label='Machine Learning Algorithm',
default='ExtraTreesClassifier',
description='Select a machine learning algorithm to construct the predictive model. \
(See scikit-learn User Guide for details.)',
choices={
'GradientBoostingClassifier': [
ChoiceField(
name='loss_gb',
label='loss',
description='Loss function to be optimized.',
default="deviance",
choices=["deviance", "exponential"],
),
FloatField(
name='learning_rate_gb',
label='learning_rate',
description='Shrinks the contribution of each tree by learning_rate.',
default=0.1,
),
IntField(
name='n_estimators_gb',
label='n_estimators',
description='Number of boosting stages to perform.',
default=100,
),
FloatField(
name='subsample_gb',
label='subsample',
description='Fraction of samples to be used for fitting individual base learners.',
default=1.0,
),
ChoiceField(
name='criterion_gb',
label='criterion',
description='Function to measure the quality of a split.',
default="friedman_mse",
choices=["friedman_mse", "mse", "mae"],
),
FloatField(
name='tol_gb',
label='tol',
description='Tolerance for early stopping.',
default=1e-4,
),
],
'RandomForestClassifier': [
IntField(
name='n_estimators_rf',
label='n_estimators',
description='Number of trees in the forest.',
default=100,
),
ChoiceField(
name='criterion_rf',
label='criterion',
description='Function to measure the quality of a split.',
default="gini",
choices=["gini", "entropy"],
),
FloatField(
name='min_samples_split_rf',
label='min_samples_split',
description='Minimum number of samples required to split an internal node. \
If int, then min_samples_split specifies the minimum number. \
If float, then min_samples_split specifies a fraction of the total number of samples.',
default=2,
),
FloatField(
name='min_samples_leaf_rf',
label='min_samples_leaf',
description='Minimum number of samples required to be at a leaf node. \
If int, then min_samples_leaf specifies the minimum number. \
If float, then min_samples_leaf specifies a fraction of the total number of samples.',
default=1,
),
ChoiceField(
name='max_features_rf',
label='max_features',
description='The number of features to consider when looking for the best split.',
default="None",
choices=["None", '"auto"', '"sqrt"', '"log2"'],
),
FloatField(
name='min_impurity_decrease_rf',
label='min_impurity_decrease',
description='A node will be split if this split induces a decrease of the impurity greater than or equal to this value.',
default=0.0,
),
ChoiceField(
name='class_weight_rf',
label='class_weight',
description='Weights associated with classes. If None, then all classes have weight one. \
The balanced mode adjusts weights inversely proportional to class frequencies in the input data. \
The balanced_subsample mode is the same as balanced except weights are computed based on the bootstrap sample for each tree.',
default='"balanced"',
choices=["None", '"balanced"', '"balanced_subsample"'],
),
FloatField(
name='ccp_alpha_rf',
label='ccp_alpha',
description='Complexity parameter used for Minimal Cost-Complexity Pruning. \
The subtree with the largest cost complexity that is smaller than ccp_alpha will be chosen. \
By default, no pruning is performed.',
default=0.0,
),
],
'AdaBoostClassifier': [
IntField(
name='max_depth_ab',
label='max_depth',
description='Maximum depth of the decision tree used as the base estimator.',
default=1,
),
IntField(
name='n_estimators_ab',
label='n_estimators',
description='Maximum number of estimators at which boosting is terminated.',
default=50,
),
FloatField(
name='learning_rate_ab',
label='learning_rate',
description='Shrinks the contribution of each classifier by learning_rate.',
default=1.0,
),
ChoiceField(
name='algorithm_ab',
label='algorithm',
description='Select the real or discrete boosting algorithm to use.',
default="SAMME.R",
choices=["SAMME", "SAMME.R"],
),
],
'ExtraTreesClassifier': [
IntField(
name='n_estimators_et',
label='n_estimators',
description='Number of trees in the forest.',
default=100,
),
ChoiceField(
name='criterion_et',
label='criterion',
description='Function to measure the quality of a split.',
default="gini",
choices=["gini", "entropy"],
),
FloatField(
name='min_samples_split_et',
label='min_samples_split',
description='Minimum number of samples required to split an internal node. \
If int, then min_samples_split specifies the minimum number. \
If float, then min_samples_split specifies a fraction of the total number of samples.',
default=2,
),
FloatField(
name='min_samples_leaf_et',
label='min_samples_leaf',
description='Minimum number of samples required to be at a leaf node. \
If int, then min_samples_leaf specifies the minimum number. \
If float, then min_samples_leaf specifies a fraction of the total number of samples.',
default=1,
),
ChoiceField(
name='max_features_et',
label='max_features',
description='The number of features to consider when looking for the best split.',
default="None",
choices=["None", '"auto"', '"sqrt"', '"log2"'],
),
FloatField(
name='min_impurity_decrease_et',
label='min_impurity_decrease',
description='A node will be split if this split induces a decrease of the impurity greater than or equal to this value.',
default=0.0,
),
ChoiceField(
name='class_weight_et',
label='class_weight',
description='Weights associated with classes. If None, then all classes have weight one. \
The balanced mode adjusts weights inversely proportional to class frequencies in the input data. \
The balanced_subsample mode is the same as balanced except weights are computed based on the bootstrap sample for each tree.',
default='"balanced"',
choices=["None", '"balanced"', '"balanced_subsample"'],
),
FloatField(
name='ccp_alpha_et',
label='ccp_alpha',
description='Complexity parameter used for Minimal Cost-Complexity Pruning. \
The subtree with the largest cost complexity that is smaller than ccp_alpha will be chosen. \
By default, no pruning is performed.',
default=0.0,
),
],
'DecisionTreeClassifier': [
ChoiceField(
name='criterion_dt',
label='criterion',
description='Function to measure the quality of a split.',
default="gini",
choices=["gini", "entropy"],
),
ChoiceField(
name='splitter_dt',
label='splitter',
description='Strategy used to choose the split at each node.',
default="best",
choices=["best", "random"],
),
FloatField(
name='min_samples_split_dt',
label='min_samples_split',
description='Minimum number of samples required to split an internal node. \
If int, then min_samples_split specifies the minimum number. \
If float, then min_samples_split specifies a fraction of the total number of samples.',
default=2,
),
FloatField(
name='min_samples_leaf_dt',
label='min_samples_leaf',
description='Minimum number of samples required to be at a leaf node. \
If int, then min_samples_leaf specifies the minimum number. \
If float, then min_samples_leaf specifies a fraction of the total number of samples.',
default=1,
),
ChoiceField(
name='max_features_dt',
label='max_features',
description='The number of features to consider when looking for the best split.',
default="None",
choices=["None", '"auto"', '"sqrt"', '"log2"'],
),
FloatField(
name='min_impurity_decrease_dt',
label='min_impurity_decrease',
description='A node will be split if this split induces a decrease of the impurity greater than or equal to this value.',
default=0.0,
),
ChoiceField(
name='class_weight_dt',
label='class_weight',
description='Weights associated with classes. If None, then all classes have weight one. \
The balanced mode adjusts weights inversely proportional to class frequencies in the input data. \
The balanced_subsample mode is the same as balanced except weights are computed based on the bootstrap sample for each tree.',
default='"balanced"',
choices=["None", '"balanced"', '"balanced_subsample"'],
),
FloatField(
name='ccp_alpha_dt',
label='ccp_alpha',
description='Complexity parameter used for Minimal Cost-Complexity Pruning. \
The subtree with the largest cost complexity that is smaller than ccp_alpha will be chosen. \
By default, no pruning is performed.',
default=0.0,
),
],
'KNeighborsClassifier': [
IntField(
name='n_neighbors_knn',
label='n_neighbors',
description='Number of neighbors to use for queries.',
default=5,
),
ChoiceField(
name='weights_knn',
label='weights',
description='Weight function used in prediction. \
If uniform, all points in each neighborhood are weighted equally. \
If distance, points are weighted by the inverse of their distance.',
default="uniform",
choices=["uniform", "distance"],
),
ChoiceField(
name='algorithm_knn',
label='algorithm',
description='Algorithm used to compute the nearest neighbors.',
default="auto",
choices=["auto", "ball_tree", "kd_tree", "brute"],
),
IntField(
name='leaf_size_knn',
label='leaf_size',
description='Leaf size passed to BallTree or KDTree.',
default=30,
),
IntField(
name='p_knn',
label='p',
description='Power parameter for the Minkowski metric.',
default=2,
),
ChoiceField(
name='metric_knn',
label='metric',
description='Distance metric to use for the tree.',
default="minkowski",
choices=["minkowski", "euclidean", "manhattan", "chebyshev"],
),
],
'RadiusNeighborsClassifier': [
FloatField(
name='radius_rn',
label='radius',
description='Range of parameter space to use for queries.',
default=1.0,
),
ChoiceField(
name='weights_rn',
label='weights',
description='Weight function used in prediction. \
If uniform, all points in each neighborhood are weighted equally. \
If distance, points are weighted by the inverse of their distance.',
default="uniform",
choices=["uniform", "distance"],
),
ChoiceField(
name='algorithm_rn',
label='algorithm',
description='Algorithm used to compute the nearest neighbors.',
default="auto",
choices=["auto", "ball_tree", "kd_tree", "brute"],
),
IntField(
name='leaf_size_rn',
label='leaf_size',
description='Leaf size passed to BallTree or KDTree.',
default=30,
),
IntField(
name='p_rn',
label='p',
description='Power parameter for the Minkowski metric.',
default=2,
),
ChoiceField(
name='metric_rn',
label='metric',
description='Distance metric to use for the tree.',
default="minkowski",
choices=["minkowski", "euclidean", "manhattan", "chebyshev"],
),
],
'MLPClassifier': [
StringField(
name='hidden_layer_sizes_mlp',
label='hidden_layer_sizes',
description='Enter a tuple, where the ith element represents the number of neurons in the ith hidden layer.',
hint='Enter a tuple: e.g. (128, 64)',
default='(100,)',
constraint='^\\(\\s*(?:\\d+,\\s*)+(?:\\d+,?\\s*)?\\)$',
),
ChoiceField(
name='activation_mlp',
label='activation',
description='Activation function for the hidden layer.',
default="relu",
choices=["identity", "logistic", "tanh", "relu"],
),
ChoiceField(
name='solver_mlp',
label='solver',
description='Solver for weight optimization.',
default="adam",
choices=["lbfgs", "sgd", "adam"],
),
FloatField(
name='alpha_mlp',
label='alpha',
description='L2 penality (regularization term) parameter.',
default=0.0001,
),
ChoiceField(
name='learning_rate_mlp',
label='learning_rate',
description='Learning rate schedule for weight updates. Only used for sgd solver.',
default="constant",
choices=["constant", "invscaling", "adaptive"],
),
FloatField(
name='learning_rate_init_mlp',
label='learning_rate_init',
description='The initial learning rate used. Controls the step-size in updating the weights. Only used for sgd or adam solver.',
default=0.001,
),
FloatField(
name='power_t_mlp',
label='power_t',
description='Exponent for inverse scaling learning rate. Only used for sgd solver with invscaling for learning_rate.',
default=0.5,
),
IntField(
name='max_iter_mlp',
label='max_iter',
description='Maximum number of iterations. The solver iterates until convergence (determined by tol) or this number of iterations.',
default=200,
),
FloatField(
name='tol_mlp',
label='tol',
description='Tolerance for the optimization.',
default=1e-4,
),
BoolField(
name='early_stopping_mlp',
label='early_stopping',
description='Whether to use early stopping to terminate training when validation score is not improving.',
default=False,
),
FloatField(
name='validation_fraction_mlp',
label='validation_fraction',
description='The proportion of training data to set aside as validation set for early stopping.',
default=0.1,
),
],
'OneClassSVM': [
ChoiceField(
name='kernel_svm',
label='kernel',
description='Specifies the kernel type to be used in the algorithm.',
default="rbf",
choices=["linear", "poly", "rbf", "sigmoid", "precomputed"],
),
IntField(
name='degree_svm',
label='degree',
description='Degree of the polynomial kernel function (‘poly’). Ignored by all other kernels.',
default=3,
),
ChoiceField(
name='gamma_svm',
label='gamma',
description='Kernel coefficient for rbf, poly and sigmoid kernels.',
default="scale",
choices=["scale", "auto"],
),
FloatField(
name='coef0_svm',
label='coef0',
description='Independent term in kernel function. It is only significant in poly and sigmoid.',
default=0.0,
),
FloatField(
name='tol_svm',
label='tol',
description='Tolerance for stopping criterion.',
default=1e-3,
),
FloatField(
name='nu_svm',
label='nu',
description='An upper bound on the fraction of training errors and a lower bound of the fraction of support vectors. \
Should be in the interval (0, 1].',
default=0.5,
),
BoolField(
name='shrinking_svm',
label='shrinking',
description='Whether to use the shrinking heuristic.',
default=True,
),
IntField(
name='max_iter_svm',
label='max_iter',
description='Hard limit on iterations within solver, or -1 for no limit.',
default=-1,
),
],
},
section='SETTINGS'
) %}
{% set calibrated = BoolField(
name='calibrated',
label='Calibrate algorithm predictions',
description='Calibrate the prediction probabilities, eliminating model-imparted bias.',
default=True,
section='SETTINGS',
) %}
{% set cv_algorithm = ChoiceField(
name='cv_algorithm',
label='Cross-Validation Algorithm',
description='Select a cross-validation method for training and evaluating the pipeline, and for making predictions. \
StratifiedGroupKFold or RepeatedStratifiedGroupKFold are recommended because they will maintain class ratios \
across train/validation splits (stratification of labels) and will group compounds by the first 14 characters of their \
InChI Keys to avoid compounds with multiple entries from appearing in both the train and validation sets.',
default='RepeatedStratifiedGroupKFold',
choices={
'KFold': 'sk.model_selection.KFold',
'GroupKFold': 'sk.model_selection.GroupKFold',
'RepeatedKFold': 'sk.model_selection.RepeatedKFold',
'StratifiedKFold': 'sk.model_selection.StratifiedKFold',
'StratifiedGroupKFold': 'StratifiedGroupKFold',
'RepeatedStratifiedKFold': 'sk.model_selection.RepeatedStratifiedKFold',
'RepeatedStratifiedGroupKFold': 'RepeatedStratifiedGroupKFold'
},
section='SETTINGS',
) %}
{% set cross_validation_n_folds = IntField(
name='cross_validation_n_folds',
label='Number of Cross-Validated Folds',
description='Cross-validation is employed as a strategy to train the model on data that the model has not seen before, more folds will ensure that the model is generalizing well.',
default=5,
min=2,
max=10,
section='SETTINGS'
) %}
{% set cross_validation_n_repeats = IntField(
name='cross_validation_n_repeats',
label='Number of Cross-Validated Repetitions',
description='Number of repetitions of cross-validation to perform. \
Only used for RepeatedKFold, RepeatedStratifiedKFold, or RepeatedStratifiedGroupKFold cross-validation algorithms, \
which repeat cross-validation with different randomizations. This yields multiple predictions per compound, which can be evaluated for consistency.',
default=5,
min=2,
section='SETTINGS'
) %}
{% set hyper_param_search = ChoiceField(
name='hyper_param_search',
label='Hyper Parameter Search Type',
default='None',
description='Hyper parameter searching is used to automatically select the best parameters (using the primary metric as the criteria).',
choices={
'None': 'None',
'RandomizedSearchCV': 'sk.model_selection.RandomizedSearchCV',
'GridSearchCV': 'sk.model_selection.GridSearchCV',
},
section='SETTINGS'
) %}
{% set primary_metric = ChoiceField(
name='primary_metric',
label='Primary Evaluation Metric',
default='roc_auc',
description='The primary evaluation metric is used for deciding how we assess the performance of our model. \
Area under the receiver operating characteristic curve (roc_auc) is recommended for most tasks.',
choices=[
'accuracy',
'adjusted_mutual_info_score',
'adjusted_rand_score',
'average_precision',
'balanced_accuracy',
'completeness_score',
'explained_variance',
'f1',
'f1_macro',
'f1_micro',
'f1_weighted',
'fowlkes_mallows_score',
'homogeneity_score',
'jaccard',
'jaccard_macro',
'jaccard_micro',
'jaccard_weighted',
'max_error',
'mutual_info_score',
'neg_brier_score',
'neg_log_loss',
'neg_mean_absolute_error',
'neg_mean_squared_error',
'neg_mean_squared_log_error',
'neg_median_absolute_error',
'neg_root_mean_squared_error',
'normalized_mutual_info_score',
'precision',
'precision_macro',
'precision_micro',
'precision_weighted',
'r2',
'recall',
'recall_macro',
'recall_micro',
'recall_weighted',
'roc_auc',
'roc_auc_ovo',
'roc_auc_ovo_weighted',
'roc_auc_ovr',
'roc_auc_ovr_weighted',
'v_measure_score'
],
section='SETTINGS'
) %}
{% set evaluation_metrics = MultiChoiceField(
name='evaluation_metrics',
label='Evaluation Metrics',
default=[],
description='Additional evaluation metrics can be specified, these metrics will also be reported for all models trained.',
value=[],
choices=[
'accuracy',
'adjusted_mutual_info_score',
'adjusted_rand_score',
'average_precision',
'balanced_accuracy',
'completeness_score',
'explained_variance',
'f1',
'f1_macro',
'f1_micro',
'f1_weighted',
'fowlkes_mallows_score',
'homogeneity_score',
'jaccard',
'jaccard_macro',
'jaccard_micro',
'jaccard_weighted',
'max_error',
'mutual_info_score',
'neg_brier_score',
'neg_log_loss',
'neg_mean_absolute_error',
'neg_mean_squared_error',
'neg_mean_squared_log_error',
'neg_median_absolute_error',
'neg_root_mean_squared_error',
'normalized_mutual_info_score',
'precision',
'precision_macro',
'precision_micro',
'precision_weighted',
'r2',
'recall',
'recall_macro',
'recall_micro',
'recall_weighted',
'roc_auc',
'roc_auc_ovo',
'roc_auc_ovo_weighted',
'roc_auc_ovr',
'roc_auc_ovr_weighted',
'v_measure_score'
],
section='SETTINGS',
) %}
{% set all_metrics = [primary_metric.value] + evaluation_metrics.value %}
%%appyter code_hide
{% set algorithm_code = {
'GradientBoostingClassifier': 'sk.ensemble.GradientBoostingClassifier(loss="{}", learning_rate={}, n_estimators={}, subsample={}, criterion="{}", tol={})',
'RandomForestClassifier': 'sk.ensemble.RandomForestClassifier(n_estimators={}, criterion="{}", min_samples_split={}, min_samples_leaf={}, max_features={}, min_impurity_decrease={}, n_jobs=-1, class_weight={}, ccp_alpha={})',
'AdaBoostClassifier': 'sk.ensemble.AdaBoostClassifier(sk.tree.DecisionTreeClassifier(max_depth={}), n_estimators={}, learning_rate={}, algorithm="{}")',
'ExtraTreesClassifier': 'sk.ensemble.ExtraTreesClassifier(n_estimators={}, criterion="{}", min_samples_split={}, min_samples_leaf={}, max_features={}, min_impurity_decrease={}, n_jobs=-1, class_weight={}, ccp_alpha={})',
'DecisionTreeClassifier': 'sk.tree.DecisionTreeClassifier(criterion="{}", splitter="{}", min_samples_split={}, min_samples_leaf={}, max_features={}, min_impurity_decrease={}, class_weight={}, ccp_alpha={})',
'KNeighborsClassifier': 'sk.neighbors.KNeighborsClassifier(n_neighbors={}, weights="{}", algorithm="{}", leaf_size={}, p={}, metric="{}", n_jobs=-1)',
'RadiusNeighborsClassifier': 'sk.neighbors.RadiusNeighborsClassifier(radius={}, weights="{}", algorithm="{}", leaf_size={}, p={}, metric="{}", outlier_label="most_frequent", n_jobs=-1)',
'MLPClassifier': 'sk.neural_network.MLPClassifier(hidden_layer_sizes={}, activation="{}", solver="{}", alpha={}, learning_rate="{}", learning_rate_init={}, power_t={}, max_iter={}, tol={}, early_stopping={}, validation_fraction={})',
'OneClassSVM': 'sk.svm.OneClassSVM(kernel="{}", degree={}, gamma="{}", coef0={}, tol={}, nu={}, shrinking={}, max_iter={})',
} %}
%%appyter markdown
We apply a {% if hyper_param_search.value != 'None' %}{{ hyper_param_search.raw_value }} search for the hyper parameters
of a {% endif %}sklearn pipeline with a dimensionality reduction step of {{ dimensionality_reduction.raw_value }}
{% if feature_selection.value != 'None' %}and a feature selection step of {{ feature_selection.raw_value }}
{% endif %} and a{% if calibrated.value %} calibrated{%endif %} {{ algorithm.raw_value }} classifier
using {{ cross_validation_n_folds.value }}-fold {{ cv_algorithm.raw_value }} cross-validation,
optimizing {{ primary_metric.value }}{% if evaluation_metrics.value %} and computing {{ ', '.join(evaluation_metrics.value) }}{% endif %}.
```
Note that training can take a long time as we are training a model for each of multiple cross-validation splits.
```
%%appyter code_exec
{% if algorithm.raw_value == 'GradientBoostingClassifier' %}
## Early stopping function
def early_stopping(n_rounds, tol=0.001):
def early_stopping_func(i, self, local):
rounds = getattr(self, '__rounds', 0)
last = getattr(self, '__last', None)
current = self.train_score_[i]
if last and current and abs(current - last) < tol:
rounds += 1
if rounds > n_rounds:
return True
else:
rounds = 0
setattr(self, '__last', current)
setattr(self, '__rounds', rounds)
return False
return early_stopping_func
{% endif %}
{#
param_grid = {
'reduce_dim__n_components': randint(2, 1024),
{% if algorithm.raw_value == 'GradientBoostingClassifier' %}
'clf__loss': ['deviance', 'exponential'],
'clf__learning_rate': randfloat(0.001, 1.),
'clf__subsample': randfloat(0.01, 1.),
{% elif algorithm.raw_value == 'RandomForestClassifier' %}
'clf__oob_score': [True],
'clf__criterion': ['gini', 'entropy'],
{% endif %}
'clf__n_estimators': randint(10, 200),
'clf__max_depth': randint(20, 50),
'clf__max_features': ['sqrt', 'log2', None],
'clf__min_impurity_decrease': randfloat(0., 0.2),
'clf__min_weight_fraction_leaf': randfloat(0., 0.5),
}
fit_params = {
{% if algorithm.raw_value == 'GradientBoostingClassifier' %}
'clf__monitor': early_stopping(5),
{% endif %}
}
#}
cv = {{ cv_algorithm }}(
n_splits={{ cross_validation_n_folds }},
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
n_repeats={{ cross_validation_n_repeats }},
{% else %}
shuffle=True,
{% endif %}
random_state=rng,
)
{% if cv_algorithm.raw_value in ['GroupKFold', 'StratifiedGroupKFold', 'RepeatedStratifiedGroupKFold'] %}
groups=[key[:14] for key in X.index] # Group compounds by atom connectivity
{% endif %}
# Scoring parameters
primary_metric = '{{ primary_metric }}'
evaluation_metrics = {{ evaluation_metrics }}
scoring_params = {k: metrics.get_scorer(k)
for k in [primary_metric, *evaluation_metrics]}
%%appyter code_exec
{% if hyper_param_search.value == 'None' %}
df_results = pd.DataFrame()
# Store performance on each split for computing ROC and PRC curves
fprs = []
tprs = []
precs = []
recs = []
# Store cross-validation test predictions and folds
y_proba_cv = [[] for _ in range(len(y))]
folds_cv = [[] for _ in range(len(y))]
# Store models
models = []
{% if cv_algorithm.raw_value in ['GroupKFold', 'StratifiedGroupKFold', 'RepeatedStratifiedGroupKFold'] %}
groups=[key[:14] for key in X.index] # Group compounds by atom connectivity
for fold, (train, test) in tqdm(enumerate(cv.split(X.values, y, groups=groups))):
{% else %}
for fold, (train, test) in tqdm(enumerate(cv.split(X.values, y))):
{% endif %}
model =
{%- if hyper_param_search.value != 'None' %} {{ hyper_param_search }}({% endif -%}
sk.pipeline.Pipeline([
{%- if dimensionality_reduction.value != 'None' %}
('reduce_dim', {{ dimensionality_reduction }}),
{% endif %}
{%- if feature_selection.value != 'None' %}
('feature_selection', {{ feature_selection }}),
{% endif %}
('clf', {% if algorithm.raw_value == 'MLPClassifier' %}{{ algorithm_code.get(algorithm.raw_value).format(algorithm.value[0].value|str_to_tuple, *algorithm.value[1:]) }}
{% elif algorithm.raw_value in ['DecisionTreeClassifier', 'RandomForestClassifier', 'ExtraTreesClassifier'] %}{{ algorithm_code.get(algorithm.raw_value).format(algorithm.value[0].value, algorithm.value[1].value, algorithm.value[2].value|int_or_float, algorithm.value[3].value|int_or_float, *algorithm.value[4:]) }}
{% else %}{{ algorithm_code.get(algorithm.raw_value).format(*algorithm.value) }}{% endif %}
),
])
{%- if hyper_param_search.value != 'None' %}){% endif %}
model.fit(X.values[train], y[train])
{% if calibrated.value %}
calibrator = sk.calibration.CalibratedClassifierCV(model, cv='prefit')
calibrator.fit(X.values[test], y[test])
model = calibrator
{% endif %}
{% for metric in all_metrics %}
df_results.loc[fold, '{{ metric }}'] = scoring_params['{{ metric }}'](model, X.values[test], y[test])
{% endfor %}
y_proba = model.predict_proba(X.values[test]) # Probability prediction will be True
for i in range(len(test)):
y_proba_cv[test[i]].append(y_proba[i, 1])
folds_cv[test[i]].append(fold % {{ cross_validation_n_folds }})
model_fpr, model_tpr, _ = metrics.roc_curve(y[test], y_proba[:, 1])
model_prec, model_rec, _ = metrics.precision_recall_curve(y[test], y_proba[:, 1])
fprs.append(model_fpr)
tprs.append(model_tpr)
precs.append(model_prec)
recs.append(model_rec)
models.append(model)
assert not(any(len(probs) == 0 for probs in y_proba_cv)), 'All probabilities should have been calculated'
display(df_results.agg(['mean', 'std']))
{% else %}
model.fit(X.values, y)
df_results = model.cv_results_
{% endif %}
```
This visualization shows the cross-validated performance of the model. Low fold variance and high AUC is desired in a well-generalized model.
* ROC curve: [roc.svg](./roc.svg)
* Precision-recall curve: [prc.svg](./prc.svg)
* Confusion matrix: [confusion_matrix.svg](./confusion_matrix.svg)
```
%%appyter code_exec
fig, ax = plt.subplots()
tprs_interp = []
aucs = []
mean_fpr = np.linspace(0, 1, 100)
for fold, (fpr, tpr) in enumerate(zip(fprs, tprs)):
tpr_interp = np.interp(mean_fpr, fpr, tpr)
tpr_interp[0] = 0.
roc_auc = metrics.auc(fpr, tpr)
tprs_interp.append(tpr_interp)
aucs.append(roc_auc)
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
ax.plot(fpr, tpr, alpha=0.4)
{% else %}
ax.plot(fpr, tpr, alpha=0.4, label='ROC Fold %d (AUC=%0.3f)' % (fold, roc_auc))
{% endif %}
mean_tpr = np.mean(tprs_interp, axis=0)
mean_tpr[-1] = 1.0
mean_auc = sk.metrics.auc(mean_fpr, mean_tpr)
std_auc = np.std(aucs)
ax.plot(mean_fpr, mean_tpr, color='b',
label=r'Mean ROC (AUC = %0.2f $\pm$ %0.2f)' % (mean_auc, std_auc),
lw=2, alpha=.8)
std_tpr = np.std(tprs_interp, axis=0)
tprs_upper = np.minimum(mean_tpr + std_tpr, 1)
tprs_lower = np.maximum(mean_tpr - std_tpr, 0)
plt.fill_between(mean_fpr, tprs_lower, tprs_upper, color='grey', alpha=.2)
ax.plot([0,1],[0,1],'--', label='Random')
ax.set_xlabel('False Positive Rate')
ax.set_ylabel('True Positive Rate')
ax.legend()
plt.savefig('roc.svg')
figure_header('Figure', 'Receiver operating characteristic (ROC) curves across cross-validation splits ({})'.format(make_clickable('roc.svg')))
plt.show()
figure_legend('Figure', 'Receiver operating characteristic (ROC) curves across cross-validation splits ({})'.format(make_clickable('roc.svg')),
'Individual curves are shown for each {{ cross_validation_n_folds }}-fold cross-validation split{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}, repeated with {{ cross_validation_n_repeats }} different randomizations{% endif %}. \
Mean ROC shows the average and standard deviation across cross-validation splits.')
z = (mean_auc - 0.5)/std_auc
cl = sp.stats.norm.cdf(z) * 100
ci = sp.stats.norm.interval(0.95, loc=mean_auc, scale=std_auc)
print('Confidence interval (95%)', ci)
print("We are %0.3f %% confident the model's results are not just chance." % (cl))
if cl > 95:
print('This is statistically significant. These results can be trusted.')
else:
print('This is not statistically significant. These results should not be trusted.')
%%appyter code_exec
fig, ax = plt.subplots()
precs_interp = []
prc_aucs = []
mean_rec = np.linspace(0, 1, 100)
for fold, (rec, prec) in enumerate(zip(recs, precs)):
prec_interp = np.interp(mean_rec, rec[::-1], prec[::-1])
prc_auc = metrics.auc(rec, prec)
precs_interp.append(prec_interp)
prc_aucs.append(prc_auc)
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
ax.plot(rec, prec, alpha=0.4)
{% else %}
ax.plot(rec, prec, alpha=0.4, label='PRC Fold %d (AUC=%0.3f)' % (fold, prc_auc))
{% endif %}
mean_prec = np.mean(precs_interp, axis=0)
mean_auc = sk.metrics.auc(mean_rec, mean_prec)
std_auc = np.std(prc_aucs)
ax.plot(mean_rec, mean_prec, color='b',
label=r'Mean PRC (AUC = %0.2f $\pm$ %0.2f)' % (mean_auc, std_auc),
lw=2, alpha=.8)
std_prec = np.std(precs_interp, axis=0)
precs_upper = np.minimum(mean_prec + std_prec, 1)
precs_lower = np.maximum(mean_prec - std_prec, 0)
plt.fill_between(mean_rec, precs_lower, precs_upper, color='grey', alpha=.2)
ax.set_xlabel('Recall')
ax.set_ylabel('Precision')
ax.legend()
plt.savefig('prc.svg')
figure_header('Figure', 'Precision-recall curves (PRC) across cross-validation splits ({})'.format(make_clickable('prc.svg')))
plt.show()
figure_legend('Figure', 'Precision-recall curves (PRC) across cross-validation splits ({})'.format(make_clickable('prc.svg')),
'Individual curves are shown for each {{ cross_validation_n_folds }}-fold cross-validation split{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}, repeated with {{ cross_validation_n_repeats }} different randomizations{% endif %}. \
Mean PRC shows the average and standard deviation across cross-validation splits.')
sns.heatmap(
metrics.confusion_matrix(y, np.array([np.mean(probs) for probs in y_proba_cv]) > 0.5),
annot=True,
cmap=plt.cm.Blues,
fmt='g'
)
plt.xlabel('Predicted')
plt.ylabel('True')
plt.savefig('confusion_matrix.svg')
figure_header('Figure', 'Confusion matrix for cross-validation predictions ({})'.format(make_clickable('confusion_matrix.svg')))
plt.show()
figure_legend('Figure', 'Confusion matrix for cross-validation predictions ({})'.format(make_clickable('confusion_matrix.svg')),
'Note that the predicted probabilities can be greatly affected by imbalanced labels and by the model choice. \
Thus, performance measures such as ROC and PRC, which evaluate performance across a range of prediction thresholds, \
are more useful than the confusion-matrix, which uses an fixed cutoff of 0.5')
```
## Examine predictions
By examining the validation-set predictions, we can rank the positive compounds and identify additional compounds that were not known to be in the positive class, but nevertheless had high predictions. These may share similar properties with the known compounds.
First, we can compare the distribution of predictions for positive and negative labels.
```
%%appyter code_exec
# Calculate mean and deviation of predictions
y_probas = np.array([np.mean(probs) for probs in y_proba_cv])
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
y_probas_std = np.array([np.std(probs) for probs in y_proba_cv])
# Find minimum non-zero standard deviation to avoid dividing by zero when computing t-statistic
min_y_probas_std = max(np.min(y_probas_std[y_probas_std != 0]), 1e-10)
t_stats = (y_probas - np.mean(y_probas)) / (np.maximum(y_probas_std, min_y_probas_std)/np.sqrt({{ cross_validation_n_repeats }}))
# Calculate p-value using one-sample t-test
p_vals_t = 1-sp.stats.t({{ cross_validation_n_repeats }}-1).cdf(t_stats)
{% endif %}
%%appyter code_exec
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
# Simulate mean predictions by
y_probas_means_{{ cross_validation_n_repeats }} = []
y_probas_values = np.array(y_proba_cv).flatten()
np.random.seed(rng)
for i in tqdm(range(100000)):
y_probas_means_{{ cross_validation_n_repeats }}.append(np.mean(np.random.choice(y_probas_values, {{ cross_validation_n_repeats }})))
y_probas_means_{{ cross_validation_n_repeats }} = np.array(sorted(y_probas_means_{{ cross_validation_n_repeats }}))
{% endif %}
%%appyter code_exec
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
y_probas_ts_{{ cross_validation_n_repeats }} = []
mean_y_probas = np.mean(y_probas)
y_probas_values = np.array(y_proba_cv).flatten()
np.random.seed(rng)
for i in tqdm(range(100000)):
sample = np.random.choice(y_probas_values, {{ cross_validation_n_repeats }})
y_probas_ts_{{ cross_validation_n_repeats }}.append((np.mean(sample) - mean_y_probas) / (np.maximum(np.std(sample), min_y_probas_std)/np.sqrt({{ cross_validation_n_repeats }})))
y_probas_ts_{{ cross_validation_n_repeats }} = np.array(sorted(y_probas_ts_{{ cross_validation_n_repeats }}))
{% endif %}
%%appyter code_exec
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
max_mean = np.max(y_probas_means_{{ cross_validation_n_repeats }})
p_vals = np.array(list(tqdm((1 - np.argwhere(y_probas_means_{{ cross_validation_n_repeats }} >= min(pred, max_mean))[0][0] / len(y_probas_means_{{ cross_validation_n_repeats }})
for pred in y_probas), total=len(y_probas))))
{% endif %}
%%appyter code_exec
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
max_t = np.max(y_probas_ts_{{ cross_validation_n_repeats }})
p_vals_t_sim = np.array(list(tqdm((1 - np.argwhere(y_probas_ts_{{ cross_validation_n_repeats }} >= min(t, max_t))[0][0] / len(y_probas_ts_{{ cross_validation_n_repeats }})
for t in t_stats), total=len(t_stats))))
{% endif %}
%%appyter code_exec
sns.distplot(y_probas[y == 0], bins=int(np.sqrt(np.sum(y == 0))*10), kde_kws={'gridsize':2000}, label='Not known positive compound')
sns.distplot(y_probas[y == 1], bins=int(np.sqrt(np.sum(y == 1))*10), kde_kws={'gridsize':2000}, label='Known positive compound')
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
sns.distplot(y_probas_means_{{ cross_validation_n_repeats }}, bins=int(np.sqrt(len(y_probas_means_{{ cross_validation_n_repeats }}))*10), kde_kws={'gridsize':2000}, label='Null distribution\n(simulated)')
{% endif %}
plt.xlabel('Mean Predicted Probability')
plt.xlim([np.min(y_probas), np.percentile(y_probas, 99)])
plt.legend()
plt.savefig('mean-prediction-distribution.svg')
figure_header('Figure', 'Distribution of{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %} mean{% endif %} cross-validation predictions ({})'.format(make_clickable('mean-prediction-distribution.svg')))
plt.show()
figure_legend('Figure', 'Distribution of{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %} mean{% endif %} cross-validation predictions ({})'.format(make_clickable('mean-prediction-distribution.svg')),
'Distribution of{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %} mean{% endif %} cross-validation predictions for all {number_of_compounds} compounds, \
including both those with known positive labels and other small molecules.\
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %} The null distribution was simulated by drawing independent samples of predictions with replacement from the distribution of all predictions.{% endif %}'.format(number_of_compounds=X.shape[0]))
%%appyter code_exec
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
sns.distplot(t_stats[y == 0], bins=int(np.sqrt(np.sum(y == 0))*10), kde_kws={'gridsize':2000}, label='Not known positive compound')
sns.distplot(t_stats[y == 1], bins=int(np.sqrt(np.sum(y == 1))*10), kde_kws={'gridsize':2000}, label='Known positive compound')
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
sns.distplot(y_probas_ts_{{ cross_validation_n_repeats }}, bins=int(np.sqrt(len(y_probas_ts_{{ cross_validation_n_repeats }}))*10), kde_kws={'gridsize':2000}, label='Null distribution\n(simulated)')
{% endif %}
plt.xlabel('t-statistic')
plt.xlim([-20,20])
plt.legend()
plt.savefig('t-statistic-distribution.svg')
figure_header('Figure', 'Distribution of t-statistics ({})'.format(make_clickable('t-statistic-distribution.svg')))
plt.show()
figure_legend('Figure', 'Distribution of t-statistics ({})'.format(make_clickable('t-statistic-distribution.svg')),
'Distributions of t-statistics for all {number_of_compounds} compounds, \
including both those with known positive labels and other small molecules. \
The null distribution was simulated by drawing independent samples of predictions with replacement from the distribution of all predictions.'.format(number_of_compounds=X.shape[0]))
{% endif %}
```
Overlaying the predictions on a visualization of the input space allows us to examine the predictions and may indicate groups of highly predicted compounds.
```
%%appyter code_exec
# Add attributes for plotting to Dataframe
X_reduced_df['Predicted Probability'] = y_probas
X_reduced_df['log10(pred)'] = np.log10(y_probas + 1e-10)
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
X_reduced_df['p-value'] = p_vals_t_sim
X_reduced_df['log10(p-value)'] = np.log10(X_reduced_df['p-value'])
X_reduced_df['Standard Deviation'] = y_probas_std
{% endif %}
X_reduced_df['Cross-validation fold'] = folds_cv
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
X_reduced_df['marker size'] = 2*np.minimum(2-np.log10(X_reduced_df['p-value']), 5)
{% else %}
max_p, min_p = np.min(-X_reduced_df['log10(pred)']), np.max(-X_reduced_df['log10(pred)'])
X_reduced_df['marker size'] = (-X_reduced_df['log10(pred)'] - min_p) / (max_p - min_p) * 6 + 4
{% endif %}
X_reduced_df['text'] = ['<br>'.join(['Drug Name: ' + str(name),
'InChI Key: ' + str(inchi),
'Predicted Probability: {:.1e}'.format(p),
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
'Standard Deviation: {:.1e}'.format(s),
'p-value: {:.1e}'.format(p_val),
{% endif %}
'Label: ' + str(label),
'Cross-validation fold: ' + str(fold)])
for name, inchi, p, {% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}s, p_val, {% endif %}label, fold in zip(X_reduced_df['Drug Name'],
X_reduced_df['InChI Key'],
X_reduced_df['Predicted Probability'],
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
X_reduced_df['Standard Deviation'],
X_reduced_df['p-value'],
{% endif %}
X_reduced_df['Label'],
X_reduced_df['Cross-validation fold'])]
X_reduced_df.to_csv('X_reduced_df.csv')
# Helper function for formatting Plotly colorbar
def colorbar_param(values_log10, **kwargs):
min_val = np.floor(np.min(values_log10))
max_val = np.ceil(np.max(values_log10))
ticks1 = 10**np.arange(min_val, max_val+1)
ticks2 = 3*10**np.arange(min_val, max_val)
ticktext = sorted(np.concatenate([ticks1, ticks2]))
tickvals = list(np.log10(ticktext))
ticktext = ['{:.0e}'.format(text) for text in ticktext]
return dict(ticktext=ticktext, tickvals=tickvals, **kwargs)
fig = go.Figure()
for label in sorted(set(X_reduced_df['Label'])):
X_plot = X_reduced_df[X_reduced_df['Label'] == label].sort_values(['Predicted Probability'])
fig.add_trace(go.Scatter(mode='markers',
x=X_plot['Component 1'], y=X_plot['Component 2'],
text=X_plot['text'],
name=label,
marker=dict(
color=X_plot['log10(pred)'],
cmin=np.percentile(X_reduced_df['log10(pred)'], 50),
cmax=np.max(X_reduced_df['log10(pred)']),
size=X_plot['marker size'],
colorbar=colorbar_param(X_plot['log10(pred)'], title='Predicted Probability'),
symbol=X_plot['marker symbol'],
line_width=1,
colorscale='plasma'
)))
fig.update_layout(height=600, width=800,
xaxis_title='Component 1',
yaxis_title='Component 2',
title_text='Predicted Probabilities ({{ visualization_reduction.raw_value }})',
legend_title_text='Target Label',
legend=dict(
yanchor="top",
y=0.98,
xanchor="left",
x=0.02
),
template='simple_white')
figure_header('Figure', '{{ visualization_reduction.raw_value }} dimensionality reduction of the input feature space overlayed with predictions')
fig.show()
figure_legend('Figure', '{{ visualization_reduction.raw_value }} dimensionality reduction of the input feature space overlayed with predictions',
f'Each point represents one of {X.shape[0]} compounds, with {X.shape[1]} features per compound, \
taken from the following datasets: {", ".join(sepl1000datasets + attribute_datasets)}. \
Compounds with known positive labels are marked by X\'s. The color and size of each point correspond to the{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %} mean{% endif %} predicted \
probability {% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}and its signficance (estimated from the simulated t-statistic null distribution), respectively{% endif %}.')
```
Full tables of top-predicted compounds with and without known positive labels are shown below.
```
%%appyter code_exec
# Obtain prediction results
results = pd.DataFrame(np.array([
querysepl1000fwd.get_drug_names(X.index),
Drugmonizome.get_drug_names(X.index),
folds_cv,
y,
y_probas,
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
y_probas_std,
t_stats,
p_vals,
p_vals_t,
p_vals_t_sim,
{% endif %}
]).T, columns=[
'Name (L1000FWD)',
'Name (Drugmonizome)',
'Cross-validation fold',
'Known',
'Prediction Probability',
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
'Prediction Probability Std. Dev.',
't statistic',
'p value (simulated mean distribution)',
'p value (one sample t test)',
'p value (simulated t distribution)',
{% endif %}
], index=X.index).astype({'Known': 'bool',
'Prediction Probability': 'float64',
{% if cv_algorithm.raw_value in ['RepeatedStratifiedKFold', 'RepeatedStratifiedGroupKFold'] %}
'Prediction Probability Std. Dev.': 'float64',
't statistic': 'float64',
'p value (simulated mean distribution)': 'float64',
'p value (one sample t test)': 'float64',
'p value (simulated t distribution)': 'float64',{% endif %}})
results.to_csv('drug_cv_predictions.csv')
# Rank predictions
figure_header('Table', 'Top-predicted compounds ({})'.format(make_clickable('drug_cv_predictions.csv')))
show(results.reset_index(), maxBytes=0, order=[[ 5, "desc" ]], columnDefs=[{'width': '110px', 'targets': [0, 1]}])
figure_legend('Table', 'Top-predicted compounds ({})'.format(make_clickable('drug_cv_predictions.csv')),
f'All {X.shape[0]} compounds ranked by cross-validation prediction probability. \
Search \'true\' or \'false\' to filter compounds with known positive labels or not, respectively. \
The table can also be sorted by other columns by selecting the column name in the header.')
```
## Examine feature importances
The relative contribution of each input feature to the final model predictions can be estimated for recursive feature selection and for a variety of tree-based models. Note that this analysis is not available if a dimensionality reduction algorithm is used.
```
%%appyter markdown
{% if feature_selection.raw_value == 'RecursiveSelectionFromExtraTrees' and dimensionality_reduction.raw_value == 'None' %}
When recursive feature selection is performed, the features are ranked by the stage at which they were removed.
Selected (i.e. estimated best) features are have importance 1. The ranks are averaged across cross-validation
splits to produce an average importance score. The full feature importance table is available at
[feature_importance.csv](./feature_importance.csv).
{% endif %}
%%appyter code_exec
{% if feature_selection.raw_value == 'RecursiveSelectionFromExtraTrees' and dimensionality_reduction.raw_value == 'None' %}
all_rankings = []
{% endif %}
{% if algorithm.raw_value in ['GradientBoostingClassifier', 'RandomForestClassifier', 'AdaBoostClassifier', 'ExtraTreesClassifier', 'DecisionTreeClassifier'] %}
all_feature_importances = []
{% endif %}
for model in models:
{% if calibrated.value %}
for calibrated_clf in model.calibrated_classifiers_:
pipeline = calibrated_clf.base_estimator
{% else %}
pipeline = model
{% endif %}
{% if feature_selection.raw_value == 'RecursiveSelectionFromExtraTrees' %}
ranking = pipeline['feature_selection'].ranking_
all_rankings.append(ranking)
{% endif %}
{% if algorithm.raw_value in ['GradientBoostingClassifier', 'RandomForestClassifier', 'AdaBoostClassifier', 'ExtraTreesClassifier', 'DecisionTreeClassifier'] %}
{% if feature_selection.raw_value != 'None' %}
feature_importances = np.zeros(pipeline['feature_selection'].get_support().shape)
feature_importances[pipeline['feature_selection'].get_support()] = pipeline['clf'].feature_importances_
{% else %}
feature_importances = pipeline['clf'].feature_importances_
{% endif %}
all_feature_importances.append(feature_importances)
{% endif %}
%%appyter code_exec
{% if dimensionality_reduction.raw_value == 'None' %}
df_feat_imp = pd.DataFrame({'Feature': X.columns,
'Dataset': reduce(lambda a,b: a+b, ([dataset]*size for dataset, size in dataset_sizes)),
{% if feature_selection.raw_value == 'RecursiveSelectionFromExtraTrees' %}
'Ranking Mean': np.mean(all_rankings, axis=0),
'Ranking Std. Dev.': np.std(all_rankings, axis=0),
{% endif %}
{% if algorithm.raw_value in ['GradientBoostingClassifier', 'RandomForestClassifier', 'AdaBoostClassifier', 'ExtraTreesClassifier', 'DecisionTreeClassifier'] %}
'Importance Mean': np.mean(all_feature_importances, axis=0),
'Importance Std. Dev.': np.std(all_feature_importances, axis=0)})
{% endif %}
df_feat_imp = df_feat_imp.set_index('Feature').sort_values('Importance Mean', ascending=False)
{% if feature_selection.raw_value == 'RecursiveSelectionFromExtraTrees' %}
figure_header('Table', 'Input features ranked by relative importance ({})'.format(make_clickable('feature_importance.csv')))
show(df_feat_imp.reset_index(), maxBytes=0, order=[[ 2, "asc"]])
figure_legend('Table', 'Input features ranked by relative importance ({})'.format(make_clickable('feature_importance.csv')),
f'All {X.shape[1]} input features are ranked by their relative importance. \
Feature ranking (Ranking Mean and Std. Dev.) specifies the round of recursive feature selection on which a given feature was eliminated. \
A feature with lower ranking is more \
important. {% if algorithm.raw_value in ['GradientBoostingClassifier', 'RandomForestClassifier', 'AdaBoostClassifier', 'ExtraTreesClassifier', 'DecisionTreeClassifier'] %}Tree-based \
models can also be used to calculate impurity-based feature importances (Importance Mean and Std. Dev.). {% endif %}Search a dataset \
name to filter features from a given dataset. \
The table can also be sorted by other columns by selecting the column name in the header.')
{% elif algorithm.raw_value in ['GradientBoostingClassifier', 'RandomForestClassifier', 'AdaBoostClassifier', 'ExtraTreesClassifier', 'DecisionTreeClassifier'] %}
figure_header('Table', 'Input features ranked by relative importance ({})'.format(make_clickable('feature_importance.csv')))
show(df_feat_imp.reset_index(), maxBytes=0, order=[[ 2, "desc"]])
figure_legend('Table', 'Input features ranked by relative importance ({})'.format(make_clickable('feature_importance.csv')),
f'All {X.shape[1]} input features are ranked by their relative importance. \
Tree-based models can be used to calculate impurity-based feature importances (Importance Mean and Std. Dev.). \
Search a dataset name to filter features from a given dataset. \
The table can also be sorted by other columns by selecting the column name in the header.')
{% else %}
figure_header('Table', 'Input features ({})'.format(make_clickable('feature_importance.csv')))
show(df_feat_imp.reset_index(), maxBytes=0)
figure_legend('Table', 'Input features ({})'.format(make_clickable('feature_importance.csv')),
f'All {X.shape[1]} input features. No ranking of features was possible for this pipeline.')
{% endif %}
df_feat_imp.to_csv('feature_importance.csv')
{% endif %}
%%appyter code_exec
{% if feature_selection.raw_value == 'RecursiveSelectionFromExtraTrees' and dimensionality_reduction.raw_value == 'None' %}
fig, axs = plt.subplots(1, 2, figsize=(15, 5))
df_feat_imp = df_feat_imp.sort_values('Ranking Mean')
for dataset in set(df_feat_imp.Dataset):
importance_scores = df_feat_imp.loc[df_feat_imp.Dataset == dataset]['Ranking Mean'].values
importance_scores_std = df_feat_imp.loc[df_feat_imp.Dataset == dataset]['Ranking Std. Dev.'].values
lower = importance_scores - importance_scores_std
upper = importance_scores + importance_scores_std
axs[0].plot(importance_scores, label=dataset)
axs[0].fill_between(np.arange(len(importance_scores)), lower, upper, alpha=.2)
axs[1].plot(np.linspace(0, 1, len(importance_scores)), importance_scores, label=dataset)
axs[1].fill_between(np.linspace(0, 1, len(importance_scores)), lower, upper, alpha=.2)
for i in [0, 1]:
axs[i].legend()
axs[i].set_title('Distribution of feature ranking from recursive feature elimination')
axs[i].set_ylabel('Average feature ranking\n(lower ranking is more important)')
axs[0].set_xlabel('Ranked features (absolute count)')
axs[1].set_xlabel('Ranked features (relative count)')
axs[0].set_xlim([0,512])
plt.tight_layout()
plt.savefig('feature_importance_rfe.svg')
figure_header('Figure', 'Distribution of feature rankings from recursive feature elimination ({})'.format(make_clickable('feature_importance_rfe.svg')))
plt.show()
figure_legend('Figure', 'Distribution of feature rankings from recursive feature elimination ({})'.format(make_clickable('feature_importance_rfe.svg')),
'The distribution of feature rankings from recursive feature elimination for each dataset. \
Features with lower scores were retained for more rounds during recursive feature selection \
and have greater relative importance.')
{% endif %}
%%appyter code_exec
{% if algorithm.raw_value in ['GradientBoostingClassifier', 'RandomForestClassifier', 'AdaBoostClassifier', 'ExtraTreesClassifier', 'DecisionTreeClassifier'] and dimensionality_reduction.raw_value == 'None' %}
fig, axs = plt.subplots(2, 2, figsize=(15, 10))
df_feat_imp = df_feat_imp.sort_values('Importance Mean', ascending=False)
for dataset in set(df_feat_imp.Dataset):
importance_scores = df_feat_imp.loc[df_feat_imp.Dataset == dataset]['Importance Mean'].values
importance_scores_std = df_feat_imp.loc[df_feat_imp.Dataset == dataset]['Importance Std. Dev.'].values
lower = importance_scores - importance_scores_std
upper = importance_scores + importance_scores_std
axs[0][0].plot(importance_scores, label=dataset)
axs[0][0].fill_between(np.arange(len(importance_scores)), lower, upper, alpha=.2)
axs[0][1].plot(np.linspace(0, 1, len(importance_scores)), importance_scores, label=dataset)
axs[0][1].fill_between(np.linspace(0, 1, len(importance_scores)), lower, upper, alpha=.2)
importance_scores = np.cumsum(df_feat_imp.loc[df_feat_imp.Dataset == dataset]['Importance Mean'].values)
axs[1][0].plot(importance_scores, label=dataset)
axs[1][1].plot(np.linspace(0, 1, len(importance_scores)), importance_scores, label=dataset)
for i in [0, 1]:
axs[0][i].legend()
axs[0][i].set_title('Distribution of feature scores from model')
axs[1][i].set_title('Cumulative distribution of feature scores from model')
axs[i][0].set_xlabel('Ranked features (absolute count)')
axs[i][1].set_xlabel('Ranked features (relative count)')
axs[0][i].set_ylabel('Average feature importance\n(higher score is more important)')
axs[1][i].set_ylabel('Cumulative sum of feature importance')
axs[i][0].set_xlim([0,512])
plt.tight_layout()
plt.savefig('feature_importance.svg')
figure_header('Figure', 'Distribution of feature scores from model ({})'.format(make_clickable('feature_importance.svg')))
plt.show()
figure_legend('Figure', 'Distribution of feature scores from model ({})'.format(make_clickable('feature_importance.svg')),
'The distribution of impurity-based feature importances for each dataset. \
Features with higher scores have greater relative contribution to the overall tree-based model.')
{% endif %}
```
| github_jupyter |
# Time-frequency behaviour
We are still looking at the two stream instability investigated in [Hawke, Andersson and Comer (2013)](http://dx.doi.org/10.1088/0264-9381/30/14/145007) - see the paper, `Figure1.ipynb` and `InstabilityAnimation.ipynb` for details. Here we are interested in the growth rate of the different modes with time - again we will only look at the linearized solution. Essentially we are reproducing parts of Figure 6 from the paper here.
```
%matplotlib inline
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
plt.rcParams.update({'font.size': 14, 'font.family': 'serif'})
import EOS
import compute_perturbation as mf
# EOS options.
options_unstable = dict([('EOS_Derivatives', EOS.EOS_Paper), \
('kappa', [0.5, 0.5]) , \
('kappa_12', 0.0) , \
('kappa_delta', 0.5) , \
('gamma', [1.6, 1.8]) , \
('m', [1.0, 1.0]) , \
('sigma', [0.0, 0.0])])
# Domain
options_unstable['L'] = 1.0
options_unstable['a_max'] = 128
# Background
rho_bg = 1.0
v_bg1 = 0.0
W_bg1 = 1.0 / np.sqrt(1.0 - v_bg1**2)
v_bg2 = 0.6
W_bg2 = 1.0 / np.sqrt(1.0 - v_bg2**2)
options_unstable['n_bg'] = np.array([[rho_bg * W_bg1, rho_bg * W_bg1 * v_bg1], \
[rho_bg * W_bg2, rho_bg * W_bg2 * v_bg2]])
# Perturbation - single sine wave
d_n = 1.0e-5
d_v = 0.0
W = 1.0 / np.sqrt(1.0 - d_v**2)
delta_nhat = np.zeros((options_unstable['a_max'], 2, 2), complex)
delta_nhat[0, 0, 0] = -0.5 * 1j * W * d_n
delta_nhat[0, 0, 1] = -0.5 * 1j * W * d_n * d_v
# This part adds adjusted floating point noise.
for a in range(1, options_unstable['a_max']):
delta_nhat[a, :, 0] += 1e-22 * 1j
options_unstable['delta_nhat'] = delta_nhat
# Now set up the time domain
t_unstable = np.linspace(0.0, 2.0, 51)
# Now the frequency space solution
FFT_linear = np.zeros((len(t_unstable), options_unstable['a_max']))
for a in range(1, options_unstable['a_max']):
inv_L_delta_F = mf.calc_inv_L_delta_F(t_unstable, a, options_unstable)
FFT_linear[:, a-1] = np.log10(np.maximum(np.abs(inv_L_delta_F[:, 0, 0]), 1.e-16))
# Now with the adjusted omega to account for the numerical differencing error.
options_adjusted = options_unstable.copy()
omega = np.pi * np.arange(options_adjusted['a_max']) / (options_adjusted['L'] * options_adjusted['a_max'])
omega_prime = (8.0 * np.sin(omega) - 2.0 * np.sin(2.0 * omega)) / 6.0 * options_adjusted['a_max']
options_adjusted['omega'] = omega_prime
# Now the adjusted frequency space solution
FFT_linear_adjusted = np.zeros((len(t_unstable), options_adjusted['a_max']))
for a in range(1, options_adjusted['a_max']):
inv_L_delta_F_adjusted = mf.calc_inv_L_delta_F(t_unstable, a, options_adjusted)
FFT_linear_adjusted[:, a-1] = np.log10(np.maximum(np.abs(inv_L_delta_F_adjusted[:, 0, 0]), 1.e-16))
fig = plt.figure(figsize = (12,6))
ax1 = fig.add_subplot(1,2,1)
ax1.pcolor(t_unstable * options_unstable['a_max'], \
np.linspace(0.0, 1.0, options_unstable['a_max']), \
FFT_linear.T)
ax1.set_xlabel("$t k_{max}$")
ax1.set_ylabel("$\omega_k / (\pi k_{max})$")
ax1.set_title("Exact")
ax2 = fig.add_subplot(1,2,2)
ax2.pcolor(t_unstable * options_adjusted['a_max'], \
np.linspace(0.0, 1.0, options_adjusted['a_max']), \
FFT_linear_adjusted.T)
ax2.set_xlabel("$t k_{max}$")
ax2.set_ylabel("$\omega_k / (\pi k_{max})$")
ax2.set_title("Adjusted")
plt.show()
```
| github_jupyter |
## ENGR 213 Content
### Daily Map
#### 2021
### Week 1
### Stress, Strain, Allowable Stress
**HW 1** HW 1 contains problems from Vable CH 1 and CH 2
[Engineering Failures](https://online-engineering.case.edu/blog/disastrous-engineering-failures-due-to-ethics) Which offends you the most and why?
V 1.20 position dependent stress
V 1.36 normal and shear
V 1.40 normal and shear
V 1.47 angled surface
V 2.5 normal strain
V 2.15 strain
V 2.33 shear strain
#### Day 1
**Lec 1** Average Shear Stress (CH 1 normal and shear)
Allowable Stress Design https://www.engineeringtoolbox.com/factors-safety-fos-d_1624.html
RP 1: Hanging Lamp
#### Day 2
**Lec 2** Strain (CH 2 normal and shear)
RP 2 Glue Joint
### Week 2
### Stress-Strain Curves and Thermal Strain
**HW 2** HW 2 contains problems from Vable CH 3
3.1 interpret stress-strain diagram
3.11 interpret stress-strain diagram
3.12 construct stress-strain diagram
3.42 modulus of elasticity
3.49 two competing materials
3.127 thermal strain
3.49 with conditions from 3.127 applied
#### Day 1:
**Lec 3:** Stress-Strain Curve (CH 3), Young's Modulus, Poisson's Ratio (CH 3)
RP 3: Stress-Strain Diagram and application
#### Day 2:
**Lec 4:** Thermal Strain (CH 3)
RP 4: Thermal Strain
### Week 3
### Axial Loads and Torsion (Drive Shafts)
**HW 3** HW 3 contains problems from
#### Day 1:
**Lec 5:** Axial Loads (CH 4)
Rp 5: Axial Loads
#### Day 2:
**Lec 6:** Torsion in Shafts (CH 5)
### Week 4
**Midterm:** Through HW 2:
#### Day 1:
Review
#### Day 2:
Midterm
### Week 5
### Project and Torsion
**HW 4** HW 5 contains problems from
#### Day 1:
Project Introduction
#### Day 2:
### Week 6
### Transverse Shear/Combined Loading
**HW 5** HW 5 contains problems from
#### Day 1:
#### Day 2:
### Week 7
### Plane Stress, Max Stresses, Mohr's Circle
**HW 6** HW 6 contains problems from sections
#### Day 1:
#### Day 2:
### Week 8
**Midterm:** Through HW 5:
#### Day 1:
Review
#### Day 2:
Midterm
### Week 9
### Beam Deflection and Design
**HW 7** HW 7 contains problems from sections
#### Day 1:
#### Day 2:
## Project Due at end of Week!
### Week 10
### Finish up and Special Topics
#### Day 1:
#### Day 2:
Review for Final
### Past syllabus based on Hibbler
Day 1
Average Shear Stress (V CH 1 normal and shear)
Allowable Stress Design https://www.engineeringtoolbox.com/factors-safety-fos-d_1624.html
RP 1
Day 2
Strain (V CH 2 normal and shear)
Stress-Strain Curve (V 3.1.1)
RP 2
Day 3
Strain energy (V 3.1.4), Poissan’s Ratio ( V 3.1.2)
RP 3
Day 4
Axial load
RP 4
HW 1 Due
Day 5
Thermal Stress
RP 5
Day 6
Shear and Moment Diagrams
RP 6
HW 2 Due
Day 7
Test Review
Day 8
Exam #1
Day 9
Project Discussion
Day 10
Torsion
RP 7
HW 3 Due
Day 11
Transverse Shear
RP 8
Day 12
Combined loadings
RP 9
HW 4 Due
Day 13
Plane Stress
Principle Stresses and Max Shear Stresses,
RP 10
Day 14
Mohr’s Circle
RP 11
HW 5 Due
Day 15
Test Review
**Project Progress Check-In
Day 16
Exam #2
Day 17
Ch.12 Beam Deflection
RP 12
Day 18
Ch.11 Beam Design
RP 13,
HW 6 Due
**Term Project Due by Midnight**
Day 19
Ch.11 Beam Design
RP 14
Day 20
Finish Course Material
RP 15
HW 7 Due
FINAL EXAM
1:00 – 3:00pm
| github_jupyter |
In order to successfully complete this assignment you need to participate both individually and in groups during class on **Wednesday March 25**.
# In-Class Assignment: Statistical Models
<img alt="Short video of a tool that demonstrates the natrual occurace of gaussian curves using a plico toy" src="https://i1.wp.com/psychbrief.com/wp-content/uploads/2017/12/normal-distribution-2.gif?resize=444%2C250&ssl=1">
Image From: https://en.wikipedia.org/wiki/Hydra
### Agenda for today's class (80 minutes)
</p>
1. [(20 minutes) pre-class_review](#pre-class_review)
1. [(20 minutes) Moving window methods](#Moving_window_methods)
2. [(20 minutes) Speedtest Example](#Speedtest_Example)
----
<a name="pre-class_review"></a>
# 1. Pre-class assignment review
* [0324--Stats-pre-class-assignment](0324--Stats-pre-class-assignment.ipynb)
---
<a name="Moving_window_methods"></a>
# 2. Moving window methods
We are going to explore "moving windows methods" used a lot in signal processing and data anlaysis to "clean up" noise in your data.
✅ **<font color=red>DO THIS:</font>** First lets download the noaa example again:
```
%matplotlib inline
import matplotlib.pylab as plt
import noaa_scraper
data = noaa_scraper.get_airtemperature_from_files()
plt.plot(data)
```
The following code makes a histogram of values within the noaa data. Notice the bimodal nature of this example:
```
h = plt.hist(data, 100, density=True);
plt.xlabel('Temperature (deg F)')
plt.ylabel('Number of measurements');
```
We are going to explore "moving windows methods" used a lot in signal processing and data anlaysis to "clean up" noise in your data.
In this example we use a window size of 300 data points. Each point new point is an average of the a window around the original data.
```
import numpy as np
data = np.array(data)
#moving average code from https://stackoverflow.com/questions/13728392/moving-average-or-running-mean
N = 300
totals, moving_aves = [0], []
for i, x in enumerate(data, 1):
totals.append(totals[i-1] + x)
if i>=N:
moving_ave = (totals[i] - totals[i-N])/N
#can do stuff with moving_ave here
moving_aves.append(moving_ave)
```
We then plot the new values here:
```
plt.plot(moving_aves)
h = plt.hist(moving_aves, 100, density=True);
plt.xlabel('Temperature (deg F)')
plt.ylabel('Number of measurements');
```
✅ **<font color=red>DO THIS:</font>** Modify the above answer to use larger and larger window sizes. Try chaning orders of magnitude (ex: 300, 3000, 30000).
✅ **<font color=red>DO THIS:</font>** Make an animated gif or mpeg video suitable for showing in a powerpoit presentation how smoothing changes when you change window size. Use a logrithmic sale for the window size $N$. I recommend $N$ in [10,100,1000,10000,100000]:
----
<a name="Speedtest_Example"></a>
# 3. Speedtest Data
Now lets look at some more statistical modeling. In the last git commit, I included a ```speedtest.dat``` file. (Note: this data was collected using this [python script in gist](https://gist.github.com/colbrydi/027a4b95399a8543b2a7c0cadb88bb0c)). The data are comma seperated values that can easily be uploaded using the following code:
```
import pandas
from IPython.display import display
filename = 'speedtest.dat'
data = pandas.read_csv(filename)
data['download'] = data['download']/1000000
data['upload'] = data['upload']/1000000
display(data)
%matplotlib inline
ax = data.plot.bar(x = 'location', y=['download'])
```
See if you can generate a bar chart that is averaged based on a location group. (Hint: if you get stuck this code is in the gist file)
-----
### Congratulations, we're done!
### Course Resources:
- [Syllabus](https://docs.google.com/document/d/e/2PACX-1vTW4OzeUNhsuG_zvh06MT4r1tguxLFXGFCiMVN49XJJRYfekb7E6LyfGLP5tyLcHqcUNJjH2Vk-Isd8/pub)
- [Preliminary Schedule](https://docs.google.com/spreadsheets/d/e/2PACX-1vRsQcyH1nlbSD4x7zvHWAbAcLrGWRo_RqeFyt2loQPgt3MxirrI5ADVFW9IoeLGSBSu_Uo6e8BE4IQc/pubhtml?gid=2142090757&single=true)
- [D2L Page](https://d2l.msu.edu/d2l/home/912152)
- [Git Repository](https://gitlab.msu.edu/colbrydi/cmse802-s20)
© Copyright 2020, Michigan State University Board of Trustees
| github_jupyter |
```
import plotly
import plotly.graph_objs as go
import numpy as np
from plotly.offline import iplot
from ipywidgets import GridspecLayout, widgets, HBox
plotly.offline.init_notebook_mode()
n_points = 8
marker_size = 3
x_shift = 3.5
opacity = 0.75
from torchvision import datasets, transforms
import torch
import torch.nn.functional as F
def place_digits_randomly(mnist_imgs):
new_img = torch.zeros([64, 64])
x_pos, y_pos = torch.randint(0, 64-28, (2, 2))
for i in range(2):
new_img[y_pos[i]:y_pos[i]+28, x_pos[i]:x_pos[i]+28] += mnist_imgs[i][0]
return new_img
SEED = 7
np.random.seed(SEED)
torch.manual_seed(SEED)
mnist = datasets.MNIST('./data', train=True, transform=transforms.ToTensor(), download=True)
i_sample = np.random.randint(len(mnist), size=2)
extended_img = place_digits_randomly([mnist[i_sample[0]][0], mnist[i_sample[1]][0]])
# flip image to get desired view
extended_img = torch.flip(extended_img, dims=[0])
def create_target(n_points, marker_size):
x, y = np.linspace(-1, 1, n_points), np.linspace(-1, 1, n_points)
X, Y = np.meshgrid(x, y)
X, Y = X.reshape(-1), Y.reshape(-1)
target = np.concatenate((X[np.newaxis, :], Y[np.newaxis, :],
np.ones([1, len(X)])), 0)
G = go.Scatter3d(x=target[0], y=target[1], z=target[2], hoverinfo='none',
mode='markers', marker=dict(size=marker_size, color='green', opacity=opacity))
return target, G
def create_init_transformation(target, n_points):
A_theta = np.array([[1, 0, 0],
[0, 1, 0]])
source_xy = A_theta @ target
source = np.concatenate((source_xy[0][np.newaxis, :],
source_xy[1][np.newaxis, :],
-4*np.ones([1, n_points**2])),
0)
G_transformed = go.Scatter3d(x=source[0], y=source[1], z=source[2],
hoverinfo='none', mode='markers', visible=True,
marker=dict(size=marker_size, color='green', opacity=opacity))
return source, G_transformed
def create_connecting_corner_lines(source, target, n_points):
corners = []
for i in [0, n_points-1, n_points*(n_points-1), n_points**2 - 1]:
corner = go.Scatter3d(x=np.array([target[0][i], source[0][i]]),
y=np.array([target[1][i], source[1][i]]),
z=np.array([target[2][i], source[2][i]]),
mode='lines', hoverinfo='none',
line=dict(color='blue', width=2), opacity=opacity)
corners.append(corner)
return corners
def create_layout(width=850, height=450):
layout = dict(
title='',
showlegend=False,
width=width,
height=height,
margin=dict(t=0, r=0, l=10, b=0),
scene=dict(
camera=dict(up=dict(x=0, y=0.5, z=0),
center=dict(x=0.9, y=-0.8, z=-0.1),
eye=dict(x=-1.3, y=0.6, z=1.8)),
#center=dict(x=0.3, y=-0.3, z=0)),
dragmode=False,
),
)
return layout
target, G_target = create_target(n_points, marker_size)
source, G_source = create_init_transformation(target, n_points)
corners = create_connecting_corner_lines(source, target, n_points)
target_2, G_target_2 = create_target(n_points, marker_size)
# update some stuff
target_2[0] = target_2[0] + x_shift
G_target_2['x'] = target_2[0]
G_target_2['marker']['color'] = 'red'
source_2, G_source_2 = create_init_transformation(target_2, n_points)
# update some stuff
source_2[2] = -(3.5/4) * source_2[2]
G_source_2['z'] = source_2[2]
G_source_2['marker']['color'] = 'red'
corners_2 = create_connecting_corner_lines(source_2, target_2, n_points)
x = np.linspace(-1, 1, 64)
y = np.linspace(-1, 1, 64)
mnist_img_original = go.Surface(x=x, y=y, z=-4*np.ones((64, 64)),
surfacecolor=extended_img, cmin=0, cmax=1,
colorscale='Gray', showscale=False)
mnist_img_resample = go.Surface(x=x, y=y, z=1*np.ones((64, 64)),
surfacecolor=extended_img, cmin=0, cmax=1,
colorscale='Gray', showscale=False)
mnist_img_resample_2 = go.Surface(x=x+x_shift, y=y, z=1*np.ones((64, 64)),
surfacecolor=extended_img, cmin=0, cmax=1,
colorscale='Gray', showscale=False)
mnist_img_reconstructed = go.Surface(x=x+x_shift, y=y, z=3.5*np.ones((64, 64)),
surfacecolor=extended_img, cmin=0, cmax=1,
colorscale='Gray', showscale=False)
data = [mnist_img_resample, mnist_img_resample_2, mnist_img_reconstructed, mnist_img_original,
G_target, G_target_2, G_source, G_source_2] + corners + corners_2
layout = create_layout()
base_figure = go.Figure(data=data, layout=layout)
base_figure
from ipywidgets import GridspecLayout, widgets, HBox
def response(change):
with fig.batch_update():
A_theta = np.array([[s.value, 0, t_x.value],
[0, s.value, t_y.value],
[0, 0, 1]])
A_theta_inv = np.linalg.pinv(A_theta)
source = A_theta @ target
# update G source
fig.data[6]['x'] = source[0]
fig.data[6]['y'] = source[1]
# update mnist image in middle
A_theta = torch.from_numpy(A_theta[0:2, :]).unsqueeze(0).type(torch.float32)
grid = F.affine_grid(A_theta, size=(1, 1, 64, 64), align_corners=False)
out = F.grid_sample(extended_img.unsqueeze(0).unsqueeze(0), grid, align_corners=False)
fig.data[0]['surfacecolor'] = out[0, 0]
fig.data[1]['surfacecolor'] = out[0, 0]
# update G source with using inverse transformation
x_rec_grid = A_theta_inv @ target
fig.data[5]['x'] = x_rec_grid[0] + x_shift
fig.data[5]['y'] = x_rec_grid[1]
# update reconstructed mnist
A_theta_inv = torch.from_numpy(A_theta_inv[0:2, :]).unsqueeze(0).type(torch.float32)
#print(A_theta_inv.shape, A_theta.shape)
grid = F.affine_grid(A_theta_inv, size=(1, 1, 64, 64), align_corners=False)
out = F.grid_sample(out, grid, align_corners=False)
fig.data[2]['surfacecolor'] = out[0, 0]
# update connecting corners for standard attention
for index, i_point in enumerate([0, n_points-1, n_points*(n_points-1), n_points**2 - 1]):
fig.data[-8 + index]['x'] = np.array([target[0][i_point], source[0][i_point]])
fig.data[-8 + index]['y'] = np.array([target[1][i_point], source[1][i_point]])
# update connecting corners for inv attention
for index, i_point in enumerate([0, n_points-1, n_points*(n_points-1), n_points**2 - 1]):
fig.data[-4 + index]['x'] = np.array([x_rec_grid[0][i_point] + x_shift,
target[0][i_point] + x_shift])
fig.data[-4 + index]['y'] = np.array([x_rec_grid[1][i_point],
target[1][i_point]])
def reset_values(b):
with fig.batch_update():
s.value, t_x.value, t_y.value = 1., 0., 0.
A_theta = np.array([[s.value, 0, t_x.value],
[0, s.value, t_y.value],
[0, 0, 1]])
A_theta_inv = np.linalg.pinv(A_theta)
source = A_theta @ target
# update G source
fig.data[6]['x'] = source[0]
fig.data[6]['y'] = source[1]
# update mnist image in middle
A_theta = torch.from_numpy(A_theta[0:2, :]).unsqueeze(0).type(torch.float32)
grid = F.affine_grid(A_theta, size=(1, 1, 64, 64), align_corners=False)
out = F.grid_sample(extended_img.unsqueeze(0).unsqueeze(0), grid, align_corners=False)
fig.data[0]['surfacecolor'] = out[0, 0]
fig.data[1]['surfacecolor'] = out[0, 0]
# update G source with using inverse transformation
x_rec_grid = A_theta_inv @ target
fig.data[5]['x'] = x_rec_grid[0] + x_shift
fig.data[5]['y'] = x_rec_grid[1]
# update reconstructed mnist
A_theta_inv = torch.from_numpy(A_theta_inv[0:2, :]).unsqueeze(0).type(torch.float32)
#print(A_theta_inv.shape, A_theta.shape)
grid = F.affine_grid(A_theta_inv, size=(1, 1, 64, 64), align_corners=False)
out = F.grid_sample(out, grid, align_corners=False)
fig.data[2]['surfacecolor'] = out[0, 0]
# update connecting corners for standard attention
for index, i_point in enumerate([0, n_points-1, n_points*(n_points-1), n_points**2 - 1]):
fig.data[-8 + index]['x'] = np.array([target[0][i_point], source[0][i_point]])
fig.data[-8 + index]['y'] = np.array([target[1][i_point], source[1][i_point]])
# update connecting corners for inv attention
for index, i_point in enumerate([0, n_points-1, n_points*(n_points-1), n_points**2 - 1]):
fig.data[-4 + index]['x'] = np.array([x_rec_grid[0][i_point] + x_shift,
target[0][i_point] + x_shift])
fig.data[-4 + index]['y'] = np.array([x_rec_grid[1][i_point],
target[1][i_point]])
fig = go.FigureWidget(base_figure)
# create sliders
s = widgets.FloatSlider(value=1, min=-1, max=1, step=0.1)
t_x = widgets.FloatSlider(value=0, min=-1, max=1, step=0.1)
t_y = widgets.FloatSlider(value=0, min=-1, max=1, step=0.1)
# add behavior to sliders
s.observe(response, names="value")
t_x.observe(response, names="value")
t_y.observe(response, names="value")
# create reset button
reset_button = widgets.Button(description = "Reset")
# add behavior
reset_button.on_click(reset_values)
# make box around figure
fig_box = widgets.Box([fig])
# create title widget
title = widgets.HTML(value="<h2 style='color:#303F5F'>Creation of Attention Crops and Inverse Transformation</h2>")
# create widget displaying formula
formula_str_1 = r'$\color{green}{\left[ \begin{array}{c} x_k^s \\ y_k^s \\ 1 \end{array} \right] = ' + \
r'\left[ \begin{array}{c} s & 0 & t_x \\' + \
r'0 & s & t_y \\ 0 & 0 & 1\end{array} \right]' + \
r'\left[\begin{array}{c} x_k^t \\ y_k^t \\ 1 \end{array} \right]}$'
formula_label_1 = widgets.Label(value=formula_str_1)
formula_str_2 = r'$\color{red}{\left[ \begin{array}{c} x_k^s \\ y_k^s \\ 1 \end{array} \right] = ' + \
r' \left( \left[\begin{array}{c} s & 0 & t_x \\' + \
r'0 & s & t_y \\ 0 & 0 & 1\end{array} \right] \right)^{+} ' + \
r'\left[\begin{array}{c} x_k^t \\ y_k^t \\ 1 \end{array} \right]}$'
formula_label_2 = widgets.Label(value=formula_str_2)
# put everything together in GridSpecLayout
n_rows, n_cols = 12, 8
grid_spec = GridspecLayout(n_rows, n_cols, height='auto', width='100%')
grid_spec[2, n_cols-2:n_cols] = HBox([widgets.Label(value=r'$s$'), s])
grid_spec[3, n_cols-2:n_cols] = HBox([widgets.Label(value=r'$t_x$'), t_x])
grid_spec[4, n_cols-2:n_cols] = HBox([widgets.Label(value=r'$t_y$'), t_y])
grid_spec[5, n_cols-2:n_cols] = reset_button
grid_spec[2:n_rows-1, 0:n_cols-2] = fig_box
grid_spec[0, 0:8] = title
grid_spec[1, 1:3] = formula_label_1
grid_spec[1, 3:6] = formula_label_2
grid_spec
```
| github_jupyter |
# SDSCH Rest
Welcome! The purpose of this exercise is to create a small lightning control system. This system will rely on a REST service to which the devices will send requests.
You can find the devices on [this website](https://cyclimse.github.io/sdsch-rest/) or you can use the embedded version below. It's up to you!
```
import IPython
url = 'https://cyclimse.github.io/sdsch-rest/'
iframe = '<iframe src=' + url + ' width=700 height=700></iframe>'
IPython.display.HTML(iframe)
```
Try to play around with the buttons on each device. You may notice that the requests of each device is being logged into the terminal.
Currently, not much is happening because we have yet to create the service the devices will talk to. Let's do it!
```
from flask import Flask, jsonify, request, Response
from flask_cors import CORS
# Flask is a very popular framework for creating HTTP servers in Python, you can find out more the project website:
# https://flask.palletsprojects.com/en/1.1.x/
app = Flask(__name__)
# Because we serve the app and send request from two distinct servers,
# we use CORS to let the browser figure it out with the backend.
cors = CORS(app)
last_temperature = 0
# In flask, you can use a variety of syntax to create new routes.
# Here, we will be using a decorator to indicate the route om which to mount our handler.
@app.route("/therm", methods=["POST"])
def thermHandler():
global last_temperature
data = request.get_json()
last_temperature = data["temperature"]
return Response(status=201)
@app.route("/lamp", methods=["GET"])
def lampHandler():
global last_temperature
if last_temperature > 20:
return jsonify({"color":"red", "intensity": 100, "frequency": 60.0/0.5})
else:
# Complete the code!
# What could you change to notify the user if the room is getting chilly?
return jsonify({})
```
Without running the code, try to figure out what it does. Can you see any problem with the current implementation ?
We will discuss this a bit latter. Right now, we will try and launch our newly created API.
```
app.run(debug=False, port=5005)
```
Let's check if it's running properly! We could use the web application, but a lot of the times when working with APIs, we work on the server before the client. In the other words, we may not always have a client to test our API with.
However, because HTTP requests are generic, we can use our browser to partially check the behavior of our API. Try loading http://localhost:5005/lamp in your browser of choice. What do you see?
We can also try to check the behavior of our API with some command line tools. We will be using the utility curl. Here we will check if the temperature gets updated when we perform a POST request on /therm.
```console
curl -X POST -H "Content-Type: application/json" http://localhost:5005/therm -d '{"temperature":25}'
```
The -X flags specify the type of request. Additionally, we set the header field "Content-type" to specify to the server that we will be expecting a json response. Finally, we pass as data the json object that we will be posting.
Try to craft a `curl` command to GET the status of the lamp!
`curl` is a very powerful piece of utility. With the right API, you can even use it to send [emails](https://sendgrid.com/)!
Let's get back to our server. Now switching between two colors is not ideal. Try to use multiple colors. You can even try to create a gradient. Here is a small [tutorial](https://bsouthga.dev/posts/color-gradients-with-python) on color gradients in Python.
```
def get_color(temperature):
# Try to come up with something beautiful!
# Dont forget to use this in your handler for /lamp.
pass
```
Now, something is not quite right! The thermometer only post the temperature every so often, while the lamp request it more frequently. As a result, the temperature shifts are quite herky jerky.
How could you adjust your code to make the transition smoother ?
Up until now, you were guided heavily. Now it's your turn to be in charge!
There's one device we haven't talked about yet: the led band. It's a collection of colored lamps.
Try and figure out how you could control it from the server by using the console on the website. Your task will be to come up with the prettiest and most elaborate animations using the LEDs.
Have fun!
PS: some mistakes are hidden in this lab! Bonus points if you can find them all. Hint: are all REST constraints that you've seen in the course properly respected?
| github_jupyter |
# Smart signatures with ASA
#### 06.3 Writing Smart Contracts
##### Peter Gruber (peter.gruber@usi.ch)
2022-01-12
* Use Smart Signatures with ASAs
* Design a contract for token burning
## Setup
See notebook 04.1, the lines below will always automatically load functions in `algo_util.py`, the five accounts and the Purestake credentials
```
# Loading shared code and credentials
import sys, os
codepath = '..'+os.path.sep+'..'+os.path.sep+'sharedCode'
sys.path.append(codepath)
from algo_util import *
cred = load_credentials()
# Shortcuts to directly access the 3 main accounts
MyAlgo = cred['MyAlgo']
Alice = cred['Alice']
Bob = cred['Bob']
Charlie = cred['Charlie']
Dina = cred['Dina']
from algosdk import account, mnemonic
from algosdk.v2client import algod
from algosdk.future import transaction
from algosdk.future.transaction import PaymentTxn
from algosdk.future.transaction import AssetConfigTxn, AssetTransferTxn, AssetFreezeTxn
from algosdk.future.transaction import LogicSig, LogicSigTransaction
import algosdk.error
import json
import base64
import hashlib
from pyteal import *
# Initialize the algod client (Testnet or Mainnet)
algod_client = algod.AlgodClient(algod_token='', algod_address=cred['algod_test'], headers=cred['purestake_token'])
print(MyAlgo['public'])
print(Alice['public'])
print(Bob['public'])
print(Charlie['public'])
```
#### Check Purestake API
```
last_block = algod_client.status()["last-round"]
print(f"Last committed block is: {last_block}")
asset_holdings_df(algod_client,MyAlgo['public'])
```
## Burning
Burning seems simple: send money to a contract that always says "no".<br>
**Question:** how can a contract that always says "no" opt into an ASA?
#### Step 1: Define conditions in Pyteal
* Want to design a Burn contract for WSC coin
* Need WSC coin index
* For all Txn fields, see https://pyteal.readthedocs.io/en/stable/accessing_transaction_field.html
```
# prepare the burn condition
WSC_idx = 71140107 # <---------- change this to your WSC coin !!!!!
burn_cond = And (
Txn.type_enum() == TxnType.AssetTransfer, # Must be an "asset transfer" transaction
Txn.amount() == Int(0), # Do not pay out ALGOs
Txn.asset_amount() == Int(0), # Do also not pay out ASAs
Txn.xfer_asset() == Int(WSC_idx) # Specific asset index
)
# prepare random condition
import random
a = Int( random.randrange(2**32-1) )
random_cond = ( a == a )
fee_cond = Txn.fee() <= Int(1000)
safety_cond = And(
# Txn.type_enum() == TxnType.Payment, # This standard payment condition makes no senese here
Txn.close_remainder_to() == Global.zero_address(),
Txn.rekey_to() == Global.zero_address(),
)
burn_pyteal = And(
burn_cond,
random_cond,
fee_cond,
safety_cond
)
```
##### Step 2: Compile PyTeal -> Teal
```
burn_teal = compileTeal(burn_pyteal, Mode.Signature, version=3)
print(burn_teal)
```
#### Step 3: Compile Teal -> Bytecode for AVM
```
Burn = algod_client.compile(burn_teal)
```
#### Step 4: Fund burning contract
* The Burn contract has to pay TX fees for the opt-in transaction
```
# Step 1: prepare transaction
sp = algod_client.suggested_params()
amt = int(0.1*1e6) + int(0.1*1e6) + int(0.001*1e6) # Min holdings + min holdings for 1 ASA + TX fee
txn = transaction.PaymentTxn(sender=Alice['public'], sp=sp, receiver=Burn['hash'], amt=amt)
# Step 2+3+4: sign and send and wait ...
stxn = txn.sign(Alice['private'])
txid = algod_client.send_transaction(stxn)
txinfo = wait_for_confirmation(algod_client, txid)
```
#### Step 5: Burn contract opts-into WSC coin to allow burning
* This is an AssetTransferTx, that is signed by the Smart Signature
* Remember, opt-in is a transfer of zero units of an ASA to oneself
```
# Step 5.1: Prepare
sp = algod_client.suggested_params()
txn = AssetTransferTxn(Burn['hash'], sp, Burn['hash'], 0, WSC_idx)
# Step 5.2: Sign
encodedProg = Burn['result'].encode()
program = base64.decodebytes(encodedProg)
lsig = LogicSig(program)
stxn = LogicSigTransaction(txn, lsig)
# Step 5.3 Send
txid = algod_client.send_transaction(stxn)
# Step 5.4 Wait for ...
txinfo = wait_for_confirmation(algod_client, txid)
print('http://testnet.algoexplorer.io/tx/'+txid)
```
## The Burn contract is now ready for use
#### Manual burn transaction
* MyAlgo burns 8 WSC coins
* Simple AssetTransferTxn
```
# WAIT a minute ... how many decimals does the WSC have?
WSC_decimals = algod_client.asset_info(WSC_idx)['params']['decimals']
print(WSC_decimals)
# Step 1: prepare transaction
sp = algod_client.suggested_params()
amt = int( 8 * 10**WSC_decimals ) # <---------8 WSC coins in SMALL unit
txn = AssetTransferTxn(sender=MyAlgo['public'], sp=sp,
receiver=Burn['hash'], amt=amt,
index=WSC_idx)
# Step 2+3+4: sign and send and wait ...
stxn = txn.sign(MyAlgo['private'])
txid = algod_client.send_transaction(stxn)
txinfo = wait_for_confirmation(algod_client, txid)
```
### QR code for burning
* Burning via QR code
```
# URL for burning WITH asset index
url = 'algorand://' + Burn['hash'] + '?amount=1000&asset='+str(WSC_idx)+'¬e=Burning'
print(url)
import qrcode
qr = qrcode.QRCode(version=1,box_size=5,border=4)
qr.add_data(url)
qr.make(fit=True)
qr.make_image(fill_color="black", back_color="white")
```
## Exercise 1
* Reconsider the **burn condition** and discuss possible safety issues.
* Espeically, discuss the fact that repeated transactions with zero ALGOs or with zero WSC are possible
```python
burn_cond = And (
Txn.type_enum() == TxnType.AssetTransfer, # Must be an "asset transfer" TX
Txn.amount() == Int(0), # Do not pay out ALGOs
Txn.asset_amount() == Int(0), # Do also not pay out ASAs
Txn.xfer_asset() == Int(WSC_idx) # Specific asset index
)
```
## Exercise 2
* Reconsider the **safety condition**
* Why was one common safety condition commented out?
```python
safety_cond = And(
# Txn.type_enum() == TxnType.Payment, # <--- why???
Txn.close_remainder_to() == Global.zero_address(),
Txn.rekey_to() == Global.zero_address(),
)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from functools import partial
from Gmodel import GProblem
```
# Visualising solutions
The code creates sample solution landscapes similar to those reported by Grim et al. - not included in paper.
```
solutions = []
for s in [0, 5, 10, 20]:
solutions.append(GProblem(2000, 2, 2, 2, s).solution)
fig, ax = plt.subplots(nrows=4, ncols=1, figsize=(20, 20), sharex=True, sharey=True)
fig.suptitle(' First 200 points of sample landscapes, created with various smoothness factors.')
ax[0].plot(range(0,200), solutions[0][:200], label='Smoothness 0')
ax[0].set_title("Smoothness 0")
ax[1].plot(range(0,200), solutions[1][:200], label='Smoothness 5')
ax[1].set_title("Smoothness 5")
ax[2].plot(range(0,200), solutions[2][:200], label='Smoothness 10')
ax[2].set_title("Smoothness 10")
ax[3].plot(range(0,200), solutions[3][:200], label='Smoothness 20')
ax[3].set_title("Smoothness 20")
solutions = []
for s in [0, 5, 10, 20]:
solutions.append(GProblem(2000, 2, 2, 2, s).solution)
ax[0].plot(range(0,200), solutions[0][:200], label='Smoothness 0')
ax[1].plot(range(0,200), solutions[1][:200], label='Smoothness 5')
ax[2].plot(range(0,200), solutions[2][:200], label='Smoothness 10')
ax[3].plot(range(0,200), solutions[3][:200], label='Smoothness 20')
solutions = []
for s in [0, 5, 10, 20]:
solutions.append(GProblem(2000, 2, 2, 2, s).solution)
ax[0].plot(range(0,200), solutions[0][:200], label='Smoothness 0')
ax[1].plot(range(0,200), solutions[1][:200], label='Smoothness 5')
ax[2].plot(range(0,200), solutions[2][:200], label='Smoothness 10')
ax[3].plot(range(0,200), solutions[3][:200], label='Smoothness 20')
fig.savefig("Figure1.png")
```
# Grim et al - replication runs - main
500 replication runs were conducted on GCE, using [pyscript2GCE](https://github.com/LukasWallrich/pyscript2gce-production). See `run_simulation_simple.py` for the code used.
```
# Analysis of results
import pickle
with open("Grimmodel_results.pkl",'rb') as f:
res = pickle.load(f)
def prep_outputs(res):
if "solution" in res.columns:
res.rename({"solution": "best_solution"}, inplace=True)
res = pd.concat([res.drop(["best_solution"], axis = 1), res.best_solution.apply(pd.Series).add_suffix("_solution")], axis = 1)
def renamer(col, prefix):
if col.endswith("agent"):
return col
else:
return prefix + col
res_random = res.agent_descriptives.apply(pd.Series).random.apply(pd.Series).rename(mapper = partial(renamer, prefix = "random_"), axis = "columns")
res_best = res.agent_descriptives.apply(pd.Series).best.apply(pd.Series).rename(mapper = partial(renamer, prefix = "best_"), axis = "columns")
res = pd.concat([res.drop(["agent_descriptives"], axis=1), res_best, res_random[res_random.columns[pd.Series(res_random.columns).str.startswith('random_')]]], axis=1)
res["run_id"] = res.reset_index().index
res = res.rename(columns={"best_agent": "top_agent"})
res["gap"] = res["random_solution"] - res["best_solution"]
return(res)
res = prep_outputs(res)
```
## Reproducing Figure 2
```
gaps = res[["smoothness", "gap"]].groupby(["smoothness"]).describe().loc[:, (slice(None), ["mean", "std"])]
gaps.columns = gaps.columns.droplevel(0)
gaps.reset_index(inplace=True)
fig, ax = plt.subplots()
tickpos = range(0, 21, 2)
ax.plot(gaps["smoothness"], gaps["mean"])
ax.fill_between(gaps["smoothness"], gaps["mean"] - gaps["std"], gaps["mean"] + gaps["std"], alpha=0.35)
ax.axhline(y=0, color='grey', linestyle='--')
ax.set_xticks(tickpos)
ax.annotate('Shaded region \n is +/- 1 SD',
xy=(10, .01), xycoords='data',
xytext=(50, 30), textcoords='offset points',
arrowprops=dict(arrowstyle = "->"),
horizontalalignment='right', verticalalignment='bottom')
fig.figure.savefig("Fig2.png")
```
Out of interest - the following replicates the table from Hong & Page. Key insight: overall performance declines with higher smoothness.
```
#Pivot so that random and best groups can be easily compared
col_names = res.columns.values.tolist()
def check_var(col_name):
return not(col_name.find("random_") != -1 or col_name.find("best_") != -1)
id_cols = list(filter(check_var, col_names))
out = pd.melt(res, id_cols)
out = out.join(out.variable.str.split("_", expand = True)).rename(columns={0:"team_type"}).pivot_table(index=id_cols + ["team_type"], columns=[1], values="value").reset_index()
out["NPdiversity"] = out["NPdiversity"] * 100 #Convert to percentages
tbl = out[["team_type", "smoothness", "solution", "NPdiversity"]].groupby(["smoothness", "team_type"]).describe().loc[:, (slice(None), ["mean", "std"])]
tbl = tbl.round(2)
sol = pd.DataFrame(tbl[('solution', 'mean')].astype(str) + " (" + tbl[('solution', 'std')].astype(str) + ")")
sol.columns = ["Solution"]
div = pd.DataFrame(tbl[('NPdiversity', 'mean')].astype(str) + " (" + tbl[('NPdiversity', 'std')].astype(str) + ")")
div.columns = ["Diversity"]
sol = sol.join(div)
sol
```
# Replication of tournament results
The comparisons between best and random groups under tournament setting was only tested with 100 runs but with l ranging up to 30. This already increased computational expense >5x compared to the range considered in the original paper.
```
import os
import pickle
#directory = os.fsencode(".")
res = []
for file in os.listdir("."):
filename = os.fsdecode(file)
if filename.startswith("GrimSweepTournament"): #Multiple files from use of spot VMs
with open(filename,'rb') as f:
res.append(pickle.load(f))
continue
else:
continue
res = pd.concat(res)
## Beware: this cell is very slow (~90 seconds)
def renamer(col, prefix):
if col.endswith("agent"):
return col
else:
return prefix + col
res = pd.concat([res.drop(["best_solution"], axis = 1), res.best_solution.apply(pd.Series).add_suffix("_solution")], axis = 1)
res_random = res.agent_descriptives.apply(pd.Series).random.apply(pd.Series).rename(mapper = partial(renamer, prefix = "random_"), axis = "columns")
res_best = res.agent_descriptives.apply(pd.Series).best.apply(pd.Series).rename(mapper = partial(renamer, prefix = "best_"), axis = "columns")
res = pd.concat([res.drop(["agent_descriptives"], axis=1), res_best, res_random[res_random.columns[pd.Series(res_random.columns).str.startswith('random_')]]], axis=1)
res["run_id"] = res.reset_index().index
res.rename(columns={"best_agent": "top_agent"}, inplace=True)
res["tournament_gap"] = res["tournament_random_solution"] - res["tournament_best_solution"]
res["relay_gap"] = res["relay_random_solution"] - res["relay_best_solution"]
res
comps = res.groupby(["l", "smoothness"]).apply(lambda s: pd.Series({
"tournament_random_advantage": s["tournament_gap"].mean(),
"relay_random_advantage": s["relay_gap"].mean(),
"tournament_advantage_random": (s["tournament_random_solution"] - s["relay_random_solution"]).mean(),
"tournament_advantage_best": (s["tournament_best_solution"] - s["relay_best_solution"]).mean()
}))
win_comps = pd.DataFrame([
{"Strategy":"Relay", "Win rate (%)": (comps["relay_random_advantage"]>0).mean().round(3) * 100,
"Mean advantage": comps["relay_random_advantage"].mean().round(2),
"random_mean": comps["relay_random_advantage"][comps["relay_random_advantage"]>0].mean().round(2),
"random_max": comps["relay_random_advantage"][comps["relay_random_advantage"]>0].max().round(2),
"best_mean": abs(comps["relay_random_advantage"][comps["relay_random_advantage"]<0].mean().round(2)),
"best_max": abs(comps["relay_random_advantage"][comps["relay_random_advantage"]<0].min().round(2))},
{"Strategy":"Tournament", "Win rate (%)": (comps["tournament_random_advantage"]>0).mean().round(3) * 100,
"Mean advantage": comps["tournament_random_advantage"].mean().round(2),
"random_mean": comps["tournament_random_advantage"][comps["tournament_random_advantage"]>0].mean().round(2),
"random_max": comps["tournament_random_advantage"][comps["tournament_random_advantage"]>0].max().round(2),
"best_mean": abs(comps["tournament_random_advantage"][comps["tournament_random_advantage"]<0].mean().round(2)),
"best_max": abs(comps["tournament_random_advantage"][comps["tournament_random_advantage"]<0].min().round(2))}])
win_comps.to_latex("win_comparisons.tex")
win_comps
import seaborn as sns; sns.set_theme()
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(8,2.8))
cbar_ax = fig.add_axes([.95, .2, .02, .6])
relay = comps[["relay_random_advantage"]].reset_index().pivot("l", "smoothness", "relay_random_advantage")
relay_neg = (relay >= 0).replace([True, False], ["", "-"])
cmap = sns.diverging_palette(150, 275, s=80, l=55, as_cmap=True)
sns.heatmap(relay, cmap = cmap, center = 0, annot = relay_neg, fmt = '', ax = ax1, vmin = -.7, vmax = 3.4, cbar_ax=cbar_ax)
ax1.invert_yaxis()
ax1.plot([2,9], [0,27])
ax1.set_title("A. Relay mode", fontweight='bold')
ax1.set_ylabel("Max step length (l)", fontweight='bold')
ax1.set_xlabel("Smoothness", fontweight='bold')
tourn = comps[["tournament_random_advantage"]].reset_index().pivot("l", "smoothness", "tournament_random_advantage")
tourn_neg = (tourn >= 0).replace([True, False], ["", "-"])
sns.heatmap(tourn, cmap = cmap, center = 0, annot = tourn_neg, fmt = '', ax = ax2, vmin = -.7, vmax = 3.4, cbar_ax=cbar_ax)
ax2.invert_yaxis()
ax2.plot([2,10], [0,16])
ax2.plot([10,16], [16,28], linestyle='dashed')
ax2.set_title("B. Tournament mode", fontweight='bold')
ax2.set_ylabel("Max step length (l)", fontweight='bold')
ax2.set_xlabel("Smoothness", fontweight='bold')
fig.figure.savefig("Fig69.png")
fig.figure.savefig("../manuscript/Fig69.png", bbox_inches='tight')
# Tournaments vs relays
pd.DataFrame([
{"Group":"Random", "Tournament wins (%)": (comps["tournament_advantage_random"]>0).mean().round(3) * 100,
"Mean advantage": comps["tournament_advantage_random"].mean().round(2)},
{"Group":"Best", "Tournament wins (%)": (comps["tournament_advantage_best"]>0).mean().round(3) * 100,
"Mean advantage": comps["tournament_advantage_best"].mean().round(2)}])
```
| github_jupyter |
# Homework 4 - Reinforcement Learning in a Smart Factory
Optimization of the robots route for pick-up and storage of items in a warehouse:
1. Implement a reinforcement-learning based algorithm
2. The robot is the agent and decides where to place the next part
3. Use the markov decision process toolbox for your solution
4. Choose the best performing MDP
```
#!pip install pymdptoolbox
## Imports
import mdptoolbox
import pandas as pd
import itertools as it
import numpy as np
import pickle
import time
from scipy import sparse
```
## Import data
```
file_path = 'Exercise 4 - Reinforcement Learning Data - warehousetraining.txt'
file_path_test= 'Exercise 4 - Reinforcement Learning Data - warehouseorder.txt'
# Name the data colums corectly
data = pd.read_csv(file_path, sep='\t', names=["action", "color_state"])
test_data = pd.read_csv(file_path_test, sep='\t', names=["action", "color_state"])
#print(data.info()) print(data.dtypes)
data.head()
data.groupby(["action", "color_state"]).count()
actions = list(np.unique(data.action)) #['restore', 'store']
item_colors = list(np.unique(data.color_state)) #['blue' 'red' 'white']
train_data = np.array( [[actions.index(v[0]), item_colors.index(v[1])] for v in data.values] , dtype=int)
```
## Reinforcement-learning based algorithm: Markov Descision Process (MDP)
A MDP is a discrete time stochastic control process where the markov property is satisfied
1. Create Transitions Matrix represeting the probabilities to lead from state *s0* another state *s1* within the action *a*
2. Generate Reward Matrix defined reward after action *a* in state *s0* to reach state *s1*
Optimize the route with following constraints:
- Size of warehouse is {1..3} x {1..3}
- Separate start/stop position outside the 3x3 storage space where the robot have to go at the end of storage and pick-up
- The first position the robot can move into is always (1,1)
- Robot can move to adjacent fields
- Robot cannot move diagonally
- There are three types of items: (white, blue, red)
```
# Set Markov Descision Process (MDP) Constrains
warehouse_size = [2,2] #2x2 grid
grid_size = np.prod(warehouse_size)
grids_cells = [(i+1,j+1) for j in range(warehouse_size[1]) for i in range(warehouse_size[0]) ]
# The actions is equal to grid size
actions_moves = grid_size.copy()
items = len(item_colors) + 1 # Consider also no item
moves = len(actions)*len(item_colors)
#Total posibilities of item in any satate on the field
items_grid = items ** grid_size
total_states = items_grid * moves
print("The total number of states is: ", total_states)
item_states_ID = dict((k,v) for v,k in enumerate( ["noitem"] + item_colors ))# dict.fromkeys(item_colors + ["noitem"], 0)
item_states_ID
# Create all the posible states indexing
def compute_grid_index(grid_size, item_states_ID):
grids = [s for s in it.product(item_states_ID.values(), repeat=grid_size)]
return np.array(grids)
grids = compute_grid_index(grid_size, item_states_ID)
print("Number of posible states: ", len(grids))
grid_states= pd.DataFrame(data=grids, columns=grids_cells)
grid_states[20:30]
def generate_warehosue_states(grid_states, item_states_ID,):
warehouse_states = pd.DataFrame()
for k,v in item_states_ID.items():
warehouse_states[k] = np.sum(grid_states ==v, axis =1)
return warehouse_states
warehouse_states = generate_warehosue_states(grid_states, item_states_ID)
warehouse_states[20:30]
```
### Transition Probability Matrix (action, state, next state)
```
def create_TPM(data, grids):
# Initialize TMP with shape (action, posible states, posible states)
P = np.zeros(( actions_moves, total_states, total_states),dtype=np.float16)
# Compute Each action probability as the count of each action on the data
move_action_probability = np.array([a*c for a in data["action"].value_counts() / len(data) for c in data["color_state"].value_counts() / len(data) ])
for action in range(actions_moves):
idx = 0
for mov in range(moves):
for s in grids:
for m in range(moves):
if m >= (moves//2): # restore actions
i = ((idx % items_grid) - (items**(actions_moves - action - 1) * (mov+1))) + (items_grid * m)
else:
i = ((idx % items_grid) - (items**(actions_moves - action - 1) * (mov+1))) + (items_grid * m)
P[action][idx][i] = move_action_probability[m]
idx += 1
return P
TMP = create_TPM(data, grids)
def create_rewards(moves, total_states, grid_states):
distances = [sum(np.array(c) - np.array(grids_cells[0])) for c in grids_cells]
rewards = dict(keys=grids_cells, values =distances )
R = np.zeros((actions_moves, total_states, ))
for action in range(actions_moves):
for idx, s in grid_states.iterrows():
next_state = idx//(len(grid_states)//moves)
try:
if(next_state < (moves//len(actions)) and s[action] == 0):
reward = rewards[str(s)]
elif (next_state > (moves//len(actions) ) and (s[action] == (next_state - len(actions)))):
reward = 10000*rewards[str(s)] #+=100
# Invalid movements
else:
reward = -10000
R[action][idx] = reward
except:
pass
return np.asarray(R).T
R = create_rewards(moves, total_states, grid_states)
assert TMP.shape[:-1] == R.shape[::-1], "The actions and states should match"
discount = 0.9
max_iter = 750
policy = mdptoolbox.mdp.PolicyIteration(TMP, R, 0.9, max_iter=max_iter)
value = mdptoolbox.mdp.ValueIteration(TMP, R, 0.9, max_iter=max_iter)
value.run()
policy.run()
p = policy.policy
iterations = policy.iter
print("Policy iterations:", iterations)
print("Value iterations:", value.iter)
```
| github_jupyter |
# 初始化模块
```
import re
import numpy as np
import pandas as pd
import pymongo
import requests
import ipywidgets
from IPython import display
from bokeh import charts
from bokeh.io import push_notebook, show, output_notebook
output_notebook()
client = pymongo.MongoClient()
db = client['hfut']
```
# 数据分析
## 数据总量
```
print('专业记录', db['major'].count(), '条')
print('学期记录', db['term'].count(), '条')
print('课程记录', db['course'].count(), '条')
print('专业计划记录', db['plan'].count(), '条')
print('教学班记录', db['class'].count(), '条')
print('学生记录', db['student'].count(), '条')
print('教学班-学生关系记录', db['class_student'].count(), '条')
# http://bokeh.pydata.org/en/latest/docs/user_guide/charts.html
student_df = pd.DataFrame(list(db['student'].find()), columns=['学号', '姓名', '性别'])
student_df['入学年份'] = student_df['学号'] // 1000000
# http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.groupby.html
_ = student_df.groupby(student_df['入学年份']).apply(lambda df: df.groupby(df['性别']).count())
_ = _.unstack()['学号']
_['比例'] = _['男'] / _['女']
_['合计'] = _['男'] + _['女']
_
# http://bokeh.pydata.org/en/latest/docs/reference/charts.html#bar
p = charts.Bar(student_df,label='入学年份', values='性别', agg='count', stack='性别')
show(p)
```
## 学生数量及人数分布
```
class StudentQueryPanel:
def __init__(self):
self.code_text = ipywidgets.Text(description='学号:')
self.code_text.on_submit(self.handle_text_submit)
self.name_text = ipywidgets.Text(
disabled=True, value='无', description='姓名:')
self.image = ipywidgets.Image(format='jpg', width=260)
self.html = ipywidgets.HTML(description='已选课程班级')
self.panel = ipywidgets.VBox(
[self.image, self.name_text, self.code_text, self.html])
display.display(self.panel)
def handle_text_submit(self, t):
# 先重置
self.image.value = b''
self.name_text.value = '无'
self.html.value = '<b>没有查询到信息</b>'
student_code = t.value
student = None
if re.match(r'201[2-6]21\d{4}', student_code):
student_code = int(student_code)
student = db['student'].find_one(filter={'学号': student_code})
if student:
self.image.value = requests.get(
'http://222.195.8.201/student/photo/{:d}/{:d}.JPG'.format(
student_code // 1000000, student_code)).content
name = student['姓名']
if student['性别'] == '女':
name += '*'
self.name_text.value = name
classes = db['class_student'].find(filter={'学号': student_code})
query = list(map(lambda v: { k: v[k] for k in ('学期代码', '课程代码', '教学班号')}, classes))
classes = list(db['class'].find({
'$or': query
}).sort([('学期代码', pymongo.DESCENDING), ('课程代码', pymongo.DESCENDING)]))
classes_df = pd.DataFrame(
classes,
columns=[
'学期代码', '课程代码', '课程名称', '教学班号', '校区', '时间地点', '起止周',
'开课单位', '任课老师', '优选范围', '性别限制', '禁选范围', '课程类型', '考核类型',
'学分', '班级容量', '选中人数', '备注'
])
self.html.value = classes_df._repr_html_()
panel = StudentQueryPanel()
db['class'].find().sort
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import yaml
import sys
sys.path.insert(0,'/Users/ruhl/code/jbolo/python/')
from jbolo_funcs import *
!ls yamls
FIXED_PSAT = True
def pwv_vary(yamlfile,site,def_pwv):
#expt_yaml = 'yamls/SAT_LFMF_20211210.yaml'
sim = yaml.safe_load(open(yamlfile))
# overload yamls, so we can easily change where a telescope is sited.
sim['sources']['atmosphere']['site']=site
sim['sources']['atmosphere']['pwv']=def_pwv #used to find Psats if not specified, in initial run.
#sim['optical_elements']['window']['thickness']=0.02
chlist = list(sim['channels'].keys())
if FIXED_PSAT:
if (sim['bolo_config']['psat_method'] != 'specified'):
run_optics(sim)
run_bolos(sim)
sim['bolo_config']['psat_method'] = 'specified'
for ch in chlist:
sim['channels'][ch]['psat']=sim['outputs'][ch]['P_sat']
out_NET = {}
out_Popt = {}
for ch in chlist:
out_NET[ch] = np.array([])
out_Popt[ch]= np.array([])
out_pwv = np.arange(0,5000,100)
for pwv in out_pwv:
sim['sources']['atmosphere']['pwv'] = pwv
run_optics(sim)
run_bolos(sim)
for ch in chlist:
out_NET[ch] = np.append(out_NET[ch],sim['outputs'][ch]['NET_NC_total'])
out_Popt[ch] = np.append(out_Popt[ch],sim['outputs'][ch]['P_opt'])
return(out_pwv, out_Popt, out_NET,chlist)
%matplotlib inline
plt.rcParams.update({'font.size': 18})
plt.rcParams['figure.figsize'] = [16, 6]
yamlfile = 'yamls/SAT_LFMF_20220216.yaml'
fig, axs = plt.subplots(1,2)
# check whether sim exists, and if it does, clear it so we don't get confused.
if "sim" in locals():
sim.clear()
pwv, Popt_chile, NET_chile, chlist = pwv_vary(yamlfile, 'Atacama',993)
pwv, Popt_pole, NET_pole, chlist = pwv_vary(yamlfile, 'Pole',321)
chcolor = {'LF_1':'r', 'LF_2':'m', 'MF_1_1':'g', 'MF_2_1':'g', 'MF_1_2':'k', 'MF_2_2':'k', 'HF_1':'c','HF_2':'b'}
for ch in chlist:
ltype = chcolor[ch]
labelstr = 'Chile, '+ch
axs[0].plot(pwv,Popt_chile[ch]*1e12, ltype,label=labelstr)
axs[1].plot(pwv,NET_chile[ch]*1e6, ltype,label=labelstr)
ltype = ltype+'--'
labelstr = 'Pole, '+ch
axs[0].plot(pwv,Popt_pole[ch]*1e12,ltype, label=labelstr)
axs[1].plot(pwv,NET_pole[ch]*1e6,ltype, label = labelstr)
axs[0].set_xlabel('pwv (mm)')
axs[0].set_ylabel('Popt (pW)')
#axs[0].legend(fontsize=10,loc='upper right')
axs[1].set_xlabel('pwv (mm)')
axs[1].set_ylabel('NET (uKrtsec)')
axs[1].legend(fontsize=10,loc='upper right')
#axs[1].text(500,500,'hey')
```
# pwv histograms
```
#### Sort the pwv's into a single time-ordered vector and plot histogram;
# This lets you plot the timestream if desired.
import pickle
pwvs = np.array([])
d = pickle.load(open("atmos/Merra2_atmos_atacama.pck", "rb"))
for ii in np.arange(0,290):
for month in np.arange(4,12): # skip jan/feb/march
for hour in np.arange(0,24):
pwvs = np.append(pwvs,d[month]['TQV'][hour][ii])
print('Atacama median pwv: {0:5.3f} mm'.format(np.median(pwvs)))
#pwv_bins = np.arange(0,10,0.1)
pwv_bins = pwv/1000
A_chile = plt.hist(pwvs, bins=pwv_bins, histtype='step',density=True, label='Atacama')
pwvs = np.array([])
d = pickle.load(open("atmos/Merra2_atmos_pole.pck", "rb"))
for ii in np.arange(0,290):
for month in np.arange(0,11): # skip december
for hour in np.arange(0,24):
pwvs = np.append(pwvs,d[month]['TQV'][hour][ii])
print('Pole median pwv: {0:5.3f} mm'.format(np.median(pwvs)))
A_pole = plt.hist(pwvs, bins=pwv_bins, histtype='step',density=True, label='Pole')
plt.xlabel('pwv')
plt.ylabel('Probability')
plt.legend()
# Histogram has fewer points than the pwv vector; its binning vector sets the left side of each bin, and right side of last bin.
# Bin centers are offset by a half a bin.
pwv_bin_centers = (A_chile[1][0:-1]+A_chile[1][1:])/2
# Linearly interpolate NET's to those same bin centers, and calculate weights
w_chile = {}
w_pole = {}
w_ratio = {}
cuts = {'LF_1':3,
'LF_2':3,
'MF_1_1':2,
'MF_2_1':2,
'MF_1_2':1.5,
'MF_2_2':1.5,
'HF_1':1.0,
'HF_2':1.0}
for ch in chlist:
cut_level = cuts[ch]
cut_vector = np.where(pwv_bin_centers<cut_level,1,0)
NET_chile_onbins = np.interp(pwv_bin_centers,pwv, NET_chile[ch])
NET_pole_onbins = np.interp(pwv_bin_centers,pwv, NET_pole[ch])
w_chile[ch] = (A_chile[0]*cut_vector)/NET_chile_onbins**2
w_pole[ch] = (A_pole[0]*cut_vector)/NET_pole_onbins**2
w_chile_tot = np.trapz(w_chile[ch],pwv_bin_centers)
w_pole_tot = np.trapz(w_pole[ch],pwv_bin_centers)
w_ratio[ch]=w_chile_tot/w_pole_tot
print(ch)
print(' Chile weight: {0:9.0f}'.format(w_chile_tot))
print(' Pole weight: {0:9.0f}'.format(w_pole_tot))
print(' Chile/Pole weight ratio: {0:6.3f}'.format(w_ratio[ch]))
#Make a plot
plt.rcParams['figure.figsize'] = [16, 4]
n_plots = len(chlist)
n_cols = 2
n_rows = int(np.ceil(n_plots/n_cols))
for ii in np.arange(0,n_plots):
plt.subplot(n_rows,n_cols,ii+1)
ch = chlist[ii]
label1 = 'Chile: '+ch
label2 = 'Pole: '+ch
plt.plot(pwv_bin_centers,w_chile[ch], label = label1)
plt.plot(pwv_bin_centers,w_pole[ch], label = label2)
plt.legend()
for ch in chlist:
if 'LF' in ch:
ntubes = 5
elif 'MF' in ch:
ntubes = 3
elif 'HF' in ch:
ntubes = 2
myvecname = {'vname':'aa'}
aa = np.array([1,2,4])
print(myvecname['vname'])
```
| github_jupyter |
---
_You are currently looking at **version 1.2** of this notebook. To download notebooks and datafiles, as well as get help on Jupyter notebooks in the Coursera platform, visit the [Jupyter Notebook FAQ](https://www.coursera.org/learn/python-social-network-analysis/resources/yPcBs) course resource._
---
# Assignment 3
In this assignment you will explore measures of centrality on two networks, a friendship network in Part 1, and a blog network in Part 2.
## Part 1
Answer questions 1-4 using the network `G1`, a network of friendships at a university department. Each node corresponds to a person, and an edge indicates friendship.
*The network has been loaded as networkx graph object `G1`.*
```
import networkx as nx
G1 = nx.read_gml('friendships.gml')
```
### Question 1
Find the degree centrality, closeness centrality, and normalized betweeness centrality (excluding endpoints) of node 100.
*This function should return a tuple of floats `(degree_centrality, closeness_centrality, betweenness_centrality)`.*
```
def answer_one():
# Your Code Here
degCent = nx.degree_centrality(G1)
closeCent = nx.closeness_centrality(G1)
btwnCent = nx.betweenness_centrality(G1, normalized = True, endpoints = False)
return degCent[100], closeCent[100], btwnCent[100]
```
<br>
#### For Questions 2, 3, and 4, assume that you do not know anything about the structure of the network, except for the all the centrality values of the nodes. That is, use one of the covered centrality measures to rank the nodes and find the most appropriate candidate.
<br>
### Question 2
Suppose you are employed by an online shopping website and are tasked with selecting one user in network G1 to send an online shopping voucher to. We expect that the user who receives the voucher will send it to their friends in the network. You want the voucher to reach as many nodes as possible. The voucher can be forwarded to multiple users at the same time, but the travel distance of the voucher is limited to one step, which means if the voucher travels more than one step in this network, it is no longer valid. Apply your knowledge in network centrality to select the best candidate for the voucher.
*This function should return an integer, the name of the node.*
```
def answer_two():
degree = nx.degree_centrality(G1)
max_node = None
max_degree = -1
for key, value in degree.items():
if value > max_degree:
max_degree = value
max_node = key
return max_node
```
### Question 3
Now the limit of the voucher’s travel distance has been removed. Because the network is connected, regardless of who you pick, every node in the network will eventually receive the voucher. However, we now want to ensure that the voucher reaches the nodes in the lowest average number of hops.
How would you change your selection strategy? Write a function to tell us who is the best candidate in the network under this condition.
*This function should return an integer, the name of the node.*
```
def answer_three():
# Your Code Here
degree = nx.closeness_centrality(G1)
max_node = None
max_degree = -1
for key, value in degree.items():
if value > max_degree:
max_degree = value
max_node = key
return max_node
```
### Question 4
Assume the restriction on the voucher’s travel distance is still removed, but now a competitor has developed a strategy to remove a person from the network in order to disrupt the distribution of your company’s voucher. Your competitor is specifically targeting people who are often bridges of information flow between other pairs of people. Identify the single riskiest person to be removed under your competitor’s strategy?
*This function should return an integer, the name of the node.*
```
def answer_four():
degree = nx.betweenness_centrality(G1)
max_node = None
max_degree = -1
for key, value in degree.items():
if value > max_degree:
max_degree = value
max_node = key
return max_node
```
## Part 2
`G2` is a directed network of political blogs, where nodes correspond to a blog and edges correspond to links between blogs. Use your knowledge of PageRank and HITS to answer Questions 5-9.
```
G2 = nx.read_gml('blogs.gml')
```
### Question 5
Apply the Scaled Page Rank Algorithm to this network. Find the Page Rank of node 'realclearpolitics.com' with damping value 0.85.
*This function should return a float.*
```
def answer_five():
pr = nx.pagerank(G2,alpha=0.85)
return pr['realclearpolitics.com']
```
### Question 6
Apply the Scaled Page Rank Algorithm to this network with damping value 0.85. Find the 5 nodes with highest Page Rank.
*This function should return a list of the top 5 blogs in desending order of Page Rank.*
```
import operator
def answer_six():
pr = nx.pagerank(G2,alpha=0.85)
pr_list = sorted(pr.items(), key=operator.itemgetter(1), reverse=True)[0:5]
ans = []
for i in pr_list:
ans.append(i[0])
return ans
```
### Question 7
Apply the HITS Algorithm to the network to find the hub and authority scores of node 'realclearpolitics.com'.
*Your result should return a tuple of floats `(hub_score, authority_score)`.*
```
def answer_seven():
hits = nx.hits(G2)
node = 'realclearpolitics.com'
print (len(hits))
return hits[0][node], hits[1][node]
```
### Question 8
Apply the HITS Algorithm to this network to find the 5 nodes with highest hub scores.
*This function should return a list of the top 5 blogs in desending order of hub scores.*
```
def answer_eight():
hits = nx.hits(G2)
hubs = hits[0]
return sorted(hubs.keys(), key=lambda key:hubs[key], reverse=True)[:5]
```
### Question 9
Apply the HITS Algorithm to this network to find the 5 nodes with highest authority scores.
*This function should return a list of the top 5 blogs in desending order of authority scores.*
```
def answer_nine():
hits = nx.hits(G2)
authorities = hits[1]
return sorted(authorities.keys(), key=lambda key:authorities[key], reverse=True)[:5]
```
| github_jupyter |
## Greedy algorithm for Jobs Scheduling Problem
Given a sequence of jobs, each has a weight `w` (priority) and a length `l`, in what order should we sequence the jobs such that the weighted sum of completion times is minimized?
