text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
```
import gym
import gym_oscillator
import oscillator_cpp
from stable_baselines.common import set_global_seeds
from stable_baselines.common.policies import MlpPolicy,MlpLnLstmPolicy,FeedForwardPolicy
from stable_baselines.common.vec_env import DummyVecEnv,SubprocVecEnv,VecNormalize, VecEnv
from stable_baselines import PPO2
from stable_baselines.common.vec_env import VecEnv
import numpy as np
from matplotlib import pyplot as plt
def make_env(env_id, rank, seed=0,s1=False,s2=False,s3=False,s4=False,s5=False):
"""
Utility function for multiprocessed env.
:param env_id: (str) the environment ID
:param num_env: (int) the number of environment you wish to have in subprocesses
:param seed: (int) the inital seed for RNG
:param rank: (int) index of the subprocess
:param s_i: (bool) reward form, only one can be true
"""
def _init():
env = gym.make(env_id)
env = DummyVecEnv([make_env_dumb(env_id,1)])
model = PPO2(MlpPolicy, env, verbose=1,tensorboard_log="MLP/")
model =model.load('trained_models/Ps6_final_3')
env = gym.make(env_id)
env.__init__(model=model)
return env
set_global_seeds(seed)
return _init
def make_env_dumb(env_id, rank, seed=0,s1=False,s2=False,s3=False,s4=False,s5=False):
"""
Utility function for multiprocessed env.
:param env_id: (str) the environment ID
:param num_env: (int) the number of environment you wish to have in subprocesses
:param seed: (int) the inital seed for RNG
:param rank: (int) index of the subprocess
:param s_i: (bool) reward form, only one can be true
"""
def _init():
env = gym.make(env_id)
env.__init__(model=None)
return env
return _init
#Our env
env_id = 'oscillator-v0'
time_steps = int(10e6)
#Number of cpus
num_cpu = 4
env = SubprocVecEnv([make_env(env_id, i,s2=True) for i in range(num_cpu)])
model = PPO2(MlpPolicy, env, verbose=1,tensorboard_log="MLP/")
# model.learn(time_steps)
# model.save('trained_models/double_extra.tf')
#env.reset()
coupling_power = 0.02
num_cpu = 4
model = model.load('trained_models/Ps6_final_3')
model2 = model.load('trained_models/double.tf')
env = gym.make('oscillator-v0',)
env.__init__(epsilon=coupling_power,model=model,initial_steps=5000,model_steps=5000)
print(env.model)
rews_ = []
obs_ = []
ss_y = env.y_state
obs = env.reset()
acs_ = []
states_x = []
states_y = []
for i in range(20000):
action, _states = model2.predict(obs)
obs, rewards, dones, info = env.step(action)
states_x.append(env.x_val)
states_y.append(env.y_val)
obs_.append(obs[0])
acs_.append(action)
rews_.append(rewards)
#Final relaxation
for i in range(5000):
obs, rewards, dones, info = env.step([0])
states_x.append(env.x_val)
states_y.append(env.y_val)
obs_.append(obs[0])
acs_.append(0)
rews_.append(rewards)
plt.figure(figsize=(25,5))
plt.title('Suppression plot')
plt.xlabel('TimeStep')
plt.ylabel('Signal Value')
plt.plot()
initial_steps = 5000
model_steps = 5000
im = initial_steps+model_steps
plt.plot(np.arange(len(env.x_states[:initial_steps])),env.x_states[:initial_steps])
plt.plot(np.arange(len(env.x_states[initial_steps:im]))+initial_steps,env.x_states[initial_steps:im])
plt.plot(np.arange(len(env.x_states[im:]))+im,env.x_states[im:])
#plt.plot(env.acs_)
plt.figure(figsize=(25,5))
plt.title('Suppression plot')
plt.xlabel('TimeStep')
plt.ylabel('Signal Value')
plt.plot()
initial_steps = 5000
model_steps = 5000
im = initial_steps+model_steps
plt.plot(np.arange(len(env.actions[:initial_steps])),env.actions[:initial_steps])
plt.plot(np.arange(len(env.actions[initial_steps:im]))+initial_steps,env.actions[initial_steps:im])
plt.plot(np.arange(len(env.actions[im:]))+im,env.actions[im:])
#plt.plot(env.acs_)
import pandas as pd
def output_to_csv(states,actions,name='35k_double_chaos.xls'):
print(len(states),len(actions))
output = pd.DataFrame([
states,actions]).T
output.columns=['States_X','Actions']
output.to_excel(name)
return True
output_to_csv(env.x_states,env.actions)
len(env.x_states),len(env.actions)
neurons_output_results = pd.DataFrame([
means_before_all_neurons,
means_after_all_neurons,
stds_before_all_neurons,
stds_after_all_neurons,[100,1000,2000,5000,10000]
]).T
neurons_output_results.columns=['Means_before','Means After','StdsBefore','StdsAfter','NeuronsNumber']
np.std(env.x_states[initial_steps:im])
np.std(env.x_states[im:55000])
q = np.std(env.x_states[:initial_steps])/np.std(env.x_states[initial_steps+5000:im])
print(np.sqrt(q))
l = np.std(env.x_states[:initial_steps])/np.std(env.x_states[im:im+20000])
print(np.sqrt(l))
l/q
im+55000
print(np.std(env.x_states[:2500])/np.std(env.x_states[2500:12500]))
```
| github_jupyter |
```
%load_ext lab_black
import os, sys
%load_ext autoreload
%autoreload 2
import pandas as pd
from os.path import join
import scanpy as sc
import numpy as np
from statsmodels.stats.multitest import multipletests
import matplotlib.pyplot as plt
DATA_PATH = "/n/holystore01/LABS/price_lab/Users/mjzhang/scDRS_data"
df_hom = pd.read_csv(
join(DATA_PATH, "gene_annotation/", "mouse_human_homologs.txt"),
sep="\t",
)
dict_hom = {row[1]: row[0] for _, row in df_hom.iterrows()}
URL_SUPP_TABLE = "https://www.dropbox.com/s/qojbzu5zln33j7f/supp_tables.xlsx?dl=1"
df_trait_info = pd.read_excel(
URL_SUPP_TABLE,
sheet_name=0,
)
list_trait = list(
df_trait_info[df_trait_info.Category == "brain"]["Trait_Identifier"].values
)
list_trait += ["UKB_460K.body_HEIGHTz"]
gs_path = join(DATA_PATH, "gs_file", "magma_10kb_1000.gs")
df_magma_gs = pd.read_csv(gs_path, sep="\t")
df_magma_gs = df_magma_gs[df_magma_gs.TRAIT.isin(list_trait)].reset_index(drop=True)
df_magma_gs["GENESET"] = df_magma_gs["GENESET"].apply(
lambda r: ",".join([dict_hom[g] for g in r.split(",") if g in dict_hom])
)
diff_expr = pd.read_csv("data/GSE67403_gene_exp.diff", sep="\t")
regions = [
"dorsal",
"intermediate",
"ventral",
"proximal",
"distal",
"superficial",
"deep",
]
dict_diff_genes = dict()
FOLD_CHANGE_THRES = 2
diff_expr = diff_expr[
diff_expr.sample_1.isin(regions) & diff_expr.sample_2.isin(regions)
]
diff_expr = diff_expr[
((diff_expr.value_1 > 10) | (diff_expr.value_2 > 10)) & (diff_expr.q_value < 0.05)
]
diff_long = diff_expr[
(diff_expr.sample_1 == "dorsal") & (diff_expr.sample_2 == "ventral")
]
diff_long = diff_long[
(np.abs(diff_long["log2(fold_change)"]) > np.log2(FOLD_CHANGE_THRES))
]
diff_transverse = diff_expr[
(diff_expr.sample_1 == "proximal") & (diff_expr.sample_2 == "distal")
]
diff_transverse = diff_transverse[
(np.abs(diff_transverse["log2(fold_change)"]) > np.log2(FOLD_CHANGE_THRES))
]
diff_radial = diff_expr[
(diff_expr.sample_1 == "superficial") & (diff_expr.sample_2 == "deep")
]
diff_radial = diff_radial[
(np.abs(diff_radial["log2(fold_change)"]) > np.log2(FOLD_CHANGE_THRES))
]
dict_diff_genes[f"ventral"] = diff_long[diff_long.test_stat > 0].gene.values
dict_diff_genes[f"dorsal"] = diff_long[diff_long.test_stat < 0].gene.values
dict_diff_genes[f"distal"] = diff_transverse[diff_transverse.test_stat > 0].gene.values
dict_diff_genes[f"proximal"] = diff_transverse[
diff_transverse.test_stat < 0
].gene.values
dict_diff_genes[f"deep"] = diff_radial[diff_radial.test_stat > 0].gene.values
dict_diff_genes[f"superficial"] = diff_radial[diff_radial.test_stat < 0].gene.values
from os.path import join
df_spatial_gs = {"TRAIT": [], "GENESET": []}
for trait in dict_diff_genes:
df_spatial_gs["TRAIT"].append("spatial_" + trait)
df_spatial_gs["GENESET"].append(
",".join([g for g in dict_diff_genes[trait] if g in dict_hom.values()])
)
df_spatial_gs = pd.DataFrame(df_spatial_gs)
```
# Spatial geneset
```
df_gs = pd.concat([df_magma_gs, df_spatial_gs])
df_mouse_gs = df_gs.copy()
df_mouse_gs.to_csv("gs_file/mouse.gs", sep="\t", index=False)
# mouse to human
dict_hom = {row[0]: row[1] for _, row in df_hom.iterrows()}
df_human_gs = df_mouse_gs.copy()
df_human_gs["GENESET"] = df_mouse_gs["GENESET"].apply(
lambda gs: ",".join([dict_hom[g] for g in gs.split(",")])
)
df_human_gs.to_csv("gs_file/human.gs", sep="\t", index=False)
# divide the gene set into several pieces for parallel submission to the cluster
def divide_gs(df_gs, out_dir, batch_size=1):
batch_dfs = np.array_split(df_gs, int(np.ceil(df_gs.shape[0] / batch_size)))
if os.path.exists(out_dir):
print(f"{out_dir} already exists. Clean up or use another directory")
return
else:
os.makedirs(out_dir)
for batch_i, batch_df in enumerate(batch_dfs):
batch_df.to_csv(join(out_dir, f"batch{batch_i}.gs"), sep="\t", index=False)
divide_gs(df_mouse_gs, "gs_file/mouse.gs.batch")
divide_gs(df_human_gs, "gs_file/human.gs.batch")
```
| github_jupyter |
# Documentation by example for `shap.plots.waterfall`
This notebook is designed to demonstrate (and so document) how to use the `shap.plots.waterfall` function. It uses an XGBoost model trained on the classic UCI adult income dataset (which is classification task to predict if people made over \\$50k in the 90s).
<hr>
<center style="color: red">
<b>Warning!</b> This notebook documents the new SHAP API, and that API is still stablizing over the coming weeks.
</center>
<hr>
```
import xgboost
import shap
# train XGBoost model
X,y = shap.datasets.adult()
model = xgboost.XGBClassifier().fit(X, y)
# compute SHAP values
explainer = shap.Explainer(model, X)
shap_values = explainer(X)
```
Waterfall plots are designed to display explanations for individual predictions, so they expect a single row of an Explanation object as input. The bottom of a waterfall plot starts as the expected value of the model output, and then each row shows how the positive (red) or negative (blue) contribution of each feature moves the value from the expected model output over the background dataset to the model output for this prediction.
Below is an example that plots the first explanation. Note that by default SHAP explains XGBoost classifer models in terms of their margin output, before the logistic link function. That means the units on the x-axis are log-odds units, so negative values imply probabilies of less than 0.5 that the person makes over $50k annually. The gray text before the feature names shows the value of each feature for this sample.
```
shap.plots.waterfall(shap_values[0])
```
Note that in the above explanation the three least impactful features have been collapsed into a single term so that we don't show more than 10 rows in the plot. The default limit of 10 rows can be changed using the `max_display` argument:
```
shap.plots.waterfall(shap_values[0], max_display=20)
```
It is interesting that having a capital gain of \\$2,174 dramatically reduces this person's predicted probability of making over \\$50k annually. Since `waterfall` plots only show a single sample worth of data, we can't see the impact of changing capital gain. To see this we can use a `scatter` plot, which shows how low values for captial gain are a more negative predictor of income that no captial gain at all. Why this happens would require a deeper dive into the data, and should also involve training a model more carefully and with bootstrap resamples to quantify any uncertainty in the model building process.
```
shap.plots.scatter(shap_values[:,"Capital Gain"])
```
<hr>
Have an idea for more helpful examples? Pull requests that add to this documentation notebook are encouraged!
| github_jupyter |
# Data Cleansing
- [Data Understanding](#Data-Understanding)
* Reading in and Exploring Data
* Dealing with Column Names
* Slicing Dataset
- [Cleaning and Exploring Columns](#Cleaning-and-Exploring-Columns)
* [Safety & Security](#Safety-&-Security)
* [Model](#Model)
* [Make](#Make)
* [Model2](#Model2)
* [Model Code](#Model-Code)
* [Country Version](#Country-Version)
* [CO2 Emission](#CO2-Emission)
* [Consumption](#Consumption)
* [Cylinders](#Cylinders)
* [Displacement](#Displacement)
* [Next Inspection](#Next-Inspection)
* [Non-smoking Vehicle](#Non-smoking-Vehicle)
* [Body Type](#Body-Type)
* [Hp](#Hp)
* [Kw](#Kw)
* [Km](#Km)
* [Offer Number](#Offer-Number)
* [Description](#Description)
---
_Additional Columns for Further Investigation_
* [Price](#Price)
* [Registration](#Registration)
## Data Understanding
```
# Import necessay modules
import json
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
# Display 100 columns max
pd.set_option('display.max_columns', 100)
# Initiate directory paths
data_folder = '/Users/stb/Documents/Github/fraud-detection/data/'
```
### Reading in and Exploring Data
```
ads_list = []
with open(data_folder + 'autos_20190626.json','r') as file:
for ad in file:
ad_obj = json.loads(ad)
ads_list.append(ad_obj)
autos = pd.DataFrame(ads_list)
autos.head(3)
autos.info()
```
### Dealing with Column Names
```
autos.columns
# Run "name_columns" function to name the columns in our convention
%run -i "/Users/stb/Documents/Github/fraud-detection/functions/name_columns.py"
```
Alternatively, we can load the file into the cell with the `%load` magic command. (`%load?` for more info)
```
# %load "/Users/stb/Documents/Github/fraud-detection/functions/name_columns.py"
autos.columns = name_columns(autos)
autos.columns
```
### Slicing Dataset
```
t_cols = ['safety_security', 'co2_emission', 'consumption', 'country_version',
'cylinders', 'displacement', 'make', 'model', 'model_code', 'next_inspection',
'non_smoking_vehicle', 'body_type', 'description', "hp", 'kw', 'km', "make_model", "offer_number"]
df = autos[t_cols]
df.info()
```
## Cleaning and Exploring Columns
### Safety & Security
```
# Number of missing values (NaNs) of a column
# print(df['safety_security'].isnull().sum())
# Change the type of NaNs from float to (empty) list
# df['safety&security'][df['safety&security'].isnull()].apply(lambda x: [])
# def NaN_to_list(data, column_name):
# '''
# When dealing with a column which consist of lists, we need to change
# the type of NaNs from 'float' to 'list' in order to perform iterative
# operations. This function detects NaNs and creates an empty list for
# missing rows.
# '''
# # Create a boolean vector for indexing
# NaN_rows = data[column_name].isnull()
# # Change the type of NaNs from 'float' to (empty) 'list'
# data.loc[NaN_rows, column_name] = data.loc[NaN_rows, column_name].apply(lambda x: [])
# NaN_to_list(df, 'safety_security')
# Check to see if there are remaining NaNs
# df['safety_security'].isnull().sum()
# def get_uniques(data, column_name):
# '''
# Get the unique elements from a column
# which consists of list of items.
# '''
# unique_vals = set()
# for row in data[column_name]:
# # Add list of items to a set
# unique_vals.update(row)
# return unique_vals
# uniques = get_uniques(df, 'safety_security')
# len(uniques), uniques
# # Create dummies using the items in the list of 'safety_security' column
# df['safety_security'].str.join('|').str.get_dummies().add_prefix('ss_').head()
# # split(",", expand=True) #hepsini ayri sutun
# Create dummies using the items in the list of 'safety&security' column
df_new = df.join(df['safety_security'].str.join('|').str.get_dummies().add_prefix('ss_'))
# Print the head of the new DataFrame
# df_new.head(3)
# df_new.loc[:, 'ABS':"Xenon headlights"].describe()
```
### Model
```
# df.model.sample(10)
# Count the number of missing values
# df.model.isnull().sum()
# Clean the model column
df_new['model'] = df.model.apply(lambda x: x[1])
# Compare auto models and new columns
# df_new.groupby('model').sum()
# Distribution of car models
# df_new.model.value_counts()
# df_new.model.value_counts().plot.bar(rot=70)
```
### Make
```
# df.make.sample(10)
# Number of missing values in this column
# df.make.isnull().sum()
# Strip "\n"s from the 'make' column
df_new['make'] = df.make.str.strip("\n")
# df_new['make'].sample(10)
# df_new.make.value_counts()
# df_new.groupby(["make", "model"]).size()
# df_new.groupby(["make", "model"]).mean()
# # Add figure size
# plt.figure(figsize=(10,5))
# # Plot heatmap of the table above
# sns.heatmap(df_new.groupby(["make", "model"]).mean(),
# linewidths=.5, cmap="RdYlGn")
```
Initial findings:
- Renault Duster has almost no extras. Interestingly however, all Dusters have ABS, Driver-side airbag and Power steering. The reason might be that Duster is a relatively new model of Renault, and that all the cars have ABS etc. (It is possible that Old Merivas, for example, might not have ABS)
- Almost none of the cars have Blind spot monitor, Emergency System, Night view Assist and Traffic Sign Recognition. This is normal since only luxury segment cars (high end models) have such characteristics.
- Only Audis have Xenon headlights (or high proportion of Audis)
- It is hard to make inference using this graph, since it shows only a portion of car "makes" and "models".
- ...
### Model2
```
# df.make_model.head()
# df.make_model.value_counts()
# df_new.groupby('make').model.value_counts()
# # Create a new Series with Make + Model
# makeModel = df_new.make + " " + df_new.model
# # Check to see whether two Series are the same
# makeModel.equals(df.model2)
# # Check to see whether two Series are the same (alternative way)
# #sum(makeModel != df.model2)
```
The column `make_model` is axactly the same as two columns `make` + `model`. Therefore we can drop the `make_model` column.
```
# Drop unnecesary column 'make_model'
df_new.drop(columns = "make_model", inplace = True)
```
### Model Code
```
# df.model_code.head()
# Proportion of missing values
# df.model_code.isnull().mean()
# Value counts of the 'model_code' column
# df.model_code.apply(lambda x: str(x)[4:-4]).head()
# Clean 'model_code' column
df_new.loc[df_new.model_code.notnull(), "model_code"] = df.model_code[df.model_code.notnull()].apply(lambda x: str(x)[4:-4])
# df_new.model_code.head()
# df_new.groupby(["make", "model"]).model_code.value_counts()
# df_new.groupby("model_code").sum().sample(10)
```
Most of the information in this column is missing. We already have make and model columns, and additionally we have extra information on cars. Thus, this column may not be necessary.
> I suggest dropping `model_code` column.
```
# df_new.drop(columns = "model_code", inplace = True)
```
### Country Version
```
# df.country_version.head()
# Proportion of missing values
# df.country_version.isnull().mean()
# Clean 'country_version' column
df_new.loc[df_new.country_version.notnull(), "country_version"] = df.country_version[df.country_version.notnull()].apply(lambda x: str(x)[4:-4])
# df_new.country_version.value_counts()
# df_new.groupby('make').country_version.value_counts()
```
Almost half of the information in this column is missing. This column may or may not be necessary.
> I suggest dropping `model_code` column if we cannot find a clever way to fill in the missing rows. Another option might be encoding this column as 1's and 0's where 1 represent information and 0 represent no information. By this way, his column may be an indication of -not fraudulent- ads.
```
# df_new.drop(columns = "country_version", inplace = True)
```
### CO2 Emission
```
# df.co2_emission.head()
# Clean 'co2_emission' column
df_new['co2_emission'] = df.co2_emission.str[0].str.extract(r'(\d+)')
# df_new.co2_emission.head()
# Change the 'co2' columns data type to numeric
df_new.co2_emission = pd.to_numeric(df_new.co2_emission)
# df_new.co2_emission.head()
# df_new.groupby(['make']).co2_emission.mean()
# df_new.boxplot("co2_emission", by="make")
# df_new.co2_emission.describe()
# # Calculate IQR for the whole column
# Q1 = df_new.co2_emission.quantile(0.25)
# Q3 = df_new.co2_emission.quantile(0.75)
# IQR = Q3 - Q1
# print(IQR)
# # Calculate the number of outliers
# sum( (df_new.co2_emission < (Q1 - 1.5 * IQR)) | (df_new.co2_emission > (Q3 + 1.5 * IQR)) )
```
There seems to be outliers, which are probably wrong entries. An Opel, for example, has a CO2 emission which is almost 800 g/km. That is impossible and such a level of emission is prohibited by the laws. (https://ec.europa.eu/clima/policies/transport/vehicles/cars_en)
We need to take care of outliers and missing values in this column. A possible solution is to take the average of each car **model**, and assign the mean to the missing/wrong entries for that model of car.
----
### Consumption
```
# df.consumption.sample(10)
# Percentage of NaNs
# autos.consumption.isnull().mean()
NaN_to_list(df, "consumption")
# Check to see how many elements each list have
df.consumption.apply(lambda x: len(x)).value_counts()
```
I expected to see at most 3 elements in a list. Interestingly, however, there are many rows with 7 and 5 elements. The reason is that, lists in those row consist of `\n`s along with consumption values (see below).
```
df.consumption[df.consumption.apply(lambda x: (len(x)== 7) | (len(x)== 5))].head()
df.consumption[0]
df.consumption.str[0].head(10)
df.consumption.str[0].str[0].head()
"comb" in str(df.consumption[0][0])
# Create a boolean for checking "comb"
comb_bool = df.consumption.str[0].str[0].str.contains("comb", na=False)
# Create a new column for 'consumption_comb'
df_new['consumption_comb'] = df[comb_bool].consumption.str[0].str[0].str.extract(r'(\d.\d|\d)')
```
### Cylinders
```
# df.cylinders.sample(10)
# Percentage of missing values
# df.cylinders.isnull().mean()
# Clean 'cylinders' column
df_new['cylinders'] = df.cylinders.str[0].str.extract(r'(\d+)')
# Change the 'cylinders' columns data type to numeric
df_new['cylinders'] = pd.to_numeric(df_new['cylinders'])
# df_new.cylinders.sample(10)
# df_new.cylinders.value_counts()
```
The values appear to be normal. We need to take care of missing values in this column though. A possible solution is to use some necessary columns such as `model` to fit a *regression* model, and to predict those values.
### Displacement
```
# df.displacement.sample(10)
# df.displacement.isnull().mean()
# df.displacement.str[0].str.replace(",","").str.extract(r'(\d+)').head()
# Extract discplacement values (and remove commas)
df_new['displacement'] = df.displacement.str[0].str.replace(",","").str.extract(r'(\d+)')
# Change the type
df_new['displacement'] = pd.to_numeric(df_new['displacement'])
# df_new['displacement'].sample(10)
# df_new.groupby(["make","model"]).displacement.mean().plot.bar(rot=75)
# df_new.displacement.describe()
# df_new.displacement.plot.box()
```
When we look at the outliers, they seem to be problematic (See below). We may need to threat them as missing and perform imputation.
```
# df_new[df_new.displacement > 3500]
```
### Next Inspection
```
# df.next_inspection.sample(10)
```
The rows are mixed with values from other columns: `CO2 Emission`, `Consumption` etc.
```
# Extract 'next_inspection' values
df_new.next_inspection = df.next_inspection.str[0].str.strip("\n")
# df_new.next_inspection.head()
# Create a boolean column from `next_inspection`
df_new['next_inspection_bool'] = df_new.next_inspection.notnull()
# Percentage of missing values
# df_new.next_inspection.isnull().mean()
```
There are so many missing values. Although this column is important, we may need to drop it. Yet, there might be information in other columns, especially in `description` column, regarding the _Next Inspection_.
### Non-smoking Vehicle
```
# df['non_smoking_vehicle'].sample(20)
# autos.iloc[10792].url
# df['non_smoking_vehicle'].isnull().mean()
# Drop 'non-smoking_vehicle' column
df_new.drop("non_smoking_vehicle", axis=1, inplace=True)
```
The information in this column is wrognly coded, and more than half is missing. We may drop this column. (" " means yes(True))
### Body Type
```
# df.body_type.head()
# df.body_type.value_counts()
```
### Hp
```
# df.hp.head()
# Extract hp from 'hp' column
df_new['hp'] = df.hp.str.extract(r'(\d+)')
# Change datatype to numeric
df_new['hp'] = pd.to_numeric(df_new['hp'])
```
### Kw
```
# df.kw.isnull().mean()
# Drop 'kw' column
df_new.drop('kw', axis=1, inplace=True)
```
### Km
```
# df.km.head()
# Clean 'km' column
df_new['km'] = df.km.str.replace(",", "").str.extract(r'(\d+)')
```
### Offer Number
```
df.offer_number.head()
# Clean "offer_number' column
df_new['offer_number'] = df.offer_number.str[0].str.replace("\n","")
```
### Description
```
df.description.head()
df.description[0]
df_new['description'] = df.description.apply(lambda x: str(x).strip("['\\n', ").replace("' ', ", ''))
df_new['description'][0]
df['description'].str.join('').str.strip("\n")[0]
```
----
### Price
```
autos.price.head()
autos.price.isnull().sum()
df_new['price'] = autos.price.apply(lambda x: x.strip("\n").strip("€ ").strip(".-").replace(",",""))
df_new.price.head()
df_new['price'] = pd.to_numeric(df_new.price)
```
### Registration
```
autos.registration.head()
# Extract years (first item) from the list
reg = autos.registration.apply(lambda x: x[0])
reg.head()
# Count the number of missing values
(reg.str.count("\d") == 0).sum()
# Count the number of missing values (alternative way)
# (reg.str.count("\d") == 0).sum()
# Count the number of missing values
reg.str.contains("-").sum()
reg_new = reg[~reg.str.contains("-")]
reg_new = pd.to_datetime(reg_new, format='%m/%Y')
reg_new.head()
reg_year = reg_new.apply(lambda x: x.year)
df_new['age'] = 2019 - reg_year
df_new.groupby("model").age.mean()
```
> There seems to be no ad for an older Duster
```
df_new.groupby("model").age.describe()
reg_year.plot.hist()
# df_new.age.plot.hist()
```
> There is no car older than 5 years. The data seems to be problematic.
```
df_new.groupby("model").age.max()
# Extract nr of "Previous owners"
autos.registration.apply(lambda x: x[1][0] if len(x) > 1 else np.nan).head()
```
| github_jupyter |
# Convolutional Neural Networks: Step by Step
Welcome to Course 4's first assignment! In this assignment, you will implement convolutional (CONV) and pooling (POOL) layers in numpy, including both forward propagation and (optionally) backward propagation.
**Notation**:
- Superscript $[l]$ denotes an object of the $l^{th}$ layer.
- Example: $a^{[4]}$ is the $4^{th}$ layer activation. $W^{[5]}$ and $b^{[5]}$ are the $5^{th}$ layer parameters.
- Superscript $(i)$ denotes an object from the $i^{th}$ example.
- Example: $x^{(i)}$ is the $i^{th}$ training example input.
- Lowerscript $i$ denotes the $i^{th}$ entry of a vector.
- Example: $a^{[l]}_i$ denotes the $i^{th}$ entry of the activations in layer $l$, assuming this is a fully connected (FC) layer.
- $n_H$, $n_W$ and $n_C$ denote respectively the height, width and number of channels of a given layer. If you want to reference a specific layer $l$, you can also write $n_H^{[l]}$, $n_W^{[l]}$, $n_C^{[l]}$.
- $n_{H_{prev}}$, $n_{W_{prev}}$ and $n_{C_{prev}}$ denote respectively the height, width and number of channels of the previous layer. If referencing a specific layer $l$, this could also be denoted $n_H^{[l-1]}$, $n_W^{[l-1]}$, $n_C^{[l-1]}$.
We assume that you are already familiar with `numpy` and/or have completed the previous courses of the specialization. Let's get started!
## 1 - Packages
Let's first import all the packages that you will need during this assignment.
- [numpy](www.numpy.org) is the fundamental package for scientific computing with Python.
- [matplotlib](http://matplotlib.org) is a library to plot graphs in Python.
- np.random.seed(1) is used to keep all the random function calls consistent. It will help us grade your work.
```
import numpy as np
import h5py
import matplotlib.pyplot as plt
%matplotlib inline
plt.rcParams['figure.figsize'] = (5.0, 4.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
%load_ext autoreload
%autoreload 2
np.random.seed(1)
```
## 2 - Outline of the Assignment
You will be implementing the building blocks of a convolutional neural network! Each function you will implement will have detailed instructions that will walk you through the steps needed:
- Convolution functions, including:
- Zero Padding
- Convolve window
- Convolution forward
- Convolution backward (optional)
- Pooling functions, including:
- Pooling forward
- Create mask
- Distribute value
- Pooling backward (optional)
This notebook will ask you to implement these functions from scratch in `numpy`. In the next notebook, you will use the TensorFlow equivalents of these functions to build the following model:
<img src="images/model.png" style="width:800px;height:300px;">
**Note** that for every forward function, there is its corresponding backward equivalent. Hence, at every step of your forward module you will store some parameters in a cache. These parameters are used to compute gradients during backpropagation.
## 3 - Convolutional Neural Networks
Although programming frameworks make convolutions easy to use, they remain one of the hardest concepts to understand in Deep Learning. A convolution layer transforms an input volume into an output volume of different size, as shown below.
<img src="images/conv_nn.png" style="width:350px;height:200px;">
In this part, you will build every step of the convolution layer. You will first implement two helper functions: one for zero padding and the other for computing the convolution function itself.
### 3.1 - Zero-Padding
Zero-padding adds zeros around the border of an image:
<img src="images/PAD.png" style="width:600px;height:400px;">
<caption><center> <u> <font color='purple'> **Figure 1** </u><font color='purple'> : **Zero-Padding**<br> Image (3 channels, RGB) with a padding of 2. </center></caption>
The main benefits of padding are the following:
- It allows you to use a CONV layer without necessarily shrinking the height and width of the volumes. This is important for building deeper networks, since otherwise the height/width would shrink as you go to deeper layers. An important special case is the "same" convolution, in which the height/width is exactly preserved after one layer.
- It helps us keep more of the information at the border of an image. Without padding, very few values at the next layer would be affected by pixels as the edges of an image.
**Exercise**: Implement the following function, which pads all the images of a batch of examples X with zeros. [Use np.pad](https://docs.scipy.org/doc/numpy/reference/generated/numpy.pad.html). Note if you want to pad the array "a" of shape $(5,5,5,5,5)$ with `pad = 1` for the 2nd dimension, `pad = 3` for the 4th dimension and `pad = 0` for the rest, you would do:
```python
a = np.pad(a, ((0,0), (1,1), (0,0), (3,3), (0,0)), 'constant', constant_values = (..,..))
```
```
# GRADED FUNCTION: zero_pad
def zero_pad(X, pad):
"""
Pad with zeros all images of the dataset X. The padding is applied to the height and width of an image,
as illustrated in Figure 1.
Argument:
X -- python numpy array of shape (m, n_H, n_W, n_C) representing a batch of m images
pad -- integer, amount of padding around each image on vertical and horizontal dimensions
Returns:
X_pad -- padded image of shape (m, n_H + 2*pad, n_W + 2*pad, n_C)
"""
### START CODE HERE ### (≈ 1 line)
X_pad = np.pad(X, ((0,0), (pad,pad), (pad,pad), (0,0)), 'constant')
### END CODE HERE ###
return X_pad
np.random.seed(1)
x = np.random.randn(4, 3, 3, 2)
x_pad = zero_pad(x, 2)
print ("x.shape =", x.shape)
print ("x_pad.shape =", x_pad.shape)
print ("x[1,1] =", x[1,1])
print ("x_pad[1,1] =", x_pad[1,1])
fig, axarr = plt.subplots(1, 2)
axarr[0].set_title('x')
axarr[0].imshow(x[0,:,:,0])
axarr[1].set_title('x_pad')
axarr[1].imshow(x_pad[0,:,:,0])
```
**Expected Output**:
<table>
<tr>
<td>
**x.shape**:
</td>
<td>
(4, 3, 3, 2)
</td>
</tr>
<tr>
<td>
**x_pad.shape**:
</td>
<td>
(4, 7, 7, 2)
</td>
</tr>
<tr>
<td>
**x[1,1]**:
</td>
<td>
[[ 0.90085595 -0.68372786]
[-0.12289023 -0.93576943]
[-0.26788808 0.53035547]]
</td>
</tr>
<tr>
<td>
**x_pad[1,1]**:
</td>
<td>
[[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]]
</td>
</tr>
</table>
### 3.2 - Single step of convolution
In this part, implement a single step of convolution, in which you apply the filter to a single position of the input. This will be used to build a convolutional unit, which:
- Takes an input volume
- Applies a filter at every position of the input
- Outputs another volume (usually of different size)
<img src="images/Convolution_schematic.gif" style="width:500px;height:300px;">
<caption><center> <u> <font color='purple'> **Figure 2** </u><font color='purple'> : **Convolution operation**<br> with a filter of 2x2 and a stride of 1 (stride = amount you move the window each time you slide) </center></caption>
In a computer vision application, each value in the matrix on the left corresponds to a single pixel value, and we convolve a 3x3 filter with the image by multiplying its values element-wise with the original matrix, then summing them up. In this first step of the exercise, you will implement a single step of convolution, corresponding to applying a filter to just one of the positions to get a single real-valued output.
Later in this notebook, you'll apply this function to multiple positions of the input to implement the full convolutional operation.
**Exercise**: Implement conv_single_step(). [Hint](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.sum.html).
```
# GRADED FUNCTION: conv_single_step
def conv_single_step(a_slice_prev, W, b):
"""
Apply one filter defined by parameters W on a single slice (a_slice_prev) of the output activation
of the previous layer.
Arguments:
a_slice_prev -- slice of input data of shape (f, f, n_C_prev)
W -- Weight parameters contained in a window - matrix of shape (f, f, n_C_prev)
b -- Bias parameters contained in a window - matrix of shape (1, 1, 1)
Returns:
Z -- a scalar value, result of convolving the sliding window (W, b) on a slice x of the input data
"""
### START CODE HERE ### (≈ 2 lines of code)
# Element-wise product between a_slice and W. Add bias.
#s = np.array(W) * np.array(a_slice_prev) + np.array(b)
s = np.multiply( np.array(W), np.array(a_slice_prev)) + np.array(b)
# Sum over all entries of the volume s
Z = np.sum(s)
### END CODE HERE ###
return Z
np.random.seed(1)
a_slice_prev = np.random.randn(4, 4, 3)
W = np.random.randn(4, 4, 3)
b = np.random.randn(1, 1, 1)
Z = conv_single_step(a_slice_prev, W, b)
print("Z =", Z)
```
**Expected Output**:
<table>
<tr>
<td>
**Z**
</td>
<td>
-23.1602122025
</td>
</tr>
</table>
### 3.3 - Convolutional Neural Networks - Forward pass
In the forward pass, you will take many filters and convolve them on the input. Each 'convolution' gives you a 2D matrix output. You will then stack these outputs to get a 3D volume:
<center>
<video width="620" height="440" src="images/conv_kiank.mp4" type="video/mp4" controls>
</video>
</center>
**Exercise**: Implement the function below to convolve the filters W on an input activation A_prev. This function takes as input A_prev, the activations output by the previous layer (for a batch of m inputs), F filters/weights denoted by W, and a bias vector denoted by b, where each filter has its own (single) bias. Finally you also have access to the hyperparameters dictionary which contains the stride and the padding.
**Hint**:
1. To select a 2x2 slice at the upper left corner of a matrix "a_prev" (shape (5,5,3)), you would do:
```python
a_slice_prev = a_prev[0:2,0:2,:]
```
This will be useful when you will define `a_slice_prev` below, using the `start/end` indexes you will define.
2. To define a_slice you will need to first define its corners `vert_start`, `vert_end`, `horiz_start` and `horiz_end`. This figure may be helpful for you to find how each of the corner can be defined using h, w, f and s in the code below.
<img src="images/vert_horiz_kiank.png" style="width:400px;height:300px;">
<caption><center> <u> <font color='purple'> **Figure 3** </u><font color='purple'> : **Definition of a slice using vertical and horizontal start/end (with a 2x2 filter)** <br> This figure shows only a single channel. </center></caption>
**Reminder**:
The formulas relating the output shape of the convolution to the input shape is:
$$ n_H = \lfloor \frac{n_{H_{prev}} - f + 2 \times pad}{stride} \rfloor +1 $$
$$ n_W = \lfloor \frac{n_{W_{prev}} - f + 2 \times pad}{stride} \rfloor +1 $$
$$ n_C = \text{number of filters used in the convolution}$$
For this exercise, we won't worry about vectorization, and will just implement everything with for-loops.
```
# GRADED FUNCTION: conv_forward
def conv_forward(A_prev, W, b, hparameters):
"""
Implements the forward propagation for a convolution function
Arguments:
A_prev -- output activations of the previous layer, numpy array of shape (m, n_H_prev, n_W_prev, n_C_prev)
W -- Weights, numpy array of shape (f, f, n_C_prev, n_C)
b -- Biases, numpy array of shape (1, 1, 1, n_C)
hparameters -- python dictionary containing "stride" and "pad"
Returns:
Z -- conv output, numpy array of shape (m, n_H, n_W, n_C)
cache -- cache of values needed for the conv_backward() function
"""
### START CODE HERE ###
# Retrieve dimensions from A_prev's shape (≈1 line)
(m, n_H_prev, n_W_prev, n_C_prev) = A_prev.shape
# Retrieve dimensions from W's shape (≈1 line)
(f, f, n_C_prev, n_C) = W.shape
# Retrieve information from "hparameters" (≈2 lines)
stride = hparameters["stride"]
pad = hparameters["pad"]
# Compute the dimensions of the CONV output volume using the formula given above. Hint: use int() to floor. (≈2 lines)
n_H = int((n_H_prev - f + 2*pad)/stride + 1)
n_W = int((n_W_prev - f + 2*pad)/stride + 1)
# Initialize the output volume Z with zeros. (≈1 line)
Z = np.zeros(shape=(m, n_H, n_W, n_C))
# Create A_prev_pad by padding A_prev
A_prev_pad = zero_pad(A_prev, pad)
for i in range(m): # loop over the batch of training examples
a_prev_pad = A_prev_pad[i] # Select ith training example's padded activation
for h in range(n_H): # loop over vertical axis of the output volume
for w in range(n_W): # loop over horizontal axis of the output volume
for c in range(n_C): # loop over channels (= #filters) of the output volume
# Find the corners of the current "slice" (≈4 lines)
vert_start = h
vert_end = vert_start + f
horiz_start = w
horiz_end = horiz_start + f
# Use the corners to define the (3D) slice of a_prev_pad (See Hint above the cell). (≈1 line)
a_slice_prev = a_prev_pad[vert_start:vert_end,horiz_start:horiz_end,:]
# Convolve the (3D) slice with the correct filter W and bias b, to get back one output neuron. (≈1 line)
Z[i, h, w, c] = conv_single_step(a_slice_prev, W[:,:,:,c], b[:,:,:,c])
### END CODE HERE ###
# Making sure your output shape is correct
assert(Z.shape == (m, n_H, n_W, n_C))
# Save information in "cache" for the backprop
cache = (A_prev, W, b, hparameters)
return Z, cache
np.random.seed(1)
A_prev = np.random.randn(10,4,4,3)
W = np.random.randn(2,2,3,8)
b = np.random.randn(1,1,1,8)
hparameters = {"pad" : 2,
"stride": 1}
Z, cache_conv = conv_forward(A_prev, W, b, hparameters)
print("Z's mean =", np.mean(Z))
print("cache_conv[0][1][2][3] =", cache_conv[0][1][2][3])
```
**Expected Output**:
<table>
<tr>
<td>
**Z's mean**
</td>
<td>
0.155859324889
</td>
</tr>
<tr>
<td>
**cache_conv[0][1][2][3]**
</td>
<td>
[-0.20075807 0.18656139 0.41005165]
</td>
</tr>
</table>
Finally, CONV layer should also contain an activation, in which case we would add the following line of code:
```python
# Convolve the window to get back one output neuron
Z[i, h, w, c] = ...
# Apply activation
A[i, h, w, c] = activation(Z[i, h, w, c])
```
You don't need to do it here.
## 4 - Pooling layer
The pooling (POOL) layer reduces the height and width of the input. It helps reduce computation, as well as helps make feature detectors more invariant to its position in the input. The two types of pooling layers are:
- Max-pooling layer: slides an ($f, f$) window over the input and stores the max value of the window in the output.
- Average-pooling layer: slides an ($f, f$) window over the input and stores the average value of the window in the output.
<table>
<td>
<img src="images/max_pool1.png" style="width:500px;height:300px;">
<td>
<td>
<img src="images/a_pool.png" style="width:500px;height:300px;">
<td>
</table>
These pooling layers have no parameters for backpropagation to train. However, they have hyperparameters such as the window size $f$. This specifies the height and width of the fxf window you would compute a max or average over.
### 4.1 - Forward Pooling
Now, you are going to implement MAX-POOL and AVG-POOL, in the same function.
**Exercise**: Implement the forward pass of the pooling layer. Follow the hints in the comments below.
**Reminder**:
As there's no padding, the formulas binding the output shape of the pooling to the input shape is:
$$ n_H = \lfloor \frac{n_{H_{prev}} - f}{stride} \rfloor +1 $$
$$ n_W = \lfloor \frac{n_{W_{prev}} - f}{stride} \rfloor +1 $$
$$ n_C = n_{C_{prev}}$$
```
# GRADED FUNCTION: pool_forward
def pool_forward(A_prev, hparameters, mode = "max"):
"""
Implements the forward pass of the pooling layer
Arguments:
A_prev -- Input data, numpy array of shape (m, n_H_prev, n_W_prev, n_C_prev)
hparameters -- python dictionary containing "f" and "stride"
mode -- the pooling mode you would like to use, defined as a string ("max" or "average")
Returns:
A -- output of the pool layer, a numpy array of shape (m, n_H, n_W, n_C)
cache -- cache used in the backward pass of the pooling layer, contains the input and hparameters
"""
# Retrieve dimensions from the input shape
(m, n_H_prev, n_W_prev, n_C_prev) = A_prev.shape
# Retrieve hyperparameters from "hparameters"
f = hparameters["f"]
stride = hparameters["stride"]
# Define the dimensions of the output
n_H = int(1 + (n_H_prev - f) / stride)
n_W = int(1 + (n_W_prev - f) / stride)
n_C = n_C_prev
# Initialize output matrix A
A = np.zeros((m, n_H, n_W, n_C))
### START CODE HERE ###
for i in range(m): # loop over the training examples
for h in range(n_H): # loop on the vertical axis of the output volume
for w in range(n_W): # loop on the horizontal axis of the output volume
for c in range (n_C): # loop over the channels of the output volume
# Find the corners of the current "slice" (≈4 lines)
vert_start = h
vert_end = h + f
horiz_start = w
horiz_end = w + f
# Use the corners to define the current slice on the ith training example of A_prev, channel c. (≈1 line)
#a_prev_slice = A_prev[i][horiz_start:horiz_end,vert_start:vert_end,c]
#a_prev_slice = A_prev[i][vert_start:vert_end, horiz_start:horiz_end,c]
a_prev_slice = A_prev[i, vert_start: vert_end, horiz_start: horiz_end, c]
# Compute the pooling operation on the slice. Use an if statment to differentiate the modes. Use np.max/np.mean.
if mode == "max":
A[i, h, w, c] = np.max(a_prev_slice)
elif mode == "average":
A[i, h, w, c] = np.mean(a_prev_slice)
### END CODE HERE ###
# Store the input and hparameters in "cache" for pool_backward()
cache = (A_prev, hparameters)
# Making sure your output shape is correct
assert(A.shape == (m, n_H, n_W, n_C))
return A, cache
np.random.seed(1)
A_prev = np.random.randn(2, 4, 4, 3)
hparameters = {"stride" : 1, "f": 4}
A, cache = pool_forward(A_prev, hparameters)
print("mode = max")
print("A =", A)
print()
A, cache = pool_forward(A_prev, hparameters, mode = "average")
print("mode = average")
print("A =", A)
```
**Expected Output:**
<table>
<tr>
<td>
A =
</td>
<td>
[[[[ 1.74481176 1.6924546 2.10025514]]] <br/>
[[[ 1.19891788 1.51981682 2.18557541]]]]
</td>
</tr>
<tr>
<td>
A =
</td>
<td>
[[[[-0.09498456 0.11180064 -0.14263511]]] <br/>
[[[-0.09525108 0.28325018 0.33035185]]]]
</td>
</tr>
</table>
Congratulations! You have now implemented the forward passes of all the layers of a convolutional network.
The remainer of this notebook is optional, and will not be graded.
## 5 - Backpropagation in convolutional neural networks (OPTIONAL / UNGRADED)
In modern deep learning frameworks, you only have to implement the forward pass, and the framework takes care of the backward pass, so most deep learning engineers don't need to bother with the details of the backward pass. The backward pass for convolutional networks is complicated. If you wish however, you can work through this optional portion of the notebook to get a sense of what backprop in a convolutional network looks like.
When in an earlier course you implemented a simple (fully connected) neural network, you used backpropagation to compute the derivatives with respect to the cost to update the parameters. Similarly, in convolutional neural networks you can to calculate the derivatives with respect to the cost in order to update the parameters. The backprop equations are not trivial and we did not derive them in lecture, but we briefly presented them below.
### 5.1 - Convolutional layer backward pass
Let's start by implementing the backward pass for a CONV layer.
#### 5.1.1 - Computing dA:
This is the formula for computing $dA$ with respect to the cost for a certain filter $W_c$ and a given training example:
$$ dA += \sum _{h=0} ^{n_H} \sum_{w=0} ^{n_W} W_c \times dZ_{hw} \tag{1}$$
Where $W_c$ is a filter and $dZ_{hw}$ is a scalar corresponding to the gradient of the cost with respect to the output of the conv layer Z at the hth row and wth column (corresponding to the dot product taken at the ith stride left and jth stride down). Note that at each time, we multiply the the same filter $W_c$ by a different dZ when updating dA. We do so mainly because when computing the forward propagation, each filter is dotted and summed by a different a_slice. Therefore when computing the backprop for dA, we are just adding the gradients of all the a_slices.
In code, inside the appropriate for-loops, this formula translates into:
```python
da_prev_pad[vert_start:vert_end, horiz_start:horiz_end, :] += W[:,:,:,c] * dZ[i, h, w, c]
```
#### 5.1.2 - Computing dW:
This is the formula for computing $dW_c$ ($dW_c$ is the derivative of one filter) with respect to the loss:
$$ dW_c += \sum _{h=0} ^{n_H} \sum_{w=0} ^ {n_W} a_{slice} \times dZ_{hw} \tag{2}$$
Where $a_{slice}$ corresponds to the slice which was used to generate the acitivation $Z_{ij}$. Hence, this ends up giving us the gradient for $W$ with respect to that slice. Since it is the same $W$, we will just add up all such gradients to get $dW$.
In code, inside the appropriate for-loops, this formula translates into:
```python
dW[:,:,:,c] += a_slice * dZ[i, h, w, c]
```
#### 5.1.3 - Computing db:
This is the formula for computing $db$ with respect to the cost for a certain filter $W_c$:
$$ db = \sum_h \sum_w dZ_{hw} \tag{3}$$
As you have previously seen in basic neural networks, db is computed by summing $dZ$. In this case, you are just summing over all the gradients of the conv output (Z) with respect to the cost.
In code, inside the appropriate for-loops, this formula translates into:
```python
db[:,:,:,c] += dZ[i, h, w, c]
```
**Exercise**: Implement the `conv_backward` function below. You should sum over all the training examples, filters, heights, and widths. You should then compute the derivatives using formulas 1, 2 and 3 above.
```
def conv_backward(dZ, cache):
"""
Implement the backward propagation for a convolution function
Arguments:
dZ -- gradient of the cost with respect to the output of the conv layer (Z), numpy array of shape (m, n_H, n_W, n_C)
cache -- cache of values needed for the conv_backward(), output of conv_forward()
Returns:
dA_prev -- gradient of the cost with respect to the input of the conv layer (A_prev),
numpy array of shape (m, n_H_prev, n_W_prev, n_C_prev)
dW -- gradient of the cost with respect to the weights of the conv layer (W)
numpy array of shape (f, f, n_C_prev, n_C)
db -- gradient of the cost with respect to the biases of the conv layer (b)
numpy array of shape (1, 1, 1, n_C)
"""
### START CODE HERE ###
# Retrieve information from "cache"
(A_prev, W, b, hparameters) = cache
# Retrieve dimensions from A_prev's shape
(m, n_H_prev, n_W_prev, n_C_prev) = A_prev.shape
# Retrieve dimensions from W's shape
(f, f, n_C_prev, n_C) = W.shape
# Retrieve information from "hparameters"
stride = hparameters['stride']
pad = hparameters['pad']
# Retrieve dimensions from dZ's shape
(m, n_H, n_W, n_C) = dZ.shape
# Initialize dA_prev, dW, db with the correct shapes
dA_prev = np.zeros(shape=(m, n_H_prev, n_W_prev, n_C_prev))
dW = np.zeros(shape=(f, f, n_C_prev, n_C))
db = np.zeros(shape=(1, 1, 1, n_C))
# Pad A_prev and dA_prev
A_prev_pad = zero_pad(A_prev, pad)
dA_prev_pad = zero_pad(dA_prev, pad)
for i in range(m): # loop over the training examples
# select ith training example from A_prev_pad and dA_prev_pad
a_prev_pad = A_prev_pad[i]
da_prev_pad = dA_prev_pad[i]
for h in range(n_H): # loop over vertical axis of the output volume
for w in range(n_W): # loop over horizontal axis of the output volume
for c in range(n_C): # loop over the channels of the output volume
# Find the corners of the current "slice"
vert_start = h
vert_end = vert_start + f
horiz_start = w
horiz_end = horiz_start + f
# Use the corners to define the slice from a_prev_pad
a_slice = a_prev_pad[vert_start:vert_end,horiz_start:horiz_end,:]
# Update gradients for the window and the filter's parameters using the code formulas given above
da_prev_pad[vert_start:vert_end, horiz_start:horiz_end, :] += W[:,:,:,c] * dZ[i, h, w, c]
dW[:,:,:,c] += a_slice * dZ[i, h, w, c]
db[:,:,:,c] += dZ[i, h, w, c]
# Set the ith training example's dA_prev to the unpaded da_prev_pad (Hint: use X[pad:-pad, pad:-pad, :])
dA_prev[i, :, :, :] = da_prev_pad[pad:-pad, pad:-pad, :]
### END CODE HERE ###
# Making sure your output shape is correct
assert(dA_prev.shape == (m, n_H_prev, n_W_prev, n_C_prev))
return dA_prev, dW, db
np.random.seed(1)
dA, dW, db = conv_backward(Z, cache_conv)
print("dA_mean =", np.mean(dA))
print("dW_mean =", np.mean(dW))
print("db_mean =", np.mean(db))
```
** Expected Output: **
<table>
<tr>
<td>
**dA_mean**
</td>
<td>
9.60899067587
</td>
</tr>
<tr>
<td>
**dW_mean**
</td>
<td>
10.5817412755
</td>
</tr>
<tr>
<td>
**db_mean**
</td>
<td>
76.3710691956
</td>
</tr>
</table>
## 5.2 Pooling layer - backward pass
Next, let's implement the backward pass for the pooling layer, starting with the MAX-POOL layer. Even though a pooling layer has no parameters for backprop to update, you still need to backpropagation the gradient through the pooling layer in order to compute gradients for layers that came before the pooling layer.
### 5.2.1 Max pooling - backward pass
Before jumping into the backpropagation of the pooling layer, you are going to build a helper function called `create_mask_from_window()` which does the following:
$$ X = \begin{bmatrix}
1 && 3 \\
4 && 2
\end{bmatrix} \quad \rightarrow \quad M =\begin{bmatrix}
0 && 0 \\
1 && 0
\end{bmatrix}\tag{4}$$
As you can see, this function creates a "mask" matrix which keeps track of where the maximum of the matrix is. True (1) indicates the position of the maximum in X, the other entries are False (0). You'll see later that the backward pass for average pooling will be similar to this but using a different mask.
**Exercise**: Implement `create_mask_from_window()`. This function will be helpful for pooling backward.
Hints:
- [np.max()]() may be helpful. It computes the maximum of an array.
- If you have a matrix X and a scalar x: `A = (X == x)` will return a matrix A of the same size as X such that:
```
A[i,j] = True if X[i,j] = x
A[i,j] = False if X[i,j] != x
```
- Here, you don't need to consider cases where there are several maxima in a matrix.
```
def create_mask_from_window(x):
"""
Creates a mask from an input matrix x, to identify the max entry of x.
Arguments:
x -- Array of shape (f, f)
Returns:
mask -- Array of the same shape as window, contains a True at the position corresponding to the max entry of x.
"""
### START CODE HERE ### (≈1 line)
mask = (x == np.max(x))
### END CODE HERE ###
return mask
np.random.seed(1)
x = np.random.randn(2,3)
mask = create_mask_from_window(x)
print('x = ', x)
print("mask = ", mask)
```
**Expected Output:**
<table>
<tr>
<td>
**x =**
</td>
<td>
[[ 1.62434536 -0.61175641 -0.52817175] <br>
[-1.07296862 0.86540763 -2.3015387 ]]
</td>
</tr>
<tr>
<td>
**mask =**
</td>
<td>
[[ True False False] <br>
[False False False]]
</td>
</tr>
</table>
Why do we keep track of the position of the max? It's because this is the input value that ultimately influenced the output, and therefore the cost. Backprop is computing gradients with respect to the cost, so anything that influences the ultimate cost should have a non-zero gradient. So, backprop will "propagate" the gradient back to this particular input value that had influenced the cost.
### 5.2.2 - Average pooling - backward pass
In max pooling, for each input window, all the "influence" on the output came from a single input value--the max. In average pooling, every element of the input window has equal influence on the output. So to implement backprop, you will now implement a helper function that reflects this.
For example if we did average pooling in the forward pass using a 2x2 filter, then the mask you'll use for the backward pass will look like:
$$ dZ = 1 \quad \rightarrow \quad dZ =\begin{bmatrix}
1/4 && 1/4 \\
1/4 && 1/4
\end{bmatrix}\tag{5}$$
This implies that each position in the $dZ$ matrix contributes equally to output because in the forward pass, we took an average.
**Exercise**: Implement the function below to equally distribute a value dz through a matrix of dimension shape. [Hint](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.ones.html)
```
def distribute_value(dz, shape):
"""
Distributes the input value in the matrix of dimension shape
Arguments:
dz -- input scalar
shape -- the shape (n_H, n_W) of the output matrix for which we want to distribute the value of dz
Returns:
a -- Array of size (n_H, n_W) for which we distributed the value of dz
"""
### START CODE HERE ###
# Retrieve dimensions from shape (≈1 line)
(n_H, n_W) = shape
# Compute the value to distribute on the matrix (≈1 line)
average = dz/(n_H * n_W)
# Create a matrix where every entry is the "average" value (≈1 line)
a = np.multiply( np.ones(shape=(n_H, n_W)), average)
### END CODE HERE ###
return a
a = distribute_value(2, (2,2))
print('distributed value =', a)
```
**Expected Output**:
<table>
<tr>
<td>
distributed_value =
</td>
<td>
[[ 0.5 0.5]
<br\>
[ 0.5 0.5]]
</td>
</tr>
</table>
### 5.2.3 Putting it together: Pooling backward
You now have everything you need to compute backward propagation on a pooling layer.
**Exercise**: Implement the `pool_backward` function in both modes (`"max"` and `"average"`). You will once again use 4 for-loops (iterating over training examples, height, width, and channels). You should use an `if/elif` statement to see if the mode is equal to `'max'` or `'average'`. If it is equal to 'average' you should use the `distribute_value()` function you implemented above to create a matrix of the same shape as `a_slice`. Otherwise, the mode is equal to '`max`', and you will create a mask with `create_mask_from_window()` and multiply it by the corresponding value of dZ.
```
def pool_backward(dA, cache, mode = "max"):
"""
Implements the backward pass of the pooling layer
Arguments:
dA -- gradient of cost with respect to the output of the pooling layer, same shape as A
cache -- cache output from the forward pass of the pooling layer, contains the layer's input and hparameters
mode -- the pooling mode you would like to use, defined as a string ("max" or "average")
Returns:
dA_prev -- gradient of cost with respect to the input of the pooling layer, same shape as A_prev
"""
### START CODE HERE ###
# Retrieve information from cache (≈1 line)
(A_prev, hparameters) = cache
# Retrieve hyperparameters from "hparameters" (≈2 lines)
stride = hparameters['stride']
f = hparameters['f']
# Retrieve dimensions from A_prev's shape and dA's shape (≈2 lines)
m, n_H_prev, n_W_prev, n_C_prev = A_prev.shape
m, n_H, n_W, n_C = dA.shape
# Initialize dA_prev with zeros (≈1 line)
dA_prev = np.zeros(shape=(m, n_H_prev, n_W_prev, n_C_prev))
for i in range(m): # loop over the training examples
# select training example from A_prev (≈1 line)
a_prev = A_prev[i]
for h in range(n_H): # loop on the vertical axis
for w in range(n_W): # loop on the horizontal axis
for c in range(n_C): # loop over the channels (depth)
# Find the corners of the current "slice" (≈4 lines)
vert_start = h
vert_end = vert_start + f
horiz_start = w
horiz_end = horiz_start + f
# Compute the backward propagation in both modes.
if mode == "max":
# Use the corners and "c" to define the current slice from a_prev (≈1 line)
#a_prev_slice = a_prev[vert_start:vert_end,horiz_start:horiz_end,c]
#a_prev_slice = A_prev[i][vert_start:vert_end, horiz_start:horiz_end,c]
a_prev_slice = A_prev[i, vert_start: vert_end, horiz_start: horiz_end, c]
# Create the mask from a_prev_slice (≈1 line)
mask = create_mask_from_window(a_prev_slice)
# Set dA_prev to be dA_prev + (the mask multiplied by the correct entry of dA) (≈1 line)
dA_prev[i, vert_start: vert_end, horiz_start: horiz_end, c] += mask * dA[i,h,w,c]
elif mode == "average":
# Get the value a from dA (≈1 line)
da = dA[i,h,w,c]
# Define the shape of the filter as fxf (≈1 line)
shape = (f,f)
# Distribute it to get the correct slice of dA_prev. i.e. Add the distributed value of da. (≈1 line)
dA_prev[i, vert_start: vert_end, horiz_start: horiz_end, c] += distribute_value(da, shape)
### END CODE ###
# Making sure your output shape is correct
assert(dA_prev.shape == A_prev.shape)
return dA_prev
np.random.seed(1)
A_prev = np.random.randn(5, 5, 3, 2)
hparameters = {"stride" : 1, "f": 2}
A, cache = pool_forward(A_prev, hparameters)
dA = np.random.randn(5, 4, 2, 2)
dA_prev = pool_backward(dA, cache, mode = "max")
print("mode = max")
print('mean of dA = ', np.mean(dA))
print('dA_prev[1,1] = ', dA_prev[1,1])
print()
dA_prev = pool_backward(dA, cache, mode = "average")
print("mode = average")
print('mean of dA = ', np.mean(dA))
print('dA_prev[1,1] = ', dA_prev[1,1])
```
**Expected Output**:
mode = max:
<table>
<tr>
<td>
**mean of dA =**
</td>
<td>
0.145713902729
</td>
</tr>
<tr>
<td>
**dA_prev[1,1] =**
</td>
<td>
[[ 0. 0. ] <br>
[ 5.05844394 -1.68282702] <br>
[ 0. 0. ]]
</td>
</tr>
</table>
mode = average
<table>
<tr>
<td>
**mean of dA =**
</td>
<td>
0.145713902729
</td>
</tr>
<tr>
<td>
**dA_prev[1,1] =**
</td>
<td>
[[ 0.08485462 0.2787552 ] <br>
[ 1.26461098 -0.25749373] <br>
[ 1.17975636 -0.53624893]]
</td>
</tr>
</table>
### Congratulations !
Congratulation on completing this assignment. You now understand how convolutional neural networks work. You have implemented all the building blocks of a neural network. In the next assignment you will implement a ConvNet using TensorFlow.
| github_jupyter |
# Converting the indications in DrugCentral to WikiData identifiers
```
import os
import requests
import pandas as pd
from pathlib import Path
from hetnet_ml.src import graph_tools as gt
```
### Drugcentral Data Dump previously extracted from postgres dump
See [here](https://github.com/mmayers12/semmed/tree/master/prepare) for more info.
```
sm_data_dir = Path('../../semmed/data/')
rel = pd.read_csv(sm_data_dir.joinpath('drugcentral_rel_06212018.csv'))
syn = pd.read_csv(sm_data_dir.joinpath('drugcentral_syn_06212018.csv'))
ids = pd.read_csv(sm_data_dir.joinpath('drugcentral_ids_06212018.csv'))
rel.head()
syn.head()
ids.head()
ids['id_type'].unique()
ids['id_type'].value_counts()
```
## Disease Xrefs
### Query WikiData for Disease X-refs
```
import functools
from wikidataintegrator.wdi_core import WDItemEngine
from tqdm import tqdm
endpoint='https://query.wikidata.org/sparql'
def parse_result_uris(result):
for c in result:
if 'Label' not in c:
idx = result[c].dropna().str.startswith('http://www.wikidata.org/entity')
if idx.sum() != 0:
idx = idx[idx].index
result.loc[idx, c] = result.loc[idx, c].apply(lambda u: u.split('/')[-1])
return result.drop_duplicates()
query_func = functools.partial(WDItemEngine.execute_sparql_query, endpoint=endpoint, as_dataframe=True)
def execute_sparql_query(query_text):
# Enforce the proper column order
col_order = query_text.split('\n')[1].split(' ?')[1:]
return parse_result_uris(query_func(query_text))[col_order]
query = """
SELECT DISTINCT ?disease ?diseaseLabel ?umlscui ?snomed_ct
WHERE {
# Initial typing for Disease
?disease wdt:P279?/wdt:P31 wd:Q12136 .
OPTIONAL {?disease wdt:P2892 ?umlscui .}
OPTIONAL {?disease wdt:P5806 ?snomed_ct. }
SERVICE wikibase:label { bd:serviceParam wikibase:language "[AUTO_LANGAGE],en" }
}
"""
disease_xrefs = execute_sparql_query(query)
disease_xrefs.head(2)
len(disease_xrefs)
```
#### Remove those with no x-ref result
```
disease_xrefs = disease_xrefs.dropna(subset=['umlscui', 'snomed_ct'], how='all').reset_index(drop=True)
len(disease_xrefs)
disease_xrefs.head(2)
```
## Compound X-refs
### Start with DrugCentral's own x-refs
```
ids.head(2)
ids['id_type'].unique()
keep_ids = ['KEGG_DRUG', 'KEGG_DRUG', 'IUPHAR_LIGAND_ID', 'CHEBI', 'DRUGBANK_ID',
'UMLSCUI', 'ChEMBL_ID', 'UNII', 'INN_ID',
'PUBCHEM_CID', 'RXNORM', 'NDFRT', 'MESH_SUPPLEMENTAL_RECORD_UI',
'MESH_DESCRIPTOR_UI', 'PDB_CHEM_ID']
ids.head(2)
dc_xrefs = (ids.query('id_type in @keep_ids')
.drop_duplicates(subset=['struct_id', 'id_type'])
.pivot(values='identifier', index='struct_id', columns='id_type'))
dc_xrefs.head(2)
```
### Now Query WikiData using the x-refs avalible to both soruces
```
id_to_wiki = {'KEGG_DRUG': 'P665',
'IUPHAR_LIGAND_ID': 'P595',
'CHEBI': 'P683',
'DRUGBANK_ID': 'P715',
'UMLSCUI': 'P2892',
'ChEMBL_ID': 'P592',
'UNII': 'P652',
'INN_ID': 'P3350',
'PUBCHEM_CID': 'P662',
'RXNORM': 'P3345',
'NDFRT': 'P2115',
'MESH_SUPPLEMENTAL_RECORD_UI': 'P6680',
'MESH_DESCRIPTOR_UI': 'P486',
'PDB_CHEM_ID': 'P3636'}
base_query = """
SELECT DISTINCT ?compound ?compoundLabel {0}
WHERE {{
# Initial typing for Compound
?compound wdt:P31 wd:Q11173 .
{1}
SERVICE wikibase:label {{ bd:serviceParam wikibase:language "[AUTO_LANGAGE],en" }}
}}
"""
line_temp = " OPTIONAL {{ ?compound wdt:{val} {key} .}}"
full_query = base_query.format(' '.join(['?'+k.lower() for k in id_to_wiki.keys()]),
'\n'.join([line_temp.format(val=v, key='?'+k.lower()) for k, v in id_to_wiki.items()]))
print(full_query)
compound_xrefs = execute_sparql_query(full_query)
compound_xrefs.head(2)
len(compound_xrefs)
```
#### Remove those with no result
```
to_drop = [c for c in compound_xrefs][2:]
compound_xrefs = compound_xrefs.dropna(subset=to_drop, how='all')
len(compound_xrefs)
compound_xrefs['compound'].nunique()
chem_xref_order = [
'unii',
'rxnorm',
'drugbank_id',
'umlscui',
'chebi',
'kegg_drug',
'iuphar_ligand_id',
'chembl_id',
'inn_id',
'pubchem_cid',
'ndfrt',
'pdb_chem_id',
'mesh_supplemental_record_ui',
'mesh_descriptor_ui']
dc_xrefs = dc_xrefs.reset_index()
dc_xrefs.columns = dc_xrefs.columns.str.lower()
```
#### Build a Final set of x-refs for the compounds
```
final_chem_xrefs = []
remaining_chems = set(dc_xrefs['struct_id'])
for x in chem_xref_order:
this_dc = dc_xrefs.query('struct_id in @remaining_chems').dropna(subset=[x])
this_wd = compound_xrefs.dropna(subset=[x])
this_xref = pd.merge(this_dc, this_wd, how='inner', on=x, suffixes=('_dc', '_wd'))
final_chem_xrefs.append(this_xref)
remaining_chems = remaining_chems - set(this_xref['struct_id'])
final_chem_xrefs = pd.concat(final_chem_xrefs, sort=False, ignore_index=True)
final_chem_xrefs.head(2)
```
#### Build a map from DrugCentral internal Structure_id to WikiData identifier
```
struct_to_wd = final_chem_xrefs.groupby('struct_id')['compound'].apply(lambda s: ';'.join(set(s))).to_dict()
disease_xrefs.head(2)
```
#### Build maps from UMLS and SNOMED to WikiData
Since Diseases have UMLS and Snomed IDs, we will build maps for both, and preferentially use the UMLS mappings
```
umls_to_wd = disease_xrefs.dropna(subset=['umlscui']).set_index('umlscui')['disease'].to_dict()
snomed_to_wd = disease_xrefs.dropna(subset=['snomed_ct']).set_index('snomed_ct')['disease'].to_dict()
rel.head(2)
inds = rel.query('relationship_name == "indication"').copy()
inds['comp_wd_id'] = inds['struct_id'].map(struct_to_wd)
# Use UMLS first, then fill missing with SNOMED mappings
inds['dis_wd_id'] = inds['umls_cui'].map(umls_to_wd).fillna(inds['snomed_conceptid'].map(snomed_to_wd))
len(inds.dropna(subset=['comp_wd_id', 'dis_wd_id']))
inds.head(2)
```
## Compare Indications to Processed WikiData Dumps
```
wd_net_dir = Path('../2_pipeline/01_querying_wikidata_for_hetnet_edges/out/').resolve()
nodes_final = pd.read_csv(wd_net_dir.joinpath('2019-09-13/nodes.csv'))
nodes_2018 = pd.read_csv(wd_net_dir.joinpath('2018-11-12/nodes.csv'))
nodes_2018_02 = pd.read_csv(wd_net_dir.joinpath('2018-02-05/nodes.csv'))
nodes_2017 = pd.read_csv(wd_net_dir.joinpath('2017-01-16/nodes.csv'))
```
### How many indications have both the compound and disease in each WikiData Hetnet?
```
nids = nodes_final[':ID']
len(inds.query('comp_wd_id in @nids and dis_wd_id in @nids'))
nids = nodes_2018[':ID']
len(inds.query('comp_wd_id in @nids and dis_wd_id in @nids'))
nids = nodes_2018_02[':ID']
len(inds.query('comp_wd_id in @nids and dis_wd_id in @nids'))
nids = nodes_2017[':ID']
len(inds.query('comp_wd_id in @nids and dis_wd_id in @nids'))
```
### How do the actual indications overlap between WikiData and DrugCentral
```
%matplotlib inline
from metapaths.tools.plot import venn2_pretty
import matplotlib.pyplot as plt
edges_final = gt.remove_colons(pd.read_csv(wd_net_dir.joinpath('2019-09-13/edges.csv'), dtype='str'))
edges_2018 = gt.remove_colons(pd.read_csv(wd_net_dir.joinpath('2018-11-12/edges.csv'), dtype='str'))
edges_2018_02 = gt.remove_colons(pd.read_csv(wd_net_dir.joinpath('2018-02-05/edges.csv'), dtype='str'))
edges_2017 = gt.remove_colons(pd.read_csv(wd_net_dir.joinpath('2017-01-16/edges.csv'), dtype='str'))
gs_wd_inds = set(inds.dropna(subset=['comp_wd_id', 'dis_wd_id'])[['comp_wd_id', 'dis_wd_id']].apply(tuple, axis=1))
len(gs_wd_inds)
edges_final.head(2)
nw_year = '2019'
treats_edges = edges_final.query('type == "TREATS_CtD"')
treats_edges = set(treats_edges[['start_id', 'end_id']].apply(tuple, axis=1))
venn2_pretty([gs_wd_inds, treats_edges], ['DrugCentral Indications\n(mapped to WD IDs)',
'WikiData Treats Edges\n{} Data Dump'.format(nw_year)])
nw_year = '2018 Nov'
treats_edges = edges_2018.query('type == "TREATS_CtD"')
treats_edges = set(treats_edges[['start_id', 'end_id']].apply(tuple, axis=1))
venn2_pretty([gs_wd_inds, treats_edges], ['DrugCentral Indications\n(mapped to WD IDs)',
'WikiData Treats Edges\n{} Data Dump'.format(nw_year)])
nw_year = '2018 Feb'
treats_edges = edges_2018_02.query('type == "TREATS_CtD"')
treats_edges = set(treats_edges[['start_id', 'end_id']].apply(tuple, axis=1))
venn2_pretty([gs_wd_inds, treats_edges], ['DrugCentral Indications\n(mapped to WD IDs)',
'WikiData Treats Edges\n{} Data Dump'.format(nw_year)])
nw_year = '2017'
treats_edges = edges_2017.query('type == "TREATS_CtD"')
treats_edges = set(treats_edges[['start_id', 'end_id']].apply(tuple, axis=1))
venn2_pretty([gs_wd_inds, treats_edges], ['DrugCentral Indications\n(mapped to WD IDs)',
'WikiData Treats Edges\n{} Data Dump'.format(nw_year)])
```
#### Looking only at Identifiers common to both DrugCentral Indications and WikiData Indications
```
nw_year = '2019'
edges = edges_final
nids = edges[['start_id', 'end_id']].stack().unique()
gs_ids = inds[['comp_wd_id', 'dis_wd_id']].stack().dropna().unique()
treats_edges = edges.query('type == "TREATS_CtD" and start_id in @gs_ids and end_id in @gs_ids')
gs_edges = set(inds.query('comp_wd_id in @nids and dis_wd_id in @nids')[['comp_wd_id', 'dis_wd_id']].apply(tuple, axis=1))
treats_edges = set(treats_edges[['start_id', 'end_id']].apply(tuple, axis=1))
venn2_pretty([gs_edges, treats_edges], ['DrugCentral Indications\n(mapped to WD IDs)',
'WikiData Treats Edges\n{} Data Dump'.format(nw_year)])
plt.title('Treats edges in DrugCentral gold standard and Wikidata:\nOnly common identifiers');
nw_year = '2018 Nov'
edges = edges_2018
nids = edges[['start_id', 'end_id']].stack().unique()
gs_ids = inds[['comp_wd_id', 'dis_wd_id']].stack().dropna().unique()
treats_edges = edges.query('type == "TREATS_CtD" and start_id in @gs_ids and end_id in @gs_ids')
gs_edges = set(inds.query('comp_wd_id in @nids and dis_wd_id in @nids')[['comp_wd_id', 'dis_wd_id']].apply(tuple, axis=1))
treats_edges = set(treats_edges[['start_id', 'end_id']].apply(tuple, axis=1))
venn2_pretty([gs_edges, treats_edges], ['DrugCentral Indications\n(mapped to WD IDs)',
'WikiData Treats Edges\n{} Data Dump'.format(nw_year)])
plt.title('Treats edges in DrugCentral gold standard and Wikidata:\nOnly common identifiers');
nw_year = '2018 Feb'
edges = edges_2018_02
nids = edges[['start_id', 'end_id']].stack().unique()
gs_ids = inds[['comp_wd_id', 'dis_wd_id']].stack().dropna().unique()
treats_edges = edges.query('type == "TREATS_CtD" and start_id in @gs_ids and end_id in @gs_ids')
gs_edges = set(inds.query('comp_wd_id in @nids and dis_wd_id in @nids')[['comp_wd_id', 'dis_wd_id']].apply(tuple, axis=1))
treats_edges = set(treats_edges[['start_id', 'end_id']].apply(tuple, axis=1))
venn2_pretty([gs_edges, treats_edges], ['DrugCentral Indications\n(mapped to WD IDs)',
'WikiData Treats Edges\n{} Data Dump'.format(nw_year)])
plt.title('Treats edges in DrugCentral gold standard and Wikidata:\nOnly common identifiers');
nw_year = '2017 January'
edges = edges_2017
nids = edges[['start_id', 'end_id']].stack().unique()
gs_ids = inds[['comp_wd_id', 'dis_wd_id']].stack().dropna().unique()
treats_edges = edges.query('type == "TREATS_CtD" and start_id in @gs_ids and end_id in @gs_ids')
gs_edges = set(inds.query('comp_wd_id in @nids and dis_wd_id in @nids')[['comp_wd_id', 'dis_wd_id']].apply(tuple, axis=1))
treats_edges = set(treats_edges[['start_id', 'end_id']].apply(tuple, axis=1))
venn2_pretty([gs_edges, treats_edges], ['DrugCentral Indications\n(mapped to WD IDs)',
'WikiData Treats Edges\n{} Data Dump'.format(nw_year)])
plt.title('Treats edges in DrugCentral gold standard and Wikidata:\nOnly common identifiers');
this_name = '05_converting_DrugCentral_indications_to_WikiData'
out_dir = Path('../2_pipeline/').joinpath(this_name).joinpath('out')
out_dir.mkdir(parents=True, exist_ok=True)
inds.to_csv(out_dir.joinpath('gold_standard.csv'), index=False)
```
| github_jupyter |
# Tutorial to zeolite graph distance
This tutorial illustrates the calculation of the graph distance between two zeolite structures with the supercell matching method.
This implementation was made by Daniel Schwalbe-Koda. It is compatible with the `pymatgen` and `networkx` packages. If you use this code or tutorial, please cite
D. Schwalbe-Koda, Z. Jensen, E. Olivetti, and R. Gómez-Bombarelli. "Graph similarity drives zeolite diffusionless transformations and intergrowth." _Nature Materials_ (2019). Link: https://www.nature.com/articles/s41563-019-0486-1
## Imports
The tools we will use in this tutorial have been written and packaged by us. They are located in the `../zeograph/` directory. To access them in this tutorial, we can simply add them to the path:
```
import sys
sys.path.append('../zeograph')
```
And then import them all:
```
import dmeasure as dm
import supercells as sc
from structure import Zeolite
```
Some extra tools for visualization:
```
import networkx as nx
import matplotlib.pyplot as plt
%matplotlib inline
```
## Computing the supercell matching
### Loading crystal structures of zeolites
In this example, we will work through the lattice matching of the SOD zeolite and the CHA one. CHA has an hexagonal lattice and SOD has a cubic lattice. To compare both through our graph-theoretical analysis, we need a transformation matrix that leads to a better match between both. Let us go over this procedure.
First, we load the crystal structures. We uploaded some sample CIF files retrieved from the International Zeolite Association website. Using our tools based on `pymatgen`, we load these files:
```
SOD = Zeolite.from_file('../data/cif/SOD.cif')
CHA = Zeolite.from_file('../data/cif/CHA.cif')
```
As mentioned before, SOD has a cubic lattice:
```
SOD.lattice
```
and CHA has a hexagonal lattice:
```
CHA.lattice
```
### Generating the transformation matrices
As discussed in the Appendix D of our paper, the implementation relies on looking for a best matching among many different transformations. First of all, we generate all transformation matrices up to a cutoff $N_\textrm{max}$, as in Eqs. D9-D11 of our SI:
```
det2matrices = sc.determinant_dict(Nmax=2)
```
`det2matrices` now have all transformation matrices with strictly positive determinants that can be found by using numbers between -2 and 2. The possible determinants for these matrices are the following:
```
print(det2matrices.keys())
```
We now have to find the determinants of the transformation matrices for each one of the zeolites. The point is to create supercells such that both structures have the same number of Si atoms inside it. This would correspond to Eq. D3 of our SI. However, as the number of determinants is limited to the list above, we have to approximate the scaling, as in Eqs. D7-8 of our SI. This corresponds to find the best possible scaling for transformations given the available determinants:
```
n_CHA, n_SOD = sc.best_scaling(
len(CHA.silicon_atoms),
len(SOD.silicon_atoms),
dets_available=list(det2matrices.keys())
)
```
We can now observe what are the best determinants for each system:
```
print('determinant for CHA: %d' % n_CHA)
print('determinant for SOD: %d' % n_SOD)
```
### Comparing the crystal structures
Now that we have the determinant for each zeolite, we can compare the lattices for a best matching. First, we take all transformation matrices and apply them to both zeolites. Therefore, the variables `A` and `B` are actually $\hat{M}^{(A)}A$ and $\hat{M}^{(B)}B$ in the article:
```
A = det2matrices[n_CHA].reshape(-1, 3, 3) @ CHA.lattice.matrix.T
B = det2matrices[n_SOD].reshape(-1, 3, 3) @ SOD.lattice.matrix.T
```
Naturally, for each system, we have many possibilities:
```
print('transformations for CHA: %d' % len(A))
print('transformations for SOD: %d' % len(B))
```
The last step is to compare the lattices. The function `supercell.compare_lattices` already implements tensor operations to make this comparison faster. It correspond to selecting transformation matrices that minimze the discrepancy between the angles of the supercell vectors (Eqs. D12-13). In this example, we will retrieve one of such transformations.
The following command can take some time to run (~5 CPU-min).
```
i, j, dist = sc.compare_lattices(A, B)
```
The discrepancy is given by the variable `dist`, and it is zero:
```
print(dist)
```
This value says we found a transformation that matches perfectly both structures. Indeed, from crystallography, we know this is possible. `i` and `j` store the indices of the transformation matrices that allow such best matching:
```
M_A = det2matrices[n_CHA][i].reshape(3, 3)
M_B = det2matrices[n_SOD][j].reshape(3, 3)
```
The transformation matrices are the following:
```
print(M_A)
print(M_B)
```
### Graph distance between supercells
Now, everything we have to do is to create the supercells specified by the matrices above and compare their graphs. Creating the supercells:
```
CHA.make_supercell(M_A)
SOD.make_supercell(M_B)
```
CHA still has the hexagonal lattice:
```
CHA.lattice
```
Now, transformed SOD also has a hexagonal lattice:
```
SOD.lattice
```
We can retrieve their periodic graphs by using the method `get_periodic_graph` of the `Zeolite` class. The `radius` specifies how far (in Å) to look for an oxygen atom during the nearest-neighbors search. This is necessary to create the right graph:
```
G_A = CHA.get_periodic_graph(radius=2)
G_B = SOD.get_periodic_graph(radius=2)
```
We can visualize such graphs:
```
fig, ax = plt.subplots(1, 2, figsize=(10, 5))
nx_draw_params = {
'node_color': '#993333',
'node_size': 100
}
nx.draw_kamada_kawai(G_A, ax=ax[0], **nx_draw_params)
nx.draw_kamada_kawai(G_B, ax=ax[1], **nx_draw_params)
ax[0].set_title('CHA')
ax[1].set_title('SOD')
plt.show()
```
Finally, we compute the D-measure between both graphs. The `DM_NORMALIZATION` is simply the normalization constant we used in our article to have the maximum distance between two zeolites equal to 1. The two zeolites with maximum supercell-matched graph distance are MRT-OSO.
Computing the D-measure for the CHA-SOD pair:
```
DM_NORMALIZATION = 0.4722216
print('D-measure for CHA-SOD: %.4f' % (dm.dmeasure(G_A, G_B) / DM_NORMALIZATION))
```
The D-measure of 0.0345 is exactly the value we provided in the article.
| github_jupyter |
# Lists
Earlier when discussing strings we introduced the concept of a *sequence* in Python. Lists can be thought of the most general version of a *sequence* in Python. Unlike strings, they are mutable, meaning the elements inside a list can be changed!
In this section we will learn about:
1.) Creating lists
2.) Indexing and Slicing Lists
3.) Basic List Methods
4.) Nesting Lists
5.) Introduction to List Comprehensions
Lists are constructed with brackets [] and commas separating every element in the list.
Let's go ahead and see how we can construct lists!
```
# Assign a list to an variable named my_list
my_list = [1,2,3]
```
We just created a list of integers, but lists can actually hold different object types. For example:
```
my_list = ['A string',23,100.232,'o']
```
Just like strings, the len() function will tell you how many items are in the sequence of the list.
```
len(my_list)
```
### Indexing and Slicing
Indexing and slicing work just like in strings. Let's make a new list to remind ourselves of how this works:
```
my_list = ['one','two','three',4,5]
# Grab element at index 0
my_list[0]
# Grab index 1 and everything past it
my_list[1:]
# Grab everything UP TO index 3
my_list[:3]
```
We can also use + to concatenate lists, just like we did for strings.
```
my_list + ['new item']
```
Note: This doesn't actually change the original list!
```
my_list
```
You would have to reassign the list to make the change permanent.
```
# Reassign
my_list = my_list + ['add new item permanently']
my_list
```
We can also use the * for a duplication method similar to strings:
```
# Make the list double
my_list * 2
# Again doubling not permanent
my_list
```
## Basic List Methods
If you are familiar with another programming language, you might start to draw parallels between arrays in another language and lists in Python. Lists in Python however, tend to be more flexible than arrays in other languages for a two good reasons: they have no fixed size (meaning we don't have to specify how big a list will be), and they have no fixed type constraint (like we've seen above).
Let's go ahead and explore some more special methods for lists:
```
# Create a new list
list1 = [1,2,3]
```
Use the **append** method to permanently add an item to the end of a list:
```
# Append
list1.append('append me!')
# Show
list1
```
Use **pop** to "pop off" an item from the list. By default pop takes off the last index, but you can also specify which index to pop off. Let's see an example:
```
# Pop off the 0 indexed item
list1.pop(0)
# Show
list1
# Assign the popped element, remember default popped index is -1
popped_item = list1.pop()
popped_item
# Show remaining list
list1
```
It should also be noted that lists indexing will return an error if there is no element at that index. For example:
```
list1[100]
```
We can use the **sort** method and the **reverse** methods to also effect your lists:
```
new_list = ['a','e','x','b','c']
#Show
new_list
# Use reverse to reverse order (this is permanent!)
new_list.reverse()
new_list
# Use sort to sort the list (in this case alphabetical order, but for numbers it will go ascending)
new_list.sort()
new_list
```
## Nesting Lists
A great feature of of Python data structures is that they support *nesting*. This means we can have data structures within data structures. For example: A list inside a list.
Let's see how this works!
```
# Let's make three lists
lst_1=[1,2,3]
lst_2=[4,5,6]
lst_3=[7,8,9]
# Make a list of lists to form a matrix
matrix = [lst_1,lst_2,lst_3]
# Show
matrix
```
We can again use indexing to grab elements, but now there are two levels for the index. The items in the matrix object, and then the items inside that list!
```
# Grab first item in matrix object
matrix[0]
# Grab first item of the first item in the matrix object
matrix[0][0]
```
# List Comprehensions
Python has an advanced feature called list comprehensions. They allow for quick construction of lists. To fully understand list comprehensions we need to understand for loops. So don't worry if you don't completely understand this section, and feel free to just skip it since we will return to this topic later.
But in case you want to know now, here are a few examples!
```
# Build a list comprehension by deconstructing a for loop within a []
first_col = [row[0] for row in matrix]
first_col
```
We used a list comprehension here to grab the first element of every row in the matrix object. We will cover this in much more detail later on!
For more advanced methods and features of lists in Python, check out the Advanced Lists section later on in this course!
| github_jupyter |
```
from plangym import AtariEnvironment, ParallelEnvironment
from plangym.montezuma import Montezuma
env = AtariEnvironment(name="MsPacman-v0", clone_seeds=True, autoreset=True)
state, obs = env.reset()
env = Montezuma(autoreset=True)
state, obs = env.reset()
states = [state.copy() for _ in range(10)]
actions = [env.action_space.sample() for _ in range(10)]
new_States, observs, rewards, ends, infos = env.step_batch(states=states, actions=actions)
state, obs
len(state)
infos
env = ParallelEnvironment(env_class=AtariEnvironment, name="MsPacman-v0",
clone_seeds=True, autoreset=True, blocking=False)
state, obs = env.reset()
states = [state.copy() for _ in range(10)]
actions = [env.action_space.sample() for _ in range(10)]
new_States, observs, rewards, ends, infos = env.step_batch(states=states, actions=actions)
import matplotlib.pyplot as plt
%matplotlib inline
mon = Montezuma()
data = mon.reset()
data[0][:1021]
len(mon.env.unwrapped.clone_full_state())
mon.step(action=0, state=data[0])
mon.step(0)
np.array(p.tuple + mon._env.room_time)
mon.pos.tuple
mon.render()
[mon.pos.x, mon.pos.y, mon.pos.objects, mon.pos.level, mon.pos.room]
import numpy as np
from plangym import AtariEnvironment, ParallelEnvironment
from plangym.montezuma import MyMontezuma, MontezumaPosLevel
class Montezuma(AtariEnvironment):
def __init__(
self,
n_repeat_action: int = 1,
min_dt: int = 1,
episodic_live: bool = False,
autoreset: bool = True,
*args, **kwargs,
):
super(Montezuma, self).__init__(name="MontezumaRevengeDeterministic-v4",
n_repeat_action=n_repeat_action,
clone_seeds=True, min_dt=min_dt, obs_ram=False)
self._env = MyMontezuma(*args, **kwargs,)
self.action_space = self._env.action_space
self.observation_space = self._env.observation_space
self.reward_range = self._env.reward_range
self.metadata = self._env.metadata
def __getattr__(self, item):
return getattr(self._env, item)
@property
def n_actions(self):
return self._env.action_space.n
def get_state(self) -> np.ndarray:
"""
Recover the internal state of the simulation. If clone seed is False the
environment will be stochastic.
Cloning the full state ensures the environment is deterministic.
"""
data = self._env.get_restore()
(
full_state,
score,
steps,
pos,
room_time,
ram_death_state,
score_objects,
cur_lives,
) = data
metadata = np.array([score, steps, room_time, ram_death_state, score_objects, cur_lives])
array = np.concatenate([full_state, metadata, np.array(pos.tuple)])
return array
def set_state(self, state: np.ndarray):
"""
Set the internal state of the simulation.
Args:
state: Target state to be set in the environment.
Returns:
None
"""
pos = MontezumaPosLevel(*state[-5:].tolist())
score, steps, room_time, ram_death_state, score_objects, cur_lives = state[-11:-5].tolist()
full_state = state[:1037].copy().astype(np.uint8)
data = (
full_state,
score,
steps,
pos,
room_time,
ram_death_state,
score_objects,
cur_lives,
)
self._env.restore(data)
def step(
self, action: np.ndarray, state: np.ndarray = None, n_repeat_action: int = None
) -> tuple:
"""
Take n_repeat_action simulation steps and make the environment evolve
in multiples of min_dt.
The info dictionary will contain a boolean called 'lost_live' that will
be true if a life was lost during the current step.
Args:
action: Chosen action applied to the environment.
state: Set the environment to the given state before stepping it.
n_repeat_action: Consecutive number of times that the action will be applied.
Returns:
if states is None returns (observs, rewards, ends, infos)
else returns(new_states, observs, rewards, ends, infos)
"""
n_repeat_action = n_repeat_action if n_repeat_action is not None else self.n_repeat_action
if state is not None:
self.set_state(state)
reward = 0
_end, lost_live = False, False
info = {"lives": -1}
terminal = False
game_end = False
for _ in range(int(n_repeat_action)):
for _ in range(self.min_dt):
obs, _reward, _end, _info = self._env.step(action)
_info["lives"] = _info.get("ale.lives", -1)
lost_live = info["lives"] > _info["lives"] or lost_live
game_end = game_end or _end
terminal = terminal or game_end
terminal = terminal or lost_live if self.episodic_life else terminal
info = _info.copy()
reward += _reward
if _end:
break
if _end:
break
# This allows to get the original values even when using an episodic life environment
info["terminal"] = terminal
info["lost_live"] = lost_live
info["game_end"] = game_end
if state is not None:
new_state = self.get_state()
data = new_state, obs, reward, terminal, info
else:
data = obs, reward, terminal, info
if _end and self.autoreset:
self._env.reset()
return data
def render(self):
"""Render the environment using OpenGL. This wraps the OpenAI render method."""
return self._env.render()
mon.cur_score
mon.room_time
data[-1]
plt.imshow(data[-1][:, :, 0])
infos
mon.pos
```
| github_jupyter |
**This notebook is an exercise in the [Time Series](https://www.kaggle.com/learn/time-series) course. You can reference the tutorial at [this link](https://www.kaggle.com/ryanholbrook/hybrid-models).**
---
# Introduction #
Run this cell to set everything up!
```
# Setup feedback system
from learntools.core import binder
binder.bind(globals())
from learntools.time_series.ex5 import *
# Setup notebook
from pathlib import Path
from learntools.time_series.style import * # plot style settings
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import LabelEncoder
from statsmodels.tsa.deterministic import DeterministicProcess
from xgboost import XGBRegressor
comp_dir = Path('../input/store-sales-time-series-forecasting')
data_dir = Path("../input/ts-course-data")
store_sales = pd.read_csv(
comp_dir / 'train.csv',
usecols=['store_nbr', 'family', 'date', 'sales', 'onpromotion'],
dtype={
'store_nbr': 'category',
'family': 'category',
'sales': 'float32',
},
parse_dates=['date'],
infer_datetime_format=True,
)
store_sales['date'] = store_sales.date.dt.to_period('D')
store_sales = store_sales.set_index(['store_nbr', 'family', 'date']).sort_index()
family_sales = (
store_sales
.groupby(['family', 'date'])
.mean()
.unstack('family')
.loc['2017']
)
```
-------------------------------------------------------------------------------
In the next two questions, you'll create a boosted hybrid for the *Store Sales* dataset by implementing a new Python class. Run this cell to create the initial class definition. You'll add `fit` and `predict` methods to give it a scikit-learn like interface.
```
# You'll add fit and predict methods to this minimal class
class BoostedHybrid:
def __init__(self, model_1, model_2):
self.model_1 = model_1
self.model_2 = model_2
self.y_columns = None # store column names from fit method
```
# 1) Define fit method for boosted hybrid
Complete the `fit` definition for the `BoostedHybrid` class. Refer back to steps 1 and 2 from the **Hybrid Forecasting with Residuals** section in the tutorial if you need.
```
def fit(self, X_1, X_2, y):
# YOUR CODE HERE: fit self.model_1
self.model_1.fit(X_1, y)
y_fit = pd.DataFrame(
# YOUR CODE HERE: make predictions with self.model_1
self.model_1.predict(X_1),
index=X_1.index, columns=y.columns,
)
# YOUR CODE HERE: compute residuals
y_resid = y - y_fit
y_resid = y_resid.stack().squeeze() # wide to long
# YOUR CODE HERE: fit self.model_2 on residuals
self.model_2.fit(X_2, y_resid)
# Save column names for predict method
self.y_columns = y.columns
# Save data for question checking
self.y_fit = y_fit
self.y_resid = y_resid
# Add method to class
BoostedHybrid.fit = fit
# Check your answer
q_1.check()
# Lines below will give you a hint or solution code
#q_1.hint()
#q_1.solution()
```
-------------------------------------------------------------------------------
# 2) Define predict method for boosted hybrid
Now define the `predict` method for the `BoostedHybrid` class. Refer back to step 3 from the **Hybrid Forecasting with Residuals** section in the tutorial if you need.
```
def predict(self, X_1, X_2):
y_pred = pd.DataFrame(
# YOUR CODE HERE: predict with self.model_1
self.model_1.predict(X_1),
index=X_1.index, columns=self.y_columns,
)
y_pred = y_pred.stack().squeeze() # wide to long
# YOUR CODE HERE: add self.model_2 predictions to y_pred
y_pred += self.model_2.predict(X_2)
return y_pred.unstack() # long to wide
# Add method to class
BoostedHybrid.predict = predict
# Check your answer
q_2.check()
# Lines below will give you a hint or solution code
#q_2.hint()
#q_2.solution()
```
-------------------------------------------------------------------------------
Now you're ready to use your new `BoostedHybrid` class to create a model for the *Store Sales* data. Run the next cell to set up the data for training.
```
# Target series
y = family_sales.loc[:, 'sales']
# X_1: Features for Linear Regression
dp = DeterministicProcess(index=y.index, order=1)
X_1 = dp.in_sample()
# X_2: Features for XGBoost
X_2 = family_sales.drop('sales', axis=1).stack() # onpromotion feature
# Label encoding for 'family'
le = LabelEncoder() # from sklearn.preprocessing
X_2 = X_2.reset_index('family')
X_2['family'] = le.fit_transform(X_2['family'])
# Label encoding for seasonality
X_2["day"] = X_2.index.day # values are day of the month
```
# 3) Train boosted hybrid
Create the hybrid model by initializing a `BoostedHybrid` class with `LinearRegression()` and `XGBRegressor()` instances.
```
# YOUR CODE HERE: Create LinearRegression + XGBRegressor hybrid with BoostedHybrid
model = BoostedHybrid(LinearRegression(), XGBRegressor())
# YOUR CODE HERE: Fit and predict
model.fit(X_1, X_2, y)
y_pred = model.predict(X_1, X_2)
y_pred = y_pred.clip(0.0)
# Check your answer
q_3.check()
# Lines below will give you a hint or solution code
#q_3.hint()
#q_3.solution()
```
-------------------------------------------------------------------------------
Depending on your problem, you might want to use other hybrid combinations than the linear regression + XGBoost hybrid you've created in the previous questions. Run the next cell to try other algorithms from scikit-learn.
```
# Model 1 (trend)
from pyearth import Earth
from sklearn.linear_model import ElasticNet, Lasso, Ridge
# Model 2
from sklearn.ensemble import ExtraTreesRegressor, RandomForestRegressor
from sklearn.neighbors import KNeighborsRegressor
from sklearn.neural_network import MLPRegressor
# Boosted Hybrid
# YOUR CODE HERE: Try different combinations of the algorithms above
model = BoostedHybrid(
model_1=Ridge(),
model_2=KNeighborsRegressor(),
)
```
These are just some suggestions. You might discover other algorithms you like in the scikit-learn [User Guide](https://scikit-learn.org/stable/supervised_learning.html).
Use the code in this cell to see the predictions your hybrid makes.
```
y_train, y_valid = y[:"2017-07-01"], y["2017-07-02":]
X1_train, X1_valid = X_1[: "2017-07-01"], X_1["2017-07-02" :]
X2_train, X2_valid = X_2.loc[:"2017-07-01"], X_2.loc["2017-07-02":]
# Some of the algorithms above do best with certain kinds of
# preprocessing on the features (like standardization), but this is
# just a demo.
model.fit(X1_train, X2_train, y_train)
y_fit = model.predict(X1_train, X2_train).clip(0.0)
y_pred = model.predict(X1_valid, X2_valid).clip(0.0)
families = y.columns[0:6]
axs = y.loc(axis=1)[families].plot(
subplots=True, sharex=True, figsize=(11, 9), **plot_params, alpha=0.5,
)
_ = y_fit.loc(axis=1)[families].plot(subplots=True, sharex=True, color='C0', ax=axs)
_ = y_pred.loc(axis=1)[families].plot(subplots=True, sharex=True, color='C3', ax=axs)
for ax, family in zip(axs, families):
ax.legend([])
ax.set_ylabel(family)
```
# 4) Fit with different learning algorithms
Once you're ready to move on, run the next cell for credit on this question.
```
# View the solution (Run this cell to receive credit!)
q_4.check()
```
# Keep Going #
[**Convert any forecasting task**](https://www.kaggle.com/ryanholbrook/forecasting-with-machine-learning) to a machine learning problem with four ML forecasting strategies.
---
*Have questions or comments? Visit the [course discussion forum](https://www.kaggle.com/learn/time-series/discussion) to chat with other learners.*
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc" style="margin-top: 1em;"><ul class="toc-item"><li><span><a href="#Choose-a-Topic" data-toc-modified-id="Choose-a-Topic-1"><span class="toc-item-num">1 </span>Choose a Topic</a></span></li><li><span><a href="#Analysis" data-toc-modified-id="Analysis-2"><span class="toc-item-num">2 </span>Analysis</a></span><ul class="toc-item"><li><span><a href="#Compare-screen-time-across-the-entire-dataset" data-toc-modified-id="Compare-screen-time-across-the-entire-dataset-2.1"><span class="toc-item-num">2.1 </span>Compare screen time across the entire dataset</a></span></li><li><span><a href="#Compare-screen-time-by-show" data-toc-modified-id="Compare-screen-time-by-show-2.2"><span class="toc-item-num">2.2 </span>Compare screen time by show</a></span><ul class="toc-item"><li><span><a href="#Including-hosts" data-toc-modified-id="Including-hosts-2.2.1"><span class="toc-item-num">2.2.1 </span>Including hosts</a></span></li><li><span><a href="#Excluding-hosts" data-toc-modified-id="Excluding-hosts-2.2.2"><span class="toc-item-num">2.2.2 </span>Excluding hosts</a></span></li></ul></li></ul></li></ul></div>
```
from esper.prelude import *
from esper.widget import *
from esper.topics import *
from esper.spark_util import *
from esper.plot_util import *
from esper.major_canonical_shows import MAJOR_CANONICAL_SHOWS
from datetime import timedelta
from collections import defaultdict
import _pickle as pickle
```
# Choose a Topic
```
topic = 'sandy hook'
lexicon = mutual_info(topic)
for word, _ in lexicon:
print(word)
selected_words = '\n'.join(x[0] for x in lexicon)
selected_words_set = set()
for line in selected_words.split('\n'):
line = line.strip()
if line == '' or line[0] == '#':
continue
selected_words_set.add(line)
filtered_lexicon = [x for x in lexicon if x[0] in selected_words_set]
segments = find_segments(filtered_lexicon, window_size=500,
threshold=100, merge_overlaps=True)
show_segments(segments[:100])
```
# Analysis
```
face_genders = get_face_genders()
face_genders = face_genders.where(
(face_genders.in_commercial == False) &
(face_genders.size_percentile >= 25) &
(face_genders.gender_id != Gender.objects.get(name='U').id)
)
intervals_by_video = defaultdict(list)
for video_id, _, interval, _, _ in segments:
intervals_by_video[video_id].append(interval)
face_genders_with_topic_overlap = annotate_interval_overlap(face_genders, intervals_by_video)
face_genders_with_topic_overlap = face_genders_with_topic_overlap.where(
face_genders_with_topic_overlap.overlap_seconds > 0)
```
## Compare screen time across the entire dataset
```
distinct_columns = ['face_id']
overlap_field = 'overlap_seconds'
z_score = 1.96
topic_screentime_with_woman = sum_distinct_over_column(
face_genders_with_topic_overlap, overlap_field, distinct_columns,
probability_column='female_probability'
)
print('Woman on screen: {:0.2f}h +/- {:0.02f}'.format(
topic_screentime_with_woman[0] / 3600, z_score * math.sqrt(topic_screentime_with_woman[1]) / 3600))
topic_screentime_with_man = sum_distinct_over_column(
face_genders_with_topic_overlap, overlap_field, distinct_columns,
probability_column='male_probability'
)
print('Man on screen: {:0.2f}h +/- {:0.02f}'.format(
topic_screentime_with_man[0] / 3600, z_score * math.sqrt(topic_screentime_with_man[1]) / 3600))
topic_screentime_with_nh_woman = sum_distinct_over_column(
face_genders_with_topic_overlap.where((face_genders_with_topic_overlap.host_probability <= 0.5)),
overlap_field, distinct_columns,
probability_column='female_probability'
)
print('Woman (non-host) on screen: {:0.2f}h +/- {:0.02f}'.format(
topic_screentime_with_nh_woman[0] / 3600, z_score * math.sqrt(topic_screentime_with_nh_woman[1]) / 3600))
topic_screentime_with_nh_man = sum_distinct_over_column(
face_genders_with_topic_overlap.where((face_genders_with_topic_overlap.host_probability <= 0.5)),
overlap_field, distinct_columns,
probability_column='male_probability'
)
print('Man (non-host) on screen: {:0.2f}h +/- {:0.02f}'.format(
topic_screentime_with_nh_man[0] / 3600, z_score * math.sqrt(topic_screentime_with_nh_man[1]) / 3600))
```
## Compare screen time by show
```
canoncal_show_map = { c.id : c.name for c in CanonicalShow.objects.all() }
distinct_columns = ['face_id']
group_by_columns = ['canonical_show_id']
overlap_field = 'overlap_seconds'
```
### Including hosts
```
CACHE_BASELINE_INCL_HOST_FILE = '/tmp/base_screentime_gender_incl_host_by_show.pkl'
try:
with open(CACHE_BASELINE_INCL_HOST_FILE, 'rb') as f:
base_screentime_with_man_by_show, base_screentime_with_woman_by_show = pickle.load(f)
print('[Base] loaded from cache')
except:
base_screentime_with_woman_by_show = {
canoncal_show_map[k[0]] : (timedelta(seconds=v[0]), v[1])
for k, v in sum_distinct_over_column(
face_genders, 'duration', distinct_columns, group_by_columns,
probability_column='female_probability'
).items() if canoncal_show_map[k[0]] in MAJOR_CANONICAL_SHOWS
}
print('[Base] Woman on screen: done')
base_screentime_with_man_by_show = {
canoncal_show_map[k[0]] : (timedelta(seconds=v[0]), v[1])
for k, v in sum_distinct_over_column(
face_genders, 'duration', distinct_columns, group_by_columns,
probability_column='male_probability'
).items() if canoncal_show_map[k[0]] in MAJOR_CANONICAL_SHOWS
}
print('[Base] Man on screen: done')
with open(CACHE_BASELINE_INCL_HOST_FILE, 'wb') as f:
pickle.dump([base_screentime_with_man_by_show, base_screentime_with_woman_by_show], f)
topic_screentime_with_woman_by_show = {
canoncal_show_map[k[0]] : (timedelta(seconds=v[0]), v[1])
for k, v in sum_distinct_over_column(
face_genders_with_topic_overlap,
overlap_field, distinct_columns, group_by_columns,
probability_column='female_probability'
).items() if canoncal_show_map[k[0]] in MAJOR_CANONICAL_SHOWS
}
print('[Topic] Woman on screen: done')
topic_screentime_with_man_by_show = {
canoncal_show_map[k[0]] : (timedelta(seconds=v[0]), v[1])
for k, v in sum_distinct_over_column(
face_genders_with_topic_overlap,
overlap_field, distinct_columns, group_by_columns,
probability_column='male_probability'
).items() if canoncal_show_map[k[0]] in MAJOR_CANONICAL_SHOWS
}
print('[Topic] Man on screen: done')
plot_binary_screentime_proportion_comparison(
['Male (incl-host)', 'Female (incl-host)'],
[topic_screentime_with_man_by_show, topic_screentime_with_woman_by_show],
'Proportion of gendered screen time by show for topic "{}"'.format(topic),
'Show name',
'Proportion of screen time',
secondary_series_names=['Baseline Male (incl-host)', 'Baseline Female (incl-host)'],
secondary_data=[base_screentime_with_man_by_show, base_screentime_with_woman_by_show]
)
```
### Excluding hosts
```
CACHE_BASELINE_NO_HOST_FILE = '/tmp/base_screentime_gender_no_host_by_show.pkl'
try:
with open(CACHE_BASELINE_NO_HOST_FILE, 'rb') as f:
base_screentime_with_nh_man_by_show, base_screentime_with_nh_woman_by_show = pickle.load(f)
print('[Base] loaded from cache')
except:
base_screentime_with_nh_woman_by_show = {
canoncal_show_map[k[0]] : (timedelta(seconds=v[0]), v[1])
for k, v in sum_distinct_over_column(
face_genders.where(face_genders.host_probability <= 0.25),
'duration', distinct_columns, group_by_columns,
probability_column='female_probability'
).items() if canoncal_show_map[k[0]] in MAJOR_CANONICAL_SHOWS
}
print('[Base] Woman (non-host) on screen: done')
base_screentime_with_nh_man_by_show = {
canoncal_show_map[k[0]] : (timedelta(seconds=v[0]), v[1])
for k, v in sum_distinct_over_column(
face_genders.where(face_genders.host_probability <= 0.25),
'duration', distinct_columns, group_by_columns,
probability_column='male_probability'
).items() if canoncal_show_map[k[0]] in MAJOR_CANONICAL_SHOWS
}
print('[Base] Man (non-host) on screen: done')
with open(CACHE_BASELINE_NO_HOST_FILE, 'wb') as f:
pickle.dump([base_screentime_with_nh_man_by_show, base_screentime_with_nh_woman_by_show], f)
topic_screentime_with_nh_woman_by_show = {
canoncal_show_map[k[0]] : (timedelta(seconds=v[0]), v[1])
for k, v in sum_distinct_over_column(
face_genders_with_topic_overlap.where(face_genders_with_topic_overlap.host_probability <= 0.25),
overlap_field, distinct_columns, group_by_columns,
probability_column='female_probability'
).items() if canoncal_show_map[k[0]] in MAJOR_CANONICAL_SHOWS
}
print('[Topic] Woman (non-host) on screen: done')
topic_screentime_with_nh_man_by_show = {
canoncal_show_map[k[0]] : (timedelta(seconds=v[0]), v[1])
for k, v in sum_distinct_over_column(
face_genders_with_topic_overlap.where(face_genders_with_topic_overlap.host_probability <= 0.25),
overlap_field, distinct_columns, group_by_columns,
probability_column='male_probability'
).items() if canoncal_show_map[k[0]] in MAJOR_CANONICAL_SHOWS
}
print('[Topic] Man (non-host) on screen: done')
plot_binary_screentime_proportion_comparison(
['Male (non-host)', 'Female (non-host)'],
[topic_screentime_with_nh_man_by_show, topic_screentime_with_nh_woman_by_show],
'Proportion of gendered screen time by show for topic "{}"'.format(topic),
'Show name',
'Proportion of screen time',
secondary_series_names=['Baseline Male (non-host)', 'Baseline Female (non-host)'],
secondary_data=[base_screentime_with_nh_man_by_show, base_screentime_with_nh_woman_by_show],
tertiary_series_names=['Male (incl-host)', 'Female (incl-host)'],
tertiary_data=[topic_screentime_with_man_by_show, topic_screentime_with_woman_by_show]
)
```
| github_jupyter |
```
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import os
plt.rcParams['axes.facecolor']='white'
plt.rcParams['figure.facecolor']='white'
```
# 0. Carga de datos
Como buena práctica para la carga de datos, es recomendado usar la función `os.path.join` de python para trabajar con directorios, ya que esta permite trabajar en distintos sistemas operativos sin mayores problemas, ya que, el manejo de directorios en Windows es distinto al de un sistema Unix-like (como macOS y linux).
```
data_path = ['..', 'm02_data_analysis', 'm02_c04_data_aggregation', 'data', 'pokemon.csv']
data_path = os.path.join(*data_path)
df = pd.read_csv(data_path)
df.head()
```
# 1. Manejo de datos y visualizaciones
Lo esencial de pandas, al fin y al cabo, es procesar datos, y el procesamiento más común que se suele hacer es agrupar datos, por eso es importante entender bien como funciona y qué es lo que hacemos con un `.groupby` por ejemplo.
En el caso de las visualizaciones, esta es la operación más frecuente, ya que estas intentan contar una historia a simple vista, y su fin es agregar la mayor cantidad de información visualmente.
En el caso de `matplotlib`, hay 2 formas de trabajar con visualizaciones:
* Usando la forma tradicional, que es pasándole los datos a las funciones de `matplotlib.pyplot`
* Usando el `wrapper` de `pandas`
## 1.1 Visualizaciones
```
pokemon_types = ((df['Type 1']
.value_counts() + df['Type 2'].value_counts())
.sort_values(ascending=False)
)
pokemon_types
fig, ax = plt.subplots()
x = pokemon_types.index
y = pokemon_types.values
ax.bar(x, y)
plt.xticks(rotation=90);
ax = (df
.pipe(lambda x: x['Type 1'].value_counts() + x['Type 2'].value_counts())
.sort_values(ascending=False)
.plot(kind='bar')
)
df.columns
stats = df.columns[4:10].tolist()
stats
axes = (df
.loc[:, ['Generation'] + stats]
.groupby('Generation')
.mean()
.plot(kind='line', layout=(1, 6), figsize=(12, 5), subplots=True, sharey=True)
)
axes = axes.ravel()
y_lim = axes[0].get_ylim()
axes[0].set_ylim((0, y_lim[1]))
# plt.tight_layout()
ax = (df
.groupby('Generation')
.Legendary.sum()
.plot(kind='bar', title='Cantidad de legendarios por generación')
)
ylims = ax.get_ylim()
ax.set_yticks(range(0, int(ylims[1]) + 1, 2));
(df
[stats]
.plot(kind='hist', subplots=True, bins=15, layout=(3, 3), figsize=(12, 10))
);
fig, axes = plt.subplots(nrows=6, ncols=3, figsize=(12, 20))
axes = axes.ravel()
for (key, group), ax in zip(df.loc[:, ['Type 1'] + stats].groupby('Type 1'), axes):
mean_group = group.mean()
data = mean_group - df.loc[:, stats].mean()
ax.bar(data.index, data.values)
ax.set_title(key)
plt.tight_layout()
(df
.loc[:, ['Type 1'] + stats]
.groupby('Type 1')
.mean()
.pipe(lambda x: x - x.mean())
.T
.plot(kind='bar', layout=(6, 3), subplots=True, figsize=(12, 20))
);
```
## Altair
```
import altair as alt
df.head()
melted = (df
.melt(id_vars='Name', value_vars=['Type 1', 'Type 2'])
.dropna()
)
melted
alt.Chart(melted).mark_bar().encode(
x=alt.X('value:N'),
y='count()'
)
melted = (df
.melt(id_vars='Generation', value_vars=stats)
)
melted.head()
alt.Chart(melted).mark_line().encode(
x='Generation',
y='mean(value)',
color='variable',
column='variable'
).properties(width=100)
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
```
# Mix of tabular + image features
```
from cape_core.tensordata import *
from cape_core.models import *
from cape_core.utils import *
from cape_core.data import *
from cape_core.ranger import *
from fastai.callbacks import SaveModelCallback
PATH = Path.cwd()
PATH.ls()
```
# Importing DataFrames
```
data = get_data(PATH); data.head()
# data = data[data.sgr_id>0]
# zooms = ['z15', 'z18', 'z20']
# zooms = ['zoom20_autoencoder', 'zoom18_autoencoder', 'zoom15_autoencoder']
# zooms = ['zoom18_256', 'zoom20_256']
zooms = ['512_zoom20_256','512_zoom18_256', '512_zoom15_256']
```
read all features DataFrames and put them on a list
```
df_list = [pd.read_pickle(f'feat_xr34_{zoom}').set_index('cad_id') for zoom in zooms]
feature_len = get_features_len(df_list[0].shape[1]); feature_len
df_list[0].head()
```
Concat the lists together
```
img_features = pd.concat([df.iloc[slice(None), -feature_len:] for df in df_list],
keys=zooms,
axis=1)
img_features.head()
img_features_flat = pd.DataFrame(data=img_features.values, index=img_features.index); img_features_flat.head()
data = data.merge(img_features_flat.reset_index(), on='cad_id'); data.shape
data.head()
data.dropna(inplace=True)
data = data.reset_index(); data.shape
val_idxs = data[~data['train']].index
cat_names = ['prop_aursrc','prop_modseg','asatMonth','prop_lastpurchMonth',
'estate_id','prop_munvalYear','prop_lastpurchYear','dont_use_rs','asatElapsed','p_90_cs_band',
'prop_bedrooms','date_3partyYear','asatYear','p_ab_final_3party_band','p_90_rs_band','p_90_3party_band',
'p_ab_rs_band','date_rsYear','p_90_final_3party_band','dont_use_cs','prop_bathrooms','dont_use_3party',
'p_ab_cs_band','p_ab_3party_band','munic_id','suburb_id']
cont_names = ['pct_offprops_cdn',
'prop_munvalinfl',
'prop_munval',
'prop_lastpurchpriceinfl',
'prop_firstlistingElapsedMonthsToDate',
'prop_minlistingprice',
'area_volotprecent',
'distancemtoairportint',
'predval_cs',
'x',
'predval_3party',
'prop_recentotp',
'estateschemedensity',
'prop_aur',
'area_maxlistingrecent',
'cad_metersfromwater',
'area_minlistingrecent',
'area_avglistingvaluerecent',
'prop_age',
'predval_final_3party',
'distancemtolargeretailcentre',
'area_avgotprecent',
'y',
'predval_rs',
'cad_metersfromcoast',
'prop_recentotpElapsedMonthsToDate',
'slope',
'cad_sqm',
'distancemtomediumretailcentre']
data[cat_names].nunique()
dep_var='trf_purchprice'
procs = [FillMissing, Categorify, Normalize]
max_log_y = np.log(np.max(data[dep_var])*1.2)
y_range = torch.tensor([12, max_log_y], device=defaults.device); y_range
features = data.iloc[slice(None), -len(zooms)*feature_len:].values; features[0:3]
# cat_names = ['estate_id',
# 'suburb_id',
# 'prop_lastpurchYear']
# cont_names = ['prop_aur',
# 'prop_age',
# 'prop_munvalinfl',
# 'predval_cs']
features_il = TensorList.from_array(features)
tabular_il = TabularList.from_df(df=data[cat_names+cont_names+[dep_var]],
cat_names=cat_names,
cont_names=cont_names,
procs=procs)
db = (MixedItemList([tabular_il, features_il], path=PATH, inner_df=tabular_il.inner_df)
.split_by_idx(val_idxs)
.label_from_df(cols=dep_var, log=True)
.databunch(no_check=True, bs=1024)
)
```
# Train
```
learn = tabular_feature_learner(db,
layers=[1000, 500],
y_range=y_range,
emb_drop=0.1,
loss_func=L1Flat(),
metrics=[exp_rmspe, exp_rmse, r2_score]).to_fp16(clip=1)
learn.model
learn.loss_func
learn.lr_find(); learn.recorder.plot()
learn.fit_one_cycle(12,
1e-3,
callbacks=[SaveModelCallback(learn, monitor='exp_rmspe', mode='min', name='best_feature_tabular')])
learn.recorder.plot_losses()
learn.load('best_feature_tabular');
p, t = learn.get_preds(DatasetType.Valid)
print_stats(p,t)
```
| github_jupyter |
```
# Imports
import os
import cPickle
from datetime import datetime
import pandas as pd
import numpy as np
from sklearn.preprocessing import Imputer
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import LabelBinarizer
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import Normalizer
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.preprocessing import MultiLabelBinarizer
from sklearn.model_selection import train_test_split
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import chi2
# Update the filename
FILENAME = 'dummy.csv'
# Constants Declaration
DATASET_DIR = './data/'
RESULT_DIR = './result/'
RANDOM_SEED = 42
EXTENSION_MAPPING = {
'read': {
'csv': 'read_csv',
'json': 'read_json',
'xlsx': 'read_excel'
},
'save': {
'csv': 'to_csv',
'json': 'to_json',
'xlsx': 'to_excel'
}
}
np.random.seed(seed=RANDOM_SEED)
# Dataset Loader
DATASET_FILE = os.path.join(DATASET_DIR, FILENAME)
file_path, file_extension = os.path.splitext(DATASET_FILE)
file_name = file_path.split(os.path.sep)[-1]
file_extension = file_extension.strip('.')
dataset_extracter = EXTENSION_MAPPING['read'].get(file_extension)
if dataset_extracter is None:
raise ValueError('Dataset type not supported')
df = getattr(pd, dataset_extracter)(DATASET_FILE)
df.head()
target_columns = list(set(['age']))
dependent_columns = list(set(df.columns) - set(target_columns))
X_train, X_test, y_train, y_test = train_test_split(
df[dependent_columns], df[target_columns],
test_size=0.2, random_state=RANDOM_SEED)
```
### Dealing with missing values
* Replace with Mean Values
* Replace with Median Values
* Replace with Most Common Values
* Replace with Specific Value
* Drop records with Missing Values
```
# Preprocessing with Sklearn, Fill with mean values for the required columns.
required_columns = []
imputer = Imputer(missing_values=np.nan, strategy="mean", axis=0)
if len(required_columns) > 0:
X_train[required_columns] = pd.DataFrame(imputer.fit_transform(X_train[required_columns]), index=X_train.index)
X_test[required_columns] = pd.DataFrame(imputer.transform(X_test[required_columns]), index=X_test.index)
# Preprocessing with Sklearn, Fill with median values for the required columns.
required_columns = []
imputer = Imputer(missing_values=np.nan, strategy="median", axis=0)
if len(required_columns) > 0:
X_train[required_columns] = pd.DataFrame(imputer.fit_transform(X_train[required_columns]), index=X_train.index)
X_test[required_columns] = pd.DataFrame(imputer.transform(X_test[required_columns]), index=X_test.index)
# Preprocessing with Sklearn, Fill with most frequent values for the required columns.
required_columns = []
imputer = Imputer(missing_values=np.nan, strategy="most_frequent", axis=0)
if len(required_columns) > 0:
X_train[required_columns] = pd.DataFrame(imputer.fit_transform(X_train[required_columns]), index=X_train.index)
X_test[required_columns] = pd.DataFrame(imputer.transform(X_test[required_columns]), index=X_test.index)
# Preprocessing with Pandas, Fill with a specific value.
value = 0
required_columns = []
if len(required_columns) > 0:
X_train[required_columns] = X_train[required_columns].fillna(value)
X_test[required_columns] = X_test[required_columns].fillna(value)
# Preprocessing with Pandas, Drop missing values
required_columns = []
if len(required_columns) > 0:
X_train.dropna(subset=required_columns, inplace=True, how='any')
X_test.dropna(subset=required_columns, inplace=True, how='any')
```
### Encoding Features
* Target Features
* Multiclass Classification
* Binary
* Non Binary
* Multilabel Classification
* Dependent Features
* Encode Classes to Labels
* One Hot Encoding of categorical data
```
# Non Binary Multiclass Classification / Encode Classes to Labels
required_columns = []
label_encoders = {}
for column in required_columns:
label_encoders[column] = LabelEncoder()
if column in X_train.columns:
X_train[column] = label_encoders[column].fit_transform(X_train[column])
X_test[column] = label_encoders[column].transform(X_test[column])
elif column in y_train.columns:
y_train[column] = label_encoders[column].fit_transform(y_train[column])
y_test[column] = label_encoders[column].transform(y_test[column])
# Multiclass Binary Classification
# Only a single column is expected
required_columns = []
label_binarizer = None
if len(required_columns) > 0:
column = required_columns[0]
if column in X_train.columns:
label_binarizer = LabelBinarizer()
X_train[column] = label_binarizer.fit_transform(X_train[column])
X_test[column] = label_binarizer.transform(X_test[column])
elif column in y_train.columns:
label_binarizer = LabelBinarizer()
y_train[column] = label_binarizer.fit_transform(y_train[column])
y_test[column] = label_binarizer.transform(y_test[column])
# Multilabel Binary Classification
# Only a single column is expected
required_columns = []
multi_label_binarizer = None
if len(required_columns) > 0:
column = required_columns[0]
if column in y_train.columns:
multi_label_binarizer = MultiLabelBinarizer()
y_train[column] = multi_label_binarizer.fit_transform(y_train[column])
y_test[column] = multi_label_binarizer.transform(y_test[column])
# One Hot Encoding of dependent features
required_columns = []
if len(required_columns) > 0:
# Avoid dummy variable trap with n-1 columns
total = pd.get_dummies(pd.concat([X_train, X_test]), columns=required_columns, drop_first=True)
X_train = total.loc[X_train.index]
X_test = total.loc[X_test.index]
#Text Preprocessing with CBOW & TFIDF Transformer
#One column expected
required_columns = []
tfidf_vect = None
if len(required_columns) > 0:
# Remove words which occur in more than 95% of the documents and should atleast have 2 occurences
tfidf_vect = TfidfVectorizer(stop_words='english', max_df=0.95, min_df=2)
column = required_columns[0]
tfidf_vect.fit(pd.concat([X_train, X_test])[column])
train_numerical_features = tfidf_vect.transform(X_train[column]).todense()
X_train = pd.concat([X_train, pd.DataFrame(train_numerical_features, index=X_train.index).add_prefix('message_')], axis=1)
test_numerical_features = tfidf_vect.transform(X_test[column]).todense()
X_test = pd.concat([X_test, pd.DataFrame(test_numerical_features, index=X_test.index).add_prefix('message_')], axis=1)
# Feature Selection with Chi2 Test
max_num_features = None
required_columns = []
selector_chi2 = None
if max_num_features is not None and len(required_columns) > 0:
selector_chi2 = SelectKBest(chi2, k=max_num_features)
print selector_chi2.fit_transform(X_train[required_columns], y_train)
print selector_chi2.transform(X_test[required_columns])
```
### Scaling Features
* Scaling
* Normalisation
```
# Utilise Standard Scaler
required_columns = []
scaler = None
if len(required_columns) > 0:
scaler = StandardScaler()
X_train[required_columns] = pd.DataFrame(scaler.fit_transform(X_train[required_columns]), index=X_train.index)
X_test[required_columns] = pd.DataFrame(scaler.transform(X_test[required_columns]), index=X_test.index)
# Utilise Normalisation
required_columns = []
normalizer = None
if len(required_columns) > 0:
normalizer = Normalizer()
X_train[required_columns] = pd.DataFrame(normalizer.fit_transform(X_train[required_columns]), index=X_train.index)
X_test[required_columns] = pd.DataFrame(normalizer.transform(X_test[required_columns]), X_test.index)
# Storage of results.
result_time = datetime.utcnow().strftime('%s')
save_dataset_fn = EXTENSION_MAPPING['save'].get(file_extension.strip('.'))
getattr(pd.concat([X_train, y_train], axis=1), save_dataset_fn)(os.path.join(RESULT_DIR, '{}.result.train.{}.{}'.format(file_name, result_time, file_extension)))
getattr(pd.concat([X_test, y_test], axis=1), save_dataset_fn)(os.path.join(RESULT_DIR, '{}.result.test.{}.{}'.format(file_name, result_time, file_extension)))
if len(label_encoders) > 0:
with open(os.path.join(RESULT_DIR, '{}.result.label_encoders.{}.{}.pkl'.format(file_name, result_time, file_extension)), 'wb') as encoder_fp:
cPickle.dump(label_encoders, encoder_fp)
if label_binarizer is not None:
with open(os.path.join(RESULT_DIR, '{}.result.label_binarizer.{}.{}.pkl'.format(file_name, result_time, file_extension)), 'wb') as encoder_fp:
cPickle.dump(label_binarizer, encoder_fp)
if multi_label_binarizer is not None:
with open(os.path.join(RESULT_DIR, '{}.result.multi_label_binarizer.{}.{}.pkl'.format(file_name, result_time, file_extension)), 'wb') as encoder_fp:
cPickle.dump(multi_label_binarizer, encoder_fp)
if scaler is not None:
with open(os.path.join(RESULT_DIR, '{}.result.scaler.{}.{}.pkl'.format(file_name, result_time, file_extension)), 'wb') as encoder_fp:
cPickle.dump(scaler, encoder_fp)
if normalizer is not None:
with open(os.path.join(RESULT_DIR, '{}.result.normalizer.{}.{}.pkl'.format(file_name, result_time, file_extension)), 'wb') as encoder_fp:
cPickle.dump(normalizer, encoder_fp)
if tfidf_vect is not None:
with open(os.path.join(RESULT_DIR, '{}.result.tfidf_vect.{}.{}.pkl'.format(file_name, result_time, file_extension)), 'wb') as encoder_fp:
cPickle.dump(tfidf_vect, encoder_fp)
if selector_chi2 is not None:
with open(os.path.join(RESULT_DIR, '{}.result.selector_chi2.{}.{}.pkl'.format(file_name, result_time, file_extension)), 'wb') as encoder_fp:
cPickle.dump(selector_chi2, encoder_fp)
```
| github_jupyter |
# *Circuitos Elétricos I - Primeiro Estágio 2020.1e*
## Gabarito da avaliação
```
m = [9,1,6] # últimos dígitos da matrícula
import numpy as np
import sympy as sp
```
### Problema 1
a. $R_{eq}=?$
```
# define valores das resistências
R1 = (m[0]+1)*1e3
R2 = (m[1]+1)*1e3
R3 = (m[2]+1)*1e3
Req = ((R1+R3)*2*R3)/(R1+3*R3)
Req = Req + 3*R2
Req = (Req*R2)/(Req+R2)
print('Req = %.2f kΩ' %(Req/1000))
```
b. Leitura do voltímetro ideal
```
# divisor de tensão
Vs = 100
Req = ((R1+R3)*2*R3)/(R1+3*R3)
Vmed1 = Vs*Req/(Req+3*R2)
print('Vmed = %.2f V' %(Vmed1))
```
c. Leitura do voltímetro de resistência interna $R_i = 20R_3$
```
# divisor de tensão
Vs = 100
Req = ((R1+R3)*2*R3)/(R1+3*R3)
Req = (Req*20*R3)/(Req+20*R3)
Vmed2 = Vs*Req/(Req+3*R2)
Erro = (Vmed1-Vmed2)/Vmed1
print('Vmed = %.2f V' %(Vmed2))
print('Erro absoluto = %.2f V' %(Vmed1-Vmed2))
print('Erro percentual = %.2f %%' %(Erro*100))
```
### Problema 2
```
# define valores das resistências
R1 = m[1]+1
R2 = m[2]+1
print('R1 = ', R1, 'Ω', ' R2 = ', R2, 'Ω',)
```
a. Correntes de malha
```
# define as variáveis
i1, i2, i3, ix = sp.symbols('i1, i2, i3, ix')
# define os sistema de equações
eq1 = sp.Eq(i1+2*ix,0)
eq2 = sp.Eq(i2+ix,0)
eq3 = sp.Eq(i3-0.5,0)
eq4 = sp.Eq(-R2*(i1-i2)-10+2*R1*(i2-i3)+3*R1*i2,0)
# resolve o sistema
soluc = sp.solve((eq1, eq2, eq3, eq4), dict=True)
print('Equações: \n\n', eq1,'\n', eq2,'\n', eq3,'\n', eq4,'\n')
i1 = np.array([sol[i1] for sol in soluc])
i2 = np.array([sol[i2] for sol in soluc])
i3 = np.array([sol[i3] for sol in soluc])
ix = np.array([sol[ix] for sol in soluc])
print('Correntes de malha:\n\n i1 = %.2f A,\n i2 = %.2f A,\n i3 = %.2f A,\n ix = %.2f A.' %(i1, i2, i3, ix))
```
b. $v_a=?$, $v_b=?$
```
va = R2*(i1-i2)
vb = 2*R1*(i2-i3)
print('va = %.2f V' %(va))
print('vb = %.2f V' %(vb))
```
c. Potências
```
# tensões desconhecidas
v_cI = R1*i1 + va
v_I = vb - 7*R2*i3
# potências
p_cI = 2*ix*v_cI
p_V = -10*i2
p_I = v_I*i3
p_R = R1*i1**2 + R2*(i1-i2)**2 + 3*R1*i2**2 + 2*R1*(i2-i3)**2 + 7*R2*i3**2
print('Potências:\n\n p_CI = %.2f W,\n p_V = %.2f W,\n p_I = %.2f W,\n p_R = %.2f W.\n' %(p_cI, p_V, p_I, p_R))
print('Soma das potências: %.2f W.'%(p_cI+p_V+p_I+p_R))
```
### Problema 3
a. $v_{th}=?$ utilizando o princípio da superposição
```
# define valores das resistências
R1 = m[0]+1
R2 = m[1]+1
R3 = m[2]+1
# define variáveis auxiliares x, y, z
x = 12
y = 2
z = 10
vth = 0
for ind in range(0,3):
# define as variáveis
v1, v2, v3 = sp.symbols('v1, v2, v3')
if ind == 0: # fonte de corrente 1A
x = 12
y = 0
z = 0
elif ind == 1:# fonte de tensão 2V
x = 0
y = 2
z = 0
elif ind == 2:# fonte de tensão 10V
x = 0
y = 0
z = 10
# define os sistema de equações
eq1 = sp.Eq(-v1/(R1+12) -v1/2 - (v2-v3)/3 - (v2-v3)/(R3+2), -x/(R1+12)+y/(R3+2))
eq2 = sp.Eq(v2-v1, z)
eq3 = sp.Eq(-v3/R2 + (v2-v3)/3 + (v2-v3)/(R3+2), -y/(R3+2))
# resolve o sistema
soluc = sp.solve((eq1, eq2, eq3), dict=True)
v1 = np.array([sol[v1] for sol in soluc])
v2 = np.array([sol[v2] for sol in soluc])
v3 = np.array([sol[v3] for sol in soluc])
vth = vth + (-v3)
print('vth %d = %.2f V' %(ind+1, -v3))
print('vth(superposição) = %.2f V' %(vth))
```
b. $R_{th}=?$
```
# Rth via resistência equivalente
Req1 = ((R1+12)*2)/(R1+14)
Req2 = ((R3+2)*3)/(R3+5)
Req = ((Req1+Req2)*R2)/(Req1+Req2+R2)
print('Via resistência equivalente:')
print('Rth = %.2f Ω\n' %(Req))
# Rth via Icc
# define variáveis auxiliares x, y, z
x = 12 # fonte de corrente 1A (transformada para fonte de tensão)
y = 2 # fonte de tensão 2V
z = 10 # fonte de tensão 10V
# define as variáveis
v1, v2 = sp.symbols('v1, v2')
# define os sistema de equações
eq1 = sp.Eq(-v1/(R1+12) -v1/2 - v2/3 - v2/(R3+2), -x/(R1+12)+y/(R3+2))
eq2 = sp.Eq(v2-v1, z)
# resolve o sistema
soluc = sp.solve((eq1, eq2), dict=True)
v1 = np.array([sol[v1] for sol in soluc])
v2 = np.array([sol[v2] for sol in soluc])
icc = -v2/3 - (v2+2)/(R3+2)
# calcula vth/icc
Rth = vth/icc
print('Via corrente de curto circuito:')
print('Rth = %.2f Ω' %(Rth))
```
c. $R_L=?$ tal que $\eta = 0.9$, onde $\eta = \frac{v_{th}i}{R_Li^2}$.
```
print('RL = %.2f Ω' %(9*Rth))
```
| github_jupyter |
# Computational and Numerical Methods
## Group 16
### Set 10 (08-10-2018): The Jacobi Iteration Method and the Gauss-Seidel Method
#### Vidhin Parmar 201601003
#### Parth Shah 201601086
```
import numpy as np
def JacobiAndGaussSeidel(A, b):
ITERATION_LIMIT = 100
print("Jacobian Method:")
print()
print("System of equations:")
for i in range(A.shape[0]):
row = ["{0:3g}*x{1}".format(A[i, j], j + 1) for j in range(A.shape[1])]
print("[{0}] = [{1:3g}]".format(" + ".join(row), b[i]))
print()
x = np.zeros_like(b)
for it_count in range(ITERATION_LIMIT):
print("Iteration {0}: {1}".format(it_count, x))
x_new = np.zeros_like(x)
for i in range(A.shape[0]):
s1 = np.dot(A[i, :i], x[:i])
s2 = np.dot(A[i, i + 1:], x[i + 1:])
x_new[i] = (b[i] - s1 - s2) / A[i, i]
if np.allclose(x, x_new, atol=1e-8, rtol=0.):
break
x = x_new
print("Analytic Solution: {0}".format(x))
# Gauss-Seidel
print()
print()
print("Gauss-Seidel:")
print()
x = np.zeros_like(b)
for it_count in range(1, ITERATION_LIMIT):
x_new = np.zeros_like(x)
print("Iteration {0}: {1}".format(it_count, x))
for i in range(A.shape[0]):
s1 = np.dot(A[i, :i], x_new[:i])
s2 = np.dot(A[i, i + 1:], x[i + 1:])
x_new[i] = (b[i] - s1 - s2) / A[i, i]
if np.allclose(x, x_new, rtol=1e-8):
break
x = x_new
print("Analytic Solution: {0}".format(x))
A = np.array([[9., 1., 1.],
[2., 10., 3.],
[3., 4., 11.]])
b = np.array([10., 19., 0.])