```
def schedule_jobs(jobs):
"""
input: a sequence of jobs in a list of tuples [(weight1, length1), (weight2, length2), ...]
output: a number "WSCT" = weighted sum of completion times,
an optimal job sequence "SEQ" = [(job, score), (job, score), ...]
time: O(nlogn), just need to sort by greedy score
"""
scores = { index:(job[0] / job[1]) for index, job in enumerate(jobs) }
SEQ = sorted(scores.items(), key=lambda x: x[1], reverse=True)
cc = 0 # completion time
WSCT = 0
for index, score in SEQ:
wt, lt = jobs[index]
cc += lt
WSCT += wt * cc
return WSCT, SEQ
jobs = [(5,2), (2,1), (8,8), (6,4), (9,3), (12,3)]
scores = { index:(job[0] / job[1]) for index, job in enumerate(jobs) }
sorted(scores.items(), key=lambda x: x[1], reverse=True)
WSCT, SEQ = schedule_jobs(jobs)
print("WSCT:", WSCT)
print("SEQ:", SEQ)
jobs = []
with open("support/jobs.txt", "rt") as fh:
for line in fh.readlines():
line = line.strip().split()
if len(line) == 2:
jobs.append((int(line[0]), int(line[1])))
WSCT, SEQ = schedule_jobs(jobs)
print("WSCT:", WSCT)
```
## LRU Cache Algorithm - Least Recently Used Cache for Redis/operating systems
### Doubly Linked List Class
```
class Node:
def __init__(self, data=None):
self.data = data
self.prev = None
self.next = None
class DoubleLinkList:
def __init__(self):
self.head = Node() # null pointer
self.tail = Node() # null pointer
self.head.next = self.tail
self.tail.prev = self.head
self._size = 0
@property
def size(self):
return self._size
def __str__(self):
curr = self.head.next
out = "[head,"
while curr.data != None:
out += (str(curr.data) + ',')
curr = curr.next
return out + "tail]"
__repr__ = __str__
def __contains__(self, node):
curr = self.head.next
while curr.data != None:
if id(curr) == id(node):
return True
curr = curr.next
return False
def append(self, node, pos="tail"):
''' append to the head/tail of the list, takes O(1) time '''
if pos == "tail":
self.tail.prev.next = node
node.prev = self.tail.prev
node.next = self.tail
self.tail.prev = node
elif pos == "head":
node.next = self.head.next
self.head.next.prev = node
self.head.next = node
node.prev = self.head
else:
return
self._size += 1
def delete(self, node):
''' delete the node in the list, takes O(1) time '''
l = node.prev
r = node.next
l.next = r
r.prev = l
self._size -= 1
def pop(self):
""" remove and return the last node in the list, takes O(1) time.
pop() should not have any arguments, otherwise it's a delete()
"""
node = self.tail.prev
node.prev.next = self.tail
self.tail.prev = node.prev
self._size -= 1
return node
def clear(self):
return self.__init__()
# l1 = SingleLinkList()
l2 = DoubleLinkList()
for data in range(10):
l2.append(Node(data))
print(l2)
Bob = Node('Bob')
l2.append(Bob)
l2
Alice = Node('Alice')
l2.append(Alice, pos="head")
l2
(Alice in l2) & (Bob in l2)
l2.delete(Alice)
l2
Alice in l2
node = l2.pop()
node.data
id(node) == id(Bob)
Bob in l2
l2
l2.size # head/tail are null pointers, not counted in
l2.clear()
l2
```
### LRU Cache Class
https://leetcode.com/problems/lru-cache/
```
class LRUCache:
'''
for fast lookups/inserts/deletes all in O(1), use "hashmap" + "doubly linked list".
(self._cache) is a doubly linked list which stores real data in different nodes.
(self._hashmap) is a dictionary that maps keys to real node objects in (self._cache).
'''
def __init__(self, capacity):
self._hashmap = {}
self._cache = DoubleLinkList()
self._capacity = capacity # maximum # of slots
self._size = 0
@property # read-only attribute
def capacity(self):
return self._capacity
@property # read-only attribute
def size(self):
return self._size
def read(self, key):
node = self._hashmap.get(key, None) # O(1)
if node and node.data:
self._cache.delete(node) # O(1)
self._cache.append(node, pos="head") # O(1)
# if the hashmap {} maps to physical addresses of node objects,
# also need to update the address: self._hashmap[key] = new_address
return node.data
else:
return -1
def write(self, key, value):
node = self._hashmap.get(key, None)
if node and node.data:
self._cache.delete(node)
node.data = value # when attributes of the node updated, the hashmap still maps to this node
self._cache.append(node, pos="head")
else:
new_node = Node(value)
self._hashmap[key] = new_node
if self._capacity <= self._size:
last = self._cache.pop()
# last = None # this will make a new copy of "last", so the mapping will persist
last.data = None # disable the hashmap mapping to this node
self._size -= 1
self._cache.append(new_node, pos="head")
self._size += 1
def view_cache(self):
for key, node in self._hashmap.items():
print(key.ljust(12), ("Node(" + str(node.data) + ")").rjust(12))
print("--------------------")
print(self._cache)
cache = LRUCache(capacity=5)
cache.capacity # read-only attribute, protected data
cache.size
cache.write("Alice", 99)
cache.write("Bob", 70)
cache.write("Carlos", 82)
cache.view_cache()
cache.read("Jack")
cache.read("Alice")
cache.view_cache()
cache.size
cache.write("Jack", 65)
cache.write("James", 79)
cache.view_cache()
cache.write("Sophia", 89) # Bob: 70 will be removed
cache.view_cache()
cache.size
cache.read("Bob")
cache.write("James", 11) # update James's score
cache.view_cache()
```
## Prim's Minimum Spanning Trees Greedy Algorithm
```
import numpy as np
import networkx as nx
import heapq as hq
import matplotlib.pyplot as plt
%matplotlib inline
def prim(graph):
"""
input: a connected undirected graph
output: MST = minimum spanning tree, a subset of the edges collection with minimum sum of edge costs,
MC = minimum total cost
time: O(m*n)
"""
source = list(graph.keys())[0] # randomly choose a source vertex
X = set([source]) # keep track of all vertices visited
T = []
MC = 0
while len(X) != len(graph):
# choose the edge that has the lowest cost
cost = np.inf
edge = None
for u in graph:
for v in graph[u]:
if u in X and v not in X:
if graph[u][v] < cost:
cost = graph[u][v]
edge = (u, v)
if edge:
T.append(edge)
X.add(edge[1])
MC += cost
return T, MC
graph = {
'a': {'b': 1, 'c': 4, 'd': 3},
'b': {'a': 1, 'd': 2},
'c': {'a': 4, 'd': 5},
'd': {'a': 3, 'c': 5, 'b': 2}
}
MST, MC = prim(graph)
MST
WG = nx.Graph()
WG.add_edge('a', 'b', weight=1)
WG.add_edge('b', 'd', weight=2)
WG.add_edge('a', 'c', weight=4)
WG.add_edge('c', 'd', weight=5)
WG.add_edge('a', 'd', weight=3)
pos = nx.fruchterman_reingold_layout(WG)
nx.draw(WG, pos=pos, with_labels=True, node_color="lightgreen")
nx.draw_networkx_edges(WG, pos, edgelist=MST, edge_color="red", style="dotted", width=3)
nx.draw_networkx_edge_labels(WG, pos, edge_labels=nx.get_edge_attributes(WG, 'weight'), font_size=15, rotate=False)
```
## Accelerate Prim's MST Algorithm using a min heap
Helper functions that support lookups/deletes in the heap
```
def search_node(heap, node):
index = 0
for item in heap:
if item[2] == node: # tuple[2] is the end node
return (item[0], item[1], item[2], index)
index += 1
return
def remove_node(heap, node):
_, _, _, index = search_node(heap, node) # 1.find the node's position in the list
heap[index] = heap[-1] # 2.swap with the last element
heap.pop() # 3.remove the last element (list.pop())
hq.heapify(heap) # 4.rearrange the heap
def prim_heap(graph):
"""
input: a connected undirected graph
output: MST = minimum spanning tree, a subset of the edges collection with minimum sum of edge costs,
MC = minimum total cost
time: O(m*logn), in each greedy step we extract-min from the heap, which takes O(logn) = O(logm)
"""
source = list(graph.keys())[0] # randomly choose a source vertex
X = set([source]) # keep track of all vertices visited
T = []
MC = 0
H = [] # initialize the heap V - X
for w in [w for w in graph if w not in X]: # run local tournament for each node w in V - X
min_cost = np.inf
winner = None
for v in graph:
if v in X and w in graph[v]:
cost = graph[v][w]
if cost < min_cost:
min_cost = cost
winner = (v, w)
if winner:
v, w = winner
hq.heappush(H, (min_cost, v, w)) # "w" is unique across the heap
else:
hq.heappush(H, (np.inf, None, w)) # for isolated nodes, key = infinity
assert len(H) == (len(graph) - len(X)), "Incorrect heap size!"
while H:
cost, v, w = hq.heappop(H) # extract-min from the heap 1 by 1, and add to X
X.add(w)
T.append((v, w))
MC += cost
for t in graph[w]: # update keys for nodes in graph[w]
if t not in X:
new_cost = graph[w][t]
cost, _, _, _ = search_node(H, t)
if new_cost < cost:
remove_node(H, t)
hq.heappush(H, (new_cost, w, t))
return T, MC
graph = {} # the undirected graph must be symmetric!
with open("support/edges.txt", "rt") as fh:
for line in fh.readlines():
line = line.strip().split()
if len(line) == 3:
u, v, cost = [int(i) for i in line]
if u not in graph:
graph[u] = {v: cost}
else:
graph[u][v] = cost
if v not in graph:
graph[v] = {u: cost}
else:
graph[v][u] = cost
graph[316] # test
%%timeit
MST, MC = prim(graph)
MC
%%timeit
MST, MC = prim_heap(graph)
MC
```
## Advanced Union-Find for handling disjoint sets
Need to design the data structure like binary search trees.
`find()`: with path compression, update parent in each run, save time for the next call
`union()`: lazy union, update parent only for 1 node, union by rank
For Advanced Union-Find, all operations = nearly constant time = `O(log*(n))`, where `log*(n)` is the number of log() operations required to make n down to <= 1. For practical values of n, `log*(n) ≤ 5`.
```
import networkx as nx
import matplotlib.pyplot as plt
%matplotlib inline
import pygraphviz
from networkx.drawing.nx_agraph import write_dot, graphviz_layout
class UnionFind:
def __init__(self, nodes):
self.parent = {v:v for v in nodes}
self.rank = {v:0 for v in nodes}
self._size = len(nodes) # number of clusters in Union-Find
@property
def size(self):
return self._size
def __str__(self):
out = "number of clusters: " + str(self.size) + "\n"
out += ("all parents: " + str(set(self.parent.values())) + "\n")
for v, p in self.parent.items():
out += ("parent(" + str(v) + "): " + str(p) + "\n")
return out
__repr__ = __str__
def find(self, v):
''' worst case O(logn)-time '''
if self.parent[v] == v:
return v
self.parent[v] = self.find(self.parent[v]) # path compression
return self.parent[v]
def union(self, u, v):
''' worst case O(logn)-time '''
if u == v:
return
pu, pv = self.find(u), self.find(v)
ru, rv = self.rank[pu], self.rank[pv]
if pu == pv:
print(u, "and", v, "are already unioned!")
return
if ru <= rv:
self.parent[pu] = pv
else:
self.parent[pv] = pu
if ru == rv:
self.rank[pv] += 1
self._size -= 1
UF = UnionFind(range(10))
UF
UF.union(3, 4)
UF
UF.union(4, 9) # {4,3} is the big tree, {9} is the small tree
UF
UF.union(4, 9)
for i in range(10):
print(UF.find(i), end=" ")
UF.size
for u, v in [(5,9),(7,3),(4,8),(6,1)]:
UF.union(u, v)
```
For easy visualization, let's plot:
```
G = nx.DiGraph({4:[3,5,7,8,9], 1:[6], 0:[], 2:[]})
plt.title('UF at this point', fontsize=16)
pos = graphviz_layout(G, prog='dot')
nx.draw_networkx(G, pos, with_labels=True, arrows=True, node_color="lightgreen")
nx.draw_networkx_nodes(G, pos, nodelist=[0,1,2,4], node_color="#E066FF", node_size=1000)
UF
UF.find(8)
for u, v in [(6,0),(2,1),(5,6)]:
UF.union(u, v)
UF.size
UF
G = nx.DiGraph({4:[3,5,7,8,9], 1:[0,2,4,6]})
plt.title('After the union, final UF looks like this', fontsize=16)
pos = graphviz_layout(G, prog='dot')
nx.draw_networkx(G, pos, with_labels=True, arrows=True, node_color="lightgreen")
nx.draw_networkx_nodes(G, pos, nodelist=[1], node_color="#E066FF", node_size=1000)
UF.rank # rank(v) = the maximum distance from v to a leaf
```
## Kruskal's MST Algorithm using Advanced Union-Find
```
import networkx as nx
import matplotlib.pyplot as plt
%matplotlib inline
G = {
'a': {'b': 1, 'c': 7},
'b': {'a': 1, 'c': 5, 'd': 4, 'e': 3},
'c': {'a': 7, 'b': 5, 'e': 6},
'd': {'b': 4, 'e': 2},
'e': {'b': 3, 'c': 6, 'd': 2}
}
def kruskal(G):
"""
input: a connected undirected graph
output: MST = minimum spanning tree, MC = minimum total cost
time: O(m*alpha(n)), where alpha(n) is the inverse Ackerman function
"""
T = set()
E = [(u,v,G[u][v]) for u in G for v in G[u]]
MC = 0
E = sorted(E, key=lambda x: x[2]) # sort edges in order of increasing cost
UF = UnionFind(list(G.keys()))
for u, v, cost in E:
if UF.find(u) != UF.find(v): # if u & v already connected, union (u,v) will make a cycle
T.add((u, v))
UF.union(u, v)
MC += cost
return T, MC
MST, MC = kruskal(G)
MST
MC
E = [('a','b',1),('a','c',7),('b','c',5),('b','d',4),('c','e',6),('d','e',2),('b','e',3)]
WG = nx.Graph()
for u, v, w in E:
WG.add_edge(u, v, weight=w)
pos = nx.fruchterman_reingold_layout(WG)
edgelist = [('a','b'),('b','c'),('b','e'),('d','e')]
nx.draw(WG, pos=pos, with_labels=True, node_color="lightgreen")
nx.draw_networkx_edges(WG, pos, edgelist=edgelist, edge_color="blue", style="dotted", width=3)
nx.draw_networkx_edges(WG, pos, edgelist=[('b','d')], edge_color="red", style="dashed", width=3)
nx.draw_networkx_edge_labels(WG, pos, edge_labels=nx.get_edge_attributes(WG, 'weight'), font_size=15, rotate=False)
graph = {} # the undirected graph must be symmetric!
with open("support/edges.txt", "rt") as fh:
for line in fh.readlines():
line = line.strip().split()
if len(line) == 3:
u, v, cost = [int(i) for i in line]
if u not in graph:
graph[u] = {v: cost}
else:
graph[u][v] = cost
if v not in graph:
graph[v] = {u: cost}
else:
graph[v][u] = cost
MST, MC = kruskal(graph)
MC # -3612829
```
Congratulations~! this result is exactly the same as we computed previously, correctness proved!
In theory, __Kruskal's + Union-Find__ is as fast as __Prim's + Min-Heap__.
## Greedy Max-spacing k-clustering using Advanced Union-Find
```
G = {}
with open("support/clustering1.txt", "rt") as fh: # make sure G is symmetric
for line in fh.readlines():
line = line.strip().split()
if len(line) == 3:
u, v, d = [int(i) for i in line]
if u in G:
G[u][v] = d
else:
G[u] = {v:d}
if v in G:
G[v][u] = d
else:
G[v] = {u:d}
def k_cluster(G):
'''
input: a weighted graph where weight is the Euclidean distance between 2 nodes
output: the minimum spacing between clusters after k-clustering
'''
E = [(u,v,G[u][v]) for u in G for v in G[u]]
E = sorted(E, key=lambda x: x[2]) # sort by increasing distances
UF = UnionFind(list(G.keys()))
for u, v, d in E:
if UF.find(u) == UF.find(v):
continue
if UF.size > 4:
UF.union(u, v)
else:
return d
MMS = k_cluster(G)
MMS
```
## Huffman codes
```
import heapq as hq
def huffman(charset, freq):
"""
input: a character set such as ['A','B','C','D','E'] where chars in the set are distinct,
a list of frequencies for each char such as [80, 50, 40, 60, 10].
output: an optimal dict of variable-length binary codes that minimizes the average encoding length,
the output encodings are ensured to be prefix-free, there's no ambiguity when decode.
note that the optimal solution is not unique, this function returns only one of them.
time: O(nlogn) if using a heap to extract-min, or O(n) if using two queues(non-trivial)
remark: the min-heap H is a list of tuples, such as [(12,['A']), (5,['B','C']), ...],
inside the tuple, we use a list ['A','B','C', ...] to store characters in the charset.
but why bother using a list inside the tuple? why not use (12,'A'), (5,'BCD'), etc?
well, of course you can just concatenate characters this way, but then, this function
would only work for single-digit charset like ['0','1',...] and single-char English charset
such as ['a','b','c',...], but it would fail for multi-digits or multi-chars charset such
as ['0','10','200'], ['Aa','b','CDe'], obviously, it would also fail for non-English
charsets like ['哈','夫','曼','码'] or ['ハ','フ','マ','ン'], for example.
"""
T = {char:'' for char in charset}
H = list(zip(freq, [[char] for char in charset]))
hq.heapify(H)
while len(H) > 1:
f1, char1 = hq.heappop(H)
f2, char2 = hq.heappop(H)
hq.heappush(H, (f1 + f2, char1 + char2))
for char in char1:
T[char] = '0' + T[char]
for char in char2:
T[char] = '1' + T[char]
return T
huffman(['A','B','C','D','E','F'], [3,2,6,8,2,6])
```
### compress/unzip and encode/decode personal data
```
def huffman_encode(data, encoding):
''' mask human-readable data as binary codes 0/1 '''
mask = ''
for char in data:
mask += encoding[char]
return mask
def huffman_decode(data, encoding):
''' translate binary codes into strings based on encoding '''
translate = {v: k for k, v in encoding.items()}
unmask = ''
curr_code = ''
for digit in data:
curr_code += digit
if curr_code in translate:
unmask += translate[curr_code]
curr_code = ''
return unmask
import string
import numpy as np
string.punctuation
string.ascii_letters
string.digits
valid_chars = string.punctuation + string.ascii_letters + string.digits
valid_chars
charset = [' ', '\n']
for char in valid_chars:
charset.append(char)
print(charset)
freq = np.random.randint(0, 1000, size=len(charset))
freq
encoding = huffman(charset, freq)
encoding
```
__text source__: https://en.wikipedia.org/wiki/Huffman_coding
```
text = '''
In computer science, a Huffman code is a particular type of optimal prefix code that is commonly used for lossless data compression. The process of finding and/or using such a code proceeds by means of Huffman coding, an algorithm developed by David A. Huffman while he was a Sc.D. student at MIT, and published in the 1952 paper "A Method for the Construction of Minimum-Redundancy Codes". The output from Huffman's algorithm can be viewed as a variable-length code table for encoding a source symbol. The algorithm derives this table from the estimated probability or frequency of occurrence (weight) for each possible value of the source symbol. However, although optimal among methods encoding symbols separately, Huffman coding is not always optimal among all compression methods.
'''
masked_data = huffman_encode(text, encoding)
masked_data
original_data = huffman_decode(masked_data, encoding)
print(original_data)
```
### trial test on Chinese character sets
```
charset = ['怎','样','证','明','该','三','次','有','理','数','系','函','的','值','不','可','能','都','是','素']
freq = np.random.randint(0, 100, len(charset))
encoding = huffman(charset, freq)
encoding
text = '怎样证明该三次有理数系数函数的值不可能都是素数'
masked_data = huffman_encode(text, encoding)
masked_data
original_data = huffman_decode(masked_data, encoding)
print(original_data)
```
### Solution for the programming assignment
```
freq = []
with open("support/huffman.txt") as fh:
for line in fh.readlines():
freq.append(int(line.strip()))
n = freq.pop(0)
charset = [str(i) for i in range(n)] # build a charset of size n
encoding = huffman(charset, freq)
encoding_length = [len(code) for code in encoding.values()]
max(encoding_length)
min(encoding_length)
```
## DP - WIS in path graphs
```
def wis(path):
"""
input: weights of the vertices in a path graph stored in a list,
weights are listed in the order in which vertices appear in the path.
output: WIS - maximum-weight independent set, W - maximum weight
remark: here we find WIS = IS[1], because every step only depends on the previous 2 states,
but this shortcut does not work in general, a step often depends on many past states.
in common, to construct the optimal set in DP, we need to trace back the path in reverse order.
"""
# initialize
if path[0] >= path[1]:
MW = [path[0], path[0]]
IS = [[1], [1]]
else:
MW = [path[0], path[1]] # max weight at every index
IS = [[1], [2]] # optimal independent set at the previous 2 indices
for i in range(2, len(path)):
if MW[i - 1] >= MW[i - 2] + path[i]:
MW.append(MW[i - 1])
IS[0] = IS[1] # the (i+1)-th vertex not in WIS
else:
MW.append(MW[i - 2] + path[i])
prev = IS[1]
IS[1] = IS[0] + [i + 1] # the (i+1)-th vertex in WIS
IS[0] = prev
return IS[1], MW[-1]
path = []
with open("support/mwis.txt", "rt") as fh:
for line in fh.readlines():
line = line.strip()
path.append(int(line))
path.pop(0)
WIS, W = wis(path)
WIS[:20]
W # 2955353732
ans = ""
for num in [1, 2, 3, 4, 17, 117, 517, 997]:
ans += str(1 if num in WIS else 0)
print(ans) # 10100110
```
## DP - The Knapsack Problem (discrete, without repetitions)
Here we are talking about the discrete knapsack problem without repetitions.
For fractional knapsack problem, simple greedy algorithm works fine.
### DP implementation for medium size dataset
```
import numpy as np
def knapsack(items, W):
"""
input: items = [(v1, w1), (v2, w2), ...], the k-th item has value "vk" and weight "wk",
W = capacity of the knapsack
output: the optimal subset of items, and the maximum value of that set
time: O(n*W)
"""
n = len(items)
A = np.zeros((W + 1, n + 1), dtype=int)
S = set()
# 1. by recurrence, find the maximum value at each (row, col) position
for w in range(1, W + 1):
for i in range(1, n + 1):
item_i = items[i - 1] # i-th item (1-indexed)
v_i, w_i = item_i[0], item_i[1]
if w - w_i < 0: # i-th item overweight
A[w, i] = A[w, i - 1]
elif A[w, i - 1] >= A[w - w_i, i - 1] + v_i: # item_i not in S
A[w, i] = A[w, i - 1]
else: # item_i in S
A[w, i] = A[w - w_i, i - 1] + v_i
# 2. from the lowest rightmost position, trace back the matrix to find the optimal S
row, col = W, n
while A[row, col] > 0:
v, w = items[col - 1]
c1 = A[row, col - 1] # case1: inherited from the left position, itm not in S
if row < w: # prevent overweight
c2 = 0
else:
c2 = A[row - w, col - 1] + v # case2: derived by adding itm to S
if c1 >= c2:
col -= 1
else:
S.add(col) # 1-base-indexed items
row -= w
col -= 1
print(A) # for debugging
return S, A[W, n] # lowest rightmost element
items = [(3,4),(2,3),(4,2),(4,3)]
W = 6
S, value = knapsack(items, W)
S # optimal subset = {item3, item4}
value # optimal value = 8
items = []
with open("support/knapsack1.txt", "rt") as fh:
for line in fh.readlines():
line = line.strip().split()
value, weight = int(line[0]), int(line[1])
items.append((value, weight))
W, n = items.pop(0) # W = 10000, n = 100
S, value = knapsack(items, W)
value # 2493893
```
__WARNING__: unit test should be followed by stress test
it is extremely important to run the __STRESS TEST__ for your function. Even if you get the correct answer for the homework with your algorithm, it could still be wrong! For a variety of random inputs, please always test your function against the brute-force benchmark before moving to production.
```
items = [(1,1), (2,2), (3,3), (4,4), (5,5), (5,5), (7,7), (7,7), (8,8), (10,10), (12,12), (19,19), (25,25)]
S, value = knapsack(items, 36)
print(S) # {2, 3, 4, 5, 7, 8, 9}
print(value) # 36
```
### Recursive implementation for very large dataset
```
def knapsack_recursive(items, i, w):
""" the previous iterative version takes O(n*W)-time using dynamic programming,
but for large n and W, the recursive version with memoization is much faster.
this is because, the recursive running time is independent of W.
if recurse without memoization, running time ~ O(2^n), exponential growth.
if recurse with memoization, running time ~ O(n^2)?, quadratic growth.
"""
M = {} # use a hash table for memoization, use tuple(i,w) as the key
def recurse(items, i, w):
''' A[w,i] = max(case1, case2 + v_i) = max(A[w,i-1], A[w-w_i,i-1] + v_i)
'''
nonlocal M
if i == 0 or w <= 0: # base case
M[(i, w)] = 0
return M[(i, w)]
item_i = items[i - 1]
v_i, w_i = item_i
if (i - 1, w) not in M: # case1
M[(i - 1, w)] = recurse(items, i - 1, w)
if (i - 1, w - w_i) not in M: # case2
M[(i - 1, w - w_i)] = recurse(items, i - 1, w - w_i)
if w - w_i < 0: # corner case: if overweight, must choose case1
M[(i, w)] = M[(i - 1, w)]
else:
M[(i, w)] = max(M[(i - 1, w)], M[(i - 1, w - w_i)] + v_i)
return M[(i, w)]
return recurse(items, i, w)
items = []
with open("support/knapsack_big.txt", "rt") as fh:
for line in fh.readlines():
line = line.strip().split()
value, weight = int(line[0]), int(line[1])
items.append((value, weight))
W, n = items.pop(0) # W = 2000000, n = 2000, NumPy does not support 2-D array of such large size
import sys, time
sys.setrecursionlimit(3000000)
start_time = time.time()
max_value = knapsack_recursive(items, n, W) # 4243395
end_time = time.time()
print("max_value =", max_value, "computed in", end_time - start_time, "seconds")
```
### Knapsack problem with repetitions (unlimited quantity of each item)
With repetitions, the subproblem and the recurrence relation are much simpler, since the `items` will not change at all. There's only 1 variation dimension - `W`, so we don't need a 2-D matrix, just use simple recursion + memoization.
`value(w) = maximize { value(w - w_i) + v_i }` over each item `i`.
```
def knapsack_repeated(items, capacity):
M = {0:0} # use a hash table for memoization, use w as the key
def recurse(W):
''' value(w) = max{ value(w - w_i) + v_i }
'''
nonlocal M
if W not in M:
for w in range(1, W + 1):
M[w] = 0
for item in items: # allow repetitions
if item[1] <= w:
value = recurse(w - item[1]) + item[0]
if value > M[w]:
M[w] = value
return M[W]
return recurse(capacity)
items = [(30,6),(14,3),(16,4),(9,2)]
knapsack_repeated(items, capacity=10)
```
## DP - Sequence Alignment (Needleman-Wunsch score)
A measure of the shortest distance between two strings.
```
import numpy as np
def needleman_wunsch(str1, str2, alpha_g=1, alpha_m=1):
"""
input: two strings "str1" and "str2",
alpha_g = penalty for a gap,
alpha_m = penalty for a mismatch.
output: print out the optimal alignment of the 2 strings that minimizes the total penalty,
return the corresponding minimum penalty (Needleman-Wunsch score, or the shortest distance)
time: O(m*n), where m, n are the length of the 2 strings
remark: when alpha_g == alpha_m, the optimal alignment might not be unique,
but in practice, a mismatch is more costly than a gap, so when dealing with ties,
we would prefer a gap insertion rather than a mismatch, which also leaves
possibility for potential future matches.
"""
m, n = len(str1), len(str2)
A = np.zeros((m + 1, n + 1), dtype=int) # minimum penalty matrix
S1, S2 = str(), str()
# 0. base case
A[:,0] = np.dot(range(m + 1), alpha_g)
A[0,:] = np.dot(range(n + 1), alpha_g)
# 1. by recurrence, find the minimum penalty at each (row, col) position
for i in range(1, m + 1):
for j in range(1, n + 1):
case1 = A[i - 1, j - 1] + (alpha_m if str1[i - 1] != str2[j - 1] else 0)
case2 = A[i - 1, j] + alpha_g
case3 = A[i, j - 1] + alpha_g
A[i, j] = min(case1, case2, case3)
# 2. trace back the matrix from the lower right corner to reconstruct the optimal alignment
# if "case1" == "case2 or case3", it's a tie, a gap insertion is preferred.
i, j = m, n
while i > 0 or j > 0:
if i == 0:
S1 = '+' + S1
S2 = str2[j - 1] + S2
j -= 1
continue
elif j == 0:
S1 = str1[i - 1] + S1
S2 = '+' + S2
i -= 1
continue
if A[i, j] == A[i - 1, j] + alpha_g: # case2: match str1[i - 1] with a gap
S1 = str1[i - 1] + S1
S2 = '+' + S2
i -= 1
elif A[i, j] == A[i, j - 1] + alpha_g: # case3: match str2[j - 1] with a gap
S1 = '+' + S1
S2 = str2[j - 1] + S2
j -= 1
else: # case1: match str1[i - 1] with str2[j - 1]
S1 = str1[i - 1] + S1
S2 = str2[j - 1] + S2
i -= 1
j -= 1
print(A) # for debugging
print(S1)
print(S2)
return A[m, n]
nw_score = needleman_wunsch('AGGGCT','AGGCA')
nw_score
needleman_wunsch('apple','pineapple')
needleman_wunsch('xix','')
needleman_wunsch('pleasant','presented')
```
## DP - Optimal Binary Search Trees
| github_jupyter |
```
%matplotlib inline
from matplotlib import style
style.use('fivethirtyeight')
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import datetime as dt
```
# Reflect Tables into SQLAlchemy ORM
```
# Python SQL toolkit and Object Relational Mapper
import sqlalchemy
from sqlalchemy.ext.automap import automap_base
from sqlalchemy.orm import Session
from sqlalchemy import create_engine, func
# create engine to hawaii.sqlite
engine = create_engine("sqlite:///Resources/hawaii.sqlite")
# reflect an existing database into a new model
db = automap_base()
# reflect the tables
db.prepare(engine, reflect=True)
# View all of the classes that automap found
db.classes.keys()
# Save references to each table
meas = db.classes.measurement
stat = db.classes.station
# Create our session (link) from Python to the DB
sess = Session(engine)
```
# Exploratory Precipitation Analysis
```
# Find the most recent date in the data set.
last_date = sess.query(meas.date).order_by(meas.date.desc()).first()
last_date
# Calculate the date one year from the last date in data set.
minus_year = dt.date(2017,8,23) - dt.timedelta(days=365)
print(f'One year before last date: {minus_year}')
#Design a query to retrieve the last 12 months of precipitation data and plot the results.
# Starting from the most recent data point in the database.
precip = sess.query(meas.date, meas.prcp).\
filter(meas.date >= minus_year).\
order_by(meas.date).all()
# Perform a query to retrieve the data and precipitation scores
all_the_rain = sess.query(meas.date, meas.prcp).order_by(meas.date.desc()).all()
# Save the query results as a Pandas DataFrame and set the index to the date column
#Sort the dataframe by date
rain_df = pd.DataFrame(precip,columns=['Date','Precipitation (in)'])
rain_df.set_index("Date", inplace=True)
rain_df.head()
df2=rain_df.sort_values("Date", ascending=False)
df2.head()
# Use Pandas Plotting with Matplotlib to plot the data
# create plot
plt.rcParams['figure.figsize']=(15,7)
rain_df.plot(title="Precipitation Over Time", xticks=(0,60,120,180,240,300,365),
figsize=(15,6), color = 'teal', linewidth = 2.0, rot = 20)
# fix axis, label, and save
plt.xlabel('Date',fontsize=16,color='black',labelpad=20 )
plt.ylabel('Precipitation (in)', fontsize=16,color='black',labelpad=20)
plt.xlim(-5,370)
plt.legend(loc='upper right')
plt.savefig("Images/precip.png")
plt.show()
# Use Pandas to calcualte the summary statistics for the precipitation data
rain_df.describe()
```
# Exploratory Station Analysis
```
# Design a query to calculate the total number stations in the dataset
s1 = sess.query(meas.station).distinct().count()
print(f'There are {s1} stations in the dataset.')
# Design a query to find the most active stations (i.e. what stations have the most rows?)
# List the stations and the counts in descending order.
most_active = sess.query(meas.station, func.count(meas.station)).\
group_by(meas.station).\
order_by(func.count(meas.station).desc()).all()
active_df = pd.DataFrame(most_active, columns=["Station ID","Observations"])
active_df.sort_values("Observations", ascending=False)
active_df.head()
# Using the most active station id from the previous query, calculate the lowest, highest, and average temperature.
most_active_station = select = [func.min(meas.tobs), func.max(meas.tobs), func.avg(meas.tobs)]
min_max = sess.query(*select).\
filter(meas.station == "USC00519281").all()
print(f'The Minimum, Maximum, and Average of the most active station\nare as follows: {min_max}')
# Using the most active station id
# Query the last 12 months of temperature observation data for this station and plot the results as a histogram
temp_obs = sess.query(meas.tobs).\
filter(meas.date >= minus_year).\
filter(meas.station == "USC00519281").\
order_by(meas.date).all()
# save as a dataframe
temp_df = pd.DataFrame(temp_obs, columns=["Observstions (F)"])
# start graphing
temp_df.plot.hist(bins=12, title="Temperature Frequency (USC00519281)",
figsize=(12,6),color = 'teal', edgecolor='xkcd:light gray')
plt.xlabel("Temperature (F)")
plt.legend(loc="upper left")
plt.savefig("Images/temp.png")
plt.show()
```
# Close session
```
# Close Session
sess.close()
```
| github_jupyter |
# Lab 4
The goals of this lab are as follows:
1. introduce criticality calculations; and
2. compare OpenMC criticality calculation for a simple geometry and material composition to 1-Group Critical Equation.
The problem is to find the critical radius of a solid sphere of pure Pu-239.
## 1-Group Critical Equation
The analytic solution is arrived at by using the 1-Group Critical Equation given by:
$$B_1^2 = \frac{k_{\infty}-1}{L^2}$$
where $B_1^2$ is the buckling, and $L^2$ is the diffusion area; and $k_{\infty}$ is the infinite multiplication factor.
The buckling depends on geometry; for this problem, the domain is spherical and buckling is given by:
$$B^{2}=\left( \frac{\pi}{\tilde{R}}\right)^{2}$$
where $\tilde{R}$ is the extrapolated radius of the sphere: $\tilde{R}=R+d$; with $d$ being the extrapolation distance. The extrapolation distance is base on the diffusion coefficient: $d = 2.13D$ and the diffusion coefficient $(D)$ is based on the macroscopic transport cross section: $D=\frac{1}{3\Sigma_{tr}}$.
The macroscopic transport cross section is, in turn, computed from the corresponding microscopic cross section $(\sigma_{tr})$ and the atom density of the Pu-239 material.
We're almost to the bottom of the rabbit hole...
The diffusion area is based on the diffusion coefficient and the macroscopic absorption cross section:
$$L^{2}=\frac{D}{\Sigma_{a}}$$
and, similar to the macroscopic transport cross section, $\Sigma_{a}$ is computed with the atom density of the Pu-239 material and the corresponding microscopic cross section $(\sigma_a)$
Lastly, $k_{\infty}$ is the infinite multiplication factor. It is based on the regeneration factor ($\eta$) which we will take to be a known material property of Pu-239 and the fuel utilization, $f$, which we will take to be unity since the domain where neutrons will be diffusing and interacting is 100% fuel.
All of this long discussion is implemented in the code below. Note that all elements of the equations above can be taken to be known *except* for the radius $R$.
```
import numpy as np
# data:
N_Pu = 0.037; #atoms/b-cm, atom density of Pu-239
sig_f = 1.85; #b, fission micro xc
sig_a = 2.11; #b, absorption micro xc
sig_tr = 6.8; #b, transport micro xc
Sig_tr = N_Pu*sig_tr; # 1/cm, transport macro xc
D = 1./(3.*Sig_tr); #cm, diffusion coefficient
d = 2.13*D; #cm, extrapolation length
f = 1.; # fuel utilization
eta_pu = 2.61; # 1-group reproduction factor for Pu-239
k_inf = f*eta_pu;
Sig_a = N_Pu*sig_a; # 1/cm, abs macro xc
L_sq = D/Sig_a; # cm**2, diffusion area
B_sq = (k_inf-1.)/L_sq; #1/cm**2, buckling
R_tild = np.pi/(np.sqrt(B_sq)); # cm, extrapolated radius
R = R_tild - d; # cm, 1-group critical radius
print(f'1-group critical radius for sphere of Pu-238: %5.4f cm'%R)
```
## OpenMC model
Below we will implment this simple model and use OpenMC features to find the critical radius.
```
import openmc
```
In order to do this analysis we will use OpenMC's criticality search machinery. The basic idea is that we need to create a function that takes one parameter - in this case $R$ - that will be varied while we search for the value to that parameter that results in $k_{eff}$ being equal to one.
```
# create the model. "R" will be the parametric variable
def build_model(R):
# create the sphere material
fuel = openmc.Material(name='fuel');
fuel.add_nuclide('Pu239',1);
fuel.set_density('atom/b-cm',N_Pu);
materials = openmc.Materials([fuel]);
fuel_sphere = openmc.Sphere(r=R,boundary_type='vacuum');
fuel_cell = openmc.Cell();
fuel_cell.fill = fuel;
fuel_cell.region = -fuel_sphere;
root_universe = openmc.Universe();
root_universe.add_cells([fuel_cell]);
geometry = openmc.Geometry(root_universe);
settings = openmc.Settings()
settings.batches = 200;
settings.inactive = 50;
settings.particles = 10000;
bounds = [-R,-R,-R,R,R,R];
uniform_dist = openmc.stats.Box(bounds[:3],bounds[3:],
only_fissionable=True);
settings.source = openmc.source.Source(space=uniform_dist);
# so we don't waste disk input/output tallies that we will not use
settings.output = {'tallies':False};
model = openmc.model.Model(geometry,materials,settings);
return model
```
### Search for Critical Radius
To perform the search we will employ a call to
<code> openmc.search_for_keff</code> function and pass in the relevant arguments.
```
crit_R, guesses, keffs = openmc.search_for_keff(build_model,
bracket=[4.,15.],
tol=1e-3,print_iterations=True)
print(f'Critical Radius: %5.4f'%crit_R);
```
## Related Problem - Critical Dimension of a Cubic Thermal Reactor
For ER362 I often assign a homework problem in which the material composition of a thermal reactor is provided (usually just fuel and moderator; no structure, poison, etc...) and modified 1-group theory is used for the calculation.
Without repeating all of the details here, one problem is a bare cubic reactor in which U-235 and graphite are combined with a relative atom abundance of $1.0\times10^{-5}$ with the graphite, of course, being in the majority. The 1-group theory result is that the side-length of the critical thermal reactor (at 20${^\circ}$C) is about 400 cm. Let's see what OpenMC says with the help of the criticality search.
```
def thermal_model(L):
fuel = openmc.Material(name='fuel');
fuel.add_nuclide('U235',1.0e-5,'ao');
fuel.add_nuclide('C0',1.,'ao');
fuel.set_density('g/cm3',1.7); #assume the entire mixture has the density of pure graphite.
fuel.add_s_alpha_beta('c_Graphite');
fuel.temperature = 273.15 + 200.; # fuel at 200C expressed in K.
materials = openmc.Materials([fuel]);
# surfaces
top = openmc.YPlane(y0=L/2.,boundary_type='vacuum');
bottom = openmc.YPlane(y0=-L/2.,boundary_type='vacuum');
front = openmc.XPlane(x0=L/2.,boundary_type='vacuum');
back = openmc.XPlane(x0=-L/2.,boundary_type='vacuum');
left = openmc.ZPlane(z0=-L/2.,boundary_type='vacuum');
right = openmc.ZPlane(z0=L/2.,boundary_type='vacuum');
core = openmc.Cell();
core.fill = fuel
core.region = -top & +bottom & -front & +back & +left & -right;
root_universe = openmc.Universe();
root_universe.add_cells([core]);
geometry = openmc.Geometry(root_universe);
settings = openmc.Settings()
settings.batches = 200;
settings.inactive = 50;
settings.particles = 10000;
settings.temperature['method']='interpolation'; # allow interpolation of temps.
settings.temperature['multipole']=False; # specify use of windowed multipole data for resolved resonances.
bounds = [-L/2.,-L/2.,-L/2.,L/2.,L/2.,L/2.];
uniform_dist = openmc.stats.Box(bounds[:3],bounds[3:],
only_fissionable=True);
settings.source = openmc.source.Source(space=uniform_dist);
# so we don't waste disk input/output tallies that we will not use
settings.output = {'tallies':False};
model = openmc.model.Model(geometry,materials,settings);
return model
crit_L, guesses, keffs = openmc.search_for_keff(thermal_model,
bracket=[200.,500.],
tol=1e-3,print_iterations=True)
print(f'Critical Side Length: %5.4f'%crit_L);
```
This is not too far off from the 1-group result above. Notice all of the gibberish below; I'm leaving this here just for posterity. It turns out that the problem was that I'd used an unrealistically high mass density for the carbon (~2.6 g/cm3) vice the recommended density (from PNNL Material Handbook) of 1.7 g/cm3. This *definitely* explains the problem and is a great example of what happens when you half-ass the QC on model inputs. The good news is that the answer reinforces the correctness of the analytic result and, further, jibes with an intuitive expectation of how the OpenMC result *should* differ from the modified 1-group result.
__________________________________________________________________________________
The first time I successfully executed this critical search, I had left the fuel temperature at 20$^{\circ}$C when in the problem I am trying to emulate the fuel temperature is at 200$^{\circ}$C; a pretty imporant detail to leave off. Nonetheless, the critical side length at 20$^{\circ}$C is about 278.4cm.
Even getting the temperature correct, the critical side length is much smaller than modified 1-group theory predicts. I'd like to know why. Once answer might be: my modified 1-group theory analysis is just wrong. Another answer might be that the neutron energy spectrum in the graphite reactor might deviate sufficiently from "thermal" spectrum that the "corrections" to diffusion parameters and reaction cross sections are incorrect. Still, if I follow that logic - which would be to assume that neutrons are more fast than thermal - then I think that I should expect critical side length to be bigger rather than smaller. Note that I added the setting specifying that windowed multipole expansions would be used for resolved resonances. This didn't have much of an effect and, anyway, the only nuclide with resonance absorption is U235. Using windowed multipole data, the critical side length is about 309.3 cm. Not using it: 309.9. Conclusion: that's not it.
The problem is actually based on a question from Lamarsh (3rd ed.)- chapter 6, question 25; except for that question the specified temperature was 250$^{\circ}$C (I'm sure I changed it to 200 so there would be no need to interpolate tabulated values for non-1/v factors and regeneration factor. I cannot use the "solution manual" for the text as backup because the "solution" for that problem is mostly missing from the PDF copy of the manual that I have. (*sigh*)
| github_jupyter |
# Practice for understanding image classification with neural network
- Single layer neural network with gradient descent
## 1) Import Packages
```
from PIL import Image
import numpy as np
import matplotlib.pyplot as plt
import random
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import OneHotEncoder
import math
import sklearn.metrics as metrics
```
## 2) Make Dataset
```
x_orig = []
y_orig = np.zeros((1,100))
for i in range(1,501):
if i <= 100 :
folder = 0
elif i <=200 :
folder = 1
elif i <=300 :
folder = 2
elif i <=400 :
folder = 3
else :
folder = 4
img = np.array(Image.open('dataset/{0}/{1}.jpg'.format(folder,i)))
img = Image.fromarray(img).convert('L') # gray
data = img.resize((64,64))
data = np.array(data)
x_orig.append(data)
for i in range(1,5):
y_orig = np.append(y_orig, np.full((1, 100),i), axis = 1)
x_orig = np.array(x_orig)
print(x_orig.shape)
print(y_orig.shape)
# Random shuffle
s = np.arange(x_orig.shape[0])
np.random.shuffle(s)
x_shuffle = x_orig[s,:]
y_shuffle = y_orig[:,s]
print(x_shuffle.shape)
print(y_shuffle.shape)
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(x_shuffle[i,:])
plt.xlabel(y_shuffle[:,i])
plt.show()
# Split train and test datasets
x_train_orig, x_test_orig, y_train_orig, y_test_orig = train_test_split(x_shuffle,y_shuffle.T,
test_size=0.2, shuffle=True, random_state=1004)
print(x_train_orig.shape)
print (y_train_orig.shape)
# Flatten the training and test images
x_train_flatten = x_train_orig.reshape(x_train_orig.shape[0], -1).T
x_test_flatten = x_test_orig.reshape(x_test_orig.shape[0], -1).T
# Normalize image vectors
x_train = x_train_flatten/255.
x_test = x_test_flatten/255.
# Convert training and test labels to one hot matrices
enc = OneHotEncoder()
y1 = y_train_orig.reshape(-1,1)
enc.fit(y1)
y_train = enc.transform(y1).toarray()
y_train = y_train.T
y2 = y_test_orig.reshape(-1,1)
enc.fit(y2)
y_test = enc.transform(y2).toarray()
y_test = y_test.T
# Explore dataset
print ("number of training examples = " + str(x_train.shape[1]))
print ("number of test examples = " + str(x_test.shape[1]))
print ("x_train shape: " + str(x_train.shape))
print ("y_train shape: " + str(y_train.shape))
print ("x_test shape: " + str(x_test.shape))
print ("y_test shape: " + str(y_test.shape))
```
## 3) Definie required functions
```
def initialize_parameters(nx, ny):
"""
Argument:
nx -- size of the input layer (4096)
ny -- size of the output layer (3)
Returns:
W -- weight matrix of shape (ny, nx)
b -- bias vector of shape (ny, 1)
"""
np.random.seed(1)
W = np.random.randn(ny,nx)*0.01
b = np.zeros((ny,1))
assert(W.shape == (ny, nx))
assert(b.shape == (ny, 1))
return W, b
def softmax(Z):
# compute the softmax activation
S = np.exp(Z + np.max(Z)) / np.sum(np.exp(Z + np.max(Z)), axis = 0)
return S
def classlabel(Z):
# probabilities back into class labels
y_hat = Z.argmax(axis=0)
return y_hat
def propagate(W, b, X, Y):
m = X.shape[1]
# Forward Propagation
Z = np.dot(W, X)+ b
A = softmax(Z) # compute activation
cost = (-1/m) * np.sum(Y * np.log(A)) # compute cost (Cross_entropy)
# Backward propagation
dW = (1/m) * (np.dot(X,(A-Y).T)).T
db = (1/m) * (np.sum(A-Y))
grads = {"dW": dW,
"db": db}
return grads, cost
```
## 4) Single-Layer Neural Network with Gradient Descent
```
def optimize(X, Y, num_iterations, learning_rate, print_cost = False):
costs = []
W, b = initialize_parameters(4096,5)
for i in range(num_iterations):
grads, cost = propagate(W,b,X,Y)
dW = grads["dW"]
db = grads["db"]
W = W - (learning_rate) * dW
b = b - (learning_rate) * db
# Record the costs for plotting
if i % 100 == 0:
costs.append(cost)
# Print the cost every 100 training iterations
if print_cost and i % 200 == 0:
print ("Cost after iteration %i: %f" %(i, cost))
# plot the cost
plt.plot(costs)
plt.ylabel('cost')
plt.xlabel('iterations (per 200)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
# Lets save the trainded parameters in a variable
params = {"W": W,
"b": b}
grads = {"dW": dW,
"db": db}
return params, grads, costs
params, grads, costs = optimize(x_train, y_train, num_iterations= 1000, learning_rate = 0.01, print_cost = True)
print ("W = " + str(params["W"]))
print ("b = " + str(params["b"]))
```
## 5) Accuracy Analysis
```
def predict(W, b, X) :
m = X.shape[1]
# Compute "A" predicting the probabilities
Z = np.dot(W, X)+ b
A = softmax(Z)
# Convert probabilities A to actual predictions
y_prediction = A.argmax(axis=0)
return y_prediction
# Predict test/train set
W1 = params['W']
b1 = params['b']
y_prediction_train = predict(W1, b1, x_train)
y_prediction_test = predict(W1, b1, x_test)
print(y_prediction_train)
print(y_prediction_test)
# Print train/test accuracy
print("train accuracy : ", metrics.accuracy_score(y_prediction_train, y_train_orig))
print("test accuracy : ", metrics.accuracy_score(y_prediction_test, y_test_orig))
```
| github_jupyter |
```
import pandas as pd
import numpy as np
```
Get the Data
```
column_names = ['user_id', 'item_id', 'rating', 'timestamp']
df = pd.read_csv("u.data", sep='\t', names=column_names)
df.head()
movie_titles = pd.read_csv("Movie_Id_Titles")
movie_titles.head()
```
We can merge them together:
```
df = pd.merge(df, movie_titles, on='item_id')
df.head()
```
EDA
Let's explore the data a bit and get a look at some of the best rated movies.
```
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('white')
%matplotlib inline
```
Let's create a ratings dataframe with average rating and number of ratings:
```
df.groupby('title')['rating'].mean().sort_values(ascending=False).head()
df.groupby('title')['rating'].count().sort_values(ascending=False).head()
ratings = pd.DataFrame(df.groupby('title')['rating'].mean())
ratings.head()
ratings['num of ratings'] = pd.DataFrame(df.groupby('title')['rating'].count())
ratings.head()
```
Now a few histograms:
```
plt.figure(figsize=(10,4))
ratings['num of ratings'].hist(bins=70)
plt.figure(figsize=(10,4))
ratings['rating'].hist(bins=70)
sns.jointplot(x='rating', y = 'num of ratings', data= ratings, alpha=0.5)
```
Recommending Similar Movies
```
moviemat = df.pivot_table(index='user_id', columns='title', values='rating')
moviemat.head()
ratings.sort_values('num of ratings', ascending=False).head(10)
ratings.head()
starwars_user_ratings = moviemat['Star Wars (1977)']
liarliar_user_ratings = moviemat['Liar Liar (1997)']
starwars_user_ratings.head()
similar_to_starwars = moviemat.corrwith(starwars_user_ratings)
similar_to_liarliar = moviemat.corrwith(liarliar_user_ratings)
corr_starwars = pd.DataFrame(similar_to_starwars, columns=['Correlation'])
corr_starwars.dropna(inplace=True)
corr_starwars.head()
```
Now if we sort the dataframe by correlation, we should get the most similar movies, however note that we get some results that don't really make sense. This is because there are a lot of movies only watched once by users who also watched star wars (it was the most popular movie).
```
corr_starwars.sort_values('Correlation',ascending=False).head(10)
corr_starwars = corr_starwars.join(ratings['num of ratings'])
corr_starwars.head()
corr_starwars[corr_starwars['num of ratings']>100].sort_values('Correlation',ascending=False).head()
```
Now the same for the comedy Liar Liar:
```
corr_liarliar = pd.DataFrame(similar_to_liarliar, columns=['Correlation'])
corr_liarliar.dropna(inplace=True)
corr_liarliar = corr_liarliar.join(ratings['num of ratings'])
corr_liarliar[corr_liarliar['num of ratings']>100].sort_values('Correlation',ascending=False).head()
```
| github_jupyter |
## Infraestructuras Computacionales para el Procesamiento de Datos Masivos
### Práctica del Módulo 3: Gestión de datos en tiempo real (Streaming)
### Autor: Jesús Galán Llano
#### Correo: jgalan279@alumno.uned.es
```
from pyspark.sql import SparkSession
from pyspark.sql.functions import split
from pyspark.sql.functions import explode
```
En primer lugar, creamos la sesión de Spark
```
spark = SparkSession \
.builder \
.appName("tp3") \
.getOrCreate()
```
Tras esto, especificamos el tipo de entrada de datos que en este caso se corresponde con Apache Kafka.
Es importante configurar correctamente el número de brokers de Kafka así como su dirección. La configuración descrita en la memoria
consiste de dos servidores, que están disponibles en los puertos 9092 y 9093 en el equipo local.
```
lines = spark \
.readStream \
.format("kafka") \
.option("kafka.bootstrap.servers", "localhost:9092,localhost:9093") \
.option("subscribe", "kafkaTwitter") \
.option("includeHeaders", "true") \
.load()
```
Creamos un DataFrame formado por todas las palabras que aparecen en los tweets. Para ello, dividimos los mensajes en espacios en blanco
con la función split(). La función explode permite devolver más de un elemento, que es la lista de palabras, por cada elemento procesado, que es un tweet.
```
words = lines.select(
explode(split(lines.value, ' ')).alias('word'),
)
```
Como solamente nos interesa almacenar los hashtags aplicamos una operación de filtrado sobre el DataFrame anterior.
Los hashtags son aquellas palabras que empiezan por el caracter '#'.
```
hashtags = words.filter(words.word.startswith('#'))
```
Tras obtener todos los hashtags, se agrupan utilizando la función groupBy() y se realiza el cálculo de cuántas veces aparece dicho hashtag con la función count().
Seguidamente, ordenamos los resultados utilizando la función orderBy en base al número de veces que aparece cada hashtags de forma descendente, empezando por los que más se repiten.
Por último, limitamos los resultados a los 10 hashtags más utilizados con la función limit().
```
hashtagsCounts = hashtags.groupBy(
hashtags.word
).count()\
.orderBy('count', ascending=False)\
.limit(10)
```
Por último, ejecutamos el proceso para procesar los tweets que van llegando desde el productor.
Se ha dejado la ejecución del programa durante un período de tiempo para validar su funcionamiento. Las conclusiones de la práctica se incluyen al final del cuaderno.
```
hashtagsCounts\
.writeStream\
.outputMode('complete')\
.format('console')\
.option('truncate', 'false')\
.start()
```
## Conclusiones
Esta segunda parte de la práctica me ha resultado muy interesante porque me ha permitido poner en práctica mis conocimientos de Kafka y de SparkStreaming. Considero que
una de las características principales de un ingeniero de datos debe ser saber conectar más de un sistema para lograr un fin aprovechando las características de cada uno, como se realiza en esta práctica.
A pesar de haberme atascado al comienzo luego he podido ir avanzando poco a poco empleando la extensa documentación que hay en internet sobre SparkStreaming
y consultando ciertos ejemplos de los libros de la bibliografía. Sin embargo, por falta de tiempo no he podido entregar el tercer ejercicio de esta parte por fala de tiempo.
Como punto a mejorar para los próximos años, comentar que los ejemplos incluidos en las transparencias respecto a Kafka están desactualizados porque funcionan con versiones de Spark inferiores a la versión 3 y requieren de Python2. Debido a esto he tenido que emplear más tiempo del deseado en preparar el entorno para la práctica y en encontrar una documentación que me fuera útil. Sería muy recomendable adaptar ese contenido al uso de Spark 3 y de Python 3 para aplicar todas las ventajas que estas versiones ofrecen.
Por último, agradecer de nuevo las facilidades del equipo docente respecto a permitir entregas después de la fecha de entrega.
| github_jupyter |
# Tests pour le tournois de 2048
Ce notebook donne un aperçu de la manière dont le tournois sera lancé. Les cellules de codes écrites en python.
Les étapes de la simulation sont les suivantes:
1. Lancer la simulation du jeu ```tournois_simulation``` (idéallement dans un terminal)
2. Lancer votre programme de joueur automatique ```2048_IA``` ou l'un des deux exécutables de démonstration de joueur automatique fournis (```2048_IA_random``` ou ```2048_IA_naive```)
3. Attendez que la simulation se termine. Si les formats d'écriture sont corrects, les fichiers ```configuration.txt``` resp. ```mouvements.txt``` seront mis à jour succéssivement par le moteur de jeu resp. votre joueur automatique.
4. Observer le résultats de la simulation dans les fichier ```configuration.txt``` et ```mouvements.txt```
Lisez attentivement les cellules suivantes et contacter votre chargé de TP si des erreurs apparaissent au lancement des cellules.
## Lancer la simulation du jeu
Ouvrez un terminal, et déplacez vous dans le répertoire ```Projet-2048/tournois```. Puis exécuter ```./tournois_simulation```. Si vous obtenez des erreurs au lancement, c'est que vous êtes sûrement sur windows, ou MacOS. En effet, pour ne pas divulger le code source (sujet du projet), nous avons compilé ```tournois_simulation.cpp``` sur le serveur jupyterHub. Réessayer de lancer cet exécutable sur votre serveur jupyterHub.
NB: Si vous voulez conserver les résultats intermédiaires de la simulation, vous pouvez toujours rediriger la sortie standard de ```./tournois_simulation``` dans un nouveau fichier avec la commande ```./tournois_simulation > resultats_intermediaires.txt```.
Si l'exécutable ne se lance toujours, pas vérifiez que les permissions du fichiers permettent bien son exécution.
Pour cela vérifiez les permissions des fichiers avec la cellule suivante:
```
import subprocess
ls_output = subprocess.check_output(['ls', '-l'])
print(ls_output.decode())
```
Si les fichiers ```tournois_simulation```, ```2048_IA_random``` et ```2048_IA_naive``` ne sont pas des exécutables (c'est à dire qu'ils ne continnennt pas de x dans ```-rw-r--r--```), alors changer les permissions avec:
```
os.system("chmod 755 tournois_simulation")
os.system("chmod 755 2048_IA_random")
os.system("chmod 755 tournois_naive")
```
## Lancer le programme d'un joueur automatique
À présent, lancez la simulation d'exemple dans un terminal (pour voir l'affichage) ou bien avec la cellule suivante (en attendant qu'elle se termine):
```
os.system("./2048_IA_random")
```
## Erreurs possibles
Si la simulation ne termine jamais ou renvoie une erreur dès les premiers déplacements, plusieurs problèmes sont possibles:
- Le format du fichier ```mouvements.txt``` n'est pas bon. Consulter le sujet ou le fichier ```mouvements.txt``` créé par les IA d'exemples ```2048_IA_random``` ou ```2048_IA_naive```.
- La lecture du fichier ```configuration.txt``` n'est pas effecutée correctement.
- Vous n'avez pas lancé la simulationn ```./tournois_simulation``` avant de lancer votre joueur automatique
- Vous n'avez pas supprimé les fichiers ```mouvements.txt``` ou ```configuration.txt``` au lancement d'une nouvelle simulation
- Vous avez lancé plusieurs fois la simulation du tournois ```./tournois_simulation``` (dans plusieurs terminnaux par exemple)
Si des erreurs persistent, demandez de l'aide à votre chargé de TD.
| github_jupyter |
# Dataoverføring og statistikk
## Lese inn en fil
`pylab.loadtxt(<filename>[, delimiter=hva man skiller dataene med (',')][, skiprows=antalllinjer å hoppe over][, dtype=datatype])`
```
import pylab
data = pylab.loadtxt("sunspots.csv", delimiter=",", skiprows=1)
nr = data[:, 0]
verdi = data[:, 1]
pylab.plot(nr, verdi)
pylab.show()
maaned = pylab.linspace(1, 12, 12)
solflekker = [0] * 12
for i in range(0, len(nr)):
month = i % 12
solflekker[month] += verdi[i]
pylab.plot(maaned, solflekker)
pylab.show()
```
## Statistisk analyse
### Gjennomsnitt
summen av alle verdiene delt på antall verdier.
$$\bar x = \frac{1}{n}\sum_{i=1}^{n}x_i$$
`pylab.mean(<liste med verdier>)`
### Standardavvik
største/minste verdi - gjennomsnittet
$$s = \sigma^2 = \sqrt{\frac{1}{n} \sum_{i=1}^{n}(x_i - \bar x)^2}$$
`pylab.std(<liste med verdier>)`
```
import pylab
alder = [15, 24, 16, 18, 19, 42, 15, 20, 16, 17, 19]
snitt = pylab.mean(alder)
print("Gjennomsnittsalder: {}".format(snitt))
avvik = pylab.std(alder)
print("StandardAvvik: {}".format(avvik))
```
## Regresjon
Finne en modell for et datasett. Detet gjøres ved å finne en funksjon (ofte en polynomfunksjon) som passer best mulig med punktene. Dette blir en modell, som kan stemme bra. Likevel kan den forutsi veldig rare verdier langt utenfor datasettet.
```
import pylab
T = [0, 20, 40, 60, 80, 100]
sol_NaCl = [35.7, 35.9, 36.4, 37.1, 38.0, 39.2]
pylab.scatter(T, sol_NaCl)
grad = 4
reg_NaCl = pylab.polyfit(T, sol_NaCl, grad)
print(reg_NaCl)
def polynomFunction(polynom, x):
ret = 0
for i in range(0, len(polynom)):
g = len(polynom) - 1 - i
ret += polynom[i] * x ** g
return ret
x = pylab.linspace(0, 150)
y1 = polynomFunction(reg_NaCl, x)
pylab.plot(x, y1)
pylab.show()
```
### R2-test
R2 score gir en verdi på hvor godt regresjonen stememr med dataene. Går fra 0 til 1, der 0 er ufattelig dårlig, og 1 er akkurat riktig.
Denne kan brukes for å finen ut om regresjonen er god, MEN den kan vise at den er veldig god fordi den passer veldig godt med dataene, men passer ikke i det heledatt mellom datapunktene.
```
import pylab
from sklearn.metrics import r2_score
T = [0, 20, 40, 60, 80, 100]
sol_NaCl = [35.7, 35.9, 36.4, 37.1, 38.0, 39.2]
pylab.scatter(T, sol_NaCl)
grad = 4
reg_NaCl = pylab.polyfit(T, sol_NaCl, grad)
def polynomFunction(polynom, x):
ret = 0
for i in range(0, len(polynom)):
g = len(polynom) - 1 - i
ret += polynom[i] * x ** g
return ret
x = pylab.linspace(0, 150)
y = polynomFunction(reg_NaCl, x)
pylab.plot(x, y)
pylab.show()
# R2 test
R2y = polynomFunction(reg_NaCl, pylab.array(T))
R2 = r2_score(sol_NaCl, R2y)
print("R2 test: {}".format(R2))
```
### Ekstrapolering
Forutsi senere verdier. Selvom R2 verdien er veldig god, trenger den ikke være god på senere verdier.
## Diagrammer
`errorbar` kan brukes for å lage et usikekrhetsplott. Det gir feilmargin barer på hvert punkt
### kakediagram og søylediagram
```
import pylab
fag = ["R2", "S2", "Kjemi 2", "Fysikk 2", "Tekforsk", "Matematikk X", "Biologi 2"]
antall = [110, 25, 74, 65, 10, 3, 45]
pylab.pie(antall, labels=fag)
pylab.show()
import pylab
hoyder = [1.79, 1.80, 1.60, 1.75, 1.65, 1.76, 1.83, 1.71, 1.72]
pylab.hist(hoyder)
pylab.show()
from pylab import *
x = linspace(-3, 3, 18)
y = 5 * exp(-x**2) * sin(6 * x)
errorbar(x, y, yerr = exp(-x**2) + .5)
show()
```
| github_jupyter |
```
# Portfolio Optimisation
# Author Sanket Karve
# Importing all required libraries
import pandas as pd
import pandas_datareader as web
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import FuncFormatter
# Importing the Portfolio Optimisation Library functions
from pypfopt.efficient_frontier import EfficientFrontier
from pypfopt import risk_models
from pypfopt import expected_returns
from pypfopt import discrete_allocation
from pypfopt.cla import CLA
import pypfopt.plotting as pplt
from matplotlib.ticker import FuncFormatter
# Get tickers of required stocks
tickers = ["BSX", "AES", "BRK-B", "SEE", "QQQ", "SPY"]
the_len = len(tickers)
price_data = []
for ticker in range(the_len):
prices = web.DataReader(tickers[ticker], start="2015-01-01", end="2020-06-06", data_source="yahoo")
price_data.append(prices.assign(ticker=ticker)[["Adj Close"]])
df_stocks = pd.concat(price_data, axis=1)
df_stocks.columns=tickers
df_stocks.head()
# Checking if there are any NAN values in the data
nullin_df = pd.DataFrame(df_stocks, columns=tickers)
print(nullin_df.isnull().sum())
# Annualised Return
mu = expected_returns.mean_historical_return(df_stocks)
mu
# Sample Variance of Portfolio
Sigma = risk_models.sample_cov(df_stocks)
Sigma
# Max Sharpe Ratio - Tangent to the EF
ef1 = EfficientFrontier(mu, Sigma, weight_bounds=(0,1)) # weight_bounds in negative allows shorting of stocks
sharpe_pfolio = ef1.max_sharpe() # May use add objective to ensure minimum zero weighting to individual stocks
sharpe_pwt = ef1.clean_weights()
print(sharpe_pwt)
# Get portfolio performance
ef1.portfolio_performance(verbose=True) # 2 year risk free rate - Treasury Yield would set risk_free_rate=0.27
# Minimum Volatility Portfolio
ef2 = EfficientFrontier(mu, Sigma, weight_bounds=(0,1)) # weight_bounds in negative allows shorting of stocks
minvol = ef2.min_volatility()
minvol_pwt = ef2.clean_weights()
print(minvol_pwt)
# Get MinVol portfolio performance
ef2.portfolio_performance(verbose=True) # 2 year risk free rate - Treasury Yield would set risk_free_rate=0.27
# Plot the Efficient Frontier for visual puposes
cl_obj = CLA(mu, Sigma)
ax = pplt.plot_efficient_frontier(cl_obj, showfig=False)
ax.xaxis.set_major_formatter(FuncFormatter(lambda x, _: "{:.0%}".format(x)))
ax.yaxis.set_major_formatter(FuncFormatter(lambda y, _: "{:.0%}".format(y)))
# Get latest prices
latest_prices = discrete_allocation.get_latest_prices(df_stocks)
latest_prices
# Sort Solvers
# import cvxpy as cp
# import cvxopt
# print(cp.installed_solvers())
# Allocate Portfolio Value in $ as required to show number of shares/stocks to buy
# Max Sharpe Ratio Portfolio Allocation $10,000
allocation_shp, rem_shp = discrete_allocation.DiscreteAllocation(sharpe_pwt, latest_prices, total_portfolio_value=10000).lp_portfolio()
print(allocation_shp)
print("Leftover fund after building Max Sharpe Portfolio is ${:.2f}".format(rem_shp))
# Min Volatility Portfolio Allocation $10,000
allocation_minv, rem_minv = discrete_allocation.DiscreteAllocation(minvol, latest_prices, total_portfolio_value=10000).lp_portfolio()
print(allocation_minv)
print("Leftover fund after building Min Volatility Portfolio is ${:.2f}".format(rem_minv))
```
| github_jupyter |
# Competition coefficient
```
# Housekeeping
library(car)
library(ggplot2)
library(MASS)
library(mgcv)
library(nlme)
library(reshape2)
library(scales)
library(tidyr)
source("../source.R")
# Read in data
species_composition = read.table("../../../data/amplicon/species_composition_relative_abundance.txt",
sep = "\t",
header = T,
row.names = 1)
metadata = read.table("../../../data/amplicon/metadata.txt",
sep = "\t",
header = T,
row.names = 1)
# Extract regime shift data without predation
x = metadata$Experiment != "FiltrateExp" & # keep only regime shift data
metadata$Predation != 1 & # exclude predation
metadata$Immigration != "stock" # exclude stock
# Subset
species_composition = species_composition[x,] # keep only species with data
species_composition = species_composition[,colSums(species_composition)>0]
metadata = metadata[x,-c(3, 6)] # remove redundant columns
# metadata$ID = paste(metadata$Streptomycin, metadata$Immigration, metadata$Replicate, sep = "_")
# metadata$ID = as.factor(metadata$ID)
species_composition$SAMPLE = rownames(species_composition)
metadata$SAMPLE = rownames(metadata)
df = merge(species_composition, metadata, all = T)
dim(df)
head(df)
df2 = melt(df[,-1], id.vars = c("Replicate", "Streptomycin", "Immigration", "Time_point"))
colnames(df2) = c("Replicate", "Streptomycin", "Immigration", "Time_point", "Species", "Abundance")
head(df2)
df3 = spread(df2, Time_point, Abundance)
colnames(df3) = c("Replicate", "Streptomycin", "Immigration", "Species", "ABUND16", "ABUND32", "ABUND48")
head(df3)
```
## Compute selection coefficient
The slope of a least square regression line (linear model) for logit-transformed allele frequency trajectories (here consisting of two time points) gives the selection coefficient
$s = \frac {logit(f(t_2)) - logit(f(t_1))} {t_2 - t_1}$
where $f(t)$ is the frequency of a mutation at time $t$ (https://www.ncbi.nlm.nih.gov/pubmed/28961717). Fitness $W$ is then given by
$W = s + 1$.
These metrics are used here for changes in the relative abundance of species and denote the competitive ability of species.
```
# Convert zero abundance to 10^-6 (order of magnitude lower than lowest detected abundance)
# to allow estimation of selection coefficients
extinction = df3[df3$ABUND32 == 0 | df3$ABUND48 == 0,]
df3 = df3[df3$ABUND16 != 0 & df3$ABUND32 != 0 & df3$ABUND48 != 0,]
#df3$ABUND16 = ifelse(df3$ABUND16 == 0, 0.000001, df3$ABUND16)
#df3$ABUND32 = ifelse(df3$ABUND32 == 0, 0.000001, df3$ABUND32)
#df3$ABUND48 = ifelse(df3$ABUND48 == 0, 0.000001, df3$ABUND48)
# Compute s
df3$ABUND_S_INTERVAL1 = (logit(df3$ABUND32) - logit(df3$ABUND16))/(32-16)
df3$ABUND_S_INTERVAL2 = (logit(df3$ABUND48) - logit(df3$ABUND32))/(48-32)
head(df3)
df3$Sample = paste(df3$Streptomycin, df3$Immigration, df3$Replicate, sep = "_")
head(df3)
library(lmerTest)
# ABUND_S_INTERVAL1
# Model with linear regression
M1 = lmer(ABUND_S_INTERVAL1 ~ Streptomycin * Immigration * Species + (1|Sample),
data = df3)
step(M1)
# Best model
M2 = lmer(ABUND_S_INTERVAL1 ~ Streptomycin * Species + (1 | Sample),
data = df3)
M3 = lmer(ABUND_S_INTERVAL1 ~ Streptomycin + Species + (1 | Sample),
data = df3)
anova(M2, M3, test = "Chi") # sig. interaction
M4 = lmer(ABUND_S_INTERVAL1 ~ Streptomycin + (1 | Sample),
data = df3)
anova(M2, M4, test = "Chi") # sig. species effect
M5 = lmer(ABUND_S_INTERVAL1 ~ Species + (1 | Sample),
data = df3)
anova(M2, M5, test = "Chi") # sig. streptomycin effect
# Streptomycin affects slope depending on species
# Plot
ggplot(df3,
aes(factor(Streptomycin), ABUND_S_INTERVAL1, colour = factor(Streptomycin))) +
geom_boxplot() +
facet_grid(Species~., scales = "free") +
ylab("Rate of change of species") +
xlab(expression(paste("Initial abundance level"))) +
theme_classic() +
theme(panel.spacing = unit(1, "lines"),
legend.title=element_blank(),
strip.background = element_blank(),
strip.text = element_text(face = "italic"))
# ABUND_S_INTERVAL2
M1 = lmer(ABUND_S_INTERVAL2 ~ Streptomycin * Immigration * Species + (1|Sample),
data = df3)
step(M1)
# Best model
M2 = lmer(ABUND_S_INTERVAL2 ~ Streptomycin * Immigration * Species + (1 | Sample),
data = df3)
M3 = lmer(ABUND_S_INTERVAL2 ~ Streptomycin + Immigration + Species + (1 | Sample),
data = df3)
anova(M2, M3, test = "Chi") # sig. interaction
M4 = lmer(ABUND_S_INTERVAL2 ~ Streptomycin * Immigration + (1 | Sample),
data = df3)
anova(M2, M4, test = "Chi") # sig. species effect
M5 = lmer(ABUND_S_INTERVAL2 ~ Streptomycin * Species + (1 | Sample),
data = df3)
anova(M2, M5, test = "Chi") # sig. immigration effect
M6 = lmer(ABUND_S_INTERVAL2 ~ Species*Immigration + (1 | Sample),
data = df3)
anova(M2, M6, test = "Chi") # sig. streptomycin effect
# Plot
ggplot(df3,
aes(factor(Streptomycin), ABUND_S_INTERVAL2)) +
geom_boxplot() +
facet_grid(Species~., scales = "free") +
ylab("Rate of change of species") +
xlab(expression(paste("Initial abundance level"))) +
theme_classic() +
theme(panel.spacing = unit(1, "lines"),
legend.title=element_blank(),
strip.background = element_blank(),
strip.text = element_text(face = "italic"))
# Slopes
M1 = lmer(ABUND_S_INTERVAL2 ~ ABUND_S_INTERVAL1 + Species + Streptomycin + Immigration + (1|Sample),
data = df3)
step(M1)
# Best model
M2 = lmer(ABUND_S_INTERVAL2 ~ ABUND_S_INTERVAL1 + Species + Streptomycin + (1 | Sample),
data = df3)
# Incorporate phenotypes
phenotypes = read.table("../../../data/amplicon/phenotypic_traits.txt",
sep = "\t",
header = T)
phenotypes$Species = rownames(phenotypes)
df4 = merge(df3, phenotypes, by = "Species", all = T)
dim(df3)
dim(df4)
head(df4)
df4$MIC_orig = exp(df4$Streptomycin_MIC)
head(df4)
# Slope 1
M1 = lmer(ABUND_S_INTERVAL1 ~ Streptomycin + Immigration + K_growth + r_growth + Aminoglycoside_resistance_gene + Carbon_sources + Streptomycin_MIC + (1|Sample),
data = df4)
step(M1)
# Best model
M2 = lmer(ABUND_S_INTERVAL1 ~ r_growth + Aminoglycoside_resistance_gene + Streptomycin_MIC + (1 | Sample),
data = df4)
summary(M2)
# Slope 2
M1 = lmer(ABUND_S_INTERVAL2 ~ Streptomycin + Immigration + K_growth + r_growth + Aminoglycoside_resistance_gene + Carbon_sources + Streptomycin_MIC + (1|Sample),
data = df4)
step(M1)
# Best model
#M2 = lmer(ABUND_S_INTERVAL2 ~ r_growth + Aminoglycoside_resistance_gene + Streptomycin_MIC + (1 | Sample),
# data = df4)
#summary(M2)
head(df4)
df5 = melt(df4[,-c(1,2,5,6,7,10,11,14,15, 16)], id.vars = c("Streptomycin", "Immigration", "r_growth", "Streptomycin_MIC"))
colnames(df5) = c("Streptomycin", "Immigration", "r_growth", "Streptomycin_MIC", "Slope_interval", "Slope")
head(df5)
df6 = melt(df5, id.vars = c("Streptomycin", "Immigration", "Slope_interval", "Slope"))
colnames(df6) = c("Streptomycin", "Immigration", "Slope_interval", "Slope", "Phenotype", "Pheno_value")
head(df6)
# Edit
df6$Slope_interval = ifelse(df6$Slope_interval == "ABUND_S_INTERVAL1", "Antibotic exposure phase", "Recovery phase")
df6$Slope_interval = factor(df6$Slope_interval, levels = c("Antibiotic exposure phase", "Recovery phase"))
ggplot(na.omit(df6),
aes(Pheno_value, Slope, colour = factor(Streptomycin), fill = factor(Streptomycin))) +
stat_smooth(method = "lm") +
facet_grid(Slope_interval~Phenotype*Streptomycin, scales = "free_x") +
ylab("Species competitive ability") +
xlab(expression(paste("Phenotype"))) +
scale_color_manual(values = c("#D3D3D3", "#cd6090", "#8f4364", "#522639")) +
scale_fill_manual(values = c("#D3D3D3", "#cd6090", "#8f4364", "#522639")) +
labs(colour="Streptomycin level", fill = "Streptomycin level") +
theme_classic() +
theme(panel.spacing = unit(1, "lines"),
strip.background = element_blank())
ggsave("../../../manuscript/figures/phenotype_response.pdf", width = 12, height = 10)
```
| github_jupyter |
# Berry Bengali: Blog Post 2
### Alex Berry, Jason Chan, Hyunjoon Lee
Brown University Data Science Initiative
DATA 2040: Deep Learning
February 29th, 2020
This project involves classifying handwritten characters of the Bengali alphabet, similar to classifying integers in the MNIST data set. In particular, the Bengali alphabet is broken down into three components for each grapheme, or character: 1) the root, 2) the vowel diacritic, 3) the consonant diacritic, where a diacritic is similar to an accent. The goal is the create a classification model that can classify each of these three components of a handwritten grapheme, and the final result is measured using the recall metrics, with double weight given to classification of the root.
Since our last blog post, we have focused our efforts on taking the given data and applying various preprocessing methods to both improve the performance of our model and decrease its training time.
# Preprocessing
Motivated by a desire for fine-tuned control over the input data's size and quality, we made the decision to apply preprocessing "outside" of the rest of the model pipeline. Our methodologies included cropping, resizing, denoising, and thresholding.
## 1. Cropping
The first preprocessing method applied to the image data is cropping. Cropping was applied first because we believed it would be most appropriate to crop the images first and then to manipulate the pixels (denoising and thresholding) of the remaining bits of the images.
Cropping is used to segment and maintain the most salient region (elements in region that stand out and attract the viewer's attention) of the image while cutting out the non-salient region. The position of the graphemes varies by image. Some graphemes are in the center, some are in top-left corner of the image. The size of the graphemes varies by image as well. Some graphemes fill the entire image, while some graphemes are slightly bigger than a dot. Our objective was to segment the graphemes and crop out the white space while maintaining as much information about the graphemes as possible.
We implemented the function below to crop our images.
```python
def crop_surrounding_whitespace(image):
"""Remove surrounding empty space around an image.
This implemenation assumes that the surrounding empty space
around the image has the same colour as the top leftmost pixel.
:param image: PIL image
:rtype: PIL image (cropped)
"""
bg = Image.new(image.mode, image.size, image.getpixel((0,0)))
diff = ImageChops.difference(image, bg)
diff = ImageChops.add(diff, diff, 2, -50)
bbox = diff.getbbox()
return image.crop(bbox)
```
## 2. Resizing
After cropping, we resized the images so that every image has the same dimension. The images must be resized to the same dimension because each sample of CNN model's input must be of the same size. We determined the dimension of our resized images based on the distribution of the cropped images. Below is the distribution of row dimension (number of pixels) of the cropped images.

Below is the distribution of column dimension (number of pixels) of the cropped images.

The distribution of row dimension was slightly right-skewed and column dimension was symmetric. However, both distribution was bimodal with images that were not cropped (given that the orignal images were $137 \times 236$). Therefore, we determined that resizing each row and column dimension by the median of their respective distribution the most appropriate. The resized dimension of the images were $106 \times 87$.
## 3. Denoising
After we resized the images, we converted the image from PIL Image to numpy array to use CV2 library tools. The first tool we used was `cv2.fastNlMeansDenoising()`. According to OpenCV docuemntation, this method performs image denoising using Non-Local Means Denoising algorithm (http://www.ipol.im/pub/algo/bcm_non_local_means_denoising/) with several compuational optimizations expected to applied to gray-scale images.
## 4. Thresholding
Lastly, we converted the cropped, resized, and denoised gray-scale image into black and white image by thresholding. Each pixel of a gray-scale image ranges from 0 to 255. We converted the pixels with 200+ pixel value as black and white for others. We chose 200 to capture the pixels that were most definitely grapheme (even after denoising, we attempted to remove any margin of graphemes with smudge/light-gray gradient). (https://opencv-python-tutroals.readthedocs.io/en/latest/py_tutorials/py_imgproc/py_thresholding/py_thresholding.html)
Below is the code we used to perform resizing, denoising, and thresholding.
```python
num_rows = int(np.median(rows)) # 106
num_cols = int(np.median(cols)) # 87
for i in range(len(train_images)):
images[i] = images[i].resize((num_rows, num_cols), Image.ANTIALIAS)
images[i] = np.array(images[i])
images[i] = cv2.fastNlMeansDenoising(images[i], h=3)
images[i] = cv2.threshold(images[i], 200, 1, cv2.THRESH_BINARY)[1]
```
### Below are the images before and after preprocessing.
#### Bottom 10 Grapheme Roots (Before)

#### Bottom 10 Grapheme Roots (After)

#### Top 5 Vowels (Before)

#### Top 5 Vowels (After)

#### All 7 Consonents (Before)

#### All 7 Consonents (After)

We have that the images were preprocessed such that the salient regions were properly cropped, maintaining the graphemes while cutting out the white spaces. The margin of the graphemes were eliminated and resulted in clearer fonts.
### Next Steps
We believe our preprocessing was satisfactory, although as you can see from above examples, not all images were preprocessed perfectly. We have one more preprocessing to be done, which is augmentation. However, we plan to use the library function provided by Keras. Our next step is to actually build the model with convolutional layers (CNN) using the preprocessed data. Stay put!
| github_jupyter |
# Now You Code 1: Address
Write a Python program to input elements of your postal address and then output them as if they were an address label. The program should use a dictionary to store the address and complete two function defintions one for inputting the address and one for printing the address.
**NOTE:** While you most certainly can write this program without using dictionaries or functions, the point of the exercise is to get used to using them!!!
Sample Run:
```
Enter Street: 314 Hinds Hall
Enter City: Syracuse
Enter State: NY
Enter Postal Zip Code: 13244
Mailing Address:
314 Hinds Hall
Syracuse , NY 13244
```
## Step 1: Problem Analysis `input_address` function
This function should get input from the user at run time and return the input address.
Inputs: None (gets input from user)
Outputs: a Python dictionary of address info (street, city, state, postal_code)
Algorithm (Steps in Program):
```
## Step 2: Write input_address_ function
#input: None (inputs from console)
#output: dictionary of the address
def input_address():
address= {}
# todo: write code here to input the street, city, state and zip code and add to dictionary at runtime and store in a dictionary
street = input('Enter a Street Address: ')
address['street'] = street
city = input('Enter a city: ')
address['city'] = city
state = input('Enter a State: ')
address['state'] = state
zipcode = int(input('Enter a Zipcode: '))
address['zipcode'] = zipcode
return address
address = input_address()
print(address)
print(address.values())
```
## Step 3: Problem Analysis `print_address` function
This function should display a mailing address using the dictionary variable
Inputs: dictionary variable of address into (street, city, state, postal_code)
Outputs: None (prints to screen)
Algorithm (Steps in Program):
```
## Step 4: write code
# input: address dictionary
# output: none (outputs to console)
def print_address(address):
# todo: write code to print the address (leave empty return at the end
print('Mailing Address: ')
print('%s' %(address['street']))
print('%s, %s, %d' % (address['city'], address['state'], address['zipcode']))
return
```
## Step 5: Problem Analysis main program
Should be trivial at this point.
Inputs:
Outputs:
Algorithm (Steps in Program):
```
## Step 6: write main program, use other 2 functions you made to solve this problem.
# main program
# todo: call input_address, then print_address
my_addresses = []
while True:
enter_address = input('Enter "y" if you wish to enter an address if not ENTER to quit')
if enter_address == '':
break
address = input_address()
my_addresses.append(address)
for address in my_addresses:
print_address(address)
```
# Step 7: Questions
1. Explain a strategy for a situation when an expected dictionary key, like 'state' for example does not exist?
2. The program as it is written is not very useful. How can we make it more useful?
## Reminder of Evaluation Criteria
1. What the problem attempted (analysis, code, and answered questions) ?
2. What the problem analysis thought out? (does the program match the plan?)
3. Does the code execute without syntax error?
4. Does the code solve the intended problem?
5. Is the code well written? (easy to understand, modular, and self-documenting, handles errors)
| github_jupyter |
# Convolutions for Images
:label:`sec_conv_layer`
Now that we understand how convolutional layers work in theory,
we are ready to see how they work in practice.
Building on our motivation of convolutional neural networks
as efficient architectures for exploring structure in image data,
we stick with images as our running example.
## The Cross-Correlation Operator
Recall that strictly speaking, *convolutional* layers
are a (slight) misnomer, since the operations they express
are more accurately described as cross correlations.
In a convolutional layer, an input array
and a *correlation kernel* array are combined
to produce an output array through a cross-correlation operation.
Let's ignore channels for now and see how this works
with two-dimensional data and hidden representations.
In :numref:`fig_correlation`,
the input is a two-dimensional array
with a height of 3 and width of 3.
We mark the shape of the array as $3 \times 3$ or ($3$, $3$).
The height and width of the kernel are both $2$.
Note that in the deep learning research community,
this object may be referred to as *a convolutional kernel*,
*a filter*, or simply the layer's *weights*.
The shape of the kernel window
is given by the height and width of the kernel
(here it is $2 \times 2$).

:label:`fig_correlation`
In the two-dimensional cross-correlation operation,
we begin with the convolution window positioned
at the top-left corner of the input array
and slide it across the input array,
both from left to right and top to bottom.
When the convolution window slides to a certain position,
the input subarray contained in that window
and the kernel array are multiplied (elementwise)
and the resulting array is summed up
yielding a single scalar value.
This result gives the value of the output array
at the corresponding location.
Here, the output array has a height of 2 and width of 2
and the four elements are derived from
the two-dimensional cross-correlation operation:
$$
0\times0+1\times1+3\times2+4\times3=19,\\
1\times0+2\times1+4\times2+5\times3=25,\\
3\times0+4\times1+6\times2+7\times3=37,\\
4\times0+5\times1+7\times2+8\times3=43.
$$
Note that along each axis, the output
is slightly *smaller* than the input.
Because the kernel has width and height greater than one,
we can only properly compute the cross-correlation
for locations where the kernel fits wholly within the image,
the output size is given by the input size $H \times W$
minus the size of the convolutional kernel $h \times w$
via $(H-h+1) \times (W-w+1)$.
This is the case since we need enough space
to 'shift' the convolutional kernel across the image
(later we will see how to keep the size unchanged
by padding the image with zeros around its boundary
such that there is enough space to shift the kernel).
Next, we implement this process in the `corr2d` function,
which accepts the input array `X` and kernel array `K`
and returns the output array `Y`.
But first we will import the relevant libraries.
```
%mavenRepo snapshots https://oss.sonatype.org/content/repositories/snapshots/
%maven ai.djl:api:0.7.0-SNAPSHOT
%maven org.slf4j:slf4j-api:1.7.26
%maven org.slf4j:slf4j-simple:1.7.26
%maven net.java.dev.jna:jna:5.3.0
%maven ai.djl.mxnet:mxnet-engine:0.7.0-SNAPSHOT
%maven ai.djl.mxnet:mxnet-native-auto:1.7.0-b
import ai.djl.*;
import ai.djl.engine.Engine;
import ai.djl.ndarray.NDArray;
import ai.djl.ndarray.NDList;
import ai.djl.ndarray.NDManager;
import ai.djl.ndarray.index.NDIndex;
import ai.djl.ndarray.types.DataType;
import ai.djl.ndarray.types.Shape;
import ai.djl.nn.Block;
import ai.djl.nn.ParameterList;
import ai.djl.nn.convolutional.Conv2d;
import ai.djl.training.GradientCollector;
import ai.djl.training.ParameterStore;
import ai.djl.training.initializer.NormalInitializer;
import ai.djl.training.loss.Loss;
public NDArray corr2d(NDArray X, NDArray K){
// Compute 2D cross-correlation.
int h = (int) K.getShape().get(0);
int w = (int) K.getShape().get(1);
NDArray Y = manager.zeros(new Shape(X.getShape().get(0) - h + 1, X.getShape().get(1) - w + 1));
for(int i=0; i < Y.getShape().get(0); i++){
for(int j=0; j < Y.getShape().get(1); j++){
Y.set(new NDIndex(i + "," + j), X.get(i + ":" + (i+h) + "," + j + ":" + (j+w)).mul(K).sum());
}
}
return Y;
}
```
We can construct the input array `X` and the kernel array `K`
from the figure above
to validate the output of the above implementation
of the two-dimensional cross-correlation operation.
```
NDManager manager = NDManager.newBaseManager();
NDArray X = manager.create(new float[]{0,1,2,3,4,5,6,7,8}, new Shape(3,3));
NDArray K = manager.create(new float[]{0,1,2,3}, new Shape(2,2));
System.out.println(corr2d(X, K));
```
## Convolutional Layers
A convolutional layer cross-correlates the input and kernels
and adds a scalar bias to produce an output.
The two parameters of the convolutional layer
are the kernel and the scalar bias.
When training models based on convolutional layers,
we typically initialize the kernels randomly,
just as we would with a fully connected layer.
We are now ready to implement a two-dimensional convolutional layer
based on the `corr2d` function defined above.
In the `ConvolutionalLayer` constructor function,
we declare `weight` and `bias` as the two class parameters.
The forward computation function `forward`
calls the `corr2d` function and adds the bias.
As with $h \times w$ cross-correlation
we also refer to convolutional layers
as $h \times w$ convolutions.
```
public class ConvolutionalLayer{
private NDArray w;
private NDArray b;
public NDArray getW(){
return w;
}
public NDArray getB(){
return b;
}
public ConvolutionalLayer(Shape shape){
NDManager manager = NDManager.newBaseManager();
w = manager.create(shape);
b = manager.randomNormal(new Shape(1));
w.attachGradient();
}
public NDArray forward(NDArray X){
return corr2d(X, w).add(b);
}
}
```
## Object Edge Detection in Images
Let's take a moment to parse a simple application of a convolutional layer:
detecting the edge of an object in an image
by finding the location of the pixel change.
First, we construct an 'image' of $6\times 8$ pixels.
The middle four columns are black (0) and the rest are white (1).
```
X = manager.ones(new Shape(6,8));
X.set(new NDIndex(":" + "," + 2 + ":" + 6), 0f);
System.out.println(X);
```
Next, we construct a kernel `K` with a height of $1$ and width of $2$.
When we perform the cross-correlation operation with the input,
if the horizontally adjacent elements are the same,
the output is 0. Otherwise, the output is non-zero.
```
K = manager.create(new float[]{1, -1}, new Shape(1,2));
```
We are ready to perform the cross-correlation operation
with arguments `X` (our input) and `K` (our kernel).
As you can see, we detect 1 for the edge from white to black
and -1 for the edge from black to white.
All other outputs take value $0$.
```
NDArray Y = corr2d(X, K);
Y
```
We can now apply the kernel to the transposed image.
As expected, it vanishes. The kernel `K` only detects vertical edges.
```
corr2d(X.transpose(), K);
```
## Learning a Kernel
Designing an edge detector by finite differences `[1, -1]` is neat
if we know this is precisely what we are looking for.
However, as we look at larger kernels,
and consider successive layers of convolutions,
it might be impossible to specify
precisely what each filter should be doing manually.
Now let us see whether we can learn the kernel that generated `Y` from `X`
by looking at the (input, output) pairs only.
We first construct a convolutional layer
and initialize its kernel as a random array.
Next, in each iteration, we will use the squared error
to compare `Y` to the output of the convolutional layer.
We can then calculate the gradient to update the weight.
For the sake of simplicity, in this convolutional layer,
we will ignore the bias.
This time, we will use the in-built `Block` and `Conv2d` class from DJL.
```
X = X.reshape(1,1,6,8);
Y = Y.reshape(1,1,6,7);
Loss l2Loss = Loss.l2Loss();
// Construct a two-dimensional convolutional layer with 1 output channel and a
// kernel of shape (1, 2). For the sake of simplicity, we ignore the bias here
Block block = Conv2d.builder()
.setKernelShape(new Shape(1, 2))
.optBias(false)
.setFilters(1)
.build();
block.setInitializer(new NormalInitializer());
block.initialize(manager, DataType.FLOAT32, X.getShape());
// The two-dimensional convolutional layer uses four-dimensional input and
// output in the format of (example, channel, height, width), where the batch
// size (number of examples in the batch) and the number of channels are both 1
ParameterList params = block.getParameters();
NDArray wParam = params.get(0).getValue().getArray();
wParam.attachGradient();
NDArray lossVal = null;
ParameterStore parameterStore = new ParameterStore(manager, false);
NDArray lossVal = null;
for (int i = 0; i < 10; i++) {
wParam.attachGradient();
try (GradientCollector gc = Engine.getInstance().newGradientCollector()) {
NDArray yHat = block.forward(parameterStore, new NDList(X), true).singletonOrThrow();
NDArray l = l2Loss.evaluate(new NDList(Y), new NDList(yHat));
lossVal = l;
gc.backward(l);
}
// Update the kernel
wParam.subi(wParam.getGradient().mul(0.40f));
if((i+1)%2 == 0){
System.out.println("batch " + (i+1) + " loss: " + lossVal.sum().getFloat());
}
}
```
Note that the error has dropped to a small value after 10 iterations. Now we will take a look at the kernel array we learned.
```
ParameterList params = block.getParameters();
NDArray wParam = params.get(0).getValue().getArray();
wParam
```
Indeed, the learned kernel array is moving close
to the kernel array `K` we defined earlier.
## Cross-Correlation and Convolution
Recall our observation from the previous section of the correspondence
between the cross-correlation and convolution operators.
The figure above makes this correspondence apparent.
Simply flip the kernel from the bottom left to the top right.
In this case, the indexing in the sum is reverted,
yet the same result can be obtained.
In keeping with standard terminology with deep learning literature,
we will continue to refer to the cross-correlation operation
as a convolution even though, strictly-speaking, it is slightly different.
## Summary
* The core computation of a two-dimensional convolutional layer is a two-dimensional cross-correlation operation. In its simplest form, this performs a cross-correlation operation on the two-dimensional input data and the kernel, and then adds a bias.
* We can design a kernel to detect edges in images.
* We can learn the kernel's parameters from data.
## Exercises
1. Construct an image `X` with diagonal edges.
* What happens if you apply the kernel `K` to it?
* What happens if you transpose `X`?
* What happens if you transpose `K`?
1. When you try to automatically find the gradient for the `Conv2d` class we created, what kind of error message do you see?
1. How do you represent a cross-correlation operation as a matrix multiplication by changing the input and kernel arrays?
1. Design some kernels manually.
* What is the form of a kernel for the second derivative?
* What is the kernel for the Laplace operator?
* What is the kernel for an integral?
* What is the minimum size of a kernel to obtain a derivative of degree $d$?
| github_jupyter |
# Iterables
Some steps in a neuroimaging analysis are repetitive. Running the same preprocessing on multiple subjects or doing statistical inference on multiple files. To prevent the creation of multiple individual scripts, Nipype has as execution plugin for ``Workflow``, called **``iterables``**.
<img src="../static/images/iterables.png" width="240">
If you are interested in more advanced procedures, such as synchronizing multiple iterables or using conditional iterables, check out the `synchronize `and `intersource` section in the [`JoinNode`](basic_joinnodes.ipynb) notebook.
## Realistic example
Let's assume we have a workflow with two nodes, node (A) does simple skull stripping, and is followed by a node (B) that does isometric smoothing. Now, let's say, that we are curious about the effect of different smoothing kernels. Therefore, we want to run the smoothing node with FWHM set to 2mm, 8mm, and 16mm.
```
from nipype import Node, Workflow
from nipype.interfaces.fsl import BET, IsotropicSmooth
# Initiate a skull stripping Node with BET
skullstrip = Node(BET(mask=True,
in_file='/data/ds000114/sub-01/ses-test/anat/sub-01_ses-test_T1w.nii.gz'),
name="skullstrip")
```
Create a smoothing Node with IsotropicSmooth
```
isosmooth = Node(IsotropicSmooth(), name='iso_smooth')
```
Now, to use ``iterables`` and therefore smooth with different ``fwhm`` is as simple as that:
```
isosmooth.iterables = ("fwhm", [4, 8, 16])
```
And to wrap it up. We need to create a workflow, connect the nodes and finally, can run the workflow in parallel.
```
# Create the workflow
wf = Workflow(name="smoothflow")
wf.base_dir = "/output"
wf.connect(skullstrip, 'out_file', isosmooth, 'in_file')
# Run it in parallel (one core for each smoothing kernel)
wf.run('MultiProc', plugin_args={'n_procs': 3})
```
**Note**, that ``iterables`` is set on a specific node (``isosmooth`` in this case), but ``Workflow`` is needed to expend the graph to three subgraphs with three different versions of the ``isosmooth`` node.
If we visualize the graph with ``exec``, we can see where the parallelization actually takes place.
```
# Visualize the detailed graph
from IPython.display import Image
wf.write_graph(graph2use='exec', format='png', simple_form=True)
Image(filename='/output/smoothflow/graph_detailed.png')
```
If you look at the structure in the workflow directory, you can also see, that for each smoothing, a specific folder was created, i.e. ``_fwhm_16``.
```
!tree /output/smoothflow -I '*txt|*pklz|report*|*.json|*js|*.dot|*.html'
```
Now, let's visualize the results!
```
from nilearn import plotting
%matplotlib inline
plotting.plot_anat(
'/data/ds000114/sub-01/ses-test/anat/sub-01_ses-test_T1w.nii.gz', title='original',
display_mode='z', dim=-1, cut_coords=(-50, -35, -20, -5), annotate=False);
plotting.plot_anat(
'/output/smoothflow/skullstrip/sub-01_ses-test_T1w_brain.nii.gz', title='skullstripped',
display_mode='z', dim=-1, cut_coords=(-50, -35, -20, -5), annotate=False);
plotting.plot_anat(
'/output/smoothflow/_fwhm_4/iso_smooth/sub-01_ses-test_T1w_brain_smooth.nii.gz', title='FWHM=4',
display_mode='z', dim=-0.5, cut_coords=(-50, -35, -20, -5), annotate=False);
plotting.plot_anat(
'/output/smoothflow/_fwhm_8/iso_smooth/sub-01_ses-test_T1w_brain_smooth.nii.gz', title='FWHM=8',
display_mode='z', dim=-0.5, cut_coords=(-50, -35, -20, -5), annotate=False);
plotting.plot_anat(
'/output/smoothflow/_fwhm_16/iso_smooth/sub-01_ses-test_T1w_brain_smooth.nii.gz', title='FWHM=16',
display_mode='z', dim=-0.5, cut_coords=(-50, -35, -20, -5), annotate=False);
```
# ``IdentityInterface`` (special use case of ``iterables``)
We often want to start our worflow from creating subgraphs, e.g. for running preprocessing for all subjects. We can easily do it with setting ``iterables`` on the ``IdentityInterface``. The ``IdentityInterface`` interface allows you to create ``Nodes`` that does simple identity mapping, i.e. ``Nodes`` that only work on parameters/strings.
For example, you want to start your workflow by collecting anatomical files for 5 subjects.
```
# First, let's specify the list of subjects
subject_list = ['01', '02', '03', '07']
```
Now, we can create the IdentityInterface Node
```
from nipype import IdentityInterface
infosource = Node(IdentityInterface(fields=['subject_id']),
name="infosource")
infosource.iterables = [('subject_id', subject_list)]
```
That's it. Now, we can connect the output fields of this ``infosource`` node to ``SelectFiles`` and ``DataSink`` nodes.
```
from os.path import join as opj
from nipype.interfaces.io import SelectFiles, DataSink
anat_file = opj('sub-{subject_id}', 'ses-test', 'anat', 'sub-{subject_id}_ses-test_T1w.nii.gz')
templates = {'anat': anat_file}
selectfiles = Node(SelectFiles(templates,
base_directory='/data/ds000114'),
name="selectfiles")
# Datasink - creates output folder for important outputs
datasink = Node(DataSink(base_directory="/output",
container="datasink"),
name="datasink")
wf_sub = Workflow(name="choosing_subjects")
wf_sub.connect(infosource, "subject_id", selectfiles, "subject_id")
wf_sub.connect(selectfiles, "anat", datasink, "anat_files")
wf_sub.run()
```
Now we can check that five anatomicl images are in ``anat_files`` directory:
```
! ls -lh /output/datasink/anat_files/
```
This was just a simple example of using ``IdentityInterface``, but a complete example of preprocessing workflow you can find in [Preprocessing Example](example_preprocessing.ipynb)).
## Exercise 1
Create a workflow to calculate various powers of ``2`` using two nodes, one for ``IdentityInterface`` with ``iterables``, and one for ``Function`` interface to calculate the power of ``2``.
```
# write your solution here
# lets start from the Identity node
from nipype import Function, Node, Workflow
from nipype.interfaces.utility import IdentityInterface
iden = Node(IdentityInterface(fields=['number']), name="identity")
iden.iterables = [("number", range(8))]
# the second node should use the Function interface
def power_of_two(n):
return 2**n
# Create Node
power = Node(Function(input_names=["n"],
output_names=["pow"],
function=power_of_two),
name='power')
#and now the workflow
wf_ex1 = Workflow(name="exercise1")
wf_ex1.connect(iden, "number", power, "n")
res_ex1 = wf_ex1.run()
# we can print the results
for i in range(8):
print(list(res_ex1.nodes())[i].result.outputs)
```
| github_jupyter |
# WeatherPy
----
#### Note
* Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
```
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import time
import scipy.stats as st
from scipy.stats import linregress
# Import API key
from api_keys import weather_api_key
# Incorporated citipy to determine city based on latitude and longitude
from citipy import citipy
# Output File (CSV)
output_data_file = "output_data/cities.csv"
# Range of latitudes and longitudes
lat_range = (-90, 90)
lng_range = (-180, 180)
```
## Generate Cities List
```
# List for holding lat_lngs and cities
lat_lngs = []
cities = []
# Create a set of random lat and lng combinations
lats = np.random.uniform(lat_range[0], lat_range[1], size=1500)
lngs = np.random.uniform(lng_range[0], lng_range[1], size=1500)
lat_lngs = zip(lats, lngs)
# Identify nearest city for each lat, lng combination
for lat_lng in lat_lngs:
city = citipy.nearest_city(lat_lng[0], lat_lng[1]).city_name
# If the city is unique, then add it to a our cities list
if city not in cities:
cities.append(city)
# Print the city count to confirm sufficient count
len(cities)
```
### Perform API Calls
* Perform a weather check on each city using a series of successive API calls.
* Include a print log of each city as it'sbeing processed (with the city number and city name).
```
# Save config information.
url = "http://api.openweathermap.org/data/2.5/weather?"
units = "imperial"
# Build partial query URL
base_url = f"{url}appid={weather_api_key}&units={units}&q="
base_url
query_url = base_url + city
query_url
response = requests.get(base_url).json()
# List of City data
lat = []
lon = []
temp = []
humidity = []
cloudy = []
windspee = []
count = 0
cityname = []
dt = []
# Get weather data
print("Beginning Data Retrival")
print("-----------------------------")
set_count = 1
record_count = 1
weather_data = {"City":[],"Lat":[],"Lng":[],"Max Temp":[],"Humidity":[],"Cloudiness":[],"Wind Speed": [],"Country":[],"Date":[]}
for city in cities:
weather_response = requests.get(base_url + city)
weather_response_json = weather_response.json()
if weather_response.status_code == 200:
weather_data["City"].append(city)
weather_data["Lat"].append(weather_response_json['coord']['lat'])
weather_data["Lng"].append(weather_response_json['coord']['lon'])
weather_data["Max Temp"].append(weather_response_json['main']['temp_max'])
weather_data["Humidity"].append(weather_response_json['main']['humidity'])
weather_data["Cloudiness"].append(weather_response_json['clouds']['all'])
weather_data["Wind Speed"].append(weather_response_json['wind']['speed'])
weather_data["Country"].append(weather_response_json['sys']['country'])
weather_data["Date"].append(weather_response_json['dt'])
if record_count <= 50:
print(f"Processing Record {record_count} of Set {set_count} | {city}")
record_count += 1
else:
record_count = 0
set_count += 1
print(f"Processing Record {record_count} of Set {set_count} | {city}")
record_count += 1
else:
print("City not found. Skipping...")
print("-----------------------------")
print("Data Retrieval Complete")
print("-----------------------------")
```
### Convert Raw Data to DataFrame
* Export the city data into a .csv.
* Display the DataFrame
```
weather_df = pd.DataFrame(weather_data)
weather_df.head()
weather_df.count()
# Export data to csv
weather_df.to_csv('weather_data.csv', index=False)
```
## Inspect the data and remove the cities where the humidity > 100%.
----
Skip this step if there are no cities that have humidity > 100%.
```
weather_df.describe()
# No Cities with Humidity >100%
```
## Plotting the Data
* Use proper labeling of the plots using plot titles (including date of analysis) and axes labels.
* Save the plotted figures as .pngs.
## Latitude vs. Temperature Plot
```
# Scatter plot
plt.scatter(weather_df["Lat"],weather_df["Max Temp"], edgecolor="black")
# Labels
plt.title("City Latitude vs. Max Temperature (06/18/20)")
plt.xlabel("Latitude")
plt.ylabel("Max Temperature (F)")
plt.grid(True)
# Save as PNG and Show
plt.savefig("LatVTemp.png")
plt.show()
```
## Latitude vs. Humidity Plot
```
# Scatter plot
plt.scatter(weather_df["Lat"],weather_df["Humidity"], edgecolor="black")
# Labels
plt.title("City Latitude vs. Humidity (06/18/20)")
plt.xlabel("Latitude")
plt.ylabel("Humidity (%)")
plt.grid(True)
# Save as PNG and Show
plt.savefig("LatVHumidity.png")
plt.show()
```
## Latitude vs. Cloudiness Plot
```
# Scatter plot
plt.scatter(weather_df["Lat"],weather_df["Cloudiness"], edgecolor="black")
# Labels
plt.title("City Latitude vs. Cloudiness (06/18/20)")
plt.xlabel("Latitude")
plt.ylabel("Cloudiness (%)")
plt.grid(True)
# Save as PNG and Show
plt.savefig("LatVCloud.png")
plt.show()
```
## Latitude vs. Wind Speed Plot
```
# Scatter plot
plt.scatter(weather_df["Lat"],weather_df["Wind Speed"], edgecolor="black")
# Labels
plt.title("City Latitude vs. Wind Speed (08/22/18)")
plt.xlabel("Latitude")
plt.ylabel("Wind Speed (mph)")
plt.grid(True)
# Save as PNG and Show
plt.savefig("LatVWindSpeed.png")
plt.show()
```
## Linear Regression
```
# OPTIONAL: Create a function to create Linear Regression plots
# Create Northern and Southern Hemisphere DataFrames
n_hemisphere = weather_df.loc[weather_df["Lat"] >= 0]
s_hemisphere = weather_df.loc[weather_df["Lat"] < 0]
n_hemisphere.head()
n_hemisphere.describe()
s_hemisphere.head()
s_hemisphere.describe()
```
#### Northern Hemisphere - Max Temp vs. Latitude Linear Regression
```
# Calculate the correlation coefficient and linear regression model
(slope, intercept,rvalue, pvalue, stderr) = linregress(n_hemisphere["Lat"], n_hemisphere["Max Temp"])
regress_values = n_hemisphere["Lat"] * slope + intercept
# Scatter Plot
x_axis = n_hemisphere["Lat"]
y_axis = n_hemisphere["Max Temp"]
plt.scatter(x_axis, y_axis, edgecolor="black")
# Regression Line
plt.plot(n_hemisphere["Lat"], regress_values, color='red')
line_eq = f"y = {round(slope, 2)}x + {round(intercept, 2)}"
r_value = f"r = {round(rvalue,2)}"
plt.annotate(line_eq, (5,35), color='red', fontsize=10)
plt.annotate(r_value, (5,32), color='red', fontsize=10)
# Labels
plt.title("Max Temp vs. Latitude (Northern Hemisphere)")
plt.xlabel("Latitude")
plt.ylabel("Max Temp (F)")
print(f"The r-squared is: {rvalue}")
plt.savefig("TempVLat-NHemisphere.png")
plt.show()
```
#### Southern Hemisphere - Max Temp vs. Latitude Linear Regression
```
# Calculate the correlation coefficient and linear regression model
(slope, intercept,rvalue, pvalue, stderr) = linregress(s_hemisphere["Lat"], s_hemisphere["Max Temp"])
regress_values = s_hemisphere["Lat"] * slope + intercept
# Scatter Plot
x_axis = s_hemisphere["Lat"]
y_axis = s_hemisphere["Max Temp"]
plt.scatter(x_axis, y_axis, edgecolor="black")
# Regression Line
plt.plot(s_hemisphere["Lat"], regress_values, color='red')
line_eq = f"y = {round(slope, 2)}x + {round(intercept, 2)}"
r_value = f"r = {round(rvalue,2)}"
plt.annotate(line_eq, (-50,80), color='red', fontsize=10)
plt.annotate(r_value, (-50,75), color='red', fontsize=10)
# Labels
plt.title("Max Temp vs. Latitude (Southern Hemisphere)")
plt.xlabel("Latitude")
plt.ylabel("Max Temp (F)")
print(f"The r-squared is: {rvalue}")
plt.savefig("TempVLat-SHemisphere.png")
plt.show()
```
#### Northern Hemisphere - Humidity (%) vs. Latitude Linear Regression
```
# Calculate the correlation coefficient and linear regression model
(slope, intercept,rvalue, pvalue, stderr) = linregress(n_hemisphere["Lat"], n_hemisphere["Humidity"])
regress_values = n_hemisphere["Lat"] * slope + intercept
# Scatter Plot
x_axis = n_hemisphere["Lat"]
y_axis = n_hemisphere["Humidity"]
plt.scatter(x_axis, y_axis, edgecolor="black")
# Regression Line
plt.plot(n_hemisphere["Lat"], regress_values, color='red')
line_eq = f"y = {round(slope, 2)}x + {round(intercept, 2)}"
r_value = f"r = {round(rvalue,2)}"
plt.annotate(line_eq, (0,20), color='red', fontsize=10)
plt.annotate(r_value, (0,10), color='red', fontsize=10)
# Labels
plt.title("Max Temp vs. Humidity (Northern Hemisphere)")
plt.xlabel("Latitude")
plt.ylabel("Max Temp (F)")
print(f"The r-squared is: {rvalue}")
plt.savefig("HumidityVLat-NHemisphere.png")
plt.show()
```
#### Southern Hemisphere - Humidity (%) vs. Latitude Linear Regression
```
# Calculate the correlation coefficient and linear regression model
(slope, intercept,rvalue, pvalue, stderr) = linregress(s_hemisphere["Lat"], s_hemisphere["Humidity"])
regress_values = s_hemisphere["Lat"] * slope + intercept
# Scatter Plot
x_axis = s_hemisphere["Lat"]
y_axis = s_hemisphere["Humidity"]
plt.scatter(x_axis, y_axis, edgecolor="black")
# Regression Line
plt.plot(s_hemisphere["Lat"], regress_values, color='red')
line_eq = f"y = {round(slope, 2)}x + {round(intercept, 2)}"
r_value = f"r = {round(rvalue,2)}"
plt.annotate(line_eq, (-55,35), color='red', fontsize=10)
plt.annotate(r_value, (-55,30), color='red', fontsize=10)
# Labels
plt.title("Max Temp vs. Humidity (Southern Hemisphere)")
plt.xlabel("Latitude")
plt.ylabel("Max Temp (F)")
print(f"The r-squared is: {rvalue}")
plt.savefig("HumidityVLat-SHemisphere.png")
plt.show()
```
#### Northern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
```
# Calculate the correlation coefficient and linear regression model
(slope, intercept,rvalue, pvalue, stderr) = linregress(n_hemisphere["Lat"], n_hemisphere["Cloudiness"])
regress_values = n_hemisphere["Lat"] * slope + intercept
# Scatter Plot
x_axis = n_hemisphere["Lat"]
y_axis = n_hemisphere["Cloudiness"]
plt.scatter(x_axis, y_axis, edgecolor="black")
# Regression Line
plt.plot(n_hemisphere["Lat"], regress_values, color='red')
line_eq = f"y = {round(slope, 2)}x + {round(intercept, 2)}"
r_value = f"r = {round(rvalue,2)}"
plt.annotate(line_eq, (25,55), color='red', fontsize=10)
plt.annotate(r_value, (25,50), color='red', fontsize=10)
# Labels
plt.title("Max Temp vs. Cloudiness (Northern Hemisphere)")
plt.xlabel("Latitude")
plt.ylabel("Max Temp (F)")
print(f"The r-squared is: {rvalue}")
plt.savefig("CloudinessVLat-NHemisphere.png")
plt.show()
```
#### Southern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
```
# Calculate the correlation coefficient and linear regression model
(slope, intercept,rvalue, pvalue, stderr) = linregress(s_hemisphere["Lat"], s_hemisphere["Cloudiness"])
regress_values = s_hemisphere["Lat"] * slope + intercept
# Scatter Plot
x_axis = s_hemisphere["Lat"]
y_axis = s_hemisphere["Cloudiness"]
plt.scatter(x_axis, y_axis, edgecolor="black")
# Regression Line
plt.plot(s_hemisphere["Lat"], regress_values, color='red')
line_eq = f"y = {round(slope, 2)}x + {round(intercept, 2)}"
r_value = f"r = {round(rvalue,2)}"
plt.annotate(line_eq, (-55,50), color='red', fontsize=10)
plt.annotate(r_value, (-55,40), color='red', fontsize=10)
# Labels
plt.title("Max Temp vs. Cloudiness (Southern Hemisphere)")
plt.xlabel("Latitude")
plt.ylabel("Max Temp (F)")
print(f"The r-squared is: {rvalue}")
plt.savefig("CloudinessVLat-SHemisphere.png")
plt.show()
```
#### Northern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
```
# Calculate the correlation coefficient and linear regression model
(slope, intercept,rvalue, pvalue, stderr) = linregress(n_hemisphere["Lat"], n_hemisphere["Wind Speed"])
regress_values = n_hemisphere["Lat"] * slope + intercept
# Scatter Plot
x_axis = n_hemisphere["Lat"]
y_axis = n_hemisphere["Wind Speed"]
plt.scatter(x_axis, y_axis, edgecolor="black")
# Regression Line
plt.plot(n_hemisphere["Lat"], regress_values, color='red')
line_eq = f"y = {round(slope, 2)}x + {round(intercept, 2)}"
r_value = f"r = {round(rvalue,2)}"
plt.annotate(line_eq, (0,35), color='red', fontsize=10)
plt.annotate(r_value, (0,32), color='red', fontsize=10)
# Labels
plt.title("Max Temp vs. Wind Speed (Northern Hemisphere)")
plt.xlabel("Latitude")
plt.ylabel("Max Temp (F)")
print(f"The r-squared is: {rvalue}")
plt.savefig("WindVLat-NHemisphere.png")
plt.show()
```
#### Southern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
```
# Calculate the correlation coefficient and linear regression model
(slope, intercept,rvalue, pvalue, stderr) = linregress(s_hemisphere["Lat"], s_hemisphere["Wind Speed"])
regress_values = s_hemisphere["Lat"] * slope + intercept
# Scatter Plot
x_axis = s_hemisphere["Lat"]
y_axis = s_hemisphere["Wind Speed"]
plt.scatter(x_axis, y_axis, edgecolor="black")
# Regression Line
plt.plot(s_hemisphere["Lat"], regress_values, color='red')
line_eq = f"y = {round(slope, 2)}x + {round(intercept, 2)}"
r_value = f"r = {round(rvalue,2)}"
plt.annotate(line_eq, (-25,23), color='red', fontsize=10)
plt.annotate(r_value, (-25,21), color='red', fontsize=10)
# Labels
plt.title("Max Temp vs. Wind Speed (Southern Hemisphere)")
plt.xlabel("Latitude")
plt.ylabel("Max Temp (F)")
print(f"The r-squared is: {rvalue}")
plt.savefig("WindVLat-SHemisphere.png")
plt.show()
```
| github_jupyter |
# Self-Driving Car Engineer Nanodegree
## Deep Learning
## Project: Build a Traffic Sign Recognition Classifier
In this notebook, a template is provided for you to implement your functionality in stages, which is required to successfully complete this project. If additional code is required that cannot be included in the notebook, be sure that the Python code is successfully imported and included in your submission if necessary.
> **Note**: Once you have completed all of the code implementations, you need to finalize your work by exporting the iPython Notebook as an HTML document. Before exporting the notebook to html, all of the code cells need to have been run so that reviewers can see the final implementation and output. You can then export the notebook by using the menu above and navigating to \n",
"**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
In addition to implementing code, there is a writeup to complete. The writeup should be completed in a separate file, which can be either a markdown file or a pdf document. There is a [write up template](https://github.com/udacity/CarND-Traffic-Sign-Classifier-Project/blob/master/writeup_template.md) that can be used to guide the writing process. Completing the code template and writeup template will cover all of the [rubric points](https://review.udacity.com/#!/rubrics/481/view) for this project.
The [rubric](https://review.udacity.com/#!/rubrics/481/view) contains "Stand Out Suggestions" for enhancing the project beyond the minimum requirements. The stand out suggestions are optional. If you decide to pursue the "stand out suggestions", you can include the code in this Ipython notebook and also discuss the results in the writeup file.
>**Note:** Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. In addition, Markdown cells can be edited by typically double-clicking the cell to enter edit mode.
---
## Step 0: Load The Data
```
# Load pickled data
import pickle
# TODO: Fill this in based on where you saved the training and testing data
training_file = "./traffic-signs-data/train.p"
validation_file= "./traffic-signs-data/valid.p"
testing_file = "./traffic-signs-data/test.p"
with open(training_file, mode='rb') as f:
train = pickle.load(f)
with open(validation_file, mode='rb') as f:
valid = pickle.load(f)
with open(testing_file, mode='rb') as f:
test = pickle.load(f)
X_train, y_train = train['features'], train['labels']
X_valid, y_valid = valid['features'], valid['labels']
X_test, y_test = test['features'], test['labels']
# Print the size of datasets
print("X_train shape:", X_train.shape)
print("y_train shape:", y_train.shape)
print("X_valid shape:", X_valid.shape)
print("y_valid shape:", y_valid.shape)
print("X_test shape:", X_test.shape)
print("y_test shape:", y_test.shape)
```
---
## Step 1: Dataset Summary & Exploration
The pickled data is a dictionary with 4 key/value pairs:
- `'features'` is a 4D array containing raw pixel data of the traffic sign images, (num examples, width, height, channels).
- `'labels'` is a 1D array containing the label/class id of the traffic sign. The file `signnames.csv` contains id -> name mappings for each id.
- `'sizes'` is a list containing tuples, (width, height) representing the original width and height the image.
- `'coords'` is a list containing tuples, (x1, y1, x2, y2) representing coordinates of a bounding box around the sign in the image. **THESE COORDINATES ASSUME THE ORIGINAL IMAGE. THE PICKLED DATA CONTAINS RESIZED VERSIONS (32 by 32) OF THESE IMAGES**
Complete the basic data summary below. Use python, numpy and/or pandas methods to calculate the data summary rather than hard coding the results. For example, the [pandas shape method](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.shape.html) might be useful for calculating some of the summary results.