JacobiAndGaussSeidel(A, b)
ITERATION_LIMIT = 100
print("Jacobian Method:")
print()
print("System of equations:")
for i in range(A.shape[0]):
row = ["{0:3g}*x{1}".format(A[i, j], j + 1) for j in range(A.shape[1])]
print("[{0}] = [{1:3g}]".format(" + ".join(row), b[i]))
print()
x = np.full_like(b, 100)
for it_count in range(ITERATION_LIMIT):
print("Iteration {0}: {1}".format(it_count, x))
x_new = np.full_like(x, 100)
for i in range(A.shape[0]):
s1 = np.dot(A[i, :i], x[:i])
s2 = np.dot(A[i, i + 1:], x[i + 1:])
x_new[i] = (b[i] - s1 - s2) / A[i, i]
if np.allclose(x, x_new, atol=1e-8, rtol=0.):
break
x = x_new
print("Analytic Solution: {0}".format(x))
# Gauss-Seidel
print()
print()
print("Gauss-Seidel:")
print()
x = np.full_like(b, 100)
for it_count in range(1, ITERATION_LIMIT):
x_new = np.full_like(x, 100)
print("Iteration {0}: {1}".format(it_count, x))
for i in range(A.shape[0]):
s1 = np.dot(A[i, :i], x_new[:i])
s2 = np.dot(A[i, i + 1:], x[i + 1:])
x_new[i] = (b[i] - s1 - s2) / A[i, i]
if np.allclose(x, x_new, rtol=1e-8):
break
x = x_new
ITERATION_LIMIT = 100
print("Jacobian Method:")
print()
print("System of equations:")
for i in range(A.shape[0]):
row = ["{0:3g}*x{1}".format(A[i, j], j + 1) for j in range(A.shape[1])]
print("[{0}] = [{1:3g}]".format(" + ".join(row), b[i]))
print()
x = np.full_like(b, 10000)
for it_count in range(ITERATION_LIMIT):
print("Iteration {0}: {1}".format(it_count, x))
x_new = np.full_like(x, 10000)
for i in range(A.shape[0]):
s1 = np.dot(A[i, :i], x[:i])
s2 = np.dot(A[i, i + 1:], x[i + 1:])
x_new[i] = (b[i] - s1 - s2) / A[i, i]
if np.allclose(x, x_new, atol=1e-8, rtol=0.):
break
x = x_new
print("Analytic Solution: {0}".format(x))
# Gauss-Seidel
print()
print()
print("Gauss-Seidel:")
print()
x = np.full_like(b, 10000)
for it_count in range(1, ITERATION_LIMIT):
x_new = np.full_like(x, 10000)
print("Iteration {0}: {1}".format(it_count, x))
for i in range(A.shape[0]):
s1 = np.dot(A[i, :i], x_new[:i])
s2 = np.dot(A[i, i + 1:], x[i + 1:])
x_new[i] = (b[i] - s1 - s2) / A[i, i]
if np.allclose(x, x_new, rtol=1e-8):
break
x = x_new
```
| github_jupyter |
```
from tensorflow.keras.applications.mobilenet import preprocess_input
from tensorflow.keras.preprocessing.image import ImageDataGenerator
import numpy as np
import tensorflow as tf
from tensorflow.keras.datasets import cifar10
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.models import Model
from tensorflow.keras import applications
from tensorflow.keras.layers import BatchNormalization,Conv2D, AveragePooling2D,TimeDistributed,Dense, Dropout, Activation, Flatten,GlobalAveragePooling2D,MaxPool2D
train_path='/home/paa/COVID-19/dataset'
train_datagen=ImageDataGenerator(rescale=1./255,preprocessing_function=preprocess_input,validation_split = 0.25) #included in our dependencies
train_generator=train_datagen.flow_from_directory(train_path,
target_size=(500,500),
color_mode='grayscale',
batch_size=32,
class_mode='binary',
subset = 'training',
shuffle=True)
validation_generator = train_datagen.flow_from_directory(train_path,
target_size=(500,500),
color_mode='grayscale',
batch_size=32,
class_mode='binary',
subset = 'validation',
shuffle=True)
step_size_train=train_generator.n//train_generator.batch_size
step_size_validation=validation_generator.n//validation_generator.batch_size
print(step_size_train)
print(step_size_validation)
img_height,img_width=500,500
base_model = applications.densenet.DenseNet121(weights= None, include_top=False, input_shape= (img_height,img_width,1))
x = base_model.output
x = GlobalAveragePooling2D()(x)
x = Dropout(0.5)(x)
predictions = Dense(1, activation= 'sigmoid')(x)
model = Model(inputs = base_model.input, outputs = predictions)
model.compile(optimizer= 'RMSprop', loss='binary_crossentropy', metrics=['accuracy'])
model.summary()
graph=model.fit_generator(generator=train_generator,
validation_data=validation_generator,
steps_per_epoch=step_size_train,
validation_steps=step_size_validation,
epochs=1)
import matplotlib.pyplot as plt
plt.plot(graph.history['acc'])
plt.plot(graph.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# summarize history for loss
plt.plot(graph.history['loss'])
plt.plot(graph.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
model=tf.keras.models.load_model("covid19 densenet02.h5")
model.summary()
def prepare(ima):
IMG_SIZE = 500 # 50 in txt-based
img_array = cv2.cvtColor(ima,cv2.COLOR_BGR2GRAY)
img_array=img_array/255.0 # filepathread in the image, convert to grayscale
new_array = cv2.resize(img_array, (IMG_SIZE, IMG_SIZE)) # resize image to match model's expected sizing
return new_array.reshape(-1,IMG_SIZE, IMG_SIZE,1)
import cv2
for i in range(1,40):
img=cv2.imread("/home/paa/COVID-19/test/"+str(i)+"N.jpeg")
a=model.predict(prepare(img))
print(a)
from sklearn.metrics import classification_report, confusion_matrix
Y_pred = model.predict_generator(validation_generator, 452 // 5+1)
# y_pred = np.argmax(Y_pred, axis=1)
Y_pred[Y_pred>0.5]=1
Y_pred[Y_pred<0.5]=0
y_pred=Y_pred
print('Confusion Matrix')
print(confusion_matrix(validation_generator.classes, y_pred))
print('Classification Report')
target_names = ['Normal', 'Positive']
print(classification_report(validation_generator.classes, y_pred, target_names=target_names))
img=cv2.imread("/home/paa/COVID-19/dataset/Normal/105.jpeg")
a=model.predict(prepare(img))
print(a)
```
| github_jupyter |
```
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import pickle
import tensorflow_probability as tfp
tfd = tfp.distributions
tf.test.is_gpu_available()
def sample_data():
count = 100000
rand = np.random.RandomState(0)
a = [[-1.5, 2.5]] + rand.randn(count // 3, 2) * 0.2
b = [[1.5, 2.5]] + rand.randn(count // 3, 2) * 0.2
c = np.c_[2 * np.cos(np.linspace(0, np.pi, count // 3)),
-np.sin(np.linspace(0, np.pi, count // 3))]
c += rand.randn(*c.shape) * 0.2
data_x = np.concatenate([a, b, c], axis=0)
data_y = np.array([0] * len(a) + [1] * len(b) + [2] * len(c))
perm = rand.permutation(len(data_x))
return data_x[perm], data_y[perm]
X, Y = sample_data()
X_train, Y_train = X[:80000,:], Y[:80000]
X_test, Y_test = X[-20000:,:], Y[-20000:]
plt.scatter(X_train[:,0], X_train[:,1], c=Y_train)
```



```
def dense(x, nrof_units, activation=None, training=True, use_batch_norm=False):
x = tf.compat.v1.layers.Dense(units=nrof_units)(x)
if use_batch_norm:
x = tf.compat.v1.layers.BatchNormalization()(x, training=training)
x = x if activation is None else activation(x)
return x
def mlp(x, nrof_units, activation, nrof_layers=1, training=True):
for _ in range(nrof_layers-1):
x = dense(x, nrof_units=nrof_units, activation=activation, training=training)
return x
def coupling_layer(x, nrof_units, nrof_layers, flip):
x1, x2 = tf.split(x, num_or_size_splits=2, axis=1)
if flip:
x1, x2 = x2, x1
y1 = x1
sx = dense(mlp(x1, nrof_units=nrof_units, nrof_layers=nrof_layers, activation=tf.nn.leaky_relu),
nrof_units=1, activation=tf.nn.tanh)
tx = dense(mlp(x1, nrof_units=nrof_units, nrof_layers=nrof_layers, activation=tf.nn.leaky_relu),
nrof_units=1, activation=None)
y2 = x2*tf.exp(sx) + tx
if flip:
y1, y2 = y2, y1
y = tf.concat([y1,y2], axis=1)
log_det_jacobian = tf.reduce_sum(sx, axis=1)
return y, log_det_jacobian
def real_nvp(x, nrof_layers, nrof_mlp_layers, nrof_mlp_units):
log_det_jacobian_list = []
# RealNVP coupling layers
for i in range(nrof_layers):
flip = i % 2 == 1
x, log_det_jacobian = coupling_layer(x, nrof_units=nrof_mlp_units, nrof_layers=nrof_mlp_layers, flip=flip)
log_det_jacobian_list += [ log_det_jacobian ]
# Element-wise sigmoid
x = tf.nn.sigmoid(x)
log_det_sigmoid = tf.reduce_sum(tf.log((1.0-x)*x + 1e-3), axis=1)
log_det_jacobian_list += [ log_det_sigmoid ]
log_det_jacobian_tot = tf.add_n(log_det_jacobian_list)
return x, log_det_jacobian_tot
batch_size=100
nrof_epochs=20
tf.reset_default_graph()
with tf.Graph().as_default():
x_ph = tf.placeholder(tf.float32, shape=(None, 2))
x, log_det_jacobian = real_nvp(x_ph, nrof_layers=6, nrof_mlp_layers=4, nrof_mlp_units=10)
base_dist = tfd.Independent(tfd.Uniform(low=[0, 0], high=[1, 1]), reinterpreted_batch_ndims=1)
log_prob = base_dist.log_prob(x) + log_det_jacobian
prob = tf.math.exp(log_prob)
nll = -tf.reduce_mean(log_prob)
loss = nll
optimizer = tf.compat.v1.train.AdamOptimizer(learning_rate=0.0001)
train_op = optimizer.minimize(loss)
sess = tf.Session()
sess.run(tf.compat.v1.global_variables_initializer())
nrof_train_batches = int(np.floor(X_train.shape[0] / batch_size))
train_loss_list = []
for epoch in range(1, nrof_epochs+1):
for i in range(nrof_train_batches):
x_batch = X_train[i*batch_size:(i+1)*batch_size]
_, loss_ = sess.run([train_op, loss], feed_dict={x_ph: x_batch})
train_loss_list += [ loss_ ]
if epoch % 1 == 0:
print('train epoch: %d loss: %.7f' % (epoch, loss_ / np.log(2)))
test_loss_ = sess.run(loss, feed_dict={x_ph: X_test})
print('Test NLL: %.3f bits/dim' % (test_loss_ / np.log(2)))
plt.plot(np.array(train_loss_list) / np.log(2))
# Learned density
x_grid = np.arange(-4, 4, 0.05)
y_grid = np.arange(-4, 4, 0.05)
x1, x2 = np.meshgrid(x_grid, y_grid)
X_mesh = np.vstack([x1.flatten(),x2.flatten()]).T
prob_ = sess.run(prob, feed_dict={x_ph: X_mesh})
sz = int(np.sqrt(prob_.shape[0]))
_ = plt.pcolormesh(x_grid, y_grid, np.reshape(prob_, (sz,sz)))
# Latent space
z_ = sess.run(x, feed_dict={x_ph: X_train[:60000,:]})
plt.scatter(z_[:,0], z_[:,1], c=Y_train[:60000])
```
| github_jupyter |
# Exemplo de uso Tensorboard com MNIST
O Tensorboard é uma ferramenta integrada ao tensorflow que permite a visualização de estatísticas de uma rede neural como parâmetros de treinamento (perda, acurácia e pesos), imagens e o grafo construído. Ele é útil para ajudar a entender o fluxo dos tensores no grafo e também corrigir e otimizar o modelo.
O Tensorboard funciona lendo *event files* escritos por uma aplicação Tensorflow que escreve as *summary data*.
Para executar o Tensoboard deve-se utilizar o comando: tensorboard --logdir=[dir] onde [dir] é o diretório onde estão localizados os *event files*.
Para escrever um *event file* é preciso criar uma instância *FileWriter*, e para isso basta chamar seu construtor *tf.summary.FileWriter([dir], [graph])*, onde [dir] é o diretório dos *event files* e [graph] é o grafo construído.
Para gerar os dados que serão observados podemos utilizar a função tf.summary.scalar(name, data)onde scalar pode ser histogram, image, audio e text, dependendo do tipo do dado a ser visualizado.
Por fim usamos writer.add_summary(summary, step) para escrever os dados no event file, onde writer é uma instância de FileWriter.
```
import numpy as np
import matplotlib.pyplot as plt
import os
import cv2
from tqdm import tqdm
import datetime, os
import pathlib, shutil
import random
#import tensorboard
import tensorflow as tf
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Activation, Flatten
from tensorflow.keras.layers import Conv2D, MaxPooling2D, BatchNormalization
from tensorflow.keras import layers, models, utils
%load_ext tensorboard
IMG_SIZE = 50
BATCH_SIZE = 2000
EPOCHS = 10
DATADIR = "/data/dataset/mnist/trainingSet/trainingSet"
TESTDIR = "/data/dataset/mnist/trainingSample/trainingSample"
CATEGORIES = ["0", "1", "2", "3", "4", "5", "6", "7", "8", "9"]
logs_base_dir = "./logs"
shutil.rmtree(logs_base_dir, ignore_errors=True, onerror=None)
os.makedirs(logs_base_dir, exist_ok=True)
data_train = pathlib.Path(DATADIR)
data_test = pathlib.Path(TESTDIR)
SIZE_OF_DATASET = len(list(data_train.glob('*/*.jpg')))
SIZE_OF_TEST = len(list(data_test.glob('*/*.jpg')))
print("Number of training images: ",SIZE_OF_DATASET)
print("Number of test images: ",SIZE_OF_TEST)
def prep_data(DATA_DIR, CATEGORIES):
data = []
for category in CATEGORIES:
path = os.path.join(DATA_DIR,category)
class_num = CATEGORIES.index(category)
for img in tqdm(os.listdir(path)):
img_array = cv2.imread(os.path.join(path,img) ,cv2.IMREAD_GRAYSCALE)
new_array = cv2.resize(img_array, (IMG_SIZE, IMG_SIZE))
data.append([new_array, class_num])
plt.figure(figsize=(1,1))
plt.imshow(new_array, cmap='gray')
plt.show()
return data
def prep_2(data):
random.shuffle(data)
X = []
y = []
for features,label in data:
X.append(features)
y.append(label)
res = np.eye(10)[y]
X = np.array(X).reshape(-1, IMG_SIZE, IMG_SIZE, 1)
return X, res
data = prep_data(TESTDIR, CATEGORIES)
tX, ty = prep_2(data)
data2 = prep_data(DATADIR, CATEGORIES)
X, y = prep_2(data2)
X=np.array(X/255.0)
y=np.array(y)
tX=np.array(tX/255.0)
ty=np.array(ty)
model = models.Sequential()
model.add(layers.Conv2D(32, (16, 16), activation='relu', input_shape=(X.shape[1:])))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(64, (5, 5), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add( layers.Flatten( ) )
model.add( layers.Dense(128, activation='relu') )
model.add( layers.Dense(10, activation='softmax') )
#opt = tf.keras.optimizers.SGD(
# learning_rate=0.01, momentum=0.1, nesterov=False, name="SGD")
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
tensorboard_callback = tf.keras.callbacks.TensorBoard(logs_base_dir, histogram_freq=1)
history = model.fit(X, y, batch_size=BATCH_SIZE, epochs=EPOCHS, validation_data=(tX, ty), callbacks=[tensorboard_callback])
plt.plot(history.history['accuracy'], label='accuracy')
plt.plot(history.history['val_accuracy'], label = 'val_accuracy')
plt.xlabel('Epoch')
plt.ylabel('Accuracy')
plt.ylim([0.5, 1])
plt.legend(loc='lower right')
#%tensorboard --logdir=logs_base_dir
%tensorboard --logdir /home/marcial/lasid-imagens/logs --bind_all
### --host 10.129.64.30
from tensorboard import notebook
notebook.list()
#notebook.display(port=6006, height=1000)
```
| github_jupyter |
```
# Insert code here.
import pandas as pd
import numpy as np
import random
import re
import time
import datetime
from nltk.corpus import stopwords
from nltk.tokenize import sent_tokenize
from sklearn import model_selection, preprocessing, linear_model, naive_bayes, metrics, svm, neighbors
from sklearn.preprocessing import LabelEncoder
from transformers import AutoTokenizer, AutoModelForSequenceClassification, AdamW, BertConfig, AutoModel
import torch
from torch.utils.data import TensorDataset, random_split, DataLoader, RandomSampler, SequentialSampler
from transformers import BertForSequenceClassification
from transformers import get_linear_schedule_with_warmup
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics import classification_report
from torch.utils.data import Dataset
from tqdm import tqdm
# from sentence_transformers import SentenceTransformer
# sent_encoder = SentenceTransformer('bert-base-nli-mean-tokens')
if torch.cuda.is_available():
# Tell PyTorch to use the GPU.
device = torch.device("cuda:2")
print('There are %d GPU(s) available.' % torch.cuda.device_count())
print('We will use the GPU:', torch.cuda.get_device_name(0))
# If not...
else:
print('No GPU available, using the CPU instead.')
device = torch.device("cpu")
torch.cuda.empty_cache()
models = ['bert-base-uncased', 'distilbert-base-uncased-finetuned-sst-2-english', 'textattack/roberta-base-SST-2','roberta-base', 'google/electra-base-discriminator', 'xlnet-base-cased', 'xlm-roberta-base', '/scratch/covid-tapt', '/scratch/covid-tapt/checkpoint-500']
model_num = 8
tokenizer = AutoTokenizer.from_pretrained(models[model_num])
train = pd.read_csv('../datasets/covid/Constraint_English_Train - Sheet1.csv')
test = pd.read_csv('../datasets/covid/Constraint_English_Val - Sheet1.csv')
import pickle
with open('train.pickle','rb') as f:
train = pickle.load(f)
train = pd.DataFrame.from_dict(train)
train.drop(train.head(1).index, inplace=True)
with open('valid.pickle','rb') as f:
valid = pickle.load(f)
valid = pd.DataFrame.from_dict(valid)
valid.drop(valid.head(1).index, inplace=True)
with open('test.pickle','rb') as f:
test = pickle.load(f)
del test['task_1']
test = pd.DataFrame.from_dict(test)
# test.drop(test.head(1).index, inplace=True)
# test = pd.DataFrame.from_dict(test)
# test = pd.read_csv('data/valid.tsv', sep='\t')
train = pd.concat([train, valid])
test.head(10)
labels = ['fake','real']
def label_encode(val):
return labels.index(val)
train['label'] = train.task_1.apply(label_encode)
train['tweet'] = train.full_tweet
test['tweet'] = test.full_tweet
valid.emoji.sample(10)
train = train.reset_index(drop=True)
REPLACE_BY_SPACE_RE = re.compile('[/(){}\[\]\|@,;]')
BAD_SYMBOLS_RE = re.compile('[^0-9a-z #+_]')
STOPWORDS = []
def clean_text(text):
"""
text: a string
return: modified initial string
"""
text = text.lower() # lowercase text
text = REPLACE_BY_SPACE_RE.sub(' ', text) # replace REPLACE_BY_SPACE_RE symbols by space in text. substitute the matched string in REPLACE_BY_SPACE_RE with space.
text = BAD_SYMBOLS_RE.sub('', text) # remove symbols which are in BAD_SYMBOLS_RE from text. substitute the matched string in BAD_SYMBOLS_RE with nothing.
# text = re.sub(r'\W+', '', text)
text = ' '.join(word for word in text.split() if word not in STOPWORDS) # remove stopwors from text
return text
train.tweet = train.tweet.apply(clean_text)
train.tweet = train.tweet.str.replace('\d+', '')
# test.label = test.label.apply(label_encode)
test = test.reset_index(drop=True)
test.tweet = test.tweet.apply(clean_text)
test.tweet = test.tweet.str.replace('\d+', '')
train.tweet.sample(10)
# split the dataset into training and validation datasets
from sklearn.model_selection import train_test_split
# train_x, valid_x, train_y, valid_y = model_selection.train_test_split(train['tweet'], train['label'])
train_x, valid_x, train_y, valid_y = train_test_split(train['tweet'], train['label'], test_size=0.2)
def count_words(text):
try:
return len(text.split())
except:
print(text)
return None
total = 0
maxw = 0
large_count = 0
for i in train_x:
temp = count_words(i)
total += temp
maxw = temp if temp > maxw else maxw
large_count += 1 if temp > 120 else 0
total/len(train_x), maxw, large_count, len(train_x)
# MAX_LENGTH = 50
posts = train.values
categories = train.values
# Sections of config
# Defining some key variables that will be used later on in the training
MAX_LEN = 128
TRAIN_BATCH_SIZE = 16
VALID_BATCH_SIZE = 32
EPOCHS = 10
LEARNING_RATE = 1e-05
import gensim.models as gsm
e2v = gsm.KeyedVectors.load_word2vec_format('emoji2vec.bin', binary=True)
# happy_vector = e2v['😂'] # Produces an embedding vector of length 300
# Download the bin file from here https://github.com/uclnlp/emoji2vec/blob/master/pre-trained/emoji2vec.bin
def getEmojiEmbeddings(emojiList,dim=300,verbose = False):
""" Generates an emoji vector by averaging the emoji representation for each emoji. If no emoji returns an empty list of dimension dim"""
if dim < 300:
raise IndexError("Dim has to be greater than 300")
result = np.zeros(dim)
if (len(emojiList) == 0):
return result
else:
embs = None
for i in emojiList:
if verbose:
if i not in e2v.vocab:
print(i)
embs = np.mean([e2v[i] for i in emojiList if i in e2v.vocab], axis=0)
if np.any(np.isnan(embs)):
return result
result[:300] = embs
return result
getEmojiEmbeddings(valid.emoji.values[0])
ids = tokenizer.encode_plus(
valid.full_tweet.values[0],
None,
truncation=True,
add_special_tokens=True,
max_length=128,
pad_to_max_length=True,
return_attention_mask = True,
return_token_type_ids=True
)['input_ids']
torch.tensor(ids, dtype=torch.long).shape, torch.tensor(getEmojiEmbeddings(valid.emoji.values[0]), dtype=torch.long).shape
class MultiLabelDataset(Dataset):
def __init__(self, dataframe, tokenizer, max_len, t = False):
self.tokenizer = tokenizer
self.data = dataframe
self.text = dataframe.tweet
self.emoji = dataframe.emoji
self.hash = dataframe.segmented_hash
self.t = t
if not self.t:
self.targets = self.data.label
self.max_len = max_len
def __len__(self):
return len(self.text)
def __getitem__(self, index):
text = str(self.text[index])
text = " ".join(text.split())
inputs = self.tokenizer.encode_plus(
text,
None,
truncation=True,
add_special_tokens=True,
max_length=self.max_len,
pad_to_max_length=True,
return_attention_mask = True,
return_token_type_ids=True
)
ids = inputs['input_ids']
mask = inputs['attention_mask']
token_type_ids = inputs["token_type_ids"]
h_text = self.hash[index]
h_text = " ".join(h_text)
inputs = self.tokenizer.encode_plus(
h_text,
None,
truncation=True,
add_special_tokens=True,
max_length=self.max_len,
pad_to_max_length=True,
return_attention_mask = True,
return_token_type_ids=True
)
h_ids = inputs['input_ids']
h_mask = inputs['attention_mask']
h_token_type_ids = inputs["token_type_ids"]
# h_inputs
emoji = getEmojiEmbeddings(self.emoji[index])
if self.t:
return {
'ids': torch.tensor(ids, dtype=torch.long),
'mask': torch.tensor(mask, dtype=torch.long),
'token_type_ids': torch.tensor(token_type_ids, dtype=torch.long),
'h_ids': torch.tensor(h_ids, dtype=torch.long),
'h_mask': torch.tensor(h_mask, dtype=torch.long),
'h_token_type_ids': torch.tensor(h_token_type_ids, dtype=torch.long),
'emoji' : torch.tensor(emoji, dtype=torch.long),
}
else:
return {
'ids': torch.tensor(ids, dtype=torch.long),
'mask': torch.tensor(mask, dtype=torch.long),
'token_type_ids': torch.tensor(token_type_ids, dtype=torch.long),
'h_ids': torch.tensor(h_ids, dtype=torch.long),
'h_mask': torch.tensor(h_mask, dtype=torch.long),
'h_token_type_ids': torch.tensor(h_token_type_ids, dtype=torch.long),
'emoji' : torch.tensor(emoji, dtype=torch.long),
'targets': torch.tensor(self.targets[index], dtype=torch.long)
}
# Creating the dataset and dataloader for the neural network
train_size = 0.8
train_data=train.sample(frac=train_size,random_state=200)
test_data=train.drop(train_data.index).reset_index(drop=True)
train_data = train_data.reset_index(drop=True)
print("FULL Dataset: {}".format(train.shape))
print("TRAIN Dataset: {}".format(train_data.shape))
print("TEST Dataset: {}".format(test_data.shape))
training_set = MultiLabelDataset(train_data, tokenizer, MAX_LEN)
testing_set = MultiLabelDataset(test_data, tokenizer, MAX_LEN)
train_params = {'batch_size': TRAIN_BATCH_SIZE,
'shuffle': True,
'num_workers': 0
}
test_params = {'batch_size': VALID_BATCH_SIZE,
'shuffle': True,
'num_workers': 0
}
training_loader = DataLoader(training_set, **train_params)
testing_loader = DataLoader(testing_set, **test_params)
class BERTClass(torch.nn.Module):
def __init__(self):
super(BERTClass, self).__init__()
self.l1 = AutoModel.from_pretrained(models[model_num])
self.l2 = AutoModel.from_pretrained(models[model_num])
self.pre_classifier_1 = torch.nn.Linear(768, 768)
self.pre_classifier_2 = torch.nn.Linear(768, 768)
self.dropout = torch.nn.Dropout(0.1)
self.pre_classifier_3 = torch.nn.Linear(1836, 1836)
# self.pre_classifier_3 = torch.nn.Linear(768, 100)
self.classifier = torch.nn.Linear(1836, 2)
def forward(self, input_ids, attention_mask, token_type_ids, h_ids, h_mask, h_token_type_ids, emoji):
output_1 = self.l1(input_ids=input_ids, attention_mask=attention_mask)
hidden_state_1 = output_1[0]
pooler_1 = hidden_state_1[:, 0]
pooler_1 = self.pre_classifier_1(pooler_1)
pooler_1 = torch.nn.Tanh()(pooler_1)
pooler_1 = self.dropout(pooler_1)
output_2 = self.l2(input_ids=h_ids, attention_mask=h_mask)
hidden_state_2 = output_2[0]
pooler_2 = hidden_state_2[:, 0]
pooler_2 = self.pre_classifier_2(pooler_2)
pooler_2 = torch.nn.Tanh()(pooler_2)
pooler_2 = self.dropout(pooler_2)
pooler_3 = torch.cat((pooler_1, pooler_2), 1)
pooler_3 = torch.cat((pooler_3, emoji), 1)
# print(pooler_1.shape,hidden_state_1.shape, pooler_2.shape, emoji.type(torch.FloatTensor).shape)
# pooler_3 = torch.nn.Tanh()(emoji.type(torch.FloatTensor))
# pooler_3 = self.dropout(pooler_3)
# print(pooler_3.shape)
pooler_3 = self.pre_classifier_3(pooler_3)
# pooler_3 = self.pre_classifier_3(pooler_2)
pooler_3 = torch.nn.Tanh()(pooler_3)
pooler_3 = self.dropout(pooler_3)
output = self.classifier(pooler_3)
return output
model = BERTClass()
model.to(device)
# from torchsummary import summary
# print(repr(model))
params = list(model.named_parameters())
print('The BERT model has {:} different named parameters.\n'.format(len(params)))
print('==== Embedding Layer ====\n')
for p in params[0:5]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
print('\n==== First Transformer ====\n')
for p in params[5:21]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
print('\n==== Output Layer ====\n')
for p in params[-4:]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
def loss_fn(outputs, targets):
return torch.nn.CrossEntropyLoss()(outputs, targets)
optimizer = torch.optim.Adam(params = model.parameters(), lr=LEARNING_RATE)
def train(epoch):
total_train_loss = 0
count = 0
model.train()
for _,data in tqdm(enumerate(training_loader, 0)):
ids = data['ids'].to(device, dtype = torch.long)
mask = data['mask'].to(device, dtype = torch.long)
token_type_ids = data['token_type_ids'].to(device, dtype = torch.long)
h_ids = data['h_ids'].to(device, dtype = torch.long)
h_mask = data['h_mask'].to(device, dtype = torch.long)
h_token_type_ids = data['h_token_type_ids'].to(device, dtype = torch.long)
targets = data['targets'].to(device, dtype = torch.long)
emoji = data['emoji'].to(device, dtype = torch.long)
outputs = model(ids, mask, token_type_ids, h_ids, h_mask, h_token_type_ids, emoji)
optimizer.zero_grad()
# loss = outputs.loss
loss = loss_fn(outputs, targets)
# if _%50==0:
# print(f'Epoch: {epoch}, Loss: {loss.item()}')
total_train_loss += loss.item()
count += 1
optimizer.zero_grad()
loss.backward()
optimizer.step()
model.eval()
fin_targets=[]
fin_outputs=[]
print(f'Epoch: {epoch}, Loss: {total_train_loss/count}')
with torch.no_grad():
for _, data in tqdm(enumerate(testing_loader, 0)):
ids = data['ids'].to(device, dtype = torch.long)
mask = data['mask'].to(device, dtype = torch.long)
token_type_ids = data['token_type_ids'].to(device, dtype = torch.long)
h_ids = data['h_ids'].to(device, dtype = torch.long)
h_mask = data['h_mask'].to(device, dtype = torch.long)
h_token_type_ids = data['h_token_type_ids'].to(device, dtype = torch.long)
targets = data['targets'].to(device, dtype = torch.long)
emoji = data['emoji'].to(device, dtype = torch.long)
outputs = model(ids, mask, token_type_ids, h_ids, h_mask, h_token_type_ids, emoji)
fin_targets.extend(targets.cpu().detach().numpy().tolist())
fin_outputs.extend(torch.sigmoid(outputs).cpu().detach().numpy().tolist())
fin_outputs = list(np.argmax(np.array(fin_outputs), axis=1).flatten())
print(classification_report(fin_outputs, fin_targets))
torch.save(model, '/scratch/epoch_'+str(epoch))
return fin_outputs, fin_targets
# final_outputs = np.array(fin_outputs) >=0.5
# final = []
# final_t = []
# final_fine = [[],[],[],[]]
# final_fine_t = [[],[],[],[]]
# for (i,j) in zip(final_outputs, fin_targets):
# output_sum = sum(i)
# target_sum = sum(j)
# if output_sum == 0:
# final.append(0)
# else:
# final.append(1)
# if target_sum == 0:
# final_t.append(0)
# else:
# final_t.append(1)
# for p in range(4):
# final_fine[p].append(int(i[p]))
# final_fine_t[p].append(int(j[p]))
# print("Coarse:")
# print(classification_report(final, final_t))
# for i in range(4):
# print("Fine", i)
# return fin_outputs, fin_targets
for epoch in range(EPOCHS):
out, tar = train(epoch)
# break
out[0:10], tar[0:10]
# Creating the dataset and dataloader for the neural network
model = torch.load('/scratch/epoch_4')
# train_size = 0.8
# test_data=test.sample(frac=1,random_state=200)
# test_data=train.drop(train_data.index).reset_index(drop=True)
test_data = test.reset_index(drop=True)
testing = MultiLabelDataset(test_data, tokenizer, MAX_LEN, t=True)
test_params = {'batch_size': VALID_BATCH_SIZE,
'shuffle': False,
'num_workers': 0
}
testing_loader = DataLoader(testing, **test_params)
model.eval()
fin_targets=[]
fin_outputs=[]
# print(f'Epoch: {epoch}, Loss: {total_train_loss/count}')
with torch.no_grad():
for _, data in tqdm(enumerate(testing_loader, 0)):
ids = data['ids'].to(device, dtype = torch.long)
mask = data['mask'].to(device, dtype = torch.long)
token_type_ids = data['token_type_ids'].to(device, dtype = torch.long)
h_ids = data['h_ids'].to(device, dtype = torch.long)
h_mask = data['h_mask'].to(device, dtype = torch.long)
h_token_type_ids = data['h_token_type_ids'].to(device, dtype = torch.long)
# targets = data['targets'].to(device, dtype = torch.long)
emoji = data['emoji'].to(device, dtype = torch.long)
outputs = model(ids, mask, token_type_ids, h_ids, h_mask, h_token_type_ids, emoji)
# fin_targets.extend(targets.cpu().detach().numpy().tolist())
fin_outputs.extend(torch.sigmoid(outputs).cpu().detach().numpy().tolist())
fin_outputs = list(np.argmax(np.array(fin_outputs), axis=1).flatten())
# print(classification_report(fin_outputs, fin_targets))
fin_outputs[0:20]
test_data.head(10)
test['label'] = np.array(fin_outputs)
len(fin_outputs)
len(test.full_tweet.values)
test.sample(10)
def label_decode(val):
return labels[val]
test.label = test.label.apply(label_decode)
test.to_csv(path_or_buf='answers2.txt', index=False, columns = ['tweet_id', 'label'] )
```
| github_jupyter |
Deep learning algorithms fail to work well if we have only one training example.
One-shot learning is a classification or object categorization task in which one or a few examples are used to classify many new examples.
The principle behind one-shot learning is Humans learn new concepts with very little supervision.
### Problem in COnvNet
- A small training set is really not enough to train a robust neural network for this task. The feature vectors trained do not contain important information that can be used for the recognition of future images.
- Retraining the ConvNet every time the number of classes or dataset is increased is way too time and resource consuming.
### Solutions to one-shot learning
#### 1. Siamese network for one-shot learning

#### Siamese networks are based on a similarity function, which does not require extensive training
#### Takes input two images—one being the actual image and other the candidate image—and outputs the similarity between the two.
The two input images, if very similar, output = lower value
The two input images, if not similar, output = Higher value
degree of difference between the two images is compared with the threshold value(𝜏) (which is a hyperparameter),
if degree of difference between 2 image < threshold -> output = same person
if degree of difference between 2 image > threshold -> output = different person
#### Both the images to be compared are passed through the same networks, called sister networks, having the same parameters and shared weights.
images are passed through a sequence of convolutional, pooling, and fully connected layers, and end up with a feature vector of a fixed size, say, 128 denoted by f(x1) as an encoding of the input image x1.
```
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Conv2D, ZeroPadding2D, Activation, Input, concatenate
from tensorflow.keras.models import Model
from tensorflow.keras.layers import BatchNormalization
from tensorflow.keras.layers import MaxPooling2D, AveragePooling2D
from tensorflow.keras.layers import Concatenate
from tensorflow.keras.layers import Lambda, Flatten, Dense
from tensorflow.keras.initializers import glorot_uniform
from tensorflow.keras.layers import Layer
from tensorflow.keras import backend as K
K.set_image_data_format('channels_last')
import os
import numpy as np
from numpy import genfromtxt
import tensorflow as tf
import PIL
from tensorflow.keras.models import model_from_json
json_file = open('faceNet/model.json', 'r')
print('json_file : \n',json_file)
loaded_model_json = json_file.read()
print('loaded_model_json : \n',loaded_model_json)
json_file.close()
model = model_from_json(loaded_model_json)
print('model: \n',model)
model.load_weights('faceNet/model.h5')
print('model weights: \n',model.load_weights('faceNet/model.h5'))
# detecting the multiple faces in the image
# detectMultiScale - Detects objects of different sizes in the input image. The detected objects are returned as a list of rectangles
faces = faceClassifier.detectMultiScale(grayImage)
# print(faces) display the detected images x axis, y axis and image width and height
# print("\nNo. of Faces Found :",len(faces)) # printing the No of faces detected using detectMultiScale
#saving every face which is detected in previous steps
for (x, y, w, h) in faces:
# drawing a rectangle on any image
cv2.rectangle(image, (x, y), (x + w, y + h), (0, 255, 0), 2)
# roi - region of intreset using slicing the y and x axis along with height and width
roi_color = image[y:y + h, x:x + w]
# saving the croped image
cv2.imwrite('./images/'+str(w) + str(h) + '_faces.jpg', roi_color)
# saving the detected image
cv2.imwrite('detected_faces.jpg', image)
```
## Croping and saving the detected faces by using the face classifier
```
def face_scrap(imagePath):
#imagePath contains the image from which the face needs to be extracted
image = cv2.imread(imagePath)
# print(image.shape) -> result -> (194, 259, 3)
# plt.imshow(image) # displaying the image
# converting the image into gray scale image
grayImage = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
# print(gray.shape) -> result -> (194, 259)
# plt.imshow(gray) # displaying the image
# using haarcascade_frontalface_default.xml for classfing(detection) the face
faceClassifier = cv2.CascadeClassifier(cv2.data.haarcascades + "haarcascade_frontalface_default.xml")
# print(faceClassfier)
# detecting the multiple faces in the image
# detectMultiScale - Detects objects of different sizes in the input image. The detected objects are returned as a list of rectangles
faces = faceClassifier.detectMultiScale(grayImage)
# print(faces) display the detected images x axis, y axis and image width and height
# print("\nNo. of Faces Found :",len(faces)) # printing the No of faces detected using detectMultiScale
#saving every face which is detected in previous steps
for (x, y, w, h) in faces:
# drawing a rectangle on any image
cv2.rectangle(image, (x, y), (x + w, y + h), (0, 255, 0), 2)
# roi - region of intreset using slicing the y and x axis along with height and width
roi_image = image[y:y + h, x:x + w]
# saving the croped image
cv2.imwrite('./images/'+str(w) + str(h) + '_faces.jpg', roi_image)
# saving the detected image
cv2.imwrite('detected_faces.jpg', image)
imagePath = './faceDetect.jpg'
face_scrap(imagePath)
```
### face embedding
FaceNet is a model that, when given a picture of a face, will extract high-quality features from it and predict a 128-element vector representation of these features, called a face embedding. ace embeddings can then be used as the basis for training classifier systems on standard face recognition benchmark datasets.
# Loading FaceNet
### Triplet loss integration

#### compare pairs of images and learn the parameters of the neural network accordingly
one “anchor” image and get the distance between it and the “positive” (matching) image
distance of the anchor image with a “negative” (non-matching) example
Triplet loss is a loss function for machine learning algorithms where a baseline (anchor) input is compared to a positive (truthy) input and a negative (falsy) input.

```
### Contrastive loss for dimensionality reduction
#### Dimensionality reduction involves reducing the dimensions of the feature vector.
If the classes are the same, loss function encourages -> output = feature vectors that are similar
If the classes are the different, loss function encourages -> output = feature vectors that are less similar
```
| github_jupyter |
# MNIST Classifier Model
## Goal
Now that we have created a model that can classify 3's and 7'2, lets create a model for the entire MNIST dataset with all the numbers 0-9.
```
#hide
!pip install -Uqq fastbook
import fastbook
fastbook.setup_book()
#hide
from fastai.vision.all import *
from fastbook import *
matplotlib.rc('image', cmap='Greys')
```
## Getting data and viewing path
```
path = untar_data(URLs.MNIST)
path.ls()
(path/"training/1").ls()
```
### This is what the data looks like
```
t = (path/"training/1").ls()
t_1 = Image.open(t[0])
t_1
show_image(tensor(t_1))
```
## Loading data
To load our data, it may be beneficial to create a method to handle this process
```
#function used to load data appropriatly
def load_data(folder):
dataList = []
labelList = []
for num in range(10):
data_path = (path/folder/f'{num}').ls().sorted() #getting path
stackedData = torch.stack([tensor(Image.open(o)) for o in data_path]) #Open each image and stack them
stackedData = stackedData.float()/255.0 #squishing between 0-1
dataList.append(stackedData) #adding to dataList
labelList.extend([num]*len(data_path))#extending labelList
#Convert so that each image data is in each row
train_x = torch.cat(dataList).view(-1, 28*28)
train_y = tensor(labelList)
return train_x, train_y
train_x, train_y = load_data("training")
test_x, test_y = load_data("testing")
```
### Creating dataloaders (Minibatches)
```
train_dset = list(zip(train_x,train_y))
valid_dset = list(zip(test_x,test_y))
dl_train = DataLoader(train_dset, batch_size=256)
dl_test = DataLoader(valid_dset, batch_size=256)
```
## Below is the functions we need to train and test the model
Most of these functions are copies and pasted from our previous MNIST model. The difference here is the loss function, which was swapped out for cross entropy (As we have multiple categories). And, our accuracy function has been adjusted due to switching out sigmoid for softmax (softmax ranges all values between 0-1).
```
def calc_grad(xb, yb, model):
preds = model(xb)
loss = F.cross_entropy(preds, yb)
loss.backward()
def train_epoch(model):
for xb,yb in dl_train:
calc_grad(xb, yb, model)
for p in params:
p.data -= p.grad.data * lr
p.grad.zero_()
def batch_accuracy(xb, yb):
pred = xb.softmax(1)
return batch_accuracy_helper(pred, yb)/float(yb.size(0))
def batch_accuracy_helper(preds, yb):
return preds.argmax(dim=1).eq(yb).sum().float()
def validate_epoch(model):
accs = [batch_accuracy(model(xb), yb) for xb,yb in dl_test]
return round(torch.stack(accs).mean().item(), 4)
def linear_layer(xb):
return xb@w + b
def init_params(x, var=1.0):
return (torch.randn(x)*var).requires_grad_()
```
## Begin by initializing parameters
```
lr = 1.
w = init_params((28*28,10))
b = init_params(10)
params = w, b
w.shape, b.shape
```
## Now lets see if our loss improves for 1 epoch
It's good practice to try training the model manually to see if the loss improves. If it doesn't this means there may be some error.
```
validate_epoch(linear_layer)
train_epoch(linear_layer)
validate_epoch(linear_layer)
```
> Our loss improved, nice!
## Training
```
def train_model(model, epochs):
for i in range(epochs):
train_epoch(model)
print(validate_epoch(model), end=' ')
train_model(linear_layer, 20)
```
> 50% acc is not that bad, given there are 10 classes
## Using FastAI toolkit
As before, we can take everything we did above and condense it using FastAI's toolkit. Additionally, I will add non-linearity this time to see how much of a performance boost it gives.
```
dls = DataLoaders(dl_train, dl_test)
simple_net = nn.Sequential(
nn.Linear(28*28,30), #30 neurons
nn.ReLU(),
nn.Linear(30, 10) # 30neurons into 10 output neurons (10 classes)
)
learn = Learner(dls, simple_net, opt_func=SGD,
loss_func=F.cross_entropy, metrics=accuracy)
learn.fit(20, .01)
```
### So it seems that adding nonlinearity increased the accuracy by 10!
# Now lets try refining the learning rate
```
learn = Learner(dls, simple_net, opt_func=SGD,
loss_func=F.cross_entropy, metrics=accuracy)
learn.fine_tune(2, base_lr=0.1)
lr_min, lr_steep = learn.lr_find() #Finding best
print(f"Minimum/10: {lr_min:.2e}, steepest point: {lr_steep:.2e}")
learn.fine_tune(20, base_lr=3e-2) #Now lets train on the steepest
```
## Adjusting the LR improved acc by 25!
| github_jupyter |
```
import cv2
from matplotlib import pyplot as plt
import numpy as np
import imutils
import easyocr
from os import listdir
from os.path import isfile, join
img = cv2.imread(r"D:\5_Integrationsseminar\Aufnahmen\still2.jpg")
#img = cv2.imread(r"D:/5_Integrationsseminar/Bilder/small/KZE_008.jpg")
dir=r"D:\5_Integrationsseminar\Bilder\small"
#images = [f for f in listdir(dir) if isfile(join(dir, f))]
#C:/Users/Kilian/Notebook/5_Semster/test_plates_kaggle/IMG_4134.jpg
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
plt.imshow(cv2.cvtColor(gray, cv2.COLOR_BGR2RGB))
bfilter = cv2.bilateralFilter(gray, 11, 17, 17) #Noise reduction
edged = cv2.Canny(bfilter, 30, 100) #Edge detection
plt.imshow(cv2.cvtColor(edged, cv2.COLOR_BGR2RGB))
plt.show()
keypoints = cv2.findContours(edged.copy(), cv2.RETR_TREE, cv2.CHAIN_APPROX_SIMPLE)
contours = imutils.grab_contours(keypoints)
contours = sorted(contours, key=cv2.contourArea, reverse=True)[:10]
location = None
for contour in contours:
approx = cv2.approxPolyDP(contour, 10, True)
if len(approx) == 4:
location = approx
break
print(location)
mask = np.zeros(gray.shape, np.uint8)
new_image = cv2.drawContours(mask, [location], 0,255, -1)
new_image = cv2.bitwise_and(img, img, mask=mask)
#ab hier nummern
plt.imshow(cv2.cvtColor(new_image, cv2.COLOR_BGR2RGB))
#plt.show()
(x,y) = np.where(mask==255)
(x1, y1) = (np.min(x), np.min(y))
(x2, y2) = (np.max(x), np.max(y))
cropped_image = gray[x1:x2+1, y1:y2+1]
plt.imshow(cv2.cvtColor(cropped_image, cv2.COLOR_BGR2RGB))
plt.show()
###########################################################
###########################################################
reader = easyocr.Reader(['en'])
result = reader.readtext(cropped_image)
print(result)
print("---------------")
text=""
for x in result:
print(x[1])
print()
print("---------------")
text=text+x[1]
text
font = cv2.FONT_HERSHEY_SIMPLEX
res = cv2.putText(img, text=text, org=(approx[0][0][0], approx[1][0][1]+60), fontFace=font, fontScale=1, color=(0,255,0), thickness=2, lineType=cv2.LINE_AA)
res = cv2.rectangle(img, tuple(approx[0][0]), tuple(approx[2][0]), (0,255,0),3)
plt.imshow(cv2.cvtColor(res, cv2.COLOR_BGR2RGB))
plt.imsave(r"D:\5_Integrationsseminar\Aufnahmen\Result2.jpg",cv2.cvtColor(res, cv2.COLOR_BGR2RGB))
# Perform text extraction
invert=image
data = pytesseract.image_to_string(invert, lang='eng', config='--psm 6')
print(data)
cv2.imshow('thresh', thresh)
cv2.imshow('opening', opening)
cv2.imshow('invert', invert)
cv2.waitKey()
for i in dic_ocr2:
if i<biggest_box_last_layer:
dic_ocr.pop(i)
import cv2
import pytesseract
img = cv2.imread('image.jpg')
# Adding custom options
custom_config = r'--oem 3 --psm 6'
pytesseract.image_to_string(img, config=custom_config)
```
| github_jupyter |
```
# Get helper_functions.py script from course GitHub
!wget https://raw.githubusercontent.com/mrdbourke/tensorflow-deep-learning/main/extras/helper_functions.py
# Import helper functions we're going to use
from helper_functions import create_tensorboard_callback, plot_loss_curves, unzip_data, walk_through_dir
import os
! kaggle datasets download -d datamunge/sign-language-mnist
unzip_data('sign-language-mnist.zip')
walk_through_dir('sign_mnist_train')
import pandas as pd
df=pd.read_csv("/content/sign_mnist_train.csv")
df.head()
df["label"].value_counts()
df.shape
df_test=pd.read_csv("/content/sign_mnist_test.csv")
df_test.head()
df_test.shape
import numpy as np
df_train = np.array(df, dtype = 'float32')
df_test = np.array(df_test, dtype='float32')
class_names = ['A', 'B', 'C', 'D', 'E', 'F', 'G', 'H', 'I', 'J', 'K', 'L', 'M',
'N', 'O', 'P', 'Q', 'R', 'S', 'T', 'U', 'V', 'W', 'X', 'Y' ]
len(class_names)
import random
import matplotlib.pyplot as plt
i = random.randint(1,df.shape[0])
fig1, ax1 = plt.subplots(figsize=(3,3))
plt.imshow(df_train[i,1:].reshape((28,28)))
print("Label for the image is: ", class_names[int(df_train[i,0])])
X_train = df_train[:, 1:] /255.
X_test = df_test[:, 1:] /255.
X_train = X_train.reshape(X_train.shape[0], *(28, 28, 1))
X_test = X_test.reshape(X_test.shape[0], *(28, 28, 1))
print(X_train.shape)
print(X_test.shape)
from tensorflow.keras.utils import to_categorical
y_train = df_train[:, 0]
y_train = to_categorical(y_train, num_classes=25)
y_test = df_test[:, 0]
y_test = to_categorical(y_test, num_classes=25)
```
TensorFlow Model
```
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Conv2D, MaxPool2D, Flatten, Dense
model_0=Sequential([
Conv2D(16,3,activation='relu',input_shape=(28,28,1)),
MaxPool2D(),
tf.keras.layers.Dropout(0.2),
Conv2D(32,3,activation='relu'),
MaxPool2D(),
tf.keras.layers.Dropout(0.2),
Conv2D(64,3,activation='relu'),
MaxPool2D(),
tf.keras.layers.Dropout(0.2),
Flatten(),
Dense(64,activation='relu'),
Dense(25,activation='softmax')])
model_0.compile(loss="categorical_crossentropy",
optimizer=tf.keras.optimizers.Adam(),
metrics='accuracy')
history_0= model_0.fit(X_train, y_train, validation_data = (X_test, y_test), epochs= 10, batch_size= 32)
from tensorflow.keras.layers import Dense,MaxPooling2D,Conv2D,Dropout,Flatten
from tensorflow.keras.models import Sequential
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.regularizers import l2,l1
# Create Model
model1 = Sequential()
model1.add(Conv2D(32, (3, 3),padding="same", input_shape = (28,28,1), kernel_regularizer=l2(0.0002), activation='relu'))
model1.add(MaxPooling2D(pool_size = (2, 2)))
model1.add(Dropout(0.2))
model1.add(Conv2D(64, (3, 3), padding="same" ,kernel_regularizer=l2(0.0002), activation='relu'))
model1.add(MaxPooling2D(pool_size = (2, 2)))
model1.add(Dropout(0.2))
model1.add(Conv2D(128, (3, 3),padding="same" ,kernel_regularizer=l2(0.0002), activation='relu'))
model1.add(MaxPooling2D(pool_size = (2, 2)))
model1.add(Dropout(0.2))
model1.add(Flatten())
model1.add(Dense(128, activation = 'relu'))
model1.add(Dense(25, activation = 'softmax'))
# Compile
model1.compile(loss ='categorical_crossentropy', optimizer='adam',metrics =['acc'])
model1.summary()
history1 = model1.fit(X_train, y_train, batch_size = 128, epochs = 10, verbose = 1, validation_data = (X_test, y_test))
model1.save('Sign_language model.h5')
class_names = np.array(class_names)
class_names
preds = model1.predict(X_test)
preds=np.array(preds)
preds
preds.shape
prediction = np.argmax(preds, axis=1)
prediction
i = random.randint(1,len(prediction))
plt.imshow(X_test[i,:,:,0])
print("Predicted Label: ", class_names[int(prediction[i])])
```
| github_jupyter |
# Image classification using CNN
## Load the data
```
import pickle
import matplotlib.pyplot as plt
import tensorflow as tf
from os.path import join
from sklearn.preprocessing import OneHotEncoder
import numpy as np
def loadCifarData(basePath):
trainX = []
testX = []
trainY = []
testY = []
"""Load training data"""
for i in range(1, 6):
with open(join(basePath, "data_batch_%d" %i), "rb") as f:
dictionary = pickle.load(f, encoding = 'bytes')
trainX.extend(dictionary[b'data'])
trainY.extend(dictionary[b'labels'])
with open(join(basePath, "test_batch"), "rb") as f:
dictionary = pickle.load(f, encoding = 'bytes')
testX.extend(dictionary[b'data'])
testY.extend(dictionary[b'labels'])
return trainX, trainY, testX, testY
def toImage(array, rows = 32, columns = 32):
return array.reshape(3, rows, columns).transpose([1, 2, 0])
def toData(img, rows = 32, columns = 32):
return img.transpose([-1, -2, 0]).flatten()
def plotImages(rows, columns, data, convert = True):
fig, ax = plt.subplots(nrows=rows, ncols=columns)
if rows == 1:
ax = [ax]
if columns == 1:
ax = [ax]
index = 0
for row in ax:
for col in row:
if convert:
col.imshow(toImage(data[index]))
else:
col.imshow(data[index])
index = index + 1
trainRawX, trainRawY, testX, testY = loadCifarData("Data")
encoder = OneHotEncoder()
trainRawY = encoder.fit_transform(np.array(trainRawY).reshape(-1,1)).todense()
testY = encoder.transform(np.array(testY).reshape(-1,1)).todense()
plotImages(3, 3, trainRawX)
```
## Data Augmentation
### Flip images
```
import numpy as np
def flipImage(srcImage):
flippedImages = []
flippedImages.append(np.fliplr(srcImage))
flippedImages.append(np.flipud(srcImage))
flippedImages.append(np.flipud(np.fliplr(srcImage)))
return flippedImages
flipped = flipImage(toImage(trainRawX[1]))
flipped.append(toImage(trainRawX[1]))
plotImages(2, 2, flipped, False)
```
### Change Brightness
```
import cv2
def changeBrightness(image):
image = cv2.cvtColor(image,cv2.COLOR_RGB2HSV)
image = np.array(image, dtype = np.float64)
randomBrightness = .5+np.random.uniform()
image[:,:,2] = image[:,:,2]*randomBrightness
image[:,:,2][image[:,:,2]>255] = 255
image = np.array(image, dtype = np.uint8)
image = cv2.cvtColor(image,cv2.COLOR_HSV2RGB)
return image
noisyImage = changeBrightness(toImage(trainRawX[1]))
plotImages(1, 2, [toImage(trainRawX[1]), noisyImage], False)
```
### Augment Image
```
def augmentImage(imageVector):
augmentedImages = []
rawImages = []
image = toImage(imageVector)
flippedImages = flipImage(image)
flippedImages.append(image)
coinTossOutcome = np.random.binomial(1, 0.5, len(flippedImages))
for img, toss in zip(flippedImages, coinTossOutcome):
if toss == 1:
img = changeBrightness(img)
augmentedImages.append(img)
rawImages.append(toData(img))
return augmentedImages, rawImages
img, imgRaw = augmentImage(trainRawX[211])
plotImages(2, 2, img, False)
```
## Batch Data Iterator
```
from random import shuffle
def batchIterator(x, y, batchSize, batchCount):
size = len(x)
if batchSize * batchCount > size:
raise ValueError("Change batch size or change batch count")
indices = list(range(0, size))
shuffle(indices)
indices = indices[0:batchSize * batchCount]
batches = np.array_split(indices, batchCount)
for batch in batches:
yield (x[batch], y[batch])
```
## Prepare data for training
```
trainX = []
trainY = []
for x, y in zip(trainRawX, trainRawY):
rawAugmentedImages = augmentImage(x)[0]
trainX.extend(rawAugmentedImages)
target = [y for i in range(0, len(rawAugmentedImages))]
trainY.extend(target)
print(len(trainRawX))
print(len(trainX))
print(trainRawY.shape)
print(len(trainY))
trainX = np.stack(trainX, axis=0)
trainY = np.stack(trainY, axis=0)
processedTestX = []
processedTestY = []
for x, y in zip(testX, testY):
processedTestY.append(y)
processedTestX.append(toImage(x))
processedTestX = np.stack(processedTestX, axis=0)
processedTestY = np.stack(processedTestY, axis=0)
```
### Helper methods
```
def createConvolutionLayer(inputLayer, kernelHeight, kernelWidth, channelSize, kernelCount, strideX, strideY):
"""This will create a four dimensional tensor
In this tensor the first and second dimension define the kernel height and width
The third dimension define the channel size. If the input layer is
first layer in neural network then the channel size will be 3 in case of RGB images
else 1 if images are grey scale. Furthermore if the input layer is Convolution layer
then the channel size should be no of kernels in previous layer"""
weights = tf.Variable(tf.truncated_normal([kernelHeight, kernelWidth, channelSize, kernelCount], stddev=0.03))
bias = tf.Variable(tf.constant(0.05, shape=[kernelCount]))
"""Stride is also 4 dimensional tensor
The first and last values should be 1 as they represent the image index and
chanel size padding. Second and Third index represent the X and Y strides"""
layer = tf.nn.conv2d(input = inputLayer, filter = weights, padding='SAME',
strides = [1, strideX, strideY, 1]) + bias
return layer
def flattenLayer(inputLayer):
"""Flatten layer. The first component is image count which is useless"""
flattenedLayer = tf.reshape(inputLayer, [-1, inputLayer.get_shape()[1:].num_elements()])
return flattenedLayer
def fullyConnectedLayer(inputLayer, outputLayerCount):
weights = tf.Variable(tf.truncated_normal(
[int(inputLayer.get_shape()[1]), outputLayerCount], stddev=0.03))
bias = tf.Variable(tf.constant(0.05, shape=[outputLayerCount]))
layer = tf.matmul(inputLayer, weights) + bias
return layer
def batchNormalization(inputLayer, isTraining):
beta = tf.Variable(tf.constant(0.0, shape=[inputLayer.get_shape()[-1]]), trainable=True)
gamma = tf.Variable(tf.constant(1.0, shape=[inputLayer.get_shape()[-1]]), name='gamma', trainable=True)
batchMean, batchVariance = tf.nn.moments(inputLayer, [0,1,2], name='moments')
ema = tf.train.ExponentialMovingAverage(decay=0.5)
def meanVarianceUpdate():
emaOp = ema.apply([batchMean, batchVariance])
with tf.control_dependencies([emaOp]):
return tf.identity(batchMean), tf.identity(batchVariance)
mean, var = tf.cond(isTraining, meanVarianceUpdate, lambda: (ema.average(batchMean), ema.average(batchVariance)))
normed = tf.nn.batch_normalization(inputLayer, mean, var, beta, gamma, 1e-3)
return normed
def log_histogram(writer, tag, values, step, bins=1000):
# Convert to a numpy array
values = np.array(values)
# Create histogram using numpy
counts, bin_edges = np.histogram(values, bins=bins)
# Fill fields of histogram proto
hist = tf.HistogramProto()
hist.min = float(np.min(values))
hist.max = float(np.max(values))
hist.num = int(np.prod(values.shape))
hist.sum = float(np.sum(values))
hist.sum_squares = float(np.sum(values**2))
# Requires equal number as bins, where the first goes from -DBL_MAX to bin_edges[1]
# See https://github.com/tensorflow/tensorflow/blob/master/tensorflow/core/framework/summary.proto#L30
# Thus, we drop the start of the first bin
bin_edges = bin_edges[1:]
# Add bin edges and counts
for edge in bin_edges:
hist.bucket_limit.append(edge)
for c in counts:
hist.bucket.append(c)
# Create and write Summary
summary = tf.Summary(value=[tf.Summary.Value(tag=tag, histo=hist)])
writer.add_summary(summary, step)
writer.flush()
```
### Define model
```
"""Input is 4 dimensional tensor -1 so that the no of images can be infered on itself"""
inputLayer = tf.placeholder(tf.float32, [None, 32, 32, 3])
yTrue = tf.placeholder(tf.float32, shape=[None, 10])
isTraining = tf.placeholder(tf.bool, [])
convolutionLayer1 = createConvolutionLayer(inputLayer, 2, 2, 3, 30, 1, 1)
seluActivatedLayer1 = tf.nn.selu(convolutionLayer1)
poolingLayer1 = tf.nn.max_pool(value=convolutionLayer1, ksize=[1, 1, 2, 1], strides = [1, 1, 1, 1], padding='SAME')
bn1 = batchNormalization(poolingLayer1, isTraining)
poolingLayer2 = tf.nn.max_pool(value=bn1, ksize=[1, 1, 2, 1], strides = [1, 1, 1, 1], padding='SAME')
flattened = flattenLayer(poolingLayer2)
fc1 = fullyConnectedLayer(flattened, 950)
seluActivatedLayer2 = tf.nn.selu(fc1)
fc2 = fullyConnectedLayer(flattened, 500)
seluActivatedLayer3 = tf.nn.selu(fc2)
fc= fullyConnectedLayer(seluActivatedLayer3, 10)
```
### Define Predictions
```
predictions = tf.argmax(tf.nn.softmax(fc), axis = 1)
actual = tf.argmax(yTrue, axis = 1)
```
### Define loss function and specify the optimizer
```
loss = tf.nn.softmax_cross_entropy_with_logits_v2(logits=fc, labels = yTrue)
costFunction = tf.reduce_mean(loss)
optimizer = tf.train.AdamOptimizer().minimize(costFunction)
accuracy = tf.reduce_mean(tf.cast(tf.equal(predictions, actual), tf.float32))
```
### Create session and initialize global variables
```
session = tf.Session()
"""Initialize the global variables"""
session.run(tf.global_variables_initializer())
summaryWriter = tf.summary.FileWriter("tensorboard/structure2/logs", graph=tf.get_default_graph())
trainAccList = []
testAccList = []
for i in range(0, 50):
print("Epoch"+str(i))
summary = tf.Summary()
for x, y in batchIterator(trainX, trainY, 500, 50):
session.run(optimizer, feed_dict={inputLayer:x, yTrue:y, isTraining:True})
loss = session.run(costFunction, feed_dict={inputLayer:x, yTrue:y, isTraining:False})
acc = session.run(accuracy, feed_dict={inputLayer:x, yTrue:y, isTraining:False})
summary.value.add(tag = "TrainingLoss", simple_value = loss)
summary.value.add(tag = "TrainingAcc", simple_value = acc)
trainAccList.append(acc)
lossTestList = []
accTestList = []
for x, y in batchIterator(processedTestX, processedTestY, 1000, 5):
lossTest = session.run(costFunction, feed_dict={inputLayer:x, yTrue:y, isTraining:False})
accTest = session.run(accuracy, feed_dict={inputLayer:x, yTrue:y, isTraining:False})
lossTestList.append(lossTest)
accTestList.append(accTest)
summary.value.add(tag = "TestLoss", simple_value = np.mean(lossTestList))
summary.value.add(tag = "TestAcc", simple_value = np.mean(accTestList))
testAccList.append(np.mean(accTestList))
summaryWriter.add_summary(summary, i)
log_histogram(summaryWriter, "TrainAccHist", trainAccList, 50)
log_histogram(summaryWriter, "TestAccHist", testAccList, 50)
session.close()
```
| github_jupyter |
# Machine learning with SPARK in SQL Server 2019 Big Data Cluster
Spark in Unified Big data compute engine that enables big data processing, Machine learning and AI
Key Spark advantages are
1. Distributed compute enging
2. Choice of langauge (Python, R, Scala, Java)
3. Single engine for Batch and Streaming jobs
In this tutorial we'll cover how we can use Spark to create and deploy machine learning models. The example is a python(PySpark) sample. The same can also be done using Scala and R ( SparkR) in Spark.
<img src = "C:\repos\sql-server-samples\samples\features\sql-big-data-cluster\spark\sparkml\Train_Score_Export_with_Spark.jpg" style="float: center;" alt="drawing" width="900">
## Steps
1. Explore your Data
2. Data Prep and split Data as Training and Test set
3. Model Training
4. Model Scoring
5. Persist as Spark Model
6. Persist as Portable Model
E2E machine learning involves several additional step e.g data exploration, feature selection and principal component analysis,model selection etc. Many of these steps are ignored here for brevity.
## Step 1 - Explore your data
### Load the data
For this example we'll use **AdultCensusIncome** data from [here]( https://amldockerdatasets.azureedge.net/AdultCensusIncome.csv ). From your Azure Data Studio connect to the HDFS/Spark gateway and create a directory called spark_data under HDFS.
Download [AdultCensusIncome.csv]( https://amldockerdatasets.azureedge.net/AdultCensusIncome.csv ) to your local machine and upload to HDFS.Upload AdultCensusIncome.csv to the folder we created.
### Exploratory Analysis
- Baisc exploration on the data
- Labels & Features
1. **Label** - This refers to predicted value. This is represented as a column in the data. Label is **income**
2. **Features** - This refers to the characteristics that are used to predict. **age** and **hours_per_week**
Note : In reality features are chosen by applying some correlations techniques to understand what best characterize the Label we are predicting.
### The Model we will build
In AdultCensusIncome.csv contains several columsn like Income range, age, hours-per-week, education, occupation etc. We'll build a model that can predict income range would be >50K or <50K.
```
datafile = "/spark_data/AdultCensusIncome.csv"
#Read the data to a spark data frame.
data_all = spark.read.format('csv').options(header='true', inferSchema='true', ignoreLeadingWhiteSpace='true', ignoreTrailingWhiteSpace='true').load(datafile)
print("Number of rows: {}, Number of coulumns : {}".format(data_all.count(), len(data_all.columns)))
data_all.printSchema()
#Replace "-" with "_" in column names
columns_new = [col.replace("-", "_") for col in data_all.columns]
data_all = data_all.toDF(*columns_new)
data_all.printSchema()
#Basic data exploration
##1. Sub set the data and print some important columns
print("Select few columns to see the data")
data_all.select(['income','age','hours_per_week']).show(10)
## Find the number of distict values
print("Number of distinct values for income")
ds_sub = data_all.select('income').distinct()
ds_sub.show()
##Add a numberic column(income_code) derived from income column
print("Added numeric column(income_code) derived from income column")
from pyspark.sql.functions import expr
df_new = data_all.withColumn("income_code", expr("case \
when income like '%<=50K%' then 0 \
when income like '%>50K%' then 1 \
else 2 end "))
df_new.select(['income','age','hours_per_week','income_code']).show(10)
##Summary statistical operations on dataframe
print("Print a statistical summary of a few columns")
df_new.select(['income','age','hours_per_week','income_code']).describe().show()
print("Calculate Co variance between a few columns to understand features to use")
mycov = df_new.stat.cov('income_code','hours_per_week')
print("Covariance between income and hours_per_week is", round(mycov,1))
mycov = df_new.stat.cov('income_code','age')
print("Covariance between income and age is", round(mycov,1))
# Choose feature columns and the label column.
label = "income"
xvars = ["age", "hours_per_week"] #all numeric
print("label = {}".format(label))
print("features = {}".format(xvars))
#Check label counts to check data bias
print("Count of rows that are <=50K", data_all[data_all.income=="<=50K"].count())
print("Count of rows that are >50K", data_all[data_all.income==">50K"].count())
select_cols = xvars
select_cols.append(label)
data = data_all.select(select_cols)
```
## Step 2 - Split as training and test set
We'll use 75% of rows to train the model and rest of the 25% to evaluate the model. Additionally we persist the train and test data sets to HDFS storage. The step is not necessary , but shown to demonstrate saving and loading with ORC format. Other format e.g. Parquet may also be used. Post this step you should see 2 directories created with the name "AdultCensusIncomeTrain" and "AdultCensusIncomeTest"
```
train, test = data.randomSplit([0.75, 0.25], seed=123)
print("train ({}, {})".format(train.count(), len(train.columns)))
print("test ({}, {})".format(test.count(), len(test.columns)))
train_data_path = "/spark_ml/AdultCensusIncomeTrain"
test_data_path = "/spark_ml/AdultCensusIncomeTest"
train.write.mode('overwrite').orc(train_data_path)
test.write.mode('overwrite').orc(test_data_path)
print("train and test datasets saved to {} and {}".format(train_data_path, test_data_path))
```
## Step 3 - Train a model
[Spark ML pipeline] ( https://spark.apache.org/docs/2.3.1/ml-pipeline.html ) allow to sequence all steps as a workflow and make it easier to experiment with various algorithms and their parameters. The following code first constructs the stages and then puts these stages together in Ml pipeline. LogisticRegression is used to create the model.
```
from pyspark.ml import Pipeline, PipelineModel
from pyspark.ml.feature import OneHotEncoder, StringIndexer, VectorAssembler
from pyspark.ml.classification import LogisticRegression
reg = 0.1
print("Using LogisticRegression model with Regularization Rate of {}.".format(reg))
# create a new Logistic Regression model.
lr = LogisticRegression(regParam=reg)
dtypes = dict(train.dtypes)
dtypes.pop(label)
si_xvars = []
ohe_xvars = []
featureCols = []
for idx,key in enumerate(dtypes):
if dtypes[key] == "string":
featureCol = "-".join([key, "encoded"])
featureCols.append(featureCol)
tmpCol = "-".join([key, "tmp"])
si_xvars.append(StringIndexer(inputCol=key, outputCol=tmpCol, handleInvalid="skip")) #, handleInvalid="keep"
ohe_xvars.append(OneHotEncoder(inputCol=tmpCol, outputCol=featureCol))
else:
featureCols.append(key)
# string-index the label column into a column named "label"
si_label = StringIndexer(inputCol=label, outputCol='label')
# assemble the encoded feature columns in to a column named "features"
assembler = VectorAssembler(inputCols=featureCols, outputCol="features")
stages = []
stages.extend(si_xvars)
stages.extend(ohe_xvars)
stages.append(si_label)
stages.append(assembler)
stages.append(lr)
pipe = Pipeline(stages=stages)
print("Pipeline Created")
model = pipe.fit(train)
print("Model Trained")
print("Model is ", model)
print("Model Stages", model.stages)
```
## Step 4 - Model scoring
Predict using the model and Evaluate the model accuracy
The code below use test data set to predict the outcome using the model created in the step above. We measure accuracy of the model with areaUnderROC and areaUnderPR metric.
```
from pyspark.ml.evaluation import BinaryClassificationEvaluator
# make prediction
pred = model.transform(test)
# evaluate. note only 2 metrics are supported out of the box by Spark ML.
bce = BinaryClassificationEvaluator(rawPredictionCol='rawPrediction')
au_roc = bce.setMetricName('areaUnderROC').evaluate(pred)
au_prc = bce.setMetricName('areaUnderPR').evaluate(pred)
print("Area under ROC: {}".format(au_roc))
print("Area Under PR: {}".format(au_prc))
pred[pred.prediction==1.0][pred.income,pred.label,pred.prediction].show()
```
## Step 5 - Persist the Spark Models
Finally we persist the model in HDFS for later use. Post this step the created model get saved as /spark_ml/AdultCensus.mml
```
model_name = "AdultCensus.mml"
model_fs = "/spark_ml/" + model_name
model.write().overwrite().save(model_fs)
print("saved model to {}".format(model_fs))
# load the model file and check its same as the in-memory model
model2 = PipelineModel.load(model_fs)
assert str(model2) == str(model)
print("Successfully loaded from {}".format(model_fs))
```
## Step 6 - Persist as Portable Model
Here we persist the Model in as Portable Mleap bundle for use outside Spark.
```
import os
from mleap.pyspark.spark_support import SimpleSparkSerializer
# serialize the model to a zip file in JSON format
model_name_export = "adult_census_pipeline.zip"
model_name_path = os.getcwd()
model_file = os.path.join(model_name_path, model_name_export)
# remove an old model file, if needed.
try:
os.remove(model_file)
except OSError:
pass
model_file_path = "jar:file:{}".format(model_file)
model.serializeToBundle(model_file_path, model.transform(train))
print("persist the mleap bundle from local to hdfs")
from subprocess import Popen, PIPE
proc = Popen(["hadoop", "fs", "-put", "-f", model_file, os.path.join("/spark_ml", model_name_export)], stdout=PIPE, stderr=PIPE)
s_output, s_err = proc.communicate()
```
| github_jupyter |
```
print('The Station {}'.format(station_id)+' has {} docks in total.'.format(station.loc[station_id,'install_dockcount']))
#Station by station
extract = trip.loc[(trip.from_station_id==station_id) | (trip.to_station_id==station_id),:]
def incrementation(row):
if (row['from_station_id']==station_id)&(row['to_station_id']==station_id):
return int(0)
if (row['from_station_id']==station_id):
return int(-1)
if (row['to_station_id']==station_id):
return int(1)
extract['incrementation'] = trip.apply(incrementation, axis=1)
extract = extract.set_index('trip_id')
#Start and Stop
temp1 = extract.loc[(extract.incrementation==0),['starttime','stoptime','bikeid','to_station_id','incrementation']]
instanteanous_variation = pd.DataFrame(columns=['trip_id','time', 'bikeid', 'destination_id', 'incrementation'])
for i in range(temp1.shape[0]):
#-1
serie1 = dict(trip_id=temp1.index[i],bikeid=temp1.bikeid.values[i],destination_id=temp1.to_station_id.values[i])
serie1['incrementation'] = -1
serie1['time'] = temp1.starttime.values[i]
#+1
serie2 = dict(trip_id=temp1.index[i],bikeid=temp1.bikeid.values[i],destination_id=temp1.to_station_id.values[i])
serie2['incrementation'] = 1
serie2['time'] = temp1.stoptime.values[i]
instanteanous_variation = instanteanous_variation.append(serie1, ignore_index=True)
instanteanous_variation = instanteanous_variation.append(serie2, ignore_index=True)
instanteanous_variation = instanteanous_variation.set_index('trip_id')
instanteanous_variation.index = instanteanous_variation.index.astype(int)
#Stop
temp2 = extract.loc[(extract.incrementation==1.0),['stoptime','bikeid','from_station_id','incrementation']]
temp2.columns=['time','bikeid','destination_id','incrementation']
instanteanous_variation=instanteanous_variation.append(temp2)
#Start
temp3 = extract.loc[(extract.incrementation==-1.0),['starttime','bikeid','to_station_id','incrementation']]
temp3.columns=['time','bikeid','destination_id','incrementation']
instanteanous_variation=instanteanous_variation.append(temp3)
#Sort by time before doing cumulative
instanteanous_variation.time = pd.to_datetime(instanteanous_variation.time)
instanteanous_variation = instanteanous_variation.sort_values('time')
#Computation of the total cumulative variation
instanteanous_variation['total_variation'] = instanteanous_variation['incrementation'].cumsum()
date_start = date(2014,10,13)
date_end = date(2016,8,31)
dates = [date_start + timedelta(days=x) for x in range((date_end-date_start).days + 1)]
daily = []
for d in dates:
temp = instanteanous_variation.loc[(instanteanous_variation.time.dt.date==d),['incrementation']].cumsum().values
daily = np.append(daily,temp)
instanteanous_variation['daily_variation'] = pd.Series(daily, index=instanteanous_variation.index)
###### Resample of instanteanous_variation towards regular time step #####
sampled_variation = instanteanous_variation.set_index('time').incrementation.groupby(pd.TimeGrouper(freq='15Min')).sum() #every 15Mins
sampled_variation = pd.DataFrame(sampled_variation, columns=['incrementation']) #to dataframe
sampled_variation = sampled_variation.fillna(value=0) #transform NaN to 0
sampled_variation['date'] = sampled_variation.index.date #faster access to date in the following
#From incrementation to daily_variation
daily = []
for d in dates:
temp = sampled_variation.loc[(sampled_variation.date==d),['incrementation']].cumsum().values
daily = np.append(daily,temp)
sampled_variation['incrementation'] = pd.Series(daily, index=sampled_variation.index)
sampled_variation.columns = ['daily_variation','date']
#Removal of first and last day to have full periods of 24h (96 by day)
sampled_variation=sampled_variation.loc[(sampled_variation.date!=date_start)&(sampled_variation.date!=date_end),:]
###### Concatenate dataframes to regress on later ####
columns_weather = ['Events','Mean_Temperature_F','Precipitation_In '] #weather data we will use for the regression
repeat_weather = pd.concat([weather.loc[(weather.index.date!=date_start)&(weather.index.date!=date_end),columns_weather]]*96).sort_index(axis=0)# repeated 96 times every day
repeat_weather.index=sampled_variation.index #same index to ease concatenation
data_to_regress = pd.concat([repeat_weather,sampled_variation],axis=1) #original data to regress on (need then to be numerized)
###### Adding a few useful features ######
data_to_regress['date'] = data_to_regress.index.month
data_to_regress['weekday'] = data_to_regress.index.dayofweek
data_to_regress['hour'] = data_to_regress.index.hour + data_to_regress.index.minute/60
data_to_regress.columns = ['Events','Mean_Temperature_F','Precipitation_In ','daily_variation','month','weekday','hour']
###### Numerizing Events #####
data_to_regress.Events = data_to_regress.Events.fillna(value=0)
to_one = ['Fog']
for k, st in enumerate(to_one):
data_to_regress.loc[(data_to_regress.Events == st),['Events']]=1
to_two = ['Rain','Fog , Rain','Fog-Rain', 'Rain-Thunderstorm','Rain , Thunderstorm']
for k, st in enumerate(to_two):
data_to_regress.loc[(data_to_regress.Events == st),['Events']]=2
to_three = ['Snow','Rain-Snow','Rain , Snow']
for k, st in enumerate(to_three):
data_to_regress.loc[(data_to_regress.Events == st),['Events']]=3
###### More cleaning ^^ #####
data_to_regress = data_to_regress.dropna(axis=0)
#print(data_to_regress.isnull().any())
#Printing
display(data_to_regress.head())
print('DataFrame shape used for regression: {}'.format(data_to_regress.shape))
```
| github_jupyter |
```
import pymc3 as pm
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
%qtconsole --colors=linux
plt.style.use('ggplot')
```
# Chapter 3 - Inferences with binomials
## 3.1 Inferring a rate
Inferring the rate $\theta$ of a binary process
$$ \theta \sim \text{Beta}(1, 1) $$
$$ k \sim \text{Binomial} ( \theta, n) $$
In the example, we set k = 5 and n = 10
```
# Data
k = np.array([5])
n = np.array([10])
with pm.Model() as model1:
# prior
theta = pm.Beta('theta', alpha=1, beta=1)
# observed
x = pm.Binomial('x', n=n, p=theta, observed=k)
# inference
trace1 = pm.sample(3e3)
pm.traceplot(trace1[:], varnames=['theta'])
```
And generate a picture that is identical to 3.2 one on page 39 of Wagenmakers, 2013:
```
from scipy.stats.kde import gaussian_kde # for plotting: to calculate a continuous
# approximation of the posterior and prior densities.
my_pdf = gaussian_kde(trace1['theta'][:])
x=np.linspace(0, 1, 100)
_, axes = plt.subplots(1, 2, figsize=(18, 5))
axes[0].plot(x, my_pdf(x), 'r') # distribution function
axes[0].hist(trace1['theta'][:], bins=100, normed=1, alpha=.3)
plt.xlabel('Rate')
plt.ylabel('Posterior Density')
pm.plot_posterior(trace1['theta'][:], ax=axes[1])
plt.show()
pm.summary(trace1, varnames=['theta'])# gives the same credible interval as in the book.
```
## 3.2 Difference between two rates
Inferring the rate $\theta$ of two binary process
$$ \theta \sim \text{Beta}(1, 1) $$
$$ k \sim \text{Binomial} ( \theta, n) $$
In the example, we set k1 = 5, n1 = 10 and k2 = 7, n2 = 10
The model involve a deterministic part in pymc3.
```
# data
k1, k2 = 5, 7
n1 = n2 = 10
with pm.Model() as model2:
# prior
theta1 = pm.Beta('theta1', alpha=1, beta=1)
theta2 = pm.Beta('theta2', alpha=1, beta=1)
# observed
x1 = pm.Binomial('x1', n=n1, p=theta1, observed=k1)
x2 = pm.Binomial('x2', n=n2, p=theta2, observed=k2)
# differences as deterministic
delta = pm.Deterministic('delta', theta1-theta2)
# inference
trace2 = pm.sample(3e3)
pm.traceplot(trace2[:])
pm.summary(trace2)# gives the credible interval
my_pdf = gaussian_kde(trace2['delta'])
x=np.linspace(-1, 1, 200)
plt.plot(x,my_pdf(x),'r') # distribution function
plt.hist(trace2['delta'],bins=100, normed=1,alpha=.3)
plt.xlabel('Difference in Rates')
plt.ylabel('Posterior Density')
plt.show()
```
## 3.3 Inferring a common rate
```
# Multiple trials
k = np.array([5, 7])
n = np.array([10, 10])
with pm.Model() as model3:
# prior
theta = pm.Beta('theta', alpha=1, beta=1)
# observed
x = pm.Binomial('x', n=n, p=theta, observed=k)
# inference
trace3 = pm.sample(3e3)
pm.traceplot(trace3, varnames=['theta'])
pm.summary(trace3)
my_pdf = gaussian_kde(trace3['theta'])
x = np.linspace(0.2, 1, 200)
plt.plot(x, my_pdf(x), 'r') # distribution function
plt.hist(trace3['theta'], bins=100, normed=1, alpha=.3)
plt.xlabel('Rate')
plt.ylabel('Posterior Density')
```
## 3.4 Prior and posterior prediction
```
k = 1
n = 15
# Uncomment for Trompetter Data
# k = 24
# n = 121
# prior only model - no observation
with pm.Model() as model_prior:
theta = pm.Beta('theta', alpha=1, beta=1)
x = pm.Binomial('x', n=n, p=theta)
trace_prior = pm.sample(3e3)
# with observation
with pm.Model() as model_prior:
theta = pm.Beta('theta', alpha=1, beta=1)
x = pm.Binomial('x', n=n, p=theta, observed=k)
trace_obs = pm.sample(3e3)
# prediction (sample from trace)
ppc = pm.sample_ppc(trace_obs, samples=500, model=model_prior)
prior_x = trace_prior['x']
pred_theta = trace_obs['theta']
plt.subplot(2, 1, 1)
my_pdf = gaussian_kde(pred_theta)
x=np.linspace(0, 1, 1000)
plt.plot(x, my_pdf(x), 'r', label='Posterior') # distribution function
from scipy.stats import beta
plt.plot(x, beta.pdf(x, 1, 1), 'b', label='Prior')
plt.xlabel('Rate')
plt.ylabel('Density')
predictx = ppc['x']
plt.subplot(2, 1, 2)
plt.hist(predictx, normed=1, bins=len(np.unique(predictx)),
alpha=.3, color='r', label='Posterior')
plt.hist(prior_x, normed=1, bins=n+1,
alpha=.3, color='b', label='Prior')
plt.xlabel('Success Count')
plt.ylabel('Mass')
plt.show()
```
## 3.5 Posterior Predictive
```
# Inferring a Common Rate, With Posterior Predictive
k1 = 2
n1 = 13
k2 = 10
n2 = 10
with pm.Model() as model5:
# prior
theta = pm.Beta('theta', alpha=1, beta=1)
# observed
x1 = pm.Binomial('x1', n=n2, p=theta, observed=k1)
x2 = pm.Binomial('x2', n=n2, p=theta, observed=k2)
# inference
trace5 = pm.sample(3e3)
pm.traceplot(trace5, varnames=['theta'])
# prediction (sample from trace)
ppc5 = pm.sample_ppc(trace5, samples=500, model=model5)
from matplotlib import gridspec
fig = plt.figure(figsize=(12, 4))
gs = gridspec.GridSpec(1,2, width_ratios=[2, 3])
ax0 = plt.subplot(gs[0])
my_pdf = gaussian_kde(trace5['theta'])
x = np.linspace(0.2, 1, 200)
ax0.plot(x, my_pdf(x), 'r') # distribution function
ax0.hist(trace5['theta'], bins=100, normed=1, alpha=.3)
plt.xlabel('Rate')
plt.ylabel('Posterior Density')
ax1 = plt.subplot(gs[1])
predx1 = ppc5['x1']
predx2 = ppc5['x2']
from scipy import sparse
A = sparse.csc_matrix((np.ones(len(predx1)), (predx1,predx2)),
shape=(n1+1,n2+1)).todense()
ax1.imshow(A, interpolation='none', alpha=.9, origin='lower')
ax1.scatter(k2, k1, s=100, c=[1,0,0])
plt.xlabel('Trial2')
plt.ylabel('Trial1')
plt.tight_layout()
```
## 3.6 Joint distributions
```
# the Survey example in the book
k = np.array([16,18,22,25,27])
nmax = 500
m = len(k)
with pm.Model() as model6:
# prior
theta = pm.Beta('theta', alpha=1,beta=1)
TotalN = pm.DiscreteUniform('TotalN', lower=1, upper=nmax)
# observed
x = pm.Binomial('x', n=TotalN, p=theta, observed=k)
# inference
trace6 = pm.sample(1e5, njobs=2)
pm.traceplot(trace6)
# First calculate MLE:
from scipy.special import *
burnin = 90000
thetapost = trace6['theta'][burnin:]
npost = trace6['TotalN'][burnin:]
cc = -float('Inf')
ind = 0
for i in range(0, len(npost)):
logL = 0
for j in k:
logL = logL+gammaln(npost[i]+1)-gammaln(j+1)-gammaln(npost[i] - j +1)
logL = logL+j*np.log(thetapost[i])+(npost[i]-j)*np.log(1-thetapost[i])
if logL > cc:
ind = i
cc = logL
# print(ind)
from matplotlib.ticker import NullFormatter
nullfmt = NullFormatter() # no labels
y = thetapost
x = npost
# definitions for the axes
left, width = 0.1, 0.65
bottom, height = 0.1, 0.65
bottom_h = left_h = left + width + 0.02
rect_scatter = [left, bottom, width, height]
rect_histx = [left, bottom_h, width, 0.2]
rect_histy = [left_h, bottom, 0.2, height]
# start with a rectangular Figure
plt.figure(1, figsize=(8, 8))
axScatter = plt.axes(rect_scatter)
axHistx = plt.axes(rect_histx)
axHisty = plt.axes(rect_histy)
# no labels
axHistx.xaxis.set_major_formatter(nullfmt)
axHisty.yaxis.set_major_formatter(nullfmt)
# the scatter plot:
axScatter.scatter(x, y, c=[1, 1, 1], alpha=.1)
axScatter.scatter(np.mean(x), np.mean(y), s=50, c=[1, 0, 0], alpha=1)
axScatter.scatter(x[ind], y[ind], s=50, c=[0, 1, 0], alpha=1)
# now determine nice limits by hand:
binwidth1 = 0.25
axScatter.set_xlim((0, nmax))
axScatter.set_ylim((0, 1))
bins1 = np.linspace(0, nmax, 20)
axHistx.hist(x, bins=bins1)
bins2 = np.linspace(0, 1, 20)
axHisty.hist(y, bins=bins2, orientation='horizontal')
axHistx.set_xlim(axScatter.get_xlim())
axHisty.set_ylim(axScatter.get_ylim())
plt.show()
plt.figure(1, figsize=(8, 8))
plt.hist2d(x, y, bins=50)
plt.show()
```
| github_jupyter |
# Introducción al Cálculo Científico
En esta clase introduciremos algunos conceptos de computación cientifica en Python, principalmente utilizando la biblioteca `NumPy`, piedra angular de otras librerías científicas.
## SciPy.org
**SciPy** es un ecosistema de software _open-source_ para matemática, ciencia y engeniería. Las principales bibliotecas son:
* NumPy: Arrays N-dimensionales. Librería base, integración con C/C++ y Fortran.
* SciPy library: Computación científica (integración, optimización, estadística, etc.)
* Matplotlib: Visualización 2D:
* IPython: Interactividad (Project Jupyter).
* SimPy: Matemática Simbólica.
* Pandas: Estructura y análisis de datos.
## Numpy
NumPy es el paquete fundamental para la computación científica en Python. Proporciona un objeto de matriz multidimensional, varios objetos derivados (como matrices y arreglos) y una variedad de rutinas para operaciones rápidas en matrices, incluida la manipulación matemática, lógica, de formas, clasificación, selección, I/O, transformadas discretas de Fourier, álgebra lineal básica, operaciones estadísticas básicas, simulación y mucho más. [Fuente.](https://numpy.org/devdocs/user/whatisnumpy.html)
Para comenzar, la forma usual de importar `NumPy` es utilizando el alias `np`. Lo verás así en una infinidad de ejmplos, libros, blogs, etc.
```
import numpy as np
```
### Lo básico
Los objetos principales de Numpy son los comúnmente conocidos como NumPy Arrays (la clase se llama `ndarray`), corresponden a una tabla de elementos, todos del mismo tipo, indexados por una tupla de enternos no-negativos. En NumPy, las dimensiones son llamadas `axes` (ejes) y su singular `axis` (eje), similar a un plano cartesiano generalizado. Esta parte de la clase está basada en el _Quickstart tutorial_ en la página oficial ([link](https://numpy.org/devdocs/user/quickstart.html)).
Instanciar un NumPy Array es simple es utilizando el constructor propio de la biblioteca.
```
a = np.array(
[
[ 0, 1, 2, 3, 4],
[ 5, 6, 7, 8, 9],
[10, 11, 12, 13, 14]
]
)
type(a)
```
Los atributos más importantes de un `ndarray` son:
```
a.shape # the dimensions of the array.
a.ndim # the number of axes (dimensions) of the array.
a.size # the total number of elements of the array.
a.dtype # an object describing the type of the elements in the array.
a.itemsize # the size in bytes of each element of the array.
```
### Crear Numpy Arrays
Hay varias formas de crear arrays, el constructor básico es el que se utilizó hace unos momentos, `np.array`. El _type_ del array resultante es inferido de los datos proporcionados.
```
a_int = np.array([2, 6, 10])
a_float = np.array([2.1, 6.1, 10.1])
print(f"a_int: {a_int.dtype.name}")
print(f"a_float: {a_float.dtype.name}")
```
También es posible utilizar otras estructuras de Python, como listas o tuplas.
```
a_list = [1, 1, 2, 3, 5]
np.array(a_list)
a_tuple = (1, 1, 1, 3, 5, 9)
np.array(a_tuple)
```
__¡Cuidado!__ Es fácil confundirse con las dimensiones o el tipo de argumento en los contructores de NumPy, por ejemplo, utilizando una lista podríamos crear un arreglo de una o dos dimensiones si no tenemos cuidado.
```
one_dim_array = np.array(a_list)
two_dim_array = np.array([a_list])
print(f"np.array(a_list) = {one_dim_array} tiene shape: {one_dim_array.shape}, es decir, {one_dim_array.ndim} dimensión(es).")
print(f"np.array([a_list]) = {two_dim_array} tiene shape: {two_dim_array.shape}, es decir, {two_dim_array.ndim} dimensión(es).")
```
Una funcionalidad útil son los constructores especiales a partir de constantes.
```
np.zeros((3, 4))
np.ones((2, 3, 4), dtype=np.int) # dtype can also be specified
np.identity(4) # Identity matrix
```
Por otro lado, NumPy proporciona una función análoga a `range`.
```
range(10)
type(range(10))
np.arange(10)
type(np.arange(10))
np.arange(3, 10)
np.arange(2, 20, 3, dtype=np.float)
np.arange(9).reshape(3, 3)
```
__Bonus:__ Utilizar `np.arange` tiene como _"ingredientes"_ el inicio (_start_), fin (_stop_) y el tamaño del espacio entre valores (_step_) y el largo (`len`) depende estos argumentos. Sin embargo, existe la función `np.linspace` que construye un `np.array` con un inicio y un fin, pero indicando la cantidad de elementos (y por lo tanto, el espaciado depende es este).
```
np.linspace(0, 100, 5)
```
Esto puede causar confusiones en ocasiones, pues recuerda que la indexación de Python (y por lo tanto NumPy) comienza en cero, por lo que si quieres replicar el `np.array` anterior con `np.arange` debes tener esto en consideración. Es decir:
```
np.arange(start=0, stop=100, step=25) # stop = 100
np.arange(start=0, stop=101, step=25) # stop = 101
```
No podía faltar la instanciación a través de elementos aleatorios
```
np.random.random(size=3) # Elementos entre 0 y 1
np.random.uniform(low=3, high=7, size=5) # Desde una distribución uniforme
np.random.normal(loc=100, scale=10, size=(2, 3)) # Desde una distribución normal indicando media y desviación estándar
```
### Acceder a los elementos de un array
Es muy probable que necesites acceder a elementos o porciones de un array, para ello NumPy tiene una sintáxis consistente con Python.
```
x1 = np.arange(0, 30, 4)
x2 = np.arange(0, 60, 3).reshape(4, 5)
print("x1:")
print(x1)
print("\nx2:")
print(x2)
x1[1] # Un elemento de un array 1D
x1[:3] # Los tres primeros elementos
x2[0, 2] # Un elemento de un array 2D
x2[0] # La primera fila
x2[:, 1] # Todas las filas y la segunda columna
x2[:, 1:3] # Todas las filas y de la segunda a la tercera columna
```
Nuevamente, recordar que Python tiene indexación partiendo desde cero. Además, la dimensión del arreglo también depende de la forma en que se haga la selección.
```
x2[:, 2]
x2[:, 2:3] # What?!
```
En el ejemplo anterior los valores son los mismos, pero las dimensiones no. En el primero se utiliza `indexing` para acceder a la tercera columna, mientras que en el segundo `slicing` para acceder desde la tercera columna a la tercera columna.
```
print(x2[:, 2].shape)
print(x2[:, 2:3].shape)
```
### Operaciones Básias
Numpy provee operaciones vectorizadas, con tal de mejorar el rendimiento de la ejecución.
Por ejemplo, pensemos en la suma de dos arreglos 2D.
```
A = np.random.random((5,5))
B = np.random.random((5,5))
```
Con los conocimientos de la clase pasada, podríamos pensar en iterar a través de dos `for`, con tal de llenar el arreglo resultando. algo así:
```
def my_sum(A, B):
n, m = A.shape
C = np.empty(shape=(n, m))
for i in range(n):
for j in range(m):
C[i, j] = A[i, j] + B[i, j]
return C
%timeit my_sum(A, B)
```
Pero la suma de `ndarray`s es simplemente con el signo de suma (`+`):
```
%timeit A + B
```
Para dos arrays tan pequeños la diferencia de tiempo es considerable, ¡Imagina con millones de datos!
Los clásicos de clásicos:
```
x = np.arange(5)
print(f"x = {x}")
print(f"x + 5 = {x + 5}")
print(f"x - 5 = {x - 5}")
print(f"x * 2 = {x * 2}")
print(f"x / 2 = {x / 2}")
print(f"x // 2 = {x // 2}")
print(f"x ** 2 = {x ** 2}")
print(f"x % 2 = {x % 2}")
```
¡Júntalos como quieras!
```
-(0.5 + x + 3) ** 2
```
Al final del día, estos son alias para funciones de Numpy, por ejemplo, la operación suma (`+`) es un _wrapper_ de la función `np.add`
```
np.add(x, 5)
```
Podríamos estar todo el día hablando de operaciones, pero básicamente, si piensas en alguna operación lo suficientemente común, es que la puedes encontrar implementada en Numpy. Por ejemplo:
```
np.abs(-(0.5 + x + 3) ** 2)
np.log(x + 5)
np.exp(x)
np.sin(x)
```
Para dimensiones mayores la idea es la misma, pero siempre hay que tener cuidado con las dimensiones y `shape` de los arrays.
```
print("A + B: \n")
print(A + B)
print("\n" + "-" * 80 + "\n")
print("A - B: \n")
print(A - B)
print("\n" + "-" * 80 + "\n")
print("A * B: \n")
print(A * B) # Producto elemento a elemento
print("\n" + "-" * 80 + "\n")
print("A / B: \n")
print(A / B) # División elemento a elemento
print("\n" + "-" * 80 + "\n")
print("A @ B: \n")
print(A @ B) # Producto matricial
```
### Operaciones Booleanas
```
print(f"x = {x}")
print(f"x > 2 = {x > 2}")
print(f"x == 2 = {x == 2}")
print(f"x == 2 = {x == 2}")
aux1 = np.array([[1, 2, 3], [2, 3, 5], [1, 9, 6]])
aux2 = np.array([[1, 2, 3], [3, 5, 5], [0, 8, 5]])
B1 = aux1 == aux2
B2 = aux1 > aux2
print("B1: \n")
print(B1)
print("\n" + "-" * 80 + "\n")
print("B2: \n")
print(B2)
print("\n" + "-" * 80 + "\n")
print("~B1: \n")
print(~B1) # También puede ser np.logical_not(B1)
print("\n" + "-" * 80 + "\n")
print("B1 | B2 : \n")
print(B1 | B2)
print("\n" + "-" * 80 + "\n")
print("B1 & B2 : \n")
print(B1 & B2)
```
### Broadcasting
¿Qué pasa si las dimensiones no coinciden? Observemos lo siguiente:
```
a = np.array([0, 1, 2])
b = np.array([5, 5, 5])
a + b
```
Todo bien, dos arrays 1D de 3 elementos, la suma retorna un array de 3 elementos.
```
a + 3
```
Sigue pareciendo normal, un array 1D de 3 elementos, se suma con un `int`, lo que retorna un array 1D de tres elementos.
```
M = np.ones((3, 3))
M
M + a
```
Magia! Esto es _broadcasting_. Una pequeña infografía para digerirlo:
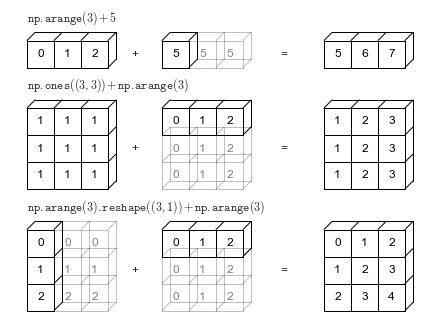
Resumen: A lo menos los dos arrays deben coincidir en una dimensión. Luego, el array de dimensión menor se extiende con tal de ajustarse a las dimensiones del otro.
La documentación oficial de estas reglas la puedes encontrar [aquí](https://numpy.org/devdocs/user/basics.broadcasting.html).
| github_jupyter |
# Article Spinning Intro
* Changing certain words of an article so it does not match the original, so a search engine can't mark it as duplicate content
* How is this done:
* take an article and slightly modify it, different terms, same meaning
* "Udemy is a **platform** or **marketplace** for online **learning**"
* "Udemy is a **podium** or **forum** for online **research**"
* Clearly context is very important!
* the idea is that you need to use the surrounding words to influence the replacement of the current word
---
# Trigram Model and Markov Models
* how can we model the probability of a word given the surrounding words?
* Lets start by taking an entire document and labeling all of the words: **w(1), w(2),...,w(n)**
* we can then model the probability of **w(i)** using the surrounding words:
* those that came before w(i): w(1)...w(i-1)
* and those that came after w(i): w(i+1)...w(n)
* Probabilistically this would look like:
$$P\Big(w(i)\;\Big|\;w(1)...w(i-1), w(i+1)...w(n)\Big)$$
* Why wouldn't this work?
* well, using this approach we are considering every word in the document, which means that only that model itself would match it exactly
* We need to do something similar to what we do with markov models and only consider the closest words
## Trigram
* we are going to use something called a trigram to accomplish this!
* we are going to create triples, where we store combinations of 3 consecutive words
* A few pieces of vocav worth knowing:
* **corpus**: collection of text
* **tokens**: words and punctuation that make up the corpus
* **Type**: distinct token
* **vocabulary**: set of all types
* **unigram**: 1 token sequence
* **bigram**: 2 token sequence
* **trigram**: 3 token sequence
* **n-gram**: n token sequence
* in the case of a trigram we are going to use the previous words and next word to predict the current word:
$$P\Big(w(i)\;\Big|\;w(i-1), w(i+1)\Big)$$
* How will we implement this?
* We are going to create a dictionary with the previous word and next word as the key, and then randomly sample the middle word **w(i)**!
* for example we could have the key ('I ', 'sports'), which would have an array of values, ['hate','love', 'enjoy', etc.]
* we would randomly sample from that array
* this is sort of like a markov model, expect a markov model is only concerned with P(w(i)|w(i-1))
* We won't replace every single word in the document, because that wouldn't give us anything useful
* so we will make the decision to replace the word based on some small probability
* Both this and latent semantic analysis are what we call unsupervised learning algorithms, because they have no labels and we just want to learn the structure of the data
* Note: spam detector and sentiment analyzer were supervised because we had labels to match to
---
# Markov Chains and Monte Carlo Methods
* Great tutorial: https://deeplearning4j.org/markovchainmontecarlo
* Markov Chain Monte Carlo (MCMC) is a mathematical method that draws samples randomly from a black-box to approximate the probability distribution of attributes over a range of objects (the height of men, the names of babies, the outcomes of events like coin tosses, the reading levels of school children, the rewards resulting from certain actions) or the futures of states.
* MCMC methods help gauge the distribution of an outcome or statistic you’re trying to predict, by randomly sampling from a complex probabilistic space.
* As with all statistical techniques, we sample from a distribution when we don’t know the function to succinctly describe the relation to two variables (actions and rewards). MCMC helps us approximate a black-box probability distribution.
## Concrete Example
Let’s say you’re a gambler in the saloon of a Gold Rush town and you roll a suspicious die without knowing if it is fair or loaded. You roll a six-sided die a hundred times, count the number of times you roll a four, and divide by a hundred. That gives you the probability of four in the total distribution. If it’s close to 16.7 (1/6 * 100), the die is probably fair.
Monte Carlo looks at the results of rolling the die many times and tallies the results to determine the probabilities of different states. It is an inductive method, drawing from experience. The die has a state space of six, one for each side.
## Systems and States
At a more abstract level, where words mean almost anything at all, a system is a set of things connected together (you might even call it a graph, where each state is a vertex, and each transition is an edge). It’s a set of states, where each state is a condition of the system. But what are states?
* Cities on a map are “states”. A road trip strings them together in transitions. The map represents the system.
* Words in a language are states. A sentence is just a series of transitions from word to word.
* Genes on a chromosome are states. To read them (and create amino acids) is to go through their transitions.
* Web pages on the Internet are states. Links are the transitions.
* Bank accounts in a financial system are states. Transactions are the transitions.
* Emotions are states in a psychological system. Mood swings are the transitions.
* Social media profiles are states in the network. Follows, likes, messages and friending are the transitions.
* Rooms in a house are states. People walking through doorways are the transitions.
So states are an abstraction used to describe these discrete, separable, things. A group of those states bound together by transitions is a system. And those systems have structure, in that some states are more likely to occur than others (ocean, land), or that some states are more likely to follow others.
We are more like to read the sequence Paris -> France than Paris -> Texas, although both series exist, just as we are more likely to drive from Los Angeles to Las Vegas than from L.A. to Slab City, although both places are nearby.
A list of all possible states is known as the “state space.” The more states you have, the larger the state space gets, and the more complex your combinatorial problem becomes.
## Markov Chains
Since states can occur one after another, it may make sense to traverse the state space, moving from one to the next. A Markov chain is a probabilistic way to traverse a system of states. It traces a series of transitions from one state to another. It’s a random walk across a graph.
Each current state may have a set of possible future states that differs from any other. For example, you can’t drive straight from Atlanta to Seattle - you’ll need to hit other states in between. We are all, always, in such corridors of probabilities; from each state, we face an array of possible future states, which in turn offer an array of future states two degrees away from the start, changing with each step as the state tree unfolds. New possibilites open up, others close behind us. Since we generally don’t have enough compute to explore every possible state of a game tree for complex games like go, one trick that organizations like DeepMind use is Monte Carlo Tree Search to narrow the beam of possibilities to only those states that promise the most likely reward.
Traversing a Markov chain, you’re not sampling with a God’s-eye view any more like a conquering alien. You are in the middle of things, groping your way toward one of several possible future states step by probabilistic step, through a Markov Chain.
While our journeys across a state space may seem unique, like road trips across America, an infinite number of road trips would slowly give us a picture of the country as a whole, and the network that links its cities together. This is known as an equilibrium distribution. That is, given infinite random walks through a state space, you can come to know how much total time would be spent in any given state. If this condition holds, you can use Monte Carlo methods to initiate randoms “draws”, or walks through the state space, in order to sample it.
## Markov Time
Markov chains have a particular property: oblivion, or forgetting.
That is, they have no long-term memory. They know nothing beyond the present, which means that the only factor determining the transition to a future state is a Markov chain’s current state. You could say the “m” in Markov stands for “memoryless”: A woman with amnesia pacing through the rooms of a house without knowing why.
Or you might say that Markov Chains assume the entirety of the past is encoded in the present, so we don’t need to know anything more than where we are to infer where we will be next. Check out a visual demo here: http://setosa.io/ev/markov-chains/
So imagine the current state as the input data, and the distribution of attributes related to those states (perhaps that attribute is reward, or perhaps it is simply the most likely future states), as the output. From each state in the system, by sampling you can determine the probability of what will happen next, doing so recursively at each step of the walk through the system’s states.
## Probability as a Spaced
When they call it a state space, they’re not joking. You can picture it, just like you can picture land and water, each one of them a probability as much as they are a physical thing. Unfold a six-sided die and you have a flattened state space in six equal pieces, shapes on a plane. Line up the letters by their frequency for 11 different languages, and you get 11 different state spaces.
Another tutorial: https://jeremykun.com/2015/04/06/markov-chain-monte-carlo-without-all-the-bullshit/
---
# Article Spinner Code
A great resource for this article spinner is found here: http://norvig.com/ngrams/ch14.pdf
Lets now write the code for our article spinner. Start with our imports.
```
import nltk
import random # needed for probabilities and sampling
import numpy as np
from bs4 import BeautifulSoup
```
### Load our positive reviews.
```
positive_reviews = BeautifulSoup(open('data/electronics/positive.review').read(), "lxml")
positive_reviews = positive_reviews.findAll('review_text')
```
### Collect all of the Trigrams
Recall, for each trigram the key is the previous and next word, and the value is going to be the possible middle words (so an array, may only contain a single value)
```
trigrams = {}
for review in positive_reviews: # loop through every review
s = review.text.lower() # don't want two versions of same word
tokens = nltk.tokenize.word_tokenize(s)
for i in range(len(tokens) - 2):
k = (tokens[i], tokens[i+2]) # the key is a tuple, tuples are immutable and can be key
if k not in trigrams:
trigrams[k] = []
trigrams[k].append(tokens[i+1]) # now we have all of the possible middle words
```
### Transform into a probability vector
Now that we have all of the possible middle words, we need to transform this into a probability vector. We need to convert these trigrams into probabilities.
```
trigrams_probabilities = {} # dictionary to hold trigram probabilities, the loop through trigrams
for k, words in trigrams.items(): # k will be the key, and words is a list of words for that key
if len(set(words)) > 1: # set gets rid of duplicates, then we need to make sure > 1 word
d = {} # another dictionary d, keyed by the middle word
n = 0
for w in words: # loop through each word, d count how many times the middle word occur
if w not in d:
d[w] = 0
d[w] += 1
n += 1 # n is going to track the total number of words
for w, c in d.items():
d[w] = float(c)/n # # of times each word occurs, divided by total number of words
trigrams_probabilities[k] = d # setting trigram prob for specific key to be that of d
```
### Function to Randomly Sample Trigram Probabilities
Now we need to create a function that will randomly sample from these trigram probabilities.
```
def random_sample(d): # function, takes dictionary (key is word, value is probability of that word)
r = random.random() # generate random number
cumulative = 0
for w, p in d.items():
cumulative += p
if r < cumulative:
return w
```
### Function to test spinner
It needs to randomly choose a review, then try to spin it and print both out so we can compare them.
```
def test_spinner():
review = random.choice(positive_reviews) # grab a random positive review
s = review.text.lower()
print('Original:', s)
tokens = nltk.tokenize.word_tokenize(s) # tokenize the positive review
for i in range(len(tokens) - 2): # loop through each token
if random.random() < 0.2: # choose with a small probability to replace (20% chance)
k = (tokens[i], tokens[i+2]) # get the word before and after our word
if k in trigrams_probabilities:
w = random_sample(trigrams_probabilities[k])
tokens[i+1] = w
print ('Spun:')
print(" ".join(tokens).replace(" .", ".").replace(" '", "'").replace(" ,", ",").replace("$ ", "$").replace(" !", "!"))
test_spinner()
```
| github_jupyter |
# Deriving a vegetation index from 4-band satellite data
A **vegetation index** is generated by combining two or more spectral bands from a satellite image. There are many different vegetation indices; in this exercise we'll learn about the most commonly-used index.
### NDVI
Researchers often use a vegetation index called NDVI to measure the "greenness" or density of vegetation across a landscape. In addition to monitoring vegetation health, NDVI _(Normalized Difference Vegetation Index)_ can be used to track climate change, agricultural production, desertification, and land cover change. Developed by NASA scientist Compton Tucker in 1977, NDVI is derived from satellite imagery and compares reflected near-infrared light to reflected visible red light. It can be expressed as following equation:

In general, healthy and/or dense vegetation reflects a lot of near-infrared light and not as much red visible light. Conversely, when vegetation is sparse or not-so-healthy, its near-infrared reflectance decreases and its red light reflectance increases. You can read more about how NDVI is used to study cyclical, seasonal, and long-term changes to the Earth's physical characteristics from [NASA](https://earthobservatory.nasa.gov/Features/MeasuringVegetation/measuring_vegetation_1.php) and [USGS](https://phenology.cr.usgs.gov/ndvi_foundation.php) researchers.
To create this vegetation index, we're going to use PlanetScope's SR _(Surface Reflectance)_ data product. You can learn [more about Surface Reflectance (SR) and Planet data here](https://support.planet.com/hc/en-us/sections/115003720348-Surface-Reflectance), but for the purposes of this exercise, all you need to know is: SR data is satellite data that has been algorithmically corrected to remove atmospheric interference.
**In this exercise, you'll learn how to perform an NDVI calculation on PlanetScope Surface Reflectance data in Python, and generate a colorized NDVI image for visual analysis. Here are the steps to follow:**
1. Download a PlanetScope SR product
2. Extract data from the red and near-infrared bands
3. Perform the NDVI calculation
4. Save the NDVI image
5. Apply a color scheme to the NDVI image
6. Generate a histogram to view NDVI values
### Requirements
- Python 2.7 or 3+
- [Planet's Python Client](https://pypi.org/project/planet/)
- [rasterio](https://github.com/mapbox/rasterio)
- [numpy](http://www.numpy.org/)
- [matplotlib](https://matplotlib.org/)
- [Planet API Key](https://developers.planet.com/docs/quickstart/getting-started/), stored as environment variable `$PL_API_KEY`.
- [Planet 4-Band Imagery](https://developers.planet.com/docs/api/psscene4band/) with the following specifications: `item-type`: `PSScene4Band`; `asset-type`: `analytic_sr`
## Step 1. Download a PlanetScope SR Product
For this exercise you'll need a 4-band PlanetScope Surface Reflectance product. You can search for & download your own data, or use the demo data provided in-class. If you choose to use the demo data, skip to **Step 2**.
To search for your own data, you'll first need to define an Area of Interest (AOI). [http://geojson.io](http://geojson.io) is a free browser-based tool that makes generating a GeoJSON-formatted AOI easy.
Once that's done, use one of the following methods to search for & download data:
- using [Planet's Python CLI](https://www.planet.com/docs/api-quickstart-examples/cli/) to interact with Planet's API from the command line
- using Planet's API directly to [search](https://developers.planet.com/docs/quickstart/searching-for-imagery/) and [download](https://developers.planet.com/docs/quickstart/downloading-imagery/)
- using the [Planet Explorer](https://www.planet.com/products/explorer/) site to visually search for & download data
With all of the above, you'll want to filter for 4-Band PlanetScope data (item type: `PSScene4Band`) and download the associated SR product (asset type: `analytic_sr`)
### Option 1: Searching & Downloading via CLI
If you choose to use Planet's CLI, you might fight these [search](https://developers.planet.com/docs/quickstart/searching-for-imagery/) and [download](https://developers.planet.com/docs/quickstart/downloading-imagery/) quickstart guides to be useful.
```
# To use Planet's CLI from this Notebook, begin your line as follows:
!planet data
# Here is an example of using Planet's CLI to search for a known item id:
# !planet data download --item-type PSScene4Band --asset-type analytic_sr --dest data --string-in id 20160831_180302_0e26
```
### Option 2: Searching & Downloading via API
If you prefer to use Planet's API directly via Python, this [search & download quickstart Notebook](../../data-api-tutorials/search_and_download_quickstart.ipynb) may be useful.
```
# To use Planet's API, you'll probably begin by importing your favorite HTTP toolkit, e.g.:
import requests
from requests.auth import HTTPBasicAuth
# Your Planet API key is available in this Notebook as an env variable, e.g.:
import os
PLANET_API_KEY = os.getenv('PL_API_KEY')
```
### Option 3: Searching & Downloading via Planet Explorer
If you prefer to browse for images visually, log in to your Planet account and use [Planet Explorer](https://www.planet.com/explorer/) to search for PlanetScope imagery. You'll want to make sure to set the Source filter to show only `4-band PlanetScope Scene` results.
You can [click here for an example search](4-band PlanetScope Scene) showing 4-band PlanetScope data in California's Central Valley.
### Success! Data Obtained
Regardless of the path you chose to obtain data for this exercise, once you have successfully aquired a 4-band PlanetScope `analytic_SR`-type GeoTIFF, place the file in the [data/](data/) directory adjacent to this Notebook.
## Step 2. Extract the data from the red and near-infrared bands
For this step, use [Rasterio](https://rasterio.readthedocs.io/en/latest/) to open the raster image you downloaded (the .tif file). After that, use Rasterio read the data from the red and near-infrared bands: this will load the band data into arrays that you can manipulate using Python's [NumPy](http://www.numpy.org/) libary.
*Note: in PlanetScope 4-band images, the band order is BGRN: (1) Blue, (2) Green, (3) Red, (4) Near-infrared.*
```
import rasterio
filename = "data/20160831_180302_0e26_3B_AnalyticMS_SR.tif"
# Load red and NIR bands - note all PlanetScope 4-band images have band order BGRN
with rasterio.open(filename) as src:
band_red = src.read(3)
with rasterio.open(filename) as src:
band_nir = src.read(4)
```
## Step 3. Perform the NDVI calculation
Next, you're going to calculate NDVI through subtraction and division of the values stored in the NumPy arrays. This calculation will give you NDVI values that range from -1 to 1. Values closer to 1 indicate a greater density of vegetation or higher level of "greenness."
As a reminder, the NDVI formula is:
\begin{equation*}
ndvi = \frac{nir-red}{(nir+red)}
\end{equation*}
Where `nir` is the Near-infrared band, and `red` is the Red band.
```
# allow division by zero without throwing a warning
import numpy
numpy.seterr(divide='ignore', invalid='ignore')
# Calculate NDVI - remember, bands read via rasterio are just numpy arrays
ndvi = (band_nir.astype(float) - band_red.astype(float)) / (band_nir + band_red)
```
As a quick check of our calculations, let's print the minimum and maximum values in our calculated `ndvi`. Because we're using the NDVI formula to normalize the input bands, we know that our expected values should fall within -1.0 to +1.0.
_(HINT: this is still a numpy array, so use numpy functions here)_.
```
# check range NDVI values, excluding NaN
print(numpy.nanmin(ndvi))
print(numpy.nanmax(ndvi))
```
Assuming your min & max values are in-range -- congratulations! You have performed what is known as *raster band math*. Well done. This skill has many applications beyond the NDVI you're calculating in this exercise: the relationship of values between different spectral bands is the basis for many kinds of remote sensing analysis.
## Step 5. Save the NDVI image
Now that you've calculated NDVI values, you're going to save the results to a new single-band image, making sure the new image file uses the geospatial metadata from the GeoTIFF you originally acquired, and the `dtype` of the new numpy array you generated above.
```
# get the metadata of original GeoTIFF:
meta = src.meta
print(meta)
# get the dtype of our NDVI array:
ndvi_dtype = ndvi.dtype
print(ndvi_dtype)
# set the source metadata as kwargs we'll use to write the new data:
kwargs = meta
# update the 'dtype' value to match our NDVI array's dtype:
kwargs.update(dtype=ndvi_dtype)
# update the 'count' value since our output will no longer be a 4-band image:
kwargs.update(count=1)
# Finally, use rasterio to write new raster file 'data/ndvi.tif':
with rasterio.open('data/ndvi.tif', 'w', **kwargs) as dst:
dst.write(ndvi, 1)
```
## Step 6. Apply a color scheme to visualize the NDVI values on the image
Now that you've created [ndvi.tif](data/ndvi.tif), you may be tempted to open it immediately & take a look at what you've accomplished. If you do, don't be disappointed when `ndvi.tif` opened in your favorite image viewer doesn't look like much at all. That's normal! Remember that this is not just any .tif but a GeoTIFF - one in which every pixel has a value of 1.0 or less.
At this point, you could open `ndvi.tif` in a Desktop GIS GUI like QGIS, and define color values for each pixel in order to get meaningful visual information out of the data. But this is a Python exercise, so let's use [Matplotlib](https://matplotlib.org/) to do the same thing.
As we verified earlier, we know the values in our NDVI will range from -1 to 1. To best visualize this, we want to use a diverging color scheme, and we want to center the colorbar at a defined midpoint. Interestingly, the best midpoint for NDVI analysis is **0.1** - not **0.0** as you might expect. You can read more about [how NDVIs are interpreted here](https://earthobservatory.nasa.gov/Features/MeasuringVegetation).
To normalize a colorbar against our custom midpoint, we're going to take advantage of the following handy class [originally created by Joe Kington](https://matplotlib.org/gallery/userdemo/colormap_normalizations_custom.html):
```
from matplotlib import colors
# Credit: Joe Kington
class MidpointNormalize(colors.Normalize):
"""
Normalize the colorbar so that diverging bars work there way either side from a prescribed midpoint value
"""
def __init__(self, vmin=None, vmax=None, midpoint=None, clip=False):
self.midpoint = midpoint
colors.Normalize.__init__(self, vmin, vmax, clip)
def __call__(self, value, clip=None):
# I'm ignoring masked values and all kinds of edge cases to make a
# simple example...
x, y = [self.vmin, self.midpoint, self.vmax], [0, 0.5, 1]
return numpy.ma.masked_array(numpy.interp(value, x, y), numpy.isnan(value))
# Begin by pulling in pyplot
import matplotlib.pyplot as plt
# Set min/max values from NDVI range for image
# HINT: refer back to earlier, when we verified our min & max values were within expected range
min=numpy.nanmin(ndvi)
max=numpy.nanmax(ndvi)
# Set our custom midpoint for most effective NDVI analysis
mid=0.1
# Set your favorite diverging color scheme
# You can use https://matplotlib.org/users/colormaps.html as a reference
colormap = plt.cm.RdYlGn
# Call MidPointNormalize with our min, max, and custom midpoint
norm = MidpointNormalize(vmin=min, vmax=max, midpoint=mid)
# Create a pyplot figure, in which we'll display our colorized NDVI
fig = plt.figure(figsize=(20,10))
# Add a subplot to our figure, which will contain the colorbar
ax = fig.add_subplot(111)
# Use 'imshow' to specify the input data, colormap, min, max, and norm for the colorbar
cbar_plot = ax.imshow(ndvi, cmap=colormap, vmin=min, vmax=max, norm=norm)
# Turn off the display of axis labels
ax.axis('off')
# Set a title
ax.set_title('Normalized Difference Vegetation Index', fontsize=18, fontweight='bold')
# Configure the colorbar
cbar = fig.colorbar(cbar_plot, orientation='horizontal', shrink=0.65)
# Call 'savefig' to save this plot to an image file
fig.savefig("data/ndvi-fig.png", dpi=200, bbox_inches='tight', pad_inches=0.7)
# Finally - let's take a look!
plt.show()
```
## 7. Generate a histogram of NDVI values
Congratulations! You've used band math to apply a well-known vegetation index formula to satellite data, and visualized it for analysis using a diverging color ramp. You're well on your way to getting meaningful information out of satellite imagery using Python.
As one last step, you use `pyplot` to generate a histogram of values in your NDVI calculation. This can be useful for quick analysis, giving visual insight into the distribution of "healthy" vs "unhealthy" vegetation values in your study area.
```
# Define a new figure
fig2 = plt.figure(figsize=(20,10))
# Give this new figure a subplot, which will contain the histogram itself
ax = fig2.add_subplot(111)
# Add a title & (x,y) labels to the plot
plt.title("NDVI Histogram", fontsize=18, fontweight='bold')
plt.xlabel("NDVI values", fontsize=14)
plt.ylabel("Number of pixels", fontsize=14)
# For the x-axis, we want to count every pixel that is not an empty value
x = ndvi[~numpy.isnan(ndvi)]
# Define the number of bins to divide the data into
bins = 20
# Define a color for the histogram
# You can use https://matplotlib.org/2.0.0/examples/color/named_colors.html as a reference
color = 'lightgreen'
# call 'hist` with our x-axis, bins, and color details
ax.hist(x,bins,color=color)
# Save the generated figure to an external image file
fig2.savefig("data/ndvi-histogram.png", dpi=200, bbox_inches='tight', pad_inches=0.7)
# Finally - let's take a look!
plt.show()
```
| github_jupyter |
<!-- Autogenerated by `scripts/make_examples.py` -->
<table align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/voxel51/fiftyone-examples/blob/master/examples/open_images_evaluation/open_images_evaluation.ipynb">
<img src="https://user-images.githubusercontent.com/25985824/104791629-6e618700-5769-11eb-857f-d176b37d2496.png" height="32" width="32">
Try in Google Colab
</a>
</td>
<td>
<a target="_blank" href="https://nbviewer.jupyter.org/github/voxel51/fiftyone-examples/blob/master/examples/open_images_evaluation/open_images_evaluation.ipynb">
<img src="https://user-images.githubusercontent.com/25985824/104791634-6efa1d80-5769-11eb-8a4c-71d6cb53ccf0.png" height="32" width="32">
Share via nbviewer
</a>
</td>
<td>
<a target="_blank" href="https://github.com/voxel51/fiftyone-examples/blob/master/examples/open_images_evaluation/open_images_evaluation.ipynb">
<img src="https://user-images.githubusercontent.com/25985824/104791633-6efa1d80-5769-11eb-8ee3-4b2123fe4b66.png" height="32" width="32">
View on GitHub
</a>
</td>
<td>
<a href="https://github.com/voxel51/fiftyone-examples/raw/master/examples/open_images_evaluation/open_images_evaluation.ipynb" download>
<img src="https://user-images.githubusercontent.com/25985824/104792428-60f9cc00-576c-11eb-95a4-5709d803023a.png" height="32" width="32">
Download notebook
</a>
</td>
</table>
# Evaluating a Detection Model on the Open Images Dataset
This tutorial demonstrates per-image evaluation of an object detection model on [the Open Images dataset](https://storage.googleapis.com/openimages/web/index.html)
that generates:
- true positives & false positives
- per-class average precision (AP)
- mean average precision (mAP)
for each image and adds this information to each [Sample](https://voxel51.com/docs/fiftyone/api/fiftyone.core.sample.html#fiftyone.core.sample.Sample)
in the [Dataset](https://voxel51.com/docs/fiftyone/api/fiftyone.core.dataset.html#fiftyone.core.dataset.Dataset).
The steps are broken down as follows:
1. [Requirements](#Requirements)
2. [Download the test data and ground-truth labels](#Download-the-test-data-and-ground-truth-labels) (optional)
3. [Generate predictions](#(optional)-Generate-predictions) (optional)
4. [Load the data into FiftyOne](#Load-the-data-into-FiftyOne)
5. [Prepare the ground-truth for evaluation](#Prepare-the-ground-truth-for-evaluation) (optional)
6. [Evaluate on a per-image granularity](#Evaluate-on-a-per-image-granularity)
7. [Explore](#Explore)
Optional steps may not be necessary depending on if you have already downloaded the data or have your own model to evaluate.
This tutorial evaluates a model on [Open Images V4](https://storage.googleapis.com/openimages/web/download_v4.html)
however this code supports later versions of Open Images as well. If using a newer version just make sure to
use the appropriate hierarchy file and class label map.
## Quickstart: Interactive visualization in under 5 minutes
The following steps demonstrate how to evaluate *your own model* on a per-image granularity using Tensorflow Object Detection API and then interactively visualize and explore true/false positive detections. If you would simply like to browse a subset of Open Images test set with evaluation on a pre-trained model, instead [download this dataset](https://voxel51.com/downloads/fiftyone/tutorials/open-images-v4-test-500.zip). You can get up and running with just 5 lines of code!
Below is the Python code to load the dataset download and visualize it:
```
import fiftyone as fo
from fiftyone import ViewField as F
# Path to the unzipped dataset you downloaded
DATASET_DIR = "/path/to/open-images-v4-test-500"
# Load the dataset
dataset = fo.Dataset.from_dir(DATASET_DIR, fo.types.FiftyOneDataset)
# Open the dataset in the App
session = fo.launch_app(dataset)
# Filter the visible detections by confidence and filter the samples
# to only those with at least one false positive
high_conf_view = (
dataset
.filter_labels("true_positives", F("confidence") > 0.4)
.filter_labels("false_positives", F("confidence") > 0.4)
.match(F("false_positives.detections").length() > 0)
.sort_by("open_images_id")
)
session.view = high_conf_view
```
## Requirements
This workflow requires a few Python packages.
First, if you haven't already, install FiftyOne:
```
%%bash
pip install fiftyone
```
Then install the appropriate version of `tensorflow` depending on whether or not you have a GPU:
```
%%bash
pip install tensorflow
```
and install other requirements:
```
%%bash
pip install numpy pandas google-api-python-client
```
### Download supporting scripts
This notebook uses a collection of helper scripts and modules. If you downloaded this notebook from the [fiftyone-examples](https://github.com/voxel51/fiftyone-examples) repository, you will also need to download the rest of the `examples/open_images_evaluation/` sudirectory.
## Download the test dataset
All of the data (images, metadata and annotations) can be found on the
[official Open Images website](https://storage.googleapis.com/openimages/web/download_v4.html).
If you are using Open Images V4 you can use the following commands to download all the necessary files.
### Download the data
**WARNING** This is 36GB of data!
```
%%bash
aws s3 --no-sign-request sync s3://open-images-dataset/test open-images-dataset/test
```
### Download the labels and metadata
```
%%bash
wget https://storage.googleapis.com/openimages/2018_04/test/test-annotations-bbox.csv
wget https://storage.googleapis.com/openimages/2018_04/test/test-annotations-human-imagelabels-boxable.csv
wget https://storage.googleapis.com/openimages/2018_04/class-descriptions-boxable.csv
wget https://storage.googleapis.com/openimages/2018_04/bbox_labels_600_hierarchy.json
```
## (optional) Generate predictions
This section steps through generating predictions using a pre-trained model publically available
on [Tensorflow Hub](https://www.tensorflow.org/hub).
The exact model used can be modified simply by changing `MODEL_HANDLE` below.
### Alternative 1: download pre-computed predictions
If you would like to skip the step of generating predictions, simply download [this predictions file](https://voxel51.com/downloads/fiftyone/tutorials/google-faster_rcnn-openimages_v4-inception_resnet_v2_predictions_3081.csv).
### Alternative 2: use your own model
If you have your own model that you would like to evaluate, make sure the outputs are saved to `csv` in
[Tensorflow Object Detection API](https://github.com/tensorflow/models/tree/master/research/object_detection) format.
Output file structure must have a single header row followed by one row per detection as follows:
```
ImageID,LabelName,Score,XMin,XMax,YMin,YMax
...,...,...,...,...,...,...
...,...,...,...,...,...,...
```
Example output for two images with two detections each:
```
ImageID,LabelName,Score,XMin,XMax,YMin,YMax
000026e7ee790996,/m/07j7r,0.1,0.071905,0.145346,0.206591,0.391306
000026e7ee790996,/m/07j7r,0.2,0.439756,0.572466,0.264153,0.435122
000062a39995e348,/m/015p6,0.4,0.205719,0.849912,0.154144,1.000000
000062a39995e348,/m/05s2s,0.5,0.137133,0.377634,0.000000,0.884185
```
### Generate predictions with a Tensorflow Hub pre-trained model
To use a Tensorflow Hub model requires the following packages:
```
%%bash
pip install Pillow tensorflow-hub
```
Populate the following environment variables and run the inference script.
This script is resumable and saves after every 10 samples are processed by default. It does
not process images in batches.
```
%%bash
IMAGES_DIR=/PATH/TO/IMAGES
OUTPUT_DIR=/PATH/TO/PREDICTIONS
MODEL_HANDLE="https://tfhub.dev/google/faster_rcnn/openimages_v4/inception_resnet_v2/1"
# MODEL_HANDLE="https://tfhub.dev/google/openimages_v4/ssd/mobilenet_v2/1"
python open_images_eval/scripts/inference.py \
--output_dir ${OUTPUT_DIR} \
--output_format tf_object_detection_api \
${IMAGES_DIR} ${MODEL_HANDLE}
```
## Load the data into FiftyOne
### Create a persistent FiftyOne dataset
The following script loads the data into a FiftyOne [Dataset](https://voxel51.com/docs/fiftyone/api/fiftyone.core.dataset.html#fiftyone.core.dataset.Dataset).
This process copies all labels and metadata to a non-relational database for rapid access and powerful querying, but only paths to the images are stored
in the database, not copies of the images themselves!
The dataset is set to [persistent](https://voxel51.com/docs/fiftyone/user_guide/using_datasets.html#dataset-persistence)
so that it remains in the database and can be loaded in a new python process.
```
%%bash
DATASET_NAME="open-images-v4-test"
IMAGES_DIR=/PATH/TO/IMAGES
BOUNDING_BOXES_EXPANDED=/PATH/TO/test-annotations-bbox_expanded.csv
IMAGE_LABELS_EXPANDED=/PATH/TO/test-annotations-human-imagelabels-boxable_expanded.csv
PREDICTIONS_PATH=/PATH/TO/PREDICTIONS.csv
CLASS_DESCRIPTIONS=/PATH/TO/class-descriptions-boxable.csv
python open_images_eval/scripts/load_data.py \
--bounding_boxes_path ${BOUNDING_BOXES_EXPANDED} \
--image_labels_path ${IMAGE_LABELS_EXPANDED} \
--predictions_path ${PREDICTIONS_PATH} \
--prediction_field_name "faster_rcnn" \
--class_descriptions_path ${CLASS_DESCRIPTIONS} \
--load_images_with_preds \
--max_num_images 1000 \
${DATASET_NAME} ${IMAGES_DIR}
```
To skip uploading predictions use the following code block. You can always add
predictions later using the function
`open_images_eval.error_analysis.load_data.add_open_images_predictions()`
```
%%bash
DATASET_NAME="open-images-v4-test"
IMAGES_DIR=/PATH/TO/IMAGES
BOUNDING_BOXES_EXPANDED=/PATH/TO/test-annotations-bbox_expanded.csv
IMAGE_LABELS_EXPANDED=/PATH/TO/test-annotations-human-imagelabels-boxable_expanded.csv
CLASS_DESCRIPTIONS=/PATH/TO/class-descriptions-boxable.csv
python open_images_eval/scripts/load_data.py \
--bounding_boxes_path ${BOUNDING_BOXES_EXPANDED} \
--image_labels_path ${IMAGE_LABELS_EXPANDED} \
--class_descriptions_path ${CLASS_DESCRIPTIONS} \
--max_num_images 1000 \
${DATASET_NAME} ${IMAGES_DIR}
```
### Visualize the data
Now that we have a Fiftyone `Dataset`, let's visualize the data before evaluating it:
```
import fiftyone as fo
from fiftyone import ViewField as F
dataset = fo.load_dataset("open-images-v4-test")
session = fo.launch_app(dataset)
# Filter the visible detections by confidence
session.view = dataset.filter_labels("faster_rcnn", F("confidence") > 0.4)
```
<img src="images/open_images_pre_eval.gif">
## Prepare the ground-truth for evaluation
Open Images requires "expanding the hierarchy" of the ground-truth labels, for
evaluation. The labels you downloaded only contain leaf node labels. So, for
example, for a bounding box labeled `Jaguar`, the hierarchy expansion would add
duplicate boxes with labels `Carnivore`, `Mammal` and `Animal`.
### Install TF Object Detection API
The first step is to install the Tensorflow Object Detection API. Instructions
on how to do so can be found
[here](https://github.com/tensorflow/models/blob/master/research/object_detection/g3doc/tf2.md).
### Create expanded hierarchy ground-truth labels
The following commands are essentially copied from [this tutorial](https://github.com/tensorflow/models/blob/master/research/object_detection/g3doc/challenge_evaluation.md).
```
%%bash
# TODO: modify these
export TF_MODELS_RESEARCH=PATH/TO/TENSORFLOW/models/research/object_detection
LABELS_DIR=PATH/TO/LABELS
HIERARCHY_FILE=${LABELS_DIR}/bbox_labels_600_hierarchy.json
BOUNDING_BOXES=${LABELS_DIR}/test-annotations-bbox
IMAGE_LABELS=${LABELS_DIR}/test-annotations-human-imagelabels-boxable
python ${TF_MODELS_RESEARCH}/object_detection/dataset_tools/oid_hierarchical_labels_expansion.py \
--json_hierarchy_file=${HIERARCHY_FILE} \
--input_annotations=${BOUNDING_BOXES}.csv \
--output_annotations=${BOUNDING_BOXES}_expanded.csv \
--annotation_type=1
python ${TF_MODELS_RESEARCH}/object_detection/dataset_tools/oid_hierarchical_labels_expansion.py \
--json_hierarchy_file=${HIERARCHY_FILE} \
--input_annotations=${IMAGE_LABELS}.csv \
--output_annotations=${IMAGE_LABELS}_expanded.csv \
--annotation_type=2
```
You should now have two new files in `LABELS_DIR`:
```
test-annotations-bbox_expanded.csv
test-annotations-human-imagelabels-boxable_expanded.csv
```
## Evaluate on a per-image granularity
This next script evaluates each image indivually using some wrapper code around the TF Object Detection API
evaluation code.
### Running evaluation
If you skipped ["Prepare the ground-truth for evaluation"](#Prepare-the-ground-truth-for-evaluation) be sure to export the `TF_MODELS_RESEARCH` environment variable.
```
%%bash
CLASS_LABEL_MAP=${TF_MODELS_RESEARCH}/object_detection/data/oid_v4_label_map.pbtxt
python open_images_eval/scripts/evaluate_model.py \
--prediction_field_name "faster_rcnn" \
--iou_threshold 0.5 \
${DATASET_NAME} ${CLASS_LABEL_MAP}
```
## Explore
At last! We can now visualize the data. Use this snippet to launch the GUI app and start browsing.
```
import fiftyone as fo
from fiftyone import ViewField as F
dataset = fo.load_dataset("open-images-v4-test")
session = fo.launch_app(dataset)
```
There so many possibilities as far as how to slice and dice this data. Let's start with high confidence predictions (`detection.confidence > 0.4`):
```
# Filter the visible detections for high confidence,
# then filter the samples to only those with at least one false positive
high_confidence_view = (
dataset
.filter_labels("faster_rcnn_TP", F("confidence") > 0.4)
.filter_labels("faster_rcnn_FP", F("confidence") > 0.4)
.match(F("faster_rcnn_FP.detections").length() > 0)
.sort_by("open_images_id")
)
session.view = high_confidence_view
```
<img src="images/open_images_eval.jpg">
On the *very first image* we see a prediction that correctly boxes a bird but is mistakenly marked as a false positive!
<img src="images/tucan.jpg">
Here are a few more ideas for how to view the data:
```
# Filter the visible detections for medium confidence,
# then filter the samples to only those with at least one false positive
medium_confidence_view = (
dataset
.filter_labels("true_positives", (F("confidence") > 0.2) & (F("confidence") < 0.4))
.filter_labels("false_positives", (F("confidence") > 0.2) & (F("confidence") < 0.4))
.match(F("false_positives.detections").length() > 0)
.sort_by("open_images_id")
)
session.view = medium_confidence_view
# Filter the visible detections for ground truth with `Bird` or `Human eye`,
# then filter the samples to only those with at least one ground truth box
bird_eye_view = (
dataset
.filter_labels("groundtruth_detections", F("label").is_in(["Bird", "Human eye"]))
.match(F("groundtruth_detections.detections").length() > 0)
.sort_by("open_images_id")
)
session.view = bird_eye_view
# Filter the visible detections for small bounding box area,
# then filter the samples to only those with at least one false positive
small_boxes_view = (
dataset
.filter_labels("false_positives", bbox_area < 0.01)
.filter_labels("true_positives", bbox_area < 0.01)
.match(F("false_positives.detections").length() > 0)
.sort_by("open_images_id")
)
session.view = small_boxes_view
```
| github_jupyter |
<h1> Preprocessing using Dataflow </h1>
This notebook illustrates:
<ol>
<li> Creating datasets for Machine Learning using Dataflow
</ol>
<p>
While Pandas is fine for experimenting, for operationalization of your workflow, it is better to do preprocessing in Apache Beam. This will also help if you need to preprocess data in flight, since Apache Beam also allows for streaming.
Apache Beam only works in Python 2 at the moment, so we're going to switch to the Python 2 kernel. In the above menu, click the dropdown arrow and select `python2`. 
Then activate a Python 2 environment and install Apache Beam.
```
%%bash
source activate py2env
conda install -y pytz
pip uninstall -y google-cloud-dataflow
pip install --upgrade apache-beam[gcp]
```
After doing a pip install, click **"Reset Session"** on the notebook so that the Python environment picks up the new packages.
```
# change these to try this notebook out
BUCKET = 'cloud-training-demos-ml'
PROJECT = 'cloud-training-demos'
REGION = 'us-central1'
import os
os.environ['BUCKET'] = BUCKET
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
%%bash
if ! gsutil ls | grep -q gs://${BUCKET}/; then
gsutil mb -l ${REGION} gs://${BUCKET}
fi
```
<h2> Save the query from earlier </h2>
The data is natality data (record of births in the US). My goal is to predict the baby's weight given a number of factors about the pregnancy and the baby's mother. Later, we will want to split the data into training and eval datasets. The hash of the year-month will be used for that.
```
# Create SQL query using natality data after the year 2000
query = """
SELECT
weight_pounds,
is_male,
mother_age,
plurality,
gestation_weeks,
ABS(FARM_FINGERPRINT(CONCAT(CAST(YEAR AS STRING), CAST(month AS STRING)))) AS hashmonth
FROM
publicdata.samples.natality
WHERE year > 2000
"""
# Call BigQuery and examine in dataframe
import google.datalab.bigquery as bq
df = bq.Query(query + " LIMIT 100").execute().result().to_dataframe()
df.head()
```
<h2> Create ML dataset using Dataflow </h2>
Let's use Cloud Dataflow to read in the BigQuery data, do some preprocessing, and write it out as CSV files.
Instead of using Beam/Dataflow, I had three other options:
* Use Cloud Dataprep to visually author a Dataflow pipeline. Cloud Dataprep also allows me to explore the data, so we could have avoided much of the handcoding of Python/Seaborn calls above as well!
* Read from BigQuery directly using TensorFlow.
* Use the BigQuery console (http://bigquery.cloud.google.com) to run a Query and save the result as a CSV file. For larger datasets, you may have to select the option to "allow large results" and save the result into a CSV file on Google Cloud Storage.
<p>
However, in this case, I want to do some preprocessing, modifying data so that we can simulate what is known if no ultrasound has been performed. If I didn't need preprocessing, I could have used the web console. Also, I prefer to script it out rather than run queries on the user interface, so I am using Cloud Dataflow for the preprocessing.
Note that after you launch this, the actual processing is happening on the cloud. Go to the GCP webconsole to the Dataflow section and monitor the running job. It took about 20 minutes for me.
<p>
If you wish to continue without doing this step, you can copy my preprocessed output:
<pre>
gsutil -m cp -r gs://cloud-training-demos/babyweight/preproc gs://your-bucket/
</pre>
```
import apache_beam as beam
import datetime, os
def to_csv(rowdict):
# Pull columns from BQ and create a line
import hashlib
import copy
CSV_COLUMNS = 'weight_pounds,is_male,mother_age,plurality,gestation_weeks'.split(',')
# Create synthetic data where we assume that no ultrasound has been performed
# and so we don't know sex of the baby. Let's assume that we can tell the difference
# between single and multiple, but that the errors rates in determining exact number
# is difficult in the absence of an ultrasound.
no_ultrasound = copy.deepcopy(rowdict)
w_ultrasound = copy.deepcopy(rowdict)
no_ultrasound['is_male'] = 'Unknown'
if rowdict['plurality'] > 1:
no_ultrasound['plurality'] = 'Multiple(2+)'
else:
no_ultrasound['plurality'] = 'Single(1)'
# Change the plurality column to strings
w_ultrasound['plurality'] = ['Single(1)', 'Twins(2)', 'Triplets(3)', 'Quadruplets(4)', 'Quintuplets(5)'][rowdict['plurality'] - 1]
# Write out two rows for each input row, one with ultrasound and one without
for result in [no_ultrasound, w_ultrasound]:
data = ','.join([str(result[k]) if k in result else 'None' for k in CSV_COLUMNS])
key = hashlib.sha224(data).hexdigest() # hash the columns to form a key
yield str('{},{}'.format(data, key))
def preprocess(in_test_mode):
import shutil, os, subprocess
job_name = 'preprocess-babyweight-features' + '-' + datetime.datetime.now().strftime('%y%m%d-%H%M%S')
if in_test_mode:
print('Launching local job ... hang on')
OUTPUT_DIR = './preproc'
shutil.rmtree(OUTPUT_DIR, ignore_errors=True)
os.makedirs(OUTPUT_DIR)
else:
print('Launching Dataflow job {} ... hang on'.format(job_name))
OUTPUT_DIR = 'gs://{0}/babyweight/preproc/'.format(BUCKET)
try:
subprocess.check_call('gsutil -m rm -r {}'.format(OUTPUT_DIR).split())
except:
pass
options = {
'staging_location': os.path.join(OUTPUT_DIR, 'tmp', 'staging'),
'temp_location': os.path.join(OUTPUT_DIR, 'tmp'),
'job_name': job_name,
'project': PROJECT,
'teardown_policy': 'TEARDOWN_ALWAYS',
'no_save_main_session': True
}
opts = beam.pipeline.PipelineOptions(flags = [], **options)
if in_test_mode:
RUNNER = 'DirectRunner'
else:
RUNNER = 'DataflowRunner'
p = beam.Pipeline(RUNNER, options = opts)
query = """
SELECT
weight_pounds,
is_male,
mother_age,
plurality,
gestation_weeks,
ABS(FARM_FINGERPRINT(CONCAT(CAST(YEAR AS STRING), CAST(month AS STRING)))) AS hashmonth
FROM
publicdata.samples.natality
WHERE year > 2000
AND weight_pounds > 0
AND mother_age > 0
AND plurality > 0
AND gestation_weeks > 0
AND month > 0
"""
if in_test_mode:
query = query + ' LIMIT 100'
for step in ['train', 'eval']:
if step == 'train':
selquery = 'SELECT * FROM ({}) WHERE MOD(ABS(hashmonth),4) < 3'.format(query)
else:
selquery = 'SELECT * FROM ({}) WHERE MOD(ABS(hashmonth),4) = 3'.format(query)
(p
| '{}_read'.format(step) >> beam.io.Read(beam.io.BigQuerySource(query = selquery, use_standard_sql = True))
| '{}_csv'.format(step) >> beam.FlatMap(to_csv)
| '{}_out'.format(step) >> beam.io.Write(beam.io.WriteToText(os.path.join(OUTPUT_DIR, '{}.csv'.format(step))))
)
job = p.run()
if in_test_mode:
job.wait_until_finish()
print("Done!")
preprocess(in_test_mode = False)
```
The above step will take 20+ minutes. Go to the GCP web console, navigate to the Dataflow section and <b>wait for the job to finish</b> before you run the follwing step.