### Provide a Basic Summary of the Data Set Using Python, Numpy and/or Pandas
```
### Replace each question mark with the appropriate value.
### Use python, pandas or numpy methods rather than hard coding the results
import numpy as np
# TODO: Number of training examples
n_train = len(X_train)
# TODO: Number of validation examples
n_validation = len(X_valid)
# TODO: Number of testing examples.
n_test = len(X_test)
# TODO: What's the shape of an traffic sign image?
image_shape = X_train[0].shape
print(y_train, y_train.shape)
# TODO: How many unique classes/labels there are in the dataset.
n_classes = len(np.unique(y_train))
print("Number of training examples =", n_train)
print("Number of validation examples", n_validation)
print("Number of testing examples =", n_test)
print("Image data shape =", image_shape)
print("Number of classes =", n_classes)
```
### Include an exploratory visualization of the dataset
Visualize the German Traffic Signs Dataset using the pickled file(s). This is open ended, suggestions include: plotting traffic sign images, plotting the count of each sign, etc.
The [Matplotlib](http://matplotlib.org/) [examples](http://matplotlib.org/examples/index.html) and [gallery](http://matplotlib.org/gallery.html) pages are a great resource for doing visualizations in Python.
**NOTE:** It's recommended you start with something simple first. If you wish to do more, come back to it after you've completed the rest of the sections. It can be interesting to look at the distribution of classes in the training, validation and test set. Is the distribution the same? Are there more examples of some classes than others?
```
### Data exploration visualization code goes here.
### Feel free to use as many code cells as needed.
import matplotlib.pyplot as plt
# Visualizations will be shown in the notebook.
%matplotlib inline
import random
# 4 images to be shown
fig, axs = plt.subplots(1, 4, figsize=(15, 6))
fig.subplots_adjust(hspace = 1, wspace=.01)
axs = axs.ravel()
for i in range(4):
index = random.randint(0, len(X_train))
image = X_train[index]
axs[i].axis('off')
axs[i].imshow(image)
axs[i].set_title(y_train[index])
# Label frequency histogram
histogram, bins = np.histogram(y_train, bins = n_classes)
width = .5 * (bins[1] - bins[0])
center = (bins[:-1] + bins[1:]) / 2
plt.bar(center, histogram, align='center', width=width)
plt.show()
```
----
## Step 2: Design and Test a Model Architecture
Design and implement a deep learning model that learns to recognize traffic signs. Train and test your model on the [German Traffic Sign Dataset](http://benchmark.ini.rub.de/?section=gtsrb&subsection=dataset).
The LeNet-5 implementation shown in the [classroom](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) at the end of the CNN lesson is a solid starting point. You'll have to change the number of classes and possibly the preprocessing, but aside from that it's plug and play!
With the LeNet-5 solution from the lecture, you should expect a validation set accuracy of about 0.89. To meet specifications, the validation set accuracy will need to be at least 0.93. It is possible to get an even higher accuracy, but 0.93 is the minimum for a successful project submission.
There are various aspects to consider when thinking about this problem:
- Neural network architecture (is the network over or underfitting?)
- Play around preprocessing techniques (normalization, rgb to grayscale, etc)
- Number of examples per label (some have more than others).
- Generate fake data.
Here is an example of a [published baseline model on this problem](http://yann.lecun.com/exdb/publis/pdf/sermanet-ijcnn-11.pdf). It's not required to be familiar with the approach used in the paper but, it's good practice to try to read papers like these.
### Pre-process the Data Set (normalization, grayscale, etc.)
Minimally, the image data should be normalized so that the data has mean zero and equal variance. For image data, `(pixel - 128)/ 128` is a quick way to approximately normalize the data and can be used in this project.
Other pre-processing steps are optional. You can try different techniques to see if it improves performance.
Use the code cell (or multiple code cells, if necessary) to implement the first step of your project.
```
### Preprocess the data here. It is required to normalize the data. Other preprocessing steps could include
### converting to grayscale, etc.
### Feel free to use as many code cells as needed.
X_train_rgb = X_train
X_train_gry = np.sum(X_train/3, axis=3, keepdims=True)
X_valid_rgb = X_valid
X_valid_gry = np.sum(X_valid/3, axis=3, keepdims=True)
X_test_rgb = X_test
X_test_gry = np.sum(X_test/3, axis=3, keepdims=True)
print('RGB: ', X_train_rgb.shape)
print('Grayscale: ', X_train_gry.shape)
X_train = X_train_gry
X_valid = X_valid_gry
X_test = X_test_gry
# Visualization
n_rows = 2
n_cols = 4
offset = 1000
fig, axs = plt.subplots(n_rows, n_cols, figsize=(18, 14))
fig.subplots_adjust(hspace = 0.01, wspace = 0.01)
axs = axs.ravel()
for j in range(0, n_rows, 2):
for i in range(n_cols):
index = i + j * n_cols
image = X_train_rgb[index + offset]
axs[index].axis('off')
axs[index].imshow(image)
for i in range(n_cols):
index = i + j * n_cols + n_cols
image = X_train_gry[index + offset - n_cols].squeeze()
axs[index].axis('off')
axs[index].imshow(image, cmap='gray')
```
### Model Architecture
```
### Define your architecture here.
### Feel free to use as many code cells as needed.
# Setup Tensorflow
import tensorflow as tf
EPOCHS = 10
BATCH_SIZE = 50
# Implementing LeNet Architecture
from tensorflow.contrib.layers import flatten
def LeNet(x):
# Arguments used for tf.truncated_normal, randomly defines variables for the weights and biases for each layer
mu = 0
sigma = 0.1
# TODO: Layer 1: Convolutional. Input = 32x32x1. Output = 28x28x6.
conv1_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 1, 6), mean = mu, stddev = sigma))
conv1_b = tf.Variable(tf.zeros(6))
conv1 = tf.nn.conv2d(x, conv1_W, strides=[1, 1, 1, 1], padding='VALID') + conv1_b
# TODO: Activation.
conv1 = tf.nn.relu(conv1)
# TODO: Pooling. Input = 28x28x6. Output = 14x14x6.
conv1 = tf.nn.max_pool(conv1, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
# TODO: Layer 2: Convolutional. Output = 10x10x16.
conv2_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 6, 16), mean = mu, stddev = sigma))
conv2_b = tf.Variable(tf.zeros(16))
conv2 = tf.nn.conv2d(conv1, conv2_W, strides=[1, 1, 1, 1], padding='VALID') + conv2_b
# TODO: Activation.
conv2 = tf.nn.relu(conv2)
# TODO: Pooling. Input = 10x10x16. Output = 5x5x16.
conv2 = tf.nn.max_pool(conv2, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
# TODO: Flatten. Input = 5x5x16. Output = 400.
fc0 = flatten(conv2)
# TODO: Layer 3: Fully Connected. Input = 400. Output = 120.
fc1_W = tf.Variable(tf.truncated_normal(shape=(400, 120), mean = mu, stddev = sigma))
fc1_b = tf.Variable(tf.zeros(120))
fc1 = tf.matmul(fc0, fc1_W) + fc1_b
# TODO: Activation.
fc1 = tf.nn.relu(fc1)
# TODO: Layer 4: Fully Connected. Input = 120. Output = 84.
fc2_W = tf.Variable(tf.truncated_normal(shape=(120, 84), mean = mu, stddev = sigma))
fc2_b = tf.Variable(tf.zeros(84))
fc2 = tf.matmul(fc1, fc2_W) + fc2_b
# TODO: Activation.
fc2 = tf.nn.relu(fc2)
# TODO: Layer 5: Fully Connected. Input = 84. Output = 10.
fc3_W = tf.Variable(tf.truncated_normal(shape=(84, 10), mean = mu, stddev = sigma))
fc3_b = tf.Variable(tf.zeros(10))
logits = tf.matmul(fc2, fc3_W) + fc3_b
return logits
tf.reset_default_graph()
x = tf.placeholder(tf.float32, (None, 32, 32, 1))
y = tf.placeholder(tf.int32, (None))
keep_prob = tf.placeholder(tf.float32)
one_hot_y = tf.one_hot(y, 43)
```
### Train, Validate and Test the Model
A validation set can be used to assess how well the model is performing. A low accuracy on the training and validation
sets imply underfitting. A high accuracy on the training set but low accuracy on the validation set implies overfitting.
```
### Train your model here.
### Calculate and report the accuracy on the training and validation set.
### Once a final model architecture is selected,
### the accuracy on the test set should be calculated and reported as well.
### Feel free to use as many code cells as needed.
rate = 0.0009
logits = LeNet(x)
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(logits, one_hot_y)
loss_operation = tf.reduce_meain(cross_entropy)
optimizer = tf.train.AdamOptimizer(learning_rate = rate)
training_operation = operation.minimize(loss_operation)
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(one_hot_y, 1))
accuracy_operation = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
saver = tf.train.saver()
def evaluate_data(X_data, y_data):
num_examples = len(X_data)
total_accuracy = 0
sess = tf.get_default_session()
for offset in range(0, num_examples, BATCH_SIZE):
batch_x, batch_y = X_data[offset: offset + BATCH_SIZE], y_data[offset: offset + BATCH_size]
accuracy = sess.run(accuracy_operation, feed_dict={x: batch_x, y: batch_y, keep_prob: 1.0})
total_accuracy += (accuracy * len(batch_x))
return total_accuracy / num_examples
print('done')
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
num_examples = len(X_train)
for i in range(EPOCHS):
X_train, y_train = shuffle(X_train, y_train)
for offset in range(0, num_examples, BATCH_SIZE):
end = offset + BATCH_SIZE
batch_x, batch_y = X_train[offset: end], y_train[offset: end]
sess.run(training_operation, feed_dict={x: batch_x, y: batch_y, keep_prob: 0.5})
validation_accuracy = evaluate(X_validation, y_validation)
saver.save(sess, 'lenet')
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
saver2 = tf.train.import_meta_graph('./lenet.meta')
saver2.restore(sess, "./lenet")
test_accuracy = evaluate(X_test_normalized, y_test)
```
---
## Step 3: Test a Model on New Images
To give yourself more insight into how your model is working, download at least five pictures of German traffic signs from the web and use your model to predict the traffic sign type.
You may find `signnames.csv` useful as it contains mappings from the class id (integer) to the actual sign name.
### Load and Output the Images
```
### Load the images and plot them here.
### Feel free to use as many code cells as needed.
import matplotlib.pyplot as plt
%matplotlib inline
import tensorflow as tf
import numpy as np
import cv2
import glob
import matplotlib.image as mpimg
fig, axs = plt.subplots(2, 4, figsize=(4, 2))
fig.subplots_adjust(hspace = 0.2, wspace = 0.001)
axs = axs.ravel()
my_images = []
for i, img in enumerate(glob.glob('./some/*x.png')):
image = cv2.imread(img)
axs[i].axis('off')
axs[i].imshow(cv2.cvtColor(image, cv2.COLOR_BGR2RGB))
my_images.append(image)
my_images = np.asarray(my_images)
my_images_gry = np.sum(my_images/3, axis=3, keepdims=True)
my_images_normalized = (my_images_gry - 128)/128
```
### Predict the Sign Type for Each Image
```
### Run the predictions here and use the model to output the prediction for each image.
### Make sure to pre-process the images with the same pre-processing pipeline used earlier.
### Feel free to use as many code cells as needed.
```
### Analyze Performance
```
### Calculate the accuracy for these 5 new images.
### For example, if the model predicted 1 out of 5 signs correctly, it's 20% accurate on these new images.
```
### Output Top 5 Softmax Probabilities For Each Image Found on the Web
For each of the new images, print out the model's softmax probabilities to show the **certainty** of the model's predictions (limit the output to the top 5 probabilities for each image). [`tf.nn.top_k`](https://www.tensorflow.org/versions/r0.12/api_docs/python/nn.html#top_k) could prove helpful here.
The example below demonstrates how tf.nn.top_k can be used to find the top k predictions for each image.
`tf.nn.top_k` will return the values and indices (class ids) of the top k predictions. So if k=3, for each sign, it'll return the 3 largest probabilities (out of a possible 43) and the correspoding class ids.
Take this numpy array as an example. The values in the array represent predictions. The array contains softmax probabilities for five candidate images with six possible classes. `tf.nn.top_k` is used to choose the three classes with the highest probability:
```
# (5, 6) array
a = np.array([[ 0.24879643, 0.07032244, 0.12641572, 0.34763842, 0.07893497,
0.12789202],
[ 0.28086119, 0.27569815, 0.08594638, 0.0178669 , 0.18063401,
0.15899337],
[ 0.26076848, 0.23664738, 0.08020603, 0.07001922, 0.1134371 ,
0.23892179],
[ 0.11943333, 0.29198961, 0.02605103, 0.26234032, 0.1351348 ,
0.16505091],
[ 0.09561176, 0.34396535, 0.0643941 , 0.16240774, 0.24206137,
0.09155967]])
```
Running it through `sess.run(tf.nn.top_k(tf.constant(a), k=3))` produces:
```
TopKV2(values=array([[ 0.34763842, 0.24879643, 0.12789202],
[ 0.28086119, 0.27569815, 0.18063401],
[ 0.26076848, 0.23892179, 0.23664738],
[ 0.29198961, 0.26234032, 0.16505091],
[ 0.34396535, 0.24206137, 0.16240774]]), indices=array([[3, 0, 5],
[0, 1, 4],
[0, 5, 1],
[1, 3, 5],
[1, 4, 3]], dtype=int32))
```
Looking just at the first row we get `[ 0.34763842, 0.24879643, 0.12789202]`, you can confirm these are the 3 largest probabilities in `a`. You'll also notice `[3, 0, 5]` are the corresponding indices.
```
### Print out the top five softmax probabilities for the predictions on the German traffic sign images found on the web.
### Feel free to use as many code cells as needed.
```
### Project Writeup
Once you have completed the code implementation, document your results in a project writeup using this [template](https://github.com/udacity/CarND-Traffic-Sign-Classifier-Project/blob/master/writeup_template.md) as a guide. The writeup can be in a markdown or pdf file.
> **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to \n",
"**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
---
## Step 4 (Optional): Visualize the Neural Network's State with Test Images
This Section is not required to complete but acts as an additional excersise for understaning the output of a neural network's weights. While neural networks can be a great learning device they are often referred to as a black box. We can understand what the weights of a neural network look like better by plotting their feature maps. After successfully training your neural network you can see what it's feature maps look like by plotting the output of the network's weight layers in response to a test stimuli image. From these plotted feature maps, it's possible to see what characteristics of an image the network finds interesting. For a sign, maybe the inner network feature maps react with high activation to the sign's boundary outline or to the contrast in the sign's painted symbol.
Provided for you below is the function code that allows you to get the visualization output of any tensorflow weight layer you want. The inputs to the function should be a stimuli image, one used during training or a new one you provided, and then the tensorflow variable name that represents the layer's state during the training process, for instance if you wanted to see what the [LeNet lab's](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) feature maps looked like for it's second convolutional layer you could enter conv2 as the tf_activation variable.
For an example of what feature map outputs look like, check out NVIDIA's results in their paper [End-to-End Deep Learning for Self-Driving Cars](https://devblogs.nvidia.com/parallelforall/deep-learning-self-driving-cars/) in the section Visualization of internal CNN State. NVIDIA was able to show that their network's inner weights had high activations to road boundary lines by comparing feature maps from an image with a clear path to one without. Try experimenting with a similar test to show that your trained network's weights are looking for interesting features, whether it's looking at differences in feature maps from images with or without a sign, or even what feature maps look like in a trained network vs a completely untrained one on the same sign image.
<figure>
<img src="visualize_cnn.png" width="380" alt="Combined Image" />
<figcaption>
<p></p>
<p style="text-align: center;"> Your output should look something like this (above)</p>
</figcaption>
</figure>
<p></p>
```
### Visualize your network's feature maps here.
### Feel free to use as many code cells as needed.
# image_input: the test image being fed into the network to produce the feature maps
# tf_activation: should be a tf variable name used during your training procedure that represents the calculated state of a specific weight layer
# activation_min/max: can be used to view the activation contrast in more detail, by default matplot sets min and max to the actual min and max values of the output
# plt_num: used to plot out multiple different weight feature map sets on the same block, just extend the plt number for each new feature map entry
def outputFeatureMap(image_input, tf_activation, activation_min=-1, activation_max=-1 ,plt_num=1):
# Here make sure to preprocess your image_input in a way your network expects
# with size, normalization, ect if needed
# image_input =
# Note: x should be the same name as your network's tensorflow data placeholder variable
# If you get an error tf_activation is not defined it may be having trouble accessing the variable from inside a function
activation = tf_activation.eval(session=sess,feed_dict={x : image_input})
featuremaps = activation.shape[3]
plt.figure(plt_num, figsize=(15,15))
for featuremap in range(featuremaps):
plt.subplot(6,8, featuremap+1) # sets the number of feature maps to show on each row and column
plt.title('FeatureMap ' + str(featuremap)) # displays the feature map number
if activation_min != -1 & activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin =activation_min, vmax=activation_max, cmap="gray")
elif activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmax=activation_max, cmap="gray")
elif activation_min !=-1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin=activation_min, cmap="gray")
else:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", cmap="gray")
```
| github_jupyter |
# Сериализация
## Обработка конфигурационных файлов
### json
JSON (JavaScript Object Notation) - простой формат обмена данными, основанный на подмножестве синтаксиса JavaScript. Модуль json позволяет кодировать и декодировать данные в удобном формате.
Некоторые возможности библиотеки **json**
**json.dump**`(obj, fp, skipkeys=False, ensure_ascii=True, check_circular=True, allow_nan=True, cls=None, indent=None, separators=None, default=None, sort_keys=False, **kw)` - сериализует obj как форматированный JSON поток в fp.
**json.dumps**`(obj, skipkeys=False, ensure_ascii=True, check_circular=True, allow_nan=True, cls=None, indent=None, separators=None, default=None, sort_keys=False, **kw)` - сериализует obj в строку JSON-формата.
**json.load**`(fp, cls=None, object_hook=None, parse_float=None, parse_int=None, parse_constant=None, object_pairs_hook=None, **kw)` - десериализует JSON из fp.
**json.loads**`(s, encoding=None, cls=None, object_hook=None, parse_float=None, parse_int=None, parse_constant=None, object_pairs_hook=None, **kw)` - десериализует s (экземпляр str, содержащий документ JSON) в объект Python.
```
import json
# Кодирование основных объектов Python
print(json.dumps(['foo', {'bar': ('baz', None, 1.0, 2)}]))
print(json.dumps({"c": 0, "b": 0, "a": 0}, sort_keys=True))
# Компактное кодирование
print(json.dumps([1,2,3,{'4': 5, '6': 7}], separators=(',', ':')))
# Красивый вывод
print(json.dumps({'4': 5, '6': 7}, sort_keys=True, indent=4))
# Декодирование (парсинг) JSON
print(json.loads('["foo", {"bar":["baz", null, 1.0, 2]}]'))
print(json.loads('"\\"foo\\bar"'))
```
### yaml
YAML (YAML Ain’t Markup Language) - еще один текстовый формат для записи данных.
YAML более приятен для восприятия человеком, чем JSON, поэтому его часто используют для описания сценариев в ПО. Например, в Ansible.
Для работы с YAML в Python используется модуль **pyyaml**. Он не входит в стандартную библиотеку модулей, поэтому его нужно установить:
`pip install pyyaml`
```
# Чтение из YAML (файл info.yaml)
import yaml
from pprint import pprint # Модуль pprint позволяет красиво отображать объекты Python
with open('info.yaml') as f:
templates = yaml.safe_load(f)
pprint(templates) # Использование функции модуля pprint для вывода
# Запись в YAML
trunk_template = [
'switchport trunk encapsulation dot1q', 'switchport mode trunk',
'switchport trunk native vlan 999', 'switchport trunk allowed vlan'
]
access_template = [
'switchport mode access', 'switchport access vlan',
'switchport nonegotiate', 'spanning-tree portfast',
'spanning-tree bpduguard enable'
]
to_yaml = {'trunk': trunk_template, 'access': access_template}
with open('sw_templates.yaml', 'w') as f:
yaml.dump(to_yaml, f)
with open('sw_templates.yaml') as f:
print(f.read())
```
### ini
Как правило, ini-файлы используют для хранения настроек приложения или операционной системы. Библиотека в ядре Python включает в себя модуль, под названием **configparser**, который вы можете использовать для создания и работы с файлами конфигурации.
```
import configparser
# Создание конфигурационного файла
config = configparser.ConfigParser()
config.add_section("Settings")
config.set("Settings", "font", "Courier")
config.set("Settings", "font_size", "10")
config.set("Settings", "font_style", "Normal")
config.set("Settings", "font_info",
"You are using %(font)s at %(font_size)s pt")
with open('my_settings.ini', 'w') as config_file:
config.write(config_file)
# ===Выведем содержимое файла===
with open('my_settings.ini', 'r') as config_file:
print(config_file.read())
# Чтение конфигурационного файла
config = configparser.ConfigParser()
config.read('my_settings.ini')
# Читаем некоторые значения из конфиг. файла.
font = config.get("Settings", "font")
font_size = config.get("Settings", "font_size")
# Меняем значения из конфиг. файла.
config.set("Settings", "font_size", "12")
# Удаляем значение из конфиг. файла.
config.remove_option("Settings", "font_style")
# Вносим изменения в конфиг. файл.
with open('my_settings.ini', "w") as config_file:
config.write(config_file)
# ===Выведем содержимое файла===
with open('my_settings.ini', 'r') as config_file:
print(config_file.read())
```
## Консервация объектов
Модуль `pickle` (англ. pickle - консервировать) реализует мощный алгоритм сериализации и десериализации объектов Python. "Pickling" - процесс преобразования объекта Python в поток байтов, а "unpickling" - обратная операция, в результате которой поток байтов преобразуется обратно в Python-объект. Так как поток байтов легко можно записать в файл, модуль `pickle` широко применяется для сохранения и загрузки сложных объектов в Python.
Модуль pickle предоставляет следующие функции для удобства сохранения/загрузки объектов:
- `pickle.dump(obj, file, protocol=None, *, fix_imports=True)`\
Записывает сериализованный объект в файл. Дополнительный аргумент protocol указывает используемый протокол. По умолчанию равен 3 и именно он рекомендован для использования в Python 3 (несмотря на то, что в Python 3.4 добавили протокол версии 4 с некоторыми оптимизациями). В любом случае, записывать и загружать надо с одним и тем же протоколом.
- `pickle.dumps(obj, protocol=None, *, fix_imports=True)`\
Возвращает сериализованный объект. Впоследствии вы его можете использовать как угодно.
- `pickle.load(file, *, fix_imports=True, encoding="ASCII", errors="strict")`\
Загружает объект из файла.
- `pickle.loads(bytes_object, *, fix_imports=True, encoding="ASCII", errors="strict")`\
Загружает объект из потока байт.
Модуль `pickle` также определяет несколько исключений:
`pickle.PickleError`
- `pickle.PicklingError` - случились проблемы с сериализацией объекта.
- `pickle.UnpicklingError` - случились проблемы с десериализацией объекта.
Этих функций вполне достаточно для сохранения и загрузки встроенных типов данных.
```
import pickle
data = {
'a': [1, 2.0, 3, 4+6j],
'b': ("character string", b"byte string"),
'c': {None, True, False}
}
with open('data.pickle', 'wb') as f:
pickle.dump(data, f)
with open('data.pickle', 'rb') as f:
data_new = pickle.load(f)
print(data_new)
```
| github_jupyter |
# JS vs PY automl: who will win?
Here an instance of a rather simplistic automl implementation in Python is pitted against implementation in JS. Does the JS version of AutoML reach the quality standards of even a simple Python version? Find out in this notebook.
```
from subprocess import call, DEVNULL
import numpy as np
import pmlb
import json
from time import time
import os
import random
```
## Reference implementation in Python
A simple grid search is employed to do automl in Python. The code below should speak for itself.
```
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.tree import DecisionTreeClassifier, DecisionTreeRegressor
from sklearn.linear_model import SGDClassifier, SGDRegressor
from sklearn.ensemble import GradientBoostingClassifier, GradientBoostingRegressor
from sklearn.dummy import DummyClassifier, DummyRegressor
from sklearn.preprocessing import StandardScaler
from sklearn.impute import SimpleImputer
from sklearn.pipeline import make_pipeline, Pipeline
from time import time
class PythonAutoML():
def __init__(self):
pass
def fit_predict(self, X_train, y_train, X_test):
# determine type of learning problem
classification = False
for v in y_train:
try:
v = float(v)
except BaseException as ex:
classification = True
break
# its a tree
tree = {
'model': [DecisionTreeClassifier() if classification else DecisionTreeRegressor()],
'model__max_leaf_nodes': [3, 5, 7, 11, 19, 31, 53, 89, 137, 179, 227, 271]
}
# its a dot product
lsgd = {
'model': [SGDClassifier() if classification else SGDRegressor()],
'model__alpha': np.logspace(-3, 6, 11),
'model__l1_ratio': [0.0, 0.5, 1.0]
}
# no its a gradient boosting
gbdt = {
'model': [GradientBoostingClassifier() if classification else GradientBoostingRegressor()],
'model__learning_rate': [0.001, 0.005, 0.01, 0.05, 0.1, 0.5]
}
model = GridSearchCV(
estimator=Pipeline([
('imputer', SimpleImputer()),
('scaler', StandardScaler()),
('model', DummyRegressor())
]),
param_grid=[
tree, lsgd, gbdt
],
cv=5,
n_jobs=-1,
verbose=0
)
start = time()
model.fit(X_train, y_train)
duration = time() - start
y_pred = model.predict(X_test)
return {
'y_pred': y_pred,
'fit_time': duration
}
```
## Implementation in JavaScript
It is given below.
```
from random import choice
import json
from subprocess import call, DEVNULL
class JSAutoML():
def __init__(self):
pass
def fit_predict(self, X_train, y_train, X_test):
puid = ''.join([choice("abcdefghijklmnopqrstuvwxyz") for _ in range(10)])
node_code = """
const puid = "%s"
const ai = require('../src/automljs')
const fs = require('fs')
var data = JSON.parse(fs.readFileSync(puid + '.data.json', 'utf8'));
async function main(){
var X = data['X'];
var y = data['y'];
var X_test = data['X_test'];
// read estimator from the serialization module
var model = new ai.automl.AutoMLModel({'max_iter':10})
var fit_start = process.hrtime();
await model.fit(X, y)
var elapsed = process.hrtime(fit_start)[1] / 1000000; // divide by a million to get nano to milli
var y_pred = await model.predict(X_test)
var res = {
'y_pred': y_pred
}
await fs.writeFile(puid + '.result.json', JSON.stringify(res), 'utf8', function(){ })
}
main()
""" % puid
jsfile = puid + '.js'
dtfile = puid + '.data.json'
rsfile = puid + '.result.json'
with open(jsfile, 'w') as s:
s.write(node_code)
with open(dtfile, 'w') as d:
json.dump({
'X': X_train.tolist(),
'y': y_train.tolist(),
'X_test': X_test.tolist()
}, d)
start = time()
call(['node ' + jsfile], shell=True)
duration = time() - start
result = None
with open(rsfile, 'r') as js:
result = json.load(js)
return {
'y_pred': result['y_pred'],
'fit_time': duration
}
```
## Benchmarks!
```
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier, DecisionTreeRegressor
from sklearn.linear_model import SGDClassifier, SGDRegressor
from sklearn.preprocessing import Imputer, StandardScaler
from sklearn.pipeline import make_pipeline
from sklearn.metrics import accuracy_score, r2_score
from tqdm import tqdm
from itertools import product
from pprint import pprint
from pmlb import classification_dataset_names, regression_dataset_names
############################################ SETTINGS ############################################
models = [PythonAutoML, JSAutoML]
max_size = 1000
data_classification = True
############################################ SETTINGS ############################################
results = []
datasets = classification_dataset_names if data_classification else regression_dataset_names
metric = accuracy_score if data_classification else r2_score
for name in datasets:
if name in {'1191_BNG_pbc', '1595_poker'}:
continue
print('Fetching data ...')
X, y = pmlb.fetch_data(name, True, local_cache_dir='/home/iaroslav/.cache/pmlb')
print(name, X.shape)
# skip too many features for now
if X.shape[-1] >= 500:
continue
# make datasets small for testing
if len(y) > max_size:
X = X[:max_size]
y = y[:max_size]
# skip datasets with constant outputs
unique_outputs = len(set(y))
if(unique_outputs < 2):
continue
print('Number of unique outputs: %s' % unique_outputs)
if data_classification:
# disambiguation with e.g. integer class labels
y = np.array(['class_' + str(v) for v in y])
# make training and testing partitions
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
result_dataset = {
'dataset_name': name,
'dataset_shape': str(X.shape),
'models': {}
}
for model in models:
inst = model()
result = inst.fit_predict(X_train, y_train, X_test)
y_pred = result['y_pred']
del result['y_pred']
score = metric(y_test, y_pred)
result['test_score'] = score
result_dataset['models'][model.__name__] = result
pprint(result_dataset)
results.append(result_dataset)
```
## Final evaluation
The results are summarized here.
```
import pandas as pd
I = [] # this will be the index of the df
rows = []
for result in results:
I.append(result['dataset_name'])
row = {}
for model_name, stats in result['models'].items():
for stat in stats:
row[model_name + "_" + stat] = stats[stat]
rows.append(row)
df = pd.DataFrame(data=rows, index=I)
display(df.describe().round(3))
```
| github_jupyter |
```
pip install tensorflow
# Import dependencies
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler,OneHotEncoder
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
import pandas as pd
import tensorflow as tf
# Import our dataset
charity_df = pd.read_csv('charity_data.csv')
charity_df.head()
# Generate our categorical variable list
charity_categorical = charity_df.dtypes[charity_df.dtypes == "object"].index.tolist()
# Check the number of unique values in each column
charity_df[charity_categorical].nunique()
charity_df.columns
# What variable(s) are considered the target for your model?
# IS_SUCCESSFUL
# What variable(s) are considered to be the features for your model?
# 'APPLICATION_TYPE', 'AFFILIATION', 'CLASSIFICATION','USE_CASE', 'ORGANIZATION', 'STATUS', 'INCOME_AMT', 'SPECIAL_CONSIDERATIONS', 'ASK_AMT'
# What variable(s) are neither and should be removed from the input data?
# 'EIN', 'NAME'
# Drop the EIN and NAME columns
charity_df.drop(columns=["EIN", "NAME"], inplace=True)
charity_df
# Evaluate APPLICATION_TYPE
application_type = charity_df.APPLICATION_TYPE.value_counts()
application_type
application_type.plot.density()
# Bucket values less than 600
application_type_bucket = list(application_type[application_type < 600].index)
for x in application_type_bucket:
charity_df.APPLICATION_TYPE = charity_df.APPLICATION_TYPE.replace(x, "Other")
charity_df.APPLICATION_TYPE.value_counts()
# Evaluate CLASSIFICATION
classification_type = charity_df.CLASSIFICATION.value_counts()
classification_type.head(30)
classification_type.plot.density()
# Bucket values less than 600
classification_type_bucket = list(classification_type[classification_type < 600].index)
for x in classification_type_bucket:
charity_df.CLASSIFICATION = charity_df.CLASSIFICATION.replace(x, "Other")
charity_df.CLASSIFICATION.value_counts()
# Generate our categorical variable list
charity_categorical = charity_df.dtypes[charity_df.dtypes == "object"].index.tolist()
# Check the number of unique values in each column
charity_df[charity_categorical].nunique()
# Create a OneHotEncoder instance
enc = OneHotEncoder(sparse=False)
# Fit and transform the OneHotEncoder using the categorical variable list
encode_df = pd.DataFrame(enc.fit_transform(charity_df[charity_categorical]))
# Add the encoded variable names to the dataframe
encode_df.columns = enc.get_feature_names(charity_categorical)
encode_df.head()
# Merge one-hot encoded features and drop the originals
charity_df = charity_df.merge(encode_df,left_index=True, right_index=True)
charity_df = charity_df.drop(charity_categorical,1)
charity_df.head()
# Remove loan status target from features data
y = charity_df["SPECIAL_CONSIDERATIONS_Y"].values
X = charity_df.drop(["SPECIAL_CONSIDERATIONS_Y","SPECIAL_CONSIDERATIONS_N"],1).values
# Split training/test datasets
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42, stratify=y)
# Create a StandardScaler instance
scaler = StandardScaler()
# Fit the StandardScaler
X_scaler = scaler.fit(X_train)
# Scale the data
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
# Define the model - deep neural net
number_input_features = len(X_train_scaled[0])
hidden_nodes_layer1 = 10
hidden_nodes_layer2 = 5
nn = tf.keras.models.Sequential()
# First hidden layer
nn.add(
tf.keras.layers.Dense(units=hidden_nodes_layer1, input_dim=number_input_features, activation="relu")
)
# Second hidden layer
nn.add(tf.keras.layers.Dense(units=hidden_nodes_layer2, activation="relu"))
# Output layer
nn.add(tf.keras.layers.Dense(units=1, activation="sigmoid"))
# Check the structure of the model
nn.summary()
# Compile the Sequential model together and customize metrics
nn.compile(loss="binary_crossentropy", optimizer="adam", metrics=["accuracy"])
# Train the model
fit_model = nn.fit(X_train,y_train,epochs=5)
# Evaluate the model using the test data
model_loss, model_accuracy = nn.evaluate(X_test_scaled,y_test,verbose=2)
print(f"Loss: {model_loss}, Accuracy: {model_accuracy}")
```
| github_jupyter |
# Custom generators
```
import tohu
from tohu.v4.primitive_generators import *
from tohu.v4.derived_generators import *
from tohu.v4.dispatch_generators import *
from tohu.v4.custom_generator import *
from tohu.v4.utils import print_generated_sequence, make_dummy_tuples
print(f'Tohu version: {tohu.__version__}')
```
## Custom generator without `__init__` method
```
class QuuxGenerator(CustomGenerator):
aa = Integer(100, 200)
bb = HashDigest(length=6)
cc = FakerGenerator(method='name')
g = QuuxGenerator()
print_generated_sequence(g, num=10, sep='\n', seed=12345)
```
### Explicitly setting the name of generated items
Let's repeat the previous example, but explicitly set the name of generated items by setting the `__tohu_items_name__` attribute inside the custom generator.
```
class SomeGeneratorWithExplicitItemsName(CustomGenerator):
__tohu_items_name__ = 'Foobar'
aa = Integer(100, 200)
bb = HashDigest(length=6)
cc = FakerGenerator(method='name')
g = SomeGeneratorWithExplicitItemsName()
```
The generated sequence is the same as above, but the name of the items has changed from `Quux` to `Foobar`.
```
print_generated_sequence(g, num=10, sep='\n', seed=12345)
```
## Custom generator with `__init__` method
```
class QuuxGenerator(CustomGenerator):
aa = Integer(100, 200)
def __init__(self, faker_method):
self.bb = FakerGenerator(method=faker_method)
# Note: the call to super().__init__() needs to be at the end,
# and it needs to be passed the same arguments as the __init__()
# method from which it is called (here: `faker_method`).
super().__init__(faker_method)
g1 = QuuxGenerator(faker_method='first_name')
g2 = QuuxGenerator(faker_method='city')
print_generated_sequence(g1, num=10, sep='\n', seed=12345); print()
print_generated_sequence(g2, num=10, sep='\n', seed=12345)
```
## Custom generator containing derived generators
```
some_tuples = make_dummy_tuples('abcdefghijklmnopqrstuvwxyz')
#some_tuples[:5]
```
### Example: extracting attributes
```
class QuuxGenerator(CustomGenerator):
aa = SelectOne(some_tuples)
bb = GetAttribute(aa, 'x')
cc = GetAttribute(aa, 'y')
g = QuuxGenerator()
print_generated_sequence(g, num=10, sep='\n', seed=12345)
```
### Example: arithmetic
```
def square(x):
return x * x
def add(x, y):
return x + y
class QuuxGenerator(CustomGenerator):
aa = Integer(0, 20)
bb = Integer(0, 20)
cc = Apply(add, aa, Apply(square, bb))
g = QuuxGenerator()
print_generated_sequence(g, num=10, sep='\n', seed=12345)
df = g.generate(num=100, seed=12345).to_df()
print(list(df['aa'][:20]))
print(list(df['bb'][:20]))
print(list(df['cc'][:20]))
all(df['aa'] + df['bb']**2 == df['cc'])
```
### Example: multi-stage dependencies
```
class QuuxGenerator(CustomGenerator):
name = FakerGenerator(method="name")
tag = SelectOne(['a', 'bb', 'ccc'])
g = QuuxGenerator()
quux_items = g.generate(num=100, seed=12345)
quux_items.to_df().head(5)
tag_lookup = {
'a': [1, 2, 3, 4, 5],
'bb': [10, 20, 30, 40, 50],
'ccc': [100, 200, 300, 400, 500],
}
class FoobarGenerator(CustomGenerator):
some_quux = SelectOne(quux_items)
number = SelectOneDerived(Lookup(GetAttribute(some_quux, 'tag'), tag_lookup))
h = FoobarGenerator()
h_items = h.generate(10000, seed=12345)
df = h_items.to_df(fields={'name': 'some_quux.name', 'tag': 'some_quux.tag', 'number': 'number'})
df.head()
print(df.query('tag == "a"')['number'].isin([1, 2, 3, 4, 5]).all())
print(df.query('tag == "bb"')['number'].isin([10, 20, 30, 40, 50]).all())
print(df.query('tag == "ccc"')['number'].isin([100, 200, 300, 400, 500]).all())
df.query('tag == "a"').head(5)
df.query('tag == "bb"').head(5)
df.query('tag == "ccc"').head(5)
```
| github_jupyter |
last edited by Claire Valva on May 13, 2019, with update and cleanup on June 24, 2019
# Test ENSO simulations and plotting
```
# import packages
import numpy as np
from scipy.fftpack import fft, ifft, fftfreq, fftshift, ifftshift
import scipy.integrate as sciint
import pandas as pd
from math import pi
from sympy import solve, Poly, Eq, Function, exp, re, im
from scipy.optimize import fsolve
from decimal import Decimal
import pickle
import time
import random
import multiprocessing as mp
from joblib import Parallel, delayed
import numpy as np
from scipy.signal import get_window, csd
from scipy.signal.windows import hann, hanning, nuttall, flattop
from scipy.fftpack import fft, ifft, fftfreq, fftshift, ifftshift
#import matplotlib
#matplotlib.use('Agg')
import matplotlib.pyplot as plt
import scipy.integrate as sciint
import pandas as pd
import datetime
import matplotlib.cm as cm
from math import pi
import matplotlib.ticker as tck
import datetime
from sympy import solve, Poly, Eq, Function, exp, re, im
from netCDF4 import Dataset, num2date # This is to read .nc files and time array
from scipy.optimize import fsolve
from decimal import Decimal
import pickle
import multiprocessing as mp
from joblib import Parallel, delayed
import matplotlib.colors as colors
from seaborn import cubehelix_palette #for contour plot colors
import seaborn as sns
from decimal import Decimal
import numpy.ma as ma
import random
#flatten season for plotting
flatten = lambda l: [item for sublist in l for item in sublist]
import scipy.stats as spyst
from os import walk
oldf = []
for (dirpath, dirnames, filenames) in walk('/scratch/midway2/clairev/enso_spectra/'):
oldf.extend(filenames)
break
f = []
for named in oldf:
if named[0:15] == "spectra_enso_02":
f.append(named)
def solve_f(X, Zofkt):
# function to solve f coeff equation for trend analysis
x,y = X
f = Zofkt - x*np.exp(1j*y)
return [np.real(f), np.imag(f)]
def real_f(X,Zofkt):
# function to wrap solve_f so that it can be used with fsolve
x,y = X
z = [x+0j,y+0j]
actual_f = solve_f(z, Zofkt)
return(actual_f)
def fwithZ(entry):
answers = fsolve(real_f, np.array([0,0]), args = entry)
return answers
# get function to generate random coeffs
def entry_fft(amp, phase = random.uniform(0, 2*pi)):
# takes amplitude and phase to give corresponding fourier coeff
entry = amp*np.exp(1j*phase)
return entry
# write functions to make a longer ifft
def ext_row(row, n):
ext_f = np.zeros(((len(row) - 1) * n + 1,), dtype="complex128")
ext_f[::n] = row * n
return ext_f
def ext_ifft_new(n, input_array):
# add the zeros onto each end
ext_f = [ext_row(entry,n) for entry in input_array]
# make up for the formulat multiplying for array length
olddim = len(input_array[5])
newdim = len(ext_f[0])
mult = newdim/olddim
ext_f = np.multiply(ext_f, mult)
adjusted_tested = np.fft.ifft2(ext_f)
return adjusted_tested
season_titles = ["Winter", "Spring", "Summer", "Fall"]
seasons = ["winter", "spring", "summer", "fall"]
# flatten season for plotting
flatten = lambda l: [item for sublist in l for item in sublist]
named = f[0]
#file_name = "/scratch/midway2/clairev/enso_spectra/averaged/01_enso_avg_" + str(named[16:21])
#file_pickle = open(file_name, "rb")
#d2_touse, d2_seasons, d2_averages = pickle.load(file_pickle)
ens = ["nino", "nina", "neutral"]
d2_names = [enso + " " + part for part in seasons for enso in ens]
d2_names
name = "01_enso_36.0N424"
name[8:13]
file_name = "/scratch/midway2/clairev/enso_sims/01_enso_36.0N424"
file_pickle = open(file_name, "rb")
pickled = pickle.load(file_pickle)
flat_sims = [flatten(entry[0]) for entry in pickled]
#make lists of el nino/regular/la nina years
nino = [1980,1983,1987,1988,1992,
1995,1998,2003,2007,2010]
neutral = [1979,1981,1982,1984,1985,1986,1990,
1991,1993,1994,1996,1997,2001,2002,
2004,2005,2006,2009,2013,2014,2015,2016]
nina = [1989,1999,2000,2008,2011,2012]
len_all = 38.0
nina_per = len(nina)/len_all
nino_per = len(nino)/len_all
neutral_per = len(neutral)/len_all
all_pers = [nina_per, nino_per, neutral_per]
all_pers
# now plot them
# weight them by years percentrage when plotting together
for j in range(4):
plt.clf();
plt.figure(figsize=(15, 5));
for k in range(3):
#print("hi")
plt.hist(x = np.real(flat_sims[j*3 + k]), bins = 100, density = True, alpha = 0.5, label = d2_names[j*3 + k])
plt.ylabel("density")
plt.legend()
plt.xlabel("geopotential height")
plt.show()
# sort them into each season
phase_all = [[[[fwithZ(entry) for entry in sublist]
for sublist in year]
for year in season]
for season in d2_seasons]
# sort them into each season
amps_all = [[[[entry[0] for entry in sublist]
for sublist in year]
for year in season]
for season in phase_all]
ps_all = [[[[entry[1] % (2 * np.pi) for entry in sublist]
for sublist in year]
for year in season]
for season in phase_all]
# adjust for winter averaging
# TO DO: come up with better procedure rather
# current: chopping off edges to make the same length for averaging
norml = 359
longl = 364
def padded(to_pad, index):
length = len(to_pad)
if index == 0:
zeros = longl - length
to_pad = list(to_pad)
for i in range(zeros):
to_pad.append(0)
return to_pad
else:
return to_pad
#pad rows with zeros to account for leap year
season_amps_adj = [[[padded(row, 0)
for row in entry]
for entry in amps_all[i]]
for i in range(len(amps_all))]
#pad rows with zeros to account for leap year
season_phase_adj = [[[padded(row, 0)
for row in entry]
for entry in ps_all[i]]
for i in range(len(ps_all))]
#get average amplitude for each season
avg_amps = [np.average(season, axis = 0)
for season in season_amps_adj]
#get std amplitude for each season
std_amps = [np.std(season, axis = 0)
for season in season_amps_adj]
#get average phases for each season
avg_phase = [spyst.circmean(season, axis = 0)
for season in season_phase_adj]
#get std phases for each season
std_phase = [spyst.circstd(season, axis = 0)
for season in season_phase_adj]
import pickle
file_name2 = "/scratch/midway2/clairev/enso_spectra/averaged/01_enso_avg_" + str(named[16:21])
file_pickle = open(file_name2,'wb')
pickle.dump([avg_amps,std_amps,avg_phase,std_phase],file_pickle)
file_pickle.close()
# get function to generate random coeffs
def entry_fft(amp,std, phase, stdphase):
# takes amplitude and phase to give corresponding fourier coeff
amp_new = np.random.normal(loc = amp, scale = std)
phase_new = np.random.normal(loc = phase, scale = stdphase)
entry = amp_new*np.exp(1j*phase_new)
return entry
# write functions to make a longer ifft
def ext_row(row, n):
ext_f = np.zeros(((len(row) - 1) * n + 1,), dtype="complex128")
ext_f[::n] = row * n
return ext_f
def ext_ifft_new(n, input_array):
# add the zeros onto each end
ext_f = [ext_row(entry,n) for entry in input_array]
# make up for the formulat multiplying for array length
olddim = len(input_array[5])
newdim = len(ext_f[0])
mult = newdim/olddim
# ext_f = np.multiply(mult, ext_f)
adjusted_tested = np.fft.ifft2(ext_f)
return adjusted_tested
def combined(amps,stds,phases,stdphases, length):
# combines generation of random phase with inverse transform
newarray = [[entry_fft(amp = amps[wave][timed],
std = stds[wave][timed],
phase = phases[wave][timed], stdphase = stdphases[wave][timed])
for timed in range(len(amps[wave]))]
for wave in range(len(amps))]
newarray = [np.array(leaf) for leaf in newarray]
iffted = ext_ifft_new(length, newarray)
return iffted
def repeater(season, stds, phases,stdphases, length, times):
# repeats the phase creation and inverse transform
newarray = [combined(season, stds, phases,stdphases,length) for leaf in range(times)]
return(newarray)
# set lims
runlen = 75
runtimes = 1
repeattimes = 20
listed_parts = []
def repeater_2(amps,stds, phases,stdphases, length, times):
#do procedure
repeated_comp = [repeater(amps[i],stds[i], phases[i], stdphases[i], length, times)
for i in range(len(amps))]
#output.put(repeated_comp)
#listed_parts.append(repeated_comp)
import pickle
file_name2 = "/scratch/midway2/clairev/enso_sims/01_enso_" + str(named[16:21]) + str(random.randint(1,1000))
file_pickle = open(file_name2,'wb')
pickle.dump(repeated_comp,file_pickle)
file_pickle.close()
return repeated_comp
toplot = repeater_2(avg_amps,std_amps, runlen, runtimes)
```
| github_jupyter |
# 2つのガウス分布を含む混合ガウス分布のためのEMアルゴリズム
(細かいコメントはもうちょっと待ってくださーい)
千葉工業大学 上田 隆一
(c) 2017 Ryuichi Ueda
This software is released under the MIT License, see LICENSE.
## はじめに
このコードは、2つの2次元ガウス分布を含む混合ガウス分布から生成されたデータについて、EMアルゴリズムでパラメータを求めるためのEMアルゴリズムの実装例です。処理の流れは、次のようなものです。
* (準備)2つのガウス分布からサンプリング
* 推定対象は、この2つのガウス分布のパラメータと、どちらからどれだけサンプリングされたかの比
* 適当なパラメータで2つガウス分布を準備し、収束するまで以下の繰り返し
* Eステップ: 各サンプルがどちらの分布から生成されたらしいかを、2つのガウス分布の確率密度関数から得られる値の比で計算
* Mステップ: Eステップで得た比を利用して、推定対象の値を計算
* 収束したら、推定値を出力
## アルゴリズムを適用される対象になるデータの生成
クラスタリングの対象となるデータを作ります。二つの2次元ガウス分布から、2:1の割合で標本抽出します。(GitHubだと行列が崩れて表示されますが、$\mathcal{N}$の二番目の引数は2x2行列です。)
* 2つの分布
* 分布A(200個抽出): $\mathcal{N}\left( \begin{bmatrix}170 \\ 70 \end{bmatrix}, \begin{bmatrix}6^2 & -30 \\ -30 & 8^2\end{bmatrix} \right)$
* 分布B(100個抽出): $\mathcal{N}\left( \begin{bmatrix}165 \\ 50 \end{bmatrix}, \begin{bmatrix}5^2 & 20 \\ 20 & 6^2\end{bmatrix} \right)$
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import math
def make_samples():
# グループAのガウス分布
muA_ans = [170,70] # 横軸、縦軸をx,y軸とすると、x=170, y=70が中心
covA_ans = [[6**2,-30],[-30,8**2]] # x軸の標準偏差6、y軸の標準偏差-30、共分散-30
samplesA = np.random.multivariate_normal(muA_ans,covA_ans,200).T #200個の点をサンプリング
# グループBのガウス分布
muB_ans = [165,50] # x=165, y=50が中心
covB_ans = [[5.**2,20],[20,6**2]] # x軸の標準偏差5、y軸の標準偏差6、共分散20
samplesB = np.random.multivariate_normal(muB_ans,covB_ans,100).T #100個の点をサンプリング
# 2つのグループのリストをくっつけて返す
return np.column_stack((samplesA,samplesB))
# データを作る
samples = make_samples()
#描画してみましょう
plt.scatter(samples[0],samples[1],color='g',marker='+') # sample[0]がx値のずらっと入ったリスト、sample[1]がy値
# このデータに関する既知のパラメータ
K = 2 # クラスタの数
N = len(samples[0]) # サンプルの数
```
以後、サンプルは$\boldsymbol{x}_n = (x_n,y_n) \quad (n=0,1,2,\dots,N)$と表現します。
## パラメータの初期設定
2つの分布のパラメータを格納する変数を準備して、このパラメータを上記の分布の式に近づけていきます。また、混合係数の変数も準備します。混合係数というのは、どっちからどれだけサンプルが生成されたかの割合のことです。上の例だと分布1で$2/3$、分布2で$1/3$となります。
* パラメータ
* 各分布(リストdistributions): $\mathcal{N}(\boldsymbol{x} | \boldsymbol{\mu}_k, \Sigma_k)\quad (k=0,1)$
* 混合係数(リストmixing_coefs): $\pi_k \quad (k=0,1; \pi_0 + \pi_1 = 1)$
```
from scipy.stats import multivariate_normal # これを使うと多次元ガウス分布のオブジェクトが生成できます
# 2つのガウス分布のオブジェクトを作る
distributions = []
distributions.append(multivariate_normal(mean=[160,80],cov= [[100,0],[0,100]]) ) # 分布1を適当な分布の中心、共分散行列で初期化
distributions.append(multivariate_normal(mean=[170,100],cov= [[100,0],[0,100]]) ) # 分布2を同様に初期化。分布1と少し値を変える必要アリ
# 混合係数のリスト
mixing_coefs = [1.0/K for k in range(K)] # 回りくどい書き方をしていますが、両方0.5で初期化されます。
```
描画の関係でサンプルの分布に重なるようにガウス分布を初期化していますが、辺鄙な値でも大丈夫です。
## 描画用の関数
```
def draw(ds,X):
# 分布を等高線で描画
x, y = np.mgrid[(min(X[0])):(max(X[0])):1, (min(X[1])):(max(X[1])):1] # 描画範囲の指定
for d in ds:
pos = np.empty(x.shape + (2,))
pos[:, :, 0] = x; pos[:, :, 1] = y
plt.contour(x, y, d.pdf(pos),alpha=0.2)
# サンプルの描画
plt.scatter(X[0],X[1],color='g',marker='+')
draw(distributions,samples)
```
以後、描かれた2つの楕円がサンプルの上に重なるように計算していきます。
## Eステップの実装
分布を固定し、各サンプルがどっちの分布に属すかを表した確率である負担率$\gamma(z_{nk})$のリストを各サンプル、各分布に対して計算して返します。
$\gamma(z_{nk}) = \dfrac{\pi_k \mathcal{N}(\boldsymbol{x}_n | \boldsymbol{\mu}_k, \Sigma_k)
}{\sum_j^K\pi_j \mathcal{N}(\boldsymbol{x}_n | \boldsymbol{\mu}_j, \Sigma_j)}$
```
def expectation_step(ds,X,pis): # 負担率の計算
ans = [] # 負担率のリスト
for n in range(N): # サンプルの数だけ繰り返し
# サンプルの地点における各分布の値(密度)を計算
ws = [ pis[k] * ds[k].pdf([X[0][n],X[1][n]]) for k in range(K) ] # 各クラスタに対して負担率の分子を計算
ans.append([ws[k]/sum(ws) for k in range(K)]) # 各クラスタの負担率の合計が1になるように正規化しているだけ
return ans # K * N 個のリスト
```
## Mステップの実装
各分布のパラメータと混合係数を更新します。次のコードの2行目の$N_k = \sum_{n=0}^{N-1} \gamma(z_{nk}) $は、各分布に関する全サンプルの負担率を合計して、各分布に「いくつ」サンプルが所属するかを求めたものです。負担率はゼロイチではないので、$N_k$は小数になります。
* 分布の中心の更新: $\boldsymbol{\mu}_k \longleftarrow \dfrac{1}{N_k} \sum_{n=0}^{N-1} \gamma(z_{nk})\boldsymbol{x}_n$
* 分布の共分散行列の更新: $\Sigma_k \longleftarrow \dfrac{1}{N_k} \sum_{n=0}^{N-1} \gamma(z_{nk}) (\boldsymbol{x}_n - \boldsymbol{\mu}_k)(\boldsymbol{x}_n - \boldsymbol{\mu}_k)^T$(更新後の$\boldsymbol{\mu}_k$を使用します。)
* 混合係数の更新: $\pi_k \longleftarrow \dfrac{N_k}{N}$
```
def maximization_step(k,X,gammas): # 引数は分布の番号、全サンプル、全サンプルと分布の負担率
N_k = sum ( [ gammas[n][k] for n in range(N) ])
# 分布の中心の更新
tmp_x = sum ( [ gammas[n][k] * X[0][n] for n in range(N) ]) / N_k # 全サンプルのx軸の値の平均値を、その分布に対する負担率で重み付き平均で計算
tmp_y = sum ( [ gammas[n][k] * X[1][n] for n in range(N) ]) / N_k # 同様にy軸の重み付き平均を計算
mu = [tmp_x,tmp_y] # 更新値
# 共分散行列の更新
ds= [ np.array([[X[0][n],X[1][n]]]) - np.array([mu]) for n in range(N) ] # 分布の中心に対するサンプルの位置のリスト
sigma = sum( [ gammas[n][k]* ds[n].T.dot(ds[n]) for n in range(N)] ) / N_k # 上のリストをかけて2x2行列を作り、負担率で重み付き平均をとる
return multivariate_normal(mean=mu,cov=sigma), N_k/N
```
### とりあえず1回ずつEステップとMステップを実行
Eステップで負担率のリストを作り、Mステップでパラメータを更新します。
```
def log_likelihood(ds,X,pis): # 収束の判断のために対数尤度を返す関数
ans = 0.0
for n in range(N):
ws = [ pis[k] * ds[k].pdf([X[0][n],X[1][n]]) for k in range(K) ]
ans += math.log1p(sum(ws) )
return ans
def one_step():
# Eステップ
gammas = expectation_step(distributions,samples,mixing_coefs)
# Mステップ
for k in range(K):
distributions[k], mixing_coefs[k] = maximization_step(k,samples,gammas)
return log_likelihood(distributions,samples,mixing_coefs)
one_step()
draw(distributions,samples)
```
少し二つの分布の位置がサンプルのある場所に近づいているのが分かります。
## 対数尤度が収束するまで繰り返し
対数尤度は次の式で与えられます。
$\ln p(\boldsymbol{x}_{0:N-1} | \boldsymbol{\mu}_{0:1}, \Sigma_{0:1}, \pi_{0:1}) = \sum_{n=0}^{N-1} \ln \left\{ \sum_{k=0}^{K-1} \pi_k \mathcal{N}(\boldsymbol{x}_n | \boldsymbol{\mu}_k, \Sigma_k) \right\}$
```
prev_log_likelihood = 0.0
for i in range(99):
after_log_likelihood = one_step()
if prev_log_likelihood/after_log_likelihood > 0.999: # 0.1%以上対数尤度が改善しなければ抜ける
break
else:
prev_log_likelihood = after_log_likelihood
if i % 3 == 0:
plt.figure()
draw(distributions,samples)
plt.figure()
draw(distributions,samples)
print("---------------------------------------------")
print("repeat: ", i+1)
for k in range(K):
print("Gauss",k,": ")
print(" share: ", mixing_coefs[k])
print(" mean: ", distributions[k].mean)
print(" cov: ", distributions[k].cov)
```
| github_jupyter |
# Description
This notebook documents allows the following on a group seven LIFX Tilechain with 5 Tiles
laid out horizontaly as following
T1 [0] [1] [2] [3] [4]
T2 [0] [1] [2] [3] [4]
T3 [0] [1] [2] [3] [4]
T4 [0] [1] [2] [3] [4]
T5 [0] [1] [2] [3] [4]
T6 [0] [1] [2] [3] [4]
T7 [0] [1] [2] [3] [4]
Care should be taken to ensure that the LIFX Tiles are all facing up to ensure that the 0,0 position is in the expected place.
Program will perform the following
- take a jpg or png located in the same folder as the notebook and create a image to display across all 4 tilechains or 20 tiles. Image will be reduced from original size to a 32x40 matrix so resolution will not be great. You've been warned.
```
!pip install pylifxtiles
#Main Program for Convert Single Image to Tiles
# Full running function with all dependencies
#imports RGB to HSBK conversion function from LIFX LAN library
from lifxlan import LifxLAN
from lifxlan.utils import RGBtoHSBK
from pylifxtiles import tiles
from pylifxtiles import actions
from matplotlib import image
from PIL import Image
# modify this variable to the name of the specific LIFX Tilechain as shown in the LIFX app
source_image = './images/Sunflowers.jpg'
def main():
lan = LifxLAN()
tilechain_lights = lan.get_tilechain_lights()
print(len(tilechain_lights))
if len(tilechain_lights) != 0:
for tile in tilechain_lights:
if tile.get_label() == 'T1':
print(tile.get_label())
T1 = tile
if tile.get_label() =='T2':
print(tile.get_label())
T2 = tile
if tile.get_label() == 'T3':
print(tile.get_label())
T3 = tile
if tile.get_label() == 'T4':
print(tile.get_label())
T4 = tile
if tile.get_label() == 'T5':
print(tile.get_label())
T5 = tile
if tile.get_label() == 'T6':
print(tile.get_label())
T6 = tile
if tile.get_label() == 'T7':
print(tile.get_label())
T7 = tile
tc_list = [ T1, T2, T3, T4, T5, T6, T7]
try:
display_image(source_image,(40,56), tc_list)
except KeyboardInterrupt:
print("Done.")
#combined function
# resize image and force a new shape and save to disk
def display_image(image_to_display,image_size, tilechain_list):
# load the image
my_image = Image.open(image_to_display)
# report the size of the image
#print(my_image.size)
# resize image and ignore original aspect ratio
img_resized = my_image.resize(image_size)
#changing the file extension from jpg to png changes output brightness. You might need to play with this.
img_resized.save('./images/resized_image.jpg')
data = image.imread('./images/resized_image.jpg')
target_tcs = []
for row in data:
temp_row = []
for pixel in row:
temp_row.append(RGBtoHSBK(pixel))
target_tcs.append(temp_row)
#print ("length of target_tcs is " + str(len(target_tcs)))
tcsplit = tiles.split_tilechains(target_tcs)
#print ("legnth of tcssplit is " + str(len(tcsplit)))
#print ("length tilelist is " + str(len(tilechain_list)))
for tile in range(len(tilechain_list)):
print (tile)
tilechain_list[tile].set_tilechain_colors(tiles.split_combined_matrix(tcsplit[tile]),rapid=True)
if __name__ == "__main__":
main()
```
# test write to three tiles
```
#Main Program for Convert Single Image to Tiles
# Full running function with all dependencies
#imports RGB to HSBK conversion function from LIFX LAN library
from lifxlan import LifxLAN
from lifxlan.utils import RGBtoHSBK
from pylifxtiles import tiles
from pylifxtiles import actions
from matplotlib import image
from PIL import Image
# modify this variable to the name of the specific LIFX Tilechain as shown in the LIFX app
source_image = './images/Youtubelogo.jpg'
def main():
lan = LifxLAN()
tilechain_lights = lan.get_tilechain_lights()
print(len(tilechain_lights))
if len(tilechain_lights) != 0:
for tile in tilechain_lights:
if tile.get_label() == 'T1':
print(tile.get_label())
T1 = tile
if tile.get_label() =='T2':
print(tile.get_label())
T2 = tile
if tile.get_label() == 'T3':
print(tile.get_label())
T3 = tile
if tile.get_label() == 'T4':
print(tile.get_label())
T4 = tile
tc_list = [T2, T3, T4]
try:
display_image(source_image,(40,24), tc_list)
except KeyboardInterrupt:
print("Done.")
#combined function
# resize image and force a new shape and save to disk
def display_image(image_to_display,image_size, tilechain_list):
# load the image
my_image = Image.open(image_to_display)
# report the size of the image
#print(my_image.size)
# resize image and ignore original aspect ratio
img_resized = my_image.resize(image_size)
#changing the file extension from jpg to png changes output brightness. You might need to play with this.
img_resized.save('./images/resized_image.jpg')
data = image.imread('./images/resized_image.jpg')
target_tcs = []
for row in data:
temp_row = []
for pixel in row:
temp_row.append(RGBtoHSBK(pixel))
target_tcs.append(temp_row)
print ("length of target_tcs is " + str(len(target_tcs)))
tcsplit = tiles.split_tilechains(target_tcs)
print ("legnth of tcssplit is " + str(len(tcsplit)))
print ("length tilelist is " + str(len(tilechain_list)))
for tile in range(len(tilechain_list)):
print (tile)
tilechain_list[tile].set_tilechain_colors(tiles.split_combined_matrix(tcsplit[tile]),rapid=True)
if __name__ == "__main__":
main()
```
| github_jupyter |
# Delicious Asian and Indian Cuisines
Install Imblearn which will enable SMOTE. This is a Scikit-learn package that helps handle imbalanced data when performing classification. (https://imbalanced-learn.org/stable/)
```
pip install imblearn
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib as mpl
import numpy as np
from imblearn.over_sampling import SMOTE
df = pd.read_csv('../../data/cuisines.csv')
```
This dataset includes 385 columns indicating all kinds of ingredients in various cuisines from a given set of cuisines.
```
df.head()
df.info()
df.cuisine.value_counts()
```
Show the cuisines in a bar graph
```
df.cuisine.value_counts().plot.barh()
thai_df = df[(df.cuisine == "thai")]
japanese_df = df[(df.cuisine == "japanese")]
chinese_df = df[(df.cuisine == "chinese")]
indian_df = df[(df.cuisine == "indian")]
korean_df = df[(df.cuisine == "korean")]
print(f'thai df: {thai_df.shape}')
print(f'japanese df: {japanese_df.shape}')
print(f'chinese df: {chinese_df.shape}')
print(f'indian df: {indian_df.shape}')
print(f'korean df: {korean_df.shape}')
```
## What are the top ingredients by class
```
def create_ingredient_df(df):
# transpose df, drop cuisine and unnamed rows, sum the row to get total for ingredient and add value header to new df
ingredient_df = df.T.drop(['cuisine','Unnamed: 0']).sum(axis=1).to_frame('value')
# drop ingredients that have a 0 sum
ingredient_df = ingredient_df[(ingredient_df.T != 0).any()]
# sort df
ingredient_df = ingredient_df.sort_values(by='value', ascending=False, inplace=False)
return ingredient_df
thai_ingredient_df = create_ingredient_df(thai_df)
thai_ingredient_df.head(10).plot.barh()
japanese_ingredient_df = create_ingredient_df(japanese_df)
japanese_ingredient_df.head(10).plot.barh()
chinese_ingredient_df = create_ingredient_df(chinese_df)
chinese_ingredient_df.head(10).plot.barh()
indian_ingredient_df = create_ingredient_df(indian_df)
indian_ingredient_df.head(10).plot.barh()
korean_ingredient_df = create_ingredient_df(korean_df)
korean_ingredient_df.head(10).plot.barh()
```
Drop very common ingredients (common to all cuisines)
```
feature_df= df.drop(['cuisine','Unnamed: 0','rice','garlic','ginger'], axis=1)
labels_df = df.cuisine #.unique()
feature_df.head()
```
Balance data with SMOTE oversampling to the highest class. Read more here: https://imbalanced-learn.org/dev/references/generated/imblearn.over_sampling.SMOTE.html
```
oversample = SMOTE()
transformed_feature_df, transformed_label_df = oversample.fit_resample(feature_df, labels_df)
print(f'new label count: {transformed_label_df.value_counts()}')
print(f'old label count: {df.cuisine.value_counts()}')
transformed_feature_df.head()
# export transformed data to new df for classification
transformed_df = pd.concat([transformed_label_df,transformed_feature_df],axis=1, join='outer')
transformed_df
transformed_df.info()
```
Save the file for future use
```
transformed_df.to_csv("../../data/cleaned_cuisine.csv")
```
| github_jupyter |
## _*LiH dissociation curve using VQE with UCCSD variational form*_
This notebook demonstrates using Qiskit Chemistry to plot graphs of the ground state energy of the Lithium Hydride (LiH) molecule over a range of inter-atomic distances using VQE and UCCSD. It is compared to the same energies as computed by the ExactEigensolver
This notebook populates a dictionary, that is a progammatic representation of an input file, in order to drive the Qiskit Chemistry stack. Such a dictionary can be manipulated programmatically and this is indeed the case here where we alter the molecule supplied to the driver in each loop.
This notebook has been written to use the PYSCF chemistry driver.
```
import numpy as np
import pylab
import copy
from qiskit.chemistry import QiskitChemistry
# Input dictionary to configure Qiskit Chemistry for the chemistry problem.
qiskit_chemistry_dict = {
'driver': {'name': 'PYSCF'},
'PYSCF': {'atom': '', 'basis': 'sto3g'},
'operator': {'name': 'hamiltonian', 'qubit_mapping': 'parity',
'two_qubit_reduction': True, 'freeze_core': True, 'orbital_reduction': [-3, -2]},
'algorithm': {'name': ''},
'optimizer': {'name': 'SLSQP', 'maxiter': 1000},
'variational_form': {'name': 'UCCSD'},
'initial_state': {'name': 'HartreeFock'}
}
molecule = 'H .0 .0 -{0}; Li .0 .0 {0}'
algorithms = ['VQE', 'ExactEigensolver']
pts = [x * 0.1 for x in range(6, 20)]
pts += [x * 0.25 for x in range(8, 16)]
pts += [4.0]
energies = np.empty([len(algorithms), len(pts)])
hf_energies = np.empty(len(pts))
distances = np.empty(len(pts))
dipoles = np.empty([len(algorithms), len(pts)])
eval_counts = np.empty(len(pts))
print('Processing step __', end='')
for i, d in enumerate(pts):
print('\b\b{:2d}'.format(i), end='', flush=True)
qiskit_chemistry_dict['PYSCF']['atom'] = molecule.format(d/2)
for j in range(len(algorithms)):
dict = copy.deepcopy(qiskit_chemistry_dict)
dict['algorithm']['name'] = algorithms[j]
if algorithms[j] == 'ExactEigensolver':
del dict['optimizer']
del dict['variational_form']
del dict['initial_state']
solver = QiskitChemistry()
result = solver.run(dict)
energies[j][i] = result['energy']
hf_energies[i] = result['hf_energy']
dipoles[j][i] = result['total_dipole_moment'] / 0.393430307
if algorithms[j] == 'VQE':
eval_counts[i] = result['algorithm_retvals']['eval_count']
distances[i] = d
print(' --- complete')
print('Distances: ', distances)
print('Energies:', energies)
print('Hartree-Fock energies:', hf_energies)
print('VQE num evaluations:', eval_counts)
pylab.plot(distances, hf_energies, label='Hartree-Fock')
for j in range(len(algorithms)):
pylab.plot(distances, energies[j], label=algorithms[j])
pylab.xlabel('Interatomic distance')
pylab.ylabel('Energy')
pylab.title('LiH Ground State Energy')
pylab.legend(loc='upper right');
pylab.plot(distances, np.subtract(hf_energies, energies[1]), label='Hartree-Fock')
pylab.plot(distances, np.subtract(energies[0], energies[1]), label='VQE')
pylab.xlabel('Interatomic distance')
pylab.ylabel('Energy')
pylab.title('Energy difference from ExactEigensolver')
pylab.legend(loc='upper left');
for j in reversed(range(len(algorithms))):
pylab.plot(distances, dipoles[j], label=algorithms[j])
pylab.xlabel('Interatomic distance')
pylab.ylabel('Moment in debye')
pylab.title('LiH Dipole Moment')
pylab.legend(loc='upper right');
pylab.plot(distances, eval_counts, '-o', color=[0.8500, 0.3250, 0.0980], label='VQE')
pylab.xlabel('Interatomic distance')
pylab.ylabel('Evaluations')
pylab.title('VQE number of evaluations')
pylab.legend(loc='upper left');
```
| github_jupyter |
## Manual publication DB insertion from raw text using syntax features
### Publications and conferences of Prof. Darabant Sergiu Adrian
#### http://www.cs.ubbcluj.ro/~dadi/
```
text = """
A Versatile 3D Face Reconstruction from Multiple Images for Face Shape Classification
Conference Paper
Sep 2019
Alexandru Ion Marinescu
Tudor Ileni
Adrian Sergiu Darabant
View
Fast In-the-Wild Hair Segmentation and Color Classification
Conference Paper
Jan 2019
Tudor Ileni
Diana Borza
Adrian Sergiu Darabant
In this paper we address the problem of hair segmentation and hair color classification in facial images using a machine learning approach based on both convolutional neural networks and classical neural networks. Hair with its color shades, shape and length represents an important feature of the human face and is used in domains like biometrics, v...
View
A Deep Learning Approach to Hair Segmentation and Color Extraction from Facial Images: 19th International Conference, ACIVS 2018, Poitiers, France, September 24–27, 2018, Proceedings
Chapter
Sep 2018
Diana Borza
Tudor Ileni
Adrian Sergiu Darabant
In this paper we tackle the problem of hair analysis in unconstrained images. We propose a fully convolutional, multi-task neural network to segment the image pixels into hair, face and background classes. The network also decides if the person is bald or not. The detected hair pixels are analyzed by a color recognition module which uses color feat...
View
Micro-Expressions Detection Based on Micro-Motions Dense Optical Flows
Conference Paper
Sep 2018
Sergiu Cosmin Nistor
Adrian Sergiu Darabant
Diana Borza
View
Automatic Skin Tone Extraction for Visagism Applications
Conference Paper
Jan 2018
Diana Borza
Adrian Sergiu Darabant
Radu Danescu
View
Figure 1. High-speed video acquisition and analysis process.
Figure 1. High-speed video acquisition and analysis process.
High-Speed Video System for Micro-Expression Detection and Recognition
Article
Full-text available
Dec 2017
Diana Borza
Radu Danescu
Razvan Itu
Adrian Sergiu Darabant
Micro-expressions play an essential part in understanding non-verbal communication and deceit detection. They are involuntary, brief facial movements that are shown when a person is trying to conceal something. Automatic analysis of micro-expression is challenging due to their low amplitude and to their short duration (they occur as fast as 1/15 to...
View
Supplementary Material
Data
Dec 2017
Diana Borza
Radu Danescu
Razvan Itu
Adrian Sergiu Darabant
View
Towards Automatic Skin Tone Classification in Facial Images
Conference Paper
Oct 2017
Diana Borza
Sergiu Cosmin Nistor
Adrian Sergiu Darabant
In this paper, we address the problem of skin tone classification in facial images, which has applications in various domains: visagisme, soft biometry and surveillance systems. We propose four skin tone classification algorithms and analyze their performance using different color spaces. The first two methods rely directly on pixel values, while t...
View
A linear approach to distributed database optimization using data reallocation
Conference Paper
Sep 2017
Adrian Sergiu Darabant
Viorica Varga
Leon Tambulea
View
Fig. 1. Flowchart of the proposed solution for gender classification
Fig. 4. Loss function of Inception-v4 trained with image distortions
Automatic gender recognition for “in the wild” facial images using convolutional neural networks
Conference Paper
Full-text available
Sep 2017
Sergiu Cosmin Nistor
Alexandra-Cristina Marina
Adrian Sergiu Darabant
Diana Borza
View
Fig. 1: The evaluation tree and the values associated to an example query.
Fig. 2: Fragment used by a binary operator-one operand is always a leaf...
Table 2 : Costs and exec times for MFRN=1 and MFRN=5, cases (a) and (b)
Fig. 4: Cost Improvements Percents for MFRN=1 and MFRN=5
Access Patterns Optimization in Distributed Databases Using Data Reallocation
Conference Paper
Full-text available
Aug 2017
Adrian Sergiu Darabant
Leon Tambulea
Viorica Varga
Large distributed databases are split into fragments stored on far distant nodes that communicate through a communication network. Query execution requires data transfers between the processing sites of the system. In this paper we propose a solution for minimizing raw data transfers by re-arranging and replicating existing data within the constrai...
View
Fast Eye Tracking and Feature Measurement using a Multi-stage Particle Filter
Conference Paper
Jan 2017
Radu Danescu
Adrian Sergiu Darabant
Diana Borza
View
Table 1 . Iris center localization accuracies compared to the...
Table 2 . Iris center localization results on the University of...
Table 3 . Iris radius computation results on the University of Michigan...
Table 4 . Performance of the eye shape segmentation algorithm the UMFD...
+3Table 5 . Mean error normalized by the inter-pupillary distance.
Real-Time Detection and Measurement of Eye Features from Color Images
Article
Full-text available
Jul 2016
Diana Borza
Adrian Sergiu Darabant
Radu Danescu
The accurate extraction and measurement of eye features is crucial to a variety of domains, including human-computer interaction, biometry, and medical research. This paper presents a fast and accurate method for extracting multiple features around the eyes: the center of the pupil, the iris radius, and the external shape of the eye. These features...
View
Magnetic Stimulation of the Spinal Cord: Evaluating the Characteristics of an Appropriate Stimulator
Article
Oct 2015
Mihaela Cretu
Adrian Sergiu Darabant
Radu V. Ciupa
This article aims to determine the necessary characteristics of a magnetic stimulator, capable of stimulating neural tracts of the spinal cord in a healthy subject. Our previous preliminary tests had shown that the commercial clinical magnetic stimulator Magstim Rapid2 was unable to reach excitable structures within the spinal cord, and only adjace...
View
Eyeglasses contour extraction using genetic algorithms
Conference Paper
Sep 2015
Diana Borza
Radu Danescu
Adrian Sergiu Darabant
This paper presents an eyeglasses contour extraction method that uses genetic algorithms to find the exact shape of the lenses. An efficient shape description, based on Fourier coefficients, is used to represent the shape of the eyeglasses, allowing a wide range of shapes to be represented with a small number of parameters. The proposed method does...
View
Figure 1. Eyeglasses detection algorithm outline.
Figure 2. Reconstruction of the rim contour using Fourier descriptors....
Table 2 . Detection rates.
Table 3 . Comparison of the proposed method with related works.
+4Figure 7. Eyeglasses region of interest (ROI). The detected position of...
Eyeglasses Lens Contour Extraction from Facial Images Using an Efficient Shape Description
Article
Full-text available
Oct 2013
Diana Borza
Adrian Sergiu Darabant
Radu Danescu
This paper presents a system that automatically extracts the position of the eyeglasses and the accurate shape and size of the frame lenses in facial images. The novelty brought by this paper consists in three key contributions. The first one is an original model for representing the shape of the eyeglasses lens, using Fourier descriptors. The seco...
View
Magnetic Stimulation of the Spinal Cord: Experimental Results and Simulations
Article
May 2013
Laura Darabant
Mihaela Cretu
Adrian Sergiu Darabant
This paper aims in interpreting the leg muscles responses recorded by electromyography during magnetic stimulation of the spinal cord by computing the electric field induced in the spinal cord and the nearby areas during this procedure. A simplified model of the spine was created and a Finite Difference Method algorithm was implemented in Matlab.
View
Fig. 4. Comparative FPE clustering results.
Fig. 5. Comparative results for small, medium and large datasets.
Clustering methods in data fragmentation
Article
Full-text available
Jan 2011
Adrian Sergiu Darabant
L. Darabant
This paper proposes an enhanced version for three clustering algorithms: hierarchical, k-means and fuzzy c-means applied in horizontal object oriented data fragmentation. The main application is focusing in distributed object oriented database (OODB) fragmentation, but the method applicability is not limited to this research area. The proposed algo...
View
Figure 1. Illuminated center of pupils
Figure 2. Auxiliary object with markers
Figure 3. Multiple reflections issue
Figure 4. Final preprocessing step: Canny Edge Detection and Closing
+2Figure 6. Center detection on binarized image of circle
Computer Vision Aided Measurement of Morphological Features in Medical Optics
Article
Full-text available
Sep 2010
Bologa Bogdana
Adrian Sergiu Darabant
This paper presents a computer vision aided method for non invasive interupupillary (IPD) distance measurement. IPD is a morphological feature requirement in any oftalmological frame prescription. A good frame prescription is highly dependent nowadays on accurate IPD estimation in order for the lenses to be eye strain free. The idea is to replace t...
View
Figure 1. Original video frame from the input video.
Figure 2. Foreground objects after subtraction.
Figure 3. Binary image(a), Eroded image(b).
Figure 4. Dilated image-blobs are well separated and compact.
+3Figure 5. Normal blobs(a), Blobs with holes(b), Fragmented blobs(c).
A Computer Vision Approach to Object Tracking and Counting
Article
Full-text available
Sep 2010
Mezei Sergiu
Adrian Sergiu Darabant
This paper, introduces a new method for counting people or more generally objects that enter or exit a certain area/building or perimeter. We propose an algorithm (method) that analyzes a video sequence, detects moving objects and their moving direction and filters them according to some criteria (ex only humans). As result one obtains in and out c...
View
Energy Efficient Coils for Magnetic Stimulation of Peripheral Nerves
Article
Apr 2009
Laura Darabant
M. Plesa
Dan Micu[...]
Adrian Sergiu Darabant
The preoccupation for improving the quality of life, for persons with different handicaps, led to extended research in the area of functional stimulation. Due to its advantages compared to electrical stimulation, magnetic stimulation of the human nervous system is now a common technique in modern medicine. A difficulty of this technique is the need...
View
Hierarchical clustering in large object datasets – a study on complexity, quality and scalability
Article
Jan 2009
Adrian Sergiu Darabant
Anca Andreica
Object database fragmentation (horizontal fragmentation) deals with splitting the extension of classes into subsets according to some criteria. The resulting fragments are then used either in distributed database processing or in parallel data processing in order to spread the computation power over multiple nodes or to increase data locality featu...
View
A medical application of electromagnetic fields: The magnetic stimulation of nerve fibers inside a cylindrical tissue
Conference Paper
Jun 2008
M. Plesa
L. Darabant
R. Ciupa
Adrian Sergiu Darabant
A model is presented that predicts the electric field induced in the arm during magnetic stimulation of a peripheral nerve. The arm is represented as a homogeneous, cylindrical volume conductor. The electric field arises from two sources: the time - varying magnetic field and the accumulation of charge on the tissue - air surface. In magnetic stimu...
View
Fig. 2-The MobMed System Architecture and Integration with the Hospital...
Fig 3-Merge Replication Architecture .
Fig. 6 MobMed's login window
Fig. 7 MobMed's main and patient form
Mobile Devices and Data Synchronization Assisting Medical Diagnosis
Article
Full-text available
Jun 2008
Adrian Sergiu Darabant
Horea Todoran
In order to be able to establish the most accurate diagnostics as quick as possible, medical doctors need fast access not only to the current patient state and test results but also to its historical medical data. With the diversity of the malady symptoms today a correct diagnostic often requires a valuable time that is not always available due to...
View
Web services for e-learning and e-recruitment
Article
Jan 2007
George Chis
Horea Grebla
D. Matis[...]
Adrian Sergiu Darabant
Mobile phone communication can no longer be conceived as a communication mean only, but also as a way to integrate voice services together with data services which are oriented towards large consumer groups. Together with voice services, mobile Internet represents the second most important component of the service packages offered in Romania. The a...
View
Fig. 3 Comparative PE costs for variant M1 on all classes.
Fig. 5-PE values for M1 on complex class fragmentation and primary...
The similarity measures and their impact on OODB fragmentation using hierarchical clustering algorithms
Article
Full-text available
Sep 2006
Adrian Sergiu Darabant
Horea Todoran
Octavian Creţ
George Chis
Class fragmentation is an essential phase in the design of Distributed Object Oriented Databases (DOODB). Due to their semantic similarity with the purpose of database fragmentation (obtaining sets of similar objects with respect to the user applications running in the system), clustering algorithms have recently begun to be investigated in the pro...
View
Building an efficient architecture for data synchronization on mobile wireless agents
Article
Aug 2006
Adrian Sergiu Darabant
H. Todoran
Nowadays, negotiation between a representative of a commercial enterprise and its clients is a pre-requisite for selling most of the industrial goods in large quantities. In many cases, it is the task of a mobile salesman to conduct the negotiation on behalf of the supplier. But this is not an easy task to accomplish, since the mobile agent must br...
View
Fig.1 Business information flow
Fig. 2 – The MobSel System Architecture and Integration with the...
Fig 5 Controlling the synchronized data.
Implementing data synchronization on mobile wireless agents
Conference Paper
Full-text available
Jul 2006
Adrian Sergiu Darabant
Horea Todoran
Mobile salesmen out in the wild and the entire commercial store with them? A while ago this would have seemed atypical. Nowadays, it has become a must have asset for any salesmen-based commercial enterprise. In the past, the Web brought the virtual store to the client's premises. While this is still enough for certain types of commerce, negotiation...
View
Table 1 . Results of the software (C++) implementation
Table 2 . Results of the software implementation
Fig. 3. The hardware architecture in the 1D case
Solving the Maximum Subsequence Problem with a Hardware Agents-based System
Article
Full-text available
Jul 2006
Octavian Creţ
Zsolt Mathe
Cristina Grama[...]
Adrian Sergiu Darabant
The maximum subsequence problem is widely encountered in various digital processing systems. Given a stream of both positive and negative integers, it consists of determining the subsequence of maximal sum inside the input stream. In its two-dimensional version, the input is an array of both positive and negative integers, and the problem consists...
View
Figure 1: The fragmentation costs for the CBAk(incremental) and...
Table 1 : Comparative results for the CBAk and k-means algorithms
Incremental Horizontal Fragmentation: A new Approach in the Design of Distributed Object Oriented Databases
Article
Full-text available
Jan 2006
Adrian Sergiu Darabant
Alina Campan
Horea Todoran
Distributed relational or more recently object-oriented databases usually employ data fragmenta-tion techniques during the design phase in order to split and allocate the database entities across the nodes of the system. Most of the design algorithms are usually static and do not take into account the system evolution: data updates and addition of...
View
A Hardware Implementation of the Kadane’s Algorithm for the Maximum Subsequence Problem
Conference Paper
Jan 2006
Octavian Creţ
Zsolt Mathe
Lucia Văcariu[...]
Levente-Karoly Gorog
View
"The School in Your Pocket": Useful PoeketPC applications for students
Article
Jan 2006
Horea Todoran
Adrian Sergiu Darabant
Much smaller than laptops and still suitable for almost all kinds of applications, hand-held devices have the potential to rapidly become interesting tools for various daily activities. They can be successfully used in education by all participants (students, educators, administrative staff), if helpful applications are carefully designed and imple...
View
Figure 2. Macroflows composed of connections originating from different...
Fine-Grained Macroflow Granularity in Congestion Control Management
Article
Full-text available
Jun 2005
Darius Bufnea
Alina Campan
Adrian Sergiu Darabant
A recent approach in Internet congestion control suggests collaboration between sets of streams that should share network resources and learn from each other about the state of the network. Currently such a set of collaborating streams – a macroflow – is organized on host pair basis. We propose in this paper a new method for grouping streams into m...
View
Figure 3 Fuzzy fragmentation vs k-means primary and k-means...
Using Fuzzy Clustering for Advanced OODB Horizontal Fragmentation with Fine-Grained Replication.
Conference Paper
Full-text available
Jan 2005
Adrian Sergiu Darabant
Alina Campan
Octavian Creţ
In this paper we present a new approach for horizontal object oriented database fragmentation combined with fine-grained object level replication in one step. We build our fragmentation/replication method using AI probabilis- tic clustering (fuzzy clustering). Fragmentation quality evaluation is provided using an evaluator function.
View
Figure 1: The database class hierarchy
Figure 2: Experimental results
CLUSTERING TECHNIQUES FOR ADAPTIVE HORIZONTAL FRAGMENTATION IN OBJECT ORIENTED DATABASES
Article
Full-text available
Jan 2005
Alina Campan
Adrian Sergiu Darabant
Gabriela Serban
Optimal application performance in a Distributed Object Ori- ented System requires class fragmentation and the development of allocation schemes to place fragments at distributed sites so data transfer is minimal. A horizontal fragmentation approach that uses data mining clustering methods for partitioning object instances into fragments has alread...
View
Figure 1: The database class hierarchy
Figure 2: The database aggregation/association graph
Figure 3: Comparative PE values for our fragmentation method,...
Figure 4: Comparative class PE values for each similarity measure
Figure 5: Comparative PE values for primary only fragmentation and our...
A NEW APPROACH IN FRAGMENTATION OF DISTRIBUTED OBJECT ORIENTED DATABASES USING CLUSTERING TECHNIQUES
Article
Full-text available
Jan 2005
Adrian Sergiu Darabant
Horizontal fragmentation plays an important role in the design phase of Distributed Databases. Complex class relationships: associations, aggregations and complex methods, require fragmentation algorithms to take into account the new problem dimensions induced by these features of the object oriented models. We propose in this paper a new method fo...
View
Table 1 .
Table 2 .
Figure 3. Parameter transmission in the SW array
Figure 4. The interface of a PE and the connections between adjacent...
FPGA-based Scalable Implementation of the General Smith-Waterman Algorithm
Conference Paper
Full-text available
Nov 2004
Octavian Creţ
Stefan Mathe
Balint Szente[...]
Adrian Sergiu Darabant
The Smith-Waterman algorithm is fundamental in Bioinformatics. This paper presents an FPGA-based systolic implementation of the Smith-Waterman algorithm that addresses a general case of it. A solution that improves the scalability of the design is proposed. The architecture is optimized for both speed and space, by reusing the hardware resources fo...
View
TABLE 2 . Allocation of Fragments to Distributed Sites
Fig. 3. Comparative quality measures for each class.
Fig. 4. Comparative PE for k-means, full replication and centralized case.
Fig. 5. Comparative PE values for our fragmentation methods.
Semi-supervised learning techniques: k-means clustering in OODB Fragmentation
Conference Paper
Full-text available
Feb 2004
Adrian Sergiu Darabant
Alina Campan
Vertical and horizontal fragmentations are central issues in the design process of distributed object based systems. A good fragmentation scheme followed by an optimal allocation could greatly enhance performance in such systems, as data transfer between distributed sites is minimized. In this paper we present a horizontal fragmentation approach th...
View
Figure 1: The database inheritance hierarchy
Figure 2: The database aggregation hierarchy
Figure 3: Partial RelGraph-CAN values and weights
Figure 4: Comparative PE values for each class
Figure 5: Comparative PE values for different fragmentation orders
A new approach for optimal fragmentation order in distributed object oriented databases
Article
Full-text available
Feb 2004
Adrian Sergiu Darabant
Alina Campan
Class fragmentation is an important task in the design of Distributed OODBs and there are many algorithms handling it. Almost none of them deals however with the class fragmentation order details. We claim that class fragmentation order can induce severe performance penalties if not considered in the frag- mentation phase. We propose here two varia...
View
Figure 1. The database inheritance hierarchy
Figure 2. The database aggregation hierarchy
OPTIMAL CLASS FRAGMENTATION ORDERING IN OBJECT ORIENTED DATABASES
Article
Full-text available
Jan 2004
Adrian Sergiu Darabant
Alina Campan
Distributed Object Oriented Databases require class fragmenta- tion, performed either horizontally or vertically. Complex class relationships like aggregation and/or association are often represented as two-way refer- ences or object-links between classes. In order to obtain a good quality horizontal fragmentation, an optimal class processing order...
View
TABLE 1.
Figure 3 Comparative PE values for our fragmentation method,...
Figure 4 Comparative class PE values for each similarity measure.
AI CLUSTERING TECHNIQUES: A NEW APPROACH IN HORIZONTAL FRAGMENTATION OF CLASSES WITH COMPLEX ATTRIBUTES AND METHODS IN OBJECT ORIENTED DATABASES
Article
Full-text available
Jan 2004
Adrian Sergiu Darabant
Alina Campan
Grigor Moldovan
Horea Grebla
Horizontal fragmentation plays an important role in the design phase of Distributed Databases. Complex class relationships: associations, aggregations and complex methods, require fragmentation algorithms to take into account the new problem dimensions induced by these features of the object oriented models. We propose in this paper a new method fo...
View
DATA ALLOCATION IN DISTRIBUTED DATABASE SYSTEMS PERFORMED BY MOBILE INTELLIGENT AGENTS
Article
Full-text available
Jan 2004
Horea Grebla
Grigor Moldovan
Adrian Sergiu Darabant
Alina Campan
As the European Union extends its boundaries the major companies have extended their presence on different markets resulting sales expansion and marketing specialization. Moreover, globalization brings a bigger impact on vital business's data because of the applications that have been developed on platforms having specific aspects by means of datab...
View
Figure 2. Comparative PE for k-means, full replication and centralized...
Figure 3. Comparison quality measures for each of our fragmentation...
Advanced Object Database Design Techniques
Article
Full-text available
Jan 2004
Adrian Sergiu Darabant
Alina Ampan
Class fragmentation is an important task in the design of Distributed Object Oriented Databases (DOOD). However, fragmentation in DOOD is still at its beginnings and mostly adapted from the relational approaches. In this paper we propose an alternative approach for horizontal fragmentation of DOOD. Our method uses two different AI clustering techni...
View
Fig. 2 . CREC development system
A hardware/software codesign method for general purpose reconfigurable computing
Conference Paper
Full-text available
Jul 2003
Octavian Creţ
Kalman Pusztai
Cristian Cosmin Vancea[...]
Adrian Sergiu Darabant
CREC is an original, low-cost general-purpose Reconfigurable Computer whose architecture is generated through a Hardware / Software CoDesign process. The main idea of the CREC computer is to generate the best-suited hardware architecture for the execution of each software application. The CREC Parallel Compiler parses the source code and generates...
View
Current Technologies in Automatic Test Suites Generation and Verification of Complex Systems
Article
Full-text available
Jan 1999
Adrian Sergiu Darabant
View
Multi-tiered client-server techniques for distributed database systems
Article
Jan 1998
Adrian Sergiu Darabant
Information explosion across all areas has determined an increase in hardware requirements for application that provide data to the users. As hardware evelopment is quite susceptible to be bound after a top barrier is reached, new technologies must be developed in the software area in order to keep up with the requirements. We present here such a t...
View
Fig. 2. The database class hierarchy
Fig. 3. The database aggregation/association graph
Hierarchical clustering in object oriented data models with complex class relationships
Article
Full-text available
Adrian Sergiu Darabant
Alina Campan
Octavian Creţ
Class fragmentation is an essential phase in the design of Distributed Object Oriented Databases (DOODB). Horizontal and vertical fragmentation are the two commonly used fragmentation techniques. We propose here two new methods for horizontal fragmentation of objects with complex attributes. They rely on AI clustering techniques for grouping object...
View
Energy Efficient Coils for Transcranial Magnetic Stimulation (TMS)
Article
Laura DARABANT
M. Plesa
Radu CIUPA[...]
Adrian Sergiu Darabant
The preoccupation for improving the quality of life, for persons with different handicaps, led to extended research in the area of functional stimulation. Due to its advantages compared to electrical stimulation, magnetic stimulation of the human nervous system is now a common technique in modern medicine. A difficulty of this technique is the need...
View
Fig.2: Web Services for E-Learning
E-Learning Services as a Recruitment Tool
Article
Full-text available
George Chis
Horea Grebla
DUMITRU MATIS[...]
Adrian Sergiu Darabant
Networks expansion and Internet provide a good platform for e-learning in the idea of connecting learners with educational resources. The various systems that are already implemented consider the learning process as a remote task to gather knowledge in order to pass some exams. In the learning process evaluation represents a final step for a course...
View
A Comparative Study of Horizontal Object Clustering-based Fragmentation Techniques
Article
Adrian Sergiu Darabant
Alina Campan
Design of modern Distributed Object Oriented Databases (DOODs) requires class fragmentation techniques. Although research has been conducted in this area, most of the developed methods are inspired from the relational fragmentation algorithms. In this paper we develop a comparative approach of two new methods for horizontal class fragmentation in a...
View
TABLE 2 . OCM -exceptional case
TABLE 3 . CVM -for OCM
Fig. 4. Comparative quality measures for fragmentation variants,...
TABLE 4 . OCM -with phantom object
TABLE 5 . CVM -with phantom object
AI Clustering Techniques: a New Approach to Object Oriented Database Fragmentation
Article
Full-text available
Adrian Sergiu Darabant
Alina Campan
Cluj Napoca
M Kogalniceanu
Optimal application performance on a Distributed Object Based System requires class fragmentation and the development of allocation schemes to place fragments at distributed sites so data transfer is minimal. In this paper we present a horizontal fragmentation approach that uses the k-means centroid based clustering method for partitioning object i...
View
A Comparative Study on the Influence of Similarity Measures in Hierarchical Clustering in Complex Distributed Object-Oriented Databases
Article
Full-text available
Adrian Sergiu Darabant
Horea Todoran
Octavian Creţ
George Chis
Class fragmentation is an essential phase in the design of Distributed Object Oriented Databases (DOODB). Due to their semantic similarity with the purpose of database fragmentation (obtaining sets of similar objects with respect to the user applications running in the system), clustering algorithms have recently begun to be investigated in the pro...
View
Figure 1. Medical information flow
Figure 2. The MobMed Architecture and Integration with the Hospital...
Figure 3. Merge Replication Architecture
EFFICIENT DATA SYNCHRONIZATION FOR MOBILE WIRELESS MEDICAL USERS
Article
Full-text available
Adrian Sergiu Darabant
Darabant And
Horea Todoran
In order to take the appropriate decisions as quick as possible, medical doctors need fast access to various pieces of information on their pa-tients. The required information should be accurate, up-to-date, and avail-able on the spot. Even more, after finishing his/her investigation, the medical doctor should be able to immediately forward the rel...
View
Implementing Efficient Data Synchronization for Mobile Wireless Medical Users
Article
Full-text available
Adrian Sergiu Darabant
In order to take the appropriate decisions as quick as possible, medical doctors need fast access to various pieces of information on their patients. The required information should be accurate, up-to-date, and available on the spot. Even more, after finishing his/her investigation, the medical doctor should be able to immediately forward the relev...
View
"""
import re
class HelperMethods:
@staticmethod
def IsDate(text):
# print("text")
# print(text)
val = re.match("(Jan|Feb|Mar|Apr|May|Jun|Jul|Aug|Sep|Oct|Nov|Dec) (1|2)(0|9)[0-9]{2}", text)
if not val:
return False
return True
mylines = []
ctr = 0
title = ""
authors = ""
affiliations = ""
date = ""
papers = []
titles = []
dates = []
for line in text.split('View')[1:-1]:
fields = []
current_date = None
print(line.split('\n'))
for field in line.split('\n'):
val = re.match("(\+[0-9])?(Figure|Fig[\.]?|Table|TABLE)( )?[0-9]+", field)
if val:
continue
if field == "":
continue
print("field: ", field)
fields.append(field)
if HelperMethods.IsDate(field):
current_date = field
title = fields[0]
papers.append((title, current_date))
print(len(papers))
print(papers)
for i, paper in enumerate(papers):
print(i, paper)
#mylines[i][0] = mylines[i][1]
```
# DB Storage (TODO)
Time to store the entries in the `papers` DB table.

```
import mariadb
import json
with open('../credentials.json', 'r') as crd_json_fd:
json_text = crd_json_fd.read()
json_obj = json.loads(json_text)
credentials = json_obj["Credentials"]
username = credentials["username"]
password = credentials["password"]
table_name = "publications_cache"
db_name = "ubbcluj"
mariadb_connection = mariadb.connect(user=username, password=password, database=db_name)
mariadb_cursor = mariadb_connection.cursor()
import datetime
from datetime import datetime
for paper in papers:
title = ""
authors = ""
pub_date = ""
affiliations = ""
try:
title = paper[0].lstrip()
except:
pass
try:
# print(paper[1])
pub_date = datetime.strptime(paper[1], "%b %Y").strftime("%Y-%m-%d")
except:
pass
insert_string = "INSERT INTO {0} SET ".format(table_name)
insert_string += "Title=\'{0}\', ".format(title)
insert_string += "ProfessorId=\'{0}\', ".format(12)
if pub_date != "":
insert_string += "PublicationDate=\'{0}\', ".format(str(pub_date))
insert_string += "Authors=\'{0}\', ".format(authors)
insert_string += "Affiliations=\'{0}\' ".format(affiliations)
print(insert_string)
# print(paper)
# continue
try:
mariadb_cursor.execute(insert_string)
except mariadb.ProgrammingError as pe:
print("Error")
raise pe
except mariadb.IntegrityError:
continue
mariadb_connection.close()
```
# Conclusion
### In the end, the DB only required ~1 manual modifications with this code.
This was first stored in a DB cache table which is a duplicate of the main, reviewed, then inserted in the main table.
| github_jupyter |
```
# General Dependencies
import os
import numpy as np
# Denoising dependencies
from trefide.pmd import batch_decompose,\
batch_recompose,\
overlapping_batch_decompose,\
overlapping_batch_recompose,\
determine_thresholds
from trefide.reformat import overlapping_component_reformat
# Plotting & Video Rendering Dependencies
import funimag
import matplotlib.pyplot as plt
from trefide.plot import pixelwise_ranks
from trefide.video import play_cv2
# Set Demo Dataset Location
ext = os.path.join("..", "example_movies")
filename = os.path.join(ext, "demoMovie.tif")
%load_ext autoreload
%autoreload 2
```
# Load Data
```
from skimage import io
mov = io.imread(filename).transpose([1,2,0])[:60,:60,:]
mov = np.asarray(mov,order='C',dtype=np.float64)
print(mov.shape)
fov_height, fov_width, num_frames = mov.shape
```
# Set Params
```
# Maximum of rank 50 blocks (safeguard to terminate early if this is hit)
max_components = 50
# Enable Decimation
max_iters_main = 10
max_iters_init = 40
d_sub=2
t_sub=2
# Defaults
consec_failures = 3
tol = 0.0005
# Set Blocksize Parameters
block_height = 20
block_width = 20
overlapping = True
```
# Compress Video
## Simulate Critical Region with Noise
```
spatial_thresh, temporal_thresh = determine_thresholds((fov_height, fov_width, num_frames),
(block_height, block_width),
consec_failures, max_iters_main,
max_iters_init, tol,
d_sub, t_sub, 5, True)
```
## Decompose Each Block Into Spatial & Temporal Components
```
# Blockwise Parallel, Single Tiling
if not overlapping:
spatial_components,\
temporal_components,\
block_ranks,\
block_indices = batch_decompose(fov_height, fov_width, num_frames,
mov, block_height, block_width,
max_components, consec_failures,
max_iters_main, max_iters_init, tol,
d_sub=d_sub, t_sub=t_sub)
# Blockwise Parallel, 4x Overlapping Tiling
else:
spatial_components,\
temporal_components,\
block_ranks,\
block_indices,\
block_weights = overlapping_batch_decompose(fov_height, fov_width, num_frames,
mov, block_height, block_width,
spatial_thresh, temporal_thresh,
max_components, consec_failures,
max_iters_main, max_iters_init, tol,
d_sub=d_sub, t_sub=t_sub)
```
# Reconstruct Denoised Video
```
# Single Tiling (No need for reqweighting)
if not overlapping:
mov_denoised = np.asarray(batch_recompose(spatial_components,
temporal_components,
block_ranks,
block_indices))
# Overlapping Tilings With Reweighting
else:
mov_denoised = np.asarray(overlapping_batch_recompose(fov_height, fov_width, num_frames,
block_height, block_width,
spatial_components,
temporal_components,
block_ranks,
block_indices,
block_weights))
```
# Produce Diagnostics
### Single Tiling Pixel-Wise Ranks
```
if overlapping:
pixelwise_ranks(block_ranks['no_skew']['full'], fov_height, fov_width, num_frames, block_height, block_width)
else:
pixelwise_ranks(block_ranks, fov_height, fov_width, num_frames, block_height, block_width)
```
### Correlation Images
```
from funimag.plots import util_plot
util_plot.comparison_plot([mov, mov_denoised + np.random.randn(np.prod(mov.shape)).reshape(mov.shape)*.01],
plot_orientation="vertical")
```
## Render Videos & Residual
```
play_cv2(np.vstack([mov, mov_denoised, mov-mov_denoised]), magnification=2)
```
# Save Results
```
U, V = overlapping_component_reformat(fov_height, fov_width, num_frames,
block_height, block_width,
spatial_components,
temporal_components,
block_ranks,
block_indices,
block_weights)
np.savez(os.path.join(ext, "demo_results.npz"), U, V,block_ranks,block_height,block_width)
```
| github_jupyter |
# Dynamic Time Warping
Goal: To identify counties which are "further along the curve" and use them to assist in prediction of other counties.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from fastdtw import fastdtw
from scipy.spatial.distance import euclidean
from sklearn.cluster import KMeans
import datetime
from fbprophet import Prophet
from fbprophet.plot import add_changepoints_to_plot
from fbprophet.diagnostics import cross_validation
from fbprophet.diagnostics import performance_metrics
from fbprophet.plot import plot_cross_validation_metric
import logging
logging.getLogger('fbprophet').setLevel(logging.WARNING)
deaths = pd.read_csv("data/us/covid/nyt_us_counties_daily.csv")
population = pd.read_csv("data/us/demographics/county_populations.csv")
```
### Los Angeles County vs. NYC County
```
la_fips = 6037
la_data = deaths.loc[deaths["fips"] == la_fips]
la_data.head()
ny_fips = 36061
ny_data = deaths.loc[deaths["fips"] == ny_fips]
ny_data.head()
# Construct datasets: deaths as a proportion of population
la = {"dates": [], "deaths": []}
la_pop = int(population.loc[population["FIPS"] == la_fips]["total_pop"])
for idx, row in la_data.iterrows():
la["dates"].append(row["date"])
la["deaths"].append(row["deaths"] / la_pop)
ny = {"dates": [], "deaths": []}
ny_pop = int(population.loc[population["FIPS"] == ny_fips]["total_pop"])
for idx, row in la_data.iterrows():
ny["dates"].append(row["date"])
ny["deaths"].append(row["deaths"] / ny_pop)
distance, path = fastdtw(np.array(la["deaths"]), np.array(ny["deaths"]), dist=euclidean)
print(distance)
print(path)
# Plot the warped data
overlay = {"dates": [], "la_deaths": [], "ny_deaths": []}
for (la_idx, ny_idx) in path:
overlay["dates"].append(la["dates"][la_idx])
overlay["la_deaths"].append(la["deaths"][la_idx])
overlay["ny_deaths"].append(ny["deaths"][ny_idx])
plt.figure(figsize=(10, 5))
plt.plot(overlay["la_deaths"], label="LA", c="blue")
plt.plot(overlay["ny_deaths"], label="NYC", c="green")
plt.legend()
plt.show()
```
### Matching a Smaller Window
```
overlay_window = 14
test_per = 14
interval = 1
ny_df = pd.DataFrame(data=ny)
ny_start = ny_df.loc[ny_df["deaths"] > 5e-7].first_valid_index()
print(ny_start)
best_j, best_dist, best_path = 0, 1, []
i = len(la["dates"]) - overlay_window - test_per
for j in range(ny_start, len(ny["dates"]) - test_per, interval):
distance, path = fastdtw(np.array(la["deaths"][i:i+overlay_window]), np.array(ny["deaths"][j:j+overlay_window]), dist=euclidean)
if distance < best_dist:
best_j = j
best_dist = distance
best_path = path
print(best_j)
# Align data by calculated indices
diff = i - best_j
overlay = pd.DataFrame(data = la)
overlay["ny"] = 0
overlay.tail()
for i, row in overlay.iterrows():
if i - diff >= 0 and i - diff < len(overlay["dates"]):
overlay.loc[i, "ny"] = ny["deaths"][i - diff]
last_date = datetime.date.fromisoformat(overlay.iloc[-1]["dates"])
date = last_date
for i in range(diff):
date += datetime.timedelta(days=1)
overlay.loc[len(overlay)] = [date.isoformat(), 0, ny["deaths"][len(ny["deaths"]) - diff + i]]
overlay.tail()
plt.figure(figsize=(10, 5))
plt.plot(overlay["deaths"][:len(overlay["deaths"]) - test_per - diff], label="LA train", c="black")
plt.plot(overlay["deaths"][len(overlay["deaths"]) - test_per - diff:len(overlay["deaths"]) - diff], label="LA test", c="red")
plt.plot(overlay["ny"][:len(overlay["ny"]) - test_per], label="NYC", c="green")
plt.legend()
plt.show()
```
## Using DTW for Prediction
```
overlay.rename(columns={'dates': 'ds', 'deaths': 'y'}, inplace=True)
m = Prophet(seasonality_mode="multiplicative", interval_width = 0.68)
m.add_regressor("ny")
m.fit(overlay.iloc[:len(overlay) - diff - test_per])
future = pd.DataFrame()
future["ds"] = 0
future["ny"] = 0
d = last_date - datetime.timedelta(days=test_per)
for i in range(test_per):
d += datetime.timedelta(days=1)
try:
idx = ny["dates"].index((d - datetime.timedelta(days=overlay_window)).isoformat())
future.loc[len(future)] = [d.isoformat(), ny["deaths"][idx]]
except ValueError as e:
future.loc[len(future)] = [d.isoformat(), 0]
future.tail()
pred = m.predict(future)
pred["yhat"] *= la_pop
la_plot = pd.DataFrame()
la_plot["date"] = 0
la_plot["deaths"] = 0
for i, row in la_data.iterrows():
la_plot.loc[len(la_plot)] = [datetime.date.fromisoformat(row["date"]), row["deaths"]]
la_plot.tail()
fig, ax = plt.subplots()
ax.plot('date', 'deaths', data=la_plot, label="Actual")
ax.plot('ds', 'yhat', data=pred, label="Predicted")
plt.legend()
plt.show()
```
### Compared with Prophet On Its Own
```
m = Prophet(seasonality_mode="multiplicative", interval_width = 0.68)
m.fit(overlay.iloc[:len(overlay) - diff - test_per])
future = pd.DataFrame()
future["ds"] = 0
d = last_date - datetime.timedelta(days=test_per)
for i in range(test_per):
d += datetime.timedelta(days=1)
future.loc[len(future)] = [d.isoformat()]
future.tail()
pred = m.predict(future)
pred["yhat"] *= la_pop
fig, ax = plt.subplots()
ax.plot('date', 'deaths', data=la_plot, label="Actual")
ax.plot('ds', 'yhat', data=pred, label="Predicted")
plt.legend()
plt.show()
```
## Finding a Best-Fit County
```
fips_list = pd.read_csv("data/us/processing_data/fips_key.csv", encoding="cp1252")
best_j, best_fips, best_dist, best_path = 0, 0, 1, []
i = len(la["dates"]) - overlay_window - test_per
for idx, rw in fips_list.iterrows():
try:
county = int(rw["FIPS"])
if (county == la_fips): continue
print("County " + str(county) + "...", end='\r', flush=True)
c_data = deaths.loc[deaths["fips"] == county]
c = {"dates": [], "deaths": []}
c_pop = int(population.loc[population["FIPS"] == county]["total_pop"])
for idx, row in c_data.iterrows():
c["dates"].append(row["date"])
c["deaths"].append(row["deaths"] / c_pop)
c_df = pd.DataFrame(data=c)
c_start = c_df.loc[c_df["deaths"] > 5e-7].first_valid_index()
if (c_start == None): continue
for j in range(c_start, len(c["dates"]) - test_per, 1):
if i - j < test_per: continue
distance, path = fastdtw(np.array(la["deaths"][i:i+overlay_window]), np.array(c["deaths"][j:j+overlay_window]), dist=euclidean, radius=3)
if distance < best_dist:
best_j = j
best_fips = county
best_dist = distance
best_path = path
except TypeError as e:
continue
print()
print(best_j)
print(best_fips)
print(best_dist)
best_data = deaths.loc[deaths["fips"] == best_fips]
best = {"dates": [], "deaths": []}
best_pop = int(population.loc[population["FIPS"] == best_fips]["total_pop"])
for idx, row in best_data.iterrows():
best["dates"].append(row["date"])
best["deaths"].append(row["deaths"] / c_pop)
best_df = pd.DataFrame(data=c)
best_start = best_df.loc[best_df["deaths"] > 5e-7].first_valid_index()
# Align data by calculated indices
diff = i - best_j
overlay = pd.DataFrame(data = la)
overlay["best_fit"] = 0
overlay["dates"]
for i, row in overlay.iterrows():
if i - diff >= 0 and i - diff < len(overlay["dates"]):
overlay.loc[i, "best_fit"] = best["deaths"][i - diff]
last_date = datetime.date.fromisoformat(overlay.iloc[-1]["dates"])
date = last_date
for i in range(diff):
date += datetime.timedelta(days=1)
overlay.loc[len(overlay)] = [date.isoformat(), 0, best["deaths"][len(best["deaths"]) - diff + i]]
overlay.tail()
overlay.rename(columns={'dates': 'ds', 'deaths': 'y'}, inplace=True)
m = Prophet(seasonality_mode="multiplicative", interval_width = 0.68)
m.add_regressor("best_fit")
m.fit(overlay.iloc[:len(overlay) - diff - test_per])
future = pd.DataFrame()
future["ds"] = 0
future["best_fit"] = 0
d = last_date - datetime.timedelta(days=test_per)
for i in range(test_per):
d += datetime.timedelta(days=1)
try:
future.loc[len(future)] = [d.isoformat(), float(overlay.loc[overlay["ds"] == d.isoformat()]["best_fit"])]
except ValueError as e:
future.loc[len(future)] = [d.isoformat(), 0]
future.tail()
pred = m.predict(future)
pred["yhat"] *= la_pop
fig, ax = plt.subplots()
ax.plot('date', 'deaths', data=la_plot, label="Actual")
ax.plot('ds', 'yhat', data=pred, label="Predicted")
plt.legend()
plt.show()
pred.head()
```
# Generating a Submission Using Dynamic Time Warping
```
class Covid_DTW:
def __init__(self, death_df, population_df, fips_df, county_to_cluster, cluster_to_counties):
"""
death_df should have columns fips, date, and deaths
population_df should have columns fips, population
fips_df should have coulumn fips
"""
self.data = death_df
self.pop = population_df
self.fips = fips_df
self.last_date = None
self.co_cl = county_to_cluster
self.cl_co = cluster_to_counties
self.num_found_fit = 0
def get_best_fit(self, target, target_county, dist_func, overlay_window, test_per, pred_per, death_lbound, log=True):
best_idx, best_fips, best_dist = -1, -1, float("inf")
target_idx = len(target["dates"]) - overlay_window - test_per
cluster = self.co_cl[str(target_county)]
cluster_fips = self.cl_co[cluster]
for county in cluster_fips:
try:
if (county == target_county): continue
if log: print("Testing best fit for " + str(target_county) + " on " + str(county) + "...", end='\r', flush=True)
c_data = self.data.loc[self.data["fips"] == int(county)]
c = {"dates": [], "deaths": []}
c_pop = int(self.pop.loc[self.pop["fips"] == int(county)]["population"])
cum_deaths = 0
for idx, row in c_data.iterrows():
c["dates"].append(row["date"])
c["deaths"].append(row["deaths"] / c_pop)
cum_deaths += row["deaths"]
#print(c["dates"])
if cum_deaths <= death_lbound: continue
c_df = pd.DataFrame(data=c)
c_start = c_df.loc[c_df["deaths"] > 5e-6].first_valid_index()
if (c_start == None): continue
for j in range(c_start, len(c["dates"]) - test_per, 1):
if target_idx - j < pred_per + test_per: continue
distance, path = fastdtw(np.array(target["deaths"][target_idx:target_idx+overlay_window]), np.array(c["deaths"][j:j+overlay_window]), dist=dist_func)
if distance < best_dist:
best_idx = j
best_fips = county
best_dist = distance
best_path = path
except TypeError as e:
continue
return best_idx, int(best_fips), best_dist
def predict_county(self, target_county, dist_func, fit_func, overlay_window, test_per, pred_per, death_lbound, log=True):
if log: print("Predicting for county " + str(target_county) + "... ", end='\r', flush=True)
target_data = self.data.loc[self.data["fips"] == target_county]
try:
target_pop = int(self.pop.loc[self.pop["fips"] == target_county]["population"])
except TypeError as e:
#print("No population found for", str(target_county))
return fit_func(pd.DataFrame(data={"dates":[], "deaths":[]}), target_county, 0, test_per, pred_per, 0, datetime.date.fromisoformat('2020-04-26'), self)
# Construct target dataset: deaths as a proportion of population
target = {"dates": [], "deaths": []}
deathcount = 0
for idx, row in target_data.iterrows():
target["dates"].append(row["date"])
target["deaths"].append(row["deaths"] / target_pop)
deathcount += row["deaths"]
#print(str(len(target["dates"])) + " ")
target_idx = len(target["dates"]) - overlay_window - test_per
if deathcount > death_lbound:
best_idx, best_fips, best_dist = self.get_best_fit(target, target_county, dist_func, overlay_window, test_per, pred_per, death_lbound, log=log)
else:
best_idx, best_fips, best_dist = -1, -1, 100
if best_fips >= 0 and best_dist < 1e-5:
# Align data by calculated indices
diff = target_idx - best_idx
overlay = pd.DataFrame(data = target)
overlay["best_fit"] = 0
best_data = self.data.loc[self.data["fips"] == best_fips]
best = {"dates": [], "deaths": []}
best_pop = int(self.pop.loc[self.pop["fips"] == best_fips]["population"])
for i, row in best_data.iterrows():
best["dates"].append(row["date"])
best["deaths"].append(row["deaths"] / best_pop)
for i, row in overlay.iterrows():
if i - diff >= 0 and i - diff < len(best["dates"]):
overlay.loc[i, "best_fit"] = best["deaths"][i - diff]
if not self.last_date:
self.last_date = datetime.date.fromisoformat(overlay.iloc[-1]["dates"])
date = self.last_date
for i in range(diff):
date += datetime.timedelta(days=1)
overlay.loc[len(overlay)] = [date.isoformat(), 0, best["deaths"][len(best["deaths"]) - diff + i]]
self.num_found_fit += 1
overlay["best_fit"] *= target_pop
else:
diff = 0
overlay = pd.DataFrame(data = target)
overlay["best_fit"] = -1
if not self.last_date:
try: self.last_date = datetime.date.fromisoformat(overlay.iloc[-1]["dates"])
except Exception as e: self.last_date = datetime.date.fromisoformat('2020-04-26')
overlay["deaths"] *= target_pop
return fit_func(overlay, target_county, target_pop, test_per, pred_per, diff, self.last_date, self)
def generate_submission(self, filename, fit_func, overlay_window, test_per, pred_per, death_lbound, dist_func=euclidean, log=True):
ids, i10, i20, i30, i40, i50, i60, i70, i80, i90 = [], [], [], [], [], [], [], [], [], []
for idx, row in self.fips.iterrows():
county = int(row["fips"])
this_id, this_10, this_20, this_30, this_40, this_50, this_60, this_70, this_80, this_90 = (
self.predict_county(county, dist_func, fit_func, overlay_window, test_per, pred_per, death_lbound, log)
)
ids += this_id
i10 += this_10
i20 += this_20
i30 += this_30
i40 += this_40
i50 += this_50
i60 += this_60
i70 += this_70
i80 += this_80
i90 += this_90
predictions = pd.DataFrame(data={"id":ids, "10":i10, "20":i20, "30":i30, "40":i40, "50":i50, "60":i60, "70":i70, "80":i80, "90":i90})
predictions.to_csv(filename, index=False)
print("Used dynamic time warping to help predictions in", self.num_found_fit, "counties! ")
return predictions
def fit_prophet_from_overlay(overlay, county, pop, test_per, pred_per, diff, last_date, dtw_instance):
overlay.rename(columns={'dates': 'ds', 'deaths': 'y'}, inplace=True)
c_row = dtw_instance.data.loc[dtw_instance.data["fips"] == county]
cum_deaths = 0
for i, item in c_row.iterrows(): cum_deaths += int(item["deaths"])
ids, i10, i20, i30, i40, i50, i60, i70, i80, i90 = [], [], [], [], [], [], [], [], [], []
z_80 = 1.28
z_60 = 0.84
z_40 = 0.525
z_20 = 0.25
if cum_deaths > 15:
try:
overlay.fillna(0, inplace=True)
m = Prophet(seasonality_mode="multiplicative", interval_width = 0.68)
if int(overlay.loc[overlay["ds"] == last_date.isoformat()]["best_fit"]) >= 0:
m.add_regressor("best_fit", prior_scale=1.0)
#print("Added regressor for", str(county))
m.fit(overlay.iloc[:len(overlay) - diff - test_per])
future = pd.DataFrame()
future["ds"] = 0
future["best_fit"] = 0
d = last_date - datetime.timedelta(days=test_per)
for i in range(test_per + pred_per):
d += datetime.timedelta(days=1)
try:
future.loc[len(future)] = [d.isoformat(), float(overlay.loc[overlay["ds"] == d.isoformat()]["best_fit"])]
except ValueError as e:
future.loc[len(future)] = [d.isoformat(), 0]
forecast = m.predict(future)
#forecast["yhat"] *= pop
#forecast["yhat_upper"] *= pop
#forecast["yhat_lower"] *= pop
for i, pred in forecast.iterrows():
date = pred["ds"]
upper = pred["yhat_upper"]
lower = pred["yhat_lower"]
mid = pred["yhat"]
sd = upper - mid
this_id = str(date)[:10] + "-" + str(county)
ids.append(this_id)
i10.append(mid - sd * z_80)
i20.append(mid - sd * z_60)
i30.append(mid - sd * z_40)
i40.append(mid - sd * z_20)
i50.append(mid)
i60.append(mid + sd * z_20)
i70.append(mid + sd * z_40)
i80.append(mid + sd * z_60)
i90.append(mid + sd * z_80)
# This county doesn't have enough historical data to train a model
except ValueError as e:
print("Not enough data for county", county, " ")
d = last_date - datetime.timedelta(days=test_per)
for i in range(test_per + pred_per):
d += datetime.timedelta(days=1)
this_id = d.isoformat() + "-" + str(county)
ids.append(this_id)
i10.append(0)
i20.append(0)
i30.append(0)
i40.append(0)
i50.append(0)
i60.append(0)
i70.append(0)
i80.append(0)
i90.append(0)
else:
d = last_date - datetime.timedelta(days=test_per)
for i in range(test_per + pred_per):
d += datetime.timedelta(days=1)
this_id = d.isoformat() + "-" + str(county)
ids.append(this_id)
i10.append(0)
i20.append(0)
i30.append(0)
i40.append(0)
i50.append(0)
i60.append(0)
i70.append(0)
i80.append(0)
i90.append(0)
return ids, i10, i20, i30, i40, i50, i60, i70, i80, i90
def cluster_counties(num_clusters):
county_data = pd.read_csv("data/us/demographics/county_land_areas.csv", encoding="cp1252", dtype={"County FIPS": str})
cluster_to_counties = {}
county_to_cluster = {}
km = KMeans(n_clusters=num_clusters)
km = km.fit(county_data[["2010 Density per square mile of land area - Population", "2010 Density per square mile of land area - Housing units"]])
county_data.loc[:, "cluster"] = km.labels_
for i, row in county_data.iterrows():
fips = row["County FIPS"]
cluster = row["cluster"]
if cluster in cluster_to_counties:
cluster_to_counties[cluster].append(fips)
else:
cluster_to_counties[cluster] = [fips]
county_to_cluster[fips] = cluster
return county_to_cluster, cluster_to_counties
```
### Using the Covid_DTW class and Prophet fit function to make a submission
```
deaths = pd.read_csv("data/us/covid/nyt_us_counties_daily.csv")
population = pd.read_csv("data/us/demographics/county_populations.csv")
fips_list = pd.read_csv("data/us/processing_data/fips_key.csv", encoding="cp1252")
population.rename(columns={"FIPS": "fips", "total_pop": "population"}, inplace=True)
fips_list.rename(columns={"FIPS": "fips"}, inplace=True)
county_to_cluster, cluster_to_counties = cluster_counties(num_clusters=12)
model = Covid_DTW(deaths, population, fips_list, county_to_cluster, cluster_to_counties)
print(str(datetime.datetime.now()) + "\n\n")
output = model.generate_submission(filename="test_submissions/dtw_prophet.csv", fit_func=fit_prophet_from_overlay, overlay_window=14, test_per=14, pred_per=0, death_lbound=15)
print("\n\n" + str(datetime.datetime.now()))
output["10"] = output["10"].apply(lambda x: x if x >= 1 else 0)
output["20"] = output["20"].apply(lambda x: x if x >= 1 else 0)
output["30"] = output["30"].apply(lambda x: x if x >= 1 else 0)
output["40"] = output["40"].apply(lambda x: x if x >= 1 else 0)
output["50"] = output["50"].apply(lambda x: x if x >= 1 else 0)
output["60"] = output["60"].apply(lambda x: x if x >= 1 else 0)
output["70"] = output["70"].apply(lambda x: x if x >= 1 else 0)
output["80"] = output["80"].apply(lambda x: x if x >= 1 else 0)
output["90"] = output["90"].apply(lambda x: x if x >= 1 else 0)
output.to_csv("test_submissions/dtw_prophet.csv", index=False)
```
### Evaluate Performance
```
# Author: Jake Will
#
# A script that allows us to locally evaluate our model's performance
import pandas as pd
import numpy as np
# Requires two NumPy arrays as input, the truth in y_true and predictions in y_pred.
# The quantile should be a number between 0 and 1. I copied this code from the
# piazza post describing how to compute the pinball loss.
def pinball_loss(y_true, y_pred, quantile = 0.5):
delta = y_true - y_pred
# Compute loss for underestimates.
loss_above = np.sum(delta[delta > 0]) * (quantile)
# Compute loss for overestimates.
loss_below = np.sum(-1 * delta[delta < 0]) * (1 - quantile)
return (loss_above + loss_below) / len(y_true)
# Input the name of the submission file to evaluate here
submission_file = "Epidemiological Models/delphi_test.csv"
# Input the desired dates into these lists - both lists
# need to be updated because the files have different
# date formats
nyt_dates = ['5/8/20', '5/9/20', '5/10/20', '5/11/20', '5/12/20', '5/13/20', '5/14/20', '5/15/20', '5/16/20', '5/17/20', '5/18/20', '5/19/20', '5/20/20', '5/21/20']
submission_dates = ["2020-05-08", "2020-05-09", "2020-05-10", "2020-05-11", "2020-05-12", "2020-05-13", "2020-05-14", "2020-05-15", "2020-05-16", "2020-05-17", "2020-05-18", "2020-05-19", "2020-05-20", "2020-05-21"]
# Because the files use different date formats, we'll use a map to
# be able to easily get the nyt dates from the submission dates.
# If things get significantly more complicated as the project
# continues, we might consider using a date-time library, but for
# now this should be sufficient.
submission_to_nyt_date = {}
for i, date in enumerate(submission_dates):
submission_to_nyt_date[date] = nyt_dates[i]
# Compute the submission predictions
submission = pd.read_csv(submission_file)
submission = submission[submission['id'].str.contains(('|'.join(submission_dates)))]
# Compute the actual results
deaths = pd.read_csv("data/us/covid/nyt_us_counties_daily.csv")
deaths = deaths[['date', 'fips', 'deaths']]
deaths = deaths[deaths['date'].str.contains(('|'.join(submission_dates)))]
# Generate a numpy array of the actual results in the same order
# as the submission. If a county has no reported deaths, we assume
# that is has 0.
truth = np.empty(len(submission['id'].values))
for i, submission_id in enumerate(submission['id'].values):
split_id = submission_id.split('-')
# Extract the FIPS and date from the id column of the submission
FIPS = int(split_id[-1])
date = '-'.join(split_id[:-1])
# Extract the relevant row of the nyt deaths data
df = deaths.loc[(deaths['fips'] == FIPS) & (deaths['date'] == date)]
# Set the truth numpy array accordingly
if df.empty:
truth[i] = 0
else:
truth[i] = df['deaths']
# Compute the pinball score using the given dates, submission, and
# truth values
score = 0.0
for column in [10, 20, 30, 40, 50, 60, 70, 80, 90]:
score = score + pinball_loss(truth, submission[str(column)].values, quantile = column / 100.0)
score = score/9.0
print(score)
```
All 0's loss: ~0.32 \
Prophet alone loss: 0.2645782822355685 \
Best prophet w/ DTW loss: 0.27197204421592697
8 clusters -> \
12 clusters -> 0.27197204421592697 (DTW used for 36 counties) \
16 clusters ->
No best distance minimum: 0.30582484582644276 \
With minimum: 0.2816424073220742
## Comparing Performance on Individual Counties With and Without DTW
```
deaths = pd.read_csv("data/us/covid/nyt_us_counties_daily.csv")
population = pd.read_csv("data/us/demographics/county_populations.csv")
fips_list = pd.read_csv("data/us/processing_data/fips_key.csv", encoding="cp1252")
population.rename(columns={"FIPS": "fips", "total_pop": "population"}, inplace=True)
fips_list.rename(columns={"FIPS": "fips"}, inplace=True)
county_to_cluster, cluster_to_counties = cluster_counties(num_clusters=12)
model = Covid_DTW(deaths, population, fips_list, county_to_cluster, cluster_to_counties)
def score_one_county(submission):
nyt_dates = ['4/22/20', '4/23/20', '4/24/20', '4/25/20', '4/26/20', '4/27/20', '4/28/20', '4/29/20', '4/30/20', '5/1/20', '5/2/20', '5/3/20', '5/4/20', '5/5/20']
submission_dates = ['2020-04-22', '2020-04-23', '2020-04-24', '2020-04-25', '2020-04-26', '2020-04-27', '2020-04-28', '2020-04-29', '2020-04-30', '2020-05-01', '2020-05-02', '2020-05-03', '2020-05-04', '2020-05-05']
submission_to_nyt_date = {}
for i, date in enumerate(submission_dates):
submission_to_nyt_date[date] = nyt_dates[i]
# Compute the submission predictions
submission = submission[submission['id'].str.contains(('|'.join(submission_dates)))]
# Compute the actual results
deaths = pd.read_csv("data/us/covid/nyt_us_counties_daily.csv")
deaths = deaths[['date', 'fips', 'deaths']]
deaths = deaths[deaths['date'].str.contains(('|'.join(submission_dates)))]
# Generate a numpy array of the actual results in the same order
# as the submission. If a county has no reported deaths, we assume
# that is has 0.
truth = np.empty(len(submission['id'].values))
for i, submission_id in enumerate(submission['id'].values):
split_id = submission_id.split('-')
# Extract the FIPS and date from the id column of the submission
FIPS = int(split_id[-1])
date = '-'.join(split_id[:-1])
# Extract the relevant row of the nyt deaths data
df = deaths.loc[(deaths['fips'] == FIPS) & (deaths['date'] == date)]
# Set the truth numpy array accordingly
if df.empty:
truth[i] = 0
else:
truth[i] = df['deaths']
# Compute the pinball score using the given dates, submission, and
# truth values
score = 0.0
for column in [10, 20, 30, 40, 50, 60, 70, 80, 90]:
score = score + pinball_loss(truth, submission[str(column)].values, quantile = column / 100.0)
score = score/9.0
return score
overlay_window = 14
test_per = 14
pred_per = 0
for idx, row in fips_list.iterrows():
target_county = row["fips"]
name = row["COUNTY"]
target_data = model.data.loc[model.data["fips"] == target_county]
try:
target_pop = int(model.pop.loc[model.pop["fips"] == target_county]["population"])
except TypeError as e:
continue
# Construct target dataset: deaths as a proportion of population
target = {"dates": [], "deaths": []}
deathcount = 0
for idx, row in target_data.iterrows():
target["dates"].append(row["date"])
target["deaths"].append(row["deaths"] / target_pop)
deathcount += row["deaths"]
target_idx = len(target["dates"]) - overlay_window - test_per
if deathcount > 15:
best_idx, best_fips, best_dist = model.get_best_fit(target, target_county, euclidean, overlay_window, test_per, pred_per, 15, log=False)
else:
best_idx, best_fips, best_dist = -1, -1, 100
if best_fips >= 0 and best_dist < 1e-5:
print("Fit found for", str(target_county), name, "with", deathcount, "cumulative deaths.")