```
%bash
gsutil ls gs://${BUCKET}/babyweight/preproc/*-00000*
```
Copyright 2017 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| github_jupyter |
```
%pylab inline
import cmath
def get_zero_sequence_impedance(sequence_impedance_matrix):
try:
return sequence_impedance_matrix[0,0]
except:
raise ValueError('sequence_impedance_matrix is not valid.')
def get_positive_sequence_impedance(sequence_impedance_matrix):
try:
return sequence_impedance_matrix[1,1]
except:
raise ValueError('sequence_impedance_matrix is not valid.')
def get_negative_sequence_impedance(sequence_impedance_matrix):
try:
return sequence_impedance_matrix[2,2]
except:
raise ValueError('sequence_impedance_matrix is not valid.')
def get_sequence_impedance_matrix(phase_impedance_matrix):
a=cmath.exp(complex(0,2./3*cmath.pi))
A=np.array([[complex(1.0,0),complex(1.0,0),complex(1.0,0)],
[complex(1.0,0),a**2,a],
[complex(1.0,0),a,a**2]])
A_inv=1./3.0*np.array([[complex(1.0,0),complex(1.0,0),complex(1.0,0)],
[complex(1.0,0),a,a**2],
[complex(1.0,0),a**2,a]])
return np.dot(A_inv,np.dot(phase_impedance_matrix,A))
def kron_reduction(primitive_impedance_matrix):
zij=primitive_impedance_matrix[:3,:3]
zin=primitive_impedance_matrix[:3,-1][:,np.newaxis]
znn=primitive_impedance_matrix[3,3]
znj=primitive_impedance_matrix[-1,:3]
return zij-np.dot(zin,np.dot(1.0/znn,znj)[np.newaxis])
def carson_equation_self(ri,GMRi):
return complex(ri+.0953,.12134*(np.log(1.0/GMRi)+7.93402))
def carson_equation(Dij):
return complex(.09530, .12134*(np.log(1.0/Dij)+7.93402))
def get_primitive_impedance_matrix(dist_matrix, GMR_list, r_list):
primitive_impedance_matrix=[]
n_rows,n_cols=dist_matrix.shape
for i in range(n_rows):
primitive_impedance_matrix.append([])
for j in range(n_cols):
if i==j:
primitive_impedance_matrix[-1].append(carson_equation_self(r_list[i],GMR_list[i]))
else:
primitive_impedance_matrix[-1].append(carson_equation(dist_matrix[i,j]))
return np.array(primitive_impedance_matrix)
def get_sequence_impedances(dist_matrix, GMR_list, r_list):
prim=get_primitive_impedance_matrix(dist_matrix, GMR_list, r_list)
phase=kron_reduction(prim)
print phase
seq=get_sequence_impedance_matrix(phase)
return get_zero_sequence_impedance(seq), get_positive_sequence_impedance(seq), get_negative_sequence_impedance(seq)
dist_matrix=np.array([[0,2.5,7.0,5.6569],
[2.5,0,4.5,4.272],
[7.0,4.5,0,5.0],
[5.6569,4.272,5.0,0]])
GMR_list=[0.0244,0.0244,0.0244,0.00814]
r_list=[0.306,0.306,0.306,0.592]
get_sequence_impedances(dist_matrix, GMR_list, r_list)
import opendssdirect as dss
dss.dss_lib.DSSPut_Command('new circuit.IEEE13Nodeckt basekv=115 pu=1.0001 phases=3 bus1=SourceBus Angle=30 MVAsc3=20000 MVASC1=21000 ')
dss.dss_lib.DSSPut_Command('New Line.650632 Phases=3 Bus1=RG60.1.2.3 Bus2=632.1.2.3 R1=0.30606998553726722 R0=0.77351087368443594 X0=1.9372561657931306 X1=0.6270002376486794 Length=2000 units=ft')
dss.Lines.First()
dss.utils.lines_to_dataframe()['RMatrix'].values
dss.utils.lines_to_dataframe()['XMatrix'].values
1.0/3.0*(2*0.30606998553726722+0.77351087368443594)
1.0/3.0*(0.77351087368443594-0.30606998553726722)
1.0/3.0*(1.9372561657931306-0.6270002376486794)
print dss.utils.lines_to_dataframe()
a,b,c=sorted(['b','a','c'])
a
a={}
for k in range(10):
a[k]={'test':k**2}
a
b={}
for k in range(4,16):
b[k]={ 'biscotte':k-2}
b
def merge_dict(d1,d2):
for k2,v2 in d2.iteritems():
if k2 in d1:
d1[k2].update(v2)
else:
d1[k2]=v2
return d1
merge_dict(a,b)
a.update(b)
a
b
class test:
def __init__(self):
self.a=1
self.b=2
self.c=3
tt=test()
dd={'a':3,'b':9}
for k,v in dd.iteritems():
setattr(tt,k,v)
tt.c
mat=[[0,1,None],[1,0,None],[None,None,0]]
mat=np.array(mat)
mat
mat[mat!=None]
pos=np.array([[1,1],[None,None],[4,4],[None,None]])
pos==np.array([None,None])
```
| github_jupyter |
here uplode ur own token from kaggle or seach on google
otherwise follow this link
https://stackoverflow.com/questions/49310470/using-kaggle-datasets-in-google-colab
```
from google.colab import files
files.upload()
!pip install -q kaggle
!mkdir -p /root/.kaggle
!cp /content/kaggle.json /root/.kaggle
#!kaggle datasets list
! kaggle competitions download -c expedia-personalized-sort
```
we want to unzip data so simply use with !unzip and followed by path
```
!unzip /content/data.zip
```
imported necessary lib
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import sklearn
import seaborn as sns
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
```
#imported training data and test data into dataframe using pandas
```
train_df=pd.read_csv('/content/train.csv')
train_df.head()
train_df.keys()
#test_Df=pd.read_csv('/content/train.csv')
train_df.isna().sum()
train_df.describe(include='all')
```
# **total no of booking_bool values which are here labels**
```
int(list(train_df['booking_bool']).count(0))+int(list(train_df['booking_bool']).count(1))
```
# **hence here are total no of clusters are 2 as seen in booking_bool column**
```
train_df.head()
plt.figure(figsize=(10,8))
sns.barplot(x='booking_bool',y='visitor_location_country_id',data=train_df[:10000])
plt.figure(figsize=(10,8))
sns.barplot(y='price_usd',x='srch_room_count',data=train_df[:10000])
plt.figure(figsize=(10,8))
sns.barplot(y='price_usd',x='srch_booking_window',data=train_df[:10000])
```
price_usd srch_booking_window srch_saturday_night_bool
```
plt.figure(figsize=(10,8))
sns.barplot(y='price_usd',x='srch_saturday_night_bool',data=train_df[:500])
plt.figure(figsize=(10,8))
sns.barplot(y='srch_booking_window',x='srch_saturday_night_bool',data=train_df[:500])
train_df.dtypes
date_time=train_df.pop('date_time')
features = StandardScaler().fit_transform(train_df[:10000][:-2].values)
from sklearn.impute import SimpleImputer
mean_imputer = SimpleImputer(missing_values=np.nan, strategy='mean')
features_mean_imputed = mean_imputer.fit_transform(features)
pca = PCA(n_components=0.90, whiten=True)
features_pca = pca.fit_transform(features_mean_imputed)
print("Original number of features:", features.shape[1])
print("Reduced number of features:", features_pca.shape[1])
temp=pd.DataFrame(features_pca)
temp
train_df.loc[train_df['prop_id'] == 104517]
train_df['date_time']=pd.Series(date_time)
df = train_df.loc[train_df['prop_id'] == 104517]
df = df.loc[df['visitor_location_country_id'] == 219]
df = df.loc[df['srch_room_count'] == 1]
df = df[['date_time', 'price_usd', 'srch_booking_window', 'srch_saturday_night_bool']]
df.describe()
#date=pd.to_datetime(train_df['date_time'])
#train_df['date']=pd.Series(date)
#train_df.pop('date_time')
#sns.barplot(y='booking_bool',x='date',data=train_df[:5000])
df = df.loc[df['price_usd'] < 5584]
df['price_usd'].describe()
print(df['date_time'].min())
print(df['date_time'].max())
df['date_time'].describe()
df['date_time'] = pd.to_datetime(df['date_time'])
df.head()
df.plot(x = 'date_time',
y = 'price_usd',
figsize = (16, 8))
plt.xlabel('dates')
plt.ylabel('USD')
plt.title('Time series of room price by date of search')
a = df.loc[df['srch_saturday_night_bool'] == 0, 'price_usd']
b = df.loc[df['srch_saturday_night_bool'] == 1, 'price_usd']
plt.figure(figsize = (16, 8))
plt.hist(a, bins = 80,
alpha = 0.3,
label = 'search w/o Sat night stay')
plt.hist(b, bins = 80,
alpha = 0.3,
label = 'search w/ Sat night stay')
plt.xlabel('Price')
plt.ylabel('Freq')
plt.legend()
plt.title('Sat night search')
plt.plot();
sns.distplot(df['price_usd'],
hist = False, label = 'USD')
sns.distplot(df['srch_booking_window'],
hist = False, label = 'booking window')
plt.xlabel('dist')
sns.despine()
sns.pairplot(df)
df = df.sort_values('date_time')
df['date_time_int'] = df.date_time.astype(np.int64)
df
from sklearn.cluster import KMeans
data = df[['price_usd', 'srch_booking_window', 'srch_saturday_night_bool']]
n_cluster = range(1, 50)
kmeans = [KMeans(n_clusters = i).fit(data) for i in n_cluster]
scores = [kmeans[i].score(data) for i in range(len(kmeans))]
kmeans,scores
fig, ax = plt.subplots(figsize = (16, 8))
ax.plot(n_cluster, scores, color = 'orange')
plt.xlabel('clusters num')
plt.ylabel('score')
plt.title('Elbow curve for K-Means')
plt.show();
test = pd.read_csv('/content/test.csv')
km = KMeans(n_clusters = 20).fit(data)
X = df[['price_usd', 'srch_booking_window', 'srch_saturday_night_bool']]
X = X.reset_index(drop = True)
s=km.predict(X)
s
df['date_time'].values.shape
new_df=pd.DataFrame()
new_df['date_time']=pd.Series(df['date_time'].values)
new_df['price in usd']=pd.Series(s)
plt.figure(figsize=(100,5))
sns.barplot(x='date_time',y='price in usd',data=new_df[:30])
```
| github_jupyter |
```
from nltk.corpus import stopwords
from nltk.tokenize import word_tokenize
from nltk.tokenize import RegexpTokenizer
from nltk import pos_tag
from nltk.stem import PorterStemmer, WordNetLemmatizer
import re
import nltk
import pprint as pp
import db_scripts
import pprint
import pickle
import json
def get_credentials():
pkl_file = open('.cred.pkl', 'rb')
data = pickle.load(pkl_file)
return data[0], data[1], data[2], data[3]
def load_stop_words(stop_word_file):
stop_words = []
fo = open(stop_word_file)
for line in fo:
if line.strip()[0:1] != "#":
for word in line.split(): # in case more than one per line
stop_words.append(word)
fo.close()
return stop_words
def load_keywords_from_bucket(keyword_file):
word_map = dict()
file = open(keyword_file)
for line in file:
key = line.split(':')
#print(key)
x, y = Keywords(key[1]).fetch()
y = list(set(y))
if key[0] in word_map:
val = word_map[key[0]]
val = [j for i in zip(val,y) for j in i]
word_map[key[0]] = val
else:
# val = list()
# val.append(y)
word_map[key[0]] = y
file.close()
return word_map
# x = load_keywords_from_bucket('keywords.txt')
# fp = open('keywords.json','w')
# import json
# json.dump(x, fp, sort_keys=True, indent=4)
# fp.close()
def load_keywords_from_json(json_file):
fp = open(json_file, 'r')
word_map = json.load(fp)
#print(word_map['Algorithm'])
fp.close()
return word_map
#load_keywords_from_json('keywords.json')
def remove_puncts(text):
tokenizer = RegexpTokenizer(r'\w+')
words_no_punct = tokenizer.tokenize(text)
data = ' '.join(words_no_punct)
return data
def is_number(s):
try:
float(s) if '.' in s else int(s)
return True
except ValueError:
return False
def generate_candidate_keywords(data, stop_words):
words = [w for w in word_tokenize(
data) if w.lower() not in stop_words and not is_number(w)]
return words
def stem_lemmatize(keywords):
lemmatizer = WordNetLemmatizer()
stemmer = PorterStemmer()
tokenizer = RegexpTokenizer(r'\w+')
x = []
y = []
for word in keywords:
x.append(stemmer.stem(word))
y.append(lemmatizer.lemmatize(word))
return x
def get_percentage_mapping(topic, lemma):
mapping = [value for value in topic if value in lemma]
#mapping = [list(filter(lambda x: x in topic, sublist)) for sublist in lemma]
#print(mapping)
return (len(mapping)/len(topic))
course_map = {}
def modify_course_map(domain, course_id):
for topic in domain:
if topic in course_map.keys():
course_map[topic].append(course_id)
else:
course_map[topic] = [course_id]
modify_course_map(['Data Science', 'Algorithms'], 'CSC522')
course_map
def database_retrieve():
username, password, db_name, collection_name = get_credentials()
all_courses = db_scripts.db_fetch_all(username, password, db_name, collection_name)
for course in all_courses:
percent = Keywords(course['description']).map_keywords()
modify_course_map(percent, course['course_id'])
database_retrieve()
class Keywords:
def __init__(self, text):
self.text = text
self.stop_words = load_stop_words("FoxStopList.txt")
def fetch(self):
candidate = remove_puncts(self.text)
keywords = generate_candidate_keywords(candidate, self.stop_words)
# table = stem_lemmatize(keywords)
st = stem_lemmatize(keywords)
return keywords, st
def map_keywords(self):
topic_map = load_keywords_from_bucket("keywords.txt")
stem, lemma = self.fetch()
percent_mapping = {}
topic = ''
maxi = 0
for key in topic_map.keys():
percent = get_percentage_mapping(topic_map[key],lemma)
percent_mapping[key] = percent
percent = sorted(percent_mapping.items(), key=lambda x:x[1])
t = [x[0] for x in percent[::-1][:2]]
return t
alda = "Introduction to the problems and techniques for automated discovery of knowledge in databases. Topics include representation, evaluation, and formalization of knowledge for discovery; classification, prediction, clustering, and association methods.Selected applications in commerce, security, and bioinformatics. Students cannot get credit for both CSC 422 and CSC 522."
algo = "Algorithm design techniques: use of data structures, divide and conquer, dynamic programming, greedy techniques, local and global search. Complexity and analysis of algorithms: asymptotic analysis, worst case and average case, recurrences, lower bounds, NP-completeness. Algorithms for classical problems including sorting, searching and graph problems (connectivity, shortest paths, minimum spanning trees)."
os = "Fundamental issues related to the design of operating systems. Process scheduling and coordination, deadlock, memory management and elements of distributed systems."
ai = "Introduction to and overview of artificial intelligence. Study of AI programming language such as LISP or PROLOG. Elements of AI problem-solving technique. State spaces and search techniques. Logic, theorem proving and associative databases. Introduction to knowledge representation, expert systems and selected topics including natural language processing, vision and robotics."
ads = "Complex and specialized data structures relevant to design and development of effective and efficient software. Hardware characteristics of storage media. Primary file organizations. Hashing functions and collision resolution techniques. Low level and bit level structures including signatures, superimposed coding, disjoint coding and Bloom filters. Tree and related structures including AVL trees, B*trees, tries and dynamic hashing techniques."
#topic, percent = Keywords(ai).map_keywords()
#topic, percent = Keywords(alda).map_keywords()
#topic, percent = Keywords(os).map_keywords()
# topic, percent = Keywords(algo).map_keywords()
percent = Keywords(ads).map_keywords()
pprint.pprint(percent)
# print(t)
import pandas as pd
df = pd.DataFrame().from_dict(d)
df
d['predict'].append('CSC501')
df = pd.DataFrame().from_dict(d,orient='index').transpose()
df
import db_scripts
all_courses = db_scripts.db_fetch_all("wolfpal", "courses")
from collections import defaultdict
d = defaultdict()
for course in all_courses:
l, p = Keywords(course['desc']).fetch()
# print(course['desc'])
p = set(p)
for i in p:
if i in d:
d[i].append(course['branch']+course['number'])
else:
d[i] = [(course['branch']+course['number'])]
import pandas as pd
df = pd.DataFrame().from_dict(d,orient='index').transpose()
df
usersays = "I like AI and systems"
p,l = Keywords(usersays).fetch()
print(l)
for i in l:
if i in d:
print("Yeh course le: ",d[i])
```
| github_jupyter |
# Visualization
## TODO: k-NN + directed version (direction = style)
```
import collections
import numpy as np
import time
import datetime
import json
from tqdm import tqdm
import os
import tensorflow as tf
import seaborn as sns
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
from bokeh.io import output_notebook
from bokeh.plotting import figure, output_file, show, ColumnDataSource
from bokeh.models import HoverTool
output_notebook()
import data_utils_LMR
from data_utils_LMR import prepare_data,read_data, EncoderDecoder
from model import Vrae as Vrae_model
from batch import Generator
training_dir = 'logs/'
training_dir += 'no_char2word'
# sentiment analyzer
from nltk.sentiment.vader import SentimentIntensityAnalyzer
sentimentAnalyzer = SentimentIntensityAnalyzer()
def getSentimentScore(sentence):
scores = sentimentAnalyzer.polarity_scores(sentence)
return (scores['neg'], scores['neu'] ,scores['pos'])
class dotdict(dict):
"""dot.notation access to dictionary attributes"""
__getattr__ = dict.get
__setattr__ = dict.__setitem__
__delattr__ = dict.__delitem__
def string2bool(st):
if st.lower() == "true":
return True
else:
return False
with open(training_dir +'/flags.json', 'r') as fp:
FLAGS = dotdict(json.loads( fp.read() ) )
for k,v in FLAGS.iteritems():
print k,':',v
n_samples = 5000#int(FLAGS.batch_size)
```
## Prepare data
```
with open(training_dir +'/training_parameters.json', 'r') as fp:
training_parameters = json.loads( fp.read() )
# vocabulary encoder-decoder
encoderDecoder = EncoderDecoder()
num_symbols = encoderDecoder.vocabularySize()
#prepare_data(1000)
sentences, ratings = read_data( max_size=None,
max_sentence_size=training_parameters['seq_max'],
min_sentence_size=int(FLAGS.sequence_min),
test=False)
print len(sentences), " sentences"
if len(sentences) < n_samples:
n_samples = len(sentences) - 1
sns.distplot( [len(sent) for sent in sentences])
plt.show()
```
## Loading models
```
space_symbol = encoderDecoder.encode("I am")[1]
word_delimiters = [ data_utils_LMR._EOS, data_utils_LMR._GO, space_symbol ]
batch_gen = Generator(sentences, ratings, n_samples, word_delimiters)
num_iters = FLAGS.epoches * batch_gen.iterations_per_epoch()
# text decoder ( text <-> ids)
encoderDecoder = EncoderDecoder()
config = tf.ConfigProto(
device_count = {'GPU': 0}, # do not use GPU for testing
)
# load model
vrae_model = Vrae_model(char2word_state_size = int(FLAGS.char2word_state_size),
char2word_num_layers = int(FLAGS.char2word_num_layers),
encoder_state_size = int(FLAGS.encoder_state_size),
encoder_num_layers = int(FLAGS.encoder_num_layers),
decoder_state_size = int(FLAGS.decoder_state_size),
decoder_num_layers = int(FLAGS.decoder_num_layers),
latent_dim=int(FLAGS.latent_dim),
batch_size=n_samples,
num_symbols=num_symbols,
latent_loss_weight=float(FLAGS.latent_loss_weight),
dtype_precision=FLAGS.dtype_precision,
cell_type=FLAGS.cell,
peephole=FLAGS.peephole,
input_keep_prob=float(FLAGS.input_keep_prob),
output_keep_prob=float(FLAGS.output_keep_prob),
sentiment_feature = string2bool(FLAGS.use_sentiment_feature),
use_char2word = string2bool(FLAGS.use_char2word)
)
def zToXdecoded(session,z_sample,s_length):
x_reconstruct = vrae_model.zToX(session,z_sample,s_length)
return encoderDecoder.prettyDecode( np.argmax(x_reconstruct[0], axis= 1) )
```
## Generating test data
```
saver = tf.train.Saver()
#print train_dir
np.random.seed(13)
batch_gen.shuffle()
samples = []
with tf.Session(config=config) as sess:
saver.restore(sess, "./"+training_dir+'/model.ckp')
print
padded_batch_xs, batch_ys, batch_lengths, batch_weights, end_of_words, batch_word_lengths, max_length = batch_gen.next_batch()
vaderSentiments = [ getSentimentScore(encoderDecoder.prettyDecode(xx)) for xx in padded_batch_xs]
x_reconstruct,z_vals,z_mean_val,z_log_sigma_sq_val, losses = vrae_model.reconstruct( sess,
padded_batch_xs,batch_lengths,
batch_weights,
end_of_words,
batch_word_lengths,
vaderSentiments)
print "Done!"
print losses
vaderSentiments = [ getSentimentScore(encoderDecoder.prettyDecode(padded_batch_xs[i])) for i in xrange(n_samples)]
```
## Reconstruction
```
#np.random.seed(13)
for i in range(10):
i = int(np.random.random()*n_samples)
i = i
print "sentiment:", vaderSentiments[i],"| rating:", batch_ys[i]
print encoderDecoder.prettyDecode( padded_batch_xs[i] )
print encoderDecoder.prettyDecode( np.argmax(x_reconstruct[i], axis= 1) )
print "------------------------------------------"
```
## Reconstruction in the Latent Space
```
# dimension reduction
from sklearn.manifold import TSNE
X = np.array(z_vals)
model = TSNE(n_components=2, random_state=0)
np.set_printoptions(suppress=True)
zs_reduced = model.fit_transform(X)
xs = [ zs_reduced[i,0] for i in xrange(n_samples) ]
ys = [ zs_reduced[i,1] for i in xrange(n_samples) ]
from bokeh.layouts import column
from bokeh.models import CustomJS, ColumnDataSource, Select
from bokeh.plotting import figure, output_file, show
!export BOKEH_LOG_LEVEL=error
output_file("latent_space.html")
inputs = [encoderDecoder.prettyDecode(x) for x in padded_batch_xs]
M =max(batch_lengths)
binary_rating = [ int(r > 5) for r in batch_ys]
colors_sent = [ "#%02x%02x%02x" % ( 100 + 150 * r[0] , 100 + 150 * r[2] , 100 + 100 * r[1] ) for r in vaderSentiments ]
color_rating = [ "#%02x%02x%02x" % (255 * (1-r) , 100, 255*r) for r in binary_rating ]
colors_lengths = [ "#%02x%02x%02x" % ( ( 255 * (float(r)/float(M))), 50, 255 - 255 * (float(r)/float(M))) for r in batch_lengths ]
hasQuestionMark = [ int("?" in x) for x in inputs]
colors_questionMark = ["#%02x%02x%02x" % (255 * (1-r) , 100, 255*r) for r in hasQuestionMark]
is_past_voice = [ int("was" in x) or int("were" in x) or int("did" in x) or int("had" in x) for x in inputs]
colors_past = ["#%02x%02x%02x" % (255 * (1-r) , 100, 255*r) for r in is_past_voice]
source = ColumnDataSource(
data=dict(
x=xs,
y=ys,
input=inputs,
output= [encoderDecoder.prettyDecode(np.argmax(y, axis= 1) ) for y in x_reconstruct],
rating=batch_ys,
sent= vaderSentiments,
rating_color=color_rating,
sentiment_color=colors_sent,
lenght_color=colors_lengths,
questionMark_color=colors_questionMark,
past_color = colors_past,
lengths=batch_lengths,
)
)
hover = HoverTool(
tooltips=[
("index", "$index"),
("(x,y)", "($x, $y)"),
("input", "@input"),
("output", "@output"),
("rating", "@rating"),
("lengths", "@lengths"),
]
)
p = figure(plot_width=800, plot_height=600, tools=[hover],title="Latent space")
cir = p.circle('x', 'y', size=9, source=source, fill_color="sentiment_color", alpha=0.8)
callback = CustomJS(args=dict(cir=cir,source=source), code="""
var selected_color = cb_obj.value;
selected_color = selected_color
console.log(selected_color);
cir.glyph.line_color.field = selected_color;
cir.glyph.fill_color.field = selected_color;
source.trigger("change")
""")
select = Select(title="Color:", value="sentiment", options=["sentiment_color", "rating_color", "lenght_color", "questionMark_color", "past_color"], callback=callback)
layout = column(select, p)
show(layout)
```
## Sentiment Distribution
```
import seaborn as sns
cols = sns.color_palette()
pos = []
neg = []
neu = []
major_sent = np.argmax(vaderSentiments, axis = 1)
for i in xrange(n_samples):
if major_sent[i] == 2:
pos.append( z_mean_val[i,:] )
elif major_sent[i] == 0:
neg.append( z_mean_val[i,:] )
else:
neu.append( z_mean_val[i,:] )
print len(pos), "positive sentences"
print len(neg), "negative sentences"
print len(neu), "neutral sentences"
pos = np.array(pos)
neg = np.array(neg)
neu = np.array(neu)
side_lenght = int(np.sqrt(int(FLAGS.latent_dim)))
if side_lenght**2 < int(FLAGS.latent_dim):
side_lenght +=1
f, axs = plt.subplots(ncols=side_lenght, nrows=side_lenght, sharey=True, figsize=(10, 10))
for i in xrange(side_lenght):
for j in xrange(side_lenght):
k = i*side_lenght+j
if k < int(FLAGS.latent_dim):
sns.distplot( neu[:,k], ax=axs[i,j], hist=False, color= cols[0] )
sns.distplot( pos[:,k], ax=axs[i,j], hist=False, color= cols[1] )
sns.distplot( neg[:,k], ax=axs[i,j], hist=False, color= cols[2])
plt.show()
```
## The Sentiment Dimension
```
import scipy
import random
KLs = []
for k in xrange(int(FLAGS.latent_dim)):
a = list(neg[:,k])
b = list(pos[:,k])
kl = np.abs( np.mean(a) - np.mean(b))
KLs.append( kl )
sorted_kls = sorted( enumerate(KLs) , key = lambda x: x[1], reverse=True )
for k,kl in sorted_kls:
print k,kl
k=sorted_kls[0][0]
sns.distplot( neu[:,k], hist=False, color= cols[0] )
sns.distplot( pos[:,k], hist=False, color= cols[1] )
sns.distplot( neg[:,k], hist=False, color= cols[2])
plt.show()
ks = sorted_kls[0][:3]
dx = 2
saver = tf.train.Saver()
with tf.Session(config=config) as sess:
saver.restore(sess, "./"+training_dir+'/model.ckp')
u0 = "I like this movie."
u1 = "I recommend this movie."
u = u0
#print u
z0 = vrae_model.XToz(sess, *encoderDecoder.encodeForTraining(u), sentiment=getSentimentScore(u))[0]
#z0 = np.zeros(int(FLAGS.latent_dim))
dz = np.zeros(int(FLAGS.latent_dim))
dz[k] = dx
z1 = z0 + dz
z2 = z0 - dz
print "distance between two points:",np.linalg.norm(z2-z1),"\n"
zs = []
for t in np.linspace(0,1,30):
zs.append( (1-t) * z1 + t * z2 )
for z_ in zs:
print zToXdecoded(sess, z_ , 45 )
```
## Dimension pairplot
cross check dimensions
```
import pandas as pd
major_sent = np.argmax(vaderSentiments, axis = 1)
sent_df = pd.DataFrame( z_mean_val )
sent_df['major_sent'] = major_sent
#sns.pairplot(sent_df, hue='major_sent', diag_kind="kde")
#plt.show()
```
## Continuous space: Homotopies
```
saver = tf.train.Saver()
with tf.Session(config=config) as sess:
saver.restore(sess, "./"+training_dir+'/model.ckp')
print
# show interpolations
u0 = ["I loved it." , "I hated this film."]
u01 = ["I loved it." , "It was terrible."]
u1 = ["I loved this movie!.", "I hated this movie!"]
u2 = ["The best movie I've seen!", "The worst movie ever made."]
u3 = ["great movie.", "terrible movie."]
u4 = ["that's actually pretty good." , "That was a failure."]
u5 = ["I didn't laugh at all.", "I wanted to love this movie."]
u6 = ["so bad that it's really bad" , "Where is the acting?"]
u7 = ["I love old movies.", "I prefer old movies."]
u8 = ["the music is very bad.", "the music is just awful."]
u9 = ["awesome!", "terrible."]
u10 = ["awful." , "pretty worthless."]
u11 = ["yes you do." , "no you don't."]
u12 = ["The acting was really bad." , "The acting was really good!"]
u13 = ["This film was fascinatingly stupid." , "This is an excellent film."]
u14 = ["I don't recommend it to anyone.", "This is a really really bad movie."]
u = u14
z1 = vrae_model.XToz(sess, *encoderDecoder.encodeForTraining(u[0]),sentiment=getSentimentScore(u[0]))[0]
z2 = vrae_model.XToz(sess, *encoderDecoder.encodeForTraining(u[1]),sentiment=getSentimentScore(u[1]))[0]
#z1 = np.zeros(16)
#z2 = 0.1 * np.ones(16)
print "distance between two points:",np.linalg.norm(z2-z1),"\n"
zs = []
for t in np.linspace(0,1,50):
zs.append( (1-t) * z1 + t * z2 )
sents = [zToXdecoded(sess, z_ , 45 ) for z_ in zs]
appeared = []
for x in sents:
if x not in appeared:
print x
appeared.append(x)
```
## Reconstruction: Reconstructing using Model's knowledge
```
saver = tf.train.Saver()
with tf.Session(config=config) as sess:
saver.restore(sess, "./"+training_dir+'/model.ckp')
u0 = "I like this movie."
u1 = "It was terrible."
u2 = "I recommend it."
u3 = "I loved it."
us = [u0,u1,u2,u3]
for u in us:
z = vrae_model.XToz(sess, *encoderDecoder.encodeForTraining(u), sentiment=getSentimentScore(u))[0]
print u, "->",
print zToXdecoded(sess,z,40)
```
## Vector Translations
```
saver = tf.train.Saver()
with tf.Session(config=config) as sess:
saver.restore(sess, "./"+training_dir+'/model.ckp')
print
a = "I liked it."
b = "I didn't like it."
c = "I recommend this movie."
#a = "I like it"
#b = "I liked it."
#c = "I recommend it."
#a = "I love this movie!"
#b = "I like this movie."
#c = "I love the acting!"
za = vrae_model.XToz(sess, *encoderDecoder.encodeForTraining(a), sentiment=getSentimentScore(a) )[0]
zb = vrae_model.XToz(sess, *encoderDecoder.encodeForTraining(b), sentiment=getSentimentScore(b))[0]
zc = vrae_model.XToz(sess, *encoderDecoder.encodeForTraining(c), sentiment=getSentimentScore(c))[0]
# translation
zd = zc + (zb - za)
print "a \t\t|", a,"|", zToXdecoded(sess, za , 40 )
print "b \t\t|", b,"|", zToXdecoded(sess, zb , 40 )
print "c \t\t|", c,"|", zToXdecoded(sess, zc , 40 )
print
print "c + b-a \t|", zToXdecoded(sess, zd , 40 )
```
## Neighborhood
```
saver = tf.train.Saver()
with tf.Session(config=config) as sess:
saver.restore(sess, "./"+training_dir+'/model.ckp')
print
# show interpolations
u1 = "That's actually pretty good."
u2 = "I love it."
u = u1
z1 = vrae_model.XToz(sess, *encoderDecoder.encodeForTraining(u), sentiment=getSentimentScore(u))[0]
print z1
print
r = 1
zs = []
for t in range(50):
z2 = [ z_ + r * np.random.random() for z_ in z1 ]
zs.append( z2)
for z_ in zs:
print zToXdecoded(sess, z_ , 40 )
```
$$\log p(x) = \sum_{i=1}^N q(z^{(i)} | x ) \log p(x)$$
$$ \sum_{i=1}^N q(z^{(i)} | x ) \left [ \frac{q_\phi(z^{(i)} | x) p_\theta(x,z)}{p_\theta(z|x^{(i)}) q_\phi(z^{(i)} | x) } \right ] $$
$$ \sum_{i=1}^N q(z^{(i)} | x ) \log \frac{q(z^{(i)} | x )}{p_\theta(z|x^{(i)})} + q(z^{(i)} | x ) \log p_\theta(x,z) - q(z^{(i)} | x ) log q(z^{(i)} | x ) $$
$$ D_{KL} (q(z^{(i)} | x ) || p_\theta(z|x^{(i)})) + E_{q(z^{(i)} | x )} [- \log q(z^{(i)} | x ) + \log p_\theta (x,z)] $$
```
& = \sum_{i=1}^N q(z^{(i)} | x ) \left [ \frac{q_\phi(z^{(i)} | x) p_\theta(x,z)}{p_\theta(z|x^{(i)}) q_\phi(z^{(i)} | x) } \right ] \\
& = \sum_{i=1}^N q(z^{(i)} | x ) \log \frac{q(z^{(i)} | x )}{p_\theta(z|x^{(i)})} + q(z^{(i)} | x ) \log p_\theta(x,z) - q(z^{(i)} | x ) log q(z^{(i)} | x ) \\
& = D_{KL} (q(z^{(i)} | x ) || p_\theta(z|x^{(i)})) + E_{q(z^{(i)} | x )} [- \log q(z^{(i)} | x ) + \log p_\theta (x,z)]
```
$$ \mathcal{L} (\theta , \phi ; \theta^{(i)}) = E_{q_\phi(z| x^{(i)} )} [- \log q_\phi( | x^{(i)} ) + \log p_\theta (x^{(i)},z)] $$
$$- D_{KL} ( q_\phi(z | x^{(i)} ) || p_\theta(z) ) + E_{q_\phi(z| x^{(i)} )} [ \log p_\theta(x^{(i)} | z)]$$
# Vader
```
vaderSentiments = [ getSentimentScore(encoderDecoder.prettyDecode(xx)) for xx in padded_batch_xs]
for k in range(15):
k = int( np.random.random() * len(padded_batch_xs) )
t = encoderDecoder.prettyDecode(padded_batch_xs[k])
s = getSentimentScore(t)
print t , ' & ', s[0] , ' & ',s[1],' & ', s[2], '\\\ \\hline'
```
bad, sad, and rubbish. & 0.767 & 0.233 & 0.0 \\ \hline
great old movie drama. & 0.0 & 0.423 & 0.577 \\ \hline
i always enjoy thomas gomez. & 0.0 & 0.484 & 0.516 \\ \hline
it is a perfect scene. & 0.0 & 0.448 & 0.552 \\ \hline
this film sucks! & 0.583 & 0.417 & 0.0 \\ \hline
the two actors are very natural. & 0.0 & 0.642 & 0.358 \\ \hline
it was hilarious. & 0.0 & 0.426 & 0.574 \\ \hline
it was dumb. & 0.623 & 0.377 & 0.0 \\ \hline
well acted and good enough plot. & 0.0 & 0.444 & 0.556 \\ \hline
it was weak, very very weak. & 0.592 & 0.408 & 0.0 \\ \hline
it' s horrible. & 0.778 & 0.222 & 0.0 \\ \hline
nothing more, nothing less. & 0.0 & 1.0 & 0.0 \\ \hline
she' s a good actress. & 0.0 & 0.408 & 0.592 \\ \hline
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
plt.rcParams.update({
"text.usetex": True,
"font.sans-serif": ["Helvetica"]})
```
# Driving forces for moving systems
In this case study, you want to accelerate a 0.1-kg flywheel with a
piston. The desired
acceleration of the flywheel is $\alpha=50~rad/s^2.$ The piston is
attached to the link and creates a horizontal driving force. This
example demonstrates how we can describe constraints in mathematical
forms.
We have two moving bodies:
1. a flywheel with radius $r=0.2~m$ and mass $m=0.1~kg$
2. a massless connecting link
Both objects move in a horizontal plane, so their positions have two
degrees of freedom to describe position and one degree of freedom that
describes orientation.
$DOF = 3 + 3 = 6$
```
from IPython.display import SVG
SVG(filename = './images/piston-flywheel.svg')
```
## Describing constraints
There are six degrees of freedom for a flywheel and a link, but there
are 6 constraints on the system's motion:
1. the flywheel is pinned to the ground in the x-dir
1. the flywheel is pinned to the ground in the y-dir
2. the top of the link is pinned to the flywheel
2. the top of the link is pinned to the flywheel
3. the bottom of the link slides along the horizontal line
4. the angle of the flywheel has acceleration, $\alpha=50~rad/s^2$
$system~DOF = 6 - 6 = 0~DOF$
> __Note:__ In general, a pin in a planar system creates 2 constraints on motion.
You should recognize at this point that there are _0 differential
equations to solve_. Once you have calculated the kinematics based upon
the system constraints, you can plug in the values and solve for force
of the piston. So, start with the kinematic description of motion.
$\mathbf{r}_2 =\mathbf{r}_{2/3} + \mathbf{r}_3$
```
SVG(filename = './images/flywheel-constraints.svg')
```
$r(\sin\theta\hat{i} - \cos\theta \hat{j}) =
-L(\cos\theta_{L}\hat{i}-\sin\theta_L\hat{j}) + d\hat{i}$
this description creates two independent equations
1. $-r\sin\theta = -L\cos\theta_{L}+ d$
2. $r\cos\theta = -L\sin\theta_L$
The constraint on $\theta$ says that $\theta(t)=\frac{\alpha t^2}{2}$.
You need to solve for $\theta_L$ and $d$ to determine the full
state of the system.
$\theta_L = \sin^{-1}\left(\frac{r}{L}\cos\theta\right)$
$d = L\cos\theta_{L}-r\sin\theta$
```
r = 0.2
L = 0.5
theta = np.linspace(0,2*np.pi,100)
thetaL = np.arcsin(r/L*np.cos(theta))
d = L*np.cos(thetaL)-r*np.sin(theta)
f, ax = plt.subplots()
ax.plot(theta*180/np.pi, thetaL*180/np.pi,'b-',label = r'\theta_L')
ax.set_xlabel(r'$\theta$ (degrees)')
ax.set_ylabel(r'$\theta_L$ (degrees)',color = 'blue')
plt.xticks(np.arange(0,360,30))
ax2 = ax.twinx()
ax2.plot(theta*180/np.pi, d, 'g--', label = 'piston d')
ax2.set_ylabel('d (meter)', color = 'green');
plt.show()
```
---
### Exercise:
What is the angular velocity and angular acceleration of the link?
$\dot{\theta}_L$ and $\ddot{\theta}_L$
---
Now, we have solved all of the kinematics we need to
solve for the applied piston force. Sometimes, you can
look at the Newton-Euler equations for the syatem as a
whole, but here we need to separate each component and
sum forces and moments.
Flywheel:
$\sum\mathbf{F}=\mathbf{0}$
$\sum M_G = I \alpha$
> __Note:__ If a body is moving, you either have to sum moments about a fixed point or its center of mass. Otherwise, the description of angular acceleration is more involved.
Link:
$\sum\mathbf{F}=\mathbf{0}$
$\sum M = 0$
> __Note:__ when links, cables, or pulleys are assumed to
> be massless they still obey Newtonian mechanics, but
> momentum is always zero. So the sum of forces or
> moments is equal to 0.
The three kinetic equations for the wheel are as such,
1. $\sum \mathbf{F}\cdot\hat{i} = F_{2x}+F_{1x} = 0$
2. $\sum \mathbf{F}\cdot\hat{j} = F_{2y}+F_{1y} = 0$
3. $\sum M_{1} = r\hat{e}_r \times (F_{2x}\hat{i}+F_{2y}\hat{j}) = \frac{mr^2}{2}\alpha$
and the three kinetic equations for the link are as such,
1. $\sum \mathbf{F}\cdot\hat{i} = R-F_{2x} = 0$
2. $\sum \mathbf{F}\cdot\hat{j} = -F_{2y} - F_{piston} = 0$
3. $\sum M_{2} = L\hat{b}_1 \times (-F_{piston}\hat{i} + R\hat{j}) = 0$
The third equation for the link is rearranged to solve for $R =
F\tan\theta_L$. The third equation for the flywheel is rearranged to
solve for $F$ as such
$rF\cos\theta -rF\tan\theta_L\sin\theta = \frac{mr^2}{2}\alpha$
finally arriving at
$F = \frac{mr\alpha}{2}\left(\cos\theta-\tan\theta_L\sin\theta
\right)^{-1}$
Plotted below as a function of flywheel angle, $\theta$,
```
m = 0.1
F = m*r/2*(np.cos(theta)-np.tan(thetaL)*np.sin(theta))**(-1)
plt.plot(theta*180/np.pi, F)
plt.ylim(-0.1,0.1)
plt.xticks(np.arange(0,360,30))
plt.xlabel(r'$\theta$ (degrees)')
plt.ylabel('Piston Force');
```
| github_jupyter |
<a href="https://colab.research.google.com/github/yohanesnuwara/66DaysOfData/blob/main/D14_EDA_NLP.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Exploratory Data Analysis for NLP
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from collections import defaultdict, Counter
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
from nltk.util import ngrams
from sklearn.feature_extraction.text import CountVectorizer
!wget 'https://github.com/yohanesnuwara/datasets/raw/master/abcnews-date-text.csv.zip'
!unzip '/content/abcnews-date-text.csv.zip'
news = pd.read_csv('/content/abcnews-date-text.csv', nrows=10000)
news.head()
```
## Character-level, word-level, frequencies, and stopwords
```
# Plot histogram of number of characters
news['headline_text'].str.len().hist()
# Plot histogram of number of words
news['headline_text'].str.split().map(lambda x: len(x)).hist()
# Plot histogram of average number of words
news['headline_text'].str.split().\
apply(lambda x : [len(i) for i in x]). \
map(lambda x: np.mean(x)).hist()
```
Stopwords are the words that are most commonly used in any language such as “the”,” a”,” an” etc. As these words are probably small in length these words may have caused the above graph to be left-skewed.
```
stop = set(stopwords.words('english'))
corpus=[]
new= news['headline_text'].str.split()
new=new.values.tolist()
corpus=[word for i in new for word in i]
dic = defaultdict(int)
for word in corpus:
if word in stop:
dic[word]+=1
top=sorted(dic.items(), key=lambda x:x[1],reverse=True)[:10]
x,y=zip(*top)
plt.bar(x,y)
```
Stopwords such as "to", "in" and "for" dominate in news headlines.
```
# which words other than these stopwords occur frequently.
counter=Counter(corpus)
most=counter.most_common()
x, y= [], []
for word,count in most[:40]:
if (word not in stop):
x.append(word)
y.append(count)
sns.barplot(x=y,y=x)
```
## N-gram exploration
```
# Bigram of a sentence. Trigram=3, and so on.
list(ngrams(['I' ,'went','to','the','river','bank'], 2))
```
Represent the vocabularies through vectorizing.
```
def get_top_ngram(corpus, n=None):
vec = CountVectorizer(ngram_range=(n, n)).fit(corpus)
bag_of_words = vec.transform(corpus)
sum_words = bag_of_words.sum(axis=0)
words_freq = [(word, sum_words[0, idx])
for word, idx in vec.vocabulary_.items()]
words_freq =sorted(words_freq, key = lambda x: x[1], reverse=True)
return words_freq[:10]
# Top bigrams
top_n_bigrams = get_top_ngram(news['headline_text'],2)[:10]
x,y=map(list,zip(*top_n_bigrams))
sns.barplot(x=y,y=x)
# Top trigrams
top_n_bigrams = get_top_ngram(news['headline_text'],3)[:10]
x,y=map(list,zip(*top_n_bigrams))
sns.barplot(x=y,y=x)
```
We see trigrams like "to face court" and "anti war protesters" occur often in the above barplots with similar redundant trigrams like "face court over" and "anti war protest". With data cleansing, we can remove redundancy.
References:
* https://neptune.ai/blog/exploratory-data-analysis-natural-language-processing-tools
| github_jupyter |
# Assignment 4
Before working on this assignment please read these instructions fully. In the submission area, you will notice that you can click the link to **Preview the Grading** for each step of the assignment. This is the criteria that will be used for peer grading. Please familiarize yourself with the criteria before beginning the assignment.
This assignment requires that you to find **at least** two datasets on the web which are related, and that you visualize these datasets to answer a question with the broad topic of **sports or athletics** (see below) for the region of **Ann Arbor, Michigan, United States**, or **United States** more broadly.
You can merge these datasets with data from different regions if you like! For instance, you might want to compare **Ann Arbor, Michigan, United States** to Ann Arbor, USA. In that case at least one source file must be about **Ann Arbor, Michigan, United States**.
You are welcome to choose datasets at your discretion, but keep in mind **they will be shared with your peers**, so choose appropriate datasets. Sensitive, confidential, illicit, and proprietary materials are not good choices for datasets for this assignment. You are welcome to upload datasets of your own as well, and link to them using a third party repository such as github, bitbucket, pastebin, etc. Please be aware of the Coursera terms of service with respect to intellectual property.
Also, you are welcome to preserve data in its original language, but for the purposes of grading you should provide english translations. You are welcome to provide multiple visuals in different languages if you would like!
As this assignment is for the whole course, you must incorporate principles discussed in the first week, such as having as high data-ink ratio (Tufte) and aligning with Cairo’s principles of truth, beauty, function, and insight.
Here are the assignment instructions:
* State the region and the domain category that your data sets are about (e.g., **Ann Arbor, Michigan, United States** and **sports or athletics**).
* You must state a question about the domain category and region that you identified as being interesting.
* You must provide at least two links to available datasets. These could be links to files such as CSV or Excel files, or links to websites which might have data in tabular form, such as Wikipedia pages.
* You must upload an image which addresses the research question you stated. In addition to addressing the question, this visual should follow Cairo's principles of truthfulness, functionality, beauty, and insightfulness.
* You must contribute a short (1-2 paragraph) written justification of how your visualization addresses your stated research question.
What do we mean by **sports or athletics**? For this category we are interested in sporting events or athletics broadly, please feel free to creatively interpret the category when building your research question!
## Tips
* Wikipedia is an excellent source of data, and I strongly encourage you to explore it for new data sources.
* Many governments run open data initiatives at the city, region, and country levels, and these are wonderful resources for localized data sources.
* Several international agencies, such as the [United Nations](http://data.un.org/), the [World Bank](http://data.worldbank.org/), the [Global Open Data Index](http://index.okfn.org/place/) are other great places to look for data.
* This assignment requires you to convert and clean datafiles. Check out the discussion forums for tips on how to do this from various sources, and share your successes with your fellow students!
## Example
Looking for an example? Here's what our course assistant put together for the **Ann Arbor, MI, USA** area using **sports and athletics** as the topic. [Example Solution File](./readonly/Assignment4_example.pdf)
### I wrote code to scrape data from the websites on my local machine because it did not work for the server notebook.
<br>
I scraped the data from these websites on December 5, 2017. The webpages may change in the future.
<br>
* Basketball (Los Angeles Lakers) https://en.wikipedia.org/wiki/List_of_Los_Angeles_Lakers_seasons
* Hockey (Los Angeles Kings) https://en.wikipedia.org/wiki/List_of_Los_Angeles_Kings_seasons
* Baseball (Los Angeles Dodgers) https://en.wikipedia.org/wiki/List_of_Los_Angeles_Dodgers_seasons
* Football (Los Angeles Rams) https://en.wikipedia.org/wiki/List_of_Los_Angeles_Rams_seasons
<br>
<br>
I cleaned the data and saved them as .csv files. For the assignment, I uploaded the cleaned .csv files into my work area and loaded from there.
### Import Modules
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
```
### Load Data
```
df_lakers_cleaned = pd.read_csv( 'lakers_data_cleaned.csv', index_col = 0)
df_kings_cleaned = pd.read_csv( 'kings_data_cleaned.csv', index_col = 0)
df_dodgers_cleaned = pd.read_csv( 'dodgers_data_cleaned.csv', index_col = 0)
df_rams_cleaned = pd.read_csv( 'Rams_data_cleaned.csv', index_col = 0)
```
### Prep data for consumption
```
# NOTE: I Googled the team color.
list_prepped_data = \
[
('Lakers', 'purple', df_lakers_cleaned),
('Kings', 'grey', df_kings_cleaned),
('Dodgers', 'blue', df_dodgers_cleaned),
('Rams', 'orange', df_rams_cleaned)
]
```
### Plot Moving Average (Rolling Mean) Graph.
```
#---------------------- Functions (Start)--------------------------
def build_graph( plt = None,
df = None,
team_name = None,
team_color = None ):
"""
Description: Creates plot on same figure.
# TODO
#
# -Set title
# -Set x label
# -Set y lable
# -Set x-axis to be the whole data but only show 10 year intervals
# -Set y-axis for 0.0 to 1.0 but have dotted lines from 0.0, 0.25, 0.75, 1.0 BUT only use the highest that contain data.
# -Set thick lines.
# -Set dotted lines at y-axis intervals
# -Set annotations for names of team next to plot lines
# -Set annotations for win%
# -Remove plot box
# -Change the name of the figure to be generic for all teams and the save image.
"""
# Create graph
plot_current_graph = plt.plot( df['Season'],
df['Rolling_Mean'],
c=team_color,
label='Lakers')
# Set line thickness and style (like dotted)
# https://matplotlib.org/examples/pylab_examples/set_and_get.html
# plt.setp(plot_current_graph,
# linestyle='--')
plt.setp( plot_current_graph,
linewidth=4 )
# x the year after the last year recorded
x_pos = 2017
# y is the height of the last value in the dataframe.
y_pos = df['Rolling_Mean'].iloc[-1]
font_size = 10
plt.text(x_pos,
y_pos,
team_name,
color = team_color,
fontsize = font_size)
#---------------------- Functions (End)--------------------------
#--------------------------------------
# Setup static features of graph.
# (Start)
#--------------------------------------
# Create new figure
fig_teams = plt.figure(figsize = (16,8))
ax = fig_teams.add_subplot(111)
# Remove axis
#plt.axis('off')
ax.spines['left'].set_visible(False)
ax.spines['bottom'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
# Title
plt.title('Los Angeles Sports Teams Win %'
'\n(10 Year Moving Average)',
fontsize=20 )
# Labels for x and y axes
plt.xlabel( 'Season',
fontsize=15 )
plt.ylabel( '10 Year Moving Average Win %',
fontsize=15 )
# Set limit on x-axis
#ax.set_xlim([datetime.date(2016, 1, 1), datetime.date(2016, 12, 31)])
ax.set_ylim(0.0, 0.85)
# https://stackoverflow.com/questions/24943991/matplotlib-change-grid-interval-and-specify-tick-labels
#
# Set x-axis to be the whole data but only show 10 year intervals
x_major_ticks = np.arange(1980, 2020, 10)
#x_minor_ticks = np.arange(1980, 2020, 1)
#
ax.set_xticks(x_major_ticks)
# ax.set_xticks(x_minor_ticks, minor=True)
#
# Set y-axis for 0.0 to 1.0 but have dotted lines from 0.0, 0.25, 0.75, 1.0 BUT only use the highest that contain data.
y_major_ticks = np.arange(0.0, 1.1, 0.25)
#
# Slice to exclude the first and last entry.
y_major_ticks = y_major_ticks[:-1]
ax.set_yticks(y_major_ticks)
# Draw horizontal lines
for num in y_major_ticks:
plt.axhline(y = num,
linestyle = '--',
color = 'grey',
alpha = 0.2 )
# Legend
plt.text(1980,
0.1,
#'Win % = Games Won\(Games Won + Games Lost)',
#r'$\frac{5 - \frac{1}{x}}{4}$',
r'Win % = $\frac{Games Won}{Games Won + Games Lost}$',
fontsize = 15,
bbox={'facecolor':'lightgrey', 'alpha':0.5, 'pad':5})
#--------------------------------------
# Setup static features of graph.
# (End)
#--------------------------------------
# Cycle through the data and graph.
for i in range( len(list_prepped_data) ):
# Current Data
team_name, team_color, df = list_prepped_data[i]
# Build the graph.
build_graph( # Pass the plot being worked on
plt = plt,
# Pass the dataframe being worked on.
df = df,
# The name of the team
team_name = team_name,
# The team color
team_color = team_color)
# Show the graph.
plt.show()
```
### Save graph to .png
```
fig_teams.savefig( 'Los_Angeles_Sports_Teams_Percent_Wins.png' )
```
| github_jupyter |
```
import os
import sys
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
from scipy.optimize import curve_fit, fminbound
from scipy import stats as st
from tableanalyser import discretize_df_columns, plotvarmen, plotcv2mean, plotoversigmacv2, getovergenes, plotoverpoints
from tacos_plot import scatterdense
working_dir = "/home/fvalle/phd/datasets/tcga/oversampling_10tissue/"
os.chdir(working_dir)
normalisation_str = "fpkm"
df = pd.read_csv(("mainTable_all.csv"), index_col=[0])
#df = df.to_sparse(fill_value=0.)
df.head()
df_files = pd.read_csv("files.dat", index_col=0)
df.info()
ngenes = len(df.index)
nfiles = len(df.columns)
print("genes:%d\trealizations:%d"%(ngenes,nfiles))
```
## Means sigmas
```
df_mv = pd.read_csv("meanVariances.csv", index_col = [0])
#type_of_gene='protein-coding'
#df_g = pd.read_csv('/Users/filippo/Developer/tesi/genes.txt', header=0, index_col=0)
#df_mv=df_mv[df_mv.index.isin(df_g[df_g['type_of_gene']=='protein-coding'].index)]
#df_mv = df_mv.loc[df_mv['type_of_gene']==type_of_gene]
if not df_mv.columns.isin(['occurrence']).any():
df_mv_occ=pd.read_csv("O.dat", header=None)
df_mv.insert(3, 'occurrence', df_mv_occ.values)
df_mv.dropna(axis=0,how='any',inplace=True)
df_mv.head()
#df_mv.round(2).to_csv("meanVariances.csv",index=True,header=True)
#df_mv.fillna(value=0.,inplace=True)
means = df_mv['mean'].values
variances = df_mv['variance'].values
occurrences = np.array(df_mv['occurrence'].values, dtype=float)
abundances = pd.read_csv("A.dat", header=None).values
len(df_mv)
```
### plot
#### **var** versus **mean**
```
fig=plt.figure(figsize=(15,8))
ax=fig.subplots()
plotvarmen(means, variances, ax=ax, normalisation_str=normalisation_str)
fig.savefig("varmean_loglog.png")
fig=plt.figure(figsize=(15,8))
ax=fig.subplots()
plotcv2mean(means, variances, ax=ax, normalisation_str=normalisation_str)
fig.savefig("cvmean_loglog.png")
import scanpy as sc
adata = sc.AnnData(df.transpose(),obs=df_files.reindex(index=df.transpose().index), var=df_mv)
sc.pp.log1p(adata)
sc.pp.highly_variable_genes(adata, n_top_genes=1000, n_bins=100)
x = adata.var['mean'].values
y = adata.var['variance'].values
cv2 = [yi/xi/xi if xi!=0 else 0 for xi, yi in zip(x,y)]
y=np.array(cv2)
HVx = adata.var[adata.var['highly_variable']==True]['mean'].values
HVy = adata.var[adata.var['highly_variable']==True]['variance'].values
HVcv2 = [yi/xi/xi if xi!=0 else 0 for xi, yi in zip(HVx,HVy)]
HVy=np.array(HVcv2)
fig=plt.figure(figsize=(15,8))
plt.scatter(x,y)
plt.scatter(HVx,HVy)
plt.ylim((y[y!=0].min()*0.95,y.max()*1.5))
plt.xlim((x[x!=0].min()*0.95,x.max()*1.5))
plt.xscale('log')
plt.yscale('log')
plt.show()
x = adata.var['means'].values
y = adata.var['dispersions'].values
HVx = adata.var[adata.var['highly_variable']==True]['means'].values
HVy = adata.var[adata.var['highly_variable']==True]['dispersions'].values
fig=plt.figure(figsize=(20,10))
plt.scatter(x,y, label='genes')
plt.scatter(HVx,HVy, label='highly variable genes')
plt.ylim((np.nanmin(y)-0.1,np.nanmax(y)))
plt.xlim((-0.1,x.max()))
plt.xlabel("Log - Mean", fontsize=18)
plt.ylabel("Log - Dispersion", fontsize=18)
plt.legend(fontsize=20)
plt.show()
fig.savefig("DispersionMean.pdf")
```
# overexpressed
```
how_many_sigmas = 1
distance = 10
fig=plt.figure(figsize=(15,8))
ax=fig.subplots()
plotoversigmacv2(means, variances, ax=ax, normalisation_str=normalisation_str, how_many_sigmas=how_many_sigmas)
fig.savefig("cvmean_loglog_%dsigma.png"%how_many_sigmas)
def get_mean_cv2(mean, cv2, knee=1., distance=distance):
if mean < knee:
return(mean, cv2, -1, -1, cv2 > distance+1./mean)
else:
return(mean, cv2, -1, -1, cv2 > 1e-1+distance)
def get_mean_cv2(mean, cv2, bin_means=[], bin_sigmas=[],bin_edges=[],how_many_sigmas=3):
bin_i = 0
for i in range(len(bin_edges[:-1])):
if mean<=bin_edges[i+1] and mean > bin_edges[i]:
bin_i = i
break
return(mean, cv2, bin_means[bin_i], bin_sigmas[bin_i], cv2>(bin_means[bin_i]+how_many_sigmas*bin_sigmas[bin_i]))
over, over_plot = getovergenes(df_mv,get_mean_cv2, how_many_sigmas=how_many_sigmas, method='sigma')
fig=plt.figure(figsize=(15,8))
ax = fig.subplots()
plotoverpoints(means, variances, over_plot, ax=ax, how_many_sigmas=how_many_sigmas)
fig.savefig("cvmean_loglog_oversigma.png")
#discretize_df_columns(df.loc[over, df.columns[np.unique(np.random.randint(len(df.columns), size=2000))]]).to_csv("main_Table_Altmann.csv",index=True, header=True)
df[df.index.isin(over)].dropna().to_csv("mainTable_over.csv",index=True, header=True)
```
### mean versus occurrence
```
fig=plt.figure(figsize=(8,5))
plt.scatter(occurrences*nfiles, means, c='b', alpha=0.8, label='genes')
if 'counts' in normalisation_str:
plt.plot(np.linspace(1,nfiles), np.linspace(1,nfiles)/(nfiles), lw=4, label='bound', c='cyan', ls='--')
bin_means, bin_edges, _ = st.binned_statistic(occurrences*nfiles, means, statistic='mean', bins=np.logspace(-3,6))
x = (bin_edges[1:]+bin_edges[:-1])/2
plt.scatter(x,bin_means, marker='x', c='r', label='binned average')
plt.ylabel("$<%s>$"%normalisation_str, fontsize=16)
#plt.xlabel("$\Sigma_j\Theta(FPKM-0.1)\Theta(1e5-FPKM)$", fontsize=16)
plt.xlabel("$\Sigma_j\Theta(%s)$"%normalisation_str, fontsize=16)
plt.xscale('log')
plt.yscale('log')
plt.ylim(means[means.nonzero()].min()/5,np.power(10,np.log10(means.max())+1))
plt.xlim(5e-1,nfiles+800)
plt.legend(fontsize=20)
plt.show()
fig.savefig("meanDiff_loglog.png")
fig=plt.figure(figsize=(8,5))
plt.scatter(means, occurrences*nfiles, c='b', alpha=0.6, label='data')
bin_means, bin_edges, _ = st.binned_statistic(means, occurrences*nfiles, statistic='mean', bins=np.logspace(-3,6))
x = (bin_edges[1:]+bin_edges[:-1])/2
plt.scatter(x,bin_means, marker='x', c='r', label='binned average')
plt.xlabel("$<%s>$"%normalisation_str, fontsize=16)
#plt.xlabel("$\Sigma_j\Theta(FPKM-0.1)\Theta(1e5-FPKM)$", fontsize=16)
plt.ylabel("$\Sigma_j\Theta(%s)$"%normalisation_str, fontsize=16)
plt.xscale('log')
plt.yscale('log')
plt.xlim(means[means.nonzero()].min()/5,np.power(10,np.log10(means.max())+1))
plt.ylim(5e-1,nfiles+800)
plt.legend(fontsize=20)
plt.show()
fig.savefig("diffMean_loglog.png")
```
## Abundance vs occurrence
```
fig=plt.figure()
plt.scatter(occurrences, abundances.T[0])
plt.xscale('log')
plt.yscale('log')
plt.xlim((1e-5,1))
plt.ylim((np.min(abundances.T[0][abundances.T[0].nonzero()]),np.max(abundances.T[0])))
plt.show()
plt.scatter(np.divide(abundances,occurrences), means)
```
### Distributions
```
fig = plt.figure()
data = means
mu = np.median(data)
s = np.std(data)
log_bins = np.logspace(-5,7)
plt.hist(data, density = False, histtype='step', bins=log_bins)
plt.title("means", fontsize=16)
plt.xlabel("$<%s>$"%normalisation_str, fontsize=16)
plt.ylabel("#", fontsize=16)
plt.yscale('log')
plt.xscale('log')
plt.xlim(5e-5,5e5)
plt.show()
fig.savefig("mean_distr.pdf")
fig = plt.figure(figsize=(7,7))
data = -np.log10(means[means.nonzero()])
minimum = data.min()
data -= minimum
mu = np.median(data)
s = np.std(data)
fit_params = st.lognorm.fit(data)
n, c, _ = plt.hist(data, density = True, histtype='step')
plt.plot(np.linspace(-1,10),st.lognorm.pdf(np.linspace(-1,10),*fit_params), label='lognormal (s:%.2f,loc:%.2f,scale:%.2f)'%(fit_params))
plt.title("means", fontsize=16)
plt.xlabel("$-log10(<%s>)+%.2f$"%(normalisation_str,-minimum), fontsize=16)
plt.ylabel("pdf", fontsize=16)
plt.yscale('log')
plt.ylim(1e-4,1)
plt.xlim(-1,10)
plt.legend(fontsize=16)
plt.show()
fig.savefig("mean_distr_scaled.pdf")
bins = 80
bins=np.logspace(np.log10(5e-1),np.log10(variances.max()))
fig = plt.figure()
n, c, _ = plt.hist(variances, density = False, histtype='step', bins=bins)
plt.title("vars", fontsize=16)
plt.xlabel("$<\sigma_{%s}^2>$"%normalisation_str, fontsize=16)
plt.ylabel("#", fontsize=16)
plt.yscale('log')
plt.xscale('log')
plt.show()
fig.savefig("var_distr.pdf")
fig = plt.figure(figsize=(7,7))
data = -np.log10(variances[variances.nonzero()])
minimum = data.min()
data -= minimum
mu = np.median(data)
s = np.std(data)
fit_params = st.lognorm.fit(data)
n, c, _ = plt.hist(data, density = True, histtype='step')
plt.plot(np.linspace(-1,data.max()),st.lognorm.pdf(np.linspace(-1,data.max()),*fit_params), label='lognormal (s:%.2f,loc:%.2f,scale:%.2f)'%(fit_params))
plt.title("means", fontsize=16)
plt.xlabel("$-log10(<%s>)+%.2f$"%(normalisation_str,-minimum), fontsize=16)
plt.ylabel("pdf", fontsize=16)
plt.yscale('log')
plt.ylim(1e-4,1)
plt.xlim(data.min()-1,data.max()+1)
plt.legend(fontsize=16)
plt.show()
fig.savefig("var_distr_scaled.pdf")
```
# null
```
df_null = pd.read_csv(("%s/nullTable.csv"%working_dir), header=None, index_col=[0])
df_null.head()
```
## meanVariances
```
df_mv_null = pd.read_csv("meanVariances_null.csv", usecols=[1,2])
df_occ_null = pd.read_csv("O_null.dat", header=None)
df_mv_null.insert(2,'occurrence', np.array(df_occ_null.values,dtype=float))
#df_mv_null.to_csv("meanVariances_null.csv", index=False, header=True)
df_mv_null.head()
means_null = np.round(df_mv_null['mean'].values,1)
variances_null = np.round(df_mv_null['variance'].values,1)
occurrences_null = np.round(np.array(df_mv_null['occurrence'].values, dtype=float),1)
len(df_mv_null)
x = means
y = variances
x_lin = np.logspace(np.log10(x[x.nonzero()].min()),np.log10(x[x.nonzero()].max()), dtype=float,num=50)
high_means = df_mv[df_mv['occurrence']==1]['mean'].values
high_var = df_mv[df_mv['occurrence']==1]['variance'].values
fig=plt.figure(figsize=(12,7))
plt.scatter(x, y, label = 'genes', marker='o', alpha=0.8, linewidths=0.1)
#plt.scatter(high_means, high_var, label = '$O_i=1$', marker='o',c='cyan', alpha=0.4, lw=0.1)
log_bins_for_x = np.logspace(-5, np.log10(np.max(x)), num=50)
bin_means, bin_edges, binnumber = st.binned_statistic(x, y, statistic='mean', bins=log_bins_for_x)
bin_centres = (bin_edges[:-1]+bin_edges[1:])/2
plt.scatter((bin_edges[:-1]+bin_edges[1:])/2., bin_means, color='r', marker='x', lw=2, label='binned average')
plt.plot(x_lin[-40:],np.power(x_lin[-40:],2), 'g-', lw=3.5, label='$<%s>^2$ (Taylor)'%normalisation_str)
plt.plot(x_lin[:20],x_lin[:20], 'r-', lw=3.5, label='$<%s>$ (Poisson)'%normalisation_str)
#popt, pcov = curve_fit(lambda x,a,b : a*np.power(x,b), bin_centres, bin_means, bounds=([1,1],[35,5]))
#plt.plot(bin_centres, popt[0]*np.power(bin_centres, popt[1]), color='y', lw=3, label='fit')
#print(popt[0],popt[1])
#bin_sigmas, bin_sigmas_edges, binsigmanumber = stats.binned_statistic(x, y, statistic=np.std, bins=log_bins_for_x)
#plt.plot((bin_edges[:-1] + bin_edges[1:])/2, bin_means+bin_sigmas*3, lw=3, color='yellow', label='binned average + $3\sigma$')
plt.scatter(means_null, variances_null, label='sampling')
plt.xlabel("$<%s>$"%normalisation_str, fontsize=16)
plt.ylabel("$\sigma^2_{%s}$"%normalisation_str, fontsize=16)
plt.legend(fontsize=20)
plt.xscale('log')
plt.yscale('log')
plt.xlim(x[x.nonzero()].min()/100,np.power(10,np.log10(x.max())+1))
plt.ylim((y[y.nonzero()].min()/100,np.power(10,np.log10(y.max())+1)))
plt.show()
fig.savefig("varmean_3sigma.png")
cv2 = np.array([variances[i]/(np.power(mean,2)) for i,mean in enumerate(means) if mean>0])
cv2_null = [variances_null[i]/(np.power(mean,2)) for i,mean in enumerate(means_null) if mean>0]
fig=plt.figure(figsize=(15,8))
plt.scatter(means[means.nonzero()], cv2, c='b', label ='genes')
plt.scatter(means_null[means_null.nonzero()], cv2_null, lw=3, c='orange', label='sampling')
plt.plot(x_lin[:30],1./x_lin[:30], 'r-', lw=1.5, label='Poisson')
plt.plot(x_lin[20:],[1e-1 for _ in x_lin[20:]], 'g-', lw=1.5, label='Taylor')
#plt.plot(x_lin,[nfiles-1 for _ in x_lin], color='cyan', lw=3, ls='--', label='bound')
bin_means, bin_edges, binnumber = st.binned_statistic(means[means.nonzero()], cv2, statistic='mean', bins=log_bins_for_x)
bin_centres = (bin_edges[:-1]+bin_edges[1:])/2
plt.scatter(np.array((bin_edges[:-1]+bin_edges[1:])/2.)[:-6], bin_means[:-6], color='r', marker='x', lw=2, label='binned average')
plt.xlabel("$<%s>$"%normalisation_str, fontsize=20)
plt.ylabel("$cv^2$", fontsize=20)
plt.xscale('log')
plt.yscale('log')
plt.xlim(means[means.nonzero()].min()/5,np.power(10,np.log10(means.max())+1))
plt.ylim((cv2[cv2.nonzero()].min()/10,np.power(10,np.log10(cv2.max())+1)))
plt.legend(fontsize=20)
plt.show()
fig.savefig("cvmean_loglog_sampling.png")
x = means
y = variances
# INIT FIGURE #################################################################
fig = plt.figure(figsize=(12, 6))
ax = fig.subplots()
# AX #########################################################################
xmin = np.log10(1e-3)
xmax = np.log10(x.max())
ymin = np.log10(1e-6)
ymax = np.log10(y.max())
nbins=80
xbins = np.logspace(xmin, xmax, nbins) # <- make a range from 10**xmin to 10**xmax
ybins = np.logspace(ymin, ymax, nbins) # <- make a range from 10**ymin to 10**ymax
counts, _, _, _ = ax.hist2d(x, y, bins=(xbins, ybins));
pcm = ax.pcolormesh(xbins, ybins, counts.T)
plt.colorbar(pcm)
#fig.colorbar(pcm, ax=ax) # this works too
ax.set_xscale("log") # <- Activate log scale on X axis
ax.set_yscale("log") # <- Activate log scale on Y axis
ax.set_xlim(xmin=xbins[0])
ax.set_xlim(xmax=xbins[-1])
ax.set_ylim(ymin=ybins[0])
ax.set_ylim(ymax=ybins[-1])
ax.set_title("")
ax.set_xlabel("$<%s>$"%normalisation_str, fontsize=16)
ax.set_ylabel("$\sigma^2_{%s}$"%normalisation_str, fontsize=16)
# SHOW AND SAVE FILE ##########################################################
plt.tight_layout()
plt.show()
fig.savefig("varmean_density.png")
fig=plt.figure(figsize=(8,5))
plt.scatter(occurrences*nfiles, means, c='b', alpha=0.6, label='genes')
log_bins_for_x = np.logspace(-5, np.log10(np.max(x)), num=50)
bin_means, bin_edges, binnumber = st.binned_statistic(occurrences*nfiles, means, statistic='mean', bins=log_bins_for_x)
bin_centres = (bin_edges[:-1]+bin_edges[1:])/2
plt.hlines(bin_means, bin_edges[:-1], bin_edges[1:], colors='r', lw=5, label='binned average')
plt.scatter(occurrences_null, means_null, c='orange', alpha=0.6, label='sampling')
plt.plot(np.linspace(1,nfiles), np.linspace(1,nfiles)/(nfiles), lw=3, label='$\\frac{O_i}{Nsamples}$', c='cyan', ls='--')
plt.ylabel("$<%s>$"%normalisation_str, fontsize=16)
#plt.xlabel("$\Sigma_j\Theta(FPKM-0.1)\Theta(1e5-FPKM)$", fontsize=16)
plt.xlabel("$\Sigma_j\Theta(%s)$"%normalisation_str, fontsize=16)
plt.xscale('log')
plt.yscale('log')
plt.ylim(5e-5,np.power(10,np.log10(means.max())+1))
plt.xlim(5e-1,nfiles+800)
plt.legend(fontsize=16, loc='upper left')
plt.show()
fig.savefig("meanDiff_binned_sampling.png")
x = occurrences
y = means
# INIT FIGURE #################################################################
fig = plt.figure(figsize=(12, 6))
ax = fig.subplots()
# AX #########################################################################
xmin = np.log10(9e-1)
xmax = np.log10(x.max())
ymin = np.log10(5e-4)
ymax = np.log10(y.max())
nbins=80
xbins = np.logspace(xmin, xmax, nbins) # <- make a range from 10**xmin to 10**xmax
ybins = np.logspace(ymin, ymax, nbins) # <- make a range from 10**ymin to 10**ymax
counts, _, _, _ = ax.hist2d(x, y, bins=(xbins, ybins));
pcm = ax.pcolormesh(xbins, ybins, counts.T)
plt.colorbar(pcm)
#fig.colorbar(pcm, ax=ax) # this works too
ax.set_xscale("log") # <- Activate log scale on X axis
ax.set_yscale("log") # <- Activate log scale on Y axis
ax.set_xlim(xmin=xbins[0])
ax.set_xlim(xmax=xbins[-1])
ax.set_ylim(ymin=ybins[0])
ax.set_ylim(ymax=ybins[-1])
ax.set_title("")
ax.set_ylabel("$<%s>$"%normalisation_str, fontsize=16)
ax.set_xlabel("$\Sigma_j\Theta(%s)$"%normalisation_str, fontsize=16)
# SHOW AND SAVE FILE ##########################################################
plt.tight_layout()
plt.show()
fig.savefig("meanDiff_density.png")
```
## Length
```
q_many = pd.read_csv("genes.txt", index_col=[0], header=[0])
q_many = q_many[q_many['type_of_gene']=='protein-coding']
lengths = q_many['exons']
df_mv.insert(3,'length',lengths)
df_mv.head()
from scipy.stats import binned_statistic
fig=plt.figure(figsize=(15,7))
lengths = df_mv['length'].values
means = df_mv['mean'].values
bin_means, bin_edges, _ = binned_statistic(lengths, means, statistic='mean', bins=np.logspace(1,7))
plt.scatter(lengths,means)
plt.scatter((bin_edges[1:]+bin_edges[:-1])/2., bin_means, marker='x')
plt.yscale('log')
plt.xscale('log')
plt.xlabel('length (bp)', fontsize=16)
plt.ylabel('mean (counts)', fontsize=16)
plt.xlim((np.nanmin(lengths)/10,np.nanmax(lengths)*10))
plt.ylim((np.nanmin(means[means.nonzero()])/10,np.nanmax(means)*10))
plt.show()
fig.savefig("meanLength.pdf")
df_mv_counts=pd.read_csv("results/counts/meanVariances.csv", index_col=[0]).dropna(axis=0, how='any')
df_mv_counts.head()
df_mv_fpkm=pd.read_csv("results/proteincoding/meanVariances.csv", index_col=[0]).dropna(axis=0, how='any')
df_mv_fpkm.head()
df_mv_rpk=pd.read_csv("results/rpk/meanVariances.csv", index_col=[0]).dropna(axis=0, how='any')
df_mv_rpk.head()
diffnormmeans = []
for g in df_mv.index:
try:
diffnormmeans.append([g,df_mv.at[g,'length'],df_mv_counts.at[g,'mean'],df_mv_fpkm.at[g,'mean'], df_mv_rpk.at[g,'mean']])
except:
print("skipping %s"%g)
fig=plt.figure(figsize=(24,8))
ax=fig.subplots(1,3)
dataset = np.array(diffnormmeans).T
lengths = np.array(dataset[1],dtype=float)
means = np.array(dataset[2],dtype=float)
bin_means, bin_edges, _ = binned_statistic(lengths, means, statistic='mean', bins=np.logspace(1,7))
ax[0].scatter(lengths,means, label='genes')
ax[0].scatter((bin_edges[1:]+bin_edges[:-1])/2., bin_means, marker='x', label='binned average')
ax[0].set_yscale('log')
ax[0].set_xscale('log')
ax[0].set_xlabel('lenght (bp)', fontsize=16)
ax[0].set_ylabel('mean (counts)', fontsize=16)
ax[0].set_xlim((np.nanmin(lengths)/10,np.nanmax(lengths)*10))
ax[0].set_ylim((means[means.nonzero()].min()/10,means.max()*10))
means = np.array(dataset[3],dtype=float)
bin_means, bin_edges, _ = binned_statistic(lengths, means, statistic='mean', bins=np.logspace(1,7))
ax[1].scatter(lengths,means, label='genes')
ax[1].scatter((bin_edges[1:]+bin_edges[:-1])/2., bin_means, marker='x',label='binned average')
ax[1].plot(lenghts,np.power(lenghts,-0.5)*1e3, label='$L[kilobp]^{-0.5}$', lw=2, color='red')
ax[1].set_yscale('log')
ax[1].set_xscale('log')
ax[1].set_xlabel('lenght (bp)', fontsize=16)
ax[1].set_ylabel('mean (FPKM)', fontsize=16)
ax[1].set_xlim((np.nanmin(lengths)/10,np.nanmax(lengths)*10))
ax[1].set_ylim((means[means.nonzero()].min()/10,means.max()*10))
ax[1].legend(fontsize=16)
means = np.array(dataset[4],dtype=float)
bin_means, bin_edges, _ = binned_statistic(lengths, means, statistic='mean', bins=np.logspace(1,7))
ax[2].scatter(lengths,means, label='genes')
ax[2].plot(lengths,np.power(lengths,-1)*1e6, label='$L[kilobp]^{-1}$', lw=2, color='red')
ax[2].scatter((bin_edges[1:]+bin_edges[:-1])/2., bin_means, marker='x',label='binned average')
ax[2].set_yscale('log')
ax[2].set_xscale('log')
ax[2].set_xlabel('length (bp)', fontsize=16)
ax[2].set_ylabel('mean (RPK)', fontsize=16)
ax[2].set_xlim((np.nanmin(lengths)/10,np.nanmax(lengths)*10))
ax[2].set_ylim((means[means.nonzero()].min()/10,means.max()*10))
ax[2].legend(fontsize=16)
plt.show()
fig.savefig("meanLenght_fpkm_rpk.pdf")
scatterdense(np.array(dataset[2],dtype=float), np.array(dataset[4],dtype=float))
plt.plot(np.array(dataset[2],dtype=float),1e-3*np.power(np.array(dataset[2],dtype=float),1), c='r')
plt.xscale('log')
plt.yscale('log')
plt.xlim(np.nanmin(np.array(dataset[2],dtype=float)), np.nanmax(np.array(dataset[2],dtype=float)))
plt.ylim(np.nanmin(np.array(dataset[4],dtype=float)), np.nanmax(np.array(dataset[4],dtype=float)))
plt.show()
```
# P_i
```
#query_g = df_mv[df_mv['mean']>1e-4].index.values
query_g = df_mv.index.values
len(query_g)
N = df.loc[query_g,:].sum(axis=0)
df_null.index=df.index
N_null = df_null.loc[query_g,:].sum(axis=0)
df_p = df.loc[query_g,:].div(N.values, axis=1)
df_p_null = df_null.loc[query_g,:].div(N_null.values, axis=1)
means = df_p.apply(np.average, axis=1).values
variances = df_p.apply(np.var, axis=1).values
o = df_p.apply(lambda x: float(len(np.nonzero(x)[0]))/len(x),axis=1).values
means_null = df_p_null.apply(np.average, axis=1).values
variances_null = df_p_null.apply(np.var, axis=1).values
o_null = df_p_null.apply(lambda x: float(len(np.nonzero(x)[0]))/len(x),axis=1).values
x = means
y = variances
log_bins_for_x = np.logspace(np.log10(x[x.nonzero()].min()),np.log10(x[x.nonzero()].max()), dtype=float,num=30)
x_null = means_null
y_null = variances_null
log_bins_for_x_null = np.logspace(np.log10(x_null[x_null.nonzero()].min()),np.log10(x_null[x_null.nonzero()].max()), dtype=float,num=30)
x_high = df_p.loc[df_mv[df_mv['occurrence']==1].index,:].mean(axis=1).values
y_high = df_p.loc[df_mv[df_mv['occurrence']==1].index,:].var(axis=1).values
fig=plt.figure(figsize=(12,7))
plt.scatter(x, y, label = 'genes', marker='o', alpha=0.8, linewidths=0.1)
plt.scatter(x_high, y_high, label = '$O_i=1$', marker='o', alpha=0.4, color='cyan', linewidths=0.1)
#plt.scatter(x_null, y_null, label = 'sampling', marker='o', alpha=0.8, linewidths=0.1)
bin_means, bin_edges, binnumber = stats.binned_statistic(x, y, statistic='mean', bins=log_bins_for_x)
bin_centres = (bin_edges[:-1]+bin_edges[1:])/2
plt.hlines(bin_means, bin_edges[:-1], bin_edges[1:], colors='r', lw=5, label='binned average')
plt.plot(log_bins_for_x[-40:],np.power(log_bins_for_x[-40:],2), 'g-', lw=3.5, label='$<p_i>^2$ (Taylor)')
plt.plot(log_bins_for_x[:],log_bins_for_x[:], 'r-', lw=3.5, label='$<p_i>$ (Poisson)')
plt.plot(log_bins_for_x[:],log_bins_for_x[:]*1e-8, ls='-', color='darkred', lw=3.5, label='$10^{-8}*<p_i>$ (Poisson)')
plt.xlabel("$<p_i>$", fontsize=16)
plt.ylabel("$\sigma^2_{p_i}$", fontsize=16)
plt.legend(fontsize=18)
plt.xscale('log')
plt.yscale('log')
plt.xlim(x[x.nonzero()].min()/100,np.power(10,np.log10(x.max())+1))
plt.ylim((y[y.nonzero()].min()/100,np.power(10,np.log10(y.max())+1)))
plt.show()
fig.savefig("pi_varmean_3sigma.png")
fig = plt.figure()
data = means
mu = np.median(data)
s = np.std(data)
log_bins = np.logspace(-12,-2)
plt.hist(data, density = False, histtype='step', bins=log_bins)
plt.title("means", fontsize=16)
plt.xlabel("$<pi>$", fontsize=16)
plt.ylabel("#", fontsize=16)
plt.yscale('log')
plt.xscale('log')
plt.xlim(data[data.nonzero()].min(),data.max())
plt.show()
fig.savefig("pi_mean_distr.pdf")
```
## over mean
```
from matplotlib.patches import Rectangle
o_min = 3e1
o_max = nfiles
m_min = 5e4
m_max = 1e6
fig=plt.figure(figsize=(8,5))
plt.scatter(occurrences, means, c='b', alpha=0.6, label='data')
plt.plot(np.linspace(1,nfiles), np.linspace(1,nfiles)/(nfiles*10), lw=2, label='', c='r')
width = o_max - o_min
height = m_max-m_min
plt.gca().add_patch(Rectangle((o_min,m_min), width=width, height=height, fill=False))
plt.ylabel("$<%s>$"%normalisation_str, fontsize=16)
#plt.xlabel("$\Sigma_j\Theta(FPKM-0.1)\Theta(1e5-FPKM)$", fontsize=16)
plt.xlabel("$\Sigma_j\Theta(%s)$"%normalisation_str, fontsize=16)
plt.xscale('log')
plt.yscale('log')
plt.ylim(5e-5,np.power(10,np.log10(means.max())+1))
plt.xlim(5e-1,nfiles+800)
plt.show()
plt.savefig("highmean.png")
up = []
for g in df_mv.index:
subdf = df_mv.loc[g,:]
mean = subdf['mean']
occ = subdf['occurrence']
if mean>m_min and mean < m_max and occ*nfiles > o_min and occ*nfiles< o_max:
up.append(g)
len(up)
for g in up:
print(g)
dat = df.loc['ENSG00000042832'].values
np.var(dat)/(np.average(dat)**2)
df_mv[df_mv['mean']>0].sort_values(by='variance', axis=0, ascending=False)
```
## set by occurrence
```
with open("o1.txt",'a') as f:
for g in df_mv[df_mv['occurrence']>4990].index:
f.write("%s\n"%g)
```
## data size Heaps check
```
col = df.loc[:,df.keys()[1]].values
np.sum(col)
len(col[col.nonzero()])
x = []
y = []
for i in range(1, nfiles):
col = df.loc[:,df.keys()[i]].values
x.append(np.sum(col))
y.append(len(col.nonzero()[0]))
plt.scatter(x,y)
i=794
x=[]
y=[]
col = df.loc[:,df.keys()[i]].values
x.append(np.sum(col))
y.append(len(col.nonzero()[0]))
x
y
col[8142:8150]
```
## Imagesave
```
bits = np.array([df.loc[g,:].values for g in df.index])
from PIL import Image
img = Image.fromarray(bits, mode='1')
img.save("mat.bmp")
```
| github_jupyter |
# Serving Deep Learning Models
```
%matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
df = pd.read_csv('../data/wifi_location.csv')
df.head()
df['location'].value_counts()
df.plot(figsize=(12, 8))
plt.axvline(500)
plt.axvline(1000)
plt.axvline(1500)
plt.title('Indoor location dataset')
plt.xlabel('Sample number')
plt.ylabel('Wifi strength (dB)');
import seaborn as sns
sns.pairplot(df, hue='location');
X = df.drop('location', axis=1).values
y = df['location'].values
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = \
train_test_split(X, y, test_size=0.25,
random_state=0)
from tensorflow.keras.models import Model
from tensorflow.keras.layers import Input, Dense, BatchNormalization
inputs = Input(shape=X_train.shape[1:])
x = BatchNormalization()(inputs)
x = Dense(50, activation='relu')(x)
x = Dense(30, activation='relu')(x)
x = Dense(10, activation='relu')(x)
predictions = Dense(4, activation='softmax')(x)
model = Model(inputs=inputs, outputs=predictions)
model.summary()
model.compile('adam',
'sparse_categorical_crossentropy',
metrics=['accuracy'])
h = model.fit(X_train, y_train,
batch_size=128,
epochs=40,
verbose=0,
validation_data=(X_test, y_test))
pd.DataFrame(h.history).plot()
plt.ylim(0, 1);
import os # Miscellaneous operating system interfaces
import json # JSON encoder and decoder
import shutil # High-level file operations
base_path = '/tmp/ztdl_models/wifi'
sub_path = 'flask'
version = 1
from os.path import join
export_path = join(base_path, sub_path, str(version))
export_path
shutil.rmtree(export_path, ignore_errors=True) # delete path, if exists
os.makedirs(export_path) # create path
json.loads(model.to_json())
with open(join(export_path, 'model.json'), 'w') as fout:
fout.write(model.to_json())
model.save_weights(join(export_path, 'weights.h5'))
os.listdir(export_path, )
from tensorflow.keras.models import model_from_json
with open(join(export_path, 'model.json')) as fin:
loaded_model = model_from_json(fin.read())
probas = loaded_model.predict(X_test)
probas
preds = np.argmax(probas, axis=1)
preds
from sklearn.metrics import accuracy_score
accuracy_score(y_test, preds)
loaded_model.load_weights(join(export_path, 'weights.h5'))
probas = loaded_model.predict(X_test) # class probabilities
preds = np.argmax(probas, axis=1) # class prediction
accuracy_score(y_test, preds) # accuracy score
```
## A simple deployment with Flask
```
!cat ./model_serving/flask_serve_model.py
```
## Exercise 1
Open a terminal and run the script with command:
```
CUDA_VISIBLE_DEVICES="" python model_serving/flask_serve_model.py
```
Then come back here and continue
```
import requests
api_url = "http://localhost:5000/"
data = X_test[:5].tolist()
data
payload = {'data': data}
headers = {'content-type': 'application/json'}
response = requests.post(api_url,
data=json.dumps(payload),
headers=headers)
response
response.json()
y_test[:5]
```
## Deployment with Tensorflow Serving
As the [documentation](https://www.tensorflow.org/serving/) says, TensorFlow Serving is a flexible, high-performance serving system for Machine Learning models, designed for production environments. TensorFlow Serving makes it easy to deploy new algorithms and experiments, while keeping the same server architecture and APIs. TensorFlow Serving provides out-of-the-box integration with TensorFlow models, but can be easily extended to serve other types of models and data.
Tensorflow Serving can accommodate both small and large deployments, and it is built for production. It is not as simple as Flask, and here we will barely scratch the surface of what it's possible with it. If you are serious about using it, we strongly recommend you take a look at the [Architecture overview](https://www.tensorflow.org/serving/architecture_overview) where many concepts like Servables, Managers and Sources are explained.
In this part of the book, we will just show you how to export a model for serving and how to ping a Tensorflow serving server. We will leave the full installation of Tensorflow serving for the end of the chapter. Installation is strongly dependent on the system you are using and is [well documented](https://www.tensorflow.org/serving/).
```
import tensorflow as tf
base_path = '/tmp/ztdl_models/wifi'
sub_path = 'tfserving'
version = 1
export_path = join(base_path, sub_path, str(version))
export_path
shutil.rmtree(export_path, ignore_errors=True)
tf.saved_model.save(model, export_path)
os.listdir(export_path)
!saved_model_cli show --dir {export_path} --all
```
## Exercise 2
Let's pull the `tensorflow/serving` docker container:
```
docker pull tensorflow/serving:latest
```
And let' run the docker container with the following command:
```
docker run \
-v /tmp/ztdl_models/wifi/tfserving/:/models/wifi \
-e MODEL_NAME=wifi \
-e MODEL_PATH=/models/wifi \
-p 8500:8500 \
-p 8501:8501 \
-t tensorflow/serving
```
Where:
- `-v`: This bind mounts a volume; it tells Docker to map the internal path `/models/wifi` to the `/tmp/ztdl_models/wifi/tfserving/` in our host computer.
- `-e`: Sets environment variables, in this case, we set the `MODEL_NAME` and `MODEL_PATH` variables
- `-p`: Publishes a container's port to the host. In this case, we are publishing port 8500 (default gRPC) and 8501 (default REST).
- `-t`: Allocate a pseudo-TTY
- `tensorflow/serving` is the name of the container we are running.
Since Tensorflow 1.8, Tensorflow serving comes with both a gRPC and REST endpoints by default, so we can test our running server by simply using curl. The correct command for this is:
```
curl -d '{"signature_name": "serving_default",
"instances": [[-62.0, -58.0, -59.0, -59.0, -67.0, -80.0, -77.0],
[-49.0, -53.0, -50.0, -48.0, -67.0, -78.0, -88.0],
[-52.0, -57.0, -49.0, -50.0, -66.0, -80.0, -80.0]]}' \
-H "Content-Type: application/json" \
-X POST http://localhost:8501/v1/models/wifi:predict
```
Go ahead and run that in a shell, you should receive an output that looks similar to the following:
```
{
"predictions": [[0.997524, 1.19462e-05, 0.00171472, 0.000749083],
[3.40611e-06, 0.00262853, 0.997005, 0.000363284],
[2.52653e-05, 0.00507444, 0.993813, 0.00108718]
]
}
```
Once you've tested the REST API, you can test the gRPC API with the following code:
```
from grpc import insecure_channel
channel = insecure_channel('localhost:8500')
channel
from tensorflow_serving.apis.prediction_service_pb2_grpc import PredictionServiceStub
stub = PredictionServiceStub(channel)
data
data_np = np.array(data)
data_pb = tf.compat.v1.make_tensor_proto(data_np,
dtype='float',
shape=data_np.shape)
data_pb
from tensorflow_serving.apis.predict_pb2 import PredictRequest
request = PredictRequest()
request.model_spec.name = 'wifi'
request.model_spec.signature_name = 'serving_default'
request.inputs['input_1'].CopyFrom(data_pb)
request
result_future = stub.Predict.future(request, 5.0)
result = result_future.result()
result
scores = tf.make_ndarray(result.outputs['dense_3'])
scores
prediction = np.argmax(scores, axis=1)
prediction
model.predict(np.array(data)).argmax(axis=1)
```
| github_jupyter |
### Load Libraries
```
import pandas as pd
import numpy as np
import os, sys, glob, json, pickle
import seaborn as sn
from sklearn.metrics import confusion_matrix
import matplotlib.pyplot as plt
from sklearn import metrics
from sklearn.metrics import accuracy_score, f1_score, recall_score, cohen_kappa_score
from sklearn.metrics import precision_score, matthews_corrcoef, roc_auc_score
import seaborn as sn
from sklearn.metrics import confusion_matrix
import matplotlib.pyplot as plt
from matplotlib import pylab
from matplotlib_venn import venn3, venn3_circles
```
### Prediction of validation and test sets using three best performing models:
### fingerprints, pharmacophore, and rdkit descriptors
```
def predict_validation_test(models):
temp = 'balanced_randomsplit7_70_15_15'
split = ['_va', '_te']
for s in split:
for m in models:
fp = results[m]
fingerprint = np.load('valid_test_features/'+\
fp+'-'+m+'-'+temp+s+'.npy')
rdkDes = np.load('valid_test_features/'+\
'rdkDes'+'-'+m+'-'+temp+s+'.npy')
pharma = np.load('valid_test_features/'+\
'tpatf'+'-'+m+'-'+temp+s+'.npy')
volsurf = np.load('valid_test_features/'+\
'volsurf'+'-'+m+'-'+temp+s+'.npy')
fingerprint_model = pickle.load(open('models_tuned_best/'\
+fp+'-'+m+'-'+temp+'.pkl', 'rb'))
rdkDes_model = pickle.load(open('models_tuned_best/'\
+'rdkDes'+'-'+m+'-'+temp+'.pkl', 'rb'))
pharma_model = pickle.load(open('models_tuned_best/'\
+'tpatf'+'-'+m+'-'+temp+'.pkl', 'rb'))
volsurf_model = pickle.load(open('models_tuned_best/'\
+'volsurf'+'-'+m+'-'+temp+'.pkl', 'rb'))
fp_list = [fingerprint, rdkDes, pharma, volsurf]
models_list = [fingerprint_model, rdkDes_model, pharma_model, volsurf_model]
df = pd.read_csv('combined_data/'+m+'/'+m+'-'+temp+\
s+'.csv')
fpnames = ['fingerprint', 'rdkDescriptors', 'pharmacophore', 'volsurf']
for finger, mod, fpname in zip(fp_list, models_list, fpnames):
X_true = finger[:,:-1]
y_true = finger[:,-1]
y_pred = mod.predict(X_true)
y_pred_label = y_pred.tolist()
y_prob = mod.predict_proba(X_true)[:,1]
df.insert(len(df.columns), fpname+'_pred_label', y_pred_label)
df.insert(len(df.columns), fpname+'_pred_prob', y_prob)
if not os.path.isdir('valid_test_csv_pred'+'/'+m):
os.mkdir('valid_test_csv_pred'+'/'+m)
df.to_csv('valid_test_csv_pred/'+m+'/'+m+s+'.csv', index=False)
```
### Make dictionary of best models and feature types among fingperint rdkit descriptors, and pharmacophore
```
def best_models_dictn():
results= {}
for f in glob.glob('reports_tuned_best/*.json'):
fp,_ = os.path.splitext(os.path.basename(f))
modelName = fp[:-42]
modelName2 = modelName.split('-')[1:]
modelName3 = "-".join(modelName2)
fpname = fp.split('-')[0]
if fpname == 'tpatf' or fpname == 'rdkDes' or fpname == "volsurf":
continue
else:
results[modelName3]=fpname
return results
results = best_models_dictn()
models=['3CL', 'ACE2', 'AlphaLISA', 'CoV1-PPE', 'CoV1-PPE_cs', 'CPE' \
,'cytotox', 'hCYTOX', 'MERS-PPE','MERS-PPE_cs', 'TruHit']
predict_validation_test(models)
def get_consensus(results):
for d in os.listdir(results):
for f in glob.glob(results+'/'+d+'/*.csv'):
consensus_label, consensus_proba, consensus_volsurf_label,\
consensus_volsurf_label_proba= [], [], [], []
df = pd.read_csv(f)
for i in range(len(df)):
if (df['fingerprint_pred_label'][i]+df['rdkDescriptors_pred_label'][i]+df['pharmacophore_pred_label'][i])>=2.0:
consensus_label.append(1.0)
else:
consensus_label.append(0.0)
if (df['fingerprint_pred_prob'][i]+df['rdkDescriptors_pred_prob'][i]+df['pharmacophore_pred_prob'][i])/3>0.5:
consensus_proba.append(1.0)
else:
consensus_proba.append(0.0)
if (df['fingerprint_pred_label'][i]+df['volsurf_pred_label'][i]+df['pharmacophore_pred_label'][i])>=2.0:
consensus_volsurf_label.append(1.0)
else:
consensus_volsurf_label.append(0.0)
if (df['fingerprint_pred_prob'][i]+df['volsurf_pred_prob'][i]+df['pharmacophore_pred_prob'][i])/3>0.5:
consensus_volsurf_label_proba.append(1.0)
else:
consensus_volsurf_label_proba.append(0.0)
df.insert(len(df.columns), 'Consensus_label', consensus_label)
df.insert(len(df.columns), 'Consensus_prob_label', consensus_proba)
df.insert(len(df.columns), 'Consensus_volsurf_label', consensus_volsurf_label)
df.insert(len(df.columns), 'Consensus_volsurf_prob_label', consensus_volsurf_label_proba)
if f.endswith('_va.csv'):
df.to_csv('valid_test_csv_pred_cons/valid/'+d+'_va.csv', index=False)
elif f.endswith('_te.csv'):
df.to_csv('valid_test_csv_pred_cons/test/'+d+'_te.csv', index=False)
get_consensus('valid_test_csv_pred')
```
### Evalution metrics
```
def evaluation_metrics(y_true, y_pred, y_prob):
Scores = dict()
roc_auc = roc_auc_score(y_true, y_prob)
acc = accuracy_score(y_true, y_pred)
f1 = f1_score(y_true, y_pred, average = 'binary')
precision = precision_score(y_true, y_pred, average= 'binary')
recall = recall_score(y_true, y_pred, average = 'binary')
kappa = cohen_kappa_score(y_true, y_pred)
mcc = matthews_corrcoef(y_true, y_pred)
tn, fp, fn, tp = confusion_matrix(y_true, y_pred).ravel()
SP = float(tn)/(tn+fp)
# Convert numpy.float64 to float using 'tolist()' and save the scores in dictonary
metrics_list = ['ACC', 'F1_Score', 'Recall', 'Precision', 'Specificity', \
'Cohen_Kappa', 'MCC', 'AUC', 'TP', 'TN', 'FP', 'FN']
scores_list = [acc.tolist(), f1.tolist(), recall.tolist(), precision.tolist(), SP.tolist(), \
kappa.tolist(), mcc, roc_auc.tolist(), tp.tolist(), tn.tolist(), fp.tolist(), fn.tolist()]
for metric, score in zip(metrics_list, scores_list):
Scores[metric] = score
return Scores
```
### Find consensus results
```
def get_consensus_scores(y_true, y_pred):
Scores = dict()
roc_auc = roc_auc_score(y_true, y_pred)
acc = accuracy_score(y_true, y_pred)
f1 = f1_score(y_true, y_pred, average = 'binary')
precision = precision_score(y_true, y_pred, average= 'binary')
recall = recall_score(y_true, y_pred, average = 'binary')
kappa = cohen_kappa_score(y_true, y_pred)
mcc = matthews_corrcoef(y_true, y_pred)
tn, fp, fn, tp = confusion_matrix(y_true, y_pred).ravel()
SP = float(tn)/(tn+fp)
metrics_list = ['ACC', 'F1_Score', 'Recall', 'Precision', 'Specificity', \
'Cohen_Kappa', 'MCC', 'AUC', 'TP', 'TN', 'FP', 'FN']
scores_list = [acc.tolist(), f1.tolist(), recall.tolist(), precision.tolist(), SP.tolist(), \
kappa.tolist(), mcc, roc_auc.tolist(), tp.tolist(), tn.tolist(), fp.tolist(), fn.tolist()]
for metric, score in zip(metrics_list, scores_list):
Scores[metric] = score
return Scores
def get_total_scores(models_dir):
total_dictn = dict()
for s in os.listdir(models_dir):
dictn_dir = dict()
for f in glob.glob(os.path.join(models_dir, s+'/*.csv')):
df = pd.read_csv(f)
name= os.path.splitext(os.path.basename(f))[0][:-3]
dictn_fp = dict()
for _type in ['fingerprint_pred_label', 'rdkDescriptors_pred_label', 'pharmacophore_pred_label',\
'volsurf_pred_label', 'Consensus_label', 'Consensus_prob_label', 'Consensus_volsurf_label', \
'Consensus_volsurf_prob_label']:
y_pred = df[_type].to_list()
y_true = df['Label'].to_list()
# y_prob is not used for consensus
if _type == 'Consensus_label' or _type=='Consensus_prob_label' or \
_type=='Consensus_volsurf_label' or _type=='Consensus_volsurf_prob_label':
scores = get_consensus_scores(y_true, y_pred)
else:
y_prob = df[_type[:-6]+'_prob'].to_list()
scores = evaluation_metrics(y_true, y_pred, y_prob)
dictn_fp[_type] = scores
dictn_dir[name] = dictn_fp
total_dictn[s] = dictn_dir
return total_dictn
results = get_total_scores('valid_test_csv_pred_cons/')
results
```
### Find the best model info:
```
# Comparision of Volsurf and RDkit Descriptors
total_models = list(results['valid'].keys())
best_f1 = []
best_model = []
model_types = ['rdkDescriptors_pred_label', 'volsurf_pred_label']
for m in total_models:
f1_m = 0
model_name = ''
for mt in model_types:
f1_score = results['valid'][m][mt]['F1_Score']
if f1_score > f1_m:
f1_m = f1_score
model_name = mt
else:
pass
best_f1.append(f1_m)
best_model.append(model_name)
print(total_models)
print(best_model)
print(best_f1)
# Comparision of Volsurf and RDkit Descriptors
total_models = list(results['valid'].keys())
f1 = []
models = []
for m in total_models:
RDK_f1 = round(results['valid'][m]['rdkDescriptors_pred_label']['F1_Score'], 3)
volsurf_f1 = round(results['valid'][m]['volsurf_pred_label']['F1_Score'], 3)
if RDK_f1 > volsurf_f1:
f1.append(m+'|'+'RDK->'+str(RDK_f1)+'|'+str(volsurf_f1)+'->VSF')
models.append(m+'|'+'RDK')
else:
f1.append(m+'|'+'VSF->'+str(volsurf_f1)+ '|'+ str(RDK_f1)+'->RDK')
models.append(m+'|'+'VSF')
# print('Models: \n', total_models)
print('F1 scores: \n', f1)
print('Best RDK or VSF: \n', models)
# Find Best models
total_models = list(results['valid'].keys())
best_f1 = []
best_model = []
model_types = list(results['valid']['cytotox'].keys())
for m in total_models:
f1_m = 0
model_name = ''
for mt in model_types:
f1_score = results['valid'][m][mt]['F1_Score']
if f1_score > f1_m:
f1_m = f1_score
model_name = mt
else:
pass
best_f1.append(round(f1_m, 3))
best_model.append(m+'|'+model_name)
print(best_model)
print(best_f1)
```
### Plots showing the evaluated metrics
```
for s in ['valid', 'test']:
models = list(results[s].keys())
for m in models:
valid_dictn = results[s][m]
df = pd.DataFrame(valid_dictn)
df = df.transpose()
df.insert(0, 'Models', list(valid_dictn.keys()))
# Select the useful metrics only
df = df.iloc[:,:-4]
label = ('fingerprint', 'rdkDescriptor', 'pharmacophore', 'volsurf', 'consensus-voting-rdkDes',\
'consensus-prob-rdkDes', 'consensus-voting-volsurf', "consensus-prob-volsurf" )
# number of xticks--> number of models
plt.xticks(np.arange(8), label, rotation=90) #, fontsize=6
plt.scatter(df['Models'].tolist(), df['F1_Score'].tolist(), c='b',s=20, label='F1')
plt.scatter(df['Models'].tolist(), df['Recall'].tolist(), c='r',s=18, label='Recall')
plt.scatter(df['Models'].tolist(), df['Precision'].tolist(), c='y',s=15, label='Precision')
plt.scatter(df['Models'].tolist(), df['ACC'].tolist(), c='g',s=13, label='Acc')
plt.scatter(df['Models'].tolist(), df['AUC'].tolist(), c='k',s=12, label='AUC')
plt.xlabel('Models')
plt.ylabel('Scores')
plt.legend(loc='center left', bbox_to_anchor=(0.983, 0.5), ncol=1)
plt.tight_layout()
if m == 'CoV1-PPE':
plt.title('CoV-PPE'+' '+s+' set'+ ' results', fontsize=6.7)
elif m == 'CoV1-PPE_cs':
plt.title('CoV-PPE_cs'+' '+s+' set'+ ' results', fontsize=6.7)
else:
plt.title(m+' '+s+' set'+ ' results', fontsize=6.7)
plt.savefig('plots/'+m+'-'+s+'.png',dpi=600)
plt.show()
plt.clf()
```
### External Set predictions for CPE and 3CL
```
def external_pred(ext_files, model):
files = glob.glob(ext_files)
for f in files:
filename = os.path.splitext(os.path.basename(f))[0]
if model == 'CPE' and filename == 'cpe_external_set_after_phys-chem-filters_stand':
fingerprint = np.load('external_set_features/'+'lfcfp4-' + filename+'.npy')
fingerprint_model = \
pickle.load(open('models_tuned_best/lfcfp4-CPE-balanced_randomsplit7_70_15_15.pkl', 'rb'))
elif model == '3CL' and filename == '3cl_external_set_stand':
fingerprint = np.load('external_set_features/'+ 'hashtt-'+ filename+'.npy')
fingerprint_model = \
pickle.load(open('models_tuned_best/hashtt-3CL-balanced_randomsplit7_70_15_15.pkl', 'rb'))
else:
continue
rdkDes = np.load('external_set_features/'+ 'rdkDes-'+ filename+'.npy')
rdkDes_model = \
pickle.load(open('models_tuned_best/rdkDes-'+ model +'-balanced_randomsplit7_70_15_15.pkl', 'rb'))
pharma = np.load('external_set_features/'+ 'tpatf-' + filename+'.npy')
pharma_model = \
pickle.load(open('models_tuned_best/tpatf-'+ model +'-balanced_randomsplit7_70_15_15.pkl', 'rb'))
fp_list = [fingerprint, rdkDes, pharma]
models_list = [fingerprint_model, rdkDes_model, pharma_model]
df_load = pd.read_csv(f)
fpnames = ['fingerprint', 'rdkDescriptor', 'pharmacophore']
for finger, mod, fpname in zip(fp_list, models_list, fpnames):
X_true = finger[:,:-1]
y_true = finger[:,-1]
y_pred = mod.predict(X_true)
y_pred_label = y_pred.tolist()
y_prob = mod.predict_proba(X_true)[:,1]
df_load.insert(len(df_load.columns), fpname, y_pred_label)
df_load.to_csv('ext_pred_cpe_3cl/'+filename+'.csv', index=False)
# External prediction
external_pred('cpe_3cl_ext_set/*.csv', 'CPE')
external_pred('cpe_3cl_ext_set/*.csv', '3CL')
```
### Venn diagram showing the distribution of predicted actives
```
def plot_venn_diagram(csvFile, n):
def get_venn_sections(sets):
num_combinations = 2 ** len(sets)
bit_flags = [2 ** n for n in range(len(sets))]
flags_zip_sets = [z for z in zip(bit_flags, sets)]
combo_sets = []
for bits in range(num_combinations - 1, 0, -1):
include_sets = [s for flag, s in flags_zip_sets if bits & flag]
exclude_sets = [s for flag, s in flags_zip_sets if not bits & flag]
combo = set.intersection(*include_sets)
combo = set.difference(combo, *exclude_sets)
tag = ''.join([str(int((bits & flag) > 0)) for flag in bit_flags])
combo_sets.append((tag, combo))
return combo_sets
df1 = pd.read_csv(csvFile)
pred_csv1_0_indices = [index for index, element in enumerate(df1['fingerprint'].to_list()) if element == 0]
pred_csv1_1_indices = [index for index, element in enumerate(df1['fingerprint'].to_list()) if element == 1]
pred_csv2_0_indices = [index for index, element in enumerate(df1['pharmacophore'].to_list()) if element == 0]
pred_csv2_1_indices = [index for index, element in enumerate(df1['pharmacophore'].to_list()) if element == 1]
pred_csv3_0_indices = [index for index, element in enumerate(df1['rdkDescriptor'].to_list()) if element == 0]
pred_csv3_1_indices = [index for index, element in enumerate(df1['rdkDescriptor'].to_list()) if element == 1]
get_common = get_venn_sections([set(pred_csv1_1_indices), set(pred_csv2_1_indices), set(pred_csv3_1_indices)])
venn3([set(pred_csv1_1_indices), set(pred_csv2_1_indices), set(pred_csv3_1_indices)], set_labels= \
['fingerprint', 'pharmacophore', 'rdkitDescriptor'])
# plt.title(n+' :summary of predicted actives of External Sets')
plt.savefig('plots/'+n+'-ext'+'.png', dpi=600)
plt.show()
# Venn diagram plot
_files = ['ext_pred_cpe_3cl/cpe_external_set_after_phys-chem-filters_stand.csv', 'ext_pred_cpe_3cl/3cl_external_set_stand.csv']
names = ['CPE', '3CL']
for _f, n in zip(_files, names) :
plot_venn_diagram(_f, n)
def external_pred_volsurf(ext_files, model):
files = glob.glob(ext_files)
for f in files:
filename = os.path.splitext(os.path.basename(f))[0]
if model == 'CPE' and filename == 'cpe_external_set_after_phys-chem-filters_stand':
fingerprint = np.load('external_set_features/'+'lfcfp4-' + filename+'.npy')
fingerprint_model = \
pickle.load(open('models_tuned_best/lfcfp4-CPE-balanced_randomsplit7_70_15_15.pkl', 'rb'))
else:
continue
volsurf = np.load('external_set_features/'+ 'volsurf-'+ filename+'.npy')
volsurf_model = \
pickle.load(open('models_tuned_best/volsurf-'+ model +'-balanced_randomsplit7_70_15_15.pkl', 'rb'))
pharma = np.load('external_set_features/'+ 'tpatf-' + filename+'.npy')
pharma_model = \
pickle.load(open('models_tuned_best/tpatf-'+ model +'-balanced_randomsplit7_70_15_15.pkl', 'rb'))
fp_list = [fingerprint, volsurf, pharma]
models_list = [fingerprint_model, volsurf_model, pharma_model]
df_load = pd.read_csv(f)
fpnames = ['fingerprint', 'volsurf', 'pharmacophore']
for finger, mod, fpname in zip(fp_list, models_list, fpnames):
X_true = finger[:,:-1]
y_true = finger[:,-1]
y_pred = mod.predict(X_true)
y_pred_label = y_pred.tolist()
y_prob = mod.predict_proba(X_true)[:,1]
df_load.insert(len(df_load.columns), fpname, y_pred_label)
df_load.to_csv('ext_pred_cpe_3cl_volsurf/'+filename+'.csv', index=False)
external_pred_volsurf('cpe_3cl_ext_set/*.csv', 'CPE')
def plot_venn_diagram(csvFile, n):
def get_venn_sections(sets):
num_combinations = 2 ** len(sets)
bit_flags = [2 ** n for n in range(len(sets))]
flags_zip_sets = [z for z in zip(bit_flags, sets)]
combo_sets = []
for bits in range(num_combinations - 1, 0, -1):
include_sets = [s for flag, s in flags_zip_sets if bits & flag]
exclude_sets = [s for flag, s in flags_zip_sets if not bits & flag]
combo = set.intersection(*include_sets)
combo = set.difference(combo, *exclude_sets)
tag = ''.join([str(int((bits & flag) > 0)) for flag in bit_flags])
combo_sets.append((tag, combo))
return combo_sets
df1 = pd.read_csv(csvFile)
pred_csv1_0_indices = [index for index, element in enumerate(df1['fingerprint'].to_list()) if element == 0]
pred_csv1_1_indices = [index for index, element in enumerate(df1['fingerprint'].to_list()) if element == 1]
pred_csv2_0_indices = [index for index, element in enumerate(df1['pharmacophore'].to_list()) if element == 0]
pred_csv2_1_indices = [index for index, element in enumerate(df1['pharmacophore'].to_list()) if element == 1]
pred_csv3_0_indices = [index for index, element in enumerate(df1['volsurf'].to_list()) if element == 0]
pred_csv3_1_indices = [index for index, element in enumerate(df1['volsurf'].to_list()) if element == 1]
get_common = get_venn_sections([set(pred_csv1_1_indices), set(pred_csv2_1_indices), set(pred_csv3_1_indices)])
venn3([set(pred_csv1_1_indices), set(pred_csv2_1_indices), set(pred_csv3_1_indices)], set_labels= \
['fingerprint', 'pharmacophore', 'volsurf'])
plt.title(n+' :summary of predicted actives of External Sets')
plt.savefig('plots/'+n+'-volsurf-ext'+'.png', dpi=600)
plt.show()
_files = ['ext_pred_cpe_3cl_volsurf/cpe_external_set_after_phys-chem-filters_stand.csv']
names = ['CPE']
for _f, n in zip(_files, names) :
plot_venn_diagram(_f, n)
```
| github_jupyter |
# Script to perform some basic data exploration
```
import pandas as pd
import numpy as np
from collections import Counter
import matplotlib.pyplot as plt
path_to_dataset = "/home/shagun/FortKnox/Quora/quora_duplicate_questions.tsv"
# Load the dataset into a pandas dataframe
df = pd.read_csv(path_to_dataset, delimiter="\t")
print("Total number of question pairs = ", str(len(df)))
# Let us look at a sample of the dataset
df_sample = df.sample(5)
df_sample
# And some basic statistics
df.describe()
print("Around", str(round(df.describe()['is_duplicate']['mean']*100)), "% of the question pairs are duplicates.")
```
## Let us dive deep into the data
```
sample_data = df.sample(10)
labels = list(sample_data['is_duplicate'].apply(lambda x: int(x)).values)
labels = np.asarray(labels)
labels
first_question_list = list(sample_data['question1'].apply(lambda x: str(x)).values)
second_question_list = list(sample_data['question2'].apply(lambda x: str(x)).values)
question_list = list(zip(first_question_list, second_question_list))
print("Question1: ",question_list[0][0])
print("Question2: ",question_list[0][1])
print("Label: ", str(labels[0]))
uniq_words_counter = Counter()
for question_tuple in question_list:
for question in question_tuple:
if(isinstance(question, str)):
for word in question.split():
# print(word)
uniq_words_counter.update([word])
uniq_words_count = str(len(uniq_words_counter))
print("Unique words in the dataset: "+ uniq_words_count)
```
## Let us compute the question statistics
```
# Create a list of all the question pairs
first_question_list = list(df['question1'].apply(lambda x: str(x)).values)
second_question_list = list(df['question2'].apply(lambda x: str(x)).values)
question_list = list(zip(first_question_list, second_question_list))
print(len(question_list))
uniq_words_counter = Counter()
for question_tuple in question_list:
for question in question_tuple:
if(isinstance(question, str)):
for word in question.split():
uniq_words_counter.update([word])
uniq_words_count = str(len(uniq_words_counter.items()))
print("Unique words in the dataset: "+ uniq_words_count)
question_list[0]
'india?' in uniq_words_counter
question_length_list = []
for question_tuple in question_list:
for question in question_tuple:
question_length_list.append(len(question.split()))
question_length_list = np.asarray(question_length_list)
print("Average question length: ", str(np.average(question_length_list)))
print("\n")
print("Median question length: ", str(np.median(question_length_list)))
print("\n")
print("Min question length: ", str(np.min(question_length_list)))
print("\n")
arg_min = int(np.argmin(question_length_list)/2)
print("Shortest question: ", question_list[arg_min])
# print("Label: ", df['is_duplicate'][arg_min])
print("\n")
print("Max question length: ", str(np.max(question_length_list)))
arg_max = int(np.argmax(question_length_list)/2)
print("\n")
print("Longest question: ", question_list[arg_max])
import matplotlib.pyplot as plt
from numpy.random import normal
# gaussian_numbers = normal(size=1000)
plt.hist(question_length_list)
plt.title("Question Length Histogram")
plt.xlabel("Question Length")
plt.ylabel("Frequency")
plt.show()
```
### It should be sufficient to pad our questions to a maximum length of 50
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#@title MIT License
#
# Copyright (c) 2017 François Chollet
#
# Permission is hereby granted, free of charge, to any person obtaining a
# copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
```
# Classificação de texto com avaliações de filmes
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/pt/r1/tutorials/keras/basic_text_classification.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Execute em Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/pt/r1/tutorials/keras/basic_text_classification.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />Veja a fonte em GitHub</a>
</td>
</table>
Este *notebook* classifica avaliações de filmes como **positiva** ou **negativa** usando o texto da avaliação. Isto é um exemplo de classificação *binária* —ou duas-classes—, um importante e bastante aplicado tipo de problema de aprendizado de máquina.
Usaremos a base de dados [IMDB](https://www.tensorflow.org/api_docs/python/tf/keras/datasets/imdb) que contém avaliaçòes de mais de 50000 filmes do bando de dados [Internet Movie Database](https://www.imdb.com/). A base é dividida em 25000 avaliações para treinamento e 25000 para teste. Os conjuntos de treinamentos e testes são *balanceados*, ou seja, eles possuem a mesma quantidade de avaliações positivas e negativas.
O notebook utiliza [tf.keras](https://www.tensorflow.org/r1/guide/keras), uma API alto-nível para construir e treinar modelos com TensorFlow. Para mais tutoriais avançados de classificação de textos usando `tf.keras`, veja em [MLCC Text Classification Guide](https://developers.google.com/machine-learning/guides/text-classification/).
```
# keras.datasets.imdb está quebrado em 1.13 e 1.14, pelo np 1.16.3
!pip install tf_nightly
import tensorflow.compat.v1 as tf
from tensorflow import keras
import numpy as np
print(tf.__version__)
```
## Baixe a base de dados IMDB
A base de dados vem empacotada com TensorFlow. Ele já vem pré-processado de forma que as avaliações (sequências de palavras) foi convertida em sequências de inteiros, onde cada inteiro representa uma palavra específica no dicionário.
O código abaixo baixa a base de dados IMDB para a sua máquina (ou usa a cópia em *cache*, caso já tenha baixado):
```
imdb = keras.datasets.imdb
(train_data, train_labels), (test_data, test_labels) = imdb.load_data(num_words=10000)
```
O argumento `num_words=10000` mantém as 10000 palavras mais frequentes no conjunto de treinamento. As palavras mais raras são descartadas para preservar o tamanho dos dados de forma maleável.
## Explore os dados
Vamos parar um momento para entender o formato dos dados. O conjunto de dados vem pré-processado: cada exemplo é um *array* de inteiros representando as palavras da avaliação do filme. Cada *label* é um inteiro com valor ou de 0 ou 1, onde 0 é uma avaliação negativa e 1 é uma avaliação positiva.
```
print("Training entries: {}, labels: {}".format(len(train_data), len(train_labels)))
```
O texto das avaliações foi convertido para inteiros, onde cada inteiro representa uma palavra específica no dicionário. Isso é como se parece a primeira revisão:
```
print(train_data[0])
```
As avaliações dos filmes têm diferentes tamanhos. O código abaixo mostra o número de palavras da primeira e segunda avaliação. Sabendo que o número de entradas da rede neural tem que ser de mesmo também, temos que resolver isto mais tarde.
```
len(train_data[0]), len(train_data[1])
```
### Converta os inteiros de volta a palavras
É util saber como converter inteiros de volta a texto. Aqui, criaremos uma função de ajuda para consultar um objeto *dictionary* que contenha inteiros mapeados em strings:
```
# Um dicionário mapeando palavras em índices inteiros
word_index = imdb.get_word_index()
# Os primeiros índices são reservados
word_index = {k:(v+3) for k,v in word_index.items()}
word_index["<PAD>"] = 0
word_index["<START>"] = 1
word_index["<UNK>"] = 2 # unknown
word_index["<UNUSED>"] = 3
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
def decode_review(text):
return ' '.join([reverse_word_index.get(i, '?') for i in text])
```
Agora, podemos usar a função `decode_review` para mostrar o texto da primeira avaliação:
```
decode_review(train_data[0])
```
## Prepare os dados
As avaliações—o *arrays* de inteiros— deve ser convertida em tensores (*tensors*) antes de alimentar a rede neural. Essa conversão pode ser feita de duas formas:
* Converter os arrays em vetores de 0s e 1s indicando a ocorrência da palavra, similar com *one-hot encoding*. Por exemplo, a sequência [3, 5] se tornaria um vetor de 10000 dimensões, onde todos seriam 0s, tirando 3 would become a 10,000-dimensional vector that is all zeros except for indices 3 and 5, which are ones. Then, make this the first layer in our network—a Dense layer—that can handle floating point vector data. This approach is memory intensive, though, requiring a `num_words * num_reviews` size matrix.
* Alternatively, we can pad the arrays so they all have the same length, then create an integer tensor of shape `max_length * num_reviews`. We can use an embedding layer capable of handling this shape as the first layer in our network.
In this tutorial, we will use the second approach.
Since the movie reviews must be the same length, we will use the [pad_sequences](https://keras.io/preprocessing/sequence/#pad_sequences) function to standardize the lengths:
```
train_data = keras.preprocessing.sequence.pad_sequences(train_data,
value=word_index["<PAD>"],
padding='post',
maxlen=256)
test_data = keras.preprocessing.sequence.pad_sequences(test_data,
value=word_index["<PAD>"],
padding='post',
maxlen=256)
```
Let's look at the length of the examples now:
```
len(train_data[0]), len(train_data[1])
```
And inspect the (now padded) first review:
```
print(train_data[0])
```
## Construindo o modelo
A rede neural é criada por camadas empilhadas —isso necessita duas decisões arquiteturais principais:
* Quantas camadas serão usadas no modelo?
* Quantas *hidden units* são usadas em cada camada?
Neste exemplo, os dados de entrada são um *array* de palavras-índices. As *labels* para predizer são ou 0 ou 1. Vamos construir um modelo para este problema:
```
# O formato de entrada é a contagem vocabulário usados pelas avaliações dos filmes (10000 palavras)
vocab_size = 10000
model = keras.Sequential()
model.add(keras.layers.Embedding(vocab_size, 16))
model.add(keras.layers.GlobalAveragePooling1D())
model.add(keras.layers.Dense(16, activation=tf.nn.relu))
model.add(keras.layers.Dense(1, activation=tf.nn.sigmoid))
model.summary()
```
As camadas são empilhadas sequencialmente para construir o classificador:
1. A primeira camada é uma camada `Embedding` layer (*`Embedding` layer*). Essa camada pega o vocabulário em inteiros e olha o vetor *embedding* em cada palavra-index. Esses vetores são aprendidos pelo modelo, ao longo do treinamento. Os vetores adicionam a dimensão ao *array* de saída. As dimensões resultantes são: `(batch, sequence, embedding)`.
2. Depois, uma camada `GlobalAveragePooling1D` retorna um vetor de saída com comprimento fixo para cada exemplo fazendo a média da sequência da dimensão. Isso permite o modelo de lidar com entradas de tamanhos diferentes da maneira mais simples possível.
3. Esse vetor de saída com tamanho fixo passa por uma camada *fully-connected* (`Dense`) layer com 16 *hidden units*.
4. A última camada é uma *densely connected* com um único nó de saída. Usando uma função de ativação `sigmoid`, esse valor é um float que varia entre 0 e 1, representando a probabilidade, ou nível de confiança.
### Hidden units
O modelo abaixo tem duas camadas intermediárias ou _"hidden"_ (hidden layers), entre a entrada e saída. O número de saídas (unidades— *units*—, nós ou neurônios) é a dimensão do espaço representacional para a camada. Em outras palavras, a quantidade de liberdade que a rede é permitida enquanto aprende uma representação interna.
Se o modelo tem mais *hidden units* (um espaço representacional de maior dimensão), e/ou mais camadas, então a rede pode aprender representações mais complexas. Entretanto, isso faz com que a rede seja computacionamente mais custosa e pode levar o aprendizado de padrões não desejados— padrões que melhoram a performance com os dados de treinamento, mas não com os de teste. Isso se chama *overfitting*, e exploraremos mais tarde.
### Função Loss e otimizadores (optimizer)
O modelo precisa de uma função *loss* e um otimizador (*optimizer*) para treinamento. Já que é um problema de classificação binário e o modelo tem com saída uma probabilidade (uma única camada com ativação sigmoide), usaremos a função loss `binary_crossentropy`.
Essa não é a única escolha de função loss, você poderia escolher, no lugar, a `mean_squared_error`. Mas, geralmente, `binary_crossentropy` é melhor para tratar probabilidades— ela mede a "distância" entre as distribuições de probabilidade, ou, no nosso caso, sobre a distribuição real e as previsões.
Mais tarde, quando explorarmos problemas de regressão (como, predizer preço de uma casa), veremos como usar outra função loss chamada *mean squared error*.
Agora, configure o modelo para usar o *optimizer* a função loss:
```
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['acc'])
```
### Crie um conjunto de validação
Quando treinando. queremos checar a acurácia do modelo com os dados que ele nunca viu. Crie uma conjunto de *validação* tirando 10000 exemplos do conjunto de treinamento original. (Por que não usar o de teste agora? Nosso objetivo é desenvolver e melhorar (tunar) nosso modelo usando somente os dados de treinamento, depois usar o de teste uma única vez para avaliar a acurácia).
```
x_val = train_data[:10000]
partial_x_train = train_data[10000:]
y_val = train_labels[:10000]
partial_y_train = train_labels[10000:]
```
## Treine o modelo
Treine o modelo em 40 *epochs* com *mini-batches* de 512 exemplos. Essas 40 iterações sobre todos os exemplos nos tensores `x_train` e `y_train`. Enquanto treina, monitore os valores do loss e da acurácia do modelo nos 10000 exemplos do conjunto de validação:
```
history = model.fit(partial_x_train,
partial_y_train,
epochs=40,
batch_size=512,
validation_data=(x_val, y_val),
verbose=1)
```
## Avalie o modelo
E vamos ver como o modelo se saiu. Dois valores serão retornados. Loss (um número que representa o nosso erro, valores mais baixos são melhores), e acurácia.
```
results = model.evaluate(test_data, test_labels, verbose=2)
print(results)
```
Está é uma aproximação ingênua que conseguiu uma acurácia de 87%. Com mais abordagens avançadas, o modelo deve chegar em 95%.
## Crie um gráfico de acurácia e loss por tempo
`model.fit()` retorna um objeto `History` que contém um dicionário de tudo o que aconteceu durante o treinamento:
```
history_dict = history.history
history_dict.keys()
```
Tem 4 entradas: uma para cada métrica monitorada durante a validação e treinamento. Podemos usá-las para plotar a comparação do loss de treinamento e validação, assim como a acurácia de treinamento e validação:
```
import matplotlib.pyplot as plt
acc = history_dict['acc']
val_acc = history_dict['val_acc']
loss = history_dict['loss']
val_loss = history_dict['val_loss']
epochs = range(1, len(acc) + 1)
# "bo" de "blue dot" ou "ponto azul"
plt.plot(epochs, loss, 'bo', label='Training loss')
# b de "solid blue line" "linha azul"
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
plt.clf() # limpa a figura
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.show()
```
No gráfico, os pontos representam o loss e acurácia de treinamento, e as linhas são o loss e a acurácia de validação.
Note que o loss de treinamento *diminui* a cada *epoch* e a acurácia *aumenta*. Isso é esperado quando usado um gradient descent optimization—ele deve minimizar a quantidade desejada a cada iteração.
Esse não é o caso do loss e da acurácia de validação— eles parecem ter um pico depois de 20 epochs. Isso é um exemplo de *overfitting*: o modelo desempenha melhor nos dados de treinamento do que quando usado com dados nunca vistos. Depois desse ponto, o modelo otimiza além da conta e aprende uma representação *especifica* para os dados de treinamento e não *generaliza* para os dados de teste.
Para esse caso particular, podemos prevenir o *overfitting* simplesmente parando o treinamento após mais ou menos 20 epochs. Depois, você verá como fazer isso automaticamente com um *callback*.
| github_jupyter |
# OpenML CC18 Metalearning Benchmark
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pandera as pa
import plotly.express as px
import re
import seaborn as sns
from pathlib import Path
# environment variables
JOB = 338
RESULTS_ROOT = Path("..") / "floyd_outputs"
```
## Training
```
training_results = (
pd.concat([
pd.read_csv(path) for path in
(RESULTS_ROOT / str(JOB)).glob("metalearn_training_results_trial_*.csv")
])
.sort_values(["trial_num", "episode"])
)
training_results.head(3)
```
### Mean Rewards per Episode
It looks like the experiment output is missing training results
data for trials 1, 2, 3, and 6. Not sure why this happened, need
to check that the trail numbers are properly recorded in the
`metalearn.experiment` module.
```
training_results.query("episode == 1")
training_results.trial_num.unique()
training_results[["trial_num", "entropy_coef_anneal_to", "learning_rate"]].drop_duplicates()
```
Roughly define regret as ${validation\ score}_{max} - {validation\ score}_{mean}$
for a particular episode. Where ${validation\ score}_{max}$ is the best validation
set found by all of the hyperparameter conditions for a particular dataset.
```
optimal_validation_scores = training_results.groupby("data_env_names").best_validation_scores.max()
training_results.assign(optimal_validation_scores=lambda df: df.data_env_names.map(optimal_validation_scores))
METRICS = [
"losses",
"aggregate_gradients",
"best_validation_scores",
"mean_rewards",
"mean_validation_scores",
"n_successful_mlfs",
"mlf_diversity",
"hyperparam_diversity",
]
agg_performance_results = (
training_results
.assign(
optimal_validation_scores=lambda df: df.data_env_names.map(
training_results.groupby("data_env_names").best_validation_scores.max())
)
.assign(regret=lambda df: df.optimal_validation_scores - df.mean_validation_scores)
.groupby([
"trial_num", "entropy_coef_anneal_to", "learning_rate",
])
.apply(lambda df: df.assign(cumulative_regret=df.regret.cumsum()))
.reset_index(drop=True)
.assign(
trial_info=lambda df: (
"trial = " + df.trial_num.astype(str) +
"; lr = " + df.learning_rate.astype(str) +
"; ec = " + df.entropy_coef_anneal_to.astype(str)
)
)
.pipe(lambda df: pd.concat([
df,
(
df
.set_index(["trial_info", "episode"])
.groupby("trial_info")
.apply(lambda df: df[METRICS].ewm(alpha=0.05).mean())
)
], sort=False))
)
fig, ax = plt.subplots(1, 1, figsize=(12, 8))
g = sns.lineplot(
data=agg_performance_results,
x="episode",
y="cumulative_regret",
hue="trial_info",
ax=ax,
)
plot_df = (
agg_performance_results
.set_index(["trial_info", "episode"])
.groupby("trial_info")
.apply(lambda df: df[METRICS].ewm(alpha=0.01).mean())
.reset_index()
)
fig, ax = plt.subplots(figsize=(12, 8))
g = sns.lineplot(
data=plot_df,
x="episode",
y="best_validation_scores",
hue="trial_info",
ax=ax,
)
plot_df = (
agg_performance_results
.set_index(["trial_info", "episode"])
.groupby("trial_info")
.apply(lambda df: df[METRICS].ewm(alpha=0.01).mean())
.reset_index()
)
fig, ax = plt.subplots(figsize=(12, 8))
g = sns.lineplot(
data=plot_df,
x="episode",
y="mean_rewards",
hue="trial_info",
ax=ax,
)
fig, ax = plt.subplots(1, 1, figsize=(12, 8))
g = sns.lineplot(
data=training_results.assign(
hue=lambda df: (
"trial = " + df.trial_num.astype(str) +
"; lr = " + df.learning_rate.astype(str)
)
),
x="episode",
y="n_unique_mlfs",
hue="hue",
ax=ax,
)
```
#### Plot Performance per Trial
```
hyperparameters = ["entropy_coef_anneal_to", "learning_rate"]
metrics = ["mean_validation_scores", "mean_rewards", "n_unique_mlfs"]
statistics = ["mean", "median"]
summary_performance_df = (
training_results
.groupby(["episode", "trial_num"])
.agg({metric: statistics for metric in metrics})
.set_axis(
labels=[
f"{stat}_of_{metric}"
for metric in metrics
for stat in statistics],
axis="columns",
inplace=False
)
# .reset_index()
)
summary_performance_df.head(3)
```
## Inference
```
check_betw_zero_one = pa.Check(lambda s: (0.0 <= s) & (s <= 1.0))
INFERENCE_RESULT_SCHEMA = pa.DataFrameSchema(
columns={
"data_env": pa.Column(pa.String),
"n_inference_steps": pa.Column(pa.Int),
"is_valid": pa.Column(pa.Bool),
"reward": pa.Column(pa.Float),
"validation_score": pa.Column(pa.Float),
},
coerce=True,
)
@pa.check_output(INFERENCE_RESULT_SCHEMA)
def data_to_longform(inference_results):
return (
inference_results
[["data_env", "n_inference_steps", "key", "value"]]
.dropna(subset=["value"])
.pivot_table(
index=["data_env", "n_inference_steps"],
columns="key",
values="value",
aggfunc=lambda x: x,
)
.reset_index()
.dropna()
# all scores should be strings
.loc[
lambda df: (
df.validation_score.map(lambda x: isinstance(x, str)) &
df.mlf.map(lambda x: isinstance(x, str)) &
df.reward.map(lambda x: isinstance(x, str)) &
df.is_valid.map(lambda x: isinstance(x, str))
)
]
.rename_axis(None, axis=1)
.reset_index(drop=True)
)
inference_results = pd.concat([
(
pd.read_csv(f"../floyd_outputs/{JOB}/{env}_env_inference_results.csv")
.pipe(data_to_longform)
.assign(data_env_partition=env)
) for env in ["training", "test"]
])
inference_results.head()
```
## Plot Validation Scores
```
px.line(
inference_results.query("data_env_partition == 'training'"),
x="n_inference_steps",
y="reward",
template="plotly_white",
color="data_env",
)
```
### Plot of Test Data Environment Validation Scores
```
px.line(
inference_results.query("data_env_partition == 'test'"),
x="n_inference_steps",
y="reward",
template="plotly_white",
color="data_env",
)
with pd.option_context("display.max_rows", 200):
display(
inference_results
.groupby(["data_env_partition", "data_env"])
.apply(lambda df: df.loc[df.validation_score.idxmax()])
.reset_index(drop=True)
.groupby("data_env_partition")
.apply(lambda df: df.sort_values("validation_score"))
.head()
)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import pandas as pd
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import math
import seaborn as sns
import matplotlib.colors as mcolors
import statsmodels.api as sm
import statsmodels.formula.api as smf
from statsmodels.formula.api import ols
from statsmodels.formula.api import mixedlm
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
import os
import matplotlib.pyplot as mpl
import matplotlib
from scipy.stats import spearmanr
colors = list(mcolors.TABLEAU_COLORS.keys())*2
parentDirectory = os.path.abspath(os.path.join(os.path.join(os.path.join(os.getcwd(), os.pardir), os.pardir),os.pardir))
DATA_DIR = parentDirectory +'/data/'
FIGURES_DIR = parentDirectory +'/figures/'
full_names = {
'AU': 'Australia',
'BR': 'Brazil',
'CA': 'Canada',
'FR': 'France',
'DE': 'Germany',
'IN': 'India',
'IT': 'Italy',
'MX': 'Mexico',
'ES': 'Spain',
'GB': 'United Kingdom',
'US': 'United States',
'DK': 'Denmark',
'KE': 'Kenya',
'NG': 'Nigeria',
'JP': 'Japan',
'SE': 'Sweden',
'ID': 'Indonesia',
'EG': 'Egypt'
}
event_dicts = [{'country': 'AU',
'end_md_1': '2020-06-07',
'start_md_1': '2020-03-27',
'start_md_2': np.nan},
{'country': 'BR',
'end_md_1': '2020-08-09',
'start_md_1': '2020-03-23',
'start_md_2': np.nan},
{'country': 'CA',
'end_md_1': '2020-06-21',
'start_md_1': '2020-03-19',
'start_md_2': '2020-10-12'},
{'country': 'DE',
'end_md_1': '2020-05-09',
'start_md_1': '2020-03-21',
'start_md_2': '2020-12-18'},
{'country': 'DK',
'end_md_1': '2020-05-07',
'start_md_1': '2020-03-17',
'start_md_2': np.nan},
{'country': 'EG',
'end_md_1': '2020-07-01',
'start_md_1': '2020-03-24',
'start_md_2': np.nan},
{'country': 'ES',
'end_md_1': '2020-06-14',
'start_md_1': '2020-03-17',
'start_md_2': '2020-11-07'},
{'country': 'FR',
'end_md_1': '2020-06-08',
'start_md_1': '2020-03-18',
'start_md_2': '2020-11-01'},
{'country': 'GB',
'end_md_1': '2020-08-03',
'start_md_1': '2020-03-23',
'start_md_2': '2020-10-21'},
{'country': 'ID',
'end_md_1': '2020-08-10',
'start_md_1': '2020-03-24',
'start_md_2': np.nan},
{'country': 'IN',
'end_md_1': '2020-10-29',
'start_md_1': '2020-03-24',
'start_md_2': np.nan},
{'country': 'IT',
'end_md_1': '2020-06-06',
'start_md_1': '2020-03-11',
'start_md_2': '2020-11-06'},
{'country': 'JP',
'end_md_1': '2020-05-30',
'start_md_1': '2020-04-12',
'start_md_2': np.nan},
{'country': 'KE',
'end_md_1': '2020-10-04',
'start_md_1': '2020-03-24',
'start_md_2': np.nan},
{'country': 'MX',
'end_md_1': '2020-10-06',
'start_md_1': '2020-03-25',
'start_md_2': np.nan},
{'country': 'NG',
'end_md_1': '2020-08-09',
'start_md_1': '2020-03-27',
'start_md_2': np.nan},
{'country': 'SE',
'end_md_1': '2020-04-09',
'start_md_1': '2020-04-03',
'start_md_2': np.nan},
{'country': 'US',
'end_md_1': '2020-06-11',
'start_md_1': '2020-03-21',
'start_md_2': '2020-11-26'}]
df_events = pd.DataFrame(event_dicts)
df_events['start_md_1'] = pd.to_datetime(df_events['start_md_1'])
df_events['end_md_1'] = pd.to_datetime(df_events['end_md_1'])
df_events['start_md_2'] = pd.to_datetime(df_events['start_md_2'])
df_agg = pd.read_pickle(DATA_DIR+'df_agg_cats.pickle')
def make_stars(val):
if val<0.0001:
return '****'
elif val<0.001:
return '***'
elif val<0.01:
return '**'
elif val<0.05:
return '*'
else:
return ''
def make_star_ste(value,ste):
if value>0 and value-2*ste>0:
return '*'
elif value<0 and value+2*ste<0:
return '*'
else:
return ''
weeks_2019 = list(df_agg.iloc[0]['volume_weekly_total'].index)[:52]
weeks_2020 = list(df_agg.iloc[0]['volume_weekly_total'].index)[52:]
l = []
for cnt, row in df_agg.iterrows():
start_md = df_events.loc[df_events['country'] == row['country']].iloc[0]['start_md_1']
end_md = df_events.loc[df_events['country'] == row['country']].iloc[0]['end_md_1']
start_md2 = df_events.loc[df_events['country'] == row['country']].iloc[0]['start_md_2']
for week in zip(row['volume_weekly_total'].index,row['volume_weekly_total'].values,row['volume_percent_weekly_total'].values):
entry = {}
entry['country'] = row['country']
entry['category'] = row['category']
if week[0] in weeks_2020:
date = pd.to_datetime(week[0])
if type(start_md2)!=pd._libs.tslibs.nattype.NaTType and date > start_md2:
continue
entry['k'] = math.floor(((date - start_md).days +7) / 7)
entry['volume_total'] = week[1]
entry['volume_percent'] = week[2]
entry['year'] = '2020'
l.append(entry)
elif week[0] in weeks_2019:
date = pd.to_datetime(weeks_2020[weeks_2019.index(week[0])])
if type(start_md2)!=pd._libs.tslibs.nattype.NaTType and date > start_md2:
continue
entry['k'] = math.floor(((date - start_md).days +7) / 7)
entry['volume_total'] = week[1]
entry['volume_percent'] = week[2]
entry['year'] = '2019'
l.append(entry)
df = pd.DataFrame(l)
df = df.loc[(df['k'] >= -30) & (df['k'] <= 30)]
df = df.loc[(df['country'].isin(list(full_names.keys())))]
df['intervention_flag'] = df['k'].apply(lambda x: 1 if x >= 0 else 0)
cats = list(df['category'].unique())
k = 30
df_temp = df.loc[(df['k'] >= -k) & (df['k'] <= k)].copy()
df_temp['volume_total'] = df_temp['volume_total'].apply(lambda x: np.log(x + 0.001))
entries_list = []
for name, group in df_temp.groupby(['category']):
print(name)
entry = {}
mod = smf.ols('volume_total ~ intervention_flag*year + C(country)', data = group)
res = mod.fit(cov_type='hc0')
entry['model_degree'] = 0
entry['category'] = name
entry['alpha'] = res.params['intervention_flag:year[T.2020]']
entry['ste'] = res.bse['intervention_flag:year[T.2020]']
entry['pval'] = res.pvalues['intervention_flag:year[T.2020]']
entry['r2'] = res.rsquared
entries_list.append(entry)
entry = {}
mod = smf.ols('volume_total ~ intervention_flag*k*year + C(country)', data = group)
res = mod.fit(cov_type='hc0')
entry['model_degree'] = 1
entry['category'] = name
entry['alpha'] = res.params['intervention_flag:year[T.2020]']
entry['ste'] = res.bse['intervention_flag:year[T.2020]']
entry['pval'] = res.pvalues['intervention_flag:year[T.2020]']
entry['r2'] = res.rsquared
entries_list.append(entry)
entry = {}
mod = smf.ols('volume_total ~ intervention_flag*k*year + intervention_flag*np.power(k,2)*year + C(country)', data = group)
res = mod.fit(cov_type='hc0')
entry['model_degree'] = 2
entry['category'] = name
entry['alpha'] = res.params['intervention_flag:year[T.2020]']
entry['ste'] = res.bse['intervention_flag:year[T.2020]']
entry['pval'] = res.pvalues['intervention_flag:year[T.2020]']
entry['r2'] = res.rsquared
entries_list.append(entry)
df_res = pd.DataFrame(entries_list)
for j in range(28):
print(str(j+1)+' &')
for i in range(3):
if i ==2:
print(df_res.loc[df_res['model_degree']==i].sort_values(by = 'alpha', ascending = False)['category'].values[j] )
else:
print(df_res.loc[df_res['model_degree']==i].sort_values(by = 'alpha', ascending = False)['category'].values[j] + ' &')
print('\\\\')
spearmanr(df_res.loc[df_res['model_degree']==0]['alpha'].values ,
df_res.loc[df_res['model_degree']==2]['alpha'].values)
spearmanr(df_res.loc[df_res['model_degree']==1]['alpha'].values ,
df_res.loc[df_res['model_degree']==2]['alpha'].values)
spearmanr(df_res.loc[df_res['model_degree']==0]['alpha'].values ,
df_res.loc[df_res['model_degree']==1]['alpha'].values)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
import sys
sys.path.append("..")
import source.explore as exp
pd.set_option('max_columns', 200)
```
From a previous run, we have the out of folds predictions over our training set. We put it together with the original set.