# Align data by calculated indices
diff = target_idx - best_idx
overlay = pd.DataFrame(data = target)
overlay["best_fit"] = 0
best_data = model.data.loc[model.data["fips"] == best_fips]
best = {"dates": [], "deaths": []}
best_pop = int(model.pop.loc[model.pop["fips"] == best_fips]["population"])
for i, row in best_data.iterrows():
best["dates"].append(row["date"])
best["deaths"].append(row["deaths"] / best_pop)
for i, row in overlay.iterrows():
if i - diff >= 0 and i - diff < len(best["dates"]):
overlay.loc[i, "best_fit"] = best["deaths"][i - diff]
if not model.last_date:
model.last_date = datetime.date.fromisoformat(overlay.iloc[-1]["dates"])
date = model.last_date
for i in range(diff):
date += datetime.timedelta(days=1)
overlay.loc[len(overlay)] = [date.isoformat(), 0, best["deaths"][len(best["deaths"]) - diff + i]]
overlay["best_fit"] *= target_pop
overlay["deaths"] *= target_pop
ids, i10, i20, i30, i40, i50, i60, i70, i80, i90 = fit_prophet_from_overlay(overlay, target_county, target_pop, test_per, pred_per, diff, model.last_date, model)
dtw_pred = pd.DataFrame(data={"id":ids, "10":i10, "20":i20, "30":i30, "40":i40, "50":i50, "60":i60, "70":i70, "80":i80, "90":i90})
overlay["best_fit"] = -1
ids, i10, i20, i30, i40, i50, i60, i70, i80, i90 = fit_prophet_from_overlay(overlay, target_county, target_pop, test_per, pred_per, diff, model.last_date, model)
nodtw_pred = pd.DataFrame(data={"id":ids, "10":i10, "20":i20, "30":i30, "40":i40, "50":i50, "60":i60, "70":i70, "80":i80, "90":i90})
# Evaluate
dtw_score = score_one_county(dtw_pred)
print("Loss was", dtw_score, "with time warping.")
nodtw_score = score_one_county(nodtw_pred)
print("Loss was", nodtw_score, "without time warping.")
print()
```
| github_jupyter |
All data credits belong to the wonderful work done by **Rekhta foundation**.
Data has been parsed into Urdu, Hindi and English translieration thanks to their excellent data organization.
Consider supporting them for their great work in pushing the urdu language.

Credits to these authors for their wonderful original creations:
*'mirza-ghalib','allama-iqbal','faiz-ahmad-faiz','sahir-ludhianvi','meer-taqi-meer',
'dagh-dehlvi','kaifi-azmi','gulzar','bahadur-shah-zafar','parveen-shakir',
'jaan-nisar-akhtar','javed-akhtar','jigar-moradabadi','jaun-eliya',
'ahmad-faraz','meer-anees','mohsin-naqvi','firaq-gorakhpuri','fahmida-riaz','wali-mohammad-wali',
'waseem-barelvi','akbar-allahabadi','altaf-hussain-hali','ameer-khusrau','naji-shakir','naseer-turabi',
'nazm-tabatabai','nida-fazli','noon-meem-rashid', 'habib-jalib'*
```
from bs4 import BeautifulSoup
from bs4.element import Comment
import urllib.request
# credits to this stackoverflow answer https://stackoverflow.com/questions/1936466/beautifulsoup-grab-visible-webpage-text
allowed_sections=['style', 'script', 'head', 'title', 'meta', '[document]']
def tag_visible(element):
if element.parent.name in allowed_sections:
return False
if isinstance(element, Comment):
return False
return True
def text_from_html(body):
soup = BeautifulSoup(body, 'html.parser')
texts = soup.findAll(text=True)
visible_texts = filter(tag_visible, texts)
return u" ".join(t.strip() for t in visible_texts)
#captures the different <p> tags as newlines
def text_with_newlines(elem):
text = ''
for e in elem.descendants:
if isinstance(e, str):
text += e
elif e.name == 'br' or e.name == 'p':
text += '\n'
return text
def parse_ghazal(url):
html = urllib.request.urlopen(url).read()
soup= BeautifulSoup(html, 'html.parser')
mydivs = soup.find("div", {"class": "pMC"})
#this section removes some of the English translations present on the webpage
# mixing language data would add noise, and make it difficult for the model to learn
#BUT in future these urdu to english translations could be a valuable resource to prepare machine translation data
for div in mydivs.find_all("div", {'class':'t'}):
div.decompose()
mydivs= text_with_newlines(mydivs)
return mydivs
url_english='https://www.rekhta.org/ghazals/sitaaron-se-aage-jahaan-aur-bhii-hain-allama-iqbal-ghazals-1'
url_urdu='https://www.rekhta.org/ghazals/sitaaron-se-aage-jahaan-aur-bhii-hain-allama-iqbal-ghazals-1?lang=ur'
url_hindi='https://www.rekhta.org/ghazals/sitaaron-se-aage-jahaan-aur-bhii-hain-allama-iqbal-ghazals-1?lang=hi'
ghazal = parse_ghazal(url_english)
print(ghazal)
'''
okay I have a problem the way this urdu font is rendered. It is absolutely terrible to read.
There have to be better font options.
'''
```
## Parsed text samples
### English transliteration
sitāroñ se aage jahāñ aur bhī haiñ abhī ishq ke imtihāñ aur bhī haiñ
tū shāhīñ hai parvāz hai kaam terā tire sāmne āsmāñ aur bhī haiñ
isī roz o shab meñ ulajh kar na rah jā ki tere zamān o makāñ aur bhī haiñ
### Urdu
ستاروں سے آگے جہاں اور بھی ہیں
تو شاہیں ہے پرواز ہے کام تیرا ترے سامنے آسماں اور بھی ہیں
اسی روز و شب میں الجھ کر نہ رہ جا کہ تیرے زمان و مکاں اور بھی ہیں
### Hindi
सितारों से आगे जहाँ और भी हैं अभी इश्क़ के इम्तिहाँ और भी हैं
तू शाहीं है परवाज़ है काम तेरा तिरे सामने आसमाँ और भी हैं
इसी रोज़ ओ शब में उलझ कर न रह जा कि तेरे ज़मान ओ मकाँ और भी हैं
### Observing the structure of urls to make sure the same script would work across different poets on the webpage
```
#The folder structure has urls as https://www.rekhta.org/ghazals/
#https://www.rekhta.org/ghazals/tire-ishq-kii-intihaa-chaahtaa-huun-allama-iqbal-ghazals
#Iqbal
url='https://www.rekhta.org/ghazals/tire-ishq-kii-intihaa-chaahtaa-huun-allama-iqbal-ghazals'
url= 'https://www.rekhta.org/ghazals/kabhii-ai-haqiiqat-e-muntazar-nazar-aa-libaas-e-majaaz-men-allama-iqbal-ghazals'
##Ghalib's ghazals
## there are currently 234 ghazals by this poet on the page
## which makes it a rich resource for training a text model
url= 'https://www.rekhta.org/ghazals/hazaaron-khvaahishen-aisii-ki-har-khvaahish-pe-dam-nikle-mirza-ghalib-ghazals'
url ='https://www.rekhta.org/ghazals/hazaaron-khvaahishen-aisii-ki-har-khvaahish-pe-dam-nikle-mirza-ghalib-ghazals?lang=ur'
url='https://www.rekhta.org/ghazals/hazaaron-khvaahishen-aisii-ki-har-khvaahish-pe-dam-nikle-mirza-ghalib-ghazals?lang=hi'
url='https://www.rekhta.org/ghazals/har-ek-baat-pe-kahte-ho-tum-ki-tuu-kyaa-hai-mirza-ghalib-ghazals'
url='https://www.rekhta.org/ghazals/ishq-mujh-ko-nahiin-vahshat-hii-sahii-mirza-ghalib-ghazals'
url='https://www.rekhta.org/ghazals/ishq-mujh-ko-nahiin-vahshat-hii-sahii-mirza-ghalib-ghazals?lang=ur'
url='https://www.rekhta.org/ghazals/ishq-mujh-ko-nahiin-vahshat-hii-sahii-mirza-ghalib-ghazals?lang=hi'
url='https://www.rekhta.org/ghazals/koii-din-gar-zindagaanii-aur-hai-mirza-ghalib-ghazals'
url='https://www.rekhta.org/ghazals/koii-din-gar-zindagaanii-aur-hai-mirza-ghalib-ghazals?lang=ur'
url='https://www.rekhta.org/ghazals/hai-bazm-e-butaan-men-sukhan-aazurda-labon-se-mirza-ghalib-ghazals'
url='https://www.rekhta.org/ghazals/hai-bazm-e-butaan-men-sukhan-aazurda-labon-se-mirza-ghalib-ghazals?lang=ur'
url='https://www.rekhta.org/ghazals/ghar-jab-banaa-liyaa-tire-dar-par-kahe-bagair-mirza-ghalib-ghazals'
url='https://www.rekhta.org/ghazals/ghar-jab-banaa-liyaa-tire-dar-par-kahe-bagair-mirza-ghalib-ghazals?lang=hi'
url='https://www.rekhta.org/ghazals/ghar-jab-banaa-liyaa-tire-dar-par-kahe-bagair-mirza-ghalib-ghazals?lang=ur'
## Sahir Ludhianvi
url='https://www.rekhta.org/ghazals/kabhii-khud-pe-kabhii-haalaat-pe-ronaa-aayaa-sahir-ludhianvi-ghazals'
url='https://www.rekhta.org/ghazals/kabhii-khud-pe-kabhii-haalaat-pe-ronaa-aayaa-sahir-ludhianvi-ghazals?lang=ur'
url= 'https://www.rekhta.org/ghazals/kabhii-khud-pe-kabhii-haalaat-pe-ronaa-aayaa-sahir-ludhianvi-ghazals?lang=hi'
## Faiz
en_url='https://www.rekhta.org/ghazals/gulon-men-rang-bhare-baad-e-nau-bahaar-chale-faiz-ahmad-faiz-ghazals'
hindi_url='https://www.rekhta.org/ghazals/gulon-men-rang-bhare-baad-e-nau-bahaar-chale-faiz-ahmad-faiz-ghazals?lang=hi'
urdu_url='https://www.rekhta.org/ghazals/gulon-men-rang-bhare-baad-e-nau-bahaar-chale-faiz-ahmad-faiz-ghazals?lang=ur'
url='https://www.rekhta.org/ghazals/hazaaron-khvaahishen-aisii-ki-har-khvaahish-pe-dam-nikle-mirza-ghalib-ghazals'#?lang=ur'
ghazal = parse_ghazal(url)
print(ghazal)
#testing the homepage for parsing all poet names now
#for ghalib
def parse_webpage_at_given_scroll(html):
ctr=0
soup= BeautifulSoup(html, 'html.parser')
mydivs = soup.find("div", {"class": "contentListBody"})
titles=[]
for a in mydivs.find_all('a', href=True):
t=a['href']
if t not in titles:
if ctr%5==0:
print("Found the URL:", t)
titles.append(t)
ctr+=1
print('=============================')
print('number of titles',len(titles))
print('=============================')
return titles
#language argument can be ur or hi for urdu or hindi
def read_and_write_web(author,language='ur'):
lang=language
#author='mirza-ghalib'
author_lan=author+'/'+lang
if not os.path.exists(author_lan):
os.makedirs(author_lan)
for url in titles:
name_poem=url.split('https://www.rekhta.org/ghazals/')[1]
path_poem= author_lan+'/'+name_poem
if os.path.exists(path_poem):
pass
else:
f= open(path_poem,"w+")
if lang=='en':
url_for_lang= url
else:
url_for_lang= url+'?lang='+lang
ghazal = parse_ghazal(url_for_lang)
f.write(ghazal)
f.close()
##Parsing based on home page of authors
url_base='https://www.rekhta.org/poets/'
## TODO : Later
## or just iterate through the list of all poets on the index, instead of hand curated list
authors=['mirza-ghalib','allama-iqbal','faiz-ahmad-faiz','sahir-ludhianvi','meer-taqi-meer',
'dagh-dehlvi','kaifi-azmi','gulzar','bahadur-shah-zafar','parveen-shakir',
'jaan-nisar-akhtar','javed-akhtar','jigar-moradabadi','jaun-eliya',
'ahmad-faraz','meer-anees','mohsin-naqvi','firaq-gorakhpuri','fahmida-riaz','wali-mohammad-wali',
'waseem-barelvi','akbar-allahabadi','altaf-hussain-hali','ameer-khusrau','naji-shakir','naseer-turabi'
,'nazm-tabatabai','nida-fazli','noon-meem-rashid','habib-jalib']
for author in authors:
url_home_page= url_base +author+ '/ghazals'
html = urllib.request.urlopen(url_home_page).read()
titles= parse_webpage_at_given_scroll(html)
read_and_write_web(author,'en')
read_and_write_web(author,'ur')
read_and_write_web(author,'hi')
```
ہم کو مٹا سکے یہ زمانے میں دم نہیں
ہم سے زمانہ خود ہے زمانے سے ہم نہیں
हम को मिटा सके ये ज़माने में दम नहीं
हम से ज़माना ख़ुद है ज़माने से हम नहीं
Jigar Moradabadi
| github_jupyter |
## Medidas de impureza
```
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
```
$$ Gini = 1 - \sum_i p_i ^ 2 $$
```
def gini(p):
return (p)*(1 - (p)) + (1 - p)*(1 - (1-p))
```
$$ Entropia = - \sum_i p_i log_2 (p_i) $$
```
def entropy(p):
return - p*np.log2(p) - (1 - p)*np.log2((1 - p))
```
$$ ErrorClasif = 1 - max (p_i) $$
```
def classification_error(p):
return 1 - np.max([p, 1 - p])
x = np.arange(0.0, 1.0, 0.01)
ent = [entropy(p) if p != 0 else None for p in x]
scaled_ent = [e*0.5 if e else None for e in ent]
c_err = [classification_error(i) for i in x]
fig = plt.figure()
ax = plt.subplot(111)
for j, lab, ls, c, in zip(
[ent, scaled_ent, gini(x), c_err],
['Entropy', 'Entropy (scaled)', 'Gini Impurity', 'Misclassification Error'],
['-', '-', '--', '-.'],
['lightgray', 'red', 'green', 'blue']):
line = ax.plot(x, j, label=lab, linestyle=ls, lw=1, color=c)
ax.legend(loc='upper left', bbox_to_anchor=(0.01, 0.85),
ncol=1, fancybox=True, shadow=False)
ax.axhline(y=0.5, linewidth=1, color='k', linestyle='--')
ax.axhline(y=1.0, linewidth=1, color='k', linestyle='--')
plt.ylim([0, 1.1])
plt.xlabel('p(j=1)')
plt.ylabel('Impurity Index')
plt.show()
```
## Árbol de decisión
Datos: [Marketing Bancario](https://archive.ics.uci.edu/ml/datasets/bank+marketing)
```
bank = pd.read_csv('datos/bank-full.csv', sep=';')
bank.head()
bank = bank.replace('yes', 1).replace('no', 0)
months = pd.DataFrame({
'month': bank.month.sort_values().unique(),
'month_no': [4, 8, 12, 2, 1, 7, 6, 3, 5, 11, 10, 9]
})
bank = bank.merge(months).drop('month', axis=1)
bank = bank.merge(pd.DataFrame({
'education': bank.education.unique(),
'edu': [1, 2, 3, np.nan]
})).drop('education', axis=1)
bank = bank.merge(pd.DataFrame({
'poutcome': bank.poutcome.sort_values().unique(),
'prev_out': [-1, np.nan, 1, 0]
})).drop('poutcome', axis=1)
bank = bank.drop('job', axis=1)
bank_dummies = pd.get_dummies(bank)
bank_dummies.sample(10)
bank_dummies = bank_dummies.dropna()
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier, export_graphviz
import graphviz
from sklearn.metrics import confusion_matrix
arbol = DecisionTreeClassifier(max_depth=2)
X_train, X_test, y_train, y_test = train_test_split(
bank_dummies.drop('y', axis=1), # X
bank_dummies.y, # y
test_size=0.2, # porcentaje que será prueba
random_state=42) # para fijar el aleatorio (reproducibilidad)
arbol_ent = arbol.fit(X_train, y_train)
confusion_matrix(y_test, arbol_ent.predict(X_test))
graf = export_graphviz(arbol_ent, out_file=None,
feature_names=X_train.columns,
filled=True, rounded=True,
special_characters=True)
graph = graphviz.Source(graf)
graph
print("Score entrenamiento:", arbol_ent.score(X_train, y_train))
print("Score prueba:", arbol_ent.score(X_test, y_test))
score_ent = []
score_pru = []
for i in range(3, 30):
arbol = DecisionTreeClassifier(max_depth=i)
arbol_ent = arbol.fit(X_train, y_train)
print("Score entrenamiento:", arbol_ent.score(X_train, y_train))
print("Score prueba:", arbol_ent.score(X_test, y_test))
score_ent.append(arbol_ent.score(X_train, y_train))
score_pru.append(arbol_ent.score(X_test, y_test))
pd.DataFrame({
'score_ent': score_ent,
'score_pru': score_pru
}, index=range(3, 30)).plot()
plt.show()
```
## Bagging
```
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import BaggingClassifier
score_ent = []
score_pru = []
for i in range(3, 30):
tree = DecisionTreeClassifier(max_depth=i)
bag = BaggingClassifier(tree, n_estimators=30, n_jobs=-1)
bag_ent = bag.fit(X_train, y_train)
print("Score entrenamiento:", bag_ent.score(X_train, y_train))
print("Score prueba:", bag_ent.score(X_test, y_test),'\n')
score_ent.append(bag_ent.score(X_train, y_train))
score_pru.append(bag_ent.score(X_test, y_test))
pd.DataFrame({
'score_ent': score_ent,
'score_pru': score_pru
}, index=range(3, 30)).plot()
plt.show()
tree = DecisionTreeClassifier()
bag = BaggingClassifier(tree, n_estimators=200, n_jobs=-1)
bag.fit(X_train, y_train)
bag_ent = bag.fit(X_train, y_train)
print("Score entrenamiento:", bag_ent.score(X_train, y_train))
print("Score prueba:", bag_ent.score(X_test, y_test))
```
| github_jupyter |
# Create a Pipeline
You can perform the various steps required to ingest data, train a model, and register the model individually by using the Azure ML SDK to run script-based experiments. However, in an enterprise environment it is common to encapsulate the sequence of discrete steps required to build a machine learning solution into a *pipeline* that can be run on one or more compute targets; either on-demand by a user, from an automated build process, or on a schedule.
In this notebook, you'll bring together all of these elements to create a simple pipeline that pre-processes data and then trains and registers a model.
## Connect to your workspace
To get started, connect to your workspace.
> **Note**: If you haven't already established an authenticated session with your Azure subscription, you'll be prompted to authenticate by clicking a link, entering an authentication code, and signing into Azure.
```
import azureml.core
from azureml.core import Workspace
# Load the workspace from the saved config file
ws = Workspace.from_config()
print('Ready to use Azure ML {} to work with {}'.format(azureml.core.VERSION, ws.name))
```
## Prepare data
In your pipeline, you'll use a dataset containing details of diabetes patients. Run the cell below to create this dataset (if you created it previously, the code will find the existing version)
```
from azureml.core import Dataset
default_ds = ws.get_default_datastore()
if 'diabetes dataset' not in ws.datasets:
default_ds.upload_files(files=['./data/diabetes.csv', './data/diabetes2.csv'], # Upload the diabetes csv files in /data
target_path='diabetes-data/', # Put it in a folder path in the datastore
overwrite=True, # Replace existing files of the same name
show_progress=True)
#Create a tabular dataset from the path on the datastore (this may take a short while)
tab_data_set = Dataset.Tabular.from_delimited_files(path=(default_ds, 'diabetes-data/*.csv'))
# Register the tabular dataset
try:
tab_data_set = tab_data_set.register(workspace=ws,
name='diabetes dataset',
description='diabetes data',
tags = {'format':'CSV'},
create_new_version=True)
print('Dataset registered.')
except Exception as ex:
print(ex)
else:
print('Dataset already registered.')
```
## Create scripts for pipeline steps
Pipelines consist of one or more *steps*, which can be Python scripts, or specialized steps like a data transfer step that copies data from one location to another. Each step can run in its own compute context. In this exercise, you'll build a simple pipeline that contains two Python script steps: one to pre-process some training data, and another to use the pre-processed data to train and register a model.
First, let's create a folder for the script files we'll use in the pipeline steps.
```
import os
# Create a folder for the pipeline step files
experiment_folder = 'diabetes_pipeline'
os.makedirs(experiment_folder, exist_ok=True)
print(experiment_folder)
```
Now let's create the first script, which will read data from the diabetes dataset and apply some simple pre-processing to remove any rows with missing data and normalize the numeric features so they're on a similar scale.
The script includes a argument named **--prepped-data**, which references the folder where the resulting data should be saved.
```
%%writefile $experiment_folder/prep_diabetes.py
# Import libraries
import os
import argparse
import pandas as pd
from azureml.core import Run
from sklearn.preprocessing import MinMaxScaler
# Get parameters
parser = argparse.ArgumentParser()
parser.add_argument("--input-data", type=str, dest='raw_dataset_id', help='raw dataset')
parser.add_argument('--prepped-data', type=str, dest='prepped_data', default='prepped_data', help='Folder for results')
args = parser.parse_args()
save_folder = args.prepped_data
# Get the experiment run context
run = Run.get_context()
# load the data (passed as an input dataset)
print("Loading Data...")
diabetes = run.input_datasets['raw_data'].to_pandas_dataframe()
# Log raw row count
row_count = (len(diabetes))
run.log('raw_rows', row_count)
# remove nulls
diabetes = diabetes.dropna()
# Normalize the numeric columns
scaler = MinMaxScaler()
num_cols = ['Pregnancies','PlasmaGlucose','DiastolicBloodPressure','TricepsThickness','SerumInsulin','BMI','DiabetesPedigree']
diabetes[num_cols] = scaler.fit_transform(diabetes[num_cols])
# Log processed rows
row_count = (len(diabetes))
run.log('processed_rows', row_count)
# Save the prepped data
print("Saving Data...")
os.makedirs(save_folder, exist_ok=True)
save_path = os.path.join(save_folder,'data.csv')
diabetes.to_csv(save_path, index=False, header=True)
# End the run
run.complete()
```
Now you can create the script for the second step, which will train a model. The script includes a argument named **--training-data**, which references the location where the prepared data was saved by the previous step.
```
%%writefile $experiment_folder/train_diabetes.py
# Import libraries
from azureml.core import Run, Model
import argparse
import pandas as pd
import numpy as np
import joblib
import os
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve
import matplotlib.pyplot as plt
# Get parameters
parser = argparse.ArgumentParser()
parser.add_argument("--training-data", type=str, dest='training_data', help='training data')
args = parser.parse_args()
training_data = args.training_data
# Get the experiment run context
run = Run.get_context()
# load the prepared data file in the training folder
print("Loading Data...")
file_path = os.path.join(training_data,'data.csv')
diabetes = pd.read_csv(file_path)
# Separate features and labels
X, y = diabetes[['Pregnancies','PlasmaGlucose','DiastolicBloodPressure','TricepsThickness','SerumInsulin','BMI','DiabetesPedigree','Age']].values, diabetes['Diabetic'].values
# Split data into training set and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=0)
# Train adecision tree model
print('Training a decision tree model...')
model = DecisionTreeClassifier().fit(X_train, y_train)
# calculate accuracy
y_hat = model.predict(X_test)
acc = np.average(y_hat == y_test)
print('Accuracy:', acc)
run.log('Accuracy', np.float(acc))
# calculate AUC
y_scores = model.predict_proba(X_test)
auc = roc_auc_score(y_test,y_scores[:,1])
print('AUC: ' + str(auc))
run.log('AUC', np.float(auc))
# plot ROC curve
fpr, tpr, thresholds = roc_curve(y_test, y_scores[:,1])
fig = plt.figure(figsize=(6, 4))
# Plot the diagonal 50% line
plt.plot([0, 1], [0, 1], 'k--')
# Plot the FPR and TPR achieved by our model
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
run.log_image(name = "ROC", plot = fig)
plt.show()
# Save the trained model in the outputs folder
print("Saving model...")
os.makedirs('outputs', exist_ok=True)
model_file = os.path.join('outputs', 'diabetes_model.pkl')
joblib.dump(value=model, filename=model_file)
# Register the model
print('Registering model...')
Model.register(workspace=run.experiment.workspace,
model_path = model_file,
model_name = 'diabetes_model',
tags={'Training context':'Pipeline'},
properties={'AUC': np.float(auc), 'Accuracy': np.float(acc)})
run.complete()
```
## Prepare a compute environment for the pipeline steps
In this exercise, you'll use the same compute for both steps, but it's important to realize that each step is run independently; so you could specify different compute contexts for each step if appropriate.
First, get the compute target you created in a previous lab (if it doesn't exist, it will be created).
> **Important**: Change *your-compute-cluster* to the name of your compute cluster in the code below before running it! Cluster names must be globally unique names between 2 to 16 characters in length. Valid characters are letters, digits, and the - character.
```
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
cluster_name = "your-compute-cluster"
try:
# Check for existing compute target
pipeline_cluster = ComputeTarget(workspace=ws, name=cluster_name)
print('Found existing cluster, use it.')
except ComputeTargetException:
# If it doesn't already exist, create it
try:
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_DS11_V2', max_nodes=2)
pipeline_cluster = ComputeTarget.create(ws, cluster_name, compute_config)
pipeline_cluster.wait_for_completion(show_output=True)
except Exception as ex:
print(ex)
```
> **Note**: Compute instances and clusters are based on standard Azure virtual machine images. For this exercise, the *Standard_DS11_v2* image is recommended to achieve the optimal balance of cost and performance. If your subscription has a quota that does not include this image, choose an alternative image; but bear in mind that a larger image may incur higher cost and a smaller image may not be sufficient to complete the tasks. Alternatively, ask your Azure administrator to extend your quota.
The compute will require a Python environment with the necessary package dependencies installed.
```
%%writefile $experiment_folder/experiment_env.yml
name: experiment_env
dependencies:
- python=3.6.2
- scikit-learn
- ipykernel
- matplotlib
- pandas
- pip
- pip:
- azureml-defaults
- pyarrow
```
Now that you have a Conda configuration file, you can create an environment and use it in the run configuration for the pipeline.
```
from azureml.core import Environment
from azureml.core.runconfig import RunConfiguration
# Create a Python environment for the experiment (from a .yml file)
experiment_env = Environment.from_conda_specification("experiment_env", experiment_folder + "/experiment_env.yml")
# Register the environment
experiment_env.register(workspace=ws)
registered_env = Environment.get(ws, 'experiment_env')
# Create a new runconfig object for the pipeline
pipeline_run_config = RunConfiguration()
# Use the compute you created above.
pipeline_run_config.target = pipeline_cluster
# Assign the environment to the run configuration
pipeline_run_config.environment = registered_env
print ("Run configuration created.")
```
## Create and run a pipeline
Now you're ready to create and run a pipeline.
First you need to define the steps for the pipeline, and any data references that need to be passed between them. In this case, the first step must write the prepared data to a folder that can be read from by the second step. Since the steps will be run on remote compute (and in fact, could each be run on different compute), the folder path must be passed as a data reference to a location in a datastore within the workspace. The **OutputFileDatasetConfig** object is a special kind of data reference that is used for interim storage locations that can be passed between pipeline steps, so you'll create one and use at as the output for the first step and the input for the second step. Note that you need to pass it as a script argument so your code can access the datastore location referenced by the data reference.
```
from azureml.data import OutputFileDatasetConfig
from azureml.pipeline.steps import PythonScriptStep
# Get the training dataset
diabetes_ds = ws.datasets.get("diabetes dataset")
# Create an OutputFileDatasetConfig (temporary Data Reference) for data passed from step 1 to step 2
prepped_data = OutputFileDatasetConfig("prepped_data")
# Step 1, Run the data prep script
prep_step = PythonScriptStep(name = "Prepare Data",
source_directory = experiment_folder,
script_name = "prep_diabetes.py",
arguments = ['--input-data', diabetes_ds.as_named_input('raw_data'),
'--prepped-data', prepped_data],
compute_target = pipeline_cluster,
runconfig = pipeline_run_config,
allow_reuse = True)
# Step 2, run the training script
train_step = PythonScriptStep(name = "Train and Register Model",
source_directory = experiment_folder,
script_name = "train_diabetes.py",
arguments = ['--training-data', prepped_data.as_input()],
compute_target = pipeline_cluster,
runconfig = pipeline_run_config,
allow_reuse = True)
print("Pipeline steps defined")
```
OK, you're ready build the pipeline from the steps you've defined and run it as an experiment.
```
from azureml.core import Experiment
from azureml.pipeline.core import Pipeline
from azureml.widgets import RunDetails
# Construct the pipeline
pipeline_steps = [prep_step, train_step]
pipeline = Pipeline(workspace=ws, steps=pipeline_steps)
print("Pipeline is built.")
# Create an experiment and run the pipeline
experiment = Experiment(workspace=ws, name = 'mslearn-diabetes-pipeline')
pipeline_run = experiment.submit(pipeline, regenerate_outputs=True)
print("Pipeline submitted for execution.")
RunDetails(pipeline_run).show()
pipeline_run.wait_for_completion(show_output=True)
```
A graphical representation of the pipeline experiment will be displayed in the widget as it runs. Keep an eye on the kernel indicator at the top right of the page, when it turns from **⚫** to **◯**, the code has finished running. You can also monitor pipeline runs in the **Experiments** page in [Azure Machine Learning studio](https://ml.azure.com).
When the pipeline has finished, you can examine the metrics recorded by it's child runs.
```
for run in pipeline_run.get_children():
print(run.name, ':')
metrics = run.get_metrics()
for metric_name in metrics:
print('\t',metric_name, ":", metrics[metric_name])
```
Assuming the pipeline was successful, a new model should be registered with a *Training context* tag indicating it was trained in a pipeline. Run the following code to verify this.
```
from azureml.core import Model
for model in Model.list(ws):
print(model.name, 'version:', model.version)
for tag_name in model.tags:
tag = model.tags[tag_name]
print ('\t',tag_name, ':', tag)
for prop_name in model.properties:
prop = model.properties[prop_name]
print ('\t',prop_name, ':', prop)
print('\n')
```
## Publish the pipeline
After you've created and tested a pipeline, you can publish it as a REST service.
```
# Publish the pipeline from the run
published_pipeline = pipeline_run.publish_pipeline(
name="diabetes-training-pipeline", description="Trains diabetes model", version="1.0")
published_pipeline
```
Note that the published pipeline has an endpoint, which you can see in the **Endpoints** page (on the **Pipeline Endpoints** tab) in [Azure Machine Learning studio](https://ml.azure.com). You can also find its URI as a property of the published pipeline object:
```
rest_endpoint = published_pipeline.endpoint
print(rest_endpoint)
```
## Call the pipeline endpoint
To use the endpoint, client applications need to make a REST call over HTTP. This request must be authenticated, so an authorization header is required. A real application would require a service principal with which to be authenticated, but to test this out, we'll use the authorization header from your current connection to your Azure workspace, which you can get using the following code:
```
from azureml.core.authentication import InteractiveLoginAuthentication
interactive_auth = InteractiveLoginAuthentication()
auth_header = interactive_auth.get_authentication_header()
print("Authentication header ready.")
```
Now we're ready to call the REST interface. The pipeline runs asynchronously, so we'll get an identifier back, which we can use to track the pipeline experiment as it runs:
```
import requests
experiment_name = 'mslearn-diabetes-pipeline'
rest_endpoint = published_pipeline.endpoint
response = requests.post(rest_endpoint,
headers=auth_header,
json={"ExperimentName": experiment_name})
run_id = response.json()["Id"]
run_id
```
Since you have the run ID, you can use it to wait for the run to complete.
> **Note**: The pipeline should complete quickly, because each step was configured to allow output reuse. This was done primarily for convenience and to save time in this course. In reality, you'd likely want the first step to run every time in case the data has changed, and trigger the subsequent steps only if the output from step one changes.
```
from azureml.pipeline.core.run import PipelineRun
published_pipeline_run = PipelineRun(ws.experiments[experiment_name], run_id)
published_pipeline_run.wait_for_completion(show_output=True)
```
## Schedule the Pipeline
Suppose the clinic for the diabetes patients collects new data each week, and adds it to the dataset. You could run the pipeline every week to retrain the model with the new data.
```
from azureml.pipeline.core import ScheduleRecurrence, Schedule
# Submit the Pipeline every Monday at 00:00 UTC
recurrence = ScheduleRecurrence(frequency="Week", interval=1, week_days=["Monday"], time_of_day="00:00")
weekly_schedule = Schedule.create(ws, name="weekly-diabetes-training",
description="Based on time",
pipeline_id=published_pipeline.id,
experiment_name='mslearn-diabetes-pipeline',
recurrence=recurrence)
print('Pipeline scheduled.')
```
You can retrieve the schedules that are defined in the workspace like this:
```
schedules = Schedule.list(ws)
schedules
```
You can check the latest run like this:
```
pipeline_experiment = ws.experiments.get('mslearn-diabetes-pipeline')
latest_run = list(pipeline_experiment.get_runs())[0]
latest_run.get_details()
```
This is a simple example, designed to demonstrate the principle. In reality, you could build more sophisticated logic into the pipeline steps - for example, evaluating the model against some test data to calculate a performance metric like AUC or accuracy, comparing the metric to that of any previously registered versions of the model, and only registering the new model if it performs better.
You can use the [Azure Machine Learning extension for Azure DevOps](https://marketplace.visualstudio.com/items?itemName=ms-air-aiagility.vss-services-azureml) to combine Azure ML pipelines with Azure DevOps pipelines (yes, it *is* confusing that they have the same name!) and integrate model retraining into a *continuous integration/continuous deployment (CI/CD)* process. For example you could use an Azure DevOps *build* pipeline to trigger an Azure ML pipeline that trains and registers a model, and when the model is registered it could trigger an Azure Devops *release* pipeline that deploys the model as a web service, along with the application or service that consumes the model.
| github_jupyter |
```
from IPython.core.display import HTML
def css_styling():
styles = open("./styles/custom.css", "r").read()
return HTML(styles)
css_styling()
```
# Approximate solutions to the Riemann Problem
## Solutions in practice
Solutions to the Riemann problem are mainly used in two contexts:
1. As reference solutions against which a numerical method is benchmarked, or
2. As part of a numerical method, such as a high resolution shock capturing method, where the flux between two numerical cells is required.
In the first case, accuracy is paramount and the complete solution (all wave speeds, and all intermediate states) is required. In the second case only one thing is required: the flux ${\bf f}^*$ between the cells, which is the flux on the characteristic line $\xi = x / t = 0$.
In this second case, the numerical method will have to repeatedly solve the Riemann problem. In a general problem, the solution may be needed tens of times *per cell, per timestep*, leading to millions (or more!) solutions in a simulation. The speed of the solution is then extremely important, and approximate solutions are often used.
## Roe-type solutions
The most obvious simplification is to reduce the nonlinear problem
\begin{equation}
\partial_t {\bf q} + \partial_x {\bf f}({\bf q}) = {\bf 0}
\end{equation}
to the *linear* problem
\begin{equation}
\partial_t {\bf q} + A \partial_x {\bf q} = {\bf 0},
\end{equation}
where $A$ is a *constant* matrix that approximates the Jacobian $\partial {\bf f} / \partial {\bf q}$. We can then solve the linear problem exactly (e.g. by diagonalising the matrix and solving the resulting uncoupled advection equations), to find
\begin{align}
{\bf q}(x, t) & = {\bf q}_l + \sum_{p: \lambda^{(p)} < \tfrac{x}{t}} \left\{ {\bf l}^{(p)} \cdot \left( {\bf q}_r - {\bf q}_l \right) \right\} {\bf r}^{(p)}, \\
& = {\bf q}_r - \sum_{p: \lambda^{(p)} > \tfrac{x}{t}} \left\{ {\bf l}^{(p)} \cdot \left( {\bf q}_r - {\bf q}_l \right) \right\} {\bf r}^{(p)}, \\
& = \frac{1}{2} \left( {\bf q}_l + {\bf q}_r \right) + \sum_{p: \lambda^{(p)} < \tfrac{x}{t}} \left\{ {\bf l}^{(p)} \cdot \left( {\bf q}_r - {\bf q}_l \right) \right\} {\bf r}^{(p)} - \sum_{p: \lambda^{(p)} > \tfrac{x}{t}} \left\{ {\bf l}^{(p)} \cdot \left( {\bf q}_r - {\bf q}_l \right) \right\} {\bf r}^{(p)}.
\end{align}
where $\lambda^{(p)}, {\bf r}^{(p)},$ and ${\bf l}^{(p)}$ are the eigenvalues and the (right and left respectively) eigenvectors of $A$, ordered such that $\lambda^{(1)} \le \dots \le \lambda^{(N)}$ as usual. All three solutions are equivalent; the last is typically used.
Given this complete solution, it is easily evaluated along $x = 0$, and the flux calculated from the result.
An even greater shortcut can be found by noting that we are approximating ${\bf f} = A {\bf q}$. Therefore the standard form is to write
\begin{equation}
{\bf f}^* = \frac{1}{2} \left( {\bf f}_l + {\bf f}_r \right) + \sum_{p} \left| \lambda^{(p)} \right| \left\{ {\bf l}^{(p)} \cdot \left( {\bf q}_r - {\bf q}_l \right) \right\} {\bf r}^{(p)},
\end{equation}
where now we are summing over all eigenvalues and eigenvectors. It should be noted that ${\bf f}^* \ne {\bf f}({\bf q}^*)$ in general, as the calculation of ${\bf f}^*$ relied on an approximation to the flux.
In order to complete this specification of the solver, we only need to say how $A$ is defined. Roe gave the suggestion that
\begin{equation}
A = A({\bf q}_{\textrm{Roe}}) = \left. \frac{\partial {\bf f}}{\partial {\bf q}} \right|_{{\bf q}_{\textrm{Roe}}},
\end{equation}
where the *Roe average* ${\bf q}_{\textrm{Roe}}$ satisfies
1. $A({\bf q}_{\textrm{Roe}}) \left( {\bf q}_r - {\bf q}_l \right) = {\bf f}_r - {\bf f}_l$,
2. $A({\bf q}_{\textrm{Roe}})$ is diagonalizable with real eigenvalues, and
3. $A({\bf q}_{\textrm{Roe}}) \to \partial {\bf f} / \partial {\bf q}$ smoothly as ${\bf q}_{\textrm{Roe}} \to {\bf q}$.
It is *possible* to construct the Roe average for many systems (such as the Euler equations, and the relativistic Euler equations). However, a simple arithmetic average is often nearly as good - in the sense that the algorithm will fail only slightly more often than the algorithm with the full Roe average!
The problem with Roe type solvers is that it approximates all waves as discontinuities. This leads to inaccuracies near rarefactions, and these can be catastrophically bad when the rarefaction fan crosses $\xi = 0$ (a *sonic rarefaction*). It is possible to detect when these problems will occur (e.g. by looking at when $\lambda^{(p)}$ changes sign between the left and right states) and change the approximation at this point, often known as an *entropy fix*. More systematic and complex methods that extend the Roe approach whilst avoiding this problem include the *Marquina* solver.
## HLL-type solutions
An alternative type of method simplifies the wave structure even more, by simplifying the number of waves. HLL (for Harten, Lax and van Leer) type solutions assume that
1. there are two waves, both discontinuities, separating a constant central state in the solution, and
2. the waves propagate at the (known) speeds $\xi_{(\pm)}$.
From these assumptions, and the Rankine-Hugoniot conditions, we have the two equations
\begin{align}
\xi_{(-)} \left[ {\bf q}_m - {\bf q}_l \right] & = {\bf f}_m - {\bf f}_l, \\
\xi_{(+)} \left[ {\bf q}_r - {\bf q}_m \right] & = {\bf f}_r - {\bf f}_m.
\end{align}
These are immediately solved to give
\begin{align}
{\bf q}_m & = \frac{\xi_{(+)} {\bf q}_r - \xi_{(-)} {\bf q}_l - {\bf f}_r + {\bf f}_l}{\xi_{(+)} - \xi_{(-)}}, \\
{\bf f}_m & = \frac{\hat{\xi}_{(+)} {\bf f}_l - \hat{\xi}_{(-)} {\bf f}_r + \hat{\xi}_{(+)} \hat{\xi}_{(-)} \left( {\bf q}_r - {\bf q}_r \right)}{\hat{\xi}_{(+)} - \hat{\xi}_{(-)}},
\end{align}
where
\begin{equation}
\hat{\xi}_{(-)} = \min(0, \xi_{(-)}), \qquad \hat{\xi}_{(+)} = \max(0, \xi_{(+)}).
\end{equation}
Again it should be noted that, in general, ${\bf f}_m \ne {\bf f}({\bf q}_m)$.
We still need some way to compute the wave speeds $\xi_{(\pm)}$. The simplest method is to make them as large as possible, compatible with stability. This means (via the CFL condition) setting
\begin{equation}
-\xi_{(-)} = \xi_{(+)} = \frac{\Delta x}{\Delta t}
\end{equation}
which implies that (as the central state is now guaranteed to include the origin, as the waves have different signs)
\begin{equation}
{\bf f}^* = \frac{1}{2} \left( {\bf f}_l + {\bf f}_r + \frac{\Delta x}{\Delta t} \left[ {\bf q}_l - {\bf q}_r \right] \right).
\end{equation}
This is the *Lax-Friedrichs* flux, as [used in HyperPython](https://github.com/ketch/HyperPython). We can also easily see how the *local* Lax-Friedrichs method, [used in lesson 3 of HyperPython](http://nbviewer.ipython.org/github/ketch/HyperPython/blob/master/Lesson_03_High-resolution_methods.ipynb), comes about: simply choose
\begin{equation}
-\xi_{(-)} = \xi_{(+)} = \alpha = \min \left( \left| \lambda \left( {\bf q}_l \right) \right|, \left| \lambda \left( {\bf q}_r \right) \right| \right)
\end{equation}
to get
\begin{equation}
{\bf f}^* = \frac{1}{2} \left( {\bf f}_l + {\bf f}_r + \alpha \left[ {\bf q}_l - {\bf q}_r \right] \right).
\end{equation}
HLL type methods are straightforward to use but typically do not capture linear waves, such as the contact wave in the Euler equations, well. Extending the HLL method by including more waves is possible (see the *HLLC* method in Toro's book as an example), but rapidly increases the complexity of the solver.
| github_jupyter |
## TFMA Notebook example
This notebook describes how to export your model for TFMA and demonstrates the analysis tooling it offers.
## Setup
Import necessary packages.
```
import apache_beam as beam
import os
import preprocess
import shutil
import tensorflow as tf
import tensorflow_data_validation as tfdv
import tensorflow_model_analysis as tfma
from google.protobuf import text_format
from tensorflow.python.lib.io import file_io
from tensorflow_transform.beam.tft_beam_io import transform_fn_io
from tensorflow_transform.coders import example_proto_coder
from tensorflow_transform.saved import saved_transform_io
from tensorflow_transform.tf_metadata import dataset_schema
from tensorflow_transform.tf_metadata import schema_utils
from trainer import task
from trainer import taxi
```
Helper functions and some constants for running the notebook locally.
```
BASE_DIR = os.getcwd()
DATA_DIR = os.path.join(BASE_DIR, 'data')
OUTPUT_DIR = os.path.join(BASE_DIR, 'chicago_taxi_output')
# Base dir containing train and eval data
TRAIN_DATA_DIR = os.path.join(DATA_DIR, 'train')
EVAL_DATA_DIR = os.path.join(DATA_DIR, 'eval')
# Base dir where TFT writes training data
TFT_TRAIN_OUTPUT_BASE_DIR = os.path.join(OUTPUT_DIR, 'tft_train')
TFT_TRAIN_FILE_PREFIX = 'train_transformed'
# Base dir where TFT writes eval data
TFT_EVAL_OUTPUT_BASE_DIR = os.path.join(OUTPUT_DIR, 'tft_eval')
TFT_EVAL_FILE_PREFIX = 'eval_transformed'
TF_OUTPUT_BASE_DIR = os.path.join(OUTPUT_DIR, 'tf')
# Base dir where TFMA writes eval data
TFMA_OUTPUT_BASE_DIR = os.path.join(OUTPUT_DIR, 'tfma')
SERVING_MODEL_DIR = 'serving_model_dir'
EVAL_MODEL_DIR = 'eval_model_dir'
def get_tft_train_output_dir(run_id):
return _get_output_dir(TFT_TRAIN_OUTPUT_BASE_DIR, run_id)
def get_tft_eval_output_dir(run_id):
return _get_output_dir(TFT_EVAL_OUTPUT_BASE_DIR, run_id)
def get_tf_output_dir(run_id):
return _get_output_dir(TF_OUTPUT_BASE_DIR, run_id)
def get_tfma_output_dir(run_id):
return _get_output_dir(TFMA_OUTPUT_BASE_DIR, run_id)
def _get_output_dir(base_dir, run_id):
return os.path.join(base_dir, 'run_' + str(run_id))
def get_schema_file():
return os.path.join(OUTPUT_DIR, 'schema.pbtxt')
```
Clean up output directories.
```
shutil.rmtree(TFT_TRAIN_OUTPUT_BASE_DIR, ignore_errors=True)
shutil.rmtree(TFT_EVAL_OUTPUT_BASE_DIR, ignore_errors=True)
shutil.rmtree(TF_OUTPUT_BASE_DIR, ignore_errors=True)
shutil.rmtree(get_schema_file(), ignore_errors=True)
```
## Compute and visualize descriptive data statistics
```
# Compute stats over training data.
train_stats = tfdv.generate_statistics_from_csv(data_location=os.path.join(TRAIN_DATA_DIR, 'data.csv'))
# Visualize training data stats.
tfdv.visualize_statistics(train_stats)
```
## Infer a schema
```
# Infer a schema from the training data stats.
schema = tfdv.infer_schema(statistics=train_stats, infer_feature_shape=False)
tfdv.display_schema(schema=schema)
```
## Check evaluation data for errors
```
# Compute stats over eval data.
eval_stats = tfdv.generate_statistics_from_csv(data_location=os.path.join(EVAL_DATA_DIR, 'data.csv'))
# Compare stats of eval data with training data.
tfdv.visualize_statistics(lhs_statistics=eval_stats, rhs_statistics=train_stats,
lhs_name='EVAL_DATASET', rhs_name='TRAIN_DATASET')
# Check eval data for errors by validating the eval data stats using the previously inferred schema.
anomalies = tfdv.validate_statistics(statistics=eval_stats, schema=schema)
tfdv.display_anomalies(anomalies)
# Update the schema based on the observed anomalies.
# Relax the minimum fraction of values that must come from the domain for feature company.
company = tfdv.get_feature(schema, 'company')
company.distribution_constraints.min_domain_mass = 0.9
# Add new value to the domain of feature payment_type.
payment_type_domain = tfdv.get_domain(schema, 'payment_type')
payment_type_domain.value.append('Prcard')
# Validate eval stats after updating the schema
updated_anomalies = tfdv.validate_statistics(eval_stats, schema)
tfdv.display_anomalies(updated_anomalies)
```
## Freeze the schema
Now that the schema has been reviewed and curated, we will store it in a file to reflect its "frozen" state.
```
file_io.recursive_create_dir(OUTPUT_DIR)
file_io.write_string_to_file(get_schema_file(), text_format.MessageToString(schema))
```
## Preprocess Inputs
transform_data is defined in preprocess.py and uses the tensorflow_transform library to perform preprocessing. The same code is used for both local preprocessing in this notebook and preprocessing in the Cloud (via Dataflow).
```
# Transform eval data
preprocess.transform_data(input_handle=os.path.join(EVAL_DATA_DIR, 'data.csv'),
outfile_prefix=TFT_EVAL_FILE_PREFIX,
working_dir=get_tft_eval_output_dir(0),
schema_file=get_schema_file(),
pipeline_args=['--runner=DirectRunner'])
print('Done')
# Transform training data
preprocess.transform_data(input_handle=os.path.join(TRAIN_DATA_DIR, 'data.csv'),
outfile_prefix=TFT_TRAIN_FILE_PREFIX,
working_dir=get_tft_train_output_dir(0),
schema_file=get_schema_file(),
pipeline_args=['--runner=DirectRunner'])
print('Done')
```
## Compute statistics over transformed data
```
# Compute stats over transformed training data.
TRANSFORMED_TRAIN_DATA = os.path.join(get_tft_train_output_dir(0), TFT_TRAIN_FILE_PREFIX + "*")
transformed_train_stats = tfdv.generate_statistics_from_tfrecord(data_location=TRANSFORMED_TRAIN_DATA)
# Visualize transformed training data stats and compare to raw training data.
# Use 'Feature search' to focus on a feature and see statistics pre- and post-transformation.
tfdv.visualize_statistics(transformed_train_stats, train_stats, lhs_name='TRANSFORMED', rhs_name='RAW')
```
## Prepare the Model
To use TFMA, export the model into an **EvalSavedModel** by calling ``tfma.export.export_eval_savedmodel``.
``tfma.export.export_eval_savedmodel`` is analogous to ``estimator.export_savedmodel`` but exports the evaluation graph as opposed to the training or inference graph. Notice that one of the inputs is ``eval_input_receiver_fn`` which is analogous to ``serving_input_receiver_fn`` for ``estimator.export_savedmodel``. For more details, refer to the documentation for TFMA on Github.
Contruct the **EvalSavedModel** after training is completed.
```
def run_experiment(hparams):
"""Run the training and evaluate using the high level API"""
# Train and evaluate the model as usual.
estimator = task.train_and_maybe_evaluate(hparams)
# Export TFMA's sepcial EvalSavedModel
eval_model_dir = os.path.join(hparams.output_dir, EVAL_MODEL_DIR)
receiver_fn = lambda: eval_input_receiver_fn(hparams.tf_transform_dir)
tfma.export.export_eval_savedmodel(
estimator=estimator,
export_dir_base=eval_model_dir,
eval_input_receiver_fn=receiver_fn)
def eval_input_receiver_fn(working_dir):
# Extract feature spec from the schema.
raw_feature_spec = schema_utils.schema_as_feature_spec(schema).feature_spec
serialized_tf_example = tf.placeholder(
dtype=tf.string, shape=[None], name='input_example_tensor')
# First we deserialize our examples using the raw schema.
features = tf.parse_example(serialized_tf_example, raw_feature_spec)
# Now that we have our raw examples, we must process them through tft
_, transformed_features = (
saved_transform_io.partially_apply_saved_transform(
os.path.join(working_dir, transform_fn_io.TRANSFORM_FN_DIR),
features))
# The key MUST be 'examples'.
receiver_tensors = {'examples': serialized_tf_example}
# NOTE: Model is driven by transformed features (since training works on the
# materialized output of TFT, but slicing will happen on raw features.
features.update(transformed_features)
return tfma.export.EvalInputReceiver(
features=features,
receiver_tensors=receiver_tensors,
labels=transformed_features[taxi.transformed_name(taxi.LABEL_KEY)])
print('Done')
```
## Train and export the model for TFMA
```
def run_local_experiment(tft_run_id, tf_run_id, num_layers, first_layer_size, scale_factor):
"""Helper method to train and export the model for TFMA
The caller specifies the input and output directory by providing run ids. The optional parameters
allows the user to change the modelfor time series view.
Args:
tft_run_id: The run id for the preprocessing. Identifies the folder containing training data.
tf_run_id: The run for this training run. Identify where the exported model will be written to.
num_layers: The number of layers used by the hiden layer.
first_layer_size: The size of the first hidden layer.
scale_factor: The scale factor between each layer in in hidden layers.
"""
hparams = tf.contrib.training.HParams(
# Inputs: are tf-transformed materialized features
train_files=os.path.join(get_tft_train_output_dir(tft_run_id), TFT_TRAIN_FILE_PREFIX + '-00000-of-*'),
eval_files=os.path.join(get_tft_eval_output_dir(tft_run_id), TFT_EVAL_FILE_PREFIX + '-00000-of-*'),
schema_file=get_schema_file(),
# Output: dir for trained model
job_dir=get_tf_output_dir(tf_run_id),
tf_transform_dir=get_tft_train_output_dir(tft_run_id),
# Output: dir for both the serving model and eval_model which will go into tfma
# evaluation
output_dir=get_tf_output_dir(tf_run_id),
train_steps=10000,
eval_steps=5000,
num_layers=num_layers,
first_layer_size=first_layer_size,
scale_factor=scale_factor,
num_epochs=None,
train_batch_size=40,
eval_batch_size=40)
run_experiment(hparams)
print('Done')
run_local_experiment(tft_run_id=0,
tf_run_id=0,
num_layers=4,
first_layer_size=100,
scale_factor=0.7)
print('Done')
```
## Run TFMA to compute metrics
For local analysis, TFMA offers a helper method ``tfma.run_model_analysis``
```
help(tfma.run_model_analysis)
```
#### You can also write your own custom pipeline if you want to perform extra transformations on the data before evaluation.
```
def run_tfma(slice_spec, tf_run_id, tfma_run_id, input_csv, schema_file, add_metrics_callbacks=None):
"""A simple wrapper function that runs tfma locally.
A function that does extra transformations on the data and then run model analysis.
Args:
slice_spec: The slicing spec for how to slice the data.
tf_run_id: An id to contruct the model directories with.
tfma_run_id: An id to construct output directories with.
input_csv: The evaluation data in csv format.
schema_file: The file holding a text-serialized schema for the input data.
add_metrics_callback: Optional list of callbacks for computing extra metrics.
Returns:
An EvalResult that can be used with TFMA visualization functions.
"""
eval_model_base_dir = os.path.join(get_tf_output_dir(tf_run_id), EVAL_MODEL_DIR)
eval_model_dir = os.path.join(eval_model_base_dir, next(os.walk(eval_model_base_dir))[1][0])
eval_shared_model = tfma.default_eval_shared_model(
eval_saved_model_path=eval_model_dir,
add_metrics_callbacks=add_metrics_callbacks)
schema = taxi.read_schema(schema_file)
print(eval_model_dir)
display_only_data_location = input_csv
with beam.Pipeline() as pipeline:
csv_coder = taxi.make_csv_coder(schema)
raw_data = (
pipeline
| 'ReadFromText' >> beam.io.ReadFromText(
input_csv,
coder=beam.coders.BytesCoder(),
skip_header_lines=True)
| 'ParseCSV' >> beam.Map(csv_coder.decode))
# Examples must be in clean tf-example format.
coder = taxi.make_proto_coder(schema)
raw_data = (
raw_data
| 'ToSerializedTFExample' >> beam.Map(coder.encode))
_ = (raw_data
| 'ExtractEvaluateAndWriteResults' >>
tfma.ExtractEvaluateAndWriteResults(
eval_shared_model=eval_shared_model,
slice_spec=slice_spec,
output_path=get_tfma_output_dir(tfma_run_id),
display_only_data_location=input_csv))
return tfma.load_eval_result(output_path=get_tfma_output_dir(tfma_run_id))
print('Done')
```
#### You can also compute metrics on slices of your data in TFMA. Slices can be specified using ``tfma.slicer.SingleSliceSpec``.
Below are examples of how slices can be specified.
```
# An empty slice spec means the overall slice, that is, the whole dataset.
OVERALL_SLICE_SPEC = tfma.slicer.SingleSliceSpec()
# Data can be sliced along a feature column
# In this case, data is sliced along feature column trip_start_hour.
FEATURE_COLUMN_SLICE_SPEC = tfma.slicer.SingleSliceSpec(columns=['trip_start_hour'])
# Data can be sliced by crossing feature columns
# In this case, slices are computed for trip_start_day x trip_start_month.
FEATURE_COLUMN_CROSS_SPEC = tfma.slicer.SingleSliceSpec(columns=['trip_start_day', 'trip_start_month'])
# Metrics can be computed for a particular feature value.
# In this case, metrics is computed for all data where trip_start_hour is 12.
FEATURE_VALUE_SPEC = tfma.slicer.SingleSliceSpec(features=[('trip_start_hour', 12)])
# It is also possible to mix column cross and feature value cross.
# In this case, data where trip_start_hour is 12 will be sliced by trip_start_day.
COLUMN_CROSS_VALUE_SPEC = tfma.slicer.SingleSliceSpec(columns=['trip_start_day'], features=[('trip_start_hour', 12)])
ALL_SPECS = [
OVERALL_SLICE_SPEC,
FEATURE_COLUMN_SLICE_SPEC,
FEATURE_COLUMN_CROSS_SPEC,
FEATURE_VALUE_SPEC,
COLUMN_CROSS_VALUE_SPEC
]
```
#### Let's run TFMA!
```
tf.logging.set_verbosity(tf.logging.INFO)
tfma_result_1 = run_tfma(input_csv=os.path.join(EVAL_DATA_DIR, 'data.csv'),
tf_run_id=0,
tfma_run_id=1,
slice_spec=ALL_SPECS,
schema_file=get_schema_file())
print('Done')
```
## Visualization: Slicing Metrics
To see the slices, either use the name of the column (by setting slicing_column) or provide a tfma.slicer.SingleSliceSpec (by setting slicing_spec). If neither is provided, the overall will be displayed.
The default visualization is **slice overview** when the number of slices is small. It shows the value of a metric for each slice sorted by the another metric. It is also possible to set a threshold to filter out slices with smaller weights.
This view also supports **metrics histogram** as an alternative visualization. It is also the defautl view when the number of slices is large. The results will be divided into buckets and the number of slices / total weights / both can be visualized. Slices with small weights can be fitlered out by setting the threshold. Further filtering can be applied by dragging the grey band. To reset the range, double click the band. Filtering can be used to remove outliers in the visualization and the metrics table below.
```
# Show data sliced along feature column trip_start_hour.
tfma.view.render_slicing_metrics(
tfma_result_1, slicing_column='trip_start_hour')
# Show metrics sliced by COLUMN_CROSS_VALUE_SPEC above.
tfma.view.render_slicing_metrics(tfma_result_1, slicing_spec=COLUMN_CROSS_VALUE_SPEC)
# Show overall metrics.
tfma.view.render_slicing_metrics(tfma_result_1)
```
## Visualization: Plots
TFMA offers a number of built-in plots. To see them, add them to ``add_metrics_callbacks``
```
tf.logging.set_verbosity(tf.logging.INFO)
tfma_vis = run_tfma(input_csv=os.path.join(EVAL_DATA_DIR, 'data.csv'),
tf_run_id=0,
tfma_run_id='vis',
slice_spec=ALL_SPECS,
schema_file=get_schema_file(),
add_metrics_callbacks=[
# calibration_plot_and_prediction_histogram computes calibration plot and prediction
# distribution at different thresholds.
tfma.post_export_metrics.calibration_plot_and_prediction_histogram(),
# auc_plots enables precision-recall curve and ROC visualization at different thresholds.
tfma.post_export_metrics.auc_plots()
])
print('Done')
```
Plots must be visualized for an individual slice. To specify a slice, use ``tfma.slicer.SingleSliceSpec``.
In the example below, we are using ``tfma.slicer.SingleSliceSpec(features=[('trip_start_hour', 0)])`` to specify the slice where trip_start_hour is 0.
Plots are interactive:
- Drag to pan
- Scroll to zoom
- Right click to reset the view
Simply hover over the desired data point to see more details.
```
tfma.view.render_plot(tfma_vis, tfma.slicer.SingleSliceSpec(features=[('trip_start_hour', 0)]))
```
#### Custom metrics
In addition to plots, it is also possible to compute additional metrics not present at export time or custom metrics metrics using ``add_metrics_callbacks``.
All metrics in ``tf.metrics`` are supported in the callback and can be used to compose other metrics:
https://www.tensorflow.org/api_docs/python/tf/metrics
In the cells below, false negative rate is computed as an example.
```
# Defines a callback that adds FNR to the result.
def add_fnr_for_threshold(threshold):
def _add_fnr_callback(features_dict, predictions_dict, labels_dict):
metric_ops = {}
prediction_tensor = tf.cast(
predictions_dict.get(tf.contrib.learn.PredictionKey.LOGISTIC), tf.float64)
fn_value_op, fn_update_op = tf.metrics.false_negatives_at_thresholds(tf.squeeze(labels_dict),
tf.squeeze(prediction_tensor),
[threshold])
tp_value_op, tp_update_op = tf.metrics.true_positives_at_thresholds(tf.squeeze(labels_dict),
tf.squeeze(prediction_tensor),
[threshold])
fnr = fn_value_op[0] / (fn_value_op[0] + tp_value_op[0])
metric_ops['FNR@' + str(threshold)] = (fnr, tf.group(fn_update_op, tp_update_op))
return metric_ops
return _add_fnr_callback
tf.logging.set_verbosity(tf.logging.INFO)
tfma_fnr = run_tfma(input_csv=os.path.join(EVAL_DATA_DIR, 'data.csv'),
tf_run_id=0,
tfma_run_id='fnr',
slice_spec=ALL_SPECS,
schema_file=get_schema_file(),
add_metrics_callbacks=[
# Simply add the call here.
add_fnr_for_threshold(0.75)
])
tfma.view.render_slicing_metrics(tfma_fnr, slicing_spec=FEATURE_COLUMN_SLICE_SPEC)
```
## Visualization: Time Series
It is important to track how your model is doing over time. TFMA offers two modes to show your model performs over time.
**Multiple model analysis** shows how model perfoms from one version to another. This is useful early on to see how the addition of new features, change in modeling technique, etc, affects the performance. TFMA offers a convenient method.
```
help(tfma.multiple_model_analysis)
```
**Multiple data analysis** shows how a model perfoms under different evaluation data set. This is useful to ensure that model performance does not degrade over time. TFMA offer a conveneient method.
```
help(tfma.multiple_data_analysis)
```
It is also possible to compose a time series manually.
```
# Create different models.
# Run some experiments with different hidden layer configurations.
run_local_experiment(tft_run_id=0,
tf_run_id=1,
num_layers=3,
first_layer_size=200,
scale_factor=0.7)
run_local_experiment(tft_run_id=0,
tf_run_id=2,
num_layers=4,
first_layer_size=240,
scale_factor=0.5)
print('Done')
tfma_result_2 = run_tfma(input_csv=os.path.join(EVAL_DATA_DIR, 'data.csv'),
tf_run_id=1,
tfma_run_id=2,
slice_spec=ALL_SPECS,
schema_file=get_schema_file())
tfma_result_3 = run_tfma(input_csv=os.path.join(EVAL_DATA_DIR, 'data.csv'),
tf_run_id=2,
tfma_run_id=3,
slice_spec=ALL_SPECS,
schema_file=get_schema_file())
print('Done')
```
Like plots, time series view must visualized for a slice too.
In the example below, we are showing the overall slice.
Select a metric to see its time series graph. Hover over each data point to get more details.
```
eval_results = tfma.make_eval_results([tfma_result_1, tfma_result_2, tfma_result_3],
tfma.constants.MODEL_CENTRIC_MODE)
tfma.view.render_time_series(eval_results, OVERALL_SLICE_SPEC)
```
Serialized results can also be used to construct a time series. Thus, there is no need to re-run TFMA for models already evaluated for a long running pipeline.
```
# Visualize the results in a Time Series. In this case, we are showing the slice specified.
eval_results_from_disk = tfma.load_eval_results([get_tfma_output_dir(1),
get_tfma_output_dir(2),
get_tfma_output_dir(3)],
tfma.constants.MODEL_CENTRIC_MODE)
tfma.view.render_time_series(eval_results_from_disk, FEATURE_VALUE_SPEC)
```
| github_jupyter |
# Think Bayes
This notebook presents code and exercises from Think Bayes, second edition.
Copyright 2018 Allen B. Downey
MIT License: https://opensource.org/licenses/MIT
```
# Configure Jupyter so figures appear in the notebook
%matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
%config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
import math
import numpy as np
from thinkbayes2 import Pmf, Cdf, Suite, Joint
import thinkplot
```
### The flea beetle problem
Different species of flea beetle can be distinguished by the width and angle of the aedeagus. The data below includes measurements and know species classification for 74 specimens.
Suppose you discover a new specimen under conditions where it is equally likely to be any of the three species. You measure the aedeagus and width 140 microns and angle 15 (in multiples of 7.5 degrees). What is the probability that it belongs to each species?
This problem is based on [this data story on DASL](https://web.archive.org/web/20160304083805/http://lib.stat.cmu.edu/DASL/Datafiles/FleaBeetles.html)
Datafile Name: Flea Beetles
Datafile Subjects: Biology
Story Names: Flea Beetles
Reference: Lubischew, A.A. (1962) On the use of discriminant functions in taxonomy. Biometrics, 18, 455-477. Also found in: Hand, D.J., et al. (1994) A Handbook of Small Data Sets, London: Chapman & Hall, 254-255.
Authorization: Contact Authors
Description: Data were collected on the genus of flea beetle Chaetocnema, which contains three species: concinna (Con), heikertingeri (Hei), and heptapotamica (Hep). Measurements were made on the width and angle of the aedeagus of each beetle. The goal of the original study was to form a classification rule to distinguish the three species.
Number of cases: 74
Variable Names:
Width: The maximal width of aedeagus in the forpart (in microns)
Angle: The front angle of the aedeagus (1 unit = 7.5 degrees)
Species: Species of flea beetle from the genus Chaetocnema
To solve this problem we have to account for two sources of uncertainty: given the data, we have some uncertainty about the actual distribution of attributes. Then, given the measurements, we have uncertainty about which species we have.
First I'll load the data.
```
measurements = (140, 15)
import pandas as pd
df = pd.read_csv('../data/flea_beetles.csv', delimiter='\t')
df.head()
def plot_cdfs(df, col):
for name, group in df.groupby('Species'):
cdf = Cdf(group[col], label=name)
thinkplot.Cdf(cdf)
thinkplot.Config(xlabel=col, legend=True, loc='lower right')
plot_cdfs(df, 'Width')
plot_cdfs(df, 'Angle')
```
The following class estimates the mean and standard deviation of a normal distribution, given the data:
```
from scipy.stats import norm
from thinkbayes2 import EvalNormalPdf
class Beetle(Suite, Joint):
def Likelihood(self, data, hypo):
"""
data: sequence of measurements
hypo: mu, sigma
"""
mu, sigma = hypo
likes = EvalNormalPdf(data, mu, sigma)
return np.prod(likes)
def PredictiveProb(self, data):
"""Compute the posterior total probability of a datum.
data: sequence of measurements
"""
total = 0
for (mu, sigma), prob in self.Items():
likes = norm.pdf(data, mu, sigma)
total += prob * np.prod(likes)
return total
```
Now we can estimate parameters for the widths, for each of the three species.
```
from itertools import product
def MakeWidthSuite(data):
mus = np.linspace(115, 160, 51)
sigmas = np.linspace(1, 10, 51)
suite = Beetle(product(mus, sigmas))
suite.Update(data)
return suite
groups = df.groupby('Species')
```
Here are the posterior distributions for mu and sigma, and the predictive probability of the width measurement, for each species.
```
for name, group in groups:
suite = MakeWidthSuite(group.Width)
thinkplot.Contour(suite)
print(name, suite.PredictiveProb(140))
```
Now we can do the same thing for the angles.
```
def MakeAngleSuite(data):
mus = np.linspace(8, 16, 101)
sigmas = np.linspace(0.1, 2, 101)
suite = Beetle(product(mus, sigmas))
suite.Update(data)
return suite
for name, group in groups:
suite = MakeAngleSuite(group.Angle)
thinkplot.Contour(suite)
print(name, suite.PredictiveProb(15))
```
These posterior distributions are used to compute the likelihoods of the measurements.
```
class Species:
def __init__(self, name, suite_width, suite_angle):
self.name = name
self.suite_width = suite_width
self.suite_angle = suite_angle
def __str__(self):
return self.name
def Likelihood(self, data):
width, angle = data
like1 = self.suite_width.PredictiveProb(width)
like2 = self.suite_angle.PredictiveProb(angle)
return like1 * like2
species = {}
for name, group in groups:
suite_width = MakeWidthSuite(group.Width)
suite_angle = MakeAngleSuite(group.Angle)
species[name] = Species(name, suite_width, suite_angle)
```
For example, here's the likelihood of the data given that the species is 'Con'
```
species['Con'].Likelihood(measurements)
```
Now we can make a `Classifier` that uses the `Species` objects as hypotheses.
```
class Classifier(Suite):
def Likelihood(self, data, hypo):
return hypo.Likelihood(data)
suite = Classifier(species.values())
for hypo, prob in suite.Items():
print(hypo, prob)
suite.Update(measurements)
for hypo, prob in suite.Items():
print(hypo, prob)
```
## Now with MCMC
Based on [this example](https://docs.pymc.io/notebooks/LKJ.html)
```
from warnings import simplefilter
simplefilter('ignore', FutureWarning)
import pymc3 as pm
N = 10000
μ_actual = np.array([1, -2])
Σ_actual = np.array([[0.5, -0.3],
[-0.3, 1.]])
x = np.random.multivariate_normal(μ_actual, Σ_actual, size=N)
df['Width10'] = df.Width / 10
observed = {}
for name, group in df.groupby('Species'):
observed[name] = group[['Width10', 'Angle']].values
print(name)
print(np.cov(np.transpose(observed[name])))
x = observed['Con']
with pm.Model() as model:
packed_L = pm.LKJCholeskyCov('packed_L', n=2,
eta=2, sd_dist=pm.HalfCauchy.dist(2.5))
with model:
L = pm.expand_packed_triangular(2, packed_L)
Σ = pm.Deterministic('Σ', L.dot(L.T))
with model:
μ = pm.Normal('μ', 0., 10., shape=2,
testval=x.mean(axis=0))
obs = pm.MvNormal('obs', μ, chol=L, observed=x)
with model:
trace = pm.sample(1000)
pm.traceplot(trace);
μ_post = trace['μ'].mean(axis=0)
Σ_post = trace['Σ'].mean(axis=0)
from statsmodels.stats.moment_helpers import cov2corr
from scipy.stats import multivariate_normal
cov2corr(Σ_post)
measured = (14, 15)
total = 0
for row in trace:
total += multivariate_normal.pdf(measured, mean=row['μ'], cov=row['Σ'])
total / len(trace)
def compute_posterior_likelihood(measured, species):
x = observed[species]
with pm.Model() as model:
packed_L = pm.LKJCholeskyCov('packed_L', n=2,
eta=2, sd_dist=pm.HalfCauchy.dist(2.5))
L = pm.expand_packed_triangular(2, packed_L)
Σ = pm.Deterministic('Σ', L.dot(L.T))
μ = pm.Normal('μ', 0., 10., shape=2,
testval=x.mean(axis=0))
obs = pm.MvNormal('obs', μ, chol=L, observed=x)
trace = pm.sample(1000)
total = 0
for row in trace:
total += multivariate_normal.pdf(measured, mean=row['μ'], cov=row['Σ'])
return total / len(trace)
suite = Suite(['Con', 'Hep', 'Hei'])
for hypo in suite:
like = compute_posterior_likelihood(measured, hypo)
print(hypo, like)
suite[hypo] *= like
suite.Normalize()
suite.Print()
```
| github_jupyter |
# Biomedical Databases: Design, Implementation and Optimisation
### WS- 2017/2018
## Team 3 lab report
**By**
Mohammed Abdelgadir Hassan
Nour Al-hanafi
Colin Jan Birkenbihl
Dejan Djukic
Faridullah Khan
Gergö Szita
Lanying Wei
# 1. Overview
In course of the Biomedical Databases Lab Course project we planned a framework of three communicating databases dedicated to protein - disease, disease - phenotype and protein - pathway interactions. Such a framework could for example be used to annotate pathway data with disease information or to compare diseases based on associated phenotypes or involved proteins (figure. 1).
As team 3, our part of project comprised mapping proteins to diseases. For this, we aimed at creating a database system that can be easily queried in order for the user to find disease-protein associations (figure. 2). The user can provide either a protein id as an input and obtain all associated diseases as an output or vice versa. Furthermore, the user can query many diseases simultaneously and obtain all proteins linked to theses diseases.
This may help find novel molecular targets for potential new therapeutics and provide new windows on disease treatments and metabolic process control.
<img src="db_Interfaces.png">
Figure. 1 Interface between databases
<img src="figure.2.jpg">
Figure. 2 Teams contributions to the databases framework
# 2. Materials & Methods
## 2. 1. Materials
We investigated many biological databases in which data may have included information about both proteins and diseases. We found that UNIPROT had the required information. Raw data existed as a text file containing proteins-ids, proteins names and disease ids (OMIM).
## 2. 2. Methods
We used python for downloading and parsing the data, and building the database with the help of SQLAlchemy. We mapped OMIM to uniprotIDs and vice versa.
### 2. 2. 1. Retrieve the raw data
The text file which includes the data is available at : http://www.uniprot.org/docs/mimtosp.txt .
It was downloaded and parsed automatically by a python script. Since the number of relations in each row was not consistent, built-in Pandas functions were not used to parse the document linked above; a custom parser had to be made which delivers the relations as a list of lists to the method which populates the database.
### 2. 2. 2. Build the data model
The dataset has many to many relationship and to accommodate this, an Association table is set up to link between the Protein and Disease tables. The association table allows for pre-defining the join conditions within the database so that cross-table queries are easily accomplished and Table objects readily obtainable, as demonstrated by two versions of our queries.
A custom SQLalchemy-based function was created to quickly (5-10 seconds of execution time) populate the database with disease - protein association data, which is also downloaded and parsed automatically by module “mapping_parser_v2” included in our package. We expect the URL of our data to be stable and we have recorded it in the constants file so it could be easily changed when or if the original URL gets broken.
A schematic of our final disease model is shown below in figure 3 and it is implemented in the “datamodel” module of our package. which holds all three tables as classes which are then imported into our database manager which utilizes our data model implementation for creating, populating and querying the database.
Even though that the data model is simple, it is easily expandable if need be and more importantly, it fills a critical gap of relating the proteins to diseases whereas most of the available tools (i.e. DisGeNET) holds the associations between genes and diseases.
Figure. 3: Data model schema; Team 3
### 2. 2. 3. Create the database manager
All of the database-related operations are handled from within the “db_manager” module of our package. It is built based on SQLalchemy session-aware design and according to best practices shown by the SQLalchemy documentation.
### 2. 2. 4. Create the database interactor
The Interactor is a class that combines the three databases of team 1, 2 and 3. It supports SQLalchemy query functions, which query across the different databases.
An example for these queries would be the annotate_pathway function:
Using a WikiPathway identifier in a query the genes interacting in the pathway will be retrieved. These genes will then be connected to the Uniprot identifiers of the associated proteins. Using the Uniprot identifiers the database of team 1 will be queried for the disease names associated with these proteins. Thus a link between WikiPathways and diseases is established.
### 2. 2. 5. Packaging
We built a project package which could be imported and worked with after installed to make the software useful and user friendly. Such all the important functions can be directly used and the user can leverage from the provided documentation and examples.
# 3. Results
## 3. 1. The processed data
The raw (relatively unstructured) text file from Uniprot was successfully processed with through our mapper_parser module and a list of list was returned for further usage in populating the database. After benchmarking the final database performance, we noticed that not all OMIM ID’s in the file actually map to diseases within OMIM and for that, a workaround for filtering out the OMIM ID’s which refer to irrelevant entities was created, based on another .tsv file holding the descriptions of all OMIM ID’s.
## 3. 2. The database
The final 3-table data model was implemented in SQLalchemy and tested through populating and querying the database on numerous examples. Using SQLalchemy makes it possible to set up the database for different SQL services (MySQL, PostGres
Uniprot accession numbers can successfully be mapped to associated diseases (in particular, their OMIM ID’s) very quickly, in a many-to-many relationship. Please see figure 3 for a detailed schematic of the final data model.
# 3. 3. The manager
The manager is a python class, which takes care of all the database related functionalities.
Our database manager is able to create and fill the database. It will create three tables (proteins, diseases and association between them). The populate_db function of the manager will take care of download the raw uniprot data, parsing the data and populating the database with it.
Functions to query the database using either uniprot id(s) or OMIM id(s) to retrieve data are also implemented in the manager.
## 3. 3. The interactor
With the interactor class we provided an object, that enables the user to perform more complicated queries across the three databases. It can be used to answer scientific questions, that could not have been answered by looking only into one data resource. Provided functionalities include for example the annotation of WikiPathway pathways with disease names.
# 3. 4. Annotating pathways with associated diseases
WikiPathway pathways are not linked to any disease information so far. Using our packages and the interactor class we provided the means necessary to accomplish this task. When passing WikiPathway identifiers to the annotate_pathways function of the interactor, the interactor will perform queries (described in more detail in 2.2.4) and then outputs the mapping of the pathway identifier to all diseases it is associated with. This can be done repeatedly and ensures that the annotation is up to date.
# 4. Conclusion
The package allows for investigating protein-disease relationships so the user can gain knowledge about which diseases relate to which proteins. This can be used as a standalone application and finding interesting proteins regards to a disease. Moreover, the package can be used with the two other packages to show the relationship between disease phenotype and protein pathways.
# 5. Limitations
The main restriction of our software is that it relies on another package within the framework (namely, group 1 which operates on the Monarch/OMIM database) to provide the user with the full names of the OMIM diseases related to a particular protein, and in a sense, vice versa; It is mainly only protein accession ID's that are returned, not the full protein names.
This is due to the fact that the package was developed in such a way that it resides in a three-package ecosystem where all packages have a role to fill in reaching the final goal. Because of that, our queries are constructed in such a way that they return the most unique pieces of information (ID's) which is the most useful to other databases and not the full names themselves.
To somewhat ameliorate the problem, a method in the “db_manager” module with the name “find_protein()” is implemented, it receives a Uniprot accession number as input and returns the name of the related protein.
Another reason for the lack of descriptiveness in our output is due to the fact that the best and most directly usable data we could find for the protein-disease associations hosted on Uniprot is delivered in such a way that we only have access to Uniprot names of the proteins, which, unfortunately, are not completely user-friendly (i.e. A4_HUMAN is Amyloid-beta A4 protein)
## 5.1 Future improvements
If we would to, for example, start a master thesis based on a similar approach and improve on what we have done here, we would do the following:
- We would parse the whole Uniprot xml raw data file and extract much more information from it;
- We would eliminate the reliance to external packages and develop our own either API handles or local database tables which contain all of the information required for solving all of the scientific questions in mind
- We would implement additional algorithms for walking the association search space
# 6. Contributors
Colin Jan Birkenbihl
Dejan Djukic
Faridullah Khan
Gergö Szita
Lanying Wei
Mohammed Abdelgadir Hassan
Nour Al-hanafi
| github_jupyter |
# Regression Week 5: LASSO (coordinate descent)
In this notebook, you will implement your very own LASSO solver via coordinate descent. You will:
* Write a function to normalize features
* Implement coordinate descent for LASSO
* Explore effects of L1 penalty
# Fire up graphlab create
Make sure you have the latest version of graphlab (>= 1.7)
```
import graphlab
```
# Load in house sales data
Dataset is from house sales in King County, the region where the city of Seattle, WA is located.
```
sales = graphlab.SFrame('kc_house_data.gl/')
# In the dataset, 'floors' was defined with type string,
# so we'll convert them to int, before using it below
sales['floors'] = sales['floors'].astype(int)
```
If we want to do any "feature engineering" like creating new features or adjusting existing ones we should do this directly using the SFrames as seen in the first notebook of Week 2. For this notebook, however, we will work with the existing features.
# Import useful functions from previous notebook
As in Week 2, we convert the SFrame into a 2D Numpy array. Copy and paste `get_num_data()` from the second notebook of Week 2.
```
import numpy as np # note this allows us to refer to numpy as np instead
```
Also, copy and paste the `predict_output()` function to compute the predictions for an entire matrix of features given the matrix and the weights:
# Normalize features
In the house dataset, features vary wildly in their relative magnitude: `sqft_living` is very large overall compared to `bedrooms`, for instance. As a result, weight for `sqft_living` would be much smaller than weight for `bedrooms`. This is problematic because "small" weights are dropped first as `l1_penalty` goes up.
To give equal considerations for all features, we need to **normalize features** as discussed in the lectures: we divide each feature by its 2-norm so that the transformed feature has norm 1.
Let's see how we can do this normalization easily with Numpy: let us first consider a small matrix.
```
X = np.array([[3.,5.,8.],[4.,12.,15.]])
print X
```
Numpy provides a shorthand for computing 2-norms of each column:
```
norms = np.linalg.norm(X, axis=0) # gives [norm(X[:,0]), norm(X[:,1]), norm(X[:,2])]
print norms
```
To normalize, apply element-wise division:
```
print X / norms # gives [X[:,0]/norm(X[:,0]), X[:,1]/norm(X[:,1]), X[:,2]/norm(X[:,2])]
```
Using the shorthand we just covered, write a short function called `normalize_features(feature_matrix)`, which normalizes columns of a given feature matrix. The function should return a pair `(normalized_features, norms)`, where the second item contains the norms of original features. As discussed in the lectures, we will use these norms to normalize the test data in the same way as we normalized the training data.
To test the function, run the following:
```
features, norms = normalize_features(np.array([[3.,6.,9.],[4.,8.,12.]]))
print features
# should print
# [[ 0.6 0.6 0.6]
# [ 0.8 0.8 0.8]]
print norms
# should print
# [5. 10. 15.]
```
# Implementing Coordinate Descent with normalized features
We seek to obtain a sparse set of weights by minimizing the LASSO cost function
```
SUM[ (prediction - output)^2 ] + lambda*( |w[1]| + ... + |w[k]|).
```
(By convention, we do not include `w[0]` in the L1 penalty term. We never want to push the intercept to zero.)
The absolute value sign makes the cost function non-differentiable, so simple gradient descent is not viable (you would need to implement a method called subgradient descent). Instead, we will use **coordinate descent**: at each iteration, we will fix all weights but weight `i` and find the value of weight `i` that minimizes the objective. That is, we look for
```
argmin_{w[i]} [ SUM[ (prediction - output)^2 ] + lambda*( |w[1]| + ... + |w[k]|) ]
```
where all weights other than `w[i]` are held to be constant. We will optimize one `w[i]` at a time, circling through the weights multiple times.
1. Pick a coordinate `i`
2. Compute `w[i]` that minimizes the cost function `SUM[ (prediction - output)^2 ] + lambda*( |w[1]| + ... + |w[k]|)`
3. Repeat Steps 1 and 2 for all coordinates, multiple times
For this notebook, we use **cyclical coordinate descent with normalized features**, where we cycle through coordinates 0 to (d-1) in order, and assume the features were normalized as discussed above. The formula for optimizing each coordinate is as follows:
```
┌ (ro[i] + lambda/2) if ro[i] < -lambda/2
w[i] = ├ 0 if -lambda/2 <= ro[i] <= lambda/2
└ (ro[i] - lambda/2) if ro[i] > lambda/2
```
where
```
ro[i] = SUM[ [feature_i]*(output - prediction + w[i]*[feature_i]) ].
```
Note that we do not regularize the weight of the constant feature (intercept) `w[0]`, so, for this weight, the update is simply:
```
w[0] = ro[i]
```
## Effect of L1 penalty
Let us consider a simple model with 2 features:
```
simple_features = ['sqft_living', 'bedrooms']
my_output = 'price'
(simple_feature_matrix, output) = get_numpy_data(sales, simple_features, my_output)
```
Don't forget to normalize features:
```
simple_feature_matrix, norms = normalize_features(simple_feature_matrix)
```
We assign some random set of initial weights and inspect the values of `ro[i]`:
```
weights = np.array([1., 4., 1.])
```
Use `predict_output()` to make predictions on this data.
```
prediction =
```
Compute the values of `ro[i]` for each feature in this simple model, using the formula given above, using the formula:
```
ro[i] = SUM[ [feature_i]*(output - prediction + w[i]*[feature_i]) ]
```
*Hint: You can get a Numpy vector for feature_i using:*
```
simple_feature_matrix[:,i]
```
***QUIZ QUESTION***
Recall that, whenever `ro[i]` falls between `-l1_penalty/2` and `l1_penalty/2`, the corresponding weight `w[i]` is sent to zero. Now suppose we were to take one step of coordinate descent on either feature 1 or feature 2. What range of values of `l1_penalty` **would not** set `w[1]` zero, but **would** set `w[2]` to zero, if we were to take a step in that coordinate?
***QUIZ QUESTION***
What range of values of `l1_penalty` would set **both** `w[1]` and `w[2]` to zero, if we were to take a step in that coordinate?
So we can say that `ro[i]` quantifies the significance of the i-th feature: the larger `ro[i]` is, the more likely it is for the i-th feature to be retained.
## Single Coordinate Descent Step
Using the formula above, implement coordinate descent that minimizes the cost function over a single feature i. Note that the intercept (weight 0) is not regularized. The function should accept feature matrix, output, current weights, l1 penalty, and index of feature to optimize over. The function should return new weight for feature i.
```
def lasso_coordinate_descent_step(i, feature_matrix, output, weights, l1_penalty):
# compute prediction
prediction = ...
# compute ro[i] = SUM[ [feature_i]*(output - prediction + weight[i]*[feature_i]) ]
ro_i = ...
if i == 0: # intercept -- do not regularize
new_weight_i = ro_i
elif ro_i < -l1_penalty/2.:
new_weight_i = ...
elif ro_i > l1_penalty/2.:
new_weight_i = ...
else:
new_weight_i = 0.
return new_weight_i
```
To test the function, run the following cell:
```
# should print 0.425558846691
import math
print lasso_coordinate_descent_step(1, np.array([[3./math.sqrt(13),1./math.sqrt(10)],[2./math.sqrt(13),3./math.sqrt(10)]]),
np.array([1., 1.]), np.array([1., 4.]), 0.1)
```
## Cyclical coordinate descent
Now that we have a function that optimizes the cost function over a single coordinate, let us implement cyclical coordinate descent where we optimize coordinates 0, 1, ..., (d-1) in order and repeat.
When do we know to stop? Each time we scan all the coordinates (features) once, we measure the change in weight for each coordinate. If no coordinate changes by more than a specified threshold, we stop.
For each iteration:
1. As you loop over features in order and perform coordinate descent, measure how much each coordinate changes.
2. After the loop, if the maximum change across all coordinates is falls below the tolerance, stop. Otherwise, go back to step 1.
Return weights
```
def lasso_cyclical_coordinate_descent(feature_matrix, output, initial_weights, l1_penalty, tolerance):
```
Using the following parameters, learn the weights on the sales dataset.
```
simple_features = ['sqft_living', 'bedrooms']
my_output = 'price'
initial_weights = np.zeros(3)
l1_penalty = 1e7
tolerance = 1.0
```
First create a normalized version of the feature matrix, `normalized_simple_feature_matrix`
```
(simple_feature_matrix, output) = get_numpy_data(sales, simple_features, my_output)
(normalized_simple_feature_matrix, simple_norms) = normalize_features(simple_feature_matrix) # normalize features
```
Then, run your implementation of LASSO coordinate descent:
```
weights = lasso_cyclical_coordinate_descent(normalized_simple_feature_matrix, output,
initial_weights, l1_penalty, tolerance)
```
***QUIZ QUESTIONS***
1. What is the RSS of the learned model on the normalized dataset?
2. Which features had weight zero at convergence?
# Evaluating LASSO fit with more features
Let us split the sales dataset into training and test sets.
```
train_data,test_data = sales.random_split(.8,seed=0)
```
Let us consider the following set of features.
```
all_features = ['bedrooms',
'bathrooms',
'sqft_living',
'sqft_lot',
'floors',
'waterfront',
'view',
'condition',
'grade',
'sqft_above',
'sqft_basement',
'yr_built',
'yr_renovated']
```
First, create a normalized feature matrix from the TRAINING data with these features. (Make you store the norms for the normalization, since we'll use them later)
First, learn the weights with `l1_penalty=1e7`, on the training data. Initialize weights to all zeros, and set the `tolerance=1`. Call resulting weights `weights1e7`, you will need them later.
***QUIZ QUESTION***
What features had non-zero weight in this case?
Next, learn the weights with `l1_penalty=1e8`, on the training data. Initialize weights to all zeros, and set the `tolerance=1`. Call resulting weights `weights1e8`, you will need them later.
***QUIZ QUESTION***
What features had non-zero weight in this case?
Finally, learn the weights with `l1_penalty=1e4`, on the training data. Initialize weights to all zeros, and set the `tolerance=5e5`. Call resulting weights `weights1e4`, you will need them later. (This case will take quite a bit longer to converge than the others above.)
***QUIZ QUESTION***
What features had non-zero weight in this case?
## Rescaling learned weights
Recall that we normalized our feature matrix, before learning the weights. To use these weights on a test set, we must normalize the test data in the same way.
Alternatively, we can rescale the learned weights to include the normalization, so we never have to worry about normalizing the test data:
In this case, we must scale the resulting weights so that we can make predictions with *original* features:
1. Store the norms of the original features to a vector called `norms`:
```
features, norms = normalize_features(features)
```
2. Run Lasso on the normalized features and obtain a `weights` vector
3. Compute the weights for the original features by performing element-wise division, i.e.
```
weights_normalized = weights / norms
```
Now, we can apply `weights_normalized` to the test data, without normalizing it!
Create a normalized version of each of the weights learned above. (`weights1e4`, `weights1e7`, `weights1e8`).
To check your results, if you call `normalized_weights1e7` the normalized version of `weights1e7`, then:
```
print normalized_weights1e7[3]
```
should return 161.31745624837794.
## Evaluating each of the learned models on the test data
Let's now evaluate the three models on the test data:
```
(test_feature_matrix, test_output) = get_numpy_data(test_data, all_features, 'price')
```
Compute the RSS of each of the three normalized weights on the (unnormalized) `test_feature_matrix`:
***QUIZ QUESTION***
Which model performed best on the test data?
| github_jupyter |
```
import pandas as pd
import warnings
warnings.filterwarnings('ignore')
from sqlalchemy.orm import sessionmaker, Session
from sqlalchemy import create_engine, select, asc, desc, func
import logging
import time
from TickerScrape.models import Security, AssetClass, Country, Currency, Industry, Exchange, Tag
# uri = 'sqlite:///databases/TickerScrape.db'
uri = 'sqlite:////Users/zenman618/Documents/git_packages/VisualStudioGit/TickerScrape/sqlite_files/TickerScrape.db'
# df = pd.read_sql_table("security", uri, schema=None, index_col='id', coerce_float=True, chunksize=None, columns=['ticker, name']) # columns=['ticker, name'], parse_dates='Inception_date',
# df.head()
def init_engine(uri):
logging.basicConfig()
logging.getLogger('sqlalchemy.engine').setLevel(logging.INFO)
engine = create_engine(uri)
return engine
def db_connect(engine):
connection = engine.connect()
logging.info("****_Ticker_Pipeline: database connected****")
return connection
def db_session(engine):
# Session = sessionmaker(bind=engine)
# session = Session()
session = Session(engine)
logging.info("****_Ticker_Pipeline: database connected****")
return session
engine = init_engine(uri)
# %%timeit
with db_connect(engine) as con:
result = con.execute("select ticker from security")
# for row in result:
# print("ticker:", row['ticker'])
df = pd.DataFrame(result)
df.tail()
# %%timeit
with db_session(engine) as session:
result = session.execute("select ticker, name from security")
df = pd.DataFrame(result, columns = ['Ticker', 'Name'])
df.tail()
# %%timeit
with db_session(engine) as session:
result = session.execute("select ticker, name from security")
# print(type(result))
# for dict_row in result.mappings():
# ticker = dict_row['ticker']
# name = dict_row['name']
# print (dict_row)
df = pd.DataFrame(result, columns = ['Ticker', 'Name'])
df.tail()
def all_tickers():
with db_session(engine) as session:
result = session.execute("select ticker, name from security")
df = pd.DataFrame(result, columns = ['Ticker', 'Name'])
df['Label'] = df['Ticker'] + " (" + df['Name'] + ")"
symbols = df['Ticker']
labels = df['Label']
return labels, symbols
labels, symbols = all_tickers()
options=[{'label': x, 'value': y} for x, y in zip(labels, symbols)]
options
df['Label'] = df['Ticker'] + " (" + df['Name'] + ")"
symbols = df['Ticker']
labels = df['Label']
df
sltd_sec = 'AAPL'
# %%timeit
with db_session(engine) as session:
stmt = select(Security.id, Security.ticker, Security.name).where(Security.ticker == sltd_sec).order_by(Security.ticker)
result = session.execute(stmt)
df = pd.DataFrame(result, columns = ['Sec_Id', 'Ticker', 'Name'])
df.tail()
# %%timeit
with db_session(engine) as session:
result = session.execute(select(Security.ticker, Security.name).order_by(Security.ticker))
df = pd.DataFrame(result, columns = ['Ticker', 'Name'])
df.tail()
df.head()
with db_session(engine) as session:
result = session.execute(select(Security.id, Security.ticker).order_by(Security.ticker))
res = result.fetchone()
print(res)
# df = pd.DataFrame(result, columns = ['Sec_Id','Ticker'])
df.tail()
with db_session(engine) as session:
# result = session.execute(select(Security.name, Country.name).join(Security.name).order_by(Security.id, Country.id))
country = session.execute(select(Country).where(Country.name == "United States"))
country = Country(name="United States")
# country = session.query(Country).filter(Country.name=="United States")
# print (country)
print (country)
result = session.execute(select(Security.ticker).where(Security.countries.contains(country)))
df = pd.DataFrame(result)
sel_country = "United States"
with db_session(engine) as session:
stmt = (select(Security.id, Security.ticker, Security.name).where(Security.countries.any(Country.name == sel_country)))
by_country = session.execute(stmt).all()
df = pd.DataFrame(by_country, columns = ['Sec_Id', 'Name', 'Ticker'])
print(df.Ticker.count())
df.tail()
sel_asset = "Stocks"
with db_session(engine) as session:
stmt = (select(Security.id, Security.ticker, Security.name).where(Security.asset_class_id == (select(AssetClass.id).where(AssetClass.name == sel_asset))))
by_asset = session.execute(stmt).all()
df = pd.DataFrame(by_asset, columns = ['Sec_Id', 'Name', 'Ticker'])
print(df.Ticker.count())
df.tail()
sel_industry = "Software"
with db_session(engine) as session:
stmt = (select(Security.id, Security.ticker, Security.name).where(Security.industries.any(Industry.name == sel_industry)))
by_industry = session.execute(stmt).all()
df = pd.DataFrame(by_industry, columns = ['Sec_Id', 'Name', 'Ticker'])
print(df.Ticker.count())
df.tail()
```
| github_jupyter |
# Simple Hello World example for IBM Cloud Functions PyWren
This is a simple Hello World example, showing how to take a function and run it with pywren. First we import the necessary libraries to run our functions.
```
import numpy as np
import os
```
It is possible to use pywren_ibm_cloud inside IBM Watson Studio or Jupyter notebooks in order to run your workloads. You must ensure that the IBM-PyWren package is installed in the environment you are using the notebook. To do so, if you can't install the package manually, we recommend to add the next lines:
```
import sys
try:
import pywren_ibm_cloud as pywren
except:
!{sys.executable} -m pip install pywren-ibm-cloud
import pywren_ibm_cloud as pywren
```
Installation supports PyWren version as an input parameter, for example:
```
# !{sys.executable} -m pip install -U pywren-ibm-cloud==1.3.0
```
Pywren is designed to run any existing python functions you have, in parallel, at scale, on the cloud. So first, we create an example python function.
```
def my_function(x):
return x + 7
```
PyWren needs the configuration to access to IBM Cloud Object Storage and IBM Cloud Functions services. If you don't have the config file in ~/.pywren/config, provide the configuration as a dictionary:
```
config = {'ibm_cf': {'endpoint': '<IBM Cloud Functions Endpoint>',
'namespace': '<NAMESPACE>',
'api_key': '<API KEY>'},
'ibm_cos': {'endpoint': '<IBM COS Endpoint>',
'private_endpoint': '<IBM COS Private Endpoint>',
'api_key' : '<API KEY>'},
'pywren' : {'storage_bucket' : '<IBM COS BUCKET>'}}
```
To start using `pywren`, we first create an executor with the previous config.
```
pw = pywren.ibm_cf_executor(config=config)
```
We can call `my_function(3)` remotely via `call_async`:
```
pw.call_async(my_function, 3)
```
Future is a placeholder for the returned value from applying `my_function` to the number `3`. We can call `result` on it and get the result. Note that this will block until the remote job has completed. Once finished it calls `close` to clean all the unnecessary data stored in COS.
```
print(pw.get_result())
pw.clean()
```
You can apply `my_function` to a list of arguments, and each will be executed remotely at the same time.
```
pw = pywren.ibm_cf_executor(config=config)
pw.map(my_function, range(10))
```
The pywren `get_all_results` function will wait until all of the futures are done and return their results
```
print(pw.get_result())
pw.clean()
```
That's it, we are now familiar how to make use of PyWren for parallelly executing a Python function across many actions in IBM Cloud Functions.
| github_jupyter |
```
import gzip
from collections import defaultdict
import random
def readGz(f):
for l in gzip.open(f):
yield eval(l)
### Rating baseline: compute averages for each user, or return the global average if we've never seen the user before
allRatings = []
userRatings = defaultdict(list)
for l in readGz("train.json.gz"):
user,business = l['reviewerID'],l['itemID']
allRatings.append(l['rating'])
userRatings[user].append(l['rating'])
globalAverage = sum(allRatings) / len(allRatings)
userAverage = {}
for u in userRatings:
userAverage[u] = sum(userRatings[u]) / len(userRatings[u])
predictions = open("predictions_Rating.txt", 'w')
for l in open("pairs_Rating.txt"):
if l.startswith("reviewerID"):
#header
predictions.write(l)
continue
u,i = l.strip().split('-')
if u in userAverage:
predictions.write(u + '-' + i + ',' + str(userAverage[u]) + '\n')
else:
predictions.write(u + '-' + i + ',' + str(globalAverage) + '\n')
predictions.close()
### Would-purchase baseline: just rank which businesses are popular and which are not, and return '1' if a business is among the top-ranked
businessCount = defaultdict(int)
totalPurchases = 0
purchases = defaultdict(set)
purchasesTest = defaultdict(set)
items = set()
count = 0
allData = defaultdict(int)
allPurchases = 0
categoryTrain = defaultdict(set)
itemCategories = defaultdict(str)
for l in readGz("train.json.gz"):
user,business = l['reviewerID'],l['itemID']
allData[business] += 1
allPurchases += 1
if count <= 100000:
businessCount[business] += 1
totalPurchases += 1
count += 1
else:
purchasesTest[l['reviewerID']].add(l['itemID'])
purchases[l['reviewerID']].add(l['itemID'])
categoryTrain[l['reviewerID']].add(l['categoryID'])
itemCategories[l['itemID']] = l['categoryID']
items.add(l['itemID'])
#Non-purchase pairs
negatives = defaultdict(set)
itemsList = list(items)
users = list(purchases.keys())
count = 0
while count < 100000:
cus = random.choice(users)
item = random.choice(itemsList)
if item not in purchases[cus] and item not in negatives[cus]:
negatives[cus].add(item)
count += 1
mostPopular = [(businessCount[x], x) for x in businessCount]
mostPopular.sort()
mostPopular.reverse()
mostPopAll = [(allData[x], x) for x in allData]
mostPopAll.sort()
mostPopAll.reverse()
validation = []
return1 = set()
for u in negatives.keys():
for i in negatives[u]:
validation.append((u,i,0))
for u in purchasesTest.keys():
for i in purchasesTest[u]:
validation.append((u,i,1))
factor = 0.5
count = 0
for ic, i in mostPopular:
count += ic
return1.add(i)
if count > (totalPurchases*factor): break
correct = 0
for p in validation:
if (p[1] in return1):
if (p[2] == 1):
correct += 1
elif (p[2] == 0):
correct += 1
print('Original model validation accuracy is: {}'.format(correct/len(validation)))
popular = 1
factor = 0.48
maxAcc = 0
maxFactor = 0
for k in range(1000):
factor += 0.0001
return1 = set()
count = 0
for ic, i in mostPopular:
count += ic
return1.add(i)
if count > (totalPurchases*factor): break
correct = 0
for p in validation:
if (p[1] in return1):
if (p[2] == 1):
correct += 1
elif (p[2] == 0):
correct += 1
if ((correct/len(validation)) > maxAcc):
maxAcc = (correct/len(validation))
maxFactor = factor
print('Validation accuracy is: {} at factor {}'.format(maxAcc, maxFactor))
popular = 1 #Choose which model to use. (Category model doesn't output accuracy as not required by the question)
#Predicting with the popularity model
if (popular == 1):
return1 = set()
count = 0
for ic, i in mostPopular:
count += ic
return1.add(i)
if count > (totalPurchases*maxFactor): break
predictions = open("predictions_Purchase.txt", 'w')
for l in open("pairs_Purchase.txt"):
if l.startswith("reviewerID"):
#header
predictions.write(l)
continue
u,i = l.strip().split('-')
if i in return1:
predictions.write(u + '-' + i + ",1\n")
else:
predictions.write(u + '-' + i + ",0\n")
predictions.close()
else:
#Predicting with the category model
predictions = open("predictions_Purchase.txt", 'w')
for l in open("pairs_Purchase.txt"):
if l.startswith("reviewerID"):
#header
predictions.write(l)
continue
u,i = l.strip().split('-')
if itemCategories[i] in categoryTrain[u]:
predictions.write(u + '-' + i + ",1\n")
else:
predictions.write(u + '-' + i + ",0\n")
predictions.close()
### Category prediction baseline: Just consider some of the most common words from each category
catDict = {
"Women": 0,
"Men": 1,
"Girls": 2,
"Boys": 3,
"Baby": 4
}
def createList():
return [0,0,0,0,0]
categoryCountsTrain = defaultdict(int)
userCategoryCountsTrain = defaultdict(createList)
categoryCountsVal = defaultdict(int)
userCategoryCountsVal = defaultdict(createList)
count = 0
for l in readGz("train.json.gz"):
if count < 100000:
categoryCountsTrain[l['categoryID']] += 1
userCategoryCountsTrain[l['reviewerID']][l['categoryID']] += 1
count += 1
else:
categoryCountsVal[l['categoryID']] += 1
userCategoryCountsVal[l['reviewerID']][l['categoryID']] += 1
user,business = l['reviewerID'],l['itemID']
allRatings.append(l['rating'])
userRatings[user].append(l['rating'])
favCategoriesTrain = defaultdict(int)
favCategoriesVal = defaultdict(int)
for cus in userCategoryCountsTrain:
userCategories = userCategoryCountsTrain[cus]
favCatVal = max(userCategories)
favCat = userCategories.index(favCatVal)
if (userCategories.count(favCatVal) > 1):
cats = []
for vals in userCategories:
if (vals == favCatVal):
cats.append(userCategories.index(vals))
for i in cats:
maxCat = 0
if (categoryCountsTrain[i] > categoryCountsTrain[maxCat]):
maxCat = i
favCat = maxCat
favCategoriesTrain[cus] = favCat
for cus in userCategoryCountsVal:
userCategories = userCategoryCountsVal[cus]
favCatVal = max(userCategories)
favCat = userCategories.index(favCatVal)
if (userCategories.count(favCatVal) > 1):
cats = []
for vals in userCategories:
if (vals == favCatVal):
cats.append(userCategories.index(vals))
for i in cats:
maxCat = 0
if (categoryCountsVal[i] > categoryCountsVal[maxCat]):
maxCat = i
favCat = maxCat
favCategoriesVal[cus] = favCat
correct = 0
total = 0
for user in favCategoriesVal:
total += 1
if user in favCategoriesTrain.keys():
predicted = favCategoriesTrain[user]
else:
predicted = 0
real = favCategoriesVal[user]
if (predicted == real):
correct += 1
print('Accuracy of category prediction on validation set: {}'.format(correct/total))
predictions = open("predictions_Category.txt", 'w')
predictions.write("reviewerID-reviewHash,category\n")
for l in readGz("test_Category.json.gz"):
cat = catDict['Women'] # If there's no evidence, just choose the most common category in the dataset
words = l['reviewText'].lower()
if 'wife' in words:
cat = catDict['Women']
if 'husband' in words:
cat = catDict['Men']
if 'daughter' in words:
cat = catDict['Girls']
if 'son' in words:
cat = catDict['Boys']
if 'baby' in words:
cat = catDict['Baby']
predictions.write(l['reviewerID'] + '-' + l['reviewHash'] + "," + str(cat) + "\n")
predictions.close()
import operator
import string
translator = str.maketrans('', '', string.punctuation)
def createDict():
return defaultdict(int)
wordCount = defaultdict(int)
categoryWordCount = defaultdict(createDict)
for l in readGz("train.json.gz"):
sentence = l['reviewText'].translate(translator).lower()
words = sentence.split()
for word in words:
wordCount[word] += 1
if 'categoryID' in l.keys():
categoryWordCount[l['categoryID']][word] += 1
topWords = sorted(wordCount.items(), key=operator.itemgetter(1))
topWords.reverse()
topWords = topWords[:500]
total = sum(pair[1] for pair in topWords)
frequencies = defaultdict(float)
for pair in topWords:
frequencies[pair[0]] = pair[1]/total
categoryFrequencies = defaultdict(list)
categoryTotals = defaultdict(int)
for categories in categoryWordCount.keys():
categoryWords = categoryWordCount[categories]
topWords = sorted(categoryWords.items(), key=operator.itemgetter(1))
topWords.reverse()
topWords = topWords[:(500 if len(topWords) > 500 else len(topWords))]
catTotal = sum(pair[1] for pair in topWords)
categoryTotals[categories] = catTotal
wordFrequencies = [(pair[0],(pair[1]/catTotal) - frequencies[pair[0]]) for pair in topWords]
wordFrequencies.sort(key=operator.itemgetter(1))
wordFrequencies.reverse()
categoryFrequencies[categories] = wordFrequencies
for cat in catDict.keys():
printThis = [pair[0] for pair in categoryFrequencies[catDict[cat]][:10]]
print("Words that are more frequent in {} category: {}".format(cat,printThis))
```
| github_jupyter |
```
####################################################
# This is a scratchpad for analysis of intermediate results in the CGMBrush library.
####################################################
# NOTE: Always clear results before checking in a jupyter notebook. This improves diff's.
%load_ext autoreload
%autoreload 2
from cgmbrush.cgmbrush import *
from cgmbrush.plots.plots import *
import matplotlib.colors as colors
from cgmbrush.cosmology import cosmology as cosmo
from cgmbrush.cosmology import halo as halo
import numpy as np
%matplotlib inline
###################################################
# Profile Mask Visualizer
###################################################
# Choose a profile and mass
# Colorbar will be in units of in DM [pc cm^3]
view_mask(PrecipitationProfile(), 2E12)
view_mask(SphericalTophatProfile(rvir_factor=3), 2E12)
view_mask(NFWProfile(), 2E12)
from cgmbrush import *
def compareRadii(mass, z):
print("Mass {:.1e}, z={}".format(mass, z))
a = halo.comoving_rvir(cosmo, mass, z)
b = halo.r200Mz(cosmo, mass, z)
print(' rVir:\t{:.4f} Mpc'.format(a))
print(' r200:\t{:.4f} Mpc'.format(b))
print(' difference:\t{:.1f}%'.format((abs(b-a)) * 100 / b))
masses = [1E10,1E13,1E15]
redshifts = [0,0.5,1,2]
for z in redshifts:
for m in masses:
compareRadii(m, z)
####################################################
# Get a feel for halo masses for the simulation
####################################################
p = BolshoiProvider()
halos = p.get_halos(0)
print('Max: {:.2e}'.format(max(halos['Mvir'])))
print(np.percentile(halos['Mvir'], [50,80,95,99,99.9,99.99,99.999]))
Mvir_avg = []
haloArray, bin_markers = create_halo_array_for_convolution(halos,DEFAULT_MIN_MASS,DEFAULT_MAX_MASS,DEFAULT_MASS_BIN_COUNT)
for j in range(0,len(bin_markers)-1):
Mvir_avg.append((np.mean((haloArray['Mvir'][bin_markers[j]:bin_markers[j+1]])) / cosmo.h, bin_markers[j+1]-bin_markers[j]))
for i in Mvir_avg:
print("{:.1e}, {}".format(i[0], i[1]))
###################################################################################################
#Simplified code that calculates the mass within the analytic profile
#Currently it does this for fire and percipitation but can be easily generalized
#
#The vertical lines are the virial radius
###################################################################################################
redshift = 0
#df = provider.get_halos(redshift)
#df, bin_markers = create_halo_array_for_convolution(df, 10**10, 9*10**15, 30)
massfactor = 1/cosmo.fb*mu*mprot/msun*(Mpc)**3
fp = FireProfile()
percip = PrecipitationProfile()
color = ['black','red', 'orange', 'blue', 'green', 'cyan', 'magenta', 'yellow', 'brown']
nummass=6
i=0
for Mvir_avg in np.logspace(10, 15, nummass):
rvir = halo.comoving_rvir(cosmo, Mvir_avg, redshift)
#rvalsFire, densityFire = fp.get_analytic_profile(Mvir_avg, redshift)
rvalsPercip, densityPercip = percip.get_analytic_profile(Mvir_avg, redshift)
#totalmassanalyticFire = 4.*np.pi*np.sum(rvalsFire[1:]**2*densityFire[1:] *(rvalsFire[1:] - rvalsFire[:-1]))
totalmassanalyticPercip = 4.*np.pi*np.sum(rvalsPercip[1:]**2*densityPercip[1:]*(rvalsPercip[1:] - rvalsPercip[:-1]))
#totalmassanalyticFire *= massfactor
totalmassanalyticPercip *= massfactor
#print(np.log10(Mvir_avg), "massFire in 1e12 = ", totalmassanalyticFire/1e12, " truth = ", Mvir_avg/1e12, " ratio =", totalmassanalyticFire/Mvir_avg)
print(np.log10(Mvir_avg), "massPercip in 1e12 = ", totalmassanalyticPercip/1e12, " truth = ", Mvir_avg/1e12, " ratio =", totalmassanalyticPercip/Mvir_avg)
#plt.loglog(rvalsFire, densityFire, color=color[i%nummass])
plt.loglog(rvalsPercip, densityPercip, '--', color=color[i%nummass])
plt.axvline(rvir, color=color[i%nummass])
i+=1
plt.xlim([.01,3])
# Analyze where mass is going in Precipitation Profile
redshift = 0
massfactor = 1/cosmo.fb*mu*mprot/msun*(Mpc)**3
p = PrecipitationProfile()
p.debug=True
Mvir = np.logspace(np.log10(DEFAULT_MIN_MASS),np.log10(DEFAULT_MAX_MASS), DEFAULT_MASS_BIN_COUNT)
for m in Mvir:
rvir_kpc = 1000*halo.comoving_rvir(cosmo, m, redshift) # comoving radius
print("Mass {:.1e}".format(m))
n1, n2, xi1, xi2, neconstant, XRvir, rmax = p.get_precipitation_params(np.log10(m), rvir_kpc, redshift, True)
# A 3Rvir tophat puts 1/27 of the tophat mass into the 1Rvir.
# So 0.037 + what is reported as being in power law is total within 1 Rvir.
# More details in needed
#print("3Rvir {:.1f}kpc".format(3*rvir_kpc))
#print("rmax {:.1f} kpc".format(rmax))
#print(" n1 = {:.1e} cm^-3; n2 = {:.1e} cm^-3".format(n1,n2))
# Mass Bin Analysis
a = np.logspace(np.log10(DEFAULT_MIN_MASS),np.log10(DEFAULT_MAX_MASS), DEFAULT_MASS_BIN_COUNT)
print(a)
#delta = 1.21164168
# Look at radial values (in kpc) for a DM vs R profile you've already made
provider = BolshoiProvider()
res = 32
date = '2022-06-03'
grid_size = 1024 * res
mass_bin_to_print = 45
DM_vs_R_total = loadArray('precipitation_and_NFW_13.3{}_512_{}_DMvsR_prof'.format(res, date))
DM_vs_R_mask_only = loadArray('precipitation_and_NFW_13.3{}_512_{}_masks'.format(res, date))
cellsize = (provider.Lbox/grid_size)
x_axis = radial_distances_kpc(cellsize, DM_vs_R_total.shape[1])
saveArray('radial_distance_kpc_res32k', x_axis)
with np.printoptions(precision=1, linewidth=1000, threshold=sys.maxsize):
print("The radial distances (in kpc) corresponding to each point int he DM vs R array.")
print(x_axis) #
print("\nTotal DM radial profile (includes mean DM of ~81)")
print(DM_vs_R_total[mass_bin_to_print]) #
print("\nApplied DM radial profile")
print(DM_vs_R_mask_only[mass_bin_to_print]) #
# # Run a config, generate DM vs R, visualize radial profiles for analysis
provider = BolshoiProvider()
series = []
resolution = 8
grid_size = provider.halofieldresolution*resolution
load = True
date = '2022-06-04'
RS_values = RS_array_gen(1,provider.Lbox)
z = RS_values[7]
config = Configuration(PrecipitationProfile(), provider=provider, resolution=resolution, RS_array=[z], den_grid_size=256, datestamp=date)
config.run(load_from_files=load)
config.generate_DM_vs_radius_profile(load_from_files=load)
config.generate_profile_of_masks(load_from_files=load)
series.append((config.DM_vs_R1, config.mask_profiles, 'Precip z={:.1f}'.format(z), 'c', '-'))
config = Configuration(NFWProfile(), provider=provider, resolution=resolution, RS_array=[z], den_grid_size=256, datestamp=date)
config.run(load_from_files=load)
config.generate_DM_vs_radius_profile(load_from_files=load)
config.generate_profile_of_masks(load_from_files=load)
series.append((config.DM_vs_R1, config.mask_profiles, 'NFW', 'b', '-'))
#config = Configuration(SphericalTophatProfile(rvir_factor=3), provider=provider, resolution=resolution, den_grid_size=256, datestamp=date)
#config.run(load_from_files=load)
#config.generate_DM_vs_radius_profile(load_from_files=load)
#config.generate_profile_of_masks(load_from_files=load)
#series.append((config.DM_vs_R1, config.mask_profiles, 'STH 3', 'y', '-'))
vir_rad_ar = config.get_virial_radii()
avg_mass_ar = config.get_halo_masses()
M_chosen = range(0, 60, 1) # plot every 3rd mass bin to see evolution
ymax = (np.arange(0, 60, 1) ** 2) + 40 # Tweak these to scale the y axis as needed
xmin = 5
xmax = 2399
make_DM_vs_Rad_profiles_plots(series, False, True, xmin, xmax, resolution, grid_size, M_chosen, vir_rad_ar, provider, avg_mass_ar, ymax, 'temp_analysis')
```
| github_jupyter |
```
import numpy as np
import tensorflow as tf
import matplotlib.pylab as plt
from modules.spectral_pool import max_pool, l2_loss_images
from modules.frequency_dropout import test_frequency_dropout
from modules.create_images import open_image, downscale_image
from modules.utils import load_cifar10
np.set_printoptions(precision=3, linewidth=200)
% matplotlib inline
% load_ext autoreload
% autoreload 2
images, _ = load_cifar10(1, get_test_data=False)
images.shape
```
### In the cell below, we choose two random images and show how the quality progressively degradees as frequency dropout is applied.