```
df = pd.read_csv('../oof_pred/for_error_analysis.csv')
df.head()
```
We can see how all the predictions are very correlated to one another (and with the target, this is good)
```
exp.plot_correlations(df[[col for col in df.columns if '_oof' in col]+['target']],
target='target', annot=True)
oof_cols = [col for col in df.columns if '_oof' in col]
for col in oof_cols:
name = col.replace('_oof', '_res')
df[name] = df['target'] - df[col]
exp.plot_correlations(df[[col for col in df.columns if '_oof' in col]+
[col for col in df.columns if '_res' in col]+
['target']], target='target', annot=True)
```
This is even more evident if we look at the following plots.
```
exp.corr_target(df, 'target',
[col for col in df.columns if '_oof' in col]+
[col for col in df.columns if '_res' in col])
```
Looking at the residual plots, it appears evident that all the models we trained so far are underestimating the price of low costs houses and overestimating the more expensive ones. This could be because we used some target encoding or simply that we are overestimating, for example, the importance of the house size.
We can try to see if there are interesting relations between the residuals and the original features.
```
exp.plot_correlations(df, target='lgb_res')
```
Or, for the categorical features, we can start focusing on the feature that was used both to stratify our folds (and test set) and then to be target encoded: Neighborhood.
```
exp.segm_target(df, 'Neighborhood', 'lgb_res')
err = exp.segm_target(df, 'Neighborhood', 'lgb_res')
tar = exp.segm_target(df, 'Neighborhood', 'target')
tot = pd.merge(err.reset_index(), tar.reset_index(), on='Neighborhood', suffixes=('_res', '_target'))
del tot['count_target']
tot
tot.corr()
```
A few unsurprising things are:
* the more houses from a neighborhood, the smaller the error on average. Which is also the pattern with the price, so we have to be mindful of that (neighborhood with more examples tend to have lower costs on average).
* We can confirm that neighborhood for which we have houses with higher average cost also get an higher error (not in absolute sense).
This makes me consider if it would be a good idea to not use the target encoding variables and see if that pattern in the error disappears.
Another possible test is to see if some variables we did not include, for example
```
exp.segm_target(df, 'Exterior1st', 'lgb_res')
```
shows how the `MetalSd` exterior leads to a particularly different pattern in the distribution of the error. A direct inspection of these houses shows the following
```
df[df.Exterior1st == 'MetalSd'].describe() - df[df.Exterior1st != 'MetalSd'].describe()
```
In other words, houses with that particular exterior
* Are less likely to have land in front of them
* Are lower in quality, in particular they never hit the perfect score. This could be interpreted as a bias in the data collection
* Were built less recently, which makes perfect sense as building techniques change with time
* Have much smaller basements and garages
* Are much smaller in general
* Are less likely to have a fireplace
* Cost less in general
We could then consider to include this feature as well and see how the model reacts.
Another approach would be to explore the entries with the biggest errors. For example
```
def high_low_errors(data, *, res_list=None, n_samples=50,
target=None, pred_list=None, mean=False,
abs_err=True, common=False):
df = data.copy()
if pred_list:
res_list = []
for col in pred_list:
name = col + '_res'
res_list.append(name)
df[name] = df[target] - df[col]
errors = {}
if mean:
df['mean_res'] = df[res_list].mean(axis=1)
res_list += ['mean_res']
for col in res_list:
if abs_err:
if col == 'abs_err':
name = 'abs_err'
else:
name = 'abs_' + col
df[name] = abs(df[col])
else:
name = col
high_err = df.sort_values(name, ascending=False).head(n_samples)
low_err = df.sort_values(name, ascending=False).tail(n_samples)
try:
errors[name] = high_err.describe(include='all').drop(index=['top', 'count', 'freq']).fillna(0) - \
low_err.describe(include='all').drop(index=['top', 'count', 'freq']).fillna(0)
except KeyError:
errors[name] = high_err.describe().fillna(0) - low_err.describe().fillna(0)
return errors
h_v_l = high_low_errors(df, res_list=[col for col in df.columns if '_res' in col], mean=True)
h_v_l.keys()
h_v_l['abs_mean_res']
```
* Alley is a low cardinality feature, that difference might be interesting
* High errors have bigger LotFrontage but much smaller LotArea
* Low errors are built more recently
* High errors have a bigger basement but also more unfinished
* High errors are much bigger in general
* It appears we are not capturing the MiscVal
* The high are negative on average, meaning that they overestimate the price
* Low errors are coming from more expensive houses.
# Using the actual model data
```
df = pd.read_csv('../oof_pred/for_error_analysis_lgb_transf.csv')
df.head()
exp.plot_correlations(df, target='lasso_oof')
h_v_l = high_low_errors(df, res_list=[col for col in df.columns if '_res' in col], mean=True)
h_v_l['abs_mean_res']
```
We indeed see how `MSSubClass` and `Neighborhood` are very different for high and low errors. This is a consequence of the lower prices of the houses with low error. The suspect that the model is following too much these 2 features is getting bigger.
| github_jupyter |
```
# default_exp models.layers
```
# Layers
> Helper function used to build PyTorch timeseries models.
```
#export
from torch.nn.init import normal_
from fastai.torch_core import Module
from fastai.layers import *
from torch.nn.utils import weight_norm, spectral_norm
from tsai.imports import *
from tsai.utils import *
#export
def noop(x): return x
#export
def init_lin_zero(m):
if isinstance(m, (nn.Linear)):
if getattr(m, 'bias', None) is not None: nn.init.constant_(m.bias, 0)
nn.init.constant_(m.weight, 0)
for l in m.children(): init_lin_zero(l)
lin_zero_init = init_lin_zero
#export
class SwishBeta(Module):
def __init__(self, beta=1.):
self.sigmoid = torch.sigmoid
self.beta = nn.Parameter(torch.Tensor(1).fill_(beta).to(default_device()))
def forward(self, x): return x.mul(self.sigmoid(x*self.beta))
#export
def same_padding1d(seq_len, ks, stride=1, dilation=1):
"Same padding formula as used in Tensorflow"
p = (seq_len - 1) * stride + (ks - 1) * dilation + 1 - seq_len
return p // 2, p - p // 2
class Pad1d(nn.ConstantPad1d):
def __init__(self, padding, value=0.):
super().__init__(padding, value)
@delegates(nn.Conv1d)
class Conv1dSame(Module):
"Conv1d with padding='same'"
def __init__(self, ni, nf, ks=3, stride=1, dilation=1, **kwargs):
self.ks, self.stride, self.dilation = ks, stride, dilation
self.conv1d_same = nn.Conv1d(ni, nf, ks, stride=stride, dilation=dilation, **kwargs)
self.weight = self.conv1d_same.weight
self.bias = self.conv1d_same.bias
self.pad = Pad1d
def forward(self, x):
self.padding = same_padding1d(x.shape[-1], self.ks, dilation=self.dilation) #stride=self.stride not used in padding calculation!
return self.conv1d_same(self.pad(self.padding)(x))
init_linear(Conv1dSame(2, 3, 3), None, init='auto', bias_std=.01)
bs = 2
c_in = 3
c_out = 5
seq_len = 6
t = torch.rand(bs, c_in, seq_len)
test_eq(Conv1dSame(c_in, c_out, ks=3, stride=1, dilation=1, bias=False)(t).shape, (bs, c_out, seq_len))
test_eq(Conv1dSame(c_in, c_out, ks=3, stride=1, dilation=2, bias=False)(t).shape, (bs, c_out, seq_len))
test_eq(Conv1dSame(c_in, c_out, ks=3, stride=2, dilation=1, bias=False)(t).shape, (bs, c_out, seq_len//2))
test_eq(Conv1dSame(c_in, c_out, ks=3, stride=2, dilation=2, bias=False)(t).shape, (bs, c_out, seq_len//2))
#export
def same_padding2d(H, W, ks, stride=(1, 1), dilation=(1, 1)):
"Same padding formula as used in Tensorflow"
if isinstance(ks, Integral): ks = (ks, ks)
if ks[0] == 1: p_h = 0
else: p_h = (H - 1) * stride[0] + (ks[0] - 1) * dilation[0] + 1 - H
if ks[1] == 1: p_w = 0
else: p_w = (W - 1) * stride[1] + (ks[1] - 1) * dilation[1] + 1 - W
return (p_w // 2, p_w - p_w // 2, p_h // 2, p_h - p_h // 2)
class Pad2d(nn.ConstantPad2d):
def __init__(self, padding, value=0.):
super().__init__(padding, value)
@delegates(nn.Conv2d)
class Conv2dSame(Module):
"Conv2d with padding='same'"
def __init__(self, ni, nf, ks=(3, 3), stride=(1, 1), dilation=(1, 1), **kwargs):
if isinstance(ks, Integral): ks = (ks, ks)
if isinstance(stride, Integral): stride = (stride, stride)
if isinstance(dilation, Integral): dilation = (dilation, dilation)
self.ks, self.stride, self.dilation = ks, stride, dilation
self.conv2d_same = nn.Conv2d(ni, nf, ks, stride=stride, dilation=dilation, **kwargs)
self.weight = self.conv2d_same.weight
self.bias = self.conv2d_same.bias
self.pad = Pad2d
def forward(self, x):
self.padding = same_padding2d(x.shape[-2], x.shape[-1], self.ks, dilation=self.dilation) #stride=self.stride not used in padding calculation!
return self.conv2d_same(self.pad(self.padding)(x))
@delegates(nn.Conv2d)
def Conv2d(ni, nf, kernel_size=None, ks=None, stride=1, padding='same', dilation=1, init='auto', bias_std=0.01, **kwargs):
"conv1d layer with padding='same', 'valid', or any integer (defaults to 'same')"
assert not (kernel_size and ks), 'use kernel_size or ks but not both simultaneously'
assert kernel_size is not None or ks is not None, 'you need to pass a ks'
kernel_size = kernel_size or ks
if padding == 'same':
conv = Conv2dSame(ni, nf, kernel_size, stride=stride, dilation=dilation, **kwargs)
elif padding == 'valid': conv = nn.Conv2d(ni, nf, kernel_size, stride=stride, padding=0, dilation=dilation, **kwargs)
else: conv = nn.Conv2d(ni, nf, kernel_size, stride=stride, padding=padding, dilation=dilation, **kwargs)
init_linear(conv, None, init=init, bias_std=bias_std)
return conv
bs = 2
c_in = 3
c_out = 5
h = 16
w = 20
t = torch.rand(bs, c_in, h, w)
test_eq(Conv2dSame(c_in, c_out, ks=3, stride=1, dilation=1, bias=False)(t).shape, (bs, c_out, h, w))
test_eq(Conv2dSame(c_in, c_out, ks=(3, 1), stride=1, dilation=1, bias=False)(t).shape, (bs, c_out, h, w))
test_eq(Conv2dSame(c_in, c_out, ks=3, stride=(1, 1), dilation=(2, 2), bias=False)(t).shape, (bs, c_out, h, w))
test_eq(Conv2dSame(c_in, c_out, ks=3, stride=(2, 2), dilation=(1, 1), bias=False)(t).shape, (bs, c_out, h//2, w//2))
test_eq(Conv2dSame(c_in, c_out, ks=3, stride=(2, 2), dilation=(2, 2), bias=False)(t).shape, (bs, c_out, h//2, w//2))
test_eq(Conv2d(c_in, c_out, ks=3, padding='same', stride=1, dilation=1, bias=False)(t).shape, (bs, c_out, h, w))
#export
class Chomp1d(nn.Module):
def __init__(self, chomp_size):
super(Chomp1d, self).__init__()
self.chomp_size = chomp_size
def forward(self, x):
return x[:, :, :-self.chomp_size].contiguous()
#export
# Modified from https://github.com/locuslab/TCN/blob/master/TCN/tcn.py
class Conv1dCausal(Module):
def __init__(self, ni, nf, ks, stride=1, dilation=1, **kwargs):
padding = (ks - 1) * dilation
self.conv_causal = nn.Conv1d(ni, nf, ks, stride=stride, padding=padding, dilation=dilation, **kwargs)
self.weight = self.conv_causal.weight
self.bias = self.conv_causal.bias
self.chomp_size = padding
def forward(self, x):
x = self.conv_causal(x)
return x[..., :-self.chomp_size].contiguous()
init_linear(Conv1dCausal(2, 3, 3), None, init='auto', bias_std=.01)
bs = 2
c_in = 3
c_out = 5
seq_len = 512
t = torch.rand(bs, c_in, seq_len)
dilation = 1
test_eq(Conv1dCausal(c_in, c_out, ks=3, dilation=dilation)(t).shape, Conv1dSame(c_in, c_out, ks=3, dilation=dilation)(t).shape)
dilation = 2
test_eq(Conv1dCausal(c_in, c_out, ks=3, dilation=dilation)(t).shape, Conv1dSame(c_in, c_out, ks=3, dilation=dilation)(t).shape)
#export
@delegates(nn.Conv1d)
def Conv1d(ni, nf, kernel_size=None, ks=None, stride=1, padding='same', dilation=1, init='auto', bias_std=0.01, **kwargs):
"conv1d layer with padding='same', 'causal', 'valid', or any integer (defaults to 'same')"
assert not (kernel_size and ks), 'use kernel_size or ks but not both simultaneously'
assert kernel_size is not None or ks is not None, 'you need to pass a ks'
kernel_size = kernel_size or ks
if padding == 'same':
if kernel_size%2==1:
conv = nn.Conv1d(ni, nf, kernel_size, stride=stride, padding=kernel_size//2 * dilation, dilation=dilation, **kwargs)
else:
conv = Conv1dSame(ni, nf, kernel_size, stride=stride, dilation=dilation, **kwargs)
elif padding == 'causal': conv = Conv1dCausal(ni, nf, kernel_size, stride=stride, dilation=dilation, **kwargs)
elif padding == 'valid': conv = nn.Conv1d(ni, nf, kernel_size, stride=stride, padding=0, dilation=dilation, **kwargs)
else: conv = nn.Conv1d(ni, nf, kernel_size, stride=stride, padding=padding, dilation=dilation, **kwargs)
init_linear(conv, None, init=init, bias_std=bias_std)
return conv
bs = 2
ni = 3
nf = 5
seq_len = 6
ks = 3
t = torch.rand(bs, c_in, seq_len)
test_eq(Conv1d(ni, nf, ks, padding=0)(t).shape, (bs, c_out, seq_len - (2 * (ks//2))))
test_eq(Conv1d(ni, nf, ks, padding='valid')(t).shape, (bs, c_out, seq_len - (2 * (ks//2))))
test_eq(Conv1d(ni, nf, ks, padding='same')(t).shape, (bs, c_out, seq_len))
test_eq(Conv1d(ni, nf, ks, padding='causal')(t).shape, (bs, c_out, seq_len))
test_error('use kernel_size or ks but not both simultaneously', Conv1d, ni, nf, kernel_size=3, ks=3)
test_error('you need to pass a ks', Conv1d, ni, nf)
conv = Conv1d(ni, nf, ks, padding='same')
init_linear(conv, None, init='auto', bias_std=.01)
conv
conv = Conv1d(ni, nf, ks, padding='causal')
init_linear(conv, None, init='auto', bias_std=.01)
conv
conv = Conv1d(ni, nf, ks, padding='valid')
init_linear(conv, None, init='auto', bias_std=.01)
weight_norm(conv)
conv
conv = Conv1d(ni, nf, ks, padding=0)
init_linear(conv, None, init='auto', bias_std=.01)
weight_norm(conv)
conv
#export
class SeparableConv1d(Module):
def __init__(self, ni, nf, ks, stride=1, padding='same', dilation=1, bias=True, bias_std=0.01):
self.depthwise_conv = Conv1d(ni, ni, ks, stride=stride, padding=padding, dilation=dilation, groups=ni, bias=bias)
self.pointwise_conv = nn.Conv1d(ni, nf, 1, stride=1, padding=0, dilation=1, groups=1, bias=bias)
if bias:
if bias_std != 0:
normal_(self.depthwise_conv.bias, 0, bias_std)
normal_(self.pointwise_conv.bias, 0, bias_std)
else:
self.depthwise_conv.bias.data.zero_()
self.pointwise_conv.bias.data.zero_()
def forward(self, x):
x = self.depthwise_conv(x)
x = self.pointwise_conv(x)
return x
bs = 64
c_in = 6
c_out = 5
seq_len = 512
t = torch.rand(bs, c_in, seq_len)
test_eq(SeparableConv1d(c_in, c_out, 3)(t).shape, (bs, c_out, seq_len))
#export
class AddCoords1d(Module):
"""Add coordinates to ease position identification without modifying mean and std"""
def forward(self, x):
bs, _, seq_len = x.shape
cc = torch.linspace(-1,1,x.shape[-1]).repeat(bs, 1, 1).to(x.device)
cc = (cc - cc.mean()) / cc.std()
x = torch.cat([x, cc], dim=1)
return x
bs = 2
c_in = 3
c_out = 5
seq_len = 50
t = torch.rand(bs, c_in, seq_len)
t = (t - t.mean()) / t.std()
test_eq(AddCoords1d()(t).shape, (bs, c_in + 1, seq_len))
new_t = AddCoords1d()(t)
test_close(new_t.mean(),0, 1e-2)
test_close(new_t.std(), 1, 1e-2)
#export
class ConvBlock(nn.Sequential):
"Create a sequence of conv1d (`ni` to `nf`), activation (if `act_cls`) and `norm_type` layers."
def __init__(self, ni, nf, kernel_size=None, ks=3, stride=1, padding='same', bias=None, bias_std=0.01, norm='Batch', zero_norm=False, bn_1st=True,
act=nn.ReLU, act_kwargs={}, init='auto', dropout=0., xtra=None, coord=False, separable=False, **kwargs):
kernel_size = kernel_size or ks
ndim = 1
layers = [AddCoords1d()] if coord else []
norm_type = getattr(NormType,f"{snake2camel(norm)}{'Zero' if zero_norm else ''}") if norm is not None else None
bn = norm_type in (NormType.Batch, NormType.BatchZero)
inn = norm_type in (NormType.Instance, NormType.InstanceZero)
if bias is None: bias = not (bn or inn)
if separable: conv = SeparableConv1d(ni + coord, nf, ks=kernel_size, bias=bias, stride=stride, padding=padding, **kwargs)
else: conv = Conv1d(ni + coord, nf, ks=kernel_size, bias=bias, stride=stride, padding=padding, **kwargs)
act = None if act is None else act(**act_kwargs)
if not separable: init_linear(conv, act, init=init, bias_std=bias_std)
if norm_type==NormType.Weight: conv = weight_norm(conv)
elif norm_type==NormType.Spectral: conv = spectral_norm(conv)
layers += [conv]
act_bn = []
if act is not None: act_bn.append(act)
if bn: act_bn.append(BatchNorm(nf, norm_type=norm_type, ndim=ndim))
if inn: act_bn.append(InstanceNorm(nf, norm_type=norm_type, ndim=ndim))
if bn_1st: act_bn.reverse()
if dropout: layers += [nn.Dropout(dropout)]
layers += act_bn
if xtra: layers.append(xtra)
super().__init__(*layers)
Conv = partial(ConvBlock, norm=None, act=None)
ConvBN = partial(ConvBlock, norm='Batch', act=None)
ConvIN = partial(ConvBlock, norm='Instance', act=None)
CoordConv = partial(ConvBlock, norm=None, act=None, coord=True)
CoordConvBN = partial(ConvBlock, norm='Batch', act=None, coord=True)
SepConv = partial(ConvBlock, norm=None, act=None, separable=True)
SepConvBN = partial(ConvBlock, norm='Batch', act=None, separable=True)
SepConvIN = partial(ConvBlock, norm='Instance', act=None, separable=True)
SepCoordConv = partial(ConvBlock, norm=None, act=None, coord=True, separable=True)
SepCoordConvBN = partial(ConvBlock, norm='Batch', act=None, coord=True, separable=True)
#export
class ResBlock1dPlus(Module):
"Resnet block from `ni` to `nh` with `stride`"
@delegates(ConvLayer.__init__)
def __init__(self, expansion, ni, nf, coord=False, stride=1, groups=1, reduction=None, nh1=None, nh2=None, dw=False, g2=1,
sa=False, sym=False, norm='Batch', zero_norm=True, act_cls=defaults.activation, ks=3,
pool=AvgPool, pool_first=True, **kwargs):
if nh2 is None: nh2 = nf
if nh1 is None: nh1 = nh2
nf,ni = nf*expansion,ni*expansion
k0 = dict(norm=norm, zero_norm=False, act=act_cls, **kwargs)
k1 = dict(norm=norm, zero_norm=zero_norm, act=None, **kwargs)
convpath = [ConvBlock(ni, nh2, ks, coord=coord, stride=stride, groups=ni if dw else groups, **k0),
ConvBlock(nh2, nf, ks, coord=coord, groups=g2, **k1)
] if expansion == 1 else [
ConvBlock(ni, nh1, 1, coord=coord, **k0),
ConvBlock(nh1, nh2, ks, coord=coord, stride=stride, groups=nh1 if dw else groups, **k0),
ConvBlock(nh2, nf, 1, coord=coord, groups=g2, **k1)]
if reduction: convpath.append(SEModule(nf, reduction=reduction, act_cls=act_cls))
if sa: convpath.append(SimpleSelfAttention(nf,ks=1,sym=sym))
self.convpath = nn.Sequential(*convpath)
idpath = []
if ni!=nf: idpath.append(ConvBlock(ni, nf, 1, coord=coord, act=None, **kwargs))
if stride!=1: idpath.insert((1,0)[pool_first], pool(stride, ndim=1, ceil_mode=True))
self.idpath = nn.Sequential(*idpath)
self.act = defaults.activation(inplace=True) if act_cls is defaults.activation else act_cls()
def forward(self, x): return self.act(self.convpath(x) + self.idpath(x))
#export
def SEModule1d(ni, reduction=16, act=nn.ReLU, act_kwargs={}):
"Squeeze and excitation module for 1d"
nf = math.ceil(ni//reduction/8)*8
assert nf != 0, 'nf cannot be 0'
return SequentialEx(nn.AdaptiveAvgPool1d(1),
ConvBlock(ni, nf, ks=1, norm=None, act=act, act_kwargs=act_kwargs),
ConvBlock(nf, ni, ks=1, norm=None, act=nn.Sigmoid), ProdLayer())
t = torch.rand(8, 32, 12)
test_eq(SEModule1d(t.shape[1], 16, act=nn.ReLU, act_kwargs={})(t).shape, t.shape)
#export
def Norm(nf, ndim=1, norm='Batch', zero_norm=False, init=True, **kwargs):
"Norm layer with `nf` features and `ndim` with auto init."
assert 1 <= ndim <= 3
nl = getattr(nn, f"{snake2camel(norm)}Norm{ndim}d")(nf, **kwargs)
if nl.affine and init:
nl.bias.data.fill_(1e-3)
nl.weight.data.fill_(0. if zero_norm else 1.)
return nl
BN1d = partial(Norm, ndim=1, norm='Batch')
IN1d = partial(Norm, ndim=1, norm='Instance')
bs = 2
ni = 3
nf = 5
sl = 4
ks = 5
t = torch.rand(bs, ni, sl)
test_eq(ConvBlock(ni, nf, ks)(t).shape, (bs, nf, sl))
test_eq(ConvBlock(ni, nf, ks, padding='causal')(t).shape, (bs, nf, sl))
test_eq(ConvBlock(ni, nf, ks, coord=True)(t).shape, (bs, nf, sl))
ConvBlock(ni, nf, ks, stride=2)(t).shape
test_eq(ConvBlock(ni, nf, ks, stride=2)(t).shape, (bs, nf, sl//2))
test_eq(BN1d(ni)(t).shape, (bs, ni, sl))
test_eq(BN1d(ni).weight.data.mean().item(), 1.)
test_eq(BN1d(ni, zero_norm=True).weight.data.mean().item(), 0.)
test_eq(ConvBlock(ni, nf, ks, norm='batch', zero_norm=True)[1].weight.data.unique().item(), 0)
test_ne(ConvBlock(ni, nf, ks, norm='batch', zero_norm=False)[1].weight.data.unique().item(), 0)
test_eq(ConvBlock(ni, nf, ks, bias=False)[0].bias, None)
ConvBlock(ni, nf, ks, act=Swish, coord=True)
#export
class LinLnDrop(nn.Sequential):
"Module grouping `LayerNorm1d`, `Dropout` and `Linear` layers"
def __init__(self, n_in, n_out, ln=True, p=0., act=None, lin_first=False):
layers = [nn.LayerNorm(n_out if lin_first else n_in)] if ln else []
if p != 0: layers.append(nn.Dropout(p))
lin = [nn.Linear(n_in, n_out, bias=not ln)]
if act is not None: lin.append(act)
layers = lin+layers if lin_first else layers+lin
super().__init__(*layers)
LinLnDrop(2, 3, p=.5)
#export
class LambdaPlus(Module):
def __init__(self, func, *args, **kwargs): self.func,self.args,self.kwargs=func,args,kwargs
def forward(self, x): return self.func(x, *self.args, **self.kwargs)
#export
class Squeeze(Module):
def __init__(self, dim=-1): self.dim = dim
def forward(self, x): return x.squeeze(dim=self.dim)
def __repr__(self): return f'{self.__class__.__name__}(dim={self.dim})'
class Unsqueeze(Module):
def __init__(self, dim=-1): self.dim = dim
def forward(self, x): return x.unsqueeze(dim=self.dim)
def __repr__(self): return f'{self.__class__.__name__}(dim={self.dim})'
class Add(Module):
def forward(self, x, y): return x.add(y)
def __repr__(self): return f'{self.__class__.__name__}'
class Concat(Module):
def __init__(self, dim=1): self.dim = dim
def forward(self, *x): return torch.cat(*x, dim=self.dim)
def __repr__(self): return f'{self.__class__.__name__}(dim={self.dim})'
class Permute(Module):
def __init__(self, *dims): self.dims = dims
def forward(self, x): return x.permute(self.dims)
def __repr__(self): return f"{self.__class__.__name__}(dims={', '.join([str(d) for d in self.dims])})"
class Transpose(Module):
def __init__(self, *dims, contiguous=False): self.dims, self.contiguous = dims, contiguous
def forward(self, x):
if self.contiguous: return x.transpose(*self.dims).contiguous()
else: return x.transpose(*self.dims)
def __repr__(self):
if self.contiguous: return f"{self.__class__.__name__}(dims={', '.join([str(d) for d in self.dims])}).contiguous()"
else: return f"{self.__class__.__name__}({', '.join([str(d) for d in self.dims])})"
class View(Module):
def __init__(self, *shape): self.shape = shape
def forward(self, x): return x.view(x.shape[0], *self.shape)
def __repr__(self): return f"{self.__class__.__name__}({', '.join(['bs'] + [str(s) for s in self.shape])})"
class Reshape(Module):
def __init__(self, *shape): self.shape = shape
def forward(self, x): return x.reshape(x.shape[0], *self.shape)
def __repr__(self): return f"{self.__class__.__name__}({', '.join(['bs'] + [str(s) for s in self.shape])})"
class Max(Module):
def __init__(self, dim=None, keepdim=False): self.dim, self.keepdim = dim, keepdim
def forward(self, x): return x.max(self.dim, keepdim=self.keepdim)[0]
def __repr__(self): return f'{self.__class__.__name__}(dim={self.dim}, keepdim={self.keepdim})'
class LastStep(Module):
def forward(self, x): return x[..., -1]
def __repr__(self): return f'{self.__class__.__name__}()'
class SoftMax(Module):
"SoftMax layer"
def __init__(self, dim=-1):
self.dim = dim
def forward(self, x):
return F.softmax(x, dim=self.dim)
def __repr__(self): return f'{self.__class__.__name__}(dim={self.dim})'
class Clamp(Module):
def __init__(self, min=None, max=None):
self.min, self.max = min, max
def forward(self, x):
return x.clamp(min=self.min, max=self.max)
def __repr__(self): return f'{self.__class__.__name__}(min={self.min}, max={self.max})'
class Clip(Module):
def __init__(self, min=None, max=None):
self.min, self.max = min, max
def forward(self, x):
if self.min is not None:
x = torch.maximum(x, self.min)
if self.max is not None:
x = torch.minimum(x, self.max)
return x
def __repr__(self): return f'{self.__class__.__name__}()'
Noop = nn.Sequential()
bs = 2
nf = 5
sl = 4
t = torch.rand(bs, nf, sl)
test_eq(Permute(0,2,1)(t).shape, (bs, sl, nf))
test_eq(Max(1)(t).shape, (bs, sl))
test_eq(Transpose(1,2)(t).shape, (bs, sl, nf))
test_eq(Transpose(1,2, contiguous=True)(t).shape, (bs, sl, nf))
test_eq(View(-1, 2, 10)(t).shape, (bs, 1, 2, 10))
test_eq(Reshape(-1, 2, 10)(t).shape, (bs, 1, 2, 10))
Transpose(1,2), Permute(0,2,1), View(-1, 2, 10), Transpose(1,2, contiguous=True), Reshape(-1, 2, 10), Noop
# export
class DropPath(nn.Module):
"""Drop paths (Stochastic Depth) per sample (when applied in main path of residual blocks).
It's similar to Dropout but it drops individual connections instead of nodes.
Original code in https://github.com/rwightman/pytorch-image-models (timm library)
"""
def __init__(self, p=None):
super().__init__()
self.p = p
def forward(self, x):
if self.p == 0. or not self.training: return x
keep_prob = 1 - self.p
shape = (x.shape[0],) + (1,) * (x.ndim - 1)
random_tensor = keep_prob + torch.rand(shape, dtype=x.dtype, device=x.device)
random_tensor.floor_()
output = x.div(keep_prob) * random_tensor
# output = x.div(random_tensor.mean()) * random_tensor # divide by the actual mean to mantain the input mean?
return output
t = torch.ones(100,2,3)
test_eq(DropPath(0.)(t), t)
assert DropPath(0.5)(t).max() >= 1
#export
class Sharpen(Module):
"This is used to increase confidence in predictions - MixMatch paper"
def __init__(self, T=.5): self.T = T
def forward(self, x):
x = x**(1. / self.T)
return x / x.sum(dim=1, keepdims=True)
n_samples = 1000
n_classes = 3
t = (torch.rand(n_samples, n_classes) - .5) * 10
probas = F.softmax(t, -1)
sharpened_probas = Sharpen()(probas)
plt.plot(probas.flatten().sort().values, color='r')
plt.plot(sharpened_probas.flatten().sort().values, color='b')
plt.show()
test_gt(sharpened_probas[n_samples//2:].max(-1).values.sum().item(), probas[n_samples//2:].max(-1).values.sum().item())
#export
class Sequential(nn.Sequential):
"""Class that allows you to pass one or multiple inputs"""
def forward(self, *x):
for i, module in enumerate(self._modules.values()):
x = module(*x) if isinstance(x, (list, tuple, L)) else module(x)
return x
#export
class TimeDistributed(nn.Module):
def __init__(self, module, batch_first=False):
super(TimeDistributed, self).__init__()
self.module = module
self.batch_first = batch_first
def forward(self, x):
if len(x.size()) <= 2:
return self.module(x)
# Squash samples and timesteps into a single axis
x_reshape = x.contiguous().view(-1, x.size(-1)) # (samples * timesteps, input_size)
y = self.module(x_reshape)
# We have to reshape Y
if self.batch_first:
y = y.contiguous().view(x.size(0), -1, y.size(-1)) # (samples, timesteps, output_size)
else:
y = y.view(-1, x.size(1), y.size(-1)) # (timesteps, samples, output_size)
return y
#export
class Temp_Scale(Module):
"Used to perform Temperature Scaling (dirichlet=False) or Single-parameter Dirichlet calibration (dirichlet=True)"
def __init__(self, temp=1., dirichlet=False):
self.weight = nn.Parameter(tensor(temp))
self.bias = None
self.log_softmax = dirichlet
def forward(self, x):
if self.log_softmax: x = F.log_softmax(x, dim=-1)
return x.div(self.weight)
class Vector_Scale(Module):
"Used to perform Vector Scaling (dirichlet=False) or Diagonal Dirichlet calibration (dirichlet=True)"
def __init__(self, n_classes=1, dirichlet=False):
self.weight = nn.Parameter(torch.ones(n_classes))
self.bias = nn.Parameter(torch.zeros(n_classes))
self.log_softmax = dirichlet
def forward(self, x):
if self.log_softmax: x = F.log_softmax(x, dim=-1)
return x.mul(self.weight).add(self.bias)
class Matrix_Scale(Module):
"Used to perform Matrix Scaling (dirichlet=False) or Dirichlet calibration (dirichlet=True)"
def __init__(self, n_classes=1, dirichlet=False):
self.ms = nn.Linear(n_classes, n_classes)
self.ms.weight.data = nn.Parameter(torch.eye(n_classes))
nn.init.constant_(self.ms.bias.data, 0.)
self.weight = self.ms.weight
self.bias = self.ms.bias
self.log_softmax = dirichlet
def forward(self, x):
if self.log_softmax: x = F.log_softmax(x, dim=-1)
return self.ms(x)
def get_calibrator(calibrator=None, n_classes=1, **kwargs):
if calibrator is None or not calibrator: return noop
elif calibrator.lower() == 'temp': return Temp_Scale(dirichlet=False, **kwargs)
elif calibrator.lower() == 'vector': return Vector_Scale(n_classes=n_classes, dirichlet=False, **kwargs)
elif calibrator.lower() == 'matrix': return Matrix_Scale(n_classes=n_classes, dirichlet=False, **kwargs)
elif calibrator.lower() == 'dtemp': return Temp_Scale(dirichlet=True, **kwargs)
elif calibrator.lower() == 'dvector': return Vector_Scale(n_classes=n_classes, dirichlet=True, **kwargs)
elif calibrator.lower() == 'dmatrix': return Matrix_Scale(n_classes=n_classes, dirichlet=True, **kwargs)
else: assert False, f'please, select a correct calibrator instead of {calibrator}'
bs = 2
c_out = 3
t = torch.rand(bs, c_out)
for calibrator, cal_name in zip(['temp', 'vector', 'matrix'], ['Temp_Scale', 'Vector_Scale', 'Matrix_Scale']):
cal = get_calibrator(calibrator, n_classes=c_out)
# print(calibrator)
# print(cal.weight, cal.bias, '\n')
test_eq(cal(t), t)
test_eq(cal.__class__.__name__, cal_name)
for calibrator, cal_name in zip(['dtemp', 'dvector', 'dmatrix'], ['Temp_Scale', 'Vector_Scale', 'Matrix_Scale']):
cal = get_calibrator(calibrator, n_classes=c_out)
# print(calibrator)
# print(cal.weight, cal.bias, '\n')
test_eq(cal(t), F.log_softmax(t, dim=1))
test_eq(cal.__class__.__name__, cal_name)
bs = 2
c_out = 3
t = torch.rand(bs, c_out)
test_eq(Temp_Scale()(t).shape, t.shape)
test_eq(Vector_Scale(c_out)(t).shape, t.shape)
test_eq(Matrix_Scale(c_out)(t).shape, t.shape)
test_eq(Temp_Scale(dirichlet=True)(t).shape, t.shape)
test_eq(Vector_Scale(c_out, dirichlet=True)(t).shape, t.shape)
test_eq(Matrix_Scale(c_out, dirichlet=True)(t).shape, t.shape)
test_eq(Temp_Scale()(t), t)
test_eq(Vector_Scale(c_out)(t), t)
test_eq(Matrix_Scale(c_out)(t), t)
bs = 2
c_out = 5
t = torch.rand(bs, c_out)
test_eq(Vector_Scale(c_out)(t), t)
test_eq(Vector_Scale(c_out).weight.data, torch.ones(c_out))
test_eq(Vector_Scale(c_out).weight.requires_grad, True)
test_eq(type(Vector_Scale(c_out).weight), torch.nn.parameter.Parameter)
bs = 2
c_out = 3
weight = 2
bias = 1
t = torch.rand(bs, c_out)
test_eq(Matrix_Scale(c_out)(t).shape, t.shape)
test_eq(Matrix_Scale(c_out).weight.requires_grad, True)
test_eq(type(Matrix_Scale(c_out).weight), torch.nn.parameter.Parameter)
#export
class LogitAdjustmentLayer(Module):
"Logit Adjustment for imbalanced datasets"
def __init__(self, class_priors):
self.class_priors = class_priors
def forward(self, x):
return x.add(self.class_priors)
LogitAdjLayer = LogitAdjustmentLayer
bs, n_classes = 16, 3
class_priors = torch.rand(n_classes)
logits = torch.randn(bs, n_classes) * 2
test_eq(LogitAdjLayer(class_priors)(logits), logits + class_priors)
#export
class PPV(Module):
def __init__(self, dim=-1):
self.dim = dim
def forward(self, x):
return torch.gt(x, 0).sum(dim=self.dim).float() / x.shape[self.dim]
def __repr__(self): return f'{self.__class__.__name__}(dim={self.dim})'
class PPAuc(Module):
def __init__(self, dim=-1):
self.dim = dim
def forward(self, x):
x = F.relu(x).sum(self.dim) / (abs(x).sum(self.dim) + 1e-8)
return x
def __repr__(self): return f'{self.__class__.__name__}(dim={self.dim})'
class MaxPPVPool1d(Module):
"Drop-in replacement for AdaptiveConcatPool1d - multiplies nf by 2"
def forward(self, x):
_max = x.max(dim=-1).values
_ppv = torch.gt(x, 0).sum(dim=-1).float() / x.shape[-1]
return torch.cat((_max, _ppv), dim=-1).unsqueeze(2)
bs = 2
nf = 5
sl = 4
t = torch.rand(bs, nf, sl)
test_eq(MaxPPVPool1d()(t).shape, (bs, nf*2, 1))
test_eq(MaxPPVPool1d()(t).shape, AdaptiveConcatPool1d(1)(t).shape)
#export
class AdaptiveWeightedAvgPool1d(Module):
'''Global Pooling layer that performs a weighted average along the temporal axis
It can be considered as a channel-wise form of local temporal attention. Inspired by the paper:
Hyun, J., Seong, H., & Kim, E. (2019). Universal Pooling--A New Pooling Method for Convolutional Neural Networks. arXiv preprint arXiv:1907.11440.'''
def __init__(self, n_in, seq_len, mult=2, n_layers=2, ln=False, dropout=0.5, act=nn.ReLU(), zero_init=True):
layers = nn.ModuleList()
for i in range(n_layers):
inp_mult = mult if i > 0 else 1
out_mult = mult if i < n_layers -1 else 1
p = dropout[i] if is_listy(dropout) else dropout
layers.append(LinLnDrop(seq_len * inp_mult, seq_len * out_mult, ln=False, p=p,
act=act if i < n_layers-1 and n_layers > 1 else None))
self.layers = layers
self.softmax = SoftMax(-1)
if zero_init: init_lin_zero(self)
def forward(self, x):
wap = x
for l in self.layers: wap = l(wap)
wap = self.softmax(wap)
return torch.mul(x, wap).sum(-1)
#export
class GAP1d(Module):
"Global Adaptive Pooling + Flatten"
def __init__(self, output_size=1):
self.gap = nn.AdaptiveAvgPool1d(output_size)
self.flatten = Flatten()
def forward(self, x):
return self.flatten(self.gap(x))
class GACP1d(Module):
"Global AdaptiveConcatPool + Flatten"
def __init__(self, output_size=1):
self.gacp = AdaptiveConcatPool1d(output_size)
self.flatten = Flatten()
def forward(self, x):
return self.flatten(self.gacp(x))
class GAWP1d(Module):
"Global AdaptiveWeightedAvgPool1d + Flatten"
def __init__(self, n_in, seq_len, n_layers=2, ln=False, dropout=0.5, act=nn.ReLU(), zero_init=False):
self.gacp = AdaptiveWeightedAvgPool1d(n_in, seq_len, n_layers=n_layers, ln=ln, dropout=dropout, act=act, zero_init=zero_init)
self.flatten = Flatten()
def forward(self, x):
return self.flatten(self.gacp(x))
# export
class GlobalWeightedAveragePool1d(Module):
""" Global Weighted Average Pooling layer
Inspired by Building Efficient CNN Architecture for Offline Handwritten Chinese Character Recognition
https://arxiv.org/pdf/1804.01259.pdf
"""
def __init__(self, n_in, seq_len):
self.weight = nn.Parameter(torch.ones(1, n_in, seq_len))
self.bias = nn.Parameter(torch.zeros(1, n_in, seq_len))
def forward(self, x):
α = F.softmax(torch.sigmoid(x * self.weight + self.bias), dim=-1)
return (x * α).sum(-1)
GWAP1d = GlobalWeightedAveragePool1d
def gwa_pool_head(n_in, c_out, seq_len, bn=True, fc_dropout=0.):
return nn.Sequential(GlobalWeightedAveragePool1d(n_in, seq_len), Flatten(), LinBnDrop(n_in, c_out, p=fc_dropout, bn=bn))
t = torch.randn(16, 64, 50)
head = gwa_pool_head(64, 5, 50)
test_eq(head(t).shape, (16, 5))
#export
class AttentionalPool1d(Module):
"""Global Adaptive Pooling layer inspired by Attentional Pooling for Action Recognition https://arxiv.org/abs/1711.01467"""
def __init__(self, n_in, c_out, bn=False):
store_attr()
self.bn = nn.BatchNorm1d(n_in) if bn else None
self.conv1 = Conv1d(n_in, 1, 1)
self.conv2 = Conv1d(n_in, c_out, 1)
def forward(self, x):
if self.bn is not None: x = self.bn(x)
return (self.conv1(x) @ self.conv2(x).transpose(1,2)).transpose(1,2)
class GAttP1d(nn.Sequential):
def __init__(self, n_in, c_out, bn=False):
super().__init__(AttentionalPool1d(n_in, c_out, bn=bn), Flatten())
def attentional_pool_head(n_in, c_out, seq_len=None, bn=True, **kwargs):
return nn.Sequential(AttentionalPool1d(n_in, c_out, bn=bn, **kwargs), Flatten())
bs, c_in, seq_len = 16, 1, 50
c_out = 3
t = torch.rand(bs, c_in, seq_len)
test_eq(GAP1d()(t).shape, (bs, c_in))
test_eq(GACP1d()(t).shape, (bs, c_in*2))
bs, c_in, seq_len = 16, 4, 50
t = torch.rand(bs, c_in, seq_len)
test_eq(GAP1d()(t).shape, (bs, c_in))
test_eq(GACP1d()(t).shape, (bs, c_in*2))
test_eq(GAWP1d(c_in, seq_len, n_layers=2, ln=False, dropout=0.5, act=nn.ReLU(), zero_init=False)(t).shape, (bs, c_in))
test_eq(GAWP1d(c_in, seq_len, n_layers=2, ln=False, dropout=0.5, act=nn.ReLU(), zero_init=False)(t).shape, (bs, c_in))
test_eq(GAWP1d(c_in, seq_len, n_layers=1, ln=False, dropout=0.5, zero_init=False)(t).shape, (bs, c_in))
test_eq(GAWP1d(c_in, seq_len, n_layers=1, ln=False, dropout=0.5, zero_init=True)(t).shape, (bs, c_in))
test_eq(AttentionalPool1d(c_in, c_out)(t).shape, (bs, c_out, 1))
bs, c_in, seq_len = 16, 128, 50
c_out = 14
t = torch.rand(bs, c_in, seq_len)
attp = attentional_pool_head(c_in, c_out)
test_eq(attp(t).shape, (bs, c_out))
#export
def create_pool_head(n_in, c_out, seq_len=None, concat_pool=False, fc_dropout=0., bn=False, y_range=None, **kwargs):
if kwargs: print(f'{kwargs} not being used')
if concat_pool: n_in*=2
layers = [GACP1d(1) if concat_pool else GAP1d(1)]
layers += [LinBnDrop(n_in, c_out, bn=bn, p=fc_dropout)]
if y_range: layers += [SigmoidRange(*y_range)]
return nn.Sequential(*layers)
pool_head = create_pool_head
average_pool_head = partial(pool_head, concat_pool=False)
setattr(average_pool_head, "__name__", "average_pool_head")
concat_pool_head = partial(pool_head, concat_pool=True)
setattr(concat_pool_head, "__name__", "concat_pool_head")
bs = 16
nf = 12
c_out = 2
seq_len = 20
t = torch.rand(bs, nf, seq_len)
test_eq(create_pool_head(nf, c_out, seq_len, fc_dropout=0.5)(t).shape, (bs, c_out))
test_eq(create_pool_head(nf, c_out, seq_len, concat_pool=True, fc_dropout=0.5)(t).shape, (bs, c_out))
create_pool_head(nf, c_out, seq_len, concat_pool=True, bn=True, fc_dropout=.5)
#export
def max_pool_head(n_in, c_out, seq_len, fc_dropout=0., bn=False, y_range=None, **kwargs):
if kwargs: print(f'{kwargs} not being used')
layers = [nn.MaxPool1d(seq_len, **kwargs), Flatten()]
layers += [LinBnDrop(n_in, c_out, bn=bn, p=fc_dropout)]
if y_range: layers += [SigmoidRange(*y_range)]
return nn.Sequential(*layers)
bs = 16
nf = 12
c_out = 2
seq_len = 20
t = torch.rand(bs, nf, seq_len)
test_eq(max_pool_head(nf, c_out, seq_len, fc_dropout=0.5)(t).shape, (bs, c_out))
#export
def create_pool_plus_head(*args, lin_ftrs=None, fc_dropout=0., concat_pool=True, bn_final=False, lin_first=False, y_range=None):
nf = args[0]
c_out = args[1]
if concat_pool: nf = nf * 2
lin_ftrs = [nf, 512, c_out] if lin_ftrs is None else [nf] + lin_ftrs + [c_out]
ps = L(fc_dropout)
if len(ps) == 1: ps = [ps[0]/2] * (len(lin_ftrs)-2) + ps
actns = [nn.ReLU(inplace=True)] * (len(lin_ftrs)-2) + [None]
pool = AdaptiveConcatPool1d() if concat_pool else nn.AdaptiveAvgPool1d(1)
layers = [pool, Flatten()]
if lin_first: layers.append(nn.Dropout(ps.pop(0)))
for ni,no,p,actn in zip(lin_ftrs[:-1], lin_ftrs[1:], ps, actns):
layers += LinBnDrop(ni, no, bn=True, p=p, act=actn, lin_first=lin_first)
if lin_first: layers.append(nn.Linear(lin_ftrs[-2], c_out))
if bn_final: layers.append(nn.BatchNorm1d(lin_ftrs[-1], momentum=0.01))
if y_range is not None: layers.append(SigmoidRange(*y_range))
return nn.Sequential(*layers)
pool_plus_head = create_pool_plus_head
bs = 16
nf = 12
c_out = 2
seq_len = 20
t = torch.rand(bs, nf, seq_len)
test_eq(create_pool_plus_head(nf, c_out, seq_len, fc_dropout=0.5)(t).shape, (bs, c_out))
test_eq(create_pool_plus_head(nf, c_out, concat_pool=True, fc_dropout=0.5)(t).shape, (bs, c_out))
create_pool_plus_head(nf, c_out, seq_len, fc_dropout=0.5)
#export
def create_conv_head(*args, adaptive_size=None, y_range=None):
nf = args[0]
c_out = args[1]
layers = [nn.AdaptiveAvgPool1d(adaptive_size)] if adaptive_size is not None else []
for i in range(2):
if nf > 1:
layers += [ConvBlock(nf, nf // 2, 1)]
nf = nf//2
else: break
layers += [ConvBlock(nf, c_out, 1), GAP1d(1)]
if y_range: layers += [SigmoidRange(*y_range)]
return nn.Sequential(*layers)
conv_head = create_conv_head
bs = 16
nf = 12
c_out = 2
seq_len = 20
t = torch.rand(bs, nf, seq_len)
test_eq(create_conv_head(nf, c_out, seq_len)(t).shape, (bs, c_out))
test_eq(create_conv_head(nf, c_out, adaptive_size=50)(t).shape, (bs, c_out))
create_conv_head(nf, c_out, 50)
#export
def create_mlp_head(nf, c_out, seq_len=None, flatten=True, fc_dropout=0., bn=False, y_range=None):
if flatten: nf *= seq_len
layers = [Flatten()] if flatten else []
layers += [LinBnDrop(nf, c_out, bn=bn, p=fc_dropout)]
if y_range: layers += [SigmoidRange(*y_range)]
return nn.Sequential(*layers)
mlp_head = create_mlp_head
bs = 16
nf = 12
c_out = 2
seq_len = 20
t = torch.rand(bs, nf, seq_len)
test_eq(create_mlp_head(nf, c_out, seq_len, fc_dropout=0.5)(t).shape, (bs, c_out))
t = torch.rand(bs, nf, seq_len)
create_mlp_head(nf, c_out, seq_len, bn=True, fc_dropout=.5)
#export
def create_fc_head(nf, c_out, seq_len=None, flatten=True, lin_ftrs=None, y_range=None, fc_dropout=0., bn=False, bn_final=False, act=nn.ReLU(inplace=True)):
if flatten: nf *= seq_len
layers = [Flatten()] if flatten else []
lin_ftrs = [nf, 512, c_out] if lin_ftrs is None else [nf] + lin_ftrs + [c_out]
if not is_listy(fc_dropout): fc_dropout = [fc_dropout]*(len(lin_ftrs) - 1)
actns = [act for _ in range(len(lin_ftrs) - 2)] + [None]
layers += [LinBnDrop(lin_ftrs[i], lin_ftrs[i+1], bn=bn and (i!=len(actns)-1 or bn_final), p=p, act=a) for i,(p,a) in enumerate(zip(fc_dropout+[0.], actns))]
if y_range is not None: layers.append(SigmoidRange(*y_range))
return nn.Sequential(*layers)
fc_head = create_fc_head
bs = 16
nf = 12
c_out = 2
seq_len = 20
t = torch.rand(bs, nf, seq_len)
test_eq(create_fc_head(nf, c_out, seq_len, fc_dropout=0.5)(t).shape, (bs, c_out))
create_mlp_head(nf, c_out, seq_len, bn=True, fc_dropout=.5)
#export
def create_rnn_head(*args, fc_dropout=0., bn=False, y_range=None):
nf = args[0]
c_out = args[1]
layers = [LastStep()]
layers += [LinBnDrop(nf, c_out, bn=bn, p=fc_dropout)]
if y_range: layers += [SigmoidRange(*y_range)]
return nn.Sequential(*layers)
rnn_head = create_rnn_head
bs = 16
nf = 12
c_out = 2
seq_len = 20
t = torch.rand(bs, nf, seq_len)
test_eq(create_rnn_head(nf, c_out, seq_len, fc_dropout=0.5)(t).shape, (bs, c_out))
create_rnn_head(nf, c_out, seq_len, bn=True, fc_dropout=.5)
# export
def imputation_head(c_in, c_out, seq_len=None, ks=1, y_range=None, fc_dropout=0.):
layers = [nn.Dropout(fc_dropout), nn.Conv1d(c_in, c_out, ks)]
if y_range is not None:
y_range = (tensor(y_range[0]), tensor(y_range[1]))
layers += [SigmoidRange(*y_range)]
return nn.Sequential(*layers)
bs = 16
nf = 12
ni = 2
seq_len = 20
t = torch.rand(bs, nf, seq_len)
head = imputation_head(nf, ni, seq_len=None, ks=1, y_range=None, fc_dropout=0.)
test_eq(head(t).shape, (bs, ni, seq_len))
head = imputation_head(nf, ni, seq_len=None, ks=1, y_range=(.3,.7), fc_dropout=0.)
test_ge(head(t).min(), .3)
test_le(head(t).max(), .7)
y_range = (tensor([0.1000, 0.1000, 0.1000, 0.1000, 0.2000, 0.2000, 0.2000, 0.2000, 0.3000,
0.3000, 0.3000, 0.3000]),
tensor([0.6000, 0.6000, 0.6000, 0.6000, 0.7000, 0.7000, 0.7000, 0.7000, 0.8000,
0.8000, 0.8000, 0.8000]))
test_ge(head(t).min(), .1)
test_le(head(t).max(), .9)
head = imputation_head(nf, ni, seq_len=None, ks=1, y_range=y_range, fc_dropout=0.)
head
# export
class create_conv_lin_3d_head(nn.Sequential):
"Module to create a 3d output head"
def __init__(self, n_in, n_out, seq_len, d=(), conv_first=True, conv_bn=True, lin_first=False, lin_bn=True, act=None, fc_dropout=0., **kwargs):
assert len(d) == 2, "you must pass a tuple of len == 2 to create a 3d output"
conv = [BatchNorm(n_in, ndim=1)] if conv_bn else []
conv.append(Conv1d(n_in, d[0], 1, padding=0, bias=not conv_bn, **kwargs))
l = [Transpose(-1, -2), BatchNorm(n_out if lin_first else seq_len, ndim=1), Transpose(-1, -2)] if lin_bn else []
if fc_dropout != 0: l.append(nn.Dropout(fc_dropout))
lin = [nn.Linear(seq_len, d[1], bias=not lin_bn)]
if act is not None: lin.append(act)
lin_layers = lin+l if lin_first else l+lin
layers = conv + lin_layers if conv_first else lin_layers + conv
super().__init__(*layers)
conv_lin_3d_head = create_conv_lin_3d_head
t = torch.randn(16, 3, 50)
head = conv_lin_3d_head(3, 20, 50, (4,5))
test_eq(head(t).shape, (16, 4, 5))
head = conv_lin_3d_head(3, 20, 50, (2, 10))
test_eq(head(t).shape, (16, 2, 10))
head
# export
class create_lin_3d_head(nn.Sequential):
"Module to create a 3d output head with linear layers"
def __init__(self, n_in, n_out, seq_len, d=(), lin_first=False, bn=True, act=None, fc_dropout=0.):
assert len(d) == 2, "you must pass a tuple of len == 2 to create a 3d output"
layers = [Flatten()]
layers += LinBnDrop(n_in * seq_len, n_out, bn=bn, p=fc_dropout, act=act, lin_first=lin_first)
layers += [Reshape(*d)]
super().__init__(*layers)
lin_3d_head = create_lin_3d_head
t = torch.randn(16, 64, 50)
head = lin_3d_head(64, 10, 50, (5,2))
test_eq(head(t).shape, (16, 5, 2))
head = lin_3d_head(64, 5, 50, (5, 1))
test_eq(head(t).shape, (16, 5, 1))
head
# export
class create_conv_3d_head(nn.Sequential):
"Module to create a 3d output head with a convolutional layer"
def __init__(self, n_in, c_out, seq_len, d=(), lin_first=False, bn=True, act=None, fc_dropout=0.):
assert len(d) == 2, "you must pass a tuple of len == 2 to create a 3d output"
assert d[1] == seq_len, 'You can only use this head when learn.dls.len == learn.dls.d'
super().__init__(Conv(n_in, d[0], 1))
conv_3d_head = create_conv_3d_head
bs = 16
c_out = 4
seq_len = 50
d = (2,50)
nf = 128
t = torch.rand(bs, nf, seq_len)
test_eq(conv_3d_head(nf, c_out, seq_len, d)(t).shape, (bs, *d))
#export
def universal_pool_head(n_in, c_out, seq_len, mult=2, pool_n_layers=2, pool_ln=True, pool_dropout=0.5, pool_act=nn.ReLU(),
zero_init=True, bn=True, fc_dropout=0.):
return nn.Sequential(AdaptiveWeightedAvgPool1d(n_in, seq_len, n_layers=pool_n_layers, mult=mult, ln=pool_ln, dropout=pool_dropout, act=pool_act),
Flatten(), LinBnDrop(n_in, c_out, p=fc_dropout, bn=bn))
bs, c_in, seq_len = 16, 128, 50
c_out = 14
t = torch.rand(bs, c_in, seq_len)
uph = universal_pool_head(c_in, c_out, seq_len)
test_eq(uph(t).shape, (bs, c_out))
uph = universal_pool_head(c_in, c_out, seq_len, 2)
test_eq(uph(t).shape, (bs, c_out))
#export
heads = [mlp_head, fc_head, average_pool_head, max_pool_head, concat_pool_head, pool_plus_head, conv_head, rnn_head,
conv_lin_3d_head, lin_3d_head, conv_3d_head, attentional_pool_head, universal_pool_head, gwa_pool_head]
bs, c_in, seq_len = 16, 128, 50
c_out = 14
d = (7, 2)
t = torch.rand(bs, c_in, seq_len)
for head in heads:
print(head.__name__)
if head.__name__ == 'create_conv_3d_head':
test_eq(head(c_in, c_out, seq_len, (d[0], seq_len))(t).shape, (bs, *(d[0], seq_len)))
elif '3d' in head.__name__:
test_eq(head(c_in, c_out, seq_len, d)(t).shape, (bs, *d))
else:
test_eq(head(c_in, c_out, seq_len)(t).shape, (bs, c_out))
#export
class SqueezeExciteBlock(Module):
def __init__(self, ni, reduction=16):
self.avg_pool = GAP1d(1)
self.fc = nn.Sequential(nn.Linear(ni, ni // reduction, bias=False), nn.ReLU(), nn.Linear(ni // reduction, ni, bias=False), nn.Sigmoid())
def forward(self, x):
y = self.avg_pool(x)
y = self.fc(y).unsqueeze(2)
return x * y.expand_as(x)
bs = 2
ni = 32
sl = 4
t = torch.rand(bs, ni, sl)
test_eq(SqueezeExciteBlock(ni)(t).shape, (bs, ni, sl))
#export
class GaussianNoise(Module):
"""Gaussian noise regularizer.
Args:
sigma (float, optional): relative standard deviation used to generate the
noise. Relative means that it will be multiplied by the magnitude of
the value your are adding the noise to. This means that sigma can be
the same regardless of the scale of the vector.
is_relative_detach (bool, optional): whether to detach the variable before
computing the scale of the noise. If `False` then the scale of the noise
won't be seen as a constant but something to optimize: this will bias the
network to generate vectors with smaller values.
"""
def __init__(self, sigma=0.1, is_relative_detach=True):
self.sigma, self.is_relative_detach = sigma, is_relative_detach
def forward(self, x):
if self.training and self.sigma not in [0, None]:
scale = self.sigma * (x.detach() if self.is_relative_detach else x)
sampled_noise = torch.empty(x.size()).normal_().to(device) * scale
x = x + sampled_noise
return x
t = torch.ones(2,3,4)
test_ne(GaussianNoise()(t), t)
test_eq(GaussianNoise()(t).shape, t.shape)
t = torch.ones(2,3)
test_ne(GaussianNoise()(t), t)
test_eq(GaussianNoise()(t).shape, t.shape)
t = torch.ones(2)
test_ne(GaussianNoise()(t), t)
test_eq(GaussianNoise()(t).shape, t.shape)
#export
def gambler_loss(reward=2):
def _gambler_loss(model_output, targets):
outputs = torch.nn.functional.softmax(model_output, dim=1)
outputs, reservation = outputs[:, :-1], outputs[:, -1]
gain = torch.gather(outputs, dim=1, index=targets.unsqueeze(1)).squeeze()
doubling_rate = (gain + reservation / reward).log()
return - doubling_rate.mean()
return _gambler_loss
model_output = torch.rand(16, 3)
targets = torch.randint(0, 2, (16,))
criterion = gambler_loss(2)
criterion(model_output, targets)
#export
def CrossEntropyLossOneHot(output, target, **kwargs):
if target.ndim == 2: _, target = target.max(dim=1)
return nn.CrossEntropyLoss(**kwargs)(output, target)
output = torch.rand(16, 2)
target = torch.randint(0, 2, (16,))
CrossEntropyLossOneHot(output, target)
from tsai.data.transforms import OneHot
output = nn.Parameter(torch.rand(16, 2))
target = torch.randint(0, 2, (16,))
one_hot_target = OneHot()(target)
CrossEntropyLossOneHot(output, one_hot_target)
#hide
def proba_certainty(output):
if output.sum(-1).mean().item() != 1: output = F.softmax(output, -1)
return (output.max(-1).values - 1. / output.shape[-1])/( 1 - 1. / output.shape[-1])
#hide
target = random_shuffle(concat(torch.zeros(5), torch.ones(7), torch.ones(4) + 1)).long()
output = nn.Parameter(5 * torch.rand((16, 3)) - 5 * torch.rand((16, 3)))
proba_certainty(output)
#hide
def CrossEntropyLossOneHotWithUncertainty():
def _CrossEntropyLossOneHotWithUncertainty(output, target, **kwargs):
return (proba_certainty(output) * CrossEntropyLossOneHot(output, target, reduction='none', **kwargs)).mean()
return _CrossEntropyLossOneHotWithUncertainty
#hide
# https://stackoverflow.com/questions/22611446/perform-2-sample-t-test
from __future__ import print_function
import numpy as np
from scipy.stats import ttest_ind, ttest_ind_from_stats
from scipy.special import stdtr
np.random.seed(1)
# Create sample data.
a = np.random.randn(40)
b = 4*np.random.randn(50)
# Use scipy.stats.ttest_ind.
t, p = ttest_ind(a, b, equal_var=False)
print("ttest_ind: t = %g p = %g" % (t, p))
# Compute the descriptive statistics of a and b.
abar = a.mean()
avar = a.var(ddof=1)
na = a.size
adof = na - 1
bbar = b.mean()
bvar = b.var(ddof=1)
nb = b.size
bdof = nb - 1
# Use scipy.stats.ttest_ind_from_stats.
t2, p2 = ttest_ind_from_stats(abar, np.sqrt(avar), na,
bbar, np.sqrt(bvar), nb,
equal_var=False)
print("ttest_ind_from_stats: t = %g p = %g" % (t2, p2))
# Use the formulas directly.
tf = (abar - bbar) / np.sqrt(avar/na + bvar/nb)
dof = (avar/na + bvar/nb)**2 / (avar**2/(na**2*adof) + bvar**2/(nb**2*bdof))
pf = 2*stdtr(dof, -np.abs(tf))
print("formula: t = %g p = %g" % (tf, pf))
a = tensor(a)
b = tensor(b)
tf = (a.mean() - b.mean()) / torch.sqrt(a.var()/a.size(0) + b.var()/b.size(0))
print("formula: t = %g" % (tf))
ttest_tensor(a, b)
#export
def ttest_bin_loss(output, target):
output = nn.Softmax(dim=-1)(output[:, 1])
return ttest_tensor(output[target == 0], output[target == 1])
def ttest_reg_loss(output, target):
return ttest_tensor(output[target <= 0], output[target > 0])
for _ in range(100):
output = torch.rand(256, 2)
target = torch.randint(0, 2, (256,))
test_close(ttest_bin_loss(output, target).item(),
ttest_ind(nn.Softmax(dim=-1)(output[:, 1])[target == 0], nn.Softmax(dim=-1)(output[:, 1])[target == 1], equal_var=False)[0], eps=1e-3)
#export
class CenterLoss(Module):
r"""
Code in Pytorch has been slightly modified from: https://github.com/KaiyangZhou/pytorch-center-loss/blob/master/center_loss.py
Based on paper: Wen et al. A Discriminative Feature Learning Approach for Deep Face Recognition. ECCV 2016.
Args:
c_out (int): number of classes.
logits_dim (int): dim 1 of the logits. By default same as c_out (for one hot encoded logits)
"""
def __init__(self, c_out, logits_dim=None):
logits_dim = ifnone(logits_dim, c_out)
self.c_out, self.logits_dim = c_out, logits_dim
self.centers = nn.Parameter(torch.randn(c_out, logits_dim).to(device=default_device()))
self.classes = torch.arange(c_out).long().to(device=default_device())
def forward(self, x, labels):
"""
Args:
x: feature matrix with shape (batch_size, logits_dim).
labels: ground truth labels with shape (batch_size).
"""
bs = x.shape[0]
distmat = torch.pow(x, 2).sum(dim=1, keepdim=True).expand(bs, self.c_out) + \
torch.pow(self.centers, 2).sum(dim=1, keepdim=True).expand(self.c_out, bs).T
distmat = torch.addmm(distmat, x, self.centers.T, beta=1, alpha=-2)
labels = labels.unsqueeze(1).expand(bs, self.c_out)
mask = labels.eq(self.classes.expand(bs, self.c_out))
dist = distmat * mask.float()
loss = dist.clamp(min=1e-12, max=1e+12).sum() / bs
return loss
class CenterPlusLoss(Module):
def __init__(self, loss, c_out, λ=1e-2, logits_dim=None):
self.loss, self.c_out, self.λ = loss, c_out, λ
self.centerloss = CenterLoss(c_out, logits_dim)
def forward(self, x, labels):
return self.loss(x, labels) + self.λ * self.centerloss(x, labels)
def __repr__(self): return f"CenterPlusLoss(loss={self.loss}, c_out={self.c_out}, λ={self.λ})"
c_in = 10
x = torch.rand(64, c_in).to(device=default_device())
x = F.softmax(x, dim=1)
label = x.max(dim=1).indices
CenterLoss(c_in)(x, label), CenterPlusLoss(LabelSmoothingCrossEntropyFlat(), c_in)(x, label)
CenterPlusLoss(LabelSmoothingCrossEntropyFlat(), c_in)
#export
class FocalLoss(Module):
def __init__(self, gamma=0, eps=1e-7):
self.gamma, self.eps, self.ce = gamma, eps, CrossEntropyLossFlat()
def forward(self, input, target):
logp = self.ce(input, target)
p = torch.exp(-logp)
loss = (1 - p) ** self.gamma * logp
return loss.mean()
c_in = 10
x = torch.rand(64, c_in).to(device=default_device())
x = F.softmax(x, dim=1)
label = x.max(dim=1).indices
FocalLoss(c_in)(x, label)
#export
class TweedieLoss(Module):
def __init__(self, p=1.5, eps=1e-10):
"""
Tweedie loss as calculated in LightGBM
Args:
p: tweedie variance power (1 < p < 2)
eps: small number to avoid log(zero).
"""
assert p > 1 and p < 2, "make sure 1 < p < 2"
self.p, self.eps = p, eps
def forward(self, inp, targ):
inp = inp.flatten()
targ = targ.flatten()
torch.clamp_min_(inp, self.eps)
a = targ * torch.exp((1 - self.p) * torch.log(inp)) / (1 - self.p)
b = torch.exp((2 - self.p) * torch.log(inp)) / (2 - self.p)
loss = -a + b
return loss.mean()
c_in = 10
output = torch.rand(64).to(device=default_device())
target = torch.rand(64).to(device=default_device())
TweedieLoss()(output, target)
# export
class GEGLU(Module):
def forward(self, x):
x, gates = x.chunk(2, dim=-1)
return x * F.gelu(gates)
class ReGLU(Module):
def forward(self, x):
x, gates = x.chunk(2, dim=-1)
return x * F.relu(gates)
class PositionwiseFeedForward(nn.Sequential):
def __init__(self, dim, dropout=0., act='reglu', mlp_ratio=1):
act = act.lower()
act_mult = 2 if act in ['geglu', 'reglu'] else 1
if act == 'relu': act_fn = nn.ReLU()
elif act == 'gelu': act_fn = nn.GELU()
elif act == 'geglu': act_fn = GEGLU()
else: act_fn = ReGLU()
super().__init__(nn.Linear(dim, dim * act_mult * mlp_ratio),
act_fn,
nn.Dropout(dropout),
nn.Linear(dim * mlp_ratio, dim),
nn.Dropout(dropout))
class TokenLayer(Module):
def __init__(self, token=True): self.token = token
def forward(self, x): return x[..., 0] if self.token is not None else x.mean(-1)
def __repr__(self): return f"{self.__class__.__name__}()"
#export
class ScaledDotProductAttention(Module):
"""Scaled Dot-Product Attention module (Vaswani et al., 2017) with optional residual attention from previous layer (He et al, 2020)"""
def __init__(self, res_attention:bool=False): self.res_attention = res_attention
def forward(self, q:Tensor, k:Tensor, v:Tensor, prev:Optional[Tensor]=None, key_padding_mask:Optional[Tensor]=None, attn_mask:Optional[Tensor]=None):
'''
Input shape:
q : [bs x n_heads x max_q_len x d_k]
k : [bs x n_heads x d_k x seq_len]
v : [bs x n_heads x seq_len x d_v]
prev : [bs x n_heads x q_len x seq_len]
key_padding_mask: [bs x seq_len]
attn_mask : [1 x seq_len x seq_len]
Output shape:
output: [bs x n_heads x q_len x d_v]
attn : [bs x n_heads x q_len x seq_len]
scores : [bs x n_heads x q_len x seq_len]
'''
# Scaled MatMul (q, k) - similarity scores for all pairs of positions in an input sequence
attn_scores = torch.matmul(q / np.sqrt(q.shape[-2]), k) # attn_scores : [bs x n_heads x max_q_len x q_len]
# Add pre-softmax attention scores from the previous layer (optional)
if prev is not None: attn_scores = attn_scores + prev
# Attention mask (optional)
if attn_mask is not None: # attn_mask with shape [q_len x seq_len] - only used when q_len == seq_len
if attn_mask.dtype == torch.bool:
attn_scores.masked_fill_(attn_mask, -np.inf)
else:
attn_scores += attn_mask
# Key padding mask (optional)
if key_padding_mask is not None: # mask with shape [q_len x q_len] (only when max_w_len == q_len)
attn_scores.masked_fill_(key_padding_mask.unsqueeze(1).unsqueeze(2), -np.inf)
# normalize the attention weights
attn_weights = F.softmax(attn_scores, dim=-1) # attn_weights : [bs x n_heads x max_q_len x q_len]
# compute the new values given the attention weights
output = torch.matmul(attn_weights, v) # output: [bs x n_heads x max_q_len x d_v]
if self.res_attention: return output, attn_weights, attn_scores
else: return output, attn_weights
B = 16
C = 10
M = 1500 # seq_len
n_heads = 1
D = 128 # model dimension
N = 512 # max_seq_len - latent's index dimension
d_k = D // n_heads
xb = torch.randn(B, C, M)
xb = (xb - xb.mean()) / xb.std()
# Attention
# input (Q)
lin = nn.Linear(M, N, bias=False)
Q = lin(xb).transpose(1,2)
test_eq(Q.shape, (B, N, C))
# q
to_q = nn.Linear(C, D, bias=False)
q = to_q(Q)
q = nn.LayerNorm(D)(q)
# k, v
context = xb.transpose(1,2)
to_kv = nn.Linear(C, D * 2, bias=False)
k, v = to_kv(context).chunk(2, dim = -1)
k = k.transpose(-1, -2)
k = nn.LayerNorm(M)(k)
v = nn.LayerNorm(D)(v)
test_eq(q.shape, (B, N, D))
test_eq(k.shape, (B, D, M))
test_eq(v.shape, (B, M, D))
output, attn, scores = ScaledDotProductAttention(res_attention=True)(q.unsqueeze(1), k.unsqueeze(1), v.unsqueeze(1))
test_eq(output.shape, (B, 1, N, D))
test_eq(attn.shape, (B, 1, N, M))
test_eq(scores.shape, (B, 1, N, M))
scores.mean(), scores.std()
#export
class MultiheadAttention(Module):
def __init__(self, d_model:int, n_heads:int, d_k:Optional[int]=None, d_v:Optional[int]=None, res_attention:bool=False,
dropout:float=0., qkv_bias:bool=True):
"""Multi Head Attention Layer
Input shape:
Q: [batch_size (bs) x max_q_len x d_model]
K, V: [batch_size (bs) x q_len x d_model]
mask: [q_len x q_len]
"""
d_k = ifnone(d_k, d_model // n_heads)
d_v = ifnone(d_v, d_model // n_heads)
self.n_heads, self.d_k, self.d_v = n_heads, d_k, d_v
self.W_Q = nn.Linear(d_model, d_k * n_heads, bias=qkv_bias)
self.W_K = nn.Linear(d_model, d_k * n_heads, bias=qkv_bias)
self.W_V = nn.Linear(d_model, d_v * n_heads, bias=qkv_bias)
# Scaled Dot-Product Attention (multiple heads)
self.res_attention = res_attention
self.sdp_attn = ScaledDotProductAttention(res_attention=self.res_attention)
# Poject output
project_out = not (n_heads == 1 and d_model == d_k)
self.to_out = nn.Sequential(nn.Linear(n_heads * d_v, d_model), nn.Dropout(dropout)) if project_out else nn.Identity()
def forward(self, Q:Tensor, K:Optional[Tensor]=None, V:Optional[Tensor]=None, prev:Optional[Tensor]=None,
key_padding_mask:Optional[Tensor]=None, attn_mask:Optional[Tensor]=None):
bs = Q.size(0)
if K is None: K = Q
if V is None: V = Q
# Linear (+ split in multiple heads)
q_s = self.W_Q(Q).view(bs, -1, self.n_heads, self.d_k).transpose(1,2) # q_s : [bs x n_heads x max_q_len x d_k]
k_s = self.W_K(K).view(bs, -1, self.n_heads, self.d_k).permute(0,2,3,1) # k_s : [bs x n_heads x d_k x q_len] - transpose(1,2) + transpose(2,3)
v_s = self.W_V(V).view(bs, -1, self.n_heads, self.d_v).transpose(1,2) # v_s : [bs x n_heads x q_len x d_v]
# Apply Scaled Dot-Product Attention (multiple heads)
if self.res_attention:
output, attn_weights, attn_scores = self.sdp_attn(q_s, k_s, v_s, prev=prev, key_padding_mask=key_padding_mask, attn_mask=attn_mask)
else:
output, attn_weights = self.sdp_attn(q_s, k_s, v_s, key_padding_mask=key_padding_mask, attn_mask=attn_mask)
# output: [bs x n_heads x q_len x d_v], attn: [bs x n_heads x q_len x q_len], scores: [bs x n_heads x max_q_len x q_len]
# back to the original inputs dimensions
output = output.transpose(1, 2).contiguous().view(bs, -1, self.n_heads * self.d_v) # output: [bs x q_len x n_heads * d_v]
output = self.to_out(output)
if self.res_attention: return output, attn_weights, attn_scores
else: return output, attn_weights
q = torch.rand([16, 3, 50, 8])
k = torch.rand([16, 3, 50, 8]).transpose(-1, -2)
v = torch.rand([16, 3, 50, 6])
attn_mask = torch.triu(torch.ones(50, 50)) # shape: q_len x q_len
key_padding_mask = torch.zeros(16, 50)
key_padding_mask[[1, 3, 6, 15], -10:] = 1
key_padding_mask = key_padding_mask.bool()
print('attn_mask', attn_mask.shape, 'key_padding_mask', key_padding_mask.shape)
output, attn = ScaledDotProductAttention()(q, k, v, attn_mask=attn_mask, key_padding_mask=key_padding_mask)
output.shape, attn.shape
t = torch.rand(16, 50, 128)
output, attn = MultiheadAttention(d_model=128, n_heads=3, d_k=8, d_v=6)(t, t, t, key_padding_mask=key_padding_mask, attn_mask=attn_mask)
output.shape, attn.shape
t = torch.rand(16, 50, 128)
att_mask = (torch.rand((50, 50)) > .85).float()
att_mask[att_mask == 1] = -np.inf
mha = MultiheadAttention(d_model=128, n_heads=3, d_k=8, d_v=6)
output, attn = mha(t, t, t, attn_mask=att_mask)
test_eq(torch.isnan(output).sum().item(), 0)
test_eq(torch.isnan(attn).sum().item(), 0)
loss = output[:2, :].sum()
test_eq(torch.isnan(loss).sum().item(), 0)
loss.backward()
for n, p in mha.named_parameters(): test_eq(torch.isnan(p.grad).sum().item(), 0)
t = torch.rand(16, 50, 128)
attn_mask = (torch.rand((50, 50)) > .85)
# True values will be masked
mha = MultiheadAttention(d_model=128, n_heads=3, d_k=8, d_v=6)
output, attn = mha(t, t, t, attn_mask=att_mask)
test_eq(torch.isnan(output).sum().item(), 0)
test_eq(torch.isnan(attn).sum().item(), 0)
loss = output[:2, :].sum()
test_eq(torch.isnan(loss).sum().item(), 0)
loss.backward()
for n, p in mha.named_parameters(): test_eq(torch.isnan(p.grad).sum().item(), 0)
# export
class MultiConcatConv1d(Module):
"""Module that applies one or multiple kernels (and optionally maxpool)"""
def __init__(self, ni, nf, kss=[3,5,7], kernel_sizes=None, maxpool=True, stride=1):
kss = ifnone(kss, kernel_sizes)
assert kss is not None, "you need to pass a kss argument"
if not is_listy(kss): kss = [kss]
_nf = nf // (len(kss) + maxpool)
_total_nf = _nf * (len(kss) + maxpool)
self.layers = nn.ModuleList()
for k in kss:
self.layers.append(Conv1d(ni, _nf, k, stride=stride))
if maxpool: self.layers.append(nn.Sequential(nn.MaxPool1d(3, stride=stride, padding=1), Conv1d(ni, _nf, 1)))
self.to_output = Conv1d(_total_nf, nf, 1) if _total_nf != nf else nn.Identity()
def forward(self, x):
for i,l in enumerate(self.layers):
out = l(x) if i == 0 else torch.cat((out, l(x)), 1)
return self.to_output(out)
t = torch.rand(16, 6, 37)
test_eq(MultiConcatConv1d(t.shape[1], 128, kernel_sizes=[3,5,7], maxpool=True)(t).shape, (t.shape[0], nf, t.shape[-1]))
test_eq(MultiConcatConv1d(t.shape[1], 128, kernel_sizes=[3,5,7], maxpool=True, stride=2)(t).shape, (t.shape[0], nf, math.ceil(t.shape[-1]/2)))
#hide
out = create_scripts(); beep(out)
```
| github_jupyter |
```
import logging
import pickle
import numpy as np
import pandas as pd
from gensim.models.word2vec import Word2Vec
from gensim.models import KeyedVectors
from tqdm import tqdm
from sklearn.mixture import GaussianMixture
from sklearn.feature_extraction.text import TfidfVectorizer,HashingVectorizer,CountVectorizer
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
class SparseCompositeDocumentVectors:
def __init__(self, glove_word_vector_file, num_clusters, pname1, pname2):
self.min_no = 0
self.max_no = 0
self.prob_wordvecs = {}
#### 読み込むファイルの設定
# GloVeの単語ベクトルファイル
self.glove_word_vector_file = glove_word_vector_file
#### 出力するファイルの設定
# GloVeの単語ベクトルに単語数とベクトルサイズを付与したファイル
self.gensim_glove_word_vector_file = "es_gensim_glove_vectors.txt"
# GMMの結果を保存するPickleファイル
self.pname1 = pname1
self.pname2 = pname2
#### その他パラメータ
# GMMのクラスタ数
self.num_clusters = num_clusters
# GloVeの次元数
self.num_features = 50
def load_glove_vector(self):
# GloVeの単語ベクトルファイルを読み込み、単語数とベクトルサイズを付与した処理用のファイルを作成する。
vectors = pd.read_csv(self.glove_word_vector_file, delimiter=' ', index_col=0, header=None)
vocab_count = vectors.shape[0] # 単語数
self.num_features = vectors.shape[1] # 次元数
with open(self.glove_word_vector_file, 'r') as original, open(self.gensim_glove_word_vector_file, 'w') as transformed:
transformed.write(f'{vocab_count} {self.num_features}\n')
transformed.write(original.read()) # 2行目以降はそのまま出力
# GloVeの単語ベクトルを読み込む
self.glove_vectors = KeyedVectors.load_word2vec_format(self.gensim_glove_word_vector_file, binary=False)
def cluster_GMM2(self):
glove_vectors = self.glove_vectors.vectors
# Initalize a GMM object and use it for clustering.
gmm_model = GaussianMixture(n_components=num_clusters, covariance_type="tied", init_params='kmeans', max_iter=100)
# Get cluster assignments.
gmm_model.fit(glove_vectors)
idx = gmm_model.predict(glove_vectors)
print ("Clustering Done...")
# Get probabilities of cluster assignments.
idx_proba = gmm_model.predict_proba(glove_vectors)
# Dump cluster assignments and probability of cluster assignments.
pickle.dump(idx, open(self.pname1,"wb"))
print ("Cluster Assignments Saved...")
pickle.dump(idx_proba,open(self.pname2, "wb"))
print ("Probabilities of Cluster Assignments Saved...")
return (idx, idx_proba)
def cluster_GMM(self):
# GMMによるクラスタリング
clf = GaussianMixture(
n_components=self.num_clusters,
#covariance_type="tied",
covariance_type="diag",
init_params="kmeans",
max_iter=50
)
glove_vectors = self.glove_vectors.vectors
# Get cluster assignments.
clf.fit(glove_vectors)
idx = clf.predict(glove_vectors)
print("Clustering Done...")
# Get probabilities of cluster assignments.
idx_proba = clf.predict_proba(glove_vectors)
# Dump cluster assignments and probability of cluster assignments.
pickle.dump(idx, open(self.pname1, "wb"))
print("Cluster Assignments Saved...")
pickle.dump(idx_proba, open(self.pname2, "wb"))
print("Probabilities of Cluster Assignments saved...")
return (idx, idx_proba)
def read_GMM(self):
# GMMモデルを読み込む。
idx = pickle.load(open(self.idx_name, "rb"))
idx_proba = pickle.load(open(self.idx_proba_name, "rb"))
print("Cluster Model Loaded...")
return (idx, idx_proba)
def get_idf_dict(self, corpus):
# IDFを算出する。
# corpus : 分かち書きした文章のリスト
# 単語の数をカウントする
count_vectorizer = CountVectorizer()
X_count = count_vectorizer.fit_transform(corpus)
# scikit-learn の TF-IDF 実装
tfidf_vectorizer = TfidfVectorizer(token_pattern="(?u)\\b\\w+\\b")
X_tfidf = tfidf_vectorizer.fit_transform(corpus)
feature_names = tfidf_vectorizer.get_feature_names()
idf = tfidf_vectorizer.idf_
word_idf_dict = {}
for pair in zip(feature_names, idf):
word_idf_dict[pair[0]] = pair[1]
return feature_names, word_idf_dict
def get_probability_word_vectors(self, corpus):
"""
corpus: 分かち書き済みの文章のリスト
"""
# GloVeの単語ベクトルを読み込む。
self.load_glove_vector()
# 単語毎のGMMクラスタの確率ベクトル
idx, idx_proba = self.cluster_GMM()
# 各単語が属する確率が高いクラスタのインデックス
word_centroid_map = dict(zip(self.glove_vectors.index2word, idx))
# 各単語が、各クラスタに属する確率
word_centroid_prob_map = dict(zip(self.glove_vectors.index2word, idx_proba))
# TF-IDFを算出する。
featurenames, word_idf_dict = self.get_idf_dict(corpus)
for word in word_centroid_map:
self.prob_wordvecs[word] = np.zeros(self.num_clusters * self.num_features, dtype="float32")
for index in range(self.num_clusters):
try:
self.prob_wordvecs[word][index*self.num_features:(index+1)*self.num_features] = \
self.glove_vectors[word] * word_centroid_prob_map[word][index] * word_idf_dict[word]
except:
continue
self.word_centroid_map = word_centroid_map
def create_cluster_vector_and_gwbowv(self, tokens, flag):
# SDV(Sparse Document Vector)を組み立てる。
bag_of_centroids = np.zeros(self.num_clusters * self.num_features, dtype="float32")
for token in tokens:
try:
temp = self.word_centroid_map[token]
except:
continue
bag_of_centroids += self.prob_wordvecs[token]
norm = np.sqrt(np.einsum('...i,...i', bag_of_centroids, bag_of_centroids))
if norm != 0:
bag_of_centroids /= norm
# 訓練で作成したベクトルをスパース化するために最小と最大を記録しておく。
if flag:
self.min_no += min(bag_of_centroids)
self.max_no += max(bag_of_centroids)
return bag_of_centroids
def make_gwbowv(self, corpus, train=True):
# ドキュメントベクトルのマトリクスを作成する。
# gwbowvには通常のドキュメントベクトルが格納される。
gwbowv = np.zeros((len(corpus), self.num_clusters*self.num_features)).astype(np.float32)
cnt = 0
for tokens in tqdm(corpus):
gwbowv[cnt] = self.create_cluster_vector_and_gwbowv(tokens, train)
cnt += 1
return gwbowv
def dump_gwbowv(self, gwbowv, path="gwbowv_matrix.npy", percentage=0.04):
# スパース化したドキュメントベクトルを保存する。
# スパース化するための閾値を算出する。
min_no = self.min_no*1.0/gwbowv.shape[0]
max_no = self.max_no*1.0/gwbowv.shape[0]
print("Average min: ", min_no)
print("Average max: ", max_no)
thres = (abs(max_no) + abs(min_no))/2
thres = thres * percentage
# 閾値未満のベクトルを0とし、スパース化する。
temp = abs(gwbowv) < thres
gwbowv[temp] = 0
np.save(path, gwbowv)
print("SDV created and dumped...")
def load_matrix(self, name):
return np.load(name)
import argparse
from sklearn.svm import SVC
from scdv import SparseCompositeDocumentVectors
def parse_args():
parser = argparse.ArgumentParser(
description="GloVeとSCDVのパラメータの設定"
)
parser.add_argument('--glove_word_vector_file', type=str)
parser.add_argument('--csv_file', type=str)
parser.add_argument(
'--num_clusters', type=int, default=20
)
parser.add_argument(
'--pname1', type=str, default="gmm_cluster.pkl"
)
parser.add_argument(
'--pname2', type=str, default="gmm_prob_cluster.pkl"
)
return parser.parse_args()
def main(args):
df = pd.read_csv(args.csv_file)
categories = df['業種(大分類)'].unique()
NUM_TOPICS = len(categories)
# 訓練データとtestデータに分ける
train_data, test_data, train_label, test_label, train_id, test_id = train_test_split(
df['分かち書き'], df['業種(大分類)'], df['ID'],
test_size=0.1, train_size=0.9, stratify=df['業種(大分類)'], shuffle=True)
vec = SparseCompositeDocumentVectors(args.glove_word_vector_file, args.num_clusters, args.pname1, args.pname2)
# 確率重み付き単語ベクトルを求める
vec.get_probability_word_vectors(train_data)
# 訓練データからSCDVを求める
train_gwbowv = vec.make_gwbowv(train_data)
# テストデータからSCDVを求める
test_gwbowv = vec.make_gwbowv(test_data, False)
print("train size:{} vector size:{}".format(len(train_gwbowv), len(train_gwbowv[0])))
print("test size:{} vector size:{}".format(len(test_gwbowv), len(test_gwbowv[0])))
print("Test start...")
start = time.time()
clf = lgb.LGBMClassifier(objective="multiclass")
clf.fit(train_gwbowv, train_label)
test_pred = clf.predict(test_gwbowv)
# print(test_pred)
print ("Report")
print (classification_report(test_label, test_pred, digits=6))
print ("Accuracy: ",clf.score(test_gwbowv, test_label))
print ("Time taken:", time.time() - start, "\n")
if __name__ == "__main__":
main(parse_args())
from sklearn.metrics import classification_report
import lightgbm as lgb
import time
num_clusters = 20
pname1 = "gmm_cluster.pkl"
pname2 = "gmm_prob_cluster.pkl"
glove_word_vector_file = "glove_word_vector_file.txt"
df = pd.read_csv('../elasticsearch/es_wakati.csv')
# df = pd.read_csv('wakati_category_all.csv')
categories = df['業種(大分類)'].unique()
NUM_TOPICS = len(categories)
# print(df.groupby(['業種(大分類)']).size())
# 訓練データとtestデータに分ける
train_data, test_data, train_label, test_label, train_id, test_id = train_test_split(
df['分かち書き'], df['業種(大分類)'], df['ID'],
test_size=0.1, train_size=0.9, stratify=df['業種(大分類)'], shuffle=True)
'''
train_id = train_id.values
train_data = train_data.values
train_label = train_label.values
test_id = test_id.values
test_data = test_data.values
test_label = test_label.values
'''
vec = SparseCompositeDocumentVectors(glove_word_vector_file, num_clusters, pname1, pname2)
# 確率重み付き単語ベクトルを求める
vec.get_probability_word_vectors(train_data)
# 訓練データからSCDVを求める
train_gwbowv = vec.make_gwbowv(train_data)
# テストデータからSCDVを求める
test_gwbowv = vec.make_gwbowv(test_data, False)
print("train size:{} vector size:{}".format(len(train_gwbowv), len(train_gwbowv[0])))
print("test size:{} vector size:{}".format(len(test_gwbowv), len(test_gwbowv[0])))
print("Test start...")
start = time.time()
clf = lgb.LGBMClassifier(objective="multiclass")
clf.fit(train_gwbowv, train_label)
test_pred = clf.predict(test_gwbowv)
# print(test_pred)
print ("Report")
print (classification_report(test_label, test_pred, digits=6))
print ("Accuracy: ",clf.score(test_gwbowv, test_label))
print ("Time taken:", time.time() - start, "\n")
from sklearn.metrics import classification_report
import lightgbm as lgb
import time
num_clusters = 20
pname1 = "gmm_cluster.pkl"
pname2 = "gmm_prob_cluster.pkl"
df = pd.read_csv('wakati_category_all.csv')
categories = df['業種(大分類)'].unique()
NUM_TOPICS = len(categories)
print(df.groupby(['業種(大分類)']).size())
all_data = df['分かち書き'].values
vec = SparseCompositeDocumentVectors(num_clusters, pname1, pname2)
# 確率重み付き単語ベクトルを求める
vec.get_probability_word_vectors(all_data)
# 訓練データからSCDVを求める
gwbowv = vec.make_gwbowv(all_data)
train_gwbowv[0]
```
| github_jupyter |
```
import folium
import folium
map_osm = folium.Map(location=[37.7549, -122.4194], zoom_start=13, detect_retina=True,
tiles='http://tile.stamen.com/watercolor/{z}/{x}/{y}.jpg', attr='Map tiles by <a href="http://stamen.com">Stamen Design</a>, under <a href="http://creativecommons.org/licenses/by/3.0">CC BY 3.0</a>. Data by <a href="http://openstreetmap.org">OpenStreetMap</a>, under <a href="http://creativecommons.org/licenses/by-sa/3.0">CC BY SA</a>.')
map_osm.add_tile_layer(tile_url='http://tile.stamen.com/toner-labels/{z}/{x}/{y}.png', attr='labels',
active=True, overlay=True)
html = r'''<div align="center"> <font size="5"><b>test2</b></font> <br><img src="http://planck.ucsc.edu/images/insight/VRGLWJryfUIlYH_woaLkHw.png" alt="NOPE" style="width:200px;height:200px;"></div>'''
iframe = folium.element.IFrame(html=html,width=250,height=250)
popup = folium.Popup(html=iframe)
#popup = folium.Popup(html, max_width=300)
icon = folium.Icon(color="blue", icon="ok")
marker1 = folium.Marker(location=[37.7549, -122.4194], popup=popup, icon=icon)
map_osm.add_children(marker1)
icon = folium.Icon(color="blue", icon="ok")
marker2 = folium.Marker(location=[37.7449, -122.4194], popup=popup, icon=icon)
map_osm.add_children(marker2)
map_osm.save('/home/carlson/web/images/insight/map.html')
map_osm
import folium
from folium import plugins
print(folium.__file__)
print(folium.__version__)
import numpy as np
data = (np.random.normal(size=(100, 3)) *
np.array([[1, 1, 1]]) +
np.array([[48, 5, 1]])).tolist()
mapa = folium.Map([48., 5.], tiles='stamentoner', zoom_start=6)
mapa.add_children(plugins.HeatMap(data))
mapa
import pandas as pd
bus_df = pd.read_pickle('../input/yelp_academic_dataset_business.pickle')
bus_df.business_id=='VRGLWJryfUIlYH_woaLkHw'
lat = bus_df.latitude[bus_df.business_id=='VRGLWJryfUIlYH_woaLkHw'].values[0]
lon = bus_df.longitude[bus_df.business_id=='VRGLWJryfUIlYH_woaLkHw'].values[0]
print lat, lon
import sys
sys.path.append('../vectorsearch/')
import vectorsearch
import pandas as pd
df_businesses = pd.read_pickle('../input/yelp_academic_dataset_business_SF.pickle')
def get_bus_ids_city_state(city, state):
bids = set(list(df_businesses.business_id[(df_businesses.city==city)
& (df_businesses.state==state)].values))
return bids
bids_in_city_state = get_bus_ids_city_state('San Francisco', 'CA')
print len(bids_in_city_state)
rev_topic = vectorsearch.GetDocTopic('Oysters, cocktails, peir, alcatraz')
bids, sims = vectorsearch.FindBusinessSimilarityLDA(rev_topic, business_ids=bids_in_city_state, top_n=30, method='Hel')
import folium
from folium import plugins
print(folium.__file__)
print(folium.__version__)
import numpy as np
# data = (np.random.normal(size=(100, 3)) *
# np.array([[1, 1, 1]]) +
# np.array([[48, 5, 1]])).tolist()
heatmap_events = [(df_businesses.latitude[df_businesses.business_id==bus_id].values[0],
df_businesses.longitude[df_businesses.business_id==bus_id].values[0],
-sims[i]+sims[0]) for i, bus_id in enumerate(bids)]
lats = sims_array = np.array(heatmap_events)[:,0]
lons = sims_array = np.array(heatmap_events)[:,1]
sims_array = np.array(heatmap_events)[:,2]
scale = sims[3]-sims[0]
sims_array = ((1-1/(np.exp(sims_array/scale)+1))*50).astype(np.int32)
heatmap = []
for i, sim in enumerate(sims_array):
for j in range(sim):
heatmap += [[lats[i]+.00001*j, lons[i]]]
#print heatmap
# heatmap_events = zip(lats,lons,sims_array)
# for event in heatmap:
# print event
mapa = folium.Map([37.7549, -122.4194], tiles='stamentoner', zoom_start=12,)
# heatmap = [[37.7549, -122.4194]]*10 + [[37.7549, -122.6194]]
# print heatmap
print heatmap
mapa.add_children(plugins.HeatMap(heatmap, max_zoom=18, radius=25, max_val=20))
mapa
import requests
from time import sleep
proxy_list = []
for i in range(60):
response = requests.get('https://www.proxicity.io/api/v1/984fee31a6c723be2c970db9df3503bf/proxy')
proxy_list.append(response.json()['ipPort'])
print response.json()['ip'], response.json()['port']
sleep(6)
```
| github_jupyter |
## <b> Scientific modules and IPython <b/>
```
%matplotlib inline
import matplotlib.pylab as plt
```
#### <b>Core scientific packages<b/>
Python is not doing your science, the packages are doing it. Some of them are here:
<img style="width:1000px;" src="core.png">
[Source of this figure](http://chris35wills.github.io/courses/pydata_stack/)
### <b> Installation <b/>
There exist various ways to install and use python and there is no perfect way. It depends on your knowledge, but most propably Miniconda and CLI interpreter IPython together with Jupyter notebooks are an easy and effective way to start.
### <b> IPython <b/>
In order to be productive you need comfortable environment, and this is what IPython provides. It was started as enhanced python interactive shell, but with time become architecture for interactive computing. It was developed by an former Mathematica user (Fernando Peréz) in 2001 and has a great web interface, called:
### <b> Jupyter notebook (we are using it already, quick intro, more later) <b/>
#### Code execution
```
print('Python is not a snake')
```
#### Text (Markdown)
IPython [website](http://ipython.org/).
List:
* [Python on Codeacademy](http://www.codecademy.com/tracks/python)
* [Google's Python Class](https://developers.google.com/edu/python/)
Code:
print('hello world')
#### $\LaTeX$ equations
$$\int_0^\infty e^{-x^2} dx=\frac{\sqrt{\pi}}{2}$$
$$
F(x,y)=0 ~~\mbox{and}~~
\left| \begin{array}{ccc}
F''_{xx} & F''_{xy} & F'_x \\
F''_{yx} & F''_{yy} & F'_y \\
F'_x & F'_y & 0
\end{array}\right| = 0
$$
#### Plots
```
x = [1,2,3,4,5]
plt.plot(x);
```
#### Rich media
```
from IPython.display import YouTubeVideo
YouTubeVideo('Bv3pB9TaWOk')
```
* [IPython website](http://ipython.org/)
* [Notebook gallery](https://github.com/jupyter/jupyter/wiki/A-gallery-of-interesting-Jupyter-Notebooks)
### <b> Run notebook <b>
In order to start Jupyter notebook you have to type:
jupyter notebook
### <b> Main IPython features <b>
### Getting help
You can use question mark in order to get help. To execute cell you have to press *Shift+Enter*
```
?
```
Question mark after a function will open pager with documentation. Double question mark will show you source code of the function.
```
plt.plot??
```
Press SHIFT+TAB after opening bracket in order to get help for the function (list of arguments, doc string).
```
sum()
```
### <b> Accessing the underlying operating system <b>
You can access system functions by typing exclamation mark.
```
!pwd
```
If you already have some netCDF file in the directory and *ncdump* is installed, you can for example look at its header.
```
!ncdump -h test_netcdf.nc
```
### <b> Magic functions <b>
The magic function system provides a series of functions which allow you to
control the behavior of IPython itself, plus a lot of system-type
features.
Let's create some set of numbers using [range](http://docs.python.org/2/library/functions.html#range) command:
```
list(range(10))
```
And find out how long does it take to run it with *%timeit* magic function:
```
%timeit list(range(10))
```
Print all interactive variables:
```
%whos
```
### <b> Cell-oriented magic <b>
Receive as argument both the current line where they are declared and the whole body of the cell.
```
%%timeit
range(10)
range(100)
```
Thre are several cell-oriented magic functions that allow you to run code in other languages:
```
%%bash
echo "My shell is:" $SHELL
%%perl
$variable = 1;
print "The variable has the value of $variable\n";
```
You can write content of the cell to a file with *%%writefile* (or *%%file* for ipython < 1.0):
```
%%writefile hello.py
#if you use ipython < 1.0, use %%file comand
#%%file
a = 'Here is your first program!'
print(a)
```
And then run it:
```
%run hello.py
```
The *%run* magic will run your python script and load all variables into your interactive namespace for further use.
```
%whos
```
In order to get information about all magic functions type:
```
%magic
```
### Links:
[The cell magics in IPython](http://nbviewer.ipython.org/urls/raw.github.com/ipython/ipython/1.x/examples/notebooks/Cell%20Magics.ipynb)
| github_jupyter |
```
# https://docs.gdc.cancer.gov/API/Users_Guide/Search_and_Retrieval/
import requests
import json
import boto3
import re
import gzip
import pandas as pd
import dask
from dask.distributed import Client
data_endpt = 'https://api.gdc.cancer.gov/data'
cases_endpt = 'https://api.gdc.cancer.gov/cases'
files_endpt = 'https://api.gdc.cancer.gov/files'
indexd_endpt = 'https://nci-crdc.datacommons.io/index/index/'
## Query Settings
# primary_site = "Breast"
project_id = "TCGA-BRCA"
data_type = "Gene Expression Quantification" # RNA-Seq
workflow_type = "HTSeq - Counts"
size = 2000
# The 'fields' parameter is passed as a comma-separated string of single names.
fields = [
"file_name"
, "cases.primary_site"
, "cases.case_id"
, "cases.project.project_id"
, "cases.days_to_lost_to_followup"
, "cases.submitter_id"
, "cases.samples.submitter_id"
, "cases.samples.sample_id"
]
fields = ','.join(fields)
#cases.project.project_id in ["TCGA-BRCA"] and files.data_type in ["Gene Expression Quantification"]
filters = {
"op":"and",
"content":[
{"op": "in",
"content":{
"field": "cases.project.project_id",
"value": [project_id]
}
},
{"op": "in",
"content":{
"field": "files.data_type",
"value": [data_type]
}
},
{"op": "in",
"content":{
"field": "files.analysis.workflow_type",
"value": [workflow_type]
}
}
]
}
# With a GET request, the filters parameter needs to be converted
# from a dictionary to JSON-formatted string
params = {
"filters": json.dumps(filters),
"fields": fields,
"format": "JSON",
"size": size
}
## Get Files
query_response = requests.get(files_endpt, params = params)
json_response = json.loads(query_response.content.decode("utf-8"))["data"]["hits"]
print (len(json_response))
##print(json_response)
files_json = json_response
## Scale out Dask Cluster
ecs = boto3.client('ecs')
resp = ecs.list_clusters()
clusters = resp['clusterArns']
if len(clusters) > 1:
print("Please manually select your cluster")
cluster = clusters[0]
numWorkers=10
ecs.update_service(cluster=cluster, service='Dask-Worker', desiredCount=numWorkers)
ecs.get_waiter('services_stable').wait(cluster=cluster, services=['Dask-Worker'])
client = Client('Dask-Scheduler.local-dask:8786')
client
@dask.delayed
def get_data(uuid, sample_submitter_id):
query_response = requests.get(indexd_endpt + "/" + uuid)
urls_response = json.loads(query_response.content.decode("utf-8"))["urls"]
url = [x for x in urls_response if x.startswith("s3://")]
if len(url) != 1:
print("Something weird with UUID " + uuid + "returned " + str(url))
url = url[0]
content = pd.read_csv(url, compression='gzip', header=None, dtype=str, sep="\t")
content.index = content[0]
content.columns = ['id', sample_submitter_id]
content = content[[sample_submitter_id]]
return content
delayed_results = []
for file_entry in files_json:
delayed_results.append(get_data(file_entry["id"], file_entry["cases"][0]["samples"][0]["submitter_id"]))
%%time
df = pd.concat(dask.compute(*delayed_results), axis=1, join="outer")
df
numWorkers=0
ecs.update_service(cluster=cluster, service='Dask-Worker', desiredCount=numWorkers)
ecs.get_waiter('services_stable').wait(cluster=cluster, services=['Dask-Worker'])
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import xarray as xr
import glob
import matplotlib
import matplotlib.pyplot as plt
%matplotlib inline
thedir = '/glade/scratch/djk2120/mini_ens/'
f = 'miniens_oaat'+'0001'+'_h0.nc'
#for use on Casper
from dask_jobqueue import SLURMCluster
from dask.distributed import Client
cluster = SLURMCluster(cores=12,
processes=12, memory="300GB",
project='P93300041',
walltime='2:00:00')
cluster.scale(12)
client = Client(cluster)
#for use on cheyenne
from dask_jobqueue import PBSCluster
from dask.distributed import Client
cluster = PBSCluster(cores=36,
processes=36, memory="109GB",
project='P93300041',
queue='regular',
resource_spec='select=5:ncpus=36:mem=109G',
walltime='01:00:00')
cluster.scale(36)
client = Client(cluster)
client
ens = range(33)
thedir = '/glade/scratch/djk2120/mini_ens/output/'
paths = [thedir+'miniens_oaat'+str(ee).zfill(4)+'_h1.nc' for ee in ens]
%time ds = xr.open_mfdataset(paths,combine='nested',concat_dim='ens',parallel='True')
ds['ens']=ens
pft = ds['pfts1d_itype_veg'][0]
pftnames = ['BG','NEMT','NEBT','NDBT','BETT','BEMT','BDTT','BDMT','BDBT','BES','BDMS','BDBS','C3ArG','C3G','C4G','C3C']
print(str(np.round(ds.nbytes/1e9,2))+' GB')
kmax = [2e-8/2**x for x in np.arange(7)]
pdir = '/glade/scratch/djk2120/mini_ens/paramfiles/'
paths = sorted(glob.glob(pdir+'*.nc'))
%time params = xr.open_mfdataset(paths,combine='nested',concat_dim='ens',parallel='True')
ds['ens']=ens
rset = []
for ee in ens:
krun = np.unique(params['rootprof_beta'].sel(ens=ee))
if krun.size==1:
rset.append(ee)
rpb = params['rootprof_beta'].sel(ens=kset).groupby('ens').mean(dim=xr.ALL_DIMS)
rpb = np.array([params['rootprof_beta'].sel(ens=ee)[0,4].values for ee in ens])
ix = rpb!=rpb[0]
rpb = rpb[ix]
rset = np.array(ens)[ix]
plt.figure(figsize=[10,8])
for ixpft in 1+np.arange(15):
f = [np.mean(ds['FPSN'].sel(ens=ee)[:,pft==ixpft]) for ee in rset]
plt.subplot(4,4,ixpft)
plt.plot(rpb,f,'-x')
#plt.plot([params['krmax'].sel(ens=0)[ixpft],params['krmax'].sel(ens=0)[ixpft]],[0,6],'r:')
if (ixpft>11):
plt.xlabel('rootprof_beta')
else:
plt.xticks([])
if (ixpft==1)|(ixpft==5)|(ixpft==9)|(ixpft==13):
plt.ylabel('FPSN')
plt.ylim([0,5.5])
#plt.xlim([-2.5e-9,2.5e-8])
plt.title(pftnames[ixpft])
kset = []
for ee in ens:
krun = np.unique(params['krmax'].sel(ens=ee))
if krun.size==1:
kset.append(ee)
krmax = params['krmax'].sel(ens=kset)[:,2]
kr_table = pd.DataFrame()
kr_table['name']=pftnames
kr_table['Krmax']=params['krmax'].sel(ens=0)[:16].values
kr_table
plt.figure(figsize=[10,8])
for ixpft in 1+np.arange(15):
f = [np.mean(ds['FPSN'].sel(ens=ee)[:,pft==ixpft]) for ee in kset]
plt.subplot(4,4,ixpft)
plt.plot(krmax,f,'-x')
plt.plot([params['krmax'].sel(ens=0)[ixpft],params['krmax'].sel(ens=0)[ixpft]],[0,6],'r:')
if (ixpft>11):
plt.xlabel('krmax')
else:
plt.xticks([])
if (ixpft==1)|(ixpft==5)|(ixpft==9)|(ixpft==13):
plt.ylabel('FPSN')
plt.ylim([0,5.5])
plt.xlim([-2.5e-9,2.5e-8])
plt.title(pftnames[ixpft])
plt.figure(figsize=[10,8])
for ixpft in 1+np.arange(15):
f = [np.mean(ds['FPSN'].sel(ens=ee)[:,pft==ixpft]) for ee in range(7)]
plt.subplot(4,4,ixpft)
plt.plot(kmax,f,'-x')
if (ixpft>11):
plt.xlabel('kmax')
else:
plt.xticks([])
if (ixpft==1)|(ixpft==5)|(ixpft==9)|(ixpft==13):
plt.ylabel('FPSN')
plt.ylim([0,5.5])
plt.title(pftnames[ixpft])
plt.figure(figsize=[10,8])
for ixpft in 1+np.arange(15):
plt.subplot(4,4,ixpft)
gpp=np.mean(ds['TLAI'].sel(ens=1)[:,pft==ixpft].values,axis=1)
ix1 = np.arange(120)
ix2 = 120+ix1
ix3 = 120+ix2
plt.plot((0.5+np.arange(120))/12,(gpp[ix2]-gpp[ix1]))
plt.plot((0.5+np.arange(120))/12,(gpp[ix3]-gpp[ix2]))
plt.ylim([-0.1,0.1])
if (ixpft==1)|(ixpft==5)|(ixpft==9)|(ixpft==13):
plt.ylabel('Delta LAI')
else:
plt.yticks([])
if (ixpft>11):
plt.xlabel('year')
else:
plt.xticks([])
plt.title(pftnames[ixpft])
plt.figure(figsize=[10,8])
for ixpft in 1+np.arange(15):
plt.subplot(4,4,ixpft)
gpp=np.mean(ds['FPSN'].sel(ens=1)[:,pft==ixpft].values,axis=1)
ix1 = np.arange(120)
ix2 = 120+ix1
ix3 = 120+ix2
plt.plot((0.5+np.arange(120))/12,(gpp[ix2]-gpp[ix1]))
plt.plot((0.5+np.arange(120))/12,(gpp[ix3]-gpp[ix2]))
plt.ylim([-0.15,0.15])
if (ixpft==1)|(ixpft==5)|(ixpft==9)|(ixpft==13):
plt.ylabel('Delta GPP')
else:
plt.yticks([])
if (ixpft>11):
plt.xlabel('year')
else:
plt.xticks([])
plt.title(pftnames[ixpft])
ens = range(13)
thedir = '/glade/scratch/djk2120/mini_ens/output/'
paths = [thedir+'miniens_oaat'+str(ee).zfill(4)+'_h0.nc' for ee in ens]
%time ds2 = xr.open_mfdataset(paths,combine='nested',concat_dim='ens',parallel='True')
ds2['ens']=ens
ix1 = np.arange(120)
ix2 = 120+ix1
ix3 = 120+ix2
plt.figure(figsize=[10,7])
for i in 1+np.arange(12):
tvc = np.mean(ds2['TOTVEGC'].sel(ens=i),axis=1).values
plt.subplot(3,4,i)
plt.plot((0.5+np.arange(120))/12,tvc[ix2]-tvc[ix1])
plt.plot((0.5+np.arange(120))/12,tvc[ix3]-tvc[ix2])
plt.ylim([-4,4])
if i>8:
plt.xlabel('year')
else:
plt.xticks([])
plt.title('ens'+str(i))
if (i==1)|(i==5)|(i==9):
plt.ylabel('Delta TOTVEGC')
else:
plt.yticks([])
ix1 = np.arange(120)
ix2 = 120+ix1
ix3 = 120+ix2
plt.figure(figsize=[10,7])
for i in 1+np.arange(12):
tvc = np.mean(ds2['TOTCOLC'].sel(ens=i),axis=1).values
plt.subplot(3,4,i)
plt.plot((0.5+np.arange(120))/12,tvc[ix2]-tvc[ix1])
plt.plot((0.5+np.arange(120))/12,tvc[ix3]-tvc[ix2])
plt.ylim([-10,10])
if i>8:
plt.xlabel('year')
else:
plt.xticks([])
plt.title('ens'+str(i))
if (i==1)|(i==5)|(i==9):
plt.ylabel('Delta TOTCOLC')
else:
plt.yticks([])
client.close()
```
| github_jupyter |
# Introduction
This notebook shows how to evaluate neural cross-lingual summarization (xls) presented in paper [A Deep Reinforced Model for Zero-Shot Cross-Lingual Summarization
with Bilingual Semantic Similarity Rewards](https://arxiv.org/pdf/2006.15454.pdf) . Their original codes are available at [zdou0830/crosslingual_summarization_semantic](https://github.com/zdou0830/crosslingual_summarization_semantic). <br><br>
XLS comes with different models, this notebook demonstrates way to evaluate 'word-level-supervised' models only. Since pinpointing the end of sentences from running Thai text is difficult, we intentionally left 'sent-level-supervised' for future works.
<br>
Make sure to use GPU runtime if it is available.
```
from google.colab import drive
drive._mount('/content/drive')
#!pip install -q torch==1.5.1 torchvision==0.6.1
!pip install -q rouge
!pip install -q bert_score
import pandas as pd
from tqdm.notebook import tqdm
import rouge
from bert_score import score
!git clone https://github.com/nakhunchumpolsathien/ThaiCrossSum_Corpora
%cd /content/ThaiCrossSum_Corpora/src/XLS
import json
config_file = open('/content/crosslingual_summarization_semantic/word-level-supervised/translate.json')
config_info = json.load(config_file)
print('Example of config file')
print(json.dumps(config_info, indent=4, sort_keys=True))
#th2en
!python '/content/ThaiCrossSum_Corpora/src/XLS/word-level-supervised/translate.json' -config '/content/ThaiCrossSum_Corpora/src/XLS/word-level-supervised/translate.json'
#th2zh (reduce batch size to 1000 if it yields 'out-of-memory'.)
!python '/content/ThaiCrossSum_Corpora/src/XLS/word-level-supervised/translate.json' -config '/content/ThaiCrossSum_Corpora/src/XLS/word-level-supervised/translate.json'
```
## Outputs Example
```
# Body Article
with open("/content/drive/MyDrive/Projects/Model_Checkpoints/XLS/dataset/th2zh-full/test.CLS.source.language1") as file:
head = list(islice(file, 1))
print(head)
```
### TH2EN
```
from pprint import pprint
from itertools import islice
with open("/content/crosslingual_summarization_semantic/xls.out") as file:
head = list(islice(file, 1))
pprint(head)
```
### TH2ZH
```
with open("/content/xls_th2zh.out") as file:
head = list(islice(file, 1))
pprint(head)
```
## Evaluate CLS output results with ROUGE
### TH2EN
```
!rouge -f '/content/crosslingual_summarization_semantic/xls.out' '/content/drive/MyDrive/Projects/th-ncls/datasets/th2en/beaver-base/test.eng.ref' --avg
```
### TH2ZH
```
!rouge -f '/content/xls_th2zh.out' '/content/drive/MyDrive/Projects/Model_Checkpoints/XLS-proposedModel/dataset/th2zh-full/test.CLS.ref.language2' --avg
```
## Evaluate CLS output results with BertScore
```
import logging
import transformers
transformers.tokenization_utils.logger.setLevel(logging.ERROR)
transformers.configuration_utils.logger.setLevel(logging.ERROR)
transformers.modeling_utils.logger.setLevel(logging.ERROR)
%matplotlib inline
```
### TH2EN
```
with open("/content/crosslingual_summarization_semantic/xls.out") as f:
cands = [line.strip() for line in f]
with open("/content/drive/MyDrive/Projects/th-ncls/datasets/th2en/beaver-base/test.eng.ref") as f:
refs = [line.strip() for line in f]
P, R, F1 = score(cands, refs, lang='en', verbose=True) #use lang='zh' if evaluate th2zh models
print(f"System level F1 score: {F1.mean():.3f}")
print(f"System level P score: {P.mean():.3f}")
print(f"System level R score: {R.mean():.3f}")
import matplotlib.pyplot as plt
plt.hist(F1, bins=30)
plt.xlabel("score")
plt.ylabel("counts")
plt.show()
```
### TH2ZH
```
with open("/content/xls_th2zh.out") as f:
cands = [line.strip() for line in f]
with open("/content/drive/MyDrive/Projects/Model_Checkpoints/XLS/dataset/th2zh-full/test.CLS.ref.language2") as f:
refs = [line.strip() for line in f]
P, R, F1 = score(cands, refs, lang='zh', verbose=True) #use lang='zh' if evaluate th2zh models
print(f"System level F1 score: {F1.mean():.3f}")
print(f"System level P score: {P.mean():.3f}")
print(f"System level R score: {R.mean():.3f}")
import matplotlib.pyplot as plt
plt.hist(F1, bins=30)
plt.xlabel("score")
plt.ylabel("counts")
plt.show()
```
## Evaluate MT Results with BLUE score
### TH2EN
```
import nltk
import codecs
from tqdm.notebook import tqdm
import pandas as pd
def get_text_list(fpath):
texts = []
with codecs.open(fpath, encoding='utf-8') as f:
for line in f:
text = line.replace('!', '').replace('.', '').replace(',', '').replace('\n', '')
text = text.replace(',', '').replace('?', '').replace('。', '')
texts.append(text)
return texts
hypos = get_text_list('/content/mt.out')
refs = get_text_list('/content/test.MT.target.EN.txt')
print(len(hypos))
print(len(refs))
scores = []
for i in tqdm(range(len(refs))):
hyp = hypos[i].split()
ref = refs[i].split()
scores.append(nltk.translate.bleu_score.sentence_bleu([ref], hyp))
th2en_mt_output_df = pd.DataFrame(list(zip(refs, hypos, scores)),
columns =['ref', 'hyp', 'bleu'])
th2en_mt_output_df.sample(n=10)
print(f'Average BLEU scores {round(th2en_mt_output_df["bleu"].mean()*100, 2)}')
```
### TH2ZH
Note: Tokenized at word level, not character level.
```
hypos = get_text_list('/content/mt_th2zh.out')
refs = get_text_list('/content/test.MT.target.ZH.txt')
scores = []
for i in tqdm(range(len(refs))):
hyp = hypos[i].split()
ref = refs[i].split()
scores.append(nltk.translate.bleu_score.sentence_bleu([ref], hyp))
th2zh_mt_output_df = pd.DataFrame(list(zip(refs, hypos, scores)),
columns =['ref', 'hyp', 'bleu'])
th2zh_mt_output_df.sample(n=10)
print(f'Average BLEU scores {round(th2zh_mt_output_df["bleu"].mean()*100, 2)}')
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# TensorFlow 2 quickstart for beginners
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/quickstart/beginner"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />Lihat di TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/id/tutorials/quickstart/beginner.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Jalankan di Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/id/tutorials/quickstart/beginner.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />Lihat sumber kode di GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/id/tutorials/quickstart/beginner.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Unduh notebook</a>
</td>
</table>
Note: Komunitas TensorFlow kami telah menerjemahkan dokumen-dokumen ini. Tidak ada jaminan bahwa translasi ini akurat, dan translasi terbaru dari [Official Dokumentasi - Bahasa Inggris](https://www.tensorflow.org/?hl=en) karena komunitas translasi ini adalah usaha terbaik dari komunitas translasi.
Jika Anda memiliki saran untuk meningkatkan terjemahan ini, silakan kirim pull request ke [tensorflow/docs](https://github.com/tensorflow/docs) repositori GitHub.
Untuk menjadi sukarelawan untuk menulis atau memeriksa terjemahan komunitas, hubungi
[daftar docs@tensorflow.org](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs).
Panduan singkat ini akan menggunakan [Keras](https://www.tensorflow.org/guide/keras/overview) untuk:
1. Membangun jaringan saraf tiruan yang mengklasifikasikan gambar.
2. Melatih jaringan saraf tiruan tersebut.
3. Dan, pada akhirnya, mengevaluasi keakuratan dari model.
Ini adalah file notebook [Google Colaboratory](https://colab.research.google.com/notebooks/welcome.ipynb). Program python akan dijalankan langsung dari browser — cara yang bagus untuk mempelajari dan menggunakan TensorFlow. Untuk mengikuti tutorial ini, jalankan notebook di Google Colab dengan mengklik tombol di bagian atas halaman ini.
1. Di halaman Colab, sambungkan ke runtime Python: Di menu sebelah kanan atas, pilih * CONNECT *.
2. Untuk menjalankan semua sel kode pada notebook: Pilih * Runtime *> * Run all *.
Download dan instal TensorFlow 2 dan impor TensorFlow ke dalam program Anda:
```
# Install TensorFlow
import tensorflow as tf
```
Siapkan [dataset MNIST](http://yann.lecun.com/exdb/mnist/). Ubah sampel dari bilangan bulat menjadi angka floating-point (desimal):
```
mnist = tf.keras.datasets.mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
```
Build model `tf.keras.Sequential` dengan cara menumpuk lapisan layer. Untuk melatih data, pilih fungsi untuk mengoptimalkan dan fungsi untuk menghitung kerugian:
```
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
```
Melatih dan mengevaluasi model:
```
model.fit(x_train, y_train, epochs=5)
model.evaluate(x_test, y_test, verbose=2)
```
Penggolong gambar tersebut, sekarang dilatih untuk akurasi ~ 98% pada dataset ini. Untuk mempelajari lebih lanjut, baca [tutorial TensorFlow](https://www.tensorflow.org/tutorials/).
| github_jupyter |
# CLUSTERING
### Importamos las librerías
```
import pandas as pd
import numpy as np
import os
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
%matplotlib inline
import seaborn as sns
from sklearn.cluster import KMeans
from sklearn.decomposition import PCA as sklearnPCA
from sklearn.preprocessing import StandardScaler
```
### Importamos el dataset y lo limpiamos
```
def clean_dataset(df):
assert isinstance(df, pd.DataFrame), "df needs to be a pd.DataFrame"
df.dropna(inplace=True)
indices_to_keep = ~df.isin([np.nan, np.inf, -np.inf]).any(1)
return df[indices_to_keep].astype(np.float64)
filename = "nasa/event/event_wind_summary/event_wind_summary.csv"
fd = pd.read_csv(filename);
fd = clean_dataset(fd);
fd.columns
```
### Normalizamos el dataset completo
```
#df_norm = ( fd - fd.min()) / ( fd.max() - fd.min() )
df_norm = StandardScaler().fit_transform(fd.astype(float))
df_norm= pd.DataFrame(df_norm, columns=fd.columns)
```
### Buscamos grupos de variables dependientes
```
f, ax = plt.subplots(figsize=(20,20))
corr = df_norm.corr()
sns.heatmap(corr,square=True ,cmap=sns.diverging_palette(220, 20, as_cmap=True), ax=ax , annot = True)
```
# Generamos los grupos con variable tiempo independiente
```
#GRUPO 1 ->
gp_1 = ['MEDIAN_X_AXIS', 'FIRST_X_AXIS',
'MAXIMUM_X_AXIS','RMS_X_AXIS_X100',
'RMS_Y_AXIS_X100', 'RMS_Z_AXIS_X100','WINDSPEED']
#GRUPO 2 ->
gp_2 = ['PRESSURE','AIR_TEMPERATURE']
#GRUPO 3 ( INDEPENDIENTES ) ->
gp_3 = ['SEISMIC_TIME_SOLS','MINIMUM_X_AXIS','MEAN_X_AXIS_CROSSINGS',
'MEAN_Y_AXIS_CROSSINGS', 'MEAN_Z_AXIS_CROSSINGS','WIND_DIRECTION']
```
### Aplicamos PCA para modelo con tiempo independiente y creamos el nucleo de entrenamiento
```
# Indicamos el numero de columnas que tiene que salir
sklearn_pca = sklearnPCA(n_components=1)
# Aplicamos PCA para los dos conjuntos que hemos hecho a partir de analisis de correlaciones
datos_pca_gp_1 = sklearn_pca.fit_transform(df_norm[gp_1])
datos_pca_gp_2 = sklearn_pca.fit_transform(df_norm[gp_2])
# Unimos para formar el dataset de entrenamiento del algoritmo de clustering
core = df_norm[gp_3];
core['sismo'] = datos_pca_gp_1;
core['pre_temp'] = datos_pca_gp_2;
core[core == np.nan].count()
f, ax = plt.subplots(figsize=(10,10))
corr = core.corr()
sns.heatmap(corr,square=True ,cmap=sns.diverging_palette(220, 20, as_cmap=True), ax=ax , annot = True)
```
### Creamos el modelo K-Means con variable tiempo independiente, lo entrenamos y sacamos las etiquetas
```
# Entrenamos el modelo
model_kmeans = KMeans(n_clusters=5).fit(core)
# Sacamos los centroides
centroids = model_kmeans.cluster_centers_
# Sacamos los tags del dataset
labels = model_kmeans.predict(core)
# Asignamos las categorias al dataset original sin normalizar
fd["TAG_KM"] = labels;
fd.groupby("TAG_KM").mean()
```
### Estudio de categorias
```
plt.scatter(X.RMS_X_AXIS_X100, X.PRESSURE, c=asignar, s=10)
plt.xlabel('RMS_X_AXIS_X100')
plt.ylabel('PRESSURE')
X = fd.copy()
fig = plt.figure()
#ax = Axes3D(fig)
colores=['blue','red','green','blue','cyan','yellow','orange','black','pink','brown','purple']
asignar=[]
for row in labels:
asignar.append(colores[row])
plt.scatter(X.RMS_X_AXIS_X100, X.SEISMIC_TIME_SOLS, c=asignar, s=10)
plt.xlabel('RMS_X_AXIS_X100')
plt.ylabel('SEISMIC_TIME_SOLS')
PROBLEMAS
* No podemos usar la variable tiempo como variable independiente a las demás, influye demasiado en el modelo.
* Opciones:
- Incluirla o meterla dentro del subgrupo variables atmosféricas ( No merece la pena ).
- Desincluirla ( EL resultado será mas fino ).
Tenemos un total de tres modelos diferentes para comparación de datos.
```
# Generamos los grupos sin tener en cuenta la variable tiempo
```
#GRUPO 1 ->
gp_1_dep = ['MEDIAN_X_AXIS', 'FIRST_X_AXIS',
'MAXIMUM_X_AXIS','RMS_X_AXIS_X100',
'RMS_Y_AXIS_X100', 'RMS_Z_AXIS_X100','WINDSPEED']
#GRUPO 2 ->
gp_2_dep = ['PRESSURE','AIR_TEMPERATURE']
#GRUPO 3 ( INDEPENDIENTES ) ->
gp_3_dep = ['MINIMUM_X_AXIS','MEAN_X_AXIS_CROSSINGS',
'MEAN_Y_AXIS_CROSSINGS', 'MEAN_Z_AXIS_CROSSINGS','WIND_DIRECTION']
# Indicamos el numero de columnas que tiene que salir
sklearn_pca = sklearnPCA(n_components=1)
# Aplicamos PCA para los dos conjuntos que hemos hecho a partir de analisis de correlaciones
datos_pca_gp_1_dep = sklearn_pca.fit_transform(df_norm[gp_1_dep])
datos_pca_gp_2_dep = sklearn_pca.fit_transform(df_norm[gp_2_dep])
# Unimos para formar el dataset de entrenamiento del algoritmo de clustering
core_dep = df_norm[gp_3_dep];
core_dep['sismo'] = datos_pca_gp_1_dep;
core_dep['pre_temp'] = datos_pca_gp_2_dep;
core_dep[core_dep == np.nan].count()
# Entrenamos el modelo
model_kmeans_dep = KMeans(n_clusters=5).fit(core_dep)
# Sacamos los centroides
centroids_dep = model_kmeans_dep.cluster_centers_
# Sacamos los tags del dataset
labels_dep = model_kmeans_dep.predict(core_dep)
# Lo incluimos en el modelo principal como variable dependiente
fd["TAG_KM_DEP"] = labels_dep;
# Agrupamos por esta ultima y vemos las medias
fd.groupby("TAG_KM_DEP").mean()
# Comparación entre
X = fd.copy()
fig = plt.figure();
#ax = Axes3D(fig)
colores=['blue','red','green','blue','cyan','yellow','orange','black','pink','brown','purple']
asignar_dep=[]
for row in labels_dep:
asignar_dep.append(colores[row])
# Clusters creados por KMeans tras aplicar variable tiempo dentro de pca
plt.scatter(X.RMS_X_AXIS_X100, X.SEISMIC_TIME_SOLS, c=asignar_dep, s=10)
plt.xlabel('RMS_X_AXIS_X100')
plt.ylabel('SEISMIC_TIME_SOLS')
plt.show()
```
# Conclusiones
* Usemos o no la variable tiempo en el modelo, estará explicitamente relacionado con los valores
* Viendo la matriz de correlaciones, vemos que no dependen del tiempo cuando se puede ver claramente que sí, esto es debido a que hay cierta relación no lineal entre variables. Una finalidad de nuestro análisis es encontrar este tipo de relaciones.
* Si no lo usamos para entrenar nuestro modelo Kmeans, el modelo es mas fino.
* Ahora tendriamos que estudiar las estadísticas para dar una categoría de los datos a cada TAG.
| github_jupyter |
```
# %%
import math
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
from matplotlib.animation import FuncAnimation
from scipy.stats import bernoulli
from svgpathtools import svg2paths
from svgpath2mpl import parse_path
# matplotlib parameters to ensure correctness of Chinese characters
plt.rcParams["font.family"] = 'sans-serif'
plt.rcParams['font.sans-serif']=['Arial Unicode MS', 'SimHei'] # Chinese font
plt.rcParams['axes.unicode_minus']=False # correct minus sign
plt.rcParams["font.size"] = 20
plt.rcParams["xtick.labelsize"] = 35
plt.rcParams["ytick.labelsize"] = 35
class UpdateDist:
def __init__(self, ax,ax1):
self.line, =ax.plot([],[],lw=5, color='r')
self.ax = ax
self.ax.set_xlim([0,0.14])
self.ax.set_ylim([0,80])
self.ax.set_ylabel("总检验平均次数z", fontsize=40)
self.ax.set_xlabel("化验阳性的概率p", fontsize=40)
self.ax.text(0.01,0.25,"k=4", transform=self.ax.transAxes, fontsize=35, color='black',)
# self.ax1.set_xlabel('Number of people tested', fontsize=20)
# self.ax1.set_ylabel('Test accuracy', fontsize=20)
self.ax.spines['top'].set_visible(False)
self.ax.spines['right'].set_visible(False)
self.line1, =ax1.plot([],[],lw=5, color='g')
self.ax1 = ax1
self.ax1.set_xlim([0,0.14])
self.ax1.set_ylim([0,35])
self.ax1.set_ylabel("分组人数k", fontsize=40)
self.ax1.set_xlabel("化验阳性的概率p", fontsize=40)
# self.ax1.set_xlabel('Number of people tested', fontsize=20)
# self.ax1.set_ylabel('Test accuracy', fontsize=20)
self.ax1.spines['top'].set_visible(False)
self.ax1.spines['right'].set_visible(False)
self.rects = ax1.barh([1,2,3], [0,0,0], )
# This vertical line represents the theoretical value, to
# which the plotted distribution should converge.
def __call__(self, i):
# This way the plot can continuously run and we just keep
# watching new realizations of the process
if i == 0:
self.success = 0
#for rect, h in zip(self.rects, [0,0]):
#rect.set_width(h)
self.line, = self.ax.plot([], [], lw=5, color='r')
self.line1, = self.ax1.plot([], [], lw=5, color='g')
# Choose success based on exceed a threshold with a uniform pick
# if np.random.rand(1,) < self.prob:
# self.success += 1
# y = beta_pdf(self.x, self.success + 1, (i - self.success) + 1)
# self.line.set_data(self.x, y)
if i <= 99:
# update curve
xdata, ydata = self.line.get_data()
xdata1, ydata1 = self.line1.get_data()
p=i*0.14/100
if len(xdata) == 0:
xdata = [0]
ydata = [25*(5-4*1**4)]
xdata1 = [0]
ydata1 = [37]
else:
xdata = np.append(xdata, p)
ydata = np.append(ydata,25*(5-4*(1-p)**4) )
xdata1 =np.append(xdata1, p)
ydata1 =np.append(ydata1, math.log(p,0.562))
self.line.set_data(xdata, ydata)
self.line1.set_data(xdata1, ydata1)
elif i==100:
self.ax1.plot([0,0.14], [4,4], lw=3, ls="--",color='black')
self.ax1.plot([0.10,0.10], [0,4], lw=3, ls="--",color='black')
self.ax1.text(0.01,0.14,"k=4", transform=self.ax1.transAxes, fontsize=35, color='black',)
self.ax1.text(0.73,0.01,"p=0.1", transform=self.ax1.transAxes, fontsize=35, color='black',)
self.ax.plot([0.1,0.1], [0,70], lw=3, ls="--",color='black')
self.ax.plot([0.1,0.14], [59,59], lw=3, ls="--",color='black')
self.ax.text(0.95,0.68,"Z=59", transform=self.ax.transAxes, fontsize=35, color='black',)
self.ax.text(0.73,0.01,"p=0.1", transform=self.ax.transAxes, fontsize=35, color='black',)
return self.rects
fig = plt.figure(figsize=(30,10),dpi=200)
#spec2 = gridspec.GridSpec(ncols=2, nrows=1, left=0.08, right=0.92, top=0.32, bottom=0.08, wspace=0.15, figure=fig)
ax1 = fig.add_subplot(1,2,2)
ax2 = fig.add_subplot(1,2,1)
ud = UpdateDist( ax1, ax2)
anim = FuncAnimation(fig, ud, frames=120, blit=True)
anim.save('curve_p.mp4', fps=10, dpi=200, codec='libx264', bitrate=-1, extra_args=['-pix_fmt', 'yuv420p'])
```
| github_jupyter |
# Image Colorization with U-Net and GAN Tutorial
**If you have already read the explanations, you can directly go to the code starting with heading: _1 - Implementing the paper - Our Baseline_**

One of the most exciting applications of deep learning is colorizing black and white images. This task needed a lot of human input and hardcoding several years ago but now the whole process can be done end-to-end with the power of AI and deep learning. You might think that you need huge amount of data or long training times to train your model from scratch for this task but in the last few weeks I worked on this and tried many different model architectures, loss functions, training strategies, etc. and finally developed an efficient strategy to train such a model, using the latest advances in deep learning, on a rather small dataset and with really short training times. In this article, I'm going to explain what I did to make this happen, including the code!, and the strategies that helped and also those that were not useful. Before that, I will explain the colorization problem and a give you a short review of what has been done in recent years. I'll assume you have basic knowledge about deep learning, GANs, and PyTorch library for the rest of the article. Let's begin!
## Introduction to colorization problem
Here I'm going to give you some basic knowledge that you may need to understand what the models do in the following codes.
### RGB vs L\*a\*b
As you might know, when we load an image, we get a rank-3 (height, width, color) array with the last axis containing the color data for our image. These data represent color in RGB color space and there are 3 numbers for each pixel indicating how much Red, Green, and Blue the pixel is. In the following image you can see that in the left part of the "main image" (the leftmost image) we have blue color so in the blue channel of the image, that part has higher values and has turned dark.

In L\*a\*b color space, we have again three numbers for each pixel but these numbers have different meanings. The first number (channel), L, encodes the Lightness of each pixel and when we visualize this channel (the second image in the row below) it appears as a black and white image. The \*a and \*b channels encode how much green-red and yellow-blue each pixel is, respectively. In the following image you can see each channel of L\*a\*b color space separately.

In all papers I studied and all codes I checked out on colorization on GitHub, people use L\*a\*b color space instead of RGB to train the models. There are a couple of reasons for this choice but I'll give you an intuition of why we make this choice. To train a model for colorization, we should give it a grayscale image and hope that it will make it colorful. When using L\*a\*b, we can give the L channel to the model (which is the grayscale image) and want it to predict the other two channels (\*a, \*b) and after its prediction, we concatenate all the channels and we get our colorful image. But if you use RGB, you have to first convert your image to grayscale, feed the grayscale image to the model and hope it will predict 3 numbers for you which is a way more difficult and unstable task due to the many more possible combinations of 3 numbers compared to two numbers. If we assume we have 256 choices (in a 8-bit unsigned integer image this is the real number of choices) for each number, predicting the three numbers for each of the pixels is choosing between 256³ combinations which is more than 16 million choices, but when predicting two numbers we have about 65000 choices (actually, we are not going to wildly choose these numbers like a classification task and I just wrote these numbers to give you an intuition).
## How to solve the problem
During the last few years, many different solutions have been proposed to colorize images by using deep learning. [_**Colorful Image Colorization**_](https://arxiv.org/abs/1603.08511) paper approached the problem as a classification task and they also considered the uncertainty of this problem (e.x. a car in the image can take on many different and valid colors and we cannot be sure about any color for it); however, another paper approached the problem as a regression task (with some more tweaks!). There are pros and cons to each approach but in this article, we are going to use a different strategy.
### The strategy we are going to use
[_**Image-to-Image Translation with Conditional Adversarial Networks**_](https://arxiv.org/abs/1611.07004) paper, which you may know by the name pix2pix, proposed a general solution to many image-to-image tasks in deep learning which one of those was colorization. In this approach two losses are used: L1 loss, which makes it a regression task, and an adversarial (GAN) loss, which helps to solve the problem in an unsupervised manner (by assigning the outputs a number indicating how "real" they look!).
In this tutorial, I will first implement what the authors did in the paper and then I will introduce a whole new generator model and some tweaks in the strategy of training which significantly helps reduce the size of needed dataset while getting amazing results. So stay tuned :)
### A deeper dive into GAN world
As mentioned earlier, we are going to build a GAN (a conditional GAN to be specific) and use an extra loss function, L1 loss. Let's start with the GAN.
As you might know, in a GAN we have a generator and a discriminator model which learn to solve a problem together. In our setting, the generator model takes a grayscale image (1-channel image) and produces a 2-channel image, a channel for \*a and another for \*b. The discriminator, takes these two produced channels and concatenates them with the input grayscale image and decides whether this new 3-channel image is fake or real. Of course the discriminator also needs to see some real images (3-channel images again in Lab color space) that are not produced by the generator and should learn that they are real.
So what about the "condition" we mentioned? Well, that grayscale image which both the generator and discriminator see is the condition that we provide to both models in our GAN and expect that the they take this condition into consideration.
Let's take a look at the math. Consider _**x**_ as the grayscale image, _**z**_ as the input noise for the generator, and _**y**_ as the 2-channel output we want from the generator (it can also represent the 2 color channels of a real image). Also, _**G**_ is the generator model and _**D**_ is the discriminator. Then the loss for our conditional GAN will be:

Notice that _**x**_ is given to both models which is the condition we introduce two both players of this game. Actually, we are not going to feed a "n" dimensional vector of random noise to the generator as you might expect but the noise is introduced in the form of dropout layers (there is something cool about it which you will read in the last section of the article) in the generator architecture.
### Loss function we optimize
The earlier loss function helps to produce good-looking colorful images that seem real, but to further help the models and introduce some supervision in our task, we combine this loss function with L1 Loss (you might know L1 loss as mean absolute error) of the predicted colors compared with the actual colors:

If we use L1 loss alone, the model still learns to colorize the images but it will be conservative and most of the time uses colors like "gray" or "brown" because when it doubts which color is the best, it takes the average and uses these colors to reduce the L1 loss as much as possible (it is similar to the blurring effect of L1 or L2 loss in super resolution task). Also, the L1 Loss is preferred over L2 loss (or mean squared error) because it reduces that effect of producing gray-ish images. So, our combined loss function will be:

where _**λ**_ is a coefficient to balance the contribution of the two losses to the final loss (of course the discriminator loss does not involve the L1 loss).
Okay. I think it's enough for theory! Let's get our hands dirty with the code! In the following section, **I first introduce the code to implement the paper** and in the section after that, **I will introduce a better strategy to get really amazing results in one or two hours of training and without needing huge amount of data!**
## 1 - Implementing the paper - Our Baseline
### 1.1- Loading Image Paths
The paper uses the whole ImageNet dataset (with 1.3 million images!) but here I'm using only 8,000 images from COCO dataset for training which I had available on my device. So our training set size is 0.6% of what was used in the paper!
You can use almost any dataset for this task as far as it contains many different scenes and locations which you hope it will learn to colorize. You can use ImageNet for example but you will only need 8000 of its images for this project.
```
import os
import glob
import time
import numpy as np
from PIL import Image
from pathlib import Path
from tqdm.notebook import tqdm
import matplotlib.pyplot as plt
from skimage.color import rgb2lab, lab2rgb
import torch
from torch import nn, optim
from torchvision import transforms
from torchvision.utils import make_grid
from torch.utils.data import Dataset, DataLoader
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
use_colab = None
```
### 1.1.x Preparing Colab for running the code
If you are opening this on **Google Colab** you can uncomment and run the following to install fastai. Almost all of the code in the tutorial is with **pure PyTorch**. We need fastai here only to download part of COCO dataset and in one other step in the second section of the tutorial.
Also make sure to set your runtime to **GPU** to be able to train the models much faster.
```
#!pip install fastai --upgrade
```
The following will download about 20,000 images from COCO dataset. Notice that **we are going to use only 8000 of them** for training. Also you can use any other dataset like ImageNet as long as it contains various scenes and locations.
```
# from fastai.data.external import untar_data, URLs
# coco_path = untar_data(URLs.COCO_SAMPLE)
# coco_path = str(coco_path) + "/train_sample"
# use_colab = True
if use_colab == True:
path = coco_path
else:
path = "Your path to the dataset"
paths = glob.glob(path + "/*.jpg") # Grabbing all the image file names
np.random.seed(123)
paths_subset = np.random.choice(paths, 10_000, replace=False) # choosing 1000 images randomly
rand_idxs = np.random.permutation(10_000)
train_idxs = rand_idxs[:8000] # choosing the first 8000 as training set
val_idxs = rand_idxs[8000:] # choosing last 2000 as validation set
train_paths = paths_subset[train_idxs]
val_paths = paths_subset[val_idxs]
print(len(train_paths), len(val_paths))
_, axes = plt.subplots(4, 4, figsize=(10, 10))
for ax, img_path in zip(axes.flatten(), train_paths):
ax.imshow(Image.open(img_path))
ax.axis("off")
```
Although we are using the same dataset and number of training samples, the exact 8000 images that you train your model on may vary (although we are seeding!) because the dataset here has only 20000 images with different ordering while I sampled 10000 images from the complete dataset.
### 1.2- Making Datasets and DataLoaders
I hope the code is self-explanatory. I'm resizing the images and flipping horizontally (flipping only if it is training set) and then I read an RGB image, convert it to Lab color space and separate the first (grayscale) channel and the color channels as my inputs and targets for the models respectively. Then I'm making the data loaders.
```
SIZE = 256
class ColorizationDataset(Dataset):
def __init__(self, paths, split='train'):
if split == 'train':
self.transforms = transforms.Compose([
transforms.Resize((SIZE, SIZE), Image.BICUBIC),
transforms.RandomHorizontalFlip(), # A little data augmentation!
])
elif split == 'val':
self.transforms = transforms.Resize((SIZE, SIZE), Image.BICUBIC)
self.split = split
self.size = SIZE
self.paths = paths
def __getitem__(self, idx):
img = Image.open(self.paths[idx]).convert("RGB")
img = self.transforms(img)
img = np.array(img)
img_lab = rgb2lab(img).astype("float32") # Converting RGB to L*a*b
img_lab = transforms.ToTensor()(img_lab)
L = img_lab[[0], ...] / 50. - 1. # Between -1 and 1
ab = img_lab[[1, 2], ...] / 110. # Between -1 and 1
return {'L': L, 'ab': ab}
def __len__(self):
return len(self.paths)
def make_dataloaders(batch_size=16, n_workers=4, pin_memory=True, **kwargs): # A handy function to make our dataloaders
dataset = ColorizationDataset(**kwargs)
dataloader = DataLoader(dataset, batch_size=batch_size, num_workers=n_workers,
pin_memory=pin_memory)
return dataloader
train_dl = make_dataloaders(paths=train_paths, split='train')
val_dl = make_dataloaders(paths=val_paths, split='val')
data = next(iter(train_dl))
Ls, abs_ = data['L'], data['ab']
print(Ls.shape, abs_.shape)
print(len(train_dl), len(val_dl))
```
### 1.3- Generator proposed by the paper
This one is a little complicated and needs explanation. This code implements a U-Net to be used as the generator of our GAN. The details of the code are out of the scope of this article but the important thing to understand is that it makes the U-Net from the middle part of it (down in the U shape) and adds down-sampling and up-sampling modules to the left and right of that middle module (respectively) at every iteration until it reaches the input module and output module. Look at the following image that I made from one of the images in the article to give you a better sense of what is happening in the code:

The blue rectangles show the order in which the related modules are built with the code. The U-Net we will build has more layers than what is depicted in this image but it suffices to give you the idea. Also notice in the code that we are going 8 layers down, so if we start with a 256 by 256 image, in the middle of the U-Net we will get a 1 by 1 (256 / 2⁸) image and then it gets up-sampled to produce a 256 by 256 image (with two channels). This code snippet is really exciting and I highly recommend to play with it to fully grasp what every line of it is doing.
```
class UnetBlock(nn.Module):
def __init__(self, nf, ni, submodule=None, input_c=None, dropout=False,
innermost=False, outermost=False):
super().__init__()
self.outermost = outermost
if input_c is None: input_c = nf
downconv = nn.Conv2d(input_c, ni, kernel_size=4,
stride=2, padding=1, bias=False)
downrelu = nn.LeakyReLU(0.2, True)
downnorm = nn.BatchNorm2d(ni)
uprelu = nn.ReLU(True)
upnorm = nn.BatchNorm2d(nf)
if outermost:
upconv = nn.ConvTranspose2d(ni * 2, nf, kernel_size=4,
stride=2, padding=1)
down = [downconv]
up = [uprelu, upconv, nn.Tanh()]
model = down + [submodule] + up
elif innermost:
upconv = nn.ConvTranspose2d(ni, nf, kernel_size=4,
stride=2, padding=1, bias=False)
down = [downrelu, downconv]
up = [uprelu, upconv, upnorm]
model = down + up
else:
upconv = nn.ConvTranspose2d(ni * 2, nf, kernel_size=4,
stride=2, padding=1, bias=False)
down = [downrelu, downconv, downnorm]
up = [uprelu, upconv, upnorm]
if dropout: up += [nn.Dropout(0.5)]
model = down + [submodule] + up
self.model = nn.Sequential(*model)
def forward(self, x):
if self.outermost:
return self.model(x)
else:
return torch.cat([x, self.model(x)], 1)
class Unet(nn.Module):
def __init__(self, input_c=1, output_c=2, n_down=8, num_filters=64):
super().__init__()
unet_block = UnetBlock(num_filters * 8, num_filters * 8, innermost=True)
for _ in range(n_down - 5):
unet_block = UnetBlock(num_filters * 8, num_filters * 8, submodule=unet_block, dropout=True)
out_filters = num_filters * 8
for _ in range(3):
unet_block = UnetBlock(out_filters // 2, out_filters, submodule=unet_block)
out_filters //= 2
self.model = UnetBlock(output_c, out_filters, input_c=input_c, submodule=unet_block, outermost=True)
def forward(self, x):
return self.model(x)
```
### 1.4- Discriminator
The architecture of our discriminator is rather straight forward. This code implements a model by stacking blocks of Conv-BatchNorm-LeackyReLU to decide whether the input image is fake or real. Notice that the first and last blocks do not use normalization and the last block has no activation function (it is embedded in the loss function we will use).
```
class PatchDiscriminator(nn.Module):
def __init__(self, input_c, num_filters=64, n_down=3):
super().__init__()
model = [self.get_layers(input_c, num_filters, norm=False)]
model += [self.get_layers(num_filters * 2 ** i, num_filters * 2 ** (i + 1), s=1 if i == (n_down-1) else 2)
for i in range(n_down)] # the 'if' statement is taking care of not using
# stride of 2 for the last block in this loop
model += [self.get_layers(num_filters * 2 ** n_down, 1, s=1, norm=False, act=False)] # Make sure to not use normalization or
# activation for the last layer of the model
self.model = nn.Sequential(*model)
def get_layers(self, ni, nf, k=4, s=2, p=1, norm=True, act=True): # when needing to make some repeatitive blocks of layers,
layers = [nn.Conv2d(ni, nf, k, s, p, bias=not norm)] # it's always helpful to make a separate method for that purpose
if norm: layers += [nn.BatchNorm2d(nf)]
if act: layers += [nn.LeakyReLU(0.2, True)]
return nn.Sequential(*layers)
def forward(self, x):
return self.model(x)
```
Let's take a look at its blocks:
```
PatchDiscriminator(3)
```
And its output shape:
```
discriminator = PatchDiscriminator(3)
dummy_input = torch.randn(16, 3, 256, 256) # batch_size, channels, size, size
out = discriminator(dummy_input)
out.shape
```
We are using a "Patch" Discriminator here. Okay, what is it?! In a vanilla discriminator, the model outputs one number (a scaler) which represents how much the model thinks the input (which is the whole image) is real (or fake). In a patch discriminator, the model outputs one number for every patch of say 70 by 70 pixels of the input image and for each of them decides whether it is fake or not separately. Using such a model for the task of colorization seems reasonable to me because the local changes that the model needs to make are really important and maybe deciding on the whole image as in vanilla discriminator cannot take care of the subtleties of this task. Here, the model's output shape is 30 by 30 but it does not mean that our patches are 30 by 30. The actual patch size is obtained when you compute the receptive field of each of these 900 (30 multiplied by 30) output numbers which in our case will be 70 by 70.
### 1.5- GAN Loss
This is a handy class we can use to calculate the GAN loss of our final model. In the __init__ we decide which kind of loss we're going to use (which will be "vanilla" in our project) and register some constant tensors as the "real" and "fake" labels. Then when we call this module, it makes an appropriate tensor full of zeros or ones (according to what we need at the stage) and computes the loss.
```
class GANLoss(nn.Module):
def __init__(self, gan_mode='vanilla', real_label=1.0, fake_label=0.0):
super().__init__()
self.register_buffer('real_label', torch.tensor(real_label))
self.register_buffer('fake_label', torch.tensor(fake_label))
if gan_mode == 'vanilla':
self.loss = nn.BCEWithLogitsLoss()
elif gan_mode == 'lsgan':
self.loss = nn.MSELoss()
def get_labels(self, preds, target_is_real):
if target_is_real:
labels = self.real_label
else:
labels = self.fake_label
return labels.expand_as(preds)
def __call__(self, preds, target_is_real):
labels = self.get_labels(preds, target_is_real)
loss = self.loss(preds, labels)
return loss
```
### 1.x Model Initialization
In the TowardsDataScince article, I didn't explain this function. Here is our logic to initialize our models. We are going to initialize the weights of our model with a mean of 0.0 and standard deviation of 0.02 which are the proposed hyperparameters in the article:
```
def init_weights(net, init='norm', gain=0.02):
def init_func(m):
classname = m.__class__.__name__
if hasattr(m, 'weight') and 'Conv' in classname:
if init == 'norm':
nn.init.normal_(m.weight.data, mean=0.0, std=gain)
elif init == 'xavier':
nn.init.xavier_normal_(m.weight.data, gain=gain)
elif init == 'kaiming':
nn.init.kaiming_normal_(m.weight.data, a=0, mode='fan_in')
if hasattr(m, 'bias') and m.bias is not None:
nn.init.constant_(m.bias.data, 0.0)
elif 'BatchNorm2d' in classname:
nn.init.normal_(m.weight.data, 1., gain)
nn.init.constant_(m.bias.data, 0.)
net.apply(init_func)
print(f"model initialized with {init} initialization")
return net
def init_model(model, device):
model = model.to(device)
model = init_weights(model)
return model
```
### 1.6- Putting everything together
This class brings together all the previous parts and implements a few methods to take care of training our complete model. Let's investigate it.
In the __init__ we define our generator and discriminator using the previous functions and classes we defined and we also initialize them with init_model function which I didn't explain here but you can refer to my GitHub repository to see how it works. Then we define our two loss functions and the optimizers of the generator and discriminator.
The whole work is being done in optimize method of this class. First and only once per iteration (batch of training set) we call the module's forward method and store the outputs in fake_color variable of the class.
Then, we first train the discriminator by using backward_D method in which we feed the fake images produced by generator to the discriminator (make sure to detach them from the generator's graph so that they act as a constant to the discriminator, like normal images) and label them as fake. Then we feed a batch of real images from training set to the discriminator and label them as real. We add up the two losses for fake and real and take the average and then call the backward on the final loss.
Now, we can train the generator. In backward_G method we feed the discriminator the fake image and try to fool it by assigning real labels to them and calculating the adversarial loss. As I mentioned earlier, we use L1 loss as well and compute the distance between the predicted two channels and the target two channels and multiply this loss by a coefficient (which is 100 in our case) to balance the two losses and then add this loss to the adversarial loss. Then we call the backward method of the loss.
```
class MainModel(nn.Module):
def __init__(self, net_G=None, lr_G=2e-4, lr_D=2e-4,
beta1=0.5, beta2=0.999, lambda_L1=100.):
super().__init__()
self.device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
self.lambda_L1 = lambda_L1
if net_G is None:
self.net_G = init_model(Unet(input_c=1, output_c=2, n_down=8, num_filters=64), self.device)
else:
self.net_G = net_G.to(self.device)
self.net_D = init_model(PatchDiscriminator(input_c=3, n_down=3, num_filters=64), self.device)
self.GANcriterion = GANLoss(gan_mode='vanilla').to(self.device)
self.L1criterion = nn.L1Loss()
self.opt_G = optim.Adam(self.net_G.parameters(), lr=lr_G, betas=(beta1, beta2))
self.opt_D = optim.Adam(self.net_D.parameters(), lr=lr_D, betas=(beta1, beta2))
def set_requires_grad(self, model, requires_grad=True):
for p in model.parameters():
p.requires_grad = requires_grad
def setup_input(self, data):
self.L = data['L'].to(self.device)
self.ab = data['ab'].to(self.device)
def forward(self):
self.fake_color = self.net_G(self.L)
def backward_D(self):
fake_image = torch.cat([self.L, self.fake_color], dim=1)
fake_preds = self.net_D(fake_image.detach())
self.loss_D_fake = self.GANcriterion(fake_preds, False)
real_image = torch.cat([self.L, self.ab], dim=1)
real_preds = self.net_D(real_image)
self.loss_D_real = self.GANcriterion(real_preds, True)
self.loss_D = (self.loss_D_fake + self.loss_D_real) * 0.5
self.loss_D.backward()
def backward_G(self):
fake_image = torch.cat([self.L, self.fake_color], dim=1)
fake_preds = self.net_D(fake_image)
self.loss_G_GAN = self.GANcriterion(fake_preds, True)
self.loss_G_L1 = self.L1criterion(self.fake_color, self.ab) * self.lambda_L1
self.loss_G = self.loss_G_GAN + self.loss_G_L1
self.loss_G.backward()
def optimize(self):
self.forward()
self.net_D.train()
self.set_requires_grad(self.net_D, True)
self.opt_D.zero_grad()
self.backward_D()
self.opt_D.step()
self.net_G.train()
self.set_requires_grad(self.net_D, False)
self.opt_G.zero_grad()
self.backward_G()
self.opt_G.step()
```
### 1.xx Utility functions
These functions were nor included in the explanations of the TDS article. These are just some utility functions to log the losses of our network and also visualize the results during training. So here you can check them out:
```
class AverageMeter:
def __init__(self):
self.reset()
def reset(self):
self.count, self.avg, self.sum = [0.] * 3
def update(self, val, count=1):
self.count += count
self.sum += count * val
self.avg = self.sum / self.count
def create_loss_meters():
loss_D_fake = AverageMeter()
loss_D_real = AverageMeter()
loss_D = AverageMeter()
loss_G_GAN = AverageMeter()
loss_G_L1 = AverageMeter()
loss_G = AverageMeter()
return {'loss_D_fake': loss_D_fake,
'loss_D_real': loss_D_real,
'loss_D': loss_D,
'loss_G_GAN': loss_G_GAN,
'loss_G_L1': loss_G_L1,
'loss_G': loss_G}
def update_losses(model, loss_meter_dict, count):
for loss_name, loss_meter in loss_meter_dict.items():
loss = getattr(model, loss_name)
loss_meter.update(loss.item(), count=count)
def lab_to_rgb(L, ab):
"""
Takes a batch of images
"""
L = (L + 1.) * 50.
ab = ab * 110.
Lab = torch.cat([L, ab], dim=1).permute(0, 2, 3, 1).cpu().numpy()
rgb_imgs = []
for img in Lab:
img_rgb = lab2rgb(img)
rgb_imgs.append(img_rgb)
return np.stack(rgb_imgs, axis=0)
def visualize(model, data, save=True):
model.net_G.eval()
with torch.no_grad():
model.setup_input(data)
model.forward()
model.net_G.train()
fake_color = model.fake_color.detach()
real_color = model.ab
L = model.L
fake_imgs = lab_to_rgb(L, fake_color)
real_imgs = lab_to_rgb(L, real_color)
fig = plt.figure(figsize=(15, 8))
for i in range(5):
ax = plt.subplot(3, 5, i + 1)
ax.imshow(L[i][0].cpu(), cmap='gray')
ax.axis("off")
ax = plt.subplot(3, 5, i + 1 + 5)
ax.imshow(fake_imgs[i])
ax.axis("off")
ax = plt.subplot(3, 5, i + 1 + 10)
ax.imshow(real_imgs[i])
ax.axis("off")
plt.show()
if save:
fig.savefig(f"colorization_{time.time()}.png")
def log_results(loss_meter_dict):
for loss_name, loss_meter in loss_meter_dict.items():
print(f"{loss_name}: {loss_meter.avg:.5f}")
```
### 1.7- Training function
I hope this code is self-explanatory. Every epoch takes about 4 minutes on not a powerful GPU as Nvidia P5000. So if you are using 1080Ti or higher, it will be much faster.
```
def train_model(model, train_dl, epochs, display_every=200):
data = next(iter(val_dl)) # getting a batch for visualizing the model output after fixed intrvals
for e in range(epochs):
loss_meter_dict = create_loss_meters() # function returing a dictionary of objects to
i = 0 # log the losses of the complete network
for data in tqdm(train_dl):
model.setup_input(data)
model.optimize()
update_losses(model, loss_meter_dict, count=data['L'].size(0)) # function updating the log objects
i += 1
if i % display_every == 0:
print(f"\nEpoch {e+1}/{epochs}")
print(f"Iteration {i}/{len(train_dl)}")
log_results(loss_meter_dict) # function to print out the losses
visualize(model, data, save=False) # function displaying the model's outputs
model = MainModel()
train_model(model, train_dl, 100)
```
Every epoch takes about 3 to 4 minutes on Colab. After about 20 epochs you should see some reasonable results.
Okay. I let the model train for some longer (about 100 epochs). Here are the results of our baseline model:

As you can see, although this baseline model has some basic understanding of some most common objects in images like sky, trees, … its output is far from something appealing and it cannot decide on the color of rare objects. It also displays some color spillovers and circle-shaped mass of color (center of first image of second row) which is not good at all. So, it seems like that with this small dataset we cannot get good results with this strategy. **Therefore, we change our strategy!**
## 2- A new strategy - the final model
Here is the focus of this article and where I'm going to explain what I did to overcome the last mentioned problem. Inspired by an idea in Super Resolution literature, I decided to pretrain the generator separately in a supervised and deterministic manner to avoid the problem of "the blind leading the blind" in the GAN game where neither generator nor discriminator knows anything about the task at the beginning of training.
Actually I use pretraining in two stages: 1- The backbone of the generator (the down sampling path) is a pretrained model for classification (on ImageNet) 2- The whole generator will be pretrained on the task of colorization with L1 loss.
In fact, I'm going to use a pretrained ResNet18 as the backbone of my U-Net and to accomplish the second stage of pretraining, we are going to train the U-Net on our training set with only L1 Loss. Then we will move to the combined adversarial and L1 loss, as we did in the previous section.
### 2.1- Using a new generator
Building a U-Net with a ResNet backbone is not something trivial so I'll use fastai library's Dynamic U-Net module to easily build one. You can simply install fastai with pip or conda (if you haven't already at the beginning of the tutorial). Here's the link to the [documentation](https://docs.fast.ai/).
```
from fastai.vision.learner import create_body
from torchvision.models.resnet import resnet18
from fastai.vision.models.unet import DynamicUnet
def build_res_unet(n_input=1, n_output=2, size=256):
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
body = create_body(resnet18, pretrained=True, n_in=n_input, cut=-2)
net_G = DynamicUnet(body, n_output, (size, size)).to(device)
return net_G
```
That's it! With just these few lines of code you can build such a complex model easily. create_body function loads the pretrained weights of the ResNet18 architecture and cuts the model to remove the last two layers (GlobalAveragePooling and a Linear layer for the ImageNet classification task). Then, DynamicUnet uses this backbone to build a U-Net with the needed output channels (2 in our case) and with an input size of 256.
### 2.2 Pretraining the generator for colorization task
```
def pretrain_generator(net_G, train_dl, opt, criterion, epochs):
for e in range(epochs):
loss_meter = AverageMeter()
for data in tqdm(train_dl):
L, ab = data['L'].to(device), data['ab'].to(device)
preds = net_G(L)
loss = criterion(preds, ab)
opt.zero_grad()
loss.backward()
opt.step()
loss_meter.update(loss.item(), L.size(0))
print(f"Epoch {e + 1}/{epochs}")
print(f"L1 Loss: {loss_meter.avg:.5f}")
net_G = build_res_unet(n_input=1, n_output=2, size=256)
opt = optim.Adam(net_G.parameters(), lr=1e-4)
criterion = nn.L1Loss()
pretrain_generator(net_G, train_dl, opt, criterion, 20)
#torch.save(net_G.state_dict(), "res18-unet.pt")
```
With this simple function, we pretrain the generator for 20 epochs and then we save its weights. This will take an hour on Colab. In the following section, we will use this model as the generator for our GAN and train the whole network as before:
### 2.3 Putting everything together, again!
If you want to train the model yourself, run the following cell. Instead, if you want to use the pretrained weights, skip the cell and run the one after that.
```
net_G = build_res_unet(n_input=1, n_output=2, size=256)
net_G.load_state_dict(torch.load("res18-unet.pt", map_location=device))
model = MainModel(net_G=net_G)
train_model(model, train_dl, 20)
```
Here I'm first loading the saved weights for the generator (which you have saved in the previous section) and then I'm using this model as the generator in our MainModel class which prevents it from randomly initializing the generator. Then we train the model for 10 to 20 epochs! (compare it to the 100 epochs of the previous section when we didn't use pretraining). Each epoch takes about 3 to 4 minutes on Colab
If you are on Colab and want to use the pretrained weights, run the following cells which download the weights from my google drive and loads it to the model:
```
# !gdown --id 1lR6DcS4m5InSbZ5y59zkH2mHt_4RQ2KV
# net_G = build_res_unet(n_input=1, n_output=2, size=256)
# net_G.load_state_dict(torch.load("res18-unet.pt", map_location=device))
# model = MainModel(net_G=net_G)
# model.load_state_dict(torch.load("final_model_weights.pt", map_location=device))
```
Now, I will show the results of this final model on the test set (the black and white images that it has never seen during training) including the main title image of this article at the very beginning:

Left: Input black & white images from test set | Right: the colorized outputs by the final model of this tutorial
---

Left: Input black & white images from test set | Right: the colorized outputs by the final model of this tutorial
---

Left: Input black & white images from test set | Right: the colorized outputs by the final model of this tutorial
---
## An accidental finding: You can safely remove Dropout!
Remember that when I was explaining the theory of conditional GAN in the beginning of this article, I said that the source of the noise in the architecture of the generator proposed by authors of the paper was the dropout layers. However, when I investigated the U-Net we built with the help of fastai, I did not find any dropout layers in there! Actually I first trained the final model and got the results and then I investigated the generator and found this out.
So, was the adversarial training useless? If there is no noise, how possibly the generator can have a creative effect on the output? Is it possible that the input grayscale image to the generator plays the role of noise as well? These were my exact questions at the time.
Therefor, I decided to email Dr. Phillip Isola, the first author of the same paper we implemented here, and he kindly answered these questions. According to what he said, this conditional GAN can still work without dropout but the outputs will be more deterministic because of the lack of that noise; however, there is still enough information in that input grayscale image which enables the generator to produce compelling outputs.
Actually, I saw this in practice that the adversarial training was helpful indeed. In the next and last section, I'm going to compare the results of the pretrained U-Net with no adversarial training against the final outputs we got with adversarial training.
## Comparing the results of the pretrained U-Net with and without adversarial training
One of the cool thing I found in my experiments was that the U-Net we built with the ResNet18 backbone is already awesome in colorizing images after pretraining with L1 Loss only (a step before the final adversarial training). But, the model is still conservative and encourages using gray-ish colors when it is not sure about what the object is or what color it should be. However, it performs really awesome for common scenes in the images like sky, tree, grass, etc.
Here I show you the outputs of the U-Net without adversarial training and U-Net with adversarial training to better depict the significant difference that the adversarial training is making in our case:

(Left: pretrained U-Net without adversarial training | Right: pretrained U-Net with adversarial training)
---
You can also see the GIF below to observe the difference between the images better:

(animation of the last two images to better see the significant difference that adversarial training is making)
---
## Final words
This project was full of important lessons for myself. I spent a lot of time during the last month to implement lots of different papers each with different strategies and it took quite a while and after A LOT of failures that I could come up with this method of training. Now you can see that how pretraining the generator significantly helped the model and improved the results.
I also learned that some observations, although at first feeling like a bad mistake of yours, are worth paying attention to and further investigation; like the case of dropout in this project. Thanks to the helpful community of deep learning and AI, you can easily ask experts and get the answer you need and become more confidant in what you were just guessing.
I want to thank the authors of this wonderful paper for their awesome work and also [the great GitHub repository of this paper](https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix) from which I borrowed some of the codes (with modification and simplification). I truly love the community of computer science and AI and all their hard work to improve the field and also make their contributions available to all. I'm happy to be a tiny part of this community.
| github_jupyter |
## 定义卷积神经网络(CNN)
查看正在使用的数据之后,了解图像与关键点的形状,接下来,就可以定义一个机器人可以从这些数据中 *学习*的卷积神经网络。
在这个notebook和`models.py`中,你的任务是:
1. 定义一个CNN,把图像作为输入,把关键点作为输出
2. 与以前一样,构造转换后的FaceKeypointsDataset
3. 使用训练数据训练这个CNN,并跟踪损失
4. 查看训练模型对测试数据的执行情况
5. 如有必要,请修改CNN结构并模拟超参数,使其*表现良好* **\***
**\*** 什么是*表现良好*?
“表现良好”意味着该模型的损失在训练期间有所降低,**而且**该模型应用于测试图像数据时,会产生与每个人脸的真实关键点紧密匹配的关键点。你会在这个notebook中看到这个例子。
---
## CNN架构
回想一下,CNN是由下列几种类型的层定义的:
* 卷积层
* 最大池化层
* 全连接层
你需要使用上述层,而且我们建议你添加多个卷积层以及可能防止过度拟合的dropout层等。此外,你还可以查看一些有关关键点检测的文献,如 [这篇论文](https://arxiv.org/pdf/1710.00977.pdf),帮助你确定该网络的结构。
### TODO: 在`models.py`文件中定义你的模型
此文件大部分为空,但其中包含预期的名称和一些用于创建模型的TODO事项。
---
## PyTorch神经网络
要在PyTorch中定义神经网络,你可以在函数`__init__`中定义一个模型的各个层,并定义一个网络的前馈行为,该网络会在函数`forward`中使用这些初始化的层,而该函数会接收输入图像张量`x`。此Net类的结构如下所示,并由你来填充。
注意:在训练期间,PyTorch能够通过跟踪网络的前馈行为并使用autograd来计算该网络中权重的更新来执行反向传播。
#### 在` __init__`中定义层
提醒一下,卷积层与池化层可以像这样来定义(在`__init__`中):
```
# 1 input image channel (for grayscale images), 32 output channels/feature maps, 3x3 square convolution kernel
self.conv1 = nn.Conv2d(1, 32, 3)
# maxpool that uses a square window of kernel_size=2, stride=2
self.pool = nn.MaxPool2d(2, 2)
```
#### 引用`forward`中的层
然后在这样的`forward`函数中引用,其中卷积1层在应用最大池化之前应用了ReLu激活函数:
```
x = self.pool(F.relu(self.conv1(x)))
```
最佳做法是把权重将在训练过程中发生变化的任何层防治在`__init__`中,并在`forward`函数中引用它们。所有始终以相同方式运行的层或函数(例如预定义的激活函数)应*只* 出现在`forward` 函数中。
#### 为什么要用models.py文件
你的任务是在`models.py`文件中定义该网络,便于在此项目目录中的不同notebook中按名称保存和加载你定义的任何模型。例如,通过在`models.py`中定义名为`Net`的CNN类,通过简单地导入该类并实例化模型,就可以在此notebook和其他notebook中创建相同的体系结构:
```
from models import Net
net = Net()
# load the data if you need to; if you have already loaded the data, you may comment this cell out
# -- DO NOT CHANGE THIS CELL -- #
!mkdir ./data
!wget -P ./data/ https://s3.amazonaws.com/video.udacity-data.com/topher/2018/May/5aea1b91_train-test-data/train-test-data.zip
!unzip -n ./data/train-test-data.zip -d /data
```
<div class="alert alert-info">**注意:**工作区会在持续30分钟的不活动状态后,自动关闭连接,包括训练时出现不活动状态。使用下面的代码段可以在训练期间保持工作区的活动状态。下面导入了active_session上下文管理器。
</div>
```
from workspace_utils import active_session
with active_session():
train_model(num_epochs)
# import the usual resources
import matplotlib.pyplot as plt
import numpy as np
# import utilities to keep workspaces alive during model training
from workspace_utils import active_session
# watch for any changes in model.py, if it changes, re-load it automatically
%load_ext autoreload
%autoreload 2
## TODO: Define the Net in models.py
import torch
import torch.nn as nn
import torch.nn.functional as F
## TODO: Once you've define the network, you can instantiate it
# one example conv layer has been provided for you
from models import Net
net = Net()
print(net)
```
## 转换数据集
为训练做准备,你还需要创建一个图像和关键点的转换数据集。
### TODO: 定义一个数据转换
在PyTorch中,卷积神经网络需要一个大小一致的torch图像作为输入。为了进行有效的训练,以及在训练过程中该模型的损失不会放大,我们还建议你对输入图像和关键点进行归一化。必要的转换已在`data_load.py`中定义,你无需再做修改。另外,你可以看一下这个文件,你会在该文件中看到Notebook 1中定义和应用的相同转换。
要定义下面的数据转换,请使用以下[组合](http://pytorch.org/tutorials/beginner/data_loading_tutorial.html#compose-transforms) :
1. 重新缩放和/或裁剪数据,最终需要一个方形图像(建议大小为224x224px)
2. 归一化图像和关键点;将每个RGB图像转换为颜色范围为[0,1]的灰度图像,并将给定关键点转换为[-1,1]的范围
3. 将这些图像和关键点转换为张量
这些转换已在`data_load.py`中定义,但是否要在下面调用它们并创建一个`data_transform`,这都取决于你。**该转换将应用于训练数据,以及稍后的测试数据**。这样将改变显示这些图像和关键点的方式,但这些步骤对于高效训练来说非常重要。
需要说明的一点是,如果你想要执行数据增强(在此项目中是可选的),并随机旋转或移动这些图像,方形图像大小将会很有用,将224x224图像旋转90度就会产生相同的输出形状。
```
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
# the dataset we created in Notebook 1 is copied in the helper file `data_load.py`
from data_load import FacialKeypointsDataset
# the transforms we defined in Notebook 1 are in the helper file `data_load.py`
from data_load import Rescale, RandomCrop, Normalize, ToTensor
## TODO: define the data_transform using transforms.Compose([all tx's, . , .])
# order matters! i.e. rescaling should come before a smaller crop
data_transform = transforms.Compose([Rescale(250),
RandomCrop(224),
Normalize(),
ToTensor()])
# testing that you've defined a transform
assert(data_transform is not None), 'Define a data_transform'
# create the transformed dataset
transformed_dataset = FacialKeypointsDataset(csv_file='data/training_frames_keypoints.csv',
root_dir='data/training/',
transform=data_transform)
print('Number of images: ', len(transformed_dataset))
# iterate through the transformed dataset and print some stats about the first few samples
for i in range(4):
sample = transformed_dataset[i]
print(i, sample['image'].size(), sample['keypoints'].size())
```
## 批处理并加载数据
定义了转换数据集之后,接下来,我们可以使用PyTorch的DataLoader类来批量加载任意大小的训练数据,也可以对训练模型的数据进行置乱处理。你可以在 [本文档](http://pytorch.org/docs/master/data.html)中阅读有关DataLoader参数的更多信息。
#### 批量大小
确定用于训练模型的最合适的批量是多少。小批量与大批量都要试一试,并注意在模型训练时损失会如何减少。批量过大可能会导致模型在训练时崩溃和/或内存不足。
**Windows用户需要注意:**请将`num_workers`改为0,否则可能会遇到DataLoader失效的问题。
```
# load training data in batches
batch_size = 16
train_loader = DataLoader(transformed_dataset,
batch_size=batch_size,
shuffle=True,
num_workers=0)
```
## 训练之前
看一下这个模型在训练之前的表现。你应该会看到,它预测的关键点从一个点开始,并且与人脸上的关键点根本不匹配!你可以把此行为可视化,并在训练后将其与模型进行比较,还可以查看该模型是如何改进的。
#### 加载测试数据集
此模型之前*没有*见过这个测试数据集,这就是说,它没有使用这些图像进行过训练。在这里,我们将加载此测试数据,并在训练前后,查看你的模型在此数据集上的表现效果如何!
为了可视化这些测试数据,我们必须要做一些非转换步骤,将图像转换为张量的python图像,并将关键点重新转换回可识别的范围。
```
# load in the test data, using the dataset class
# AND apply the data_transform you defined above
# create the test dataset
test_dataset = FacialKeypointsDataset(csv_file='data/test_frames_keypoints.csv',
root_dir='data/test/',
transform=data_transform)
# load test data in batches
batch_size = 8
test_loader = DataLoader(test_dataset,
batch_size=batch_size,
shuffle=False,
num_workers=0)
```
## 将模型应用于测试样本
要在测试数据样本上测试模型,你必须执行以下步骤:
1. 从样本中提取图像和实际真值关键点
2. 将图像隐藏在变量中,便于你的网络将其作为输入处理,并跟踪图像在该网络中移动时发生的变化。
3. 确保图像是模型所需的FloatTensor。
4. 通过网络向前传递图像,获得预测的输出关键点。
此函数测试的是该网络在第一批测试数据上的执行情况。它会返回图像、转换图像、预测由模型产生的关键点以及实际真值关键点。
```
# test the model on a batch of test images
def net_sample_output():
# iterate through the test dataset
for i, sample in enumerate(test_loader):
# get sample data: images and ground truth keypoints
images = sample['image']
key_pts = sample['keypoints']
# convert images to FloatTensors
images = images.type(torch.FloatTensor)
# forward pass to get net output
output_pts = net(images)
# reshape to batch_size x 68 x 2 pts
output_pts = output_pts.view(output_pts.size()[0], 68, -1)
# break after first image is tested
if i == 0:
return images, output_pts, key_pts
```
#### 调试技巧
如果此处出现尺寸或维度错误,请确保你的网络输出预期数量的关键点!或者,如果收到Tensor类型的错误,请考虑将数据转换为float类型的上述代码进行更改,float类型为:`images = images.type(torch.FloatTensor)`。
```
# call the above function
# returns: test images, test predicted keypoints, test ground truth keypoints
test_images, test_outputs, gt_pts = net_sample_output()
# print out the dimensions of the data to see if they make sense
print(test_images.data.size())
print(test_outputs.data.size())
print(gt_pts.size())
```
## 将预测的关键点可视化
让模型生成一些预测的输出关键点之后,就可以用一种类似于我们之前显示这些数据的方式来显示这些点,只是这一次,要显示这些点,我们必须“取消转换”图像/关键点数据。
请注意,我已经定义了一个*新*函数`show_all_keypoints`,它会显示灰度图像、其预测的关键点以及其实际真值关键点(如果提供的话)。
```
def show_all_keypoints(image, predicted_key_pts, gt_pts=None):
"""Show image with predicted keypoints"""
# image is grayscale
plt.imshow(image, cmap='gray')
plt.scatter(predicted_key_pts[:, 0], predicted_key_pts[:, 1], s=20, marker='.', c='m')
# plot ground truth points as green pts
if gt_pts is not None:
plt.scatter(gt_pts[:, 0], gt_pts[:, 1], s=20, marker='.', c='g')
```
#### 非转换
接下来,你会看到一个辅助函数,即`visualize_output`,它会接收一批图像、预测关键点以及实际真值关键点,并显示一组图像及其真实/预测关键点。
此函数的主要作用是获取批量图像和关键点数据(CNN的输入和输出),并将它们转换为numpy图像和非归一化关键点(x,y),从而进行正常显示。非转换过程将关键点和图像转换为来自Tensors的numpy数组,*此外*, 它撤消了Normalize()转换中完成的关键点归一化。但前提是我们假设,你在载测试数据时应用了这些转换。
```
# visualize the output
# by default this shows a batch of 10 images
def visualize_output(test_images, test_outputs, gt_pts=None, batch_size=8):
for i in range(batch_size):
plt.figure(figsize=(20,10))
ax = plt.subplot(1, batch_size, i+1)
# un-transform the image data
image = test_images[i].data # get the image from it's Variable wrapper
image = image.numpy() # convert to numpy array from a Tensor
image = np.transpose(image, (1, 2, 0)) # transpose to go from torch to numpy image
# un-transform the predicted key_pts data
predicted_key_pts = test_outputs[i].data
predicted_key_pts = predicted_key_pts.numpy()
# undo normalization of keypoints
predicted_key_pts = predicted_key_pts*50.0+100
# plot ground truth points for comparison, if they exist
ground_truth_pts = None
if gt_pts is not None:
ground_truth_pts = gt_pts[i]
ground_truth_pts = ground_truth_pts*50.0+100
# call show_all_keypoints
show_all_keypoints(np.squeeze(image), predicted_key_pts, ground_truth_pts)
plt.axis('off')
plt.show()
# call it
visualize_output(test_images, test_outputs, gt_pts, batch_size=8)
```
## 训练
#### 损失函数
训练一个用于预测关键点的网络与训练一个用于预测类的网络不同。你可能希望选择适合回归的损失函数,而不是输出类的分布并使用交交叉熵损失函数,因为损失函数可以用于直接比较预测值和目标值。有关各种损失函数(如MSE或L1 / SmoothL1损失),请阅读 [本文档](http://pytorch.org/docs/master/_modules/torch/nn/modules/loss.html)中的内容。
### TODO: 定义损失与优化
接下来,你需要通过定义损失函数和优化程序来定义模型的训练方式。
---
```
## TODO: Define the loss and optimization
import torch.optim as optim
criterion = nn.SmoothL1Loss()
optimizer = optim.Adam(net.parameters(), lr=0.001)
scheduler = optim.lr_scheduler.ExponentialLR(optimizer, gamma=0.9)
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
device
```
## 训练与初步观察
现在,你要使用大量epoch,从`train_loader`中训练你的批量训练数据。
为了快速观察你的模型是如何训练并决定是否应该修改它的结构或超参数,我们建议你最开始的时候使用一个或两个epoch。训练时,请注意观察模型的损失会如何随着时间的推移而变化:例如,它会先快速减少然后再减慢吗?或者起初会在一段时间后出现减少?如果更改了训练数据的批量大小或修改损失函数,会发生什么变化?
在使用多个epoch进行训练并创建最终模型之前,使用这些初始观察值对模型进行更改并确定一个最佳架构。
```
def train_net(net, train_loader, n_epochs, criterion, optimizer, scheduler, device='cpu'):
# prepare the net for training
net.to(device)
net.train()
for epoch in range(n_epochs): # loop over the dataset multiple times
running_loss = 0.0
# train on batches of data, assumes you already have train_loader
for batch_i, data in enumerate(train_loader):
# get the input images and their corresponding labels
images = data['image']
key_pts = data['keypoints']
# flatten pts
key_pts = key_pts.view(key_pts.size(0), -1)
# convert variables to floats for regression loss
key_pts = key_pts.type(torch.FloatTensor)
images = images.type(torch.FloatTensor)
key_pts = key_pts.to(device)
images = images.to(device)
# forward pass to get outputs
output_pts = net(images)
# calculate the loss between predicted and target keypoints
loss = criterion(output_pts, key_pts)
# zero the parameter (weight) gradients
optimizer.zero_grad()
# backward pass to calculate the weight gradients
loss.backward()
# update the weights
optimizer.step()
#scheduler.step()
# print loss statistics
running_loss += loss.item()
if batch_i % 25 == 24: # print every 50 batches
print('Epoch: {}, Batch: {}, Avg. Loss: {}'.format(epoch + 1, batch_i+1, running_loss/10))
running_loss = 0.0
scheduler.step()
print('epoch:{},lr:{}'.format(epoch + 1, scheduler.get_lr()[0]))
print('Finished Training')
from models import Net
import time
# train your network
batch_size = 16
train_loader = DataLoader(transformed_dataset,
batch_size=batch_size,
shuffle=True,
num_workers=0)
net = Net()
criterion = nn.SmoothL1Loss()
optimizer = optim.Adam(net.parameters(), lr=0.001)
scheduler = optim.lr_scheduler.ExponentialLR(optimizer, gamma=0.9)
n_epochs = 20 # start small, and increase when you've decided on your model structure and hyperparams
start = time.time()
# this is a Workspaces-specific context manager to keep the connection
# alive while training your model, not part of pytorch
#with active_session():
# train_net(n_epochs)
train_net(net, train_loader, n_epochs, criterion, optimizer, scheduler, device)
end = time.time()
runing_time = end - start
print('Train time is {:.0f}m {:.0f}s'.format(runing_time//60,runing_time%60))
```
## 测试数据
了解你的模型在之前未见过的测试数据上的表现如何。我们已经对测试数据进行加载与转换,这一点类似于与训练数据时的做法类似。接下来,在这些图像上运行已被训练的模型,查看其生成的关键点类型。你应该能够观察到你的模型是否拟合了它看到的每个新人脸,这些点是否是随机分布的,以及这些点实际上是否过度拟合了训练数据而没有进行归纳。
```
# get a sample of test data again
net.eval()
net.to('cpu')
test_images, test_outputs, gt_pts = net_sample_output()
print(test_images.data.size())
print(test_outputs.data.size())
print(gt_pts.size())
## TODO: visualize your test output
# you can use the same function as before, by un-commenting the line below:
visualize_output(test_images, test_outputs, gt_pts, batch_size=8)
```
找到了一个或两个表现良好的模型后,保存你的模型,这样你就可以加载它并在以后使用它了!
在这里,你需要保存模型,但请**在提交项目之前删除任何检查点和已保存的模型**,否则你的工作区可能会因为太大而无法提交。
```
## TODO: change the name to something uniqe for each new model
model_dir = 'saved_models/'
model_name = 'keypoints_model_1.pth'
# after training, save your model parameters in the dir 'saved_models'
torch.save(net.state_dict(), model_dir+model_name)
```
完成对一个表现良好的模型的训练后,请回答以下问题,以便我们对你的训练和架构选择过程有一些了解。要通过此项目,你需要回答下列所有问题。
### 问题1:你选择了哪些优化和损失函数?为什么会这样选择?
**答案**:优化器选择的是Adam,损失函数是SmoothL1Loss
优化器的选择:Adam在随机梯度算法基础上,增加了动量计算;结合了Adagrad善于处理稀疏梯度和RMSprop善于处理非平稳目标的优点;对内存需求较小;为不同的参数计算不同的自适应学习率<br>
损失函数的选择:首先此回归模型最后拟合的是136个关键点,为稀疏数据,故选择L1Loss;而SmoothL1Loss相比于L1损失函数,可以收敛得更快
### 问题2:最开始,你的网络架构是什么样的?在尝试不同的架构时,又做了怎样的修改?为避免过度拟合数据,你是否决定添加了更多卷积层或其他层?
**答案**: 最开始网络架构为:<br>
Net(<br>
  (conv1): Conv2d(1, 32, kernel_size=(5, 5), stride=(1, 1))<br>
  (pool): MaxPool2d(kernel_size=2, stride=2, padding=0, dilation=1, ceil_mode=False)<br>
  (conv2): Conv2d(32, 64, kernel_size=(5, 5), stride=(1, 1))<br>
  (conv3): Conv2d(64, 128, kernel_size=(5, 5), stride=(1, 1))<br>
  (conv4): Conv2d(128, 256, kernel_size=(5, 5), stride=(1, 1))<br>
  (conv5): Conv2d(256, 512, kernel_size=(5, 5), stride=(1, 1))<br>
  (fc1): Linear(in_features=4608, out_features=1024, bias=True)<br>
  (fc1_drop): Dropout(p=0.4, inplace=False)<br>
  (fc2): Linear(in_features=1024, out_features=136, bias=True)<br>
)<br>
各卷积层后接最大化池化层pool,全连接层fc1后接fc1_drop<br>
修改后的架构为:<br>
Net(<br>
  (conv1): Conv2d(1, 32, kernel_size=(5, 5), stride=(1, 1))<br>
  (pool): MaxPool2d(kernel_size=2, stride=2, padding=0, dilation=1, ceil_mode=False)<br>
  (conv2): Conv2d(32, 64, kernel_size=(5, 5), stride=(1, 1))<br>
  (conv3): Conv2d(64, 128, kernel_size=(5, 5), stride=(1, 1))<br>
  (conv4): Conv2d(128, 256, kernel_size=(5, 5), stride=(1, 1))<br>
  (conv5): Conv2d(256, 512, kernel_size=(5, 5), stride=(1, 1))<br>
  (conv1_drop): Dropout(p=0.1, inplace=False)<br>
  (conv2_drop): Dropout(p=0.2, inplace=False)<br>
  (conv3_drop): Dropout(p=0.25, inplace=False)<br>
  (conv4_drop): Dropout(p=0.3, inplace=False)<br>
  (conv5_drop): Dropout(p=0.35, inplace=False)<br>
  (fc1): Linear(in_features=4608, out_features=1024, bias=True)<br>
  (fc1_drop): Dropout(p=0.4, inplace=False)<br>
  (fc2): Linear(in_features=1024, out_features=136, bias=True)<br>
)<br>
各卷积层后接最大池化层以外,各添加了dropout,避免过拟合
### 问题3:你是如何决定训练模型的epoch数量和batch_size的?
**答案**:
1.epoch设定为5的情况下(lr为0.001),通过最终的train loss数值确定batch_size,<br>
batch_size为10时对应的train loss为0.11059900186955929,运行时间4分6秒<br>
batch_size为16时对应的train loss为0.08914702665060759,运行时间3分25秒<br>
batch_size为32时对应的train loss为0.19058764688670635,运行时间2分56秒<br>
batch_size为64时对应的train loss为0.2119064297527075,运行时间2分42秒<br>
根据以上情况,同样是5个epoch,batch_size为32和64时,train loss收敛幅度不如batch_size为10和16,再考虑到时间因素,故batch_size确定为16
2.确定了batch_size为16的情况下,跑20个epoch,观察每个epoch执行完毕之后的train loss值,<br>
Epoch: 1, Batch: 200, Avg. Loss: 0.24211961589753628<br>
Epoch: 2, Batch: 200, Avg. Loss: 0.19592789337038993<br>
Epoch: 3, Batch: 200, Avg. Loss: 0.12666836492717265<br>
Epoch: 4, Batch: 200, Avg. Loss: 0.10287750419229269<br>
Epoch: 5, Batch: 200, Avg. Loss: 0.08058780394494533<br>
Epoch: 6, Batch: 200, Avg. Loss: 0.06290266215801239<br>
Epoch: 7, Batch: 200, Avg. Loss: 0.05217911237850785<br>
Epoch: 8, Batch: 200, Avg. Loss: 0.06019743625074625<br>
Epoch: 9, Batch: 200, Avg. Loss: 0.047731098253279924<br>
Epoch: 10, Batch: 200, Avg. Loss: 0.05877945628017187<br>
Epoch: 11, Batch: 200, Avg. Loss: 0.03298123115673661<br>
Epoch: 12, Batch: 200, Avg. Loss: 0.044879808742553<br>
Epoch: 13, Batch: 200, Avg. Loss: 0.03970537865534425<br>
Epoch: 14, Batch: 200, Avg. Loss: 0.03596739610657096<br>
Epoch: 15, Batch: 200, Avg. Loss: 0.03561003883369267<br>
Epoch: 16, Batch: 200, Avg. Loss: 0.028288140986114742<br>
Epoch: 17, Batch: 200, Avg. Loss: 0.029539932357147337<br>
Epoch: 18, Batch: 200, Avg. Loss: 0.026936942432075738<br>
Epoch: 19, Batch: 200, Avg. Loss: 0.03204792528413236<br>
Epoch: 20, Batch: 200, Avg. Loss: 0.03614617483690381<br>
由上结果可以看出,epoch在达到18之后,train loss不降反升,故epoch设为18就可以(为节约计算资源,不再重复训练,以epoch20结果保存模型)
## 特征可视化
有时,神经网络会被当做是一个黑盒子,给定一些输入,它就会学习产生一些输出。 事实上,CNN正在学习识别各种空间模式,你可以通过查看构成每个卷积核的权重并将这些一次性应用于样本图像来可视化每个卷积层已被训练识别的内容。这种技术称为特征可视化,它对于理解CNN的内部工作方式很有帮助。
在下面的单元格中,你可以看到如何从第一个卷积层中按索引提取单个滤波器。滤波器应显示为灰度网格。
```
# Get the weights in the first conv layer, "conv1"
# if necessary, change this to reflect the name of your first conv layer
weights1 = net.conv1.weight.data
w = weights1.numpy()
filter_index = 0
print(w[filter_index][0])
print(w[filter_index][0].shape)
# display the filter weights
plt.imshow(w[filter_index][0], cmap='gray')
```
## 特征映射
每个CNN至少包含一个由堆叠滤波器(也称为卷积核)组成的卷积层。CNN在进行训练时,它要学习在卷积内核中包含哪些权重,当这些内核应用于某些输入图像时,它们会产生一组**特征映射**。因此,特征映射只是过滤图像的集合,它们是通过将卷积核应用于输入图像而产生的图像。这些映射向我们展示了神经网络不同层学习提取的特征。例如,你可以想象一个卷积内核,它可以检测到脸部的垂直边缘,而另一个可以检测到眼角的边缘。通过将这些内核应用于图像,你可以看到每个内核检测到了哪些特征。具体请看以下示例,从它在图像中显示线条的方式,你可以将其表征为边缘检测滤波。
<img src='images/feature_map_ex.png' width=50% height=50%/>
接下来,选择一个测试图像并使用已被训练的CNN中的一个卷积内核对其进行过滤。查看过滤后的输出,了解该内核检测到的内容。
### TODO: 过滤图像,查看卷积内核的效果
---
```
##TODO: load in and display any image from the transformed test dataset
sample_num = np.random.randint(len(test_dataset))
sample_img = test_dataset[sample_num]['image']
#print(sample_img)
sample_img = sample_img.data.numpy()
sample_img = np.transpose(sample_img, (1, 2, 0))
sample_img = np.squeeze(sample_img)
plt.imshow(sample_img, cmap='gray')
## TODO: Using cv's filter2D function,
## apply a specific set of filter weights (like the one displayed above) to the test image
weights = net.conv1.weight.data
w = weights.numpy()
#plt.imshow(w[0][0], cmap='gray')
import cv2
fig=plt.figure(figsize=(30, 10))
columns = 5*2
rows = 2
for i in range(0, columns*rows):
fig.add_subplot(rows, columns, i+1)
if ((i%2)==0):
plt.imshow(w[int(i/2)][0], cmap='gray')
else:
c = cv2.filter2D(sample_img, -1, w[int((i-1)/2)][0])
plt.imshow(c, cmap='gray')
plt.show()
```
### 问题4:从已被训练的CNN中选择一个滤波器并将其应用于测试图像。你认为它会起到什么作用?你认为它会检测到哪种特征?
**答案**:观察上图第一个过滤器的特征映射图,检测为水平方向的边缘检测器
---
## 继续加油吧!
现在,你已经定义并训练了模型,最终也保存了一个最佳模型。接下来,就是最后一个notebook,它会将人脸检测器与你保存的模型相结合,创建一个人脸关键点检测系统,用于预测一种图像中*任何一个* 人脸的关键点!
| github_jupyter |
## Python not in the Notebook
We will often want to save our Python classes, for use in multiple Notebooks.
We can do this by writing text files with a .py extension, and then `importing` them.
### Writing Python in Text Files
You can use a text editor like [VS Code](https://code.visualstudio.com/) or [Spyder](https://www.spyder-ide.org/). If you create your own Python files ending in `.py`, then you can import them with `import` just like external libraries.
You can also maintain your library code in a Notebook, and use `%%writefile` to create your library, though this is not encouraged!
Libraries are usually structured with multiple files, one for each class.
We will be turning the code we have written for the maze into a library, so that other code can reuse it.
We group our modules into packages, by putting them together into a folder. You can do this with explorer, or using a shell, or even with Python:
```
import os
if 'mazetool' not in os.listdir(os.getcwd()):
os.mkdir('mazetool')
%%writefile mazetool/maze.py
from .room import Room
from .person import Person
class Maze(object):
def __init__(self, name):
self.name = name
self.rooms = []
self.occupants = []
def add_room(self, name, capacity):
result = Room(name, capacity)
self.rooms.append(result)
return result
def add_exit(self, name, source, target, reverse= None):
source.add_exit(name, target)
if reverse:
target.add_exit(reverse, source)
def add_occupant(self, name, room):
self.occupants.append(Person(name, room))
room.occupancy += 1
def wander(self):
"Move all the people in a random direction"
for occupant in self.occupants:
occupant.wander()
def describe(self):
for occupant in self.occupants:
occupant.describe()
def step(self):
house.describe()
print()
house.wander()
print()
def simulate(self, steps):
for _ in range(steps):
self.step()
%%writefile mazetool/room.py
from .exit import Exit
class Room(object):
def __init__(self, name, capacity):
self.name = name
self.capacity = capacity
self.occupancy = 0
self.exits = []
def has_space(self):
return self.occupancy < self.capacity
def available_exits(self):
return [exit for exit in self.exits if exit.valid() ]
def random_valid_exit(self):
import random
if not self.available_exits():
return None
return random.choice(self.available_exits())
def add_exit(self, name, target):
self.exits.append(Exit(name, target))
%%writefile mazetool/person.py
class Person(object):
def __init__(self, name, room = None):
self.name=name
self.room=room
def use(self, exit):
self.room.occupancy -= 1
destination=exit.target
destination.occupancy +=1
self.room=destination
print(self.name, "goes", exit.name, "to the", destination.name)
def wander(self):
exit = self.room.random_valid_exit()
if exit:
self.use(exit)
def describe(self):
print(self.name, "is in the", self.room.name)
%%writefile mazetool/exit.py
class Exit(object):
def __init__(self, name, target):
self.name = name
self.target = target
def valid(self):
return self.target.has_space()
```
In order to tell Python that our "mazetool" folder is a Python package,
we have to make a special file called `__init__.py`. If you import things in there, they are imported as part of the package:
```
%%writefile mazetool/__init__.py
from .maze import Maze # Python 3 relative import
```
In this case we are making it easier to import `Maze` as we are making it available one level above.
### Loading Our Package
We just wrote the files, there is no "Maze" class in this notebook yet:
```
myhouse = Maze('My New House')
```
But now, we can import Maze, (and the other files will get imported via the chained Import statements, starting from the `__init__.py` file.
```
import mazetool
```
Let's see how we can access the files we created:
```
mazetool.exit.Exit
from mazetool import Maze
house = Maze('My New House')
living = house.add_room('livingroom', 2)
```
Note the files we have created are on the disk in the folder we made:
```
import os
os.listdir(os.path.join(os.getcwd(), 'mazetool') )
```
You may get also `.pyc` files. Those are "Compiled" temporary python files that the system generates to speed things up. They'll be regenerated
on the fly when your `.py` files change. They may appear inside the `__pycache__` directory.
### The Python Path
We want to `import` these from notebooks elsewhere on our computer:
it would be a bad idea to keep all our Python work in one folder.
The best way to do this is to learn how to make our code
into a proper module that we can install. We'll see more on that in a [few lectures' time](./03Packaging.html) ([notebook](./03Packaging.ipynb)).
Alternatively, we can add a folder to the "`PYTHONPATH`", where python searches for modules:
```
import sys
print('\n'.join(sys.path[-3:]))
from pathlib import Path
sys.path.append(os.path.join(Path.home(), 'devel', 'libraries', 'python'))
print(sys.path[-1])
```
I've thus added a folder to the list of places searched. If you want to do this permanently, you should set the `PYTHONPATH` Environment Variable,
which you can learn about in a shell course, or can read about online for your operating system.
| github_jupyter |
You may want to make use of parts of .net that aren't default opened
```
System.Windows.Forms.DataVisualization //WebClient / System.NET
#r "System.Windows.Forms.DataVisualization.dll"
System.Windows.Forms.DataVisualization.Charting.Point3D()
```
You can also use this with your own libraries:
```
#r "../../somewhere/on/yourlocalmachine/amazing.dll"
```
Paket https://fsprojects.github.io/Paket/ (NuGet client)
```
#load "Paket.fsx"
//Paket.Package(["FsLab"])
Paket.Version(["FsLab", "1.1.3"]) //Note not using 1.1.5 to ease connection to XPlot
```
So we could #r into the FsLab
```
//#r "/packages/FsLab/FsLab.fsx"
```
But it's much safer to just let Paket handle this as well (we activate this by default: https://fsprojects.github.io/Paket/paket-generate-load-scripts.html)
```
#load "Paket.Generated.Refs.fsx"
//#load @".paket/load/main.group.fsx"
open System
open Deedle
```
http://fslaborg.github.io/Deedle/tutorial.html#Creating-series-and-frames
```
let dates =
[ DateTime(2013,1,1);
DateTime(2013,1,4);
DateTime(2013,1,8) ]
let values =
[ 10.0; 20.0; 30.0 ]
let first = Series(dates, values)
// Create from a single list of observations
Series.ofObservations
[ DateTime(2013,1,1) => 10.0
DateTime(2013,1,4) => 20.0
DateTime(2013,1,8) => 30.0 ]
//#load "Deedle.fsx" //Doesn't help, we'll return to this later
Series.ofObservations
[ DateTime(2013,1,1) => 10.0
DateTime(2013,1,4) => 20.0
DateTime(2013,1,8) => 30.0 ]
```
Getting formatting? GitHub dependencies https://github.com/fsprojects/IfSharp/pull/179 thanks to Людмила Мухаметдинова (Lucy Mu) and of course Paket itself
```
Paket.GitHub ["mndrake/IfSharpLab src/DeedleFormat.fs"]
```
https://github.com/mndrake/IfSharpLab thanks to David Carlson
```
#load "paket-files/mndrake/IfSharpLab/src/DeedleFormat.fs"
Series.ofObservations
[ DateTime(2013,1,1) => 10.0
DateTime(2013,1,4) => 20.0
DateTime(2013,1,8) => 30.0 ]
```
Don't want to hard code all our data in!
https://datahelpdesk.worldbank.org/knowledgebase/articles/902061-climate-data-api
http://climatedataapi.worldbank.org/climateweb/rest/v1/country/cru/tas/month/USA.csv
Ignore this I guess? Can a Deedle expert explain?
https://stackoverflow.com/questions/42671973/why-frame-x-static-methods-from-deedle-are-generating-warnings-in-vs-2017
https://github.com/fslaborg/Deedle/blob/d4acfa54f4112ac8143db9ba55b138fcff2b8c10/src/Deedle/FrameExtensions.fs#L91
```
Frame.ReadCsv("USA.csv")
Frame.ReadCsv("USA.csv", hasHeaders=true)
```
| github_jupyter |
```
import os,sys
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import pylab as P
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.cm as cm, matplotlib.font_manager as fm
sns.set(style="darkgrid")
import matplotlib.patheffects as PathEffects
from matplotlib.ticker import FuncFormatter
%pylab inline
#nets = pd.read_csv(os.path.join(drop,'Data/nets2015_long.csv'))
from pandas.api.types import CategoricalDtype
from matplotlib.backends.backend_pdf import PdfPages
#from shapely.geometry import Point
#from fiona.crs import from_epsg
#import geopandas as gpd
#from pyproj import Proj
#from geopandas.tools import sjoin
#from shapely.geometry import Point
#from fiona.crs import from_epsg
from textwrap import wrap
#import shapely
#from platform import python_version
#print(python_version())
from matplotlib.backends.backend_pdf import PdfPages
import matplotlib.ticker as ticker
import matplotlib.colors as colors
import matplotlib.patheffects as PathEffects
def comma_format(x, p):
return '$'+format(x, "6,.0f").replace(",", ".")
def percent_format(x, p):
return format(x, "6,.02f").replace(",", ".")
percent_format = lambda x,pos: '{:,.0f}%'.format(x)
comma_format = lambda x,pos: '{:,.0f}'.format(x)
# Get current size
fig_size = plt.rcParams["figure.figsize"]
print("Current size:%s" %fig_size)
# Set figure width to 12 and height to 9
fig_size[0] = 11
fig_size[1] = 8.5
plt.rcParams["figure.figsize"] = fig_size
print("Current size:%s" %fig_size)
plt.style.use('ggplot')
pct=lambda x: x/x.sum()
def easylabeler(breaks,currency=True,prefix='AGE'):
"""
turn list of breaks into list of strings for binning / cutting.
Parameters
----------
breaks : list-like
Returns
-------
out : list of named strings
"""
if currency:
ptrn_main="${fr:,.0f} - ${to:,.0f}"
ptrn_first='Less than ${dt:,.0f}'
ptrn_last='Greater than ${dt:,.0f}'
else:
ptrn_main="{fr:,.0f} - {to:,.0f}"
ptrn_first='Less than {dt:,.0f}'
ptrn_last='Greater than {dt:,.0f}'
difflabels=[]
for f in range(len(breaks)-1):
difflabels.append(ptrn_main.format(fr=breaks[f],to=breaks[f+1]-1))
## fix first, last boundary entries
difflabels[0]=ptrn_first.format(dt=breaks[1])
difflabels[-1]=ptrn_last.format(dt=breaks[-2])
return difflabels
```
### Load / define a few mapping files
#### Get SOC, NAICS mappings
```
BOX_DIR = '/Users/aolsen/Box'
if (os.environ["USERNAME"])=="lzorn":
BOX_DIR = 'C:\\Users\\lzorn\\Box'
INPUT_PATH = os.path.join(BOX_DIR, 'Horizon and Plan Bay Area 2050/Blueprint/Final Blueprint Modeling/Protesting Polar Bears/NETS analysis/inputs')
OUTPUT_PATH = os.path.join(BOX_DIR, 'Horizon and Plan Bay Area 2050/Blueprint/Final Blueprint Modeling/Protesting Polar Bears/NETS analysis')
NETS_PATH = os.path.join(BOX_DIR, 'DataViz Projects/Data Analysis and Visualization/National Establishment Time Series')
THIS_SCRIPT = 'https://github.com/BayAreaMetro/petrale/blob/master/applications/travel_model_lu_inputs/2015/Employment/NETS_2015_work_from_home_analysis.ipynb'
soc = pd.read_excel(os.path.join(INPUT_PATH,'soc_structure_2010.xls'),skiprows=11,
names=[u'Major Group', u'Minor Group', u'Broad Group', u'Detailed Occupation', u'Description'])
soc['maj']=soc['Major Group'].fillna('').str.split('-').apply(lambda x: x[0])
def classifier(df):
"""
we need to classify each row with the appropriate hierarchy level.
for each row, we want to get the rightmost value available--
that represents the finest grained detail class
"""
x = df.iloc[:4].tolist()
try:
out = next(s for s in x if not s is np.NaN)
except:
out = None
return out
def classlevel(s):
try:
if s[3:]=='0000':
return 'major'
elif np.float64(s[3:]) % 100==0:
return 'minor'
elif np.float64(s[3:]) % 10==0:
return 'broad'
else:
return 'detail'
except:
return 'none'
soc['class']=soc.iloc[:,:4].apply(classifier,axis=1)
soc['hierarchy']=soc['class'].fillna('-1').map(classlevel)
soc[['class','hierarchy','Description']].iloc[1200:1202]
soc[['class','hierarchy','Description']].groupby('hierarchy').size()
soc['Description']=soc.Description.fillna('0').str.lower()
#soc.index=SOC['SOCP_2']
soc['SOCP_2']=soc['class'].apply(lambda x: str(x)[:2])
majorclass = soc.loc[soc['Major Group'].notnull(),['SOCP_2','Description']].copy()
majorclass.index=majorclass.SOCP_2
majorclass=majorclass.Description.to_dict()
soc=soc[soc['class'].notnull()]
soc['class']=soc['class'].str.replace('-','')
#soc[soc.hierarchy=='major'].to_csv(os.path.join(drop,'Documents/Data/_BLS/SOC/soc_structure_2010_major.csv'))
soc_map=soc[soc.hierarchy=='detail'].set_index('Detailed Occupation').Description
soc
## get abag-naics mapping
names=['NAICS','ABAGSector','mtc11','mtc6','EDDSector','acsnames','acscensus']
naics_mappings = pd.read_excel(os.path.join(INPUT_PATH, 'NAICS_to_ABAG_SECTORS.xlsx'),sheet_name='both')
naics_mappings.columns=names
naics_mappings['NAICS']=naics_mappings['NAICS'].astype(str)
#naics_abag = pd.read_csv(os.path.join(drop, r'Documents\Data\BayArea\Projections 2013\NAICS_to_ABAG_SECTORS.csv'),sep='\t',dtype=object)
naics_abag = naics_mappings.set_index('NAICS').mtc11.to_dict()
naics_mtc = naics_mappings.set_index('NAICS').mtc6.to_dict()
naics_abag_11 = naics_mappings.set_index('NAICS')['mtc11'].to_dict()
naics_mappings.mtc11=naics_mappings.mtc11.str.strip()
naics_mappings.mtc6=naics_mappings.mtc6.str.strip()
naics_mappings.groupby(['acsnames','mtc11']).size().reset_index()
naics_mappings
# taz = gpd.read_file(os.path.join(drop,'Documents/Data/GIS/zones1454.shp'),crs=from_epsg(3740)).to_crs(from_epsg(3740))
# #taz['county']=taz.county_mtc.map(countymap)
# taz=taz[taz.geometry.notnull()]
# taz['geometry']=taz.buffer(.001)
# ## read naics-3 values from file
# naicsmap3 = pd.read_csv(os.path.join(drop,'Documents/Data/Classification/naics_3.csv'),sep='\t',
# dtype={'Naics_3': object, 'description': object}).set_index('Naics_3').description.to_dict()
## naics-2 dict
naicsmap = {'11':'Agriculture, Forestry, Fishing and Hunting',
'21': 'Mining, Quarrying, and Oil and Gas Extraction',
'22': 'Utilities',
'23': 'Construction',
'31': 'Manufacturing',
'32': 'Manufacturing',
'33': 'Manufacturing',
'42': 'Wholesale Trade',
'44': 'Retail Trade',
'45': 'Retail Trade',
'48': 'Transportation and Warehousing',
'49': 'Transportation and Warehousing',
'51': 'Information',
'52': 'Finance and Insurance',
'53': 'Real Estate and Rental and Leasing',
'54': 'Professional, Scientific, and Technical Services',
'55': 'Management of Companies and Enterprises',
'56': 'Administrative and Support and Waste Management and Remediation Services',
'61': 'Educational Services',
'62': 'Health Care and Social Assistance',
'71': 'Arts, Entertainment, and Recreation',
'72': 'Accommodation and Food Services',
'81': 'Other Services except Public Administration',
'92': 'Public Administration',
'99': 'Unclassified Establishments'
}
# ## define dicts relating the field name in the data to the year it represents (e.g. NAICS01: 2001)
# ## we use later to extract the year
# fipstypemap = dict([('fips{:02d}'.format(int(str(x)[2:])),np.dtype('a5')) for x in range(1990,2016)]+\
# [('naics{:02d}'.format(int(str(x)[2:])),np.dtype('a6')) for x in range(1990,2016)]+\
# [('sales{:02d}'.format(int(str(x)[2:])),str) for x in range(1990,2016)]+\
# [('emp{:02d}'.format(int(str(x)[2:])),np.int32) for x in range(1990,2016)])
# fipsmap = dict([('fips{:02d}'.format(int(str(x)[2:])),x) for x in range(1990,2016)]+\
# [('sales{:02d}'.format(int(str(x)[2:])),x) for x in range(1990,2016)]+\
# [('naics{:02d}'.format(int(str(x)[2:])),x) for x in range(1990,2016)]+\
# [('emp{:02d}'.format(int(str(x)[2:])),x) for x in range(1990,2016)])
# fipstypemap['dunsnumber']=str
```
### Load naics to occupation mappings
```
## load naics-to-soc crosswalk for naics 92 public adm data from PUMS 1-year data
naics92_soc_pct = pd.read_csv(os.path.join(INPUT_PATH,'pums_2016_naics92_soc_share.csv'),dtype={'NAICSP_2':str}).rename(columns={'NAICSP_2':'naics_2','soc_x':'occ_code'}).set_index(['naics_2','occ_code']).Total
naics92_soc_pct
## define 'overflow' industries at 2 digit level
naics_exp={'31': '31-33',
'32': '31-33',
'33': '31-33',
'44': u'44-45',
'45': u'44-45',
'48': u'48-49',
'49': u'48-49'}
## bay area counties
bayareafips_full = {'06001':'Alameda', '06013':'Contra Costa', '06041':'Marin', '06055':'Napa', '06075':'San Francisco', '06081':'San Mateo', '06085':'Santa Clara', '06097':'Sonoma', '06095':'Solano'}
bayareamsas={'06001': u'San Francisco-Oakland-Hayward, CA',
'06013': u'San Francisco-Oakland-Hayward, CA',
'06041': u'San Francisco-Oakland-Hayward, CA',
'06055': u'Napa, CA',
#'06069': u'San Jose-Sunnyvale-Santa Clara, CA',
'06075': u'San Francisco-Oakland-Hayward, CA',
#'06077': u'Stockton-Lodi, CA',
'06081': u'San Francisco-Oakland-Hayward, CA',
'06085': u'San Jose-Sunnyvale-Santa Clara, CA',
'06087': u'Santa Cruz-Watsonville, CA',
'06095': u'Vallejo-Fairfield, CA',
'06097': u'Santa Rosa, CA'}
## load naics sector to occupation crosswalk from OES research estimates, subsetted to CA only
naics_to_occ=pd.read_excel(os.path.join(INPUT_PATH,'oes_research_data_2019_naics_to_occ_share_ca.xlsx'),dtype={'naics':str,'occ_code':str})
naics_to_occ=naics_to_occ.set_index(['naics','occ_code']).tot_emp
naics_to_occ.index=naics_to_occ.index.set_names('naics_2',level=0)
naics_to_occ.head(2)
naics_to_occ_w_naics_92 = naics_to_occ.append(naics92_soc_pct)#.reset_index(name='tot_emp')
naics_to_occ_w_naics_92.name='tot_emp'
naics_to_occ_w_naics_92 = naics_to_occ_w_naics_92.reset_index()
naics_to_occ_w_naics_92
# ## load Dingel's (2020) occupational propensities
# workfromhomeocc_onet_bls=pd.read_csv('https://raw.githubusercontent.com/jdingel/DingelNeiman-workathome/master/occ_onet_scores/output/occupations_workathome.csv')
# workfromhomeocc_onet_bls['OCC_CODE']=workfromhomeocc_onet_bls.onetsoccode.str.split('\.').apply(lambda x: x[0])
# workfromhomeocc_onet_bls=workfromhomeocc_onet_bls.groupby(['OCC_CODE']).teleworkable.mean()
## load Dingel's (2020) occupational propensities, from O*NET scoring, and mapped to OES categories
# 'https://raw.githubusercontent.com/jdingel/DingelNeiman-workathome/master/onet_to_BLS_crosswalk/output/onet_teleworkable_blscodes.csv')
#workfromhomeocc_onet=pd.read_csv(os.path.join(INPUT_PATH,'onet_teleworkable_blscodes.csv'),sep='\t')
workfromhomeocc_onet=pd.read_csv('https://raw.githubusercontent.com/jdingel/DingelNeiman-workathome/master/onet_to_BLS_crosswalk/output/onet_teleworkable_blscodes.csv')
workfromhomeocc=workfromhomeocc_onet.set_index('OCC_CODE').teleworkable
```
### Load the employment data
Get from postgres whenever there are query changes. Otherwise, grab from csv made each time postgres is queried.
```
float_int = lambda x: pd.to_numeric('{:.0f}'.format(pd.to_numeric(x)),errors='coerce')
%%time
## large file - get just a few cols
nets = pd.read_csv(os.path.join(NETS_PATH,'nets2015wide.csv'),
usecols=['dunsnumber','firstyear','lastyear','fips15','naics15','emp15'],
na_values=[''],
engine='python')
nets = nets.set_index(['dunsnumber','firstyear','lastyear'])
print("nets length:{} rows size: {} MB".format(len(nets), nets.memory_usage(index=True).sum()*1e-6))
nets
diffbreaks_5 =[0,25,50,100,250,500,1000,np.inf]
difflabels_5 = easylabeler(breaks=diffbreaks_5,currency=False)
difflabels_5
# nets_2015=nets.filter(regex='15') # lmz commented out since this appears to do nothing
nets_2015=nets.loc[nets.fips15.notnull()].copy()
nets_2015['naics_2']=nets_2015.naics15.astype(int).apply(lambda x: '{:0<6}'.format(x)).str.slice(0,2)
nets_2015['naics_2_desc']=nets_2015.naics_2.map(naicsmap)
nets_2015['naics_abag']=nets_2015.naics_2.map(naics_abag)
nets_2015['naics_mtc']=nets_2015.naics_2.map(naics_mtc)
nets_2015['naics_2']=nets_2015['naics_2'].replace(naics_exp)
nets_2015['emp_size_cat']=pd.cut(nets_2015.emp15,bins=diffbreaks_5,labels=difflabels_5)
nets_2015['emp_bucket']=pd.cut(nets_2015.emp15,bins=[0,25,np.inf],labels=['0-25 employees','25+ employees'])
nets_2015['STCOUNTY']=nets_2015.fips15.astype(int).apply(lambda x: '{:0>5}'.format(x))
nets_2015['CBSA']=nets_2015.STCOUNTY.map(bayareamsas)
nets_2015=nets_2015[nets_2015.STCOUNTY.isin(bayareafips_full)]
print("nets_2015 length: {} rows; size: {} MB".format(len(nets_2015), nets_2015.memory_usage(index=True).sum()*1e-6))
nets_2015
## basic summary by indus
nets_2015_by_indus_size = nets_2015.groupby(['CBSA','STCOUNTY','naics_2','naics_2_desc','naics_abag','naics_mtc','emp_bucket']).emp15.sum()
nets_2015_by_indus_size = nets_2015_by_indus_size.reset_index()
print("nets_2015_by_indus_size has {} rows with emp15=null, {} rows with emp15 not null".format(
pd.isnull(nets_2015_by_indus_size.emp15).sum(),
pd.notnull(nets_2015_by_indus_size.emp15).sum()))
# select only those few rows with emp15 not null
nets_2015_by_indus_size = nets_2015_by_indus_size.loc[pd.notnull(nets_2015_by_indus_size.emp15),]
print("nets_2015_by_indus_size total emp15: {}".format(nets_2015_by_indus_size.emp15.sum()))
nets_2015_by_indus_size
## merge emp data with naics-to-occ crosswalk
nets_2015_occ_exp=nets_2015_by_indus_size.merge(naics_to_occ_w_naics_92,on=['naics_2'])
nets_2015_occ_exp['reg']='Bay Area'
## weigh employment by telecommute propensity from Dingel (2020)
nets_2015_occ_exp['emp15_occ']=nets_2015_occ_exp.emp15*nets_2015_occ_exp.tot_emp
print("nets_2015_occ_exp length: {} rows size: {} MB".format(len(nets_2015_occ_exp), sys.getsizeof(nets_2015_occ_exp)*1e-6))
nets_2015_occ_exp
## codes in NETS data but *not* in the Dingel work from home matrix
nets_occ_unmatched=list(set(nets_2015_occ_exp.occ_code)-set(workfromhomeocc.index))
#oes_occ_unmatched=list(set(naics_to_occ_w_naics_92.index.get_level_values(1).unique())-set(workfromhomeocc.index))
#onet_occ_unmatched=list(set(workfromhomeocc.index)-set(naics_to_occ_w_naics_92.index.get_level_values(1).unique()))
# nets_occ_unmatched
soc_det=soc.loc[soc['Detailed Occupation'].notnull()]
soc_det_missing_from_onet_WFH=soc_det[soc_det['Detailed Occupation'].isin(nets_occ_unmatched)].set_index('Detailed Occupation').Description.index
## take the codes that do *not* have a wfh flag and assign the most common flag
## for occupations in the containing major group. Many of them are "other xxx"
onet_missing_group_imputed=pd.Series(data=soc_det_missing_from_onet_WFH.str.slice(0,2).map(workfromhomeocc.groupby(lambda x: x[:2]).median()),
index=soc_det_missing_from_onet_WFH)
## ADD work from home share
nets_2015_occ_exp['wfh_flag']=nets_2015_occ_exp.occ_code.map(workfromhomeocc.append(onet_missing_group_imputed))
nets_2015_occ_exp['emp15_wfh']=nets_2015_occ_exp.emp15_occ*nets_2015_occ_exp.wfh_flag
```
### Quick summaries
```
nets_2015_occ_exp.groupby(['emp_bucket'])['emp15_occ','emp15_wfh'].sum().plot(kind='barh',figsize=[8,5])
title('Bay Area employment, total and work from home potential\nSources: Work from home potential from Dingel (2020)\nIndustry - Occupation matrix from BLS OES 2019 Research Estimates (CA subset)\nEmployment data from National Establishment Timeseries (NETS) 2015')
fig=plt.figure(figsize=[8,6])
ax=sns.heatmap((nets_2015_occ_exp.groupby(['CBSA','naics_mtc','emp_bucket'])['emp15_wfh'].sum()/\
nets_2015_occ_exp.groupby(['CBSA','naics_mtc','emp_bucket'])['emp15_occ'].sum()).unstack([0]).loc(0)[:,'25+ employees'].reset_index(1,drop=True).T
,
annot=True,fmt=',.2f',linewidths=.5,cmap=cm.coolwarm,
annot_kws={'fontsize':12})
title('Employment susceptible to telecommuting, using NETS data\nclassified using Dingel (2020) after mapping industry to occupation data\nBay Area CBSAs shown')
plt.tight_layout()
plt.yticks(rotation=0,size=12)
plt.xticks(rotation=45,size=12)
fig=plt.figure(figsize=[8,6])
ax=sns.heatmap((nets_2015_occ_exp.groupby(['STCOUNTY','naics_mtc','emp_bucket'])['emp15_wfh'].sum()/\
nets_2015_occ_exp.groupby(['STCOUNTY','naics_mtc','emp_bucket'])['emp15_occ'].sum()).unstack([0]).loc(0)[:,'25+ employees'].reset_index(1,drop=True).T
,
annot=True,fmt=',.2f',linewidths=.5,cmap=cm.coolwarm,
annot_kws={'fontsize':12})
title('Employment susceptible to telecommuting, using NETS data\nclassified using Dingel (2020) after mapping industry to occupation data\nBay Area CBSAs shown')
plt.tight_layout()
plt.yticks(rotation=0,size=12)
plt.xticks(rotation=45,size=12)
#savefig(os.path.join(box, 'RHNA/Analyses/equity/divergence_opportunity_corr.pdf'))
fig=plt.figure(figsize=[8,6])
ax=sns.heatmap((nets_2015_occ_exp.groupby(['naics_mtc','emp_bucket'])['emp15_wfh'].sum()/\
nets_2015_occ_exp.groupby(['naics_mtc','emp_bucket'])['emp15_occ'].sum()).unstack(),
annot=True,fmt=',.2f',linewidths=.5,cmap=cm.coolwarm,
annot_kws={'fontsize':12})
title('Employment susceptible to telecommuting, using NETS data\nclassified using Dingel (2020) after mapping industry to occupation data')
plt.tight_layout()
plt.yticks(rotation=0,size=12)
plt.xticks(rotation=45,size=12)
#savefig(os.path.join(box, 'RHNA/Analyses/equity/divergence_opportunity_corr.pdf'))
```
### Write out spreadsheet with WFH potential share based on industry / occupation and establishment size alone
```
pd.options.display.float_format = '{:,.2f}'.format
xl4=pd.ExcelWriter(os.path.join(OUTPUT_PATH,'WFH_By_Sector_V4.xlsx'))
header = pd.DataFrame(data=["Source: {}".format(THIS_SCRIPT),"Table"], columns=["col1"])
## WORK FROM HOME POTENTIAL SHARE, by ESTAB SIZE
header.loc[1] = "Table 1: Work from home potential, by establishment size"
header.to_excel(xl4, 'wfhshare_by_estabsize',index=False,merge_cells=False,header=False)
(nets_2015_occ_exp.groupby(['emp_bucket'])['emp15_wfh'].sum()/\
nets_2015_occ_exp.groupby(['emp_bucket'])['emp15_occ'].sum()).reset_index(name='value').to_excel(xl4,'wfhshare_by_estabsize',index=False,merge_cells=False,startrow=3)
## WORK FROM HOME POTENTIAL SHARE, by ESTAB SIZE, SECTOR
header.loc[1] = "Table 2: Work from home potential, by establishment size and sector"
header.to_excel(xl4, 'wfhshare_by_estabsize_sector',index=False,merge_cells=False,header=False)
(nets_2015_occ_exp.groupby(['naics_mtc','emp_bucket'])['emp15_wfh'].sum()/\
nets_2015_occ_exp.groupby(['naics_mtc','emp_bucket'])['emp15_occ'].sum()).unstack(1).to_excel(xl4,'wfhshare_by_estabsize_sector',index=True,merge_cells=False,startrow=3)
## Work from home potential, by establishment size, sector and MSA
header.loc[1] = "Table 3: Work from home potential, by establishment size, sector and MSA"
header.to_excel(xl4, 'wfhshare_by_cbsa_sector',index=False,merge_cells=False,header=False)
(nets_2015_occ_exp.groupby(['CBSA','naics_mtc'])['emp15_wfh'].sum()/\
nets_2015_occ_exp.groupby(['CBSA','naics_mtc'])['emp15_occ'].sum()).reset_index(name='share').to_excel(xl4,'wfhshare_by_cbsa_sector',index=False,merge_cells=False,startrow=3)
## Work from home potential, by sector
header.loc[1] = "Table 4: Work from home potential, by sector"
header.to_excel(xl4, 'wfhshare_by_sector',index=False,merge_cells=False,header=False)
(nets_2015_occ_exp.groupby(['naics_mtc'])['emp15_wfh'].sum()/\
nets_2015_occ_exp.groupby(['naics_mtc'])['emp15_occ'].sum()).reset_index(name='share').to_excel(xl4,'wfhshare_by_sector',index=False,merge_cells=False,startrow=3)
## Table 5 Work from home potential, by establishment size, sector and county fips
header.loc[1] = "Table 5: Work from home potential, by establishment size, sector and county fips"
header.to_excel(xl4, 'wfhshare_by_county_sector',index=False,merge_cells=False,header=False)
(nets_2015_occ_exp.groupby(['STCOUNTY','naics_mtc'])['emp15_wfh'].sum()/\
nets_2015_occ_exp.groupby(['STCOUNTY','naics_mtc'])['emp15_occ'].sum()).reset_index(name='share').to_excel(xl4,'wfhshare_by_county_sector',index=False,merge_cells=False,startrow=3)
## Table 6 Employees by firm size
header.loc[1] = "Table 6: Employment by Establishment Size"
header.to_excel(xl4, 'emp_by_size',index=False,merge_cells=False,header=False)
emp_by_size = nets_2015.groupby(['emp_bucket'])['emp15'].sum().reset_index()
emp_by_size["emp15_share"] = emp_by_size.emp15/emp_by_size.emp15.sum()
emp_by_size.to_excel(xl4, 'emp_by_size', index=False,merge_cells=False,startrow=3)
xl4.close()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import time
import operator
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.metrics import log_loss, f1_score, accuracy_score
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
trn = pd.read_csv("../input/train_clean.csv")
target = pd.read_csv("../input/train.csv", usecols=["target"])
tst = pd.read_csv("../input/test_clean.csv")
test_id = tst["ncodpers"]
tst.drop(["ncodpers"], axis=1, inplace=True)
trn.drop(["ncodpers"], axis=1, inplace=True)
print(trn.shape, target.shape, tst.shape)
```
Train 데이터와 Test 데이터의 컬럼이 동일해야 하므로 확인을 해봐야 합니다 :)
```
trn.columns == tst.columns
```
Scikit-learn의 경우 수치형 데이터가 아니면 들어가질 않기 때문에
수치형이 아닌 친구를 확인해봐요
```
for col in trn.columns:
if trn[col].dtype == "object":
print(col)
for col in trn.columns:
if trn[col].dtype == "object":
lb = LabelEncoder()
lb.fit(pd.concat([trn[col], tst[col]])) # 테스트와 트레인 데이터를 아래로 합침
trn[col] = lb.transform(trn[col]) # 컬럼을 덮어씌움
tst[col] = lb.transform(tst[col])
# 이제 변수 타입들 다시 확인
for col in trn.columns:
print(col, trn[col].dtype, tst[col].dtype)
# target을 이제 확인해봐요
for t in np.unique(target):
print(t, sum(target["target"]==t))
# 빈도가 적은 데이터 제거
rem_targets = [2, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 15, 17, 18, 19, 21, 22, 23] # 18 classes
trn = trn[target["target"].isin(rem_targets)]
target = target[target["target"].isin(rem_targets)]
target = LabelEncoder().fit_transform(target)
def evaluate(x, y, model):
trn_scores = dict(); vld_scores = dict()
sss = StratifiedShuffleSplit(n_splits=3, test_size=0.1, random_state=777) # 10% 테스트, random_state 는 시드값
for t_ind, v_ind in sss.split(x,y):
# split data
x_trn, x_vld = x.iloc[t_ind], x.iloc[v_ind]
y_trn, y_vld = y[t_ind], y[v_ind]
# fit model
model.fit(x_trn, y_trn) # train만 학습시킵니다!
# eval _ trn
preds = model.predict(x_trn)
acc_scores = trn_scores.get('accuracy', [])
acc_scores.append(accuracy_score(y_trn, preds))
trn_scores['accuracy'] = acc_scores
f1_scores = trn_scores.get('f1 score', [])
f1_scores.append(f1_score(y_trn, preds, average='weighted'))
trn_scores['f1 score'] = f1_scores
preds = model.predict_proba(x_trn)
log_scores = trn_scores.get('log loss', [])
log_scores.append(log_loss(y_trn, preds))
trn_scores['log loss'] = log_scores
# eval _ vld
preds = model.predict(x_vld) # predice된 값을 y_vld랑 비교
acc_scores = vld_scores.get('accuracy', [])
acc_scores.append(accuracy_score(y_vld, preds))
vld_scores['accuracy'] = acc_scores
f1_scores = vld_scores.get('f1 score', [])
f1_scores.append(f1_score(y_vld, preds, average='weighted'))
vld_scores['f1 score'] = f1_scores
preds = model.predict_proba(x_vld)
log_scores = vld_scores.get('log loss', [])
log_scores.append(log_loss(y_vld, preds))
vld_scores['log loss'] = log_scores
return trn_scores, vld_scores
def print_scores(trn_scores, vld_scores):
prefix = ' '
cols = ['accuracy', 'f1 score','log loss']
print('='*50)
print('TRAIN EVAL')
for col in cols:
print('-'*50)
print('# {}'.format(col))
print('# {} Mean : {}'.format(prefix, np.mean(trn_scores[col])))
print('# {} Raw : {}'.format(prefix, trn_scores[col]))
print('='*50)
print('VALID EVAL')
for col in cols:
print('-'*50)
print('# {}'.format(col))
print('# {} Mean : {}'.format(prefix, np.mean(vld_scores[col])))
print('# {} Raw : {}'.format(prefix, vld_scores[col]))
def print_time(end, start):
print('='*50)
elapsed = end - start
print('{} secs'.format(round(elapsed)))
def fit_and_eval(trn, target, model):
trn_scores, vld_scores = evaluate(trn,target,model)
print_scores(trn_scores, vld_scores)
print_time(time.time(), st)
st = time.time()
from sklearn.linear_model import LogisticRegression
model = LogisticRegression(n_jobs=-1, random_state=777) #n_jobs= -1 인 경우 자싱니 가진 모든 cpu를 사용함
fit_and_eval(trn, target, model)
# 58 sec
```
훈련 데이터와 검증 데이터의 평가 척도가 비슷한지 확인을 해야합니다-!
너무 다른 경우 오버피팅의 가능성이 존재해요
로지스틱 regression에서 C의 값은 regularization의 역수임
지금 과적합 상태라면 c의 값을 낮추면 정규성이 증가됨~!!
모델은 복잡도 기준으로 바라볼 것
sroted Feature importance 기준으로 볼 경우 변수의 scale이 다른데 그냥 한번에 돌림..! normalize를 하고 다시 돌려볼 것
```
# Utility
def observe_model_lr(model):
target_num = 0
print('='*50)
print(model)
print('='*50)
print('# Coefficients for target_num == {}'.format(target_num))
print(model.coef_[target_num])
print('-'*50)
print('# Mapped to Column Name')
prefix = ' '
coefs = dict()
for i, coef in enumerate(model.coef_[target_num]):
print('{} {} \t {}'.format(prefix, round(coef,5), trn.columns[i]))
coefs[trn.columns[i]] = np.absolute(coef)
print('-'*50)
print('# Sorted Feature Importance')
coefs_sorted = sorted(coefs.items(), key=operator.itemgetter(1), reverse=True)
for item in coefs_sorted:
print('{} {} \t {}'.format(prefix, round(item[1],5), item[0]))
return coefs_sorted
def plot_coef(coef):
x = []; y = []
for item in coef:
x.append(item[0])
y.append(item[1])
f, ax = plt.subplots(figsize=(20, 15))
sns.barplot(x,y,alpha=0.5)
ax.set_title('Feature Importance for Model : Logistic Regression')
ax.set(xlabel='Column Name', ylabel='Feature Importance')
# 모델 상세 보기
coef = observe_model_lr(model)
```
# 캐글 결과물 출력을 위한 코드
```
from datetime import datetime
import os
print('='*50)
print('# Test shape : {}'.format(tst.shape))
model = LogisticRegression(n_jobs=-1, random_state=777)
model.fit(trn,target)
preds = model.predict_proba(tst)
preds = np.fliplr(np.argsort(preds, axis=1)) # 왼쪽이 제일 큰 값이 나오게 설정
cols = ['ind_ahor_fin_ult1', 'ind_aval_fin_ult1', 'ind_cco_fin_ult1',
'ind_cder_fin_ult1', 'ind_cno_fin_ult1', 'ind_ctju_fin_ult1',
'ind_ctma_fin_ult1', 'ind_ctop_fin_ult1', 'ind_ctpp_fin_ult1',
'ind_deco_fin_ult1', 'ind_deme_fin_ult1', 'ind_dela_fin_ult1',
'ind_ecue_fin_ult1', 'ind_fond_fin_ult1', 'ind_hip_fin_ult1',
'ind_plan_fin_ult1', 'ind_pres_fin_ult1', 'ind_reca_fin_ult1',
'ind_tjcr_fin_ult1', 'ind_valo_fin_ult1', 'ind_viv_fin_ult1',
'ind_nomina_ult1', 'ind_nom_pens_ult1', 'ind_recibo_ult1']
target_cols = [cols[i] for i, col in enumerate(cols) if i in rem_targets]
final_preds = []
for pred in preds:
top_products = []
for i, product in enumerate(pred):
top_products.append(target_cols[product])
if i == 6:
break
final_preds.append(' '.join(top_products))
out_df = pd.DataFrame({'ncodpers':test_id, 'added_products':final_preds})
file_name = datetime.now().strftime("result_%Y%m%d%H%M%S") + '.csv'
out_df.to_csv(os.path.join('../output',file_name), index=False)
```
| github_jupyter |
# Preferential Bayesian Optimization: Dueling-Thompson Sampling
Implementation of the algorithm by Gonzalez et al (2017).
```
import numpy as np
import gpflow
import tensorflow as tf
import tensorflow_probability as tfp
import matplotlib.pyplot as plt
import sys
import os
import datetime
import pickle
from gpflow.utilities import set_trainable, print_summary
gpflow.config.set_default_summary_fmt("notebook")
sys.path.append(os.path.split(os.path.split(os.path.split(os.getcwd())[0])[0])[0]) # Move 3 levels up directory to import project files as module
import importlib
PBO = importlib.import_module("Top-k-Ranking-Bayesian-Optimization")
print("Num GPUs Available: ", len(tf.config.experimental.list_physical_devices('GPU')))
gpu_to_use = 0
print("Num GPUs Available: ", len(tf.config.experimental.list_physical_devices('GPU')))
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
# Restrict TensorFlow to only use the first GPU
try:
for gpu in gpus:
tf.config.experimental.set_memory_growth(gpu, True)
tf.config.experimental.set_visible_devices(gpus[gpu_to_use], 'GPU')
logical_gpus = tf.config.experimental.list_logical_devices('GPU')
print(len(gpus), "Physical GPUs,", len(logical_gpus), "Logical GPU")
except RuntimeError as e:
# Visible devices must be set before GPUs have been initialized
print(e)
def log(message):
print(str(datetime.datetime.now()) + ': ' + message)
objective = PBO.objectives.six_hump_camel
objective_low = -1.5
objective_high = 1.5
objective_name = "SHC"
acquisition_name = "DTS"
experiment_name = acquisition_name + "_" + objective_name
num_runs = 10
num_evals = 35
num_samples = 100
num_choices = 2
input_dims = 2
num_maximizers = 20
num_init_points = 3
num_inducing_init = 3
num_discrete_per_dim = 20
num_fourier_features = 200
results_dir = os.getcwd() + '/results/' + experiment_name + '/'
try:
# Create target Directory
os.makedirs(results_dir)
print("Directory " , results_dir , " created ")
except FileExistsError:
print("Directory " , results_dir , " already exists")
def visualize_model(query_points, y, m, title="Model", cmap="Spectral"):
if query_points.shape[-1] != 2:
return
pos_vals = []
neg_vals = []
for i in range(len(y)):
if y[i]:
pos_vals.append(query_points[i])
else:
neg_vals.append(query_points[i])
pos_vals = np.array(pos_vals)
neg_vals = np.array(neg_vals)
num_discrete_points = num_discrete_per_dim
side = np.linspace(0, 1, num_discrete_points)
X,Y = np.meshgrid(side,side)
preds = tf.transpose(tf.reshape(m.predict_y(combs)[0], [num_discrete_points, num_discrete_points]))
variances = tf.transpose(tf.reshape(PBO.acquisitions.dts.variance_logistic_f(m, combs), [num_discrete_points, num_discrete_points]))
fig, (ax1, ax2) = plt.subplots(1, 2)
fig.suptitle(title)
fig.set_size_inches(18.5, 6.88)
fig.set_dpi((200))
ax1.axis('equal')
if len(pos_vals) != 0:
ax1.scatter(*pos_vals.T, c="black", marker="o")
if len(neg_vals) != 0:
ax1.scatter(*neg_vals.T, c="black", marker="x")
im1 = ax1.imshow(preds, interpolation='nearest', extent=(0.0, 1.0, 0.0, 1.0), origin='lower', cmap=cmap)
ax1.set_title("Mean of y(x, x')")
ax1.set_xlabel("x")
ax1.set_ylabel("x'")
ax1.axvline(x=0.757, linestyle='--')
fig.colorbar(im1, ax=ax1)
ax2.axis('equal')
if len(pos_vals) != 0:
ax2.scatter(*pos_vals.T, c="black", marker="o")
if len(neg_vals) != 0:
ax2.scatter(*neg_vals.T, c="black", marker="x")
im2 = ax2.imshow(variances, interpolation='nearest', extent=(0.0, 1.0, 0.0, 1.0), origin='lower', cmap=cmap)
ax2.set_title("Variance of y(x, x')")
ax2.set_xlabel("x")
ax2.set_ylabel("x'")
fig.colorbar(im2, ax=ax2)
plt.savefig(fname=results_dir + title + ".png")
plt.show()
def std_representation(X, num_choices):
"""
:param X: tensor of shape (num_data, input_dims * num_choices)
:return: tensor of shape (num_data, num_choices, input_dims)
"""
input_dims = X.shape[-1] // num_choices
ret_val = np.zeros((X.shape[0], num_choices, input_dims))
for i in range(num_choices):
ret_val[:, i, :] = X[:, input_dims*i:input_dims*(i+1)]
return ret_val
def visualize_f_sample(f_vals, cmap="Spectral"):
fig, (ax1) = plt.subplots(1)
fig.suptitle('Sampled f values')
fig.set_size_inches(4, 3.3)
fig.set_dpi((100))
ax1.axis('equal')
im1 = ax1.imshow(tf.transpose(tf.reshape(f_vals, [num_discrete_points, num_discrete_points])),
interpolation='nearest', extent=(0.0, 1.0, 0.0, 1.0), origin='lower', cmap=cmap)
ax1.set_xlabel("x")
ax1.set_ylabel("x'")
ax1.axvline(x=0.757, linestyle='--')
fig.colorbar(im1, ax=ax1)
def get_noisy_observation_dts(X, objective):
"""
:param X: tensor of shape (num_data, input_dims * 2)
:param objective: objective function
"""
num_data = X.shape[0]
X_std = std_representation(X, num_choices) # (num_data, num_choices, input_dims)
f = PBO.objectives.objective_get_f_neg(X_std, objective)
obs = np.array(PBO.observation_model.gen_observation_from_f(X_std, f, 1)) # (num_data, 1, input_dims)
ret_val = np.zeros((num_data, 1), dtype=np.int8)
for i in range(num_data):
if np.allclose(X_std[i, 0], obs[i, 0]):
ret_val[i] = 1
return ret_val
regularizer_lengthscale_mean_over_range = 0.2
regularizer_lengthscale_std_over_range = 0.5
input_range = objective_high - objective_low
lengthscale_mean_regularizer = input_range * regularizer_lengthscale_mean_over_range
lengthscale_std_regularizer = input_range * regularizer_lengthscale_std_over_range
lengthscale = lengthscale_mean_regularizer
@tf.function
def lengthscale_regularizer(kernel): # for product kernel
loss = 0
for k in kernel.kernels:
loss += 0.5 * tf.reduce_sum(tf.square((k.lengthscale - lengthscale_mean_regularizer) / lengthscale_std_regularizer))
return loss
def train_and_visualize(X, y, lengthscale, title, num_steps=3000):
kernel = gpflow.kernels.Product([gpflow.kernels.RBF(lengthscale=lengthscale,
active_dims=[i, i+input_dims])
for i in range(input_dims)])
m = gpflow.models.SVGP(kernel=kernel,
likelihood=gpflow.likelihoods.Bernoulli(invlink=tf.math.sigmoid),
inducing_variable=X,
whiten=False)
m.inducing_variable.Z.trainable = False
optimizer = tf.keras.optimizers.RMSprop(rho=0.0)
loss = lambda: -m.log_likelihood(X, y) + lengthscale_regularizer(m.kernel)
prev_loss = loss().numpy()
for i in range(num_steps):
optimizer.minimize(loss, m.trainable_variables)
current_loss = loss().numpy()
if i % 500 == 0:
print('Loss at step %s: %s' % (i, current_loss))
if abs((current_loss-prev_loss) / prev_loss) < 1e-7:
print('Loss at step %s: %s' % (i, current_loss))
break
prev_loss = current_loss
visualize_model(X, y, m, title=title)
return m
def uniform_grid(input_dims, num_discrete_per_dim, low, high):
"""
Returns an array with all possible permutations of discrete values in input_dims number of dimensions.
:param input_dims: int
:param num_discrete_per_dim: int
:param low: int
:param high: int
:return: tensor of shape (num_discrete_per_dim ** input_dims, input_dims)
"""
num_points = num_discrete_per_dim ** input_dims
out = np.zeros([num_points, input_dims])
discrete_points = np.linspace(low, high, num_discrete_per_dim)
for i in range(num_points):
for dim in range(input_dims):
val = num_discrete_per_dim ** (dim)
out[i, dim] = discrete_points[int((i // val) % num_discrete_per_dim)]
return out
def best_guess(m, discrete_space, combs):
return PBO.acquisitions.dts.soft_copeland_maximizer(m.predict_y(combs)[0], discrete_space)
def flip(X):
"""
:param X: tensor of shape (num_data, input_dims * 2)
:return: tensor of shape (num_data, input_dims * 2), where the first input_dims is swapped with the second
"""
input_dims = X.shape[-1] // 2
ret_val = np.zeros((X.shape))
for i in range(X.shape[0]):
ret_val[i, :input_dims] = X[i, input_dims:]
ret_val[i, input_dims:] = X[i, :input_dims]
return ret_val
def flip_y(y):
"""
:param y: tensor of shape (num_data, 1), with int values either 0 or 1
"""
return (y + 1) % 2
```
Create the initial values for each run:
```
np.random.seed(0)
init_points = np.random.uniform(low=objective_low, high=objective_high, size=[num_runs, num_init_points, input_dims])
num_combs = int((num_init_points-1) * num_init_points / 2)
init_vals = np.zeros([num_runs, num_combs, num_choices, input_dims])
for run in range(num_runs):
cur_idx = 0
for init_point in range(num_init_points-1):
for next_point in range(init_point+1, num_init_points):
init_vals[run, cur_idx, 0] = init_points[run, init_point]
init_vals[run, cur_idx, 1] = init_points[run, next_point]
cur_idx += 1
init_vals = np.reshape(init_vals, [num_runs, num_combs, num_choices * input_dims])
discrete_space = uniform_grid(input_dims, num_discrete_per_dim, objective_low, objective_high)
combs = PBO.acquisitions.dts.combinations(discrete_space)
```
Store the results in these arrays:
```
num_data_at_end = (num_combs + num_evals) * 2
X_results = np.zeros([num_runs, num_data_at_end, input_dims * num_choices])
y_results = np.zeros([num_runs, num_data_at_end, 1])
best_guess_results = np.zeros([num_runs, num_evals, input_dims])
for run in range(num_runs):
#Fit a GP with kernel k to Dn
X = init_vals[run]
y = get_noisy_observation_dts(X, objective)
X = np.vstack([X, flip(X)])
y = np.vstack([y, flip_y(y)])
model = train_and_visualize(X, y, lengthscale=lengthscale, title="Run_{}_Initial_model".format(run)) #TODO: CHECK LENGTHSCALE
for evaluation in range(num_evals):
log("Starting evaluation " + str(evaluation))
# Sample f using RFF
f_vals = PBO.acquisitions.dts.sample_f(model, X, combs, num_fourier_features)
# 2 and 3. Compute the acquisition for duels alpha and get next duel
log("Computing acquisition function")
x_next = PBO.acquisitions.dts.soft_copeland_maximizer(f_vals, discrete_space)
all_pairs = np.concatenate([np.tile(x_next, (discrete_space.shape[0], 1)), discrete_space], axis=1)
next_vars = np.squeeze(PBO.acquisitions.dts.variance_logistic_f(model, all_pairs),
axis=1)
xprime_next = discrete_space[np.argmax(next_vars)]
x_xprime_next = np.expand_dims(np.concatenate([x_next, xprime_next]), axis=0)
# Change by random small values otherwise Fourier features matrix becomes non-invertible
if np.all(np.equal(x_xprime_next, flip(x_xprime_next))) or x_xprime_next in X:
for i in range(len(x_xprime_next[0])):
if x_xprime_next[0][i] < 0:
x_xprime_next[0][i] += np.random.uniform(low=0., high=1e-3)
else:
x_xprime_next[0][i] -= np.random.uniform(low=0., high=1e-3)
log("x and x_prime: \n" + str(x_xprime_next))
# 4. Run the duel and get y
y_next = get_noisy_observation_dts(x_xprime_next, objective)
log("y_next: \n" + str(y_next))
# 5. Augment X and Y, and add symmetric points
X = np.vstack([X, x_xprime_next, flip(x_xprime_next)])
y = np.vstack([y, y_next, flip_y(y_next)])
# Fit a GP with kernel k to Dj and learn pi(x).
model = train_and_visualize(X, y, lengthscale=lengthscale, title="Run_{}_Evaluation_{}".format(run, evaluation))
# Save model
kernels_variance = []
kernels_lengthscale = []
for k in model.kernel.kernels:
kernels_variance.append(k.variance.numpy())
kernels_lengthscale.append(k.lengthscale.numpy())
pickle.dump((X, y,
tuple(kernels_variance),
tuple(kernels_lengthscale),
model.q_mu.numpy(),
model.q_sqrt.numpy()),
open(results_dir + "Model_Run_{}_Evaluation_{}.p".format(run, evaluation), "wb"))
# Get current best guess
best_guess_results[run, evaluation] = best_guess(model, discrete_space, combs)
X_results[run] = X
y_results[run] = y
pickle.dump((X_results, y_results, best_guess_results), open(results_dir + "Xybestguess.p", "wb"))
global_min = np.min(objective(discrete_space))
metric = best_guess_results
ir = objective(metric) - global_min
mean = np.mean(ir, axis=0)
std_dev = np.std(ir, axis=0)
std_err = std_dev / np.sqrt(ir.shape[0])
print("Mean immediate regret at each evaluation averaged across all runs:")
print(mean)
print("Standard error of immediate regret at each evaluation averaged across all runs:")
print(std_err)
with open(results_dir + acquisition_name + "_" + objective_name + "_" + "mean_sem" + ".txt", "w") as text_file:
print("Mean immediate regret at each evaluation averaged across all runs:", file=text_file)
print(mean, file=text_file)
print("Standard error of immediate regret at each evaluation averaged across all runs:", file=text_file)
print(std_err, file=text_file)
pickle.dump((mean, std_err), open(results_dir + acquisition_name + "_" + objective_name + "_" + "mean_sem.p", "wb"))
```
| github_jupyter |
Welcome back, folks! In this series of 3 blog post, we will be discussing pandas which one of my favorite python libraries. We will go through 74 exercises to solidify your skills with pandas and as usual, I will explain the WHY behind every single exercise.
Pandas is a powerful open-source library for data analysis and data manipulation. The library is packed with a ton of feature, well supported and documented by the community. It is built on top of NumPy and integrate well with all the main machine learning libraries like Scikit-learn and Matplotlib.
Pandas already come bundles in the Anaconda distribution. If you don't have it installed already, please refer to my other blog [here](https://semasuka.github.io/blog/2019/01/06/introduction-to-jupyter-notebook.html) to get you started.
These exercises are inspired from [this](https://www.machinelearningplus.com/python/101-pandas-exercises-python/) amazing blog post.
Remember there is always different ways we can achieve the same result, so if your code does not look like mine. No worries! if you got the same result, then you are good to go.
Now let's jump right in into the exercises.
### Ex 1: How to import pandas and check the version?
Q: As a warm up, we will import pandas and print it's version
#### Solution
```
import pandas as pd
pd.__version__
```
We import pandas as pd which is the common way to refer to pandas and use the dot notation to print its version.
### Ex 2: How to create a series from a list, numpy array and dict?
Q: Create a pandas series from each of the items below: a list, numpy and a dictionary and print the first 5 elements.
```
import numpy as np
mylist = list('abcedfghijklmnopqrstuvwxyz')
myarr = np.arange(26)
mydict = dict(zip(mylist, myarr))
```
#### Desired output
```
# List
# 0 a
# 1 b
# 2 c
# 3 e
# 4 d
# dtype: object
#
# Array
# 0 0
# 1 1
# 2 2
# 3 3
# 4 4
# dtype: int64
#
# Dictionary
# a 0
# b 1
# c 2
# e 3
# d 4
# dtype: int64
```
#### Solution
```
pd.Series(mylist).head()
pd.Series(myarr).head()
pd.Series(mydict).head()
```
Let's first explain what is a series in pandas, as I said at the beginning of this post, pandas is a tool for data manipulation and most of the data is in form of tables and tables are comprised of columns. In pandas, the data are represented in a dataframe comprised of columns and rows and the basic data structure of a dataframe is a series comprised of one column and an index column.
Coming back to our exercise, we are casting(changing the datatype) the list, array and the dictionary into a Series comprised of only one column of data and another column of indexes by using the Series method and print only the first 5 elements.
### Ex 3: How to convert the index of a series into a column of a dataframe?
Q: Convert the series ser into a dataframe with its index as another column on the dataframe.
```
mylist = list('abcedfghijklmnopqrstuvwxyz')
myarr = np.arange(26)
mydict = dict(zip(mylist, myarr))
ser = pd.Series(mydict)
```
#### Desired output

#### Solution
```
ser.to_frame().reset_index().head()
```
To convert a series into a dataframe, we use the to_frame method and to change the ser's index to number, we use the reset_index. We finally print the first 5 elements in the dataframe.
### Ex 4: How to combine many series to form a dataframe?
Q: Combine ser1 and ser2 to form a dataframe.
```
import numpy as np
ser1 = pd.Series(list('abcedfghijklmnopqrstuvwxyz'),name="ser1")
ser2 = pd.Series(np.arange(26),name="ser2")
```
#### Desired output

#### Solution
#### 1st Method
```
pd.concat([ser1,ser2],axis=1).head()
```
We can concatenate the two series into on a Dataframe using the concat method and set the axis equal to 1 to concatenate column-wise. We finally print the first 5 elements.
#### 2nd Method
```
pd.DataFrame({"ser1":ser1,"ser2":ser2}).head()
```
An alternative way to solve this issue, would be to use DataFrame method and passed in a dictionary where the keys are the column's names and the value are the series and then print the first 5 elements.
### Ex 5: How to assign name to the series’ index?
Q: Give a name to the series ser calling it ‘alphabets’.
```
ser = pd.Series(list('abcedfghijklmnopqrstuvwxyz'))
```
#### Desired output
```
# a 0
# b 1
# c 2
# e 3
# d 4
# Name: alphabets, dtype: int64
```
#### Solution
```
ser.name = "alphabets"
ser.head()
```
We give a name to a series through the dot operator by calling the name method and assign it the actual name, in this example "alphabets"
### Ex 6: How to get the items of series A not present in series B?
Q: From ser1 remove items present in ser2.
```
ser1 = pd.Series([1, 2, 3, 4, 5])
ser2 = pd.Series([4, 5, 6, 7, 8])
```
#### Desired output
```
# 0 1
# 1 2
# 2 3
# dtype: int64
```
#### Solution
```
ser1[~ser1.isin(ser2)]
```
We first find which elements present both in ser1 and ser2 using isin method, a boolean DataFrame is returned where True is the position of elements present in ser1 and ser2 and where False is the position of elements only in ser1.
So to get the elements unique to ser1, we use the ~ to reverse the boolean DataFrame and then use indexing to get the actual value.
### Ex 7: How to get the items not common to both series A and series B?
Q: Get all items of ser1 and ser2 not common to both.
```
ser1 = pd.Series([1, 2, 3, 4, 5])
ser2 = pd.Series([4, 5, 6, 7, 8])
```
#### Desired output
```
# 0 1
# 1 2
# 2 3
# 3 6
# 4 7
# 5 8
# dtype: int64
```
### Solution
#### 1st Method
```
unique_ser1 = ser1[~ser1.isin(ser2)]
unique_ser2 = ser2[~ser2.isin(ser1)]
```
We get elements that are not common in both series just like we did in exercise 6
```
unique_ser1
unique_ser2
uniques = pd.Series(np.union1d(unique_ser1,unique_ser2))
uniques
```
At last, we merge the two series unique_ser1 and unique_ser2 using the NumPy function union1d and cast the array into a series.
#### 2nd Method
```
series_u = pd.Series(np.union1d(ser1,ser2))
series_i = pd.Series(np.intersect1d(ser1,ser2))
series_u[~series_u.isin(series_i)]
```
The second method is quite similar to the first one, the difference is that this time we get first the intersection and the union separately using NumPy function and then use indexing on the union series to get the unique element in the two series just like we did in exercise 6.
### Ex 8: How to get the minimum, 25th percentile, median, 75th, and max of a numeric series?
Q: Compute the minimum, 25th percentile, median, 75th, and maximum of ser.
```
ser = pd.Series(np.random.normal(10, 5, 25))
```
#### Desired output
```
#the minimum is :1.63, the 25th percentile is: 7.27, the median is: 10.21, the 75th percentile is: 15.29 and the maximum is: 22.64
```
#### Solution
#### 1st Method
```
print("the minimum is :{0:.2f}, the 25th percentile is: {1:.2f}, the median is: {2:.2f}, the 75th percentile is: {3:.2f} and the maximum is: {4:.2f}".format(ser.min(),ser.quantile(q=0.25),ser.median(),ser.quantile(q=0.75),ser.max()))
```
#### 2nd Method
```
print("the minimum is :{0:.2f}, the 25th percentile is: {1:.2f}, the median is: {2:.2f}, the 75th percentile is: {3:.2f} and the maximum is: {4:.2f}".format(ser.quantile(q=0),ser.quantile(q=0.25),ser.quantile(q=0.50),ser.quantile(q=0.75),ser.quantile(q=1)))
```
We can get the different percentile using the quantile method and pass as argument q the percentile, so for the 0th percentile (which is the min) q will be 0, for the 25th percentile q will be 0.25, for the 50th percentile (which is the median) q will be 0.5, for the 75th percentile q will be 0.75 and last for the 100th percentile (which is the max) q will be 1.
The min, median and max have their functions too in case you don't wanna use the quantile function.
### Ex 9: How to get frequency counts of unique items of a series?
Q: Calculate the frequency counts of each unique value ser.
```
ser = pd.Series(np.take(list('abcdefgh'), np.random.randint(8, size=30)))
```
#### Desired output
```
# b 7
# a 5
# e 5
# f 4
# d 4
# c 2
# h 2
# g 1
# dtype: int64
```
#### Solution
```
ser.value_counts()
```
To get the count of how many times a value is repeated, we use the value_count function on the series.
### Ex 10: How to keep only the top 2 most frequent values as it is and replace everything else as ‘Other’?
Q: From ser, keep the top 2 most frequent items as it is and replace everything else as ‘Other’.
```
np.random.RandomState(100)
ser = pd.Series(np.random.randint(1, 5, [12]))
```
#### Desired output
```
# 0 2
# 1 4
# 2 2
# 3 other
# 4 2
# 5 4
# 6 other
# 7 2
# 8 2
# 9 other
# 10 other
# 11 4
# dtype: object
```
#### Solution
```
most_freq_el = ser.value_counts()[:2].index
ser[~ser.isin(most_freq_el)] = "other"
ser
```
We get first the two most frequent element in ser using the value_count, which will return the series with the values as indexes and the count of how many times those values are repeated. We only need the value so we called the index function.
We use isin and indexing to select all the values other than the two most frequent value and assign it to the string "other".
### Ex 11: How to bin a numeric series to 10 groups of equal size?
Q: Bin the series ser into 10 equal deciles and replace the values with the bin name.
```
ser = pd.Series(np.random.random(20))
```
#### Desired output
```
# 0 4th
# 1 9th
# 2 6th
# 3 2nd
# 4 2nd
# dtype: category
# Categories (10, object): [1st < 2nd < 3rd < 4th ... 7th < 8th < 9th < 10th]
```
#### Solution
```
pd.cut(ser,bins=10,labels=["1st","2nd","3rd","4th","5th","6th","7th","8th","9th","10th"]).head()
```
To get the segment of the series, we use the cut function pass the series, specify how many bins or basket we want to use and give them a label.
### Ex 12: How to convert a numpy array to a dataframe of given shape? (L1)
Q: Reshape the series ser into a dataframe with 7 rows and 5 columns
```
ser = pd.Series(np.random.randint(1, 10, 35))
```
#### Desired output
```
# array([[9, 1, 4, 8, 3],
# [2, 3, 9, 7, 2],
# [2, 9, 6, 6, 5],
# [2, 2, 7, 8, 5],
# [7, 3, 3, 9, 6],
# [2, 3, 4, 3, 3],
# [1, 6, 6, 3, 1]])
```
#### Solution
#### 1st Method
```
ser.values.reshape((7,5))
```
To reshape the ser, we call the reshape function on the series and pass a tuple with the first element the number of rows and the second element the number of columns.
#### 2nd Method
```
ser.values.reshape((-1,5))
ser.values.reshape((7,-1))
```
The other way to go about this would be to populate the tuple with only one element (row or column) and let Pandas figure out the other element to be used by placing -1 in the tuple.
### Ex 13: How to find the positions of numbers that are multiples of 3 from a series?
Q: Find the positions of numbers that are multiples of 3 from ser.
```
ser = pd.Series(np.random.randint(1, 10, 7))
```
#### Desired output
```
# 0 1
# 1 6
# dtype: int64
```
#### Solution
#### 1st Method
```
pd.Series(ser[ser%3 == 0].index)
```
We index the series and pass in the condition to return all the values that have a remainder of 0 when divided by 3. It means that those values are multiples of 3.
We then extract the indexes(positions) and cast them to a series.
#### 2nd Method
```
pd.Series(np.argwhere(ser%3==0).flatten())
```
Alternately, we could use NumPy function argwhere which returns all the values that have a remainder of 0 when divided by 3. We then flatten the array and cast it to a series.
### Ex 14: How to extract items at given positions from a series
Q: From ser, extract the items at positions in list pos.
```
ser = pd.Series(list('abcdefghijklmnopqrstuvwxyz'))
pos = [0, 4, 8, 14, 20]
```
#### Desired output
```
# 0 a
# 4 e
# 8 i
# 14 o
# 20 u
# dtype: object
```
#### Solution
#### 1st Method
```
pd.Series(ser.iloc[pos])
```
We use the iloc function to get the element at a specific index and cast to a series.
#### 2nd Method
```
ser.take(pos)
```
Alternatively, we can use the take function to achieve the same result.
### Ex 15: How to stack two series vertically?
Q: Stack ser1 and ser2 vertically to form a dataframe.
```
ser1 = pd.Series(range(5))
ser2 = pd.Series(list('abcde'))
```
#### Desired output
```
# 0 0
# 1 1
# 2 2
# 3 3
# 4 4
# 0 a
# 1 b
# 2 c
# 3 d
# 4 e
# dtype: object
```
#### Solution
```
pd.concat((ser1,ser2),axis=0)
```
To combine the two series into one, we use the concat function and pass in as a tuple the two series and set the axis to 0 to tell Pandas that we want to concatenate row-wise(vertically).
### Ex 16: How to get the positions of items of series A in another series B?
Q: Get the positions of items of ser2 in ser1 as a list.
```
ser1 = pd.Series([10, 9, 6, 5, 3, 1, 12, 8, 13])
ser2 = pd.Series([1, 3, 10, 13])
```
#### Desired output
```
# [0, 4, 5, 8]
```
#### Solution
```
list(ser1[ser1.isin(ser2)].index)
```
We use the isin function on ser1 in ser2. We get back the indexes that correspond to the positions and cast them to a list.
### Ex 17: How to compute the mean squared error on series A and predicted series B?
Q: Compute the mean squared error of truth and pred series.
```
truth = pd.Series(range(10))
pred = pd.Series(range(10)) + np.random.random(10)
```
#### Desired output
```
# Since we are generating random variable, your result will be different
#0.34688071383011976
```
#### Solution
```
np.square(np.subtract(truth,pred)).mean()
np.mean((truth-pred)**2)
```
The two notation is the same, to find the mean squared error we use its formula which pretty much translates into the code above.
Visit the [Wikipedia page](https://en.wikipedia.org/wiki/Mean_squared_error) to learn more about the mean squared error.
### Ex 18: How to convert the first character of each element in a series to uppercase?
Q: Change the first character of each word to upper case in each word of ser.
```
ser = pd.Series(['how', 'to', 'kick', 'ass?'])
```
#### Desired output
```
# 0 How
# 1 To
# 2 Kick
# 3 Ass?
# dtype: object
```
#### Solution
#### 1st Method: The pythonic way (least recommended)
```
def uppercase(the_series):
capitalized_ser = []
for word in the_series:
capitalized_ser.append(word.capitalize())
print(pd.Series(capitalized_ser))
uppercase(ser)
```
One way to solve this would be to use the vanilla Python code. We build a function that takes the series and create a new list to store the words that we will be capitalizing. We loop through the series and capitalize each word and place it in the list. We finally cast the list to a series and print it.
The reason why this is the least recommended of the bunch, it is because to achieve the result by writing five lines of code which make our code verbose.
#### 2nd Method: Using May (recommended)
```
ser.map(lambda x: x.title())
```
Ban! a much simpler method in one line of code, is to use map with lambda expression. we use the title function to capitalize each first letter of each word. We can use the capitalize function instead of the title function.
#### 3rd Method: Using Pandas built-in function (most recommended)
```
ser.str.capitalize()
```
We can call the pandas's capitalize function write away series'string.
### Ex 19: How to calculate the number of characters in each word in a series?
Q: Get the number of characters in each word in a series
```
ser = pd.Series(['how', 'to', 'kick', 'ass?'])
```
#### Desired output
```
# 0 3
# 1 2
# 2 4
# 3 4
# dtype: int64
```
#### Solution
#### 1st Method
```
ser.str.count(pat=".")
```
We can get the length of each word in the series by calling the string and the count function. We pass in the count function the pattern "." (it is a regular expression) to select any character in the word.
#### 2nd Method
```
ser.map(lambda x: len(x))
```
We can also use map with lambda expression by getting the length of each word by using len(x).
### Ex 20: How to compute the difference of differences between consecutive numbers of a series?
Q: Find the difference of differences between the consecutive numbers of ser.
```
ser = pd.Series([1, 3, 6, 10, 15, 21, 27, 35])
```
#### Desired output
```
# [nan, 2.0, 3.0, 4.0, 5.0, 6.0, 6.0, 8.0]
# [nan, nan, 1.0, 1.0, 1.0, 1.0, 0.0, 2.0]
```
#### Solution
```
ser.diff().tolist()
ser.diff().diff().tolist()
```
To calculate the difference of a series element compared with another element in the series, we use the diff function.
The first line of code we use it on the element in the series and the second time we use it on the difference list. So we have performed a difference of difference on that series.
### Ex 21: How to convert a series of date-strings to a timeseries?
Q: How to convert a series of date-strings to a timeseries?
```
ser = pd.Series(['01 Jan 2010', '02-02-2011', '20120303', '2013/04/04', '2014-05-05', '2015-06-06T12:20'])
```
#### Desired output
```
# 0 2010-01-01 00:00:00
# 1 2011-02-02 00:00:00
# 2 2012-03-03 00:00:00
# 3 2013-04-04 00:00:00
# 4 2014-05-05 00:00:00
# 5 2015-06-06 12:20:00
# dtype: datetime64[ns]
```
#### Solution
```
pd.to_datetime(ser)
```
To get the timeseries of the corresponding series, we use the function to_datetime and pass the series as the argument.
### Ex 22: How to get the day of the month, week number, day of year and day of the week from a series of date strings?
Q: Get the day of the month, week number, day of year and day of the week from ser.
```
ser = pd.Series(['01 Jan 2010', '02-02-2011', '20120303', '2013/04/04', '2014-05-05', '2015-06-06T12:20'])
```
#### Desired output
```
# Date: [1, 2, 3, 4, 5, 6]
# Week number: [53, 5, 9, 14, 19, 23]
# Day num of year: [1, 33, 63, 94, 125, 157]
# Day of week: ['Friday', 'Wednesday', 'Saturday', 'Thursday', 'Monday', 'Saturday']
```
#### Solution
```
ser_dt = pd.to_datetime(ser)
date = list(ser_dt.dt.day)
week_number = list(ser_dt.dt.week)
day_num = list(ser_dt.dt.dayofyear)
day_name = list(ser_dt.dt.day_name())
print("Date: {}\nWeek number: {}\nDay num of year: {}\nDay of week: {}".format(date,week_number,day_num,day_name))
```
We start by changing the series into a datetime, then access its dt function to get the dates, week number, day of the year and day name. Finally, we cast them to a list and print those variables.
### Ex 23: How to convert year-month string to dates corresponding to the 4th day of the month?
Q: Change ser to dates that start with 4th of the respective months.
```
ser = pd.Series(['Jan 2010', 'Feb 2011', 'Mar 2012'])
```
#### Desired output
```
# 0 2010-01-04
# 1 2011-02-04
# 2 2012-03-04
# dtype: datetime64[ns]
```
#### Solution
```
from dateutil.parser import parse
ser.map(lambda d: parse(d+" 4"))
```
For this exercise, we will need to install the parse function from the dateutile package to parse most known formats representing a date and/or time.
Then we will use the map function with a lambda expression, and parse the series concatenated with the date we want to add to the series.
### Ex 24: How to filter words that contain atleast 2 vowels from a series?
Q: From ser, extract words that contain atleast 2 vowels.
```
ser = pd.Series(['Apple', 'Orange', 'Plan', 'Python', 'Money'])
```
#### Desired output
```
# 0 Apple
# 1 Orange
# 4 Money
# dtype: object
```
#### Solution
```
vowel_count = ser.str.count(pat="(?i)[aeiou]")
vowel_count
ser[np.argwhere(vowel_count.values >= 2).flatten()]
```
We use the count function to get the count of vowels in each word by using a regular expression pattern. We get back a series with positions and the corresponding count of vowel at those positions.
We use the Numpy argwhere function to return the indexes where the condition in the paratheses is satisfied. In this example, the condition is a vowel count greater than 2. we get back 3 arrays of the indexes where the word has 2 or more vowels and then we flatten the 3 arrays into one 1D array. We use indexing to get back the words from the original series.
### Ex 25: How to filter valid emails from a series?
Extract the valid emails from the series emails. The regex pattern for valid emails is provided as reference.
```
emails = pd.Series(['buying books at amazom.com', 'rameses@egypt.com', 'matt@t.co', 'narendra@modi.com','dad@comp'])
pattern ='[A-Za-z0-9._%+-]+@[A-Za-z0-9.-]+\\.[A-Za-z]{2,4}'
```
#### Desired output
```
# 1 rameses@egypt.com
# 2 matt@t.co
# 3 narendra@modi.com
# dtype: object
```
#### Solution
```
emails[emails.str.match(pat=pattern)]
```
This exercise is similar to the previous one. This time, we use the match function to get back all the words that match the pattern and use the indexing to get those words.
### Conclusion
Have you noticed how easy Pandas is? That is why it is my favorite libraries. It is easy to grasp, and there are a plethora of resources online if you are stuck. Do not forget to visit StackOverflow too, and ask questions. There is always someone ready to help.
This post is exclusively focused on series, the primary data structure of Pandas. In the next two posts, we will explore the dataframe which is the most popular Pandas data structure. Find the jupyter notebook of this post at my GitHub profile [here.]()
Thank you again for doing these exercises with me. I hope you have learned one or two things. If you like this post, please subscribe to stay updated with new posts, and if you have a thought or a question, I would love to hear it by commenting below. Cheers, and keep learning!
| github_jupyter |
```
# default_exp data
```
# Data
> This module contains functions to download and preprocess the data
```
#hide
from nbdev.export import notebook2script
#export
import ee
import os
import requests
import rasterio
import pandas as pd
import numpy as np
import zipfile
import json
from IPython.core.debugger import set_trace
from pathlib import Path
import warnings
from fastprogress.fastprogress import progress_bar
from banet.geo import open_tif, merge, Region
from banet.geo import downsample
#export
class RegionST(Region):
"Defines a region in space and time with a name, a bounding box and the pixel size."
def __init__(self, name:str, bbox:list, pixel_size:float=None, scale_meters:int=None,
time_start:str=None, time_end:str=None, time_freq:str='D', time_margin:int=0,
shape:tuple=None, epsg=4326):
if scale_meters is not None and pixel_size is not None:
raise Exception('Either pixel_size or scale_meters must be set to None.')
self.name = name
self.bbox = rasterio.coords.BoundingBox(*bbox) # left, bottom, right, top
if pixel_size is not None:
self.pixel_size = pixel_size
else:
self.pixel_size = scale_meters/111000
self.epsg = epsg
self.scale_meters = scale_meters
self._shape = shape
self.time_start = pd.Timestamp(str(time_start))
self.time_end = pd.Timestamp(str(time_end))
self.time_margin = time_margin
self.time_freq = time_freq
@property
def shape(self):
"Shape of the region (height, width)"
if self._shape is None:
return (self.height, self.width)
else: return self._shape
@property
def times(self):
"Property that computes the date_range for the region."
tstart = self.time_start - pd.Timedelta(days=self.time_margin)
tend = self.time_end + pd.Timedelta(days=self.time_margin)
return pd.date_range(tstart, tend, freq=self.time_freq)
@classmethod
def load(cls, file, time_start=None, time_end=None):
"Loads region information from json file"
with open(file, 'r') as f:
args = json.load(f)
if time_start is None:
time_start = args['time_start']
if time_end is None:
time_end = args['time_end']
return cls(args['name'], args['bbox'], args['pixel_size'],
time_start=time_start, time_end=time_end)
def extract_region(df_row, cls=Region):
"Create Region object from a row of the metadata dataframe."
if issubclass(cls, RegionST):
return cls(df_row.event_id, df_row.bbox, df_row.pixel_size,
df_row.time_start, df_row.time_end)
elif issubclass(cls, Region):
return cls(df_row.event_id, df_row.bbox, df_row.pixel_size)
else: raise NotImplemented('cls must be one of the following [Region, RegionST]')
#export
def coords2bbox(lon, lat, pixel_size):
return [lon.min(), lat.min(), lon.max()+pixel_size, lat.max()+pixel_size]
def split_region(region:RegionST, size:int, cls=Region):
lon, lat = region.coords()
Nlon = (len(lon)//size)*size
Nlat = (len(lat)//size)*size
lons = [*lon[:Nlon].reshape(-1, size), lon[Nlon:][None]]
lats = [*lat[:Nlat].reshape(-1, size), lat[Nlat:][None]]
if len(lats[-1].reshape(-1)) == 0 and len(lons[-1].reshape(-1)) == 0:
lons = lons[:-1]
lats = lats[:-1]
#lons = lon.reshape(-1, size)
#lats = lat.reshape(-1, size)
if issubclass(cls, RegionST):
return [cls('', coords2bbox(ilon, ilat, region.pixel_size),
pixel_size=region.pixel_size, time_start=region.time_start,
time_end=region.time_end, time_freq=region.time_freq,
time_margin=region.time_margin) for ilon in lons for ilat in lats]
elif issubclass(cls, Region):
return [cls('', coords2bbox(ilon, ilat, region.pixel_size), pixel_size=region.pixel_size)
for ilon in lons for ilat in lats]
else: raise NotImplemented('cls must be one of the following [Region, RegionST]')
return
def merge_tifs(files:list, fname:str, delete=False):
data, tfm = merge([open_tif(str(f)) for f in files])
data = data.squeeze()
fname = Path(files[0]).parent/fname
profile = open_tif(str(files[0])).profile
with rasterio.Env():
height, width = data.shape
profile.update(width=width, height=height, transform=tfm, compress='lzw')
with rasterio.open(str(fname), 'w', **profile) as dst:
dst.write(data, 1)
if delete:
for f in files: os.remove(f)
#export
def filter_region(image_collection:ee.ImageCollection, region:RegionST, times:tuple, bands=None):
image_collection = image_collection.filterDate(times[0], times[1])
geometry = ee.Geometry.Rectangle(region.bbox)
image_collection = image_collection.filterBounds(geometry)
if bands is not None:
image_collection = image_collection.select(bands)
return image_collection
def filter_cloudy(image_collection:ee.ImageCollection, max_cloud_fraction=0.2):
return image_collection.filterMetadata(
'CLOUDY_PIXEL_PERCENTAGE', 'not_greater_than', max_cloud_fraction)
def n_least_cloudy(image_collection:ee.ImageCollection, n=5):
image_collection = image_collection.sort(prop='CLOUDY_PIXEL_PERCENTAGE')
image_collection = image_collection.toList(image_collection.size())
colsize = image_collection.size().getInfo()
if colsize < n:
warnings.warn(f'Total number of images in the collection {colsize} less than n={n}. Setting n={colsize}')
n = colsize
image_collection = ee.ImageCollection([ee.Image(image_collection.get(i)) for i in range(n)])
return image_collection
def download_topography_data(R:RegionST, path_save=Path('.'), scale=None,
download_crop_size=1000, show_progress=False):
if scale is None: scale = R.scale_meters
ee.Initialize()
image = ee.Image('srtm90_v4')
path_save.mkdir(exist_ok=True, parents=True)
sR = [R] if min(R.shape) <= download_crop_size else split_region(R, size=download_crop_size)
if not (path_save/'srtm90_v4.elevation.tif').is_file():
files = []
loop = enumerate(sR) if not show_progress else progress_bar(enumerate(sR),total=len(sR))
for j, R in loop:
region = (f"[[{R.bbox.left}, {R.bbox.bottom}], [{R.bbox.right}, {R.bbox.bottom}], " +
f"[{R.bbox.right}, {R.bbox.top}], [{R.bbox.left}, {R.bbox.top}]]")
url = image.getDownloadUrl(
{'scale': scale, 'crs': 'EPSG:4326', 'region': f'{region}'})
r = requests.get(url)
with open(str(path_save/'data.zip'), 'wb') as f:
f.write(r.content)
with zipfile.ZipFile(str(path_save/'data.zip'), 'r') as f:
f.extractall(str(path_save))
os.rename(str(path_save/'srtm90_v4.elevation.tif'),
str(path_save/f'srtm90_v4.elevation_{j}.tif'))
files.append(str(path_save/f'srtm90_v4.elevation_{j}.tif'))
os.remove(str(path_save/'data.zip'))
merge_tifs(files, 'srtm90_v4.elevation.tif', delete=True)
def download_data(R:RegionST, times, products, bands, path_save, scale=None, max_cloud_fraction=None,
use_least_cloudy=None, download_crop_size=1000, show_progress=False):
if scale is None: scale = R.scale_meters
ee.Initialize()
path_save.mkdir(exist_ok=True, parents=True)
if not ((path_save/f'download.{bands[0]}.tif').is_file() and
(path_save/f'download.{bands[1]}.tif').is_file() and
(path_save/f'download.{bands[2]}.tif').is_file()):
sR = [R] if min(R.shape) <= download_crop_size else split_region(R, size=download_crop_size, cls=RegionST)
fsaves = []
#for j, R in tqdm(enumerate(sR), total=len(sR)):
loop = enumerate(sR) if not show_progress else progress_bar(enumerate(sR),total=len(sR))
for j, R in loop:
region = (f"[[{R.bbox.left}, {R.bbox.bottom}], [{R.bbox.right}, {R.bbox.bottom}], " +
f"[{R.bbox.right}, {R.bbox.top}], [{R.bbox.left}, {R.bbox.top}]]")
if not ((path_save/f'download.{bands[0]}_{j}.tif').is_file() and
(path_save/f'download.{bands[1]}_{j}.tif').is_file() and
(path_save/f'download.{bands[2]}_{j}.tif').is_file()):
# Merge products to single image collection
imCol = ee.ImageCollection(products[0])
for i in range(1, len(products)):
imCol = imCol.merge(ee.ImageCollection(products[i]))
imCol = filter_region(imCol, R, times=times, bands=bands)
if max_cloud_fraction is not None:
imCol = filter_cloudy(imCol, max_cloud_fraction=max_cloud_fraction)
if use_least_cloudy is not None:
imCol = n_least_cloudy(imCol, n=use_least_cloudy)
im = imCol.median()
imCol = ee.ImageCollection([im])
colList = imCol.toList(imCol.size())
# info = colList.getInfo()
# data_times = [pd.to_datetime(o['properties']['system:time_start'], unit='ms') for o in info]
# data_cloudy = [o['properties']['CLOUDY_PIXEL_PERCENTAGE'] for o in info]
# Download each image
for i in range(colList.size().getInfo()):
image = ee.Image(colList.get(i))
fname = 'download'
#fname = image.get('system:id').getInfo().split('/')[-1]
fnames_full = [f'{fname}.{b}.tif' for b in bands]
fnames_partial0 = [f'{fname}.{b}_{j}.tif' for b in bands]
fnames_full = all([(path_save/f).is_file() for f in fnames_full])
fnames_partial = all([(path_save/f).is_file() for f in fnames_partial0])
if not fnames_full:
fsaves.append([path_save/f for f in fnames_partial0])
if not fnames_partial:
zip_error = True
for i in range(10): # Try 10 times
if zip_error:
try:
url = image.getDownloadURL(
{'scale': scale, 'crs': 'EPSG:4326',
'region': f'{region}'})
r = requests.get(url)
with open(str(path_save/'data.zip'), 'wb') as f:
f.write(r.content)
with zipfile.ZipFile(str(path_save/'data.zip'), 'r') as f:
files = f.namelist()
f.extractall(str(path_save))
os.remove(str(path_save/'data.zip'))
zip_error = False
except:
zip_error = True
os.remove(str(path_save/'data.zip'))
time.sleep(10)
if zip_error: raise Exception(f'Failed to process {url}')
for f in files:
f = path_save/f
os.rename(str(f), str(path_save/f'{f.stem}_{j}{f.suffix}'))
# Merge files
suffix = '.tif'
files = path_save.ls(include=[suffix])
#files = np.unique(fsaves)
files = [o.stem for o in files]
ref = np.unique(['_'.join(o.split('_')[:-1])
for o in files if len(o.split('_')[-1]) < 6])
ids = np.unique([int(o.split('_')[-1])
for o in files if len(o.split('_')[-1]) < 6])
#file_groups = [[path_save/f'{r}_{i}{suffix}' for i in ids] for r in ref]
file_groups = [[path_save/f'{r}_{i}{suffix}' for i in ids
if f'{r}_{i}' in files] for r in ref]
for fs in file_groups:
if len(fs) < 500:
fsave = '_'.join(fs[0].stem.split('_')[:-1]) + suffix
merge_tifs(fs, fsave, delete=True)
else:
fs_break = np.array(fs)[:(len(fs)//500)*500].reshape(len(fs)//500,-1).tolist()
if len(fs[(len(fs)//500)*500:]) > 0:
fs_break.append(fs[(len(fs)//500)*500:])
for fsi, fs2 in enumerate(fs_break):
fsave = '_'.join(fs2[0].stem.split('_')[:-1]) + f'_break{fsi}' + suffix
merge_tifs(fs2, fsave, delete=True)
files = path_save.ls(include=[suffix, '_break'])
files = [o.stem for o in files]
ref = np.unique(['_'.join(o.split('_')[:-1])
for o in files if len(o.split('_')[-1]) < 11])
ids = np.unique([o.split('_')[-1]
for o in files if len(o.split('_')[-1]) < 11])
#file_groups = [[path_save/f'{r}_{i}{suffix}' for i in ids] for r in ref]
file_groups = [[path_save/f'{r}_{i}{suffix}' for i in ids
if f'{r}_{i}' in files] for r in ref]
for fs in file_groups:
fsave = '_'.join(fs[0].stem.split('_')[:-1]) + suffix
merge_tifs(fs, fsave, delete=True)
def download_data_ts(R:RegionST, products, bands, path_save, scale=None,
download_crop_size=1000, show_progress=False):
if scale is None: scale = R.scale_meters
ee.Initialize()
times = (R.times[0], R.times[-1])
path_save.mkdir(exist_ok=True, parents=True)
sR = [R] if min(R.shape) <= download_crop_size else split_region(R, size=download_crop_size, cls=RegionST)
loop = enumerate(sR) if not show_progress else progress_bar(enumerate(sR),total=len(sR))
for j, R in loop:
region = (f"[[{R.bbox.left}, {R.bbox.bottom}], [{R.bbox.right}, {R.bbox.bottom}], " +
f"[{R.bbox.right}, {R.bbox.top}], [{R.bbox.left}, {R.bbox.top}]]")
# Merge products to single image collection
imCol = ee.ImageCollection(products[0])
for i in range(1, len(products)):
imCol = imCol.merge(ee.ImageCollection(products[i]))
imCol = filter_region(imCol, R, times=times, bands=bands)
imCol = ee.ImageCollection(imCol)
colList = imCol.toList(imCol.size())
# Download each image
for i in range(colList.size().getInfo()):
image = ee.Image(colList.get(i))
zip_error = True
for i in range(10): # Try 10 times
if zip_error:
try:
url = image.getDownloadURL(
{'scale': scale, 'crs': 'EPSG:4326',
'region': f'{region}'})
r = requests.get(url)
with open(str(path_save/'data.zip'), 'wb') as f:
f.write(r.content)
with zipfile.ZipFile(str(path_save/'data.zip'), 'r') as f:
files = f.namelist()
f.extractall(str(path_save))
os.remove(str(path_save/'data.zip'))
zip_error = False
except:
zip_error = True
os.remove(str(path_save/'data.zip'))
time.sleep(10)
if zip_error: raise Exception(f'Failed to process {url}')
for f in files:
f = path_save/f
os.rename(str(f), str(path_save/f'{f.stem}_{j}{f.suffix}'))
# Merge files
suffix = '.tif'
files = path_save.ls(include=[suffix])
files = [o.stem for o in files]
ref = np.unique(['_'.join(o.split('_')[:-1])
for o in files if len(o.split('_')[-1]) < 6])
ids = np.unique([int(o.split('_')[-1])
for o in files if len(o.split('_')[-1]) < 6])
file_groups = [[path_save/f'{r}_{i}{suffix}' for i in ids
if f'{r}_{i}' in files] for r in ref]
for fs in file_groups:
if len(fs) < 500:
fsave = '_'.join(fs[0].stem.split('_')[:-1]) + suffix
merge_tifs(fs, fsave, delete=True)
else:
fs_break = np.array(fs)[:(len(fs)//500)*500].reshape(len(fs)//500,-1).tolist()
if len(fs[(len(fs)//500)*500:]) > 0:
fs_break.append(fs[(len(fs)//500)*500:])
for fsi, fs2 in enumerate(fs_break):
fsave = '_'.join(fs2[0].stem.split('_')[:-1]) + f'_break{fsi}' + suffix
merge_tifs(fs2, fsave, delete=True)
files = path_save.ls(include=[suffix, '_break'])
files = [o.stem for o in files]
ref = np.unique(['_'.join(o.split('_')[:-1])
for o in files if len(o.split('_')[-1]) < 11])
ids = np.unique([o.split('_')[-1]
for o in files if len(o.split('_')[-1]) < 11])
file_groups = [[path_save/f'{r}_{i}{suffix}' for i in ids
if f'{r}_{i}' in files] for r in ref]
for fs in file_groups:
fsave = '_'.join(fs[0].stem.split('_')[:-1]) + suffix
merge_tifs(fs, fsave, delete=True)
```
Download median composite for any region example:
```python
R = RegionST('test_region1', [-8.0,39.95,-7.9,40.05], 0.001,
time_start='2020-07-01', time_end='2020-07-15')
R.time_margin=1
products = ["COPERNICUS/S2"]
bands = ['B4', 'B8', 'B12']
path = Path('temp')
before = (R.times[0]-pd.Timedelta(days=120), R.times[0])
after = (R.times[-1], R.times[-1]+pd.Timedelta(days=120))
for mode, time_window in zip(['before', 'after'], [before, after]):
path_save = path/R.name/mode
download_data(R, time_window, products, bands, path_save,
use_least_cloudy=5)
```
Download all data for any region example:
```python
R = RegionST('test_region2', [-8.0,39.95,-7.9,40.05], 0.001,
time_start='2020-07-01', time_end='2020-08-31')
R.time_margin=0
products = ["COPERNICUS/S2"]
bands = ['B4', 'B8', 'B12']
path_save = Path('temp')/R.name
download_data_ts(R, products, bands, path_save)
```
```
#export
def get_event_data(event_id, year, coarse_mask_file, path=Path('.'),
coarse_mask_doy_layer=2, products=['COPERNICUS/S2'],
bands=['B4', 'B8', 'B12'], scale_factor=1e-4, composite_days=[60,60],
max_cloud_fraction=None, use_least_cloudy=None, scale=10,
topography=False, banet_pixel_size=0.001):
rst_ba100 = open_tif(coarse_mask_file)
doys = rst_ba100.read(coarse_mask_doy_layer).astype(np.float16)
doys[doys==0] = np.nan
doy_start, doy_end = np.nanmin(doys), np.nanmax(doys)
del doys
time_start = pd.Timestamp(f'{year}-01-01') + pd.Timedelta(days=doy_start-1)
time_end = pd.Timestamp(f'{year}-01-01') + pd.Timedelta(days=doy_end-1)
print('Event time_start:', str(time_start))
print('Event time_end:', str(time_end))
R = RegionST(event_id, list(rst_ba100.bounds), scale_meters=scale,
time_start=time_start, time_end=time_end, time_margin=1)
R_banet = R.new(pixel_size=banet_pixel_size)
before = (R.times[0]-pd.Timedelta(days=composite_days[0]), R.times[0])
after = (R.times[-1], R.times[-1]+pd.Timedelta(days=composite_days[1]))
for mode, time_window in zip(['before', 'after'], [before, after]):
path_save = path/R.name/mode
print('Downloading GEE median composite for:', ' to '.join([str(o) for o in time_window]))
download_data(R, time_window, products, bands, path_save,
max_cloud_fraction=max_cloud_fraction, use_least_cloudy=use_least_cloudy,
scale=scale)
if topography:
print('Downloading topography data.')
download_topography_data(R, path/event_id/'topography', scale=scale)
rst_ba100 = rst_ba100.read(coarse_mask_doy_layer)
s10before_files = np.array((path/R.name/'before').ls(exclude=['.xml']))[[1,2,0]].tolist()
s10after_files = np.array((path/R.name/'after').ls(exclude=['.xml']))[[1,2,0]].tolist()
transform = rasterio.open(str(s10before_files[0])).transform
crs = rasterio.open(str(s10before_files[0])).crs
rst_s10before = np.concatenate(
[rasterio.open(str(f)).read() for f in s10before_files]).astype(np.float16)*scale_factor
rst_s10after = np.concatenate(
[rasterio.open(str(f)).read() for f in s10after_files]).astype(np.float16)*scale_factor
rst_ba100 = downsample(rst_ba100, src_tfm=R_banet.transform, dst_tfm=transform,
dst_shape=(1, *rst_s10before.shape[-2:]), resampling='bilinear').astype(np.float32)
im = np.concatenate([rst_s10before, rst_s10after, rst_ba100], axis=0).transpose(1,2,0)
return im, transform, crs
#local
im, transform, crs = get_event_data('temp', 2020, 'temp/banet100m.tif', topography=True)
im.shape, transform, crs
#hide
notebook2script()
```
| github_jupyter |
# GraphRNN
```
!git clone --single-branch --branch colab https://github.com/joaopedromattos/GraphRNN
!pip install gdown
!gdown --id 1RF_bIo5ndxPhu9SJw-T8HBcuHyaGQGL0 && tar -xzvf datasets.tar.gz
!mv GraphRNN/* .