```
batch_size=2
random_selection_indices = np.random.choice(len(images), size=batch_size)
for cutoff in range(16,1,-2):
minibatch_cutoff = tf.cast(tf.constant(cutoff), dtype=tf.float32)
random_selection = images[random_selection_indices]
downsampled_images = np.moveaxis(
test_frequency_dropout(
np.moveaxis(random_selection, 3, 1),
minibatch_cutoff
), 1, 3
)
print('Cutoff = {0}'.format(cutoff))
for i in range(batch_size):
plt.imshow(np.clip(downsampled_images[i],0,1), cmap='gray')
plt.show()
```
### The next cell demonstrates how the random cutoff is applied to all images in a minibatch, but changes from batch to batch.
```
batch_size = 2
minibatch_cutoff = tf.random_uniform([], 2, 12)
for iter_idx in range(3):
random_selection_indices = np.random.choice(len(images), size=batch_size)
random_selection = images[random_selection_indices]
downsampled_images = np.moveaxis(
test_frequency_dropout(
np.moveaxis(random_selection, 3, 1),
minibatch_cutoff
), 1, 3
)
print('Minibatch {0}'.format(iter_idx+1))
for i in range(batch_size):
plt.imshow(random_selection[i], cmap='gray')
plt.show()
plt.imshow(np.clip(downsampled_images[i],0,1), cmap='gray')
plt.show()
```
### max pool test
```
images_pool = max_pool(images, 2)
images_pool.shape
plt.imshow(images_pool[1], cmap='gray')
```
### spectral pool test
```
cutoff_freq = int(32 / (2 * 2))
tf_cutoff_freq = tf.cast(tf.constant(cutoff_freq), tf.float32)
images_spectral_pool = np.clip(np.moveaxis(
test_frequency_dropout(
np.moveaxis(images, 3, 1),
tf_cutoff_freq
), 1, 3
), 0, 1)
images_spectral_pool.shape
plt.imshow(images_spectral_pool[1], cmap='gray')
```
## Iterate and plot
```
images_sample = images[np.random.choice(len(images), size=256)]
# calculate losses for max_pool:
pool_size_mp = [2, 4, 8, 16, 32]
max_pool_errors = []
for s in pool_size_mp:
images_pool = max_pool(images_sample, s)
loss = l2_loss_images(images_sample, images_pool)
max_pool_errors.append(loss)
# calculate losses for spectral_pool:
filter_size_sp = np.arange(16)
spec_pool_errors = []
for s in filter_size_sp:
tf_cutoff_freq = tf.cast(tf.constant(s), tf.float32)
images_sp = np.moveaxis(
test_frequency_dropout(
np.moveaxis(images_sample, 3, 1),
tf_cutoff_freq
), 1, 3
)
loss = l2_loss_images(images_sample, images_sp)
spec_pool_errors.append(loss)
pool_frac_kept = [1/x**2 for x in pool_size_mp]
sp_frac_kept = [(x/16)**2 for x in filter_size_sp]
fig, ax = plt.subplots(1, 1)
ax.semilogy(pool_frac_kept, max_pool_errors, basey=2,
marker='o', linestyle='--', color='r', label='Max Pooling')
ax.semilogy(sp_frac_kept, spec_pool_errors, basey=2,
marker='o', linestyle='--', color='b', label='Spectral Pooling')
ax.legend()
ax.grid(linestyle='--', alpha=0.5)
ax.set_xlabel('Fraction of Parameters Kept')
ax.set_ylabel('Relative Loss')
fig.savefig('../Images/Figure4_Approximation_Loss.png')
```
| github_jupyter |
```
#read file
import pandas as pd
import requests
import json
from config import gkey
filepath = "pollution_us_2000_2016.csv"
us_pollution_data = pd.read_csv(filepath)
us_pollution_data.head()
#Create Year column. which we will use to group the data by year.
Date_data = []
#run a loop on the 'Date Local' Column
for i in us_pollution_data['Date Local']:
#Split each row (which is date in string format) into a list
date = i.split("-")
#The first element of the list will be the year.
year= date[0]
#Append the year to a list of dates
Date_data.append(year)
#Create a year column using Date data
us_pollution_data['Year'] = Date_data
#clean data by dropping rows with null values.
us_pollution_data_clean = us_pollution_data.dropna(how='any')
#Create a collection of columns of interest
Chem_Columns_data = us_pollution_data_clean[['State', 'Date Local', 'Year','NO2 Units', 'NO2 Mean', 'O3 Units', 'O3 Mean','SO2 Units', 'SO2 Mean', 'CO Units', 'CO Mean']]
#reindex data to set indices starting from 0
Chem_Columns_data = Chem_Columns_data.reset_index()
#drop the index column
Chem_Columns_data = Chem_Columns_data.drop(columns=['index'])
Chem_Columns_data.head()
#convert mean values to same units.
#In this case we convert all parts per billion to parts per million.
#To do this we divide all parts per billion observations by 1000
Chem_Columns_data['NO2 Mean'] = Chem_Columns_data['NO2 Mean']/1000
Chem_Columns_data['SO2 Mean'] = Chem_Columns_data['SO2 Mean']/1000
Chem_by_ST_Year = Chem_Columns_data.groupby(['Year', 'State']).mean()
Chem_by_ST_Year
#Create a column for the average of the mean measurements
#note still need to take conversions into account
Mean_us_pol = []
for index, row in Chem_by_ST_Year.iterrows():
avg_pol = (row[0] + row[1] + row[2] + row[3])/4
Mean_us_pol.append(avg_pol)
Chem_by_ST_Year['State Average'] = Mean_us_pol
Chem_by_ST_Year.head()
Chem_by_Year = Chem_Columns_data.groupby(['Year']).mean()
Chem_by_Year.head()
Year_Mean_us_pol = []
for index, row in Chem_by_Year.iterrows():
avg_pol = (row[0] + row[1] + row[2] + row[3])/4
Year_Mean_us_pol.append(avg_pol)
Chem_by_Year['Year Average'] = Year_Mean_us_pol
Chem_by_Year.head()
#save DataFrame as Csv for easy access to Dataframe data
Chem_by_ST_Year.to_csv("../DataFrames/Chem_mean_by_State.csv", header=True)
#Are we sure we have the correct interpretation of a summation of the pullutant means
#save second DataFrame as Csv for easy access to Dataframe data
Chem_by_Year.to_csv("../DataFrames/Chem_mean_by_Year.csv", header=True)
#Create a new Groupby object which will be help for acquiring State pollutant means by year
Group_State_then_Year = Chem_Columns_data.groupby(['State', 'Year']).mean()
Year_Mean_us_pol2 = []
#Create an average column like we did before
for index, row in Group_State_then_Year.iterrows():
avg_pol = (row[0] + row[1] + row[2] + row[3])/4
Year_Mean_us_pol2.append(avg_pol)
Group_State_then_Year['Year Average'] = Year_Mean_us_pol2
Group_State_then_Year.head()
Group_State_then_Year.to_csv("../DataFrames/Group_State_then_Year.csv", header=True)
AQI_data = us_pollution_data_clean[['State Code', 'County Code', 'State', 'County', 'City','Year','NO2 AQI', 'O3 AQI', 'SO2 AQI', 'CO AQI']]
AQI_data.head()
#plot the average state values for a year versus number of cases in states for that year. or
#plot the average us_val for a year versus number of case for that year
#might have to merge on state or produce an overall average mean (of year) for the US based on an average of all states (for a year)
target_city = "Phoenix, Arizona"
# Build the endpoint URL
target_url = ('https://maps.googleapis.com/maps/api/geocode/json?'
'address={0}&key={1}').format(target_city, gkey)
geo_data = requests.get(target_url).json()
lat = geo_data["results"][0]["geometry"]["location"]["lat"]
lng = geo_data["results"][0]["geometry"]["location"]["lng"]
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import frame as fr
import algo_result as alg_res
import os
from scipy.fft import fft, fftn, fftfreq, fftshift
import xgboost as xgb
from xgboost import plot_importance, plot_tree
import graphviz
%run algo_process.ipynb
def peak_search_dopp(spectrum,Lo_thresh,Hi_thresh,peak_relevance):
counter = 0
peak_strength = np.max(spectrum)
peak_bin = np.argmax(spectrum)
doppler_details = {"peak_count":0,"total_harmonic_relative_strength":0,"peak_bin":[],"harmonic_relative_strength":[],"max_peak_strength":peak_strength}
harmonic_strength = 0
fbc = 2
fbl2 = fbc - 2
fbl1 = fbc - 1
fbr1 = fbc + 1
fbr2 = fbc + 2
for idx in range(len(spectrum)):
if fbr2 >= len(spectrum):
continue
pvl2 = spectrum[fbl2]
pvl1 = spectrum[fbl1]
pvc = spectrum[fbc]
pvr1 = spectrum[fbr1]
pvr2 = spectrum[fbr2]
if pvl2+peak_relevance < pvl1 and pvr1 > pvr2+peak_relevance and pvc > Lo_thresh and pvc <= Hi_thresh and pvc >= pvl1 and pvc >= pvr1 and fbc != peak_bin:
harmonic_strength += pvc[0]
counter += 1
doppler_details["peak_bin"].append(fbc)
doppler_details["harmonic_relative_strength"].append(pvc[0]/peak_strength)
fbc += 1
fbl2 = fbc - 2
fbl1 = fbc - 1
fbr1 = fbc + 1
fbr2 = fbc + 2
doppler_details["peak_count"] = counter
doppler_details["total_harmonic_relative_strength"] = harmonic_strength/peak_strength
return doppler_details
def doppler_details_extraction(spectrum,T_Lo_thr,T_Mi_thr,T_Hi_thr):
doppler_details = peak_search_dopp(spectrum,0,np.Inf,0)
Hi_details={"total_harmonic_relative_strength":0,"peak_count":0,"peak_bin":[],"harmonic_relative_strength":[]}
Mi_details={"total_harmonic_relative_strength":0,"peak_count":0,"peak_bin":[],"harmonic_relative_strength":[]}
Lo_details={"total_harmonic_relative_strength":0,"peak_count":0,"peak_bin":[],"harmonic_relative_strength":[]}
for peak_idx in range(doppler_details["peak_count"]):
if doppler_details["harmonic_relative_strength"][peak_idx] > 1/T_Lo_thr and doppler_details["harmonic_relative_strength"][peak_idx] <= 1/T_Mi_thr:
Lo_details["peak_count"] += 1
Lo_details["peak_bin"].append(doppler_details["peak_bin"][peak_idx])
Lo_details["harmonic_relative_strength"].append(doppler_details["harmonic_relative_strength"][peak_idx])
elif doppler_details["harmonic_relative_strength"][peak_idx] > 1/T_Mi_thr and doppler_details["harmonic_relative_strength"][peak_idx] <= 1/T_Hi_thr:
Mi_details["peak_count"] += 1
Mi_details["peak_bin"].append(doppler_details["peak_bin"][peak_idx])
Mi_details["harmonic_relative_strength"].append(doppler_details["harmonic_relative_strength"][peak_idx])
elif doppler_details["harmonic_relative_strength"][peak_idx] > 1/T_Hi_thr:
Hi_details["peak_count"] += 1
Hi_details["peak_bin"].append(doppler_details["peak_bin"][peak_idx])
Hi_details["harmonic_relative_strength"].append(doppler_details["harmonic_relative_strength"][peak_idx])
Lo_details["total_harmonic_relative_strength"] = sum(Lo_details["harmonic_relative_strength"])
Mi_details["total_harmonic_relative_strength"] = sum(Mi_details["harmonic_relative_strength"])
Hi_details["total_harmonic_relative_strength"] = sum(Hi_details["harmonic_relative_strength"])
return Hi_details,Mi_details,Lo_details,doppler_details
SAMPLES_PER_CHIRP = 64
CHIRPS_PER_FRAME = 128
T = 300e-6
SPEED_OF_LIGHT = 3e8
START_FREQUENCY = 24.025e9
B = 200e6
PULSE_REPETITION_INTERVAL = 500e-6
SAMPLE_PERIOD = T/SAMPLES_PER_CHIRP
SAMPLE_FREQUENCY = 1/SAMPLE_PERIOD
LAMBDA = SPEED_OF_LIGHT/START_FREQUENCY
RANGE_PAD = 256
DOPPLER_PAD = 512
ANTENNA_SPACING = 6.22e-3
PEAK_THRESHOLD = 0.005 # normalized FFT absolute minimum strength
PEAK_SLICE = 2 #meters around target
PEAK_WIDTH = 1 #integer
PEAK_RELEVANCE = 0 #minimum distance between pvl1,pvr1 and pvc
SEARCH_ZONE = 25 #split spectrum in slices of SEARCH_ZONE meters to find a single peak
MIN_DIST = 5 #minimum distance for detection
ANGLE_CALIBRATION = -150
ANGLE_PRECISION = 1
ANTENNA_NUMBER = 1
FRAME_REP_INTERVAL = 0.2
classifier = xgb.XGBClassifier()
classifier.load_model('../statistics_data_processing/code/Trained_stuff/boresight_diagonal_azimuth_model.model')
asd = pd.read_csv("../statistics_data_processing/code/Trained_stuff/Boresight_Diagonal_azimuth.csv",delimiter='\t')
d = asd.to_dict("split")
###Get dictionary with performance for each number of feature
my_dictionary_list = []
for row_idx in range(len(d['data'])):
for col_idx in range(len(d['columns'])):
if d['columns'][col_idx] == 'support':
split_bools = d['data'][row_idx][col_idx] = d['data'][row_idx][col_idx].replace("\n", "").replace(" "," ").replace("[","").replace("]","").split(" ")
d['data'][row_idx][col_idx] = []
for elem in split_bools:
if elem == 'True':
d['data'][row_idx][col_idx].append(True)
elif elem == 'False':
d['data'][row_idx][col_idx].append(False)
best_params= dict(zip(d["columns"],d["data"][row_idx]))
best_params[d['columns'][col_idx]] = d['data']
my_dictionary_list.append(best_params)
###Get best performance
max_score = 0
support = []
feature_names = []
for elem in my_dictionary_list:
if elem['score'] > max_score:
max_score = elem['score']
support = elem['support']
feature_names = elem['features'].replace("'","").replace('[','').replace(']','').replace('\n','').split(" ")
###Get feature importance
importance_type='weight'
fscores = classifier.get_booster().get_score(importance_type=importance_type)
feat_importances = []
for ft, score in fscores.items():
feat_importances.append({'Feature': ft, 'Importance': score, 'Name': feature_names[int(ft[1:])]})
feat_importances = pd.DataFrame(feat_importances)
feat_importances = feat_importances.sort_values(
by='Importance', ascending=False).reset_index(drop=True)
print(feat_importances)
###Sort labels with feature importance
feat_labels = []
for elem in feat_importances.values:
feat_labels.append(elem[2])
feat_labels = np.flip(feat_labels)
fig,ax = plt.subplots(1,1,figsize=(10,10))
###Plot importance
importance_plot=plot_importance(classifier,ax=ax,importance_type=importance_type,show_values=False)
importance_plot.set_title(f"Feature importance (by {importance_type})")
importance_plot.set_yticklabels(feat_labels)
normalization_factors = pd.read_csv("../statistics_data_processing/code/Trained_stuff/boresight_diagonal_azimuth_norm_factors.csv",delimiter='\t').values
norm_mean = normalization_factors[0]
norm_scale = normalization_factors[1]
#fig,ax = plt.subplots(1,1,figsize=(10,20))
#plot_tree(classifier,ax=ax)
#plt.savefig('Tree',format='png')
directory = '../../data/'
folders = []
folders.append('Dataset_2')
folders.append('Dataset_1')
for folder in folders:
###Calibration Data
calibration_data = pd.read_csv(directory + folder+ '/environment_1.txt', sep='\t', header=None)
calibration = calibration_data.select_dtypes(include = ['float']).values
CALIBRATION_FRAME_NUMBER = len(calibration)//(SAMPLES_PER_CHIRP*CHIRPS_PER_FRAME)
calibration_frames = []
###Create dataset
for frame in range(CALIBRATION_FRAME_NUMBER):
calibration_frames.append(fr.Frame(calibration[frame*CHIRPS_PER_FRAME*SAMPLES_PER_CHIRP:(frame+1)*CHIRPS_PER_FRAME*SAMPLES_PER_CHIRP,:],\
SAMPLES_PER_CHIRP, CHIRPS_PER_FRAME, ANTENNA_NUMBER, T))
average_calib_chirp = np.zeros((SAMPLES_PER_CHIRP,ANTENNA_NUMBER),dtype=complex)
for frame in range(1):
for chirp in range(CHIRPS_PER_FRAME):
average_calib_chirp += calibration_frames[frame].get_chirp(chirp)
average_calib_chirp /= CHIRPS_PER_FRAME
###Target Data
data_directory = os.fsencode(directory + folder + '/')
for file in os.listdir(data_directory):
filename = os.fsdecode(file)
if filename.find('environment') == -1 and filename.endswith('.txt') and filename.find('azimuth') != -1:
actual_filename = filename
path = os.path.join(os.fsdecode(data_directory), filename)
print(path)
data = pd.read_csv(path, sep='\t', header=None)
data.columns = ["idx","I_RX1","Q_RX1"]
recording = data.select_dtypes(include = ['float']).values
FRAME_NUMBER = len(data)//(SAMPLES_PER_CHIRP*CHIRPS_PER_FRAME)
Hu_bi_frame = []
###Create dataset
for frame in range(FRAME_NUMBER):
Hu_bi_frame.append(fr.Frame(recording[frame*CHIRPS_PER_FRAME*SAMPLES_PER_CHIRP:(frame+1)*CHIRPS_PER_FRAME*SAMPLES_PER_CHIRP,:],\
SAMPLES_PER_CHIRP, CHIRPS_PER_FRAME, ANTENNA_NUMBER, T))
###Calibrate frames
calibrate = True
if calibrate:
for frame in range(FRAME_NUMBER):
Hu_bi_frame[frame].calibrate(average_calib_chirp)
xf = np.arange(0,RANGE_PAD)
range_bin = xf*T*SPEED_OF_LIGHT/(2*B)/(T/SAMPLES_PER_CHIRP)/RANGE_PAD
range2bin = 1/(T/SAMPLES_PER_CHIRP)/RANGE_PAD*T*SPEED_OF_LIGHT/(2*B)
vel_bin = fftshift(fftfreq(DOPPLER_PAD,PULSE_REPETITION_INTERVAL))*SPEED_OF_LIGHT/(2*START_FREQUENCY)
Target_observations = []
Range_spectrum_history = []
Unfiltered_spectrum_history = []
Target_info_list = []
Target_presence_list = []
###Process frames
for frame in range(FRAME_NUMBER):
data_out,target_info, MTI_out = algo_process(Hu_bi_frame[frame],RANGE_PAD,CHIRPS_PER_FRAME,DOPPLER_PAD,PEAK_THRESHOLD,PEAK_SLICE,PEAK_WIDTH,PEAK_RELEVANCE,SEARCH_ZONE,ANGLE_CALIBRATION,ANGLE_PRECISION,round(LAMBDA/ANTENNA_SPACING),range2bin)
Range_spectrum_history.append(MTI_out)
Unfiltered_spectrum_history.append(abs(data_out[:,0,0]))
if(target_info.num_targets > 0 and target_info.location[0] < SEARCH_ZONE and target_info.location[0] > MIN_DIST):
Target_info_list.append(target_info)
Target_presence_list.append(1)
target_doppler_spec = target_info.doppler_spectrum[:,0]
Target_observations.append(np.append(np.roll(target_doppler_spec,DOPPLER_PAD//2-np.argmax(target_doppler_spec)), target_info.location[0]))
else:
Target_presence_list.append(0)
df = pd.DataFrame(Target_observations).T
output_path = "../statistics_data_processing/data_NN/"
output_filename = actual_filename.split('.')[0]
extension = '_statistics'
df.T.to_csv(output_path+output_filename+'_'+folder+extension+'.txt', sep='\t',index=False, header=False)
predictions_converted = []
for pred in predictions:
if pred == 0 or pred == 3 or pred == 6:
predictions_converted.append(0)
elif pred == 1 or pred == 4 or pred == 7:
predictions_converted.append(1)
elif pred == 2 or pred == 5 or pred == 8:
predictions_converted.append(2)
elif pred == -1:
predictions_converted.append(-1)
predictions = predictions_converted
print(predictions)
import matplotlib as mpl
from matplotlib.ticker import FormatStrFormatter
%matplotlib inline
### PLOT DOPPLER VS TIME
fig,[ax,cax] = plt.subplots(1,2, gridspec_kw={"width_ratios":[10,1],'wspace':0.01}, figsize=[20,10])
ax.clear()
cmap = mpl.cm.get_cmap('turbo')
norm = mpl.colors.Normalize(vmin=0, vmax=np.max(target_doppler_spec))
cb1 = mpl.colorbar.ColorbarBase(cax, cmap=cmap,
norm=norm,
orientation='vertical')
levels = np.linspace(0.0, np.max(target_doppler_spec), 100)
ax.pcolormesh(np.arange(0,counter)*FRAME_REP_INTERVAL,vel_bin,target_doppler_spec, cmap=cmap, norm=norm, shading='nearest')
ax.set_xlabel('seconds',fontsize=30)
ax.set_ylabel('velocity', fontsize=30)
_ = ax.set_xticks(np.arange(0,FRAME_NUMBER,20)*FRAME_REP_INTERVAL)
_ = ax.set_xticklabels((np.arange(0,FRAME_NUMBER,20)*FRAME_REP_INTERVAL).astype('int'),fontsize=20)
#_ = ax.set_yticks(vel_bin[np.arange(0,DOPPLER_PAD,64)])
_ = ax.set_yticklabels(ax.get_yticks(),fontsize=20)
plt.savefig('Thesis_figures/'+'all_doppler_driving_diagonal_1.png')
%matplotlib inline
rolled_doppler = np.zeros(target_doppler_spec.shape)
for frame in range(FRAME_NUMBER):
if max(target_doppler_spec[:,frame]) > 0:
max_idx = np.argmax(target_doppler_spec[:,frame])
#round(sum([(i)*target_doppler_spec[i,frame] for i in range(DOPPLER_PAD)])/sum(target_doppler_spec[:,frame]))
rolled_doppler[:,frame] = np.roll(target_doppler_spec[:,frame],(DOPPLER_PAD//2 - max_idx))
fig,[ax,cax] = plt.subplots(1,2, gridspec_kw={"width_ratios":[10,1],'wspace':0.01}, figsize=[20,10])
ax.clear()
cmap = mpl.cm.get_cmap('turbo')
norm = mpl.colors.Normalize(vmin=0, vmax=np.max(rolled_doppler))
cb1 = mpl.colorbar.ColorbarBase(cax, cmap=cmap,
norm=norm,
orientation='vertical')
levels = np.linspace(0.0, np.max(rolled_doppler), 100)
ax.pcolormesh(np.arange(0,FRAME_NUMBER)*FRAME_REP_INTERVAL,vel_bin,rolled_doppler, cmap=cmap, norm=norm, shading='nearest')
#ax.contourf(np.arange(0,FRAME_NUMBER),vel_bin,rolled_doppler, levels, cmap=cmap, norm=norm)
ax.set_xlabel('seconds',fontsize=30)
ax.set_ylabel('velocity', fontsize=30)
_ = ax.set_xticks(np.arange(0,FRAME_NUMBER,20)*FRAME_REP_INTERVAL)
_ = ax.set_xticklabels((np.arange(0,FRAME_NUMBER,20)*FRAME_REP_INTERVAL).astype('int'),fontsize=20)
#_ = ax.set_yticks(vel_bin[np.arange(0,DOPPLER_PAD,64)])
_ = ax.set_yticklabels(ax.get_yticks(),fontsize=20)
#plt.savefig('Thesis_figures/'+'centered_all_doppler_biking_boresight_1.png')
column_plots = 1
row_plots = 8
fig,ax = plt.subplots(row_plots,column_plots,figsize=[12,5])
from_second_number = 100#round(32/FRAME_REP_INTERVAL)
for col in range(column_plots):
for row in range(row_plots):
data = target_doppler_spec[:,row + row_plots*col + from_second_number].reshape((1,-1))[0,:]
ax[row].plot(vel_bin,data)
plt.show()
%matplotlib widget
fig,ax = plt.subplots(3,1, figsize=[12,5])
from_second_number = 105#round(32/FRAME_REP_INTERVAL)+5
#for frame in np.arange(from_second_number,from_second_number+1):
ax[0].plot(np.sum(target_doppler_spec[:,from_second_number:from_second_number+1],axis=1))
if(sum(target_doppler_spec[:,from_second_number:from_second_number+1][:] > 0)):
actual_dopp = target_doppler_spec[:,from_second_number:from_second_number+1]
weighted_avg_1 = sum([i*actual_dopp[i] for i in range(length)])/sum(actual_dopp[:])
ax[0].plot(weighted_avg_1,np.max(actual_dopp),'ro')
low_tresh = np.max(actual_dopp)/T_Lo_thr*np.ones(length)
mid_tresh = np.max(actual_dopp)/T_Mi_thr*np.ones(length)
high_tresh = np.max(actual_dopp)/T_Hi_thr*np.ones(length)
ax[0].plot(low_tresh)
ax[0].plot(mid_tresh)
ax[0].plot(high_tresh)
ax[0].set_ylim((0,high_tresh[0]*11/10))
actual_dopp = np.roll(actual_dopp,DOPPLER_PAD//2 - round(weighted_avg_1[0]))
ax[1].plot(actual_dopp)
weighted_avg_1 = sum([i*actual_dopp[i] for i in range(length)])/sum(actual_dopp[:])
ax[1].plot(DOPPLER_PAD//2,np.max(actual_dopp),'ro')
ax[1].plot(low_tresh)
ax[1].plot(mid_tresh)
ax[1].plot(high_tresh)
weighted_std_1 = ([(i-DOPPLER_PAD//2)**2*actual_dopp[i] for i in np.arange(DOPPLER_PAD//4,3*DOPPLER_PAD//4)])/sum(tmp_roll[DOPPLER_PAD//4:3*DOPPLER_PAD//4])/(DOPPLER_PAD//4)**2
ax[2].plot(np.arange(0,len(weighted_std_1))+DOPPLER_PAD//4,weighted_std_1,'bo')
print(np.sqrt(sum(weighted_std_1)))
print(round(weighted_avg_1[0]))
%matplotlib inline
fig,ax = plt.subplots(1,1,figsize=(20,10))
ax.plot(np.sum(target_doppler_spec[:,from_second_number:from_second_number+1],axis=1))
if(sum(target_doppler_spec[:,from_second_number:from_second_number+1][:] > 0)):
actual_dopp = target_doppler_spec[:,from_second_number:from_second_number+1]
weighted_avg_1 = sum([i*actual_dopp[i] for i in range(length)])/sum(actual_dopp[:])
ax.plot(weighted_avg_1,np.max(actual_dopp),'ro')
low_tresh = np.max(actual_dopp)/T_Lo_thr*np.ones(length)
mid_tresh = np.max(actual_dopp)/T_Mi_thr*np.ones(length)
high_tresh = np.max(actual_dopp)/T_Hi_thr*np.ones(length)
ax.plot(low_tresh)
ax.plot(mid_tresh)
ax.plot(high_tresh)
#ax.set_ylim((0,high_tresh[0]*11/10))
def peak_search_details(spectrum,Lo_thresh,Hi_thresh,peak_relevance):
counter = 0
peak_power = np.max(spectrum)
peak_bin = np.argmax(spectrum)
harmonic_power = 0
fbc = 2
fbl2 = fbc - 2
fbl1 = fbc - 1
fbr1 = fbc + 1
fbr2 = fbc + 2
peak_info = {"peak_bin":[],"peak_strength":[],"max_peak_strength":peak_power}
for idx in range(len(spectrum)):
if fbr2 >= len(spectrum):
continue
pvl2 = spectrum[fbl2]
pvl1 = spectrum[fbl1]
pvc = spectrum[fbc]
pvr1 = spectrum[fbr1]
pvr2 = spectrum[fbr2]
if pvl2+peak_relevance < pvl1 and pvr1 > pvr2+peak_relevance and pvc > Lo_thresh and pvc < Hi_thresh and pvc >= pvl1 and pvc >= pvr1 and fbc != peak_bin:
peak_info["peak_bin"].append(fbc)
peak_info["peak_strength"].append(pvc/peak_power)
fbc += 1
fbl2 = fbc - 2
fbl1 = fbc - 1
fbr1 = fbc + 1
fbr2 = fbc + 2
return peak_info
frame_doppler_peaks_dict = []
for target in Target_info_list:
max_bin = np.argmax(target.doppler_spectrum[:,0])
frame_doppler_peaks_dict.append(peak_search_details(np.roll(target.doppler_spectrum[:,0]*target.location[0]**2,DOPPLER_PAD//2 - max_bin),0,np.Inf,0))
%matplotlib widget
fig,ax = plt.subplots(1,1, figsize=[12,5])
all_doppler_peaks = np.array([])
for frame in frame_doppler_peaks_dict:
all_doppler_peaks = np.append(all_doppler_peaks,np.array(frame["peak_strength"]))
n,bins=np.histogram(all_doppler_peaks,5000)
cumulative_n = [0]
for idx in range(len(n)):
cumulative_n.append(n[idx] + cumulative_n[idx])
ax.plot(bins,cumulative_n/cumulative_n[-1])
ax.set_xlim((0,0.8))
peak_presence_frequency = np.zeros(len(bins)-1)
for frame_peaks in frame_doppler_peaks_dict:
for bin_idx in range(len(bins)-1):
for peak in frame_peaks['peak_strength']:
if bins[bin_idx] <= peak and bins[bin_idx+1] >= peak:
peak_presence_frequency[bin_idx] += 1
break
fig,ax = plt.subplots(1,1, figsize=[12,5])
ax.plot(bins[:-1],peak_presence_frequency/sum(Target_presence_list))
fig,ax = plt.subplots(1,1, figsize=[12,5])
ax.plot(bins[:-1],peak_presence_frequency/sum(Target_presence_list)**2*n)
ax.set_xlim((0,0.04))
fig,[ax,cax] = plt.subplots(1,2, gridspec_kw={"width_ratios":[10,1],'wspace':0.01}, figsize=[20,10])
ax.clear()
cmap = mpl.cm.get_cmap('turbo')
norm = mpl.colors.Normalize(vmin=0, vmax=np.max(Range_spectrum_history))
cb1 = mpl.colorbar.ColorbarBase(cax, cmap=cmap,
norm=norm,
orientation='vertical')
levels = np.linspace(0.0, np.max(Range_spectrum_history), 100)
ax.pcolormesh(np.arange(0,FRAME_NUMBER)*FRAME_REP_INTERVAL,range_bin,np.array(Range_spectrum_history).T, cmap=cmap, norm=norm, shading='nearest')
ax.set_xlabel('seconds',fontsize=30)
ax.set_ylabel('range', fontsize=30)
_ = ax.set_xticks(np.arange(0,FRAME_NUMBER,20)*FRAME_REP_INTERVAL)
_ = ax.set_xticklabels((np.arange(0,FRAME_NUMBER,20)*FRAME_REP_INTERVAL).astype('int'),fontsize=20)
#_ = ax.set_yticks(vel_bin[np.arange(0,DOPPLER_PAD,64)])
_ = ax.set_yticklabels(ax.get_yticks(),fontsize=20)
#plt.savefig('Thesis_figures/'+'MTI_range_biking_boresight_1.png')
fig,[ax,cax] = plt.subplots(1,2, gridspec_kw={"width_ratios":[10,1],'wspace':0.01}, figsize=[20,10])
ax.clear()
cmap = mpl.cm.get_cmap('turbo')
norm = mpl.colors.Normalize(vmin=0, vmax=np.max(Unfiltered_spectrum_history))
cb1 = mpl.colorbar.ColorbarBase(cax, cmap=cmap,
norm=norm,
orientation='vertical')
levels = np.linspace(0.0, np.max(Unfiltered_spectrum_history), 100)
ax.pcolormesh(np.arange(0,FRAME_NUMBER)*FRAME_REP_INTERVAL,range_bin,np.array(Unfiltered_spectrum_history).T, cmap=cmap, norm=norm, shading='nearest')
ax.set_xlabel('seconds',fontsize=30)
ax.set_ylabel('range', fontsize=30)
_ = ax.set_xticks(np.arange(0,FRAME_NUMBER,20)*FRAME_REP_INTERVAL)
_ = ax.set_xticklabels((np.arange(0,FRAME_NUMBER,20)*FRAME_REP_INTERVAL).astype('int'),fontsize=20)
#_ = ax.set_yticks(vel_bin[np.arange(0,DOPPLER_PAD,64)])
_ = ax.set_yticklabels(ax.get_yticks(),fontsize=20)
#plt.savefig('Thesis_figures/'+'Unfiltered_range_biking_boresight_1.png')
%matplotlib widget
column_plots = 1
row_plots = 2
fig,ax = plt.subplots(row_plots,column_plots,figsize=[12,5])
from_second_number = 0
separator = np.zeros(len(Target_info_list[0].print[:,0,0]))
separator[-1] = 0.05
separator_list = []
for i in range(len(Target_info_list[0].print[0,:,0])):
separator_list.append(separator)
separator_list = np.array(separator_list).reshape((1,-1))[0,:]
for col in range(column_plots):
for row in range(row_plots):
data = Target_info_list[row + row_plots*col + from_second_number].print[:,:,0].T.reshape((1,-1))[0,:]
ax[row].set_ylim(0,0.2)
ax[row].plot(data)
ax[row].plot(separator_list[:],'ro',markersize=0.5)
plt.show()
peak_avg = sum(np.max(Target_info_list[from_second_number].print[:,:,0],axis=0))/CHIRPS_PER_FRAME
column_plots = 1
row_plots = 8
fig,ax = plt.subplots(row_plots,column_plots,figsize=[20,10])
from_second_number = 9
separator = np.zeros(len(Target_info_list[0].print[:,0,0]))
separator[-1] = 0.05
separator_list = []
for i in range(len(Target_info_list[0].print[0,:,0])):
separator_list.append(separator)
separator_list = np.array(separator_list).reshape((1,-1))[0,:]
for col in range(column_plots):
for row in range(row_plots):
data = np.max(Target_info_list[row + row_plots*col + from_second_number].print[:,:,0],axis=0).T.reshape((1,-1))[0,:]
ax[row].hist(data,bins=100,range=(0,0.2),density=False)
plt.show()
print(Target_info_list[from_second_number].location[0])
peak_collection = []
for target in Target_info_list:
peak_strength = np.max(target.print[:,:,0],axis=0)*target.location[0]**2
peak_strength_mean = np.sum(peak_strength)/CHIRPS_PER_FRAME
peak_collection.append(peak_strength-peak_strength_mean)
peak_collection = np.array(peak_collection).reshape((1,-1))
_=plt.hist(peak_collection[0,:],bins=100)
from matplotlib.animation import FuncAnimation, writers
fig,[ax1,ax2] = plt.subplots(2,1,figsize=(25, 25))
classes = ['Pedestrian','Cyclist','Car']
ax1.title.set_text('Range')
ax1.title.set_fontsize(40)
ax2.title.set_fontsize(40)
ax2.title.set_text('Doppler')
ax1.set_xlim(range_bin[0], range_bin[-1])
ax1.set_ylim(0,np.max(Range_spectrum_history)*8/10)
ax2.set_xlim(vel_bin[0], vel_bin[-1])
ax1.tick_params(labelsize=30)
ax2.tick_params(labelsize=30)
#ax2.set_ylim(0,np.max(target_doppler_spec)*0.5)
ax1.axvline(MIN_DIST, lw=3, linestyle='--', color='black')
ax1.axvline(SEARCH_ZONE, lw=3, linestyle='--', color='black', label='Search Region')
ax1.plot(range_bin,np.ones(len(range_bin))*PEAK_THRESHOLD, lw=3, linestyle='dotted', color='gray', label='Detection Threshold')
# intialize two line objects (one in each axes)
line1, = ax1.plot([], [], lw=4, color='r', label='Filtered Range FFT Spectrum')
line2, = ax2.plot([], [], lw=4, color='r', label='Doppler FFT Spectrum')
line11, = ax1.plot([], [], 'D', color='black', markersize=15,label='Target location')
line21, = ax2.plot([],[], lw=3, linestyle='dashdot',color='limegreen', label='Feature Extractor')
line22, = ax2.plot([],[], lw=3, linestyle='dashdot',color='limegreen')
line23, = ax2.plot([],[], lw=3, linestyle='dashdot',color='limegreen')
line24, = ax2.plot([],[], lw=3, linestyle='dashdot',color='royalblue', label='Feature Extractor')
line25, = ax2.plot([],[], lw=3, linestyle='dashdot',color='royalblue')
line = [line1, line2, line21,line22,line23,line24,line25, line11]
ax1.legend(fontsize=20,loc=1)
ax2.legend(fontsize=20,loc=1)
#plt.xlabel(r'meters')
#plt.ylabel(r'fft magnitude')
text_axis = ax2.text(.2, -.2, 'Class:'+'No prediction available', style='italic',fontsize=60,
bbox={'facecolor': 'bisque', 'alpha': 0.5, 'pad': 10},visible=True,transform=ax2.transAxes)
#
# animation function
def animate(i):
line[0].set_data(range_bin, Range_spectrum_history[i])
if predictions[i] == -1:
text_axis.set_text('Class:'+'No prediction available')
elif predictions[i] == 0:
text_axis.set_text('Class:'+classes[predictions[i]] + ' at ' + str(round(range_bin[np.argmax(Range_spectrum_history[i])],2))+ ' m')
elif predictions[i] == 1:
text_axis.set_text('Class:'+classes[predictions[i]] + ' at ' + str(round(range_bin[np.argmax(Range_spectrum_history[i])],2))+ ' m')
elif predictions[i] == 2:
text_axis.set_text('Class:'+classes[predictions[i]] + ' at ' + str(round(range_bin[np.argmax(Range_spectrum_history[i])],2))+ ' m')
if(Target_presence_list[i]>0):
#ax1.set_ylim(0,np.max(Range_spectrum_history[i]*11/10))
ax2.set_ylim(0,np.max(target_doppler_spec[:,i])*11/10)
line[1].set_data(vel_bin,target_doppler_spec[:,i])
line[2].set_data(vel_bin,np.ones(DOPPLER_PAD)*np.max(target_doppler_spec[:,i])/T_Lo_thr)
line[3].set_data(vel_bin,np.ones(DOPPLER_PAD)*np.max(target_doppler_spec[:,i])/T_Mi_thr)
line[4].set_data(vel_bin,np.ones(DOPPLER_PAD)*np.max(target_doppler_spec[:,i])/T_Hi_thr)
line[5].set_data(vel_bin,np.roll(np.array(bi_quadratic)*np.max(target_doppler_spec[:,i]),np.argmax(target_doppler_spec[:,i])-DOPPLER_PAD//2))
line[6].set_data(vel_bin,np.roll(np.array(hor_quadratic)*np.max(target_doppler_spec[:,i]),np.argmax(target_doppler_spec[:,i])-DOPPLER_PAD//2))
line[7].set_data(range_bin[np.argmax(Range_spectrum_history[i])],np.max(Range_spectrum_history[i]))
else:
for i in np.arange(1,8):
line[i].set_data([],[])
return line
anim = FuncAnimation(fig, animate, frames=FRAME_NUMBER)
Writer = writers['ffmpeg']
writer = Writer(fps=1/(FRAME_REP_INTERVAL), metadata={'artist':'Me'}, bitrate=3600)
anim.save('../../videos/'+folder+'_'+actual_filename + '_complete_model' +'.mp4',writer)
print(len(Range_spectrum_history[0]))
print(Target_presence_list)
```
| github_jupyter |
```
%matplotlib inline
# Importing standard Qiskit libraries
import qiskit
from qiskit import QuantumCircuit, execute, Aer, IBMQ
from qiskit.compiler import transpile, assemble
from qiskit.tools.jupyter import *
from qiskit.visualization import *
from ibm_quantum_widgets import *
from qiskit import QuantumRegister
from qiskit import ClassicalRegister
from qiskit.circuit import Parameter
# Loading your IBM Q account(s)
provider = IBMQ.load_account()
import numpy as np
numOfInputs = 2
numOfOutputs = 2
# Function: Creates Controlled-NOT Gates (CNOT) and Barriers for aesthetics
# Parameters: 'circuit', 'channels'
# 'circuit' : quantum circuit
# 'channels' : number of active states in circuit (generally same as 'numIn')
def CNOT_and_Barriers(circuit, channels):
for channel in range(channels-1):
circuit.cx(channel, channel+1)
for count in range(channel):
circuit.barrier(count)
def clusterStateQC(numIn, numOut):
# Define Cluster State Quantum Circuit (abbreviated: cs_qc)
# numIn = number of initial quantum states
# numOut = number of output states of cs_qc (likely same as numIn)
cs_qc = QuantumCircuit(numIn, numOut)
#cs_qc.add_register(quReg)
# Hadamard Gates for all incoming feature inputs to achieve superposition of states
#for num in range(numIn):
# cs_qc.h(quReg[num])
cs_qc.h(range(numIn))
# Place Controlled-NOT Gates (CNOT) for entanglement and barriers for aesthetics
CNOT_and_Barriers(cs_qc, numIn)
# Place Pauli-Y Gates (Y) for rotation of states about y-axis
#for num in range(numIn):
# cs_qc.y(quReg[num])
cs_qc.y(range(numIn))
# Place Controlled-NOT Gates (CNOT) for entanglement and barriers for aesthetics
CNOT_and_Barriers(cs_qc, numIn)
cs_qc.barrier()
return cs_qc
def qConvLayer1(shift, prm1, prm2, prm3, prm4, inputs):
qConv = QuantumCircuit(inputs, inputs)
for i in range(inputs):
index = int(i)
qConv.u(prm1, prm2, prm3, index)
for i in range(inputs-1):
if (i%2==0):
qConv.cx(i,i+1)
return qConv
def qConvLayer2(shift, prm1, prm2, prm3, prm4, inputs):
qConv = QuantumCircuit(inputs, inputs)
for i in range(0+shift,inputs-1,2):
index = int(i)
if (i%2==shift):
qConv.cu(prm1, prm2, prm3, prm4, control_qubit=index, target_qubit=index+1)
return qConv
def qPoolLayer(inputs, outputs, shift=0):
qPool = QuantumCircuit(inputs, outputs)
if shift == 0:
for i in range(inputs):
qPool.measure(i, i)
return qPool
for i in range(inputs):
if (i%2==shift):
qPool.measure(i, i)
return qPool
class MyQuantumCircuit:
"""
This class provides a simple interface for interaction
with the quantum circuit
"""
def __init__(self, n_qubits, backend, shots):
# --- Circuit definition ---
self._circuit = QuantumCircuit(n_qubits)
self.theta = Parameter('θ')
self.phi = Parameter('Φ')
self.lam = Parameter('λ')
self.gamma = Parameter('γ')
nIn = n_qubits
nOut = nIn
shift = 0
self._circuit += clusterStateQC(numIn=nIn, numOut=nOut)
self._circuit += qConvLayer1(0, self.theta, self.phi, self.lam, self.gamma, nIn)
self._circuit += qConvLayer2(0, self.theta, self.phi, self.lam, self.gamma, nIn)
#self._circuit += qConvLayer2(1, self.theta, self.phi, self.lam, self.gamma, nIn)
self._circuit.barrier()
self._circuit += qPoolLayer(nIn, nOut, 1)
self._circuit.barrier()
self._circuit += qConvLayer2(0, self.theta, self.phi, self.lam, self.gamma, nIn)
self._circuit.barrier()
self._circuit += qPoolLayer(nIn, nOut, 0)
# ---------------------------
self.backend = backend
self.shots = shots
def run(self, val_range):
t_qc = transpile(self._circuit,
self.backend)
qobj = assemble(t_qc,
shots=self.shots,
parameter_binds = [{self.theta: val1, self.phi: val2, self.lam: val3, self.gamma: val4}
for (val1, val2, val3, val4) in [np.linspace(0,np.pi/2,4,dtype=float)]])
#[np.linspace(0,np.pi,4,dtype=int)]
#[{self.theta: val, self.phi: val, self.lam: val, self.gamma: val}
# for val in val_range]
job = self.backend.run(qobj)
result = job.result().get_counts()
counts = np.array(list(result.values()))
states = np.array(list(result.keys())).astype(float)
# Compute probabilities for each state
probabilities = counts / self.shots
# Get state expectation
expectation = np.sum(states * probabilities)
return np.array([expectation])
simulator = qiskit.Aer.get_backend('qasm_simulator')
circuit = MyQuantumCircuit(2, simulator, 100)
print('Expected value for rotation pi {}'.format(circuit.run([np.linspace(0,np.pi / 2,4,dtype=float)])[0]))
circuit._circuit.draw()
print(circuit._circuit.parameters)
import matplotlib.pyplot as plt
import torch
from torch.autograd import Function
from torchvision import datasets, transforms
import torch.optim as optim
import torch.nn as nn
import torch.nn.functional as F
class HybridFunction(Function):
""" Hybrid quantum - classical function definition """
@staticmethod
def forward(ctx, input, quantum_circuit, shift):
""" Forward pass computation """
ctx.shift = shift
ctx.quantum_circuit = quantum_circuit
expectation_z = ctx.quantum_circuit.run(input.tolist())
#print(len(input.tolist()))
#print(input)
result = torch.tensor([expectation_z])
ctx.save_for_backward(input, result)
return result
@staticmethod
def backward(ctx, grad_output):
""" Backward pass computation """
input, expectation_z = ctx.saved_tensors
#print(input.shape)
input_list = np.array(input.tolist())
shift_right = input_list + np.ones(input_list.shape) * ctx.shift
shift_left = input_list - np.ones(input_list.shape) * ctx.shift
gradients = []
for i in range(len(input_list)):
expectation_right = ctx.quantum_circuit.run(shift_right[i])
expectation_left = ctx.quantum_circuit.run(shift_left[i])
gradient = torch.tensor([expectation_right]) - torch.tensor([expectation_left])
gradients.append(gradient)
gradients = np.array([gradients]).T
return torch.tensor([gradients]).float() * grad_output.float(), None, None
class Hybrid(nn.Module):
""" Hybrid quantum - classical layer definition """
def __init__(self, backend, shots, shift):
super(Hybrid, self).__init__()
self.quantum_circuit = MyQuantumCircuit(numOfInputs, backend, shots)
self.shift = shift
def forward(self, input):
#print(self.parameters)
return HybridFunction.apply(input, self.quantum_circuit, self.shift)
# Concentrating on the first 100 samples
n_samples = 20
X_train = datasets.MNIST(root='./data', train=True, download=True, transform=transforms.Compose([transforms.ToTensor()]))
# Leaving only labels 0 and 1
idx = np.append(np.where(X_train.targets == 0)[0][:n_samples],
np.where(X_train.targets == 1)[0][:n_samples])
X_train.data = X_train.data[idx]
X_train.targets = X_train.targets[idx]
train_loader = torch.utils.data.DataLoader(X_train, batch_size=1, shuffle=True)
n_samples_show = 6
data_iter = iter(train_loader)
fig, axes = plt.subplots(nrows=1, ncols=n_samples_show, figsize=(10, 3))
while n_samples_show > 0:
images, targets = data_iter.__next__()
axes[n_samples_show - 1].imshow(images[0].numpy().squeeze(), cmap='gray')
axes[n_samples_show - 1].set_xticks([])
axes[n_samples_show - 1].set_yticks([])
axes[n_samples_show - 1].set_title("Labeled: {}".format(targets.item()))
n_samples_show -= 1
n_samples = 50
X_test = datasets.MNIST(root='./data', train=False, download=True,
transform=transforms.Compose([transforms.ToTensor()]))
idx = np.append(np.where(X_test.targets == 0)[0][:n_samples],
np.where(X_test.targets == 1)[0][:n_samples])
X_test.data = X_test.data[idx]
X_test.targets = X_test.targets[idx]
test_loader = torch.utils.data.DataLoader(X_test, batch_size=1, shuffle=True)
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 6, kernel_size=5)
self.conv2 = nn.Conv2d(6, 16, kernel_size=5)
self.dropout = nn.Dropout2d()
self.fc1 = nn.Linear(256, 64)
self.fc2 = nn.Linear(64, 1)
self.hybrid = Hybrid(qiskit.Aer.get_backend('qasm_simulator'), 100, np.pi)
def forward(self, x):
x = F.relu(self.conv1(x))
x = F.max_pool2d(x, 2)
x = F.relu(self.conv2(x))
x = F.max_pool2d(x, 2)
x = self.dropout(x)
x = x.view(1, -1)
x = F.relu(self.fc1(x))
x = self.fc2(x)
x = self.hybrid(x)
return torch.cat((x, 1 - x), -1)
model = Net()
optimizer = optim.Adam(model.parameters(), lr=0.001)
#from qiskit.aqua.components.optimizers import AQGD
#optimizer = AQGD
print(model.parameters)
loss_func = nn.NLLLoss()
epochs = 20
loss_list = []
print(model)
qc = QuantumCircuit()
model.train()
for epoch in range(epochs):
total_loss = []
for batch_idx, (data, target) in enumerate(train_loader):
optimizer.zero_grad()
#optimizer.optimize(4, )
# Forward pass
output = model(data)
# Calculating loss
loss = loss_func(output, target)
# Backward pass
loss.backward()
# Optimize the weights
optimizer.step()
total_loss.append(loss.item())
loss_list.append(sum(total_loss)/len(total_loss))
print('Training [{:.0f}%]\tLoss: {:.4f}'.format(100. * (epoch + 1) / epochs, loss_list[-1]))
plt.plot(loss_list)
plt.title('Hybrid NN Training Convergence')
plt.xlabel('Training Iterations')
plt.ylabel('Neg Log Likelihood Loss')
model.eval()
with torch.no_grad():
correct = 0
for batch_idx, (data, target) in enumerate(test_loader):
output = model(data)
pred = output.argmax(dim=1, keepdim=True)
correct += pred.eq(target.view_as(pred)).sum().item()
loss = loss_func(output, target)
total_loss.append(loss.item())
print('Performance on test data:\n\tLoss: {:.4f}\n\tAccuracy: {:.1f}%'.format(
sum(total_loss) / len(total_loss),
correct / len(test_loader) * 100)
)
n_samples_show = 6
count = 0
fig, axes = plt.subplots(nrows=1, ncols=n_samples_show, figsize=(10, 3))
model.eval()
with torch.no_grad():
for batch_idx, (data, target) in enumerate(test_loader):
if count == n_samples_show:
break
output = model(data)
pred = output.argmax(dim=1, keepdim=True)
axes[count].imshow(data[0].numpy().squeeze(), cmap='gray')
axes[count].set_xticks([])
axes[count].set_yticks([])
axes[count].set_title('Predicted {}'.format(pred.item()))
count += 1
```
| github_jupyter |
# How to use OpenNMT-py as a Library
The example notebook (available [here](https://github.com/OpenNMT/OpenNMT-py/blob/master/docs/source/examples/Library.ipynb)) should be able to run as a standalone execution, provided `onmt` is in the path (installed via `pip` for instance).
Some parts may not be 100% 'library-friendly' but it's mostly workable.
### Import a few modules and functions that will be necessary
```
import yaml
import torch
import torch.nn as nn
from argparse import Namespace
from collections import defaultdict, Counter
import onmt
from onmt.inputters.inputter import _load_vocab, _build_fields_vocab, get_fields, IterOnDevice
from onmt.inputters.corpus import ParallelCorpus
from onmt.inputters.dynamic_iterator import DynamicDatasetIter
from onmt.translate import GNMTGlobalScorer, Translator, TranslationBuilder
from onmt.utils.misc import set_random_seed
```
### Enable logging
```
# enable logging
from onmt.utils.logging import init_logger, logger
init_logger()
```
### Set random seed
```
is_cuda = torch.cuda.is_available()
set_random_seed(1111, is_cuda)
```
### Retrieve data
To make a proper example, we will need some data, as well as some vocabulary(ies).
Let's take the same data as in the [quickstart](https://opennmt.net/OpenNMT-py/quickstart.html):
```
!wget https://s3.amazonaws.com/opennmt-trainingdata/toy-ende.tar.gz
!tar xf toy-ende.tar.gz
ls toy-ende
```
### Prepare data and vocab
As for any use case of OpenNMT-py 2.0, we can start by creating a simple YAML configuration with our datasets. This is the easiest way to build the proper `opts` `Namespace` that will be used to create the vocabulary(ies).
```
yaml_config = """
## Where the vocab(s) will be written
save_data: toy-ende/run/example
# Corpus opts:
data:
corpus:
path_src: toy-ende/src-train.txt
path_tgt: toy-ende/tgt-train.txt
transforms: []
weight: 1
valid:
path_src: toy-ende/src-val.txt
path_tgt: toy-ende/tgt-val.txt
transforms: []
"""
config = yaml.safe_load(yaml_config)
with open("toy-ende/config.yaml", "w") as f:
f.write(yaml_config)
from onmt.utils.parse import ArgumentParser
parser = ArgumentParser(description='build_vocab.py')
from onmt.opts import dynamic_prepare_opts
dynamic_prepare_opts(parser, build_vocab_only=True)
base_args = (["-config", "toy-ende/config.yaml", "-n_sample", "10000"])
opts, unknown = parser.parse_known_args(base_args)
opts
from onmt.bin.build_vocab import build_vocab_main
build_vocab_main(opts)
ls toy-ende/run
```
We just created our source and target vocabularies, respectively `toy-ende/run/example.vocab.src` and `toy-ende/run/example.vocab.tgt`.
### Build fields
We can build the fields from the text files that were just created.
```
src_vocab_path = "toy-ende/run/example.vocab.src"
tgt_vocab_path = "toy-ende/run/example.vocab.tgt"
# initialize the frequency counter
counters = defaultdict(Counter)
# load source vocab
_src_vocab, _src_vocab_size = _load_vocab(
src_vocab_path,
'src',
counters)
# load target vocab
_tgt_vocab, _tgt_vocab_size = _load_vocab(
tgt_vocab_path,
'tgt',
counters)
# initialize fields
src_nfeats, tgt_nfeats = 0, 0 # do not support word features for now
fields = get_fields(
'text', src_nfeats, tgt_nfeats)
fields
# build fields vocab
share_vocab = False
vocab_size_multiple = 1
src_vocab_size = 30000
tgt_vocab_size = 30000
src_words_min_frequency = 1
tgt_words_min_frequency = 1
vocab_fields = _build_fields_vocab(
fields, counters, 'text', share_vocab,
vocab_size_multiple,
src_vocab_size, src_words_min_frequency,
tgt_vocab_size, tgt_words_min_frequency)
```
An alternative way of creating these fields is to run `onmt_train` without actually training, to just output the necessary files.
### Prepare for training: model and optimizer creation
Let's get a few fields/vocab related variables to simplify the model creation a bit:
```
src_text_field = vocab_fields["src"].base_field
src_vocab = src_text_field.vocab
src_padding = src_vocab.stoi[src_text_field.pad_token]
tgt_text_field = vocab_fields['tgt'].base_field
tgt_vocab = tgt_text_field.vocab
tgt_padding = tgt_vocab.stoi[tgt_text_field.pad_token]
```
Next we specify the core model itself. Here we will build a small model with an encoder and an attention based input feeding decoder. Both models will be RNNs and the encoder will be bidirectional
```
emb_size = 100
rnn_size = 500
# Specify the core model.
encoder_embeddings = onmt.modules.Embeddings(emb_size, len(src_vocab),
word_padding_idx=src_padding)
encoder = onmt.encoders.RNNEncoder(hidden_size=rnn_size, num_layers=1,
rnn_type="LSTM", bidirectional=True,
embeddings=encoder_embeddings)
decoder_embeddings = onmt.modules.Embeddings(emb_size, len(tgt_vocab),
word_padding_idx=tgt_padding)
decoder = onmt.decoders.decoder.InputFeedRNNDecoder(
hidden_size=rnn_size, num_layers=1, bidirectional_encoder=True,
rnn_type="LSTM", embeddings=decoder_embeddings)
device = "cuda" if torch.cuda.is_available() else "cpu"
model = onmt.models.model.NMTModel(encoder, decoder)
model.to(device)
# Specify the tgt word generator and loss computation module
model.generator = nn.Sequential(
nn.Linear(rnn_size, len(tgt_vocab)),
nn.LogSoftmax(dim=-1)).to(device)
loss = onmt.utils.loss.NMTLossCompute(
criterion=nn.NLLLoss(ignore_index=tgt_padding, reduction="sum"),
generator=model.generator)
```
Now we set up the optimizer. This could be a core torch optim class, or our wrapper which handles learning rate updates and gradient normalization automatically.
```
lr = 1
torch_optimizer = torch.optim.SGD(model.parameters(), lr=lr)
optim = onmt.utils.optimizers.Optimizer(
torch_optimizer, learning_rate=lr, max_grad_norm=2)
```
### Create the training and validation data iterators
Now we need to create the dynamic dataset iterator.
This is not very 'library-friendly' for now because of the way the `DynamicDatasetIter` constructor is defined. It may evolve in the future.
```
src_train = "toy-ende/src-train.txt"
tgt_train = "toy-ende/tgt-train.txt"
src_val = "toy-ende/src-val.txt"
tgt_val = "toy-ende/tgt-val.txt"
# build the ParallelCorpus
corpus = ParallelCorpus("corpus", src_train, tgt_train)
valid = ParallelCorpus("valid", src_val, tgt_val)
# build the training iterator
train_iter = DynamicDatasetIter(
corpora={"corpus": corpus},
corpora_info={"corpus": {"weight": 1}},
transforms={},
fields=vocab_fields,
is_train=True,
batch_type="tokens",
batch_size=4096,
batch_size_multiple=1,
data_type="text")
# make sure the iteration happens on GPU 0 (-1 for CPU, N for GPU N)
train_iter = iter(IterOnDevice(train_iter, 0))
# build the validation iterator
valid_iter = DynamicDatasetIter(
corpora={"valid": valid},
corpora_info={"valid": {"weight": 1}},
transforms={},
fields=vocab_fields,
is_train=False,
batch_type="sents",
batch_size=8,
batch_size_multiple=1,
data_type="text")
valid_iter = IterOnDevice(valid_iter, 0)
```
### Training
Finally we train.
```
report_manager = onmt.utils.ReportMgr(
report_every=50, start_time=None, tensorboard_writer=None)
trainer = onmt.Trainer(model=model,
train_loss=loss,
valid_loss=loss,
optim=optim,
report_manager=report_manager,
dropout=[0.1])
trainer.train(train_iter=train_iter,
train_steps=1000,
valid_iter=valid_iter,
valid_steps=500)
```
### Translate
For translation, we can build a "traditional" (as opposed to dynamic) dataset for now.
```
src_data = {"reader": onmt.inputters.str2reader["text"](), "data": src_val}
tgt_data = {"reader": onmt.inputters.str2reader["text"](), "data": tgt_val}
_readers, _data = onmt.inputters.Dataset.config(
[('src', src_data), ('tgt', tgt_data)])
dataset = onmt.inputters.Dataset(
vocab_fields, readers=_readers, data=_data,
sort_key=onmt.inputters.str2sortkey["text"])
data_iter = onmt.inputters.OrderedIterator(
dataset=dataset,
device="cuda",
batch_size=10,
train=False,
sort=False,
sort_within_batch=True,
shuffle=False
)
src_reader = onmt.inputters.str2reader["text"]
tgt_reader = onmt.inputters.str2reader["text"]
scorer = GNMTGlobalScorer(alpha=0.7,
beta=0.,
length_penalty="avg",
coverage_penalty="none")
gpu = 0 if torch.cuda.is_available() else -1
translator = Translator(model=model,
fields=vocab_fields,
src_reader=src_reader,
tgt_reader=tgt_reader,
global_scorer=scorer,
gpu=gpu)
builder = onmt.translate.TranslationBuilder(data=dataset,
fields=vocab_fields)
```
**Note**: translations will be very poor, because of the very low quantity of data, the absence of proper tokenization, and the brevity of the training.
```
for batch in data_iter:
trans_batch = translator.translate_batch(
batch=batch, src_vocabs=[src_vocab],
attn_debug=False)
translations = builder.from_batch(trans_batch)
for trans in translations:
print(trans.log(0))
break
```
| github_jupyter |
# Pessimistic Neighbourhood Aggregation for States in Reinforcement Learning
*Author: Maleakhi Agung Wijaya
Supervisors: Marcus Hutter, Sultan Javed Majeed
Date Created: 21/12/2017*
```
import random
import math
import sys
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from IPython.display import display, clear_output
# Set grid color for seaborn
sns.set(style="whitegrid")
```
## Mountain Car Environment
**Mountain Car** is a standard testing domain in Reinforcement Learning, in which an under-powered car must drive up a steep hill. Since gravity is stronger than the car's engine, even at full throttle, the car cannot simply accelerate up the steep slope. The car is situated in a valley and must learn to leverage potential energy by driving up the opposite hill before the car is able to make it to the goal at the top of the rightmost hill.
**Technical Details**
- *State:* feature vectors consisting of velocity and position represented by an array [velocity, position]
- *Reward:* -1 for every step taken, 0 for achieving the goal
- *Action:* (left, neutral, right) represented by (-1, 0, 1)
- *Initial state:* velocity = 0.0, position = -0.5 represented by [0.0, -0.5]
- *Terminal state:* position >= 0.6
- *Boundaries:* velocity = (-0.07, 0.07), position = (-1.2, 0.6)
- *Update function:* velocity = velocity + (Action) \* 0.001 + cos(3\*Position) * (-0.0025), position = position + velocity
```
class MountainCarEnvironment:
"""
Description: Environment for Mountain Car problem, adapted from Sutton and Barto's Introduction to Reinforcement Learning.
Author: Maleakhi Agung Wijaya
"""
VELOCITY_BOUNDARIES = (-0.07, 0.07)
POSITION_BOUNDARIES = (-1.2, 0.6)
INITIAL_VELOCITY = 0.0
INITIAL_POSITION = -0.5
REWARD_STEP = -1
REWARD_TERMINAL = 0
# Constructor for MountainCarEnvironment
# Input: agent for the MountainCarEnvironment
# Output: MountainCarEnvironment object
def __init__(self, car):
self.car = car
self.reset()
# Compute next state (feature)
# Output: [new velocity, new position]
def nextState(self, action):
# Get current state (velocity, position) and the action chosen by the agent
velocity = self.car.state[0]
position = self.car.state[1]
# Calculate the new velocity and new position
velocity += action * 0.001 + math.cos(3*position) * (-0.0025)
# Consider boundary for velocity
if (velocity < MountainCarEnvironment.VELOCITY_BOUNDARIES[0]):
velocity = MountainCarEnvironment.VELOCITY_BOUNDARIES[0]
elif (velocity > MountainCarEnvironment.VELOCITY_BOUNDARIES[1]):
velocity = MountainCarEnvironment.VELOCITY_BOUNDARIES[1]
position += velocity
# Consider boundary for position
if (position < MountainCarEnvironment.POSITION_BOUNDARIES[0]):
position = MountainCarEnvironment.POSITION_BOUNDARIES[0]
velocity = 0
elif (position > MountainCarEnvironment.POSITION_BOUNDARIES[1]):
position = MountainCarEnvironment.POSITION_BOUNDARIES[1]
new_state = [velocity, position]
return(new_state)
# Reset to the initial state
def reset(self):
self.car.state[0] = MountainCarEnvironment.INITIAL_VELOCITY
self.car.state[1] = MountainCarEnvironment.INITIAL_POSITION
# Give reward for each of the chosen action, depending on what the next state that the agent end up in
# Output: terminal state = 0, non-terminal state = -1
def calculateReward(self):
# Get current position of the agent
position = self.car.state[1]
# Determine the reward given
if (position >= MountainCarEnvironment.POSITION_BOUNDARIES[1]):
return(MountainCarEnvironment.REWARD_TERMINAL)
else:
return(MountainCarEnvironment.REWARD_STEP)
```
## KNN-TD Agent
**kNN-TD** combines the concept of *K-Nearest Neighbours* and *TD-Learning* to learn and evaluate Q values in both continuous and discrete state space RL problems. This method is especially useful in continuous states RL problems as the number of (state, action) pairs is very large and thus impossible to store and learn this information. By choosing a particular k-values and decided some initial points over continuous states, one can estimate Q values based on calculated the weighted average of Q values of the k-nearest neighbours for the state that the agent are currently in and use that values to decide the next move using some decision methods (i.e. UCB or epsilon-greedy). As for the learning process, one can update all of the k-nearest neighbours that contribute for the Q calculation.
**Algorithm:**
1. Cover the whole state space by some initial Q(s,a) pairs, possibly scatter it uniformly across the whole state space and give an initial value of 0/ -1
2. When an agent in a particular state, get the feature vectors representing the state and possible actions from the state
3. For each possible action from the state, calculate Q(s,a) pairs by taking the expected value from previous Q values based on k-nearest neighbours of a particular action.
*Steps for k-nearest neighbours:*
- Standardise every feature in the feature vectors to (-1, 1) or other ranges to make sure that one feature scale not dominate the distance calculation (i.e. if position ranges between (-50, 50) and velocity (-0.7, 0.7) position will dominate distance calculation).
- Calculate the distance between current state and all of other points with the same action using distance formula (i.e. Euclidean distance) and store the k-nearest neighbours to knn vector, and it's distance (for weight) in weight vector
- Determine the probability p(x) for the expected value by using weight calculation (i.e. weight = 1/distance). To calculate weight, one can use other formula as long as that formula gives more weight to closer point. To calculate p(x) just divide individual weight with sum of all weights to get probability
- Estimate the Q(s,a) pairs using expectation formula from kNN previous Q values
4. Using epsilon greedy/ UCB/ other decision methods to choose the next move
5. Observe the reward and update the Q values for all of the neighbours on knn vector using SARSA or Q Learning. (on the code below, I use Q Learning)
6. Repeat step 2-5
```
class KNNAgent:
"""
Description: Mountain Car problem agent based on kNN-TD(0) algorithm
Author: Maleakhi Agung Wijaya
"""
INITIAL_VELOCITY = 0.0
INITIAL_POSITION = -0.5
INITIAL_VALUE = -1
ACTIONS = [-1, 0, 1]
GAMMA = 0.995
EPSILON = 0.05
INDEX_DISTANCE = 0
INDEX_ORIGINAL = 1
INDEX_WEIGHT = 2
REWARD_STEP = -1
REWARD_TERMINAL = 0
# Constructor
# Input: size of the storage for previous Q values, parameters for how many neighbours which the agent will choose
def __init__(self, size, k):
self.state = [KNNAgent.INITIAL_VELOCITY, KNNAgent.INITIAL_POSITION]
self.q_storage = []
self.k = k # fixed number of nearest neighbours that we will used
self.alpha = 1 # will be decaying and change later
# Storage of the k nearest neighbour (data) and weight (inverse of distance) for a particular step
self.knn = []
self.weight = []
# Initialise the storage with random point
for i in range(size):
initial_action = random.randint(-1, 1)
initial_state = [random.uniform(-0.07, 0.07), random.uniform(-1.2, 0.6)]
# Each data on the array will consist of state, action pair + value
data = {"state": initial_state, "value": KNNAgent.INITIAL_VALUE, "action": initial_action}
self.q_storage.append(data)
# Find all index for a given value
# Input: value, list to search
# Output: list of all index where you find that value on the list
def findAllIndex(self, value, list_value):
indices = []
for i in range(len(list_value)):
if (value == list_value[i]):
indices.append(i)
return indices
# Standardise feature vector given
# Input: feature vector to be standardised
# Output: standardised feature vector
def standardiseState(self, state):
standardised_state = []
# The number is taken from VELOCITY_BOUNDARIES and POSITION_BOUNDARIES using normal standardisation formula
standardised_velocity = 2 * ((state[0]+0.07) / (0.07+0.07)) - 1
standardised_position = 2 * ((state[1]+1.2) / (0.6+1.2)) - 1
standardised_state.append(standardised_velocity)
standardised_state.append(standardised_position)
return(standardised_state)
# Calculate Euclidean distance between 2 vectors
# Input: 2 feature vectors
# Output: distance between them
def calculateDistance(self, vector1, vector2):
return(math.sqrt((vector1[0]-vector2[0])**2 + (vector1[1]-vector2[1])**2))
# Calculate total weight
# Input: list of weights
# Output: total weight
def calculateTotalWeight(self, weight_list):
total_weight = 0
for i in range(len(weight_list)):
total_weight += weight_list[i][KNNAgent.INDEX_WEIGHT]
return(total_weight)
# Apply the kNN algorithm for feature vector and store the data point on the neighbours array
# Input: feature vector of current state, actions array consisting of all possible actions, list that will store knn data and weights data
# Output: vector containing the value of taking each action (left, neutral, right)
def kNNTD(self, state, actions, knn_list, weight_list):
approximate_action = []
# Get the standardised version of state
standardised_state = self.standardiseState(state)
# Loop through every element in the storage array and only calculate for particular action
for action in actions:
temp = [] # array consisting of tuple (distance, original index, weight) for each point in the q_storage
for i in range(len(self.q_storage)):
data = self.q_storage[i]
# Only want to calculate the nearest neighbour state which has the same action
if (data["action"] == action):
vector_2 = data["state"]
standardised_vector_2 = self.standardiseState(vector_2)
distance = self.calculateDistance(standardised_state, standardised_vector_2)
index = i
weight = 1 / (1+distance**2) # weight formula
# Create the tuple and append that to temp
temp.append(tuple((distance, index, weight)))
else:
continue
# After we finish looping through all of the point and calculating the standardise distance,
# Sort the tuple based on the distance and only take k of it and append that to the neighbours array
# We also need to calculate the total weight to make it into valid probability that we can compute it's expectation
sorted_temp = sorted(temp, key=lambda x: x[0])
for i in range(self.k):
try:
weight_list.append(sorted_temp[i])
knn_list.append(self.q_storage[sorted_temp[i][KNNAgent.INDEX_ORIGINAL]])
except IndexError:
sys.exit(0)
# Calculate the expected value of the action and append it to the approximate_action array
expected_value = 0
total_weight = self.calculateTotalWeight(weight_list[(action+1)*self.k:(action+1)*self.k + self.k])
for i in range((action+1)*self.k, (action+1)*self.k + self.k):
weight = weight_list[i][KNNAgent.INDEX_WEIGHT]
probability = weight / total_weight
expected_value += probability * knn_list[i]["value"]
approximate_action.append(expected_value)
return(approximate_action)
# Select which action to choose, whether left, neutral, or right (using epsilon greedy)
# Output: -1 (left), 0 (neutral), 1 (right)
def selectAction(self):
# First call the knn-td algorithm to determine the value of each Q(s,a) pairs
action_value = self.kNNTD(self.state, KNNAgent.ACTIONS, self.knn, self.weight)
# Use the epsilon-greedy method to choose value
random_number = random.uniform(0.0, 1.0)
if (random_number <= KNNAgent.EPSILON):
action_chosen = random.randint(-1, 1)
else:
# Return the action with highest Q(s,a)
possible_index = self.findAllIndex(max(action_value), action_value)
action_chosen = possible_index[random.randrange(len(possible_index))] - 1
# Only store chosen data in the knn and weight list
# Clearance step
chosen_knn = []
chosen_weight = []
for i in range(self.k*(action_chosen+1), self.k*(action_chosen+1) + self.k):
chosen_knn.append(self.knn[i])
chosen_weight.append(self.weight[i])
self.knn = chosen_knn
self.weight = chosen_weight
return action_chosen
# Calculate TD target based on Q Learning/ SARSAMAX
# Input: Immediate reward based on what the environment gave
# Output: TD target based on off policy Q learning
def calculateTDTarget(self, immediate_reward):
# Consider condition on the final state, return 0 immediately
if (immediate_reward == KNNAgent.REWARD_TERMINAL):
return(immediate_reward)
knn_prime = []
weight_prime = []
action_value = self.kNNTD(self.state, KNNAgent.ACTIONS, knn_prime, weight_prime)
return(immediate_reward + KNNAgent.GAMMA*max(action_value))
# Q learning TD updates on every neighbours on the kNN based on the contribution that are calculated using probability weight
# Input: Immediate reward based on what the environment gave
def TDUpdate(self, immediate_reward, alpha):
self.alpha = alpha
# First, calculate the TD target
td_target = self.calculateTDTarget(immediate_reward)
# Iterate every kNN and update using Q learning method based on the weighting
total_weight = self.calculateTotalWeight(self.weight)
for i in range(len(self.weight)):
index = self.weight[i][KNNAgent.INDEX_ORIGINAL]
probability = self.weight[i][KNNAgent.INDEX_WEIGHT] / total_weight
# Begin updating
td_error = td_target - self.q_storage[index]["value"]
self.q_storage[index]["value"] = self.q_storage[index]["value"] + self.alpha*td_error*probability
self.cleanList() # clean list to prepare for another step
# Clear the knn list and also the weight list
def cleanList(self):
self.knn = []
self.weight = []
```
## KNN Main Function
**KNN Main function** is responsible for initiating the KNN agent, environment and handling agent-environment interaction. It consists of a non-terminate inner loop that direct agent decision while also giving reward and next state from the environment. This inner loop will only break after the agent successfully get out of the environment, which in this case the mountain. The outer loop can also be created to control the number of episodes which the agent will perform before the main function ends.
Apart from handling agent-environment interaction, main function also responsible to display three kinds of visualisation which will be explain below the appropriate graph.
```
# Generate decaying alphas
# Input: minimum alpha, number of episodes
# Output: list containing alpha
def generateAlphas(minimum_alpha, n_episodes):
return(np.linspace(1.0, MIN_ALPHA, N_EPISODES))
N_EPISODES = 1000
MIN_ALPHA = 0.02
alphas = generateAlphas(MIN_ALPHA, N_EPISODES)
# Initialise the environment and the agent
size = 1000 # size of the q_storage
k = 6 # knn parameter (this is the best k so far that we have)
agent = KNNAgent(size, k)
mountain_car_environment = MountainCarEnvironment(agent)
# Used for graphing purposes
count_step = [] # counter for how many step in each episodes
# Iterate the process, train the agent (training_iteration episodes)
training_iteration = N_EPISODES
for i in range(training_iteration):
step = 0
alpha = alphas[i]
mountain_car_environment.reset()
while (True):
action = agent.selectAction()
next_state = mountain_car_environment.nextState(action)
# Change agent current state and getting reward
agent.state = next_state
immediate_reward = mountain_car_environment.calculateReward()
# Used for graphing
step += 1
# Test for successful learning
if (immediate_reward == MountainCarEnvironment.REWARD_TERMINAL):
agent.TDUpdate(immediate_reward, alpha)
count_step.append(step)
clear_output(wait=True) # clear previous output
# Create table
d = {"Steps": count_step}
episode_table = pd.DataFrame(data=d, index=np.arange(1, len(count_step)+1))
episode_table.index.names = ['Episodes']
display(episode_table)
break
# Update using Q Learning and kNN
agent.TDUpdate(immediate_reward, alpha)
```
The table above displays total step data taken from 1000 episodes simulation. The first column represents episode and the second column represents total steps taken in a particular episode. It can be seen from the table that during the first few episodes, the agent hasn't learned the environment and hence it chose action unoptimally represented by huge number of steps taken to get to goal. Despite that, after experiencing hundred of episodes the agent have learnt the environment and Q values which enable it to reach the goal in just 200-400 steps.
```
# Create graph for step vs episodes
y = count_step
x = np.arange(1, len(y) + 1)
plt.plot(x, y)
plt.title("Steps vs Episodes (Log Scale)", fontsize=16)
plt.xlabel("Episodes")
plt.ylabel("Steps")
plt.xscale('log')
plt.yscale('log')
plt.show()
```
The line plot visualise the table that are explained above. On the y axis, the plot displays steps taken on each episode, while on the x axis the number of episodes (1000 in the simulation). The line plot is displayed in log-log scale to make it easy to visualise small fluctuation within episode and making sure that large steps in first few episodes don't dominate the graph. From the plot we can see that the overall trend is going downward. This result implies that over many episodes the Q values is getting better and better which eventually will converge to true Q values. Consequently, the agent perform better and better and the step taken to get out of the mountain will decrease with respect to number of episodes.
```
# Create heatmap for Q values
data = pd.DataFrame()
data_left = []
data_neutral = []
data_right = []
position_left = []
position_neutral = []
position_right = []
velocity_left = []
velocity_neutral = []
velocity_right = []
# Sort q_storage based on position and velocity
q_storage_sorted = sorted(agent.q_storage, key=lambda k: k['state'][0])
# Separate action left, neutral, and right
for elem in q_storage_sorted:
if (elem["action"] == -1):
data_left.append(elem["value"])
position_left.append(elem["state"][1])
velocity_left.append(elem["state"][0])
elif (elem["action"] == 0):
data_neutral.append(elem["value"])
position_neutral.append(elem["state"][1])
velocity_neutral.append(elem["state"][0])
else:
data_right.append(elem["value"])
position_right.append(elem["state"][1])
velocity_right.append(elem["state"][0])
# Make scatter plot for 3 actions (left, neutral, right)
# Left
plt.scatter(x=velocity_left, y=position_left, c=data_left, cmap="YlGnBu")
plt.title("Q Values (Action Left)", fontsize=16)
plt.xlabel("Velocity")
plt.ylabel("Position")
plt.colorbar()
plt.show()
# Neutral
plt.scatter(x=velocity_neutral, y=position_neutral, c=data_neutral, cmap="YlGnBu")
plt.title("Q Values (Action Neutral)", fontsize=16)
plt.xlabel("Velocity")
plt.ylabel("Position")
plt.colorbar()
plt.show()
# Right
plt.scatter(x=velocity_right, y=position_right, c=data_right, cmap="YlGnBu")
plt.title("Q Values (Action Right)", fontsize=16)
plt.xlabel("Velocity")
plt.ylabel("Position")
plt.colorbar()
plt.show()
```
Three scatter plots above display Q values for every action on the last episode (1000). Y axis represents position and x axis represents velocity of the 1000 points that we scattered random uniformly initially. To represent Q values for every point, these scatter plots use color indicating value that can be seen from the color bar. When the point is darker, the Q value is around -20. On the other hand, if the point is lighter the Q value is around -100. These Q values are later used for comparison with PNA Algorithm
## PNA Agent
**PNA** may be viewed as a refinement for kNN, with k adapting to the situation. On the one hand, it is beneficial to use large k since that means large data can be learn from. On the other hand, it is beneficial to learn only from the most similar past experiences (small k), as the data they provide should be the most relevant.
PNA suggests that when predicting the value of an action a in a state s, k should be chosen dynamically to minimise:

where c = 1 and Var(Nsa) is the variance of observed rewards in the neighbourhood Nsa. This is a negative version of the term endorsing exploration in the UCB algorithm. Here it promotes choosing neighbourhoods that contain as much data as possible but with small variation between rewards. For example, in the ideal choice of k, all k nearest neighbours of (s, a) behave similarly, but actions farther away behave very differently.
Action are chosen optimistically according to the UCB:

with c > 0 a small constant. The upper confidence bound is composed of two terms: The first terms is the estimated value, and the second term is an exploration bonus for action whose value is uncertain. Actions can have uncertain value either because they have rarely been selected or have a high variance among previous returns. Meanwhile, the neighbourhoods are chosen "pessimistically" for each action to minimise the exploration bonus.
**Algorithm:**
1. Cover the whole state space by some initial Q(s,a) pairs, possibly scatter it uniformly across the whole state space and give an initial value of 0/ -1
2. When an agent in a particular state, get the feature vectors representing the state and possible actions from the state
3. For each possible action from the state, calculate Q(s,a) pairs by taking the expected value from previous Q values based on k-nearest neighbours of a particular action. With PNA, we also need to dynamically consider the k values
*Steps for PNA:*
- Standardise every feature in the feature vectors to (-1, 1) or other ranges to make sure that one feature scale not dominate the distance calculation (i.e. if position ranges between (-50, 50) and velocity (-0.7, 0.7) position will dominate distance calculation).
- Calculate the distance between current state and all of other points with the same action using distance formula (i.e. Euclidean distance) and sort based on the closest distance
- Determine k by minimising the variance function described above
- Store the k-nearest neighbours to knn vector, and it's distance (for weight) in weight vector
- Determine the probability p(x) for the expected value by using weight calculation (i.e. weight = 1/distance). To calculate weight, one can use other formula as long as that formula gives more weight to closer point. To calculate p(x) just divide individual weight with sum of all weights to get probability
- Estimate the Q(s,a) pairs using expectation formula from kNN previous Q values
4. Using epsilon greedy/ UCB/ other decision methods to choose the next move
5. Observe the reward and update the Q values for only the closest neighbour (1 point or chosen by hyperparametric) from KNN array using SARSA or Q Learning. (on the code below, I use Q Learning)
6. Repeat step 2-5
```
class PNAAgent:
"""
Description: Mountain Car problem agent based on PNA algorithm adapted from Marcus Hutter's literatures
Author: Maleakhi Agung Wijaya
"""
INITIAL_VELOCITY = 0.0
INITIAL_POSITION = -0.5
INITIAL_VALUE = -1
ACTIONS = [-1, 0, 1]
GAMMA = 0.995
C = 0.01 # UCB constant
EPSILON = 0.05
RADIUS = 1
INDEX_DISTANCE = 0
INDEX_ORIGINAL = 1
INDEX_WEIGHT = 2
REWARD_STEP = -1
REWARD_TERMINAL = 0
# Constructor
# Input: size of the storage for previous Q values
def __init__(self, size):
self.state = [PNAAgent.INITIAL_VELOCITY, PNAAgent.INITIAL_POSITION]
self.q_storage = []
self.alpha = 1 # choose fixed alpha, but we will vary alpha later
# Storage of the k nearest neighbour (data) and weight (inverse of distance) for a particular step
self.knn = []
self.weight = []
self.k_history = [] # used to store history of k chosen for each action
# For plotting expected PNA function graph
self.var_function_left = []
self.var_function_neutral = []
self.var_function_right = []
self.converge_function_left = []
self.converge_function_neutral = []
self.converge_function_right = []
self.episode = 0 # keep count of how many episodes for plotting purposes as well
# Initialise the storage with random point
for i in range(size):
initial_value = PNAAgent.INITIAL_VALUE
initial_action = random.randint(-1, 1)
initial_state = [random.uniform(-0.07, 0.07), random.uniform(-1.2, 0.6)]
# Fill the graph with all possible k
if (initial_action == -1):
self.var_function_left.append(0)
self.converge_function_left.append(0)
elif (initial_action == 0):
self.var_function_neutral.append(0)
self.converge_function_neutral.append(0)
else:
self.var_function_right.append(0)
self.converge_function_right.append(0)
# Each data on the array will consist of state, action pair + value
data = {"state": initial_state, "value": initial_value, "action": initial_action}
self.q_storage.append(data)
# Since the k start at 2 that we want to calculate, just pop 1
self.var_function_left.pop()
self.var_function_neutral.pop()
self.var_function_right.pop()
self.converge_function_left.pop()
self.converge_function_neutral.pop()
self.converge_function_right.pop()
# Standardise feature vector given
# Input: feature vector to be standardised
# Output: standardised feature vector
def standardiseState(self, state):
standardised_state = []
standardised_velocity = 2 * ((state[0]+0.07) / (0.07+0.07)) - 1
standardised_position = 2 * ((state[1]+1.2) / (0.6+1.2)) - 1
standardised_state.append(standardised_velocity)
standardised_state.append(standardised_position)
return(standardised_state)
# Find all index for a given value
# Input: value, list to search
# Output: list of all index where you find that value on the list
def findAllIndex(self, value, list_value):
indices = []
for i in range(len(list_value)):
if (value == list_value[i]):
indices.append(i)
return indices
# Calculate Euclidean distance between 2 vectors
# Input: 2 feature vectors
# Output: distance between them
def calculateDistance(self, vector1, vector2):
return(math.sqrt((vector1[0]-vector2[0])**2 + (vector1[1]-vector2[1])**2))
# Calculate total weight
# Input: list of weights
# Output: total weight
def calculateTotalWeight(self, weight_list):
total_weight = 0
for i in range(len(weight_list)):
total_weight += weight_list[i][PNAAgent.INDEX_WEIGHT]
return(total_weight)
# Clear the knn list, k_history, and also the weight list
def cleanList(self):
self.knn = []
self.weight = []
self.k_history = []
# Choose the appropriate k by minimising variance and maximising the number of data to learn
# Input: sorted neighbourhood list based on distance (distance, index, weight)
# Output: k (numbers of nearest neighbour) that minimise neighbourhood variance function
def chooseK(self, neighbourhood_list):
data_list = []
# Extract the data (Q value from the neighbourhood_list) and append it to the data_list
for data in neighbourhood_list:
data_list.append(self.q_storage[data[PNAAgent.INDEX_ORIGINAL]]["value"])
action = self.q_storage[data[PNAAgent.INDEX_ORIGINAL]]["action"]
# Initialise minimum variance
minimum_k = 2 # Variable that will be return that minimise the variance of the neighbourhood
minimum_function = self.neighbourhoodVariance(1, data_list[:2])
# For plotting variance function graph
list_var = []
if (action == -1):
list_var = self.var_function_left
elif (action == 0):
list_var = self.var_function_neutral
else:
list_var = self.var_function_right
list_var[0] += minimum_function
if (self.episode > 900):
list_var_converge = []
if (action == -1):
list_var_converge = self.converge_function_left
elif (action == 0):
list_var_converge = self.converge_function_neutral
else:
list_var_converge = self.converge_function_right
list_var_converge[0] += minimum_function
previous_sum_variance = np.var(data_list[:2]) * 2
previous_mean = np.mean(data_list[:2])
k = 2
# Iterate to find optimal k that will minimise the neighbourhood variance function
for i in range(2, len(neighbourhood_list)):
target_x = data_list[i]
mean = (previous_mean * k + target_x) / (k + 1)
current_sum_variance = previous_sum_variance + (target_x - previous_mean) * (target_x - mean)
# Update for next iteration
k = k + 1
previous_sum_variance = current_sum_variance
previous_mean = mean
function = self.neighbourhoodVariance(1, [], previous_sum_variance / k, k)
list_var[k-2] += function
if (self.episode > 900):
list_var_converge[k-2] += function
# Update the k value and minimum var value if find parameter which better minimise than the previous value
if (function <= minimum_function):
minimum_k = k
minimum_function = function
return(minimum_k)
# PNA variance function that needed to be minimise
# Input: constant c, list containing data points
# Output: calculation result from the neighbourhood variance function
def neighbourhoodVariance(self, c, data_list, var = None, k = None):
if (var == None):
return(math.sqrt(c * np.var(data_list) / len(data_list)))
else:
return(math.sqrt(c * var / k))
# Get starting index for the weight list
# Input: action, k_history
# Output: starting index for the weight list
def getStartingIndex(self, action, k_history):
count_action = action + 1
if (count_action == 0):
return(0)
else:
index = 0
for i in range(count_action):
index += k_history[i]
return(index)
# Apply the PNA algorithm for feature vector and store the data point on the neighbours array
# Input: feature vector of current state, actions array consisting of all possible actions, list that will store knn data and weights data, k_history
# Output: vector containing the value of taking each action (left, neutral, right)
def PNA(self, state, actions, knn_list, weight_list, k_history):
approximate_action = []
# Get the standardised version of state
standardised_state = self.standardiseState(state)
# Loop through every element in the storage array and only calculate for particular action
for action in actions:
temp = [] # array consisting of tuple (distance, original index, weight) for each point in the q_storage
for i in range(len(self.q_storage)):
data = self.q_storage[i]
# Only want to calculate the nearest neighbour state which has the same action
if (data["action"] == action):
vector_2 = data["state"]
standardised_vector_2 = self.standardiseState(vector_2)
distance = self.calculateDistance(standardised_state, standardised_vector_2)
index = i
weight = 1 / (1+distance**2)
# Create the tuple and append that to temp
temp.append(tuple((distance, index, weight)))
else:
continue
# After we finish looping through all of the point and calculating the standardise distance,
# Sort the tuple based on the distance and only take k of it and append that to the neighbours array
sorted_temp = sorted(temp, key=lambda x: x[0])
# Get the value of the k dynamically
k = self.chooseK(sorted_temp)
k_history.append(k)
for i in range(k):
try:
weight_list.append(sorted_temp[i])
knn_list.append(self.q_storage[sorted_temp[i][PNAAgent.INDEX_ORIGINAL]])
except IndexError:
sys.exit(0)
# Calculate the expected value of the action and append it to the approximate_action array
expected_value = 0
# We also need to calculate the total weight to make it into valid probability that we can compute it's expectation
total_weight = self.calculateTotalWeight(weight_list[self.getStartingIndex(action, k_history):self.getStartingIndex(action, k_history)+k])
for i in range(self.getStartingIndex(action, k_history), self.getStartingIndex(action, k_history) + k):
try:
weight = weight_list[i][PNAAgent.INDEX_WEIGHT]
probability = weight / total_weight
expected_value += probability * knn_list[i]["value"]
except IndexError:
sys.exit(0)
approximate_action.append(expected_value)
return(approximate_action)
# Calculate TD target based on Q Learning/ SARSAMAX
# Input: Immediate reward based on what the environment gave
# Output: TD target based on off policy Q learning
def calculateTDTarget(self, immediate_reward):
# Condition if final state
if (immediate_reward == PNAAgent.REWARD_TERMINAL):
return(immediate_reward)
k_history = []
knn_prime = []
weight_prime = []
action_value = self.PNA(self.state, PNAAgent.ACTIONS, knn_prime, weight_prime, k_history)
return(immediate_reward + PNAAgent.GAMMA * max(action_value))
# Q learning TD updates on every neighbours on the kNN based on the contribution that are calculated using probability weight
# Input: Immediate reward based on what the environment gave
def TDUpdate(self, immediate_reward, alpha):
self.alpha = alpha
# First, calculate the TD target
td_target = self.calculateTDTarget(immediate_reward)
try:
# Update only the #radius closest point
total_weight = self.calculateTotalWeight(self.weight[0:PNAAgent.RADIUS])
for i in range(PNAAgent.RADIUS):
index = self.weight[i][PNAAgent.INDEX_ORIGINAL]
probability = self.weight[i][PNAAgent.INDEX_WEIGHT] / total_weight
td_error = td_target - self.q_storage[index]["value"]
self.q_storage[index]["value"] += self.alpha * td_error * probability
except IndexError:
total_weight = self.calculateTotalWeight(self.weight)
for i in range(len(self.weight)):
index = self.weight[i][PNAAgent.INDEX_ORIGINAL]
probability = self.weight[i][PNAAgent.INDEX_WEIGHT] / total_weight
# Begin updating
td_error = td_target - self.q_storage[index]["value"]
self.q_storage[index]["value"] += self.alpha * td_error * probability
self.cleanList() # clean list to prepare for another step
# Choosing based on Epsilon Greedy method
# Input: action_value array consisting the Q value of every action
# Output: action chosen (-1, 0, 1)
def epsilonGreedy(self, action_value):
# Use the epsilon-greedy method to choose value
random_number = random.uniform(0.0, 1.0)
if (random_number <= PNAAgent.EPSILON):
action_chosen = random.randint(-1, 1)
else:
# Return the action with highest Q(s,a)
possible_index = self.findAllIndex(max(action_value), action_value)
action_chosen = possible_index[random.randrange(len(possible_index))] - 1
return action_chosen
# Getting the maximum of the ucb method
# Input: action_value list, bonus_variance list
# Output: action which maximise
def maximumUCB(self, action_value, bonus_variance):
max_index = 0
max_value = action_value[0] + bonus_variance[0]
# Check 1, 2 (all possible action)
for i in range(1, 3):
value = action_value[i] + bonus_variance[i]
if (value >= max_value):
max_value = value
max_index = i
return(max_index - 1) # return the action which maximise
# Select which action to choose, whether left, neutral, or right (using UCB)
# Output: -1 (left), 0 (neutral), 1 (right)
def selectAction(self):
action_value = self.PNA(self.state, PNAAgent.ACTIONS, self.knn, self.weight, self.k_history)
# Second term of ucb, calculate the bonus variance
start_index = [] # used to calculate start index for each action
finish_index = [] # used to calculate end index for each action
for action in PNAAgent.ACTIONS:
# Prevent index out of bound
if (action != 1):
# Data extraction
start_index.append(self.getStartingIndex(action, self.k_history))
finish_index.append(self.getStartingIndex(action+1, self.k_history))
else:
# Data extraction
start_index.append(self.getStartingIndex(action, self.k_history))
finish_index.append(len(self.weight))
# Choose the action based on ucb method
action_chosen = self.epsilonGreedy(action_value)
# Only store chosen data in the knn and weight list
# Clearance step
chosen_knn = []
chosen_weight = []
for i in range(start_index[action_chosen+1], finish_index[action_chosen+1]):
chosen_knn.append(self.knn[i])
chosen_weight.append(self.weight[i])
self.knn = chosen_knn
self.weight = chosen_weight
return action_chosen
```
## PNA Main Function
**PNA Main function** is responsible for initiating the PNA agent, environment and handling agent-environment interaction. It consists of a non-terminate inner loop that direct agent decision while also giving reward and next state from the environment. This inner loop will only break after the agent successfully get out of the environment, which in this case the mountain or if it is taking too long to converge. The outer loop can also be created to control the number of episodes which the agent will perform before the main function ends.
Apart from handling agent-environment interaction, main function also responsible to display five kinds of visualisation. First, table/ DataFrame displaying episodes and step that are required by the agent to get out of the mountain on each episode. Second, scatter plot displaying steps on the y axis and episodes on the x axis to learn about algorithm convergence property. Third, expected standard error function for every actions. Fourth, heatmap of the Q value for the last episode. Lastly, as the k is dynamically changing each steps, I have created a heatmap indicating k chosen each steps for first episode and last episode.
```
# Generate decaying alphas
# Input: minimum alpha, number of episodes
# Output: list containing alpha
def generateAlphas(minimum_alpha, n_episodes):
return(np.linspace(1.0, MIN_ALPHA, N_EPISODES))
N_EPISODES = 1000
MIN_ALPHA = 0.02
alphas = generateAlphas(MIN_ALPHA, N_EPISODES)
# Initialise the environment and the agent
size = 1000 # size of the q_storage
agent = PNAAgent(size)
mountain_car_environment = MountainCarEnvironment(agent)
convergence = 100 # used to extract data when agent has converges
# Used for graphing purposes
count_step = [] # counter for how many step in each episodes
k_first_left = []
k_first_neutral = []
k_first_right = []
k_last_left = []
k_last_neutral = []
k_last_right = []
k_convergence_left = agent.var_function_left[:]
k_convergence_neutral = agent.var_function_neutral[:]
k_convergence_right = agent.var_function_right[:]
# Iterate the process, train the agent (training_iteration episodes)
total_step = 0
training_iteration = N_EPISODES
for i in range(training_iteration):
step = 0
alpha = alphas[i]
mountain_car_environment.reset()
agent.episode = i + 1
while (True):
action = agent.selectAction()
next_state = mountain_car_environment.nextState(action)
# Change agent current state and getting reward
agent.state = next_state
immediate_reward = mountain_car_environment.calculateReward()
# Used for graphing
step += 1
total_step += 1
# Only append first and last episode (for the k)
if (i == 1):
k_first_left.append(agent.k_history[0])
k_first_neutral.append(agent.k_history[1])
k_first_right.append(agent.k_history[2])
if (i == (training_iteration - 1)):
k_last_left.append(agent.k_history[0])
k_last_neutral.append(agent.k_history[1])
k_last_right.append(agent.k_history[2])
# Count how many k chosen after converge
if (agent.episode > 900):
# Increment count when a particular k is chosen, 2 is just scaling factor since the k starts from 2 in the array
k_convergence_left[agent.k_history[0]-2] += 1
k_convergence_neutral[agent.k_history[1]-2] += 1
k_convergence_right[agent.k_history[2]-2] += 1
# Test for successful learning
if (immediate_reward == MountainCarEnvironment.REWARD_TERMINAL):
agent.TDUpdate(immediate_reward, alpha)
count_step.append(step)
clear_output(wait=True) # clear previous output
# Create table
d = {"Steps": count_step}
episode_table = pd.DataFrame(data=d, index=np.arange(1, len(count_step)+1))
episode_table.index.names = ['Episodes']
display(episode_table)
break
# Update using Q Learning and kNN
agent.TDUpdate(immediate_reward, alpha)
```
The table above displays total step data taken from 1000 episodes simulation. The first column represents episode and the second column represents total steps taken in a particular episode. It can be seen from the table that during the first few episodes, the agent hasn't learned the environment and hence it chose action unoptimally represented by huge number of steps taken to get to goal. Despite that, after experiencing hundred of episodes the agent have learnt the environment and Q values which enable it to reach the goal in around 300-600 steps.
```
# Create graph for step vs episodes
y = count_step
x = np.arange(1, len(y) + 1)
plt.plot(x, y)
plt.title("Steps vs Episodes (Log Scale)", fontsize=16)
plt.xlabel("Episodes (Log)")
plt.ylabel("Steps (Log)")
plt.xscale('log')
plt.yscale('log')
plt.show()
```
The line plot visualise the table that are explained above. On the y axis, the plot displays steps taken on each episode, while on the x axis the number of episodes (1000 in the simulation). The line plot is displayed in log-log scale to make it easy to visualise small fluctuation within episode and making sure that large steps in first few episodes don't dominate the graph. From the plot we can see that the overall trend is going downward. The result implies that over many episodes the Q values is getting better and better which eventually will converge to true Q values. Consequently, the agent perform better and better and the step taken to get out of the mountain will decrease with respect to number of episodes.
```
# Create plot for the average standard error function
average_var_left = []
average_var_neutral = []
average_var_right = []
for elem in agent.var_function_left:
average_var_left.append(elem / total_step)
for elem in agent.var_function_neutral:
average_var_neutral.append(elem / total_step)
for elem in agent.var_function_right:
average_var_right.append(elem / total_step)
# Make a scatter plot
# Left
y = average_var_left
x = np.arange(2, len(y)+2)
plt.plot(x, y, color="#55A868")
plt.title("Average Standard Error Function vs K (Action Left)", fontsize=16)
plt.xlabel("K")
plt.ylabel("Average f(K)")
plt.xticks(np.arange(2, len(y) + 2, 50))
plt.show()
# Make a scatter plot
# Neutral
y = average_var_neutral
x = np.arange(2, len(y)+2)
plt.plot(x, y, color="#55A868")
plt.title("Average Standard Error Function vs K (Action Neutral)", fontsize=16)
plt.xlabel("K")
plt.ylabel("Average f(K)")
plt.xticks(np.arange(2, len(y) + 2, 50))
plt.show()
# Make a scatter plot
# Right
y = average_var_right
x = np.arange(2, len(y)+2)
plt.plot(x, y, color="#55A868")
plt.title(" Average Standard Error Function vs K (Action Right)", fontsize=16)
plt.xlabel("K")
plt.ylabel("Average f(K)")
plt.xticks(np.arange(2, len(y) + 2, 50))
plt.show()
# Now plot the standard error function after convergence
reverse_count_step = count_step[::-1]
total_last_step = 0
for i in range(convergence):
total_last_step += reverse_count_step[i]
average_converge_left = []
average_converge_neutral = []
average_converge_right = []
for elem in agent.converge_function_left:
average_converge_left.append(elem / total_last_step)
for elem in agent.converge_function_neutral:
average_converge_neutral.append(elem / total_last_step)
for elem in agent.converge_function_right:
average_converge_right.append(elem / total_last_step)
# Make a scatter plot
# Left
y = average_converge_left
x = np.arange(2, len(y)+2)
plt.plot(x, y, color="#B14C4D")
plt.title("Average Standard Error Function vs K After Convergence (Action Left)", fontsize=16)
plt.xlabel("K")
plt.ylabel("Average f(K)")
plt.xticks(np.arange(2, len(y) + 2, 50))
plt.show()
# Make a scatter plot
# Neutral
y = average_converge_neutral
x = np.arange(2, len(y)+2)
plt.plot(x, y, color="#B14C4D")
plt.title("Average Standard Error Function vs K After Convergence (Action Neutral)", fontsize=16)
plt.xlabel("K")
plt.ylabel("Average f(K)")
plt.xticks(np.arange(2, len(y) + 2, 50))
plt.show()
# Make a scatter plot
# Right
y = average_converge_right
x = np.arange(2, len(y)+2)
plt.plot(x, y, color="#B14C4D")
plt.title(" Average Standard Error Function vs K After Convergence (Action Right)", fontsize=16)
plt.xlabel("K")
plt.ylabel("Average f(K)")
plt.xticks(np.arange(2, len(y) + 2, 50))
plt.show()
```
The first 3 graphs display average standard error function calculated for every steps from episode 1 - episode 1000. X axis display the possible k for every actions, while y axis display the average standard error function for each k. From both the plot above and bar plot below, it can be seen that k = 2 is chosen most of the time since it's mostly minimise the standard error function compare to other k. Even though 2 is the most frequent k chosen, if we dissect the plot for every episodes, it is not always the case. On some steps/ episodes, the graph are dominated by the number of neighbourhood which makes the graph looks like 1/sqrt(n) resulted in large amount of k (200-300) chosen.
The last 3 graphs display average standard error function calculated for the last 100 episodes out of 1000 episodes (converges). These graphs have similar value with the first 3 graphs and hence the explanation is similar.
```
# Create heatmap for Q values
data = pd.DataFrame()
data_left = []
data_neutral = []
data_right = []
position_left = []
position_neutral = []
position_right = []
velocity_left = []
velocity_neutral = []
velocity_right = []
# Sort q_storage based on position and velocity
q_storage_sorted = sorted(agent.q_storage, key=lambda k: k['state'][0])
# Separate action left, neutral, and right
for elem in q_storage_sorted:
if (elem["action"] == -1):
data_left.append(elem["value"])
position_left.append(elem["state"][1])
velocity_left.append(elem["state"][0])
elif (elem["action"] == 0):
data_neutral.append(elem["value"])
position_neutral.append(elem["state"][1])
velocity_neutral.append(elem["state"][0])
else:
data_right.append(elem["value"])
position_right.append(elem["state"][1])
velocity_right.append(elem["state"][0])
# Make scatter plot for 3 actions (left, neutral, right)
# Left
plt.scatter(x=velocity_left, y=position_left, c=data_left, cmap="YlGnBu")
plt.title("Q Values (Action Left)", fontsize=16)
plt.xlabel("Velocity")
plt.ylabel("Position")
plt.colorbar()
plt.show()
# Neutral
plt.scatter(x=velocity_neutral, y=position_neutral, c=data_neutral, cmap="YlGnBu")
plt.title("Q Values (Action Neutral)", fontsize=16)
plt.xlabel("Velocity")
plt.ylabel("Position")
plt.colorbar()
plt.show()
# Right
plt.scatter(x=velocity_right, y=position_right, c=data_right, cmap="YlGnBu")
plt.title("Q Values (Action Right)", fontsize=16)
plt.xlabel("Velocity")
plt.ylabel("Position")
plt.colorbar()
plt.show()
```
Three scatter plots above display Q values for every action on the last episode (1000). Y axis represents position and x axis represents velocity of the 1000 points that we scattered random uniformly initially. To represent Q values for every point, these scatter plots use color indicating the value that can be seen from the color bar. When the point is darker, the Q value is around -20. On the other hand, if the point is lighter the Q value is around -100.
If we observe the Q values for both KNN-TD and PNA, it can be seen that the Q values are roughly similar. This result implies that both of the algorithm converges for the Mountain Car problem and eventually after numerous episodes, the agent Q values will converge to the true Q values.
```
# Create heatmap showing the k (first episode)
data = pd.DataFrame()
data["Action Left"] = k_first_left
data["Action Neutral"] = k_first_neutral
data["Action Right"] = k_first_right
data["Steps"] = np.arange(1, len(k_first_left) + 1)
data.set_index("Steps", inplace=True)
grid_kws = {"height_ratios": (.9, .05), "hspace": .3}
f, (ax, cbar_ax) = plt.subplots(2, gridspec_kw=grid_kws)
ax = sns.heatmap(data, ax=ax, cbar_ax=cbar_ax, cbar_kws={"orientation": "horizontal"}, yticklabels=False)
ax.set_title("Number of K Chosen Each Step (First Episode)", fontsize=16)
plt.show()
# Create heatmap showing the k (last episode)
data = pd.DataFrame()
data["Action Left"] = k_last_left
data["Action Neutral"] = k_last_neutral
data["Action Right"] = k_last_right
data["Steps"] = np.arange(1, len(k_last_left) + 1)
data.set_index("Steps", inplace=True)
grid_kws = {"height_ratios": (.9, .05), "hspace": .3}
f, (ax, cbar_ax) = plt.subplots(2, gridspec_kw=grid_kws)
ax = sns.heatmap(data, ax=ax, cbar_ax=cbar_ax, cbar_kws={"orientation": "horizontal"}, yticklabels=False)
ax.set_title("Number of K Chosen Each Step (Last Episode)", fontsize=16)
plt.show()
```
The heatmap displayed above represents k chosen each step for every actions. Each strip on the heatmap represents the k chosen on a step. The first heatmap displayed data from the first episode. Based on the heatmap, it can be seen that the k chosen each step during the first episode is large (around 120-180). This occurs because all points are initialise to have a uniform value of -1. As the Q values is uniformly/ roughly -1 across the whole space, this will make the variance approximately 0 and hence resulted in the standard error function depends largely on the number of k chosen/ neighbourhoods. As the algorithm prefer to minimise the standard error function, it will chooses as many point as possible.
The second heatmap displayed data from the last episode. Based on the heatmap, it can be seen that the k chosen each step is relatively small (around 2-60). This occurs since the agent has gain large amount of experience and Q values greatly differ in different region. As a result, if the agent choose a really large k to learn, it will make the variance very high and hence really large standard error. Consequently, the agent will minimise standard error function by repeatedly choose k around 2-60.
```
# Plot bar chart displaying number of k chosen after convergence
y_bar = k_convergence_left
x_bar = np.arange(2, len(y_bar) + 2)
plt.bar(x_bar, y_bar, color="#FFD700")
plt.yscale('log')
plt.title("Number of K Chosen vs K After Convergence (Action Left)", fontsize=16)
plt.xlabel("K")
plt.ylabel("Number of K Chosen")
plt.xticks(np.arange(2, len(y_bar) + 2, 50))
plt.show()
y_bar = k_convergence_neutral
x_bar = np.arange(2, len(y_bar) + 2)
plt.bar(x_bar, y_bar, color="#FFD700")
plt.yscale('log')
plt.title("Number of K Chosen vs K After Convergence (Action Neutral)", fontsize=16)
plt.xlabel("K")
plt.ylabel("Number of K Chosen")
plt.xticks(np.arange(2, len(y_bar) + 2, 50))
plt.show()
y_bar = k_convergence_right
x_bar = np.arange(2, len(y_bar) + 2)
plt.bar(x_bar, y_bar, color="#FFD700")
plt.yscale('log')
plt.title("Number of K Chosen vs K After Convergence (Action Right)", fontsize=16)
plt.xlabel("K")
plt.ylabel("Number of K Chosen")
plt.xticks(np.arange(2, len(y_bar) + 2, 50))
plt.show()
```
These bar plots represent the number of k chosen for each k after convergence for every actions. X axis represents possible k for each action, while y axis represents how many times the k chosen for a particular k after convergence. The convergence defined in the code is the last 100 episodes out of 1000 episodes.
In all of bar plots, we can see that after convergence the agent mostly choose k equals 2 and relatively small k such as from 2 - 150. This condition occurs because the agent has lots of experience which thus make the Q values highly differ between region. Based on the result, we can see that mostly variance dominates the standard error function which force the agent to choose small k to minimise the standard error function.
| github_jupyter |
```
from aide_design.play import*
#Below are the items that were imported by the code above so that you know what abbreviations to use in your code.
# Third-party imports
#import numpy as np
#import pandas as pd
#import matplotlib.pyplot as plt
#import matplotlib
# AIDE imports
#import aide_design
#import aide_design.pipedatabase as pipe
#from aide_design.units import unit_registry as u
#from aide_design import physchem as pc
#import aide_design.expert_inputs as exp
#import aide_design.materials_database as mat
#import aide_design.utility as ut
#import aide_design.k_value_of_reductions_utility as k
#import aide_design.pipeline_utility as pipeline
#import warnings
```
## Prelim 2
<div class="alert alert-block alert-info">
# Name this file: Lastname_Firstname_Prelim2
# Multiple Choice (4 points each - 12 points)
Make your answer **bold** by typing `**` before and after the correct answer.
1. In class we talked about the floc volcano phenomenon of the San Nicolas plant. The floc volcano phenomenon occurs when water entering the sed tank is warmer than the water in the sed tank, and stops when the water entering the sed tank becomes colder than the water in the sed tank. Why does colder water stop the floc volcano phenomenon?
1. Colder influent water has a higher viscosity and thus reduces the velocity of the floc volcanoes
1. Colder influent water is more dense than the water in the sedimentation tank and thus the density stratification in the sedimentation tank is stable
1. Flow rate through the plant decreases when the water is colder, and thus floc volcano formation is averted
<br>
<br>
1. Which of the following changes could AguaClara do to further reduce the **depth** of the sedimentation tanks? For each change assume that the other design parameters are held constant while that one change is made.
1. Increase the vertical velocity
1. Increase the capture velocity
1. Increase the spacing between plate settlers (hold capture velocity constant)
1. Increase the water velocity exiting the diffusers
<br>
<br>
1. What does the volume of water in a vertical flow hydraulic flocculator do when the flow rate inceases?
1. Decreases
1. Remains constant
1. Increases
***
# Short Answer (3 points each - 18 points)
Explain your answers as thoroughly as possible.
## Double click on this cell to write your answers
1. Give at least 3 reasons why water treatment plants should be enclosed in a building
1.
1.
1.
<br>
<br>
1. List at least 3 problems that can cause sedimentation tanks to perform poorly
1.
1.
1.
<br>
<br>
1. Re-sort the following list according to the amount of solids each unit process removes from highly turbid (for example, 500 NTU) raw water. A is the most solids removal and C is the least solids removal: <br>
1. Flocculation
1. Sedimentation
1. Filtration
<br>
<br>
1. List three reasons why the Sedimentation tank inlet manifold has diffuser pipes.
1.
1.
1.
<br>
<br>
1. How does accumulated sludge in a sedimentation tank cause poor performance?
1.
<br>
<br>
1. Why is it important to be able to send poorly flocculated water to waste BEFORE it enters the sedimentation tank?
1.
# Design Challenges (70 points)
Assign the given inputs to variables. Write the equation in python using variables. ** Make sure to use print statements which put your answer in context, and to display your answer in appropriate units **
## 1) Flocculator (27 points)
A 60 L/s water treatment plant has a vertical flow hydraulic flocculator. The channels are 50 cm wide, 6 m long, and 2 m deep at the end of the flocculator. The head loss through the flocculator is 40 cm. The minimum water temperature is 15°C.
There are 4 channels. The minor loss coefficient for each 180° bend is 2.5.
Answer A through F.
State any assumptions that you make.
```
# define your parameters and variables given in the problem statement here:
```
### A. (5 points)
What is the hydraulic residence time in the flocculator if you neglect the volume change due to head loss?
### B. (5 points)
What is the average velocity gradient in the flocculator?
### C. (5 points)
What is the collision potential $\big(G\theta\big)$ for this flocculator?
### D. (5 points)
What is the maximum distance between expansions for this flocculator? The maximum ratio of H/S ($\Pi _{H{S_{Max}}}$) is 6
### E. (2 points)
Is it necessary to include additional expansions in this design to maintain high efficiency?
**Your Response Here:**
### F. (5 points)
What is the spacing of the baffles in this flocculator?
## 2) Sedimentation tank area (5 points)
The average water use per person in the US is 3 mL/s. How much floc blanket plan view area does one person require? The floc blanket upflow velocity is 1 mm/s.
## 3) Plate settler design (15 points)
Assume the upflow velocity in AguaClara sedimentation tanks is increased to 3 mm/s,
the capture velocity is increased to 0.2 mm/s, the spacing between plates is maintained at 2.5 cm, and the plates are inclined at 60$^{\circ}$ from the horizontal. You may assume that the vertical velocity entering the plate settlers is 3 mm/s.
Answer questions A and B.
### A. (10 points)
How long would the plate settlers be? You may neglect the effect of the thickness of the
plate settlers. **Create a function** for the equation that you use to solve this problem. Include all the necessary inputs in your function definition.
### B. (5 points)
What is the ** *vertical* ** height of the plate settlers? This will tell us how much of the sedimentation tank depth these plates occupy.
## 4) Head loss through a floc blanket (8 points)
### A. (5 points)
What is the head loss through a 1 m deep floc blanket with a clay concentration of 3 g/L and an upflow velocity of 1 mm/s? You may assume the water temperature is 20°C. The density of clay is 2650 $\frac{kg}{m^3}$. This head loss helps ensure a uniform velocity of water leaving the water - floc blanket interface at the top of the floc blanket.
### B. (3 points)
Which head loss is greater, the head loss through the floc blanket or the head loss through the plate settlers?
**Your Response Here:**
## 5) Floc Hopper (10 points)
The AguaClara 1 L/s sedimentation tank has a floc hopper that is a 3" PVC pipe that is angled at 60$^{\circ}$C from the horizontal. The raw water is 1000 NTU and the settled water turbidity is 3 NTU.
A sample from the consolidated sludge in the bottom of the floc hopper was diluted by a factor of 100 and the turbidity of the diluted sample was 150 NTU. (Remember that NTU is an approximate measure of concentration and the unit NTU has been defined for you).
What flow rate must the floc hopper drain valve be set at to maintain a constant level of the consolidated sludge in the floc hopper?
## 6) Multiple stage filtration plan view area (5 points)
How much area would the unit processes of a multiple stage filtration plant require to provide the water for one person from the US?
<div class="alert alert-block alert-info">
### When you finish the exam:
** * Double check to ensure that your file is named correctly* **
Make sure to 'Restart and Run All' before you save for the final time to ensure your outputs show your latest work. Once you have done that, email the .ipynb file to cee4540@gmail.com. Congratulations on finishing the last exam for this course!
| github_jupyter |
# Credit Card Fraud Detection
The dataset can be found in Kaggle: https://www.kaggle.com/mlg-ulb/creditcardfraud
The datasets contains transactions made by credit cards in September 2013 by european cardholders. \
This dataset presents transactions that occurred in two days, where we have 492 frauds out of 284,807 transactions. \
The dataset is highly unbalanced, the positive class (frauds) account for 0.172% of all transactions.