!mkdir ./dataset/EVENT
```
## Preparing our graph
```
import networkx as nx
import numpy as np
G = nx.read_gpickle('./datasets_runs/run_1_gold_standard_5w1h_graph_hin.nx') # selecting the graph
len(G.nodes), len(G.edges)
```
### Indicator file
```
# GraphRNN receives a file with a number in every ith line,
# that represents the graph to which the ith node belongs to.
# E.g.: line 85824 with a value 222 means that the node 85824 belongs to
# the graph number 222.
node_mapper = {i : v for i, v in enumerate(G.nodes)}
node_mapper.keys()
np.savetxt("EVENT_graph_indicator.txt", np.ones(shape=len(G.nodes)), fmt='%d')
```
### Adj Matrix
```
# GraphRNN receives an edgelist to mount an adjacency matrix
# inside data.py file on Graph_load_batch method.
G_relabel = nx.relabel_nodes(G, {v : k for k, v in node_mapper.items()})
nx.write_edgelist(G_relabel, "EVENT_A.txt", data=False, delimiter=', ')
```
### Node Labels
```
# Here we get a list of all labels of all nodes.
# In case of non-labeled nodes, we manually label them with "no_label"
labels = [G.nodes[v]['label'] if 'label' in G.nodes[v] else 'no_label' for i, v in enumerate(G.nodes)]
# We'll give a unique natural number to each label of our graph.
label_mapper = dict()
count = 0
for i, v in enumerate(labels):
if (not (v in label_mapper) ):
label_mapper[v] = count
count += 1
print(label_mapper)
# Mapping our labels to natural numbers and writing them to a file.
node_labels_list = list(map(lambda x: label_mapper[x], labels))
print(node_labels_list)
np.savetxt("EVENT_node_labels.txt", node_labels_list, fmt='%d')
!mv EVENT_* dataset/EVENT/
```
## Running GraphRNN
```
!pip install -r requirements.txt
!python main.py
```
## Converting our outputs
```
import pickle
G_pred_list = pickle.load( open( "graphs/GraphRNN_RNN_EVENT_4_128_pred_10_1_4000_nodes.dat", "rb" ) )
G_pred_list
graph_list = []
for i in G_pred_list:
cur_graph_edges = [(j, k, i.edge[j][k]['weight']) for j in i.edge.keys() for k in i.edge[j]]
test_graph = nx.DiGraph()
test_graph.add_nodes_from(i.node)
test_graph.add_weighted_edges_from(cur_graph_edges)
graph_list.append(test_graph)
print("Nodes, edges:", len(i.node.keys()), len(cur_graph_edges))
pickle.dump(graph_list, open('graph_list.dat', 'wb'))
```
| github_jupyter |
## 神经网络实现翻译
- 参考链接 : https://pytorch.org/tutorials/intermediate/seq2seq_translation_tutorial.html
- 论文参考链接 : https://arxiv.org/abs/1409.3215
In this project we will be teaching a neural network to translate from French to English.
最终实现的目标如下
```python
[KEY: > input, = target, < output]
> il est en train de peindre un tableau .
= he is painting a picture .
< he is painting a picture .
> pourquoi ne pas essayer ce vin delicieux ?
= why not try that delicious wine ?
< why not try that delicious wine ?
> elle n est pas poete mais romanciere .
= she is not a poet but a novelist .
< she not not a poet but a novelist .
> vous etes trop maigre .
= you re too skinny .
< you re all alone .
```
### 主要思想
An encoder network condenses an input sequence into a vector, and a decoder network unfolds that vector into a new sequence.
```
from __future__ import unicode_literals, print_function, division
from io import open
import unicodedata
import string
import re
import random
import torch
import torch.nn as nn
from torch import optim
import torch.nn.functional as F
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
```
## 数据预处理&读取数据
```
SOS_token = 0
EOS_token = 1
class Lang:
"""word → index (word2index) and index → word (index2word) dictionaries
A count of each word word2count to use to later replace rare words.
"""
def __init__(self, name):
self.name = name
self.word2index = {}
self.word2count = {}
self.index2word = {0: "SOS", 1: "EOS"}
self.n_words = 2 # Count SOS and EOS
def addSentence(self, sentence):
for word in sentence.split(' '):
self.addWord(word)
def addWord(self, word):
if word not in self.word2index:
self.word2index[word] = self.n_words
self.word2count[word] = 1
self.index2word[self.n_words] = word
self.n_words += 1
else:
self.word2count[word] += 1
"""
we will turn Unicode characters to ASCII, make everything lowercase,
and trim most punctuation.
"""
# Turn a Unicode string to plain ASCII, thanks to
# https://stackoverflow.com/a/518232/2809427
def unicodeToAscii(s):
return ''.join(
c for c in unicodedata.normalize('NFD', s)
if unicodedata.category(c) != 'Mn'
)
# Lowercase, trim, and remove non-letter characters
def normalizeString(s):
s = unicodeToAscii(s.lower().strip())
s = re.sub(r"([.!?])", r" \1", s)
s = re.sub(r"[^a-zA-Z.!?]+", r" ", s)
return s
# 转换为ASCII, 大写变小写, 留下重要的标点, 去掉大部分的标点
normalizeString('I am a Boy!~$%^&')
def readLangs(lang1, lang2, reverse=False):
"""逐行读取file, 并将每行分为pair, 并做标准化
"""
print("Reading lines...")
# Read the file and split into lines
lines = open('./data/%s-%s.txt' % (lang1, lang2), encoding='utf-8').\
read().strip().split('\n')
# Split every line into pairs and normalize
pairs = [[normalizeString(s) for s in l.split('\t')] for l in lines]
# Reverse pairs, make Lang instances
if reverse:
pairs = [list(reversed(p)) for p in pairs]
input_lang = Lang(lang2)
output_lang = Lang(lang1)
else:
input_lang = Lang(lang1)
output_lang = Lang(lang2)
return input_lang, output_lang, pairs
```
为了加快训练的速度, 我们把句子长度最大设置为10, 同时我们过滤句子后使得其开头变为如i am, he is等词汇
```
MAX_LENGTH = 10
eng_prefixes = (
"i am ", "i m ",
"he is", "he s ",
"she is", "she s ",
"you are", "you re ",
"we are", "we re ",
"they are", "they re "
)
def filterPair(p):
return len(p[0].split(' ')) < MAX_LENGTH and \
len(p[1].split(' ')) < MAX_LENGTH and \
p[1].startswith(eng_prefixes)
def filterPairs(pairs):
return [pair for pair in pairs if filterPair(pair)]
# 会去掉单词个数超过10个的句子
# 会去掉不是以特定开头的句子
filterPairs([['i am a girl','i am a boy'],
['how are you','how are you'],
['i am a girl i am a girl i am a girl','i am a girl i am a girl']])
```
The full process for preparing the data is:
- Read text file and split into lines, split lines into pairs
- Normalize text, filter by length and content
- Make word lists from sentences in pairs
```
def prepareData(lang1, lang2, reverse=False):
"""开始读取语言的文件
"""
# 读取文件, 返回的是句子对
input_lang, output_lang, pairs = readLangs(lang1, lang2, reverse)
print("Read %s sentence pairs" % len(pairs))
# 过滤掉句子对中较长的句子, 和
pairs = filterPairs(pairs)
print("Trimmed to %s sentence pairs" % len(pairs))
print("Counting words...")
for pair in pairs:
input_lang.addSentence(pair[0])
output_lang.addSentence(pair[1])
print("Counted words:")
print(input_lang.name, input_lang.n_words)
print(output_lang.name, output_lang.n_words)
return input_lang, output_lang, pairs
# 开始读取数据
input_lang, output_lang, pairs = prepareData('eng', 'fra', True)
print(random.choice(pairs))
# 不同单词出现的次数
output_lang.word2index
```
### Preparing Training Data
To train, for each pair we will need an input tensor (indexes of the words in the input sentence) and target tensor (indexes of the words in the target sentence). While creating these vectors we will append the EOS token to both sequences.
```
def indexesFromSentence(lang, sentence):
return [lang.word2index[word] for word in sentence.split(' ')]
def tensorFromSentence(lang, sentence):
indexes = indexesFromSentence(lang, sentence)
indexes.append(EOS_token)
return torch.tensor(indexes, dtype=torch.long, device=device).view(-1, 1)
def tensorsFromPair(pair):
input_tensor = tensorFromSentence(input_lang, pair[0])
target_tensor = tensorFromSentence(output_lang, pair[1])
return (input_tensor, target_tensor)
# 将一句话中的每个字母转为Index, 并在结尾加上终止符
tensorFromSentence(output_lang, 'i am a boy')
```
## The Seq2Seq Model
### Seq2Seq的好处
对比传统的单层的RNN来说, 可以不需要输入和输出是相同的长度的.
下面是完整的思想, 这里的原文还是很不错的.
Unlike sequence prediction with a single RNN, where every input corresponds to an output, the seq2seq model frees us from sequence length and order, which makes it ideal for translation between two languages.
Consider the sentence “Je ne suis pas le chat noir” → “I am not the black cat”. Most of the words in the input sentence have a direct translation in the output sentence, but are in slightly different orders, e.g. “chat noir” and “black cat”. Because of the “ne/pas” construction there is also one more word in the input sentence. It would be difficult to produce a correct translation directly from the sequence of input words.
With a seq2seq model the encoder creates a single vector which, in the ideal case, encodes the “meaning” of the input sequence into a single vector — a single point in some N dimensional space of sentences.
### The Encoder
The encoder of a seq2seq network is a RNN that outputs some value for every word from the input sentence. For every input word the encoder outputs a vector and a hidden state, and uses the hidden state for the next input word.
```
class EncoderRNN(nn.Module):
def __init__(self, input_size, hidden_size, batch_size=1, n_layers=1):
super(EncoderRNN, self).__init__()
self.input_size = input_size
self.hidden_size = hidden_size
self.batch_size = batch_size # 输入的时候batch_size
self.n_layers = n_layers # RNN中的层数
self.embedding = nn.Embedding(self.input_size, self.hidden_size)
self.gru = nn.GRU(self.hidden_size, self.hidden_size)
def forward(self, x):
self.sentence_length = x.size(0) # 获取一句话的长度
embedded = self.embedding(x).view(self.sentence_length, 1, -1) # seq_len * batch_size * word_size
output = embedded
self.hidden = self.initHidden()
output, hidden = self.gru(output, self.hidden)
return output, hidden
def initHidden(self):
return torch.zeros(self.n_layers, self.batch_size, self.hidden_size).to(device)
test_data = tensorsFromPair(random.choice(pairs))
test_data[0]
# encoder测试
encoder1 = EncoderRNN(input_lang.n_words, 256).to(device)
output, hidden = encoder1(test_data[0].unsqueeze(1))
output.shape
hidden.shape
```
### The Decoder
The decoder is another RNN that takes the encoder output vector(s) and outputs a sequence of words to create the translation.
#### Simple Decoder
In the simplest seq2seq decoder we use only last output of the encoder. This last output is sometimes called the context vector as it encodes context from the entire sequence. This context vector is used as the initial hidden state of the decoder.
At every step of decoding, the decoder is given an input token and hidden state. The initial input token is the start-of-string <SOS> token, and the first hidden state is the context vector (the encoder’s last hidden state).
### 问题
- decoder下面每次出入的x是什么=>上一次的output
- 如何计算误差
### The Attention Decoder
If only the context vector is passed betweeen the encoder and decoder, that single vector carries the burden of encoding the entire sentence.(词向量会有整个句子的含义)
Attention allows the decoder network to “focus” on a different part of the encoder’s outputs for every step of the decoder’s own outputs. First we calculate a set of attention weights. These will be multiplied by the encoder output vectors to create a weighted combination. The result (called attn_applied in the code) should contain information about that specific part of the input sequence, and thus help the decoder choose the right output words.

```
class AttenDecoder(nn.Module):
def __init__(self, hidden_size, output_size, dropout_p=0.1, max_length=MAX_LENGTH):
super(AttenDecoder, self).__init__()
self.hidden_size = hidden_size
self.output_size = output_size
self.dropout_p = dropout_p
self.max_length = max_length
self.embedding = nn.Embedding(self.output_size, self.hidden_size)
self.attn = nn.Linear(self.hidden_size*2, self.max_length)
self.attn_combine = nn.Linear(self.hidden_size*2, self.hidden_size)
self.dropout = nn.Dropout(self.dropout_p)
self.gru = nn.GRU(self.hidden_size, self.hidden_size)
self.out = nn.Linear(self.hidden_size, self.output_size)
def forward(self, x, hidden, encoder_outputs):
# x是输入, 这里有两种类型
# hidden是上一层中隐藏层的内容
# encoder_outputs里面是encoder的RNN的每一步的输出(不是最后一个的输出)
embedded = self.embedding(x).view(1,1,-1)
embedded = self.dropout(embedded)
# print('embedded.shape',embedded.shape)
# ----------------------------
# 下面的attention weight表示:
# 连接输入的词向量和上一步的hide state并建立bp训练,
# 他们决定了attention权重
# -----------------------------
attn_weights = torch.cat((embedded[0],hidden[0]),1)
# print('attn_weights1',attn_weights.shape)
attn_weights = self.attn(attn_weights)
attn_weights = F.softmax(attn_weights,dim=1)
# print('attn_weights2',attn_weights.shape)
# 这是做矩阵乘法
# 施加权重到所有的语义向量上
attn_applied = torch.bmm(attn_weights.unsqueeze(0), encoder_outputs.unsqueeze(0))
# print('attn_applied',attn_applied.shape)
# 加了attention的语义向量和输入的词向量共同作为输
# 此处对应解码方式三+attention
output = torch.cat((embedded[0],attn_applied[0]),1)
# print('output1',output.shape)
# 进入RNN之前,先过了一个全连接层
output = self.attn_combine(output).unsqueeze(0)
# print('output2',output.shape)
output = F.relu(output)
output, hidden = self.gru(output, hidden)
# print('output3',output.shape)
# 输出分类结果
output = F.log_softmax(self.out(output[0]),dim=1)
# print('output4',output.shape)
return output, hidden, attn_weights
# output是encoder的输出, hidden是encoder的最后的hidden state
output.shape, hidden.shape
# 这个是第一个输入, 代表起始符
test_data = torch.tensor([[SOS_token]]).to(device)
test_data
# decoder测试
decoder1 = AttenDecoder(256, output_lang.n_words, max_length=output.size(0)).to(device)
output, hidden, attn_weights = decoder1(test_data, hidden, output.squeeze(1))
test_output = torch.tensor([[1,2,3],[4,5,6]]).float()
test_outputs = torch.zeros((5,3))
test_outputs[:test_output.size(0)]=test_output
test_outputs
```
## Training
### Training the Model
To train we run the input sentence through the encoder, and keep track of every output and the latest hidden state. Then the decoder is given the <SOS> token as its first input, and the last hidden state of the encoder as its first hidden state.(总体训练流程)
```
# helper function
import time
import math
def asMinutes(s):
"""将秒转换为分钟
"""
m = math.floor(s / 60)
s -= m * 60
return '%dm %ds' % (m, s)
def timeSince(since, percent):
"""打印已经花费的时间和预计花费的时间
预计花费的时间, 用 完成百分比的时间/现在完成的百分比 来预测
"""
now = time.time()
s = now - since
es = s / (percent)
rs = es - s
return '%s (- %s)' % (asMinutes(s), asMinutes(rs))
teacher_forcing_ratio = 0.5 # 50%的概率使用teacher_forcing的模式
def train(input_tensor, target_tensor, encoder, decoder, encoder_optimizer, decoder_optimizer, criterion, max_length=MAX_LENGTH):
encoder_optimizer.zero_grad()
decoder_optimizer.zero_grad()
input_length = input_tensor.size(0)
target_length = target_tensor.size(0)
# encoder_outputs = torch.zeros(max_length, encoder.hidden_size).to(device)
# Encoder
encoder_output, encoder_hidden = encoder1(input_tensor.unsqueeze(1))
# 因为一个Decoder的MAX_LENGTH是固定长度的, 所以我们需要将encoder_output变为一样长的
encoder_output = encoder_output.squeeze(1)
encoder_outputs = torch.zeros(max_length, encoder_output.size(1)).to(device)
encoder_outputs[:encoder_output.size(0)] = encoder_output
# Decoder
loss = 0
decoder_hidden = encoder_hidden # encoder最后的hidden作为decoder的hidden
decoder_input = torch.tensor([[SOS_token]]).to(device)
# 判断是使用哪一种模式
use_teacher_forcing = True if random.random() < teacher_forcing_ratio else False
if use_teacher_forcing:
# Teacher forcing: Feed the target as the next input
for di in range(target_length):
decoder_output, decoder_hidden, attn_weights = decoder(decoder_input, decoder_hidden, encoder_outputs)
loss = loss + criterion(decoder_output, target_tensor[di])
decoder_input = target_tensor[di] # Teacher Forcing
else:
# Without teacher forcing: use its own predictions as the next input
for di in range(target_length):
decoder_output, decoder_hidden, attn_weights = decoder(decoder_input, decoder_hidden, encoder_outputs)
topv, topi = decoder_output.topk(1)
decoder_input = topi.squeeze().detach() # detach from history as input
loss = loss + criterion(decoder_output, target_tensor[di])
if decoder_input.item() == EOS_token:
break
# 反向传播, 进行优化
loss.backward()
encoder_optimizer.step()
decoder_optimizer.step()
return loss.item() / target_length
```
The whole training process looks like this:
- Start a timer
- Initialize optimizers and criterion
- Create set of training pairs
- Start empty losses array for plotting
Then we call train many times and occasionally print the progress (% of examples, time so far, estimated time) and average loss.
```
def trainIters(encoder, decoder, n_iters, print_every=1000, plot_every=100, learning_rate=0.01):
start = time.time()
plot_losses = []
print_loss_total = 0 # Reset every print_every
plot_loss_total = 0 # Reset every plot_every
# 初始化优化器
encoder_optimizer = optim.SGD(encoder.parameters(), lr=learning_rate)
decoder_optimizer = optim.SGD(decoder.parameters(), lr=learning_rate)
# 初始化样本
training_pairs = [tensorsFromPair(random.choice(pairs)) for i in range(n_iters)]
criterion = nn.NLLLoss()
for iter in range(1, n_iters+1):
training_pair = training_pairs[iter-1]
input_tensor = training_pair[0]
target_tensor = training_pair[1]
loss = train(input_tensor, target_tensor, encoder, decoder, encoder_optimizer, decoder_optimizer, criterion)
print_loss_total = print_loss_total + loss
plot_loss_total = plot_loss_total + loss
if iter % print_every == 0:
print_loss_avg = print_loss_total / print_every
print_loss_total = 0
print('%s (%d %d%%) %.4f' % (timeSince(start, iter / n_iters),
iter, iter / n_iters * 100, print_loss_avg))
if iter % plot_every == 0:
plot_loss_avg = plot_loss_total / plot_every
plot_losses.append(plot_loss_avg)
plot_loss_total = 0
showPlot(plot_losses)
```
## Ploting results
```
import matplotlib.pyplot as plt
plt.switch_backend('agg')
import matplotlib.ticker as ticker
import numpy as np
def showPlot(points):
plt.figure(figsize=(14,7))
fig, ax = plt.subplots()
# this locator puts ticks at regular intervals
loc = ticker.MultipleLocator(base=0.2)
ax.yaxis.set_major_locator(loc)
plt.plot(points)
```
## Start Training
```
hidden_size = 256
encoder1 = EncoderRNN(input_lang.n_words, hidden_size).to(device)
decoder1 = AttenDecoder(hidden_size, output_lang.n_words).to(device)
trainIters(encoder1, decoder1, n_iters=100000, print_every=500, plot_every=10)
```
## Evaluation
Evaluation is mostly the same as training, but there are no targets so we simply feed the decoder’s predictions back to itself for each step. Every time it predicts a word we add it to the output string, and if it predicts the EOS token we stop there. We also store the decoder’s attention outputs for display later.
```
def evaluate(encoder, decoder, sentence, max_length=MAX_LENGTH):
with torch.no_grad():
input_tensor = tensorFromSentence(input_lang, sentence)
input_length = input_tensor.size()[0]
# Encoder
encoder_output, encoder_hidden = encoder1(input_tensor.unsqueeze(1))
encoder_output = encoder_output.squeeze(1)
encoder_outputs = torch.zeros(max_length, encoder_output.size(1)).to(device)
encoder_outputs[:encoder_output.size(0)] = encoder_output
# Decoder
decoder_input = torch.tensor([[SOS_token]], device=device) # SOS
decoder_hidden = encoder_hidden
decoded_words = []
decoder_attentions = torch.zeros(max_length, max_length) # 记录输入每个词时的attention weight
for di in range(max_length):
decoder_output, decoder_hidden, attn_weights = decoder(decoder_input, decoder_hidden, encoder_outputs)
decoder_attentions[di] = attn_weights.data
topv, topi = decoder_output.data.topk(1)
if topi.item() == EOS_token:
"""遇到终止符就停止
"""
decoded_words.append('<EOS>')
break
else:
"""把decode的word加入数组中
"""
decoded_words.append(output_lang.index2word[topi.item()])
# 下一个的输入是上一个的输出
decoder_input = topi.squeeze().detach()
return decoded_words,decoder_attentions[:di+1]
def evaluateRandomly(encoder, decoder, n=10):
for i in range(n):
pair = random.choice(pairs)
print('>', pair[0])
print('=', pair[1])
output_words,_ = evaluate(encoder, decoder, pair[0])
output_sentence = ' '.join(output_words)
print('<', output_sentence)
print('')
evaluateRandomly(encoder1, decoder1, n=10)
```
## Visualizing Attention
```
_, attentions = evaluate(encoder1, decoder1, "je suis trop froid .")
plt.matshow(attentions.cpu().numpy())
```
For a better viewing experience we will do the extra work of adding axes and labels:
```
# 更好的可视化
def showAttention(input_sentence, output_words, attentions):
# Set up figure with colorbar
fig = plt.figure()
ax = fig.add_subplot(111)
cax = ax.matshow(attentions.numpy(), cmap='bone')
fig.colorbar(cax)
# Set up axes
ax.set_xticklabels([''] + input_sentence.split(' ') +
['<EOS>'], rotation=90)
ax.set_yticklabels([''] + output_words)
# Show label at every tick
ax.xaxis.set_major_locator(ticker.MultipleLocator(1))
ax.yaxis.set_major_locator(ticker.MultipleLocator(1))
plt.show()
def evaluateAndShowAttention(input_sentence):
output_words, attentions = evaluate(encoder1, decoder1, input_sentence)
print('input =', input_sentence)
print('output =', ' '.join(output_words))
showAttention(input_sentence, output_words, attentions)
evaluateAndShowAttention("c est un jeune directeur plein de talent .")
```
| github_jupyter |
## Python File Operations
# Binary Files
```
with open("myfile.bin", "wb") as f:
f.write(b'\x30\x31\x09\x32\x20\x52\x43\x53\x0A\x51\xFE\x00\xFF') # notice b prefix!!
with open("myfile.bin", "r") as f:
lines=f.readlines()
f.seek(0)
text=f.read()
print(lines)
print(lines[0])
print(text)
```
ASCII Codes
https://en.wikipedia.org/wiki/ASCII
```
with open("myfile.bin", "rb") as f:
blines=f.readlines()
print(blines)
blines
```
## For more complicated binary writing and reading of binaries
### pickle standard library is recommended
https://docs.python.org/3/library/pickle.html
```
import pickle
with open("myfile.bin", "wb") as f:
myint = 42
mystring = "Hello, RCS!"
mylist = ["sun", "moon", "earth"]
mydict = { "name": "Val", "job": "Teacher" }
pickle.dump(myint, f)
pickle.dump(mystring, f)
pickle.dump(mylist, f)
pickle.dump(mydict, f)
with open("myfile.bin", "r", encoding=None) as f:
mf=f.read()
mf
```
### Not very helpful is it? Better to use pickle to retrieve the data and "unpickle" it
```
with open("myfile.bin", "rb") as f:
myint = pickle.load(f)
mystring = pickle.load(f)
mylist = pickle.load(f)
mydict = pickle.load(f)
myint,mystring
mylist
mydict
# Recipe for opening a pickled file with unknown number of variables
with open('myfile.bin', "rb") as f:
mylist = []
while True:
try:
mylist.append(pickle.load(f))
except EOFError:
print("End of file reached!")
break
print("Going to close file now")
len(mylist)
mylist[3]
# write a for loop printing all data types in mylist
for item in mylist:
print(type(item))
print(myint,mystring,mylist,mydict)
import os
print(os.getcwd()) # cwd - current working directory
mycwd = os.getcwd()
mycwd
%pwd
!dir
os.getlogin()
os.getcwd()
os.rename('numbers.txt', 'bignumbers.txt')
myfiles = os.listdir()
myfiles[:5]
myfiles
myfiles[4]
'.txt' in myfiles[4]
'.txt' in myfiles[2]
# How do we select only text files
# We can create a new list of only text files (with extension .txt)
mytextfiles = [file for file in myfiles if '.txt' in file]
mytextfiles
mytxtlist = []
for item in myfiles:
if '.txt' in item:
mytxtlist.append(item)
# we could do more stuff here not just make a list
mytxtlist
os
result = []
for file in mytextfiles:
with open(file) as f:
txt = f.read() # careful here, we might not want to read the full file
result.append(len(txt))
result
filesizes = []
for file in mytextfiles:
filesizes.append((file, os.path.getsize(file)))
filesizes
# list comprehension will be shorter
filesizes = [(file, os.path.getsize(file)) for file in mytextfiles]
filesizes
filedict = {}
for file in mytextfiles:
filedict[file] = os.path.getsize(file)
filedict
print('mytextfiles is a ', type(mytextfiles))
# one line dictionary comprehension
fdict = {f:os.path.getsize(f) for f in mytextfiles}
fdict
result
os.listdir('C:\\')
os.chdir("c:\\")
os.getcwd()
os.chdir(mycwd)
os.getcwd()
os.path.expanduser('~')
os.chdir(os.path.expanduser('~')+"\\Github\\RCS_Data_Analysis_Python_2019_July")
os.getcwd()
```
The os.path.join() function constructs a pathname out of one or more partial pathnames
**Don’t fuss with slashes; always use os.path.join() and let Python do the right thing.**
```
# OS neutral path join should work on all OSes
os.path.join(os.getcwd(), "README.md")
os.path.join("C:\\Users\\vsd\\Documents\\Github\\RCS_Python","README.md")
print(os.path.join(os.path.expanduser('~'), 'Github', 'RCS_Data_Analysis_Python_2019_July', 'README.md'))
print(os.path.join(os.getcwd(),'Documents', 'Github', 'RCS_Python', 'README.md'))
newpath=os.path.join(os.path.expanduser('~'), 'Github')
newpath
print(newpath) ## Aha pretty print!!
mypath=os.getcwd()
mypath
mysplit = os.path.split(mypath)
mysplit
mydir, myfname = os.path.split(mypath)
print(mydir,":",myfname)
```
The glob module finds all the pathnames matching a specified pattern according to the rules used by the Unix shell, although results are returned in arbitrary order.
```
glob.
# we can get a list of all files in current directory matching certaing wildcards
from glob import glob as gl
ifiles=gl('Python*.ipynb')
ifiles
ipyth=glob.glob('*Python*.ipynb')
ipyth
ifile2=glob.glob('*File*.*,*.md')
ifile2
# Find Matching Files recursively
from pathlib import Path
# We find all matching text files with path and split the path from the filename
for filename in Path('').glob('**/*.txt'):
print(os.path.split(filename))
```
### New in version 3.4.
For example, consider a directory containing the following files: 1.gif, 2.txt, card.gif and a subdirectory sub which contains only the file 3.txt. glob() will produce the following results. Notice how any leading components of the path are preserved.
```
glob.glob('./*V*.*')
! mkdir Text # ! is Jupyter command for running OS commands b
```
# File Operations directly from Python
https://docs.python.org/3/library/subprocess.html#module-subprocess
subprocess.run(args, *, stdin=None, input=None, stdout=None, stderr=None, shell=False, cwd=None, timeout=None, check=False, encoding=None, errors=None)
```
%pwd
import sys
sys.path
import subprocess
print(subprocess.run("calculator", shell=True, stdout=subprocess.PIPE))
import subprocess
print(subprocess.run("dir", shell=True, stdout=subprocess.PIPE)) ## Without pipe we will get no output
print(subprocess.run("C:\\Program Files (x86)\\Google\\Chrome\\Application\\chrome.exe"))
print(subprocess.run("chrome.exe"))
subprocess.run(["mkdir", "testdir"], shell=True, stdout=subprocess.PIPE)
import sys
! dir
%%writefile ./Text/Test.txt
Just a simple text file
Nothing special
! dir Text
glob.glob('**/*.txt', recursive=True) #We should also get subdirectory name
glob.glob('./*.md')
glob.glob('./?.md') # requires a single char only so we wont get a match for longer file names
meta = os.stat('README.md')
print(type(meta),meta) # os.stat returns a class containing file meta information
import time
time.localtime(meta.st_mtime) #mtime last modified tiem
## Homework 2 for file operations
# Process 'resources\\cleaned.txt',
# Generate a dictionary of words and their frequency - "Un" : 76
# Save this dictionary in a text file, each word and frequency in a new line
# Bonus for pickling the dictionary
```
| github_jupyter |
```
import os
import pandas as pd
import matplotlib.pyplot as plt
from keras.applications.vgg16 import VGG16
from keras.models import Model
from keras.callbacks import ModelCheckpoint,EarlyStopping
from keras.preprocessing.image import ImageDataGenerator
from keras.utils import np_utils
from keras.models import Sequential
from keras.callbacks import EarlyStopping, History, ModelCheckpoint
from keras.layers.core import Flatten, Dense, Dropout, Reshape, Lambda
from keras.layers.normalization import BatchNormalization
from sklearn.preprocessing import LabelEncoder
from keras.utils.np_utils import to_categorical
from sklearn.metrics import log_loss
from sklearn.model_selection import train_test_split
import numpy as np
train_features = np.load('train_preprocesed.npy')
valid_features = np.load('valid_preprocessed.npy')
train_dir = "new_train/"
valid_dir = "new_valid/"
classes = os.listdir(train_dir)
# Get the labels
train_labels = []
for c in classes:
l = [c]*len(os.listdir(train_dir+c+'/'))
train_labels.extend(l)
len(train_labels)
valid_labels = []
for c in classes:
l = [c]*len(os.listdir(valid_dir+c+'/'))
valid_labels.extend(l)
onehot_train = to_categorical(LabelEncoder().fit_transform(train_labels))
onehot_valid = to_categorical(LabelEncoder().fit_transform(valid_labels))
vgg16_base = VGG16(include_top=False, weights='imagenet',
input_tensor=None, input_shape=(150, 150,3))
# Note that the preprocessing of InceptionV3 is:
# (x / 255 - 0.5) x 2
print('Adding new layers...')
output = vgg16_base.get_layer(index = -1).output
output = Flatten()(output)
# let's add a fully-connected layer
output = Dense(4096,activation = "relu")(output)
output = BatchNormalization()(output)
output = Dropout(0.5)(output)
output = Dense(512,activation = "relu")(output)
output = BatchNormalization()(output)
output = Dropout(0.5)(output)
# and a logistic layer -- let's say we have 200 classes
output = Dense(8, activation='softmax')(output)
vgg16_model = Model(vgg16_base.input, output)
#InceptionV3_model.summary()
for layer in vgg16_model.layers[:19]:
layer.trainable = False
vgg16_model.compile(optimizer="adam",loss="categorical_crossentropy",metrics =["accuracy"])
vgg16_model.summary()
train_datagen = ImageDataGenerator(
shear_range=0.1,
zoom_range=0.1,
rotation_range=10.,
width_shift_range=0.1,
height_shift_range=0.1,
horizontal_flip=True)
val_datagen = ImageDataGenerator()
callbacks = EarlyStopping(monitor='val_loss', patience=2, verbose=1, mode='auto')
# autosave best Model
best_model_file = "./data_augmented_weights.h5"
best_model = ModelCheckpoint(best_model_file, monitor='val_acc', verbose = 1, save_best_only = True)
history = vgg16_model.fit_generator(train_datagen.flow(train_features, onehot_train, batch_size=10), nb_epoch=5,
samples_per_epoch = 3019,
validation_data=val_datagen.flow(valid_features,onehot_valid,batch_size=10,shuffle=False),
nb_val_samples=758,callbacks = [callbacks,best_model])
#model.load_weights("batch_normalized_weights.h5")
vgg16_model.load_weights("data_augmented_weights.h5")
# summarize history for accuracy
plt.figure(figsize=(15, 5))
plt.subplot(1, 2, 1)
plt.plot(history.history['acc']); plt.plot(history.history['val_acc']);
plt.title('model accuracy'); plt.ylabel('accuracy');
plt.xlabel('epoch'); plt.legend(['train', 'valid'], loc='upper left');
# summarize history for loss
plt.subplot(1, 2, 2)
plt.plot(history.history['loss']); plt.plot(history.history['val_loss']);
plt.title('model loss'); plt.ylabel('loss');
plt.xlabel('epoch'); plt.legend(['train', 'valid'], loc='upper left');
plt.show()
test_features = np.load("test_preprocessed.npy")
test_preds = vgg16_model.predict(test_features, batch_size=5, verbose=1)
test_preds[0:5]
submission1 = pd.DataFrame(test_preds, columns= os.listdir(train_dir))
test_files = os.listdir("test_stg1/test_stg1/")
submission1.insert(0, 'image', test_files)
submission1.head()
clipped_preds = np.clip(test_preds,(1-0.82)/7,0.82)
submission2 = pd.DataFrame(clipped_preds, columns= os.listdir("train/train/"))
submission2.insert(0, 'image', test_files)
submission2.head()
submission2.to_csv("data_augmented_batch_normalized.csv",index = False)
valid_preds = vgg16_model.predict_classes(valid_features, batch_size=5, verbose=1)
```
| github_jupyter |
# TPR : From symbols to tensors
__(Cho, Goldrick & Smolensky 2016)__
## Data
This notebook tries to illustrate how to use Tensor Product Representation (TPR) to represent discrete or gradient blend structures. The concrete examples apply TPR to root allomorphy. In Sanskrit and Greek, for instance, we have a phenomenon known as Grassmann's law : no two aspirated stops are allowed in the same root. If a root happens to have two aspirates, we typically observe alternations:
The root $bud^h$ (*a.o.* 'awake', 'know') can surface either as $b^hud-$ or $bud^h$ (e.g. 3sg pres. _bodhati_, 3sg future _bhotsyati_).
The problem has attracted the attention of many scholars due to impossibility to model this phenomenon through phonological rules, which would lead to ordering paradoxes and a number of other open issues related to this phenomenon.
GSC offers a solution if we take into account the possibility that both the aspirated and plain variants of the consonants were underlying represented and the UR as whole is a gradient blend of the two segments/or possibly features.
$$b^hud-/bud^h = [\alpha \cdot b^h + \beta \cdot b] \, u \, [\gamma \cdot d + \delta \cdot d^h]$$
```
# Imports
import torch
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# Set seed for reproducibility
# torch.manual_seed(111)
```
## The TPR Representation
In matrix language the **Tensor Product Representation** of a structure $s$ can be expressed as:
$$T_s = F \times B \times R^T$$
the chain product of the filler matrix, times the binding matrix times the role matrix.
**Ex**:
Let's suppose we have the input *budh*. This can be decomposed into
$$b \otimes r_1 + u \otimes r_2 + dh \otimes r_3$$
Now suppose that our language also consists of the additional fillers "bh" and "d". The set of fillers could be:
["bh", "b", "u", "d", "dh", "_"]
(the last filler representing the empty filler (I've used it occasionally to pad shorter strings or to explicitely prefer simple over complex codas)
The binding matrix for *budh* would be:
```
fillers = ["bh", "b", "u", "d", "dh", "_"]
roles = ["pos1", "pos2", "pos3"]
# 3 cols : 3 positions for the roots (initial, middle, final) and 6 rows, one for each filler
budh = torch.tensor([[0., 0., 0.],
[1., 0., 0.],
[0., 1., 0.],
[0., 0., 0.],
[0., 0., 1.],
[0., 0., 0.]])
print(pd.DataFrame(budh.numpy(), index=fillers, columns=roles))
df = pd.DataFrame(budh.numpy(), index=fillers, columns=roles)
sns.heatmap(df, annot=True, cmap="Greens")
```
The role and the filler matrices can be built in different ways. The following examples were built using random components, chosen so that the column vectors build a set of linearly independent vectors and their pairwise dotproduct is 0 (maximally different).
```
R = torch.tensor([[ 0.4311, 0.8892, -0.1533],[-0.3264, 0.3121, 0.8922],[ 0.8412, -0.3346, 0.4248]])
F = torch.tensor([[ 0.2958, -0.3054, 0.0164, 0.2841, 0.4751, 0.7159],
[ 0.1613, -0.2552, 0.7667, 0.1514, 0.3030, -0.4542],
[ 0.5245, 0.4364, 0.4019, 0.0709, -0.5345, 0.2867],
[-0.3085, 0.6143, 0.2897, -0.4018, 0.4824, 0.2222],
[ 0.6668, 0.3079, -0.4022, -0.0627, 0.3952, -0.3724],
[-0.2674, 0.4232, -0.0681, 0.8520, 0.0888, -0.1045]])
print(f"Roles matrix:\n{R}\n\n")
print(f"Fillers matrix:\n{F}")
```
*budh* can be then represented as:
```
tpr_budh = F.matmul(budh).matmul(R.T)
tpr_budh
df = pd.DataFrame(tpr_budh.numpy(), index=fillers, columns=roles)
sns.heatmap(df, annot=True)
```
Notice that even though this representation is now distributed, we can always retrieve (unbind) the original fillers using matrix multiplication:
$$TPR \times (R^T)^{-1} = F$$
```
tpr_budh.matmul(torch.pinverse(R.T))
```
Compare this matrix with the Filler matrix:
```
F
```
The matrix we obtained by multiplying the TP representation with the inverse of the role matrix is a (6,3) matrix, where each column represent a role (in this example the position of the filler in the string) and the component of each column are exactly those of the second, third and fifth column of the filler matrix, corresponding resp. to the fillers "b", "u" and "dh".
## Blend Representations
The same procedure can be applied if the Binding matrix doesn't contain just 0s and 1s but some values between 0s and 1s, corresponding to partial activations of specific fillers.
So for instance the blend $(0.8 \cdot b + 0.7 \cdot bh)udh$ can be represented with the following binding matrix:
```
bbhudh = torch.tensor([[.7, 0, 0], [.8, 0, 0], [0,1,0],[0,0,0],[0,0,1],[0,0,0]])
bbhudh_df = pd.DataFrame(bbhudh.numpy(), index=fillers, columns=roles)
print(bbhudh_df)
sns.heatmap(bbhudh_df, annot=True, cmap="GnBu")
```
It's representation in the neural space will then be:
```
tpr_bbhudh = F @ bbhudh @ R.T
tpr_bbhudh
blend_df = pd.DataFrame(tpr_bbhudh.numpy(), index=fillers, columns=roles)
sns.heatmap(blend_df, annot=True, cmap="GnBu")
```
The same holds here as before. We can go back to the local representation using the matrix multiplication.
```
bbhudh_fillers = tpr_bbhudh.matmul(torch.pinverse(R.T))
print(f"bbhudh_fillers :\n{bbhudh_fillers}\n\n")
print(f"Filler Matrix :\n{F}\n")
```
Notice that nothing changed as for the representation of the second and third position in the string:
```
assert torch.allclose(F[:,2],bbhudh_fillers[:,1])
assert torch.allclose(F[:,4],bbhudh_fillers[:,2])
```
But the first column in our representation is now a blend of the first and second column in the Matrix of fillers. How much of the first and second filler went into the representation of the first element of the blend representation is revealed using the dot product between those vectors
```
torch.dot(F[:,0],bbhudh_fillers[:,0])
torch.dot(F[:,1],bbhudh_fillers[:,0])
```
Notice also that the similarity with other fillers is close to 0, which guarantees that we can always unbind the representations.
```
torch.dot(F[:,2],bbhudh_fillers[:,0])
torch.dot(F[:,3],bbhudh_fillers[:,0])
torch.dot(F[:,4],bbhudh_fillers[:,0])
```
## Heat map visualization of blends and pure representations
```
sns.set_theme(style="darkgrid")
roles = ["pos1", "pos2", "pos3"]
fillers = ["bh", "b", "u", "d", "dh", "_"]
```
## Discrete structure
```
# Purely local representation
budh_heat = sns.heatmap(budh,
cmap="GnBu",
annot=True,
xticklabels=roles,
yticklabels=fillers)
# Distributed representation
sns.heatmap(tpr_budh.reshape((18,1)),
cmap="GnBu")
```
Each row in this distributed representation represent a binding (1 role, 1 filler). We had 6 possible fillers and 3 possible positions, hence we get 18 possible bindings (rows 0-17).
### Gradient structures
```
# Blend representation (discrete)
sns.heatmap(bbhudh,
cmap="GnBu",
annot=True,
xticklabels=roles,
yticklabels=fillers)
# Blend representation (distributed)
sns.heatmap(tpr_bbhudh.reshape((18,1)),
cmap="GnBu",
annot=True)
```
| github_jupyter |
Handling models in GPflow
--
*James Hensman November 2015, January 2016*,
*Artem Artemev December 2017*
One of the key ingredients in GPflow is the model class, which allows the user to carefully control parameters. This notebook shows how some of these parameter control features work, and how to build your own model with GPflow. First we'll look at
- How to view models and parameters
- How to set parameter values
- How to constrain parameters (e.g. variance > 0)
- How to fix model parameters
- How to apply priors to parameters
- How to optimize models
Then we'll show how to build a simple logistic regression model, demonstrating the ease of the parameter framework.
GPy users should feel right at home, but there are some small differences.
First, let's deal with the usual notebook boilerplate and make a simple GP regression model. See the Regression notebook for specifics of the model: we just want some parameters to play with.
```
import gpflow
import numpy as np
```
Create a very simple GPR model without building it in TensorFlow graph.
```
np.random.seed(1)
X = np.random.rand(20, 1)
Y = np.sin(12 * X) + 0.66 * np.cos(25 * X) + np.random.randn(20,1) * 0.01
with gpflow.defer_build():
m = gpflow.models.GPR(X, Y, kern=gpflow.kernels.Matern32(1) + gpflow.kernels.Linear(1))
```
### Viewing, getting and setting parameters
You can display the state of the model in a terminal with `print(m)`, and by simply returning it in a notebook:
```
m
```
This model has four parameters. The kernel is made of the sum of two parts: the first (counting from zero) is an RBF kernel that has a variance parameter and a lengthscale parameter; the second is a linear kernel that only has a variance parameter. There is also a parameter controlling the variance of the noise, as part of the likelihood.
All of the model variables have been initialized at one. Individual parameters can be accessed in the same way as they are displayed in the table: to see all the parameters that are part of the likelihood, do
```
m.likelihood
```
This gets more useful with more complex models!
To set the value of a parameter, just assign.
```
m.kern.kernels[0].lengthscales = 0.5
m.likelihood.variance = 0.01
m
```
### Constraints and trainable variables
GPflow helpfully creates an unconstrained representation of all the variables. Above, all the variables are constrained positive (see right hand table column), the unconstrained representation is given by $\alpha = \log(\exp(\theta)-1)$. `read_trainables()` returns the constrained values:
```
m.read_trainables()
```
Each parameter has an `unconstrained_tensor` attribute that allows accessing the unconstrained value as a tensorflow Tensor (though only after the model has been compiled). We can also check the unconstrained value as follows:
```
p = m.kern.kernels[0].lengthscales
p.transform.backward(p.value)
```
Constraints are handled by the `Transform` classes. You might prefer the constraint $\alpha = \log(\theta)$: this is easily done by changing the transform attribute on a parameter, with one simple condition - the model has not been compiled yet:
```
m.kern.kernels[0].lengthscales.transform = gpflow.transforms.Exp()
```
Though the lengthscale itself remains the same, the unconstrained lengthscale has changed:
```
p.transform.backward(p.value)
```
Another helpful feature is the ability to fix parameters. This is done by simply setting the `trainable` attribute to False: this is shown in the 'trainable' column of the representation, and the corresponding variable is removed from the free state.
```
m.kern.kernels[1].variance.trainable = False
m
m.read_trainables()
```
To unfix a parameter, just flip the boolean back and set the parameter to be trainable again.
```
m.kern.kernels[1].variance.trainable = True
m
```
### Priors
Priors are set just like transforms and trainability, using members of the `gpflow.priors` module. Let's set a Gamma prior on the RBF-variance.
```
m.kern.kernels[0].variance.prior = gpflow.priors.Gamma(2, 3)
m
```
### Optimization
Optimization is done by creating an instance of optimizer, in our case it is `gpflow.train.ScipyOptimizer`, which has optional arguments that are passed through to `scipy.optimize.minimize` (we minimize the negative log-likelihood) and calling `minimize` method of that optimizer with model as optimization target. Variables that have priors are MAP-estimated, i.e. we add the log prior to the log likelihood, otherwise using Maximum Likelihood.
```
m.compile()
opt = gpflow.train.ScipyOptimizer()
opt.minimize(m)
```
### Building new models
To build new models, you'll need to inherit from `gpflow.models.Model`. Parameters are instantiated with `gpflow.Param`. You may also be interested in `gpflow.params.Parameterized` which acts as a 'container' of `Param`s (e.g. kernels are Parameterized).
In this very simple demo, we'll implement linear multiclass classification. There will be two parameters: a weight matrix and a 'bias' (offset). The key thing to implement is the private `_build_likelihood` method, which should return a tensorflow scalar representing the (log) likelihood. By decorating the function with `@gpflow.params_as_tensors`, Param objects can be used inside `_build_likelihood`: they will appear as appropriate (constrained) tensors.
```
import tensorflow as tf
class LinearMulticlass(gpflow.models.Model):
def __init__(self, X, Y, name=None):
super().__init__(name=name) # always call the parent constructor
self.X = X.copy() # X is a numpy array of inputs
self.Y = Y.copy() # Y is a 1-of-k (one-hot) representation of the labels
self.num_data, self.input_dim = X.shape
_, self.num_classes = Y.shape
#make some parameters
self.W = gpflow.Param(np.random.randn(self.input_dim, self.num_classes))
self.b = gpflow.Param(np.random.randn(self.num_classes))
# ^^ You must make the parameters attributes of the class for
# them to be picked up by the model. i.e. this won't work:
#
# W = gpflow.Param(... <-- must be self.W
@gpflow.params_as_tensors
def _build_likelihood(self): # takes no arguments
p = tf.nn.softmax(tf.matmul(self.X, self.W) + self.b) # Param variables are used as tensorflow arrays.
return tf.reduce_sum(tf.log(p) * self.Y) # be sure to return a scalar
```
...and that's it. Let's build a really simple demo to show that it works.
```
np.random.seed(123)
X = np.vstack([np.random.randn(10,2) + [2,2],
np.random.randn(10,2) + [-2,2],
np.random.randn(10,2) + [2,-2]])
Y = np.repeat(np.eye(3), 10, 0)
from matplotlib import pyplot as plt
plt.style.use('ggplot')
%matplotlib inline
import matplotlib
matplotlib.rcParams['figure.figsize'] = (12,6)
plt.scatter(X[:,0], X[:,1], 100, np.argmax(Y, 1), lw=2, cmap=plt.cm.viridis);
m = LinearMulticlass(X, Y)
m
opt = gpflow.train.ScipyOptimizer()
opt.minimize(m)
m
xx, yy = np.mgrid[-4:4:200j, -4:4:200j]
X_test = np.vstack([xx.flatten(), yy.flatten()]).T
f_test = np.dot(X_test, m.W.read_value()) + m.b.read_value()
p_test = np.exp(f_test)
p_test /= p_test.sum(1)[:,None]
plt.figure(figsize=(12, 6))
for i in range(3):
plt.contour(xx, yy, p_test[:,i].reshape(200,200), [0.5], colors='k', linewidths=1)
plt.scatter(X[:,0], X[:,1], 100, np.argmax(Y, 1), lw=2, cmap=plt.cm.viridis);
```
That concludes the new model example and this notebook. You might want to convince yourself that the `LinearMulticlass` model and its parameters have all the functionality demonstrated above. You could also add some priors and run Hamiltonian Monte Carlo using the HMC optimizer `gpflow.train.HMC` and its `sample` method. See the sparse_MCMC notebook for details of running the sampler.
| github_jupyter |
```
import scanpy as sc
import squidpy as sq
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
from squidpy.pl._utils import save_fig
from time import process_time
sc.logging.print_header()
sc.set_figure_params(facecolor="white", figsize=(8, 8))
sc.settings.verbosity = 1
sc.settings.dpi = 300
sq.__version__
sc.settings.figdir = "./figures"
%load_ext autoreload
%autoreload 2
%load_ext lab_black
adata_visium = sq.datasets.visium_fluo_adata()
adata_slideseq = sq.datasets.slideseqv2()
adata_seqfish = sq.datasets.seqfish()
adata_fouri = sq.datasets.four_i()
adata_imc = sq.datasets.imc()
adata_merfish = sq.datasets.merfish()
adata_mibitof = sq.datasets.mibitof()
adata_mibitof = adata_mibitof[adata_mibitof.obs.batch == "0"].copy()
adata_merfish = adata_merfish[adata_merfish.obs.batch == "0"].copy()
adata_seqfish.obs["cluster"] = adata_seqfish.obs.celltype_mapped_refined
adata_imc.obs["cluster"] = adata_imc.obs["cell type"]
adata_slideseq.obs["cluster"] = adata_slideseq.obs["cluster"]
adata_merfish.obs["cluster"] = adata_merfish.obs["Cell_class"]
adata_mibitof.obs["cluster"] = adata_mibitof.obs["Cluster"]
dic_list = []
for i in np.arange(5):
for adata, data_id in zip(
[
adata_visium,
adata_seqfish,
adata_fouri,
adata_imc,
adata_merfish,
adata_mibitof,
adata_slideseq,
],
["visium", "seqfish", "4i", "imc", "merfish", "mibitof", "slideseq"],
):
cluster_id = "cluster"
start_t = process_time()
sq.gr.spatial_neighbors(adata, coord_type="generic")
duration_graph = process_time() - start_t
start_t = process_time()
sq.gr.nhood_enrichment(adata, cluster_key=cluster_id)
duration_nhood = process_time() - start_t
dic_list.append(
{
"dataset": data_id,
"time_nhood": duration_nhood,
"time_graph": duration_graph,
"n_obs": adata.shape[0],
"n_cluster": adata.obs[cluster_id].cat.categories.shape[0],
"idx": i,
}
)
df = pd.DataFrame(dic_list)
df = df.groupby(["dataset", "n_obs"]).mean()
df.reset_index(drop=False, inplace=True)
df = df[["dataset", "time_nhood", "time_graph", "n_obs"]].copy()
df.rename(
columns={"time_nhood": "nhood_enrichment", "time_graph": "graph"}, inplace=True
)
df = df.melt(
id_vars="dataset",
value_vars=["nhood_enrichment", "graph"],
value_name="mean",
var_name="method",
)
df["tool"] = "squidpy"
df
obs_df = pd.DataFrame(dic_list)
obs_df = obs_df[["n_obs", "n_cluster", "dataset"]].copy()
obs_df.drop_duplicates(inplace=True)
obs_df.reset_index(inplace=True, drop=True)
giotto = pd.read_csv("./benchmark_giotto_results.csv", index_col=0)
giotto.rename(columns={"expr": "method"}, inplace=True)
giotto = giotto[["method", "mean", "dataset"]].copy()
giotto = giotto[giotto.method != "net_delaunay"].copy()
giotto.replace({"net_knn": "graph", "cellproxy": "nhood_enrichment"}, inplace=True)
giotto["tool"] = "giotto"
giotto
giotto.dataset.replace("slideseqv2", "slideseq", inplace=True)
giotto
final_df = pd.concat([df, giotto], axis=0)
final_df = final_df.merge(obs_df, on="dataset")
final_df["log_mean"] = np.log10(1 + final_df["mean"].values)
final_df["log_n_obs"] = np.log10(1 + final_df["n_obs"].values)
final_df
fig, ax = plt.subplots(tight_layout=True, dpi=180, figsize=(5, 3))
data = final_df[final_df.method == "graph"]
sns.scatterplot(
data=data, x="log_n_obs", y="log_mean", hue="dataset", style="tool", s=100, ax=ax
)
plt.xticks(data.log_n_obs.values, data.n_obs.values, rotation=90)
plt.yticks(
np.round(data.log_mean.values[[0, 1]], 2),
np.round(data["mean"].values[[0, 1]], 2),
)
ax.set_ylabel("runtime (s)")
ax.set_xlabel("#observations")
ax.legend(loc="center left", bbox_to_anchor=(1, 0.5))
_ = ax.set_title("comparisons of runtimes")
fig, ax = plt.subplots(tight_layout=True, dpi=180, figsize=(5, 3))
data = final_df[final_df.method == "nhood_enrichment"]
sns.scatterplot(
data=data, x="log_n_obs", y="log_mean", hue="dataset", style="tool", s=100, ax=ax
)
plt.xticks(data.log_n_obs.values, data.n_obs.values, rotation=90)
plt.yticks(
np.round(data.log_mean.values[[0, 1]], 2),
np.round(data["mean"].values[[0, 1]], 2),
)
ax.set_ylabel("runtime (s)")
ax.set_xlabel("#observations")
ax.legend(loc="center left", bbox_to_anchor=(1, 0.5))
_ = ax.set_title("comparisons of runtimes")
fig, ax = plt.subplots(tight_layout=True, dpi=180, figsize=(5, 3))
data = final_df[final_df.method == "nhood_enrichment"]
sns.scatterplot(
data=data, x="log_n_obs", y="log_mean", hue="dataset", s=100, style="tool", ax=ax
)
plt.xticks(data.log_n_obs.values, data.n_obs.values, rotation=90)
ax.set_ylabel(r"runtime $\log_{10}(s+1)$")
ax.set_xlabel("#observations")
ax.legend(loc="center left", bbox_to_anchor=(1, 0.5))
_ = ax.set_title("comparisons of runtimes")
```
| github_jupyter |
<a href="https://colab.research.google.com/github/kartikgill/The-GAN-Book/blob/main/Skill-01/Pixel-CNN-for-MNIST.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Importing useful libraries
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
%matplotlib inline
import tensorflow
print (tensorflow.__version__)
```
# Load and show the Dataset
```
from tensorflow.keras.datasets import mnist
(trainX, trainy), (testX, testy) = mnist.load_data()
print('Training data shapes: X=%s, y=%s' % (trainX.shape, trainy.shape))
print('Testing data shapes: X=%s, y=%s' % (testX.shape, testy.shape))
for k in range(9):
plt.figure(figsize=(9,6))
for j in range(9):
i = np.random.randint(0, 10000)
plt.subplot(990 + 1 + j)
plt.imshow(trainX[i], cmap='gray_r')
plt.axis('off')
#plt.title(trainy[i])
plt.show()
```
# Prepare data for the model
```
trainX = np.where(trainX < (0.33 * 256), 0, 1)
train_data = trainX.astype(np.float32)
testX = np.where(testX < (0.33 * 256), 0, 1)
test_data = testX.astype(np.float32)
train_data = np.reshape(train_data, (60000, 28, 28, 1))
test_data = np.reshape(test_data, (10000, 28, 28, 1))
print (train_data.shape, test_data.shape)
```
# Define Masked CNN layers for Pixel CNN
### Following code is inspired from : https://keras.io/examples/generative/pixelcnn/
```
# The first layer is the PixelCNN layer. This layer simply
# builds on the 2D convolutional layer, but includes masking.
import tensorflow
class PixelConvLayer(tensorflow.keras.layers.Layer):
def __init__(self, mask_type, **kwargs):
super(PixelConvLayer, self).__init__()
self.mask_type = mask_type
self.conv = tensorflow.keras.layers.Conv2D(**kwargs)
def build(self, input_shape):
# Build the conv2d layer to initialize kernel variables
self.conv.build(input_shape)
# Use the initialized kernel to create the mask
kernel_shape = self.conv.kernel.get_shape()
self.mask = np.zeros(shape=kernel_shape)
self.mask[: kernel_shape[0] // 2, ...] = 1.0
self.mask[kernel_shape[0] // 2, : kernel_shape[1] // 2, ...] = 1.0
if self.mask_type == "B":
self.mask[kernel_shape[0] // 2, kernel_shape[1] // 2, ...] = 1.0
def call(self, inputs):
self.conv.kernel.assign(self.conv.kernel * self.mask)
return self.conv(inputs)
# Next, we build our residual block layer.
# This is just a normal residual block, but based on the PixelConvLayer.
class ResidualBlock(tensorflow.keras.layers.Layer):
def __init__(self, filters, **kwargs):
super(ResidualBlock, self).__init__(**kwargs)
self.conv1 = tensorflow.keras.layers.Conv2D(
filters=filters, kernel_size=1, activation="relu"
)
self.pixel_conv = PixelConvLayer(
mask_type="B",
filters=filters // 2,
kernel_size=3,
activation="relu",
padding="same",
)
self.conv2 = tensorflow.keras.layers.Conv2D(
filters=filters, kernel_size=1, activation="relu"
)
def call(self, inputs):
x = self.conv1(inputs)
x = self.pixel_conv(x)
x = self.conv2(x)
return tensorflow.keras.layers.add([inputs, x])
```
# Define Pixel-CNN
```
inputs = tensorflow.keras.Input(shape=(28,28,1))
x = PixelConvLayer(
mask_type="A", filters=128, kernel_size=7, activation="relu", padding="same"
)(inputs)
for _ in range(5):
x = ResidualBlock(filters=128)(x)
for _ in range(2):
x = PixelConvLayer(
mask_type="B",
filters=128,
kernel_size=1,
strides=1,
activation="relu",
padding="valid",
)(x)
out = tensorflow.keras.layers.Conv2D(
filters=1, kernel_size=1, strides=1, activation="sigmoid", padding="valid"
)(x)
pixel_cnn = tensorflow.keras.Model(inputs, out)
pixel_cnn.summary()
```
# Compiling PixelCNN
```
adam = tensorflow.keras.optimizers.Adam(learning_rate=0.0005)
pixel_cnn.compile(optimizer=adam, loss="binary_crossentropy")
```
# Training Pixel-CNN
```
pixel_cnn.fit(
x=train_data, y=train_data, batch_size=128, epochs=50, validation_data=(test_data, test_data), verbose=1
)
```
# Display Results 81 images
```
from IPython.display import Image, display
from tqdm import tqdm_notebook
# Create an empty array of pixels.
batch = 81
pixels = np.zeros(shape=(batch,) + (pixel_cnn.input_shape)[1:])
batch, rows, cols, channels = pixels.shape
# Iterate over the pixels because generation has to be done sequentially pixel by pixel.
for row in tqdm_notebook(range(rows)):
for col in range(cols):
for channel in range(channels):
# Feed the whole array and retrieving the pixel value probabilities for the next
# pixel.
probs = pixel_cnn.predict(pixels)[:, row, col, channel]
# Use the probabilities to pick pixel values and append the values to the image
# frame.
pixels[:, row, col, channel] = tensorflow.math.ceil(
probs - tensorflow.random.uniform(probs.shape)
)
counter = 0
for i in range(9):
plt.figure(figsize=(9,6))
for j in range(9):
plt.subplot(990 + 1 + j)
plt.imshow(pixels[counter,:,:,0], cmap='gray_r')
counter += 1
plt.axis('off')
plt.show()
```
| github_jupyter |
```
import os
import sys
import glob
import itertools
import random
from IPython.display import Image
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.mlab as mlab
from matplotlib.colors import ListedColormap
from scipy.stats import multivariate_normal
import numpy as np
import pandas as pd
from scipy.stats import beta
random.seed(1234)
%matplotlib inline
```
## Mondrian Processes
### Various Functions for Mondrian Processes
Sampling...
```
### SAMPLE MONDRIAN PROCESS ###
def draw_Mondrian(theta_space, budget=5):
return draw_Mondrian_at_t(theta_space, 0, budget)
def draw_Mondrian_at_t(theta_space, t, budget):
dists = theta_space[:,1] - theta_space[:,0]
lin_dim = np.sum(dists)
T = np.random.exponential(scale=1./lin_dim)
if t+T > budget:
return (theta_space, None, None)
d = np.argmax(np.random.multinomial(n=1, pvals=dists/lin_dim))
x = np.random.uniform(low=theta_space[d,0], high=theta_space[d,1])
theta_left = np.copy(theta_space)
theta_left[d][1] = x
M_left = draw_Mondrian_at_t(theta_left, t+T, budget)
theta_right = np.copy(theta_space)
theta_right[d][0] = x
M_right = draw_Mondrian_at_t(theta_right, t+T, budget)
return (theta_space, M_left, M_right)
def comp_log_p_sample(theta_space, data):
if theta_space[1] == None and theta_space[2] == None:
if data.shape[0] == 0:
return 0
else:
mu = np.mean(data, axis = 0)
residual = data - mu
cov = np.dot(residual.T , residual) / data.shape[0] + np.identity(data.shape[1])*0.001
return np.log(multivariate_normal.pdf(data, mean=mu, cov=cov)).sum()
# find the dimension and location of first cut
root_rec = theta_space[0]
left_rec = theta_space[1][0]
for _ in range(root_rec.shape[0]):
if root_rec[_,1] != left_rec[_,1]:
break
dim, pos = _, left_rec[_,1]
idx_left = data[:,dim] < pos
idx_right = data[:,dim] >= pos
log_len_left = np.log(pos - root_rec[dim,0])
log_len_right = np.log(root_rec[dim,1] - pos)
return comp_log_p_sample(theta_space[1], data[idx_left]) + comp_log_p_sample(theta_space[2], data[idx_right])
```
Visualization...
```
### VISUALIZE 2D MONDRIAN PROCESS ###
def print_partitions(p, trans_level=1., color='k'):
if not p[1] and not p[2]:
plt.plot([p[0][0,0], p[0][0,0]], [p[0][1,0], p[0][1,1]], color+'-', linewidth=5, alpha=trans_level)
plt.plot([p[0][0,1], p[0][0,1]], [p[0][1,0], p[0][1,1]], color+'-', linewidth=5, alpha=trans_level)
plt.plot([p[0][0,0], p[0][0,1]], [p[0][1,0], p[0][1,0]], color+'-', linewidth=5, alpha=trans_level)
plt.plot([p[0][0,0], p[0][0,1]], [p[0][1,1], p[0][1,1]], color+'-', linewidth=5, alpha=trans_level)
else:
print_partitions(p[1], trans_level, color)
print_partitions(p[2], trans_level, color)
### VISUALIZE 2D POSTERIOR WITH DATA###
def print_posterior(data, samples, trans_level=.05, color='k'):
plt.figure()
plt.scatter(data[:,0], data[:,1], c='k', edgecolors='k', s=5, alpha=.5)
#print all samples
for sample in samples:
print_partitions(sample, trans_level, color)
def print_tree_at_leaf(mp_tree, table):
if mp_tree[1] == None and mp_tree[2] == None:
print table.shape
return 1
# find the dimension and location of first cut
root_rec = mp_tree[0]
left_rec = mp_tree[1][0]
for _ in range(root_rec.shape[0]):
if root_rec[_,1] != left_rec[_,1]:
break
d, pos = _, left_rec[_,1]
cut_type = ' '.join([str(int(x)) for x in sorted(set(table[table.columns[d]]))])
if cut_type in {"-1 0 1", '-1 1'}:
idx_table_left = table[table.columns[d]] != 1
table_left = table.loc[idx_table_left]
idx_table_right = table[table.columns[d]] != -1
table_right = table.loc[idx_table_right]
if cut_type == '-1 0':
idx_table_left = table[table.columns[d]] == -1
table_left = table.loc[idx_table_left]
idx_table_right = table[table.columns[d]] == 0
table_right = table.loc[idx_table_right]
if cut_type == '0 1':
idx_table_left = table[table.columns[d]] == 0
table_left = table.loc[idx_table_left]
idx_table_right = table[table.columns[d]] == 1
table_right = table.loc[idx_table_right]
return print_tree_at_leaf(mp_tree[1], table_left) + print_tree_at_leaf(mp_tree[2], table_right)
```
## Mondrian Process Generative Model
We apply Mondrian Processes (MPs) to flow cytometry data, using the prior information in the table above to guide the axis-aligned cuts. Instead of uniformly, we draw the cut proportion from $w \sim \text{Beta}(a_{0}, b_{0})$.
Now let's re-implement the MP sampling function, accounting for the prior information...
```
### SAMPLE MONDRIAN PROCESS WITH PRIOR INFORMATION ###
def draw_informed_Mondrian(theta_space, table, budget=5):
# INFORMATIVE PRIORS
upper_cut = (5., 2.)
lower_cut = (2., 5.)
middle_cut = (5., 5.)
neutral_cut = (2., 2.)
priors_dict = { '-1':lower_cut, '0':neutral_cut, '1':upper_cut,
'-1 0':lower_cut, '-1 1':middle_cut, '0 1':upper_cut,
'-1 0 1': middle_cut, '': neutral_cut
}
cut_history = [1] * theta_space.shape[0]
return draw_informed_Mondrian_at_t(theta_space, table, priors_dict, cut_history)
def draw_informed_Mondrian_at_t(theta_space, table, priors_dict, cut_history):
if sum(cut_history) == 0 or table.shape[0] == 1:
return (theta_space, None, None)
types_str = [' '.join([str(int(x)) for x in sorted(set(table[table.columns[d]]))])
for d in range(table.shape[1])]
if set([types_str[d] for d in range(table.shape[1]) if cut_history[d] == 1]).issubset({'0','1','-1'}):
return (theta_space, None, None)
low, medium, high, very_high = 0, 1, 100, 1000
priority_dict = {'-1': low , '0': low, '1': low,
'-1 0': medium, '0 1': medium,
'-1 0 1': high, '-1 1':very_high
}
types = np.array([priority_dict[_] for _ in types_str])
dists = (theta_space[:,1] - theta_space[:,0])* types
lin_dim = np.sum(dists)
# draw dimension to cut
dim_probs = ((dists/lin_dim) * np.array(cut_history))
dim_probs /= np.sum(dim_probs)
d = np.argmax(np.random.multinomial(n=1, pvals=dim_probs))
cut_history[d] = 0
prior_type_str = ' '.join([str(int(x)) for x in sorted(set(table[table.columns[d]]))])
prior_params = priors_dict[prior_type_str]
# make scaled cut
x = (theta_space[d,1] - theta_space[d,0]) * np.random.beta(prior_params[0], prior_params[1]) + theta_space[d,0]
cut_type = types_str[d]
if cut_type in {"-1 0 1", '-1 1'}:
idx_table_left = table[table.columns[d]] != 1
table_left = table.loc[idx_table_left]
idx_table_right = table[table.columns[d]] != -1
table_right = table.loc[idx_table_right]
if cut_type == '-1 0':
idx_table_left = table[table.columns[d]] == -1
table_left = table.loc[idx_table_left]
idx_table_right = table[table.columns[d]] == 0
table_right = table.loc[idx_table_right]
if cut_type == '0 1':
idx_table_left = table[table.columns[d]] == 0
table_left = table.loc[idx_table_left]
idx_table_right = table[table.columns[d]] == 1
table_right = table.loc[idx_table_right]
# make lower partition
theta_left = np.copy(theta_space)
theta_left[d][1] = x
M_left = draw_informed_Mondrian_at_t(theta_left, table_left, priors_dict, list(cut_history))
# make upper partition
theta_right = np.copy(theta_space)
theta_right[d][0] = x
M_right = draw_informed_Mondrian_at_t(theta_right, table_right, priors_dict,list(cut_history))
return (theta_space, M_left, M_right)
def Mondrian_Gaussian_perturbation(theta_space, old_sample, stepsize):
"""
Input:
theta_space: a rectangle
old_sample: partioned theta_space of a mondrian process
stepsize: gaussian std
"""
if old_sample[1] == None and old_sample[2] == None:
return (theta_space, None, None)
# find the dimension and location of first cut in the old_sample
for _ in range(old_sample[0].shape[0]):
if old_sample[0][_,1] > old_sample[1][0][_,1]:
break
dim, pos = _, old_sample[1][0][_,1]
# propose position of new cut
good_propose = False
while good_propose == False:
new_pos = pos + np.random.normal(0,(old_sample[0][dim,1] - old_sample[0][dim,0])*stepsize,1)[0]
if new_pos < theta_space[dim,1] and new_pos > theta_space[dim,0]:
good_propose = True
theta_left = np.copy(theta_space)
theta_left[dim,1] = new_pos
theta_right = np.copy(theta_space)
theta_right[dim,0] = new_pos
new_M_left= Mondrian_Gaussian_perturbation(theta_left, old_sample[1], stepsize)
new_M_right = Mondrian_Gaussian_perturbation(theta_right, old_sample[2], stepsize)
return (theta_space, new_M_left, new_M_right)
def comp_log_p_prior(theta_space, table, cut_history):
"""
This function returns prior probability of a Mondrian process theta_space
"""
if theta_space[1] == None and theta_space[2] == None:
return 0
log_prior = 0
# INFORMATIVE PRIORS
upper_cut = (5., 2.)
lower_cut = (2., 5.)
middle_cut = (5., 5.)
neutral_cut = (2., 2.)
priors_dict = { '-1':lower_cut, '0':neutral_cut, '1':upper_cut,
'-1 0':lower_cut, '-1 1':middle_cut, '0 1':upper_cut,
'-1 0 1': middle_cut, '': neutral_cut
}
# find the dimension and location of first cut
root_rec = theta_space[0]
left_rec = theta_space[1][0]
for _ in range(root_rec.shape[0]):
if root_rec[_,1] != left_rec[_,1]:
break
dim = _
beta_pos = (left_rec[_,1] - left_rec[dim,0])/(root_rec[dim,1] - root_rec[dim, 0])
prior_params = priors_dict[' '.join([str(int(x)) \
for x in sorted(set(table[table.columns[dim]]))])]
# compute the log likelihood of the first cut
types_str = [' '.join([str(int(x)) for x in sorted(set(table[table.columns[d]]))])
for d in range(table.shape[1])]
low_priority, medium_priority, high_priority, very_high_priority = 0, 1, 100, 1000
priority_dict = {'-1': low_priority , '0': low_priority, '1': low_priority,
'-1 0': medium_priority, '0 1': medium_priority,
'-1 0 1': high_priority, '-1 1':very_high_priority
}
types = np.array([priority_dict[_] for _ in types_str])
dists = (root_rec[:,1] - root_rec[:,0])* types
lin_dim = np.sum(dists)
# probability of dim
dim_probs = ((dists/lin_dim) * np.array(cut_history))
dim_probs /= np.sum(dim_probs)
log_prior += np.log(dim_probs[dim])
# probability of pos
log_prior += np.log(beta.pdf(beta_pos, prior_params[0], prior_params[1]))
# split the table
cut_history[dim] = 0
cut_type = types_str[dim]
if cut_type in {"-1 0 1", '-1 1'}:
idx_table_left = table[table.columns[dim]] != 1
table_left = table.loc[idx_table_left]
idx_table_right = table[table.columns[dim]] != -1
table_right = table.loc[idx_table_right]
if cut_type == '-1 0':
idx_table_left = table[table.columns[dim]] == -1
table_left = table.loc[idx_table_left]
idx_table_right = table[table.columns[dim]] == 0
table_right = table.loc[idx_table_right]
if cut_type == '0 1':
idx_table_left = table[table.columns[dim]] == 0
table_left = table.loc[idx_table_left]
idx_table_right = table[table.columns[dim]] == 1
table_right = table.loc[idx_table_right]
return log_prior + comp_log_p_prior(theta_space[1], table_left, list(cut_history)) \
+ comp_log_p_prior(theta_space[2], table_right, list(cut_history))
```
# Classification
```
def classify_cells(data, mp_tree, table, cell_type_name2idx):
Y = np.array([1]*data.shape[0])
if data.shape[0] == 0:
return Y
if mp_tree[1] == None and mp_tree[2] == None:
if table.shape[0] > 1:
# print "more than one clusters, number of data points:", data.shape[0]
labels = [cell_type_name2idx[table.index[_]] for _ in range(table.shape[0])]
return np.array(np.random.choice(labels, data.shape[0],replace = True))
else:
return Y * cell_type_name2idx[table.index[0]]
# find the dimension and location of first cut
root_rec = mp_tree[0]
left_rec = mp_tree[1][0]
for _ in range(root_rec.shape[0]):
if root_rec[_,1] != left_rec[_,1]:
break
dim, pos = _, left_rec[_,1]
# find labels that match dim info from table
idx_table_left = table[table.columns[dim]] != 1
table_left = table.loc[idx_table_left]
idx_table_right = table[table.columns[dim]] != -1
table_right = table.loc[idx_table_right]
# find data INDICIES that go high / low on cut position in dimension dim
idx_left = data[:,dim] < pos
idx_right = data[:,dim] >= pos
Y[idx_left] = classify_cells(data[idx_left],mp_tree[1],table_left, cell_type_name2idx)
Y[idx_right] = classify_cells(data[idx_right],mp_tree[2],table_right, cell_type_name2idx)
return Y
```
## Flow Cytometry Data
Load BMMC dataset from [ACDC paper](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5447237/pdf/btx054.pdf)...
```
# load BMMC data and table
##### X: np.array, flow cytometry data, arcsin transformed
##### T: table of expert knowledge
np.random.seed(1234)
PATH = '/home/disij/projects/acdc/data/'
### LOAD DATA ###
path = PATH + 'BMMC_benchmark/'
df = pd.read_csv( path + 'BMMC_benchmark.csv.gz', sep=',', header = 0, compression = 'gzip', engine='python')
table = pd.read_csv(path + 'BMMC_table.csv', sep=',', header=0, index_col=0)
print table.shape
### PROCESS: discard ungated events ###
channels = ['CD45','CD45RA', 'CD19', 'CD11b', 'CD4', 'CD8', 'CD34',
'CD20', 'CD33', 'CD123', 'CD38', 'CD90', 'CD3']
df.columns = channels + ['cell_type']
df = df[df.cell_type != 'NotGated']
df = df.loc[df['cell_type'] != 'NotDebrisSinglets']
df = df.loc[df['cell_type'] != 'Megakaryocyte']
df = df.loc[df['cell_type'] != 'CD11bmid Monocyte']
df = df.loc[df['cell_type'] != 'Platelet']
df = df.loc[df['cell_type'] != 'Myelocyte']
df = df.loc[df['cell_type'] != 'Erythroblast']
table = table.fillna(0)
X = df[channels].values
### transform data
data = np.arcsinh((X-1.)/5.)
theta_space = np.array([[data[:,d].min(), data[:,d].max()] for d in range(data.shape[1])])
cell_type_name2idx = {x:i for i,x in enumerate(table.index)}
cell_type_name2idx['unknown'] = len(cell_type_name2idx)
Y = np.array([cell_type_name2idx[_]
if _ in cell_type_name2idx else cell_type_name2idx['unknown']
for _ in df.cell_type])
print table
print cell_type_name2idx
from sklearn.utils import shuffle
N = data.shape[0]
new_df = shuffle(df)[:N]
X_subset = new_df[channels].values
data_subset = np.arcsinh((X_subset-1.)/5.)
Y_subset = np.array([cell_type_name2idx[_]
if _ in cell_type_name2idx else cell_type_name2idx['unknown']
for _ in new_df.cell_type])
N, d = data_subset.shape
print N,d
emp_bounds = np.array([(data_subset[:,i].min(), data_subset[:,i].max()) for i in range(d)])
%%time
n_mcmc_chain = 50
n_mcmc_sample = 2000
mcmc_gaussin_std = 0.1 # tune step size s.t. acceptance rate ~50%
accepts = [[] for _ in range(n_mcmc_chain)]
rejects = [[] for _ in range(n_mcmc_chain)]
logl_accepted_trace = [[] for _ in range(n_mcmc_chain)]
logl_complete_trace = [[] for _ in range(n_mcmc_chain)]
Y_predict_accepted_trace = [[] for _ in range(n_mcmc_chain)]
accuracy_accepted_trace = [[] for _ in range(n_mcmc_chain)]
for chain in range(n_mcmc_chain):
print "Drawing Chain %d ..." % chain
sample = draw_informed_Mondrian(emp_bounds, table)
log_p_sample = comp_log_p_sample(sample, data_subset)
accepts[chain].append(sample)
logl_accepted_trace[chain].append(log_p_sample)
logl_complete_trace[chain].append(log_p_sample)
Y_predict = classify_cells(data_subset, sample, table, cell_type_name2idx)
accuracy = sum(Y_subset == Y_predict)*1.0/ data_subset.shape[0]
accuracy_accepted_trace[chain].append(accuracy)
Y_predict_accepted_trace[chain].append(Y_predict)
for idx in range(n_mcmc_sample):
new_sample = Mondrian_Gaussian_perturbation(emp_bounds,sample, mcmc_gaussin_std)
new_log_p_sample = comp_log_p_sample(new_sample, data_subset)
logl_complete_trace[chain].append(new_log_p_sample)
if new_log_p_sample < log_p_sample and \
np.log(np.random.uniform(low=0, high=1.)) > (new_log_p_sample - log_p_sample):
rejects[chain].append(new_sample)
else:
if new_log_p_sample < log_p_sample:
print "accepted some bad samples"
sample = new_sample
log_p_sample = new_log_p_sample
accepts[chain].append(sample)
logl_accepted_trace[chain].append(log_p_sample)
Y_predict = classify_cells(data_subset, sample, table, cell_type_name2idx)
accuracy = sum(Y_subset == Y_predict)*1.0/ data_subset.shape[0]
accuracy_accepted_trace[chain].append(accuracy)
Y_predict_accepted_trace[chain].append(Y_predict)
if (idx+1) % 500 == 0:
print "Iteration %d, cummulative accepted sample size is %d" %(idx+1, len(accepts[chain]))
if (chain + 1) % 10 == 0:
# prediction and visualization
Y_predict = classify_cells(data_subset, accepts[chain][-1], table, cell_type_name2idx)
accuracy = sum(Y_subset == Y_predict)*1.0/ data_subset.shape[0]
print "Chain % d accuracy on subset data: %.3f" % (chain+1,accuracy)
print "Total number of accepted samples: %d" %(sum([len(accepts[chain]) for chain in range(n_mcmc_chain)]))
# plot 5 chains
fig, axs = plt.subplots(5, 3, figsize=(10,10) )
for chain in range(5):
axs[chain, 0].plot(logl_complete_trace[chain])
axs[chain, 1].plot(logl_accepted_trace[chain])
axs[chain, 2].plot(accuracy_accepted_trace[chain])
axs[chain, 0].set_title('Trace of likelihood Chain %d, all samples' % chain, fontsize=8)
axs[chain, 1].set_title('Trace of likelihood Chain %d, accepted samples' % chain, fontsize=8)
axs[chain, 2].set_title('Trace of accuracy Chain %d, accepted samples' % chain, fontsize=8)
fig.tight_layout()
L = min(len(_) for _ in accuracy_accepted_trace)
res = []
for i in range(L):
res.append(np.array([_[i] for _ in accuracy_accepted_trace]).mean())
plt.plot(res)
# vote, and compute accuracy
# keep last 2 samples
burnt_samples = []
burnt_predictions = []
for i in range(len(accepts)):
accepted_chain = accepts[i]
likelihoods = logl_accepted_trace[i]
predictions = Y_predict_accepted_trace[i]
burnt_samples += [accepted_chain[_] for _
in range(len(accepted_chain) - 2,len(accepted_chain))]
burnt_predictions += [predictions[_] for _
in range(len(accepted_chain) - 2,len(accepted_chain))]
# vote
votes = np.zeros([data_subset.shape[0], table.shape[0]])
for Y_predict in burnt_predictions:
for _ in range(len(Y_predict)):
votes[_,Y_predict[_]] += 1
Y_predict_majority = np.argmax(votes, axis=1)
print votes.sum()
accuracy = sum(Y_subset == Y_predict_majority)*1.0/ data_subset.shape[0]
print "Accuracy on subset data: %.3f" % (accuracy)
bins = table.shape[0]
plt.hist(Y_subset, bins, alpha=0.5, label='Y:cell type')
plt.hist(Y_predict_majority, bins, alpha=0.5, label='Z:prediction')
plt.legend(loc='upper right')
plt.show()
# vote, and compute accuracy
# keep the first sample
burnt_samples = []
burnt_predictions = []
for i in range(len(accepts)):
accepted_chain = accepts[i]
likelihoods = logl_accepted_trace[i]
predictions = Y_predict_accepted_trace[i]
burnt_samples += [accepted_chain[_] for _
in range(0,1)]
burnt_predictions += [predictions[_] for _
in range(0,1)]
# vote
votes = np.zeros([data_subset.shape[0], table.shape[0]])
for Y_predict in burnt_predictions:
for _ in range(len(Y_predict)):
votes[_,Y_predict[_]] += 1
Y_predict_majority = np.argmax(votes, axis=1)
print votes.sum()
accuracy = sum(Y_subset == Y_predict_majority)*1.0/ data_subset.shape[0]
print "Accuracy on subset data: %.3f" % (accuracy)
bins = table.shape[0]
plt.hist(Y_subset, bins, alpha=0.5, label='Y:cell type')
plt.hist(Y_predict_majority, bins, alpha=0.5, label='Z:prediction')
plt.legend(loc='upper right')
plt.show()
```
| github_jupyter |
- V1 : LGBM STACKING
- V2 : LGBM, MLP16 STACKING
- V3 : V2 + pred 5 score 3
```
import warnings
warnings.filterwarnings('ignore')
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from tqdm import tqdm_notebook
from sklearn import svm, neighbors, linear_model, neural_network
from sklearn.metrics import roc_auc_score
from sklearn.preprocessing import StandardScaler, MinMaxScaler, QuantileTransformer
from sklearn.model_selection import StratifiedKFold
from sklearn.feature_selection import VarianceThreshold
from sklearn.decomposition import PCA, TruncatedSVD, KernelPCA
from sklearn.mixture import GaussianMixture as GMM
from sklearn.metrics import silhouette_score
import lightgbm as lgb
from sklearn.metrics import roc_auc_score
from sklearn.model_selection import StratifiedKFold, KFold
from sklearn import preprocessing
from sklearn import svm, neighbors, linear_model
import gc
warnings.filterwarnings('ignore')
from sklearn.neural_network import MLPClassifier
from sklearn.naive_bayes import GaussianNB, BernoulliNB
from sklearn.gaussian_process import GaussianProcessClassifier
from sklearn.gaussian_process.kernels import Matern, RationalQuadratic
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis, QuadraticDiscriminantAnalysis
from sklearn.decomposition import FastICA, TruncatedSVD, PCA
from sklearn.ensemble import RandomForestClassifier
import lightgbm as lgb
import xgboost as xgb
import catboost as cat
from tqdm import *
%%time
train_df = pd.read_csv('../input/train.csv')
test_df = pd.read_csv('../input/test.csv')
train_columns = [c for c in train_df.columns if c not in ['id','target','wheezy-copper-turtle-magic']]
magic_variance_over2 = {}
for magic in sorted(train_df['wheezy-copper-turtle-magic'].unique()):
temp = train_df.loc[train_df['wheezy-copper-turtle-magic']==magic]
std = temp[train_columns].std()
magic_variance_over2[magic] = list(std.index.values[np.where(std >2)])
class hist_model(object):
def __init__(self, bins=50):
self.bins = bins
def fit(self, X):
bin_hight, bin_edge = [], []
for var in X.T:
# get bins hight and interval
bh, bedge = np.histogram(var, bins=self.bins)
bin_hight.append(bh)
bin_edge.append(bedge)
self.bin_hight = np.array(bin_hight)
self.bin_edge = np.array(bin_edge)
def predict(self, X):
scores = []
for obs in X:
obs_score = []
for i, var in enumerate(obs):
# find wich bin obs is in
bin_num = (var > self.bin_edge[i]).argmin()-1
obs_score.append(self.bin_hight[i, bin_num]) # find bin hitght
scores.append(np.mean(obs_score))
return np.array(scores)
from sklearn.neighbors import KNeighborsClassifier, RadiusNeighborsClassifier
from sklearn.gaussian_process import GaussianProcessClassifier
from sklearn.gaussian_process.kernels import RBF
from sklearn.naive_bayes import GaussianNB, MultinomialNB, BernoulliNB
from sklearn.linear_model import SGDClassifier
from sklearn.linear_model import Lasso, LassoLars
random_state = 42
debug = True
debug = False
svnu_params = {'probability':True, 'kernel':'poly','degree':4,'gamma':'auto','nu':0.4,'coef0':0.08, 'random_state':4}
svnu2_params = {'probability':True, 'kernel':'poly','degree':2,'gamma':'auto','nu':0.4,'coef0':0.08, 'random_state':4}
svc_params = {'probability':True,'kernel':'poly','degree':4,'gamma':'auto', 'random_state':4}
lr_params = {'solver':'liblinear','penalty':'l1','C':0.05,'n_jobs':-1, 'random_state':42}
mlp16_params = {'activation':'relu','solver':'lbfgs','tol':1e-06, 'hidden_layer_sizes':(16, ), 'random_state':42}
mlp128_params = {'activation':'relu','solver':'lbfgs','tol':1e-06, 'hidden_layer_sizes':(128, ), 'random_state':42}
def get_oofs(random_state):
oof_nusvc = np.zeros(len(train_df))
preds_nusvc = np.zeros(len(test_df))
oof_nusvc2 = np.zeros(len(train_df))
preds_nusvc2 = np.zeros(len(test_df))
oof_qda = np.zeros(len(train_df))
preds_qda = np.zeros(len(test_df))
oof_svc = np.zeros(len(train_df))
preds_svc = np.zeros(len(test_df))
oof_knn = np.zeros(len(train_df))
preds_knn = np.zeros(len(test_df))
oof_lr = np.zeros(len(train_df))
preds_lr = np.zeros(len(test_df))
cols = [c for c in train_df.columns if c not in ['id', 'target', 'wheezy-copper-turtle-magic']]
for i in tqdm_notebook(range(512)):
# each magic
train = train_df[train_df['wheezy-copper-turtle-magic'] == i]
test = test_df[test_df['wheezy-copper-turtle-magic'] == i]
# for oof
train_idx_origin = train.index
test_idx_origin = test.index
# start point
# new cols
cols = magic_variance_over2[i]
X_train = train.reset_index(drop=True)[cols].values
y_train = train.reset_index(drop=True).target
X_test = test[cols].values
# vstack
data = np.vstack([X_train, X_test])
# PCA
data = KernelPCA(n_components=len(cols), kernel='cosine', random_state=random_state).fit_transform(data)
# Bad
'''
gmm_pred = np.zeros((len(data), 5))
for j in range(5):
gmm = GMM(n_components=4, random_state=random_state + j, max_iter=1000).fit(data)
gmm_pred[:, j] += gmm.predict(data)
'''
# original
gmm = GMM(n_components=5, random_state=random_state, max_iter=1000).fit(data)
gmm_pred = gmm.predict_proba(data)
gmm_score = gmm.score_samples(data)
gmm_label = gmm.predict(data)
hist = hist_model(); hist.fit(data)
hist_pred = hist.predict(data).reshape(-1, 1)
data = np.hstack([data, gmm_pred])
# HOXI
data = np.hstack([data, gmm_pred])
data = np.hstack([data, gmm_pred])
data = np.hstack([data, gmm_pred])
# Add Some Features
data = np.hstack([data, gmm_pred])
data = np.hstack([data, hist_pred, gmm_score.reshape(-1, 1)])
data = np.hstack([data, gmm_score.reshape(-1, 1)])
data = np.hstack([data, gmm_score.reshape(-1, 1)])
# STANDARD SCALER
data = StandardScaler().fit_transform(data)
# new train/test
X_train = data[:X_train.shape[0]]
X_test = data[X_train.shape[0]:]
fold = StratifiedKFold(n_splits=5, random_state=random_state)
for tr_idx, val_idx in fold.split(X_train, gmm_label[:X_train.shape[0]]):
# NuSVC 1
clf = svm.NuSVC(**svnu_params)
clf.fit(X_train[tr_idx], y_train[tr_idx])
oof_nusvc[train_idx_origin[val_idx]] = clf.predict_proba(X_train[val_idx])[:,1]
preds_nusvc[test_idx_origin] += clf.predict_proba(X_test)[:,1] / fold.n_splits
# NuSVC 2
clf = svm.NuSVC(**svnu2_params)
clf.fit(X_train[tr_idx], y_train[tr_idx])
oof_nusvc2[train_idx_origin[val_idx]] = clf.predict_proba(X_train[val_idx])[:,1]
preds_nusvc2[test_idx_origin] += clf.predict_proba(X_test)[:,1] / fold.n_splits
# qda 3
clf = QuadraticDiscriminantAnalysis(reg_param=0.111)
clf.fit(X_train[tr_idx], y_train[tr_idx])
oof_qda[train_idx_origin[val_idx]] = clf.predict_proba(X_train[val_idx])[:,1]
preds_qda[test_idx_origin] += clf.predict_proba(X_test)[:,1] / fold.n_splits
# SVC 4
clf = svm.SVC(**svc_params)
clf.fit(X_train[tr_idx], y_train[tr_idx])
oof_svc[train_idx_origin[val_idx]] = clf.predict_proba(X_train[val_idx])[:,1]
preds_svc[test_idx_origin] += clf.predict_proba(X_test)[:,1] / fold.n_splits
# knn 8
clf = KNeighborsClassifier(n_neighbors=16)
clf.fit(X_train[tr_idx], y_train[tr_idx])
oof_knn[train_idx_origin[val_idx]] = clf.predict_proba(X_train[val_idx])[:,1]
preds_knn[test_idx_origin] += clf.predict_proba(X_test)[:,1] / fold.n_splits
# LR 5
clf = linear_model.LogisticRegression(**lr_params)
clf.fit(X_train[tr_idx], y_train[tr_idx])
oof_lr[train_idx_origin[val_idx]] = clf.predict_proba(X_train[val_idx])[:,1]
preds_lr[test_idx_origin] += clf.predict_proba(X_test)[:,1] / fold.n_splits
oof_train = pd.DataFrame()
oof_train['nusvc'] = oof_nusvc
oof_train['nusvc2'] = oof_nusvc2
oof_train['qda'] = oof_qda
oof_train['svc'] = oof_svc
oof_train['knn'] = oof_knn
oof_train['lr'] = oof_lr
oof_test = pd.DataFrame()
oof_test['nusvc'] = preds_nusvc
oof_test['nusvc2'] = preds_nusvc2
oof_test['qda'] = preds_qda
oof_test['svc'] = preds_svc
oof_test['knn'] = preds_knn
oof_test['lr'] = preds_lr
print('nusvc', roc_auc_score(train_df['target'], oof_nusvc))
print('nusvc2', roc_auc_score(train_df['target'], oof_nusvc2))
print('qda', roc_auc_score(train_df['target'], oof_qda))
print('svc', roc_auc_score(train_df['target'], oof_svc))
print('knn', roc_auc_score(train_df['target'], oof_knn))
print('knn', roc_auc_score(train_df['target'], oof_lr))
return oof_train, oof_test
def get_oofs_2(random_state):
oof_nusvc = np.zeros(len(train_df))
preds_nusvc = np.zeros(len(test_df))
oof_nusvc2 = np.zeros(len(train_df))
preds_nusvc2 = np.zeros(len(test_df))
oof_qda = np.zeros(len(train_df))
preds_qda = np.zeros(len(test_df))
oof_svc = np.zeros(len(train_df))
preds_svc = np.zeros(len(test_df))
oof_knn = np.zeros(len(train_df))
preds_knn = np.zeros(len(test_df))
oof_lr = np.zeros(len(train_df))
preds_lr = np.zeros(len(test_df))
cols = [c for c in train_df.columns if c not in ['id', 'target', 'wheezy-copper-turtle-magic']]
for i in tqdm_notebook(range(512)):
# each magic
train = train_df[train_df['wheezy-copper-turtle-magic'] == i]
test = test_df[test_df['wheezy-copper-turtle-magic'] == i]
# for oof
train_idx_origin = train.index
test_idx_origin = test.index
# start point
# new cols
cols = magic_variance_over2[i]
X_train = train.reset_index(drop=True)[cols].values
y_train = train.reset_index(drop=True).target
X_test = test[cols].values
# vstack
data = np.vstack([X_train, X_test])
# PCA
data = KernelPCA(n_components=len(cols), kernel='cosine', random_state=random_state).fit_transform(data)
# Bad
'''
gmm_pred = np.zeros((len(data), 5))
for j in range(5):
gmm = GMM(n_components=4, random_state=random_state + j, max_iter=1000).fit(data)
gmm_pred[:, j] += gmm.predict(data)
'''
# original
gmm = GMM(n_components=5, random_state=random_state, max_iter=1000, init_params='random').fit(data)
gmm_pred = gmm.predict_proba(data)
gmm_score = gmm.score_samples(data)
gmm_label = gmm.predict(data)
hist = hist_model(); hist.fit(data)
hist_pred = hist.predict(data).reshape(-1, 1)
data = np.hstack([data, gmm_pred])
# HOXI
data = np.hstack([data, gmm_pred])
data = np.hstack([data, gmm_pred])
data = np.hstack([data, gmm_pred])
# Add Some Features
data = np.hstack([data, gmm_pred])
data = np.hstack([data, hist_pred, gmm_score.reshape(-1, 1)])
data = np.hstack([data, gmm_score.reshape(-1, 1)])
data = np.hstack([data, gmm_score.reshape(-1, 1)])
# STANDARD SCALER
data = StandardScaler().fit_transform(data)
# new train/test
X_train = data[:X_train.shape[0]]
X_test = data[X_train.shape[0]:]
fold = StratifiedKFold(n_splits=5, random_state=random_state)
for tr_idx, val_idx in fold.split(X_train, gmm_label[:X_train.shape[0]]):
# NuSVC 1
clf = svm.NuSVC(**svnu_params)
clf.fit(X_train[tr_idx], y_train[tr_idx])
oof_nusvc[train_idx_origin[val_idx]] = clf.predict_proba(X_train[val_idx])[:,1]
preds_nusvc[test_idx_origin] += clf.predict_proba(X_test)[:,1] / fold.n_splits
# NuSVC 2
clf = svm.NuSVC(**svnu2_params)
clf.fit(X_train[tr_idx], y_train[tr_idx])
oof_nusvc2[train_idx_origin[val_idx]] = clf.predict_proba(X_train[val_idx])[:,1]
preds_nusvc2[test_idx_origin] += clf.predict_proba(X_test)[:,1] / fold.n_splits
# qda 3
clf = QuadraticDiscriminantAnalysis(reg_param=0.111)
clf.fit(X_train[tr_idx], y_train[tr_idx])
oof_qda[train_idx_origin[val_idx]] = clf.predict_proba(X_train[val_idx])[:,1]
preds_qda[test_idx_origin] += clf.predict_proba(X_test)[:,1] / fold.n_splits
# SVC 4
clf = svm.SVC(**svc_params)
clf.fit(X_train[tr_idx], y_train[tr_idx])
oof_svc[train_idx_origin[val_idx]] = clf.predict_proba(X_train[val_idx])[:,1]
preds_svc[test_idx_origin] += clf.predict_proba(X_test)[:,1] / fold.n_splits
# knn 8
clf = KNeighborsClassifier(n_neighbors=16)
clf.fit(X_train[tr_idx], y_train[tr_idx])
oof_knn[train_idx_origin[val_idx]] = clf.predict_proba(X_train[val_idx])[:,1]
preds_knn[test_idx_origin] += clf.predict_proba(X_test)[:,1] / fold.n_splits
# LR 5
clf = linear_model.LogisticRegression(**lr_params)
clf.fit(X_train[tr_idx], y_train[tr_idx])
oof_lr[train_idx_origin[val_idx]] = clf.predict_proba(X_train[val_idx])[:,1]
preds_lr[test_idx_origin] += clf.predict_proba(X_test)[:,1] / fold.n_splits
oof_train = pd.DataFrame()
oof_train['nusvc'] = oof_nusvc
oof_train['nusvc2'] = oof_nusvc2
oof_train['qda'] = oof_qda
oof_train['svc'] = oof_svc
oof_train['knn'] = oof_knn
oof_train['lr'] = oof_lr
oof_test = pd.DataFrame()
oof_test['nusvc'] = preds_nusvc
oof_test['nusvc2'] = preds_nusvc2
oof_test['qda'] = preds_qda
oof_test['svc'] = preds_svc
oof_test['knn'] = preds_knn
oof_test['lr'] = preds_lr
print('nusvc', roc_auc_score(train_df['target'], oof_nusvc))
print('nusvc2', roc_auc_score(train_df['target'], oof_nusvc2))
print('qda', roc_auc_score(train_df['target'], oof_qda))
print('svc', roc_auc_score(train_df['target'], oof_svc))
print('knn', roc_auc_score(train_df['target'], oof_knn))
print('knn', roc_auc_score(train_df['target'], oof_lr))
return oof_train, oof_test
oof_train_1, oof_test_1 = get_oofs(1)
oof_train_2, oof_test_2 = get_oofs(2)
oof_train_3, oof_test_3 = get_oofs_2(1)
oof_train_4, oof_test_4 = get_oofs_2(2)
x_train_second_layer = oof_train_1 + oof_train_2 + oof_train_3 + oof_train_4
x_test_second_layer = oof_test_1 + oof_test_2 + oof_test_3 + oof_test_4
print('Ensemble', roc_auc_score(train_df['target'], x_train_second_layer.mean(1)))
```
아래 SEED 수정 된 거로 바꿔야 함.
```
def time_decorator(func):
@wraps(func)
def wrapper(*args, **kwargs):
print("\nStartTime: ", datetime.now() + timedelta(hours=9))
start_time = time.time()
df = func(*args, **kwargs)
print("EndTime: ", datetime.now() + timedelta(hours=9))
print("TotalTime: ", time.time() - start_time)
return df
return wrapper
class SklearnWrapper(object):
def __init__(self, clf, params=None, **kwargs):
"""
params['random_state'] = kwargs.get('seed', 0)
self.clf = clf(**params)
self.is_classification_problem = True
"""
if 'seed' in kwargs:
params['random_state'] = kwargs.get('seed', 0)
self.clf = clf(**params)
self.is_classification_problem = True
#@time_decorator
def train(self, x_train, y_train, x_cross=None, y_cross=None):
if len(np.unique(y_train)) > 30:
self.is_classification_problem = False
self.clf.fit(x_train, y_train)
def predict(self, x):
if self.is_classification_problem is True:
return self.clf.predict_proba(x)[:,1]
else:
return self.clf.predict(x)
class LgbmWrapper(object):
def __init__(self, params=None, **kwargs):
self.param = params
if 'seed' in kwargs:
self.param['seed'] = kwargs.get('seed', 0)
self.num_rounds = kwargs.get('num_rounds', 1000)
self.early_stopping = kwargs.get('ealry_stopping', 100)
self.eval_function = kwargs.get('eval_function', None)
self.verbose_eval = kwargs.get('verbose_eval', 100)
self.best_round = 0
self.feature_importance = pd.DataFrame()
#@time_decorator
def train(self, x_train, y_train, x_cross=None, y_cross=None):
"""
x_cross or y_cross is None
-> model train limted num_rounds
x_cross and y_cross is Not None
-> model train using validation set
"""
if isinstance(y_train, pd.DataFrame) is True:
y_train = y_train[y_train.columns[0]]
if y_cross is not None:
y_cross = y_cross[y_cross.columns[0]]
if x_cross is None:
dtrain = lgb.Dataset(x_train, label=y_train, silent= True)
train_round = self.best_round
if self.best_round == 0:
train_round = self.num_rounds
self.clf = lgb.train(self.param, train_set=dtrain, num_boost_round=train_round)
del dtrain
else:
dtrain = lgb.Dataset(x_train, label=y_train, silent=True)
dvalid = lgb.Dataset(x_cross, label=y_cross, silent=True)
self.clf = lgb.train(self.param, train_set=dtrain, num_boost_round=self.num_rounds, valid_sets=[dtrain, dvalid],
feval=self.eval_function, early_stopping_rounds=self.early_stopping,
verbose_eval=self.verbose_eval)
try:
self.feature_importance = pd.DataFrame()
self.feature_importance["Feature"] = x_train.columns
self.feature_importance["Importance"] = self.clf.feature_importance()
except:
pass
self.best_round = max(self.best_round, self.clf.best_iteration)
del dtrain, dvalid
gc.collect()
def get_importance_df(self):
return self.feature_importance
def predict(self, x):
return self.clf.predict(x, num_iteration=self.clf.best_iteration)
def plot_importance(self):
lgb.plot_importance(self.clf, max_num_features=50, height=0.7, figsize=(10,30))
plt.show()
def get_params(self):
return self.param
#@time_decorator
def get_oof(clf, x_train, y_train, x_test, eval_func, **kwargs):
nfolds = kwargs.get('NFOLDS', 5)
kfold_shuffle = kwargs.get('kfold_shuffle', True)
kfold_random_state = kwargs.get('kfold_random_state', 0)
stratified_kfold_ytrain = kwargs.get('stratifed_kfold_y_value', None)
inner_predict = kwargs.get('inner_predict', True)
export_feature_importance = kwargs.get('export_feature_importance', True)
ntrain = x_train.shape[0]
ntest = x_test.shape[0]
kf_split = None
if stratified_kfold_ytrain is None:
kf = KFold(n_splits=nfolds, shuffle=kfold_shuffle, random_state=kfold_random_state)
kf_split = kf.split(x_train)
else:
kf = StratifiedKFold(n_splits=nfolds, shuffle=kfold_shuffle, random_state=kfold_random_state)
kf_split = kf.split(x_train, stratified_kfold_ytrain)
oof_train = np.zeros((ntrain,))
oof_test = np.zeros((ntest,))
cv_sum = 0
# before running model, print model param
# lightgbm model and xgboost model use get_params()
"""
try:
if clf.clf is not None:
print(clf.clf)
except:
print(clf)
print(clf.get_params())
"""
feature_importance_df = pd.DataFrame()
for i, (train_index, cross_index) in enumerate(kf_split):
x_tr, x_cr = None, None
y_tr, y_cr = None, None
if isinstance(x_train, pd.DataFrame):
x_tr, x_cr = x_train.iloc[train_index], x_train.iloc[cross_index]
y_tr, y_cr = y_train.iloc[train_index], y_train.iloc[cross_index]
else:
x_tr, x_cr = x_train[train_index], x_train[cross_index]
y_tr, y_cr = y_train[train_index], y_train[cross_index]
clf.train(x_tr, y_tr, x_cr, y_cr)
if isinstance(clf, LgbmWrapper) is True:
feature_importance_df = pd.concat([feature_importance_df, clf.get_importance_df()], axis=0)
oof_train[cross_index] = clf.predict(x_cr)
if inner_predict is True:
oof_test += clf.predict(x_test)
cv_score = eval_func(y_cr, oof_train[cross_index])
#print('Fold %d / ' % (i+1), 'CV-Score: %.6f' % cv_score)
cv_sum = cv_sum + cv_score
del x_tr, x_cr, y_tr, y_cr
gc.collect()
score = cv_sum / nfolds
#print("Average CV-Score: ", score)
#print("OOF CV-Score: ", eval_func(y_train, oof_train))
if export_feature_importance is True:
print("Export Feature Importance")
filename = '{}_cv{:.6f}'.format(datetime.now().strftime('%Y%m%d_%H%M%S'), score)
if os.path.isdir("importance/") is True:
feature_importance_df.to_csv('importance/importance_{}.csv'.format(filename),index=False)
else:
feature_importance_df.to_csv('importance_{}.csv'.format(filename),index=False)
if inner_predict is True:
oof_test = oof_test/nfolds
else:
# Using All Dataset, retrain
clf.train(x_train, y_train)
oof_test = clf.predict(x_test)
return oof_train, oof_test, score
x_train_second_layer1 = pd.DataFrame(x_train_second_layer)
x_test_second_layer1 = pd.DataFrame(x_test_second_layer)
import time
from datetime import datetime, timedelta,date
import warnings
import itertools
from functools import wraps
import functools
import seaborn as sns
import matplotlib.pyplot as plt
import lightgbm as lgb
from sklearn.metrics import roc_auc_score
from sklearn.model_selection import StratifiedKFold, KFold
from sklearn import preprocessing
from sklearn import svm, neighbors, linear_model
import gc
warnings.filterwarnings('ignore')
from sklearn.mixture import GaussianMixture as GMM
param = {
#'bagging_freq': 5,
#'bagging_fraction': 0.8,
'min_child_weight':6.790,
"subsample_for_bin":50000,
'bagging_seed': 0,
'boost_from_average':'true',
'boost': 'gbdt',
'feature_fraction': 0.450,
'bagging_fraction': 0.343,
'learning_rate': 0.025,
'max_depth': 10,
'metric':'auc',
'min_data_in_leaf': 78,
'min_sum_hessian_in_leaf': 8,
'num_leaves': 18,
'num_threads': 8,
'tree_learner': 'serial',
'objective': 'binary',
'verbosity': 1,
'lambda_l1': 7.961,
'lambda_l2': 7.781
#'reg_lambda': 0.3,
}
mlp16_params = {'activation':'relu','solver':'lbfgs','tol':1e-06, 'hidden_layer_sizes':(16, ), 'random_state':42}
lgbm_meta_model = LgbmWrapper(params=param, num_rounds = 2000, ealry_stopping=100)
mlp_meta_model = SklearnWrapper(neural_network.MLPClassifier,mlp16_params)
third_number = 4
oof_train_5 = pd.DataFrame()
oof_test_5 = pd.DataFrame()
third_oof = np.zeros(len(train_df))
third_pred = np.zeros(len(test_df))
third_oof1 = np.zeros(len(train_df))
third_pred1 = np.zeros(len(test_df))
for SEED in np.arange(third_number):
second_oof, second_pred, second_score = get_oof(lgbm_meta_model, x_train_second_layer1, train_df['target'], x_test_second_layer1, eval_func=roc_auc_score, NFOLDS=5, kfold_random_sate= SEED )
second_oof1, second_pred1, second_score1 = get_oof(mlp_meta_model, x_train_second_layer1, train_df['target'], x_test_second_layer1, eval_func=roc_auc_score, NFOLDS=5, kfold_random_sate= SEED )
third_oof += second_oof
third_pred += second_pred
print(second_score)
third_oof1 += second_oof1
third_pred1 += second_pred1
print(second_score1)
print("")
oof_train_5['lgb'] = third_oof
oof_test_5['lgb'] = third_pred
oof_train_5['mlp'] = third_oof1
oof_test_5['mlp'] = third_pred1
x_train_second_layer = oof_train_1 + oof_train_2 + oof_train_3 + oof_train_4
x_test_second_layer = oof_test_1 + oof_test_2 + oof_test_3 + oof_test_4
x_train_second_layer = pd.concat([x_train_second_layer,oof_train_5],axis=1)
x_test_second_layer = pd.concat([x_test_second_layer,oof_test_5],axis=1)
print('Ensemble', roc_auc_score(train_df['target'], x_train_second_layer.mean(1)))
submit = pd.read_csv('../input/sample_submission.csv')
submit["target"] = x_test_second_layer.mean(1)
submit.to_csv("submission.csv", index=False)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/W2D1-postcourse-bugfix/tutorials/W2D1_BayesianStatistics/W2D1_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Neuromatch Academy: Week 2, Day 1, Tutorial 1
# Bayes rule with Gaussians
__Content creators:__ Vincent Valton, Konrad Kording, with help from Matt Krause
__Content reviewers:__ Matt Krause, Jesse Livezey, Karolina Stosio, Saeed Salehi, Michael Waskom
# Tutorial Objectives
This is the first in a series of three main tutorials (+ one bonus tutorial) on Bayesian statistics. In these tutorials, we will develop a Bayesian model for localizing sounds based on audio and visual cues. This model will combine **prior** information about where sounds generally originate with sensory information about the **likelihood** that a specific sound came from a particular location. As we will see in subsequent lessons, the resulting **posterior distribution** not only allows us to make optimal decision about the sound's origin, but also lets us quantify how uncertain that decision is. Bayesian techniques are therefore useful **normative models**: the behavior of human or animal subjects can be compared against these models to determine how efficiently they make use of information.
This notebook will introduce two fundamental building blocks for Bayesian statistics: the Gaussian distribution and the Bayes Theorem. You will:
1. Implement a Gaussian distribution
2. Use Bayes' Theorem to find the posterior from a Gaussian-distributed prior and likelihood.
3. Change the likelihood mean and variance and observe how posterior changes.
4. Advanced (*optional*): Observe what happens if the prior is a mixture of two gaussians?
```
#@title Video 1: Introduction to Bayesian Statistics
from IPython.display import YouTubeVideo
video = YouTubeVideo(id='K4sSKZtk-Sc', width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
```
## Setup
Please execute the cells below to initialize the notebook environment.
```
import numpy as np
import matplotlib.pyplot as plt
#@title Figure Settings
import ipywidgets as widgets
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
#@title Helper functions
def my_plot_single(x, px):
"""
Plots normalized Gaussian distribution
Args:
x (numpy array of floats): points at which the likelihood has been evaluated
px (numpy array of floats): normalized probabilities for prior evaluated at each `x`
Returns:
Nothing.
"""
if px is None:
px = np.zeros_like(x)
fig, ax = plt.subplots()
ax.plot(x, px, '-', color='C2', LineWidth=2, label='Prior')
ax.legend()
ax.set_ylabel('Probability')
ax.set_xlabel('Orientation (Degrees)')
def posterior_plot(x, likelihood=None, prior=None, posterior_pointwise=None, ax=None):
"""
Plots normalized Gaussian distributions and posterior
Args:
x (numpy array of floats): points at which the likelihood has been evaluated
auditory (numpy array of floats): normalized probabilities for auditory likelihood evaluated at each `x`
visual (numpy array of floats): normalized probabilities for visual likelihood evaluated at each `x`
posterior (numpy array of floats): normalized probabilities for the posterior evaluated at each `x`
ax: Axis in which to plot. If None, create new axis.
Returns:
Nothing.
"""
if likelihood is None:
likelihood = np.zeros_like(x)
if prior is None:
prior = np.zeros_like(x)
if posterior_pointwise is None:
posterior_pointwise = np.zeros_like(x)
if ax is None:
fig, ax = plt.subplots()
ax.plot(x, likelihood, '-C1', LineWidth=2, label='Auditory')
ax.plot(x, prior, '-C0', LineWidth=2, label='Visual')
ax.plot(x, posterior_pointwise, '-C2', LineWidth=2, label='Posterior')
ax.legend()
ax.set_ylabel('Probability')
ax.set_xlabel('Orientation (Degrees)')
return ax
def plot_visual(mu_visuals, mu_posteriors, max_posteriors):
"""
Plots the comparison of computing the mean of the posterior analytically and
the max of the posterior empirically via multiplication.
Args:
mu_visuals (numpy array of floats): means of the visual likelihood
mu_posteriors (numpy array of floats): means of the posterior, calculated analytically
max_posteriors (numpy array of floats): max of the posteriors, calculated via maxing the max_posteriors.
posterior (numpy array of floats): normalized probabilities for the posterior evaluated at each `x`
Returns:
Nothing.
"""
fig_w, fig_h = plt.rcParams.get('figure.figsize')
fig, ax = plt.subplots(nrows=2, ncols=1, figsize=(fig_w, 2 * fig_h))
ax[0].plot(mu_visuals, max_posteriors, '-C2', label='mean')
ax[0].set_xlabel('Visual stimulus position')
ax[0].set_ylabel('Multiplied posterior mean')
ax[0].set_title('Sample output')
ax[1].plot(mu_visuals, mu_posteriors, '--', color='xkcd:gray', label='argmax')
ax[1].set_xlabel('Visual stimulus position')
ax[1].set_ylabel('Analytical posterior mean')
fig.tight_layout()
ax[1].set_title('Hurray for math!')
def multimodal_plot(x, example_prior, example_likelihood,
mu_visuals, posterior_modes):
"""Helper function for plotting Section 4 results"""
fig_w, fig_h = plt.rcParams.get('figure.figsize')
fig, ax = plt.subplots(nrows=2, ncols=1, figsize=(fig_w, 2*fig_h), sharex=True)
# Plot the last instance that we tried.
posterior_plot(x,
example_prior,
example_likelihood,
compute_posterior_pointwise(example_prior, example_likelihood),
ax=ax[0]
)
ax[0].set_title('Example combination')
ax[1].plot(mu_visuals, posterior_modes, '-C2', label='argmax')
ax[1].set_xlabel('Visual stimulus position\n(Mean of blue dist. above)')
ax[1].set_ylabel('Posterior mode\n(Peak of green dist. above)')
fig.tight_layout()
```
# Section 1: The Gaussian Distribution
Bayesian analysis operates on probability distributions. Although these can take many forms, the Gaussian distribution is a very common choice. Because of the central limit theorem, many quantities are Gaussian-distributed. Gaussians also have some mathematical properties that permit simple closed-form solutions to several important problems.
In this exercise, you will implement a Gaussian by filling in the missing portion of `my_gaussian` below. Gaussians have two parameters. The **mean** $\mu$, which sets the location of its center. Its "scale" or spread is controlled by its **standard deviation** $\sigma$ or its square, the **variance** $\sigma^2$. (Be careful not to use one when the other is required).
The equation for a Gaussian is:
$$
\mathcal{N}(\mu,\sigma^2) = \frac{1}{\sqrt{2\pi\sigma^2}}\exp\left(\frac{-(x-\mu)^2}{2\sigma^2}\right)
$$
Also, don't forget that this is a probability distribution and should therefore sum to one. While this happens "automatically" when integrated from $-\infty$ to $\infty$, your version will only be computed over a finite number of points. You therefore need to explicitly normalize it yourself.
Test out your implementation with a $\mu = -1$ and $\sigma = 1$. After you have it working, play with the parameters to develop an intuition for how changing $\mu$ and $\sigma$ alter the shape of the Gaussian. This is important, because subsequent exercises will be built out of Gaussians.
## Exercise 1: Implement a Gaussian
```
def my_gaussian(x_points, mu, sigma):
"""
Returns normalized Gaussian estimated at points `x_points`, with parameters:
mean `mu` and std `sigma`
Args:
x_points (numpy array of floats): points at which the gaussian is
evaluated
mu (scalar): mean of the Gaussian
sigma (scalar): std of the gaussian
Returns:
(numpy array of floats) : normalized Gaussian evaluated at `x`
"""
###################################################################
## Add code to calcualte the gaussian px as a function of mu and sigma,
## for every x in x_points
## Function Hints: exp -> np.exp()
## power -> z**2
## remove the raise below to test your function
raise NotImplementedError("You need to implement the Gaussian function!")
###################################################################
px = ...
return px
x = np.arange(-8, 9, 0.1)
# Uncomment to plot the results
# px = my_gaussian(x, -1, 1)
# my_plot_single(x, px)
# to_remove solution
def my_gaussian(x_points, mu, sigma):
"""
Returns normalized Gaussian estimated at points `x_points`, with parameters:
mean `mu` and std `sigma`
Args:
x_points (numpy array of floats): points at which the gaussian is
evaluated
mu (scalar): mean of the Gaussian
sigma (scalar): std of the gaussian
Returns:
(numpy array of floats) : normalized Gaussian evaluated at `x`
"""
px = np.exp(- 1/2/sigma**2 * (mu - x_points) ** 2)
px = px / px.sum() # this is the normalization: this part ensures the sum of
# the individual probabilities at each `x` add up to one.
# It makes a very strong assumption though:
# That the `x_points` cover the big portion of
# probability mass around the mean.
# Please think/discuss when this would be a dangerous
# assumption.
# E.g.: What do you think will happen to the values on
# the y-axis
# if the `x` values (x_point) range from -1 to 8 instead
# of -8 to 8?
return px
x = np.arange(-8, 9, 0.1)
px = my_gaussian(x, -1, 1)
with plt.xkcd():
my_plot_single(x, px)
```
# Section 2. Bayes' Theorem and the Posterior
```
#@title Video 2: Bayes' theorem
from IPython.display import YouTubeVideo
video = YouTubeVideo(id='ewQPHQMcdBs', width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
```
Bayes' rule tells us how to combine two sources of information: the prior (e.g., a noisy representation of our expectations about where the stimulus might come from) and the likelihood (e.g., a noisy representation of the stimulus position on a given trial), to obtain a posterior distribution taking into account both pieces of information. Bayes' rule states:
\begin{eqnarray}
\text{Posterior} = \frac{ \text{Likelihood} \times \text{Prior}}{ \text{Normalization constant}}
\end{eqnarray}
When both the prior and likelihood are Gaussians, this translates into the following form:
$$
\begin{array}{rcl}
\text{Likelihood} &=& \mathcal{N}(\mu_{likelihood},\sigma_{likelihood}^2) \\
\text{Prior} &=& \mathcal{N}(\mu_{prior},\sigma_{prior}^2) \\
\text{Posterior} &\propto& \mathcal{N}(\mu_{likelihood},\sigma_{likelihood}^2) \times \mathcal{N}(\mu_{prior},\sigma_{prior}^2) \\
&&= \mathcal{N}\left( \frac{\sigma^2_{likelihood}\mu_{prior}+\sigma^2_{prior}\mu_{likelihood}}{\sigma^2_{likelihood}+\sigma^2_{prior}}, \frac{\sigma^2_{likelihood}\sigma^2_{prior}}{\sigma^2_{likelihood}+\sigma^2_{prior}} \right)
\end{array}
$$
In these equations, $\mathcal{N}(\mu,\sigma^2)$ denotes a Gaussian distribution with parameters $\mu$ and $\sigma^2$:
$$
\mathcal{N}(\mu, \sigma) = \frac{1}{\sqrt{2 \pi \sigma^2}} \; \exp \bigg( \frac{-(x-\mu)^2}{2\sigma^2} \bigg)
$$
In Exercise 2A, we will use the first form of the posterior, where the two distributions are combined via pointwise multiplication. Although this method requires more computation, it works for any type of probability distribution. In Exercise 2B, we will see that the closed-form solution shown on the line below produces the same result.
## Exercise 2A: Finding the posterior computationally
Imagine an experiment where participants estimate the location of a noise-emitting object. To estimate its position, the participants can use two sources of information:
1. new noisy auditory information (the likelihood)
2. prior visual expectations of where the stimulus is likely to come from (visual prior).
The auditory and visual information are both noisy, so participants will combine these sources of information to better estimate the position of the object.
We will use Gaussian distributions to represent the auditory likelihood (in red), and a Gaussian visual prior (expectations - in blue). Using Bayes rule, you will combine them into a posterior distribution that summarizes the probability that the object is in each location.
We have provided you with a ready-to-use plotting function, and a code skeleton.
* Use `my_gaussian`, the answer to exercise 1, to generate an auditory likelihood with parameters $\mu$ = 3 and $\sigma$ = 1.5
* Generate a visual prior with parameters $\mu$ = -1 and $\sigma$ = 1.5
* Calculate the posterior using pointwise multiplication of the likelihood and prior. Don't forget to normalize so the posterior adds up to 1.
* Plot the likelihood, prior and posterior using the predefined function `posterior_plot`
```
def compute_posterior_pointwise(prior, likelihood):
##############################################################################
# Write code to compute the posterior from the prior and likelihood via
# pointwise multiplication. (You may assume both are defined over the same x-axis)
#
# Comment out the line below to test your solution
raise NotImplementedError("Finish the simulation code first")
##############################################################################
posterior = ...
return posterior
def localization_simulation(mu_auditory=3.0, sigma_auditory=1.5,
mu_visual=-1.0, sigma_visual=1.5):
##############################################################################
## Using the x variable below,
## create a gaussian called 'auditory' with mean 3, and std 1.5
## create a gaussian called 'visual' with mean -1, and std 1.5
#
#
## Comment out the line below to test your solution
raise NotImplementedError("Finish the simulation code first")
###############################################################################
x = np.arange(-8, 9, 0.1)
auditory = ...
visual = ...
posterior = compute_posterior_pointwise(auditory, visual)
return x, auditory, visual, posterior
# Uncomment the lines below to plot the results
# x, auditory, visual, posterior_pointwise = localization_simulation()
# posterior_plot(x, auditory, visual, posterior_pointwise)
# to_remove solution
def compute_posterior_pointwise(prior, likelihood):
##############################################################################
# Write code to compute the posterior from the prior and likelihood via
# pointwise multiplication. (You may assume both are defined over the same x-axis)
#
# Comment out the line below to test your solution
# raise NotImplementedError("Finish the simulation code first")
##############################################################################
posterior = prior * likelihood
posterior /= posterior.sum()
return posterior
def localization_simulation(mu_auditory=3.0, sigma_auditory=1.5,
mu_visual=-1.0, sigma_visual=1.5):
##############################################################################
## Using the x variable below,
## create a gaussian called 'auditory' with mean 3, and std 1.5
## create a gaussian called 'visual' with mean -1, and std 1.5
#
## Comment out the line below to test your solution
#raise NotImplementedError("Finish the simulation code first")
###############################################################################
x = np.arange(-8, 9, 0.1)
auditory = my_gaussian(x, mu_auditory, sigma_auditory)
visual = my_gaussian(x, mu_visual, sigma_visual)
posterior = compute_posterior_pointwise(auditory, visual)
return x, auditory, visual, posterior
# Uncomment the lines below to plot the results
x, auditory, visual, posterior_pointwise = localization_simulation()
with plt.xkcd():
posterior_plot(x, auditory, visual, posterior_pointwise)
```
## Interactive Demo: What affects the posterior?
Now that we can compute the posterior of two Gaussians with *Bayes rule*, let's vary the parameters of those Gaussians to see how changing the prior and likelihood affect the posterior.
**Hit the Play button or Ctrl+Enter in the cell below** and play with the sliders to get an intuition for how the means and standard deviations of prior and likelihood influence the posterior.
When does the prior have the strongest influence over the posterior? When is it the weakest?
```
#@title
#@markdown Make sure you execute this cell to enable the widget!
x = np.arange(-10, 11, 0.1)
import ipywidgets as widgets
def refresh(mu_auditory=3, sigma_auditory=1.5, mu_visual=-1, sigma_visual=1.5):
auditory = my_gaussian(x, mu_auditory, sigma_auditory)
visual = my_gaussian(x, mu_visual, sigma_visual)
posterior_pointwise = visual * auditory
posterior_pointwise /= posterior_pointwise.sum()
w_auditory = (sigma_visual** 2) / (sigma_auditory**2 + sigma_visual**2)
theoretical_prediction = mu_auditory * w_auditory + mu_visual * (1 - w_auditory)
ax = posterior_plot(x, auditory, visual, posterior_pointwise)
ax.plot([theoretical_prediction, theoretical_prediction],
[0, posterior_pointwise.max() * 1.2], '-.', color='xkcd:medium gray')
ax.set_title(f"Gray line shows analytical mean of posterior: {theoretical_prediction:0.2f}")
plt.show()
style = {'description_width': 'initial'}
_ = widgets.interact(refresh,
mu_auditory=widgets.FloatSlider(value=2, min=-10, max=10, step=0.5, description="mu_auditory:", style=style),
sigma_auditory=widgets.FloatSlider(value=0.5, min=0.5, max=10, step=0.5, description="sigma_auditory:", style=style),
mu_visual=widgets.FloatSlider(value=-2, min=-10, max=10, step=0.5, description="mu_visual:", style=style),
sigma_visual=widgets.FloatSlider(value=0.5, min=0.5, max=10, step=0.5, description="sigma_visual:", style=style)
)
```
## Video 3: Multiplying Gaussians
```
#@title
from IPython.display import YouTubeVideo
video = YouTubeVideo(id='AbXorOLBrws', width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
```
## Exercise 2B: Finding the posterior analytically
[If you are running short on time, feel free to skip the coding exercise below].
As you may have noticed from the interactive demo, the product of two Gaussian distributions, like our prior and likelihood, remains a Gaussian, regardless of the parameters. We can directly compute the parameters of that Gaussian from the means and variances of the prior and likelihood. For example, the posterior mean is given by:
$$ \mu_{posterior} = \frac{\mu_{auditory} \cdot \frac{1}{\sigma_{auditory}^2} + \mu_{visual} \cdot \frac{1}{\sigma_{visual}^2}}{1/\sigma_{auditory}^2 + 1/\sigma_{visual}^2}
$$
This formula is a special case for two Gaussians, but is a very useful one because:
* The posterior has the same form (here, a normal distribution) as the prior, and
* There is simple, closed-form expression for its parameters.
When these properties hold, we call them **conjugate distributions** or **conjugate priors** (for a particular likelihood). Working with conjugate distributions is very convenient; otherwise, it is often necessary to use computationally-intensive numerical methods to combine the prior and likelihood.
In this exercise, we ask you to verify that property. To do so, we will hold our auditory likelihood constant as an $\mathcal{N}(3, 1.5)$ distribution, while considering visual priors with different means ranging from $\mu=-10$ to $\mu=10$. For each prior,
* Compute the posterior distribution using the function you wrote in Exercise 2A. Next, find its mean. The mean of a probability distribution is $\int_x p(x) dx$ or $\sum_x x\cdot p(x)$.
* Compute the analytical posterior mean from auditory and visual using the equation above.
* Use the provided plotting code to plot both estimates of the mean.
Are the estimates of the posterior mean the same in both cases?
Using these results, try to predict the posterior mean for the combination of a $\mathcal{N}(-4,4)$ prior and and $\mathcal{N}(4, 2)$ likelihood. Use the widget above to check your prediction. You can enter values directly by clicking on the numbers to the right of each slider; $\sqrt{2} \approx 1.41$.
```
def compare_computational_analytical_means():
x = np.arange(-10, 11, 0.1)
# Fixed auditory likelihood
mu_auditory = 3
sigma_auditory = 1.5
likelihood = my_gaussian(x, mu_auditory, sigma_auditory)
# Varying visual prior
mu_visuals = np.linspace(-10, 10)
sigma_visual = 1.5
# Accumulate results here
mus_by_integration = []
mus_analytical = []
for mu_visual in mu_visuals:
prior = my_gaussian(x, mu_visual, sigma_visual)
posterior = compute_posterior_pointwise(prior, likelihood)
############################################################################
## Add code that will find the posterior mean via numerical integration
#
############################################################################
mu_integrated = ...
############################################################################
## Add more code below that will calculate the posterior mean analytically
#
# Comment out the line below to test your solution
raise NotImplementedError("Please add code to find the mean both ways first")
############################################################################
mu_analytical = ...
mus_by_integration.append(mu_integrated)
mus_analytical.append(mu_analytical)
return mu_visuals, mus_analytical, mus_by_integration
# Uncomment the lines below to visualize your results
# mu_visuals, mu_analytical, mu_computational = compare_computational_analytical_means()
# plot_visual(mu_visuals, mu_analytical, mu_computational)
#to_remove solution
def compare_computational_analytical_means():
x = np.arange(-10, 11, 0.1)
# Fixed auditory likelihood
mu_auditory = 3
sigma_auditory = 1.5
likelihood = my_gaussian(x, mu_auditory, sigma_auditory)
# Varying visual prior
mu_visuals = np.linspace(-10, 10)
sigma_visual = 1.5
# Accumulate results here
mus_by_integration = []
mus_analytical = []
for mu_visual in mu_visuals:
prior = my_gaussian(x, mu_visual, sigma_visual)
posterior = compute_posterior_pointwise(prior, likelihood)
############################################################################
## Add code that will find the posterior mean via numerical integration
#
############################################################################
mu_integrated = np.sum(x*posterior)
############################################################################
## Add more code below that will calculate the posterior mean analytically
#
# Comment out the line below to test your solution
#raise NotImplementedError("Please add code to find the mean both ways first")
############################################################################
mu_analytical = ((mu_auditory / sigma_auditory ** 2 + mu_visual / sigma_visual ** 2) /
(1 / sigma_auditory ** 2 + 1 / sigma_visual ** 2))
mus_by_integration.append(mu_integrated)
mus_analytical.append(mu_analytical)
return mu_visuals, mus_analytical, mus_by_integration
# Uncomment the lines below to visualize your results
mu_visuals, mu_analytical, mu_computational = compare_computational_analytical_means()
with plt.xkcd():
plot_visual(mu_visuals, mu_analytical, mu_computational)
```
# Section 3: Conclusion
This tutorial introduced the Gaussian distribution and used Bayes' Theorem to combine Gaussians representing priors and likelihoods. In the next tutorial, we will use these concepts to probe how subjects integrate sensory information.
```
#@title Video 4: Conclusion
from IPython.display import YouTubeVideo
video = YouTubeVideo(id='YC8GylOAAHs', width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
```
# Bonus Section: Multimodal Priors
**Only do this if the first half-hour has not yet passed.**
The preceeding exercises used a Gaussian prior, implying that participants expected the stimulus to come from a single location, though they might not know precisely where. However, suppose the subjects actually thought that sound might come from one of two distinct locations. Perhaps they can see two speakers (and know that speakers often emit noise).
We could model this using a Gaussian prior with a large $\sigma$ that covers both locations, but that would also make every point in between seem likely too.A better approach is to adjust the form of the prior so that it better matches the participants' experiences/expectations. In this optional exercise, we will build a bimodal (2-peaked) prior out of Gaussians and examine the resulting posterior and its peaks.
## Exercise 3: Implement and test a multimodal prior
* Complete the `bimodal_prior` function below to create a bimodal prior, comprised of the sum of two Gaussians with means $\mu = -3$ and $\mu = 3$. Use $\sigma=1$ for both Gaussians. Be sure to normalize the result so it is a proper probability distribution.
* In Exercise 2, we used the mean location to summarize the posterior distribution. This is not always the best choice, especially for multimodal distributions. What is the mean of our new prior? Is it a particularly likely location for the stimulus? Instead, we will use the posterior **mode** to summarize the distribution. The mode is the *location* of the most probable part of the distribution. Complete `posterior_mode` below, to find it. (Hint: `np.argmax` returns the *index* of the largest element in an array).
* Run the provided simulation and plotting code. Observe what happens to the posterior as the likelihood gets closer to the different peaks of the prior.
* Notice what happens to the posterior when the likelihood is exactly in between the two modes of the prior (i.e., $\mu_{Likelihood} = 0$)
```
def bimodal_prior(x, mu_1=-3, sigma_1=1, mu_2=3, sigma_2=1):
################################################################################
## Finish this function so that it returns a bimodal prior, comprised of the
# sum of two Gaussians
#
# Comment out the line below to test out your solution
raise NotImplementedError("Please implement the bimodal prior")
################################################################################
prior = ...
return prior
def posterior_mode(x, posterior):
################################################################################
## Finish this function so that it returns the location of the mode
#
# Comment out the line below to test out your solution
raise NotImplementedError("Please implement the posterior mode")
################################################################################
mode = ...
return mode
def multimodal_simulation(x, mus_visual, sigma_visual=1):
"""
Simulate an experiment where bimodal prior is held constant while
a Gaussian visual likelihood is shifted across locations.