We will use both Decision Tree and SVM algorithms for Detection
```
import numpy as np
import pandas as pd
import sklearn
import scipy
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics import classification_report,accuracy_score
from sklearn.ensemble import IsolationForest
from sklearn.neighbors import LocalOutlierFactor
from sklearn.svm import OneClassSVM
import matplotlib.pyplot as plt
%matplotlib inline
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.svm import SVC
from sklearn.svm import SVR
from sklearn.naive_bayes import GaussianNB
from sklearn.tree import DecisionTreeClassifier
import math
from pylab import rcParams
rcParams['figure.figsize'] = 14, 8
RANDOM_SEED = 42
LABELS = ["Normal", "Fraud"]
from sklearn.preprocessing import StandardScaler
dataset = pd.read_csv('creditcard.csv')
dataset.head()
dataset.info()
x = dataset.iloc[: , 1:30].values
y = dataset.iloc[:, 30].values
print("Input Range : ", x.shape)
print("Output Range : ", y.shape)
print ("Class Labels : \n", y)
dataset.isnull().values.any()
set_class = pd.value_counts(dataset['Class'], sort = True)
set_class.plot(kind = 'bar', rot=0)
plt.title("Class Distribution of Transaction")
plt.xticks(range(2), LABELS)
plt.xlabel("Classes")
plt.ylabel("No of occurences")
fraud_data = dataset[dataset['Class']==1]
normal_data = dataset[dataset['Class']==0]
print(fraud_data.shape,normal_data.shape)
fraud_data.Amount.describe()
normal_data.Amount.describe()
## Correlation
import seaborn as sns
#get correlations of each features in dataset
corrmat = dataset.corr()
top_corr_features = corrmat.index
plt.figure(figsize=(20,20))
#plot heat map
g=sns.heatmap(dataset[top_corr_features].corr(),annot=True,cmap="RdYlGn")
xtrain, xtest, ytrain, ytest = train_test_split(x, y, test_size=0.25, random_state = 0)
print("xtrain.shape : ", xtrain.shape)
print("xtest.shape : ", xtest.shape)
print("ytrain.shape : ", ytrain.shape)
print("ytest.shape : ", ytest.shape)
stdsc = StandardScaler()
xtrain = stdsc.fit_transform(xtrain)
xtest = stdsc.transform(xtest)
print("Training Set after Standardised : \n", xtrain[0])
dt_classifier = DecisionTreeClassifier(criterion = 'entropy', random_state = 0)
dt_classifier.fit(xtrain, ytrain)
y_pred_decision_tree = dt_classifier.predict(xtest)
print("y_pred_decision_tree : \n", y_pred_decision_tree)
com_decision = confusion_matrix(ytest, y_pred_decision_tree)
print("confusion Matrix : \n", com_decision)
Accuracy_Model = ((com_decision[0][0] + com_decision[1][1]) / com_decision.sum()) *100
print("Accuracy_Decison : ", Accuracy_Model)
Error_rate_Model= ((com_decision[0][1] + com_decision[1][0]) / com_decision.sum()) *100
print("Error_rate_Decison : ", Error_rate_Model)
# True Fake Rate
Specificity_Model= (com_decision[1][1] / (com_decision[1][1] + com_decision[0][1])) *100
print("Specificity_Decison : ", Specificity_Model)
# True Genuine Rate
Sensitivity_Model = (com_decision[0][0] / (com_decision[0][0] + com_decision[1][0])) *100
print("Sensitivity_Decison : ", Sensitivity_Model)
svc_classifier = SVC(kernel = 'rbf', random_state =0)
svc_classifier.fit(xtrain, ytrain)
y_pred2 = svc_classifier.predict(xtest)
print("y_pred_randomforest : \n", y_pred2)
cm2 = confusion_matrix(ytest, y_pred2)
print("Confusion Matrix : \n\n", cm2)
# Validating the Prediction
Accuracy_Model = ((cm2[0][0] + cm2[1][1]) / cm2.sum()) *100
print("Accuracy_svc : ", Accuracy_Model)
Error_rate_Model = ((cm2[0][1] + cm2[1][0]) / cm2.sum()) *100
print("Error_rate_svc : ", Error_rate_Model)
# True Fake Rate
Specificity_Model= (cm2[1][1] / (cm2[1][1] + cm2[0][1])) *100
print("Specificity_svc : ", Specificity_Model)
# True Genuine Rate
Sensitivity_Model= (cm2[0][0] / (cm2[0][0] + cm2[1][0])) *100
print("Sensitivity_svc : ", Sensitivity_Model)
```
| github_jupyter |
# Decisions
This notebook is based on materials kindly provided by the [IN1900]( https://www.uio.no/studier/emner/matnat/ifi/IN1900/h19/) team.
How can we use Python to automatically recognize different features in our data, and take a different action for each?
Here, we will learn how to write code that executes only when certain conditions are true.
We can tell Python to take an action depending on the value of a variable:
```
length = 42
if length > 100:
print('greater')
```
We can also include an alternate path, `else`:
```
length = 42
if length > 100:
print('greater')
else:
print('smaller')
print('done')
```
This code can be illustrated with a flowchart:

## `elif`
We can chain multiple `if`-tests with
`elif`, short for "else if".
```
length = 42
if length > 100:
print('greater')
elif length < 0:
print('Oops, negative length?')
else:
print('smaller')
print('done')
```
### <span style="color:green"> Exercise: multiple hits </span>
With `elif`, only the first test that yields `True` is executed.
The code below is supposed to show a warning for temperatures above 70, but there is a bug.
Find two different ways to fix the code, so that the warning is displayed.
```
temperature = 120
if temperature > 0:
print("it's warm")
elif temperature <= 0:
print("it's freezing")
elif temperature > 70:
print("WARNING: dangerously hot")
```
## `boolean` Expressions
The comparisons that are part of the if statements in the examples are Boolean expressions.
Boolean expressions include comparisons (`>`, `<`), equality (`==`) and inequality (`!=`).
Boolean expressions evaluate to `True` or `False`.
### `boolean` Connectors
We can use the `boolean` connectors or operators to build larger expressions.
The boolean connectors in Python are `and`, `or` and `not`.
```
warm = True
cloudy = False
print(warm and cloudy)
print(warm or cloudy)
if warm and not cloudy:
print("Remember sunscreen!")
```
### <span style="color:green"> Exercise: Boolean Operators </span>
Again we look at the temperature test.
This time, use a Boolean operator to fix this test so that the warning is displayed.
```
temperature = 120
if temperature > 0:
print("it's warm")
elif temperature <= 0:
print("it's freezing")
elif temperature > 70:
print("WARNING: dangerously hot")
```
### <span style="color:green"> Case Law Exercise: count dissenting opinions </span>
In the code below, we loop through a list of cases from the Case Law Api, then
loop through the opinions for each of those cases. Each `opinion` has a `"type"`
field which describes if it's a majority opinion, dissenting opinion or concurring opinion.
First, try to run the code below to check if you can print out the value of this field for each opinion:
```
import requests
import json
URL = "https://api.case.law/v1/cases/?jurisdiction=ill&full_case=true&decision_date_min=2011-01-01&page_size=20"
data = requests.get(URL).json()
cases = data["results"]
for case in cases:
opinions = case["casebody"]["data"]["opinions"]
for opinion in opinions:
print(opinion["type"])
```
Now, try to modify the code below to count the number of dissenting opinions by using an `if` test with `opinion["type"]`.
If you find a dissent, you will need to increase the variable `dissent_count`:
```
import requests
import json
URL = "https://api.case.law/v1/cases/?jurisdiction=ill&full_case=true&decision_date_min=2011-01-01&page_size=20"
data = requests.get(URL).json()
dissent_count = 0
cases = data["results"]
for case in cases:
opinions = case["casebody"]["data"]["opinions"]
for opinion in opinions:
'Your code here'
print("Number of dissents:", dissent_count)
```
### <span style="color:green"> Library Data Exercise: Count Fulltext Documents </span>
In the code below, we loop through a list of items from the National Library API.
Each `item` has a dictionary `accessInfo`, containing a key `isDigital`.
The corresponding value is a Boolean which is `True` if the document is available digitally in fulltext.
First, try to run the code below to check if you can print out the value of `isDigital` for each item:
```
import requests
import json
URL = "https://api.nb.no/catalog/v1/items?size=20&filter=mediatype:b%C3%B8ker&q=Bing,Jon"
data = requests.get(URL).json()
embedded = data['_embedded']
items = embedded['items']
for item in items:
accessInfo = item['accessInfo']
isDigital = accessInfo['isDigital']
print(isDigital)
```
Now, try to modify the code below to count the number of digital fulltext documents by using an `if` test with `isDigital`.
If you find a digital document, you will need to increase the variable `fulltext_count`:
```
import requests
import json
URL = "https://api.nb.no/catalog/v1/items?size=20&filter=mediatype:b%C3%B8ker&q=Bing,Jon"
data = requests.get(URL).json()
embedded = data['_embedded']
items = embedded['items']
fulltext_count = 0
for item in items:
accessInfo = item['accessInfo']
isDigital = accessInfo['isDigital']
# your code here
```
## <span style="color:blue">Key Points</span>
- We use `if`-statements to control program flow
- `if`-statements can have an `else`-part
- We can chain multiple `if`-statements with `elif`
- `if`-statements use Boolean expressions, which can be `True` or `False`
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
```
# **COVID-19 Twitter Sentiments**
# A. **Problem**: Do Twitter-tweet sentiments have any correlations with COVID19 death counts? That is, do states with higher death counts have a particular sentiment correlated to its tweets?
# **B. Datasets used**
## Tweet Source:
I constructed the twextual dataset by using a guide on twitter webscraping. I used the Twint library to construct a twitter webscraper that did not need to use Twitter's API.
https://pypi.org/project/twint/
Twint allowed me to filter by tweet date, querey (keyword being COIVD19), number of tweets that are to be scraped, location of the tweet (state), and finally an output file in `.csv` of the scarped data.
The code can be found on my github. The code may be ran in an UNIX-Based OS via terminal. If that's not possible, one could make an args data class and delete the argparse part from the code.
Code: https://github.com/kwxk/twitter-textual-scraper with comments for each line.
Here is the general format for the crawler within the argparse of the code:
`python tweet_crawler -q [write here query] -d [write here since date] -c [write here how many tweets you want from each state] -s [provide here a list of states each state between quotation marks] -o [write here output file name]`
So for example: `python tweet_crawler -q covid19 -d 2020-01-01 -c 100 -s "New Jersey" "Florida" -o output.csv`
Tweets were collected from a year to date (Decemeber 01, 2021).
**I treated this as if it were an ETL pipeline.**
## **Tweet Dataset**
The main dataset must be split between states and english (en) tweets must be preserved in each dataset.
### **Main tweet data frame**
```
df = pd.read_csv('covid19.csv')
df.head()
```
## **Split Tweet Dataframe (split by states)**
```
flp = df[df['near'].str.contains('Florida',na=False)]
fl = flp[flp['language'].str.contains('en',na=False)]
fl
txp = df[df['near'].str.contains('Texas',na=False)]
tx = txp[txp['language'].str.contains('en',na=False)]
tx
njp = df[df['near'].str.contains('New Jersey',na=False)]
njp = njp[njp['language'].str.contains('en',na=False)]
nj
nyp = df[df['near'].str.contains('New York',na=False)]
ny = nyp[nyp['language'].str.contains('en',na=False)]
ny
```
## **Stopwords**
```
### Stopwords List
stop= open("stopwords.txt").read().replace("\n",' ').split(" ")[:-1]
stat = pd.read_csv('us-states.csv')
stat
ny_stat = stat[stat['state'].str.contains('New York',na=False)]
nystat2 = ny_stat.drop(['fips','cases','state'], axis = 1)
nj_stat = stat[stat['state'].str.contains('New Jersey',na=False)]
njstat2 = nj_stat.drop(['fips','cases','state'], axis = 1)
tx_stat = stat[stat['state'].str.contains('Texas',na=False)]
txstat2 = tx_stat.drop(['fips','cases','state'], axis = 1)
fl_stat = stat[stat['state'].str.contains('Florida',na=False)]
flstat2 = fl_stat.drop(['fips','cases','state'], axis = 1)
fl_stat = stat[stat['state'].str.contains('Florida',na=False)]
fl_stat
```
## **Sentiment Analysis**
```
from nltk.stem.wordnet import WordNetLemmatizer
from gensim import corpora, models
from nltk.tokenize import word_tokenize
import gensim
#import pyLDAvis.gensim_models as gensimvis
from gensim import corpora
from matplotlib.patches import Rectangle
import pandas as pd
import numpy as np
import nltk
nltk.downloader.download('vader_lexicon')
nltk.downloader.download('stopwords')
from nltk.corpus import stopwords
from nltk.tokenize import RegexpTokenizer
from wordcloud import WordCloud, STOPWORDS
import matplotlib.colors as mcolors
import string
import matplotlib.pyplot as plt
import seaborn as sns
from wordcloud import WordCloud
import itertools
import collections
import re
import nltk
from nltk.corpus import stopwords
import re
import networkx
from textblob import TextBlob
df1= df.copy()
df1.drop(["Unnamed: 0","username","link","language"],axis=1, inplace=True)
df1
def cleaner(text):
text = re.sub("@[A-Za-z0-9]+","",text)
text = re.sub("#[A-Za-z0-9]+","",text)#Remove @ sign
text = re.sub(r"(?:\@|http?\://|https?\://|www)\S+", "", text) #Remove http links
text = text.replace("\n", "")
text=text.lower()
return text
df1['tweet'] = df1['tweet'].map(lambda x: cleaner(x))
punct = "\n\r"+string.punctuation
df1['tweet'] = df1['tweet'].str.translate(str.maketrans('','',punct))
def clean_sentence(val):
"remove chars that are not letters or number or down case then remove"
regex=re.compile('([^\sa-zA-Z0-9]|_)+')
sentence=regex.sub('',val).lower()
sentence = sentence.replace('\n'," ")
sentence = sentence.replace(','," ")
sentence = sentence.replace('\\~'," ")
sentence = sentence.replace('QAQ\\~\\~'," ")
sentence=sentence.split(" ")
for word in list(sentence):
if word in stop:
sentence.remove(word)
sentence=" ".join(sentence)
return sentence
def clean_dataframe(data):
"drop nans,thenb apply clean sentence function to description"
# data=data.dropna(how="any")
for col in ['tweet']:
df1[col]=df1[col].apply(clean_sentence)
return df1
cleaned_text = clean_dataframe(df1)
# Create textblob objects of the tweets
sentiment_objects = [TextBlob(text) for text in df1['tweet']]
sentiment_objects[2].polarity, sentiment_objects[0]
sentiment_values = [[text.sentiment.polarity, str(text)] for text in sentiment_objects]
sentiment_df = pd.DataFrame(sentiment_values, columns=["polarity", "tweet"])
a=sentiment_df['polarity'].round(2)
b=list(a)
sentiment_df['Polar']=b
new_list=[]
for i in range(len(sentiment_df['Polar'])):
a = sentiment_df['Polar'][i]
if a == 0:
new_list.append("Neutral")
continue
if a >0:
new_list.append("Positive")
continue
if a <0:
new_list.append("Negative")
continue
sentiment_df['Sentiments']=new_list
sentiment_df
df1['Sentiments']=sentiment_df['Sentiments']
df1['Polar']=sentiment_df['Polar']
df1
```
## **Florida Sentiments Analysis**
```
df_fl= df1[df1['near']=="Florida"].reset_index(drop=True)
df_fl
```
## **New York Sentiments Analysis**
```
df_ny= df1[df1['near']=="New York"].reset_index(drop=True)
df_ny
```
## **New Jersy Sentiments Analysis**
```
df_nj= df1[df1['near']=="New Jersey"].reset_index(drop=True)
df_nj
```
## **Texas Sentiments Analysis**
```
df_tx= df1[df1['near']=="Texas"].reset_index(drop=True)
df_tx
```
# **C. Findings:**
## **Overall Sentiments among all states**
```
## Visualizing the Text sentiments
pos=df1[df1['Sentiments']=='Positive']
neg=df1[df1['Sentiments']=='Negative']
neu=df1[df1['Sentiments']=='Neutral']
import plotly.express as px
#Frist_Day = Frist_Day
fig = px.pie(df1, names='Sentiments')
fig.show()
plt.title('Total number of tweets and sentiments')
plt.xlabel('Emotions')
plt.ylabel('Number of Tweets')
sns.countplot(x='Sentiments', data=df1)
```
**Finding: Neutral Sentiments are the most prevalent of sentiments from the combine dataframe of NJ, NY, FL, TX tweets. There are slightly more positive sentiments than negative sentiments.**
```
df1['near'].unique()
```
## **Barplot for the Sentiments (New Jersey)**
```
b=df_nj['Sentiments'].value_counts().reset_index()
plt.title('NJ number of tweets and sentiments')
plt.xlabel('Emotions')
plt.ylabel('Number of Tweets')
plt.bar(x=b['index'], height=b['Sentiments'])
```
**Findings: New Jersey has a majority neutral sentiment tweets from the dataframe. It has slightly more positive sentiment tweets than there are negative sentiment tweets.**
## **Barplot for the Sentiments (New York)**
```
b=df_ny['Sentiments'].value_counts().reset_index()
plt.title('NY number of tweets and sentiments')
plt.xlabel('Emotions')
plt.ylabel('Number of Tweets')
plt.bar(x=b['index'], height=b['Sentiments'])
```
**Findings: New York has a majority neutral sentiment tweets from the dataframe. It has more positive sentiment tweets than there are negative sentiment tweets.**
```
stat
df1
import datetime
lst=[]
#df1['date'] = datetime.datetime.strptime(df1['date'], '%Y-%m-%d' )
for i in range(len(df1)):
dat= datetime.datetime.strptime(df1['date'][i], '%Y-%m-%d %H:%M:%S')
df1['date'][i]= dat.date()
df1.sort_values(by='date').reset_index(drop=True)
a= ['New Jersey', 'Florida', 'Texas', 'New York']
lst=[]
for i in range(len(stat)):
if stat['state'][i] in a:
lst.append(i)
df_stat= stat.iloc[lst].reset_index(drop=True)
df_stat
```
## **Barplot for the Sentiments (Texas)**
```
b=df_tx['Sentiments'].value_counts().reset_index()
plt.title('TX number of tweets and sentiments')
plt.xlabel('Emotions')
plt.ylabel('Number of Tweets')
plt.bar(x=b['index'], height=b['Sentiments'])
```
**Findings: Texas has a majority neutral sentiment tweets from the dataframe. It has slightly more negative sentiment tweets than there are positive sentiment tweets.**
## **Barplot for the Sentiments (Florida)**
```
b=df_fl['Sentiments'].value_counts().reset_index()
plt.title('FL number of tweets and sentiments')
plt.xlabel('Emotions')
plt.ylabel('Number of Tweets')
plt.bar(x=b['index'], height=b['Sentiments'])
```
**Findings: Florida has a majority neutral sentiment tweets from the dataframe. It has slightly more positive sentiment tweets than there are negative sentiment tweets.**
# **Total Covid Deaths Year to Date (Decemeber 4th)**
---
Source: https://github.com/nytimes/covid-19-data
```
import plotly.express as px
fig = px.line(df_stat, x='date', y='deaths', color='state')
fig.show()
```
The above graph shows the total covid deaths from 02/13/2020 until 12/04/2021 for the states of Texas, Florida, New York, and New Jersey. Texas and Florida have the two most deaths with Texas leading. New York and New Jersey have the least deaths with New York leading New Jersey with the most deaths out of the two.
## **Initial Questions ▶**
1. **Would it stand to reason that the states with more positive-neutral sentiments toward COVID-19 had lower total deaths?**
There are no correlations between tweet sentiments and total deaths according to the curated dataset. Looking at the CSV dataset from the New York Times' Github on the total COVID-19 deaths for the states of Texas, Florida, New York, and New Jersey, it shows that Texas and Florida are top out of the states in terms of the total death count. Texas and Florida had different Positive to Negative sentiments as is apparent from the graph.
2. **Which state had a higher infection death count?**
Texas has the higher death count out of all of the states. New Jersey has the least.
3. **Which states had more negative than positive twitter sentiment to 'COVID-19' in their dataset?**
Texas was the only state that had more negative twitter sentiments in its dataset than positive.
4. **What was the most common sentiment in all datasets?**
Neutral sentiment tweet was the most popular category having much of the tweets in the total dataset: 45.7% of textual data was neutral. 28% of the total textual dataset was positive in sentiment and 26.3% was negative.
5. **Are the sentiment results correlated or related to total death count?**
No. There are no correlations/realtions between sentiment and total death counts. In the Texas dataset, its graph observed more negative sentiments than positive. In the Florida dataset, its graph observed more positive sentiments than negative.
If we look at New Jersey and New York, both datasets have more positive than negative sentiment tweets. New York has considerably more positive tweets than negative tweets. NJ has slightly more positive tweets than negative tweets.
If we wanted to make a statement that states that have more positive tweets to negative tweets have higher total death counts, Texas would have to have that same trend. Texas breaks this trend such that there are more negative tweets than positive tweets in its dataset despite it having the highest total death count out of all of the states.
```
## Visualizing the Text sentiments
pos=df1[df1['Sentiments']=='Positive']
neg=df1[df1['Sentiments']=='Negative']
neu=df1[df1['Sentiments']=='Neutral']
import plotly.express as px
#Frist_Day = Frist_Day
fig = px.pie(df1, names='Sentiments')
fig.show()
```
# **D. Implications**
**For the hiring firm:**
According to the dataset, twitter sentiments alone cannot give any meaningful indication as to whether or not tweets and their emotions have any bearing on COVID-19's death total death count. Better methodologies must be made: perhaps tweets of a certain popularity (perhaps a ratio between likes, retweets, sharing, etc) should be curated into a dataset. Simply looking at tweets at random is a good measure against bias however there is too much statistical noise within the dataset to make any meaningful correlations.
**For Social Science:**
Better methodologies in general should be developed when looking at social media posts. Considerable weight should be given to popular/viral content when curating a dataset as that is a category of data that inherently has the most interaction and 'social proof' due to its popularity on the website.
# **E. Learning Outcomes**
The more I developed my analytical skills, the more I realized that my project had a lot of statsitcal noise. First, I should have developed a better methodolgy for curating tweets. I simply used TWINT to currated 1300+ tweets randomly according to a fixed criteria. I did not add factors such as popularity of a tweet or its general social-media interaction score (primarily because I do not know how to do that yet).
If I were to do this project again, I would start off by curating textual data that had a certain virality to it. I would alone curate tweets with specific likes, shares, and comments.
This would be a difficult task, as I don't know if twitter has an ELO score for tweets: If twitter had a virality ratio for a tweet I would likely curate on that factor as it would come from a class of textual data that has genderated a certain amount of influence.
However, this would add additional questions that would have to be considered as well: How much of the virality score would be coming from a particular state?
For instance, if a score of 10 is VERY viral and that tweet comes from New York, are New York twitter users responsible for that tweet being a score of 10 or could it be users from another geographic location? This is a fair question because I would want to know how much influence the tweet has in its geographic location. It may be possible to develope a webscraper capable of achieving this goal, but It may involve many calculations that still would not gaurantee the results being adequately parsed.
| github_jupyter |
Data source: https://www.kaggle.com/c/emvic/discussion/1730
# Imports
```
import matplotlib.pyplot as plt
import numpy as np
import math
from fixation_saccade_classifier import *
```
# Get data
```
def read_data(file_name):
f = open(file_name)
f.readline()
samples = []
target = []
for line in f:
line = line.strip().split(",")
sample = [float(x) for x in line[1:]]
samples.append(sample)
target.append(line[0])
return (samples, target)
training, target = read_data("train.csv")
lx, ly, rx, ry = [], [], [], []
for i in range(len(training)):
lx.append(training[i][:2048])
ly.append(training[i][2048:4096])
rx.append(training[i][4096:6144])
ry.append(training[i][6144:8192])
target = np.array(target)
lx = np.array(lx[0])
ly = np.array(ly[0])
rx = np.array(rx[0])
ry = np.array(ry[0])
```
# I-DT
```
classifier = IDTFixationSaccadeClassifier(threshold = 100.0, win_len = 50)
fixations, saccades, fixation_colors, saccades_colors = classifier.fit_predict(lx, ly)
plt.scatter(lx, ly, alpha = 0.1)
plt.scatter(lx[fixations], ly[fixations], c=fixation_colors, alpha = 0.5)
plt.scatter(lx[saccades], ly[saccades], c=saccades_colors, alpha = 0.5)
plt.figure(figsize = (10, 10))
plt.plot(saccades, lx[saccades], marker='o', color = 'red', alpha = 0.25)
plt.scatter(fixations, lx[fixations], marker='+')
```
# I-VT
```
classifier = IVTFixationSaccadeClassifier(threshold = 15.0)
fixations, saccades, fixation_colors, saccades_colors = classifier.fit_predict(lx, ly)
plt.scatter(lx, ly, alpha = 0.1)
plt.scatter(lx[fixations], ly[fixations], c=fixation_colors, alpha = 0.5)
plt.scatter(lx[saccades], ly[saccades], c=saccades_colors, alpha = 0.5)
plt.figure(figsize = (10, 10))
plt.plot(saccades, lx[saccades], marker='o', color = 'red', alpha = 0.25)
plt.scatter(fixations, lx[fixations], marker='+')
```
# I-HMM
```
classifier = IHMMFixationSaccadeClassifier(fix_median = 1.0,
fix_variance = 10.0,
sacc_median = 80.0,
sacc_variance = 60.0,
prob_fix_fix = math.log(0.95),
prob_sacc_sacc = math.log(0.95),
prob_fix_sacc = math.log(0.95),
prob_sacc_fix = math.log(0.95))
fixations, saccades, fixation_colors, saccades_colors = classifier.fit_predict(lx, ly)
plt.scatter(lx, ly, alpha = 0.1)
plt.scatter(lx[fixations], ly[fixations], c=fixation_colors, alpha = 0.5)
plt.scatter(lx[saccades], ly[saccades], c=saccades_colors, alpha = 0.5)
plt.figure(figsize = (10, 10))
plt.plot(saccades, lx[saccades], marker='o', color = 'red', alpha = 0.25)
plt.scatter(fixations, lx[fixations], marker='+')
```
# I-AOI
```
classifier = IAOIFixationSaccadeClassifier(threshold = 15.0, areas = [[-100.0, -100.0, 100.0, 100.0]])
fixations, fixation_colors = classifier.fit_predict(lx, ly)
plt.scatter(lx, ly, alpha = 0.1)
plt.xlim([-200, 200])
plt.ylim([-200, 200])
plt.scatter(lx[fixations], ly[fixations], c=fixation_colors, alpha = 0.5)
plt.show()
plt.figure(figsize = (10, 10))
plt.plot(saccades, lx[saccades], marker='o', color = 'red', alpha = 0.25)
plt.scatter(fixations, lx[fixations], marker='+')
```
# I-VT (window)
```
classifier = IWVTFixationSaccadeClassifier(threshold = 15.0, win_len = 10)
fixations, saccades, fixation_colors, saccades_colors = classifier.fit_predict(lx, ly)
plt.scatter(lx, ly, alpha = 0.1)
plt.scatter(lx[fixations], ly[fixations], c=fixation_colors, alpha = 0.5)
plt.scatter(lx[saccades], ly[saccades], c=saccades_colors, alpha = 0.5)
plt.figure(figsize = (10, 10))
plt.plot(saccades, lx[saccades], marker='o', color = 'red', alpha = 0.25)
plt.scatter(fixations, lx[fixations], marker='+')
```
# How to transform result to separate fixations/saccades
```
fixations = np.array(fixations)
fixation_colors = np.array(fixation_colors)
ans = []
for elem in np.unique(fixation_colors):
ans.append(fixations[fixation_colors == elem])
ans
```
| github_jupyter |
### Integrate plot
Qarpo is a library to build a jupyter notebook user interface to submit jobs to job scheduler, display output interface to display accomplished jobs' outputs and plot its results.
This notebook provides a recipe to integrate plot displaying the results of accomplished jobs in the jupyter notebook
To start using the qarpo, run the following import line
```
import qarpo
```
The plot in qarpo UI interface consists of 2 main parts. The first part is the backend, which writes the resulted output to a file, in our example here, we are writing the time, fps and total number of frames into stats.json file. These code lines are integarted into the python script running.
```python
import json
import time
t1 = time.time()
//Inference execution
infer_time = time.time()-t1
stats = {}
stats['time'] = str(infer_time)
stats['frame'] = str(num_frames)
stats['fps'] = str(num_frames / infer_time)
stats_file = "results/{}/stats.json".format(job_id)
with open(stats_file, 'w') as f:
json.dump(stats, f)
```
The second part is defined in the UI configuration, this UI configuration is an input to the class constructor Interface.
To add the plot configuratoion to the UI configuration, use the following format:
{
"job": # Define how to launch the job and interpret results
{
"output_type": ".txt", # The type of input (text/video)
"results_path": "app/results/", # Path to job result files
"plots": #list of dictionaries, each dictionary represents a plot configuration
[
{
"title" : < plot title >,
"type" : <plot type, "time" or "fps" or any different value specified in the json file in the backend part>,
"xlabel" : <x-axis label>,
"ylabel" : <y-axis label>
}
]
}
}
```
job_interface = qarpo.Interface( {
"job": # Define how to launch the job and interpret results
{
"output_type": ".png", # The type of input (text/video)
"results_path": "app/results/", # Path to job result files
"plots":[
{
"title" : "",
"type" : "time",
"xlabel" : "Job ID",
"ylabel" : "Time in seconds"
}
]
}
} )
job_interface.displayUI()
job_interface.submitJob("qsub app/example_job.sh -l nodes=1:idc001skl:i5-6500te -F 'app/results/'")
```
| github_jupyter |
# Popular Data Science Questions
The goal in this project is to use [Data Science Stack Exchange](https://datascience.stackexchange.com) to determine what the most popular data science categories are being searched and talked about.
## Stack Exchange
# Exploring the Data
We can read in the data while immediately making sure `CreationDate` will be stored as a datetime object:
```
# We import everything that we'll use
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
questions = pd.read_csv("2019_questions.csv", parse_dates=["CreationDate"])
```
Running [`questions.info()`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.info.html) should gives a lot of useful information.
```
questions.info()
```
We see that only `FavoriteCount` has missing values. A missing value on this column probably means that the question was is not present in any users' favorite list, so we can replace the missing values with zero.
The types seem adequate for every column, however, after we fill in the missing values on `FavoriteCount`, there is no reason to store the values as floats.
Since the `object` dtype is a catch-all type, let's see what types the objects in `questions["Tags"]` are.
```
questions["Tags"].apply(lambda value: type(value)).unique()
```
We see that every value in this column is a string. On Stack Exchange, each question can only have a maximum of five tags ([source](https://meta.stackexchange.com/a/18879)), so one way to deal with this column is to create five columns in `questions` called `Tag1`, `Tag2`, `Tag3`, `Tag4`, and `Tag5` and populate the columns with the tags in each row.
However, since doesn't help is relating tags from one question to another, we'll just keep them as a list.
# Cleaning the Data
We'll begin by fixing `FavoriteCount`.
```
questions.fillna(value={"FavoriteCount": 0}, inplace=True)
questions["FavoriteCount"] = questions["FavoriteCount"].astype(int)
questions.dtypes
```
Let's now modify `Tags` to make it easier to work with.
```
questions["Tags"] = questions["Tags"].str.replace("^<|>$", "").str.split("><")
questions.sample(3)
```
# Most Used and Most Viewed
We'll begin by counting how many times each tag was used
```
tag_count = dict()
for tags in questions["Tags"]:
for tag in tags:
if tag in tag_count:
tag_count[tag] += 1
else:
tag_count[tag] = 1
```
For improved aesthetics, let's transform `tag_count` in a dataframe.
```
tag_count = pd.DataFrame.from_dict(tag_count, orient="index")
tag_count.rename(columns={0: "Count"}, inplace=True)
tag_count.head(10)
```
Let's now sort this dataframe by `Count` and visualize the top 20 results.
```
most_used = tag_count.sort_values(by="Count").tail(20)
most_used
```
The threshold of `20` is somewhat arbitrary and we can experiment with others, however, popularity of the tags rapidly declines, so looking at these tags should be enough to help us with our goal. Let's visualize these data.
```
most_used.plot(kind="barh", figsize=(16,8))
```
Some tags are very, very broad and are unlikely to be useful; e.g.: `python`, `dataset`, `r`. Before we investigate the tags a little deeper, let's repeat the same process for views.
```
some_iterable = "Iterate this!"
for i,c in enumerate(some_iterable):
print(i,c)
```
In addition to the elements of `some_iterable`, `enumerate` gives us the index of each of them.
```
tag_view_count = dict()
for idx, tags in enumerate(questions["Tags"]):
for tag in tags:
if tag in tag_view_count:
tag_view_count[tag] += questions["ViewCount"].iloc[idx]
else:
tag_view_count[tag] = questions["ViewCount"].iloc[idx]
tag_view_count = pd.DataFrame.from_dict(tag_view_count, orient="index")
tag_view_count.rename(columns={0: "ViewCount"}, inplace=True)
most_viewed = tag_view_count.sort_values(by="ViewCount").tail(20)
most_viewed.plot(kind="barh", figsize=(16,8))
```
Let's see them side by side.
```
fig, axes = plt.subplots(nrows=1, ncols=2)
fig.set_size_inches((24, 10))
most_used.plot(kind="barh", ax=axes[0], subplots=True)
most_viewed.plot(kind="barh", ax=axes[1], subplots=True)
in_used = pd.merge(most_used, most_viewed, how="left", left_index=True, right_index=True)
in_viewed = pd.merge(most_used, most_viewed, how="right", left_index=True, right_index=True)
```
# Relations Between Tags
One way of trying to gauge how pairs of tags are related to each other, is to count how many times each pair appears together. Let's do this.
We'll begin by creating a list of all tags.
```
all_tags = list(tag_count.index)
```
We'll now create a dataframe where each row will represent a tag, and each column as well. Something like this:
<table class="dataframe" border="1">
<thead>
<tr style="text-align: right;">
<th></th>
<th>tag1</th>
<th>tag2</th>
<th>tag3</th>
</tr>
</thead>
<tbody>
<tr>
<th>tag1</th>
<td></td>
<td></td>
<td></td>
</tr>
<tr>
<th>tag2</th>
<td></td>
<td></td>
<td></td>
</tr>
<tr>
<th>tag3</th>
<td></td>
<td></td>
<td></td>
</tr>
</tbody>
</table>
```
associations = pd.DataFrame(index=all_tags, columns=all_tags)
associations.iloc[0:4,0:4]
```
We will now fill this dataframe with zeroes and then, for each lists of tags in `questions["Tags"]`, we will increment the intervening tags by one. The end result will be a dataframe that for each pair of tags, it tells us how many times they were used together.
```
associations.fillna(0, inplace=True)
for tags in questions["Tags"]:
associations.loc[tags, tags] += 1
```
This dataframe is quite large. Let's focus our attention on the most used tags. We'll add some colors to make it easier to talk about the dataframe. (At the time of this writing, GitHub's renderer does not display the colors, we suggest you use this solution notebook together with [JupyterLab](https://jupyterlab.readthedocs.io/en/stable/)).
```
relations_most_used = associations.loc[most_used.index, most_used.index]
def style_cells(x):
helper_df = pd.DataFrame('', index=x.index, columns=x.columns)
helper_df.loc["time-series", "r"] = "background-color: yellow"
helper_df.loc["r", "time-series"] = "background-color: yellow"
for k in range(helper_df.shape[0]):
helper_df.iloc[k,k] = "color: blue"
return helper_df
relations_most_used.style.apply(style_cells, axis=None)
```
The cells highlighted in yellow tell us that `time-series` was used together with `r` 22 times. The values in blue tell us how many times each of the tags was used. We saw earlier that `machine-learning` was used 2693 times and we confirm it in this dataframe.
```
for i in range(relations_most_used.shape[0]):
relations_most_used.iloc[i,i] = pd.np.NaN
plt.figure(figsize=(12,8))
sns.heatmap(relations_most_used, cmap="Greens", annot=False)
```
The most used tags also seem to have the strongest relationships, as given by the dark concentration in the bottom right corner. However, this could simply be because each of these tags is used a lot, and so end up being used together a lot without possibly even having any strong relation between them.
# Enter Domain Knowledge
[Keras](https://keras.io/), [scikit-learn](https://scikit-learn.org/), [TensorFlow](https://www.tensorflow.org/) are all Python libraries that allow their users to employ deep learning (a type of neural network).
Most of the top tags are all intimately related with one central machine learning theme: deep learning. If we want to be very specific, we can suggest the creation of Python content that uses deep learning for classification problems (and other variations of this suggestion).
# Just a Fad?
Let's read in the file into a dataframe called `all_q`. We'll parse the dates at read-time.
```
all_q = pd.read_csv("all_questions.csv", parse_dates=["CreationDate"])
```
We can use the same technique as before to clean the tags column.
```
all_q["Tags"] = all_q["Tags"].str.replace("^<|>$", "").str.split("><")
```
Before deciding which questions should be classified as being deep learning questions, we should decide what tags are deep learning tags.
The definition of what constitutes a deep learning tag we'll use is: a tag that belongs to the list `["lstm", "cnn", "scikit-learn", "tensorflow", "keras", "neural-network", "deep-learning"]`.
This list was obtained by looking at all the tags in `most_used` and seeing which ones had any relation to deep learning. You can use Google and read the tags descriptions to reach similar results.
We'll now create a function that assigns `1` to deep learning questions and `0` otherwise; and we use it.
```
def class_deep_learning(tags):
for tag in tags:
if tag in ["lstm", "cnn", "scikit-learn", "tensorflow",
"keras", "neural-network", "deep-learning"]:
return 1
return 0
all_q["DeepLearning"] = all_q["Tags"].apply(class_deep_learning)
all_q.sample(5)
```
Looks good!
The data-science-techonology landscape isn't something as dynamic to merit daily, weekly, or even monthly tracking. Let's track it quarterly.
Since we don't have all the data for the first quarter of 2020, we'll get rid of those dates:
```
all_q = all_q[all_q["CreationDate"].dt.year < 2020]
```
Let's create a column that identifies the quarter in which a question was asked.
```
def fetch_quarter(datetime):
year = str(datetime.year)[-2:]
quarter = str(((datetime.month-1) // 3) + 1)
return "{y}Q{q}".format(y=year, q=quarter)
all_q["Quarter"] = all_q["CreationDate"].apply(fetch_quarter)
all_q.head()
```
For the final stretch of this screen, we'll group by quarter and:
* Count the number of deep learning questions.
* Count the total number of questions.
* Compute the ratio between the two numbers above.
```
quarterly = all_q.groupby('Quarter').agg({"DeepLearning": ['sum', 'size']})
quarterly.columns = ['DeepLearningQuestions', 'TotalQuestions']
quarterly["DeepLearningRate"] = quarterly["DeepLearningQuestions"]\
/quarterly["TotalQuestions"]
# The following is done to help with visualizations later.
quarterly.reset_index(inplace=True)
quarterly.sample(5)
ax1 = quarterly.plot(x="Quarter", y="DeepLearningRate",
kind="line", linestyle="-", marker="o", color="orange",
figsize=(24,12)
)
ax2 = quarterly.plot(x="Quarter", y="TotalQuestions",
kind="bar", ax=ax1, secondary_y=True, alpha=0.7, rot=45)
for idx, t in enumerate(quarterly["TotalQuestions"]):
ax2.text(idx, t, str(t), ha="center", va="bottom")
xlims = ax1.get_xlim()
ax1.get_legend().remove()
handles1, labels1 = ax1.get_legend_handles_labels()
handles2, labels2 = ax2.get_legend_handles_labels()
ax1.legend(handles=handles1 + handles2,
labels=labels1 + labels2,
loc="upper left", prop={"size": 12})
for ax in (ax1, ax2):
for where in ("top", "right"):
ax.spines[where].set_visible(False)
ax.tick_params(right=False, labelright=False)
```
| github_jupyter |
# Наработки
```
import open3d as o3d
import numpy as np
def convert_from_bin_to_pcd(path_to_binary_file: str, path_to_new_pcd_file: str):
bin_pcd = np.fromfile(path_to_binary_file, dtype=np.float32)
points = bin_pcd.reshape((-1, 4))[:, 0:3]
o3d_pcd = o3d.geometry.PointCloud(o3d.utility.Vector3dVector(points))
o3d.io.write_point_cloud(path_to_new_pcd_file, o3d_pcd)
def convert_from_pcd_to_ply(path_to_pcd_file: str, path_to_new_ply_file: str):
pcd = o3d.io.read_point_cloud(path_to_pcd_file)
o3d.io.write_point_cloud(path_to_new_ply_file, pcd)
def scan_next_plane_from_point_cloud(
point_cloud: o3d.geometry.PointCloud, distance: float = 0.1
):
try:
_, inliers = point_cloud.segment_plane(
distance_threshold=distance, ransac_n=3, num_iterations=5000
)
except Exception:
return (point_cloud, point_cloud.clear())
inlier_cloud = point_cloud.select_by_index(inliers)
inlier_cloud.paint_uniform_color([1.0, 0, 0])
outlier_cloud = point_cloud.select_by_index(inliers, invert=True)
return (inlier_cloud, outlier_cloud)
def select_points_by_label_id(
path_to_pcd_file: str, path_to_label_file: str, label_id: int
) -> o3d.geometry.PointCloud:
pcd = o3d.io.read_point_cloud(path_to_pcd_file)
labels = np.fromfile(path_to_label_file, dtype=np.uint32)
labels = labels.reshape((-1))
pcd_point_by_id = pcd.select_by_index(np.where(labels == label_id)[0])
return pcd_point_by_id
def segment_all_planes_from_point_cloud(point_cloud: o3d.geometry.PointCloud) -> list:
all_planes = []
inlier_cloud, outlier_cloud = scan_next_plane_from_point_cloud(point_cloud)
while outlier_cloud.has_points():
all_planes.append(inlier_cloud)
inlier_cloud, outlier_cloud = scan_next_plane_from_point_cloud(outlier_cloud)
if inlier_cloud.has_points():
all_planes.append(inlier_cloud)
return all_planes
def segment_all_planes_by_list_of_labels(
path_to_pcd_file: str, path_to_label_file: str, list_of_labels: list
) -> dict:
result_dict = {}
for label in list_of_labels:
point_cloud = select_points_by_label_id(
path_to_pcd_file, path_to_label_file, label
)
planes = extract_all_planes_from_point_cloud(point_cloud)
result_dict[label] = planes
return result_dict
path_to_bin_file = "/home/pavel/dataset/sequences/00/velodyne/000000.bin"
path_to_pcd_file = "/home/pavel/Point-Cloud/src/test.pcd"
path_to_label_file = "/home/pavel/dataset/sequences/00/labels/000000.label"
BUILDING_LABEL = 50
OTHER_STRUCTURE_LABEL = 52
ROAD_LABEL = 40
list_of_planes = [BUILDING_LABEL, OTHER_STRUCTURE_LABEL, ROAD_LABEL]
segmented_planes = segment_all_planes_by_list_of_labels(
path_to_pcd_file, path_to_label_file, list_of_planes
)
len(segmented_planes)
```
| github_jupyter |

# **Amazon SageMaker in Practice - Workshop**
## **Click-Through Rate Prediction**
This lab covers the steps for creating a click-through rate (CTR) prediction pipeline. The source code of the workshop prepared by [Pattern Match](https://pattern-match.com) is available on the [company's Github account](https://github.com/patternmatch/amazon-sagemaker-in-practice).
You can reach authors us via the following emails:
- [Sebastian Feduniak](mailto:sebastian.feduniak@pattern-match.com)
- [Wojciech Gawroński](mailto:wojciech.gawronski@pattern-match.com)
- [Paweł Pikuła](mailto:pawel.pikula@pattern-match.com)
Today we use the [Criteo Labs](http://labs.criteo.com/) dataset, used for the old [Kaggle competition](https://www.kaggle.com/c/criteo-display-ad-challenge) for the same purpose.
**WARNING**: First you need to update `pandas` to 0.23.4 for the `conda_python3` kernel.
# Background
In advertising, the most critical aspect when it comes to revenue is the final click on the ad. It is one of the ways to compensate for ad delivery for the provider. In the industry, an individual view of the specific ad is called an *impression*.
To compare different algorithms and heuristics of ad serving, "clickability" of the ad is measured and presented in the form of [*click-through rate* metric (CTR)](https://en.wikipedia.org/wiki/Click-through_rate):

If you present randomly sufficient amount of ads to your user base, you get a baseline level of clicks. It is the easiest and simple solution. However, random ads have multiple problems - starting with a lack of relevance, causing distrust and annoyance.
**Ad targeting** is a crucial technique for increasing the relevance of the ad presented to the user. Because resources and a customer's attention is limited, the goal is to provide an ad to most interested users. Predicting those potential clicks based on readily available information like device metadata, demographics, past interactions, and environmental factors is a universal machine learning problem.
# Steps
This notebook presents an example problem to predict if a customer clicks on a given advertisement. The steps include:
- Prepare your *Amazon SageMaker* notebook.
- Download data from the internet into *Amazon SageMaker*.
- Investigate and transforming the data for usage inside *Amazon SageMaker* algorithms.
- Estimate a model using the *Gradient Boosting* algorithm (`xgboost`).
- Leverage hyperparameter optimization for training multiple models with varying hyperparameters in parallel.
- Evaluate and compare the effectiveness of the models.
- Host the model up to make on-going predictions.
# What is *Amazon SageMaker*?
*Amazon SageMaker* is a fully managed machine learning service. It enables discovery and exploration with use of *Jupyter* notebooks and then allows for very easy industrialization on a production-grade, distributed environment - that can handle and scale to extensive datasets.
It provides solutions and algorithms for existing problems, but you can bring your algorithms into service without any problem. Everything mentioned above happens inside your *AWS infrastructure*. That includes secure and isolated *VPC* (*Virtual Private Cloud*), supported by the full power of the platform.
[Typical workflow](https://docs.aws.amazon.com/sagemaker/latest/dg/how-it-works-mlconcepts.html) for creating machine learning models:
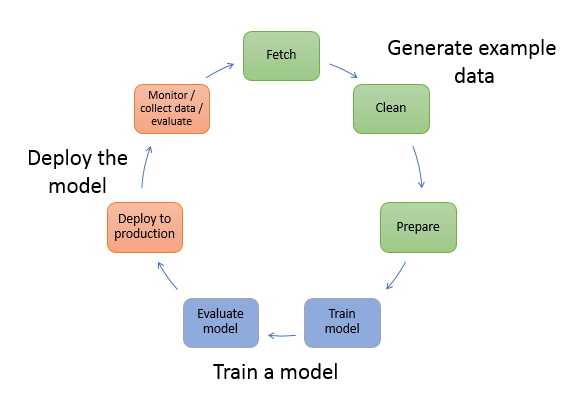
## Note about *Amazon* vs. *AWS* prefix
Why *Amazon* and not *AWS*?
Some services available in *Amazon Web Services* portfolio are branded by *AWS* itself, and some by Amazon.
Everything depends on the origin and team that maintains it - in that case, it originated from the core of the Amazon, and they maintain this service inside the core division.
## Working with *Amazon SageMaker* locally
It is possible to fetch *Amazon SageMaker SDK* library via `pip` and use containers provided by *Amazon* locally, and you are free to do it. The reason why and when you should use *Notebook Instance* is when your datasets are far more significant than you want to store locally and they are residing on *S3* - for such cases it is very convenient to have the *Amazon SageMaker* notebooks available.
# Preparation
The primary way for interacting with *Amazon SageMaker* is to use *S3* as storage for input data and output results.
For our workshops, we have prepared two buckets. One is a dedicated bucket for each user (see the credentials card you have received at the beginning of the workshop) - you should put the name of that bucket into `output_bucket` variable. That bucket is used for storing output models and transformed and split input datasets.
We have also prepared a shared bucket called `amazon-sagemaker-in-practice-workshop` which contains the input dataset inside a path presented below.
```
data_bucket = 'amazon-sagemaker-in-practice-workshop'
user_number = 'CHANGE_TO_YOUR_NUMBER'
user_name = 'user-{}'.format(user_number)
output_bucket = 'amazon-sagemaker-in-practice-bucket-{}'.format(user_name)
path = 'criteo-display-ad-challenge'
key = 'sample.csv'
data_location = 's3://{}/{}/{}'.format(data_bucket, path, key)
```
*Amazon SageMaker* as a service runs is a specific security context applied via *IAM role*. You have created that role when creating *notebook instance* before we have uploaded this content.
Each *notebook* instance provides a *Jupyter* environment with preinstalled libraries and *AWS SDKs*. One of such *SDKs* is *Amazon SageMaker SDK* available from the *Python* environment. With the use of that *SDK* we can check which security context we can use:
```
import boto3
from sagemaker import get_execution_role
role = get_execution_role()
print(role)
```
As a next, we need to import some stuff. It includes *IPython*, *Pandas*, *numpy*, commonly used libraries from *Python's* Standard Library and *Amazon SageMaker* utilities:
```
import numpy as np # For matrix operations and numerical processing
import pandas as pd # For munging tabular data
import matplotlib.pyplot as plt # For charts and visualizations
from IPython.display import Image # For displaying images in the notebook
from IPython.display import display # For displaying outputs in the notebook
from time import gmtime, strftime # For labeling SageMaker models, endpoints, etc.
import sys # For writing outputs to notebook
import math # For ceiling function
import json # For parsing hosting outputs
import os # For manipulating filepath names
import sagemaker # Amazon SageMaker's Python SDK provides helper functions
from sagemaker.predictor import csv_serializer # Converts strings for HTTP POST requests on inference
from sagemaker.tuner import IntegerParameter # Importing HPO elements.
from sagemaker.tuner import CategoricalParameter
from sagemaker.tuner import ContinuousParameter
from sagemaker.tuner import HyperparameterTuner
```
Now we are ready to investigate the dataset.
# Data
The training dataset consists of a portion of Criteo's traffic over a period of 7 days. Each row corresponds to a display ad served by Criteo and the first column indicates whether this ad was clicked or not. The positive (clicked) and negative (non-clicked) examples have both been subsampled (but at different rates) to reduce the dataset size.
There are 13 features taking integer values (mostly count features) and 26 categorical features. Authors hashed values of the categorical features onto 32 bits for anonymization purposes. The semantics of these features is undisclosed. Some features may have missing values (represented as a `-1` for integer values and empty string for categorical ones). Order of the rows is chronological.
You may ask, why in the first place we are investigating such *obfuscated* dataset. In *ad tech* it is not unusual to deal with anonymized, or pseudonymized data, which are not semantical - mostly due to privacy and security reasons.
The test set is similar to the training set but, it corresponds to events on the day following the training period. For that dataset author removed *label* (the first column).
Unfortunately, because of that, it is hard to guess for sure which feature means what, but we can infer that based on the distribution - as we can see below.
## Format
The columns are tab separeted with the following schema:
```
<label> <integer feature 1> ... <integer feature 13> <categorical feature 1> ... <categorical feature 26>
```
When a value is missing, the field is just empty. There is no label field in the test set.
Sample dataset (`sample.csv`) contains *100 000* random rows which are taken from a training dataset to ease the exploration.
## How to load the dataset?
Easy, if it is less than 5 GB - as the disk available on our Notebook instance is equal to 5 GB.
However, there is no way to increase that. :(
It is because of that EBS volume size is fixed at 5GB. As a workaround, you can use the `/tmp` directory for storing large files temporarily. The `/tmp` directory is on the root drive that has around 20GB of free space. However, data stored there cannot be persisted across stopping and restarting of the notebook instance.
What if we need more? We need to preprocess the data in another way (e.g., using *AWS Glue*) and store it on *S3* available for *Amazon SageMaker* training machines.
To read a *CSV* correctly we use *Pandas*. We need to be aware that dataset uses tabs as separators and we do not have the header:
```
data = pd.read_csv(data_location, header = None, sep = '\t')
pd.set_option('display.max_columns', 500) # Make sure we can see all of the columns.
pd.set_option('display.max_rows', 20) # Keep the output on one page.
```
## Exploration
Now we would like to explore our data, especially that we do not know anything about the semantics. How can we do that?
We can do that by reviewing the histograms, frequency tables, correlation matrix, and scatter matrix. Based on that we can try to infer and *"sniff"* the meaning and semantics of the particular features.
### Integer features
First 13 features from the dataset are represented as an integer features, let's review them:
```
# Histograms for each numeric features:
display(data.describe())
%matplotlib inline
hist = data.hist(bins = 30, sharey = True, figsize = (10, 10))
display(data.corr())
pd.plotting.scatter_matrix(data, figsize = (12, 12))
plt.show()
```
### Categorical features
Next 26 features from the dataset are represented as an categorical features. Now it's time to review those:
```
# Frequency tables for each categorical feature:
for column in data.select_dtypes(include = ['object']).columns:
display(pd.crosstab(index = data[column], columns = '% observations', normalize = 'columns'))
categorical_feature = data[14]
unique_values = data[14].unique()
print("Number of unique values in 14th feature: {}\n".format(len(unique_values)))
print(data[14])
```
As for *integer features*, we can push them as-is to the *Amazon SageMaker* algorithms. We cannot do the same thing for *categorical* one.
As you can see above, we have many unique values inside the categorical column. They hashed that into a *32-bit number* represented in a hexadecimal format - as a *string*.
We need to convert that into a number, and we can leverage *one-hot encoding* for that.
#### One-Hot Encoding
It is a way of converting categorical data (e.g., type of animal - *dog*, *cat*, *bear*, and so on) into a numerical one, one-hot encoding means that for a row we create `N` additional columns and we put a `1` if that category is applicable for such row.
#### Sparse Vectors
It is the more efficient way to store data points which are not dense and do not contain all features. It is possible to efficiently compute various operations between those two forms - dense and sparse.
### Problem with *one-hot encoding* in this dataset
Unfortunately, we cannot use *OHE* as-is for this dataset. Why?
```
for column in data.select_dtypes(include=['object']).columns:
size = data.groupby([column]).size()
print("Column '{}' - number of categories: {}".format(column, len(size)))
for column in data.select_dtypes(include=['number']).columns:
size = data.groupby([column]).size()
print("Column '{}' - number of categories: {}".format(column, len(size)))
```
We have too many distinct categories per feature! In the worst case, for an individual feature, we create couple hundred thousands of new columns. Even with the sparse representation it significantly affects memory usage and execution time.
What kind of features are represented by that? Examples of such features are *Device ID*, *User Agent* strings and similar.
How to workaround that? We can use *indexing*.
```
for column in data.select_dtypes(include = ['object']).columns:
print("Converting '{}' column to indexed values...".format(column))
indexed_column = "{}_index".format(column)
data[indexed_column] = pd.Categorical(data[column])
data[indexed_column] = data[indexed_column].cat.codes
categorical_feature = data['14_index']
unique_values = data['14_index'].unique()
print("Number of unique values in 14th feature: {}\n".format(len(unique_values)))
print(data['14_index'])
for column in data.select_dtypes(include=['object']).columns:
data.drop([ column ], axis = 1, inplace = True)
display(data)
```
It is another way of representing a categorical feature in *encoded* form. It is not friendly for *Linear Learner* and classical logistic regression, but we use `xgboost` library - which can leverage such a column without any problems.
## Finishing Touches
Last, but not least - we need to unify the values that are pointing out a missing value `NaN` and `-1`. We use `NaN` everywhere:
```
# Replace all -1 to NaN:
for column in data.columns:
data[column] = data[column].replace(-1, np.nan)
testing = data[2]
testing_unique_values = data[2].unique()
print("Number of unique values in 2nd feature: {}\n".format(len(testing_unique_values)))
print(testing)
```
## Splitting the dataset
We need to split the dataset. We decided to randomize the dataset, and split into 70% for training, 20% for validation and 10% for the test.
```
# Randomly sort the data then split out first 70%, second 20%, and last 10%:
data_len = len(data)
sampled_data = data.sample(frac = 1)
train_data, validation_data, test_data = np.split(sampled_data, [ int(0.7 * data_len), int(0.9 * data_len) ])
```
After splitting, we need to save new training and validation dataset as *CSV* files. After saving, we upload them to the `output_bucket`.
```
train_data.to_csv('train.sample.csv', index = False, header = False)
validation_data.to_csv('validation.sample.csv', index = False, header = False)
s3client = boto3.Session().resource('s3')
train_csv_file = os.path.join(path, 'train/train.csv')
validation_csv_file = os.path.join(path, 'validation/validation.csv')
s3client.Bucket(output_bucket).Object(train_csv_file).upload_file('train.sample.csv')
s3client.Bucket(output_bucket).Object(validation_csv_file).upload_file('validation.sample.csv')
```
Now we are ready to leverage *Amazon SageMaker* for training.
# Training
## Preparation
As a first step, we need to point which libraries we want to use. We do that by fetching the container name based on the name of the library we want to use. In our case, it is `xgboost`.
```
from sagemaker.amazon.amazon_estimator import get_image_uri
container = get_image_uri(boto3.Session().region_name, 'xgboost')
```
Then, we need to point out where to look for input data. In our case, we use *CSV* files uploaded in the previous section to `output_bucket`.
```
train_csv_key = 's3://{}/{}/train/train.csv'.format(output_bucket, path)
validation_csv_key = 's3://{}/{}/validation/validation.csv'.format(output_bucket, path)
s3_input_train = sagemaker.s3_input(s3_data = train_csv_key, content_type = 'csv')
s3_input_validation = sagemaker.s3_input(s3_data = validation_csv_key, content_type = 'csv')
```
## Differences from usual workflow and frameworks usage
Even that *Amazon SageMaker* supports *CSV* files, most of the algorithms work best when you use the optimized `protobuf` `recordIO` format for the training data.
Using this format allows you to take advantage of *pipe mode* when training the algorithms that support it. File mode loads all of your data from *Amazon S3* to the training instance volumes. In *pipe mode*, your training job streams data directly from *Amazon S3*. Streaming can provide faster start times for training jobs and better throughput.
With this mode, you also reduce the size of the *Amazon EBS* volumes for your training instances. *Pipe mode* needs only enough disk space to store your final model artifacts. File mode needs disk space to store both your final model artifacts and your full training dataset.
For our use case - we leverage *CSV* files.
## Single training job
```
sess = sagemaker.Session()
xgb = sagemaker.estimator.Estimator(container,
role,
train_instance_count = 1,
train_instance_type = 'ml.m4.xlarge',
base_job_name = user_name,
output_path = 's3://{}/{}/output'.format(output_bucket, path),
sagemaker_session = sess)
xgb.set_hyperparameters(eval_metric = 'logloss',
objective = 'binary:logistic',
eta = 0.2,
max_depth = 10,
colsample_bytree = 0.7,
colsample_bylevel = 0.8,
min_child_weight = 4,
rate_drop = 0.3,
num_round = 75,
gamma = 0.8)
xgb.fit({'train': s3_input_train, 'validation': s3_input_validation})
```
Now, we are ready to create *Amazon SageMaker session* and `xgboost` framework objects.
For a single training job, we need to create *Estimator*, where we point the container and *security context*. In this step, we are specifying the instance type and amount of those used for learning. Last, but not least - we need to specify `output_path` and pass the session object.
For the created *Estimator* instance we need to specify the `objective`, `eval_metric` and other hyperparameters used for that training session.
As the last step, we need to start the training process passing the training and validation datasets. Whole training job takes approximately 1-2 minutes at most for the following setup.
## FAQ
**Q**: I see a strange error: `ClientError: Hidden file found in the data path! Remove that before training`. What is that?
**A**: There is something wrong with your input files, probably you messed up the *S3* path passed into training job.
## Hyperparameter Tuning (HPO)
The single job is just one way. We can automate the whole process with use of *hyperparameter tuning*.
As in the case of a single training job, we need to create *Estimator* with the specification for an individual job and set up initial and fixed values for *hyperparameters*. However, outside those - we are setting up the ranges in which algorithm automatically tune in, inside the process of the *HPO*.
Inside the *HyperparameterTuner* specification we are specifying how many jobs we want to run and how many of them we want to run in parallel.
```
hpo_sess = sagemaker.Session()
hpo_xgb = sagemaker.estimator.Estimator(container,
role,
train_instance_count = 1,
train_instance_type = 'ml.m4.xlarge',
output_path = 's3://{}/{}/output_hpo'.format(output_bucket, path),
sagemaker_session = hpo_sess)
hpo_xgb.set_hyperparameters(eval_metric = 'logloss',
objective = 'binary:logistic',
colsample_bytree = 0.7,
colsample_bylevel = 0.8,
num_round = 75,
rate_drop = 0.3,
gamma = 0.8)
hyperparameter_ranges = {
'eta': ContinuousParameter(0, 1),
'min_child_weight': ContinuousParameter(1, 10),
'alpha': ContinuousParameter(0, 2),
'max_depth': IntegerParameter(1, 10),
}
objective_metric_name = 'validation:logloss'
objective_type = 'Minimize'
tuner = HyperparameterTuner(hpo_xgb,
objective_metric_name,
hyperparameter_ranges,
base_tuning_job_name = user_name,
max_jobs = 20,
max_parallel_jobs = 5,
objective_type = objective_type)
tuner.fit({'train': s3_input_train, 'validation': s3_input_validation})
```
Another thing that is different is how we see the progress of that particular type of the job. In the previous case, logs were shipped automatically into a *notebook*. For *HPO*, we need to fetch job status via *Amazon SageMaker SDK*. Unfortunately, it allows fetching the only status - logs are available in *Amazon CloudWatch*.
**Beware**, that with current setup whole *HPO* job may take 20-30 minutes.
```
smclient = boto3.client('sagemaker')
job_name = tuner.latest_tuning_job.job_name
hpo_job = smclient.describe_hyper_parameter_tuning_job(HyperParameterTuningJobName = job_name)
hpo_job['HyperParameterTuningJobStatus']
```
# Hosting the single model
After finishing the training, *Amazon SageMaker* by default saves the model inside *S3* bucket we have specified. Moreover, based on that model we can either download the archive and use inside our source code and services when deploying, or we can leverage the hosting mechanism available in the *Amazon SageMaker* service.
## How it works?
After you deploy a model into production using *Amazon SageMaker* hosting services, it creates the endpoint with its configuration.
Your client applications use `InvokeEndpoint` API to get inferences from the model hosted at the specified endpoint. *Amazon SageMaker* strips all `POST` headers except those supported by the *API*. Service may add additional headers.
Does it mean that everyone can call our model? No, calls to `InvokeEndpoint` are authenticated by using *AWS Signature Version 4*.
A customer's model containers must respond to requests within 60 seconds. The model itself can have a maximum processing time of 60 seconds before responding to the /invocations. If your model is going to take 50-60 seconds of processing time, the SDK socket timeout should be set to be 70 seconds.
```
xgb_predictor = xgb.deploy(initial_instance_count = 1, instance_type = 'ml.m4.xlarge')
```
**Beware**, the '!' in the output after hosting model means that it deployed with success.
# Hosting the best model from HPO
Hosting *HPO* model is no different from a single job. *Amazon SageMaker SDK* in very convenient way selects the best model automatically and uses that as a back-end for the endpoint.
```
xgb_predictor_hpo = tuner.deploy(initial_instance_count = 1, instance_type = 'ml.m4.xlarge')
```
# Evaluation
After training and hosting the best possible model, we would like to evaluate its performance with `test_data` subset prepared when splitting data.
As a first step, we need to prepare our hosted predictors to expect `text/csv` payload, which deserializes via *Amazon SageMaker SDK* entity `csv_serializer`.
```
xgb_predictor.content_type = 'text/csv'
xgb_predictor.serializer = csv_serializer
xgb_predictor_hpo.content_type = 'text/csv'
xgb_predictor_hpo.serializer = csv_serializer
```
As a next step, we need to prepare a helper function that split `test_data` into smaller chunks and serialize them before passing it to predictors.
```
def predict(predictor, data, rows = 500):
split_array = np.array_split(data, int(data.shape[0] / float(rows) + 1))
predictions = ''
for array in split_array:
predictions = ','.join([predictions, predictor.predict(array).decode('utf-8')])
return np.fromstring(predictions[1:], sep =',')
predictions = predict(xgb_predictor, test_data.drop([0], axis=1).values)
hpo_predictions = predict(xgb_predictor_hpo, test_data.drop([0], axis=1).values)
```
As a final step, we would like to compare how many clicks available in `test_data` subset were predicted correctly for job trained individually and with *HPO* jobs.
```
rows = ['actuals']
cols = ['predictions']
clicks = np.round(predictions)
result = pd.crosstab(index = test_data[0], columns = clicks, rownames = rows, colnames = cols)
display("Single job results:")
display(result)
display(result.apply(lambda r: r/r.sum(), axis = 1))
hpo_clicks = np.round(hpo_predictions)
result_hpo = pd.crosstab(index = test_data[0], columns = hpo_clicks, rownames = rows, colnames = cols)
display("HPO job results:")
display(result_hpo)
display(result_hpo.apply(lambda r: r/r.sum(), axis = 1))
```
As you may expect, the model trained with the use of *HPO* works better.
What is interesting - without any tuning and significant improvements, we were able to be classified in the first 25-30 results of the leaderboard from the old [Kaggle competition](https://www.kaggle.com/c/criteo-display-ad-challenge/leaderboard). Impressive!
# Clean-up
To avoid incurring unnecessary charges, use the *AWS Management Console* to delete the resources that you created for this exercise.
Open the *Amazon SageMaker* console at and delete the following resources:
1. The endpoint - that also deletes the ML compute instance or instances.
2. The endpoint configuration.
3. The model.
4. The notebook instance. You need to stop the instance before deleting it.
Keep in mind that *you can not* delete the history of trained individual and hyperparameter optimization jobs, but that do not incur any charges.
Open the Amazon S3 console at and delete the bucket that you created for storing model artifacts and the training dataset. Remember, that before deleting you need to empty it, by removing all objects.
Open the *IAM* console at and delete the *IAM* role. If you created permission policies, you could delete them, too.
Open the *Amazon CloudWatch* console at and delete all of the log groups that have names starting with `/aws/sagemaker`.
When it comes to *endpoints* you can leverage the *Amazon SageMaker SDK* for that operation:
```
sagemaker.Session().delete_endpoint(xgb_predictor.endpoint)
sagemaker.Session().delete_endpoint(xgb_predictor_hpo.endpoint)
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Forecasting with an RNN
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l08c06_forecasting_with_rnn.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l08c06_forecasting_with_rnn.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
## Setup
```
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
keras = tf.keras
def plot_series(time, series, format="-", start=0, end=None, label=None):
plt.plot(time[start:end], series[start:end], format, label=label)
plt.xlabel("Time")
plt.ylabel("Value")
if label:
plt.legend(fontsize=14)
plt.grid(True)
def trend(time, slope=0):
return slope * time
def seasonal_pattern(season_time):
"""Just an arbitrary pattern, you can change it if you wish"""
return np.where(season_time < 0.4,
np.cos(season_time * 2 * np.pi),
1 / np.exp(3 * season_time))
def seasonality(time, period, amplitude=1, phase=0):
"""Repeats the same pattern at each period"""
season_time = ((time + phase) % period) / period
return amplitude * seasonal_pattern(season_time)
def white_noise(time, noise_level=1, seed=None):
rnd = np.random.RandomState(seed)
return rnd.randn(len(time)) * noise_level
def window_dataset(series, window_size, batch_size=32,
shuffle_buffer=1000):
dataset = tf.data.Dataset.from_tensor_slices(series)
dataset = dataset.window(window_size + 1, shift=1, drop_remainder=True)
dataset = dataset.flat_map(lambda window: window.batch(window_size + 1))
dataset = dataset.shuffle(shuffle_buffer)
dataset = dataset.map(lambda window: (window[:-1], window[-1]))
dataset = dataset.batch(batch_size).prefetch(1)
return dataset
def model_forecast(model, series, window_size):
ds = tf.data.Dataset.from_tensor_slices(series)
ds = ds.window(window_size, shift=1, drop_remainder=True)
ds = ds.flat_map(lambda w: w.batch(window_size))
ds = ds.batch(32).prefetch(1)
forecast = model.predict(ds)
return forecast
time = np.arange(4 * 365 + 1)
slope = 0.05
baseline = 10
amplitude = 40
series = baseline + trend(time, slope) + seasonality(time, period=365, amplitude=amplitude)
noise_level = 5
noise = white_noise(time, noise_level, seed=42)
series += noise
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
split_time = 1000
time_train = time[:split_time]
x_train = series[:split_time]
time_valid = time[split_time:]
x_valid = series[split_time:]
```
## Simple RNN Forecasting
```
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
window_size = 30
train_set = window_dataset(x_train, window_size, batch_size=128)
model = keras.models.Sequential([
keras.layers.Lambda(lambda x: tf.expand_dims(x, axis=-1),
input_shape=[None]),
keras.layers.SimpleRNN(100, return_sequences=True),
keras.layers.SimpleRNN(100),
keras.layers.Dense(1),
keras.layers.Lambda(lambda x: x * 200.0)
])
lr_schedule = keras.callbacks.LearningRateScheduler(
lambda epoch: 1e-7 * 10**(epoch / 20))
optimizer = keras.optimizers.SGD(lr=1e-7, momentum=0.9)
model.compile(loss=keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
history = model.fit(train_set, epochs=100, callbacks=[lr_schedule])
plt.semilogx(history.history["lr"], history.history["loss"])
plt.axis([1e-7, 1e-4, 0, 30])
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
window_size = 30
train_set = window_dataset(x_train, window_size, batch_size=128)
valid_set = window_dataset(x_valid, window_size, batch_size=128)
model = keras.models.Sequential([
keras.layers.Lambda(lambda x: tf.expand_dims(x, axis=-1),
input_shape=[None]),
keras.layers.SimpleRNN(100, return_sequences=True),
keras.layers.SimpleRNN(100),
keras.layers.Dense(1),
keras.layers.Lambda(lambda x: x * 200.0)
])
optimizer = keras.optimizers.SGD(lr=1.5e-6, momentum=0.9)
model.compile(loss=keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
early_stopping = keras.callbacks.EarlyStopping(patience=50)
model_checkpoint = keras.callbacks.ModelCheckpoint(
"my_checkpoint", save_best_only=True)
model.fit(train_set, epochs=500,
validation_data=valid_set,
callbacks=[early_stopping, model_checkpoint])
model = keras.models.load_model("my_checkpoint")
rnn_forecast = model_forecast(
model,
series[split_time - window_size:-1],
window_size)[:, 0]
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, rnn_forecast)
keras.metrics.mean_absolute_error(x_valid, rnn_forecast).numpy()
```
## Sequence-to-Sequence Forecasting
```
def seq2seq_window_dataset(series, window_size, batch_size=32,
shuffle_buffer=1000):
series = tf.expand_dims(series, axis=-1)
ds = tf.data.Dataset.from_tensor_slices(series)
ds = ds.window(window_size + 1, shift=1, drop_remainder=True)
ds = ds.flat_map(lambda w: w.batch(window_size + 1))
ds = ds.shuffle(shuffle_buffer)
ds = ds.map(lambda w: (w[:-1], w[1:]))
return ds.batch(batch_size).prefetch(1)
for X_batch, Y_batch in seq2seq_window_dataset(tf.range(10), 3,
batch_size=1):
print("X:", X_batch.numpy())
print("Y:", Y_batch.numpy())
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
window_size = 30
train_set = seq2seq_window_dataset(x_train, window_size,
batch_size=128)
model = keras.models.Sequential([
keras.layers.SimpleRNN(100, return_sequences=True,
input_shape=[None, 1]),
keras.layers.SimpleRNN(100, return_sequences=True),
keras.layers.Dense(1),
keras.layers.Lambda(lambda x: x * 200)
])
lr_schedule = keras.callbacks.LearningRateScheduler(
lambda epoch: 1e-7 * 10**(epoch / 30))
optimizer = keras.optimizers.SGD(lr=1e-7, momentum=0.9)
model.compile(loss=keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
history = model.fit(train_set, epochs=100, callbacks=[lr_schedule])
plt.semilogx(history.history["lr"], history.history["loss"])
plt.axis([1e-7, 1e-4, 0, 30])
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
window_size = 30
train_set = seq2seq_window_dataset(x_train, window_size,
batch_size=128)
valid_set = seq2seq_window_dataset(x_valid, window_size,
batch_size=128)
model = keras.models.Sequential([
keras.layers.SimpleRNN(100, return_sequences=True,
input_shape=[None, 1]),
keras.layers.SimpleRNN(100, return_sequences=True),
keras.layers.Dense(1),
keras.layers.Lambda(lambda x: x * 200.0)
])
optimizer = keras.optimizers.SGD(lr=1e-6, momentum=0.9)
model.compile(loss=keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
early_stopping = keras.callbacks.EarlyStopping(patience=10)
model.fit(train_set, epochs=500,
validation_data=valid_set,
callbacks=[early_stopping])
rnn_forecast = model_forecast(model, series[..., np.newaxis], window_size)
rnn_forecast = rnn_forecast[split_time - window_size:-1, -1, 0]
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, rnn_forecast)
keras.metrics.mean_absolute_error(x_valid, rnn_forecast).numpy()
```
| github_jupyter |
```
%matplotlib inline
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import numpy as np
from scipy import integrate
import scipy.linalg as la
import scipy.sparse.linalg as sla
k = 9
m = 1
T = 2 * np.pi * ((k/m)**0.5)
omega = 2 * np.pi / T
N = 1e4
R0 = [0.5, 1.]
T = 2 * np.pi * ((k/m)**0.5)
t0, t1 = 0, 5 * T
t = np.linspace(t0, t1, 10000)
R = np.zeros((len(t), len(R0)), dtype=np.float64)
En = np.array([1/2 * k * R[:, 1][i]**2 + 1/2 * m * R[:, 0][i]**2 for i in range(len(R[:, 0]))])
E0 = 1/2 * k * R0[1]**2 + 1/2 * m * R0[0]**2
plt.plot(np.linspace(0, len(En), len(En)), (En - E0) / E0)
def osc1(t, z):
omega = 3
gamma = 0.5
dy = np.zeros(2)
dy[0] = z[1]
dy[1] = (-1 * (omega)**2) * z[0] - gamma * z[1]
return dy
def osc2(t, z):
omega = 3
gamma = 0.5
A0 = 1
omegaF = 2
F = A0 * np.cos(omegaF * t)
dy = np.zeros(2)
dy[0] = z[1]
dy[1] = (-1 * (omega)**2) * z[0] - gamma * z[1] + F
return dy
def f1(R0, oscillator):
N = 1e4
t0, t1 = 0, 5 * T
t = np.linspace(t0, t1, 10000)
R = np.zeros((len(t), len(R0)), dtype=np.float64)
R[0, :] = R0
r = integrate.ode(oscillator).set_integrator("dopri5")
r.set_initial_value(R0, t0)
for i in range(1, t.size):
R[i, :] = r.integrate(t[i])
if not r.successful():
raise RuntimeError("Could not integrate")
fig, axs = plt.subplots(3, 1, figsize=(8,8))
axs[0].plot(t, R[:,0])
axs[1].plot(t, R[:,1])
axs[2].plot(R[:, 0], R[:,1])
R0 = [1, 0]
f1(R0, osc1)
R0 = [1, 0]
f1(R0, oscillator2)
N = 2
m = [1, 1]
k = [1, 0.8, 1]
R0 = np.array([-0.5, 0])
v0 = np.array([0, 0])
omega = np.zeros((N+1, N), dtype=np.float64)
for alpha in range(N+1):
for beta in range(N):
omega[alpha, beta] = k[alpha] / m[beta]
Omega = np.zeros((N, N), dtype=np.float64)
for i in range(N):
if i == 0:
Omega[i, i] = omega[0, 0] + omega[1, 0]
Omega[0, 1] = -omega[1, 0]
if i > 0:
if i < N-1:
Omega[i, i-1] = -omega[i, i]
Omega[i,i] = omega[i, i] + omega[i + 1, i]
Omega[i, i+1] = -omega[i+1, i]
else:
Omega[i, i-1] = -omega[i, i]
Omega[i, i] = omega[i, i] + omega[i + 1, i]
Theta, Sigma = np.linalg.eig(Omega)
Theta = np.sqrt(Theta)
SigmaV = np.zeros((N, N), dtype=np.float64)
for i in range(N):
for j in range(N):
SigmaV[j, i] = -Theta[i] * Sigma[j, i]
C1 = np.dot(np.linalg.inv(Sigma),R0[None].T.conj())
C2 = np.dot(np.linalg.inv(SigmaV),v0[None].T.conj())
C = np.sqrt(C1**2 + C2**2)
alpha = np.zeros(N, dtype=np.float64)
for i in range(N):
if C[i] == 0:
alpha[i] = 0
else:
alpha[i] = np.arctan(C2[i]/C1[i])
if C1[i] < 0:
alpha[i] = np.pi + alpha[i]
if C1[i] > 0:
if C2[i] < 0:
alpha[i] = 2 * np.pi + alpha[i]
N = len(Omega)
N1 = int(1000)
Tmax = 80
t = np.zeros(N1, dtype=np.float64)
X = np.zeros((N, N1), dtype=np.float64)
Xv = np.zeros((N, N1), dtype=np.float64)
for j in range(N1):
t[j] = (j-1)/(N1-1)*Tmax
for j in range(N1):
s = np.zeros(N, dtype=np.float64)
for i in range(N):
s = s + C[i] * Sigma[:,i] * np.cos(Theta[i]*t[j] + alpha[i])
X[:, j] = s
for j in range(N1):
s = np.zeros(N, dtype=np.float64)
for i in range(N):
s = s + C[i] * Sigma[:,i] * Theta[i] * np.sin(Theta[i]*t[j] + alpha[i])
Xv[:, j] = -s
c1 = np.fft.fft(X[0])
c2 = np.fft.fft(X[1])
Cm1 = np.zeros(N1 // 2, dtype=np.float64)
Cm2 = np.zeros(N1 // 2, dtype=np.float64)
Freq = np.zeros(N1 // 2, dtype=np.float64)
for j in range(1, N1//2):
Cm1[j-1] = abs(c1[j-1]) / (N1/2)
Cm2[j-1] = abs(c2[j-1]) / (N1/2)
Freq[j-1] = (j-1) / Tmax
fig, axs = plt.subplots(5, 1, figsize=(10,10))
axs[0].plot(t, X[0],'blue', t, X[1],'green')
axs[1].plot(t, Xv[0],'blue', t, Xv[1],'green')
axs[2].plot(X[0], Xv[0])
axs[3].plot(X[1], Xv[1])
axs[4].set_xscale('log')
axs[4].set_yscale('log')
axs[4].plot(Freq, Cm1,'blue', Freq, Cm2,'green')
N = 2
m = [1, 1]
k = [1, 1, 1]
R0 = np.array([-0.5, 0])
v0 = np.array([0, 0])
omega = np.zeros((N+1, N), dtype=np.float64)
for alpha in range(N+1):
for beta in range(N):
omega[alpha, beta] = k[alpha] / m[beta]
Omega = np.zeros((N, N), dtype=np.float64)
for i in range(N):
if i == 0:
Omega[i, i] = omega[0, 0] + omega[1, 0]
Omega[0, 1] = -omega[1, 0]
if i > 0:
if i < N-1:
Omega[i, i-1] = -omega[i, i]
Omega[i,i] = omega[i, i] + omega[i+1, i]
Omega[i, i+1] = -omega[i+1, i]
else:
Omega[i, i-1] = -omega[i, i]
Omega[i, i] = omega[i, i] + omega[i+1, i]
Theta, Sigma = np.linalg.eig(Omega)
Theta = np.sqrt(Theta)
SigmaV = np.zeros((N, N), dtype=np.float64)
for i in range(N):
for j in range(N):
SigmaV[j, i] = -Theta[i] * Sigma[j, i]
C1 = np.dot(np.linalg.inv(Sigma),R0[None].T.conj())
C2 = np.dot(np.linalg.inv(SigmaV),v0[None].T.conj())
C = np.sqrt(C1**2 + C2**2)
alpha = np.zeros(N, dtype=np.float64)
for i in range(N):
if C[i] == 0:
alpha[i] = 0
else:
alpha[i] = np.arctan(C2[i] / C1[i])
if C1[i] < 0:
alpha[i] = np.pi + alpha[i]
if C1[i] > 0:
if C2[i] < 0:
alpha[i] = 2 * np.pi + alpha[i]
N = len(Omega)
N1 = int(500)
Tmax = 80
t = np.zeros(N1, dtype=np.float64)
X = np.zeros((N, N1), dtype=np.float64)
Xv = np.zeros((N, N1), dtype=np.float64)
for j in range(N1):
t[j] = (j-1) / (N1-1) * Tmax
for j in range(N1):
s = np.zeros(N, dtype=np.float64)
for i in range(N):
s = s + C[i] * Sigma[:,i] * np.cos(Theta[i]*t[j] + alpha[i])
X[:, j] = s
for j in range(N1):
s = np.zeros(N, dtype=np.float64)
for i in range(N):
s = s+ C[i] * Sigma[:,i] * Theta[i] * np.sin(Theta[i]*t[j] + alpha[i])
Xv[:, j] = -s
A1 = np.fft.fft(X[0])
A2 = np.fft.fft(X[1])
C1 = np.zeros(N1 // 2, dtype=np.float64)
C2 = np.zeros(N1 // 2, dtype=np.float64)
Freq = np.zeros(N1 // 2, dtype=np.float64)
for j in range(1, N1 // 2):
C1[j-1] = abs(A1[j-1]) / (N1/2)
C2[j-1] = abs(A2[j-1]) / (N1/2)
Freq[j-1] = (j-1) / Tmax
fig, axs = plt.subplots(5, 1, figsize=(10,10))
axs[0].plot(t, X[0],'blue', t, X[1],'green')
axs[1].plot(t, Xv[0],'blue', t, Xv[1],'green')
axs[2].plot(X[0], Xv[0])
axs[3].plot(X[1], Xv[1])
axs[4].set_xscale('log')
axs[4].set_yscale('log')
axs[4].plot(Freq, Cm1,'blue', Freq, Cm2,'green')
```
| github_jupyter |
# NLP Task - Classify real news from fake news
### Importing libraries and importing Datasets
```
import pandas as pd
import numpy as np
import os, time, gc, re
import nltk
nltk.download('punkt')
nltk.download('wordnet')
from nltk.corpus import stopwords
from nltk.util import ngrams
from nltk.tokenize import word_tokenize,sent_tokenize
from keras.preprocessing.text import Tokenizer
from gensim.models import KeyedVectors
import torch
from torch import nn, Tensor, device
from torch.optim import Optimizer
import torch.multiprocessing as mp
from torch.utils.data import DataLoader
from torch.utils.tensorboard import SummaryWriter
from keras.preprocessing.text import Tokenizer
import torch.nn.functional as F
from torch.nn.utils.rnn import pack_padded_sequence , pad_sequence
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics import roc_auc_score,confusion_matrix, classification_report
from datetime import datetime
import random
import json
import requests
import math
import pickle
import sys
import time
from tqdm.autonotebook import trange
from tqdm._tqdm_notebook import tqdm_notebook
from tqdm import tqdm
tqdm_notebook.pandas()
# Loading Sentence Transformer to Load pretrained RoBERTa for sentence Embeddings
from sentence_transformers import SentenceTransformer
model = SentenceTransformer('paraphrase-distilroberta-base-v1')
from IPython.display import clear_output
clear_output()
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
import warnings
warnings.filterwarnings("ignore")
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
from IPython.display import display
from IPython.core.display import HTML
import matplotlib.pyplot as plt
import seaborn as sns
plt.style.use('ggplot')
real = pd.read_csv('data/True.csv')
fake = pd.read_csv('data/Fake.csv')
fake['Fake'] = True
real['Fake'] = False
data = pd.concat([real , fake])
data['news'] = data['title'] + ' - ' + data['text']
data['length'] = data.news.apply(lambda x: len(x.split(' ')))
data = data.sample(frac = 1.0)
display(HTML('<h3>Sample Data From Fake Dataset</h3>'))
fake.sample(5)
display(HTML('<h3>Sample Data From Real Dataset</h3>'))
real.sample(5)
display(HTML(f'<b>Fake Dataset Shape</b> : {fake.shape[0]} Rows * {fake.shape[1]} Columns'))
display(HTML(f'<b>Real Dataset Shape</b> : {real.shape[0]} Rows * {real.shape[1]} Columns'))
display(HTML(f"Count plot of Length of Text in Dataset"))
fig, axs = plt.subplots(figsize=(15, 8))
sns.histplot(data, x = 'length', hue = 'Fake',ax = axs, kde = True, log_scale=True)
# sns.histplot(fake.text.apply(lambda x : len(x.split(' '))) , color = 'red', ax = axs)
# sns.histplot(real.text.apply(lambda x : len(x.split(' '))) , color = 'blue' , ax = axs)
```
# Text Preprocessing
```
lemma = nltk.WordNetLemmatizer()
def normalize(text):
text = re.sub(r'U.S.|US|U.S.A','US',text)
text = re.sub(r'(@[A-Za-z0-9]+)|(#[A-Za-z0-9]+)|([^\x00-\x7F]+)|([0-9])|(\w+:\/\/\S+)|([^\w\s])|(\s+)', ' ', text)
text = text.strip().lower()
text = re.sub(r'\s+', ' ', text)
text = nltk.word_tokenize(text) # Tokenizing
text = [lemma.lemmatize(word) for word in text] # Lemmatizing
text = " ".join(text)
return text
data['news'] = data['news'].apply(normalize)
data.sample(10)
```
# Possible ways to solve this Problem
* RNN Based Models : Training Any DNN Model from Scratch and Training that over Complete Dataset.
* Machine Learning Approch : Tokenizing the Dataset and using Word2Vec followed by Any Machine Learning Model like Random Forest or KNN to classify sentences.
* Using Any pretrained Sentence Embedding Like BERT or GPT-2 followed by ANN to find Fake News in Dataset
## Advantages of using Pretrained Sentence Embedding
* Faster Compution due to Simpler Artitecture
* Can Take Advantage of Pretrained BERT Network
* Sentence sizes are Long and required 16+ Gb of RAM if not truncated while implementing RNN Based Network
* Vocab size is 1,00,000+ Making Embedding Matrix Very large and Computationaly Expensive
# Neural Network Model
```
class Model(nn.Module):
def __init__(self, ntoken = 768, nout = 1):
super(Model, self).__init__()
self.model_type = 'ANN Model'
self.encoder = nn.Linear(ntoken , ntoken*2)
self.relu = nn.ReLU()
self.norm = nn.LayerNorm(ntoken*2)
self.decoder = nn.Linear(ntoken*2, nout)
self.sigmoid = nn.Sigmoid()
self.dropout = nn.Dropout(.25)
def forward(self, x):
x = self.encoder(x)
x = self.norm(self.relu(x))
x = self.dropout(x)
x = self.decoder(x)
return self.sigmoid(x.squeeze())
```
# Dataloader
```
class TrainerDataset:
def __init__(self, dataset ,target):
self.dataset = dataset
self.target = target
def __len__(self):
return len(self.dataset)
def __getitem__(self, item):
target = self.target[item]
item = self.dataset[item]
# print(item)
return torch.tensor(item) , torch.tensor(target, dtype=torch.float32)
def create_dataloader():
train_data = TrainerDataset(embeddings[:40000] , data.Fake.values[:40000])
train_data = DataLoader(train_data , batch_size = 64 , shuffle = True , drop_last = True)
valid_data = TrainerDataset(embeddings[40000:], data.Fake.values[40000:])
valid_data = DataLoader(valid_data , batch_size = 64 , shuffle = True , drop_last = True)
return train_data , valid_data
```
# Encoding Sentence Embeddings for News
```
embeddings = model.encode(data.news.values , normalize_embeddings= True ,batch_size= 128, device= 'cuda', show_progress_bar= True )
class Pytorch_Trainer():
"""
Custom Pytorch Trainer Model Which makes easy to Train Pytorch Model
BY: KAUSTUBH PATHAK
"""
def __init__(self):
super().__init__()
self._BT_Avg_Loss_Train = []
self._BT_Avg_Loss_Valid = []
self._Loss_df = []
self.max_grad_norm = 1
self.__Use_CLR = False
self.model_save_path = '/content/drive/MyDrive/LambdaTest/models_2'
os.makedirs(self.model_save_path, exist_ok=True)
self.device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# To Save Model after Training (Trainer dont have Automatic Saving to Save Best Model while Training { To Implement})
def _Save_Model(self , model , optimizer , path):
checkpoint = {
'Training Loss': self._BT_Avg_Loss_Train,
'Validation Loss': self._BT_Avg_Loss_Valid,
'Loss df': self._Loss_df,
'Optimizer': optimizer.state_dict(),
'State_Dict': model.state_dict()
}
torch.save(checkpoint , path)
print('Model Saved Successfully....')
# Load the Saved Model and Optimizer
def _Load_Model(self, model, optimizer ,path):
checkpoint = torch.load(path)
model.load_state_dict(checkpoint['State_Dict'])
self._BT_Avg_Loss_Train = checkpoint['Training Loss']
self._BT_Avg_Loss_Valid = checkpoint['Validation Loss']
self._Loss_df = checkpoint['Loss df']
optimizer.load_state_dict(checkpoint['Optimizer'])
print('Loading Completed....')
return model, optimizer
# Switch Between Training and Evaluation modes.
def _Train_Eval(self , model , mode):
if mode.lower() == 'train':
model.train()
torch.set_grad_enabled(True)
elif mode.lower() == 'eval':
model.eval()
torch.set_grad_enabled(False)
else:
raise Exception("Unknown Type....!! Set to 'Train' or 'Eval'")
# Predict Function (Correctly {Not for Large Datasets.})
def _Predict(self , model, x):
model._Train_Eval(mode = 'Eval')
x = torch.tensor(x).to(device)
return model(x).detach()
# Run/Train the Model with Automatic Mixed Precision
def fit(self , model , optimizer , loss , dataloader , validloader ,scheduler = None,ex_flag = 0, epochs = 'Default' , use_grad = False):
print('Abbreviations : \n\tV.S. -> Validation Size \tV.L. -> Validation Loss \tT.S. -> Training Size \tT.L. -> Training Loss \tT.T. -> Training Time\nResults : ')
loss = loss.to(self.device)
model = model.to(self.device)
if epochs == 'Default':
epochs = 5
if use_grad:
scaler = GradScaler()
for j in range(epochs):
time_now = time.time()
self._Train_Eval(model, mode = 'Train')
Train_Total_Loss = 0 #For Epoch Loss
for x_train , target in dataloader:
x_train , target = x_train.to(device) , target.to(device)
optimizer.zero_grad()
if use_grad:
with autocast():
embed_x = model.__call__(x_train )
loss_value = loss(embed_x, target)
scale_before_step = scaler.get_scale()
scaler.scale(loss_value).backward()
scaler.unscale_(self.optimizer)
torch.nn.utils.clip_grad_norm_(model.parameters(), self.max_grad_norm)
scaler.step(self.optimizer)
scaler.update()
skip_scheduler = scaler.get_scale() != scale_before_step
else:
embed_x = model.__call__(x_train )
loss_value = loss(embed_x, target)
loss_value.backward()
torch.nn.utils.clip_grad_norm_(model.parameters(), self.max_grad_norm)
optimizer.step()
if scheduler != None:
scheduler.step() # Stepping If using Cyclic Learning Rate ()
self._BT_Avg_Loss_Train.append([j,ex_flag,loss_value.item()]) # Otherwise Simple Append
Train_Total_Loss += loss_value.item() #For Epoch Loss
ex_flag += 1 #Flag
#### For Evaluation ....
self._Train_Eval(model , mode = 'Eval') # Switching to Evaluation Mode
ex_flag = 0 #Flag
Valid_Total_Loss = 0 # For Calculating Validation Loss
for x_train , target in validloader:
x_train , target = x_train.to(device) , target.to(device)
if use_grad:
with autocast():
embed_x = model.__call__(x_train )
loss_value = loss(embed_x, target)
else:
embed_x = model.__call__(x_train ) # Calling to get Prediction
loss_value = loss(embed_x, target) # Calculating the Loss
self._BT_Avg_Loss_Valid.append([j,ex_flag , loss_value.item()]) # Otherwise Simple Append
Valid_Total_Loss += loss_value.item() #Epoch Validation Loss
ex_flag += 1 #Flag
# self._Save_Model(model , optimizer , f'{self.model_save_path}/epochs{j+1}_ex_flag_{ex_flag}.pt')
print('Epoch : {} | V.S. : {} | V.L. : {:07.5f} | T.S. : {} | T.L : {:07.5f} | T.T. : {:05.3f} Sec ||'.\
format(\
j+1 , \
len(validloader),round(Valid_Total_Loss / len(validloader) , 5) ,
len(dataloader) , round(Train_Total_Loss / len(dataloader) , 5) ,
round(time.time() - time_now,3)
)\
)
self._Loss_df.append(
{
'Epoch':j ,
'Training Loss': Train_Total_Loss/len(dataloader) ,
'Validation Loss': Valid_Total_Loss / len(validloader)
}
)
```
# Model Training
```
ann_model = Model()
trainer = Pytorch_Trainer()
loss = nn.BCELoss()
train_loader , valid_loader = create_dataloader()
optimizer = torch.optim.Adam(ann_model.parameters(), lr=1.0*0.001, betas=(0.9, 0.999), eps=1e-08, weight_decay=0)
trainer.fit(ann_model , optimizer, loss, train_loader , valid_loader)
```
# Testing and Evaluation
```
pred = []
target = []
ann_model.eval()
for embedss , targets in train_loader:
preds = ann_model(embedss.to(device))
pred.append(preds.detach().cpu().numpy())
target.append(targets.cpu().numpy())
pred , target = np.hstack(pred) , np.hstack(target)
result = pred > .5
result = [int(i) for i in result]
accuracy_value = roc_auc_score(target,pred)
display(HTML(f"<b>Accuracy Over Training Dataset</b> - {round(accuracy_value*100 , 3)}% <br><br><b>Classification Report</b>"))
print(classification_report(target, result, target_names=['Real News', 'Fake News']))
display(HTML(f"<br><b>Confusion Matrix</b>"))
cm = confusion_matrix(target, result)
plt.figure(figsize = (10, 10))
sns.heatmap(pd.DataFrame(cm , index = ['Real News' , 'Fake News'] , columns = ['Real News', 'Fake News']), annot = True,fmt='g')
pred = []
target = []
ann_model.eval()
for embedss , targets in valid_loader:
preds = ann_model(embedss.to(device))
pred.append(preds.detach().cpu().numpy())
target.append(targets.cpu().numpy())
pred , target = np.hstack(pred) , np.hstack(target)
result = pred > .5
result = [int(i) for i in result]
accuracy_value = roc_auc_score(target,pred)
display(HTML(f"<b>Accuracy Over Testing Dataset</b> - {round(accuracy_value*100 , 3)}% <br><br><b>Classification Report</b>"))
print(classification_report(target, result, target_names=['Real News', 'Fake News']))
display(HTML(f"<br><b>Confusion Matrix</b>"))
cm = confusion_matrix(target, result)
plt.figure(figsize = (10, 10))
sns.heatmap(pd.DataFrame(cm , index = ['Real News' , 'Fake News'] , columns = ['Real News', 'Fake News']), annot = True,fmt='g')
```
# Result
* Using Pretrained Encoding decrease model training time
* Pretrained Sentence Embeddings increase model Accuracy to 99.98 % over Training Set and 99.75 %
| github_jupyter |
```
!pip install pymongo
!pip install pymongo[srv]
!pip install dnspython
!pip install tweepy
!pip install twitter
import pymongo
from pymongo import MongoClient
import json
import tweepy
import twitter
from pprint import pprint
import configparser
import pandas as pd
config = configparser.ConfigParser()
config.read('config.ini')
CONSUMER_KEY = config['mytwitter']['api_key']
CONSUMER_SECRET = config['mytwitter']['api_secrete']
OAUTH_TOKEN = config['mytwitter']['access_token']
OATH_TOKEN_SECRET = config['mytwitter']['access_secrete']
mongod_connect = config['mymongo']['connection']
client = MongoClient(mongod_connect)
db = client.lab9 # use or create a database named demo
tweet_collection = db.tweet_collection #use or create a collection named tweet_collection
tweet_collection.create_index([("id", pymongo.ASCENDING)],unique = True) # make sure the collected tweets are unique
rest_auth = twitter.oauth.OAuth(OAUTH_TOKEN,OATH_TOKEN_SECRET,CONSUMER_KEY,CONSUMER_SECRET)
rest_api = twitter.Twitter(auth=rest_auth)
count = 100 #number of returned tweets, default and max is 100
geocode = "38.4392897,-78.9412224,50mi" # defin the location, in Harrisonburg, VA
q = "covid19" #define the keywords, tweets contain election
search_results = rest_api.search.tweets( count=count,q=q) #you can use both q and geocode
statuses = search_results["statuses"]
since_id_new = statuses[-1]['id']
for statuse in statuses:
try:
tweet_collection.insert_one(statuse)
pprint(statuse['created_at'])# print the date of the collected tweets
except:
pass
since_id_old = 0
while(since_id_new != since_id_old):
since_id_old = since_id_new
search_results = rest_api.search.tweets( count=count,q=q,
max_id= since_id_new)
statuses = search_results["statuses"]
since_id_new = statuses[-1]['id']
for statuse in statuses:
try:
tweet_collection.insert_one(statuse)
pprint(statuse['created_at']) # print the date of the collected tweets
except:
pass
print(tweet_collection.estimated_document_count())# number of tweets collected
user_cursor = tweet_collection.distinct("user.id")
print (len(user_cursor)) # number of unique Twitter users
tweet_collection.create_index([("text", pymongo.TEXT)], name='text_index', default_language='english') # create a text index
tweet_cursor = tweet_collection.find({"$text": {"$search": "covid"}}) # return tweets contain covid
for document in tweet_cursor[0:10]: # display the first 10 tweets from the query
try:
print ('----')
# pprint (document) # use pprint to print the entire tweet document
print ('name:', document["user"]["name"]) # user name
print ('text:', document["text"]) # tweets
except:
print ("***error in encoding")
pass
tweet_cursor = tweet_collection.find({"$text": {"$search": "vaccine"}}) # return tweets contain vaccine
tweet_df = pd.DataFrame(list(tweet_cursor ))
tweet_df[:10] #display the first 10 tweets
tweet_df["favorite_count"].hist() # create a histogram show the favorite count
```
| github_jupyter |
<a href="https://colab.research.google.com/github/wlsgud623/vaccine_tweet_analysis/blob/main/VaccineHesistant.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
!pip install transformers==3
import transformers
from transformers import BertTokenizer
import torch
import torch.nn.functional as F
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
!pip install kaggle
from google.colab import files
files.upload()
!mkdir -p ~/.kaggle
!cp kaggle.json ~/.kaggle/
!chmod 600 ~/.kaggle/kaggle.json
! kaggle datasets download -d kaushiksuresh147/covidvaccine-tweets
! kaggle datasets download -d gpreda/all-covid19-vaccines-tweets
!unzip all-covid19-vaccines-tweets.zip
!unzip covidvaccine-tweets.zip
first_vaccine_data = pd.read_csv("/content/covidvaccine.csv", nrows = 50000)
second_vaccine_data = pd.read_csv("/content/vaccination_all_tweets.csv", nrows = 50000)
data_column = ['user_name', 'user_description','user_location','user_followers','text']
first_vaccine_data = first_vaccine_data[data_column]
second_vaccine_data = second_vaccine_data[data_column]
import re
def con(x):
return ' '.join(re.sub("(#[A-Za-z0-9]+)|(@[A-Za-z0-9]+)|([^0-9A-Za-z \t])|(\w+:\/\/\S+)"," ",str(x)).split())
first_vaccine_data['text']=first_vaccine_data['text'].apply(con)
second_vaccine_data['text']=second_vaccine_data['text'].apply(con)
analysis_data = pd.concat([first_vaccine_data,second_vaccine_data], ignore_index=True)
analysis_data['text'].dropna()
analysis_data.drop_duplicates()
analysis_data.head()
from google.colab import drive
drive.mount('/gdrive', force_remount=True)
sentiment_data = pd.read_csv("/gdrive/My Drive/sentiment.csv", header=None)
del sentiment_data[0]
sentiment_data.rename(columns={1:'label'}, inplace=True)
sentiment_data.head()
print(analysis_data.shape)
print(sentiment_data.shape)
positive_tweet = pd.DataFrame(columns=['user_name', 'user_description','user_location','user_followers','text'])
neutral_tweet = pd.DataFrame(columns=['user_name', 'user_description','user_location','user_followers','text'])
negative_tweet = pd.DataFrame(columns=['user_name', 'user_description','user_location','user_followers','text'])
for index, values in analysis_data.iterrows():
if sentiment_data['label'][index] == 0:
negative_tweet = negative_tweet.append(values, ignore_index=True)
elif sentiment_data['label'][index] == 1:
neutral_tweet = neutral_tweet.append(values, ignore_index=True)
elif sentiment_data['label'][index] == 2:
positive_tweet = positive_tweet.append(values, ignore_index=True)
#positive_tweet = positive_tweet.reset_index(drop=True)
#positive_tweet.head()
#neutral_tweet = neutral_tweet.drop(index)
#neutral_tweet = neutral_tweet.reset_index(drop=True)
#neutral_tweet.head()
#negative_tweet = negative_tweet.drop(index)
#negative_tweet = negative_tweet.reset_index(drop=True)
negative_tweet.head()
tweet_number =[len(positive_tweet),len(neutral_tweet),len(negative_tweet)]
tweet_label = ['positive','neutral','negative']
plt.bar(tweet_label, tweet_number)
plt.title('Number of tweet sentiments')
plt.show()
```
Tweet analysis by follwers
```
%matplotlib inline
sns.displot(x=negative_tweet['user_followers'], color='red', rug=True)
plt.title('Number of tweet followers')
plt.show()
```
Tweet analysis by location
```
positive_loc = positive_tweet['user_location'].str.split(',',expand=True)
negative_loc = negative_tweet['user_location'].str.split(',',expand=True)
postive_loc=positive_loc.rename(columns={0:'fst_loc',1:'snd_loc'})
negative_loc=negative_loc.rename(columns={0:'fst_loc',1:'snd_loc'})
postive_loc['snd_loc'] = postive_loc['snd_loc'].str.strip()
negative_loc['snd_loc'] = negative_loc['snd_loc'].str.strip()
state_fix = {'Ontario': 'Canada','United Arab Emirates': 'UAE','TX': 'USA','NY': 'USA'
,'FL': 'USA','England': 'UK','Watford': 'UK','GA': 'USA','IL': 'USA'
,'Alberta': 'Canada','WA': 'USA','NC': 'USA','British Columbia': 'Canada','MA': 'USA','ON':'Canada'
,'OH':'USA','MO':'USA','AZ':'USA','NJ':'USA','CA':'USA','DC':'USA','AB':'USA','PA':'USA','SC':'USA'
,'VA':'USA','TN':'USA','New York':'USA','Dubai':'UAE','CO':'USA', 'Gujarat':'India', 'darkest Victoria':'UK'}
postive_loc = postive_loc.replace({"snd_loc": state_fix})
negative_loc = negative_loc.replace({"snd_loc": state_fix})
#sns.countplot(postive_loc["snd_loc"][:100])
positive_loc.value_counts()[:20]
sns.countplot(negative_loc["snd_loc"])
plt.show()
```
Tweet Topic
```
!pip install yake
from wordcloud import WordCloud, STOPWORDS, ImageColorGenerator
import yake
kw_extracter = yake.KeywordExtractor(lan='eng', n=2, dedupLim=0.9, top=3, features=None)
positive_keyword = ''
for p in positive_tweet['text']:
keywords = kw_extracter.extract_keywords(p)
for key in keywords:
positive_keyword = positive_keyword + str(key)
positive_wordcloud = WordCloud(collocations=False, background_color='white',mode='RGBA', max_words=20).generate(positive_keyword)
plt.figure()
plt.imshow(positive_wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
negative_keyword = ''
for n in negative_tweet['text']:
keywords = kw_extracter.extract_keywords(n)
for key in keywords:
negative_keyword = negative_keyword + str(key)
negative_wordcloud = WordCloud(collocations=False, background_color='white',mode='RGBA', max_words=20).generate(negative_keyword)
plt.figure()
plt.imshow(negative_wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
```
# Reason of Hesistant
**Reference paper** : Psychological characteristics and the mediating role of the 5C Model in explaining students’ COVID-19 vaccination intention, Wismans A, Thurik R, Baptista R, Dejardin M, Janssen F
5C Model : Confidence, Calculation, Complacency, Constraints, Collective Responsibility
Get associative words from https://www.visualthesaurus.com/
```
five_C_model = ['Confidence', 'Calculation', 'Complacency', 'Constraints','Collective Responsibility']
# Confidence : 백신 효능과 안전성, 이를 제공하는 보건 서비스, 그리고 정부의 배포 결정에 대한 신뢰 여부
# Calculation : 질병 자체를 건강에 심각한 위험으로 생각하는지 여부
# Complacency : 손익을 따지기 위해 광범위한 정보 검색에 참여하는지 여부
# Constraints : 백신을 접하기가 얼마나 쉬운지 여부
# Colllective Responsibility : 스스로 예방 접종을 통해 타인을 보호하려는 의지가 있는지
tokenizer = BertTokenizer.from_pretrained('bert-base-cased')
```
C-19 Vaccine Realated
```
# 백신의 효과 인지
perceived_effectiveness = ['effect', 'set up', 'outcome','useless', 'futile', 'vain']
pe = []
for key in perceived_effectiveness:
tokens = tokenizer.tokenize(key)
for token in tokens:
pe.append(token)
# 백신의 위험성 인지
perceived_risk_vaccine = ['aftereffect', 'fear', 'risk', 'dangerous']
pv = []
for key in perceived_risk_vaccine:
tokens = tokenizer.tokenize(key)
for token in tokens:
pv.append(token)
# 가족, 사회적 규범
normative_beliefs = ['family', 'friend', 'relative', 'social', 'religion', 'chruch']
nb = []
for key in normative_beliefs:
tokens = tokenizer.tokenize(key)
for token in tokens:
nb.append(token)
# 접종을 통해 얻을 수 있는 이익
perceived_benefits = ['benefit', 'cost', 'expensive', 'price']
pb = []
for key in perceived_benefits:
tokens = tokenizer.tokenize(key)
for token in tokens:
pb.append(token)
```
C-19 Related
```
# covid-19의 위험성 인지
perceived_risk_c19 = ['symptom', 'terror', 'worry', 'unease']
pr = []
for key in perceived_risk_vaccine:
tokens = tokenizer.tokenize(key)
for token in tokens:
pr.append(token)
# 감염된 경험
c19_infection = ['ill', 'suffer', 'taken']
ci = []
for key in c19_infection:
tokens = tokenizer.tokenize(key)
for token in tokens:
ci.append(token)
```
Personality
```
# 낙관론
optimism = ['optimism', 'positive', 'termperament', 'hope', 'disposition']
opt = []
for key in optimism:
tokens = tokenizer.tokenize(key)
for token in tokens:
opt.append(token)
# 충동 정도
impulsivity = ['impulse', 'impel', 'hotheaded', 'madcap', 'impetuos', 'driving', 'capricious', 'early', 'fast']
imp = []
for key in impulsivity:
tokens = tokenizer.tokenize(key)
for token in tokens:
imp.append(token)
# 이타주의
altruism = ['altruism', 'unselfishness', 'philanthropy', 'selflessness']
alt = []
for key in altruism:
tokens = tokenizer.tokenize(key)
for token in tokens:
alt.append(token)
# 소속감
need_to_belong = ['belong', 'our', 'nation', 'happy','pertain', 'inhere', 'appertain']
ntb = []
for key in need_to_belong:
tokens = tokenizer.tokenize(key)
for token in tokens:
ntb.append(token)
```
General
```
# 정부, 기관에 대한 신뢰
trust_in_goverment = ['goverment','president', 'leader', 'political', 'distrust', 'concern', 'doubt', 'suspect']
tig = []
for key in trust_in_goverment:
tokens = tokenizer.tokenize(key)
for token in tokens:
tig.append(token)
```
# Score measurement
```
def check_word_in_list(target, keywords):
NUM_WORD = 0
for word in keywords:
NUM_WORD = NUM_WORD + target.count(word)
return NUM_WORD
def Five_C_Score(text):
# 텍스트를 bert tokenizer로 분리
tokenized_text = tokenizer.tokenize(text)
score_pe = check_word_in_list(tokenized_text, perceived_effectiveness)
score_prv = check_word_in_list(tokenized_text, perceived_risk_vaccine)
score_nb = check_word_in_list(tokenized_text, normative_beliefs)
score_pb = check_word_in_list(tokenized_text, perceived_benefits)
score_rc = check_word_in_list(tokenized_text, perceived_risk_c19)
score_ci = check_word_in_list(tokenized_text, c19_infection)
score_opt = check_word_in_list(tokenized_text, optimism)
score_imp = check_word_in_list(tokenized_text, impulsivity)
score_alt = check_word_in_list(tokenized_text, altruism)
score_ntb = check_word_in_list(tokenized_text, need_to_belong)
score_tig = check_word_in_list(tokenized_text, trust_in_goverment)
C_score = [0,0,0,0,0]
C_score[0] = score_pe + score_prv + score_nb + score_opt - score_tig
C_score[1] = score_prv + score_rc + score_opt + score_imp
C_score[2] = score_nb + score_rc + score_ci
C_score[3] = score_opt + score_imp + score_alt
C_score[4] = -score_pb + score_rc + score_alt + score_ntb
c_max = C_score.index(max(C_score))
if C_score == [0,0,0,0,0]:
c_max = None
total_score = C_score[0] - C_score[1] - C_score[2] - C_score[3] + C_score[4]
return total_score,c_max
best_reason = []
score_list = []
for text in negative_tweet['text']:
c_score,c_index = Five_C_Score(text)
best_reason.append(c_index)
score_list.append(c_score)
print(negative_tweet['text'][15])
print(best_reason[15])
reason_list = [best_reason.count(0),best_reason.count(1),best_reason.count(2),best_reason.count(3),best_reason.count(4)]
plt.bar(five_C_model, reason_list)
plt.title('Number of Reasons')
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/patprem/IMDb-SentimentAnalysis/blob/main/SentimentAnalysis.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
**Sentiment Analysis of IMDb Movie Reviews**
Importing the basic and required libraries used in this project
```
import torch
from torchtext.legacy import data
from torchtext.legacy import datasets
import torchvision
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import torch.nn as nn
import torch.nn.functional as F
import random
```
Mounting personal Google Drive to load the dataset. **IMPORTANT: Change the directory and root path variable accordingly to yours.**
```
from google.colab import drive
import sys
#Mount your Google drive to the VM
drive.mount('/content/gdrive')
sys.path.append("/content/gdrive/My Drive/ECE4179 S1 2021 Prathik")
#set a root path variable to use
ROOT = "/content/gdrive/My Drive/ECE4179 S1 2021 Prathik/Final Project"
#Follow link and give permission, copy code and paste in text box
#You only have to do this once per session
```
Reading the data from the loaded dataset
**IMPORTANT:**
1. Download the dataset provided under Datasets section on README.md or download from this links: [IMDB Dataset (csv)](https://www.kaggle.com/lakshmi25npathi/sentiment-analysis-of-imdb-movie-reviews/data) and [Large Movie Review Dataset](https://ai.stanford.edu/~amaas/data/sentiment/).
2. Import the downloaded datasets onto your local Google Drive and **change the path variable** accordingly.
```
#from google.colab import files
#uploaded = files.upload()
#import io
#dataset = pd.read_csv(io.BytesIO(uploaded['IMDB Dataset.csv']))
# Dataset is now stored in a Pandas Dataframe
# Reading the data from the dataset.
dataset = pd.read_csv('gdrive/My Drive/ECE4179 S1 2021 Prathik/Final Project/IMDB Dataset.csv')
```
### If you have successfully executed all cells upto this point, then just simply click *Run all* under Runtime tab or press *Ctrl+F9* to execute the remanining cells or follow through the comments besides each cell below to get an understanding of the methodology of this project.
Exploring the loaded dataset
```
pd.set_option('display.max_colwidth',2000) # set the column width to 2000 so that we can read the complete review.
pd.set_option('max_rows', 200)
dataset.head(10) # setting .head(10) to read just the first 10 reviews from the dataset.
dataset.info() # information about the dataset; two columns: review and sentiment,
# where sentiment is the target column or the column that we need to predict.
# number of positive and negative reviews in the dataset.
# dataset is completely balanced and has equal number of positive and negative
# sentiments.
dataset['sentiment'].value_counts()
# reading second review from the dataset and checking how the contents of the review is
# and why we need to use NLP (Natural Language Processing) tasks on this dataset.
review = dataset['review'].loc[10]
review
```
From the above review (output), we can see that there HTML contents, punctuations, special characters, stopwords and others which do not offer much insight into the prediction of our model. The following NLP tasks (text cleaning technqiues) are implemented.
1. Eliminating HTML tags/contents like 'br"
2. Removing punctuations and special characters like |, /, apostrophes, commas and other punctuation marks and etc.
3. Remove stopwords that do not affect the prediction of our outcome and does not offer much insight such as 'are', 'is', 'the' and etc.
4. Use Lemmatization to bring back multiple forms of the same word to their common/base root. For example, words like 'ran', 'running', 'runs' to 'run'.
5. Using Text Tokenization and Vectorization to encode numerical values to our data after the above text cleaning techniques.
6. Lastly, fit these data to a deep learning model like Convolutional Neural Network (CNN) and LinearSVC model and compare the discrepancies between them
```
# Removing HTML contents like "<br>"
# BeautifulSoup is a Python library for extracting data out of HTML and XML files,
# by omitting HTML contents such as "<br>"
from bs4 import BeautifulSoup
soup = BeautifulSoup(review, "html.parser")
review = soup.get_text()
review
# notice that the HTML tags are eliminated.
# Removal of other special characters or punctuations except upper or lower case
# letters using Regular Expressions (Regex)
import re # importing Regex
review = re.sub('\[[^]]*\]', ' ', review) # removing punctuations
review = re.sub('[^a-zA-Z]', ' ', review) # regex; removing strings that contains a non-letter
# i.e., remove except a-z to A-Z
review
# set all characters to lower case for simplicity
review = review.lower()
review
```
Tokenization of reviews in the dataset
```
# Tokenization of reviews
# Stopwords removal: Split the text into tokens since stopwords removal
# works on every word in the text.
review = review.split()
review
```
Removal of Stopwords
```
# importing nltk library to remove stopwords
# Stopwords are words (English language words) that does not add much
# meaning to a sentence. Could be safely ignored without sacrificing the
# meaning of the sentence or review in this case. Words like 'he', 'have',
# 'the' does not provide any insights.
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
review = [word for word in review if not word in set(stopwords.words('english'))]
review
```
**Stemming technique**
Stemming is a process to extract the base form of the words by removing affixes from the words.
Both Stemming and Lemmatization technqiues are implemented on a sample review here to observe the discrepancies between them and why Lemmatization is a better algorithm.
```
# importing PorterStemmer library to perform stemming
from nltk.stem.porter import PorterStemmer
p_stem = PorterStemmer()
review_p_stem = [p_stem.stem(word) for word in review]
review_p_stem
```
**Lemmatization technique**
Lemmatization has the same objective as Stemming, however, it takes into consideration the morphological analysis of the words, i.e., it ensures that the root word is a valid English word alphabetically and meaningfully.
```
from nltk.stem import WordNetLemmatizer
nltk.download('wordnet')
lemma = WordNetLemmatizer()
review = [lemma.lemmatize(word) for word in review]
review
```
From the above results, we can notice that there is a huge difference between the techniques used. For example, 'little' has become 'littl' after Stemming, whereas it remained as 'little' after Lemmatization. Stemming tries to achieve a reduction in words to their root form but the stem itself is not a valid English word. Hence, Lemmatization is used in this project.
```
# merging the words to form a cleaned up version of the text.
review = ' '.join(review)
review
```
We can now see that the text is all cleaned up with no HTML tags, punctuations, special characters and stopwords, and it is ready for vectorization and training the model.
**Vectorization of reviews in the dataset**
```
# create a corpus to convert the text to mathematical forms or numeric values
corpus = [] # empty vector
corpus.append(review)
```
Two Vectorization techniques are applied to check the discrepancy between them and the technique with the highest accuracy will be choosen.
1. CountVectorizer (Bag of Words (BoW) Model)
2. Tfidf Vectorizer (Bag of Words (BoW) Model)
CountVectorizer (Bag of Words (BoW) Model)
```
# importing CountVectorizer to perform vectorization
# Data becomes numeric with 1,2,3s based on the number of times
# they appear in the text
from sklearn.feature_extraction.text import CountVectorizer
count_vect = CountVectorizer()
review_count_vect = count_vect.fit_transform(corpus) # fitting this technique
# onto the corpus
review_count_vect.toarray()
```
Tfidf Vectorizer (Bag of Words (BoW) Model)
1. Text Frequency (TF): how many times a word appears in a review
2. Inverse Document Frequency (IDF): log(total number of reviews/# reviews with that particular word)
TF-IDF score = TF*IDF
```
# importing TfidfVectorizer to perform vectorization
from sklearn.feature_extraction.text import TfidfVectorizer
# IDF acts as a diminishing factor and diminishes the weights of terms that
# occurs frequently in the text and increases the weights of the terms
# that occurs rarely.
tfidf_vect = TfidfVectorizer()
review_tfidf_vect = tfidf_vect.fit_transform(corpus)
review_tfidf_vect.toarray()
```
So far, the techniques mentioned above have been implemented on only one sample review. Now, the above techniques will be applied on all the reviews in the dataset. As there is no test dataset, the dataset is split into 25% of the data as test dataset to test the performance of the model.
```
# splitting the dataset into training and test data
# 25% of the data as test dataset and pseudo random generator
# to randomly distribute the reviews to each dataset
from sklearn.model_selection import train_test_split
train_dataset, test_dataset, traindata_label, testdata_label = train_test_split(dataset['review'], dataset['sentiment'], test_size=0.25, random_state=42)
# Convert the sentiments (target column) to numeric forms (1s and 0s) for simplicity
traindata_label = (traindata_label.replace({'positive': 1, 'negative': 0})).values
testdata_label = (testdata_label.replace({'positive': 1, 'negative': 0})).values
```
Implementation of text cleaning techniques discussed above on the whole dataset and build the train and test corpus.