Args:
x: array of points at which prior/likelihood/posterior are evaluated
mus_visual: array of means for the Gaussian likelihood
sigma_visual: scalar standard deviation for the Gaussian likelihood
Returns:
posterior_modes: array containing the posterior mode for each mean in mus_visual
"""
prior = bimodal_prior(x, -3, 1, 3, 1)
posterior_modes = []
for mu in mus_visual:
likelihood = my_gaussian(x, mu, 3)
posterior = compute_posterior_pointwise(prior, likelihood)
p_mode = posterior_mode(x, posterior)
posterior_modes.append(p_mode)
return posterior_modes
x = np.arange(-10, 10, 0.1)
mus = np.arange(-8, 8, 0.05)
# Uncomment the lines below to visualize your results
# posterior_modes = multimodal_simulation(x, mus, 1)
# multimodal_plot(x,
# bimodal_prior(x, -3, 1, 3, 1),
# my_gaussian(x, 1, 1),
# mus, posterior_modes)
#to_remove solution
def bimodal_prior(x, mu_1=-3, sigma_1=1, mu_2=3, sigma_2=1):
################################################################################
## Finish this function so that it returns a bimodal prior, comprised of the
# sum of two Gaussians
#
# Comment out the line below to test out your solution
#raise NotImplementedError("Please implement the bimodal prior")
################################################################################
prior = my_gaussian(x, mu_1, sigma_1) + my_gaussian(x, mu_2, sigma_2)
prior /= prior.sum()
return prior
def posterior_mode(x, posterior):
################################################################################
## Finish this function so that it returns the location of the mode
#
# Comment out the line below to test out your solution
#raise NotImplementedError("Please implement the posterior mode")
################################################################################
mode = x[np.argmax(posterior)]
return mode
def multimodal_simulation(x, mus_visual, sigma_visual=1):
"""
Simulate an experiment where bimodal prior is held constant while
a Gaussian visual likelihood is shifted across locations.
Args:
x: array of points at which prior/likelihood/posterior are evaluated
mus_visual: array of means for the Gaussian likelihood
sigma_visual: scalar standard deviation for the Gaussian likelihood
Returns:
posterior_modes: array containing the posterior mode for each mean in mus_visual
"""
prior = bimodal_prior(x, -3, 1, 3, 1)
posterior_modes = []
for mu in mus_visual:
likelihood = my_gaussian(x, mu, 3)
posterior = compute_posterior_pointwise(prior, likelihood)
p_mode = posterior_mode(x, posterior)
posterior_modes.append(p_mode)
return posterior_modes
x = np.arange(-10, 10, 0.1)
mus = np.arange(-8, 8, 0.05)
# Uncomment the lines below to visualize your results
posterior_modes = multimodal_simulation(x, mus, 1)
with plt.xkcd():
multimodal_plot(x,
bimodal_prior(x, -3, 1, 3, 1),
my_gaussian(x, 1, 1),
mus, posterior_modes)
```
| github_jupyter |
# 1. Run-Length Encoding
See Answer for Lab 04
# 2. Weave 1
```
weave_first_series = [i for i in range(1, 11)]
weave_second_series = [i for i in range(10, 0, -1)]
weave_answer = [1, 10, 2, 9, 3, 8, 4, 7, 5, 6, 6, 5, 7, 4, 8, 3, 9, 2, 10, 1]
def weave(first_series, second_series):
output = []
# We check for a minimum length here so we don't go out of bounds with lists of different sizes
max_range = min(len(first_series), len(second_series))
for i in range(max_range):
output.append(first_series[i])
output.append(second_series[i])
return output
print(weave_answer == weave(weave_first_series, weave_second_series))
```
# 3. Weave 2
```
weave_third_series = [i for i in range(1, 21)]
weave_first_answer = [1, 10, 2, 9, 3, 8, 4, 7, 5, 6, 6, 5, 7, 4, 8, 3, 9, 2, 10, 1]
weave_second_answer = [1, 1, 10, 2, 2, 3, 9, 4, 3, 5, 8, 6, 4, 7, 7, 8, 5, 9, 6, 10, 6, 11, 5, 12, 7, 13, 4, 14, 8, 15, 3, 16, 9, 17, 2, 18, 10, 19, 1, 20]
class Number_Row:
def __init__(self, starting_list = []):
self.current_weave = starting_list
def weave(self, new_list):
new_weave = []
max_range = min(len(self.current_weave), len(new_list))
for i in range(max_range):
new_weave.append(self.current_weave[i])
new_weave.append(new_list[i])
self.current_weave = new_weave
def get_weave(self):
return self.current_weave
nr = Number_Row(weave_first_series)
nr.weave(weave_second_series)
print(weave_first_answer == nr.get_weave())
nr.weave(weave_third_series)
print(weave_second_answer == nr.get_weave())
```
# 4. Body Mass Index
```
# These strings could be taken from user input, a CSV file, an XML file etc.
patient_01_data = "Dean Johnson M 1.78 83"
patient_02_data = "Sophia Miller V 1.69 60"
class Patient:
def __init__(self, name, gender, height, weight):
self.first_name = name[0]
self.last_name = name[-1]
self.gender = gender
self.height = height
self.weight = weight
self.bmi = self.calculate_bmi()
self.status = self.determine_status()
self.title = self.determine_title()
def calculate_bmi(self):
return (float(self.weight) / (float(self.height) ** 2))
def determine_status(self):
return "healthy" if (self.bmi >= 18.5 and self.bmi <= 25) else "unhealthy"
def determine_title(self):
return "Mr." if self.gender == "M" else "Ms." if self.gender == "V" else "Patient"
def process_patient_data(patient_data):
# Python doesn't have the concept of "constants" (values that you cannot change), but in other languages it is customary to make constant values uppercase to indicate their status
# It is generally a good idea to give any "magic numbers" names - the index "-1" means nothing, but "WEIGHT_INDEX" is pretty clear
WEIGHT_INDEX = -1
HEIGHT_INDEX = -2
GENDER_INDEX = -3
NAME_START_INDEX = 0
NAME_END_INDEX = GENDER_INDEX
patient_data = patient_data.split(" ")
name = patient_data[NAME_START_INDEX : NAME_END_INDEX]
gender = patient_data[GENDER_INDEX]
height = patient_data[HEIGHT_INDEX]
weight = patient_data[WEIGHT_INDEX]
return Patient(name, gender, height, weight)
def print_patient_data(patient):
print("{} {}'s bmi is {:.1f} and is {}".format(patient.title, patient.last_name, patient.bmi, patient.status))
print_patient_data(process_patient_data(patient_01_data))
print_patient_data(process_patient_data(patient_02_data))
```
# 5. Body Mass Index 2
```
all_patient_data = ["Dean Johnson M 1.78 83 Yes", "Sophia Miller V 1.69 60 No", "Stacey Williams V 1.52 53 Yes", "Arwa Leon V 1.67 62 Yes", "Agata Rosales V 1.75 62 Yes"]
class Patient:
def __init__(self, name, gender, height, weight, hk):
self.first_name = name[0]
self.last_name = name[-1]
self.gender = gender
self.height = height
self.weight = weight
self.bmi = self.calculate_bmi()
self.status = self.determine_status()
self.title = self.determine_title()
self.hk = hk
def calculate_bmi(self):
return (float(self.weight) / (float(self.height) ** 2))
def determine_status(self):
return "healthy" if (self.bmi >= 18.5 and self.bmi <= 25) else "unhealthy"
def determine_title(self):
return "Mr." if self.gender == "M" else "Ms." if self.gender == "V" else "Patient"
def process_patient_data(patient_data):
HK_INDEX = -1
WEIGHT_INDEX = -2
HEIGHT_INDEX = -3
GENDER_INDEX = -4
NAME_START_INDEX = 0
NAME_END_INDEX = GENDER_INDEX
patient_data = patient_data.split(" ")
name = patient_data[NAME_START_INDEX : NAME_END_INDEX]
gender = patient_data[GENDER_INDEX]
height = patient_data[HEIGHT_INDEX]
weight = patient_data[WEIGHT_INDEX]
hk = True if patient_data[HK_INDEX].lower() == "yes" else False
return Patient(name, gender, height, weight, hk)
def average_bmi(patients):
bmi_sum = 0
for patient in patients:
bmi_sum += patient.bmi
return bmi_sum / len(patients)
def hk_incidence_above_bmi(threshold_bmi, patients):
hk_incidence = 0
for patient in patients:
if patient.hk == True and patient.bmi >= threshold_bmi:
hk_incidence += 1
return hk_incidence
def hk_incidence_below_bmi(threshold_bmi, patients):
hk_incidence = 0
for patient in patients:
if patient.hk == True and patient.bmi < threshold_bmi:
hk_incidence += 1
return hk_incidence
def process_all_patients(patient_data_list):
patient_list = []
for patient in patient_data_list:
patient_list.append(process_patient_data(patient))
threshold_bmi = average_bmi(patient_list)
print("The average BMI of the test subjects is {:.1f}".format(threshold_bmi))
print("There are {} cases of the syndrome amongst people with a BMI >= {:.1f}".format(hk_incidence_above_bmi(threshold_bmi, patient_list), threshold_bmi))
print("There are {} cases of the syndrome amongst people with a BMI < {:.1f}".format(hk_incidence_below_bmi(threshold_bmi, patient_list), threshold_bmi))
process_all_patients(all_patient_data)
```
| github_jupyter |
```
import os, glob
import numpy as np
import pandas as pd
from calendar import monthrange,month_name
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
%matplotlib inline
fs = 18
plt.rc('font', family='serif')
plt.rc('font', size=18)
# date parser for pandas
dp = lambda x: pd.datetime.strptime(x,'%d-%m-%Y %H:%M:%S')
# paths (must mount volume smb://nrel.gov/shared/wind/WindWeb/MetData/135mData/M5Twr)
# metPathLoHz = '/Volumes/M5Twr/10min/txt/'
metPathLoHz = '/Volumes/shared/Wind/WindWeb/MetData/135mData/M5Twr/10min/txt/'
# time range
# year1 = [2012]
# year2 = [2017]
# years = [np.arange(year1,year2+1,1)]
years = [2017]
months = [ int(a) for a in np.arange(1,12.1,1) ]
# read in data
filecount = 0
for year in years:
for month in months:
fName = glob.glob(os.path.join(metPathLoHz,'{0}_{1}.txt'.format(year,month_name[month])))
if len(fName)>0:
fName = fName[0] ; print(fName)
df_lo = pd.read_csv(fName,index_col=[0],parse_dates=[0],date_parser=dp,skiprows=[0,1,2,3,4,5,6,8,9],
low_memory=False)
if filecount==0:
df = df_lo.copy()
else:
df = df.append(df_lo.copy())
filecount += 1
df.index = df.index.tz_localize('UTC').tz_convert('America/Denver')
```
## Implement QC masking on input data (here on 10 minute bins)
Each data field is associated with a QC field <br>
- Extract data: dfField = [date, field, field QC]
- Create mask: mask = field QC == 1
- Filter data: dfFiltField = dfField[mask]
```
temp = [name for name in list(df.columns.values) if ' QC' not in name]
qcNames = [name for name in list(df.columns.values) if ' QC' in name]
fNames = [name for name in temp if name + ' QC' in qcNames]
print('number of data columns:', len(fNames))
print('number of QC columns:', len(qcNames))
# initialize filtered dataframe with 'record', 'version'
dfFilt = df[df.columns.values[[0,1]]].copy()
# apply QC mask to each set of columns individually
for f, q in zip(fNames, qcNames):
# print(fname, qname)
temp = df[[f, q]]
mask = temp[q] == 1
temp = temp[mask]
# dfFilt = pd.concat([dfFilt,temp], axis = 1)
dfFilt[f] = temp[f]
# find and replace missing data (-999.0) with consistent nan values
dfFilt = dfFilt.replace(to_replace=-999.0, value=np.nan)
dfFilt.head()
# Sort data by hour, produces a single diurnal cycle
diurnal_cycle = dfFilt.groupby(dfFilt.index.hour).mean()
diurnal_cycle.columns
monthly_diurnal_cycle = dfFilt.groupby([dfFilt.index.month, dfFilt.index.hour]).mean()
# find all data produced by cup anemometers from the
# spdcols = diurnal_cycle.columns
spdcols = [col for col in diurnal_cycle.columns.values if'Speed' in col and 'cup' in col]
print(spdcols)
diurnal_cycle[spdcols].describe()
ax = diurnal_cycle[spdcols].plot()
plt.legend( loc='center left', bbox_to_anchor=(1,0.5))
plt.xlabel('Day of month')
plt.ylabel('Average velocity (m/s)')
handles, labels = ax.get_legend_handles_labels()
plt.title('Hourly average')
#temp = monthly_diurnal_cycle['Speed (cup_ 80 m)'].xs(month)
#temp.describe()
monthly_diurnal_cycle.index.values
fig = plt.figure(figsize=(12,12))
for month in months:
plt.subplot(4, 3, month)
ax = plt.plot( monthly_diurnal_cycle[spdcols].xs(month))
plt.xlabel('Time of day')
plt.ylabel('Average Velocity')
plt.title(month_name[month])
# handles, labels = ax.get_legend_handles_labels()
fig.legend(handles, labels, loc='center right', bbox_to_anchor=(1.15, 0.5), borderaxespad=0., ncol=1)
fig.tight_layout()
varName = 'd(u)/d(t) (sonic_61m)'
fig = plt.figure(figsize=(6,5))
ax = fig.add_subplot(111)
ax.plot(diurnal_cycle[varName],'-.k')
ax.set_xlabel('Hour of Day')
ax.set_ylabel(varName)
```
| github_jupyter |
<script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
<script>
window.dataLayer = window.dataLayer || [];
function gtag(){dataLayer.push(arguments);}
gtag('js', new Date());
gtag('config', 'UA-59152712-8');
</script>
# Start-to-Finish Example: Head-On Black Hole Collision
## Author: Zach Etienne
### Formatting improvements courtesy Brandon Clark
## This module implements a basic numerical relativity code to merge two black holes in *spherical coordinates*
### Here we place the black holes initially on the $z$-axis, so the entire simulation is axisymmetric about the $\phi$-axis. Not sampling in the $\phi$ direction greatly speeds up the simulation.
**Notebook Status:** <font color = green><b> Validated </b></font>
**Validation Notes:** This module has been validated to exhibit convergence to zero of the Hamiltonian constraint violation at the expected order to the exact solution *after a short numerical evolution of the initial data* (see [plots at bottom](#convergence)), and all quantities have been validated against the [original SENR code](https://bitbucket.org/zach_etienne/nrpy).
### NRPy+ Source Code & Tutorials for this module:
* [BSSN/BrillLindquist.py](../edit/BSSN/BrillLindquist.py); [\[**tutorial**\]](Tutorial-ADM_Initial_Data-Brill-Lindquist.ipynb): Brill-Lindquist initial data; sets all ADM variables in Cartesian basis:
* [BSSN/ADM_Exact_Spherical_or_Cartesian_to_BSSNCurvilinear.py](../edit/BSSN/ADM_Exact_Spherical_or_Cartesian_to_BSSNCurvilinear.py); [\[**tutorial**\]](Tutorial-ADM_Initial_Data-Converting_Exact_ADM_Spherical_or_Cartesian_to_BSSNCurvilinear.ipynb): Spherical/Cartesian ADM$\to$Curvilinear BSSN converter function, for which exact expressions are given for ADM quantities.
* [BSSN/BSSN_ID_function_string.py](../edit/BSSN/BSSN_ID_function_string.py): Sets up the C code string enabling initial data be set up in a point-by-point fashion
* [BSSN/BSSN_Ccodegen_library.py](../edit/BSSN/BSSN_Ccodegen_library.py); [\[**tutorial**\]](Tutorial-BSSN_time_evolution-C_codegen_library.ipynb): Implements a number of helper functions for generating C codes from symbolic expressions generated in the following modules/tutorials:
* [BSSN/BSSN_constraints.py](../edit/BSSN/BSSN_constraints.py); [\[**tutorial**\]](Tutorial-BSSN_constraints.ipynb): Hamiltonian constraint in BSSN curvilinear basis/coordinates
* [BSSN/BSSN_RHSs.py](../edit/BSSN/BSSN_RHSs.py); [\[**tutorial**\]](Tutorial-BSSN_time_evolution-BSSN_RHSs.ipynb): Generates the right-hand sides for the BSSN evolution equations in singular, curvilinear coordinates
* [BSSN/BSSN_gauge_RHSs.py](../edit/BSSN/BSSN_gauge_RHSs.py); [\[**tutorial**\]](Tutorial-BSSN_time_evolution-BSSN_gauge_RHSs.ipynb): Generates the right-hand sides for the BSSN gauge evolution equations in singular, curvilinear coordinates
## Introduction:
Here we use NRPy+ to generate the C source code necessary to set up initial data for two black holes (Brill-Lindquist, [Brill & Lindquist, Phys. Rev. 131, 471, 1963](https://journals.aps.org/pr/abstract/10.1103/PhysRev.131.471); see also Eq. 1 of [Brandt & Brügmann, arXiv:gr-qc/9711015v1](https://arxiv.org/pdf/gr-qc/9711015v1.pdf)). Then we use it to generate the RHS expressions for [Method of Lines](https://reference.wolfram.com/language/tutorial/NDSolveMethodOfLines.html) time integration based on an [explicit Runge-Kutta fourth-order scheme](https://en.wikipedia.org/wiki/Runge%E2%80%93Kutta_methods) (RK4 is chosen below, but multiple options exist).
The entire algorithm is outlined as follows, with links to the relevant NRPy+ tutorial notebooks listed at each step:
1. Allocate memory for gridfunctions, including temporary storage for the Method of Lines time integration
* [**NRPy+ tutorial on Method of Lines algorithm**](Tutorial-Method_of_Lines-C_Code_Generation.ipynb).
1. Set gridfunction values to initial data
* [**NRPy+ tutorial on Brill-Lindquist initial data**](Tutorial-ADM_Initial_Data-Brill-Lindquist.ipynb)
* [**NRPy+ tutorial on validating Brill-Lindquist initial data**](Tutorial-Start_to_Finish-BSSNCurvilinear-Setting_up_Exact_Initial_Data.ipynb).
1. Next, integrate the initial data forward in time using the Method of Lines coupled to a Runge-Kutta explicit timestepping algorithm:
1. At the start of each iteration in time, output the Hamiltonian constraint violation
* [**NRPy+ tutorial on BSSN constraints**](Tutorial-BSSN_constraints.ipynb).
1. At each RK time substep, do the following:
1. Evaluate BSSN RHS expressions
* [**NRPy+ tutorial on BSSN right-hand sides**](Tutorial-BSSN_time_evolution-BSSN_RHSs.ipynb) ([**BSSN Introduction Notebook**](Tutorial-BSSN_formulation.ipynb))
* [**NRPy+ tutorial on BSSN gauge condition right-hand sides**](Tutorial-BSSN_time_evolution-BSSN_gauge_RHSs.ipynb)
1. Apply singular, curvilinear coordinate boundary conditions [*a la* the SENR/NRPy+ paper](https://arxiv.org/abs/1712.07658)
* [**NRPy+ tutorial on setting up singular, curvilinear boundary conditions**](Tutorial-Start_to_Finish-Curvilinear_BCs.ipynb)
1. Enforce constraint on conformal 3-metric: $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$
* [**NRPy+ tutorial on enforcing $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$ constraint**](Tutorial-BSSN_enforcing_determinant_gammabar_equals_gammahat_constraint.ipynb)
1. Repeat above steps at two numerical resolutions to confirm convergence to zero.
<a id='toc'></a>
# Table of Contents
$$\label{toc}$$
This notebook is organized as follows
1. [Step 1](#initializenrpy): Set core NRPy+ parameters for numerical grids and reference metric
1. [Step 2](#ccodegen): Generate C code kernels for BSSN expressions, in parallel if possible
1. [Step 2.a](#rfm_ccodegen): Generate C code kernels for reference metric
1. [Step 3](#cparams_rfm_and_domainsize): Set `free_parameters.h`; also output C codes needed for declaring and setting Cparameters
1. [Step 4](#bc_functs): Set up boundary condition functions for chosen singular, curvilinear coordinate system
1. [Step 5](#mainc): `BrillLindquist_Playground`: The C code `main()` function
1. [Step 6](#compileexec): Compile generated C codes & perform the black hole collision calculation
1. [Step 7](#visualize): Visualize the output!
1. [Step 7.a](#installdownload): Install `scipy` and download `ffmpeg` if they are not yet installed/downloaded
1. [Step 7.b](#genimages): Generate images for visualization animation
1. [Step 7.c](#genvideo): Generate visualization animation
1. [Step 8](#convergence): Plot the numerical error at the end of the simulation, and confirm that it converges to zero with increasing numerical resolution (sampling)
1. [Step 9](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
<a id='initializenrpy'></a>
# Step 1: Set core NRPy+ parameters for numerical grids and reference metric \[Back to [top](#toc)\]
$$\label{initializenrpy}$$
```
# Step P1: Import needed NRPy+ core modules:
from outputC import lhrh, outCfunction, outC_function_dict, add_to_Cfunction_dict # NRPy+: Core C code output module
import finite_difference as fin # NRPy+: Finite difference C code generation module
import NRPy_param_funcs as par # NRPy+: Parameter interface
import grid as gri # NRPy+: Functions having to do with numerical grids
import reference_metric as rfm # NRPy+: Reference metric support
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
from pickling import pickle_NRPy_env, unpickle_NRPy_env # NRPy+: Pickle/unpickle NRPy+ environment, for parallel codegen
import shutil, os, sys, time # Standard Python modules for multiplatform OS-level functions, benchmarking
# Step P2: Create C code output directory:
Ccodesrootdir = os.path.join("BSSN_Two_BHs_Collide_Ccodes_new_way")
# First remove C code output directory if it exists
# Courtesy https://stackoverflow.com/questions/303200/how-do-i-remove-delete-a-folder-that-is-not-empty
# !rm -r ScalarWaveCurvilinear_Playground_Ccodes
shutil.rmtree(Ccodesrootdir, ignore_errors=True)
# Then create a fresh directory
cmd.mkdir(Ccodesrootdir)
# Step P3: Create executable output directory:
outdir = os.path.join(Ccodesrootdir, "output")
cmd.mkdir(outdir)
# Step 1.a: Enable SIMD-optimized code?
# I.e., generate BSSN and Ricci C code kernels using SIMD-vectorized
# compiler intrinsics, which *greatly improve the code's performance*,
# though at the expense of making the C-code kernels less
# human-readable.
# * Important note in case you wish to modify the BSSN/Ricci kernels
# here by adding expressions containing transcendental functions
# (e.g., certain scalar fields):
# Note that SIMD-based transcendental function intrinsics are not
# supported by the default installation of gcc or clang (you will
# need to use e.g., the SLEEF library from sleef.org, for this
# purpose). The Intel compiler suite does support these intrinsics
# however without the need for external libraries.
enable_SIMD = True
# Step 1.b: Enable reference metric precomputation.
enable_rfm_precompute = True
if enable_SIMD and not enable_rfm_precompute:
print("ERROR: SIMD does not currently handle transcendental functions,\n")
print(" like those found in rfmstruct (rfm_precompute).\n")
print(" Therefore, enable_SIMD==True and enable_rfm_precompute==False\n")
print(" is not supported.\n")
sys.exit(1)
# Step 1.c: Enable "FD functions". In other words, all finite-difference stencils
# will be output as inlined static functions. This is essential for
# compiling highly complex FD kernels with using certain versions of GCC;
# GCC 10-ish will choke on BSSN FD kernels at high FD order, sometimes
# taking *hours* to compile. Unaffected GCC versions compile these kernels
# in seconds. FD functions do not slow the code performance, but do add
# another header file to the C source tree.
# With gcc 7.5.0, enable_FD_functions=True decreases performance by 10%
enable_FD_functions = False
# Step 2: Set some core parameters, including CoordSystem MoL timestepping algorithm,
# FD order, floating point precision, and CFL factor:
# Choices are: Spherical, SinhSpherical, SinhSphericalv2, Cylindrical, SinhCylindrical,
# SymTP, SinhSymTP
CoordSystem = "Spherical"
par.set_parval_from_str("reference_metric::CoordSystem", CoordSystem)
rfm.reference_metric()
# Step 2.a: Set defaults for Coordinate system parameters.
# These are perhaps the most commonly adjusted parameters,
# so we enable modifications at this high level.
# domain_size sets the default value for:
# * Spherical's params.RMAX
# * SinhSpherical*'s params.AMAX
# * Cartesians*'s -params.{x,y,z}min & .{x,y,z}max
# * Cylindrical's -params.ZMIN & .{Z,RHO}MAX
# * SinhCylindrical's params.AMPL{RHO,Z}
# * *SymTP's params.AMAX
domain_size = 7.5 # Needed for all coordinate systems.
# sinh_width sets the default value for:
# * SinhSpherical's params.SINHW
# * SinhCylindrical's params.SINHW{RHO,Z}
# * SinhSymTP's params.SINHWAA
sinh_width = 0.4 # If Sinh* coordinates chosen
# sinhv2_const_dr sets the default value for:
# * SinhSphericalv2's params.const_dr
# * SinhCylindricalv2's params.const_d{rho,z}
sinhv2_const_dr = 0.05 # If Sinh*v2 coordinates chosen
# SymTP_bScale sets the default value for:
# * SinhSymTP's params.bScale
SymTP_bScale = 0.5 # If SymTP chosen
# Step 2.b: Set the order of spatial and temporal derivatives;
# the core data type, and the CFL factor.
# RK_method choices include: Euler, "RK2 Heun", "RK2 MP", "RK2 Ralston", RK3, "RK3 Heun", "RK3 Ralston",
# SSPRK3, RK4, DP5, DP5alt, CK5, DP6, L6, DP8
RK_method = "RK4"
FD_order = 4 # Finite difference order: even numbers only, starting with 2. 12 is generally unstable
REAL = "double" # Best to use double here.
default_CFL_FACTOR= 0.5 # (GETS OVERWRITTEN IF SPECIFIED AT COMMAND LINE.)
# In pure axisymmetry (symmetry_axes = 2 below) 1.0 works fine. Otherwise 0.5 or lower.
# Step 5: Set the finite differencing order to FD_order (set above).
par.set_parval_from_str("finite_difference::FD_CENTDERIVS_ORDER", FD_order)
# Directory for reference_metric precomputation header files:
rfm_precompute_Ccode_outdir = os.path.join(Ccodesrootdir, "rfm_files/")
if enable_rfm_precompute:
cmd.mkdir(os.path.join(Ccodesrootdir, "rfm_files/"))
par.set_parval_from_str("reference_metric::rfm_precompute_Ccode_outdir", rfm_precompute_Ccode_outdir)
# Step 6: Copy SIMD/SIMD_intrinsics.h to $Ccodesrootdir/SIMD/SIMD_intrinsics.h
if enable_SIMD:
cmd.mkdir(os.path.join(Ccodesrootdir,"SIMD"))
shutil.copy(os.path.join("SIMD/")+"SIMD_intrinsics.h",os.path.join(Ccodesrootdir,"SIMD/"))
# Step 7: Set finite_difference::enable_FD_functions appropriately. Defaults to False
if enable_FD_functions:
par.set_parval_from_str("finite_difference::enable_FD_functions", enable_FD_functions)
# Step 8: If enable_SIMD, then copy SIMD/SIMD_intrinsics.h to $Ccodesrootdir/SIMD/SIMD_intrinsics.h
cmd.mkdir(os.path.join(Ccodesrootdir,"SIMD"))
if enable_SIMD:
shutil.copy(os.path.join("SIMD", "SIMD_intrinsics.h"), os.path.join(Ccodesrootdir, "SIMD"))
# Step 9: Set the direction=2 (phi) axis to be the symmetry axis; i.e.,
# axis "2", corresponding to the i2 direction.
# This sets all spatial derivatives in the phi direction to zero.
par.set_parval_from_str("indexedexp::symmetry_axes","2")
OMP_pragma_on = "i1" # structure OpenMP loops to parallelize, not over i2 (phi direction), but i1 (theta direction)
# Step 10: Generate Runge-Kutta-based (RK-based) timestepping code.
# As described above the Table of Contents, this is a 3-step process:
# 3.A: Evaluate RHSs (RHS_string)
# 3.B: Apply boundary conditions (post_RHS_string, pt 1)
# 3.C: Enforce det(gammabar) = det(gammahat) constraint (post_RHS_string, pt 2)
import MoLtimestepping.MoL_new_way as MoL
# from MoLtimestepping.RK_Butcher_Table_Dictionary import Butcher_dict
# RK_order = Butcher_dict[RK_method][1]
RHS_string = """
Ricci_eval(params, rfmstruct, RK_INPUT_GFS, auxevol_gfs);
rhs_eval(params, rfmstruct, auxevol_gfs, RK_INPUT_GFS, RK_OUTPUT_GFS);"""
post_RHS_string = """
apply_bcs_curvilinear(params, bcstruct, NUM_EVOL_GFS, evol_gf_parity, RK_OUTPUT_GFS);
enforce_detgammahat_constraint(params, rfmstruct, RK_OUTPUT_GFS);\n"""
if not enable_rfm_precompute:
RHS_string = RHS_string.replace("rfmstruct", "xx")
post_RHS_string = post_RHS_string.replace("rfmstruct", "xx")
MoL.register_C_functions_and_NRPy_basic_defines(RK_method,
RHS_string=RHS_string, post_RHS_string=post_RHS_string,
enable_rfm=enable_rfm_precompute, enable_curviBCs=True)
```
<a id='ccodegen'></a>
# Step 2: Generate C code kernels for BSSN expressions, in parallel if possible \[Back to [top](#toc)\]
$$\label{ccodegen}$$
In the following code cell, we create a list of Python functions, which each registers a single C code function in `outputC`'s `outC_function_dict` dictionary. These Python functions are defined in
1. the [`BSSN.BrillLindquist`](../edit/BSSN/BrillLindquist.py) NRPy+ module, which does the following:
1. Sets up Brill-Lindquist initial data [ADM](https://en.wikipedia.org/wiki/ADM_formalism) quantities in the **Cartesian basis**, as [documented here](Tutorial-ADM_Initial_Data-Brill-Lindquist.ipynb).
1. Converts the ADM **Cartesian quantities** to **BSSN quantities in the desired Curvilinear basis** (set by reference_metric::CoordSystem), as [documented here](Tutorial-ADM_Initial_Data-Converting_ADMCartesian_to_BSSNCurvilinear.ipynb).
1. Sets up the standardized C function for setting all BSSN Curvilinear gridfunctions in a pointwise fashion, as [written here](../edit/BSSN/BSSN_ID_function_string.py), and returns the C function as a Python string.
1. the [`BSSN.BSSN_Ccodegen_library`](../edit/BSSN/BSSN_Ccodegen_library.py) NRPy+ module [\[**tutorial**\]](Tutorial-BSSN_time_evolution-C_codegen_library.ipynb), which contains Python functions for generating C code from symbolic expressions constructed within the following NRPy+ modules/tutorials:
1. [BSSN/BSSN_constraints.py](../edit/BSSN/BSSN_constraints.py); [\[**tutorial**\]](Tutorial-BSSN_constraints.ipynb): Hamiltonian constraint in BSSN curvilinear basis/coordinates
1. [BSSN/BSSN_RHSs.py](../edit/BSSN/BSSN_RHSs.py); [\[**tutorial**\]](Tutorial-BSSN_time_evolution-BSSN_RHSs.ipynb): Generates the right-hand sides for the BSSN evolution equations in singular, curvilinear coordinates
1. [BSSN/BSSN_gauge_RHSs.py](../edit/BSSN/BSSN_gauge_RHSs.py); [\[**tutorial**\]](Tutorial-BSSN_time_evolution-BSSN_gauge_RHSs.ipynb): Generates the right-hand sides for the BSSN gauge evolution equations in singular, curvilinear coordinates
1. [BSSN/Enforce_Detgammahat_Constraint.py](../edit/BSSN/Enforce_Detgammahat_Constraint.py); [**tutorial**](Tutorial-BSSN_enforcing_determinant_gammabar_equals_gammahat_constraint.ipynb): Generates symbolic expressions for enforcing the $\det{\bar{\gamma}}=\det{\hat{\gamma}}$ constraint
Next, from within a `multiprocessing` environment, we then call all the Python C-code generation functions in this list in parallel (if `multiprocessing` is supported). This is quite useful, as these functions take several seconds to complete.
Within each `multiprocessing` process, the current NRPy+ environment is cloned, and a new function is registered to the `outC_function_dict` dictionary. Thus when each process completes, it contains a unique NRPy+ environment, with only its function registered. We address this by saving each process' NRPy+ environment and sending it back in a common binary format known as a `pickle`, using NRPy+'s [`pickling`](../edit/pickling.py) module. The environments are combined in an unpickling such that all functions exist within the same `outC_function_dict` dictionary.
To make the current environment fully consistent, we call `reference_metric.py` to register all its associated C functions (stored in globals) and contributions to `NRPy_basic_defines.h`.
```
# Step 2: Generate C code kernels for BSSN expressions, in parallel if possible;
import BSSN.BSSN_Ccodegen_library as BCL
import BSSN.BrillLindquist as bl
# Step 2.a: Create a list of functions we wish to evaluate in parallel (if possible)
# Create lists for all BSSN functions
BSSN_funcs = [bl.BrillLindquist]
BSSN_funcs.append(BCL.add_rhs_eval_to_Cfunction_dict)
BSSN_funcs.append(BCL.add_Ricci_eval_to_Cfunction_dict)
BSSN_funcs.append(BCL.add_BSSN_constraints_to_Cfunction_dict)
BSSN_funcs.append(BCL.add_enforce_detgammahat_constraint_to_Cfunction_dict)
# Step 2.b: Define master functions for parallelization.
# Note that lambdifying this doesn't work in Python 3
def master_func(arg):
if BSSN_funcs[arg] == bl.BrillLindquist:
ret = BSSN_funcs[arg](include_NRPy_basic_defines_and_pickle=True)
else:
if enable_rfm_precompute:
# We use rfm_precompute for all BSSN functions:
par.set_parval_from_str("reference_metric::enable_rfm_precompute", "True")
rfm.reference_metric()
if BSSN_funcs[arg].__name__ == "add_BSSN_constraints_to_Cfunction_dict":
ret = BSSN_funcs[arg](includes=["NRPy_basic_defines.h"],
rel_path_to_Cparams=os.path.join("."), output_H_only=True,
enable_rfm_precompute=enable_rfm_precompute, enable_SIMD=enable_SIMD,
OMP_pragma_on=OMP_pragma_on)
elif BSSN_funcs[arg].__name__ == "add_rhs_eval_to_Cfunction_dict" or \
BSSN_funcs[arg].__name__ == "add_Ricci_eval_to_Cfunction_dict":
ret = BSSN_funcs[arg](includes=["NRPy_basic_defines.h"],
rel_path_to_Cparams=os.path.join("."),
enable_rfm_precompute=enable_rfm_precompute, enable_SIMD=enable_SIMD,
OMP_pragma_on=OMP_pragma_on)
elif BSSN_funcs[arg].__name__ == "add_enforce_detgammahat_constraint_to_Cfunction_dict":
ret = BSSN_funcs[arg](includes=["NRPy_basic_defines.h"],
rel_path_to_Cparams=os.path.join("."),
enable_rfm_precompute=enable_rfm_precompute, OMP_pragma_on=OMP_pragma_on)
else:
print("ERROR: DID NOT RECOGNIZE FUNCTION " + BSSN_funcs[arg].__name__ + "\n")
sys.exit(1)
if enable_rfm_precompute:
par.set_parval_from_str("reference_metric::enable_rfm_precompute", "False")
rfm.ref_metric__hatted_quantities()
return ret
NRPyEnvVars = []
raised_exception = False
try:
if os.name == 'nt':
# It's a mess to get working in Windows, so we don't bother. :/
# https://medium.com/@grvsinghal/speed-up-your-python-code-using-multiprocessing-on-windows-and-jupyter-or-ipython-2714b49d6fac
raise Exception("Parallel codegen currently not available in certain environments, e.g., Windows")
# Step 2.d: Import the multiprocessing module.
import multiprocessing
# Step 2.e: Evaluate list of functions in parallel if possible;
# otherwise fallback to serial evaluation:
pool = multiprocessing.Pool()
NRPyEnvVars.append(pool.map(master_func, range(len(BSSN_funcs))))
pool.terminate()
pool.join()
except:
print("FAILED PARALLEL CODEGEN!")
NRPyEnvVars = [] # Reset, as pickling/unpickling unnecessary for serial codegen (see next line)
# Steps 2.d-e, alternate: As fallback, evaluate functions in serial.
# This will happen on Android and Windows systems
for i, func in enumerate(BSSN_funcs):
master_func(i)
raised_exception = True
if not raised_exception:
unpickle_NRPy_env(NRPyEnvVars)
```
<a id='rfm_ccodegen'></a>
## Step 2.a: Generate C code kernels for reference metric \[Back to [top](#toc)\]
$$\label{rfm_ccodegen}$$
In the [reference_metric](../edit/reference_metric.py) NRPy+ module, `register_C_functions_and_NRPy_basic_defines()` registers the following C functions to `outC_Cfunction_dict`:
1. `find_timestep()`: Finds the minimum spacing between adjacent gridpoints on our numerical grid $\min(ds_i)$, and sets the timestep according to the [CFL](https://en.wikipedia.org/w/index.php?title=Courant%E2%80%93Friedrichs%E2%80%93Lewy_condition&oldid=806430673) condition: $\Delta t \le \frac{\min(ds_i)}{c}$, where $c$ is the wavespeed, and $ds_i = h_i \Delta x^i$ is the proper distance between neighboring gridpoints in the $i$th direction (in 3D, there are 3 directions), $h_i$ is the $i$th reference metric scale factor, and $\Delta x^i$ is the uniform grid spacing in the $i$th direction.
1. `xx_to_Cart()`: Input = uniformly sampled coordinate xx0,xx1,xx2 (e.g., r,theta,phi in Spherical coordinates). Output = Cartesian coordinate (x,y,z).
1. `set_Nxx_dxx_invdx_params__and__xx()`: Sets `Nxx{0,1,2}`, `Nxx_plus_2NGHOSTS{0,1,2}`, `dxx{0,1,2}`, and `invdx{0,1,2}`; and defines `xx[3][]`.
1. `Cart_to_xx_and_nearest_i0i1i2()`: Input = Cartesian coordinate (x,y,z). Output = uniformly sampled coordinate xx0,xx1,xx2 (e.g., r,theta,phi in Spherical coordinates), as well as corresponding grid index `i0,i1,i2`.
```
# Generate & register C function set_Nxx_dxx_invdx_params__and__xx()
# Generate & register C function xx_to_Cart() for
# (the mapping from xx->Cartesian) for the chosen
# CoordSystem:
# Generate & register the find_timestep() function
# Sets reference_metric globals: NRPy_basic_defines_str, rfm_struct__malloc, rfm_struct__define, rfm_struct__freemem
if enable_rfm_precompute:
par.set_parval_from_str("reference_metric::rfm_precompute_Ccode_outdir", rfm_precompute_Ccode_outdir)
par.set_parval_from_str("reference_metric::enable_rfm_precompute", "True")
par.set_parval_from_str("reference_metric::rfm_precompute_to_Cfunctions_and_NRPy_basic_defines", "True")
rfm.reference_metric()
rfm.register_C_functions_and_NRPy_basic_defines(enable_rfm_precompute=enable_rfm_precompute,
use_unit_wavespeed_for_find_timestep=True)
if enable_rfm_precompute:
par.set_parval_from_str("reference_metric::enable_rfm_precompute", "False")
rfm.ref_metric__hatted_quantities()
```
<a id='cparams_rfm_and_domainsize'></a>
# Step 3: Set `free_parameters.h`; also output C codes needed for declaring and setting Cparameters \[Back to [top](#toc)\]
$$\label{cparams_rfm_and_domainsize}$$
First we output `free_parameters.h`, which sets initial data parameters, as well as grid domain & reference metric parameters, applying `domain_size` and `sinh_width`/`SymTP_bScale` (if applicable) as set above.
```
# Step 3.e.i: Set free_parameters.h
with open(os.path.join(Ccodesrootdir,"free_parameters.h"),"w") as file:
file.write("""
// Set free-parameter values.
// Set the default CFL Factor. Can be overwritten at command line.
REAL CFL_FACTOR = """+str(default_CFL_FACTOR)+r""";
// Set free-parameter values for BSSN evolution:
params.eta = 1.0;
// Set free parameters for the (Brill-Lindquist) initial data
params.BH1_posn_x = 0.0; params.BH1_posn_y = 0.0; params.BH1_posn_z =+0.5;
params.BH2_posn_x = 0.0; params.BH2_posn_y = 0.0; params.BH2_posn_z =-0.5;
params.BH1_mass = 0.5; params.BH2_mass = 0.5;
""")
# Append to $Ccodesrootdir/free_parameters.h reference metric parameters based on generic
# domain_size,sinh_width,sinhv2_const_dr,SymTP_bScale,
# parameters set above.
rfm.out_default_free_parameters_for_rfm(os.path.join(Ccodesrootdir,"free_parameters.h"),
domain_size,sinh_width,sinhv2_const_dr,SymTP_bScale)
```
<a id='bc_functs'></a>
# Step 4: Set up boundary condition functions for chosen singular, curvilinear coordinate system \[Back to [top](#toc)\]
$$\label{bc_functs}$$
Next apply singular, curvilinear coordinate boundary conditions [as documented in the corresponding NRPy+ tutorial notebook](Tutorial-Start_to_Finish-Curvilinear_BCs_new_way.ipynb)
```
import CurviBoundaryConditions.CurviBoundaryConditions_new_way as CBC
CBC.CurviBoundaryConditions_register_C_functions_and_NRPy_basic_defines()
```
<a id='mainc'></a>
# Step 5: The C code `main()` function for `BrillLindquist_Playground` \[Back to [top](#toc)\]
$$\label{mainc}$$
```
def add_to_Cfunction_dict_main__BrillLindquist_Playground():
includes = ["NRPy_basic_defines.h", "NRPy_function_prototypes.h", "time.h"]
desc = """// main() function:
// Step 0: Read command-line input, set up grid structure, allocate memory for gridfunctions, set up coordinates
// Step 1: Set up initial data to an exact solution
// Step 2: Start the timer, for keeping track of how fast the simulation is progressing.
// Step 3: Integrate the initial data forward in time using the chosen RK-like Method of
// Lines timestepping algorithm, and output periodic simulation diagnostics
// Step 3.a: Output 2D data file periodically, for visualization
// Step 3.b: Step forward one timestep (t -> t+dt) in time using
// chosen RK-like MoL timestepping algorithm
// Step 3.c: If t=t_final, output conformal factor & Hamiltonian
// constraint violation to 2D data file
// Step 3.d: Progress indicator printing to stderr
// Step 4: Free all allocated memory
"""
c_type = "int"
name = "main"
params = "int argc, const char *argv[]"
body = r"""
paramstruct params;
set_Cparameters_to_default(¶ms);
// Step 0a: Set free parameters, overwriting Cparameters defaults
// by hand or with command-line input, as desired.
#include "free_parameters.h"
// Step 0b: Read command-line input, error out if nonconformant
if((argc != 4 && argc != 5) || atoi(argv[1]) < NGHOSTS || atoi(argv[2]) < NGHOSTS || atoi(argv[3]) < 2 /* FIXME; allow for axisymmetric sims */) {
fprintf(stderr,"Error: Expected three command-line arguments: ./BrillLindquist_Playground Nx0 Nx1 Nx2,\n");
fprintf(stderr,"where Nx[0,1,2] is the number of grid points in the 0, 1, and 2 directions.\n");
fprintf(stderr,"Nx[] MUST BE larger than NGHOSTS (= %d)\n",NGHOSTS);
exit(1);
}
if(argc == 5) {
CFL_FACTOR = strtod(argv[4],NULL);
if(CFL_FACTOR > 0.5 && atoi(argv[3])!=2) {
fprintf(stderr,"WARNING: CFL_FACTOR was set to %e, which is > 0.5.\n",CFL_FACTOR);
fprintf(stderr," This will generally only be stable if the simulation is purely axisymmetric\n");
fprintf(stderr," However, Nx2 was set to %d>2, which implies a non-axisymmetric simulation\n",atoi(argv[3]));
}
}
// Step 0c: Set up numerical grid structure, first in space...
const int Nxx[3] = { atoi(argv[1]), atoi(argv[2]), atoi(argv[3]) };
if(Nxx[0]%2 != 0 || Nxx[1]%2 != 0 || Nxx[2]%2 != 0) {
fprintf(stderr,"Error: Cannot guarantee a proper cell-centered grid if number of grid cells not set to even number.\n");
fprintf(stderr," For example, in case of angular directions, proper symmetry zones will not exist.\n");
exit(1);
}
// Step 0d: Uniform coordinate grids are stored to *xx[3]
REAL *xx[3];
// Step 0d.i: Set bcstruct
bc_struct bcstruct;
{
int EigenCoord = 1;
// Step 0d.ii: Call set_Nxx_dxx_invdx_params__and__xx(), which sets
// params Nxx,Nxx_plus_2NGHOSTS,dxx,invdx, and xx[] for the
// chosen Eigen-CoordSystem.
set_Nxx_dxx_invdx_params__and__xx(EigenCoord, Nxx, ¶ms, xx);
// Step 0e: Find ghostzone mappings; set up bcstruct
driver_bcstruct(¶ms, &bcstruct, xx);
// Step 0e.i: Free allocated space for xx[][] array
for(int i=0;i<3;i++) free(xx[i]);
}
// Step 0f: Call set_Nxx_dxx_invdx_params__and__xx(), which sets
// params Nxx,Nxx_plus_2NGHOSTS,dxx,invdx, and xx[] for the
// chosen (non-Eigen) CoordSystem.
int EigenCoord = 0;
set_Nxx_dxx_invdx_params__and__xx(EigenCoord, Nxx, ¶ms, xx);
// Step 0g: Set all C parameters "blah" for params.blah, including
// Nxx_plus_2NGHOSTS0 = params.Nxx_plus_2NGHOSTS0, etc.
#include "set_Cparameters-nopointer.h"
// Step 0h: Time coordinate parameters
const REAL t_final = domain_size; /* Final time is set so that at t=t_final,
* data at the origin have not been corrupted
* by the approximate outer boundary condition */
// Step 0i: Set timestep based on smallest proper distance between gridpoints and CFL factor
REAL dt = find_timestep(¶ms, xx, CFL_FACTOR);
//fprintf(stderr,"# Timestep set to = %e\n",(double)dt);
int N_final = (int)(t_final / dt + 0.5); // The number of points in time.
// Add 0.5 to account for C rounding down
// typecasts to integers.
int output_every_N = (int)((REAL)N_final/800.0);
if(output_every_N == 0) output_every_N = 1;
// Step 0j: Error out if the number of auxiliary gridfunctions outnumber evolved gridfunctions.
// This is a limitation of the RK method. You are always welcome to declare & allocate
// additional gridfunctions by hand.
if(NUM_AUX_GFS > NUM_EVOL_GFS) {
fprintf(stderr,"Error: NUM_AUX_GFS > NUM_EVOL_GFS. Either reduce the number of auxiliary gridfunctions,\n");
fprintf(stderr," or allocate (malloc) by hand storage for *diagnostic_output_gfs. \n");
exit(1);
}
// Step 0k: Declare struct for gridfunctions and allocate memory for y_n_gfs gridfunctions
MoL_gridfunctions_struct gridfuncs;
MoL_malloc_y_n_gfs(¶ms, &gridfuncs);
"""
if enable_rfm_precompute:
body += """
// Step 0l: Set up precomputed reference metric arrays
// Step 0l.i: Allocate space for precomputed reference metric arrays.
rfm_struct rfmstruct;
rfm_precompute_rfmstruct_malloc(¶ms, &rfmstruct);
// Step 0l.ii: Define precomputed reference metric arrays.
rfm_precompute_rfmstruct_define(¶ms, xx, &rfmstruct);\n"""
body += r"""
// Step 1: Set up initial data to an exact solution
initial_data(¶ms, xx, gridfuncs.y_n_gfs);
// Step 1a: Allocate memory for non-y_n_gfs gridfunctions
MoL_malloc_non_y_n_gfs(¶ms, &gridfuncs);
// Step 1b: Apply boundary conditions, as initial data
// are sometimes ill-defined in ghost zones.
// E.g., spherical initial data might not be
// properly defined at points where r=-1.
apply_bcs_curvilinear(¶ms, &bcstruct, NUM_EVOL_GFS,evol_gf_parity, gridfuncs.y_n_gfs);
enforce_detgammahat_constraint(¶ms, &rfmstruct, gridfuncs.y_n_gfs);
// Step 2: Start the timer, for keeping track of how fast the simulation is progressing.
#ifdef __linux__ // Use high-precision timer in Linux.
struct timespec start, end;
clock_gettime(CLOCK_REALTIME, &start);
#else // Resort to low-resolution, standards-compliant timer in non-Linux OSs
// http://www.cplusplus.com/reference/ctime/time/
time_t start_timer,end_timer;
time(&start_timer); // Resolution of one second...
#endif
// Step 3: Integrate the initial data forward in time using the chosen RK-like Method of
// Lines timestepping algorithm, and output periodic simulation diagnostics
for(int n=0;n<=N_final;n++) { // Main loop to progress forward in time.
// Step 3.a: Output 2D data file periodically, for visualization
if(n%100 == 0) {
// Evaluate BSSN constraints (currently only Hamiltonian constraint violation computed).
Ricci_eval(¶ms, &rfmstruct, gridfuncs.y_n_gfs, gridfuncs.auxevol_gfs); // <- depends on Ricci.
BSSN_constraints(¶ms, &rfmstruct, gridfuncs.y_n_gfs, gridfuncs.auxevol_gfs, gridfuncs.diagnostic_output_gfs);
char filename[100]; sprintf(filename,"out%d-%08d.txt",Nxx[0],n);
FILE *out2D = fopen(filename, "w");
LOOP_REGION(NGHOSTS,Nxx_plus_2NGHOSTS0-NGHOSTS,
NGHOSTS,Nxx_plus_2NGHOSTS1-NGHOSTS,
NGHOSTS,Nxx_plus_2NGHOSTS2-NGHOSTS) {
const int idx = IDX3S(i0,i1,i2);
REAL xCart[3]; xx_to_Cart(¶ms,xx,i0,i1,i2, xCart);
fprintf(out2D,"%e %e %e %e\n",
xCart[1],xCart[2],
gridfuncs.y_n_gfs[IDX4ptS(CFGF,idx)],log10(fabs(gridfuncs.diagnostic_output_gfs[IDX4ptS(HGF,idx)])));
}
fclose(out2D);
}
// Step 3.b: Step forward one timestep (t -> t+dt) in time using
// chosen RK-like MoL timestepping algorithm
MoL_step_forward_in_time(¶ms, &rfmstruct, &bcstruct, &gridfuncs, dt);
// Step 3.c: If t=t_final, output conformal factor & Hamiltonian
// constraint violation to 2D data file
if(n==N_final-1) {
// Evaluate BSSN constraints (currently only Hamiltonian constraint violation computed).
Ricci_eval(¶ms, &rfmstruct, gridfuncs.y_n_gfs, gridfuncs.auxevol_gfs); // <- depends on Ricci.
BSSN_constraints(¶ms, &rfmstruct, gridfuncs.y_n_gfs, gridfuncs.auxevol_gfs, gridfuncs.diagnostic_output_gfs);
char filename[100]; sprintf(filename,"out%d.txt",Nxx[0]);
FILE *out2D = fopen(filename, "w");
LOOP_REGION(NGHOSTS,Nxx_plus_2NGHOSTS0-NGHOSTS,
NGHOSTS,Nxx_plus_2NGHOSTS1-NGHOSTS,
NGHOSTS,Nxx_plus_2NGHOSTS2-NGHOSTS) {
int idx = IDX3S(i0,i1,i2);
REAL xCart[3]; xx_to_Cart(¶ms,xx,i0,i1,i2, xCart);
fprintf(out2D,"%e %e %e %e\n",
xCart[1],xCart[2],
gridfuncs.y_n_gfs[IDX4ptS(CFGF,idx)],log10(fabs(gridfuncs.diagnostic_output_gfs[IDX4ptS(HGF,idx)])));
}
fclose(out2D);
}
// Step 3.d: Progress indicator printing to stderr
// Step 3.d.i: Measure average time per iteration
#ifdef __linux__ // Use high-precision timer in Linux.
clock_gettime(CLOCK_REALTIME, &end);
const long long unsigned int time_in_ns = 1000000000L * (end.tv_sec - start.tv_sec) + end.tv_nsec - start.tv_nsec;
#else // Resort to low-resolution, standards-compliant timer in non-Linux OSs
time(&end_timer); // Resolution of one second...
REAL time_in_ns = difftime(end_timer,start_timer)*1.0e9+0.5; // Round up to avoid divide-by-zero.
#endif
const REAL s_per_iteration_avg = ((REAL)time_in_ns / (REAL)n) / 1.0e9;
const int iterations_remaining = N_final - n;
const REAL time_remaining_in_mins = s_per_iteration_avg * (REAL)iterations_remaining / 60.0;
const REAL num_RHS_pt_evals = (REAL)(Nxx[0]*Nxx[1]*Nxx[2]) * 4.0 * (REAL)n; // 4 RHS evals per gridpoint for RK4
const REAL RHS_pt_evals_per_sec = num_RHS_pt_evals / ((REAL)time_in_ns / 1.0e9);
// Step 3.d.ii: Output simulation progress to stderr
if(n % 10 == 0) {
fprintf(stderr,"%c[2K", 27); // Clear the line
fprintf(stderr,"It: %d t=%.2f dt=%.2e | %.1f%%; ETA %.0f s | t/h %.2f | gp/s %.2e\r", // \r is carriage return, move cursor to the beginning of the line
n, n * (double)dt, (double)dt, (double)(100.0 * (REAL)n / (REAL)N_final),
(double)time_remaining_in_mins*60, (double)(dt * 3600.0 / s_per_iteration_avg), (double)RHS_pt_evals_per_sec);
fflush(stderr); // Flush the stderr buffer
} // End progress indicator if(n % 10 == 0)
} // End main loop to progress forward in time.
fprintf(stderr,"\n"); // Clear the final line of output from progress indicator.
// Step 4: Free all allocated memory
"""
if enable_rfm_precompute:
body += " rfm_precompute_rfmstruct_freemem(¶ms, &rfmstruct);\n"
body += r"""
freemem_bcstruct(¶ms, &bcstruct);
MoL_free_memory_y_n_gfs(¶ms, &gridfuncs);
MoL_free_memory_non_y_n_gfs(¶ms, &gridfuncs);
for(int i=0;i<3;i++) free(xx[i]);
return 0;
"""
# As rfmstruct stores functions of xx, when rfm_precompute is disabled,
# we always pass xx to a function instead of &rfmstruct.
if not enable_rfm_precompute:
body = body.replace("&rfmstruct", "xx")
add_to_Cfunction_dict(
includes=includes,
desc=desc,
c_type=c_type, name=name, params=params,
body=body,
rel_path_to_Cparams=os.path.join("."), enableCparameters=False)
```
<a id='compileexec'></a>
# Step 6: Compile generated C codes & perform the black hole collision calculation \[Back to [top](#toc)\]
$$\label{compileexec}$$
First we register remaining C functions and contributions to `NRPy_basic_defines.h`, then we output `NRPy_basic_defines.h` and `NRPy_function_prototypes.h`.
```
add_to_Cfunction_dict_main__BrillLindquist_Playground()
import outputC as outC
outC.outputC_register_C_functions_and_NRPy_basic_defines() # #define M_PI, etc.
# Declare paramstruct, register set_Cparameters_to_default(),
# and output declare_Cparameters_struct.h and set_Cparameters[].h:
outC.NRPy_param_funcs_register_C_functions_and_NRPy_basic_defines(os.path.join(Ccodesrootdir))
gri.register_C_functions_and_NRPy_basic_defines() # #define IDX3S(), etc.
fin.register_C_functions_and_NRPy_basic_defines(NGHOSTS_account_for_onezone_upwind=True,
enable_SIMD=enable_SIMD) # #define NGHOSTS, and UPWIND() macro if SIMD disabled
# Output functions for computing all finite-difference stencils.
# Must be called after defining all functions depending on FD stencils.
if enable_FD_functions:
fin.output_finite_difference_functions_h(path=Ccodesrootdir)
# Call this last: Set up NRPy_basic_defines.h and NRPy_function_prototypes.h.
outC.construct_NRPy_basic_defines_h(Ccodesrootdir, enable_SIMD=enable_SIMD)
outC.construct_NRPy_function_prototypes_h(Ccodesrootdir)
```
Finally, we output all the C codes in `outC_function_dict` to files in the directory `Ccodesrootdir`, generate a `Makefile`, and compile the project using a parallel `make` command. If the `make` command fails, a backup serial compilation script is run.
To aid in the cross-platform-compatible (with Windows, MacOS, & Linux) compilation and execution, we make use of `cmdline_helper` [(**Tutorial**)](Tutorial-cmdline_helper.ipynb).
```
import cmdline_helper as cmd
cmd.new_C_compile(Ccodesrootdir, os.path.join("output", "BrillLindquist_Playground"),
uses_free_parameters_h=True, compiler_opt_option="fast") # fastdebug or debug also supported
# Change to output directory
os.chdir(os.path.join(Ccodesrootdir, "output"))
# Clean up existing output files
cmd.delete_existing_files("out*.txt")
cmd.delete_existing_files("out*.png")
# Run executable with CFL_FACTOR = 1.0, which is allowed since
# simulation is axisymmetric and all phi derivs are set to zero.
CFL_FACTOR=1.0
cmd.Execute("BrillLindquist_Playground", "72 12 2 "+str(CFL_FACTOR))
cmd.Execute("BrillLindquist_Playground", "96 16 2 "+str(CFL_FACTOR))
os.chdir(os.path.join("..", ".."))
```
<a id='visualize'></a>
# Step 7: Visualize the output! \[Back to [top](#toc)\]
$$\label{visualize}$$
In this section we will generate a movie, plotting the conformal factor of these initial data on a 2D grid, such that darker colors imply stronger gravitational fields. Hence, we see the two black holes initially centered at $z/M=\pm 0.5$, where $M$ is an arbitrary mass scale (conventionally the [ADM mass](https://en.wikipedia.org/w/index.php?title=ADM_formalism&oldid=846335453) is chosen), and our formulation of Einstein's equations adopt $G=c=1$ [geometrized units](https://en.wikipedia.org/w/index.php?title=Geometrized_unit_system&oldid=861682626).
<a id='installdownload'></a>
## Step 7.a: Install `scipy` and download `ffmpeg` if they are not yet installed/downloaded \[Back to [top](#toc)\]
$$\label{installdownload}$$
Note that if you are not running this within `mybinder`, but on a Windows system, `ffmpeg` must be installed using a separate package (on [this site](http://ffmpeg.org/)), or if running Jupyter within Anaconda, use the command: `conda install -c conda-forge ffmpeg`.
```
!pip install scipy > /dev/null
check_for_ffmpeg = !which ffmpeg >/dev/null && echo $?
if check_for_ffmpeg != ['0']:
print("Couldn't find ffmpeg, so I'll download it.")
# Courtesy https://johnvansickle.com/ffmpeg/
!wget http://astro.phys.wvu.edu/zetienne/ffmpeg-static-amd64-johnvansickle.tar.xz
!tar Jxf ffmpeg-static-amd64-johnvansickle.tar.xz
print("Copying ffmpeg to ~/.local/bin/. Assumes ~/.local/bin is in the PATH.")
!mkdir ~/.local/bin/
!cp ffmpeg-static-amd64-johnvansickle/ffmpeg ~/.local/bin/
print("If this doesn't work, then install ffmpeg yourself. It should work fine on mybinder.")
```
<a id='genimages'></a>
## Step 7.b: Generate images for visualization animation \[Back to [top](#toc)\]
$$\label{genimages}$$
Here we loop through the data files output by the executable compiled and run in [the previous step](#mainc), generating a [png](https://en.wikipedia.org/wiki/Portable_Network_Graphics) image for each data file.
**Special thanks to Terrence Pierre Jacques. His work with the first versions of these scripts greatly contributed to the scripts as they exist below.**
```
## VISUALIZATION ANIMATION, PART 1: Generate PNGs, one per frame of movie ##
import numpy as np
from scipy.interpolate import griddata
import matplotlib.pyplot as plt
from matplotlib.pyplot import savefig
from IPython.display import HTML
import matplotlib.image as mgimg
import glob
import sys
from matplotlib import animation
globby = glob.glob(os.path.join(outdir,'out96-00*.txt'))
file_list = []
for x in sorted(globby):
file_list.append(x)
bound=1.4
pl_xmin = -bound
pl_xmax = +bound
pl_ymin = -bound
pl_ymax = +bound
for filename in file_list:
fig = plt.figure()
x,y,cf,Ham = np.loadtxt(filename).T #Transposed for easier unpacking
plotquantity = cf
plotdescription = "Numerical Soln."
plt.title("Black Hole Head-on Collision (conf factor)")
plt.xlabel("y/M")
plt.ylabel("z/M")
grid_x, grid_y = np.mgrid[pl_xmin:pl_xmax:300j, pl_ymin:pl_ymax:300j]
points = np.zeros((len(x), 2))
for i in range(len(x)):
# Zach says: No idea why x and y get flipped...
points[i][0] = y[i]
points[i][1] = x[i]
grid = griddata(points, plotquantity, (grid_x, grid_y), method='nearest')
gridcub = griddata(points, plotquantity, (grid_x, grid_y), method='cubic')
im = plt.imshow(gridcub, extent=(pl_xmin,pl_xmax, pl_ymin,pl_ymax))
ax = plt.colorbar()
ax.set_label(plotdescription)
savefig(os.path.join(filename+".png"),dpi=150)
plt.close(fig)
sys.stdout.write("%c[2K" % 27)
sys.stdout.write("Processing file "+filename+"\r")
sys.stdout.flush()
```
<a id='genvideo'></a>
## Step 7.c: Generate visualization animation \[Back to [top](#toc)\]
$$\label{genvideo}$$
In the following step, [ffmpeg](http://ffmpeg.org) is used to generate an [mp4](https://en.wikipedia.org/wiki/MPEG-4) video file, which can be played directly from this Jupyter notebook.
```
## VISUALIZATION ANIMATION, PART 2: Combine PNGs to generate movie ##
# https://stackoverflow.com/questions/14908576/how-to-remove-frame-from-matplotlib-pyplot-figure-vs-matplotlib-figure-frame
# https://stackoverflow.com/questions/23176161/animating-pngs-in-matplotlib-using-artistanimation
fig = plt.figure(frameon=False)
ax = fig.add_axes([0, 0, 1, 1])
ax.axis('off')
myimages = []
for i in range(len(file_list)):
img = mgimg.imread(file_list[i]+".png")
imgplot = plt.imshow(img)
myimages.append([imgplot])
ani = animation.ArtistAnimation(fig, myimages, interval=100, repeat_delay=1000)
plt.close()
ani.save(os.path.join(outdir,'BH_Head-on_Collision.mp4'), fps=5,dpi=150)
## VISUALIZATION ANIMATION, PART 3: Display movie as embedded HTML5 (see next cell) ##
# https://stackoverflow.com/questions/18019477/how-can-i-play-a-local-video-in-my-ipython-notebook
# Embed video based on suggestion:
# https://stackoverflow.com/questions/39900173/jupyter-notebook-html-cell-magic-with-python-variable
HTML("""
<video width="480" height="360" controls>
<source src=\""""+os.path.join(outdir,"BH_Head-on_Collision.mp4")+"""\" type="video/mp4">
</video>
""")
```
<a id='convergence'></a>
# Step 8: Plot the numerical error at the end of the simulation, and confirm that it converges to zero with increasing numerical resolution (sampling) \[Back to [top](#toc)\]
$$\label{convergence}$$
First we plot the log10-absolute value Hamiltonian constraint violation on the $x$-$z$ plane, near the black hole (at $x=y=z=0$). Notice the constraint violation is largest inside the horizon, as expected.
```
x96,y96,valuesCF96,valuesHam96 = np.loadtxt(os.path.join(outdir,'out96.txt')).T #Transposed for easier unpacking
pl_xmin = -2.5
pl_xmax = +2.5
pl_ymin = -2.5
pl_ymax = +2.5
grid_x, grid_y = np.mgrid[pl_xmin:pl_xmax:100j, pl_ymin:pl_ymax:100j]
points96 = np.zeros((len(x96), 2))
for i in range(len(x96)):
points96[i][0] = x96[i]
points96[i][1] = y96[i]
grid96 = griddata(points96, valuesCF96, (grid_x, grid_y), method='nearest')
grid96cub = griddata(points96, valuesCF96, (grid_x, grid_y), method='cubic')
grid96 = griddata(points96, valuesHam96, (grid_x, grid_y), method='nearest')
grid96cub = griddata(points96, valuesHam96, (grid_x, grid_y), method='cubic')
# fig, ax = plt.subplots()
plt.clf()
plt.title("96x16 Num. Err.: log_{10}|Ham|")
plt.xlabel("x/M")
plt.ylabel("z/M")
fig96cub = plt.imshow(grid96cub.T, extent=(pl_xmin,pl_xmax, pl_ymin,pl_ymax))
cb = plt.colorbar(fig96cub)
```
Next we check that indeed the numerical errors converge to zero as expected, using the fact that the Hamiltonian constraint violation should converge to zero with increasing resolution. See [the Scalar Wave Curvilinear tutorial notebook](Tutorial-Start_to_Finish-ScalarWaveCurvilinear_new_way.ipynb) for more documentation on measuring numerical convergence.
```
x72,y72,valuesCF72,valuesHam72 = np.loadtxt(os.path.join(outdir,'out72.txt')).T #Transposed for easier unpacking
points72 = np.zeros((len(x72), 2))
for i in range(len(x72)):
points72[i][0] = x72[i]
points72[i][1] = y72[i]
grid72 = griddata(points72, valuesHam72, (grid_x, grid_y), method='nearest')
griddiff_72_minus_96 = np.zeros((100,100))
griddiff_72_minus_96_1darray = np.zeros(100*100)
gridx_1darray_yeq0 = np.zeros(100)
grid72_1darray_yeq0 = np.zeros(100)
grid96_1darray_yeq0 = np.zeros(100)
count = 0
for i in range(100):
for j in range(100):
griddiff_72_minus_96[i][j] = grid72[i][j] - grid96[i][j]
griddiff_72_minus_96_1darray[count] = griddiff_72_minus_96[i][j]
if j==49:
gridx_1darray_yeq0[i] = grid_x[i][j]
grid72_1darray_yeq0[i] = grid72[i][j] + np.log10((72./96.)**4)
grid96_1darray_yeq0[i] = grid96[i][j]
count = count + 1
plt.clf()
fig, ax = plt.subplots()
plt.title("4th-order Convergence, at t/M=7.5 (post-merger; horiz at x/M=+/-1)")
plt.xlabel("x/M")
plt.ylabel("log10(Relative error)")
ax.plot(gridx_1darray_yeq0, grid96_1darray_yeq0, 'k-', label='Nr=96')
ax.plot(gridx_1darray_yeq0, grid72_1darray_yeq0, 'k--', label='Nr=72, mult by (72/96)^4')
ax.set_ylim([-8.5,0.5])
legend = ax.legend(loc='lower right', shadow=True, fontsize='x-large')
legend.get_frame().set_facecolor('C1')
plt.show()
```
<a id='latex_pdf_output'></a>
# Step 9: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
$$\label{latex_pdf_output}$$
The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
[Tutorial-Start_to_Finish-BSSNCurvilinear-Two_BHs_Collide_new_way.pdf](Tutorial-Start_to_Finish-BSSNCurvilinear-Two_BHs_Collide_new_way.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
```
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-Start_to_Finish-BSSNCurvilinear-Two_BHs_Collide_new_way")
```
| github_jupyter |
```
%pylab inline
import seaborn as sns
sns.set_style('white')
import pandas as pd
import torch
import torch.nn as nn
from src.data.cmnist_dist import make_joint_distribution
from src.discrete.distribution import DiscreteDistribution, compute_ce, compute_kl
from src.discrete.distribution.plot import plot_data
from src.discrete.models.training import train
from src.discrete.models.encoders import DiscreteEncoder
```
# Data distribution
We will use the y-CMNIST data distribution as an example in this notebook, which defines a joint distribution of digits $d$, color $c$, pictures $x$, environments $e$, labels $y$ and selection $t$.
The binary selection variable $t$ indicates if the data is selected for training $t=1$ or not $t=0$.
```
# Create the joint distribution for y-CMNIST (CMNIST or d-CMNIST are also available)
dataset = 'y-CMNIST'
dist = make_joint_distribution(dataset)
print('Joint distribution: %s' %dist)
# We visualize some of the pair-wise joint distributions
f, ax = plt.subplots(1,3, figsize=(15, 5))
for i, joint in enumerate([
dist.marginal(['y','e']),
dist.marginal(['y','d']),
dist.marginal(['y','c'])]):
ax[i].set_title(joint, size=18)
ax[i].imshow(joint.p, clim=(0,1), cmap='viridis')
ax[i].set_xlabel(joint.indices[1], size=18)
ax[i].set_ylabel(joint.indices[0], size=18)
ax[i].set_xticks(range(joint.p.shape[1]))
ax[i].set_yticks(range(joint.p.shape[0]))
```
The joint distribution factorizes as $p(y,e,d,x,c,t) = p(y|d)p(e)p(d)p(c|y,e)p(x|c,d)p(t|e)$.
Train and test distributions are created by conditioning on $t=1$ and $t=0$, respectively.
```
# Create the training distribution by selecting t=1
train_dist = dist.condition_on('t',1)
# Create the test distribution by selecting t=0
test_dist = dist.condition_on('t',0)
print('Train distribution: %s' %train_dist)
print('Test distribution: %s' %test_dist)
```
We can describe how distant the train and test distributions are by computing $I(xy;t)$, which represents how much information the selection carries about the joint distribution of raw features (pictures $x$) and targets (labels $y$). The distribution shift can be seen as the joint effect of covariate shift $I(t;x)$ and concept shift $I(y;t|x)$:
$$\underbrace{I(xy;t)}_{\text{distribution shift}} = \underbrace{I(x;t)}_{\text{covariate shift}} + \underbrace{I(y;t|x)}_{\text{concept shift}}$$
```
# Compute the amout of distribution shift I(xy;t)
print('Distribution shift: %f nats' %(dist.compute('I(x,y;t)')))
# Compute the amout of concept shift I(y;t|x)
print('Concept shift: %f nats' %(dist.compute('I(y;t|x)')))
# Compute the amout of concept shift I(x;t)
print('Covariate shift: %f nats' %(dist.compute('I(x;t)')))
```
Note that the selection $t$ does induce concept shift, therefore a maximum likelihood solution must result in positive Test error.
Given a model $q(y|x)$ that is optimal on the training distribution $p(y|x,t=1)$ we have:
$$KL(p(y|x,t=0)||q(y|x))\ge \frac{1}{1-\alpha} I(y;t|x) > 0.237813 \text{ nats}$$
# Introducing a Latent representation
The test error for a model that is composed by an encoder $q(z|x)$ and classifier $q(y|z)$ is upper bounded by the *Test Information Loss*, which represents the amount of information lost trhough the encoding procedure (in red), and *Latent Test error* (in blue) which represents the error when using the latent features $z$ instead of the original observations $x$:
$$\underbrace{KL(p(y|x,t=0)||q(y|x))}_{\text{Test Error}}\le \underbrace{I_{t=0}(x;y|z)}_{\text{Test Information Loss}} + \underbrace{KL(p(y|z,t=0)||q(y|z))}_{\text{Latent Test Error}}$$
Clearly, considering an identity encoder, which maps each observation into itself, the Test error is equivalent to the Latent Test error. This also happens for any encoder $q(z|x)$ that retains all the predictive information.
```
data = []
# define alpha as the minimum probability between p(t=0) and p(t=1)
alpha = dist.marginal('t').p.min()
# Consider a model q(y|x) that matches the train distribution p(y|x,t=1)
q_y_x = train_dist.conditional('y','x')
print('q(y|x)=%s' % q_y_x)
# Compute the lower bound I(y;t|x)/(1-alpha)
l_bound = dist.compute('I(y;t|x)')/(1-alpha)
# Compute the OOD error for q(y|x): KL(p(y|x,t=0)||q(y|x))
test_error = compute_kl(test_dist, q_y_x)
print('Lower-bound I(y;x|t): %f nats' % l_bound)
print('Test error: %f nats' % test_error)
data.append({
'Model': 'Picture',
'Test Information Loss': 0,
'Latent Test error': test_error.item()
})
f, ax = plt.subplots(1,1)
plot_data(data, ax)
```
We compute and visualize the components of the Test error for models that build representations based on color $c$, digit $d$ or no information (prior).
Since the two marginal label distributions $p(y|t=0)$ and $p(y|t=1)$ are equivalent, the error of a model that discards all the information is entirely due to information loss, and the corresponding Latent Test error is zero.
On the other hand, any model that relies exclusively on color information (discarding anying about the digits) incurs in a higher Test error since the dependency between color and label is inverted at test time ($t=0$).
A model that discards color information while keeping digit information is able to minimize the latent Test error, while retaining most of the predictive information. The corresponding representation contains only digit information which is reliable to predict the label $y$ across all the environments.
```
models = []
# Color
data.append({
'Model': 'Color',
'Latent Test error': compute_kl(test_dist, train_dist.conditional('y','c')),
'Test Information Loss': test_dist.mi('y','x','c')
})
# Digit
data.append({
'Model': 'Digit',
'Latent Test error': compute_kl(test_dist, train_dist.conditional('y','d')),
'Test Information Loss': test_dist.mi('y','x','d')
})
# Prior
data.append({
'Model': 'Prior',
'Latent Test error': compute_kl(test_dist, train_dist.marginal('y')),
'Test Information Loss': test_dist.mi('y','x')
})
f, ax = plt.subplots(1,1)
sns.despine()
plot_data(data, ax)
```
As a next step, we randomly initialize an encoder $q(z|x)$ and measure the compoments of the Test error for the untrained encoder. Usually most of the information is discarded and the latent OOD error is small.
```
# Create an encoder which maps each 'x' into a latent 'z' which consists of 64 different values
encoder = DiscreteEncoder(z_dim=64)
# Define an classifier q(y|z)=p(y|z,\t=1) that is optimal on training
q_y_z = encoder(train_dist).conditional('y','z')
latent_test_dist = encoder(test_dist).marginal(['y','z','x'])
print('q(y|z)=%s' % q_y_z)
# Compute the Test error of the overall model q(y|x)
test_error = compute_kl(latent_test_dist, q_y_z, cond_1='x')
print('Test error: %f' % test_error)
# Compute the latent Test error of the model q(y|z)
lat_test_error = compute_kl(latent_test_dist, q_y_z, cond_1='z')
print('Latent Test error: %f' % lat_test_error)
# Compute the amount of test information loss
test_info_loss = encoder(test_dist).mi('y','x','z')
print('Test Infomation Loss: %f' % test_info_loss)
data.append({
'Model': 'Random',
'Latent Test error': lat_test_error.item(),
'Test Information Loss': test_info_loss.item()
})
f, ax = plt.subplots(1,1)
plot_data(data, ax)
sns.despine()
```
The goal is to train the encoder $q(z|x)$ to minimize the Latent Test error while retaining maximal predictive information. We analyze different families of objectives in literature. Each of them present a loss in the form:
$$\mathcal{L}(\lambda) = \mathbb{E}_{t=1}[-\log q(y|z)]+ \lambda \mathcal{R}(q(z|x)),$$
in which the first term aims to maximize the amount of predictive information in the representation, while the second is a regularization term that aims to reduce the Latent Test error.
The trade-off between the two objectives is regulated by the hyper-parameter $\lambda$. Different regularization will be based on the observation of an environment variable $e$, which represents the factor(s) on which the selection is based on.
# Model Training
Here we explore the effect of training a latent representation using the following criteria:
- Independence $\mathcal{R}(q(z|x))=I(e;z)$
- Sufficiency $\mathcal{R}(q(z|x))=I(e;y|z)$
- Separation $\mathcal{R}(q(z|x))=I(e;z|y)$
## Independence Criterion
We train the model using the regularization prescribed by the independence criterion until convergence
```
from src.discrete.models.criteria import IndependenceCriterion
# re-initialize the encoder
encoder = DiscreteEncoder(z_dim=64)
# Use the Independence criterion for training with a strong regularization $\lambda=10^6$
criterion = IndependenceCriterion(reg=1e6)
# Train until convergence, logging train and test cross entropy
logs = train(encoder,
criterion,
train_dist=train_dist.marginal(['x','y','e']), # train distribution
test_dist=test_dist.marginal(['x','y']), # test distribution
verbose=False)
# Visualize the logs
pd.DataFrame(logs).plot(x='iteration')
```
We can compute the value of the regularization term $I(e;z)$ at the end of the training. One can notice that the independence constraint is correctly enforced on training but there is a small dependency left when considering the unselected (original) distribution.
```
# Compute the value of I(e;z) on for the trained model for the train distribution
print('I(e;z|t=1) = %f nats' % encoder(train_dist).mi('e','z'))
# Compute the value of I(e;z) on for the trained model
print('I(e;z) = %f nats' % encoder(dist).mi('e','z'))
```
We visualize the error compoments after convergence. Even if constraint is correctly enforced on train, the separation criterion does not result in minimal Test error
```
q_y_z = encoder(train_dist).conditional('y','z')
latent_test_dist = encoder(test_dist).marginal(['y','z','x'])
# Compute the Test error of the overall model q(y|x)
test_error = compute_kl(latent_test_dist, q_y_z, cond_1='x')
print('Test error: %f' % test_error)
# Compute the latent Test error of the model q(y|z)
lat_test_error = compute_kl(latent_test_dist, q_y_z, cond_1='z')
print('Latent Test error: %f' % lat_test_error)
# Compute the amount of test information loss
test_info_loss = encoder(test_dist).mi('y','x','z')
print('Test Infomation Loss: %f' % test_info_loss)
data.append({
'Model': 'Independence',
'Latent Test error': lat_test_error.item(),
'Test Information Loss': test_info_loss.item()
})
# Plot the results
f, ax = plt.subplots(1,1)
plot_data(data, ax)
sns.despine()
```
## Sufficiency Criterion
We re-initialize the encoder and train it using the sufficiency criterion
```
from src.discrete.models.criteria import SufficiencyCriterion
# re-initialize the encoder
encoder = DiscreteEncoder(z_dim=64)
# Use the Sufficiency criterion for training with a strong regularization $\lambda=10^6$
criterion = SufficiencyCriterion(reg=1e6)
# Train until convergence, logging train and test cross entropy
logs = train(encoder,
criterion,
train_dist=train_dist.marginal(['x','y','e']), # train distribution
test_dist=test_dist.marginal(['x','y']), # test distribution
verbose=False)
# Visualize the logs
pd.DataFrame(logs).plot(x='iteration')
```
Note that despite the long training, the model is not able to create a representation that satisfies the sufficiency constraint $ I(e;y|z)=0 $ on training (such representation does not exist for the d-CMNIST dataset
```
# Compute the value of I(e;z) on for the trained model for the train distribution
print('I(e;y|z, t=1) = %f' % encoder(train_dist).mi('e','y','z'))
# Compute the value of I(e;z) on for the trained model
print('I(e;y|z) = %f' % encoder(dist).mi('e','y','z'))
```
Once again the resulting model is not optimal in terms of Test error
```
q_y_z = encoder(train_dist).conditional('y','z')
latent_test_dist = encoder(test_dist).marginal(['y','z','x'])
# Compute the Test error of the overall model q(y|x)
test_error = compute_kl(latent_test_dist, q_y_z, cond_1='x')
print('Test error: %f' % test_error)
# Compute the latent Test error of the model q(y|z)
lat_test_error = compute_kl(latent_test_dist, q_y_z, cond_1='z')
print('Latent Test error: %f' % lat_test_error)
# Compute the amount of test information loss
test_info_loss = encoder(test_dist).mi('y','x','z')
print('Test Infomation Loss: %f' % test_info_loss)
data.append({
'Model': 'Sufficiency',
'Latent Test error': lat_test_error.item(),
'Test Information Loss': test_info_loss.item()
})
# Plot the results
f, ax = plt.subplots(1,1)
plot_data(data, ax)
sns.despine()
```
## Separation Criterion
We repeat the same procedure by applying the separation criterion instead
```
from src.discrete.models.criteria import SeparationCriterion
# re-initialize the encoder
encoder = DiscreteEncoder(z_dim=64)
# Use the Sufficiency criterion for training with a strong regularization $\lambda=10^6$
criterion = SeparationCriterion(reg=1e6)
# Train until convergence, logging train and test cross entropy
logs = train(encoder,
criterion,
train_dist=train_dist.marginal(['x','y','e']), # train distribution
test_dist=test_dist.marginal(['x','y']), # test distribution
verbose=False)
# Visualize the logs
pd.DataFrame(logs).plot(x='iteration')
```
Once again, we check if the model manages to enforce the separation constraint $I(e;z|y)=0$ for the overall (unselected) distribution
```
# Compute the value of I(e;z) on for the trained model for the train distribution
print('I(e;z|y, t=1) = %f' % encoder(train_dist).mi('e','z','y'))
# Compute the value of I(e;z) on for the trained model
print('I(e;z|y) = %f' % encoder(dist).mi('e','z','y'))
```
The model that enforces the separation criterion is the only one that minimizes the overall OOD error on the y-CMNIST dataset
```
q_y_z = encoder(train_dist).conditional('y','z')
latent_test_dist = encoder(test_dist).marginal(['y','z','x'])
# Compute the Test error of the overall model q(y|x)
test_error = compute_kl(latent_test_dist, q_y_z, cond_1='x')
print('Test error: %f' % test_error)
# Compute the latent Test error of the model q(y|z)
lat_test_error = compute_kl(latent_test_dist, q_y_z, cond_1='z')
print('Latent Test error: %f' % lat_test_error)
# Compute the amount of test information loss
test_info_loss = encoder(test_dist).mi('y','x','z')
print('Test Infomation Loss: %f' % test_info_loss)
data.append({
'Model': 'Separation',
'Latent Test error': lat_test_error.item(),
'Test Information Loss': test_info_loss.item()
})
# Plot the results
f, ax = plt.subplots(1,1)
plot_data(data, ax)
sns.despine()
```
The other experiments reported in the paper can be repliated by changing the data-generating distribution reported at the beginning of this notebook. The `train_discrete.py` training script used to produce the trajectories shown in the paper. Here we show the trajectories obtained by training with different values of $\lambda$
```
# Create the axis for plotting
f, ax = plt.subplots(1,3, figsize=(15,5))
# Load the data from files
disc_data = pd.read_csv('results/trajectories.csv')
th_data = pd.read_csv('results/points.csv')
# Plot the points corresponding to models using picture, digit, color or prior information
for i, (dataset, dataset_data) in enumerate(th_data.groupby('dataset')):
for _, entry in dataset_data.iterrows():
ax[i].plot(
entry['CrossEntropy(T=1)'],
entry['CrossEntropy(T=0)'],
'o',
label = entry['name']
)
ax[i].set_xlabel('Train Cross-entropy', fontsize=15)
# Plot the trajectories obtained using different values of lambda for the different criteria
for i, (dataset_name, dataset_results) in enumerate(disc_data.groupby('dataset')):
ax[i].set_title(dataset_name, size=15)
for j, (criterion_name, criterion_data) in enumerate(dataset_results.groupby('criterion')):
ax[i].plot(criterion_data['CrossEntropy(t=1)'],
criterion_data['CrossEntropy(t=0)'],
'.-', label=criterion_name, zorder=-j)
ax[0].set_ylabel('Test Cross-entropy', fontsize=15)
ax[-1].legend(fontsize=12)
sns.despine()
```
| github_jupyter |
# Principle Component Analysis (PCA)
* Unsupervised learning method
* Difficult to understand components beyond which have highest variance
* Good step to do at end of processing because of way data gets transformed and reshaped
References:
* [Dimensionality Reduction in Python](https://campus.datacamp.com/courses/dimensionality-reduction-in-python/feature-extraction)
```
import numpy as np
import pandas as pd
from sklearn.decomposition import PCA
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler
import seaborn as sns
from sklearn.pipeline import Pipeline
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
```
## Made up example
```
df = pd.DataFrame({
'low_variance': np.random.normal(100, 1, 10),
'high_variance': np.random.normal(100, 50, 10),
'medium_variance': np.random.normal(100, 10, 10)
})
print("Original:\n")
print(df)
pca = PCA()
arr = pca.fit_transform(df)
print("\nAfter PCA:\n")
print(arr)
print("\nExplained Variance Ratio:\n")
print(pca.explained_variance_ratio_)
print("\nExplained Variance:\n")
print(pca.explained_variance_)
```
* Notice how most of the variance is explained by one feature (the high variance one).
* The order of `explained_variance_` and `explained_variance_ratio_` _do not_ necessarilly line up with the order of the features in the original data frame. They're ordered by the variance of the original features.
* By default PCA uses the same number of components as features, but can be reduced by setting `n_components`
* Explained Variance Ratio: Percentage of variance explained by each of the selected components (adds up to 1)
* Explained Variance: The amount of variance explained by each of the selected components.
## Charting the variance
```
features = range(pca.n_components_)
plt.bar(features, pca.explained_variance_)
plt.xlabel('PCA feature')
plt.ylabel('variance')
plt.xticks(features)
plt.show()
```
## Titantic data
```
df = pd.read_csv("data/titantic-train.csv").dropna()
display(df.head())
y = df['Survived']
X = df[['Age', 'Fare', 'Pclass', 'SibSp']]
display(X.head())
```
## Pair plot of original data
```
sns.pairplot(X)
plt.show()
```
## Scaling the data
```
scaler = StandardScaler()
X_std = scaler.fit_transform(X)
display(X_std[:3,:])
```
## PCA analyais
```
pca = PCA()
pc = pca.fit_transform(X_std)
pc_df = pd.DataFrame(pc, columns=["PC1", "PC2", "PC3", "PC4"])
display(pc_df.head())
print("\nExplained Variance Ratio:\n")
print(pca.explained_variance_ratio_)
print("\nCumulative Explained Variance Ratio:\n")
print(pca.explained_variance_ratio_.cumsum())
```
## Pair plot of principle components
```
# "Notice how, in contrast to the input features, none of the principal components are correlated to one another."
sns.pairplot(pc_df)
plt.show()
```
## Pipeline
```
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
pipe = Pipeline([
('scaler', StandardScaler()),
('reducer', PCA(n_components=2)),
('rf', RandomForestClassifier(max_depth=2))
])
pipe.fit(X_train, y_train)
print("Accuracy score: {:.2f}".format(pipe.score(X_test, y_test)))
vectors = pipe.steps[1][1].components_.round(2)
# Print feature effects
print('\nPC 1 effects = ' + str(dict(zip(X.columns, vectors[0]))))
print('PC 2 effects = ' + str(dict(zip(X.columns, vectors[1]))))
# PC 1 component quantifies a trade-off between class and age, for example
pipe.steps[1][1]
```
## Segmenting a scatter plot by a categorical variable
```
pipe = Pipeline([
('scaler', StandardScaler()),
('reducer', PCA(n_components=2))
])
pc = pipe.fit_transform(X)
df1 = df[["Sex"]].copy()
df1["PC1"] = pc[:, 0]
df1["PC2"] = pc[:, 1]
display(df1.head())
sns.scatterplot(data=df1, x="PC1", y="PC2", hue="Sex")
plt.show()
```
| github_jupyter |
# Bytecode Processing
```
import os;
os.getpid()
import hybridcuda
import json
def inspection(f):
hc = hybridcuda.disassemble(f)
print('=== hybrid ===')
print(hc['hybrid'])
print('=== inspect ===')
print(hc['inspect'])
def validate(f):
hc = hybridcuda.disassemble(f)
parsedinspect = json.loads(hc['inspect'])
parsedhybrid = json.loads(hc['hybrid'])
ppinspect = json.dumps(parsedinspect, indent=4, sort_keys=True)
pphybrid = json.dumps(parsedhybrid, indent=4, sort_keys=True)
linesinspect = ppinspect.splitlines()
lineshybrid = pphybrid.splitlines()
if (linesinspect.__len__() != lineshybrid.__len__()):
raise Exception('NOT SAME NUMBER OF LINES')
samecount = 0
for i in range(linesinspect.__len__()):
if (linesinspect[i] == lineshybrid[i]):
samecount = samecount + 1
continue
if ('lineno' in linesinspect[i]):
continue
if ('col_offset' in linesinspect[i]):
continue
print('=== INSPECT ===')
print(linesinspect[i])
print('=== HYBRID ===')
print(lineshybrid[i])
raise Exception('LINE DELTA')
return samecount
```
<a href="#HERE">here</a>
```
def disasm_001():
return 42
inspection(disasm_001)
validate(disasm_001)
def disasm_002(x):
return x
inspection(disasm_002)
validate(disasm_002)
def disasm_003(x,y):
return x+y
inspection(disasm_003)
validate(disasm_003)
def disasm_004(x,y):
return x*y
inspection(disasm_004)
validate(disasm_004)
def disasm_005(x,y):
x[0] = y
inspection(disasm_005)
#print(json.dumps(json.loads(hybridcuda.disassemble(disasm_005)['hybrid']), indent=4, sort_keys=True))
validate(disasm_005)
def disasm_006(x,y):
return x[0]
inspection(disasm_006)
#print(json.dumps(json.loads(hybridcuda.disassemble(disasm_006)['inspect']), indent=4, sort_keys=True))
validate(disasm_006)
def disasm_007(x,y):
# pass INSERTED IN INSPECT BUT NOT VISIBLE IN BYTE CODE !!!
return x[0]
inspection(disasm_007)
#print(json.dumps(json.loads(hybridcuda.disassemble(disasm_007)['hybrid']), indent=4, sort_keys=True))
validate(disasm_007)
def disasm_008(x,y):
pass
inspection(disasm_008)
#print(json.dumps(json.loads(hybridcuda.disassemble(disasm_007)['hybrid']), indent=4, sort_keys=True))
# NOT SAME NUMBER OF CELLS IS EXPECTED FOR PASS => CANNOT BE RECONSTRUCTED... validate(disasm_008)
```
https://docs.python.org/3.6/library/dis.html#python-bytecode-instructions
## Unary operations
https://docs.python.org/3.6/library/dis.html#opcode-UNARY_POSITIVE
```
def disasm_101(x,y):
x = +x
inspection(disasm_101)
validate(disasm_101)
#import dis
#dis.dis(disasm_101)
def disasm_102(x,y):
x = -x
inspection(disasm_102)
validate(disasm_102)
def disasm_103(x,y):
x = not x
inspection(disasm_103)
validate(disasm_103)
def disasm_104(x,y):
x = ~ x
inspection(disasm_104)
validate(disasm_104)
```
TODO: https://docs.python.org/3.6/library/dis.html#opcode-GET_ITER
TODO: https://docs.python.org/3.6/library/dis.html#opcode-GET_YIELD_FROM_ITER
## Binary operations
https://docs.python.org/3.6/library/dis.html#opcode-BINARY_POWER
```
def disasm_201(x,y):
return x ** y
inspection(disasm_201)
validate(disasm_201)
def disasm_202(x,y):
return x * y
inspection(disasm_202)
validate(disasm_202)
# TODO ??? MATRIX MULTIPLY ?
def disasm_204(x,y):
return x // y
inspection(disasm_204)
validate(disasm_204)
def disasm_205(x,y):
return x / y
inspection(disasm_205)
validate(disasm_205)
def disasm_206(x,y):
return x % y
inspection(disasm_206)
validate(disasm_206)
def disasm_207(x,y):
return x + y
inspection(disasm_207)
validate(disasm_207)
def disasm_208(x,y):
return x - y
inspection(disasm_208)
validate(disasm_208)
def disasm_209(x,y):
return x [y]
inspection(disasm_209)
validate(disasm_209)
def disasm_210(x,y):
return x << y
inspection(disasm_210)
validate(disasm_210)
def disasm_211(x,y):
return x >> y
inspection(disasm_211)
validate(disasm_211)
def disasm_212(x,y):
return x & y
inspection(disasm_212)
validate(disasm_212)
def disasm_213(x,y):
return x ^ y
inspection(disasm_213)
validate(disasm_213)
def disasm_214(x,y):
return x | y
inspection(disasm_214)
validate(disasm_214)
def disasm_215(a,b):
return a and b
inspection(disasm_215)
validate(disasm_215)
import dis
dis.dis(disasm_215)
def disasm_216(a,b):
return a or b
inspection(disasm_216)
validate(disasm_216)
def disasm_217(a,b):
return not b
inspection(disasm_217)
validate(disasm_217)
def disasm_218(a,b, c, d):
return a and b and c and d
inspection(disasm_218)
validate(disasm_218)
def disasm_219(a,b, c, d):
return a or b or c or d
inspection(disasm_219)
validate(disasm_219)
def disasm_220(a,b, c):
return a or b and c
inspection(disasm_220)
validate(disasm_220)
```
## In-place operations
https://docs.python.org/3.6/library/dis.html#opcode-INPLACE_POWER
https://docs.python.org/3.6/reference/simple_stmts.html#augmented-assignment-statements
```
def disasm_251(a,b):
a += b
inspection(disasm_251)
validate(disasm_251)
def disasm_252(a,b):
a -= b
inspection(disasm_252)
validate(disasm_252)
def disasm_253(a,b):
a *= b
inspection(disasm_253)
validate(disasm_253)
def disasm_254(a,b):
a /= b
inspection(disasm_254)
validate(disasm_254)
def disasm_255(a,b):
a //= b
inspection(disasm_255)
validate(disasm_255)
def disasm_256(a,b):
a %= b
inspection(disasm_256)
validate(disasm_256)
def disasm_257(a,b):
a **= b
inspection(disasm_257)
validate(disasm_257)
def disasm_258(a,b):
a >>= b
inspection(disasm_258)
validate(disasm_258)
def disasm_259(a,b):
a <<= b
inspection(disasm_259)
validate(disasm_259)
def disasm_260(a,b):
a &= b
inspection(disasm_260)
validate(disasm_260)
def disasm_261(a,b):
a ^= b
inspection(disasm_261)
validate(disasm_261)
def disasm_262(a,b):
a |= b
inspection(disasm_262)
validate(disasm_262)
```
## General instructions
https://docs.python.org/3.6/library/dis.html#opcode-NOP
```
def disasm_301(a,b):
b[0] += a
import dis
dis.dis(disasm_301)
inspection(disasm_301)
validate(disasm_301)
def disasm_302(x):
print(x)
inspection(disasm_302)
#print(json.dumps(json.loads(hybridcuda.disassemble(disasm_005)['hybrid']), indent=4, sort_keys=True))
import dis
dis.dis(disasm_302)
validate(disasm_302)
```
## Coroutine opcodes
https://docs.python.org/3.6/library/dis.html#opcode-GET_AWAITABLE
*SUPPORT ?*
## Miscellaneous opcodes
https://docs.python.org/3.6/library/dis.html#opcode-PRINT_EXPR
```
def disasm_501(a,b):
a = 1 if b else 2
inspection(disasm_501)
validate(disasm_501)
def disasm_502(a,b):
a = 1 if b else 2 if a else 3
inspection(disasm_502)
validate(disasm_502)
def disasm_503(a,b):
if b:
a = 1
else:
a = 2
inspection(disasm_503)
validate(disasm_503)
def disasm_504(a,b):
if b:
a = 1
inspection(disasm_504)
validate(disasm_504)
def disasm_505(a,b,c):
if a:
if b:
c = 1
else:
c = 2
else:
if b:
c = 3
else:
c = 4
#import dis
#dis.dis(disasm_505)
#hc = hybridcuda.disassemble(disasm_505)
#import json
#parsed = json.loads(hc['hybrid'])
#json.dumps(parsed, indent=4, sort_keys=True).splitlines()
inspection(disasm_505)
validate(disasm_505)
def disasm_506(a,b,c):
if a:
if b:
c = 1
else:
if b:
c = 3
else:
c = 4
inspection(disasm_506)
validate(disasm_506)
def disasm_507(a,b,c):
if a:
if b:
c = 1
else:
if b:
c = 3
inspection(disasm_507)
validate(disasm_507)
def disasm_508(a,b,c):
if a:
if b:
c = 1
else:
c = 2
else:
if b:
c = 3
inspection(disasm_508)
validate(disasm_508)
def disasm_509(a,b,c):
c[0] = a < b
c[1] = a > b
c[2] = a != b
c[3] = a == b
c[4] = a <= b
c[5] = a >= b
#import dis
#dis.dis(disasm_509)
inspection(disasm_509)
validate(disasm_509)
# CAVEAT : multiple compares are not supported
def disasm_510(a,b):
x = 0
while (x < a):
x = x + 1
import dis
dis.dis(disasm_510)
inspection(disasm_510)
validate(disasm_510)
# hc = hybridcuda.disassemble(disasm_510)
# import json
# parsed = json.loads(hc['inspect'])
# print(json.dumps(parsed, indent=4, sort_keys=True))
def disasm_511(a,b):
x = 0
while (x < a):
x = x + 1
x = x - 1
import dis
dis.dis(disasm_511)
inspection(disasm_511)
validate(disasm_511)
def disasm_512(a,b):
x = 0
while (x < a):
x = x + 1
break
x = x - 1
import dis
dis.dis(disasm_512)
inspection(disasm_512)
validate(disasm_512)
# TODO ? print_expr
def disasm_513():
for i in range(1):
x = x+i
import dis
dis.dis(disasm_513)
inspection(disasm_513)
validate(disasm_513)
def disasm_514():
for i in range(1):
x = x+i
if (x > 3):
continue
import dis
dis.dis(disasm_514)
inspection(disasm_514)
validate(disasm_514)
def disasm_601(a,b):
assert (a == b[0])
import dis
dis.dis(disasm_601)
inspection(disasm_601)
validate(disasm_601)
def disasm_602(a,b):
assert (a == b[0]), 'delta'
import dis
dis.dis(disasm_602)
inspection(disasm_602)
validate(disasm_602)
```
# HERE
<a href="#Bytecode-Processing">head</a>
```
import hybridcuda
import json
def inspection(f):
hc = hybridcuda.disassemble(f)
print('=== hybrid ===')
print(hc['hybrid'])
print('=== inspect ===')
print(hc['inspect'])
def validate(f):
hc = hybridcuda.disassemble(f)
parsedinspect = json.loads(hc['inspect'])
parsedhybrid = json.loads(hc['hybrid'])
ppinspect = json.dumps(parsedinspect, indent=4, sort_keys=True)
pphybrid = json.dumps(parsedhybrid, indent=4, sort_keys=True)
linesinspect = ppinspect.splitlines()
lineshybrid = pphybrid.splitlines()
if (linesinspect.__len__() != lineshybrid.__len__()):
raise Exception('NOT SAME NUMBER OF LINES')
samecount = 0
for i in range(linesinspect.__len__()):
if (linesinspect[i] == lineshybrid[i]):
samecount = samecount + 1
continue
if ('lineno' in linesinspect[i]):
continue
if ('col_offset' in linesinspect[i]):
continue
print('=== INSPECT ===')
print(linesinspect[i])
print('=== HYBRID ===')
print(lineshybrid[i])
raise Exception('LINE DELTA')
return samecount
import os;
os.getpid()
def disasm_701(x,y):
w = lambda a,b : x + y
return w
w = disasm_701(30,12)
inspection(w)
# -- INSPECT FAILS FOR LAMBDAS => would not match anyway since names are different -- validate(w)
```
# SANDBOX
```
import dis
dis.dis(disasm_007)
'hell' in 'hello'
hc = hybridcuda.disassemble(disasm_003)
import json
parsed = json.loads(hc['inspect'])
json.dumps(parsed, indent=4, sort_keys=True).splitlines().__len__()
print(hc['inspect'])
import dis
dis.dis(disasm_001)
def f(x,y):
a = x+y
k = "hello"
w = 42
return a
def g():
return 42;
print(getattr(getattr(f, '__code__'), 'co_code'))
print(getattr(getattr(f, '__code__'), 'co_consts'))
print(getattr(getattr(f, '__code__'), 'co_argcount'))
print(getattr(getattr(f, '__code__'), 'co_freevars'))
print(getattr(getattr(f, '__code__'), 'co_cellvars'))
print(getattr(getattr(f, '__code__'), 'co_varnames'))
print(getattr(getattr(f, '__code__'), 'co_nlocals'))
print(getattr(getattr(f, '__code__'), 'co_lnotab'))
print(getattr(getattr(f, '__code__'), 'co_stacksize'))
print(getattr(getattr(f, '__code__'), 'co_firstlineno'))
print(getattr(getattr(g, '__code__'), 'co_firstlineno'))
def arithmetic_convert_001(a,b,c):
c[0] = a + b
import dis
dis.dis(arithmetic_convert_001)
def multiassign(a,b,c):
x,y = a,b
c[0] = (x,y)
import dis
dis.dis(multiassign)
import dis
dis.dis(f)
type(getattr(getattr(f, '__code__'), 'co_varnames')[0])
def g():
return 'hello'
print(getattr(getattr(g, '__code__'), 'co_code'))
dis.dis(g)
dir(getattr(g, '__code__'))
2+2
def somme(x,y):
return x+y
somme(30,12)
somme(10,8)
def f(N : int, a):
return a + N
print(getattr(getattr(f, '__code__'), 'co_code'))
print(getattr(getattr(f, '__code__'), 'co_consts'))
print(getattr(getattr(f, '__code__'), 'co_argcount'))
print(getattr(getattr(f, '__code__'), 'co_freevars'))
print(getattr(getattr(f, '__code__'), 'co_cellvars'))
print(getattr(getattr(f, '__code__'), 'co_varnames'))
print(getattr(getattr(f, '__code__'), 'co_nlocals'))
print(getattr(getattr(f, '__code__'), 'co_lnotab'))
print(getattr(getattr(f, '__code__'), 'co_stacksize'))
print(getattr(getattr(f, '__code__'), 'co_flags'))
print(getattr(getattr(f, '__code__'), '__repr__')())
dir(getattr(f, '__code__'))
#type(getattr(getattr(f, '__code__'), 'co_flags'))
```
| github_jupyter |
# How to perform aperture photometry with custom apertures?
We have discussed in previous tutorials how Simple Aperture Photometry works. We choose a set of pixels in the image and sum those to produce a single flux value. We sum the same pre-selected pixels for every image at each time slice to produce a light curve.
The [Kepler Data Pipeline](https://github.com/nasa/kepler-pipeline) produces an aperture, which is used by default by lightkurve. However, there are some cases where you might want to produce your own aperture. The field may be crowded, or you may wish to change the aperture size to change the relative contribution of the background. K2 data generally needs larger apertures than the default pipeline mask. Lightkurve offers tools to select pixels programmatically.
First, let's load a target pixel file. Let's choose Kepler planet canidate [KIC 6679295](https://exoplanetarchive.ipac.caltech.edu/cgi-bin/DisplayOverview/nph-DisplayOverview?objname=KOI-2862.01&type=KEPLER_CANDIDATE). We'll use the `search_targetpixelfile` function to download every target pixel file available for each quarter of this data set.
```
%matplotlib inline
from lightkurve import search_targetpixelfile
import matplotlib.pyplot as plt
tpfs = search_targetpixelfile('KIC 6679295').download_all()
```
We've now created a list of `KeplerTargetPixelFile` objects, where each item is a different quarter. We're going to be able to combine these just like in the [stitching tutorial](03-appending-lightcurves.html).
Let's take a look at just one of those target pixel files.
```
# Build the light curve
pipeline_lc = tpfs[0].to_lightcurve().flatten()
for tpf in tpfs[1:]:
pipeline_lc = pipeline_lc.append(tpf.to_lightcurve().flatten())
# Clean the light curve
pipeline_lc = pipeline_lc.remove_nans().remove_outliers()
```
Above we have created the light curve from the target pixel files, stitched them all together in the same way as in the [stitching tutorial] using lightkurves `append` function. To recap the steps we:
* Convert to a `KeplerLightCurve` object with `to_lightcurve()`
* Remove NaNs with `remove_nans()`
* Remove long term trends with `flatten()`
* Remove outliers with simple sigma clipping using `remove_outliers()`
The period for this planet candidate is 24.57537 days. Let's plot it up and take a look.
```
pipeline_lc.fold(period=24.57537, t0=21.3).bin().errorbar()
plt.xlim(-0.015, 0.015)
plt.ylim(0.998, 1.0015);
```
Looks like a great candidate. However, we might just want to check on the pixels. Let's plot one of the target pixel files.
```
tpf.plot(frame=100, aperture_mask=tpf.pipeline_mask, mask_color='red');
```
The Kepler Pipeline aperture is in red. It looks like there is a nearby contaminate star! We might want to check that the signal is not really coming from the bright, nearby contaminant, rather than our target star. Let's use the top right corner four pixels as our new mask.
```
import numpy as np
aper = np.zeros(tpf.shape[1:], dtype=np.int)
aper[-2:, 0:2] = 1
tpf.plot(aperture_mask=aper, mask_color='red');
```
The new mask covers the bright star. Now we can iterate through the target pixel files and build the light curve in the same way as before, but this time
```
# Build the NEW aperture, and the light curve
aper = np.zeros(tpfs[0].shape[1:])
aper[-2:, 0:2] = 1
user_lc = tpfs[0].to_lightcurve(aperture_mask=aper.astype(bool)).flatten()
for tpf in tpfs[1:]:
aper = np.zeros(tpf.shape[1:])
aper[-2:, 0:2]=1
user_lc = user_lc.append(tpf.to_lightcurve(aperture_mask=aper.astype(bool)).flatten())
# Clean the light curve
user_lc = user_lc.remove_nans().remove_outliers()
```
Now we have our new light curve we can plot it up again and find out if there is still a planet signal.
```
user_lc.fold(period=24.57537, t0=-0.133).bin().errorbar();
plt.xlim(-0.015,0.015)
plt.ylim(0.998,1.0015)
```
Looks like the planet signal is only in the target star and doesn't belong to the contaminant. This is just one of many checks you might want to perform to validate your planet candidates!
| github_jupyter |
# Bytes Data Type
**ToDo**:
- Add an illustration and explain the concept of UTF-8, Unicode, Bytes, ASCII - Similar to [this](https://blog.finxter.com/wp-content/uploads/2020/06/byte-1024x576.jpg)
- Add relevant resources at the end
---
Most cryptographic functions require [Bytes](https://docs.python.org/3/library/stdtypes.html#bytes-objects) objects. In the case of strings, the [`encode`](https://docs.python.org/3/library/stdtypes.html#str.encode) and [`decode`](https://docs.python.org/3/library/stdtypes.html#bytes.decode) methods can be used, however for custom objects, two possible ways are:
1. Converting the object to JSON and then convert the JSON string to bytes
1. Implementing a `__bytes__` method
However, usually hashes are used with plain types like strings which are easily convertible to bytes.
## Data Conversions
```
data_string = "Hello World!"
data_bytes = data_string.encode("utf-8")
data_hex = data_bytes.hex()
data_decoded = data_bytes.decode("utf-8")
data_hex_bytes = bytes.fromhex(data_hex)
print(f" Original String: {data_string}")
print(f"From String to Bytes: {data_bytes}")
print(f" From Bytes to Hex: {data_hex}")
print(f"From Bytes to String: {data_decoded}")
print(f" From Hex to Bytes: {data_hex_bytes}")
```
### Using the `binascii` module
The [binascii module](https://docs.python.org/3/library/binascii.html) exposes two utility functions, one to convert from bytes to hex called [hexlify](https://docs.python.org/3/library/binascii.html#binascii.hexlify) and another to do the reverse conversion called [unhexlify](https://docs.python.org/3/library/binascii.html#binascii.unhexlify)
**Important note:** the hexlify function returns a bytes object whereas the .hex() method of bytes returns a string. See the `b` before the `'`
```
import binascii
data_string = "Hello World!"
data_bytes = data_string.encode("utf-8")
data_hex = binascii.hexlify(data_bytes)
data_hex_string = binascii.unhexlify(data_hex)
print(f" Original String: {data_string}")
print(f"From String to Bytes: {data_bytes}")
print(f" From Bytes to Hex: {data_hex}")
print(f" From Hex to Bytes: {data_hex_bytes}")
```
## Examples
```
import hashlib
```
#### Example with plain strings
```
data = "Hello World!"
data_bytes = data.encode("utf-8")
data_decoded = data_bytes.decode("utf-8")
data_hashed = hashlib.sha256(data_bytes).hexdigest()
print(f"Original: {data}")
print(f" Encoded: {data_bytes}")
print(f" Decoded: {data_decoded}")
print(f" Hashed: {data_hashed}")
```
#### Example with Custom objects
```
from dataclasses import dataclass, asdict
import json
@dataclass
class Person:
first_name: str
last_name: str
@property
def fullname(self):
return f"{self.last_name}, {self.first_name}"
def __bytes__(self):
dictionary_representation = asdict(self)
json_representation = json.dumps(dictionary_representation)
return json_representation.encode("utf-8")
@classmethod
def from_bytes(cls, bytes_object):
string_representation = bytes_object.decode("utf-8")
dictionary_representation = json.loads(string_representation)
return cls(**dictionary_representation)
person = Person("John", "Doe")
person_bytes = bytes(person)
person_decoded = Person.from_bytes(person_bytes)
person_hashed = hashlib.sha256(person_bytes).hexdigest()
print(f" Original: {person}")
print(f" Encoded: {person_bytes}")
print(f" Decoded: {person_decoded}")
print(f"Full Name: {person_decoded.fullname}")
print(f" Hashed: {person_hashed}")
```
#### Example with Mixins
In bigger projects, it may be against best practice to duplicate the `__bytes__` and `from_bytes` methods. A [Mixin](https://en.wikipedia.org/wiki/Mixin?oldformat=true) class can be used and then inherit from. Mixins are special class, similar to a [Protocol](https://www.python.org/dev/peps/pep-0544/) which define methods to be used in child classes. Mixins as opposed to Interfaces or Abstract Classes are meant to be incomplete and it should not make sense to instanciate them directly.
**Note**: In this case, it would have been possible to make `PersonBase` inherit from `ByteConvertibleMixin` and thus avoiding the multiple-inheritance. However, if the bytes conversion is desirable only to a subset of children classes of `PersonBase`, then the multiple-inheritance approach is the most idiomatic in Python.
```
from abc import ABC
from dataclasses import dataclass, asdict
import json
@dataclass
class ByteConvertibleMixin(ABC):
def __bytes__(self):
dictionary_representation = asdict(self)
json_representation = json.dumps(dictionary_representation)
return json_representation.encode("utf-8")
@classmethod
def from_bytes(cls, bytes_object):
string_representation = bytes_object.decode("utf-8")
dictionary_representation = json.loads(string_representation)
return cls(**dictionary_representation)
@dataclass
class PersonBase: # Name changed to avoid overwriting
first_name: str
last_name: str
@property
def fullname(self):
return f"{self.last_name}, {self.first_name}"
@dataclass
class Customer(PersonBase, ByteConvertibleMixin): # Multiple-Inheritance
address: str
customer = Customer("John", "Doe", "Neverland 10")
customer_bytes = bytes(customer)
customer_decoded = Customer.from_bytes(customer_bytes)
customer_hashed = hashlib.sha256(customer_bytes).hexdigest()
print(f" Original: {customer}")
print(f" Encoded: {customer_bytes}")
print(f" Decoded: {customer_decoded}")
print(f"Full Name: {customer_decoded.fullname}")
print(f" Hashed: {customer_hashed}")
```
## Special Case: Files
When working with files, it is possible to work in bytes format out of the box. The easiest way is through the [`read_bytes`](https://docs.python.org/3/library/pathlib.html#pathlib.Path.read_bytes) method of the [`pathlib`](https://docs.python.org/3/library/pathlib.html) module.
```
from pathlib import Path
image_bytes = Path("../_static/images/certificate_details_dns.png").read_bytes()
image_bytes = image_bytes[:150] # Trimmed to improve display
print(f"Byte Representation: \n{image_bytes}\n")
print(f"Hex Representation: \n{image_bytes.hex()}")
```
## Random Bytes
There are many ways to generate randomness or pseudo-randomness, some of which are considered **insecured** and other **secure**.
### Using the `secrets` module
Python 3.6 introduced the [`secrets`](https://docs.python.org/3/library/secrets.html) module to conveniently generate several types of **secure** random bytes.
The relevant methods are the `token_*` methods, each receives a lenght parameter. The more bytes, the safer the token, see [this resource](https://docs.python.org/3/library/secrets.html#how-many-bytes-should-tokens-use) for more information. Moreover, this [video](https://www.youtube.com/watch?v=S9JGmA5_unY) illustrates how secure 32 bytes (256bits) randomness is.
When using only hexadecimal, there will be 2 characters per byte, to generate shorter strings but at the same time being able to insert them in URL (e.g. for password reset tokens), the `token_urlsafe` can be used, which will yield a string approximately 25% shorter
There are other ways to generate random bytes in Python but using secrets is common practice since Python 3.6. For other options see this [detailed answer](https://stackoverflow.com/questions/42190663/questions-about-python3-6-os-urandom-os-getrandom-secrets).
```
import secrets
lenght = 15
print(f" Secure Random Bytes: {secrets.token_bytes(lenght)}")
print(f" Secure Random Bytes (Hex): {secrets.token_hex(lenght)}")
print(f"Secure Random Bytes (Hex URLSafe): {secrets.token_urlsafe(lenght)}")
```
### Comparing Secrets
To avoid [timming attacks](https://www.wikiwand.com/en/Timing_attack), it is important to **NOT** use `==` when comparing secrets. For that the `secrets` module exposes a method [`compare_digest`](https://docs.python.org/3/library/secrets.html#secrets.compare_digest) which is actually an alias of the [`hmac`](https://docs.python.org/3/library/hmac.htm) module [homonymous method](https://docs.python.org/3/library/hmac.html#hmac.compare_digest).
For a demonstration of this type of attack, see this [demo](https://www.youtube.com/watch?v=XThL0LP3RjY).
```
# Excesively large lenght for better illustration
lenght = 1000
real_token = secrets.token_bytes(lenght)
guess_token_all_wrong = secrets.token_bytes(lenght)
guess_token_all_but_one = real_token[:-1] + secrets.token_bytes(1)
print(f"Is short guess the real? {secrets.compare_digest(real_token, guess_token_all_wrong)}")
print(f"Is long guess the real? {secrets.compare_digest(real_token, guess_token_all_but_one)}")
print(f"Is real guess the real? {secrets.compare_digest(real_token, real_token)}")
```
## Conclusion
To avoid duplicated work, it is important to work with standards, in the case of security and encryption, that standard is the Bytes format. All methods and algorithms work with bytes objects and therefore it is important to know how to handle them while programming.
Python has several tools like `bytes`, `binascii` and `secrets` to work, generate and convert bytes. It is also possible to define conversion for custom objects through the `__bytes__` magic method. The `pathlib` module also allows to read files as bytes out of the box.
| github_jupyter |
# Process the Unsplash dataset with CLIP
This notebook processes all the downloaded photos using OpenAI's [CLIP neural network](https://github.com/openai/CLIP). For each image we get a feature vector containing 512 float numbers, which we will store in a file. These feature vectors will be used later to compare them to the text feature vectors.
This step will be significantly faster if you have a GPU, but it will also work on the CPU.
## Load the photos
Load all photos from the folder they were stored.
```
from pathlib import Path
# Set the path to the photos
dataset_version = "lite" # Use "lite" or "full"
photos_path = Path("unsplash-dataset") / dataset_version / "photos"
# List all JPGs in the folder
photos_files = list(photos_path.glob("*.jpg"))
# Print some statistics
print(f"Photos found: {len(photos_files)}")
```
## Load the CLIP net
Load the CLIP net and define the function that computes the feature vectors
```
import clip
import torch
from PIL import Image
# Load the open CLIP model
device = "cuda" if torch.cuda.is_available() else "cpu"
model, preprocess = clip.load("ViT-B/32", device=device)
# Function that computes the feature vectors for a batch of images
def compute_clip_features(photos_batch):
# Load all the photos from the files
photos = [Image.open(photo_file) for photo_file in photos_batch]
# Preprocess all photos
photos_preprocessed = torch.stack([preprocess(photo) for photo in photos]).to(device)
with torch.no_grad():
# Encode the photos batch to compute the feature vectors and normalize them
photos_features = model.encode_image(photos_preprocessed)
photos_features /= photos_features.norm(dim=-1, keepdim=True)
# Transfer the feature vectors back to the CPU and convert to numpy
return photos_features.cpu().numpy()
```
## Process all photos
Now we need to compute the features for all photos. We will do that in batches, because it is much more efficient. You should tune the batch size so that it fits on your GPU. The processing on the GPU is fairly fast, so the bottleneck will probably be loading the photos from the disk.
In this step the feature vectors and the photo IDs of each batch will be saved to a file separately. This makes the whole process more robust. We will merge the data later.
```
import math
import numpy as np
import pandas as pd
# Define the batch size so that it fits on your GPU. You can also do the processing on the CPU, but it will be slower.
batch_size = 16
# Path where the feature vectors will be stored
features_path = Path("unsplash-dataset") / dataset_version / "features"
# Compute how many batches are needed
batches = math.ceil(len(photos_files) / batch_size)
# Process each batch
for i in range(batches):
print(f"Processing batch {i+1}/{batches}")
batch_ids_path = features_path / f"{i:010d}.csv"
batch_features_path = features_path / f"{i:010d}.npy"
# Only do the processing if the batch wasn't processed yet
if not batch_features_path.exists():
try:
# Select the photos for the current batch
batch_files = photos_files[i*batch_size : (i+1)*batch_size]
# Compute the features and save to a numpy file
batch_features = compute_clip_features(batch_files)
np.save(batch_features_path, batch_features)
# Save the photo IDs to a CSV file
photo_ids = [photo_file.name.split(".")[0] for photo_file in batch_files]
photo_ids_data = pd.DataFrame(photo_ids, columns=['photo_id'])
photo_ids_data.to_csv(batch_ids_path, index=False)
except:
# Catch problems with the processing to make the process more robust
print(f'Problem with batch {i}')
```
Merge the features and the photo IDs. The resulting files are `features.npy` and `photo_ids.csv`. Feel free to delete the intermediate results.
```
import numpy as np
import pandas as pd
# Load all numpy files
features_list = [np.load(features_file) for features_file in sorted(features_path.glob("*.npy"))]
# Concatenate the features and store in a merged file
features = np.concatenate(features_list)
np.save(features_path / "features.npy", features)
# Load all the photo IDs
photo_ids = pd.concat([pd.read_csv(ids_file) for ids_file in sorted(features_path.glob("*.csv"))])
photo_ids.to_csv(features_path / "photo_ids.csv", index=False)
```
| github_jupyter |
```
import requests, datetime, time, pytz
from pyquery import PyQuery as pq
from dataflows import Flow, printer, dump_to_path, sort_rows
def get_messages(before_id=None):
url = 'https://t.me/s/MOHreport'
if before_id:
url += '?before=' + str(before_id)
print('loading ' + url)
for message in pq(requests.get(url).text)('[data-post]'):
message_id = int(message.attrib['data-post'].replace('MOHreport/', ''))
date_elts = message.find_class('tgme_widget_message_date')
assert len(date_elts) == 1
date_elt = date_elts[0]
message_datetime = next(date_elt.iterchildren()).attrib['datetime']
message_datetime = "".join(reversed("".join(reversed(message_datetime)).replace(':','',1)))
message_datetime = datetime.datetime.strptime(message_datetime, '%Y-%m-%dT%H:%M:%S%z').astimezone(pytz.timezone('Asia/Jerusalem'))
content_elts = message.find_class('tgme_widget_message_bubble')
assert len(content_elts) == 1
content_elt = content_elts[0]
message_htmls = []
image_urls = []
for child in content_elt.iterchildren():
if 'tgme_widget_message_text' in list(child.classes):
message_htmls.append(pq(child).html())
elif 'tgme_widget_message_photo_wrap' in list(child.classes):
image_urls.append(child.attrib['style'].split("url('")[1].split("'")[0])
message_html = "<br/><br/>".join(message_htmls)
message_text = message_html.replace('<br/>', "\n")
image_urls = ",".join(image_urls)
yield {'id': message_id, 'date': message_datetime, 'text': message_text, 'images': image_urls}
def get_all_messages(min_message_id=2525):
last_message_id = None
num_messages = 0
while True:
if num_messages > 0 and num_messages % 500 == 0: print('Loaded ' + str(num_messages) + ' messages..')
if last_message_id and last_message_id <= min_message_id: break
for message in get_messages(last_message_id):
if not last_message_id or message['id'] < last_message_id:
last_message_id = message['id']
yield message
num_messages += 1
print('sleeping .1 seconds..')
time.sleep(.1)
Flow(
get_all_messages(),
sort_rows('{date}', reverse=True),
printer(tablefmt='html', num_rows=1),
dump_to_path('data/MOHReport')
).process()
import os
CKAN_URL = 'https://www.odata.org.il'
if os.environ.get('CKAN_API_KEY'):
CKAN_API_KEY = os.environ['CKAN_API_KEY']
else:
import getpass
CKAN_API_KEY = getpass.getpass('CKAN_API_KEY')
from dataflows import load
import json
data = Flow(
load('data/MOHReport/datapackage.json')
).results()[0][0]
def format_row(row):
row['date'] = row['date'].strftime('%Y-%m-%dT%H:%M:%S')
row['images'] = '' if not row['images'] else row['images']
return row
records = [format_row(row) for row in data]
print(records[0])
res = requests.post('https://www.odata.org.il/api/3/action/datastore_create', json={
'resource_id': 'ce4c9482-cd3a-485b-af56-d3d7118a7552',
'force': True,
'primary_key': ['id'],
}, headers={'Authorization':CKAN_API_KEY})
print(res.status_code)
print(res.text)
assert res.status_code == 200
res = requests.post('https://www.odata.org.il/api/3/action/datastore_upsert', json={
'resource_id': 'ce4c9482-cd3a-485b-af56-d3d7118a7552',
'records': records,
'method': 'upsert',
'force': True
}, headers={'Authorization':CKAN_API_KEY})
print(res.status_code)
# print(res.text)
assert res.status_code == 200
if os.environ.get('SERVICE_ACCOUNT_FILE'):
SERVICE_ACCOUNT_FILE = os.environ['SERVICE_ACCOUNT_FILE']
else:
import getpass
SERVICE_ACCOUNT_FILE = getpass.getpass('SERVICE_ACCOUNT_FILE')
from google.oauth2 import service_account
import googleapiclient.discovery
SCOPES = ['https://www.googleapis.com/auth/drive']
credentials = service_account.Credentials.from_service_account_file(SERVICE_ACCOUNT_FILE, scopes=SCOPES)
spreadsheets = googleapiclient.discovery.build('sheets', 'v4', credentials=credentials).spreadsheets()
sheets_values = spreadsheets.values().get(spreadsheetId='19pKanFwuABaNyPGISihcqcPuFcIj5e3svcnr5LLm1ns', range='res_1!A:D').execute()['values']
sheets_data = {}
for rownum, row in enumerate(sheets_values):
if rownum == 0: continue
sheets_data[int(row[0])] = {
'id': int(row[0]),
'date': row[1],
'text': row[2] if len(row) == 3 else '',
'images': row[3] if len(row) == 4 else ''
}
for row in sorted(data, key=lambda row: row['id']):
if row['id'] not in sheets_data:
value_input_option = 'RAW'
insert_data_option = 'INSERT_ROWS'
value_range_body = {
"values": [[row['id'], row['date'], row['text'], row['images']]]
}
request = spreadsheets.values().append(spreadsheetId='19pKanFwuABaNyPGISihcqcPuFcIj5e3svcnr5LLm1ns',
range='res_1!A:D',
valueInputOption=value_input_option,
insertDataOption=insert_data_option,
body=value_range_body)
request.execute()
time.sleep(2)
```
| github_jupyter |
# LUSD Pool Model
```
import numpy as np
from matplotlib import pyplot as plt
from matplotlib import dates as md
from matplotlib import ticker
import scipy as scp
import scipy.optimize as opt
import csv
import math
import random
import pandas as pd
import copy
from datetime import datetime, timedelta
```
## Core Idea
The core idea of a pool is a LUSD : staBAL pool, which utilizes StableSwap invariant and provides low slippage tool for swapping tokens equal in price (as LUSD and staBAL are both close to 1\\$: LUSD = 1.01\\$ and staBAL = 0.99$). staBAL is a BPT token of Balancer stable pool with 3 stable tokens: USDC, USDT and DAI.
Such pool has a potential to be a high-profitable mining/income tool. That is because Balacner architecture allows to stake liquidity tokens of the pool into other pools and investing mechanisms, though still maintaining the pool as if all the tokens still remain in the pool. In other words, LUSD tokens from the pool could be staked into Liquidity Stability Pool for additional profit (with some reserve left in the original pool to maintain small swaps) and perform liquidity mining for the liquidity providers of the original pool. This profits will sum with the conventional profits from swap fees and liquidity provision in the original pool, effectively adding APRs. At the time of the document creation, expected APR of Liquity Stability Pool staking is ~13%.
## Math
The following code block is Balancer's stablepool logic
```
#pool functions
def calculateInvariant_b(A, pool, roundUp):
n = len(pool)
S = 0
for i in range(0, n):
if (not(isinstance(pool[i], int))):
raise TypeError('Not int in pool')
S = S + pool[i]
prevD = 0
D = S
An = mul(A, pow(n, n))
for i in range(0, 255):
P_D = pool[0]*n
for j in range (1, n):
P_D = div(mul(mul(P_D, pool[j]), n), D, roundUp)
prevD = D
D = div(
add(mul(mul(n, D), D), mul(mul(An, S), P_D)
),
add(mul(n + 1, D), mul(An - 1, P_D)
),
roundUp
)
if (D > prevD):
if (D - prevD <= 1):
return D
elif (prevD - D <= 1):
return D
raise NoConvergenceException('D didn\'t converge, error = ', abs(D - prevD))
def calculateInvariant(A, pool):
n = len(pool)
S = sum(pool)
P = 1
for i in range(0, n):
P = P*pool[i]
t = pow(n, n)
D = S
T = A*t
for i in range(0, 255):
prevD = D
K = pow(D, n)/(t*P)
D = (n*K*D + T*S)/(T + (n+1)*K - 1)
if (D > prevD):
if (D - prevD <= 1):
return D
elif (prevD - D <= 1):
return D
raise NoConvergenceException('D didn\'t converge')
def getTokenBalanceGivenInvariantAndAllOtherBalances_b(A, pool, D, tokenindex):
n = len(pool)
S = pool[0]
P_D = pool[0]*n
for j in range(1, n):
P_D = divDown(mul(mul(P_D, pool[j]), n), D)
S = add(S, pool[j])
S = sub(S, pool[tokenindex])
D2 = mul(D, D)
An = int(A*pow(n, n))
c = mul(
divUp(D2, mul(An, P_D)),
pool[tokenindex]
)
b = add(S,divDown(D, An))
prevTokenBalance = 0
tokenBalance = divUp(add(D2, c), add(D, b))
for i in range(0, 255):
prevTokenBalance = tokenBalance
tokenBalance = divUp(
add(mul(tokenBalance, tokenBalance), c),
sub(add(mul(tokenBalance, 2), b), D)
)
if (tokenBalance > prevTokenBalance):
if (tokenBalance - prevTokenBalance <= 1000):
return tokenBalance
elif (prevTokenBalance - tokenBalance <= 1000):
return tokenBalance
raise NoConvergenceException('balance didn\'t converge, error = ', abs(tokenBalance - prevTokenBalance))
def getTokenBalanceGivenInvariantAndAllOtherBalances(A, pool, D, tokenindex):
n = len(pool)
S = 0
P = 1
for i in range(0, n):
if (i != tokenindex):
S = S + pool[i]
P = P*pool[i]
t = pow(n, n) #N
G = A*t #A
K = pow(D, n+1)/(P*t)
L = G*S + D - G*D
x = D
for i in range(0, 255):
x = (pow(x, 2)*G + K)/(L + 2*x*G)
return x
def calcOutGivenIn_b(A, pool, tokenin, tokenout, amountin):
D = calculateInvariant_b(A, pool, True)
pool[tokenin] = add(pool[tokenin], amountin)
finalBalanceOut = getTokenBalanceGivenInvariantAndAllOtherBalances_b(A, pool, D, tokenout)
pool[tokenin] = pool[tokenin] - amountin
return sub(sub(pool[tokenout], finalBalanceOut), 1)
def calcOutGivenIn(A, pool, tokenin, tokenout, amountin):
D = calculateInvariant(A, pool)
pool[tokenin] = pool[tokenin] + amountin
xout = getTokenBalanceGivenInvariantAndAllOtherBalances(A, pool, D, tokenout)
pool[tokenin] = pool[tokenin] - amountin
return pool[tokenout] - xout
def calcInGivenOut_b(A, pool, tokenin, tokenout, amountout):
D = calculateInvariant_b(A, pool, True)
pool[tokenout] = sub(pool[tokenout], amountout)
finalBalanceIn = getTokenBalanceGivenInvariantAndAllOtherBalances_b(A, pool, D, tokenin)
pool[tokenout] = pool[tokenout] + amountout
return add(sub(finalBalanceIn, pool[tokenin]), 1)
def calcInGivenOut(A, pool_, tokenin, tokenout, amountout):
pool = copy.deepcopy(pool_)
D = calculateInvariant(A, pool)
pool[tokenout] = pool[tokenout] - amountout
finalBalanceIn = getTokenBalanceGivenInvariantAndAllOtherBalances(A, pool, D, tokenin)
return finalBalanceIn - pool[tokenin]
def calcBptOutGivenExactTokensIn(A, pool, amountsin, bpttotalsupply, swapfeepercentage):
n = len(pool)
n2 = len(amountsin)
if (n != n2):
print('pool and amounts_in have different dimensions: ', n, n2)
return
S = pool[0]
for j in range(1, n):
S = add(S, pool[j])
balanceRatiosWithFee = [0 for i in range(n)]
invariantRatioWithFees = 0
for i in range(0, n):
currentWeight = divDown_f(pool[i], S)
balanceRatiosWithFee[i] = divDown_f(add(pool[i], amountsin[i]), pool[i])
invariantRatioWithFees = add(invariantRatioWithFees, mulDown_f(balanceRatiosWithFee[i], currentWeight))
print(balanceRatiosWithFee)
print(invariantRatioWithFees)
newpool = [0 for i in range(n)]
for i in range(0, n):
if (balanceRatiosWithFee[i] > invariantRatioWithFees):
nonTaxableAmount = mulDown_f(pool[i], sub(invariantRatioWithFees, ONE))
taxableAmount = sub(amountsin[i], nonTaxableAmount)
amountInWithoutFee = add(nonTaxableAmount, mulDown_f(taxableAmount, ONE - swapfeepercentage))
else:
amountInWithoutFee = amountsin[i]
newpool[i] = add(pool[i], amountInWithoutFee)
currentD = calculateInvariant_b(A, pool, True)
newD = calculateInvariant_b(A, newpool, False)
invariantRatio = divDown_f(newD, currentD)
if (invariantRatio > 1):
return mulDown_f(bpttotalsupply, (invariantRatio - ONE))
else:
return 0
def calcTokenInGivenExactBptOut(A, pool, token, bptamountout, bpttotalsupply):
D = calculateInvariant(A, pool)
newD = (bpttotalsupply + bptamountout)/(bpttotalsupply)*D
newBalanceToken = getTokenBalanceGivenInvariantAndAllOtherBalances(A, pool, newD, token)
amountInWithoutFee = newBalanceToken - pool[token]
S = sum[pool]
currentWeight = pool[token]/S
taxablePercentage = 1 - currentWeight
taxableAmount = amountInWithoutFee*taxablePercentage
nonTaxableAmount = amountInWithoutFee - taxableAmount
return nonTaxableAmount + taxableAmount/(1-swapfeepercentage)
def calcBptInGivenExactTokensOut(A, pool, amountsout, bpttotalsupply):
S = sum(pool)
balanceRatiosWithoutFee = []
invariantRatioWithoutFees = 0
for i in range(0, n):
currentWeight = pool[i]/S
balanceRatiosWithoutFee[i] = (pool[i]-amountsout[i])/pool[i]
invariantRatioWithoutFees = invariantRatioWithoutFees + balanceRatiosWithoutFee[i]*currentWeight
newpool = []
for i in range(0, n):
if (invariantRatioWithoutFees > balanceRatiosWithoutFee[i]):
nonTaxableAmount = pool[i]*(1-invariantRatioWithoutFees)
taxableAmount = amountsout[i] - nonTaxableAmount
amountOutWithFee = nonTaxableAmount + taxableAmount/(1-swapfeepercentage)
else:
amountOutWithFee = amountsout[i]
newpool[i] = pool[i] - amountOutWithFee
D = calculateInvariant(A, pool)
newD = calculateInvariant(A, newpool)
invariantRatio = newD/D
return bpttotalsupply*(1-invariantRatio)
def calcTokenOutGivenExactBptIn(A, pool, token, bptAmountIn, bptTotalSupply, swapfeepercentage):
D = calculateInvariant(A, pool)
newD = (bptTotalSupply - bptAmountIn)/bptTotalSupply*D
newBalanceToken = getTokenBalanceGivenInvariantAndAllOtherBalances(A, pool, newD, token)
amountOutWithoutFee = pool[token] - newBalanceToken
S = sum[pool]
currentWeight = pool[token]/S
taxablePercentage = (1-currentWeight)
taxableAmount = amountOutWithoutFee*taxablePercentage
nonTaxableAmount = amountOutWithoutFee - taxableAmount
return nonTaxableAmount + taxableAmount*(1-swapfeepercentage)
def calcTokensOutGivenExactBptIn(pool, bptAmountIn, bptTotalSupply):
bptRatio = bptAmountIn/bptTotalSupply
amountsOut = []
for i in range(0, n):
amountsOut[i] = pool[i]*bptRatio
return amountsOut
def calcDueTokenProtocolSwapFeeAmount(A, pool, lastD, token, protocolSwapFeePercentage):
finalBalanceFeeToken = getTokenBalanceGivenInvariantAndAllOtherBalances(A, pool, lastD, token)
if (pool[token] <= finalBalanceFeeToken):
return 0
accumulatedTokenSwapFees = pool[token] - finalBalanceFeeToken
return accumulatedTokenSwapFees*protocolSwapFeePercentage/(1)
```
# Pool Model Based on Previous Data
For LUSD part of the Pool, we take the existing data on operation of *Liquity* protocol from July 1st, 2021 to September 9th, 2021. The pool acts as if "it has always been there".
### LUSD operations
* LQTY rewards are calculated based on the Pool's share of LUSD in the Stability Pool. The amount of LQTY rewards is calculated as follows:
$$
SP_{share} = \frac{LUSD_{Pool}^{i}}{LUSD_{StabilityPool}^{i}};\\
\\
LQTY_{gained}^{i} = (LQTY_{total}^{i} - LQTY_{total}^{i-1})\times SP_{share},
$$
where upper index $i$ denotes the timestamp of the calculation. The timestamps are exactly 1 hour difference, except the events of liquidations.
* Liquidation rewards are calculated basically the same way, taking into consideration the mathematics behind the collateral distibution and LUSD burning.
$$
COL_{distributed}^{i} = COL_{liquidated}^{i}\times(1 - 0.05)\cdot100\%;\\
COL_{gained}^{i} = COL_{distributed}^{i}\times SP_{share}^{i};\\
LUSD_{lost}^{i} = LUSD_{burned}^{i}\times SP_{share}^{i}.
$$
* Trades of LQTY token occur when a minimum resonable amount is collected in the Pool. It is almost impossible to account for slippage and trading fees, thus every trading operation is multiplyed by loss coefficient:
$$
LUSD_{bought}^{i} = \frac{LQTY_{sold}^{i}\times{LQTY_price}^{i}}{LUSD_{price}^{i}}\times L; \hspace{3mm} L = 0.9
$$
### staBAL operations
Due to lack of the data on staBAL, following assumptions have been made:
* staBAL price rises linearly at a rate of 1% per year from initial start at 1\$:
$$
staBAL_{price}^{i} = staBAL_{price}^{i-1}\cdot \frac{(1 + 0.01)}{365\times 24}
$$
* staBAL APR for BAL rewards is roughly estimated at 7\%.
* BAL prices have been obtained. The logic of selling BAL tokens for staBAL is the same as for LQTY tokens.
### Pool Initial data
We start as a Pool of 50/50 LUSD/staBAL.