```
# test and training corpus
train_corpus = []
test_corpus = []
# text cleaning techniques for training dataset
for i in range(train_dataset.shape[0]):
soup = BeautifulSoup(train_dataset.iloc[i], "html.parser")
review = soup.get_text()
review = re.sub('\[[^]]*\]', ' ', review)
review = re.sub('[^a-zA-Z]', ' ', review)
review = review.lower()
review = review.split()
review = [word for word in review if not word in set(stopwords.words('english'))]
lemma = WordNetLemmatizer()
review = [lemma.lemmatize(word) for word in review]
review = ' '.join(review)
train_corpus.append(review)
# text cleaning techniques for test dataset
for j in range(test_dataset.shape[0]):
soup = BeautifulSoup(test_dataset.iloc[j], "html.parser")
review = soup.get_text()
review = re.sub('\[[^]]*\]', ' ', review)
review = re.sub('[^a-zA-Z]', ' ', review)
review = review.lower()
review = review.split()
review = [word for word in review if not word in set(stopwords.words('english'))]
lemma = WordNetLemmatizer()
review = [lemma.lemmatize(word) for word in review]
review = ' '.join(review)
test_corpus.append(review)
```
Validate one sample entry
```
# training corpus
train_corpus[1]
# test corpus
test_corpus[1]
```
Vectorize the training and test corpus using TFIDF technique
```
# lower and upper boundary of the range of n-values for different word n-grams to be extracted.
# (1,3) means unigrams and trigrams.
tfidf_vect = TfidfVectorizer(ngram_range=(1, 3))
# fitting training corpus and test corpus onto TFIDF Vectorizer
tfidf_vect_train = tfidf_vect.fit_transform(train_corpus)
tfidf_vect_test = tfidf_vect.transform(test_corpus)
```
**First model: LinearSVC**
```
# importing LinearSVC library and fitting the data onto the model
from sklearn.svm import LinearSVC
# C: float; regularization parameter, must be positive.
# random_state: controls pseudo random number generation for
# shuffling data for dual coordinate descent.
linear_SVC = LinearSVC(C = 0.5, random_state = 42)
linear_SVC.fit(tfidf_vect_train, traindata_label)
predict = linear_SVC.predict(tfidf_vect_test)
```
LinearSVC with TFIDF Vectorization
```
# Check the performance of the model
from sklearn.metrics import classification_report, confusion_matrix, accuracy_score
print("Classification Report of LinearSVC model with TFIDF: \n", classification_report(testdata_label, predict,target_names=['Negative','Positive']))
print("Confusion Matrix of LinearSVC with TFIDF: \n", confusion_matrix(testdata_label, predict))
print("Accuracy of LinearSVC with TFIDF: \n", accuracy_score(testdata_label, predict))
import seaborn as sns
con_matrix = confusion_matrix(testdata_label, predict)
plt.figure(figsize = (10,10))
sns.heatmap(con_matrix, cmap= "Blues", linecolor = 'black', linewidth = 1, annot = True, fmt= '', xticklabels = ['Negative Reviews','Positive Reviews'], yticklabels = ['Negative Reviews','Positive Reviews'])
plt.xlabel("Predicted Sentiment")
plt.ylabel("Actual Sentiment")
```
LinearSVC with CountVectorizer (binary=False) Vectorization
```
# fitting the data onto the model using CountVectorizer technique
# binary = False -> If you set binary=True then CountVectorizer no longer uses the counts of terms/tokens.
# If a token is present in a document, it is 1, if absent it is 0 regardless of its frequency of occurrence.
# So you will be dealing with just binary values. By default, binary=False.
# If True, all non zero counts are set to 1. This is useful for discrete probabilistic models that model binary events rather than integer counts.
count_vect = CountVectorizer(ngram_range=(1, 3), binary = False) # lower and upper boundary
# of the range of n-values for different word n-grams to be extracted.
# (1,3) means unigrams and trigrams.
count_vect_train = count_vect.fit_transform(train_corpus)
count_vect_test = count_vect.transform(test_corpus)
linear_SVC_count = LinearSVC(C = 0.5, random_state = 42, max_iter = 5000)
linear_SVC_count.fit(count_vect_train, traindata_label)
predict_count = linear_SVC_count.predict(count_vect_test)
# Check the performance of the model
print("Classification Report of LinearSVC with CountVectorizer: \n", classification_report(testdata_label, predict_count,target_names=['Negative','Positive']))
print("Confusion Matrix of LinearSVC with CountVectorizer: \n", confusion_matrix(testdata_label, predict_count))
print("Accuracy of LinearSVC with CountVectorizer: \n", accuracy_score(testdata_label, predict_count))
con_matrix = confusion_matrix(testdata_label, predict_count)
plt.figure(figsize = (10,10))
sns.heatmap(con_matrix,cmap= "Blues", linecolor = 'black' , linewidth = 1 , annot = True, fmt='' , xticklabels = ['Negative Reviews','Positive Reviews'] , yticklabels = ['Negative Reviews','Positive Reviews'])
plt.xlabel("Predicted Sentiment")
plt.ylabel("Actual Sentiment")
```
From the above results, we can observe that **LinearSVC with TFIDF vectorization** gives the maximum accuracy and the outcome on our test dataset can be observed.
```
# prediction of data using the above model
predict_dataset = test_dataset.copy()
predict_dataset = pd.DataFrame(predict_dataset)
# setting columns of the predicted outcomes on the dataset
predict_dataset.columns = ['Review']
predict_dataset = predict_dataset.reset_index()
predict_dataset = predict_dataset.drop(['index'], axis=1)
# set the maximum column width to 100000 or more to view the complete review
pd.set_option('display.max_colwidth',100000)
pd.set_option('max_rows', 200)
predict_dataset.head(10)
# comparing the actual/original label with the predicted label
testactual_label = testdata_label.copy()
testactual_label = pd.DataFrame(testactual_label)
testactual_label.columns = ['Sentiment']
# replacing back the numeric forms of the sentiments to positive and negative respectively
testactual_label['Sentiment'] = testactual_label['Sentiment'].replace({1: 'positive', 0: 'negative'})
# predicted sentiments
testpredicted_label = predict.copy()
testpredicted_label = pd.DataFrame(testpredicted_label)
testpredicted_label.columns = ['Predicted Sentiment']
testpredicted_label['Predicted Sentiment'] = testpredicted_label['Predicted Sentiment'].replace({1: 'positive', 0: 'negative'})
# concatenate the original and predicted labels along with its corresponding review
test_result = pd.concat([predict_dataset, testactual_label, testpredicted_label], axis=1)
pd.set_option('display.max_colwidth',100000)
pd.set_option('max_rows', 200)
test_result.head(10)
```
**Second model: Convolutional Neural Network (CNN)**
Using CNN to conduct sentiment analysis
Preparing the data using a different dataset
```
n = 1234
random.seed(n)
np.random.seed(n)
torch.manual_seed(n)
torch.backends.cudnn.deterministic = True
# for convolutional layers
# batch dimension is first
# 'batch_first = true' argument used to tell torchtext to return the permuted data
# in CNN, batch dimension is first, so no need to permute data as 'batch_first' is set to true in TEXT field
TEXT = data.Field(tokenize = 'spacy', tokenizer_language = 'en_core_web_sm', batch_first = True)
LABEL = data.LabelField(dtype = torch.float)
# splitting the dataset into training and test data
train_dataset, test_dataset = datasets.IMDB.splits(TEXT, LABEL)
train_dataset, valid_dataset = train_dataset.split(random_state = random.seed(n))
# building the vocabulary and loading the pre-trained word embeddings
MAX_VOCAB_SIZE = 25_000
TEXT.build_vocab(train_dataset, max_size = MAX_VOCAB_SIZE, vectors = "glove.6B.100d", unk_init = torch.Tensor.normal_)
LABEL.build_vocab(train_dataset)
# creating the iterators
# batch size of 64 is used
BATCH_SIZE = 64
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
train_iterator, valid_iterator, test_iterator = data.BucketIterator.splits((train_dataset, valid_dataset, test_dataset), batch_size = BATCH_SIZE, device = device)
# checking the number of reviews in training, test and validation datasets
print(f'Training reviews: {len(train_dataset)}')
print(f'Validation reviews: {len(valid_dataset)}')
print(f'Testing reviews : {len(test_dataset)}')
```
Building a CNN for the dataset (Text is 1 dimensional)
1. Convert words into word embeddings to visualize words in 2 dimensions, each word along one axis and other axis for the elements of vectors.
2. Use a filter size of [n*width]. 'n' is the number of sequential words (n-grams, number of tokens in the review) and width is the dimensions of the word or dimensional embeddings (depth of filter).
3. Bi-grams are filters that covers two words at a time, tri-grams covers three words and so on. And each element of the filter has a weight associated with it.
4. The output of this filter is the weighted sum of all elements covered by the filter (single real number). Similarly, the filter moves to cover the next bi-gram and another output is calculated and so on.
5. This is an example of one such filter. CNNs has a plethora of these filters. The main idea is that each filter will learn a different feature to extract. For example, each of the [2*width] filters looks for the occurence of different bi-grams that are relevant for analysing sentiment of movie reviews. And the same goes for different sizes of filters (n-grams) with heights of 3,4,5 etc.
6. Then, use max pooling on the output of the convolutional layers, which takes the maximum value over a dimension.
7. The maximum value is the most important feature for determining the sentiment of the review, which corresponds to the most essential n-gram within the review. Through backpropagation, the weights of the filters are updated so that whenever certain n-grams that are highly indicative of the sentiment are seen, the output of the filter is a high or the highest value amongst all. This high value is then passed through the max pooling layer if it is the maximum value in the output.
8. This model has 100 filters of 3 different sizes (n-grams), i.e., 300 different n-grams. Later, these are concatenated into a single vector and passed through a linear layer to predict the sentiment.
9. Most importantly, input review has to be atleast as long as the largest filter height used.
```
import torch.nn as nn
import torch.nn.functional as F
# implementing the convolutional layers (nn.Conv2d)
class CNN(nn.Module):
def __init__(self, vocab_size, embedding_dim, n_filters, filter_sizes, output_dim, dropout, pad_idx):
# in_channels: number of channels in text/image fed into convolutional layer
# in text, only one single channel
# in_channels: number of filters
# kernel_size: size of filters (n*emb_dim); n is the size of n-grams
# and emb_dim is the dimensional embedding or width of the text
super().__init__()
self.embedding = nn.Embedding(vocab_size, embedding_dim, padding_idx = pad_idx)
self.conv_0 = nn.Conv2d(in_channels = 1, out_channels = n_filters, kernel_size = (filter_sizes[0], embedding_dim))
self.conv_1 = nn.Conv2d(in_channels = 1, out_channels = n_filters, kernel_size = (filter_sizes[1], embedding_dim))
self.conv_2 = nn.Conv2d(in_channels = 1, out_channels = n_filters, kernel_size = (filter_sizes[2], embedding_dim))
self.fc = nn.Linear(len(filter_sizes) * n_filters, output_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, text):
# pass review to an embedding layer to get embeddings
# second dimension of the input to nn.Conv2d is the channel dimension
embed_done = self.embedding(text)
# text has no channel dimension, so unsqueeze to make one
# and matches with in_channels (=1) dimension
embed_done = embed_done.unsqueeze(1)
# pass tensors through convolutional and pooling layers using ReLU
# (non-linearity) activation function after the conv layers
conv_layer0 = F.relu(self.conv_0(embed_done).squeeze(3))
conv_layer1 = F.relu(self.conv_1(embed_done).squeeze(3))
conv_layer2 = F.relu(self.conv_2(embed_done).squeeze(3))
# pooling layers handles reviews of different lengths
# with max pooling, input to linear layer is the total no. of filters
max_pool0 = F.max_pool1d(conv_layer0, conv_layer0.shape[2]).squeeze(2)
max_pool1 = F.max_pool1d(conv_layer1, conv_layer1.shape[2]).squeeze(2)
max_pool2 = F.max_pool1d(conv_layer2, conv_layer2.shape[2]).squeeze(2)
# output size of conv layers depends on the input size
# different batches contains reviews of different lengths
# lastly, apply dropout on the concatenated filter outputs
concatenation = self.dropout(torch.cat((max_pool0, max_pool1, max_pool2), dim = 1))
# pass through a linear layer (fully-connected layer) to make predictions
return self.fc(concatenation)
```
The above CNN uses only 3 different sized filters. The below code is a generic CNN that takes in any number of filters.
```
# place all conv layers in a nn.ModuleList - function in PyTorch to hold a list
# of PyTorch nn.Module
# pass arbitrary sized list of filter sizes (generic model)
# creates a conv layer for each
class CNN(nn.Module):
def __init__(self, vocab_size, embedding_dim, n_filters, filter_sizes, output_dim, dropout, pad_idx):
super().__init__()
self.embedding = nn.Embedding(vocab_size, embedding_dim, padding_idx = pad_idx)
self.convs = nn.ModuleList([nn.Conv2d(in_channels = 1, out_channels = n_filters, kernel_size = (fs, embedding_dim)) for fs in filter_sizes])
self.fc = nn.Linear(len(filter_sizes) * n_filters, output_dim)
self.dropout = nn.Dropout(dropout)
# iterate through the list applying each conv layer to get a list of
# conv outputs which is fed into max pooling layer in a list
# comprehension before concatenation and passing through dropout
# and linear layers
def forward(self, text):
embed_done = self.embedding(text)
embed_done = embed_done.unsqueeze(1)
conv_layer_relu = [F.relu(conv(embed_done)).squeeze(3) for conv in self.convs]
max_pool_drop = [F.max_pool1d(conv, conv.shape[2]).squeeze(2) for conv in conv_layer_relu]
concatenation = self.dropout(torch.cat(max_pool_drop, dim = 1))
return self.fc(concatenation)
```
Creating an instance of our CNN model
```
dimension_input = len(TEXT.vocab)
# dimensional embeddings
dimn_embedding = 100
# number of filters
number_filters = 100
# size of the filters
size_filter = [3,4,5]
# output size
dimension_output = 1
# dropout (value of 'p')
p = 0.5
# padding
padding = TEXT.vocab.stoi[TEXT.pad_token]
# applying all these to the CNN
model = CNN(dimension_input, dimn_embedding, number_filters, size_filter, dimension_output, p, padding)
# check number of parameters in CNN model
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters')
```
Loading the pre-trained embeddings
```
embed_pretrain = TEXT.vocab.vectors
# weights
model.embedding.weight.data.copy_(embed_pretrain)
# zero the initial weights of the unknown and padding tokens
token = TEXT.vocab.stoi[TEXT.unk_token]
model.embedding.weight.data[token] = torch.zeros(dimn_embedding)
model.embedding.weight.data[padding] = torch.zeros(dimn_embedding)
```
Next, now it is ready to train our model. The optimizer and loss function (criterion) are initialized. Here, I have used the ADAM optimizer and Binary Cross Entropy with Logits Loss function.
```
# importing ADAM optimizer
import torch.optim as optim
# set ADAM optimizer
optimizer = optim.Adam(model.parameters())
# set the loss function
criterion = nn.BCEWithLogitsLoss()
# set model and criterion on GPU
model = model.to(device)
criterion = criterion.to(device)
```
Implementing a function to calculate accuracy in order to check the performance of the model
```
# returns accuracy per batch, will return, for example, 0.8 instead of 8.
def binary_accuracy(preds, y):
# rounds predictions to the closest integer
predictions_rounded = torch.round(torch.sigmoid(preds))
true_prediction = (predictions_rounded == y).float() # float better for division purposes
accuracy = true_prediction.sum() / len(true_prediction)
return accuracy
# function for training the model
def train(model, iterator, optimizer, criterion):
# initialise the epoch loss and accuracy
epoch_accuracy = 0
epoch_loss = 0
model.train() # to ensure dropout is turned ON while training
for batch in iterator:
optimizer.zero_grad()
predictions = model(batch.text).squeeze(1)
loss = criterion(predictions, batch.label)
accuracy = binary_accuracy(predictions, batch.label)
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_accuracy += accuracy .item()
return epoch_loss / len(iterator), epoch_accuracy / len(iterator)
# function for testing the model
def evaluate(model, iterator, criterion):
# initialise the epoch loss and accuracy
epoch_loss = 0
epoch_accuracy = 0
model.eval() # to ensure dropout is turned OFF while evaluating/testing
with torch.no_grad():
for batch in iterator:
predictions = model(batch.text).squeeze(1)
loss = criterion(predictions, batch.label)
accuracy = binary_accuracy(predictions, batch.label)
epoch_loss += loss.item()
epoch_accuracy += accuracy.item()
return epoch_loss / len(iterator), epoch_accuracy / len(iterator)
# importing time library to define function to tell the time taken of our
# epochs
import time
def epoch_time(start_time, end_time):
time_taken = end_time - start_time
time_taken_mins = int(time_taken / 60)
time_taken_secs = int(time_taken - (time_taken_mins * 60))
return time_taken_mins, time_taken_secs
```
**Training the CNN model**
```
# 5 epochs are enough to view the values of loss and accuracy
number_epochs = 5
good_validationloss = float('inf') # set to float
for epoch in range(number_epochs):
start_time = time.time()
# calculating the training loss and accuracy and the validation loss
# and accuracy
train_loss, train_accuracy = train(model, train_iterator, optimizer, criterion)
valid_loss, valid_accuracy = evaluate(model, valid_iterator, criterion)
end_time = time.time()
epoch_minutes, epoch_secs = epoch_time(start_time, end_time)
if valid_loss < good_validationloss:
good_validationloss = valid_loss
torch.save(model.state_dict(), 'tut4-model.pt')
# print the training loss and accuracy and validation loss and accuracy
print(f'Epoch: {epoch+1:02} | Epoch Time: {epoch_minutes}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Train Acc: {train_accuracy*100:.2f}%')
print(f'\t Val. Loss: {valid_loss:.3f} | Val. Acc: {valid_accuracy*100:.2f}%')
```
This function will prompt the user to input their reviews. Based on the review, the model will the predict whether the sentiment of the review is positive or negative along with how accurate the model predicts the sentiment.
```
import spacy
prompt = spacy.load('en_core_web_sm')
# minimum_length is set to 10000 so that utmost 10000 tokens are accepted for computing
# the outcome, i.e., 10000 words in a review which is more than enough
def classify_predict_sentiment(model, sentence, minimum_length = 10000):
model.eval()
tokenization_done = [tok.text for tok in prompt.tokenizer(sentence)]
# classify_predict_sentiment function accepts minimum length argument also by changing
# minimum_length
# If tokenization_done input sentence is less than minimum_length tokens, then we append
# padding tokens ('<pad>') to make it minimum_length tokens
if len(tokenization_done) < minimum_length:
tokenization_done += ['<pad>'] * (minimum_length - len(tokenization_done))
indexing = [TEXT.vocab.stoi[t] for t in tokenization_done]
box = torch.LongTensor(indexing).to(device)
box = box.unsqueeze(0)
prediction = torch.sigmoid(model(box))
# if the accuracy of the review is less than 0.5, it shall be considered
# a negative review and anything above 0.5 shall be considered a positive
# review
if prediction.item() < 0.5:
print(f'Negative Review')
else:
print(f'Positive Review')
return print(f'Accuracy of this review: {prediction.item():.8f}')
```
The following positive and negative reviews are fed into the model and the outcome is displayed along with the accuracy from the model, i.e., how accurate the model predicts whether it is a positive or negative review.
```
classify_predict_sentiment(model, "I thought this was a wonderful way to spend time on a too hot summer weekend, sitting in the air conditioned theater and watching a light-hearted comedy. The plot is simplistic, but the dialogue is witty and the characters are likable (even the well bread suspected serial killer). While some may be disappointed when they realize this is not Match Point 2: Risk Addiction, I thought it was proof that Woody Allen is still fully in control of the style many of us have grown to love.<br /><br />This was the most I'd laughed at one of Woody's comedies in years (dare I say a decade?). While I've never been impressed with Scarlet Johanson, in this she managed to tone down her sexy image and jumped right into a average, but spirited young woman.<br /><br />This may not be the crown jewel of his career, but it was wittier than Devil Wears Prada and more interesting than Superman a great comedy to go see with friends.")
classify_predict_sentiment(model, "This show was an amazing, fresh & innovative idea in the 70's when it first aired. The first 7 or 8 years were brilliant, but things dropped off after that. By 1990, the show was not really funny anymore, and it's continued its decline further to the complete waste of time it is today.<br /><br />It's truly disgraceful how far this show has fallen. The writing is painfully bad, the performances are almost as bad - if not for the mildly entertaining respite of the guest-hosts, this show probably wouldn't still be on the air. I find it so hard to believe that the same creator that hand-selected the original cast also chose the band of hacks that followed. How can one recognize such brilliance and then see fit to replace it with such mediocrity? I felt I must give 2 stars out of respect for the original cast that made this show such a huge success. As it is now, the show is just awful. I can't believe it's still on the air.")
classify_predict_sentiment(model, "This a fantastic movie of three prisoners who become famous. One of the actors is george clooney and I'm not a fan but this roll is not bad. Another good thing about the movie is the soundtrack (The man of constant sorrow). I recommand this movie to everybody. Greetings Bart")
classify_predict_sentiment(model,"I saw this movie when I was about 12 when it came out. I recall the scariest scene was the big bird eating men dangling helplessly from parachutes right out of the air. The horror. The horror.<br /><br />As a young kid going to these cheesy B films on Saturday afternoons, I still was tired of the formula for these monster type movies that usually included the hero, a beautiful woman who might be the daughter of a professor and a happy resolution when the monster died in the end. I didn't care much for the romantic angle as a 12 year old and the predictable plots. I love them now for the unintentional humor.<br /><br />But, about a year or so later, I saw Psycho when it came out and I loved that the star, Janet Leigh, was bumped off early in the film. I sat up and took notice at that point. Since screenwriters are making up the story, make it up to be as scary as possible and not from a well-worn formula. There are no rules.")
classify_predict_sentiment(model,"The Karen Carpenter Story shows a little more about singer Karen Carpenter's complex life. Though it fails in giving accurate facts, and details.<br /><br />Cynthia Gibb (portrays Karen) was not a fine election. She is a good actress , but plays a very naive and sort of dumb Karen Carpenter. I think that the role needed a stronger character. Someone with a stronger personality.<br /><br />Louise Fletcher role as Agnes Carpenter is terrific, she does a great job as Karen's mother.<br /><br />It has great songs, which could have been included in a soundtrack album. Unfortunately they weren't, though this movie was on the top of the ratings in USA and other several countries.")
classify_predict_sentiment(model,"I watched this film not really expecting much, I got it in a pack of 5 films, all of which were pretty terrible in their own way for under a fiver so what could I expect? and you know what I was right, they were all terrible, this movie has a few (and a few is stretching it) interesting points, the occasional camcorder view is a nice touch, the drummer is very like a drummer, i.e damned annoying and, well thats about it actually, the problem is that its just so boring, in what I can only assume was an attempt to build tension, a whole lot of nothing happens and when it does its utterly tedious (I had my thumb on the fast forward button, ready to press for most of the movie, but gave it a go) and seriously is the lead singer of the band that great looking, coz they don't half mention how beautiful he is a hell of a lot, I thought he looked a bit like a meercat, all this and I haven't even mentioned the killer, I'm not even gonna go into it, its just not worth explaining. Anyway as far as I'm concerned Star and London are just about the only reason to watch this and with the exception of London (who was actually quite funny) it wasn't because of their acting talent, I've certainly seen a lot worse, but I've also seen a lot better. Best avoid unless your bored of watching paint dry.")
```
| github_jupyter |
### Linear Problem
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torch
from torch.utils.data import DataLoader, Dataset
import seaborn as sns
from torch import nn
from torch.nn import functional as F
```
### Data Preparation
```
data = pd.read_csv('data/test.csv')
data.head()
sns.scatterplot(data=data, x='x', y='y',hue='color')
class Data(Dataset):
def __init__(self, path, transform=None, shuffle=True):
self.dataFrame = pd.read_csv(path)
self.xy = pd.read_csv(path).values
if shuffle:
np.random.shuffle(self.xy)
self.len = self.xy.shape[0]
self.x = self.xy[:, :-1]
self.y = self.xy[:, -1]
self.transform = transform
print(self.x.shape)
def __getitem__(self, index):
sample = self.x[index], self.y[index]
if self.transform:
sample = self.transform(sample)
return sample
def __len__(self):
return self.len
def plot(self):
sns.scatterplot(data=self.dataFrame, x='x', y='y',hue='color')
plt.show()
```
### Transformers on our data
```
class ToTensor:
def __call__(self, samples):
x, y = samples
return torch.from_numpy(x.astype('float32')) ,torch.from_numpy(np.array(y, dtype='float32'))
train = Data(path='data/train.csv', transform=ToTensor(), shuffle=True)
test = Data(path='data/test.csv', transform=ToTensor(),shuffle=True )
train.plot()
test.plot()
train_set = DataLoader(dataset=train,
batch_size =5,
shuffle=True)
test_set = DataLoader(dataset=test,
batch_size =5,
shuffle=False)
```
### Predicting the Color
```
class Net(nn.Module):
def __init__(self):
super().__init__()
self.fc1 = nn.Linear(2,32)
self.fc2 = nn.Linear(32,64)
self.fc3 = nn.Linear(64, 1)
def forward(self,x):
x = torch.relu(self.fc1(x))
x = torch.relu(self.fc2(x))
x = torch.sigmoid(self.fc3(x))
return x
net = Net()
net
criterion = torch.nn.BCELoss()
optimizer = torch.optim.SGD(net.parameters(), lr=0.01)
EPOCHS = 5
for epoch in range(EPOCHS):
print(f'Epochs: {epoch+1}/{EPOCHS}')
for data in train_set:
X, y = data
optimizer.zero_grad()
# forward pass
output = net(X.view(-1, 2))
#calculate loss
loss = criterion(output, y.unsqueeze(1))
## backward pass
loss.backward()
# update the weights
optimizer.step()
print("loss: ", loss.item())
total, correct = 0, 0
with torch.no_grad():
for data in test_set:
X, y = data
outputs = net(X.view(-1, 2))
for i in range(len(torch.round(outputs))):
if y[i] == torch.round(outputs[i]):
correct +=1
total +=1
print(correct/total)
total, correct = 0, 0
with torch.no_grad():
for data in train_set:
X, y = data
outputs = net(X.view(-1, 2))
for i in range(len(torch.round(outputs))):
if y[i] == torch.round(outputs[i]):
correct +=1
total +=1
print(correct/total)
```
### Making Predictions
```
test.plot()
test[0]
torch.Tensor([1., 0.])
torch.round(net(torch.Tensor([1., 2.]))).item()
```
> Done
| github_jupyter |
# NBAiLab - Finetuning and Evaluating a BERT model for NER and POS
<img src="https://raw.githubusercontent.com/NBAiLab/notram/master/images/nblogo_2.png">
In this notebook we will finetune the [NB-BERTbase Model](https://github.com/NBAiLab/notram) released by the National Library of Norway. This is a model trained on a large corpus (110GB) of Norwegian texts.
We will finetune this model on the [NorNE dataset](https://github.com/ltgoslo/norne). for Named Entity Recognition (NER) and Part of Speech (POS) tags using the [Transformers Library by Huggingface](https://huggingface.co/transformers/). After training the model should be able to accept any text string input (up to 512 tokens) and return POS or NER-tags for this text. This is useful for a number of NLP tasks, for instance for extracting/removing names/places from a document. After training, we will save the model, evaluate it and use it for predictions.
The Notebook is intended for experimentation with the pre-release NoTram models from the National Library of Norway, and is made for educational purposes. If you just want to use the model, you can instead initiate one of our finetuned models.
## Before proceeding
Create a copy of this notebook by going to "File - Save a Copy in Drive"
# Install Dependencies and Define Helper Functions
You need to run the code below to install some libraries and initiate some helper functions. Click "Show Code" if you later want to examine this part as well.
```
#@title
#The notebook is using some functions for reporting that are only available in Transformers 4.2.0. Until that is released, we are installing from the source.
!pip -q install https://github.com/huggingface/transformers/archive/0ecbb698064b94560f24c24fbfbd6843786f088b.zip
!pip install -qU scikit-learn datasets seqeval conllu pyarrow
import logging
import os
import sys
from dataclasses import dataclass
from dataclasses import field
from typing import Optional
import numpy as np
import pandas as pd
import tensorflow as tf
import tensorflow_datasets as tfds
import transformers
from datasets import load_dataset
from seqeval.metrics import accuracy_score
from seqeval.metrics import f1_score
from seqeval.metrics import precision_score
from seqeval.metrics import recall_score
from seqeval.metrics import classification_report
from transformers.training_args import TrainingArguments
from tqdm import tqdm
from transformers import (
AutoConfig,
AutoModelForTokenClassification,
AutoTokenizer,
DataCollatorForTokenClassification,
PreTrainedTokenizerFast,
Trainer,
TrainingArguments,
pipeline,
set_seed
)
from google.colab import output
from IPython.display import Markdown
from IPython.display import display
# Helper Funtions - Allows us to format output by Markdown
def printm(string):
display(Markdown(string))
## Preprocessing the dataset
# Tokenize texts and align the labels with them.
def tokenize_and_align_labels(examples):
tokenized_inputs = tokenizer(
examples[text_column_name],
max_length=max_length,
padding=padding,
truncation=True,
# We use this argument because the texts in our dataset are lists of words (with a label for each word).
is_split_into_words=True,
)
labels = []
for i, label in enumerate(examples[label_column_name]):
word_ids = tokenized_inputs.word_ids(batch_index=i)
previous_word_idx = None
label_ids = []
for word_idx in word_ids:
# Special tokens have a word id that is None. We set the label to -100 so they are automatically
# ignored in the loss function.
if word_idx is None:
label_ids.append(-100)
# We set the label for the first token of each word.
elif word_idx != previous_word_idx:
label_ids.append(label_to_id[label[word_idx]])
# For the other tokens in a word, we set the label to either the current label or -100, depending on
# the label_all_tokens flag.
else:
label_ids.append(label_to_id[label[word_idx]] if label_all_tokens else -100)
previous_word_idx = word_idx
labels.append(label_ids)
tokenized_inputs["labels"] = labels
return tokenized_inputs
# Metrics
def compute_metrics(pairs):
predictions, labels = pairs
predictions = np.argmax(predictions, axis=2)
# Remove ignored index (special tokens)
true_predictions = [
[label_list[p] for (p, l) in zip(prediction, label) if l != -100]
for prediction, label in zip(predictions, labels)
]
true_labels = [
[label_list[l] for (p, l) in zip(prediction, label) if l != -100]
for prediction, label in zip(predictions, labels)
]
return {
"accuracy_score": accuracy_score(true_labels, true_predictions),
"precision": precision_score(true_labels, true_predictions),
"recall": recall_score(true_labels, true_predictions),
"f1": f1_score(true_labels, true_predictions),
"report": classification_report(true_labels, true_predictions, digits=4)
}
```
# Settings
Try running this with the default settings first. The default setting should give you a pretty good result. If you want training to go even faster, reduce the number of epochs. The first variables you should consider changing are the one in the dropdown menus. Later you can also experiment with the other settings to get even better results.
```
#Model, Dataset, and Task
#@markdown Set the main model that the training should start from
model_name = 'NbAiLab/nb-bert-base' #@param ["NbAiLab/nb-bert-base", "bert-base-multilingual-cased"]
#@markdown ---
#@markdown Set the dataset for the task we are training on
dataset_name = "NbAiLab/norne" #@param ["NbAiLab/norne", "norwegian_ner"]
dataset_config = "bokmaal" #@param ["bokmaal", "nynorsk"]
task_name = "ner" #@param ["ner", "pos"]
#General
overwrite_cache = False #@#param {type:"boolean"}
cache_dir = ".cache" #param {type:"string"}
output_dir = "./output" #param {type:"string"}
overwrite_output_dir = False #param {type:"boolean"}
seed = 42 #param {type:"number"}
set_seed(seed)
#Tokenizer
padding = False #param ["False", "'max_length'"] {type: 'raw'}
max_length = 512 #param {type: "number"}
label_all_tokens = False #param {type:"boolean"}
# Training
#@markdown ---
#@markdown Set training parameters
per_device_train_batch_size = 8 #param {type: "integer"}
per_device_eval_batch_size = 8 #param {type: "integer"}
learning_rate = 3e-05 #@param {type: "number"}
weight_decay = 0.0 #param {type: "number"}
adam_beta1 = 0.9 #param {type: "number"}
adam_beta2 = 0.999 #param {type: "number"}
adam_epsilon = 1e-08 #param {type: "number"}
max_grad_norm = 1.0 #param {type: "number"}
num_train_epochs = 4.0 #@param {type: "number"}
num_warmup_steps = 750 #@param {type: "number"}
save_total_limit = 1 #param {type: "integer"}
load_best_model_at_end = True #@param {type: "boolean"}
```
# Load the Dataset used for Finetuning
The default setting is to use the NorNE dataset. This is currently the largest (and best) dataset with annotated POS/NER tags that are available today. All sentences is tagged both for POS and NER. The dataset is available as a Huggingface dataset, so loading it is very easy.
```
#Load the dataset
dataset = load_dataset(dataset_name, dataset_config)
#Getting some variables from the dataset
column_names = dataset["train"].column_names
features = dataset["train"].features
text_column_name = "tokens" if "tokens" in column_names else column_names[0]
label_column_name = (
f"{task_name}_tags" if f"{task_name}_tags" in column_names else column_names[1]
)
label_list = features[label_column_name].feature.names
label_to_id = {i: i for i in range(len(label_list))}
num_labels = len(label_list)
#Look at the dataset
printm(f"###Quick Look at the NorNE Dataset")
print(dataset["train"].data.to_pandas()[[text_column_name, label_column_name]])
printm(f"###All labels ({num_labels})")
print(label_list)
if task_name == "ner":
mlabel_list = {label.split("-")[-1] for label in label_list}
printm(f"###Main labels ({len(mlabel_list)})")
print(mlabels)
```
# Initialize Training
We are here using the native Trainer interface provided by Huggingface. Huggingface also has an interface for Tensorflow and PyTorch. To see an example of how to use the Tensorflow interface, please take a look at our notebook about classification.
```
config = AutoConfig.from_pretrained(
model_name,
num_labels=num_labels,
finetuning_task=task_name,
cache_dir=cache_dir,
)
tokenizer = AutoTokenizer.from_pretrained(
model_name,
cache_dir=cache_dir,
use_fast=True,
)
model = AutoModelForTokenClassification.from_pretrained(
model_name,
from_tf=bool(".ckpt" in model_name),
config=config,
cache_dir=cache_dir,
)
data_collator = DataCollatorForTokenClassification(tokenizer)
tokenized_datasets = dataset.map(
tokenize_and_align_labels,
batched=True,
load_from_cache_file=not overwrite_cache,
num_proc=os.cpu_count(),
)
training_args = TrainingArguments(
output_dir=output_dir,
overwrite_output_dir=overwrite_output_dir,
do_train=True,
do_eval=True,
do_predict=True,
per_device_train_batch_size=per_device_train_batch_size,
per_device_eval_batch_size=per_device_eval_batch_size,
learning_rate=learning_rate,
weight_decay=weight_decay,
adam_beta1=adam_beta1,
adam_beta2=adam_beta2,
adam_epsilon=adam_epsilon,
max_grad_norm=max_grad_norm,
num_train_epochs=num_train_epochs,
warmup_steps=num_warmup_steps,
load_best_model_at_end=load_best_model_at_end,
seed=seed,
save_total_limit=save_total_limit,
)
# Initialize our Trainer
trainer = Trainer(
model=model,
args=training_args,
train_dataset=tokenized_datasets["train"],
eval_dataset=tokenized_datasets["validation"],
tokenizer=tokenizer,
data_collator=data_collator,
compute_metrics=compute_metrics,
)
```
# Start Training
Training for the default 4 epochs should take around 10-15 minutes if you have access to GPU.
```
%%time
train_result = trainer.train()
trainer.save_model() # Saves the tokenizer too for easy upload
# Need to save the state, since Trainer.save_model saves only the tokenizer with the model
trainer.state.save_to_json(os.path.join(training_args.output_dir, "trainer_state.json"))
#Print Results
output_train_file = os.path.join(output_dir, "train_results.txt")
with open(output_train_file, "w") as writer:
printm("**Train results**")
for key, value in sorted(train_result.metrics.items()):
printm(f"{key} = {value}")
writer.write(f"{key} = {value}\n")
```
# Evaluate the Model
The model is now saved on your Colab disk. This is a temporary disk that will disappear when the Colab is closed. You should copy it to another place if you want to keep the result. Now we can evaluate the model and play with it. Expect some UserWarnings since there might be errors in the training file.
```
printm("**Evaluate**")
results = trainer.evaluate()
output_eval_file = os.path.join(output_dir, "eval_results.txt")
with open(output_eval_file, "w") as writer:
printm("**Eval results**")
for key, value in results.items():
printm(f"{key} = {value}")
writer.write(f"{key} = {value}\n")
```
# Run Preditions on the Test Dataset
You should be able to end up with a result not far from what we have reported for the NB-BERT-model:
<table align="left">
<tr><td></td><td>Bokmål</td><td>Nynorsk</td></tr>
<tr><td>POS</td><td>98.86</td><td>98.77</td></tr>
<tr><td>NER</td><td>93.66</td><td>92.02</td></tr>
</table>
```
printm("**Predict**")
test_dataset = tokenized_datasets["test"]
predictions, labels, metrics = trainer.predict(test_dataset)
predictions = np.argmax(predictions, axis=2)
output_test_results_file = os.path.join(output_dir, "test_results.txt")
with open(output_test_results_file, "w") as writer:
printm("**Predict results**")
for key, value in sorted(metrics.items()):
printm(f"{key} = {value}")
writer.write(f"{key} = {value}\n")
# Remove ignored index (special tokens)
true_predictions = [
[label_list[p] for (p, l) in zip(prediction, label) if l != -100]
for prediction, label in zip(predictions, labels)
]
# Save predictions
output_test_predictions_file = os.path.join(output_dir, "test_predictions.txt")
with open(output_test_predictions_file, "w") as writer:
for prediction in true_predictions:
writer.write(" ".join(prediction) + "\n")
```
# Use the model
This model will assign labels to the different word/tokens. B-TAG marks the beginning of the entity, while I-TAG is a continuation of the entity. In the example below the model should be able to pick out the individual names as well as understand how many places and organisations that are mentioned.
```
text = "Svein Arne Brygfjeld, Freddy Wetjen, Javier de la Rosa og Per E Kummervold jobber alle ved AILABen til Nasjonalbiblioteket. Nasjonalbiblioteket har lokaler b\xE5de i Mo i Rana og i Oslo. " #@param {type:"string"}
group_entities = True #param {type:"boolean"}
#Load the saved model in the pipeline, and run some predicions
model = AutoModelForTokenClassification.from_pretrained(output_dir)
try:
tokenizer = AutoTokenizer.from_pretrained(output_dir)
except TypeError:
tokenizer = AutoTokenizer.from_pretrained(model_name)
ner_model = pipeline(
"ner", model=model, tokenizer=tokenizer, grouped_entities=group_entities
)
result = ner_model(text)
output = []
for token in result:
entity = int(token['entity_group'].replace("LABEL_", ""))
output.append({
"word": token['word'],
"entity": label_list[entity],
"score": token['score'],
})
pd.DataFrame(output).style.hide_index()
```
---
##### Copyright 2020 © National Library of Norway
| github_jupyter |
# Table of Contents
<p><div class="lev1"><a href="#Introduction-to-Pandas"><span class="toc-item-num">1 </span>Introduction to Pandas</a></div><div class="lev2"><a href="#Pandas-Data-Structures"><span class="toc-item-num">1.1 </span>Pandas Data Structures</a></div><div class="lev3"><a href="#Series"><span class="toc-item-num">1.1.1 </span>Series</a></div><div class="lev3"><a href="#DataFrame"><span class="toc-item-num">1.1.2 </span>DataFrame</a></div><div class="lev3"><a href="#Exercise-1"><span class="toc-item-num">1.1.3 </span>Exercise 1</a></div><div class="lev3"><a href="#Exercise-2"><span class="toc-item-num">1.1.4 </span>Exercise 2</a></div><div class="lev2"><a href="#Importing-data"><span class="toc-item-num">1.2 </span>Importing data</a></div><div class="lev3"><a href="#Microsoft-Excel"><span class="toc-item-num">1.2.1 </span>Microsoft Excel</a></div><div class="lev2"><a href="#Pandas-Fundamentals"><span class="toc-item-num">1.3 </span>Pandas Fundamentals</a></div><div class="lev3"><a href="#Manipulating-indices"><span class="toc-item-num">1.3.1 </span>Manipulating indices</a></div><div class="lev2"><a href="#Indexing-and-Selection"><span class="toc-item-num">1.4 </span>Indexing and Selection</a></div><div class="lev3"><a href="#Exercise-3"><span class="toc-item-num">1.4.1 </span>Exercise 3</a></div><div class="lev2"><a href="#Operations"><span class="toc-item-num">1.5 </span>Operations</a></div><div class="lev2"><a href="#Sorting-and-Ranking"><span class="toc-item-num">1.6 </span>Sorting and Ranking</a></div><div class="lev3"><a href="#Exercise-4"><span class="toc-item-num">1.6.1 </span>Exercise 4</a></div><div class="lev2"><a href="#Hierarchical-indexing"><span class="toc-item-num">1.7 </span>Hierarchical indexing</a></div><div class="lev2"><a href="#Missing-data"><span class="toc-item-num">1.8 </span>Missing data</a></div><div class="lev3"><a href="#Exercise-5"><span class="toc-item-num">1.8.1 </span>Exercise 5</a></div><div class="lev2"><a href="#Data-summarization"><span class="toc-item-num">1.9 </span>Data summarization</a></div><div class="lev2"><a href="#Writing-Data-to-Files"><span class="toc-item-num">1.10 </span>Writing Data to Files</a></div><div class="lev3"><a href="#Advanced-Exercise:-Compiling-Ebola-Data"><span class="toc-item-num">1.10.1 </span>Advanced Exercise: Compiling Ebola Data</a></div><div class="lev2"><a href="#References"><span class="toc-item-num">1.11 </span>References</a></div>
# Introduction to Pandas
**pandas** is a Python package providing fast, flexible, and expressive data structures designed to work with *relational* or *labeled* data both. It is a fundamental high-level building block for doing practical, real world data analysis in Python.
pandas is well suited for:
- Tabular data with heterogeneously-typed columns, as in an SQL table or Excel spreadsheet
- Ordered and unordered (not necessarily fixed-frequency) time series data.
- Arbitrary matrix data (homogeneously typed or heterogeneous) with row and column labels
- Any other form of observational / statistical data sets. The data actually need not be labeled at all to be placed into a pandas data structure
Key features:
- Easy handling of **missing data**
- **Size mutability**: columns can be inserted and deleted from DataFrame and higher dimensional objects
- Automatic and explicit **data alignment**: objects can be explicitly aligned to a set of labels, or the data can be aligned automatically
- Powerful, flexible **group by functionality** to perform split-apply-combine operations on data sets
- Intelligent label-based **slicing, fancy indexing, and subsetting** of large data sets
- Intuitive **merging and joining** data sets
- Flexible **reshaping and pivoting** of data sets
- **Hierarchical labeling** of axes
- Robust **IO tools** for loading data from flat files, Excel files, databases, and HDF5
- **Time series functionality**: date range generation and frequency conversion, moving window statistics, moving window linear regressions, date shifting and lagging, etc.
```
import pandas as pd
import numpy as np
pd.options.mode.chained_assignment = None # default='warn'
```
## Pandas Data Structures
### Series
A **Series** is a single vector of data (like a NumPy array) with an *index* that labels each element in the vector.
```
counts = pd.Series([632, 1638, 569, 115])
counts
```
If an index is not specified, a default sequence of integers is assigned as the index. A NumPy array comprises the values of the `Series`, while the index is a pandas `Index` object.
```
counts.values
counts.index
```
We can assign meaningful labels to the index, if they are available:
```
bacteria = pd.Series([632, 1638, 569, 115],
index=['Firmicutes', 'Proteobacteria', 'Actinobacteria', 'Bacteroidetes'])
bacteria
```
These labels can be used to refer to the values in the `Series`.
```
bacteria['Actinobacteria']
bacteria[[name.endswith('bacteria') for name in bacteria.index]]
[name.endswith('bacteria') for name in bacteria.index]
```
Notice that the indexing operation preserved the association between the values and the corresponding indices.
We can still use positional indexing if we wish.
```
bacteria[0]
```
We can give both the array of values and the index meaningful labels themselves:
```
bacteria.name = 'counts'
bacteria.index.name = 'phylum'
bacteria
```
NumPy's math functions and other operations can be applied to Series without losing the data structure.
```
# natural logarithm
np.log(bacteria)
# log base 10
np.log10(bacteria)
```
We can also filter according to the values in the `Series`:
```
bacteria[bacteria>1000]
```
A `Series` can be thought of as an ordered key-value store. In fact, we can create one from a `dict`:
```
bacteria_dict = {'Firmicutes': 632, 'Proteobacteria': 1638, 'Actinobacteria': 569,
'Bacteroidetes': 115}
pd.Series(bacteria_dict)
```
Notice that the `Series` is created in key-sorted order.
If we pass a custom index to `Series`, it will select the corresponding values from the dict, and treat indices without corrsponding values as missing. Pandas uses the `NaN` (not a number) type for missing values.
```
bacteria2 = pd.Series(bacteria_dict,
index=['Cyanobacteria','Firmicutes',
'Proteobacteria','Actinobacteria'])
bacteria2
bacteria2.isnull()
```
Critically, the labels are used to **align data** when used in operations with other Series objects:
```
bacteria + bacteria2
```
Contrast this with NumPy arrays, where arrays of the same length will combine values element-wise; adding Series combined values with the same label in the resulting series. Notice also that the missing values were propogated by addition.
### DataFrame
Inevitably, we want to be able to store, view and manipulate data that is *multivariate*, where for every index there are multiple fields or columns of data, often of varying data type.
A `DataFrame` is a tabular data structure, encapsulating multiple series like columns in a spreadsheet. Data are stored internally as a 2-dimensional object, but the `DataFrame` allows us to represent and manipulate higher-dimensional data.
```
data = pd.DataFrame({'value':[632, 1638, 569, 115, 433, 1130, 754, 555],
'patient':[1, 1, 1, 1, 2, 2, 2, 2],
'phylum':['Firmicutes', 'Proteobacteria', 'Actinobacteria',
'Bacteroidetes', 'Firmicutes', 'Proteobacteria', 'Actinobacteria', 'Bacteroidetes']})
data
```
Notice the `DataFrame` is sorted by column name. We can change the order by indexing them in the order we desire:
```
data[['phylum','value','patient']]
```
A `DataFrame` has a second index, representing the columns:
```
data.columns
```
The `dtypes` attribute reveals the data type for each column in our DataFrame.
- `int64` is numeric integer values
- `object` strings (letters and numbers)
- `float64` floating-point values
```
data.dtypes
```
If we wish to access columns, we can do so either by dict-like indexing or by attribute:
```
data['patient']
data.patient
type(data.value)
data[['value']]
```
Notice this is different than with `Series`, where dict-like indexing retrieved a particular element (row).
If we want access to a row in a `DataFrame`, we index its `loc` attribute.
```
data.loc[3]
```
### Exercise 1
Try out these commands to see what they return:
- `data.head()`
- `data.tail(3)`
- `data.shape`
```
data.head() # returns the first (5 by default) rows of data.
data.tail(3) # returns the 3 last rows of data
data.shape # returns the dimension of data (nbr rows, nbr cols)
```
An alternative way of initializing a `DataFrame` is with a list of dicts:
```
data = pd.DataFrame([{'patient': 1, 'phylum': 'Firmicutes', 'value': 632},
{'patient': 1, 'phylum': 'Proteobacteria', 'value': 1638},
{'patient': 1, 'phylum': 'Actinobacteria', 'value': 569},
{'patient': 1, 'phylum': 'Bacteroidetes', 'value': 115},
{'patient': 2, 'phylum': 'Firmicutes', 'value': 433},
{'patient': 2, 'phylum': 'Proteobacteria', 'value': 1130},
{'patient': 2, 'phylum': 'Actinobacteria', 'value': 754},
{'patient': 2, 'phylum': 'Bacteroidetes', 'value': 555}])
data
```
Its important to note that the Series returned when a DataFrame is indexted is merely a **view** on the DataFrame, and not a copy of the data itself. So you must be cautious when manipulating this data:
```
vals = data.value
vals
vals[5] = 0
vals
```
If we plan on modifying an extracted Series, its a good idea to make a copy.
```
vals = data.value.copy()
vals[5] = 1000
vals
```
We can create or modify columns by assignment:
```
data.value[[3,4,6]] = [14, 21, 5]
data
data['year'] = 2013
data
```
But note, we cannot use the attribute indexing method to add a new column:
```
data.treatment = 1
data
data.treatment
```
### Exercise 2
From the `data` table above, create an index to return all rows for which the phylum name ends in "bacteria" and the value is greater than 1000.
----------------------------
Find the values of 'phylum' ending in 'bacteria'
```
colwitbacteria = [col for col in data['phylum'] if col.endswith('bacteria')]
colwitbacteria
```
then filter the rows having one of the 'bacteria' values
```
rowswithbacteria = data[data['phylum'].isin(colwitbacteria)]
```
then take the values bigger than 1000
```
rowswithbacteria[rowswithbacteria.value > 1000]
```
Note that it is probably faster to first filter the values bigger than 1000 as it filters more values out.
Specifying a `Series` as a new columns cause its values to be added according to the `DataFrame`'s index:
```
treatment = pd.Series([0]*4 + [1]*2)
treatment
data['treatment'] = treatment
data
```
Other Python data structures (ones without an index) need to be the same length as the `DataFrame`:
```
month = ['Jan', 'Feb', 'Mar', 'Apr']
# data['month'] = month # throws error (done on puropse)
data['month'] = ['Jan']*len(data)
data
```
We can use the `drop` method to remove rows or columns, which by default drops rows. We can be explicit by using the `axis` argument:
```
data_nomonth = data.drop('month', axis=1)
data_nomonth
```
We can extract the underlying data as a simple `ndarray` by accessing the `values` attribute:
```
data.values
```
Notice that because of the mix of string and integer (and `NaN`) values, the dtype of the array is `object`. The dtype will automatically be chosen to be as general as needed to accomodate all the columns.
```
df = pd.DataFrame({'foo': [1,2,3], 'bar':[0.4, -1.0, 4.5]})
df.values
```
Pandas uses a custom data structure to represent the indices of Series and DataFrames.
```
data.index
```
Index objects are immutable:
```
# data.index[0] = 15 # throws error
```
This is so that Index objects can be shared between data structures without fear that they will be changed.
```
bacteria2.index = bacteria.index
bacteria2
```
## Importing data
A key, but often under-appreciated, step in data analysis is importing the data that we wish to analyze. Though it is easy to load basic data structures into Python using built-in tools or those provided by packages like NumPy, it is non-trivial to import structured data well, and to easily convert this input into a robust data structure:
genes = np.loadtxt("genes.csv", delimiter=",", dtype=[('gene', '|S10'), ('value', '<f4')])
Pandas provides a convenient set of functions for importing tabular data in a number of formats directly into a `DataFrame` object. These functions include a slew of options to perform type inference, indexing, parsing, iterating and cleaning automatically as data are imported.
Let's start with some more bacteria data, stored in csv format.
```
!cat Data/microbiome.csv
```
This table can be read into a DataFrame using `read_csv`:
```
mb = pd.read_csv("Data/microbiome.csv")
mb
```
Notice that `read_csv` automatically considered the first row in the file to be a header row.
We can override default behavior by customizing some the arguments, like `header`, `names` or `index_col`.
```
pd.read_csv("Data/microbiome.csv", header=None).head()
```
`read_csv` is just a convenience function for `read_table`, since csv is such a common format:
```
mb = pd.read_table("Data/microbiome.csv", sep=',')
```
The `sep` argument can be customized as needed to accomodate arbitrary separators. For example, we can use a regular expression to define a variable amount of whitespace, which is unfortunately very common in some data formats:
sep='\s+'
For a more useful index, we can specify the first two columns, which together provide a unique index to the data.
```
mb = pd.read_csv("Data/microbiome.csv", index_col=['Patient','Taxon'])
mb.head()
```
This is called a *hierarchical* index, which we will revisit later in the section.
If we have sections of data that we do not wish to import (for example, known bad data), we can populate the `skiprows` argument:
```
pd.read_csv("Data/microbiome.csv", skiprows=[3,4,6]).head()
```
If we only want to import a small number of rows from, say, a very large data file we can use `nrows`:
```
pd.read_csv("Data/microbiome.csv", nrows=4)
```
Alternately, if we want to process our data in reasonable chunks, the `chunksize` argument will return an iterable object that can be employed in a data processing loop. For example, our microbiome data are organized by bacterial phylum, with 14 patients represented in each:
```
pd.read_csv("Data/microbiome.csv", chunksize=14)
data_chunks = pd.read_csv("Data/microbiome.csv", chunksize=14)
mean_tissue = pd.Series({chunk.Taxon[0]: chunk.Tissue.mean() for chunk in data_chunks})
mean_tissue
```
Most real-world data is incomplete, with values missing due to incomplete observation, data entry or transcription error, or other reasons. Pandas will automatically recognize and parse common missing data indicators, including `NA` and `NULL`.
```
!cat Data/microbiome_missing.csv
pd.read_csv("Data/microbiome_missing.csv").head(20)
```
Above, Pandas recognized `NA` and an empty field as missing data.
```
pd.isnull(pd.read_csv("Data/microbiome_missing.csv")).head(20)
```
Unfortunately, there will sometimes be inconsistency with the conventions for missing data. In this example, there is a question mark "?" and a large negative number where there should have been a positive integer. We can specify additional symbols with the `na_values` argument:
```
pd.read_csv("Data/microbiome_missing.csv", na_values=['?', -99999]).head(20)
```
These can be specified on a column-wise basis using an appropriate dict as the argument for `na_values`.
### Microsoft Excel
Since so much financial and scientific data ends up in Excel spreadsheets (regrettably), Pandas' ability to directly import Excel spreadsheets is valuable. This support is contingent on having one or two dependencies (depending on what version of Excel file is being imported) installed: `xlrd` and `openpyxl` (these may be installed with either `pip` or `easy_install`).
The read_excel convenience function in pandas imports a specific sheet from an Excel file
```
mb = pd.read_excel('Data/microbiome/MID2.xls', sheetname='Sheet 1', header=None)
mb.head()
```
There are several other data formats that can be imported into Python and converted into DataFrames, with the help of buitl-in or third-party libraries. These include JSON, XML, HDF5, relational and non-relational databases, and various web APIs. These are beyond the scope of this tutorial, but are covered in [Python for Data Analysis](http://shop.oreilly.com/product/0636920023784.do).
## Pandas Fundamentals
This section introduces the new user to the key functionality of Pandas that is required to use the software effectively.
For some variety, we will leave our digestive tract bacteria behind and employ some baseball data.
```
baseball = pd.read_csv("Data/baseball.csv", index_col='id')
baseball.head()
```
Notice that we specified the `id` column as the index, since it appears to be a unique identifier. We could try to create a unique index ourselves by combining `player` and `year`:
```
player_id = baseball.player + baseball.year.astype(str)
baseball_newind = baseball.copy()
baseball_newind.index = player_id
baseball_newind.head()
```
This looks okay, but let's check:
```
baseball_newind.index.is_unique
```
So, indices need not be unique. Our choice is not unique because some players change teams within years.
```
pd.Series(baseball_newind.index).value_counts()
```
The most important consequence of a non-unique index is that indexing by label will return multiple values for some labels:
```
baseball_newind.loc['wickmbo012007']
```
We will learn more about indexing below.
We can create a truly unique index by combining `player`, `team` and `year`:
```
player_unique = baseball.player + baseball.team + baseball.year.astype(str)
baseball_newind = baseball.copy()
baseball_newind.index = player_unique
baseball_newind.head()
baseball_newind.index.is_unique
```
We can create meaningful indices more easily using a hierarchical index; for now, we will stick with the numeric `id` field as our index.
### Manipulating indices
**Reindexing** allows users to manipulate the data labels in a DataFrame. It forces a DataFrame to conform to the new index, and optionally, fill in missing data if requested.
A simple use of `reindex` is to alter the order of the rows:
```
baseball.reindex(baseball.index[::-1]).head()
```
Notice that the `id` index is not sequential. Say we wanted to populate the table with every `id` value. We could specify and index that is a sequence from the first to the last `id` numbers in the database, and Pandas would fill in the missing data with `NaN` values:
```
id_range = range(baseball.index.values.min(), baseball.index.values.max())
baseball.reindex(id_range).head()
```
Missing values can be filled as desired, either with selected values, or by rule:
```
baseball.reindex(id_range, method='ffill', columns=['player','year']).head()
baseball.reindex(id_range, fill_value='charliebrown', columns=['player']).head()
```
Keep in mind that `reindex` does not work if we pass a non-unique index series.
We can remove rows or columns via the `drop` method:
```
baseball.shape
baseball.drop([89525, 89526])
baseball.drop(['ibb','hbp'], axis=1)
```
## Indexing and Selection
Indexing works analogously to indexing in NumPy arrays, except we can use the labels in the `Index` object to extract values in addition to arrays of integers.
```
# Sample Series object
hits = baseball_newind.h
hits
# Numpy-style indexing
hits[:3]
# Indexing by label
hits[['womacto01CHN2006','schilcu01BOS2006']]
```
We can also slice with data labels, since they have an intrinsic order within the Index:
```
hits['womacto01CHN2006':'gonzalu01ARI2006']
hits['womacto01CHN2006':'gonzalu01ARI2006'] = 5
hits
```
In a `DataFrame` we can slice along either or both axes:
```
baseball_newind[['h','ab']]
baseball_newind[baseball_newind.ab>500]
```
For a more concise (and readable) syntax, we can use the new `query` method to perform selection on a `DataFrame`. Instead of having to type the fully-specified column, we can simply pass a string that describes what to select. The query above is then simply:
```
baseball_newind.query('ab > 500')
```
The `DataFrame.index` and `DataFrame.columns` are placed in the query namespace by default. If you want to refer to a variable in the current namespace, you can prefix the variable with `@`:
```
min_ab = 450
baseball_newind.query('ab > @min_ab')
```
The indexing field `loc` allows us to select subsets of rows and columns in an intuitive way:
```
baseball_newind.loc['gonzalu01ARI2006', ['h','X2b', 'X3b', 'hr']]
baseball_newind.loc[:'myersmi01NYA2006', 'hr']
```
In addition to using `loc` to select rows and columns by **label**, pandas also allows indexing by **position** using the `iloc` attribute.
So, we can query rows and columns by absolute position, rather than by name:
```
baseball_newind.iloc[:5, 5:8]
```
### Exercise 3
You can use the `isin` method query a DataFrame based upon a list of values as follows:
data['phylum'].isin(['Firmacutes', 'Bacteroidetes'])
Use `isin` to find all players that played for the Los Angeles Dodgers (LAN) or the San Francisco Giants (SFN). How many records contain these values?
```
baseball[baseball['team'].isin(['LAN', 'SFN'])]
```
15 records contains those values
## Operations
`DataFrame` and `Series` objects allow for several operations to take place either on a single object, or between two or more objects.
For example, we can perform arithmetic on the elements of two objects, such as combining baseball statistics across years. First, let's (artificially) construct two Series, consisting of home runs hit in years 2006 and 2007, respectively:
```
hr2006 = baseball.loc[baseball.year==2006, 'hr']
hr2006.index = baseball.player[baseball.year==2006]
hr2007 = baseball.loc[baseball.year==2007, 'hr']
hr2007.index = baseball.player[baseball.year==2007]
hr2007
```
Now, let's add them together, in hopes of getting 2-year home run totals:
```
hr_total = hr2006 + hr2007
hr_total
```
Pandas' data alignment places `NaN` values for labels that do not overlap in the two Series. In fact, there are only 6 players that occur in both years.
```
hr_total[hr_total.notnull()]
```
While we do want the operation to honor the data labels in this way, we probably do not want the missing values to be filled with `NaN`. We can use the `add` method to calculate player home run totals by using the `fill_value` argument to insert a zero for home runs where labels do not overlap:
```
hr2007.add(hr2006, fill_value=0)
```
Operations can also be **broadcast** between rows or columns.
For example, if we subtract the maximum number of home runs hit from the `hr` column, we get how many fewer than the maximum were hit by each player:
```
baseball.hr - baseball.hr.max()
```
Or, looking at things row-wise, we can see how a particular player compares with the rest of the group with respect to important statistics
```
baseball.loc[89521, "player"]
stats = baseball[['h','X2b', 'X3b', 'hr']]
diff = stats - stats.loc[88641]
diff[:10]
```
We can also apply functions to each column or row of a `DataFrame`
```
stats.apply(np.median)
def range_calc(x):
return x.max() - x.min()
stat_range = lambda x: x.max() - x.min()
stats.apply(stat_range)
```
Lets use apply to calculate a meaningful baseball statistics, [slugging percentage](https://en.wikipedia.org/wiki/Slugging_percentage):
$$SLG = \frac{1B + (2 \times 2B) + (3 \times 3B) + (4 \times HR)}{AB}$$
And just for fun, we will format the resulting estimate.
```
def slugging(x):
bases = x['h']-x['X2b']-x['X3b']-x['hr'] + 2*x['X2b'] + 3*x['X3b'] + 4*x['hr']
ab = x['ab']+1e-6
return bases/ab
baseball.apply(slugging, axis=1).round(3)
```
## Sorting and Ranking
Pandas objects include methods for re-ordering data.
```
baseball_newind.sort_index().head()
baseball_newind.sort_index(ascending=False).head()
```
Try sorting the **columns** instead of the rows, in ascending order:
```
baseball_newind.sort_index(axis=1).head()
```
We can also use `sort_values` to sort a `Series` by value, rather than by label.
```
baseball.hr.sort_values(ascending=False)
```
For a `DataFrame`, we can sort according to the values of one or more columns using the `by` argument of `sort_values`:
```
baseball[['player','sb','cs']].sort_values(ascending=[False,True],
by=['sb', 'cs']).head(10)
```
**Ranking** does not re-arrange data, but instead returns an index that ranks each value relative to others in the Series.
```
baseball.hr.rank()
```
Ties are assigned the mean value of the tied ranks, which may result in decimal values.
```
pd.Series([100,100]).rank()
```
Alternatively, you can break ties via one of several methods, such as by the order in which they occur in the dataset:
```
baseball.hr.rank(method='first')
```
Calling the `DataFrame`'s `rank` method results in the ranks of all columns:
```
baseball.rank(ascending=False).head()
baseball[['r','h','hr']].rank(ascending=False).head()
```
### Exercise 4
Calculate **on base percentage** for each player, and return the ordered series of estimates.
$$OBP = \frac{H + BB + HBP}{AB + BB + HBP + SF}$$
define the function and appy it.
```
def on_base_perc(pl):
nominator = pl['h'] + pl['bb'] + pl['hbp'] #H+BB+HBP
denom = pl['ab'] + pl['bb'] +pl['hbp'] +pl['sf']
if(denom == 0) : # If the denom == 0 we can not compute nominator/denom, hence we retrun NaN
return np.NaN
return nominator / denom
baseball.apply(on_base_perc, axis=1).round(3)
```
and again but ordered
```
baseball.apply(on_base_perc, axis=1).round(3).sort_values(ascending=False)
```
## Hierarchical indexing
In the baseball example, I was forced to combine 3 fields to obtain a unique index that was not simply an integer value. A more elegant way to have done this would be to create a hierarchical index from the three fields.
```
baseball_h = baseball.set_index(['year', 'team', 'player'])
baseball_h.head(10)
```
This index is a `MultiIndex` object that consists of a sequence of tuples, the elements of which is some combination of the three columns used to create the index. Where there are multiple repeated values, Pandas does not print the repeats, making it easy to identify groups of values.
```
baseball_h.index[:10]
baseball_h.index.is_unique
```
Try using this hierarchical index to retrieve Julio Franco (`francju01`), who played for the Atlanta Braves (`ATL`) in 2007:
```
baseball_h.loc[(2007, 'ATL', 'francju01')]
```
Recall earlier we imported some microbiome data using two index columns. This created a 2-level hierarchical index:
```
mb = pd.read_csv("Data/microbiome.csv", index_col=['Taxon','Patient'])
mb.head(10)
```
With a hierachical index, we can select subsets of the data based on a *partial* index:
```
mb.loc['Proteobacteria']
```
Hierarchical indices can be created on either or both axes. Here is a trivial example:
```
frame = pd.DataFrame(np.arange(12).reshape(( 4, 3)),
index =[['a', 'a', 'b', 'b'], [1, 2, 1, 2]],
columns =[['Ohio', 'Ohio', 'Colorado'], ['Green', 'Red', 'Green']])
frame
```
If you want to get fancy, both the row and column indices themselves can be given names:
```
frame.index.names = ['key1', 'key2']
frame.columns.names = ['state', 'color']
frame
```
With this, we can do all sorts of custom indexing:
```
frame.loc['a', 'Ohio']
```
Try retrieving the value corresponding to `b2` in `Colorado`:
---------------------------
fetch b2 and then Colorado
```
frame.loc['b', 2]['Colorado']
```
Additionally, the order of the set of indices in a hierarchical `MultiIndex` can be changed by swapping them pairwise:
```
mb.swaplevel('Patient', 'Taxon').head()
```
Data can also be sorted by any index level, using `sortlevel`:
```
mb.sortlevel('Patient', ascending=False).head()
```
## Missing data
The occurence of missing data is so prevalent that it pays to use tools like Pandas, which seamlessly integrates missing data handling so that it can be dealt with easily, and in the manner required by the analysis at hand.
Missing data are represented in `Series` and `DataFrame` objects by the `NaN` floating point value. However, `None` is also treated as missing, since it is commonly used as such in other contexts (*e.g.* NumPy).
```
foo = pd.Series([np.nan, -3, None, 'foobar'])
foo
foo.isnull()
```
Missing values may be dropped or indexed out:
```
bacteria2
bacteria2.dropna()
bacteria2.isnull()
bacteria2[bacteria2.notnull()]
```
By default, `dropna` drops entire rows in which one or more values are missing.
```
data.dropna()
```
This can be overridden by passing the `how='all'` argument, which only drops a row when every field is a missing value.
```
data.dropna(how='all')
```
This can be customized further by specifying how many values need to be present before a row is dropped via the `thresh` argument.
```
data.loc[7, 'year'] = np.nan
data
data.dropna(thresh=5)
```
This is typically used in time series applications, where there are repeated measurements that are incomplete for some subjects.
### Exercise 5
Try using the `axis` argument to drop columns with missing values:
```
data.dropna(axis=1)
```
Rather than omitting missing data from an analysis, in some cases it may be suitable to fill the missing value in, either with a default value (such as zero) or a value that is either imputed or carried forward/backward from similar data points. We can do this programmatically in Pandas with the `fillna` argument.
```
bacteria2.fillna(0)
data.fillna({'year': 2013, 'treatment':2})
```
Notice that `fillna` by default returns a new object with the desired filling behavior, rather than changing the `Series` or `DataFrame` in place (**in general, we like to do this, by the way!**).
We can alter values in-place using `inplace=True`.
```
data.year.fillna(2013, inplace=True)
data
```
Missing values can also be interpolated, using any one of a variety of methods:
```
bacteria2.fillna(method='bfill')
```
## Data summarization
We often wish to summarize data in `Series` or `DataFrame` objects, so that they can more easily be understood or compared with similar data. The NumPy package contains several functions that are useful here, but several summarization or reduction methods are built into Pandas data structures.
```
baseball.sum()
```
Clearly, `sum` is more meaningful for some columns than others. For methods like `mean` for which application to string variables is not just meaningless, but impossible, these columns are automatically exculded:
```
baseball.mean()
```
The important difference between NumPy's functions and Pandas' methods is that the latter have built-in support for handling missing data.
```
bacteria2
bacteria2.mean()
```
Sometimes we may not want to ignore missing values, and allow the `nan` to propagate.
```
bacteria2.mean(skipna=False)
```
Passing `axis=1` will summarize over rows instead of columns, which only makes sense in certain situations.
```
extra_bases = baseball[['X2b','X3b','hr']].sum(axis=1)
extra_bases.sort_values(ascending=False)
```
A useful summarization that gives a quick snapshot of multiple statistics for a `Series` or `DataFrame` is `describe`:
```
baseball.describe()
```
`describe` can detect non-numeric data and sometimes yield useful information about it.
```
baseball.player.describe()
```
We can also calculate summary statistics *across* multiple columns, for example, correlation and covariance.
$$cov(x,y) = \sum_i (x_i - \bar{x})(y_i - \bar{y})$$
```
baseball.hr.cov(baseball.X2b)
```
$$corr(x,y) = \frac{cov(x,y)}{(n-1)s_x s_y} = \frac{\sum_i (x_i - \bar{x})(y_i - \bar{y})}{\sqrt{\sum_i (x_i - \bar{x})^2 \sum_i (y_i - \bar{y})^2}}$$
```
baseball.hr.corr(baseball.X2b)
baseball.ab.corr(baseball.h)
```
Try running `corr` on the entire `baseball` DataFrame to see what is returned:
----------------------------
```
baseball.corr()
```
it returns the correlation matrix for all features
----------------------------
If we have a `DataFrame` with a hierarchical index (or indices), summary statistics can be applied with respect to any of the index levels:
```
mb.head()
mb.sum(level='Taxon')
```
## Writing Data to Files
As well as being able to read several data input formats, Pandas can also export data to a variety of storage formats. We will bring your attention to just a couple of these.
```
mb.to_csv("mb.csv")
```
The `to_csv` method writes a `DataFrame` to a comma-separated values (csv) file. You can specify custom delimiters (via `sep` argument), how missing values are written (via `na_rep` argument), whether the index is writen (via `index` argument), whether the header is included (via `header` argument), among other options.
An efficient way of storing data to disk is in binary format. Pandas supports this using Python’s built-in pickle serialization.
```
baseball.to_pickle("baseball_pickle")
```
The complement to `to_pickle` is the `read_pickle` function, which restores the pickle to a `DataFrame` or `Series`:
```
pd.read_pickle("baseball_pickle")
```
As Wes warns in his book, it is recommended that binary storage of data via pickle only be used as a temporary storage format, in situations where speed is relevant. This is because there is no guarantee that the pickle format will not change with future versions of Python.
### Advanced Exercise: Compiling Ebola Data
The `Data/ebola` folder contains summarized reports of Ebola cases from three countries during the recent outbreak of the disease in West Africa. For each country, there are daily reports that contain various information about the outbreak in several cities in each country.
From these data files, use pandas to import them and create a single data frame that includes the daily totals of new cases and deaths for each country.
### Our solution is in a seperate notebook
| github_jupyter |
# Scalar and vector
> Marcos Duarte, Renato Naville Watanabe
> [Laboratory of Biomechanics and Motor Control](http://pesquisa.ufabc.edu.br/bmclab)
> Federal University of ABC, Brazil
<h1>Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Python-setup" data-toc-modified-id="Python-setup-1"><span class="toc-item-num">1 </span>Python setup</a></span></li><li><span><a href="#Scalar" data-toc-modified-id="Scalar-2"><span class="toc-item-num">2 </span>Scalar</a></span><ul class="toc-item"><li><span><a href="#Scalar-operations-in-Python" data-toc-modified-id="Scalar-operations-in-Python-2.1"><span class="toc-item-num">2.1 </span>Scalar operations in Python</a></span></li></ul></li><li><span><a href="#Vector" data-toc-modified-id="Vector-3"><span class="toc-item-num">3 </span>Vector</a></span><ul class="toc-item"><li><span><a href="#Magnitude-(length-or-norm)-of-a-vector" data-toc-modified-id="Magnitude-(length-or-norm)-of-a-vector-3.1"><span class="toc-item-num">3.1 </span>Magnitude (length or norm) of a vector</a></span></li><li><span><a href="#Vecton-addition-and-subtraction" data-toc-modified-id="Vecton-addition-and-subtraction-3.2"><span class="toc-item-num">3.2 </span>Vecton addition and subtraction</a></span></li></ul></li><li><span><a href="#Dot-product" data-toc-modified-id="Dot-product-4"><span class="toc-item-num">4 </span>Dot product</a></span></li><li><span><a href="#Vector-product" data-toc-modified-id="Vector-product-5"><span class="toc-item-num">5 </span>Vector product</a></span><ul class="toc-item"><li><span><a href="#Gram–Schmidt-process" data-toc-modified-id="Gram–Schmidt-process-5.1"><span class="toc-item-num">5.1 </span>Gram–Schmidt process</a></span></li></ul></li><li><span><a href="#Further-reading" data-toc-modified-id="Further-reading-6"><span class="toc-item-num">6 </span>Further reading</a></span></li><li><span><a href="#Video-lectures-on-the-Internet" data-toc-modified-id="Video-lectures-on-the-Internet-7"><span class="toc-item-num">7 </span>Video lectures on the Internet</a></span></li><li><span><a href="#Problems" data-toc-modified-id="Problems-8"><span class="toc-item-num">8 </span>Problems</a></span></li><li><span><a href="#References" data-toc-modified-id="References-9"><span class="toc-item-num">9 </span>References</a></span></li></ul></div>
Python handles very well all mathematical operations with numeric scalars and vectors and you can use [Sympy](http://sympy.org) for similar stuff but with abstract symbols. Let's briefly review scalars and vectors and show how to use Python for numerical calculation.
For a review about scalars and vectors, see chapter 2 of [Ruina and Rudra's book](http://ruina.tam.cornell.edu/Book/index.html).
## Python setup
```
from IPython.display import IFrame
import math
import numpy as np
```
## Scalar
>A **scalar** is a one-dimensional physical quantity, which can be described by a single real number.
For example, time, mass, and energy are examples of scalars.
### Scalar operations in Python
Simple arithmetic operations with scalars are indeed simple:
```
a = 2
b = 3
print('a =', a, ', b =', b)
print('a + b =', a + b)
print('a - b =', a - b)
print('a * b =', a * b)
print('a / b =', a / b)
print('a ** b =', a ** b)
print('sqrt(b) =', math.sqrt(b))
```
If you have a set of numbers, or an array, it is probably better to use Numpy; it will be faster for large data sets, and combined with Scipy, has many more mathematical funcions.
```
a = 2
b = [3, 4, 5, 6, 7, 8]
b = np.array(b)
print('a =', a, ', b =', b)
print('a + b =', a + b)
print('a - b =', a - b)
print('a * b =', a * b)
print('a / b =', a / b)
print('a ** b =', a ** b)
print('np.sqrt(b) =', np.sqrt(b)) # use numpy functions for numpy arrays
```
Numpy performs the arithmetic operations of the single number in `a` with all the numbers of the array `b`. This is called broadcasting in computer science.
Even if you have two arrays (but they must have the same size), Numpy handles for you:
```
a = np.array([1, 2, 3])
b = np.array([4, 5, 6])
print('a =', a, ', b =', b)
print('a + b =', a + b)
print('a - b =', a - b)
print('a * b =', a * b)
print('a / b =', a / b)
print('a ** b =', a ** b)
```
## Vector
>A **vector** is a quantity with magnitude (or length) and direction expressed numerically as an ordered list of values according to a coordinate reference system.
For example, position, force, and torque are physical quantities defined by vectors.
For instance, consider the position of a point in space represented by a vector:
<br>
<figure><img src="./../images/vector3D.png" width=300/><figcaption><center><i>Figure. Position of a point represented by a vector in a Cartesian coordinate system.</i></center></figcaption></figure>
The position of the point (the vector) above can be represented as a tuple of values:
$$ (x,\: y,\: z) \; \Rightarrow \; (1, 3, 2) $$
or in matrix form:
$$ \begin{bmatrix} x \\y \\z \end{bmatrix} \;\; \Rightarrow \;\; \begin{bmatrix} 1 \\3 \\2 \end{bmatrix}$$
We can use the Numpy array to represent the components of vectors.
For instance, for the vector above is expressed in Python as:
```
a = np.array([1, 3, 2])
print('a =', a)
```
Exactly like the arrays in the last example for scalars, so all operations we performed will result in the same values, of course.
However, as we are now dealing with vectors, now some of the operations don't make sense. For example, for vectors there are no multiplication, division, power, and square root in the way we calculated.
A vector can also be represented as:
<span class="notranslate">
$$ \overrightarrow{\mathbf{a}} = a_x\hat{\mathbf{i}} + a_y\hat{\mathbf{j}} + a_z\hat{\mathbf{k}} $$
</span>
<br>
<figure><img src="./../images/vector3Dijk.png" width=300/><figcaption><center><i>Figure. A vector representation in a Cartesian coordinate system. The versors <span class="notranslate"> $\hat{\mathbf{i}},\, \hat{\mathbf{j}},\, \hat{\mathbf{k}}\,$ </span> are usually represented in the color sequence <b>rgb</b> (red, green, blue) for easier visualization.</i></center></figcaption></figure>
Where <span class="notranslate"> $\hat{\mathbf{i}},\, \hat{\mathbf{j}},\, \hat{\mathbf{k}}\,$ </span> are unit vectors, each representing a direction and <span class="notranslate"> $ a_x\hat{\mathbf{i}},\: a_y\hat{\mathbf{j}},\: a_z\hat{\mathbf{k}} $ </span> are the vector components of the vector $\overrightarrow{\mathbf{a}}$.
A unit vector (or versor) is a vector whose length (or norm) is 1.
The unit vector of a non-zero vector $\overrightarrow{\mathbf{a}}$ is the unit vector codirectional with $\overrightarrow{\mathbf{a}}$:
<span class="notranslate">
$$ \mathbf{\hat{u}} = \frac{\overrightarrow{\mathbf{a}}}{||\overrightarrow{\mathbf{a}}||} = \frac{a_x\,\hat{\mathbf{i}} + a_y\,\hat{\mathbf{j}} + a_z\, \hat{\mathbf{k}}}{\sqrt{a_x^2+a_y^2+a_z^2}} $$
</span>
### Magnitude (length or norm) of a vector
The magnitude (length) of a vector is often represented by the symbol $||\;||$, also known as the norm (or Euclidean norm) of a vector and it is defined as:
<span class="notranslate">
$$ ||\overrightarrow{\mathbf{a}}|| = \sqrt{a_x^2+a_y^2+a_z^2} $$
</span>
The function `numpy.linalg.norm` calculates the norm:
```
a = np.array([1, 2, 3])
np.linalg.norm(a)
```
Or we can use the definition and compute directly:
```
np.sqrt(np.sum(a*a))
```
Then, the versor for the vector <span class="notranslate"> $ \overrightarrow{\mathbf{a}} = (1, 2, 3) $ </span> is:
```
a = np.array([1, 2, 3])
u = a/np.linalg.norm(a)
print('u =', u)
```
And we can verify its magnitude is indeed 1:
```
np.linalg.norm(u)
```
But the representation of a vector as a tuple of values is only valid for a vector with its origin coinciding with the origin $ (0, 0, 0) $ of the coordinate system we adopted.
For instance, consider the following vector:
<br>
<figure><img src="./../images/vector2.png" width=260/><figcaption><center><i>Figure. A vector in space.</i></center></figcaption></figure>
Such a vector cannot be represented by $ (b_x, b_y, b_z) $ because this would be for the vector from the origin to the point B. To represent exactly this vector we need the two vectors <span class="notranslate"> $ \mathbf{a} $ </span> and <span class="notranslate"> $ \mathbf{b} $ </span>. This fact is important when we perform some calculations in Mechanics.
### Vecton addition and subtraction
The addition of two vectors is another vector:
<span class="notranslate">
$$ \overrightarrow{\mathbf{a}} + \overrightarrow{\mathbf{b}} = (a_x\hat{\mathbf{i}} + a_y\hat{\mathbf{j}} + a_z\hat{\mathbf{k}}) + (b_x\hat{\mathbf{i}} + b_y\hat{\mathbf{j}} + b_z\hat{\mathbf{k}}) =
(a_x+b_x)\hat{\mathbf{i}} + (a_y+b_y)\hat{\mathbf{j}} + (a_z+b_z)\hat{\mathbf{k}} $$
</span>
<figure><img src="http://upload.wikimedia.org/wikipedia/commons/2/28/Vector_addition.svg" width=300 alt="Vector addition"/><figcaption><center><i>Figure. Vector addition (image from Wikipedia).</i></center></figcaption></figure>
The subtraction of two vectors is also another vector:
<span class="notranslate">
$$ \overrightarrow{\mathbf{a}} - \overrightarrow{\mathbf{b}} = (a_x\hat{\mathbf{i}} + a_y\hat{\mathbf{j}} + a_z\hat{\mathbf{k}}) + (b_x\hat{\mathbf{i}} + b_y\hat{\mathbf{j}} + b_z\hat{\mathbf{k}}) =
(a_x-b_x)\hat{\mathbf{i}} + (a_y-b_y)\hat{\mathbf{j}} + (a_z-b_z)\hat{\mathbf{k}} $$
</span>
<figure><img src="http://upload.wikimedia.org/wikipedia/commons/2/24/Vector_subtraction.svg" width=160 alt="Vector subtraction"/><figcaption><center><i>Figure. Vector subtraction (image from Wikipedia).</i></center></figcaption></figure></div>
Consider two 2D arrays (rows and columns) representing the position of two objects moving in space. The columns represent the vector components and the rows the values of the position vector in different instants.
Once again, it's easy to perform addition and subtraction with these vectors:
```
a = np.array([[1, 2, 3], [1, 1, 1]])
b = np.array([[4, 5, 6], [7, 8, 9]])
print('a =', a, '\nb =', b)
print('a + b =', a + b)
print('a - b =', a - b)
```
Numpy can handle a N-dimensional array with the size limited by the available memory in your computer.
And we can perform operations on each vector, for example, calculate the norm of each one.
First let's check the shape of the variable `a` using the method `shape` or the function `numpy.shape`:
```
print(a.shape)
print(np.shape(a))
```
This means the variable `a` has 2 rows and 3 columns.
We have to tell the function `numpy.norm` to calculate the norm for each vector, i.e., to operate through the columns of the variable `a` using the paraneter `axis`:
```
np.linalg.norm(a, axis=1)
```
## Dot product
Dot product (or scalar product or inner product) between two vectors is a mathematical operation algebraically defined as the sum of the products of the corresponding components (maginitudes in each direction) of the two vectors. The result of the dot product is a single number (a scalar).
The dot product between vectors <span class="notranslate">$\overrightarrow{\mathbf{a}}$</span> and $\overrightarrow{\mathbf{b}}$ is:
<span class="notranslate">
$$ \overrightarrow{\mathbf{a}} \cdot \overrightarrow{\mathbf{b}} = (a_x\,\hat{\mathbf{i}}+a_y\,\hat{\mathbf{j}}+a_z\,\hat{\mathbf{k}}) \cdot (b_x\,\hat{\mathbf{i}}+b_y\,\hat{\mathbf{j}}+b_z\,\hat{\mathbf{k}}) = a_x b_x + a_y b_y + a_z b_z $$
</span>
Because by definition:
<span class="notranslate">
$$ \hat{\mathbf{i}} \cdot \hat{\mathbf{i}} = \hat{\mathbf{j}} \cdot \hat{\mathbf{j}} = \hat{\mathbf{k}} \cdot \hat{\mathbf{k}}= 1 \quad \text{and} \quad \hat{\mathbf{i}} \cdot \hat{\mathbf{j}} = \hat{\mathbf{i}} \cdot \hat{\mathbf{k}} = \hat{\mathbf{j}} \cdot \hat{\mathbf{k}} = 0 $$
</span>
The geometric equivalent of the dot product is the product of the magnitudes of the two vectors and the cosine of the angle between them:
<span class="notranslate">
$$ \overrightarrow{\mathbf{a}} \cdot \overrightarrow{\mathbf{b}} = ||\overrightarrow{\mathbf{a}}||\:||\overrightarrow{\mathbf{b}}||\:cos(\theta) $$
</span>
Which is also equivalent to state that the dot product between two vectors $\overrightarrow{\mathbf{a}}$ and $\overrightarrow{\mathbf{b}}$ is the magnitude of $\overrightarrow{\mathbf{a}}$ times the magnitude of the component of $\overrightarrow{\mathbf{b}}$ parallel to $\overrightarrow{\mathbf{a}}$ (or the magnitude of $\overrightarrow{\mathbf{b}}$ times the magnitude of the component of $\overrightarrow{\mathbf{a}}$ parallel to $\overrightarrow{\mathbf{b}}$).
The dot product between two vectors can be visualized in this interactive animation:
```
IFrame('https://www.geogebra.org/classic/ncdf2jsw?embed',
width='100%', height=500)
```
The Numpy function for the dot product is `numpy.dot`:
```
a = np.array([1, 2, 3])
b = np.array([4, 5, 6])
print('a =', a, '\nb =', b)
print('np.dot(a, b) =', np.dot(a, b))
```
Or we can use the definition and compute directly:
```
np.sum(a*b)
```
For 2D arrays, the `numpy.dot` function performs matrix multiplication rather than the dot product; so let's use the `numpy.sum` function:
```
a = np.array([[1, 2, 3], [1, 1, 1]])
b = np.array([[4, 5, 6], [7, 8, 9]])
np.sum(a*b, axis=1)
```
## Vector product
Cross product or vector product between two vectors is a mathematical operation in three-dimensional space which results in a vector perpendicular to both of the vectors being multiplied and a length (norm) equal to the product of the perpendicular components of the vectors being multiplied (which is equal to the area of the parallelogram that the vectors span).
The cross product between vectors $\overrightarrow{\mathbf{a}}$ and $\overrightarrow{\mathbf{b}}$ is:
<span class="notranslate">
$$ \overrightarrow{\mathbf{a}} \times \overrightarrow{\mathbf{b}} = (a_x\,\hat{\mathbf{i}} + a_y\,\hat{\mathbf{j}} + a_z\,\hat{\mathbf{k}}) \times (b_x\,\hat{\mathbf{i}}+b_y\,\hat{\mathbf{j}}+b_z\,\hat{\mathbf{k}}) = (a_yb_z-a_zb_y)\hat{\mathbf{i}} + (a_zb_x-a_xb_z)\hat{\mathbf{j}}+(a_xb_y-a_yb_x)\hat{\mathbf{k}} $$
</span>
Because by definition:
<span class="notranslate">
$$ \begin{array}{l l}
\hat{\mathbf{i}} \times \hat{\mathbf{i}} = \hat{\mathbf{j}} \times \hat{\mathbf{j}} = \hat{\mathbf{k}} \times \hat{\mathbf{k}} = 0 \\
\hat{\mathbf{i}} \times \hat{\mathbf{j}} = \hat{\mathbf{k}}, \quad \hat{\mathbf{k}} \times \hat{\mathbf{k}} = \hat{\mathbf{i}}, \quad \hat{\mathbf{k}} \times \hat{\mathbf{i}} = \hat{\mathbf{j}} \\
\hat{\mathbf{j}} \times \hat{\mathbf{i}} = -\hat{\mathbf{k}}, \quad \hat{\mathbf{k}} \times \hat{\mathbf{j}}= -\hat{\mathbf{i}}, \quad \hat{\mathbf{i}} \times \hat{\mathbf{k}} = -\hat{\mathbf{j}}
\end{array} $$
</span>
The direction of the vector resulting from the cross product between the vectors $\overrightarrow{\mathbf{a}}$ and $\overrightarrow{\mathbf{b}}$ is given by the right-hand rule.
The geometric equivalent of the cross product is:
The geometric equivalent of the cross product is the product of the magnitudes of the two vectors and the sine of the angle between them:
<span class="notranslate">
$$ \overrightarrow{\mathbf{a}} \times \overrightarrow{\mathbf{b}} = ||\overrightarrow{\mathbf{a}}||\:||\overrightarrow{\mathbf{b}}||\:sin(\theta) $$
</span>
Which is also equivalent to state that the cross product between two vectors $\overrightarrow{\mathbf{a}}$ and $\overrightarrow{\mathbf{b}}$ is the magnitude of $\overrightarrow{\mathbf{a}}$ times the magnitude of the component of $\overrightarrow{\mathbf{b}}$ perpendicular to $\overrightarrow{\mathbf{a}}$ (or the magnitude of $\overrightarrow{\mathbf{b}}$ times the magnitude of the component of $\overrightarrow{\mathbf{a}}$ perpendicular to $\overrightarrow{\mathbf{b}}$).
The definition above, also implies that the magnitude of the cross product is the area of the parallelogram spanned by the two vectors:
<br>
<figure><img src="http://upload.wikimedia.org/wikipedia/commons/4/4e/Cross_product_parallelogram.svg" width=160 alt="Vector subtraction"/><figcaption><center><i>Figure. Area of a parallelogram as the magnitude of the cross product (image from Wikipedia).</i></center></figcaption></figure>
The cross product can also be calculated as the determinant of a matrix:
<span class="notranslate">
$$ \overrightarrow{\mathbf{a}} \times \overrightarrow{\mathbf{b}} = \left| \begin{array}{ccc}
\hat{\mathbf{i}} & \hat{\mathbf{j}} & \hat{\mathbf{k}} \\
a_x & a_y & a_z \\
b_x & b_y & b_z
\end{array} \right|
= a_y b_z \hat{\mathbf{i}} + a_z b_x \hat{\mathbf{j}} + a_x b_y \hat{\mathbf{k}} - a_y b_x \hat{\mathbf{k}}-a_z b_y \hat{\mathbf{i}} - a_x b_z \hat{\mathbf{j}} \\
\overrightarrow{\mathbf{a}} \times \overrightarrow{\mathbf{b}} = (a_yb_z-a_zb_y)\hat{\mathbf{i}} + (a_zb_x-a_xb_z)\hat{\mathbf{j}} + (a_xb_y-a_yb_x)\hat{\mathbf{k}} $$
</span>
The same result as before.
The cross product between two vectors can be visualized in this interactive animation:
```
IFrame('https://www.geogebra.org/classic/cz6v2U99?embed',
width='100%', height=500)
```
The Numpy function for the cross product is `numpy.cross`:
```
print('a =', a, '\nb =', b)
print('np.cross(a, b) =', np.cross(a, b))
```
For 2D arrays with vectors in different rows:
```
a = np.array([[1, 2, 3], [1, 1, 1]])
b = np.array([[4, 5, 6], [7, 8, 9]])
np.cross(a, b, axis=1)
```
### Gram–Schmidt process
The [Gram–Schmidt process](http://en.wikipedia.org/wiki/Gram%E2%80%93Schmidt_process) is a method for orthonormalizing (orthogonal unit versors) a set of vectors using the scalar product. The Gram–Schmidt process works for any number of vectors.
For example, given three vectors, $\overrightarrow{\mathbf{a}}, \overrightarrow{\mathbf{b}}, \overrightarrow{\mathbf{c}}$, in the 3D space, a basis $\{\hat{e}_a, \hat{e}_b, \hat{e}_c\}$ can be found using the Gram–Schmidt process by:
The first versor is in the $\overrightarrow{\mathbf{a}}$ direction (or in the direction of any of the other vectors):
$$ \hat{e}_a = \frac{\overrightarrow{\mathbf{a}}}{||\overrightarrow{\mathbf{a}}||} $$
The second versor, orthogonal to $\hat{e}_a$, can be found considering we can express vector $\overrightarrow{\mathbf{b}}$ in terms of the $\hat{e}_a$ direction as:
$$ \overrightarrow{\mathbf{b}} = \overrightarrow{\mathbf{b}}^\| + \overrightarrow{\mathbf{b}}^\bot $$
Then:
$$ \overrightarrow{\mathbf{b}}^\bot = \overrightarrow{\mathbf{b}} - \overrightarrow{\mathbf{b}}^\| = \overrightarrow{\mathbf{b}} - (\overrightarrow{\mathbf{b}} \cdot \hat{e}_a ) \hat{e}_a $$
Finally:
$$ \hat{e}_b = \frac{\overrightarrow{\mathbf{b}}^\bot}{||\overrightarrow{\mathbf{b}}^\bot||} $$
The third versor, orthogonal to $\{\hat{e}_a, \hat{e}_b\}$, can be found expressing the vector $\overrightarrow{\mathbf{C}}$ in terms of $\hat{e}_a$ and $\hat{e}_b$ directions as:
$$ \overrightarrow{\mathbf{c}} = \overrightarrow{\mathbf{c}}^\| + \overrightarrow{\mathbf{c}}^\bot $$
Then:
$$ \overrightarrow{\mathbf{c}}^\bot = \overrightarrow{\mathbf{c}} - \overrightarrow{\mathbf{c}}^\| $$
Where:
$$ \overrightarrow{\mathbf{c}}^\| = (\overrightarrow{\mathbf{c}} \cdot \hat{e}_a ) \hat{e}_a + (\overrightarrow{\mathbf{c}} \cdot \hat{e}_b ) \hat{e}_b $$
Finally:
$$ \hat{e}_c = \frac{\overrightarrow{\mathbf{c}}^\bot}{||\overrightarrow{\mathbf{c}}^\bot||} $$
Let's implement the Gram–Schmidt process in Python.
For example, consider the positions (vectors) $\overrightarrow{\mathbf{a}} = [1,2,0], \overrightarrow{\mathbf{b}} = [0,1,3], \overrightarrow{\mathbf{c}} = [1,0,1]$:
```
a = np.array([1, 2, 0])
b = np.array([0, 1, 3])
c = np.array([1, 0, 1])
```
The first versor is:
```
ea = a/np.linalg.norm(a)
print(ea)
```
The second versor is:
```
eb = b - np.dot(b, ea)*ea
eb = eb/np.linalg.norm(eb)
print(eb)
```
And the third version is:
```
ec = c - np.dot(c, ea)*ea - np.dot(c, eb)*eb
ec = ec/np.linalg.norm(ec)
print(ec)
```
Let's check the orthonormality between these versors:
```
print('Versors:', '\nea =', ea, '\neb =', eb, '\nec =', ec)
print('\nTest of orthogonality (scalar product between versors):',
'\nea x eb:', np.dot(ea, eb),
'\neb x ec:', np.dot(eb, ec),
'\nec x ea:', np.dot(ec, ea))
print('\nNorm of each versor:',
'\n||ea|| =', np.linalg.norm(ea),
'\n||eb|| =', np.linalg.norm(eb),
'\n||ec|| =', np.linalg.norm(ec))
```
Or, we can simply use the built-in QR factorization function from NumPy:
```
vectors = np.vstack((a,b,c)).T
Q, R = np.linalg.qr(vectors)
print(Q)
ea, eb, ec = Q[:, 0], Q[:, 1], Q[:, 2]
print('Versors:', '\nea =', ea, '\neb =', eb, '\nec =', ec)
print('\nTest of orthogonality (scalar product between versors):')
print(np.dot(Q.T, Q))
print('\nTest of orthogonality (scalar product between versors):',
'\nea x eb:', np.dot(ea, eb),
'\neb x ec:', np.dot(eb, ec),
'\nec x ea:', np.dot(ec, ea))
print('\nNorm of each versor:',
'\n||ea|| =', np.linalg.norm(ea),
'\n||eb|| =', np.linalg.norm(eb),
'\n||ec|| =', np.linalg.norm(ec))
```
Which results in the same basis with exception of the changed signals.
## Further reading
- Read pages 44-92 of the first chapter of the [Ruina and Rudra's book](http://ruina.tam.cornell.edu/Book/index.html) about scalars and vectors in Mechanics.
## Video lectures on the Internet
- Khan Academy: [Vectors](https://www.khanacademy.org/math/algebra-home/alg-vectors)
- [Vectors, what even are they?](https://youtu.be/fNk_zzaMoSs)
## Problems
1. Given the vectors, $\overrightarrow{\mathbf{a}}=[1, 0, 0]$ and $\overrightarrow{\mathbf{b}}=[1, 1, 1]$, calculate the dot and cross products between them.
2. Calculate the unit vectors for $[2, −2, 3]$ and $[3, −3, 2]$ and determine an orthogonal vector to these two vectors.
3. Given the vectors $\overrightarrow{\mathbf{a}}$=[1, 0, 0] and $\overrightarrow{\mathbf{b}}$=[1, 1, 1], calculate $ \overrightarrow{\mathbf{a}} \times \overrightarrow{\mathbf{b}} $ and verify that this vector is orthogonal to vectors $\overrightarrow{\mathbf{a}}$ and $\overrightarrow{\mathbf{b}}$. Also, calculate $\overrightarrow{\mathbf{b}} \times \overrightarrow{\mathbf{a}}$ and compare it with $\overrightarrow{\mathbf{a}} \times \overrightarrow{\mathbf{b}}$.
4. Given the vectors $[1, 1, 0]; [1, 0, 1]; [0, 1, 1]$, calculate a basis using the Gram–Schmidt process.
5. Write a Python function to calculate a basis using the Gram–Schmidt process (implement the algorithm!) considering that the input are three variables where each one contains the coordinates of vectors as columns and different positions of these vectors as rows. For example, sample variables can be generated with the command `np.random.randn(5, 3)`.
6. Study the sample problems **1.1** to **1.9**, **1.11** (using Python), **1.12**, **1.14**, **1.17**, **1.18** to **1.24** of Ruina and Rudra's book
7. From Ruina and Rudra's book, solve the problems **1.1.1** to **1.3.16**.
If you are new to scalars and vectors, you should solve these problems first by hand and then use Python to check the answers.
## References
- Ruina A, Rudra P (2019) [Introduction to Statics and Dynamics](http://ruina.tam.cornell.edu/Book/index.html). Oxford University Press.
| github_jupyter |
## Chapter 3.5 뉴스 기사 분류: 다중 분류 문제
### Loading Reuters Data Set
```
from keras.datasets import reuters
```
### data split
```
(train_data, train_labels), (test_data, test_labels) = reuters.load_data(num_words= 10000)
```
- 원래 텍스트로 디코딩하기
```
word_index = reuters.get_word_index()
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
decoded_newwire = ' '.join([reverse_word_index.get(i-3, '?') for i in train_data[0]])
decoded_newwire
```
### data preprocessing
```
import numpy as np
def vectorize_sequences(sequences, dimension=10000):
results = np.zeros((len(sequences), dimension))
for i, sequence in enumerate(sequences):
results[i, sequence] = 1
return results
x_train = vectorize_sequences(train_data)
x_test = vectorize_sequences(test_data)
```
#### multi- label :: one hot encoidng
```
def to_one_hot(labels, dimension =46):
results = np.zeros((len(labels), dimension))
for i, label in enumerate(labels):
results[i, label] = i
return results
one_hot_train_labels = to_one_hot(train_labels)
one_hot_test_labels = to_one_hot(test_labels)
```
- 16차원의 공간으로 46개의 클래스를 구분하기엔 너무 제약이 많다. // 한 layer 에서 분류 문제에 필요한 일부 정보를 누락하면 그 다음 충에서 이를 복원할 수 없다.
- 64개의 유닛으로 시도
### Model Construction
```
from keras import models
from keras import layers
model = models.Sequential()
model.add(layers.Dense(64, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(64, activation='relu'))
model.add(layers.Dense(46, activation='softmax'))
```
#### model compile
```
from keras import losses
model.compile(optimizer='rmsprop',
loss=losses.categorical_crossentropy,
metrics=['accuracy'])
```
#### model validation
```
x_val = x_train[:1000]
partial_x_train = x_test[1000:]
y_val = one_hot_train_labels[:1000]
partial_y_train = one_hot_test_labels[1000:]
history = model.fit(partial_x_train,
partial_y_train,
epochs=20,
batch_size=512,
validation_data=(x_val, y_val))
```
### visualization
```
import matplotlib.pyplot as plt
%matplotlib inline
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(loss) + 1)
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
import matplotlib.pyplot as plt
%matplotlib inline
loss = history.history['acc']
val_loss = history.history['val_acc']
epochs = range(1, len(loss) + 1)
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.show()
```
### 과대적합을 피하기 위해 9th epoch 까지 진행
```
model = models.Sequential()
model.add(layers.Dense(64, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(64, activation='relu'))
model.add(layers.Dense(46, activation='softmax'))
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
model.fit(partial_x_train,
partial_y_train,
epochs=9,
batch_size=512,
validation_data=(x_val, y_val))
results = model.evaluate(x_test, one_hot_test_labels)
results
```
### model predict
```
prediction = model.predict(x_test)
```
### categorical_crossentropy & sparse_categorical_crossentropy
- 정수 레이블을 사용할 때에는 sparse_categorical_crossentropy 를 사용해야 한다.
```
y_train = np.array(train_labels)
y_test = np.array(test_labels)
model.compile(optimizer='rmsprop',
loss='sparse_categorical_crossentropy',
metrcis=['acc'])
```
### 46차원보다 작은 중간층을 두었을때의 문제
```
model = models.Sequential()
model.add(layers.Dense(64, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(4, activation='relu'))
model.add(layers.Dense(46, activation='softmax'))
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
model.fit(partial_x_train,
partial_y_train,
epochs=9,
batch_size=512,
validation_data=(x_val, y_val))
```
### 추가 실험
#### 더 큰 은닉 유닛 사용
```
model = models.Sequential()
model.add(layers.Dense(64, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(128, activation='relu'))
model.add(layers.Dense(46, activation='softmax'))
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
model.fit(partial_x_train,
partial_y_train,
epochs=9,
batch_size=512,
validation_data=(x_val, y_val))
```
#### 3개의 hidden layer 사용
```
model = models.Sequential()
model.add(layers.Dense(64, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(128, activation='relu'))
model.add(layers.Dense(128, activation='relu'))
model.add(layers.Dense(46, activation='softmax'))
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
model.fit(partial_x_train,
partial_y_train,
epochs=9,
batch_size=512,
validation_data=(x_val, y_val))
```
| github_jupyter |
<img src="images/logo.jpg" style="display: block; margin-left: auto; margin-right: auto;" alt="לוגו של מיזם לימוד הפייתון. נחש מצויר בצבעי צהוב וכחול, הנע בין האותיות של שם הקורס: לומדים פייתון. הסלוגן המופיע מעל לשם הקורס הוא מיזם חינמי ללימוד תכנות בעברית.">
# <span style="text-align: right; direction: rtl; float: right;">לולאת for</span>
## <span style="text-align: right; direction: rtl; float: right; clear: both;">הקדמה</span>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
פעמים רבות אנחנו מנסים לפתור בעיה, שכדי להגיע לפתרונה נידרש לעבור על כל הערכים במבנה מסוים:
</p>
<ul style="text-align: right; direction: rtl; float: right; clear: both;">
<li>קבל את רשימת גובהי התלמידים בכיתה, והחזר את גובהו של התלמיד הגבוה ביותר.</li>
<li>קבל את רשימת הקלפים שבידי, והחזר את הקלף המתאים ביותר לשליפה עכשיו.</li>
<li>קבל רשימת השמעה (Playlist), והחזר את כל השירים של הלהקה Led Zeppelin.</li>
<li>קבל את רשימת המסעדות בצרפת ואת הדירוגים שלהן, והחזר את 3 המסעדות בעלות הדירוג הגבוה ביותר.</li>
</ul>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
מה משותף לכל הבעיות שהוצגו למעלה?<br>
דרך ראויה לפתור אותן היא בעזרת לולאה שתעבור על כל האיברים שהוצגו בבעיה, ותבצע על כל איבר סדרת פעולות.<br>
נכתוב בפסאודו־קוד דוגמה לפתרון הבעיה הראשונה – מציאת הגובה של התלמיד הגבוה ביותר בכיתה:
</p>
<ol style="text-align: right; direction: rtl; float: right; clear: both;">
<li><strong>קלוט</strong> <em>רשימת גבהים</em> לתוך המשתנה <var>גבהים</var>.</li>
<li><strong>אפס</strong> את המשתנה ששמו <var>הכי_גבוה_שמצאנו</var> כך שיכיל את הערך <em>0</em>.</li>
<li>
<strong>עבור כל</strong> <var>גובה</var> שנמצא בתוך <var>גבהים</var>:
<ol>
<li style="list-style: numbers;">
<strong>אם</strong> הערך של <var>גובה</var> גדול יותר מ<var>הכי_גבוה_שמצאנו</var>:
<ol>
<li>
<strong>שים</strong> בתוך <var>הכי_גבוה_שמצאנו</var> את הערך של <var>גובה</var>.
</li>
</ol>
</li>
</ol>
</li>
<li><strong>החזר</strong> את <var>הכי_גבוה_שמצאנו</var>.</li>
</ol>
<div class="align-center" style="display: flex; text-align: right; direction: rtl; clear: both;">
<div style="display: flex; width: 10%; float: right; clear: both;">
<img src="images/exercise.svg" style="height: 50px !important;" alt="תרגול">
</div>
<div style="width: 70%">
<p style="text-align: right; direction: rtl; float: right; clear: both;">
ממשו פונקציה שמקבלת רשימת גבהים של האנשים בכיתה, ומחזירה את הגובה של התלמיד הגבוה ביותר.<br>
לדוגמה, עבור הרשימה <code dir="ltr" style="direction: ltr;">[1.50, 1.84, 1.73, 1.51]</code> החזירו <samp>1.84</samp>.
</p>
</div>
<div style="display: flex; width: 20%; border-right: 0.1rem solid #A5A5A5; padding: 1rem 2rem;">
<p style="text-align: center; direction: rtl; justify-content: center; align-items: center; clear: both;">
<strong>חשוב!</strong><br>
פתרו לפני שתמשיכו!
</p>
</div>
</div>
### <span style="text-align: right; direction: rtl; float: right; clear: both;">לולאת while</span>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
עד כה, אחד מהשימושים הנפוצים שעשינו בלולאת <code>while</code> היה לעבור על איברים ברשימה באמצעות מיקום.<br>
הקוד שכתבתם בתרגול למעלה, שבו הייתם צריכים לעבור על גובהי התלמידים, זה אחר זה, הוא דוגמה טובה לכך.<br>
ייתכן שהקוד שלכם דומה לקוד שאני כתבתי לפתרון התרגיל הזה:
</p>
```
def get_tallest(student_heights):
max_height_so_far = 0
current_student_index = 0
while current_student_index < len(student_heights):
current_student_height = student_heights[current_student_index]
if current_student_height > max_height_so_far:
max_height_so_far = current_student_height
current_student_index = current_student_index + 1
return max_height_so_far
heights = [1.50, 1.84, 1.73, 1.51]
print(get_tallest(heights))
```
<p style="text-align: right; direction: rtl; float: right; clear: both;">
הקוד בתא האחרון עושה את הפעולות הבאות:
</p>
<ul style="text-align: right; direction: rtl; float: right; clear: both;">
<li>איפוס המשתנה <var>max_height_so_far</var>, ששומר את הגובה המרבי שמצאנו עד כה ברשימה.</li>
<li>איפוס המשתנה <var>current_student_index</var>, שמצביע על מיקום התלמיד שאנחנו בודקים באיטרציה הנוכחית של הלולאה.</li>
<li>בכל איטרציה, הביטוי <code>student_heights[current_student_index]</code> ישיג את גובהו של אחד התלמידים, לפי הסדר.</li>
<li>אם התלמיד הנבדק גבוה יותר מהתלמיד הכי גבוה שמצאנו עד עכשיו, שמור את הגובה המרבי החדש בתוך <var>max_height_so_far</var>.</li>
<li>קדם את <var>current_student_index</var> כך שיצביע לתא שבו מופיע התלמיד הבא.</li>
<li>בסיום המעבר על כל הערכים, החזר את <var>max_height_so_far</var>.</li>
</ul>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
הלולאה שמופיעה תעבור על מספרי התאים ברשימה ותבדוק את התוכן שלהם.<br>
עד כה, פעמים רבות השימוש שלנו בלולאות היה לצורך <mark>מעבר על כל האיברים של iterable כלשהו</mark>.
</p>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
למעשה, בדוגמה שלמעלה <mark>אנחנו מבצעים פעולה עבור כל איבר בתוך <var>student_heights</var></mark>.
</p>
## <span style="text-align: right; direction: rtl; float: right; clear: both;">שימוש ב־for</span>
### <span style="text-align: right; direction: rtl; float: right; clear: both;">שימוש בסיסי</span>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
בפעמים שבהן אנחנו רוצים לבצע דבר מה <em>עבור</em> כל אחד מהאיברים במבנה כלשהו, נשתמש ב<dfn>לולאת <code>for</code></dfn>.<br>
נראה, לדוגמה, איך נשתמש בלולאת <code>while</code> כדי להדפיס את שמותיהן של כל התלמידות בכיתה:
</p>
```
names_of_students_in_class = ['Galia', 'Hadas', 'Hen', 'Ilana', 'Ivria', 'Karin', 'Maya', 'Noa']
student_index = 0
while student_index < len(names_of_students_in_class):
student_name = names_of_students_in_class[student_index]
print(student_name)
student_index = student_index + 1
```
<p style="text-align: right; direction: rtl; float: right; clear: both;">
בעברית, היינו משתמשים בנוסח דומה לזה: <mark>עבור כל שם של תלמידה בתוך רשימת שמות התלמידות שבכיתה, הדפס את שם התלמידה</mark>.</p>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
נכתוב בפייתון, הפעם בעזרת לולאת <code>for</code>:
</p>
```
for student_name in names_of_students_in_class:
print(student_name)
```
<p style="text-align: right; direction: rtl; float: right; clear: both;">
השוו את האלגנטיות של הקוד הזה לאלגנטיות של הקוד שמשתמש בלולאת <code>while</code>, ואת הדמיון בין כל אחת מהן לבין הנוסח המילולי שכתבנו.
</p>
### <span style="text-align: right; direction: rtl; float: right; clear: both;">איך זה עובד?</span>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
לולאת ה־<code>for</code> שראיתם מתחלקת ל־3 חלקים:
</p>
<ol style="text-align: right; direction: rtl; float: right; clear: both;">
<li>iterable כלשהו – נסתכל על כל האיברים שבו לפי הסדר שלהם.</li>
<li>שם משתנה חדש שנמציא – פייתון תיצור אותו במיוחד עבור הלולאה. המשתנה הזה יצביע בכל פעם על איבר אחד, לפי הסדר, מתוך ה־iterable.</li>
<li>הפעולה או הפעולות שאנחנו רוצים לבצע על כל אחד מהאיברים האלו.</li>
</ol>
```
names_of_students_in_class = ['Galia', 'Hadas', 'Hen', 'Ilana', 'Ivria', 'Karin', 'Maya', 'Noa']
# השם שנמציא
# V Iterable, ערך שאפשר לפרק לכלל איבריו
for student_name in names_of_students_in_class:
print(student_name) # <---- הפעולות לביצוע
```
<p style="text-align: right; direction: rtl; float: right; clear: both;">
נבחן את הלולאה שמדפיסה את שמות התלמידות בכיתה, שלב אחר שלב, כדי להבין איך לולאת <code>for</code> פועלת.
</p>
<div class="align-center" style="display: flex; text-align: right; direction: rtl;">
<div style="display: flex; width: 10%; float: right; ">
<img src="images/warning.png" style="height: 50px !important;" alt="אזהרה!">
</div>
<div style="width: 90%">
<p style="text-align: right; direction: rtl;">
כמה מקטעי הקוד הבאים לא ירוצו, כיוון שחלק מהקוד מסומן בהערה.<br>
המטרה של קטעי הקוד בחלק הזה של המחברת היא להדגיש איזה קוד רץ באותו רגע.
</p>
</div>
</div>
```
names_of_students_in_class = ['Galia', 'Hadas', 'Hen', 'Ilana', 'Ivria', 'Karin', 'Maya', 'Noa']
```
<table style="font-size: 1.5rem; border: 0px solid black; border-spacing: 0px;">
<caption style="direction: rtl; text-align: center;">תוכן המשתנה <var>names_of_students_in_class</var></caption>
<tr>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">2</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">3</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">4</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">5</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">6</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">7</td>
</tr>
<tbody>
<tr>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Galia"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Hadas"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Hen"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Ilana"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Ivria"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Karin"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Maya"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Noa"</td>
</tr>
<tr style="background: #f5f5f5;">
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-8</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-7</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-6</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-5</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-4</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-3</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-2</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
</tr>
</tbody>
</table>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
בחזרור הראשון, המשתנה החדש שיצרנו, בשם <var>student_name</var>, יצביע על הערך הראשון ברשימה, <samp>Galia</samp>.<br>
נשים לב שהמשתנה <var>student_name</var> לא היה קיים לפני הלולאה, והלולאה היא המבנה הקסום שיוצר את המשתנה וגורם לו להצביע לערכים:
</p>
```
for student_name in names_of_students_in_class: # <--- אנחנו פה
# print(student_name)
```
<table style="font-size: 1.5rem; border: 0px solid black; border-spacing: 0px;">
<caption style="direction: rtl; text-align: center;">חזרור ראשון, <var>student_name</var> מצביע על "Galia"</caption>
<tr>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">2</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">3</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">4</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">5</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">6</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">7</td>
</tr>
<tbody>
<tr>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: yellow">"Galia"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Hadas"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Hen"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Ilana"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Ivria"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Karin"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Maya"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Noa"</td>
</tr>
<tr style="background: #f5f5f5;">
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: center;"><var>student_name</var> <span style="font-size: 1.8rem;">↑</span></td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-7</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-6</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-5</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-4</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-3</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-2</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
</tr>
</tbody>
</table>
<div>
</div>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
מייד לאחר מכן יודפס התוכן שאליו <var>student_name</var> מצביע:</p>
```
#for student_name in names_of_students_in_class:
print(student_name) # <--- אנחנו פה
```
<p style="text-align: right; direction: rtl; float: right; clear: both;">
סיימנו את האיטרציה! מה עכשיו?<br>
עולים חזרה לראש הלולאה כדי לבדוק אם נשארו עוד איברים לעבור עליהם:</p>
```
for student_name in names_of_students_in_class: # <--- אנחנו פה
# print(student_name)
```
<p style="text-align: right; direction: rtl; float: right; clear: both;">
המשתנה <var>student_name</var> יעבור להצביע על האיבר הבא, <samp>Hadas</samp>:
</p>
<table style="font-size: 1.5rem; border: 0px solid black; border-spacing: 0px;">
<caption style="direction: rtl; text-align: center;">חזרור שני, <var>student_name</var> מצביע על "Hadas"</caption>
<tr>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">2</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">3</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">4</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">5</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">6</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">7</td>
</tr>
<tbody>
<tr>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Galia"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: yellow">"Hadas"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Hen"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Ilana"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Ivria"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Karin"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Maya"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Noa"</td>
</tr>
<tr style="background: #f5f5f5;">
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-8</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;"><var>student_name</var> <span style="font-size: 1.8rem;">↑</span></td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-6</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-5</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-4</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-3</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-2</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
</tr>
</tbody>
</table>
<div>
</div>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
ושוב, נדפיס את המחרוזת ש־<var>student_name</var> מצביע עליה:
</p>
```
#for student_name in names_of_students_in_class:
print(student_name) # <---- עכשיו אנחנו פה
```
<p style="text-align: right; direction: rtl; float: right; clear: both;">
כך נמשיך לבצע את הלולאה, עד שנגיע לאיבר האחרון ברשימה, התלמידה <samp>Noa</samp>:
</p>
```
for student_name in names_of_students_in_class: # אנחנו פה, אחרי שעברנו על שמות כללל התלמידות, פרט לנועה
# print(student_name)
```
<table style="font-size: 1.5rem; border: 0px solid black; border-spacing: 0px;">
<caption style="direction: rtl; text-align: center;">חזרור אחרון, <var>student_name</var> מצביע על "Noa"</caption>
<tr>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">2</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">3</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">4</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">5</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">6</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">7</td>
</tr>
<tbody>
<tr>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Galia"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Hadas"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Hen"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Ilana"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Ivria"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Karin"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Maya"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: yellow;">"Noa"</td>
</tr>
<tr style="background: #f5f5f5;">
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-8</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-7</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-6</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-5</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-4</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-3</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-2</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;"><var>student_name</var> <span style="font-size: 1.8rem;">↑</span></td>
</tr>
</tbody>
</table>
<div>
</div>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
נדפיס בפעם האחרונה את המחרוזת שעליה אנחנו מצביעים:
</p>
```
#for student_name in names_of_students_in_class:
print(student_name) # <---- עכשיו אנחנו פה
```
<p style="text-align: right; direction: rtl; float: right; clear: both;">
כשנחזור לראש הלולאה, נראה שעברנו על כל האיברים.<br>
הלולאה תיפסק, וירוץ הקוד שנמצא אחרי הלולאה.
</p>
<table style="text-align: right; direction: rtl; clear: both; font-size: 1.3rem">
<caption style="text-align: center; direction: rtl; clear: both; font-size: 2rem; padding-bottom: 2rem;">החלקים בלולאת for</caption>
<thead>
<tr>
<th>החלק בלולאה</th>
<th>איפה בקוד</th>
<th>דוגמה</th>
</tr>
</thead>
<tbody>
<tr>
<td>המבנה שאנחנו רוצים לעבור על כלל איבריו – חייב להיות iterable</td>
<td>אחרי המילה <em>in</em></td>
<td><var>names_of_students_in_class</var></td>
</tr>
<tr>
<td>שם שנמציא למשתנה – בכל חזרור יכיל איבר מתוך ה־iterable</td>
<td>אחרי המילה <em>for</em>, לפני המילה <em>in</em></td>
<td><var>student_name</var></td>
</tr>
<tr>
<td>תוכן הלולאה – הפעולות שנרצה לבצע על כל איבר</td>
<td>אחרי הנקודתיים, בשורה חדשה (אחת או יותר), בהזחה</td>
<td><code>print(student_name)</code></td>
</tr>
</tbody>
</table>
<div class="align-center" style="display: flex; text-align: right; direction: rtl; clear: both;">
<div style="display: flex; width: 10%; float: right; clear: both;">
<img src="images/exercise.svg" style="height: 50px !important;" alt="תרגול">
</div>
<div style="width: 70%">
<p style="text-align: right; direction: rtl; float: right; clear: both;">
שנו את הקוד שכתבתם למציאת האדם הגבוה ביותר בכיתה, כך שישתמש ב־<code>for</code> ולא ב־<code>while</code>.<br>
טיפ: השתמשו בפסאודו־קוד שהוצג לפני כן, והשוו בין ה־<code>for</code> לבין ה־<code>while</code> בדוגמה של הדפסת שמות התלמידים.<br>
</p>
</div>
<div style="display: flex; width: 20%; border-right: 0.1rem solid #A5A5A5; padding: 1rem 2rem;">
<p style="text-align: center; direction: rtl; justify-content: center; align-items: center; clear: both;">
<strong>חשוב!</strong><br>
פתרו לפני שתמשיכו!
</p>
</div>
</div>
### <span style="text-align: right; direction: rtl; float: right;">דוגמאות מילוליות נוספות</span>
<ul style="text-align: right; direction: rtl; float: right; clear: both;">
<li>עבור כל עמוד בספר – קרא את העמוד.</li>
<li>עבור כל צלחת – שטוף אותה במים חמים, קרצף אותה היטב בסקוץ' ספוג בסבון, נגב אותה במגבת יבשה ואחסן אותה בארון.</li>
<li>בהינתן רשימה של 1,000 תלמידים, חשב את הגובה הממוצע של תלמיד.</li>
<li>בליל כל הקדושים, התחפש, צא החוצה, ועבור כל בית ברחוב: גש לדלת, צלצל בפעמון, אמור "ממתק או תעלול", קח ממתק ואמור תודה.</li>
</ul>
### <span style="text-align: right; direction: rtl; float: right;">מתי להשתמש?</span>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
לולאת <code>for</code> יוצרת מבנה אלגנטי וקריא, ומתכנתים רבים מעדיפים אותה על פני לולאת <code>while</code>.<br>
ננסה לעמוד על ההבדלים בין הלולאות:<br>
</p>
<table style="text-align: right; direction: rtl; clear: both; font-size: 1.3rem">
<caption style="text-align: center; direction: rtl; clear: both; font-size: 2rem; padding-bottom: 2rem;">הבדלים בין while ל־for</caption>
<thead>
<tr>
<th>נתון להשוואה</th>
<th>לולאת <code>for</code></th>
<th>לולאת <code>while</code></th>
</tr>
</thead>
<tbody>
<tr>
<td>מה "מניע" את הלולאה?</td>
<td>iterable שהלולאה תעבור על כל האיברים שלו</td>
<td>ביטוי שערכו הבוליאני שקול ל־<code>True</code> או ל־<code>False</code></td>
</tr>
<tr>
<td>מתי הלולאה מפסיקה</td>
<td>כשהלולאה עברה על כל האיברים של ה־iterable</td>
<td>כשמגיעים לתנאי של הפונקציה וערכו שקול ל־<code>False</code></td>
</tr>
<tr>
<td>שימושים עיקריים</td>
<td>ביצוע פעולה עבור כל ערך בסדרת ערכים,
כמו איברי רשימה או תווים במחרוזת</td>
<td>חזרה על פעולה כל עוד מצאנו שהמשימה לא הושלמה</td>
</tr>
</tbody>
</table>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
בשלב הנוכחי בקורס, תמיד נוכל להשתמש בלולאת <code>while</code> במקום בלולאת <code>for</code>, אך לא תמיד נוכל להחליף לולאות <code>for</code> בלולאות <code>while</code>.<br>
באופן כללי, לולאות <code>while</code> יכולות להוכיח את עצמן כשימושיות מאוד מפעם לפעם.<br>
חשבו, לדוגמה, על מצב שבו אתם צריכים לקבל מהמשתמש קלט חדש כל עוד הקלט שלו לא תקין.
</p>
<div class="align-center" style="display: flex; text-align: right; direction: rtl; clear: both;">
<div style="display: flex; width: 10%; float: right; clear: both;">
<img src="images/exercise.svg" style="height: 50px !important;" alt="תרגול">
</div>
<div style="width: 70%">
<p style="text-align: right; direction: rtl; float: right; clear: both;">
כתבו 3 דוגמאות מילוליות נוספות ללולאות <code>for</code>.
</p>
</div>
<div style="display: flex; width: 20%; border-right: 0.1rem solid #A5A5A5; padding: 1rem 2rem;">
<p style="text-align: center; direction: rtl; justify-content: center; align-items: center; clear: both;">
<strong>חשוב!</strong><br>
פתרו לפני שתמשיכו!
</p>
</div>
</div>
### <span style="text-align: right; direction: rtl; float: right;">תרגיל ביניים: פורלולה 1</span>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
בתחרות המרוצים "פורלולה 1", שבה משתתפות בקביעות 6 מכוניות מרוץ, אפשר להמר על הסדר שבו יגיעו המכוניות לקו הסיום.<br>
משתתף זוכה הוא משתתף שהצליח לנחש נכונה את סדר ההגעה של המכוניות לקו הסיום, עם לא יותר מ־2 טעויות.<br>
כתבו פונקציה שמקבלת הימור בודד ואת סדר ההגעה של המכוניות במרוץ, והחזירו אם ההימור זכה או הפסיד.
</p>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
לדוגמה, במרוץ האחרון סדר ההגעה לקו הסיום היה:<br>
<samp dir="ltr">[1, 2, 3, 4, 5, 6]</samp>
</p>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
הנה דוגמאות להימורים של משתתפים ולתוצאתם:<br>
</p>
<ul style="text-align: right; direction: rtl; float: right; clear: both;">
<li><samp dir="ltr">[1, 2, 3, 4, 5, 6]</samp> – הימור זוכה (0 טעויות)</li>
<li><samp dir="ltr">[2, 1, 3, 4, 5, 6]</samp> – הימור זוכה (2 טעויות)</li>
<li><samp dir="ltr">[1, 2, 6, 4, 5, 3]</samp> – הימור זוכה (2 טעויות)</li>
<li><samp dir="ltr">[1, 2, 4, 4, 5, 6]</samp> – הימור זוכה (טעות אחת)</li>
<li><samp dir="ltr">[1, 6, 2, 4, 5, 3]</samp> – הימור מפסיד (3 טעויות)</li>
<li><samp dir="ltr">[5, 3, 2, 4, 6, 1]</samp> – הימור מפסיד (5 טעויות)</li>
<li><samp dir="ltr">[6, 5, 4, 3, 2, 1]</samp> – הימור מפסיד (6 טעויות)</li>
</ul>
### <span style="text-align: right; direction: rtl; float: right;">מבנים מורכבים</span>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
הרשימה הבאה מכילה tuple־ים בגודל 2 איברים:
</p>
```
words = [('star', 'rats'), ('wolf', 'flow'), ('racecar', 'racecar'), ('ekans', 'snake')]
```
<div class="align-center" style="display: flex; text-align: right; direction: rtl; clear: both;">
<div style="display: flex; width: 10%; float: right; clear: both;">
<img src="images/exercise.svg" style="height: 50px !important;" alt="תרגול">
</div>
<div style="width: 70%">
<p style="text-align: right; direction: rtl; float: right; clear: both;">
כתבו לולאת <code>for</code> שתדפיס עבור כל זוג מחרוזות ברשימה: <samp dir="ltr" style="direction: ltr">Flip "X" to get "Y"</samp>.<br>
לדוגמה, עבור הזוג האחרון מתוך 4 הזוגות, היא תדפיס: <samp dir="ltr" style="direction: ltr">Flip "ekans" to get "snake"</samp>.
</p>
</div>
<div style="display: flex; width: 20%; border-right: 0.1rem solid #A5A5A5; padding: 1rem 2rem;">
<p style="text-align: center; direction: rtl; justify-content: center; align-items: center; clear: both;">
<strong>חשוב!</strong><br>
פתרו לפני שתמשיכו!
</p>
</div>
</div>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
כדי לפתור את התרגיל, כתבתם קוד שהשתמש במיקום של הערך הראשון (0) ושל הערך השני (1).<br>
כך אני פתרתי אותו:
</p>
```
for word_and_reversed_word in words:
word = word_and_reversed_word[0]
reversed_word = word_and_reversed_word[1]
print(f'Flip "{word}" to get "{reversed_word}".')
```
<p style="text-align: right; direction: rtl; float: right; clear: both;">
נבחן דרך נוספת לפתור את התרגיל, רק שהפעם נשתמש בטריק שנקרא <dfn>unpacking</dfn> (או "<dfn>פירוק</dfn>").<br>
כיוון שכל tuple ברשימת <var>words</var> מכיל בדיוק 2 איברים, נוכל לתת להם שמות כבר בראש הלולאה ולחלץ אותם מה־tuple:
</p>
```
for word, reversed_word in words:
print(f'Flip "{word}" to get "{reversed_word}".')
```
<p style="text-align: right; direction: rtl; float: right; clear: both;">
בכל חזרור של הלולאה שכתובה למעלה, ה־<code>for</code> יעבור על tuple בודד מתוך <var>words</var>, בדיוק כמו שקרה עד עכשיו.<br>
השינוי הוא שבמקום שה־tuple יישמר כמו שהוא במשתנה בודד שהוגדר בראש הלולאה, הערכים שבו "יחולצו" למשתנים בראש הלולאה.<br>
הטריק הזה עובד כיוון שבראש הלולאה כתבנו מספר משתנים שזהה למספר הערכים שנמצאים בכל tuple.
</p>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
לפניכם תרשים המתאר את תוכן המשתנה <var>words</var>.<br>
הטבלה הגדולה מייצגת את הרשימה <var>words</var>, וכל אחד מהתאים שבה מייצג tuple בתוך הרשימה.<br>
בכל חזרור של ה־<code>for</code> המופיע למעלה, המשתנה <var>word</var> יקבל ערך שמסומן באדום, והמשתנה <var>reversed_word</var> יקבל את הערך הירוק התואם לו.
</p>
<table style="font-size: 1.5rem; border: 0px solid black; border-spacing: 0px;">
<caption style="direction: rtl; text-align: center;">תצוגה של המשתנה <var>words</var> ושל צורת הפירוק שלו</caption>
<tr>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">2</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">3</td>
</tr>
<tbody>
<tr>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">
<table style="font-size: 1.5rem; border: 0px solid black; border-spacing: 0px;">
<tr>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
</tr>
<tbody>
<tr>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: #FF8578;">"star"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: #98FB98;">"rats"</td>
</tr>
<tr style="background: #f5f5f5;">
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-2</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
</tr>
</tbody>
</table>
</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">
<table style="font-size: 1.5rem; border: 0px solid black; border-spacing: 0px;">
<tr>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
</tr>
<tbody>
<tr>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: #FF8578;">"wolf"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: #98FB98;">"flow"</td>
</tr>
<tr style="background: #f5f5f5;">
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-2</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
</tr>
</tbody>
</table>
</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">
<table style="font-size: 1.5rem; border: 0px solid black; border-spacing: 0px;">
<tr>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
</tr>
<tbody>
<tr>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: #FF8578;">"racecar"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: #98FB98;">"racecar"</td>
</tr>
<tr style="background: #f5f5f5;">
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-2</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
</tr>
</tbody>
</table>
</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">
<table style="font-size: 1.5rem; border: 0px solid black; border-spacing: 0px;">
<tr>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
</tr>
<tbody>
<tr>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: #FF8578;">"ekans"</td>
<td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: #98FB98;">"snake"</td>
</tr>
<tr style="background: #f5f5f5;">
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-2</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
</tr>
</tbody>
</table>
</td>
</tr>
<tr style="background: #f5f5f5;">
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-4</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-3</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-2</td>
<td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
</tr>
</tbody>
</table>
### <span style="text-align: right; direction: rtl; float: right;">שינויים בתוך הלולאה</span>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
בדרך כלל נעדיף להימנע משינוי רשימה בזמן ביצוע לולאת <code>for</code>.<br>
שינויים כאלו עלולים לגרום להתנהגות בלתי צפויה, ללולאות אין־סופיות ולקוד קשה במיוחד לקריאה.
</p>
#### <span style="text-align: right; direction: rtl; float: right;">שינוי של מספר האיברים ברשימה בזמן ריצת הלולאה</span>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
נבחן את קטע הקוד הבא, שעבור כל איבר ברשימה, מוציא איבר מסוף הרשימה:
</p>
```
numbers = ['a', 'b', 'c', 'd', 'e']
print(f"The reader expects {len(numbers)} iterations.")
for i in numbers:
j = numbers.pop()
print(i, j)
```
<p style="text-align: right; direction: rtl; float: right; clear: both;">
הלולאה הסתיימה מוקדם מהרגיל, כיוון שכשניסתה להגיע לתא שערכו <code>'d'</code> הוא כבר לא היה שם.<br>
קוד שכזה אינו צפוי, קשה לקריאה ויוצר תקלים. מומלץ מאוד לא לשנות את מספר האיברים ב־iterable בזמן הריצה.
</p>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
פתרון אפשרי הוא ליצור עותק של הרשימה באמצעות הפעולה <code dir="ltr" style="direction: ltr;">list.copy()</code> ולהשתמש בו במקום:
</p>
```
numbers = ['a', 'b', 'c', 'd', 'e']
numbers_to_pop = numbers.copy()
print(f"The reader expects {len(numbers)} iterations.")
for i in numbers:
j = numbers_to_pop.pop()
print(i, j)
```
#### <span style="text-align: right; direction: rtl; float: right;">עריכת הערכים שנמצאים ב־iterable</span>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
ננסה להכפיל כל תא ברשימה שלנו פי 2:
</p>
```
numbers = [1, 3, 5]
print(f'This code will multiply every item in {numbers} by 2.')
print(f'The user expects:')
print(f'[{numbers[0] * 2}, {numbers[1] * 2}, {numbers[2] * 2}]')
for num in numbers:
num = num * 2
print("The final result:")
print(numbers)
```
<p style="text-align: right; direction: rtl; float: right; clear: both;">
נוכל לראות שהרשימה נותרה ללא שינוי, למרות הלולאה שתכליתה היה להכפיל את איברי הרשימה פי 2.
</p>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
כדי להבין למה זה קרה, ננסה להיזכר בשיעור על mutability.<br>
במהלך כל חזרור, המשתנה <var>num</var> מקבל ערך כלשהו <strong>להצביע</strong> עליו.<br>
לדוגמה, בחזרור הראשון <var>num</var> <strong>מצביע</strong> על <code>numbers[0]</code>, המקום הראשון ברשימה:
</p>
<img src="images/mutability1.svg" style="max-width:100%; margin-right: auto; margin-left: auto; text-align: center;" alt="משתנה בשם numbers מצביע על רשימת מספרים שבה יש את האיברים 1, 3 ו־5. מתחתיו יש עוד משתנה שמצביע לאיבר הראשון בתוך הרשימה, 1."/>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
בעצם הפעולה <code>num * 2</code> אנחנו יוצרים ערך חדש שמאוחסן בכתובת שונה.<br>
</p>
<img src="images/mutability2.svg" style="max-width:100%; margin-right: auto; margin-left: auto; text-align: center;" alt="משתנה בשם numbers מצביע על רשימת מספרים שבה יש את האיברים 1, 3 ו־5. מתחתיו יש עוד משתנה שמצביע לאיבר הראשון בתוך הרשימה, 1. נוסף ערך חדש על המסך, 2, שעליו לא מצביע אף משתנה. לידו כתוב: 'התוצאה של num * 2:'"/>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
הפעולה <code>num = num * 2</code> לא באמת "תשנה ערך בתוך <var>num</var>", אלא תגרום לו להצביע על ערך אחר.
</p>
<img src="images/mutability3.svg" style="max-width:100%; margin-right: auto; margin-left: auto; text-align: center;" alt="משתנה בשם numbers מצביע על רשימת מספרים שבה יש את האיברים 1, 3 ו־5. מתחתיו יש עוד משתנה שמצביע לאיבר הראשון בתוך הרשימה, 1. הפעם num מצביע על הערך 2 שנוסף למסך בתמונה הקודמת. הוא כבר לא מצביע על 1."/>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
נסכם כך: <mark>המשתנה <var>num</var> הצביע תחילה על מספר שנמצא בתוך הרשימה, ועכשיו הוא מצביע על מספר אחר.</mark><br>
ההשמה, כרגיל, שינתה את המקום שעליו המשתנה מצביע, ולא את הערך שהמשתנה מצביע עליו.<br>
בקוד שמתואר למעלה, לא שינינו את האיבר בתוך הרשימה.
</p>
### <span style="text-align: right; direction: rtl; float: right;">דוגמאות נוספות</span>
#### <span style="text-align: right; direction: rtl; float: right;">סכום רשימה</span>
```
def total(numbers):
total = 0
for number in numbers:
total = total + number
return total
print(total([1, 2, 3]))
```
#### <span style="text-align: right; direction: rtl; float: right;">ראשי תיבות</span>
```
def acronym(sentence):
acronym_word = ''
for word in sentence.split():
if len(word) >= 1:
acronym_word = acronym_word + word[0]
return acronym_word
print(acronym(''))
```
#### <span style="text-align: right; direction: rtl; float: right;">סכום איברים חיוביים</span>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
קלוט מהמשתמש מספרים. אם צריך, המר את הקלט כך שיהיה מסוג רשימה, ובה יהיו מספרים שלמים. סכום את האיברים הגדולים מ־0.
</p>
```
def to_numbers(strings):
numbers = []
for semi_number in strings:
if semi_number.isdecimal():
numbers.append(int(semi_number))
return numbers
def sum_positives(numbers):
total = 0
for number in numbers:
if number > 0:
total = total + number
return total
user_numbers = input("Enter numbers seperated by comma: ")
stringy_numbers = user_numbers.replace(' ', '').split(',')
numbers = to_numbers(stringy_numbers)
print(sum_positives(numbers))
```
#### <span style="text-align: right; direction: rtl; float: right;">7 בום</span>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
סכום את האיברים שמופיעים בכל מקום שביעי ברשימה.
</p>
```
def sum_only_7th_places(numbers):
total = 0
for i in numbers[6::7]:
total = total + i
return total
print(sum_only_7th_places([0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20]))
```
## <span style="align: right; direction: rtl; float: right; clear: both;">תרגילים</span>
### <span style="align: right; direction: ltr; float: right; clear: both;">אקרוסטיכון</span>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
<dfn>אקרוסטיכון</dfn> הוא אמצעי ספרותי שבו משתמשים בכתיבת שירה.<br>
בשיר שבו יש אקרוסטיכון, כשנחבר את האות הראשונה בכל השורות, נקבל מסר מסוים.<br>
ניקח לדוגמה את שירו של אדגר אלן פו, "אקרוסטיכון", שאותו הוא הקדיש ללטישיה אליזבת' לאנדון ובו מופיע האקרוסטיכון <strong>ELIZABETH</strong>:
</p>
<blockquote style="border-right: none !important; position: relative; padding-left: 0.5em; padding: 1.5em; line-height: 1.5em; min-height: 2em; border-left: 3px solid #a93226; background-color: #fbe7e6; font-size: 1.4rem; direction: ltr; text-align: left; clear: both;">
<strong>E</strong>lizabeth it is in vain you say<br>
<strong>L</strong>ove not — thou sayest it in so sweet a way:<br>
<strong>I</strong>n vain those words from thee or L.E.L.<br>
<strong>Z</strong>antippe's talents had enforced so well:<br>
<strong>A</strong>h! if that language from thy heart arise,<br>
<strong>B</strong>reath it less gently forth — and veil thine eyes.<br>
<strong>E</strong>ndymion, recollect, when Luna tried<br>
<strong>T</strong>o cure his love — was cured of all beside —<br>
<strong>H</strong>is follie — pride — and passion — for he died.<br>
</blockquote>
<p style="text-align: right; direction: rtl; float: right; clear: both;">
כתבו פונקציה שמקבלת שיר כמחרוזת.<br>
החזירו את האקרוסטיכון שנוצר אם ניקח את האות הראשונה מכל שורה בשיר.<br>
</p>
### <span style="align: right; direction: ltr; float: right; clear: both;">שעורה תרבותית</span>
<p style="text-align: right; direction: rtl; clear: both;">
שנים רבות חלפו מאז אותם ימי הקיץ בשדות השעורה, והזמרת סיגנט שכחה היכן החביאה את הזהב שלה.<br>
בידי סיגנט יש מפה, לפיה היא נמצאת כרגע בנקודה <span dir="ltr">(0, 0)</span> בשדה החיטה.<br>
המפה מתארת אילו צעדים היא צריכה לעשות כדי להגיע למטמון.<br>
עזרו לסיגנט לחשב: בהינתן שהיא תלך לפי כל הצעדים שמופיעים במפה – באיזו נקודה יימצא המטמון?
</p>
<p style="text-align: right; direction: rtl; clear: both;">
לדוגמה: עבור הצעדים <code dir="ltr" style="direction: ltr;">[(1, 5), (6, -2), (4, 3)]</code> יוחזר שהמטמון נמצא בנקודה <code dir="ltr" style="direction: ltr;">(11, 6)</code>.
</p>
<p style="text-align: right; direction: rtl; clear: both;">
<strong>הסבר מורחב</strong>: קבלו רשימה של צעדים המורים לסיגנט כיצד ללכת.<br>
כל "צעד" מורכב מזוג מספרים שלמים, שיכולים להיות שליליים – הראשון מסמל כמה צעדים ללכת ימינה, והשני מסמל כמה צעדים ללכת למעלה.<br>
אם המספר הראשון שלילי, עליה ללכת את מספר הצעדים הזה שמאלה. אם המספר השני שלילי, עליה ללכת את מספר הצעדים הזה למטה.<br>
כתבו פונקציה שמקבלת רשימה של צעדים ומחזירה את מיקום המטמון.
</p>
### <span style="align: right; direction: ltr; float: right; clear: both;">גבעת ווטרשיפ</span>
<p style="text-align: right; direction: rtl; clear: both;">
בגבעת ווטרשיפ קצב ההתרבות גבוה. בכל שנה נוספים עוד ועוד ארנבים לארנבייה.<br>
חומש הארנב החליט לנהל מעקב דמוגרפי אחרי הגידול.<br>
הוא מעוניין שתבנו לו פונקציה שמקבלת כפרמטר רשימה של מספר הארנבים שנולדו בכל שנה.<br>
הפונקציה תחזיר רשימה שבה כל תא מייצג את הכמות הנצברת של הארנבים בארנבייה עד כה.<br>
לדוגמה: עבור הרשימה <code dir="ltr" style="direction: ltr;">[1, 2, 3, 4]</code>, הפונקציה תחזיר <code dir="ltr" style="direction: ltr;">[1, 3, 6, 10]</code>.
</p>
| github_jupyter |
```
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import datetime
import tensorflow as tf
import tflearn
import numpy as np
from sklearn.model_selection import train_test_split
import drqn
import student as st
import data_generator as dg
import concept_dependency_graph as cdg
from experience_buffer import ExperienceBuffer
import dataset_utils as d_utils
import utils
import models_dict_utils
from drqn_tests import *
%load_ext autoreload
%autoreload 2
%reload_ext autoreload
```
# General Workflow
## 1. Create Data Set
```
n_concepts = 4
use_student2 = True
student2_str = '2' if use_student2 else ''
learn_prob = 0.15
lp_str = '-lp{}'.format(int(learn_prob*100)) if not use_student2 else ''
n_students = 100000
seqlen = 7
filter_mastery = False
filter_str = '' if not filter_mastery else '-filtered'
policy = 'expert'
filename = 'test{}-n{}-l{}{}-{}{}.pickle'.format(student2_str, n_students, seqlen,
lp_str, policy, filter_str)
```
#### only run the next two cells if dataset hasn't been created yet
```
#concept_tree = sm.create_custom_dependency()
concept_tree = cdg.ConceptDependencyGraph()
concept_tree.init_default_tree(n_concepts)
if not use_student2:
test_student = st.Student(n=n_concepts,p_trans_satisfied=learn_prob, p_trans_not_satisfied=0.0, p_get_ex_correct_if_concepts_learned=1.0)
else:
test_student = st.Student2(n_concepts)
print(filename)
print ("Initializing synthetic data sets...")
dg.generate_data(concept_tree, student=test_student, n_students=n_students, filter_mastery=filter_mastery, seqlen=seqlen, policy=policy, filename="{}{}".format(dg.SYN_DATA_DIR, filename))
print ("Data generation completed. ")
data = d_utils.load_data(filename="../synthetic_data/{}".format(filename))
dqn_data = d_utils.preprocess_data_for_dqn(data, reward_model="semisparse")
dqn_data_train, dqn_data_test = train_test_split(dqn_data, test_size=0.2)
# Creating training and validation data
train_buffer = ExperienceBuffer()
train_buffer.buffer = dqn_data_train
train_buffer.buffer_sz = len(train_buffer.buffer)
val_buffer = ExperienceBuffer()
val_buffer.buffer = dqn_data_test
val_buffer.buffer_sz = len(val_buffer.buffer)
print (train_buffer.sample(1))
```
## 2. Create Model and Train
```
model_id = "test2_model_drqn_mid_expert"
model = drqn.DRQNModel(model_id, timesteps=seqlen-1)
model.init_trainer()
# train the model (uses the previously initialized trainer object)
date_time_string = datetime.datetime.now().strftime("%m-%d-%Y_%H-%M-%S")
run_id = "{}".format(date_time_string)
model.train(train_buffer, val_buffer, n_epoch=60,
run_id=run_id, load_checkpoint=True)
test_drqn(model_id=model_id, DEBUG=True)
model_id = "test2_model_drqn_mid"
model = drqn.DRQNModel(model_id, timesteps=seqlen-1)
model.init_trainer()
# train the model (uses the previously initialized trainer object)
date_time_string = datetime.datetime.now().strftime("%m-%d-%Y_%H-%M-%S")
run_id = "{}".format(date_time_string)
model.train(train_buffer, val_buffer, n_epoch=1,
run_id=run_id, load_checkpoint=True)
a = np.array([[1,2,3], [0,5,6]])
np.argmax(a, axis=1)
```
| github_jupyter |
# Image Compression and Decompression
## Downloading the data and preprocessing it
```
from keras.datasets import mnist
import numpy as np
(x_train, _), (x_test, _) = mnist.load_data()
x_train = x_train.astype('float32') / 255.
x_test = x_test.astype('float32') / 255.
print(x_train.shape,x_test.shape)
x_train = np.reshape(x_train, (len(x_train), 28, 28, 1))
x_test = np.reshape(x_test, (len(x_test), 28, 28, 1))
x_train.shape,x_test.shape
```
## Visualising training data image
```
from matplotlib import pyplot as plt
import numpy as np
first_image = x_train[0]
first_image = np.array(first_image, dtype='float')
pixels = first_image.reshape((28, 28))
plt.imshow(pixels, cmap='gray')
plt.show()
```
## Creating the Autoencoder
```
import keras
from keras import layers
input_img = keras.Input(shape=(28, 28, 1))
x = layers.Conv2D(16, (3, 3), activation='relu', padding='same')(input_img)
x = layers.MaxPooling2D((2, 2), padding='same')(x)
x = layers.Conv2D(8, (3, 3), activation='relu', padding='same')(x)
x = layers.MaxPooling2D((2, 2), padding='same')(x)
x = layers.Conv2D(8, (3, 3), activation='relu', padding='same')(x)
encoded = layers.MaxPooling2D((2, 2), padding='same')(x)
# at this point the representation is (4, 4, 8) i.e. 128-dimensional
x = layers.Conv2D(8, (3, 3), activation='relu', padding='same')(encoded)
x = layers.UpSampling2D((2, 2))(x)
x = layers.Conv2D(8, (3, 3), activation='relu', padding='same')(x)
x = layers.UpSampling2D((2, 2))(x)
x = layers.Conv2D(16, (3, 3), activation='relu')(x)
x = layers.UpSampling2D((2, 2))(x)
decoded = layers.Conv2D(1, (3, 3), activation='sigmoid', padding='same')(x)
autoencoder = keras.Model(input_img, decoded)
autoencoder.compile(optimizer='adam', loss='binary_crossentropy')
autoencoder.summary()
from keras.utils import plot_model
plot_model(autoencoder, to_file='model_plot.png', show_shapes=True, show_layer_names=True)
```
## Training the autoencoder
```
history = autoencoder.fit(x_train, x_train,
epochs=5,
batch_size=128,
shuffle=True,
validation_data=(x_test, x_test))
autoencoder.save("autoencoder.h5")
from keras.models import load_model
autoencoder=load_model("autoencoder.h5")
```
## Testing the trained model and comparing it with the original data
```
decoded_imgs = autoencoder.predict(x_test)
n = 10
plt.figure(figsize=(20, 4))
for i in range(1, n + 1):
# Display original
ax = plt.subplot(2, n, i)
plt.imshow(x_test[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
# Display reconstruction
ax = plt.subplot(2, n, i + n)
plt.imshow(decoded_imgs[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
```
## Visualising the states of a image through the autoencoder
```
from tensorflow.keras import Sequential
import tensorflow as tf
#encoder model
model=tf.keras.models.Sequential([
tf.keras.layers.Conv2D(16,(3,3),activation ='relu', input_shape=(28,28,1)),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Conv2D(8,(3,3),activation ='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Conv2D(8,(3,3),activation ='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Flatten(),
])
def visualize(img,encoder):
code = encoder.predict(img[None])[0]
# Display original
plt.title("Original Image")
plt.imshow(x_test[0].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
#Display compressed
plt.subplot(1,3,2)
plt.title("Compressed Image")
plt.imshow(code.reshape([code.shape[-1]//2,-1]))
plt.show()
# Display reconstruction
plt.title("Decompressed Image")
plt.imshow(decoded_imgs[0].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
visualize(x_test[0],model)
```
## Analysing the loss wrt epoch
```
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
```
# Denoising model for the Decompressed Image
## Adding noise to the train and test data
```
# Adding random noise to the images
noise_factor = 0.5
x_train_noisy = x_train + noise_factor * np.random.normal(loc=0.0, scale=1.0, size=x_train.shape)
x_test_noisy = x_test + noise_factor * np.random.normal(loc=0.0, scale=1.0, size=x_test.shape)
x_train_noisy = np.clip(x_train_noisy, 0., 1.)
x_test_noisy = np.clip(x_test_noisy, 0., 1.)
```
## Visualising the training data
```
n = 10
plt.figure(figsize=(20, 2))
for i in range(1, n + 1):
ax = plt.subplot(1, n, i)
plt.imshow(x_test_noisy[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
print("Training Data:")
plt.show()
```
## Creating the encoder model
```
input_img = keras.Input(shape=(28, 28, 1))
x = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(input_img)
x = layers.MaxPooling2D((2, 2), padding='same')(x)
x = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(x)
encoded = layers.MaxPooling2D((2, 2), padding='same')(x)
# At this point the representation is (7, 7, 32)
x = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(encoded)
x = layers.UpSampling2D((2, 2))(x)
x = layers.Conv2D(32, (3, 3), activation='relu', padding='same')(x)
x = layers.UpSampling2D((2, 2))(x)
decoded = layers.Conv2D(1, (3, 3), activation='sigmoid', padding='same')(x)
autoencoder = keras.Model(input_img, decoded)
autoencoder.compile(optimizer='adam', loss='binary_crossentropy')
```
## Training the model
```
history2 = autoencoder.fit(x_train_noisy, x_train,
epochs=50,
batch_size=128,
shuffle=True,
validation_data=(x_test_noisy, x_test))
from keras import models
autoencoder = models.load_model('denoising_model.h5')
```
## Visualising the results of denoising the decompressed data
```
denoised_imgs = autoencoder.predict(decoded_imgs)
n = 10
plt.figure(figsize=(20, 4))
for i in range(1, n + 1):
# Display original
ax = plt.subplot(2, n, i)
plt.imshow(decoded_imgs[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
# Display reconstruction
ax = plt.subplot(2, n, i + n)
plt.imshow(denoised_imgs[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
```
## Analysing the loss wrt epoch
```
plt.plot(history2.history['loss'])
plt.plot(history2.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
```
## Quality Metrics - PSNR
```
from math import log10, sqrt
import cv2
import numpy as np
def PSNR(original, compressed):
mse = np.mean((original - decompressed) ** 2)
if(mse == 0): # MSE is zero means no noise is present in the signal .
# Therefore PSNR have no importance.
return 100
max_pixel = 255.0
psnr = 20 * log10(max_pixel / sqrt(mse))
return psnr
psnr=0
for i in range(0,50):
original = x_test[i].reshape(28, 28)
decompressed =denoised_imgs[i].reshape(28,28)
value = PSNR(original, decompressed)
psnr+=value
psnr=psnr/50
print(f"PSNR value is {psnr} dB")
```
| github_jupyter |
```
from theano.sandbox import cuda
%matplotlib inline
import utils; reload(utils)
from utils import *
from __future__ import division, print_function
path = "data/"
#path = "data/sample/"
batch_size=64
```
# Setup Directories
```
%cd data
%mkdir valid
%cd train
## make the categories directories under valid
g = glob('*')
for d in g: os.mkdir('../valid/'+d)
## randomly copy 500 over to the valid directory
g = glob('*/*.jpg')
shuf = np.random.permutation(g)
for i in range(500): os.rename(shuf[i], '../valid/' + shuf[i])
## make the sample set directories
%mkdir ../sample
%mkdir ../sample/train
%mkdir ../sample/valid
## make the category directories under sample
g = glob('*')
for d in g:
os.mkdir('../sample/train/'+d)
os.mkdir('../sample/valid/'+d)
# make the sample set
from shutil import copyfile
g = glob('*/*.jpg')
shuf = np.random.permutation(g)
for i in range(400): copyfile(shuf[i], '../sample/train/' + shuf[i])
%cd ../valid
g = glob('*/*.jpg')
shuf = np.random.permutation(g)
for i in range(200): copyfile(shuf[i], '../sample/valid/' + shuf[i])
%mkdir ../results
%mkdir ../sample/results
%cd ../..
```
# Basic VGG
Start with VGG approach (with batch norm). VGG with batch normalization is implemented in [vgg_bn.py](https://github.com/fastai/courses/blob/master/deeplearning1/nbs/vgg16bn.py), and there is a version of ``vgg_ft`` (our fine tuning function) with batch norm called ``vgg_ft_bn`` in [utils.py](https://github.com/fastai/courses/blob/master/deeplearning1/nbs/utils.py).
```
batches = get_batches(path+'train', batch_size=batch_size)
val_batches = get_batches(path+'valid', batch_size=batch_size*2, shuffle=False)
(val_classes, trn_classes, val_labels, trn_labels,
val_filenames, filenames, test_filenames) = get_classes(path)
raw_test_filenames = [f.split('/')[-1] if "test_stg2" not in f else f for f in test_filenames]
raw_test_filenames[13000]
raw_filenames = [f.split('/')[-1] for f in filenames]
#raw_test_filenames = [f.split('/')[-1] for f in test_filenames]
raw_val_filenames = [f.split('/')[-1] for f in val_filenames]
from vgg16bn import Vgg16BN
model = vgg_ft_bn(8)
trn = get_data(path+'train')
val = get_data(path+'valid')
test = get_data(path+'test')
save_array(path+'results/trn.data',trn)
save_array(path+'results/val.data', val)
save_array(path+'results/test.data', test)
trn = load_array(path+'results/trn.data')
val = load_array(path+'results/val.data')
test = load_array(path+'results/test.data')
gen = image.ImageDataGenerator()
model.compile(optimizer=Adam(1e-3), loss='categorical_crossentropy', metrics=['accuracy'])
model.fit(trn, trn_labels, batch_size=batch_size, nb_epoch=3, validation_data=(val, val_labels))
model.save_weights(path+'results/ft1.h5')
```
### Precomput convolutional output
pre-comput the output of the last convolution layer of VGG. (won't fine-tune those layers.
```
model.load_weights(path+'results/ft1.h5')
conv_layers, fc_layers = split_at(model, Convolution2D)
conv_model = Sequential(conv_layers)
conv_feat = conv_model.predict(trn)
conv_val_feat = conv_model.predict(val)
conv_test_feat = conv_model.predict(test)
save_array(path+'results/conv_val_feat.dat', conv_val_feat)
save_array(path+'results/conv_feat.dat', conv_feat)
save_array(path+'results/conv_test_feat.dat', conv_test_feat)
conv_feat = load_array(path+'results/conv_feat.dat')
conv_val_feat = load_array(path+'results/conv_val_feat.dat')
conv_test = load_array(path+'results/conv_test.dat')
```
### Train model
create the baseline model - simple 3-layer FC net
```
def get_bn_layers(p):
return [
MaxPooling2D(input_shape=conv_layers[-1].output_shape[1:]),
BatchNormalization(axis=1),
Dropout(p/4),
Flatten(),
Dense(512, activation='relu'),
BatchNormalization(),
Dropout(p),
Dense(512, activation='relu'),
BatchNormalization(),
Dropout(p/2),
Dense(8, activation='softmax')
]
p=0.6
bn_model = Sequential (get_bn_layers(p))
bn_model.compile(Adam(lr=0.001), loss='categorical_crossentropy', metrics=['accuracy'])
bn_model.fit(conv_feat, trn_labels, batch_size=batch_size, nb_epoch=3,
validation_data=(conv_val_feat, val_labels))
bn_model.optimizer.lr = 1e-4
bn_model.fit(conv_feat, trn_labels, batch_size=batch_size, nb_epoch=7,
validation_data=(conv_val_feat, val_labels))
bn_model.save_weights(path+'models/conv_512_6.h5')
bn_model.load_weights(path+'models/conv_512_6.h5')
```
## Submit
```
def do_clip(arr,mx): return np.clip(arr, (1-mx)/7, mx)
bn_model.evaluate(conv_val_feat, val_labels, batch_size*2)
preds = bn_model.predict(conv_test_feat, batch_size=batch_size)
preds
preds[1]
##preds = preds[1]
##preds = conv_model.predict(test, batch_size=32)
subm = do_clip(preds, 0.82)
classes = ['ALB', 'BET', 'DOL', 'LAG', 'NoF', 'OTHER', 'SHARK', 'YFT']
subm_name = path+'results/subm_bb.gz'
submission = pd.DataFrame(subm, columns=classes)
submission.insert(0, 'image', raw_test_filenames)
submission.head()
submission.to_csv(subm_name, index=False, compression='gzip')
FileLink(subm_name)
```
## Bounding boxes & multi output
```
#get the size of the images because photos with the same sizes should be from the same boat
sizes = [PIL.Image.open(path+'train/'+f).size for f in filenames]
id2size = list(set(sizes))
size2id = {o:i for i,o in enumerate(id2size)}
import collections
collections.Counter(sizes)
trn_sizes_orig = to_categorical([size2id[o] for o in sizes], len(id2size))
raw_val_sizes = [PIL.Image.open(path+'valid/'+f).size for f in val_filenames]
val_sizes = to_categorical([size2id[o] for o in raw_val_sizes], len(id2size))
trn_sizes = trn_sizes_orig-trn_sizes_orig.mean(axis=0)/trn_sizes_orig.std(axis=0)
val_sizes = val_sizes-trn_sizes_orig.mean(axis=0)/trn_sizes_orig.std(axis=0)
import ujson as json
anno_classes = ['alb', 'bet', 'dol', 'lag', 'other', 'shark', 'yft']
bb_json={}
for c in anno_classes:
j = json.load(open('{}annos/{}_labels.json'.format(path,c), 'r'))
for l in j:
if 'annotations' in l.keys() and len(l['annotations'])>0:
bb_json[l['filename'].split('/')[-1]] = sorted(
l['annotations'], key=lambda x:x['height']*x['width'])[-1]
bb_json['img_04908.jpg']
file2idx = {o:i for i,o in enumerate(raw_filenames)}
val_file2idx = {o:i for i,o in enumerate(raw_val_filenames)}
## for any images without annotations, create an empty bounding box
empty_bbox = {'height':0., 'width': 0., 'x':0., 'y': 0.}
for f in raw_filenames:
if not f in bb_json.keys(): bb_json[f] = empty_bbox
for f in raw_val_filenames:
if not f in bb_json.keys(): bb_json[f] = empty_bbox
## convert the dictionary into an array, convert the coordinates to our resized 224x224 images
bb_params = ['height', 'width', 'x', 'y']
def convert_bb(bb, size):
bb = [bb[p] for p in bb_params]
conv_x = (224. / size[0])
conv_y = (224. / size[1])
bb[0] = bb[0]*conv_y
bb[1] = bb[1]*conv_x
bb[2] = max(bb[2]*conv_x, 0)
bb[3] = max(bb[3]*conv_y, 0)
return bb
trn_bbox = np.stack([convert_bb(bb_json[f], s) for f,s in zip(raw_filenames, sizes)],
).astype(np.float32)
val_bbox = np.stack([convert_bb(bb_json[f], s)
for f,s in zip(raw_val_filenames, raw_val_sizes)]).astype(np.float32)
## check our work by drawing one of the annotations
def create_rect(bb, color='red'):
return plt.Rectangle((bb[2], bb[3]), bb[1], bb[0], color=color, fill=False, lw=3)
def show_bb(i):
bb = val_bbox[i]
plot(val[i])
plt.gca().add_patch(create_rect(bb))
show_bb(3)
```
## Create & train model
Not allowed to manually annotate test set.
Create a model that predicts the locations of the bounding box on each image.
It will predict
- type of fish
- 4 bounding box coordinates
```
p = 0.6
inp = Input(conv_layers[-1].output_shape[1:])
x = MaxPooling2D()(inp)
x = BatchNormalization(axis=1)(x)
x = Dropout(p/4)(x)
x = Flatten()(x)
x = Dense(512, activation='relu')(x)
x = BatchNormalization()(x)
x = Dropout(p/4)(x)
x = Dense(512, activation='relu')(x)
x = BatchNormalization()(x)
x = Dropout(p/2)(x)
x_bb = Dense(4, name='bb')(x)
x_class = Dense(8, activation='softmax', name='class')(x)
model = Model([inp], [x_bb, x_class])
model.compile(Adam(lr=0.001), loss=['mse', 'categorical_crossentropy'], metrics=['accuracy'],
loss_weights=[.001, 1.])
model.fit(conv_feat, [trn_bbox, trn_labels], batch_size=batch_size, nb_epoch=3,
validation_data=(conv_val_feat, [val_bbox, val_labels]))
model.optimizer.lr=1e-5
model.fit(conv_feat, [trn_bbox, trn_labels], batch_size=batch_size, nb_epoch=10,
validation_data=(conv_val_feat, [val_bbox, val_labels]))
## check to see how well the bounding box prediction do.
pred = model.predict(conv_val_feat[0:10])
def show_bb_pred(i):
bb = val_bbox[i]
bb_pred = pred[0][i]
plt.figure(figsize=(6,6))
plot(val[i])
ax=plt.gca()
ax.add_patch(create_rect(bb_pred, 'yellow'))
ax.add_patch(create_rect(bb))
show_bb_pred(6)
model.evaluate(conv_val_feat, [val_bbox, val_labels])
model.save_weights(path+'models/bn_anno.h5')
model.load_weights(path+'models/bn_anno.h5')
```
## Larger size
Try to use larger size images. (640x360)
### Set up image size
```
trn = get_data(path+'train', (360,640))
val = get_data(path+'valid', (360,640))
plot(trn[1])
test = get_data(path+'test', (360,640))
save_array(path+'results/trn_640.dat', trn)
save_array(path+'results/val_640.dat', val)
save_array(path+'results/test_640.dat', test)
trn = load_data(path+'results/trn_640.dat')
val = load_data(path+'results/val_640.dat')
## create VGG model. Tell that it is not using the normal 224 x 224
## will not use the fully connected layers
## remove the last max pooling layer
vgg640 = Vgg16BN((360, 640)).model
vgg640.pop()
vgg640.input_shape, vgg640.output_shape
vgg640.compile(Adam(), 'categorical_crossentropy', metrics=['accuracy'])
## pre-compute the convolutional part of VGG
conv_val_feat = vgg640.predict(val, batch_size=32, verbose=1)
conv_trn_feat = vgg640.predict(trn, batch_size=32, verbose=1)
save_array(path+'results/conv_trn_640.dat', conv_trn_feat)
save_array(path+'results/conv_val_640.dat', conv_val_feat)
conv_test_feat = vgg640.predict(test, batch_size=32, verbose=1)
save_array(path+'results/conv_test_640.dat', conv_test_feat)
conv_trn_feat = load_data(path+'results/conv_trn_640.dat')
conv_val_feat = load_data(path+'results/conv_val_640.dat')
conv_test_feat = load_data(path+'results/conv_test_640.dat')
```
| github_jupyter |
This notebook shows the use of `PowerLawRates`. This is a class meant to make the use of
rates that depend on the redshift through a power law easier. It follows the API set out
by `population_param_abstracts.BaseRateDistributions`.
```
from varpop import PowerLawRates
import varpop
print(varpop.__version__)
import numpy as np
%matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('whitegrid')
```
## Instantiate the class
```
rng = np.random.RandomState(1)
pl = PowerLawRates(rng, sky_area=5, zlower=1.0e-8, zhigher=1.2, num_bins=200,
sky_fraction=None, zbin_edges=None, beta_rate=1.0)
```
The settings can be acccessed throughi the following attributes
```
pl.randomState
# This is the area of the sky used in sq degrees
pl.sky_area
pl.sky_fraction
```
An alternative way of specifying the same class is through the other parameters
```
pl2 = PowerLawRates(rng, sky_area=None,
sky_fraction=pl.sky_fraction, zbin_edges=pl.zbin_edges, beta_rate=1.0)
```
## Bin Edges :
This is the actual set of the bin edges used in the calculation.
These are the fundamental quantities irrespective of whether the edges are set
using `zbin_edges` or `zlower`, `zhigher`, `num_bins`
```
pl.zbin_edges
```
The rate at each of these bins (at the mid point) of the redshifts defining the edges is given by the `volumetric_rate` and used for calculating the number of sources in the bin.
```
pl.volumetric_rate(np.array([0.1, 0.5, 1.0]))
```
### z_sample_size:
The expected number of TDA sources in each of the redshift bins is given by the following expression. This could be improved by first sampling `z_samples` and then binning which I intend to add in time. The problem with the current method is demonstrated below, but it is fine for small bins
```
pl.z_sample_sizes
```
## z_samples
The actual samples of redshift can be obtained through
```
pl.z_samples
```
One can compare the size of `z_samples` and the expected size, `z_sample_sizes.sum()`
```
pl.z_sample_sizes.sum()
```
# A Comparison for SN
Let us choose the the canonical values for SNIa rates `alpha_rate` = 2.6e-5, and `beta_rate` = 1.0 (there are cases where this number is reported to be `1.5`, but we will ignore that for the simplicity of the limiting case. This means that the number density in each redshift bin is
```n(z) = volumetric rate(z) * survey Time / (1 + z) = alpha_rate * survey Time * (h/0.7)^3```
Thus the expected number at z < 1.2 is Comoving Vol (z=1.2) * sky_fraction * alpha * (h/0.7)**3 and should match `pl.z_sample_size.sum()` for a sanity check
```
pl.cosmology.comoving_volume(1.2).value * ((5* np.radians(1)**2)/np.pi/4.) * pl.alpha_rate * 10. * (pl.cosmology.h/0.7)**3
pl.z_sample_sizes.sum()
```
| github_jupyter |
```
import warnings
warnings.filterwarnings("ignore")
from IPython.core.display import display, HTML
display(HTML("<style>.container {width: 80% !important; }</style>"))
import sys
import time
import scanpy as sc
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import matplotlib
from matplotlib import colors
myColors = ['#e6194b', '#3cb44b', '#ffe119', '#4363d8', '#f58231',
'#911eb4', '#46f0f0', '#f032e6', '#bcf60c', '#fabebe',
'#008080', '#e6beff', '#9a6324', '#fffac8', '#800000',
'#aaffc3', '#808000', '#ffd8b1', '#000075', '#808080',
'#307D7E', '#000000', "#DDEFFF", "#000035", "#7B4F4B",
"#A1C299", "#300018", "#C2FF99", "#0AA6D8", "#013349",
"#00846F", "#8CD0FF", "#3B9700", "#04F757", "#C8A1A1",
"#1E6E00", "#DFFB71", "#868E7E", "#513A01", "#CCAA35"]
colors2 = plt.cm.Reds(np.linspace(0, 1, 128))
colors3 = plt.cm.Greys_r(np.linspace(0.7,0.8,20))
colorsComb = np.vstack([colors3, colors2])
mymap = colors.LinearSegmentedColormap.from_list('my_colormap', colorsComb)
sys.path.append("../../functions")
from SMaSH_functions import SMaSH_functions
sf = SMaSH_functions()
sys.path.append("/home/ubuntu/Taneda/Functions")
from scRNA_functions import scRNA_functions
fc = scRNA_functions()
```
# Loading annData object
```
obj = sc.read_h5ad('../../../../External_datasets/Healthy_foetal_organ_20pct.h5ad')
print("%d genes across %s cells"%(obj.n_vars, obj.n_obs))
```
#### Removing general genes
```
s1 = time.time()
obj = sf.remove_general_genes(obj)
```
#### Removing house-keeping genes
http://www.housekeeping.unicamp.br/?homePageGlobal
```
obj = sf.remove_housekeepingenes(obj, path="../../data/house_keeping_genes_human_foetal_HSC.txt")
obj = sf.remove_housekeepingenes(obj, path="../../data/house_keeping_genes_human_foetal_liver.txt")
```
#### Removing genes expressed in less than 30% within groups
```
obj = sf.remove_features_pct(obj, group_by="organ", pct=0.3)
```
#### Removing genes expressed in more than 50% in a given group where genes are expressed for more 75% within a given group
```
obj = sf.remove_features_pct_2groups(obj, group_by="organ", pct1=0.75, pct2=0.5)
```
#### Revert PCA
```
obj = sf.scale_filter_features(obj, n_components=None, filter_expression=True)
```
#### ensemble_learning
```
s2 = time.time()
clf = sf.ensemble_learning(obj, group_by="organ", classifier="BalancedRandomForest", balance=True, verbose=True)
```
#### gini_importance
```
selectedGenes, selectedGenes_dict = sf.gini_importance(obj, clf, group_by="organ", verbose=True, restrict_top=("local", 20))
e2 = time.time()
```
#### Classifiers
```
sf.run_classifiers(obj, group_by="organ", genes=selectedGenes, classifier="KNN", balance=True, title="BRF-KNN")
```
#### Sorting genes per cluster
```
axs, selectedGenes_top_dict = sf.sort_and_plot(obj, selectedGenes, group_by="organ", group_by2=None, top=5, figsize=(4,12))
e1 = time.time()
axs.savefig("Figures/BRF_top5_perGroup.pdf")
```
# Elapsed time
```
print("%d genes across %s cells"%(obj.n_vars, obj.n_obs))
print('Elapsed time (s): ', e1-s1)
print('Elapsed time (s): ', e2-s2)
```
| github_jupyter |
# Publications markdown generator for academicpages
Takes a TSV of publications with metadata and converts them for use with [academicpages.github.io](academicpages.github.io). This is an interactive Jupyter notebook ([see more info here](http://jupyter-notebook-beginner-guide.readthedocs.io/en/latest/what_is_jupyter.html)). The core python code is also in `publications.py`. Run either from the `markdown_generator` folder after replacing `publications.tsv` with one containing your data.
TODO: Make this work with BibTex and other databases of citations, rather than Stuart's non-standard TSV format and citation style.
## Data format
The TSV needs to have the following columns: pub_date, title, venue, excerpt, citation, site_url, and paper_url, with a header at the top.
- `excerpt` and `paper_url` can be blank, but the others must have values.
- `pub_date` must be formatted as YYYY-MM-DD.
- `url_slug` will be the descriptive part of the .md file and the permalink URL for the page about the paper. The .md file will be `YYYY-MM-DD-[url_slug].md` and the permalink will be `https://[yourdomain]/publications/YYYY-MM-DD-[url_slug]`
This is how the raw file looks (it doesn't look pretty, use a spreadsheet or other program to edit and create).
```
!cat publications.csv
```
## Import pandas
We are using the very handy pandas library for dataframes.
```
import pandas as pd
```
## Import TSV
Pandas makes this easy with the read_csv function. We are using a TSV, so we specify the separator as a tab, or `\t`.
I found it important to put this data in a tab-separated values format, because there are a lot of commas in this kind of data and comma-separated values can get messed up. However, you can modify the import statement, as pandas also has read_excel(), read_json(), and others.
```
publications = pd.read_csv("publications.csv", sep=",", header=0)
publications
```
## Escape special characters
YAML is very picky about how it takes a valid string, so we are replacing single and double quotes (and ampersands) with their HTML encoded equivilents. This makes them look not so readable in raw format, but they are parsed and rendered nicely.
```
html_escape_table = {
"&": "&",
'"': """,
"'": "'"
}
def html_escape(text):
"""Produce entities within text."""
return "".join(html_escape_table.get(c,c) for c in text)
```
## Creating the markdown files
This is where the heavy lifting is done. This loops through all the rows in the TSV dataframe, then starts to concatentate a big string (```md```) that contains the markdown for each type. It does the YAML metadata first, then does the description for the individual page.
```
import os
for row, item in publications.iterrows():
print(item.pub_date)
md_filename = str(item.pub_date) + "-" + item.url_slug + ".md"
html_filename = str(item.pub_date) + "-" + item.url_slug
year = str(item.pub_date)[:4]
## YAML variables
md = "---\ntitle: \"" + item.title + '"\n'
md += """collection: publications"""
md += """\npermalink: /publication/""" + html_filename
# if len(str(item.excerpt)) > 5:
# md += "\nexcerpt: '" + html_escape(item.excerpt) + "'"
md += "\ndate: " + str(item.pub_date)
md += "\nvenue: '" + html_escape(item.venue) + "'"
if len(str(item.paper_url)) > 5:
md += "\npaperurl: '" + item.paper_url + "'"
# md += "\ncitation: '" + html_escape(item.citation) + "'"
md += "\nauthors: '" + item.authors + "'"
md += "\n---"
## Markdown description for individual page
# if len(str(item.excerpt)) > 5:
# md += "\n" + html_escape(item.excerpt) + "\n"
# if len(str(item.paper_url)) > 5:
# md += "\n[Download paper here](" + item.paper_url + ")\n"
# md += "\nRecommended citation: " + item.citation
md_filename = os.path.basename(md_filename)
with open("../_publications/" + md_filename, 'w') as f:
f.write(md)
```
These files are in the publications directory, one directory below where we're working from.
```
!ls ../_publications/
!cat ../_publications/2018-09-01-icde.md
```
| github_jupyter |
# 10.1 Defining a Function
The keyword `def` introduces a [function definition](https://docs.python.org/3.5/tutorial/controlflow.html#defining-functions). It's followed by the function name, parenthesized list of formal parameters, and ends with a colon. The indented statements below it are executed with the function name is called.
```
def fib(n):
result = []
a, b = 0, 1
while a < n:
result.append(a) # see below
a, b = b, a+b
return result
```
The __`fib()`__ function is defined above. Now let's call this function. Calling a function is simple.
```
fib(10) # oops
```
## 10.2 Positional Arguments
The function requires a positional argument: "__`n`__". This is a good time to mention that naming things descriptively really helps. Coupled with Python's helpful error messages, descriptive variable, function, and class names make it easy to understand and debug errors. In this case, 'n' is a number. Specifically, this function returns a fibonacci sequence for as long as the numbers in the squence are less than the given max number.
Let's give it a better name and then call the function properly.
```
def fib(max_number):
"""Return a list containing the Fibonacci series up to max_number."""
result = []
a, b = 0, 1
while a < max_number:
result.append(a) # see below
a, b = b, a+b
return result
fib(17)
```
## 10.3 Keyword Arguments
Arguments can be made optional when default values are provided. These are known as keyword arguments.
Let's make our argument optional with a default max_number then let's call our function without any arguments.
```
def fib(max_number=17):
"""Return a list containing the Fibonacci series up to max_number."""
result = []
a, b = 0, 1
while a < max_number:
result.append(a) # see below
a, b = b, a+b
return result
fib()
```
Now let's try calling our function with a different argument.
```
fib(3) # still works!
```
## 10.4 Argument Syntax
There can be any number of positional arguments and any number of optional arguments. They can appear together in a function definition for as long as required positional arguments come before optional defaulted arguments.
```
def foo(p=1, q):
return p, q
foo(1)
def foo(p, q, r=1, s=2):
return p, q, r, s
foo(-1, 0)
def foo(p, q, r=1, s=2):
return p, q, r, s
foo(0, 1, s=3, r=2) # the order of defaulted arguments doesn't matter
```
## 10.5 Starred Arguments
In Python, there's a third way of passing arguments to a function. If you wanted to pass a list with an unknown length, even empty, you could pass them in starred arguments.
```
args = [1, 2, 3, 4, 5]
def arguments(*args):
for a in args:
print(a)
return args
arguments(args)
```
We could have specified each argument and it would have worked but that would mean our arguments are fixed. Starred arguments give us flexibility by making the positional arguments optional and of any length.
```
arguments() # still works!
```
For keyword arguments, the only difference is to use `**`. You could pass a dictionary and it would be treated as an arbitrary number of keyword arguments.
```
kwargs = {'a': 1, 'b': 2, 'c': 3, 'd': 4, 'e': 5}
def keywords(**kwargs):
for key, value in kwargs.items():
print(key, value)
return kwargs
keywords(**kwargs)
keywords() # still works!
```
## 10.6 Packing and Unpacking Arguments
### `def function(*args, **kwargs):`
This pattern allows you to change functionality while avoiding breaking your code by just checking the arguments if certain parameters exist and then adding a conditional statement based on that.
Class methods that use this pattern allow data to be passed between objects without loss, transforming the data as needed without needing to know about other objects.
Let's look at more exmaples to illustrate the differences.
```
greeting = 'hello'
def echo(arg):
return arg
echo(greeting)
echo('hello renzo') # it's required...
greeting = 'hello'
def echo(*arg):
return arg
echo(greeting)
greeting = 'hello'
def echo(*arg):
return arg
echo(*greeting) # asterisk unpacks iterables
greeting = ['hello'] # it's now a list
def echo(*arg):
return arg
echo(*greeting)
greeting = [
'hello',
'hi',
'ohayou',
]
def echo(*arg):
return arg
echo(*greeting) # accepts lists
echo() # still works!
```
Let's try it with keyword arguments.
```
kwargs = {
'greeting1': 'Hello',
'greeting2': 'Hi',
'greeting3': 'Ohayou',
}
def echo(kwarg=None, **kwargs):
print(kwarg)
return kwargs
echo(kwargs) # the dictionary data type is unordered unlike lists
echo(**kwargs)
kwargs = {
'greeting1': 'Hello',
'greeting2': 'Hi',
'greeting3': 'Ohayou',
'kwarg': 'World!', # we have a default value for this, which is None
}
def echo(kwarg=None, **kwargs):
print(kwarg)
return kwargs
echo(**kwargs)
```
The dictionary we passed was unpacked and considered as if it was a bunch of keyword arguments passed to the function.
Notice how the keyword argument with a default value was overridden.
| github_jupyter |
# CONTROL DE FLUJOS
### IF
- Toma de decisión simple. Sólo si se cumple la condición dada, entonces, genera la acción deseada.
- Con la siguiente estructura:
if (condición1) entonces:
Acción1
Fin
```
x = 53
y = 400
if y > x:
print("'y' es mayor que 'x'")
#algoritmo de aprobación de curso
nota= float (input("indique nota de estudiante: "))
#si nota es mayor que 3.0 (condición)...
if (nota>3.0):
print("Aprueba curso")
```
### ELSE
- Toma de decisión doble. (De lo contrario...) ... todo lo que no esté afirmado por las condiciones anteriores.
- Con la siguiente estructura:
if (condición1) entonces:
Acción1
else
Acción2
Fin
```
#algoritmo de pico y cédula
x= int (input("indique número de documento: "))
#si x termina en par (condición)...
if (x%2)==0:
print("Puede salir los días lunes, miércoles, viernes y domingo")
#de lo contrario...
else:
print("Puede salir los días martes, jueves, sábado y domingo")
#En este caso el joven puede salir a jugar, solo si ordenó la casa o sacó 5 en el examen.
x= input("conteste si ordenó la casa: ").upper() #convierte a mayúsculas
y= float (input("ponga su nota: "))
if x=="SI" or y==5:
print("Puede salir")
else:
print("no puede salir")
```
### ELIF
- Es la abreviatura de 'else if' ..." si la condición anterior no es cierta, entonces pruebe esta siguiente condición".
- Con la siguiente estructura:
if (condición1) entonces:
Acción 1
elif (condición2) entonces:
Acción 2
elif (condición n) entonces:
Acción n
Fin
Ejemplos:
```
x = 10
y = 10
if y > x:
print("'y' es mayor que 'x'")
elif x == y:
print("'x' y 'y' son iguales")
x = 30
y = 10
if y > x:
print("'y' es mayor que 'x'")
elif x == y:
print("'x' y 'y' son iguales")
else:
print ("'x' es mayor que 'y'")
```
- Se pueden enlazar varias condiciones 'else if'...
```
#Algoritmo para determinar si el número ingresado es negativo, positivo o cero.
x = int (input("Por favor ingresa un número: "))
if x < 0:
print('X es Negativo')
elif x > 0:
print('X es positivo')
elif x == 0:
print('X es cero')
```
### Ejercicio.
La luz de un semáforo nos indica las siguientes opciones:
- Verde = Siga
- Amarillo = Precaución
- Rojo = Pare
Escriba un código para esta situación utilizando el condicional.
```
luz = input("Digite el color de la luz").upper()
print(luz)
if luz == "VERDE":
print ("siga")
elif luz == "AMARILLO":
print("precaucion")
else:
print("pare")
```
### Ejercicio 2
Utilice ahora condiciones anidadas.
Si tenemos una variable booleana que nos indica si hay peaton o no, las condiciones cambian de esta forma:
- Verde -------- Si hay peaton= Pare, Sino = Siga
- Amarillo ----------- Si hay peaton =Pare, Sino= Precaución
- Rojo = Pare
```
peaton= (input("""hay peatón?, indique:
si o no: """)).upper()
#entrada luz
luz= input("Digite el color de la luz").upper()
#entrada booleana si hay o no peatón.
if peaton=="SI":
x=True
else:
x=False
if luz=="VERDE":
if not x:
print("Siga")
else:
print("Pare")
elif luz=="AMARILLO":
if not x:
print("Precaucion")
else:
print("Pare")
elif luz=="ROJO":
print("Pare")
```
### Ejercicio 3
Si el mercado me cuesta hasta 100, pago con dinero en efectivo.
Si me cuesta más que 100 pero menos de 300, pago con tarjeta de débito.
De lo contrario, si me cuesta aún más pago con tarjeta de crédito.
Escriba el código correspondiente, solicitando el valor de la compra para saber cómo procedo a pagar.
```
valor_compra= int (input("Digite el valor de la compra: "))
if valor_compra <= 100:
print ("Pago con dinero en efectivo")
elif valor_compra > 100 and valor_compra < 300:
print ("Pago con débito")
else:
print("Pago con crédito")
```
# LOOPS (BUCLES)
### FOR
- El bucle "for" se puede usar para iterar sobre una secuencia (lista, diccionario, cadena), en el orden en que aparezcan en la secuencia.
Ejemplo:
```
# Ver todos los elementos de la lista.
comprar = ["manzanas", "vegetales", "queso", "gaseosa", "servilletas"]
#for itera sobre la lista comprar e imprime cada uno de los elementos:
for i in comprar:
print(i)
# Ver todos los elementos de una lista de números.
valores = [2,34,11,87,55,23,12,23,12,66,8,3,0,1,3,5,66]
#for itera sobre la lista comprar e imprime cada uno de los elementos:
for i in valores:
print(i)
```
- Incluso, como un string es una cadena iterable, podría usar un *for* para ver todas las letras del primer elemento de la lista
```
for j in "manzanas":
print(j)
```
### WHILE
- (mientras que...)
Con while podremos ejecutar una serie de sentencias, siempre y cuando una condición se cumpla.
```
i = 1
while i < 6:
print(i)
i += 1
```
¿Por qué solo imprime hasta 5?
¿Qué quiere decir i+=1? ¿Cuáles son otras formas de escribir esto?
```
#Ejemplo.
# dado un conjunto de numeros ingresado por teclado, determine, cuántos fueron positivos y cuántos, fueron negativos...
# hasta que se ingrese el número 0.
numero= ""
pos=0
neg=0
while numero !=0:
numero= int (input ("digite numero"))
if numero>=1:
pos+= 1
else:
neg+= 1
print("""La cantidad de numeros positivos es {0}
y la cantidad de negativos es {1}""".format(pos,neg))
#Digitar cualquier palabra hasta que... escriba: salir.
palabra = " "
while palabra != "Salir":
palabra = input("Digite la palabra: ").capitalize()
```
| github_jupyter |
# Problem: IMDB Dataset
### Problem class: Recommender system
### Problem dataset link: https://bit.ly/33IAohl
### Problem description:
Creating a simple recommendation system using IMDB dataset.
### Problem Task:
Recommend top 25 movies.
# Importing libraries
```
import warnings
warnings.filterwarnings('ignore')
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
sns.set()
```
# Load the dataset into a pandas dataframe
```
# """movies_metadata.csv: The main Movies Metadata file.
# Contains information on 45,000 movies featured in the
# Full MovieLens dataset. Features include posters, backdrops,
# budget, revenue, release dates, languages, production countries
# and companies.
# """
df = pd.read_csv("data/movies_metadata.csv")
df.head()
```
## The Metric
```
# TODO: Choose a metric (or score) to rate the movies on
# The metric is the numeric quantity based on which we rank movies
# A movie is considered to be better than another movie if it has a higher metric score than the other movie
# IMDB uses these formula for metric: Weighted Rating (WR) = ((v /(v + m )) * R) + ((m / (v + m)) * C)
# v is the number of votes garnered by the movie
# m is the minimum number of votes required for the movie to be in the chart (the prerequisite)
# R is the mean rating of the movie
# C is the mean rating of all the movies in the dataset
## v and R for every movie in the form of the vote_count and vote_average features
## The number of votes garnered by the 80th percentile movie as our value for m
## Calculating the number of votes garnered by the 80th percentile movie
m = df['vote_count'].quantile(0.80)
m ## Different values of m will change the output of the recommender.
## Insight: Only 20% of the movies have gained more than 50 votes
## TODO: Decide on the prerequisites for the movie to be featured on the chart
# only consider movies that are greater than 45 minutes and less than 300 minutes in length
q_movies = df[(df['runtime'] >= 45) & (df['runtime'] <= 300)]
# Only consider movies that have garnered more than m votes
q_movies = q_movies[q_movies['vote_count'] >= m]
# Inspect the number of movies that made the cut
q_movies.shape
## Insight: Dataset of 45,000 movies approximately 9,000 movies (or 20%) made the cut
## TODO: Calculate the score for every movie that satisfies the conditions
# Calculate C
C = df['vote_average'].mean()
C
## Insight: The average rating of a movie is approximately 5.6/10
# Compute the IMDB weighted rating for each movie
def weighted_rating(df: pd.DataFrame, m: float=m, C: float=C)-> float:
v = df['vote_count']
R = df['vote_average']
# Compute the weighted score
return (v / (v + m) * R) + (m / (m + v) * C)
# Compute the score using the weighted_rating function and added score(new feature) to dataset
## Help Note : axis=1 means rowwise operation
q_movies['score'] = q_movies.apply(weighted_rating, axis=1)
# TODO: Output the list of movies in decreasing order of their scores
# Sort movies in descending order of their scores
q_movies = q_movies.sort_values('score', ascending=False)
# Print the top 25 movies
q_movies[['title', 'vote_count', 'vote_average', 'score', 'runtime']].head(25)
```
| github_jupyter |
## Geospatial Data
### Part 2 of n
# Structured Query Language (SQL)
### Talking to a Database through Queries
## Reminder
<a href="#/slide-2-0" class="navigate-right" style="background-color:blue;color:white;padding:8px;margin:2px;font-weight:bold;">Continue with the lesson</a>
<br>
</br>
<font size="+1">
By continuing with this lesson you are granting your permission to take part in this research study for the Hour of Cyberinfrastructure: Developing Cyber Literacy for GIScience project. In this study, you will be learning about cyberinfrastructure and related concepts using a web-based platform that will take approximately one hour per lesson. Participation in this study is voluntary.
Participants in this research must be 18 years or older. If you are under the age of 18 then please exit this webpage or navigate to another website such as the Hour of Code at https://hourofcode.com, which is designed for K-12 students.
If you are not interested in participating please exit the browser or navigate to this website: http://www.umn.edu. Your participation is voluntary and you are free to stop the lesson at any time.
For the full description please navigate to this website: <a href="../../gateway-lesson/gateway/gateway-1.ipynb">Gateway Lesson Research Study Permission</a>.
</font>
```
# This code cell starts the necessary setup for Hour of CI lesson notebooks.
# First, it enables users to hide and unhide code by producing a 'Toggle raw code' button below.
# Second, it imports the hourofci package, which is necessary for lessons and interactive Jupyter Widgets.
# Third, it helps hide/control other aspects of Jupyter Notebooks to improve the user experience
# This is an initialization cell
# It is not displayed because the Slide Type is 'Skip'
from IPython.display import HTML, IFrame, Javascript, display
from ipywidgets import interactive
import ipywidgets as widgets
from ipywidgets import Layout
import getpass # This library allows us to get the username (User agent string)
# import package for hourofci project
import sys
sys.path.append('../../supplementary') # relative path (may change depending on the location of the lesson notebook)
# sys.path.append('supplementary')
import hourofci
try:
import os
os.chdir('supplementary')
except:
pass
# load javascript to initialize/hide cells, get user agent string, and hide output indicator
# hide code by introducing a toggle button "Toggle raw code"
HTML('''
<script type="text/javascript" src=\"../../supplementary/js/custom.js\"></script>
<style>
.output_prompt{opacity:0;}
</style>
<input id="toggle_code" type="button" value="Toggle raw code">
''')
```
### Database languages :
#### 1) Data definition language(DDL) :
DDL is used to define database objects .The conceptual schema is
specified by a set of definitions expressed by this language. It also give some
details about how to implement this schema in the physical devices used to store
the data. This definition includes all the entity sets and their associated attributes
and their relation ships. The result of DDL statements will be a set of tables that
are stored in special file called data dictionary.
#### 2) Data manipulation language(DML) :
A DML is a language that enables users to access or manipulate data stored in
the database. Data manipulation involves retrieval of data from the database,
insertion of new data into the database and deletion of data or modification of
existing data.
#### 3) Data control language(DCL):
This language enables users to grant authorization and canceling
authorization of database objects.
#### 4) Data Query Language (DQL):
DQL statements are used for performing queries on the data within schema objects. The purpose of DQL commands is to get the schema relation based on the query passed to it. Although often considered part of DML, the SQL `SELECT` statement is strictly speaking an example of DQL. When adding `FROM` or `WHERE` data manipulators to the `SELECT` statement the statement is then considered part of the DML.
### Here we will focus on DDL and DQL languages!
### DDL: Creating and Deleting Databases and Tables
#### Creating a new Database
The general syntax for creating a new database is
```mysql
CREATE DATABASE databasename;
```
#### Dropping/Deleting a Database
The general syntax for dropping database is
```mysql
DROP DATABASE databasename;
```
#### Creating a new Table
The general syntax for creating a Table
```mysql
CREATE TABLE Persons (
PersonID int,
LastName varchar(255),
FirstName varchar(255),
Address varchar(255),
City varchar(255)
)
```
#### Dropping existing Table
The general syntax for dropping an existing Table
```mysql
drop table table_name
```
### DQL: Querying a database
```
import displaydatabases
from questiondisplay import QueryWindow
disp = displaydatabases.Display()
disp.displayDatabases
```
### Select Statement
>**Select statement** is used to **retrieve data from a Database**
The **syntax** for **select statement** is
```mysql
select column1,column2..columnN from table_name
```
Where column1, column2 are the columns that you want to select from the table (with name table_name)
##### Select All Columns from a Table
The syntax for selecting all columns from a table is
```mysql
select * from table_name
```
The <b>*</b> symbol indicates that we would want all the columns from the table.
Let's see some concrete example.
**1. Select all columns from actor table**
```
QueryWindow(1).display()
```
**2. Select all columns from staff table**
```
QueryWindow(2).display()
```
##### Select subset of columns from a table
The syntax for selecting subset of columns from a table is
```mysql
select column1,column2...columnN from table_name
```
Let's see some concrete example.
**3. Select staff_id,first_name and last_name from staff table**
```
QueryWindow(3).display()
```
**4. Select first_name and last_name from actor table**
```
QueryWindow(4).display()
```
##### Select distinct values from a column
The **syntax** for **select distinct statement** is
```mysql
select distinct column1,column2..columnN from table_name
```
Where column1, column2, columnN are the columns that you want to select from the table (with name table_name) and only distinct values for column1 will be selected
Let's see some concrete example.
**5. Select the unique set of cities from employees table**
```
QueryWindow(5).display()
```
<font size="+1"><a style="background-color:blue;color:white;padding:12px;margin:10px;font-weight:bold;" href="dbms-3.ipynb">Click here to go to the next notebook.</a></font>
| github_jupyter |
# Causal Discovery using a Perfect Oracle
A perfect oracle is a conditional independence (CI) test that always yields the true answer.
For that, the oracle requires access to the true underlying graph from which it can read-off the true conditional independence relation.
Although this is impractical, access to the true graph is granted in a simulated environment for evaluating the number of CI tests required by an algorithm in the large sample limit.
For demonstrating how to use a perfect oracle we follow these steps:
1. Create a true underlying DAG
2. Learn a PAG using ICD and a perfect oracle
3. Plot the results
Let's start by importing the required classes and methods.
```
import sys
sys.path.append('..')
from causal_discovery_algs import LearnStructICD # import ICD algorithm
from causal_discovery_utils.cond_indep_tests import DSep # import the perfect oracle
from graphical_models import DAG # import a DAG class
from plot_utils import draw_graph # for plotting the graphs
```
Next, let's create the true underlying DAG, a graph with 7 nodes, and define which nodes are observed.
```
nodes_of_dag = set(range(7))
dag = DAG(nodes_of_dag)
dag.add_edges(parents_set={5, 0}, target_node=1)
dag.add_edges(parents_set={6, 0}, target_node=2)
dag.add_edges(parents_set={5, 2}, target_node=3)
dag.add_edges(parents_set={6, 1}, target_node=4)
observed_set = {0, 1, 2, 3, 4} # latents set is {5, 6}
fig_dag = draw_graph(dag, latent_nodes=nodes_of_dag - observed_set) # plot the DAG
```
Now, instantiate a perfect oracle, set it to count the performed CI tests, and turn on caching.
```
ci_test_dsep = DSep(true_dag=dag, # the DAG from which to read-off the true conditional independence relations
count_tests=True, # count CI tests per conditioning set size
use_cache=True) # make sure the same CI test is not performed and counted more than once
```
Then, learn the casual structure using the perfect oracle.
```
icd = LearnStructICD(nodes_set=observed_set, ci_test=ci_test_dsep) # instantiate ICD
icd.learn_structure() # learn the structure
fig_pag = draw_graph(icd.graph) # plot the PAG
```
Finally, let's plot the number of CI tests per conditioning set size.
```
num_ci_tests = ci_test_dsep.test_counter
str_cs_size = 'Condition set size: '
str_num_cit = 'Number of CI tests: '
max_str_len = len(str(max(num_ci_tests))) + 3
for cs_size, num_cit in enumerate(num_ci_tests):
str_cs_size += str(cs_size).ljust(max_str_len)
str_num_cit += str(num_cit).ljust(max_str_len)
print('Number of estimated CI tests')
print('----------------------------')
print(str_cs_size)
print(str_num_cit)
print('Total number of CI tests: ', sum(num_ci_tests))
```
| github_jupyter |
```
import pandas as pd
import pyspark.sql.functions as F
from datetime import datetime
from pyspark.sql.types import *
from pyspark import StorageLevel
import numpy as np
pd.set_option("display.max_rows", 1000)
pd.set_option("display.max_columns", 1000)
pd.set_option("mode.chained_assignment", None)
from pyspark.ml import Pipeline
from pyspark.ml.classification import GBTClassifier
from pyspark.ml.feature import IndexToString, StringIndexer, VectorIndexer
# from pyspark.ml.evaluation import MulticlassClassificationEvaluator
from pyspark.ml.feature import OneHotEncoderEstimator, StringIndexer, VectorAssembler
from pyspark.ml.evaluation import BinaryClassificationEvaluator
from pyspark.ml.tuning import CrossValidator, ParamGridBuilder
from pyspark.sql import Row
from pyspark.ml.linalg import Vectors
# !pip install scikit-plot
import sklearn
import scikitplot as skplt
from sklearn.metrics import classification_report, confusion_matrix, precision_score
```
<hr />
<hr />
<hr />
```
result_schema = StructType([
StructField('experiment_filter', StringType(), True),
StructField('undersampling_method', StringType(), True),
StructField('undersampling_column', StringType(), True),
StructField('filename', StringType(), True),
StructField('experiment_id', StringType(), True),
StructField('n_covid', IntegerType(), True),
StructField('n_not_covid', IntegerType(), True),
StructField('model_name', StringType(), True),
StructField('model_seed', StringType(), True),
StructField('model_maxIter', IntegerType(), True),
StructField('model_maxDepth', IntegerType(), True),
StructField('model_maxBins', IntegerType(), True),
StructField('model_minInstancesPerNode', IntegerType(), True),
StructField('model_minInfoGain', FloatType(), True),
StructField('model_featureSubsetStrategy', StringType(), True),
StructField('model_n_estimators', IntegerType(), True),
StructField('model_learning_rate', FloatType(), True),
StructField('model_impurity', StringType(), True),
StructField('model_AUC_ROC', StringType(), True),
StructField('model_AUC_PR', StringType(), True),
StructField('model_covid_precision', StringType(), True),
StructField('model_covid_recall', StringType(), True),
StructField('model_covid_f1', StringType(), True),
StructField('model_not_covid_precision', StringType(), True),
StructField('model_not_covid_recall', StringType(), True),
StructField('model_not_covid_f1', StringType(), True),
StructField('model_avg_precision', StringType(), True),
StructField('model_avg_recall', StringType(), True),
StructField('model_avg_f1', StringType(), True),
StructField('model_avg_acc', StringType(), True),
StructField('model_TP', StringType(), True),
StructField('model_TN', StringType(), True),
StructField('model_FN', StringType(), True),
StructField('model_FP', StringType(), True),
StructField('model_time_exec', StringType(), True),
StructField('model_col_set', StringType(), True)
])
```
<hr />
<hr />
<hr />
```
# undersamp_col = ['03-STRSAMP-AG', '04-STRSAMP-EW']
# dfs = ['ds-1', 'ds-2', 'ds-3']
# cols_sets = ['cols_set_1', 'cols_set_2', 'cols_set_3']
undersamp_col = ['02-KMODES']
dfs = ['ds-1']
cols_sets = ['cols_set_1']
# lists of params
model_maxIter = [20, 50, 100]
model_maxDepth = [3, 5, 7]
model_maxBins = [32, 64]
# model_learningRate = [0.01, 0.1, 0.5]
# model_loss = ['logLoss', 'leastSquaresError', 'leastAbsoluteError']
list_of_param_dicts = []
for maxIter in model_maxIter:
for maxDepth in model_maxDepth:
for maxBins in model_maxBins:
params_dict = {}
params_dict['maxIter'] = maxIter
params_dict['maxDepth'] = maxDepth
params_dict['maxBins'] = maxBins
list_of_param_dicts.append(params_dict)
print("There is {} set of params.".format(len(list_of_param_dicts)))
# list_of_param_dicts
prefix = 'gs://ai-covid19-datalake/trusted/experiment_map/'
```
<hr />
<hr />
<hr />
```
# filename = 'gs://ai-covid19-datalake/trusted/experiment_map/03-STRSAMP-AG/ds-1/cols_set_1/experiment0.parquet'
# df = spark.read.parquet(filename)
# df.limit(2).toPandas()
# params_dict = {'maxIter': 100,
# 'maxDepth': 3,
# 'maxBins': 32,
# 'learningRate': 0.5,
# 'loss': 'leastAbsoluteError'}
# cols = 'cols_set_1'
# experiment_filter = 'ds-1'
# undersampling_method = '03-STRSAMP-AG',
# experiment_id = 0
# run_gbt(df, params_dict, cols, filename, experiment_filter, undersampling_method, experiment_id)
```
<hr />
<hr />
<hr />
```
def run_gbt(exp_df, params_dict, cols, filename, experiment_filter,
undersampling_method, experiment_id):
import time
start_time = time.time()
n_covid = exp_df.filter(F.col('CLASSI_FIN') == 1.0).count()
n_not_covid = exp_df.filter(F.col('CLASSI_FIN') == 0.0).count()
id_cols = ['NU_NOTIFIC', 'CLASSI_FIN']
labelIndexer = StringIndexer(inputCol="CLASSI_FIN", outputCol="indexedLabel").fit(exp_df)
input_cols = [x for x in exp_df.columns if x not in id_cols]
assembler = VectorAssembler(inputCols = input_cols, outputCol= 'features')
exp_df = assembler.transform(exp_df)
# Automatically identify categorical features, and index them.
# Set maxCategories so features with > 4 distinct values are treated as continuous.
featureIndexer = VectorIndexer(inputCol="features", outputCol="indexedFeatures", maxCategories=30).fit(exp_df)
# Split the data into training and test sets (30% held out for testing)
(trainingData, testData) = exp_df.randomSplit([0.7, 0.3])
trainingData = trainingData.persist(StorageLevel.MEMORY_ONLY)
testData = testData.persist(StorageLevel.MEMORY_ONLY)
# Train a RandomForest model.
gbt = GBTClassifier(labelCol = "indexedLabel", featuresCol = "indexedFeatures",
maxIter = params_dict['maxIter'],
maxDepth = params_dict['maxDepth'],
maxBins = params_dict['maxBins'])
# Convert indexed labels back to original labels.
labelConverter = IndexToString(inputCol="prediction", outputCol="predictedLabel",
labels=labelIndexer.labels)
# Chain indexers and forest in a Pipeline
pipeline = Pipeline(stages=[labelIndexer, featureIndexer, gbt, labelConverter])
# Train model. This also runs the indexers.
model = pipeline.fit(trainingData)
# Make predictions.
predictions = model.transform(testData)
pred = predictions.select(['CLASSI_FIN', 'predictedLabel'])\
.withColumn('predictedLabel', F.col('predictedLabel').cast('double'))\
.withColumn('predictedLabel', F.when(F.col('predictedLabel') == 1.0, 'covid').otherwise('n-covid'))\
.withColumn('CLASSI_FIN', F.when(F.col('CLASSI_FIN') == 1.0, 'covid').otherwise('n-covid'))\
.toPandas()
y_true = pred['CLASSI_FIN'].tolist()
y_pred = pred['predictedLabel'].tolist()
report = classification_report(y_true, y_pred, output_dict=True)
evaluator_ROC = BinaryClassificationEvaluator(labelCol="indexedLabel", rawPredictionCol="prediction", metricName="areaUnderROC")
accuracy_ROC = evaluator_ROC.evaluate(predictions)
evaluator_PR = BinaryClassificationEvaluator(labelCol="indexedLabel", rawPredictionCol="prediction", metricName="areaUnderPR")
accuracy_PR = evaluator_PR.evaluate(predictions)
conf_matrix = confusion_matrix(y_true, y_pred)
result_dict = {}
result_dict['experiment_filter'] = experiment_filter
result_dict['undersampling_method'] = undersampling_method
result_dict['filename'] = filename
result_dict['experiment_id'] = experiment_id
result_dict['n_covid'] = n_covid
result_dict['n_not_covid'] = n_not_covid
result_dict['model_name'] = 'GBT'
result_dict['params'] = params_dict
result_dict['model_AUC_ROC'] = accuracy_ROC
result_dict['model_AUC_PR'] = accuracy_PR
result_dict['model_covid_precision'] = report['covid']['precision']
result_dict['model_covid_recall'] = report['covid']['recall']
result_dict['model_covid_f1'] = report['covid']['f1-score']
result_dict['model_not_covid_precision'] = report['n-covid']['precision']
result_dict['model_not_covid_recall'] = report['n-covid']['recall']
result_dict['model_not_covid_f1'] = report['n-covid']['f1-score']
result_dict['model_avg_precision'] = report['macro avg']['precision']
result_dict['model_avg_recall'] = report['macro avg']['recall']
result_dict['model_avg_f1'] = report['macro avg']['f1-score']
result_dict['model_avg_acc'] = report['accuracy']
result_dict['model_TP'] = conf_matrix[0][0]
result_dict['model_TN'] = conf_matrix[1][1]
result_dict['model_FN'] = conf_matrix[0][1]
result_dict['model_FP'] = conf_matrix[1][0]
result_dict['model_time_exec'] = time.time() - start_time
result_dict['model_col_set'] = cols
return result_dict
```
<hr />
<hr />
<hr />
# Running GBT on 10 samples for each experiment
### 3x col sets -> ['cols_set_1', 'cols_set_2', 'cols_set_3']
### 3x model_maxIter -> [100, 200, 300]
### 3x model_maxDepth -> [5, 10, 15]
### 3x model_maxBins -> [16, 32, 64]
Total: 10 * 3 * 3 * 3 * 3 = 810
```
experiments = []
```
### Datasets: strat_samp_lab_agegrp
```
for uc in undersamp_col:
for ds in dfs:
for col_set in cols_sets:
for params_dict in list_of_param_dicts:
for id_exp in range(50):
filename = prefix + uc + '/' + ds + '/' + col_set + '/' + 'experiment' + str(id_exp) + '.parquet'
exp_dataframe = spark.read.parquet(filename)
print('read {}'.format(filename))
undersampling_method = uc
experiment_filter = ds
experiment_id = id_exp
try:
model = run_gbt(exp_dataframe, params_dict, col_set, filename, experiment_filter, undersampling_method, experiment_id)
experiments.append(model)
print("Parameters ==> {}\n Results: \n AUC_PR: {} \n Precision: {} \n Time: {}".format(str(params_dict), str(model['model_AUC_PR']), str(model['model_avg_precision']), str(model['model_time_exec'])))
print('=========================== \n')
except:
print('=========== W A R N I N G =========== \n')
print('Something wrong with the exp: {}, {}, {}'.format(filename, params_dict, col_set))
```
<hr />
<hr />
<hr />
```
for i in range(len(experiments)):
for d in list(experiments[i].keys()):
experiments[i][d] = str(experiments[i][d])
# experiments
cols = ['experiment_filter', 'undersampling_method', 'filename', 'experiment_id', 'n_covid', 'n_not_covid', 'model_name', 'params', 'model_AUC_ROC', 'model_AUC_PR', 'model_covid_precision', 'model_covid_recall', 'model_covid_f1', 'model_not_covid_precision', 'model_not_covid_recall', 'model_not_covid_f1', 'model_avg_precision', 'model_avg_recall', 'model_avg_f1', 'model_avg_acc', 'model_TP', 'model_TN', 'model_FN', 'model_FP', 'model_time_exec', 'model_col_set']
intermed_results = spark.createDataFrame(data=experiments).select(cols)
intermed_results.toPandas()
intermed_results.write.parquet('gs://ai-covid19-datalake/trusted/intermed_results/KMODES/GBT_experiments-kmodes-ds1-cs1.parquet', mode='overwrite')
df = spark.read.parquet('gs://ai-covid19-datalake/trusted/intermed_results/KMODES/GBT_experiments-kmodes-ds1-cs1.parquet')
df.limit(2).toPandas()
print('finished')
intermed_results.show()
```
| github_jupyter |
## Kernel Principal Component Analysis (KPCA)
1. Apply the KPCA to reduce the complexity of the dataset to recommend a wine to a customer
2. **Input** = wine.csv
## Importing the libraries
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
```
##Importing the dataset
```
dataset = pd.read_csv('Wine.csv')
X = dataset.iloc[:, :-1].values
y = dataset.iloc[:, -1].values
```
##Splitting the dataset into the Training set and Test set
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
```
##Feature Scaling
```
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
```
##Aplying KPCA
```
from sklearn.decomposition import KernelPCA as KPCA
kpca = KPCA(n_components = 2, kernel='rbf') #poly also performs as 1.0
X_train = kpca.fit_transform(X_train)
X_test = kpca.transform(X_test)
```
##Training the Logistic Regression model on the Training set
```
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression(random_state = 0)
classifier.fit(X_train, y_train)
```
## Making the Confusion Matrix
```
from sklearn.metrics import confusion_matrix, accuracy_score
y_pred = classifier.predict(X_test)
cm = confusion_matrix(y_test, y_pred)
print(cm)
accuracy_score(y_test, y_pred)
```
## Visualising the Training set results
```
from matplotlib.colors import ListedColormap
X_set, y_set = X_train, y_train
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 1 , stop = X_set[:, 0].max() + 1, step = 0.01),
np.arange(start = X_set[:, 1].min() - 1, stop = X_set[:, 1].max() + 1, step = 0.01))
plt.contourf(X1, X2, classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green', 'blue')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i,j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1], c = ListedColormap(('red','green','blue'))(i), label = j)
plt.title('Logistic Regression (Training set)')
plt.xlabel('PC1')
plt.ylabel('PC2')
plt.legend()
plt.show()
```
## Visualising the Test set results
```
from matplotlib.colors import ListedColormap
X_set, y_set = X_test, y_test
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 1 , stop = X_set[:, 0].max() + 1, step = 0.01),
np.arange(start = X_set[:, 1].min() - 1, stop = X_set[:, 1].max() + 1, step = 0.01))
plt.contourf(X1, X2, classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green', 'blue')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i,j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1], c = ListedColormap(('red','green','blue'))(i), label = j)
plt.title('Logistic Regression (Test set)')
plt.xlabel('PC1')
plt.ylabel('PC2')
plt.legend()
plt.show()
```
| github_jupyter |
This application demonstrates how to build a simple neural network using the Graph mark.
Interactions can be enabled by adding event handlers (click, hover etc) on the nodes of the network.
See the [Mark Interactions notebook](../Interactions/Mark Interactions.ipynb) and the [Scatter Notebook](../Marks/Scatter.ipynb) for details.
```
from itertools import chain, product
import numpy as np
from bqplot import *
class NeuralNet(Figure):
def __init__(self, **kwargs):
self.height = kwargs.get('height', 600)
self.width = kwargs.get('width', 960)
self.directed_links = kwargs.get('directed_links', False)
self.num_inputs = kwargs['num_inputs']
self.num_hidden_layers = kwargs['num_hidden_layers']
self.nodes_output_layer = kwargs['num_outputs']
self.layer_colors = kwargs.get('layer_colors',
['Orange'] * (len(self.num_hidden_layers) + 2))
self.build_net()
super(NeuralNet, self).__init__(**kwargs)
def build_net(self):
# create nodes
self.layer_nodes = []
self.layer_nodes.append(['x' + str(i+1) for i in range(self.num_inputs)])
for i, h in enumerate(self.num_hidden_layers):
self.layer_nodes.append(['h' + str(i+1) + ',' + str(j+1) for j in range(h)])
self.layer_nodes.append(['y' + str(i+1) for i in range(self.nodes_output_layer)])
self.flattened_layer_nodes = list(chain(*self.layer_nodes))
# build link matrix
i = 0
node_indices = {}
for layer in self.layer_nodes:
for node in layer:
node_indices[node] = i
i += 1
n = len(self.flattened_layer_nodes)
self.link_matrix = np.empty((n,n))
self.link_matrix[:] = np.nan
for i in range(len(self.layer_nodes) - 1):
curr_layer_nodes_indices = [node_indices[d] for d in self.layer_nodes[i]]
next_layer_nodes = [node_indices[d] for d in self.layer_nodes[i+1]]
for s, t in product(curr_layer_nodes_indices, next_layer_nodes):
self.link_matrix[s, t] = 1
# set node x locations
self.nodes_x = np.repeat(np.linspace(0, 100,
len(self.layer_nodes) + 1,
endpoint=False)[1:],
[len(n) for n in self.layer_nodes])
# set node y locations
self.nodes_y = np.array([])
for layer in self.layer_nodes:
n = len(layer)
ys = np.linspace(0, 100, n+1, endpoint=False)[1:]
self.nodes_y = np.append(self.nodes_y, ys[::-1])
# set node colors
n_layers = len(self.layer_nodes)
self.node_colors = np.repeat(np.array(self.layer_colors[:n_layers]),
[len(layer) for layer in self.layer_nodes]).tolist()
xs = LinearScale(min=0, max=100)
ys = LinearScale(min=0, max=100)
self.graph = Graph(node_data=[{'label': d,
'label_display': 'none'} for d in self.flattened_layer_nodes],
link_matrix=self.link_matrix,
link_type='line',
colors=self.node_colors,
directed=self.directed_links,
scales={'x': xs, 'y': ys},
x=self.nodes_x,
y=self.nodes_y,
# color=2 * np.random.rand(len(self.flattened_layer_nodes)) - 1
)
self.graph.hovered_style = {'stroke': '1.5'}
self.graph.unhovered_style = {'opacity': '0.4'}
self.graph.selected_style = {'opacity': '1',
'stroke': 'red',
'stroke-width': '2.5'}
self.marks = [self.graph]
self.title = 'Neural Network'
self.layout.width = str(self.width) + 'px'
self.layout.height = str(self.height) + 'px'
NeuralNet(num_inputs=3, num_hidden_layers=[10, 10, 8, 5], num_outputs=1)
```
| github_jupyter |
# TF-Slim Walkthrough
This notebook will walk you through the basics of using TF-Slim to define, train and evaluate neural networks on various tasks. It assumes a basic knowledge of neural networks.
## Table of contents
<a href="#Install">Installation and setup</a><br>
<a href='#MLP'>Creating your first neural network with TF-Slim</a><br>
<a href='#ReadingTFSlimDatasets'>Reading Data with TF-Slim</a><br>
<a href='#CNN'>Training a convolutional neural network (CNN)</a><br>
<a href='#Pretained'>Using pre-trained models</a><br>
## Installation and setup
<a id='Install'></a>
Since the stable release of TF 1.0, the latest version of slim has been available as `tf.contrib.slim`.
To test that your installation is working, execute the following command; it should run without raising any errors.
```
python -c "import tensorflow.contrib.slim as slim; eval = slim.evaluation.evaluate_once"
```
Although, to use TF-Slim for image classification (as we do in this notebook), you also have to install the TF-Slim image models library from [here](https://github.com/tensorflow/models/tree/master/research/slim). Let's suppose you install this into a directory called TF_MODELS. Then you should change directory to TF_MODELS/research/slim **before** running this notebook, so that these files are in your python path.
To check you've got these two steps to work, just execute the cell below. If it complains about unknown modules, restart the notebook after moving to the TF-Slim models directory.
```
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import matplotlib
%matplotlib inline
import matplotlib.pyplot as plt
import math
import numpy as np
import tensorflow.compat.v1 as tf
import time
from datasets import dataset_utils
# Main slim library
import tf_slim as slim
```
## Creating your first neural network with TF-Slim
<a id='MLP'></a>
Below we give some code to create a simple multilayer perceptron (MLP) which can be used
for regression problems. The model has 2 hidden layers.
The output is a single node.
When this function is called, it will create various nodes, and silently add them to whichever global TF graph is currently in scope. When a node which corresponds to a layer with adjustable parameters (eg., a fully connected layer) is created, additional parameter variable nodes are silently created, and added to the graph. (We will discuss how to train the parameters later.)
We use variable scope to put all the nodes under a common name,
so that the graph has some hierarchical structure.
This is useful when we want to visualize the TF graph in tensorboard, or if we want to query related
variables.
The fully connected layers all use the same L2 weight decay and ReLu activations, as specified by **arg_scope**. (However, the final layer overrides these defaults, and uses an identity activation function.)
We also illustrate how to add a dropout layer after the first fully connected layer (FC1). Note that at test time,
we do not drop out nodes, but instead use the average activations; hence we need to know whether the model is being
constructed for training or testing, since the computational graph will be different in the two cases
(although the variables, storing the model parameters, will be shared, since they have the same name/scope).
```
def regression_model(inputs, is_training=True, scope="deep_regression"):
"""Creates the regression model.
Args:
inputs: A node that yields a `Tensor` of size [batch_size, dimensions].
is_training: Whether or not we're currently training the model.
scope: An optional variable_op scope for the model.
Returns:
predictions: 1-D `Tensor` of shape [batch_size] of responses.
end_points: A dict of end points representing the hidden layers.
"""
with tf.variable_scope(scope, 'deep_regression', [inputs]):
end_points = {}
# Set the default weight _regularizer and acvitation for each fully_connected layer.
with slim.arg_scope([slim.fully_connected],
activation_fn=tf.nn.relu,
weights_regularizer=slim.l2_regularizer(0.01)):
# Creates a fully connected layer from the inputs with 32 hidden units.
net = slim.fully_connected(inputs, 32, scope='fc1')
end_points['fc1'] = net
# Adds a dropout layer to prevent over-fitting.
net = slim.dropout(net, 0.8, is_training=is_training)
# Adds another fully connected layer with 16 hidden units.
net = slim.fully_connected(net, 16, scope='fc2')
end_points['fc2'] = net
# Creates a fully-connected layer with a single hidden unit. Note that the
# layer is made linear by setting activation_fn=None.
predictions = slim.fully_connected(net, 1, activation_fn=None, scope='prediction')
end_points['out'] = predictions
return predictions, end_points
```
### Let's create the model and examine its structure.
We create a TF graph and call regression_model(), which adds nodes (tensors) to the graph. We then examine their shape, and print the names of all the model variables which have been implicitly created inside of each layer. We see that the names of the variables follow the scopes that we specified.
```
with tf.Graph().as_default():
# Dummy placeholders for arbitrary number of 1d inputs and outputs
inputs = tf.placeholder(tf.float32, shape=(None, 1))
outputs = tf.placeholder(tf.float32, shape=(None, 1))
# Build model
predictions, end_points = regression_model(inputs)
# Print name and shape of each tensor.
print("Layers")
for k, v in end_points.items():
print('name = {}, shape = {}'.format(v.name, v.get_shape()))
# Print name and shape of parameter nodes (values not yet initialized)
print("\n")
print("Parameters")
for v in slim.get_model_variables():
print('name = {}, shape = {}'.format(v.name, v.get_shape()))
```
### Let's create some 1d regression data .
We will train and test the model on some noisy observations of a nonlinear function.
```
def produce_batch(batch_size, noise=0.3):
xs = np.random.random(size=[batch_size, 1]) * 10
ys = np.sin(xs) + 5 + np.random.normal(size=[batch_size, 1], scale=noise)
return [xs.astype(np.float32), ys.astype(np.float32)]
x_train, y_train = produce_batch(200)
x_test, y_test = produce_batch(200)
plt.scatter(x_train, y_train)
```
### Let's fit the model to the data
The user has to specify the loss function and the optimizer, and slim does the rest.
In particular, the slim.learning.train function does the following:
- For each iteration, evaluate the train_op, which updates the parameters using the optimizer applied to the current minibatch. Also, update the global_step.
- Occasionally store the model checkpoint in the specified directory. This is useful in case your machine crashes - then you can simply restart from the specified checkpoint.
```
def convert_data_to_tensors(x, y):
inputs = tf.constant(x)
inputs.set_shape([None, 1])
outputs = tf.constant(y)
outputs.set_shape([None, 1])
return inputs, outputs
# The following snippet trains the regression model using a mean_squared_error loss.
ckpt_dir = '/tmp/regression_model/'
with tf.Graph().as_default():
tf.logging.set_verbosity(tf.logging.INFO)
inputs, targets = convert_data_to_tensors(x_train, y_train)
# Make the model.
predictions, nodes = regression_model(inputs, is_training=True)
# Add the loss function to the graph.
loss = tf.losses.mean_squared_error(labels=targets, predictions=predictions)
# The total loss is the user's loss plus any regularization losses.
total_loss = slim.losses.get_total_loss()
# Specify the optimizer and create the train op:
optimizer = tf.train.AdamOptimizer(learning_rate=0.005)
train_op = slim.learning.create_train_op(total_loss, optimizer)
# Run the training inside a session.
final_loss = slim.learning.train(
train_op,
logdir=ckpt_dir,
number_of_steps=5000,
save_summaries_secs=5,
log_every_n_steps=500)
print("Finished training. Last batch loss:", final_loss)
print("Checkpoint saved in %s" % ckpt_dir)
```
### Training with multiple loss functions.
Sometimes we have multiple objectives we want to simultaneously optimize.
In slim, it is easy to add more losses, as we show below. (We do not optimize the total loss in this example,
but we show how to compute it.)
```
with tf.Graph().as_default():
inputs, targets = convert_data_to_tensors(x_train, y_train)
predictions, end_points = regression_model(inputs, is_training=True)
# Add multiple loss nodes.
mean_squared_error_loss = tf.losses.mean_squared_error(labels=targets, predictions=predictions)
absolute_difference_loss = slim.losses.absolute_difference(predictions, targets)
# The following two ways to compute the total loss are equivalent
regularization_loss = tf.add_n(slim.losses.get_regularization_losses())
total_loss1 = mean_squared_error_loss + absolute_difference_loss + regularization_loss
# Regularization Loss is included in the total loss by default.
# This is good for training, but not for testing.
total_loss2 = slim.losses.get_total_loss(add_regularization_losses=True)
init_op = tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(init_op) # Will initialize the parameters with random weights.
total_loss1, total_loss2 = sess.run([total_loss1, total_loss2])
print('Total Loss1: %f' % total_loss1)
print('Total Loss2: %f' % total_loss2)
print('Regularization Losses:')
for loss in slim.losses.get_regularization_losses():
print(loss)
print('Loss Functions:')
for loss in slim.losses.get_losses():
print(loss)
```
### Let's load the saved model and use it for prediction.
```
with tf.Graph().as_default():
inputs, targets = convert_data_to_tensors(x_test, y_test)
# Create the model structure. (Parameters will be loaded below.)
predictions, end_points = regression_model(inputs, is_training=False)
# Make a session which restores the old parameters from a checkpoint.
sv = tf.train.Supervisor(logdir=ckpt_dir)
with sv.managed_session() as sess:
inputs, predictions, targets = sess.run([inputs, predictions, targets])
plt.scatter(inputs, targets, c='r');
plt.scatter(inputs, predictions, c='b');
plt.title('red=true, blue=predicted')
```
### Let's compute various evaluation metrics on the test set.
In TF-Slim termiology, losses are optimized, but metrics (which may not be differentiable, e.g., precision and recall) are just measured. As an illustration, the code below computes mean squared error and mean absolute error metrics on the test set.
Each metric declaration creates several local variables (which must be initialized via tf.initialize_local_variables()) and returns both a value_op and an update_op. When evaluated, the value_op returns the current value of the metric. The update_op loads a new batch of data, runs the model, obtains the predictions and accumulates the metric statistics appropriately before returning the current value of the metric. We store these value nodes and update nodes in 2 dictionaries.
After creating the metric nodes, we can pass them to slim.evaluation.evaluation, which repeatedly evaluates these nodes the specified number of times. (This allows us to compute the evaluation in a streaming fashion across minibatches, which is usefulf for large datasets.) Finally, we print the final value of each metric.
```
with tf.Graph().as_default():
inputs, targets = convert_data_to_tensors(x_test, y_test)
predictions, end_points = regression_model(inputs, is_training=False)
# Specify metrics to evaluate:
names_to_value_nodes, names_to_update_nodes = slim.metrics.aggregate_metric_map({
'Mean Squared Error': slim.metrics.streaming_mean_squared_error(predictions, targets),
'Mean Absolute Error': slim.metrics.streaming_mean_absolute_error(predictions, targets)
})
# Make a session which restores the old graph parameters, and then run eval.
sv = tf.train.Supervisor(logdir=ckpt_dir)
with sv.managed_session() as sess:
metric_values = slim.evaluation.evaluation(
sess,
num_evals=1, # Single pass over data
eval_op=names_to_update_nodes.values(),
final_op=names_to_value_nodes.values())
names_to_values = dict(zip(names_to_value_nodes.keys(), metric_values))
for key, value in names_to_values.items():
print('%s: %f' % (key, value))
```
# Reading Data with TF-Slim
<a id='ReadingTFSlimDatasets'></a>
Reading data with TF-Slim has two main components: A
[Dataset](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/contrib/slim/python/slim/data/dataset.py) and a
[DatasetDataProvider](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/contrib/slim/python/slim/data/dataset_data_provider.py). The former is a descriptor of a dataset, while the latter performs the actions necessary for actually reading the data. Lets look at each one in detail:
## Dataset
A TF-Slim
[Dataset](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/contrib/slim/python/slim/data/dataset.py)
contains descriptive information about a dataset necessary for reading it, such as the list of data files and how to decode them. It also contains metadata including class labels, the size of the train/test splits and descriptions of the tensors that the dataset provides. For example, some datasets contain images with labels. Others augment this data with bounding box annotations, etc. The Dataset object allows us to write generic code using the same API, regardless of the data content and encoding type.
TF-Slim's Dataset works especially well when the data is stored as a (possibly sharded)
[TFRecords file](https://www.tensorflow.org/versions/r0.10/how_tos/reading_data/index.html#file-formats), where each record contains a [tf.train.Example protocol buffer](https://github.com/tensorflow/tensorflow/blob/r0.10/tensorflow/core/example/example.proto).
TF-Slim uses a consistent convention for naming the keys and values inside each Example record.
## DatasetDataProvider
A
[DatasetDataProvider](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/contrib/slim/python/slim/data/dataset_data_provider.py) is a class which actually reads the data from a dataset. It is highly configurable to read the data in various ways that may make a big impact on the efficiency of your training process. For example, it can be single or multi-threaded. If your data is sharded across many files, it can read each files serially, or from every file simultaneously.
## Demo: The Flowers Dataset
For convenience, we've include scripts to convert several common image datasets into TFRecord format and have provided
the Dataset descriptor files necessary for reading them. We demonstrate how easy it is to use these dataset via the Flowers dataset below.
### Download the Flowers Dataset
<a id='DownloadFlowers'></a>
We've made available a tarball of the Flowers dataset which has already been converted to TFRecord format.
```
import tensorflow.compat.v1 as tf
from datasets import dataset_utils
url = "http://download.tensorflow.org/data/flowers.tar.gz"
flowers_data_dir = '/tmp/flowers'
if not tf.gfile.Exists(flowers_data_dir):
tf.gfile.MakeDirs(flowers_data_dir)
dataset_utils.download_and_uncompress_tarball(url, flowers_data_dir)
```
### Display some of the data.
```
from datasets import flowers
import tensorflow.compat.v1 as tf
from tensorflow.contrib import slim
with tf.Graph().as_default():
dataset = flowers.get_split('train', flowers_data_dir)
data_provider = slim.dataset_data_provider.DatasetDataProvider(
dataset, common_queue_capacity=32, common_queue_min=1)
image, label = data_provider.get(['image', 'label'])
with tf.Session() as sess:
with slim.queues.QueueRunners(sess):
for i in range(4):
np_image, np_label = sess.run([image, label])
height, width, _ = np_image.shape
class_name = name = dataset.labels_to_names[np_label]
plt.figure()
plt.imshow(np_image)
plt.title('%s, %d x %d' % (name, height, width))
plt.axis('off')
plt.show()
```
# Convolutional neural nets (CNNs).
<a id='CNN'></a>
In this section, we show how to train an image classifier using a simple CNN.
### Define the model.
Below we define a simple CNN. Note that the output layer is linear function - we will apply softmax transformation externally to the model, either in the loss function (for training), or in the prediction function (during testing).
```
def my_cnn(images, num_classes, is_training): # is_training is not used...
with slim.arg_scope([slim.max_pool2d], kernel_size=[3, 3], stride=2):
net = slim.conv2d(images, 64, [5, 5])
net = slim.max_pool2d(net)
net = slim.conv2d(net, 64, [5, 5])
net = slim.max_pool2d(net)
net = slim.flatten(net)
net = slim.fully_connected(net, 192)
net = slim.fully_connected(net, num_classes, activation_fn=None)
return net
```
### Apply the model to some randomly generated images.
```
import tensorflow as tf
with tf.Graph().as_default():
# The model can handle any input size because the first layer is convolutional.
# The size of the model is determined when image_node is first passed into the my_cnn function.
# Once the variables are initialized, the size of all the weight matrices is fixed.
# Because of the fully connected layers, this means that all subsequent images must have the same
# input size as the first image.
batch_size, height, width, channels = 3, 28, 28, 3
images = tf.random_uniform([batch_size, height, width, channels], maxval=1)
# Create the model.
num_classes = 10
logits = my_cnn(images, num_classes, is_training=True)
probabilities = tf.nn.softmax(logits)
# Initialize all the variables (including parameters) randomly.
init_op = tf.global_variables_initializer()
with tf.Session() as sess:
# Run the init_op, evaluate the model outputs and print the results:
sess.run(init_op)
probabilities = sess.run(probabilities)
print('Probabilities Shape:')
print(probabilities.shape) # batch_size x num_classes
print('\nProbabilities:')
print(probabilities)
print('\nSumming across all classes (Should equal 1):')
print(np.sum(probabilities, 1)) # Each row sums to 1
```
### Train the model on the Flowers dataset.
Before starting, make sure you've run the code to <a href="#DownloadFlowers">Download the Flowers</a> dataset. Now, we'll get a sense of what it looks like to use TF-Slim's training functions found in
[learning.py](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/contrib/slim/python/slim/learning.py). First, we'll create a function, `load_batch`, that loads batches of dataset from a dataset. Next, we'll train a model for a single step (just to demonstrate the API), and evaluate the results.
```
from preprocessing import inception_preprocessing
import tensorflow as tf
from tensorflow.contrib import slim
def load_batch(dataset, batch_size=32, height=299, width=299, is_training=False):
"""Loads a single batch of data.
Args:
dataset: The dataset to load.
batch_size: The number of images in the batch.
height: The size of each image after preprocessing.
width: The size of each image after preprocessing.
is_training: Whether or not we're currently training or evaluating.
Returns:
images: A Tensor of size [batch_size, height, width, 3], image samples that have been preprocessed.
images_raw: A Tensor of size [batch_size, height, width, 3], image samples that can be used for visualization.
labels: A Tensor of size [batch_size], whose values range between 0 and dataset.num_classes.
"""
data_provider = slim.dataset_data_provider.DatasetDataProvider(
dataset, common_queue_capacity=32,
common_queue_min=8)
image_raw, label = data_provider.get(['image', 'label'])
# Preprocess image for usage by Inception.
image = inception_preprocessing.preprocess_image(image_raw, height, width, is_training=is_training)
# Preprocess the image for display purposes.
image_raw = tf.expand_dims(image_raw, 0)
image_raw = tf.image.resize_images(image_raw, [height, width])
image_raw = tf.squeeze(image_raw)
# Batch it up.
images, images_raw, labels = tf.train.batch(
[image, image_raw, label],
batch_size=batch_size,
num_threads=1,
capacity=2 * batch_size)
return images, images_raw, labels
from datasets import flowers
# This might take a few minutes.
train_dir = '/tmp/tfslim_model/'
print('Will save model to %s' % train_dir)
with tf.Graph().as_default():
tf.logging.set_verbosity(tf.logging.INFO)
dataset = flowers.get_split('train', flowers_data_dir)
images, _, labels = load_batch(dataset)
# Create the model:
logits = my_cnn(images, num_classes=dataset.num_classes, is_training=True)
# Specify the loss function:
one_hot_labels = slim.one_hot_encoding(labels, dataset.num_classes)
slim.losses.softmax_cross_entropy(logits, one_hot_labels)
total_loss = slim.losses.get_total_loss()
# Create some summaries to visualize the training process:
tf.summary.scalar('losses/Total Loss', total_loss)
# Specify the optimizer and create the train op:
optimizer = tf.train.AdamOptimizer(learning_rate=0.01)
train_op = slim.learning.create_train_op(total_loss, optimizer)
# Run the training:
final_loss = slim.learning.train(
train_op,
logdir=train_dir,
number_of_steps=1, # For speed, we just do 1 epoch
save_summaries_secs=1)
print('Finished training. Final batch loss %d' % final_loss)
```
### Evaluate some metrics.
As we discussed above, we can compute various metrics besides the loss.
Below we show how to compute prediction accuracy of the trained model, as well as top-5 classification accuracy. (The difference between evaluation and evaluation_loop is that the latter writes the results to a log directory, so they can be viewed in tensorboard.)
```
from datasets import flowers
# This might take a few minutes.
with tf.Graph().as_default():
tf.logging.set_verbosity(tf.logging.DEBUG)
dataset = flowers.get_split('train', flowers_data_dir)
images, _, labels = load_batch(dataset)
logits = my_cnn(images, num_classes=dataset.num_classes, is_training=False)
predictions = tf.argmax(logits, 1)
# Define the metrics:
names_to_values, names_to_updates = slim.metrics.aggregate_metric_map({
'eval/Accuracy': slim.metrics.streaming_accuracy(predictions, labels),
'eval/Recall@5': slim.metrics.streaming_recall_at_k(logits, labels, 5),
})
print('Running evaluation Loop...')
checkpoint_path = tf.train.latest_checkpoint(train_dir)
metric_values = slim.evaluation.evaluate_once(
master='',
checkpoint_path=checkpoint_path,
logdir=train_dir,
eval_op=names_to_updates.values(),
final_op=names_to_values.values())
names_to_values = dict(zip(names_to_values.keys(), metric_values))
for name in names_to_values:
print('%s: %f' % (name, names_to_values[name]))
```
# Using pre-trained models
<a id='Pretrained'></a>
Neural nets work best when they have many parameters, making them very flexible function approximators.
However, this means they must be trained on big datasets. Since this process is slow, we provide various pre-trained models - see the list [here](https://github.com/tensorflow/models/tree/master/research/slim#pre-trained-models).
You can either use these models as-is, or you can perform "surgery" on them, to modify them for some other task. For example, it is common to "chop off" the final pre-softmax layer, and replace it with a new set of weights corresponding to some new set of labels. You can then quickly fine tune the new model on a small new dataset. We illustrate this below, using inception-v1 as the base model. While models like Inception V3 are more powerful, Inception V1 is used for speed purposes.
Take into account that VGG and ResNet final layers have only 1000 outputs rather than 1001. The ImageNet dataset provied has an empty background class which can be used to fine-tune the model to other tasks. VGG and ResNet models provided here don't use that class. We provide two examples of using pretrained models: Inception V1 and VGG-19 models to highlight this difference.
### Download the Inception V1 checkpoint
```
from datasets import dataset_utils
url = "http://download.tensorflow.org/models/inception_v1_2016_08_28.tar.gz"
checkpoints_dir = '/tmp/checkpoints'
if not tf.gfile.Exists(checkpoints_dir):
tf.gfile.MakeDirs(checkpoints_dir)
dataset_utils.download_and_uncompress_tarball(url, checkpoints_dir)
```
### Apply Pre-trained Inception V1 model to Images.
We have to convert each image to the size expected by the model checkpoint.
There is no easy way to determine this size from the checkpoint itself.
So we use a preprocessor to enforce this.
```
import numpy as np
import os
import tensorflow as tf
try:
import urllib2 as urllib
except ImportError:
import urllib.request as urllib
from datasets import imagenet
from nets import inception
from preprocessing import inception_preprocessing
from tensorflow.contrib import slim
image_size = inception.inception_v1.default_image_size
with tf.Graph().as_default():
url = 'https://upload.wikimedia.org/wikipedia/commons/7/70/EnglishCockerSpaniel_simon.jpg'
image_string = urllib.urlopen(url).read()
image = tf.image.decode_jpeg(image_string, channels=3)
processed_image = inception_preprocessing.preprocess_image(image, image_size, image_size, is_training=False)
processed_images = tf.expand_dims(processed_image, 0)
# Create the model, use the default arg scope to configure the batch norm parameters.
with slim.arg_scope(inception.inception_v1_arg_scope()):
logits, _ = inception.inception_v1(processed_images, num_classes=1001, is_training=False)
probabilities = tf.nn.softmax(logits)
init_fn = slim.assign_from_checkpoint_fn(
os.path.join(checkpoints_dir, 'inception_v1.ckpt'),
slim.get_model_variables('InceptionV1'))
with tf.Session() as sess:
init_fn(sess)
np_image, probabilities = sess.run([image, probabilities])
probabilities = probabilities[0, 0:]
sorted_inds = [i[0] for i in sorted(enumerate(-probabilities), key=lambda x:x[1])]
plt.figure()
plt.imshow(np_image.astype(np.uint8))
plt.axis('off')
plt.show()
names = imagenet.create_readable_names_for_imagenet_labels()
for i in range(5):
index = sorted_inds[i]
print('Probability %0.2f%% => [%s]' % (probabilities[index] * 100, names[index]))
```
### Download the VGG-16 checkpoint
```
from datasets import dataset_utils
import tensorflow as tf
url = "http://download.tensorflow.org/models/vgg_16_2016_08_28.tar.gz"
checkpoints_dir = '/tmp/checkpoints'
if not tf.gfile.Exists(checkpoints_dir):
tf.gfile.MakeDirs(checkpoints_dir)
dataset_utils.download_and_uncompress_tarball(url, checkpoints_dir)
```
### Apply Pre-trained VGG-16 model to Images.
We have to convert each image to the size expected by the model checkpoint.
There is no easy way to determine this size from the checkpoint itself.
So we use a preprocessor to enforce this. Pay attention to the difference caused by 1000 classes instead of 1001.
```
import numpy as np
import os
import tensorflow as tf
try:
import urllib2
except ImportError:
import urllib.request as urllib
from datasets import imagenet
from nets import vgg
from preprocessing import vgg_preprocessing
from tensorflow.contrib import slim
image_size = vgg.vgg_16.default_image_size
with tf.Graph().as_default():
url = 'https://upload.wikimedia.org/wikipedia/commons/d/d9/First_Student_IC_school_bus_202076.jpg'
image_string = urllib.urlopen(url).read()
image = tf.image.decode_jpeg(image_string, channels=3)
processed_image = vgg_preprocessing.preprocess_image(image, image_size, image_size, is_training=False)
processed_images = tf.expand_dims(processed_image, 0)
# Create the model, use the default arg scope to configure the batch norm parameters.
with slim.arg_scope(vgg.vgg_arg_scope()):
# 1000 classes instead of 1001.
logits, _ = vgg.vgg_16(processed_images, num_classes=1000, is_training=False)
probabilities = tf.nn.softmax(logits)
init_fn = slim.assign_from_checkpoint_fn(
os.path.join(checkpoints_dir, 'vgg_16.ckpt'),
slim.get_model_variables('vgg_16'))
with tf.Session() as sess:
init_fn(sess)
np_image, probabilities = sess.run([image, probabilities])
probabilities = probabilities[0, 0:]
sorted_inds = [i[0] for i in sorted(enumerate(-probabilities), key=lambda x:x[1])]
plt.figure()
plt.imshow(np_image.astype(np.uint8))
plt.axis('off')
plt.show()
names = imagenet.create_readable_names_for_imagenet_labels()
for i in range(5):
index = sorted_inds[i]
# Shift the index of a class name by one.
print('Probability %0.2f%% => [%s]' % (probabilities[index] * 100, names[index+1]))
```
### Fine-tune the model on a different set of labels.
We will fine tune the inception model on the Flowers dataset.
```
# Note that this may take several minutes.
import os
from datasets import flowers
from nets import inception
from preprocessing import inception_preprocessing
from tensorflow.contrib import slim
image_size = inception.inception_v1.default_image_size
def get_init_fn():
"""Returns a function run by the chief worker to warm-start the training."""
checkpoint_exclude_scopes=["InceptionV1/Logits", "InceptionV1/AuxLogits"]
exclusions = [scope.strip() for scope in checkpoint_exclude_scopes]
variables_to_restore = []
for var in slim.get_model_variables():
for exclusion in exclusions:
if var.op.name.startswith(exclusion):
break
else:
variables_to_restore.append(var)
return slim.assign_from_checkpoint_fn(
os.path.join(checkpoints_dir, 'inception_v1.ckpt'),
variables_to_restore)
train_dir = '/tmp/inception_finetuned/'
with tf.Graph().as_default():
tf.logging.set_verbosity(tf.logging.INFO)
dataset = flowers.get_split('train', flowers_data_dir)
images, _, labels = load_batch(dataset, height=image_size, width=image_size)
# Create the model, use the default arg scope to configure the batch norm parameters.
with slim.arg_scope(inception.inception_v1_arg_scope()):
logits, _ = inception.inception_v1(images, num_classes=dataset.num_classes, is_training=True)
# Specify the loss function:
one_hot_labels = slim.one_hot_encoding(labels, dataset.num_classes)
slim.losses.softmax_cross_entropy(logits, one_hot_labels)
total_loss = slim.losses.get_total_loss()
# Create some summaries to visualize the training process:
tf.summary.scalar('losses/Total Loss', total_loss)
# Specify the optimizer and create the train op:
optimizer = tf.train.AdamOptimizer(learning_rate=0.01)
train_op = slim.learning.create_train_op(total_loss, optimizer)
# Run the training:
final_loss = slim.learning.train(
train_op,
logdir=train_dir,
init_fn=get_init_fn(),
number_of_steps=2)
print('Finished training. Last batch loss %f' % final_loss)
```
### Apply fine tuned model to some images.
```
import numpy as np
import tensorflow as tf
from datasets import flowers
from nets import inception
from tensorflow.contrib import slim
image_size = inception.inception_v1.default_image_size
batch_size = 3
with tf.Graph().as_default():
tf.logging.set_verbosity(tf.logging.INFO)
dataset = flowers.get_split('train', flowers_data_dir)
images, images_raw, labels = load_batch(dataset, height=image_size, width=image_size)
# Create the model, use the default arg scope to configure the batch norm parameters.
with slim.arg_scope(inception.inception_v1_arg_scope()):
logits, _ = inception.inception_v1(images, num_classes=dataset.num_classes, is_training=True)
probabilities = tf.nn.softmax(logits)
checkpoint_path = tf.train.latest_checkpoint(train_dir)
init_fn = slim.assign_from_checkpoint_fn(
checkpoint_path,
slim.get_variables_to_restore())
with tf.Session() as sess:
with slim.queues.QueueRunners(sess):
sess.run(tf.initialize_local_variables())
init_fn(sess)
np_probabilities, np_images_raw, np_labels = sess.run([probabilities, images_raw, labels])
for i in range(batch_size):
image = np_images_raw[i, :, :, :]
true_label = np_labels[i]
predicted_label = np.argmax(np_probabilities[i, :])
predicted_name = dataset.labels_to_names[predicted_label]
true_name = dataset.labels_to_names[true_label]
plt.figure()
plt.imshow(image.astype(np.uint8))
plt.title('Ground Truth: [%s], Prediction [%s]' % (true_name, predicted_name))
plt.axis('off')
plt.show()
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.