* $TVL = \$ 60M$
### Creating Dataframes of Different Statistics
##### Opening files
```
Total_LUSD_Supply = pd.read_csv('data/Total LUSD Supply.csv')
LUSD_Utilization = pd.read_csv('data/LUSD Utilization.csv')
Total_LQTY_Staked = pd.read_csv('data/Total LQTY Staked.csv')
Liquidations_ = pd.read_csv('data/Recent Liquidations.csv')
Redeems_ = pd.read_csv('data/Recently Redeemed Troves.csv')
start_date = datetime(2021, 7, 1, 0, 0, 0) #1th of July, 2021, 00:00
start_date = pd.to_datetime(start_date, utc=True)
end_date = datetime(2021, 9, 29, 11, 0, 0) #29th of September, 2021, 11:00
end_date = pd.to_datetime(end_date, utc=True)
dateColumn = pd.date_range(start_date, end_date, freq='H')
dateColumn = pd.DataFrame(dateColumn, columns=['date',])
dateColumn
```
##### Creating LUSD dataframe
```
LUSD = pd.DataFrame()
LUSD['date'] = pd.to_datetime(LUSD_Utilization['hour'])
LUSD['LUSD in SP'] = LUSD_Utilization['stabilityPool']
LUSD['LUSD other'] = LUSD_Utilization.iloc[:, 1:].sum(1, numeric_only=True) - LUSD_Utilization['stabilityPool']
LUSD = LUSD.merge(right=dateColumn, on='date', how='right')
LUSD.head()
```
##### Creating LQTY dataframe
```
LQTY = pd.DataFrame()
LQTY['date'] = pd.to_datetime(Total_LQTY_Staked['hour'])
LQTY['LQTY staked'] = Total_LQTY_Staked['totalLQTYStaked']
LQTY['LQTY circulating'] = Total_LQTY_Staked['totalLQTYClaimed'] - Total_LQTY_Staked['totalLQTYStaked']
LQTY['LQTY total'] = Total_LQTY_Staked['totalLQTYClaimed']
LQTY = LQTY.merge(right=dateColumn, on='date', how='right')
LQTY.head()
```
##### Creating Liquidations dataframe
```
Liquidations = pd.DataFrame()
Liquidations['date'] = pd.to_datetime(Liquidations_['timestamp'])
Liquidations['LIQ col'] = Liquidations_['collateral']
Liquidations['LIQ debt'] = Liquidations_['debt']
Liquidations['LIQ price'] = Liquidations_['price']
Liquidations['LIQ CR'] = Liquidations_['collateralRatio']
Liquidations['LIQ mode'] = Liquidations_['mode']
Liquidations = Liquidations[Liquidations['date'] >= start_date].merge(right=dateColumn, on='date', how='outer')
Liquidations.sort_values(by='date', ignore_index=True, inplace=True)
Liquidations.loc[:, 'LIQ col':'LIQ CR'] = Liquidations.loc[:, 'LIQ col':'LIQ CR'].fillna(value=0)
Liquidations.loc[:, 'LIQ mode'] = Liquidations.loc[:, 'LIQ mode'].fillna(value = 'none')
Liquidations.head()
```
##### Creating Redemption dataframe
```
Redeems = pd.DataFrame()
Redeems['date'] = pd.to_datetime(Redeems_['timestamp'])
Redeems['col'] = Redeems_['collateral']
Redeems['debt'] = Redeems_['debt']
Redeems['price'] = Redeems_['price']
Redeems['CR'] = Redeems_['collateralRatio']
Redeems['ETH Redeemed'] = Redeems_['ethRedeemed']
Redeems['LUSD Redeemed'] = Redeems_['lusdRedeemed']
Redeems = Redeems[Redeems['date'] >= start_date].merge(right=dateColumn, on='date', how='outer')
Redeems.sort_values(by='date', ignore_index=True, inplace=True)
Redeems.fillna(value=0, inplace=True)
Redeems.head()
```
## Loading Prices
##### Loading ETH price
```
ETHprice = pd.read_csv('./data/ETH price.csv')
ETHprice.drop(ETHprice.loc[:, 'high':'close'], axis=1, inplace=True)
ETHprice['timestamp'] = pd.to_datetime(ETHprice['timestamp'], unit = 's', utc=True )
ETHprice = ETHprice.sort_values(by='timestamp', ascending=True, ignore_index=True)
ETHprice.rename(columns = {'timestamp':'date', 'open':'ETH price'}, inplace=True)
ETHprice
```
##### Loading BAL price
```
BALprice = pd.read_csv('./data/BAL price.csv')
BALprice.rename(columns={'timestamp': 'date', 'close':'BAL price'}, inplace=True)
BALprice = BALprice.loc[:, ['date','BAL price']]
BALprice.loc[:, 'date'] = pd.to_datetime(BALprice.loc[:, 'date'])
BALprice.sort_values(by='date', ascending=True, inplace=True, ignore_index=True)
BALprice = BALprice.merge(right=dateColumn, how='inner')
BALprice
```
##### Loading staBAL prices
```
staBAL_price_initial = 1
staBALdata = {
'APR' : 0.07
}
staBAL = copy.deepcopy(dateColumn)
#staBAL['date'] = dateColumn
staBAL['staBAL price'] = staBAL_price_initial
for i in range(1, len(staBAL)):
staBAL.loc[i, 'staBAL price'] = staBAL['staBAL price'][i-1]*(1 + 0.01/365/24)
staBAL
```
### Merging All Data into Single Dataframe
##### Creating main dataframe and merging info
```
Data = pd.DataFrame()
Data = copy.deepcopy(dateColumn)
Data = Data.merge(LUSD, how='outer', on='date')
Data = Data.merge(LQTY, how='outer', on='date')
Data = Data.merge(Liquidations, how='outer', on='date')
Data = Data.merge(ETHprice.loc[:, ['date', 'ETH price']], on='date', how='outer')
Data = Data.merge(BALprice, on='date', how='outer')
Data = Data.merge(staBAL, on='date', how='outer')
Data.sort_values(by=['date'], ignore_index=True, inplace=True)
Data[['ETH price', 'BAL price']] = Data[['ETH price', 'BAL price']].fillna(method='ffill')
Data[Data.columns.drop(['ETH price', 'LIQ mode'])] = Data[Data.columns.drop(['ETH price', 'LIQ mode'])].fillna(method='ffill')
Data
```
## Initializing Pool with 50/50 $ 60M Total TVL
### Creating a Pool Dataframe
```
Pool = pd.DataFrame(
columns = ['date', 'LUSD', 'staBAL', 'LQTY', 'BAL', 'ETH', 'SP share', 'ETH received', 'BAL received']
)
Pool['date'] = Data['date']
Pool.loc[:, 'LUSD'] = 30e6
Pool.loc[:, 'staBAL'] = 30e6
Pool.loc[:, 'LQTY'] = 0
Pool.loc[:, 'BAL'] = 0
Pool.loc[:, 'SP share'] = 0
Pool.loc[:, 'ETH'] = 0
Pool.loc[:, 'ETH received'] = 0
Pool.loc[:, 'BAL received'] = 0
Pool.sort_values('date', ignore_index=True, inplace=True)
Pool
for i in range(len(Pool)):
if (i > 0):
Pool.loc[i, 'LUSD'] = Pool['LUSD'][i-1]
Pool.loc[i, 'ETH'] = Pool['ETH'][i-1]
Pool.loc[i, 'staBAL'] = Pool['staBAL'][i-1]
Pool.loc[i, 'LQTY'] = Pool['LQTY'][i-1]
Pool.loc[i, 'BAL'] = Pool['BAL'][i-1]
Pool.loc[i, 'SP share'] = Pool['LUSD'][i] / Data['LUSD in SP'][i]
if not(Data['LIQ mode'][i] == 'none'): #Processing liquidation gains
ETH_received = Data['LIQ col'][i]*(1-0.005)*Pool['SP share'][i]
LUSD_lost = Data['LIQ debt'][i]*Pool['SP share'][i]
Pool.loc[i, 'ETH received'] = ETH_received
Pool.loc[i, 'ETH'] += ETH_received
Pool.loc[i, 'LUSD'] -= LUSD_lost
else:
#checking if any ETH is on the account
if (Pool['ETH'][i] >= 0):
#selling ETH for LUSD
LQTY_sold = 0
LUSD_bought = 0
if (Pool['ETH'][i] >= 10):
ETH_sold = 10
LUSD_bought = ETH_sold*Data['ETH price'][i]*(0.9)
else:
ETH_sold = Pool['ETH'][i]
LUSD_bought = ETH_sold*Data['ETH price'][i]*(0.9)
Pool.loc[i, 'ETH'] -= ETH_sold
Pool.loc[i, 'LUSD'] += LUSD_bought
if (i > 0):
#calculating LQTY reward
LQTY_minted = Data['LQTY total'][i] - Data['LQTY total'][i-1]
Pool.loc[i, 'LQTY'] += LQTY_minted*Pool['SP share'][i]
#checking if any LQTY is on the account
if (Pool['LQTY'][i] > 0):
#selling LQTY for LUSD
LQTY_sold = 0
LUSD_bought = 0
if (Pool['LQTY'][i] > 1000):
LQTY_sold = 1000
LUSD_bought = LQTY_sold*6*(0.9) #Average $6 per LQTY
elif (Pool['LQTY'][i] > 50):
LQTY_sold = Pool['LQTY'][i]
LUSD_bought = LQTY_sold*6*(0.9)
Pool.loc[i, 'LQTY'] -= LQTY_sold
Pool.loc[i, 'LUSD'] += LUSD_bought
#calculating BAL reward
dT = Pool['date'][i] - Pool['date'][i-1]
PR = staBALdata['APR'] * dT/timedelta(days = 365)
BAL_minted = Pool['staBAL'][i]*Data['staBAL price'][i]*PR/Data['BAL price'][i]
Pool.loc[i, 'BAL'] += BAL_minted
Pool.loc[i, 'BAL received'] = BAL_minted
#checking if any BAL is on the account
if (Pool['BAL'][i] > 0):
#selling BAL for stable token, then restaking to staBAL
BAL_sold = 0
staBAL_staked = 0
if (Pool['BAL'][i] > 1000):
BAL_sold = 1000
staBAL_staked = BAL_sold*Data['BAL price'][i]/Data['staBAL price'][i]*(0.9)
elif (Pool['BAL'][i] > 50):
BAL_sold = Pool['BAL'][i]
staBAL_staked = BAL_sold*Data['BAL price'][i]/Data['staBAL price'][i]*(0.9)
Pool.loc[i, 'BAL'] -= BAL_sold
Pool.loc[i, 'staBAL'] += staBAL_staked
Pool
```
### Results
Let us calculate the results of the model:
```
LUSD_gain = Pool.iloc[-1]['LUSD'] - Pool['LUSD'][0]
staBAL_gain = (Pool.iloc[-1]['staBAL'] - Pool['staBAL'][0])*Data.iloc[-1]['staBAL price']
LUSD_percentage = LUSD_gain/Pool.iloc[0]['LUSD']
staBAL_percentage = staBAL_gain/Pool.iloc[0]['staBAL']
timeDelta = Pool.iloc[-1]['date'] - Pool.iloc[0]['date']
year = pd.to_timedelta(timedelta(days=365))
coef = timeDelta
LUSD_APR = LUSD_percentage*year/timeDelta
staBAL_APR = staBAL_percentage*year/timeDelta
print('LUSD APR: {:0,.3f}'.format(LUSD_APR))
print('staBAL APR: {:0,.3f}'.format(staBAL_APR))
```
* **LUSD APR** resulted in 0.223 (**22.3%**)
* **staBAL APR** resulted in 0.063 (**6.3%**)
Let's see the behaviour of the Pool on graphs:
```
%config InlineBackend.figure_format='svg'
dates = np.arange(0, len(Pool), 72)
date_ticks = [Pool['date'][i] for i in dates]
fig1, ax1 = plt.subplots()
plot = ax1.plot(Pool['date'], Pool['LUSD'], color='blue')
plot = ax1.plot(Pool['date'], Pool['staBAL'], color='brown')
ax1.legend(('LUSD', 'staBAL'))
ax1.grid()
ax1.set_title('Pool Composition')
ax1.set_xlabel('Date')
ax1.set_ylabel('Millions of $')
scale_y = 1e6
ticks_y = ticker.FuncFormatter(lambda x, pos: '{0:g}'.format(x/scale_y))
ax1.yaxis.set_major_formatter(ticks_y)
fig1.set_size_inches(8, 5)
ax2 = ax1.twinx()
ax2.plot(Pool['date'], Data['LIQ col'], color='red')
ax2.legend(('Liquidated ETH',))
ax2.set_ylabel('ETH')
```
The gap we see in the LUSD balance is a massive liquidation (most likely occuring during the recovery mode). Corresponding to what was stated by Liquity, liquidations increase the LP's total networh, which is seen by the jump in the LUSD balance after the received collateral ETH was traded back to LUSD.
| github_jupyter |
# Tutorial 08: Creating Custom Environments
This tutorial walks you through the process of creating custom environments in Flow. Custom environments contain specific methods that define the problem space of a task, such as the state and action spaces of the RL agent and the signal (or reward) that the RL algorithm will optimize over. By specifying a few methods within a custom environment, individuals can use Flow to design traffic control tasks of various types, such as optimal traffic light signal timing and flow regulation via mixed autonomy traffic (see the figures below). Finally, these environments are compatible with OpenAI Gym.
The rest of the tutorial is organized as follows: in section 1 walks through the process of creating an environment for mixed autonomy vehicle control where the autonomous vehicles perceive all vehicles in the network, and section two implements the environment in simulation.
<img src="img/sample_envs.png">
## 1. Creating an Environment Class
In this exercise we will create an environment in which the accelerations of a handful of vehicles in the network are specified by a single centralized agent, with the objective of the agent being to improve the average speed of all vehicle in the network. In order to create this environment, we begin by inheriting the base environment class located in *flow.envs*:
```
# import the base environment class
from flow.envs import Env
# define the environment class, and inherit properties from the base environment class
class myEnv(Env):
pass
```
`Env` provides the interface for running and modifying a SUMO simulation. Using this class, we are able to start sumo, provide a network to specify a configuration and controllers, perform simulation steps, and reset the simulation to an initial configuration.
By inheriting Flow's base environment, a custom environment for varying control tasks can be created by adding the following functions to the child class:
* **action_space**
* **observation_space**
* **apply_rl_actions**
* **get_state**
* **compute_reward**
Each of these components are covered in the next few subsections.
### 1.1 ADDITIONAL_ENV_PARAMS
The features used to parametrize components of the state/action space as well as the reward function are specified within the `EnvParams` input, as discussed in tutorial 1. Specifically, for the sake of our environment, the `additional_params` attribute within `EnvParams` will be responsible for storing information on the maximum possible accelerations and decelerations by the autonomous vehicles in the network. Accordingly, for this problem, we define an `ADDITIONAL_ENV_PARAMS` variable of the form:
```
ADDITIONAL_ENV_PARAMS = {
"max_accel": 1,
"max_decel": 1,
}
```
All environments presented in Flow provide a unique `ADDITIONAL_ENV_PARAMS` component containing the information needed to properly define some environment-specific parameters. We assume that these values are always provided by the user, and accordingly can be called from `env_params`. For example, if we would like to call the "max_accel" parameter, we simply type:
max_accel = env_params.additional_params["max_accel"]
### 1.2 action_space
The `action_space` method defines the number and bounds of the actions provided by the RL agent. In order to define these bounds with an OpenAI gym setting, we use several objects located within *gym.spaces*. For instance, the `Box` object is used to define a bounded array of values in $\mathbb{R}^n$.
```
from gym.spaces.box import Box
```
In addition, `Tuple` objects (not used by this exercise) allow users to combine multiple `Box` elements together.
```
from gym.spaces import Tuple
```
Once we have imported the above objects, we are ready to define the bounds of our action space. Given that our actions consist of a list of n real numbers (where n is the number of autonomous vehicles) bounded from above and below by "max_accel" and "max_decel" respectively (see section 1.1), we can define our action space as follows:
```
class myEnv(myEnv):
@property
def action_space(self):
num_actions = self.initial_vehicles.num_rl_vehicles
accel_ub = self.env_params.additional_params["max_accel"]
accel_lb = - abs(self.env_params.additional_params["max_decel"])
return Box(low=accel_lb,
high=accel_ub,
shape=(num_actions,))
```
### 1.3 observation_space
The observation space of an environment represents the number and types of observations that are provided to the reinforcement learning agent. For this example, we will be observe two values for each vehicle: its position and speed. Accordingly, we need a observation space that is twice the size of the number of vehicles in the network.
```
class myEnv(myEnv): # update my environment class
@property
def observation_space(self):
return Box(
low=0,
high=float("inf"),
shape=(2*self.initial_vehicles.num_vehicles,),
)
```
### 1.4 apply_rl_actions
The function `apply_rl_actions` is responsible for transforming commands specified by the RL agent into actual actions performed within the simulator. The vehicle kernel within the environment class contains several helper methods that may be of used to facilitate this process. These functions include:
* **apply_acceleration** (list of str, list of float) -> None: converts an action, or a list of actions, into accelerations to the specified vehicles (in simulation)
* **apply_lane_change** (list of str, list of {-1, 0, 1}) -> None: converts an action, or a list of actions, into lane change directions for the specified vehicles (in simulation)
* **choose_route** (list of str, list of list of str) -> None: converts an action, or a list of actions, into rerouting commands for the specified vehicles (in simulation)
For our example we consider a situation where the RL agent can only specify accelerations for the RL vehicles; accordingly, the actuation method for the RL agent is defined as follows:
```
class myEnv(myEnv): # update my environment class
def _apply_rl_actions(self, rl_actions):
# the names of all autonomous (RL) vehicles in the network
rl_ids = self.k.vehicle.get_rl_ids()
# use the base environment method to convert actions into accelerations for the rl vehicles
self.k.vehicle.apply_acceleration(rl_ids, rl_actions)
```
### 1.5 get_state
The `get_state` method extracts features from within the environments and provides then as inputs to the policy provided by the RL agent. Several helper methods exist within flow to help facilitate this process. Some useful helper method can be accessed from the following objects:
* **self.k.vehicle**: provides current state information for all vehicles within the network
* **self.k.traffic_light**: provides state information on the traffic lights
* **self.k.network**: information on the network, which unlike the vehicles and traffic lights is static
* More accessor objects and methods can be found within the Flow documentation at: http://berkeleyflow.readthedocs.io/en/latest/
In order to model global observability within the network, our state space consists of the speeds and positions of all vehicles (as mentioned in section 1.3). This is implemented as follows:
```
import numpy as np
class myEnv(myEnv): # update my environment class
def get_state(self, **kwargs):
# the get_ids() method is used to get the names of all vehicles in the network
ids = self.k.vehicle.get_ids()
# we use the get_absolute_position method to get the positions of all vehicles
pos = [self.k.vehicle.get_x_by_id(veh_id) for veh_id in ids]
# we use the get_speed method to get the velocities of all vehicles
vel = [self.k.vehicle.get_speed(veh_id) for veh_id in ids]
# the speeds and positions are concatenated to produce the state
return np.concatenate((pos, vel))
```
### 1.6 compute_reward
The `compute_reward` method returns the reward associated with any given state. These value may encompass returns from values within the state space (defined in section 1.5) or may contain information provided by the environment but not immediately available within the state, as is the case in partially observable tasks (or POMDPs).
For this exercise, we choose the reward function to be the average speed of all vehicles currently in the network. In order to extract this information from the environment, we use the `get_speed` method within the Vehicle kernel class to collect the current speed of all vehicles in the network, and return the average of these speeds as the reward. This is done as follows:
```
import numpy as np
class myEnv(myEnv): # update my environment class
def compute_reward(self, rl_actions, **kwargs):
# the get_ids() method is used to get the names of all vehicles in the network
ids = self.k.vehicle.get_ids()
# we next get a list of the speeds of all vehicles in the network
speeds = self.k.vehicle.get_speed(ids)
# finally, we return the average of all these speeds as the reward
return np.mean(speeds)
```
## 2. Testing the New Environment
### 2.1 Testing in Simulation
Now that we have successfully created our new environment, we are ready to test this environment in simulation. We begin by running this environment in a non-RL based simulation. The return provided at the end of the simulation is indicative of the cumulative expected reward when jam-like behavior exists within the netowrk.
```
from flow.controllers import IDMController, ContinuousRouter
from flow.core.experiment import Experiment
from flow.core.params import SumoParams, EnvParams, \
InitialConfig, NetParams
from flow.core.params import VehicleParams
from flow.networks.ring import RingNetwork, ADDITIONAL_NET_PARAMS
sumo_params = SumoParams(sim_step=0.1, render=True)
vehicles = VehicleParams()
vehicles.add(veh_id="idm",
acceleration_controller=(IDMController, {}),
routing_controller=(ContinuousRouter, {}),
num_vehicles=22)
env_params = EnvParams(additional_params=ADDITIONAL_ENV_PARAMS)
additional_net_params = ADDITIONAL_NET_PARAMS.copy()
net_params = NetParams(additional_params=additional_net_params)
initial_config = InitialConfig(bunching=20)
network = RingNetwork(name="sugiyama",
vehicles=vehicles,
net_params=net_params,
initial_config=initial_config)
#############################################################
######## using my new environment for the simulation ########
#############################################################
env = myEnv(env_params, sumo_params, network)
#############################################################
exp = Experiment(env)
_ = exp.run(1, 1500)
```
### 2.2 Training the New Environment
Next, we wish to train this environment in the presence of the autonomous vehicle agent to reduce the formation of waves in the network, thereby pushing the performance of vehicles in the network past the above expected return.
In order for an environment to be trainable in either RLLib for rllab (as we have shown in tutorials 2 and 3), the environment must be acccessable via import from *flow.envs*. In order to do so, copy the above envrionment onto a .py and import the environment in `flow.envs.__init__.py`. You can ensure that the process was successful by running the following command:
```
# NOTE: only runs if the above procedure have been performed
from flow.envs import myEnv
```
Once this is done, the below code block may be used to train the above environment using the Trust Region Policy Optimization (TRPO) algorithm provided by rllab. We do not recommend training this environment to completion within a jupyter notebook setting; however, once training is complete, visualization of the resulting policy should show that the autonomous vehicle learns to dissipate the formation and propagation of waves in the network.
```
from rllab.envs.normalized_env import normalize
from rllab.misc.instrument import run_experiment_lite
from rllab.algos.trpo import TRPO
from rllab.baselines.linear_feature_baseline import LinearFeatureBaseline
from rllab.policies.gaussian_gru_policy import GaussianGRUPolicy
from flow.networks.ring import RingNetwork
from flow.controllers import RLController, IDMController, ContinuousRouter
from flow.core.params import VehicleParams
from flow.core.params import SumoParams, EnvParams, NetParams, InitialConfig
from rllab.envs.gym_env import GymEnv
HORIZON = 1500
def run_task(*_):
sumo_params = SumoParams(sim_step=0.1, render=False)
vehicles = VehicleParams()
vehicles.add(veh_id="rl",
acceleration_controller=(RLController, {}),
routing_controller=(ContinuousRouter, {}),
num_vehicles=1)
vehicles.add(veh_id="idm",
acceleration_controller=(IDMController, {}),
routing_controller=(ContinuousRouter, {}),
num_vehicles=21)
env_params = EnvParams(horizon=HORIZON,
additional_params=ADDITIONAL_ENV_PARAMS)
additional_net_params = ADDITIONAL_NET_PARAMS.copy()
net_params = NetParams(additional_params=additional_net_params)
initial_config = InitialConfig(bunching=20)
network = RingNetwork(name="sugiyama-training",
vehicles=vehicles,
net_params=net_params,
initial_config=initial_config)
#######################################################
######## using my new environment for training ########
#######################################################
env_name = "myEnv"
#######################################################
pass_params = (env_name, sumo_params, vehicles, env_params, net_params,
initial_config, network)
env = GymEnv(env_name, record_video=False, register_params=pass_params)
horizon = env.horizon
env = normalize(env)
policy = GaussianGRUPolicy(
env_spec=env.spec,
hidden_sizes=(5,),
)
baseline = LinearFeatureBaseline(env_spec=env.spec)
algo = TRPO(
env=env,
policy=policy,
baseline=baseline,
batch_size=30000,
max_path_length=horizon,
n_itr=500,
discount=0.999,
)
algo.train(),
exp_tag = "stabilizing-the-ring"
for seed in [5]: # , 20, 68]:
run_experiment_lite(
run_task,
n_parallel=1,
snapshot_mode="all",
seed=seed,
mode="local",
exp_prefix=exp_tag,
)
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Save and serialize models with Keras
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/guide/keras/save_and_serialize"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/keras/save_and_serialize.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/keras/save_and_serialize.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/keras/save_and_serialize.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
The first part of this guide covers saving and serialization for Keras models built using the Functional and Sequential APIs. Saving and serialization is exactly same for both of these model APIs.
The second part of this guide covers "[saving and loading subclassed models](save_and_serialize.ipynb#saving-subclassed-models)". The subclassing API differs from the Keras sequential and functional API.
## Setup
```
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
tf.keras.backend.clear_session() # For easy reset of notebook state.
```
## Part I: Saving Sequential models or Functional models
Let's consider the following model:
```
from tensorflow import keras
from tensorflow.keras import layers
inputs = keras.Input(shape=(784,), name='digits')
x = layers.Dense(64, activation='relu', name='dense_1')(inputs)
x = layers.Dense(64, activation='relu', name='dense_2')(x)
outputs = layers.Dense(10, name='predictions')(x)
model = keras.Model(inputs=inputs, outputs=outputs, name='3_layer_mlp')
model.summary()
```
Optionally, let's train this model, just so it has weight values to save, as well as an optimizer state.
Of course, you can save models you've never trained, too, but obviously that's less interesting.
```
(x_train, y_train), (x_test, y_test) = keras.datasets.mnist.load_data()
x_train = x_train.reshape(60000, 784).astype('float32') / 255
x_test = x_test.reshape(10000, 784).astype('float32') / 255
model.compile(loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
optimizer=keras.optimizers.RMSprop())
history = model.fit(x_train, y_train,
batch_size=64,
epochs=1)
# Reset metrics before saving so that loaded model has same state,
# since metric states are not preserved by Model.save_weights
model.reset_metrics()
# Save predictions for future checks
predictions = model.predict(x_test)
```
### Whole-model saving
You can save a model built with the Functional API into a single file. You can later recreate the same model from this file, even if you no longer have access to the code that created the model.
This file includes:
- The model's architecture
- The model's weight values (which were learned during training)
- The model's training config (what you passed to `compile`), if any
- The optimizer and its state, if any (this enables you to restart training where you left)
```
# Save the model
model.save('path_to_my_model.h5')
# Recreate the exact same model purely from the file
new_model = keras.models.load_model('path_to_my_model.h5')
import numpy as np
# Check that the state is preserved
new_predictions = new_model.predict(x_test)
np.testing.assert_allclose(predictions, new_predictions, rtol=1e-6, atol=1e-6)
# Note that the optimizer state is preserved as well:
# you can resume training where you left off.
```
### Export to SavedModel
You can also export a whole model to the TensorFlow `SavedModel` format. `SavedModel` is a standalone serialization format for TensorFlow objects, supported by TensorFlow serving as well as TensorFlow implementations other than Python.
```
# Export the model to a SavedModel
model.save('path_to_saved_model', save_format='tf')
# Recreate the exact same model
new_model = keras.models.load_model('path_to_saved_model')
# Check that the state is preserved
new_predictions = new_model.predict(x_test)
np.testing.assert_allclose(predictions, new_predictions, rtol=1e-6, atol=1e-6)
# Note that the optimizer state is preserved as well:
# you can resume training where you left off.
```
The `SavedModel` files that were created contain:
- A TensorFlow checkpoint containing the model weights.
- A `SavedModel` proto containing the underlying TensorFlow graph.
### Architecture-only saving
Sometimes, you are only interested in the architecture of the model, and you don't need to save the weight values or the optimizer. In this case, you can retrieve the "config" of the model via the `get_config()` method. The config is a Python dict that enables you to recreate the same model -- initialized from scratch, without any of the information learned previously during training.
```
config = model.get_config()
reinitialized_model = keras.Model.from_config(config)
# Note that the model state is not preserved! We only saved the architecture.
new_predictions = reinitialized_model.predict(x_test)
assert abs(np.sum(predictions - new_predictions)) > 0.
```
You can alternatively use `to_json()` from `from_json()`, which uses a JSON string to store the config instead of a Python dict. This is useful to save the config to disk.
```
json_config = model.to_json()
reinitialized_model = keras.models.model_from_json(json_config)
```
### Weights-only saving
Sometimes, you are only interested in the state of the model -- its weights values -- and not in the architecture. In this case, you can retrieve the weights values as a list of Numpy arrays via `get_weights()`, and set the state of the model via `set_weights`:
```
weights = model.get_weights() # Retrieves the state of the model.
model.set_weights(weights) # Sets the state of the model.
```
You can combine `get_config()`/`from_config()` and `get_weights()`/`set_weights()` to recreate your model in the same state. However, unlike `model.save()`, this will not include the training config and the optimizer. You would have to call `compile()` again before using the model for training.
```
config = model.get_config()
weights = model.get_weights()
new_model = keras.Model.from_config(config)
new_model.set_weights(weights)
# Check that the state is preserved
new_predictions = new_model.predict(x_test)
np.testing.assert_allclose(predictions, new_predictions, rtol=1e-6, atol=1e-6)
# Note that the optimizer was not preserved,
# so the model should be compiled anew before training
# (and the optimizer will start from a blank state).
```
The save-to-disk alternative to `get_weights()` and `set_weights(weights)`
is `save_weights(fpath)` and `load_weights(fpath)`.
Here's an example that saves to disk:
```
# Save JSON config to disk
json_config = model.to_json()
with open('model_config.json', 'w') as json_file:
json_file.write(json_config)
# Save weights to disk
model.save_weights('path_to_my_weights.h5')
# Reload the model from the 2 files we saved
with open('model_config.json') as json_file:
json_config = json_file.read()
new_model = keras.models.model_from_json(json_config)
new_model.load_weights('path_to_my_weights.h5')
# Check that the state is preserved
new_predictions = new_model.predict(x_test)
np.testing.assert_allclose(predictions, new_predictions, rtol=1e-6, atol=1e-6)
# Note that the optimizer was not preserved.
```
But remember that the simplest, recommended way is just this:
```
model.save('path_to_my_model.h5')
del model
model = keras.models.load_model('path_to_my_model.h5')
```
### Weights-only saving using TensorFlow checkpoints
Note that `save_weights` can create files either in the Keras HDF5 format,
or in the [TensorFlow Checkpoint format](https://www.tensorflow.org/api_docs/python/tf/train/Checkpoint). The format is inferred from the file extension you provide: if it is ".h5" or ".keras", the framework uses the Keras HDF5 format. Anything else defaults to Checkpoint.
```
model.save_weights('path_to_my_tf_checkpoint')
```
For total explicitness, the format can be explicitly passed via the `save_format` argument, which can take the value "tf" or "h5":
```
model.save_weights('path_to_my_tf_checkpoint', save_format='tf')
```
## Part II: Saving and Loading of Subclassed Models
Sequential models and Functional models are datastructures that represent a DAG of layers. As such,
they can be safely serialized and deserialized.
A subclassed model differs in that it's not a datastructure, it's a piece of code. The architecture of the model
is defined via the body of the `call` method. This means that the architecture of the model cannot be safely serialized. To load a model, you'll need to have access to the code that created it (the code of the model subclass). Alternatively, you could be serializing this code as bytecode (e.g. via pickling), but that's unsafe and generally not portable.
For more information about these differences, see the article ["What are Symbolic and Imperative APIs in TensorFlow 2.0?"](https://medium.com/tensorflow/what-are-symbolic-and-imperative-apis-in-tensorflow-2-0-dfccecb01021).
Let's consider the following subclassed model, which follows the same structure as the model from the first section:
```
class ThreeLayerMLP(keras.Model):
def __init__(self, name=None):
super(ThreeLayerMLP, self).__init__(name=name)
self.dense_1 = layers.Dense(64, activation='relu', name='dense_1')
self.dense_2 = layers.Dense(64, activation='relu', name='dense_2')
self.pred_layer = layers.Dense(10, name='predictions')
def call(self, inputs):
x = self.dense_1(inputs)
x = self.dense_2(x)
return self.pred_layer(x)
def get_model():
return ThreeLayerMLP(name='3_layer_mlp')
model = get_model()
```
First of all, *a subclassed model that has never been used cannot be saved*.
That's because a subclassed model needs to be called on some data in order to create its weights.
Until the model has been called, it does not know the shape and dtype of the input data it should be
expecting, and thus cannot create its weight variables. You may remember that in the Functional model from the first section, the shape and dtype of the inputs was specified in advance (via `keras.Input(...)`) -- that's why Functional models have a state as soon as they're instantiated.
Let's train the model, so as to give it a state:
```
(x_train, y_train), (x_test, y_test) = keras.datasets.mnist.load_data()
x_train = x_train.reshape(60000, 784).astype('float32') / 255
x_test = x_test.reshape(10000, 784).astype('float32') / 255
model.compile(loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
optimizer=keras.optimizers.RMSprop())
history = model.fit(x_train, y_train,
batch_size=64,
epochs=1)
# Reset metrics before saving so that loaded model has same state,
# since metric states are not preserved by Model.save_weights
model.reset_metrics()
```
There are three different approaches to save and restore a subclassed model. The following sections provides more details on those three approaches.
### Approach 1:
The recommended way to save a subclassed model is to use `save_weights` to create a TensorFlow SavedModel checkpoint, which will contain the value of all variables associated with the model:
- The layers' weights
- The optimizer's state
- Any variables associated with stateful model metrics (if any)
```
model.save_weights('path_to_my_weights', save_format='tf')
# Save predictions for future checks
predictions = model.predict(x_test)
# Also save the loss on the first batch
# to later assert that the optimizer state was preserved
first_batch_loss = model.train_on_batch(x_train[:64], y_train[:64])
```
To restore your model, you will need access to the code that created the model object.
Note that in order to restore the optimizer state and the state of any stateful metric, you should
compile the model (with the exact same arguments as before) and call it on some data before calling `load_weights`:
```
# Recreate the model
new_model = get_model()
new_model.compile(loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
optimizer=keras.optimizers.RMSprop())
# This initializes the variables used by the optimizers,
# as well as any stateful metric variables
new_model.train_on_batch(x_train[:1], y_train[:1])
# Load the state of the old model
new_model.load_weights('path_to_my_weights')
# Check that the model state has been preserved
new_predictions = new_model.predict(x_test)
np.testing.assert_allclose(predictions, new_predictions, rtol=1e-6, atol=1e-6)
# The optimizer state is preserved as well,
# so you can resume training where you left off
new_first_batch_loss=new_model.train_on_batch(x_train[:64], y_train[:64])
assert first_batch_loss == new_first_batch_loss
```
### Approach 2:
Second approach is by using `model.save` to save whole model and by using `load_model` to restore previously stored subclassed model. The following code snippets describe how to implement them.
```
# Save the model
model.save('path_to_my_model',save_format='tf')
# Recreate the exact same model purely from the file
new_model = keras.models.load_model('path_to_my_model')
```
### Approach 3:
Third approach is by using `tf.saved_model.save`. This is equivalent to the `tf` format in `model.save`. You can once again call `load_model` to restore the previously saved subclassed model. The following code snippets describe how to implement them.
```
# Save the model
tf.saved_model.save(model,'my_saved_model')
# Restoring the model
restored_saved_model = keras.models.load_model('my_saved_model')
```
| github_jupyter |
# Science User Case - Inspecting a Candidate List
Ogle et al. (2016) mined the NASA/IPAC Extragalactic Database (NED) to identify a new type of galaxy: Superluminous Spiral Galaxies.
Here's the paper: https://ui.adsabs.harvard.edu//#abs/2016ApJ...817..109O/abstract
Table 1 lists the positions of these Super Spirals. Based on those positions, let's create multiwavelength cutouts for each super spiral to see what is unique about this new class of objects.
## 1. Import the Python modules we'll be using.
```
# Suppress unimportant warnings.
import warnings
warnings.filterwarnings("ignore", module="astropy.io.votable.*")
warnings.filterwarnings("ignore", module="pyvo.utils.xml.*")
warnings.filterwarnings('ignore', '.*RADECSYS=*', append=True)
import matplotlib.pyplot as plt
import numpy as np
from astropy.coordinates import SkyCoord
from astropy.io import fits
from astropy.nddata import Cutout2D
import astropy.visualization as vis
from astropy.wcs import WCS
from astroquery.ned import Ned
import pyvo as vo
```
## 2. Search NED for objects in this paper.
Consult QuickReference.md to figure out how to use astroquery to search NED for all objects in a paper, based on the refcode of the paper. Inspect the resulting astropy table.
## 3. Filter the NED results.
The results from NED will include galaxies, but also other kinds of objects. Print the 'Type' column to see the full range of classifications. Next, print the 'Type' of just the first source in the table, in order to determine its data type (since Python 3 distinguishes between strings and byte strings). Finally, use the data type information to filter the results so that we only keep the galaxies in the list.
## 4. Search the NAVO Registry for image resources.
The paper selected super spirals using WISE, SDSS, and GALEX images. Search the NAVO registry for all image resources, using the 'service_type' search parameter. How many image resources are currently available?
## 5. Search the NAVO Registry for image resources that will allow you to search for AllWISE images.
There are hundreds of image resources...too many to quickly read through. Try adding the 'keyword' search parameter to your registry search, and find the image resource you would need to search the AllWISE images. Remember from the Known Issues that 'keywords' must be a list.
## 6. Select the AllWISE image service that you are interested in.
Hint: there should be only one service after searching with ['allwise']
## 7. Make a SkyCoord from the first galaxy in the NED list.
Helpful code snippet:
```
ra = galaxies['RA'][0]
dec = galaxies['DEC'][0]
pos = SkyCoord(ra, dec, unit = 'deg')
```
## 8. Search for a list of AllWISE images that cover this galaxy.
How many images are returned? Which are you most interested in?
## 9. Use the .to_table() method to view the results as an Astropy table.
## 10. From the result in 8., select the first record for an image taken in WISE band W1 (3.6 micron)
Hints:
* Loop over records and test on the `.bandpass_id` attribute of each record
* Print the `.title` and `.bandpass_id` of the record you find, to verify it is the right one.
## 11. Visualize this AllWISE image.
Helpful code snippet:
```
allwise_w1_image = fits.open(allwise_image_record.getdataurl())
```
For plotting:
```
fig = plt.figure()
wcs = WCS(allwise_w1_image[0].header)
ax = fig.add_subplot(1, 1, 1, projection=wcs)
ax.imshow(allwise_w1_image[0].data, cmap='gray_r', origin='lower', vmax = 10)
ax.scatter(ra, dec, transform=ax.get_transform('fk5'), s=500, edgecolor='red', facecolor='none')
```
## 12. Plot a cutout of the AllWISE image, centered on your position.
Try a 60 arcsecond cutout. Use `Cutout2D` that we imported earlier.
## 13. Try visualizing a cutout of a GALEX image that covers your position.
Repeat steps 4, 5, 6, 8 through 12 for GALEX.
## 14. Try visualizing a cutout of an SDSS image that covers your position.
Hints:
* Search the registry using `keywords=['sloan']
* Find the service with a `short_name` of `b'SDSS SIAP'`
* From Known Issues, recall that an empty string must be specified to the `format` parameter dues to a bug in the service.
* After obtaining your search results, select r-band images using the `.title` attribute of the records that are returned, since `.bandpass_id` is not populated.
## 15. Try looping over the first few positions and plotting multiwavelength cutouts.
| github_jupyter |
```
# default_exp solvers
```
# solvers
> algorithms to solve the MAP problems
```
#export
from thompson_sampling.abstractions import AbstractSolver, AbstractContextualSolver,AbstractContextualSolverSingleModel
import numpy as np
import scipy.stats as stats
import matplotlib.cm as cm
import matplotlib.pyplot as plt
import matplotlib.pyplot as plt
%matplotlib inline
%load_ext autoreload
%autoreload 2
```
## categorical
### non-contextual
#### AB test
```
#export
class AB(AbstractSolver):
def __init__(self, n_experiments=1000, num_options=2):
self.trials = np.zeros(shape=(num_options,))
self.successes = np.zeros(shape=(num_options,))
self.experiments_done = 0
self.n_experiments = n_experiments
def choose_arm(self):
"""we choose to either randomly sample an arm
or play the previously determined best choice"""
# if we need more experimentation, we explore
if self.experiments_done < self.n_experiments:
arm = self.explore()
# otherwise, we exploit
else:
arm = self.exploit()
return arm
def update(self, arm, reward):
"""Updates the arms about being played and about receiving a reward"""
# simply count the number of trials and successes for each arm
self.trials[arm] += 1
if reward:
self.successes[arm] += 1
self.experiments_done += 1
def explore(self):
"""returns arm 0 or arm 1 depending on a draw from interval [0,1]
NOTE: this isn't necessarilyhow you'd do it in real life,
please consult the sources for that case"""
# literally choose by random which arm to return
if np.random.random() <.5:
return 0
else:
return 1
def exploit(self):
"""returns arm with highest expected payoff
Always the same arm after exploration phase"""
# return the arm with the highest success rate
return np.argmax(self.successes/self.trials)
#export
class BetaBandit(AbstractSolver):
def __init__(self, num_options = 2, prior = None):
"""initialize BetaBandit"""
self.num_options = num_options
#setting the prior, either uninformative or user generated
if prior == None:
self.prior = np.ones(shape=(num_options,2))
else:
assert prior.shape == (num_options,2), f"the prior seems to have wrong dimensionality, please conform to (num_options, 2){(num_options,2)}"
self.prior = prior
self.trials = np.zeros(shape=(num_options,))
self.successes = np.zeros(shape=(num_options,))
def choose_arm(self):
"""draw from arms.
arm with the highest expected outcome wins.
expected outcome is determined stochastically, so even an arm with bad
outcome until now will have a chance of being drawn"""
sampled_theta=[]
for i in range(self.num_options):
dist = stats.beta(self.prior[i,0]+self.successes[i],
self.prior[i,1]+self.trials[i] - self.successes[i])
sampled_theta += [dist.rvs()]
return(sampled_theta.index(max(sampled_theta)))
def update(self,arm,success):
"""update beta-parameters of specific arm"""
#count times arm has been drawn"""
self.trials[arm] = self.trials[arm] +1
#count number of successes on that arm"""
# for decay factors: self.successes = self.successes *.99
if success:
self.successes[arm] = self.successes[arm]+ 1
# helpers
def choose_arm_and_plot(self):
sampled_theta = []
dist_heights = []
for i in range(self.num_options):
dist = stats.beta(self.prior[i,0]+self.successes[i],
self.prior[i,1]+self.trials[i] - self.successes[i])
sample = dist.rvs()
sampled_theta += [sample]
dist_heights += [dist.pdf(sample)]
w = 10
z = 5
colors = iter(cm.rainbow(np.linspace(0, 1, self.num_options)))
for k,i in enumerate(range(self.num_options)):
color = next(colors)
dist = stats.beta(self.prior[i,0] + self.successes[i],
self.prior[i,1] + self.trials[i] - self.successes[i])
x = np.linspace(0,1,100)
y = dist.pdf(x)
plt.plot(x,y,color=color,label="arm #%i"%(i+1),alpha=0.8)
plt.scatter(sampled_theta[i],dist_heights[i], s = 200,label=f'sample drawn from arm {i}')
plt.fill_between(x,0,y,alpha=1/(self.num_options+1),color=color)
leg = plt.legend()
plt.tight_layout
return(sampled_theta.index(max(sampled_theta)))
def plot_params(self):
"""plot the distributions that underly the arms"""
w = 10
z = 5
colors = iter(cm.rainbow(np.linspace(0, 1, self.num_options)))
for k,i in enumerate(range(self.num_options)):
color = next(colors)
dist = stats.beta(self.prior[i,0] + self.successes[i],
self.prior[i,1] + self.trials[i] - self.successes[i])
x = np.linspace(0,1,100)
y = dist.pdf(x)
plt.plot(x,y,color=color,label="arm #%i"%(i+1))
plt.fill_between(x,0,y,alpha=1/self.num_options,color=color)
leg = plt.legend()
plt.tight_layout
bb = BetaBandit()
np.mean([bb.choose_arm() for x in range(100)])
bb.choose_arm_and_plot() #uniform binomials, no observations yet
bb.choose_arm_and_plot() #uniform binomials, no observations yet
bb.update(0,0)
bb.update(0,1)
bb.update(0,1)
np.mean([bb.choose_arm() for x in range(100)])
bb.choose_arm_and_plot()
bb.update(1,0)
bb.update(1,0)
bb.update(1,0)
bb.choose_arm_and_plot()
```
## Contextual
```
#export
class LogisticThompsonSampler(AbstractContextualSolver):
def update(self,arm,context,reward):
model = self.model_list[arm]
#X = np.atleast_1d(np.append(arm, context))
X = np.atleast_2d(context)
# print(f'X {X}')
reward = np.atleast_1d(reward)
#model.observe(X.reshape(1,-1), reward.reshape(1,-1))
model.observe(X, reward)
def choose_arm(self,context):
reward_list = []
for arm in range(self.num_arms):
model = self.model_list[arm]
X = np.atleast_2d(context)
probas = model.predict_proba(X)
reward_sample = probas[0][1]
# print(reward_sample)
reward_list += [reward_sample]
return np.argmax(reward_list)
# helpers
def plot_params(self):
colors = iter(cm.rainbow(np.linspace(0, 1, self.num_arms)))
for arm in range(len(self.model_list)):
color = next(colors)
model = self.model_list[arm]
X_pdf = np.linspace(-2, 2, 1000)
pdf = stats.norm(loc=model.m, scale=model.q**(-1.0)).pdf(X_pdf)
# plotting distriution of weights
plt.plot(X_pdf, pdf, color=color, linewidth=2, alpha=0.5, label=f'estimated parameter arm {arm}')
plt.fill_between(X_pdf, pdf, 0, color=color, alpha=0.2)
plt.legend()
def choose_arm_and_plot(self,context):
colors = iter(cm.rainbow(np.linspace(0, 1, self.num_arms)))
reward_list = []
for arm in range(self.num_arms):
model = self.model_list[arm]
X = np.atleast_1d(context)
reward_sample = model.predict_proba(X)[0]
reward_list += [reward_sample]
# plot
color = next(colors)
model = self.model_list[arm]
X_pdf = np.linspace(-2, 2, 1000)
dist = stats.norm(loc=model.m, scale=model.alpha**(-1.0))
pdf = dist.pdf(X_pdf)
height = dist.pdf(reward_sample)
# plotting distriution of weights
plt.plot(X_pdf, pdf, color=color, linewidth=2, alpha=0.5, label=f'estimated parameter arm {arm}')
plt.fill_between(X_pdf, pdf, 0, color=color, alpha=0.2)
plt.scatter(reward_sample, height, s = 200,label=f'sample drawn from arm {arm}')
plt.legend()
return np.argmax(reward_list)
from thompson_sampling.models import OnlineLogisticRegression
from thompson_sampling.multi_armed_bandits import contextual_categorical_bandit
lts = LogisticThompsonSampler(OnlineLogisticRegression, num_arms=2, num_context = 1)
lts.model_list[0].get_weights()
lts.choose_arm_and_plot(np.atleast_2d([0.9]))
arm0 = 0
arm1 = 1
theta = [0.1, 1.9]
noise = 0.0
lts.update(arm1, .1, contextual_categorical_bandit(context = np.array(0.1),choice = arm1, theta = theta, noise = noise)[0])
lts.update(arm1, .1, contextual_categorical_bandit(context = np.array(.1),choice = arm1, theta = theta, noise = noise)[0])
lts.update(arm1, .5, contextual_categorical_bandit(context = np.array(0.5),choice = arm1, theta = theta, noise = noise)[0])
lts.update(arm1, .5, contextual_categorical_bandit(context = np.array(.5),choice = arm1, theta = theta, noise = noise)[0])
lts.update(arm1, .9, contextual_categorical_bandit(context = np.array(0.9),choice = arm1, theta = theta, noise = noise)[0])
lts.update(arm1, .9, contextual_categorical_bandit(context = np.array(.9),choice = arm1, theta = theta, noise = noise)[0])
lts.choose_arm_and_plot(0.9)
lts.choose_arm(0.9)
lts2 = LogisticThompsonSampler(OnlineLogisticRegression, num_arms=2, num_context = 2)
context = np.atleast_1d(np.array([1,1]))#.T#.reshape(1,-1)#.T
lts2.choose_arm(context)
context.shape
lts2.update(0,context,np.atleast_1d(1))
```
# Numerical
## no context
```
#export
class GaussianBandit(AbstractSolver):
def __init__(self, num_options = 2, mean_prior = 0, std_prior = 1):
"""initialize BetaBandit"""
self.num_options = num_options
#setting the prior, either uninformative or user generated
# if prior == None:
# self.prior = np.ones(shape=(num_options,2))
# else:
# assert prior.shape == (num_options,2), f"the prior seems to have wrong dimensionality, please conform to (num_options, 2){(num_options,2)}"
# self.prior = prior
self.trials = np.zeros(shape=(num_options,))
self.sum_x = np.zeros(shape=(num_options,))
self.sum_x2 = np.zeros(shape=(num_options,))
self.mean_prior = mean_prior
self.std_prior = std_prior
def choose_arm(self):
"""draw from arms.
arm with the highest expected outcome wins.
expected outcome is determined stochastically, so even an arm with bad
outcome until now will have a chance of being drawn"""
sampled_outcomes = []
for i in range(self.num_options):
if self.trials[i] > 1:
mean = self.compute_online_mean(i)
stdev = self.compute_online_std(i, mean)
else:
mean = 0
stdev = 1
dist = stats.norm(mean,stdev)
sampled_outcome = dist.rvs()
#print(sampled_outcome)
sampled_outcomes += [sampled_outcome]
return np.argmax(sampled_outcomes)
#return(sampled_outcomes.argmax(sampled_outcomes))
def update(self, arm, outcome):
"""update parameters of specific arm"""
#count times arm has been drawn"""
self.trials[arm] = self.trials[arm] +1
#count number of successes on that arm"""
# for decay factors: self.successes = self.successes *.99
self.sum_x[arm] += outcome
self.sum_x2[arm] += outcome*outcome
def compute_online_mean(self, arm):
return self.sum_x[arm] / (self.trials[arm])
def compute_online_std(self, arm, mean = None):
mean = mean or self.compute_online_mean(arm)
#np max against degeneration)
return np.max([np.sqrt((self.sum_x2[arm] / (self.trials[arm])) - (mean * mean)), 0.00001])
def plot_params(self):
"""plot the distributions that underly the arms"""
w = 10
z = 5
colors = iter(cm.rainbow(np.linspace(0, 1, self.num_options)))
for k,i in enumerate(range(self.num_options)):
color = next(colors)
if self.trials[i] > 1:
mean = self.compute_online_mean(i)
stdev = self.compute_online_std(i, mean)
else:
mean = 0
stdev = 1
dist = stats.norm(mean,stdev)
x = np.linspace(-6,6,100)
y = dist.pdf(x)
plt.plot(x,y,color=color,label="arm #%i"%(i+1))
plt.fill_between(x,0,y,alpha=1/self.num_options,color=color)
leg = plt.legend()
plt.tight_layout
num_arms = 10
gb = GaussianBandit(num_arms)
gb.plot_params()
np.mean([gb.choose_arm() for x in range(100)]), np.histogram([gb.choose_arm() for x in range(100)], bins=list(range(num_arms)))
gb.update(0, 10)
gb.update(0, 1)
gb.update(1, -3)
gb.update(1, 0)
# degenerate gaussian
gb.plot_params()
np.mean([gb.choose_arm() for x in range(100)]), np.histogram([gb.choose_arm() for x in range(100)], bins=list(range(num_arms)))
gb.trials
gb.compute_online_mean(0), gb.compute_online_mean(1)
gb.compute_online_std(0), gb.compute_online_std(1)
gb.update(1, 5)
gb.update(0, 1)
gb.update(0, -10)
gb.update(1, 10)
np.mean([gb.choose_arm() for x in range(100)]), np.histogram([gb.choose_arm() for x in range(100)], bins=list(range(num_arms)))
gb.compute_online_mean(0), gb.compute_online_mean(1)
gb.compute_online_std(0), gb.compute_online_std(1)
gb.plot_params()
```
## context
```
#export
class GaussianContextualSampler(AbstractContextualSolver):
def update(self,arm,context,reward):
model = self.model_list[arm]
X = np.atleast_2d(context)
reward = np.atleast_1d(reward)
model.observe(X, reward)
def choose_arm(self,context):
reward_list = []
for arm in range(self.num_arms):
model = self.model_list[arm]
X = np.atleast_2d(context)
arm_dist = model.predict(X)
reward_sample = arm_dist.rvs()
reward_list += [reward_sample]
return np.argmax(reward_list)
def choose_arm_and_plot(self,context):
colors = iter(cm.rainbow(np.linspace(0, 1, self.num_arms)))
reward_list = []
for arm in range(self.num_arms):
model = self.model_list[arm]
X = np.atleast_2d(context)
arm_dist = model.predict(X)
reward_sample = arm_dist.rvs()
reward_list += [reward_sample]
# plot
color = next(colors)
model = self.model_list[arm]
X_pdf = np.linspace(-10, 10, 1000)
pdf = arm_dist.pdf(X_pdf)
height = arm_dist.pdf(reward_sample)
# plotting distriution of weights
plt.plot(X_pdf, pdf, color=color, linewidth=2, alpha=0.5, label=f'estimated parameter arm {arm}')
plt.fill_between(X_pdf, pdf, 0, color=color, alpha=0.2)
plt.scatter(reward_sample, height, s = 200,label=f'sample drawn from arm {arm}')
plt.legend()
return np.argmax(reward_list)
from thompson_sampling.models import BatchBayesLinReg
from thompson_sampling.multi_armed_bandits import contextual_numerical_bandit
gcs = GaussianContextualSampler(BatchBayesLinReg,num_arms=2, num_context = 1,model_params={'alpha':1, 'beta':4})
arm0 = 0
arm1 = 1
theta = [0.1, 1.9]
noise = 1
X = np.linspace(-10,10, 10)
y0 = [contextual_numerical_bandit(x,0,theta,noise) for x in X]
y1 = [contextual_numerical_bandit(x,1,theta,noise) for x in X]
plt.scatter(X,y1)
plt.scatter(X,y0)
contexts = [10,-10,-1,9.5,-9,1]
a = [gcs.update(arm1, context, contextual_numerical_bandit(context = np.array(context),choice = arm1, theta = theta, noise = noise)) for context in contexts]
a
a = [contextual_numerical_bandit(context = np.array(context),choice = arm0, theta = theta, noise = noise) for context in contexts]
a
contextual_numerical_bandit(context = np.array(10),choice = arm1, theta = theta, noise = noise)
#plt.plot(context,a)
contextual_numerical_bandit(context = np.array(context),choice = arm1, theta = theta, noise = noise)
gcs.update(arm1, 10, contextual_numerical_bandit(context = np.array(context),choice = arm1, theta = theta, noise = noise))
gcs.choose_arm_and_plot(5)
gcs.choose_arm_and_plot(-5)
```
# numerical or categorical reward, categorical arm
## no context
```
#export
class GaussianCategoricalBandit(GaussianBandit):
def choose_arm(self):
"""draw from arms.
arm with the highest expected outcome wins.
expected outcome is determined stochastically, so even an arm with bad
outcome until now will have a chance of being drawn"""
sampled_outcomes = []
arm_samples = np.zeros(shape=(self.num_options,100))
for arm in range(self.num_options):
for i in range(100):
if self.trials[arm] > 1:
mean = self.compute_online_mean(arm)
stdev = self.compute_online_std(arm, mean)
else:
mean = 0
stdev = 1
dist = stats.norm(mean,stdev)
arm_samples[arm,i] = dist.rvs()
winning_ratio = np.argmax(arm_samples,0)
winning_arm = np.bincount(winning_ratio).argmax()
# ToDo sample from ratio of winning arms, i.e. nb.bincount(winning_ratio).mean()
return winning_arm
def choose_arm_and_plot(self):
print('this is not implemented, will not plot the distribution, please also call plot arm')
return self.choose_arm()
gcb = GaussianCategoricalBandit(2)
gcb.plot_params()
[i for i in range(gcb.num_options)]
gcb.update(1,0)
gcb.update(1,0)
gcb.update(1,0)
gcb.update(1,1)
gcb.update(0,0)
gcb.update(0,1)
gcb.update(0,1)
gcb.update(0,1)
gcb.plot_params()
gcb.update(1, 5)
gcb.update(0, 1)
gcb.update(0, -10)
gcb.update(1, 10)
gcb.plot_params()
```
## contextual
```
#export
class GaussianUniversalContextualSampler(GaussianContextualSampler):
def update(self,arm,context,reward):
model = self.model_list[arm]
X = np.atleast_2d(context)
reward = np.atleast_1d(reward)
model.observe(X, reward)
def choose_arm(self, context, num_samples = 10):
"""draw from arms.
arm with the highest expected outcome wins.
expected outcome is determined stochastically, so even an arm with bad
outcome until now will have a chance of being drawn"""
sampled_outcomes = []
arm_samples = np.zeros(shape=(self.num_arms,num_samples))
X = np.atleast_2d(context)
for arm in range(self.num_arms):
model = self.model_list[arm]
arm_dist = model.predict(X)
for i in range(num_samples):
arm_samples[arm,i] = arm_dist.rvs()
winning_ratio = np.argmax(arm_samples,0)
winning_arm = np.bincount(winning_ratio).argmax()
# ToDo sample from ratio of winning arms, i.e. nb.bincount(winning_ratio).mean()
return winning_arm
#gucs = GaussianUniversalContextualSampler
gucs = GaussianUniversalContextualSampler(BatchBayesLinReg,num_arms=2, num_context = 1,model_params={'alpha':1, 'beta':4})
arm0 = 0
arm1 = 1
theta = [0.1, 1.9]
noise = 0.0
contexts = [10,-10,-1,9.5,-9,1]
a = [gucs.update(arm1, context, contextual_numerical_bandit(context = np.array(context),choice = arm1, theta = theta, noise = noise)) for context in contexts]
a = [gucs.update(arm1, context, contextual_numerical_bandit(context = np.array(context),choice = arm1, theta = theta, noise = noise)) for context in contexts]
a = [gucs.update(arm1, context, contextual_numerical_bandit(context = np.array(context),choice = arm1, theta = theta, noise = noise)) for context in contexts]
gcs.choose_arm_and_plot(1)
gcs.choose_arm_and_plot(0)
gcs.choose_arm_and_plot(0)
from nbdev.export import *
notebook2script()
#export
from collections import OrderedDict
class GaussianContextualSamplerSingleModel(AbstractContextualSolverSingleModel):
def update(self,arm_ix,context,reward):
arm = self.arms[arm_ix]
X = np.append(arm,context)
reward = np.atleast_1d(reward)
#print(X, reward)
self.errors.append(self.model.observe(X, reward))
def choose_arm(self,context):
reward_dict = {}
for arm_ix in self.arms:
arm = self.arms[arm_ix]
X = np.append(arm,context)
#print(X)
arm_dist = self.model.predict(X)
reward_sample = arm_dist.rvs()
#print(reward_sample)
#print(arm)
reward_dict[arm_ix] = reward_sample
max_ix = max(reward_dict, key=lambda key: reward_dict[key])
return {'arm' :self.arms[max_ix], 'arm_ix':max_ix}
def choose_arm_and_plot(self,context):
colors = iter(cm.rainbow(np.linspace(0, 1, self.num_arms)))
reward_list = []
for arm in range(self.num_arms):
X = np.atleast_2d(np.append(arm,context))
print(X)
arm_dist = self.model.predict(X)
reward_sample = arm_dist.rvs()
print(reward_sample)
reward_list += [reward_sample]
# plot
color = next(colors)
X_pdf = np.linspace(-10, 10, 1000)
pdf = arm_dist.pdf(X_pdf)
height = arm_dist.pdf(reward_sample)
# plotting distriution of weights
plt.plot(X_pdf, pdf, color=color, linewidth=2, alpha=0.5, label=f'estimated parameter arm {arm}')
plt.fill_between(X_pdf, pdf, 0, color=color, alpha=0.2)
plt.scatter(reward_sample, height, s = 200,label=f'sample drawn from arm {arm}')
plt.legend()
return np.argmax(reward_list)
gcssm = GaussianContextualSamplerSingleModel(BatchBayesLinReg,num_arms=5, num_context = 1,model_params={'alpha':1, 'beta':4})
gcssm.model, gcssm.arms.values()
gcssm.choose_arm(np.array([9]))
gcssm.choose_arm(np.array([9]))
gcssm.update(arm_ix=0,context=np.array([9]),reward=1)
gcssm.arms
```
| github_jupyter |
# Excercises Electric Machinery Fundamentals
## Chapter 4
## Problem 4-29
```
%pylab notebook
```
### Description
A 100-MVA, 14.4-kV 0.8-PF-lagging, Y-connected synchronous generator has a negligible armature
resistance and a synchronous reactance of 1.0 per-unit. The generator is connected in parallel with a 60-
Hz, 14.4-kV infinite bus that is capable of supplying or consuming any amount of real or reactive power
with no change in frequency or terminal voltage.
```
Sbase = 100e6 # [VA]
Vbase = 14.4e3 # [V]
ra = 0.0 # pu
xs = 1.0 # pu
PF = 0.8
```
#### (a)
* What is the synchronous reactance of the generator in ohms?
#### (b)
* What is the internal generated voltage $E_A$ of this generator under rated conditions?
#### (c)
* What is the armature current $I_A$ in this machine at rated conditions?
#### (d)
Suppose that the generator is initially operating at rated conditions. If the internal generated voltage $E_A$ is decreased by 5 percent
* What will the new armature current $I_A$ be?
#### (e)
* Repeat part (d) for 10, 15, 20, and 25 percent reductions in $E_A$ .
#### (f)
* Plot the magnitude of the armature current $I_A$ as a function of $E_A$ .
### SOLUTION
#### (a)
The rated phase voltage of this generator is:
```
Vphi_base = Vbase / sqrt(3)
print('Vphi = {:.0f} V'.format(Vphi_base))
```
The base impedance of this generator is:
$$Z_\text{base} = \frac{3V^2_{\phi,\text{base}}}{S_\text{base}}$$
```
Zbase = (3*Vphi_base**2) / Sbase
print('Zbase = {:.2f} Ω'.format(Zbase))
```
Therefore,
```
Ra = ra * Zbase
Xs = xs * Zbase
print('''
Ra = {:.1f} Ω Xs = {:.1f} Ω
========================'''.format(Ra, Xs))
```
#### (b)
The rated armature current is:
$$I_A = I_F = \frac{S}{\sqrt{3}V_T}$$
```
ia = Sbase / (sqrt(3)*Vbase)
print('ia = {:.0f} A'.format(ia))
```
The power factor is 0.8 lagging, so
```
Ia_angle = -arccos(PF)
Ia = ia * (cos(Ia_angle) + sin(Ia_angle) * 1j)
print('Ia = {:.0f} A ∠{:.2f}°'.format(abs(Ia), Ia_angle/pi*180))
```
Therefore, the internal generated voltage is:
$$\vec{E}_A = \vec{V}_\phi + R_A\vec{I}_A + jX_S\vec{I}_A$$
```
EA = Vphi_base + Ra*Ia + Xs*1j *Ia
EA_angle = arctan(EA.imag/EA.real)
print('''
EA = {:.0f} V ∠{:.2f}°
===================='''.format(abs(EA), EA_angle/pi*180))
```
#### (c)
From the above calculations
```
print('''
Ia = {:.0f} V ∠{:.2f}°
===================='''.format(abs(Ia), Ia_angle/pi*180))
```
#### (d)
If $E_A$ is decreased by 5%, the armature current will change as shown below. Note that the infinite bus will keep $V_\phi$ and $\omega_m$ constant. Also, since the prime mover hasn’t changed, the power supplied by the generator will be constant.
<img src="figs/Problem_4-29.png" width="60%">
$P = \frac{3V_\phi E_A}{X_S}\sin{\delta} =$ constant, so: $E_{A1}\sin{\delta_1} = E_{A2}\sin{\delta_2}$
With a **5%** decrease,
```
Ea1 = abs(EA)
Ea2 = Ea1 * 0.95
print('Ea1 = {:.0f} V Ea2 = {:.0f} V'.format(Ea1, Ea2))
```
$$\delta_2 = \arcsin\left(\frac{E_{A1}}{E_{A2}}\sin{\delta_1}\right)$$
```
delta1 = EA_angle
delta2 = arcsin(Ea1/Ea2 * sin(delta1))
print('delta2 = {:.1f}°'.format(delta2/pi*180))
EA2 = Ea2 * exp(1j*delta2)
```
Therefore, the new armature current is:
$$\vec{I}_A = \frac{\vec{E}_{A2} - \vec{V}_\phi}{jX_S}$$
```
Ia2 = (EA2 - Vphi_base) / (Xs*1j)
Ia2_angle = arctan(Ia2.imag/Ia2.real)
print('''
Ia2 = {:.0f} V ∠{:.1f}°
===================='''.format(abs(Ia2), Ia2_angle/pi*180))
```
#### (e)
Repeating part (d):
With a **10%** decrease,
```
Ea1 = abs(EA)
Ea3 = Ea1 * 0.9
print('Ea1 = {:.0f} V Ea3 = {:.0f} V'.format(Ea1, Ea3))
delta1 = EA_angle
delta3 = arcsin(Ea1/Ea3 * sin(delta1))
print('delta3 = {:.1f}°'.format(delta3/pi*180))
EA3 = Ea3 * exp(1j*delta3)
```
Therefore, the new armature current is:
```
Ia3 = (EA3 - Vphi_base) / (Xs*1j)
Ia3_angle = arctan(Ia3.imag/Ia3.real)
print('''
Ia3 = {:.0f} A ∠{:.1f}°
====================='''.format(abs(Ia3), Ia3_angle/pi *180))
```
With a **15%** decrease,
```
Ea1 = abs(EA)
Ea4 = Ea1 * 0.85
print('Ea1 = {:.0f} V Ea4 = {:.0f} V'.format(Ea1, Ea4))
delta1 = EA_angle
delta4 = arcsin(Ea1/Ea4 * sin(delta1))
print('delta4 = {:.1f}°'.format(delta4/pi*180))
EA4 = Ea4 * exp(1j*delta4)
```
Therefore, the new armature current is:
```
Ia4 = (EA4 - Vphi_base) / (Xs*1j)
Ia4_angle = arctan(Ia4.imag/Ia4.real)
print('''
Ia4 = {:.0f} A ∠{:.1f}°
====================='''.format(abs(Ia4), Ia4_angle/pi *180))
```
With a **20%** decrease,
```
Ea1 = abs(EA)
Ea5 = Ea1 * 0.80
print('Ea1 = {:.0f} V Ea5 = {:.0f} V'.format(Ea1, Ea5))
delta1 = EA_angle
delta5 = arcsin(Ea1/Ea5 * sin(delta1))
print('delta5 = {:.1f}°'.format(delta5/pi*180))
EA5 = Ea5 * exp(1j*delta5)
```
Therefore, the new armature current is:
```
Ia5 = (EA5 - Vphi_base) / (Xs*1j)
Ia5_angle = arctan(Ia5.imag/Ia5.real)
print('''
Ia5 = {:.0f} A ∠{:.1f}°
====================='''.format(abs(Ia5), Ia5_angle/pi *180))
```
With a **25%** decrease,
```
Ea1 = abs(EA)
Ea6 = Ea1 * 0.75
print('Ea1 = {:.0f} V Ea6 = {:.0f} V'.format(Ea1, Ea6))
delta1 = EA_angle
delta6 = arcsin(Ea1/Ea6 * sin(delta1))
print('delta6 = {:.1f}°'.format(delta6/pi*180))
EA6 = Ea6 * exp(1j*delta6)
```
Therefore, the new armature current is:
```
Ia6 = (EA6 - Vphi_base) / (Xs*1j)
Ia6_angle = arctan(Ia6.imag/Ia6.real)
print('''
Ia6 = {:.0f} A ∠{:.1f}°
====================='''.format(abs(Ia6), Ia6_angle/pi *180))
```
#### (f)
We are going to plot the magnitude of the armature current $I_A$ as a function of $E_A$ below.
Define values for this generator:
```
Ea = linspace(0.55, 1.00, 46) * abs(EA)
d1 = EA_angle
```
Calculate delta for each $E_A$
```
d_ = arcsin( abs(EA) / Ea * sin(d1))
```
Calculate Ia for each flux:
```
Ea_ = Ea * exp(1j*d_)
Ia_ = ( Ea_ - Vphi_base ) / (Xs*1j)
```
Plot the armature current versus Ea:
```
rc('text', usetex=True) # enable LaTeX commands for plot
title(r'Armature current versus $E_A$')
xlabel(r'$E_A$ [kV]')
ylabel(r'$I_A$ [A]')
plot(abs(Ea_)/1000,abs(Ia_), linewidth = 2)
grid()
```
| github_jupyter |
```
import sys
import tensorflow as tf
from tensorflow.keras import layers, activations, losses, Model, Input
from tensorflow.nn import leaky_relu
import numpy as np
from itertools import combinations
from tensorflow.keras.utils import plot_model, Progbar
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
# In case your sys.path does not contain the base repo, go there.
print(sys.path)
%cd '~/ml-solr-course'
```
The idea behind RankNet is to model the **joint probability** that `document i` comes before `document j` as the following:
$P_{ij} = 1$ if $s_i > s_j$
$P_{ij} = 0.5$ if $s_i = s_j$
$P_{ij} = 0$ if $s_i < s_j$
So for *every pair of inputs* we will calculate both outputs, substract them, pass a logistic function to model the probability:
<img src="files/ranknet.png">
```
# model architecture
class RankNet(Model):
def __init__(self):
super().__init__()
self.dense = [layers.Dense(16, activation=leaky_relu), layers.Dense(8, activation=leaky_relu)]
self.o = layers.Dense(1, activation='linear')
self.oi_minus_oj = layers.Subtract()
def call(self, inputs):
xi, xj = inputs
densei = self.dense[0](xi)
densej = self.dense[0](xj)
for dense in self.dense[1:]:
densei = dense(densei)
densej = dense(densej)
oi = self.o(densei)
oj= self.o(densej)
oij = self.oi_minus_oj([oi, oj])
output = layers.Activation('sigmoid')(oij)
return output
def build_graph(self):
x = [Input(shape=(10)), Input(shape=(10))]
return Model(inputs=x, outputs=self.call(x))
nb_query = 20
query = np.array([i+1 for i in range(nb_query) for x in range(int(np.ceil(np.abs(np.random.normal(0,scale=15))+2)))])
doc_features = np.random.random((len(query), 10))
doc_scores = np.random.randint(5, size=len(query)).astype(np.float32)
query
print(doc_scores)
doc_features
# put data into pairs
xi = []
xj = []
pij = []
pair_id = []
pair_query_id = []
for q in np.unique(query):
query_idx = np.where(query == q)[0]
for pair_idx in combinations(query_idx, 2):
pair_query_id.append(q)
pair_id.append(pair_idx)
i = pair_idx[0]
j = pair_idx[1]
xi.append(doc_features[i])
xj.append(doc_features[j])
if doc_scores[i] == doc_scores[j]:
_pij = 0.5
elif doc_scores[i] > doc_scores[j]:
_pij = 1
else:
_pij = 0
pij.append(_pij)
xi = np.array(xi)
xj = np.array(xj)
pij = np.array(pij)
pair_query_id = np.array(pair_query_id)
xi_train, xi_test, xj_train, xj_test, pij_train, pij_test, pair_id_train, pair_id_test = train_test_split(
xi, xj, pij, pair_id, test_size=0.2, stratify=pair_query_id)
# train model using compile and fit
ranknet = RankNet()
ranknet.compile(optimizer='adam', loss='binary_crossentropy')
history = ranknet.fit([xi_train, xj_train], pij_train, epochs=50, batch_size=1, validation_data=([xi_test, xj_test], pij_test))
# function for plotting loss
def plot_metrics(train_metric, val_metric=None, metric_name=None, title=None, ylim=5):
plt.title(title)
plt.ylim(0,ylim)
plt.plot(train_metric,color='blue',label=metric_name)
if val_metric is not None: plt.plot(val_metric,color='green',label='val_' + metric_name)
plt.legend(loc="upper right")
# plot loss history
plot_metrics(history.history['loss'], history.history['val_loss'], "Loss", "Loss", ylim=1.0)
new_doci = [np.random.random(10), np.random.random(10)]
new_docj = [np.random.random(10), np.random.random(10)]
inputs = tf.constant(np.array([new_doci, new_docj]))
ranknet(inputs)
```
| github_jupyter |
```
import logging
logging.basicConfig(level="INFO", format="[%(name)s - %(levelname)s] %(message)s")
ROOT = logging.getLogger()
import pandas as pd
import sanger_sequencing
from pandas import read_excel
def tube_samples(filepath):
"""Read the particular excel file into a pandas.DataFrame."""
df = read_excel(filepath, converters={
"Primer ID": str,
"Plasmid ID": str
})
df.dropna(how="any", inplace=True)
return df
old_template = tube_samples("../sanger-service/cfb/tests/data/Mix2Seq_SA00360224_sample.xls")
old_template.head()
from glob import glob
from Bio import SeqIO
from os.path import splitext, basename
def genbank_db(samples):
db = dict()
for plasmid_id in samples["Plasmid ID"].unique():
path = glob(f"../sanger-service/cfb/tests/data/pCfB{plasmid_id}*.gbk")[0]
db[plasmid_id] = splitext(basename(path))[0], SeqIO.read(path, "gb")
return db
plasmids = genbank_db(old_template)
plasmids
from os.path import basename, splitext
def ab1_filter(filepath) -> bool:
"""Select non-hidden ab1 files."""
if basename(filepath).startswith('.'):
return False
return splitext(filepath)[1].lower().endswith('.ab1')
def get_tube_code(filepath) -> str:
"""Extract the tube code part from a filename."""
return splitext(basename(filepath))[0].split("_")[-1]
from zipfile import ZipFile
from io import BytesIO
def parse_tube_ab1(archive: ZipFile,
tube_codes):
"""Read particular ab1 sequences from an archive."""
sequences = dict()
# Create a tube code -> file name map from the archive's contents.
names = {get_tube_code(f): f for f in filter(
ab1_filter, archive.namelist())}
# Try to extract all desired sequences.
for code in tube_codes:
if code not in names:
LOGGER.error("No ab1 file found for tube with code '%s'.", code)
continue
with archive.open(names[code]) as file_handle:
# We use `SeqIO.read` because ab1 files contain only a single
# sequence record.
sequences[code] = SeqIO.read(BytesIO(file_handle.read()), 'abi')
return sequences
def extract_sequences(template, archive):
return parse_tube_ab1(archive, template['Tube Code'].unique())
zip_path = "../sanger-service/cfb/tests/data/11104089228-1_SCF_SEQ_ABI.zip"
with ZipFile(zip_path) as archive:
samples = extract_sequences(old_template, archive)
from pandas import DataFrame
template = DataFrame({
"plasmid": old_template["Plasmid ID"],
"primer": old_template["Primer ID"],
"sample": old_template["Tube Code"]
})
new_plasmids = {pid: seq for pid, (name, seq) in plasmids.items()}
template.head()
from sanger_sequencing.api import plasmid_report
ROOT.setLevel(logging.INFO)
report = plasmid_report("7138", new_plasmids["7138"], template[:5], samples)
report["samples"][0]["conflicts"]
for r in report["samples"]:
print(r["conflicts"])
```
| github_jupyter |
```
%matplotlib inline
import pandas as pd
import numpy as np
import seaborn as sns; sns.set()
import matplotlib.pyplot as plt
from scipy import stats
# dataset test
from sklearn.datasets import make_blobs
X, y =make_blobs(n_samples=50, centers = 2,random_state = 0, cluster_std = 0.60)
plt.scatter(X[:,0],X[:,1],c=y,cmap='autumn')
```
# Possible seprators
```
xfit = np.linspace(-1,3.5)
plt.scatter(X[:,0],X[:,1],c=y,cmap='autumn')
for m , b in [(1,0.65), (0.5,1.6), (-0.2,2.9)]:
yfit = m*xfit + b
plt.plot(xfit, yfit,'-k')
plt.xlim(1-1,3.5)
```
# Margins
```
xfit = np.linspace(-1,3.5)
plt.scatter(X[:,0],X[:,1],c=y,cmap='autumn')
for m , b , d in [(1,0.65,0.33), (0.5,1.6,0.5), (-0.2,2.9,0.2)]:
yfit = m*xfit + b
plt.plot(xfit, yfit,'-k')
plt.fill_between(xfit,yfit-d, yfit+d, edgecolor='None', color='#AAAAAA',alpha=0.4)
plt.xlim(1-1,3.5)
```
# Training SVM
```
from sklearn.svm import SVC
model = SVC(kernel='linear',C=1).fit(X,y)
#plotting decision boundary with maximum margin
def plot_decision_boundaries(plot, ax=None, plot_support=True):
if ax is None:
ax = plt.gca()
xlim= ax.get_xlim()
ylim = ax.get_ylim()
x = np.linspace(xlim[0], xlim[1],30)
y= np.linspace(ylim[0],ylim[1],30)
Y, X = np.meshgrid(y,x)
xy= np.vstack([X.ravel(),Y.ravel()]).T
P= model.decision_function(xy).reshape(X.shape)
#plot db and margins
ax.contour(X,Y,P, colors='k',levels=[-1,0,1], alpha = 0.5,linestyles=['--','-','--'])
#plotting support vectors
if plot_support:
ax.scatter(model.support_vectors_[:,0],model.support_vectors_[:,1],s=300,linewidth=1,facecolor='None')
ax.set_xlim(xlim)
ax.set_ylim(ylim)
#plotting decision boundary for the model
plt.scatter(X[:,0],X[:,1],c=y,cmap='autumn')
plot_decision_boundaries(model)
```
# Face recognition model project
```
from sklearn.datasets import fetch_lfw_people
faces= fetch_lfw_people(min_faces_per_person= 60)
print(faces.target_names)
print(faces.images.shape)
fig,ax = plt.subplots(3,3)
for i, axi in enumerate(ax.flat):
axi.imshow(faces.images[i], cmap='bone')
axi.set(xticks=[],yticks=[], xlabel = faces.target_names[faces.target[i]])
# create pipeline for principal component analysis and support vector classifier
from sklearn.svm import SVC
from sklearn.decomposition import PCA
from sklearn.pipeline import make_pipeline
pca= PCA(n_components=150, random_state=42,whiten=True)
svc = SVC(kernel='rbf',class_weight='balanced',random_state=42)
model1 = make_pipeline(pca, svc)
# splitting the dataset
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(faces.data, faces.target , random_state=42)
print(y_train)
# find best model
from sklearn.model_selection import GridSearchCV
param_grid = {'svc__C':[1,5,7,10,50],
'svc__gamma':[0.0001,0.0005,0.001,0.005,0.01]}
grid_search = GridSearchCV(model1, param_grid=param_grid)
%time grid_search.fit(X_train,y_train)
print (grid_search.best_params_)
# predict
model = grid_search.best_estimator_
yfit = model.predict(X_test)
fig, ax = plt.subplots(4,6)
for i , axi in enumerate(ax.flat):
axi.imshow(X_test[i].reshape(62,47), cmap='bone')
axi.set(xticks=[],yticks=[])
axi.set_ylabel(faces.target_names[yfit[i]].split()[-1],
color='black' if yfit[i] == y_test[i] else 'red' )
fig.suptitle("predicted names; incorrect labels in Red",size =14)
```
# Evaluation
```
from sklearn.metrics import classification_report
print(classification_report(y_test, yfit, target_names=faces.target_names))
#plotting confusion matrix
from sklearn.metrics import confusion_matrix
mat=confusion_matrix(y_test, yfit)
sns.heatmap(mat.T, square=True, annot= True, fmt='d', cbar=False,
xticklabels=faces.target_names,
yticklabels= faces.target_names)
```
| github_jupyter |
```
import numpy as np
from matplotlib import pyplot as plt
import baseline
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
import scipy.fftpack as F
%pylab inline
data = baseline.prepare_data('/Users/daphne/Dropbox (MIT)/pd-mlhc/CIS')
subject_ids, measurement_ids, all_data, all_n_data, on_off_labels, dyskinesia_labels, tremor_labels = data
X_tre, y_tre = baseline.cleaned_data(all_data, tremor_labels)
X_med, y_med = baseline.cleaned_data(all_data, on_off_labels)
X_dys, y_dys = baseline.cleaned_data(all_data, dyskinesia_labels)
#############################################################
chosen = 'medication'
X_chosen = X_med
y_chosen = y_med
# there are 13 patients. Which go in train? which go in val?
np.random.seed(123)
rate = 0.7
train = []
val = []
for subj in subject_ids :
r = np.random.rand()
if r < rate :
train.append(subj)
else :
val.append(subj)
#############################################################
avgs_train = []
var_train = []
y_train = []
avgs_validate = []
var_validate = []
y_validate = []
both_train = []
both_validate = []
for i in range(len(X_chosen)) :
s = X_chosen[i]
ident = subject_ids[i]
for m in s :
a = np.mean(m,axis=0)
v = np.var(m,axis=0)
if ident in train :
avgs_train.append(a)
var_train.append(v)
both_train.append(np.hstack((a,v)))
elif ident in val :
avgs_validate.append(a)
var_validate.append(v)
both_validate.append(np.hstack((a,v)))
if ident in train :
y_train += y_chosen[i]
elif ident in val :
y_validate += y_chosen[i]
# do fourier transform... the above is not good enough
# don't forget - sliding window, regressing against time from event!
clf = LogisticRegression(solver='lbfgs',multi_class='auto').fit(both_train, y_train)
print('Performance on training data for {} labels is {:.3f}'.format(chosen,clf.score(both_train,y_train)))
print('Performance on validation data for {} labels is {:.3f}'.format(chosen,clf.score(both_validate,y_validate)))
clf = SVC(gamma='scale',kernel='sigmoid').fit(both_train, y_train)
print('Performance on training data for {} labels is {:.3f}'.format(chosen, clf.score(both_train,y_train)))
print('Performance on validation data for {} labels is {:.3f}'.format(chosen, clf.score(both_validate,y_validate)))
def do_fft(signal) :
N = signal.shape[0]
T = .02 #seconds per sample
x = np.linspace(0.0, N*T, N)
yf = F.fft(signal)
xf = np.linspace(0.0, 1.0/(2.0*T), N/2)
return xf, 2.0/N * np.abs(yf[:N//2])
'''
window is in seconds
'''
def window(signal, time_window=30, fs=1/0.02) :
n = signal.shape[0]
window = int(time_window * fs)
num_splits = np.floor(n/window)
sigs_raw_trimmed = signal[:(-1*(n%window)),:]
sigs_list = np.split(sigs_raw_trimmed, num_splits, axis=0)
return sigs_list
sigs_list = window(X_tre[0][0])
avg_segs = [np.mean(seg,axis=0) for seg in sigs_list]
var_segs = [np.var(seg,axis=0) for seg in sigs_list]
```
| github_jupyter |
<script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
<script>
window.dataLayer = window.dataLayer || [];
function gtag(){dataLayer.push(arguments);}
gtag('js', new Date());
gtag('config', 'UA-59152712-8');
</script>
# Start-to-Finish Example: Setting up Polytropic [TOV](https://en.wikipedia.org/wiki/Tolman%E2%80%93Oppenheimer%E2%80%93Volkoff_equation) Initial Data, in Curvilinear Coordinates
## Authors: Zach Etienne, Phil Chang, and Leo Werneck
### Formatting improvements courtesy Brandon Clark
## This module sets up initial data for a TOV star in *spherical, isotropic coordinates*, using the *Numerical* ADM Spherical to BSSN Curvilinear initial data module (numerical = BSSN $\lambda^i$'s are computed using finite-difference derivatives instead of exact expressions).
**Notebook Status:** <font color='green'><b> Validated </b></font>
**Validation Notes:** This module has been validated to exhibit convergence to zero of the Hamiltonian constraint violation at the expected order to the exact solution (see [plots](#convergence) at bottom). Note that convergence at the surface of the star will be lower order due to the sharp drop to zero in $T^{\mu\nu}$.</font>
### NRPy+ Source Code for this module:
* [TOV/TOV_Solver.py](../edit/TOV/TOV_Solver.py); ([**NRPy+ Tutorial module reviewing mathematical formulation and equations solved**](Tutorial-ADM_Initial_Data-TOV.ipynb)); ([**start-to-finish NRPy+ Tutorial module demonstrating that initial data satisfy Hamiltonian constraint**](Tutorial-Start_to_Finish-BSSNCurvilinear-Setting_up_TOV_initial_data.ipynb)): Tolman-Oppenheimer-Volkoff (TOV) initial data; defines all ADM variables and nonzero $T^{\mu\nu}$ components in Spherical basis.
* [BSSN/ADM_Numerical_Spherical_or_Cartesian_to_BSSNCurvilinear.py](../edit/BSSN/ADM_Numerical_Spherical_or_Cartesian_to_BSSNCurvilinear.py); [\[**tutorial**\]](Tutorial-ADM_Initial_Data-Converting_Numerical_ADM_Spherical_or_Cartesian_to_BSSNCurvilinear.ipynb): *Numerical* Spherical ADM$\to$Curvilinear BSSN converter function
* [BSSN/BSSN_constraints.py](../edit/BSSN/BSSN_constraints.py); [\[**tutorial**\]](Tutorial-BSSN_constraints.ipynb): Hamiltonian constraint in BSSN curvilinear basis/coordinates
## Introduction:
Here we use NRPy+ to set up initial data for a [simple polytrope TOV star](https://en.wikipedia.org/wiki/Tolman%E2%80%93Oppenheimer%E2%80%93Volkoff_equation).
The entire algorithm is outlined as follows, with links to the relevant NRPy+ tutorial notebooks listed at each step:
1. Allocate memory for gridfunctions, including temporary storage for the Method of Lines time integration [(**NRPy+ tutorial on NRPy+ Method of Lines algorithm**)](Tutorial-Method_of_Lines-C_Code_Generation.ipynb).
1. Set gridfunction values to initial data
* [**NRPy+ tutorial on TOV initial data**](Tutorial-ADM_Initial_Data-TOV.ipynb)
* [**NRPy+ tutorial on validating TOV initial data**](Tutorial-Start_to_Finish-BSSNCurvilinear-Setting_up_TOV_initial_data.ipynb).
1. Evaluate the Hamiltonian constraint violation
* [**NRPy+ tutorial on BSSN constraints**](Tutorial-BSSN_constraints.ipynb)
1. Repeat above steps at two numerical resolutions to confirm convergence of Hamiltonian constraint violation to zero.
<a id='toc'></a>
# Table of Contents
$$\label{toc}$$
This notebook is organized as follows
1. [Step 1](#initializenrpy): Set core NRPy+ parameters for numerical grids and reference metric
1. [Step 2](#adm_id_tov): Set up ADM initial data for polytropic TOV Star
1. [Step 2.a](#tov_interp): Interpolating the TOV data file as needed
1. [Step 2.b](#source): Compute source terms $S_{ij}$, $S_{i}$, $S$, and $\rho$
1. [Step 2.c](#jacobian): Jacobian transformation on the ADM/BSSN source terms
1. [Step 2.d](#tensor): Rescale tensorial quantities
1. [Step 3](#adm_id_spacetime): Convert ADM spacetime quantity initial data from Spherical to BSSN Curvilinear coordinates
1. [Step 4](#validate): Validating that the TOV initial data satisfy the Hamiltonian constraint
1. [Step 4.a](#ham_const_output): Output the Hamiltonian Constraint
1. [Step 4.b](#apply_bcs): Apply singular, curvilinear coordinate boundary conditions
1. [Step 4.c](#enforce3metric): Enforce conformal 3-metric $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$
1. [Step 5](#mainc): `TOV_Playground.c`: The Main C Code
1. [Step 6](#plot): Plotting the single-neutron-star initial data
1. [Step 7](#convergence): Validation: Convergence of numerical errors (Hamiltonian constraint violation) to zero
1. [Step 8](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
<a id='initalizenrpy'></a>
# Step 1: Set core NRPy+ parameters for numerical grids and reference metric \[Back to [top](#toc)\]
$$\label{initializenrpy}$$
```
# Step P1: Import needed NRPy+ core modules:
from outputC import lhrh,outCfunction,outputC # NRPy+: Core C code output module
import NRPy_param_funcs as par # NRPy+: Parameter interface
import sympy as sp # SymPy: The Python computer algebra package upon which NRPy+ depends
import finite_difference as fin # NRPy+: Finite difference C code generation module
import grid as gri # NRPy+: Functions having to do with numerical grids
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import reference_metric as rfm # NRPy+: Reference metric support
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
import shutil, os, sys # Standard Python modules for multiplatform OS-level functions
# Step P2: Create C code output directory:
Ccodesdir = os.path.join("TOVID_Ccodes/")
# First remove C code output directory if it exists
# Courtesy https://stackoverflow.com/questions/303200/how-do-i-remove-delete-a-folder-that-is-not-empty
# !rm -r ScalarWaveCurvilinear_Playground_Ccodes
shutil.rmtree(Ccodesdir, ignore_errors=True)
# Then create a fresh directory
cmd.mkdir(Ccodesdir)
# Step P3: Create executable output directory:
outdir = os.path.join(Ccodesdir,"output/")
cmd.mkdir(outdir)
# Step 1: Set the spatial dimension parameter
# to three this time, and then read
# the parameter as DIM.
par.set_parval_from_str("grid::DIM",3)
DIM = par.parval_from_str("grid::DIM")
# Step 2: Set some core parameters, including CoordSystem MoL timestepping algorithm,
# FD order, floating point precision, and CFL factor:
# Choices are: Spherical, SinhSpherical, SinhSphericalv2, Cylindrical, SinhCylindrical,
# SymTP, SinhSymTP
CoordSystem = "Spherical"
# Step 2.a: Set defaults for Coordinate system parameters.
# These are perhaps the most commonly adjusted parameters,
# so we enable modifications at this high level.
# domain_size = 7.5 # SET BELOW BASED ON TOV STELLAR RADIUS
# sinh_width sets the default value for:
# * SinhSpherical's params.SINHW
# * SinhCylindrical's params.SINHW{RHO,Z}
# * SinhSymTP's params.SINHWAA
sinh_width = 0.4 # If Sinh* coordinates chosen
# sinhv2_const_dr sets the default value for:
# * SinhSphericalv2's params.const_dr
# * SinhCylindricalv2's params.const_d{rho,z}
sinhv2_const_dr = 0.05# If Sinh*v2 coordinates chosen
# SymTP_bScale sets the default value for:
# * SinhSymTP's params.bScale
SymTP_bScale = 0.5 # If SymTP chosen
# Step 2.b: Set the order of spatial finite difference derivatives;
# and the core data type.
FD_order = 4 # Finite difference order: even numbers only, starting with 2. 12 is generally unstable
REAL = "double" # Best to use double here.
# Step 3: Set the coordinate system for the numerical grid
par.set_parval_from_str("reference_metric::CoordSystem",CoordSystem)
rfm.reference_metric() # Create ReU, ReDD needed for rescaling B-L initial data, generating BSSN RHSs, etc.
# Step 4: Set the finite differencing order to FD_order (set above).
par.set_parval_from_str("finite_difference::FD_CENTDERIVS_ORDER", FD_order)
# Step 5: Set the direction=2 (phi) axis to be the symmetry axis; i.e.,
# axis "2", corresponding to the i2 direction.
# This sets all spatial derivatives in the phi direction to zero.
par.set_parval_from_str("indexedexp::symmetry_axes","2")
# Step 6: The MoLtimestepping interface is only used for memory allocation/deallocation
import MoLtimestepping.C_Code_Generation as MoL
from MoLtimestepping.RK_Butcher_Table_Dictionary import Butcher_dict
RK_method = "Euler" # DOES NOT MATTER; Again MoL interface is only used for memory alloc/dealloc.
RK_order = Butcher_dict[RK_method][1]
cmd.mkdir(os.path.join(Ccodesdir,"MoLtimestepping/"))
MoL.MoL_C_Code_Generation(RK_method, RHS_string = "", post_RHS_string = "",
outdir = os.path.join(Ccodesdir,"MoLtimestepping/"))
# Step 7: Polytropic EOS setup
# For EOS_type, choose either "SimplePolytrope" or "PiecewisePolytrope"
EOS_type = "SimplePolytrope"
# If "PiecewisePolytrope" is chosen as EOS_type, you
# must also choose the name of the EOS, which can
# be any of the following:
# 'PAL6', 'SLy', 'APR1', 'APR2', 'APR3', 'APR4',
# 'FPS', 'WFF1', 'WFF2', 'WFF3', 'BBB2', 'BPAL12',
# 'ENG', 'MPA1', 'MS1', 'MS2', 'MS1b', 'PS', 'GS1',
# 'GS2', 'BGN1H1', 'GNH3', 'H1', 'H2', 'H3', 'H4',
# 'H5', 'H6', 'H7', 'PCL2', 'ALF1', 'ALF2', 'ALF3',
# 'ALF4'
EOS_name = 'SLy' # <-- IGNORED IF EOS_type is not PiecewisePolytrope.
```
<a id='adm_id_tov'></a>
# Step 2: Set up ADM initial data for polytropic TOV Star \[Back to [top](#toc)\]
$$\label{adm_id_tov}$$
As documented [in the TOV Initial Data NRPy+ Tutorial Module](Tutorial-TOV_Initial_Data.ipynb) ([older version here](Tutorial-GRMHD_UnitConversion.ipynb)), we will now set up TOV initial data, storing the densely-sampled result to file (***Courtesy Phil Chang***).
The TOV solver uses an ODE integration routine provided by scipy, so we first make sure that scipy is installed:
```
!pip install scipy > /dev/null
```
Next we call the [`TOV.TOV_Solver()` function](../edit/TOV/TOV_Solver.py) ([NRPy+ Tutorial module](Tutorial-ADM_Initial_Data-TOV.ipynb)) to set up the initial data, using the default parameters for initial data. This function outputs the solution to a file named "outputTOVpolytrope.txt".
```
##########################
# Polytropic EOS example #
##########################
import TOV.Polytropic_EOSs as ppeos
if EOS_type == "SimplePolytrope":
# Set neos = 1 (single polytrope)
neos = 1
# Set rho_poly_tab (not needed for a single polytrope)
rho_poly_tab = []
# Set Gamma_poly_tab
Gamma_poly_tab = [2.0]
# Set K_poly_tab0
K_poly_tab0 = 1. # ZACH NOTES: CHANGED FROM 100.
# Set the eos quantities
eos = ppeos.set_up_EOS_parameters__complete_set_of_input_variables(neos,rho_poly_tab,Gamma_poly_tab,K_poly_tab0)
rho_baryon_central = 0.129285
elif EOS_type == "PiecewisePolytrope":
eos = ppeos.set_up_EOS_parameters__Read_et_al_input_variables(EOS_name)
rho_baryon_central=2.0
else:
print("""Error: unknown EOS_type. Valid types are 'SimplePolytrope' and 'PiecewisePolytrope' """)
sys.exit(1)
import TOV.TOV_Solver as TOV
M_TOV, R_Schw_TOV, R_iso_TOV = TOV.TOV_Solver(eos,
outfile="outputTOVpolytrope.txt",
rho_baryon_central=rho_baryon_central,
return_M_RSchw_and_Riso = True,
verbose = True)
# domain_size sets the default value for:
# * Spherical's params.RMAX
# * SinhSpherical*'s params.AMAX
# * Cartesians*'s -params.{x,y,z}min & .{x,y,z}max
# * Cylindrical's -params.ZMIN & .{Z,RHO}MAX
# * SinhCylindrical's params.AMPL{RHO,Z}
# * *SymTP's params.AMAX
domain_size = 2.0 * R_iso_TOV
```
<a id='tov_interp'></a>
## Step 2.a: Interpolate the TOV data file as needed to set up ADM spacetime quantities in spherical basis (for input into the `Converting_Numerical_ADM_Spherical_or_Cartesian_to_BSSNCurvilinear` module) and $T^{\mu\nu}$ in the chosen reference metric basis \[Back to [top](#toc)\]
$$\label{tov_interp}$$
The TOV data file just written stored $\left(r,\rho(r),P(r),M(r),e^{\nu(r)}\right)$, where $\rho(r)$ is the total mass-energy density (cf. $\rho_{\text{baryonic}}$).
**METRIC DATA IN TERMS OF ADM QUANTITIES**
The [TOV line element](https://en.wikipedia.org/wiki/Tolman%E2%80%93Oppenheimer%E2%80%93Volkoff_equation) in *Schwarzschild coordinates* is written (in the $-+++$ form):
$$
ds^2 = - c^2 e^\nu dt^2 + \left(1 - \frac{2GM}{rc^2}\right)^{-1} dr^2 + r^2 d\Omega^2.
$$
In *isotropic coordinates* with $G=c=1$ (i.e., the coordinate system we'd prefer to use), the ($-+++$ form) line element is written:
$$
ds^2 = - e^{\nu} dt^2 + e^{4\phi} \left(d\bar{r}^2 + \bar{r}^2 d\Omega^2\right),
$$
where $\phi$ here is the *conformal factor*.
The ADM 3+1 line element for this diagonal metric in isotropic spherical coordinates is given by:
$$
ds^2 = (-\alpha^2 + \beta_k \beta^k) dt^2 + \gamma_{\bar{r}\bar{r}} d\bar{r}^2 + \gamma_{\theta\theta} d\theta^2+ \gamma_{\phi\phi} d\phi^2,
$$
from which we can immediately read off the ADM quantities:
\begin{align}
\alpha &= e^{\nu(\bar{r})/2} \\
\beta^k &= 0 \\
\gamma_{\bar{r}\bar{r}} &= e^{4\phi}\\
\gamma_{\theta\theta} &= e^{4\phi} \bar{r}^2 \\
\gamma_{\phi\phi} &= e^{4\phi} \bar{r}^2 \sin^2 \theta \\
\end{align}
**STRESS-ENERGY TENSOR $T^{\mu\nu}$**
We will also need the stress-energy tensor $T^{\mu\nu}$. [As discussed here](https://en.wikipedia.org/wiki/Tolman%E2%80%93Oppenheimer%E2%80%93Volkoff_equation), the stress-energy tensor is diagonal:
\begin{align}
T^t_t &= -\rho \\
T^i_j &= P \delta^i_j \\
\text{All other components of }T^\mu_\nu &= 0.
\end{align}
Since $\beta^i=0$ the inverse metric expression simplifies to (Eq. 4.49 in [Gourgoulhon](https://arxiv.org/pdf/gr-qc/0703035.pdf)):
$$
g^{\mu\nu} = \begin{pmatrix}
-\frac{1}{\alpha^2} & \frac{\beta^i}{\alpha^2} \\
\frac{\beta^i}{\alpha^2} & \gamma^{ij} - \frac{\beta^i\beta^j}{\alpha^2}
\end{pmatrix} =
\begin{pmatrix}
-\frac{1}{\alpha^2} & 0 \\
0 & \gamma^{ij}
\end{pmatrix},
$$
and since the 3-metric is diagonal we get
\begin{align}
\gamma^{\bar{r}\bar{r}} &= e^{-4\phi}\\
\gamma^{\theta\theta} &= e^{-4\phi}\frac{1}{\bar{r}^2} \\
\gamma^{\phi\phi} &= e^{-4\phi}\frac{1}{\bar{r}^2 \sin^2 \theta}.
\end{align}
Thus raising $T^\mu_\nu$ yields a diagonal $T^{\mu\nu}$
\begin{align}
T^{tt} &= -g^{tt} \rho = \frac{1}{\alpha^2} \rho = e^{-\nu(\bar{r})} \rho \\
T^{\bar{r}\bar{r}} &= g^{\bar{r}\bar{r}} P = \frac{1}{e^{4 \phi}} P \\
T^{\theta\theta} &= g^{\theta\theta} P = \frac{1}{e^{4 \phi}\bar{r}^2} P\\
T^{\phi\phi} &= g^{\phi\phi} P = \frac{1}{e^{4\phi}\bar{r}^2 \sin^2 \theta} P
\end{align}
```
thismodule = "TOVID"
rbar,theta,rho,P,expnu,exp4phi = par.Cparameters("REAL",thismodule,
["rbar","theta","rho","P","expnu","exp4phi"],1e300)
IDalpha = sp.sqrt(expnu)
gammaSphDD = ixp.zerorank2(DIM=3)
gammaSphDD[0][0] = exp4phi
gammaSphDD[1][1] = exp4phi*rbar**2
gammaSphDD[2][2] = exp4phi*rbar**2*sp.sin(theta)**2
T4SphUU = ixp.zerorank2(DIM=4)
T4SphUU[0][0] = rho/expnu
T4SphUU[1][1] = P/exp4phi
T4SphUU[2][2] = P/(exp4phi*rbar**2)
T4SphUU[3][3] = P/(exp4phi*rbar**2*sp.sin(theta)**2)
expr_list = [IDalpha]
name_list = ["*alpha"]
for i in range(3):
for j in range(i,3):
expr_list.append(gammaSphDD[i][j])
name_list.append("*gammaDD"+str(i)+str(j))
desc = """This function takes as input either (x,y,z) or (r,th,ph) and outputs
all ADM quantities in the Cartesian or Spherical basis, respectively."""
name = "ID_TOV_ADM_quantities"
outCparams = "preindent=1,outCverbose=False,includebraces=False"
outCfunction(
outfile=os.path.join(Ccodesdir, name + ".h"), desc=desc, name=name,
params=""" const REAL xyz_or_rthph[3],
const ID_inputs other_inputs,
REAL *gammaDD00,REAL *gammaDD01,REAL *gammaDD02,REAL *gammaDD11,REAL *gammaDD12,REAL *gammaDD22,
REAL *KDD00,REAL *KDD01,REAL *KDD02,REAL *KDD11,REAL *KDD12,REAL *KDD22,
REAL *alpha,
REAL *betaU0,REAL *betaU1,REAL *betaU2,
REAL *BU0,REAL *BU1,REAL *BU2""",
body="""
// Set trivial metric quantities:
*KDD00 = *KDD01 = *KDD02 = 0.0;
/**/ *KDD11 = *KDD12 = 0.0;
/**/ *KDD22 = 0.0;
*betaU0 = *betaU1 = *betaU2 = 0.0;
*BU0 = *BU1 = *BU2 = 0.0;
// Next set gamma_{ij} in spherical basis
const REAL rbar = xyz_or_rthph[0];
const REAL theta = xyz_or_rthph[1];
const REAL phi = xyz_or_rthph[2];
REAL rho,rho_baryon,P,M,expnu,exp4phi;
TOV_interpolate_1D(rbar,other_inputs.Rbar,other_inputs.Rbar_idx,other_inputs.interp_stencil_size,
other_inputs.numlines_in_file,
other_inputs.r_Schw_arr,other_inputs.rho_arr,other_inputs.rho_baryon_arr,other_inputs.P_arr,other_inputs.M_arr,
other_inputs.expnu_arr,other_inputs.exp4phi_arr,other_inputs.rbar_arr,
&rho,&rho_baryon,&P,&M,&expnu,&exp4phi);\n"""+
outputC(expr_list,name_list, "returnstring",outCparams),
opts="DisableCparameters")
```
As all input quantities are functions of $r$, we will simply read the solution from file and interpolate it to the values of $r$ needed by the initial data.
1. First we define functions `ID_TOV_ADM_quantities()` and `ID_TOV_TUPMUNU()` that call the [1D TOV interpolator function](../edit/TOV/tov_interp.h) to evaluate the ADM spacetime quantities and $T^{\mu\nu}$, respectively, at any given point $(r,\theta,\phi)$ in the Spherical basis. All quantities are defined as above.
1. Next we will construct the BSSN/ADM source terms $\{S_{ij},S_{i},S,\rho\}$ in the Spherical basis
1. Then we will perform the Jacobian transformation on $\{S_{ij},S_{i},S,\rho\}$ to the desired `(xx0,xx1,xx2)` basis
1. Next we call the *Numerical* Spherical ADM$\to$Curvilinear BSSN converter function to conver the above ADM quantities to the rescaled BSSN quantities in the desired curvilinear coordinate system: [BSSN/ADM_Numerical_Spherical_or_Cartesian_to_BSSNCurvilinear.py](../edit/BSSN/ADM_Numerical_Spherical_or_Cartesian_to_BSSNCurvilinear.py); [\[**tutorial**\]](Tutorial-ADM_Initial_Data-Converting_Numerical_ADM_Spherical_or_Cartesian_to_BSSNCurvilinear.ipynb).
$$
{\rm Jac\_dUSph\_dDrfmUD[mu][nu]} = \frac{\partial x^\mu_{\rm Sph}}{\partial x^\nu_{\rm rfm}},
$$
via exact differentiation (courtesy SymPy), and the inverse Jacobian
$$
{\rm Jac\_dUrfm\_dDSphUD[mu][nu]} = \frac{\partial x^\mu_{\rm rfm}}{\partial x^\nu_{\rm Sph}},
$$
using NRPy+'s `generic_matrix_inverter3x3()` function. In terms of these, the transformation of BSSN tensors from Spherical to `"reference_metric::CoordSystem"` coordinates may be written:
$$
T^{\mu\nu}_{\rm rfm} =
\frac{\partial x^\mu_{\rm rfm}}{\partial x^\delta_{\rm Sph}}
\frac{\partial x^\nu_{\rm rfm}}{\partial x^\sigma_{\rm Sph}} T^{\delta\sigma}_{\rm Sph}
$$
```
r_th_ph_or_Cart_xyz_oID_xx = []
CoordType_in = "Spherical"
if CoordType_in == "Spherical":
r_th_ph_or_Cart_xyz_oID_xx = rfm.xxSph
elif CoordType_in == "Cartesian":
r_th_ph_or_Cart_xyz_oID_xx = rfm.xxCart
else:
print("Error: Can only convert ADM Cartesian or Spherical initial data to BSSN Curvilinear coords.")
exit(1)
# Next apply Jacobian transformations to convert into the (xx0,xx1,xx2) basis
# rho and S are scalar, so no Jacobian transformations are necessary.
Jac4_dUSphorCart_dDrfmUD = ixp.zerorank2(DIM=4)
Jac4_dUSphorCart_dDrfmUD[0][0] = sp.sympify(1)
for i in range(DIM):
for j in range(DIM):
Jac4_dUSphorCart_dDrfmUD[i+1][j+1] = sp.diff(r_th_ph_or_Cart_xyz_oID_xx[i],rfm.xx[j])
Jac4_dUrfm_dDSphorCartUD, dummyDET = ixp.generic_matrix_inverter4x4(Jac4_dUSphorCart_dDrfmUD)
# Perform Jacobian operations on T^{mu nu} and gamma_{ij}
T4UU = ixp.register_gridfunctions_for_single_rank2("AUXEVOL","T4UU","sym01",DIM=4)
IDT4UU = ixp.zerorank2(DIM=4)
for mu in range(4):
for nu in range(4):
for delta in range(4):
for sigma in range(4):
IDT4UU[mu][nu] += \
Jac4_dUrfm_dDSphorCartUD[mu][delta]*Jac4_dUrfm_dDSphorCartUD[nu][sigma]*T4SphUU[delta][sigma]
lhrh_list = []
for mu in range(4):
for nu in range(mu,4):
lhrh_list.append(lhrh(lhs=gri.gfaccess("auxevol_gfs","T4UU"+str(mu)+str(nu)),rhs=IDT4UU[mu][nu]))
desc = """This function takes as input either (x,y,z) or (r,th,ph) and outputs
all ADM quantities in the Cartesian or Spherical basis, respectively."""
name = "ID_TOV_TUPMUNU_xx0xx1xx2"
outCparams = "preindent=1,outCverbose=False,includebraces=False"
outCfunction(
outfile=os.path.join(Ccodesdir, name + ".h"), desc=desc, name=name,
params="""const paramstruct *restrict params,REAL *restrict xx[3],
const ID_inputs other_inputs,REAL *restrict auxevol_gfs""",
body=outputC([rfm.xxSph[0],rfm.xxSph[1],rfm.xxSph[2]],
["const REAL rbar","const REAL theta","const REAL ph"],"returnstring",
"CSE_enable=False,includebraces=False")+"""
REAL rho,rho_baryon,P,M,expnu,exp4phi;
TOV_interpolate_1D(rbar,other_inputs.Rbar,other_inputs.Rbar_idx,other_inputs.interp_stencil_size,
other_inputs.numlines_in_file,
other_inputs.r_Schw_arr,other_inputs.rho_arr,other_inputs.rho_baryon_arr,other_inputs.P_arr,other_inputs.M_arr,
other_inputs.expnu_arr,other_inputs.exp4phi_arr,other_inputs.rbar_arr,
&rho,&rho_baryon,&P,&M,&expnu,&exp4phi);\n"""+
fin.FD_outputC("returnstring",lhrh_list,params="outCverbose=False,includebraces=False").replace("IDX4","IDX4S"),
loopopts="AllPoints,Read_xxs")
```
<a id='adm_id_spacetime'></a>
# Step 3: Convert ADM initial data to BSSN-in-curvilinear coordinates \[Back to [top](#toc)\]
$$\label{adm_id_spacetime}$$
This is an automated process, taken care of by [`BSSN.ADM_Numerical_Spherical_or_Cartesian_to_BSSNCurvilinear`](../edit/BSSN.ADM_Numerical_Spherical_or_Cartesian_to_BSSNCurvilinear.py), and documented [in this tutorial notebook](Tutorial-ADM_Initial_Data-Converting_Numerical_ADM_Spherical_or_Cartesian_to_BSSNCurvilinear.ipynb).
```
import BSSN.ADM_Numerical_Spherical_or_Cartesian_to_BSSNCurvilinear as AtoBnum
AtoBnum.Convert_Spherical_or_Cartesian_ADM_to_BSSN_curvilinear("Spherical","ID_TOV_ADM_quantities",
Ccodesdir=Ccodesdir,loopopts="")
```
<a id='validate'></a>
# Step 4: Validating that the TOV initial data satisfy the Hamiltonian constraint \[Back to [top](#toc)\]
$$\label{validate}$$
We will validate that the TOV initial data satisfy the Hamiltonian constraint, modulo numerical finite differencing error
<a id='ham_const_output'></a>
## Step 4.a: Output the Hamiltonian constraint \[Back to [top](#toc)\]
$$\label{ham_const_output}$$
First output the Hamiltonian constraint [as documented in the corresponding NRPy+ tutorial notebook](Tutorial-BSSN_constraints.ipynb)
```
# Enable rfm_precompute infrastructure, which results in
# BSSN RHSs that are free of transcendental functions,
# even in curvilinear coordinates, so long as
# ConformalFactor is set to "W" (default).
cmd.mkdir(os.path.join(Ccodesdir,"rfm_files/"))
par.set_parval_from_str("reference_metric::enable_rfm_precompute","True")
par.set_parval_from_str("reference_metric::rfm_precompute_Ccode_outdir",os.path.join(Ccodesdir,"rfm_files/"))
import BSSN.Enforce_Detgammabar_Constraint as EGC
enforce_detg_constraint_symb_expressions = EGC.Enforce_Detgammabar_Constraint_symb_expressions()
# Now register the Hamiltonian as a gridfunction.
H = gri.register_gridfunctions("AUX","H")
# Then define the Hamiltonian constraint and output the optimized C code.
import BSSN.BSSN_constraints as bssncon
import BSSN.BSSN_stress_energy_source_terms as Bsest
bssncon.BSSN_constraints(add_T4UUmunu_source_terms=False)
Bsest.BSSN_source_terms_for_BSSN_constraints(T4UU)
bssncon.H += Bsest.sourceterm_H
# Now that we are finished with all the rfm hatted
# quantities in generic precomputed functional
# form, let's restore them to their closed-
# form expressions.
par.set_parval_from_str("reference_metric::enable_rfm_precompute","False") # Reset to False to disable rfm_precompute.
rfm.ref_metric__hatted_quantities()
desc="Evaluate the Hamiltonian constraint"
name="Hamiltonian_constraint"
outCfunction(
outfile = os.path.join(Ccodesdir,name+".h"), desc=desc, name=name,
params = """rfm_struct *restrict rfmstruct,const paramstruct *restrict params,
REAL *restrict in_gfs, REAL *restrict auxevol_gfs, REAL *restrict aux_gfs""",
body = fin.FD_outputC("returnstring",lhrh(lhs=gri.gfaccess("aux_gfs", "H"), rhs=bssncon.H),
params="outCverbose=False").replace("IDX4","IDX4S"),
loopopts = "InteriorPoints,Enable_rfm_precompute")
```
<a id='bc_functs'></a>
## Step 4.b: Set up boundary condition functions for chosen singular, curvilinear coordinate system \[Back to [top](#toc)\]
$$\label{bc_functs}$$
Next apply singular, curvilinear coordinate boundary conditions [as documented in the corresponding NRPy+ tutorial notebook](Tutorial-Start_to_Finish-Curvilinear_BCs.ipynb)
```
import CurviBoundaryConditions.CurviBoundaryConditions as cbcs
cbcs.Set_up_CurviBoundaryConditions(os.path.join(Ccodesdir,"boundary_conditions/"),Cparamspath=os.path.join("../"))
```
<a id='enforce3metric'></a>
## Step 4.c: Enforce conformal 3-metric $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$ constraint \[Back to [top](#toc)\]
$$\label{enforce3metric}$$
Then enforce conformal 3-metric $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$ constraint (Eq. 53 of [Ruchlin, Etienne, and Baumgarte (2018)](https://arxiv.org/abs/1712.07658)), as [documented in the corresponding NRPy+ tutorial notebook](Tutorial-BSSN-Enforcing_Determinant_gammabar_equals_gammahat_Constraint.ipynb)
Applying curvilinear boundary conditions should affect the initial data at the outer boundary, and will in general cause the $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$ constraint to be violated there. Thus after we apply these boundary conditions, we must always call the routine for enforcing the $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$ constraint:
```
# Set up the C function for the det(gammahat) = det(gammabar)
EGC.output_Enforce_Detgammabar_Constraint_Ccode(Ccodesdir,
exprs=enforce_detg_constraint_symb_expressions)
```
<a id='cparams_rfm_and_domainsize'></a>
## Step 4.d: Output C codes needed for declaring and setting Cparameters; also set `free_parameters.h` \[Back to [top](#toc)\]
$$\label{cparams_rfm_and_domainsize}$$
Based on declared NRPy+ Cparameters, first we generate `declare_Cparameters_struct.h`, `set_Cparameters_default.h`, and `set_Cparameters[-SIMD].h`.
Then we output `free_parameters.h`, which sets initial data parameters, as well as grid domain & reference metric parameters, applying `domain_size` and `sinh_width`/`SymTP_bScale` (if applicable) as set above
```
# Step 3.d.i: Generate declare_Cparameters_struct.h, set_Cparameters_default.h, and set_Cparameters[-SIMD].h
par.generate_Cparameters_Ccodes(os.path.join(Ccodesdir))
# Step 3.d.ii: Set free_parameters.h
# Output to $Ccodesdir/free_parameters.h reference metric parameters based on generic
# domain_size,sinh_width,sinhv2_const_dr,SymTP_bScale,
# parameters set above.
rfm.out_default_free_parameters_for_rfm(os.path.join(Ccodesdir,"free_parameters.h"),
domain_size,sinh_width,sinhv2_const_dr,SymTP_bScale)
# Step 3.d.iii: Generate set_Nxx_dxx_invdx_params__and__xx.h:
rfm.set_Nxx_dxx_invdx_params__and__xx_h(Ccodesdir)
# Step 3.d.iv: Generate xxCart.h, which contains xxCart() for
# (the mapping from xx->Cartesian) for the chosen
# CoordSystem:
rfm.xxCart_h("xxCart","./set_Cparameters.h",os.path.join(Ccodesdir,"xxCart.h"))
# Step 3.d.v: Generate declare_Cparameters_struct.h, set_Cparameters_default.h, and set_Cparameters[-SIMD].h
par.generate_Cparameters_Ccodes(os.path.join(Ccodesdir))
```
<a id='mainc'></a>
# Step 5: `TOV_Playground.c`: The Main C Code \[Back to [top](#toc)\]
$$\label{mainc}$$
```
# Part P0: Define REAL, set the number of ghost cells NGHOSTS (from NRPy+'s FD_CENTDERIVS_ORDER)
with open(os.path.join(Ccodesdir,"TOV_Playground_REAL__NGHOSTS.h"), "w") as file:
file.write("""
// Part P0.a: Set the number of ghost cells, from NRPy+'s FD_CENTDERIVS_ORDER
#define NGHOSTS """+str(int(FD_order/2)+1)+"""
// Part P0.b: Set the numerical precision (REAL) to double, ensuring all floating point
// numbers are stored to at least ~16 significant digits
#define REAL """+REAL+"""
// Part P0.c: Set TOV stellar parameters
#define TOV_Mass """+str(M_TOV)+"""
#define TOV_Riso """+str(R_iso_TOV)+"\n")
%%writefile $Ccodesdir/TOV_Playground.c
// Step P0: Define REAL and NGHOSTS. This header is generated by NRPy+.
#include "TOV_Playground_REAL__NGHOSTS.h"
#include "rfm_files/rfm_struct__declare.h"
#include "declare_Cparameters_struct.h"
// Step P1: Import needed header files
#include "stdio.h"
#include "stdlib.h"
#include "math.h"
#ifndef M_PI
#define M_PI 3.141592653589793238462643383279502884L
#endif
#ifndef M_SQRT1_2
#define M_SQRT1_2 0.707106781186547524400844362104849039L
#endif
// Step P2: Declare the IDX4S(gf,i,j,k) macro, which enables us to store 4-dimensions of
// data in a 1D array. In this case, consecutive values of "i"
// (all other indices held to a fixed value) are consecutive in memory, where
// consecutive values of "j" (fixing all other indices) are separated by
// Nxx_plus_2NGHOSTS0 elements in memory. Similarly, consecutive values of
// "k" are separated by Nxx_plus_2NGHOSTS0*Nxx_plus_2NGHOSTS1 in memory, etc.
#define IDX4S(g,i,j,k) \
( (i) + Nxx_plus_2NGHOSTS0 * ( (j) + Nxx_plus_2NGHOSTS1 * ( (k) + Nxx_plus_2NGHOSTS2 * (g) ) ) )
#define IDX4ptS(g,idx) ( (idx) + (Nxx_plus_2NGHOSTS0*Nxx_plus_2NGHOSTS1*Nxx_plus_2NGHOSTS2) * (g) )
#define IDX3S(i,j,k) ( (i) + Nxx_plus_2NGHOSTS0 * ( (j) + Nxx_plus_2NGHOSTS1 * ( (k) ) ) )
#define LOOP_REGION(i0min,i0max, i1min,i1max, i2min,i2max) \
for(int i2=i2min;i2<i2max;i2++) for(int i1=i1min;i1<i1max;i1++) for(int i0=i0min;i0<i0max;i0++)
#define LOOP_ALL_GFS_GPS(ii) _Pragma("omp parallel for") \
for(int (ii)=0;(ii)<Nxx_plus_2NGHOSTS_tot*NUM_EVOL_GFS;(ii)++)
// Step P3: Set UUGF and VVGF macros, as well as xxCart()
#include "boundary_conditions/gridfunction_defines.h"
// Step P4: Set xxCart(const paramstruct *restrict params,
// REAL *restrict xx[3],
// const int i0,const int i1,const int i2,
// REAL xCart[3]),
// which maps xx->Cartesian via
// {xx[0][i0],xx[1][i1],xx[2][i2]}->{xCart[0],xCart[1],xCart[2]}
#include "xxCart.h"
// Step P5: Defines set_Nxx_dxx_invdx_params__and__xx(const int EigenCoord, const int Nxx[3],
// paramstruct *restrict params, REAL *restrict xx[3]),
// which sets params Nxx,Nxx_plus_2NGHOSTS,dxx,invdx, and xx[] for
// the chosen Eigen-CoordSystem if EigenCoord==1, or
// CoordSystem if EigenCoord==0.
#include "set_Nxx_dxx_invdx_params__and__xx.h"
// Step P6: Include basic functions needed to impose curvilinear
// parity and boundary conditions.
#include "boundary_conditions/CurviBC_include_Cfunctions.h"
// Step P8: Include function for enforcing detgammabar constraint.
#include "enforce_detgammabar_constraint.h"
// Step P4: Declare initial data input struct:
// stores data from initial data solver,
// so they can be put on the numerical grid.
typedef struct __ID_inputs {
REAL Rbar;
int Rbar_idx;
int interp_stencil_size;
int numlines_in_file;
REAL *r_Schw_arr,*rho_arr,*rho_baryon_arr,*P_arr,*M_arr,*expnu_arr,*exp4phi_arr,*rbar_arr;
} ID_inputs;
// Part P11: Declare all functions for setting up TOV initial data.
/* Routines to interpolate the TOV solution and convert to ADM & T^{munu}: */
#include "../TOV/tov_interp.h"
#include "ID_TOV_ADM_quantities.h"
#include "ID_TOV_TUPMUNU_xx0xx1xx2.h"
/* Next perform the basis conversion and compute all needed BSSN quantities */
#include "ID_ADM_xx0xx1xx2_to_BSSN_xx0xx1xx2__ALL_BUT_LAMBDAs.h"
#include "ID_BSSN__ALL_BUT_LAMBDAs.h"
#include "ID_BSSN_lambdas.h"
// Step P10: Declare function necessary for setting up the initial data.
// Step P10.a: Define BSSN_ID() for BrillLindquist initial data
// Step P10.b: Set the generic driver function for setting up BSSN initial data
void initial_data(const paramstruct *restrict params,const bc_struct *restrict bcstruct,
const rfm_struct *restrict rfmstruct,
REAL *restrict xx[3], REAL *restrict auxevol_gfs, REAL *restrict in_gfs) {
#include "set_Cparameters.h"
// Step 1: Set up TOV initial data
// Step 1.a: Read TOV initial data from data file
// Open the data file:
char filename[100];
sprintf(filename,"./outputTOVpolytrope.txt");
FILE *in1Dpolytrope = fopen(filename, "r");
if (in1Dpolytrope == NULL) {
fprintf(stderr,"ERROR: could not open file %s\n",filename);
exit(1);
}
// Count the number of lines in the data file:
int numlines_in_file = count_num_lines_in_file(in1Dpolytrope);
// Allocate space for all data arrays:
REAL *r_Schw_arr = (REAL *)malloc(sizeof(REAL)*numlines_in_file);
REAL *rho_arr = (REAL *)malloc(sizeof(REAL)*numlines_in_file);
REAL *rho_baryon_arr = (REAL *)malloc(sizeof(REAL)*numlines_in_file);
REAL *P_arr = (REAL *)malloc(sizeof(REAL)*numlines_in_file);
REAL *M_arr = (REAL *)malloc(sizeof(REAL)*numlines_in_file);
REAL *expnu_arr = (REAL *)malloc(sizeof(REAL)*numlines_in_file);
REAL *exp4phi_arr = (REAL *)malloc(sizeof(REAL)*numlines_in_file);
REAL *rbar_arr = (REAL *)malloc(sizeof(REAL)*numlines_in_file);
// Read from the data file, filling in arrays
// read_datafile__set_arrays() may be found in TOV/tov_interp.h
if(read_datafile__set_arrays(in1Dpolytrope, r_Schw_arr,rho_arr,rho_baryon_arr,P_arr,M_arr,expnu_arr,exp4phi_arr,rbar_arr) == 1) {
fprintf(stderr,"ERROR WHEN READING FILE %s!\n",filename);
exit(1);
}
fclose(in1Dpolytrope);
REAL Rbar = -100;
int Rbar_idx = -100;
for(int i=1;i<numlines_in_file;i++) {
if(rho_arr[i-1]>0 && rho_arr[i]==0) { Rbar = rbar_arr[i-1]; Rbar_idx = i-1; }
}
if(Rbar<0) {
fprintf(stderr,"Error: could not find rbar=Rbar from data file.\n");
exit(1);
}
ID_inputs TOV_in;
TOV_in.Rbar = Rbar;
TOV_in.Rbar_idx = Rbar_idx;
const int interp_stencil_size = 12;
TOV_in.interp_stencil_size = interp_stencil_size;
TOV_in.numlines_in_file = numlines_in_file;
TOV_in.r_Schw_arr = r_Schw_arr;
TOV_in.rho_arr = rho_arr;
TOV_in.rho_baryon_arr = rho_baryon_arr;
TOV_in.P_arr = P_arr;
TOV_in.M_arr = M_arr;
TOV_in.expnu_arr = expnu_arr;
TOV_in.exp4phi_arr = exp4phi_arr;
TOV_in.rbar_arr = rbar_arr;
/* END TOV INPUT ROUTINE */
// Step 1.b: Interpolate data from data file to set BSSN gridfunctions
ID_BSSN__ALL_BUT_LAMBDAs(params,xx,TOV_in, in_gfs);
apply_bcs_curvilinear(params, bcstruct, NUM_EVOL_GFS, evol_gf_parity, in_gfs);
enforce_detgammabar_constraint(rfmstruct, params, in_gfs);
ID_BSSN_lambdas(params, xx, in_gfs);
apply_bcs_curvilinear(params, bcstruct, NUM_EVOL_GFS, evol_gf_parity, in_gfs);
enforce_detgammabar_constraint(rfmstruct, params, in_gfs);
ID_TOV_TUPMUNU_xx0xx1xx2(params,xx,TOV_in,auxevol_gfs);
free(rbar_arr);
free(rho_arr);
free(rho_baryon_arr);
free(P_arr);
free(M_arr);
free(expnu_arr);
}
// Step P11: Declare function for evaluating Hamiltonian constraint (diagnostic)
#include "Hamiltonian_constraint.h"
// main() function:
// Step 0: Read command-line input, set up grid structure, allocate memory for gridfunctions, set up coordinates
// Step 1: Set up initial data to an exact solution
// Step 2: Start the timer, for keeping track of how fast the simulation is progressing.
// Step 3: Integrate the initial data forward in time using the chosen RK-like Method of
// Lines timestepping algorithm, and output periodic simulation diagnostics
// Step 3.a: Output 2D data file periodically, for visualization
// Step 3.b: Step forward one timestep (t -> t+dt) in time using
// chosen RK-like MoL timestepping algorithm
// Step 3.c: If t=t_final, output conformal factor & Hamiltonian
// constraint violation to 2D data file
// Step 3.d: Progress indicator printing to stderr
// Step 4: Free all allocated memory
int main(int argc, const char *argv[]) {
paramstruct params;
#include "set_Cparameters_default.h"
// Step 0a: Read command-line input, error out if nonconformant
if((argc != 4) || atoi(argv[1]) < NGHOSTS || atoi(argv[2]) < NGHOSTS || atoi(argv[3]) < 2 /* FIXME; allow for axisymmetric sims */) {
fprintf(stderr,"Error: Expected three command-line arguments: ./BrillLindquist_Playground Nx0 Nx1 Nx2,\n");
fprintf(stderr,"where Nx[0,1,2] is the number of grid points in the 0, 1, and 2 directions.\n");
fprintf(stderr,"Nx[] MUST BE larger than NGHOSTS (= %d)\n",NGHOSTS);
exit(1);
}
// Step 0b: Set up numerical grid structure, first in space...
const int Nxx[3] = { atoi(argv[1]), atoi(argv[2]), atoi(argv[3]) };
if(Nxx[0]%2 != 0 || Nxx[1]%2 != 0 || Nxx[2]%2 != 0) {
fprintf(stderr,"Error: Cannot guarantee a proper cell-centered grid if number of grid cells not set to even number.\n");
fprintf(stderr," For example, in case of angular directions, proper symmetry zones will not exist.\n");
exit(1);
}
// Step 0c: Set free parameters, overwriting Cparameters defaults
// by hand or with command-line input, as desired.
#include "free_parameters.h"
// Step 0d: Uniform coordinate grids are stored to *xx[3]
REAL *xx[3];
// Step 0d.i: Set bcstruct
bc_struct bcstruct;
{
int EigenCoord = 1;
// Step 0d.ii: Call set_Nxx_dxx_invdx_params__and__xx(), which sets
// params Nxx,Nxx_plus_2NGHOSTS,dxx,invdx, and xx[] for the
// chosen Eigen-CoordSystem.
set_Nxx_dxx_invdx_params__and__xx(EigenCoord, Nxx, ¶ms, xx);
// Step 0d.iii: Set Nxx_plus_2NGHOSTS_tot
#include "set_Cparameters-nopointer.h"
const int Nxx_plus_2NGHOSTS_tot = Nxx_plus_2NGHOSTS0*Nxx_plus_2NGHOSTS1*Nxx_plus_2NGHOSTS2;
// Step 0e: Find ghostzone mappings; set up bcstruct
#include "boundary_conditions/driver_bcstruct.h"
// Step 0e.i: Free allocated space for xx[][] array
for(int i=0;i<3;i++) free(xx[i]);
}
// Step 0f: Call set_Nxx_dxx_invdx_params__and__xx(), which sets
// params Nxx,Nxx_plus_2NGHOSTS,dxx,invdx, and xx[] for the
// chosen (non-Eigen) CoordSystem.
int EigenCoord = 0;
set_Nxx_dxx_invdx_params__and__xx(EigenCoord, Nxx, ¶ms, xx);
// Step 0g: Set all C parameters "blah" for params.blah, including
// Nxx_plus_2NGHOSTS0 = params.Nxx_plus_2NGHOSTS0, etc.
#include "set_Cparameters-nopointer.h"
const int Nxx_plus_2NGHOSTS_tot = Nxx_plus_2NGHOSTS0*Nxx_plus_2NGHOSTS1*Nxx_plus_2NGHOSTS2;
// Step 0j: Error out if the number of auxiliary gridfunctions outnumber evolved gridfunctions.
// This is a limitation of the RK method. You are always welcome to declare & allocate
// additional gridfunctions by hand.
if(NUM_AUX_GFS > NUM_EVOL_GFS) {
fprintf(stderr,"Error: NUM_AUX_GFS > NUM_EVOL_GFS. Either reduce the number of auxiliary gridfunctions,\n");
fprintf(stderr," or allocate (malloc) by hand storage for *diagnostic_output_gfs. \n");
exit(1);
}
// Step 0k: Allocate memory for gridfunctions
#include "MoLtimestepping/RK_Allocate_Memory.h"
REAL *restrict auxevol_gfs = (REAL *)malloc(sizeof(REAL) * NUM_AUXEVOL_GFS * Nxx_plus_2NGHOSTS_tot);
// Step 0l: Set up precomputed reference metric arrays
// Step 0l.i: Allocate space for precomputed reference metric arrays.
#include "rfm_files/rfm_struct__malloc.h"
// Step 0l.ii: Define precomputed reference metric arrays.
{
#include "set_Cparameters-nopointer.h"
#include "rfm_files/rfm_struct__define.h"
}
// Step 1: Set up initial data to an exact solution
initial_data(¶ms,&bcstruct, &rfmstruct, xx, auxevol_gfs, y_n_gfs);
// Step 1b: Apply boundary conditions, as initial data
// are sometimes ill-defined in ghost zones.
// E.g., spherical initial data might not be
// properly defined at points where r=-1.
apply_bcs_curvilinear(¶ms, &bcstruct, NUM_EVOL_GFS,evol_gf_parity, y_n_gfs);
enforce_detgammabar_constraint(&rfmstruct, ¶ms, y_n_gfs);
// Evaluate Hamiltonian constraint violation
Hamiltonian_constraint(&rfmstruct, ¶ms, y_n_gfs,auxevol_gfs, diagnostic_output_gfs);
char filename[100];
sprintf(filename,"out%d.txt",Nxx[0]);
FILE *out2D = fopen(filename, "w");
LOOP_REGION(NGHOSTS,Nxx_plus_2NGHOSTS0-NGHOSTS,
NGHOSTS,Nxx_plus_2NGHOSTS1-NGHOSTS,
NGHOSTS,Nxx_plus_2NGHOSTS2-NGHOSTS) {
REAL xx0 = xx[0][i0];
REAL xx1 = xx[1][i1];
REAL xx2 = xx[2][i2];
REAL xCart[3];
xxCart(¶ms,xx,i0,i1,i2,xCart);
int idx = IDX3S(i0,i1,i2);
fprintf(out2D,"%e %e %e %e\n",xCart[1]/TOV_Mass,xCart[2]/TOV_Mass, y_n_gfs[IDX4ptS(CFGF,idx)],
log10(fabs(diagnostic_output_gfs[IDX4ptS(HGF,idx)])));
}
fclose(out2D);
// Step 4: Free all allocated memory
#include "rfm_files/rfm_struct__freemem.h"
#include "boundary_conditions/bcstruct_freemem.h"
#include "MoLtimestepping/RK_Free_Memory.h"
free(auxevol_gfs);
for(int i=0;i<3;i++) free(xx[i]);
return 0;
}
import cmdline_helper as cmd
cmd.C_compile(os.path.join(Ccodesdir,"TOV_Playground.c"), "TOV_Playground")
cmd.delete_existing_files("out96.txt")
cmd.Execute("TOV_Playground", "96 96 2", "out96.txt")
```
<a id='plot'></a>
# Step 6: Plotting the single-neutron-star initial data \[Back to [top](#toc)\]
$$\label{plot}$$
Here we plot the conformal factor of these initial data on a 2D grid, such that darker colors imply stronger gravitational fields. Hence, we see the single neutron star centered at the origin: $x/M=y/M=z/M=0$, where $M$ is an arbitrary mass scale (conventionally the [ADM mass](https://en.wikipedia.org/w/index.php?title=ADM_formalism&oldid=846335453) is chosen), and our formulation of Einstein's equations adopt $G=c=1$ [geometrized units](https://en.wikipedia.org/w/index.php?title=Geometrized_unit_system&oldid=861682626).
```
import numpy as np
from scipy.interpolate import griddata
from pylab import savefig
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from IPython.display import Image
x96,y96,valuesCF96,valuesHam96 = np.loadtxt('out96.txt').T #Transposed for easier unpacking
bounds = 7.5
pl_xmin = -bounds
pl_xmax = +bounds
pl_ymin = -bounds
pl_ymax = +bounds
grid_x, grid_y = np.mgrid[pl_xmin:pl_xmax:100j, pl_ymin:pl_ymax:100j]
points96 = np.zeros((len(x96), 2))
for i in range(len(x96)):
points96[i][0] = x96[i]
points96[i][1] = y96[i]
grid96 = griddata(points96, valuesCF96, (grid_x, grid_y), method='nearest')
grid96cub = griddata(points96, valuesCF96, (grid_x, grid_y), method='cubic')
plt.clf()
plt.title("Neutron Star: log10( max(1e-6,Energy Density) )")
plt.xlabel("x/M")
plt.ylabel("y/M")
# fig, ax = plt.subplots()
# ax.plot(grid96cub.T, extent=(pl_xmin,pl_xmax, pl_ymin,pl_ymax))
# plt.close(fig)
fig96cf = plt.imshow(grid96.T, extent=(pl_xmin,pl_xmax, pl_ymin,pl_ymax))
cb = plt.colorbar(fig96cf)
savefig("BHB.png")
from IPython.display import Image
Image("BHB.png")
# # interpolation='nearest', cmap=cm.gist_rainbow)
```
<a id='convergence'></a>
# Step 7: Validation: Convergence of numerical errors (Hamiltonian constraint violation) to zero \[Back to [top](#toc)\]
$$\label{convergence}$$
The equations behind these initial data solve Einstein's equations exactly, at a single instant in time. One reflection of this solution is that the Hamiltonian constraint violation should be exactly zero in the initial data.
However, when evaluated on numerical grids, the Hamiltonian constraint violation will *not* generally evaluate to zero due to the associated numerical derivatives not being exact. However, these numerical derivatives (finite difference derivatives in this case) should *converge* to the exact derivatives as the density of numerical sampling points approaches infinity.
In this case, all of our finite difference derivatives agree with the exact solution, with an error term that drops with the uniform gridspacing to the fourth power: $\left(\Delta x^i\right)^4$.
Here, as in the [Start-to-Finish Scalar Wave (Cartesian grids) NRPy+ tutorial](Tutorial-Start_to_Finish-ScalarWave.ipynb) and the [Start-to-Finish Scalar Wave (curvilinear grids) NRPy+ tutorial](Tutorial-Start_to_Finish-ScalarWaveCurvilinear.ipynb) we confirm this convergence.
First, let's take a look at what the numerical error looks like on the x-y plane at a given numerical resolution, plotting $\log_{10}|H|$, where $H$ is the Hamiltonian constraint violation:
```
grid96 = griddata(points96, valuesHam96, (grid_x, grid_y), method='nearest')
grid96cub = griddata(points96, valuesHam96, (grid_x, grid_y), method='cubic')
# fig, ax = plt.subplots()
plt.clf()
plt.title("96^3 Numerical Err.: log_{10}|Ham|")
plt.xlabel("x/M")
plt.ylabel("y/M")
fig96cub = plt.imshow(grid96cub.T, extent=(pl_xmin,pl_xmax, pl_ymin,pl_ymax))
cb = plt.colorbar(fig96cub)
```
Next, we set up the same initial data but on a lower-resolution, $48\times 8\times 2$ grid (axisymmetric in the $\phi$ direction). Since the constraint violation (numerical error associated with the fourth-order-accurate, finite-difference derivatives) should converge to zero with the uniform gridspacing to the fourth power: $\left(\Delta x^i\right)^4$, we expect the constraint violation will increase (relative to the $96\times 16\times 2$ grid) by a factor of $\left(96/48\right)^4$. Here we demonstrate that indeed this order of convergence is observed as expected, *except* at the star's surface where the stress-energy tensor $T^{\mu\nu}$ sharply drops to zero.
```
# Now rerun TOV_Playground with twice lower resolution.
cmd.delete_existing_files("out48.txt")
cmd.Execute("TOV_Playground", "48 48 2", "out48.txt")
x48,y48,valuesCF48,valuesHam48 = np.loadtxt('out48.txt').T #Transposed for easier unpacking
points48 = np.zeros((len(x48), 2))
for i in range(len(x48)):
points48[i][0] = x48[i]
points48[i][1] = y48[i]
grid48 = griddata(points48, valuesHam48, (grid_x, grid_y), method='cubic')
griddiff_48_minus_96 = np.zeros((100,100))
griddiff_48_minus_96_1darray = np.zeros(100*100)
gridx_1darray_yeq0 = np.zeros(100)
grid48_1darray_yeq0 = np.zeros(100)
grid96_1darray_yeq0 = np.zeros(100)
count = 0
outarray = []
for i in range(100):
for j in range(100):
griddiff_48_minus_96[i][j] = grid48[i][j] - grid96[i][j]
griddiff_48_minus_96_1darray[count] = griddiff_48_minus_96[i][j]
if j==49:
gridx_1darray_yeq0[i] = grid_x[i][j]
grid48_1darray_yeq0[i] = grid48[i][j] + np.log10((48./96.)**4)
grid96_1darray_yeq0[i] = grid96[i][j]
count = count + 1
plt.clf()
fig, ax = plt.subplots()
plt.title("Plot Demonstrating 4th-order Convergence")
plt.xlabel("x/M")
plt.ylabel("log10(Relative error)")
ax.plot(gridx_1darray_yeq0, grid96_1darray_yeq0, 'k-', label='Nr=96')
ax.plot(gridx_1darray_yeq0, grid48_1darray_yeq0, 'k--', label='Nr=48, mult by (48/96)^4')
ax.set_ylim([-12.5,1.5])
legend = ax.legend(loc='lower right', shadow=True, fontsize='x-large')
legend.get_frame().set_facecolor('C1')
plt.show()
```
<a id='latex_pdf_output'></a>
# Step 8: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
$$\label{latex_pdf_output}$$
The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
[Tutorial-Start_to_Finish-BSSNCurvilinear-Setting_up_TOV_initial_data.pdf](Tutorial-Start_to_Finish-BSSNCurvilinear-Setting_up_TOV_initial_data.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
```
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-Start_to_Finish-BSSNCurvilinear-Setting_up_TOV_initial_data")
```
| github_jupyter |
# Custom Display Logic
## Overview
As described in the [Rich Output](Rich Output.ipynb) tutorial, the IPython display system can display rich representations of objects in the following formats:
* JavaScript
* HTML
* PNG
* JPEG
* SVG
* LaTeX
* PDF
* Markdown
This Notebook shows how you can add custom display logic to your own classes, so that they can be displayed using these rich representations. There are two ways of accomplishing this:
1. Implementing special display methods such as `_repr_html_` when you define your class.
2. Registering a display function for a particular existing class.
This Notebook describes and illustrates both approaches.
Import the IPython display functions.
```
from IPython.display import (
display, display_html, display_png, display_svg
)
```
Parts of this notebook need the matplotlib inline backend:
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
```
## Background: classes
Classes let you define new types of objects to use in your code. Most of the code in a larger Python application, like Jupyter, is typically in classes.
Here's how you define a class:
```
import random
class DiceSet:
# The special name __init__ makes a 'constructor', which sets up new
# instances of the class. Instances of this class store two pieces of data.
def __init__(self, n, sides=6):
self.n = n
self.sides = sides
# Functions on classes are called 'methods'. The first argument is an instance
# of the class.
def roll(self):
r = []
for i in range(self.n):
r.append(random.randint(1, self.sides))
return r
```
And here's how to use our new class:
```
monopoly_dice = DiceSet(2) # two six-sided dice
# monopoly_dice is an instance of DiceSet
monopoly_dice.roll()
strange_dice = DiceSet(n=5, sides=11) # another instance of the same class
strange_dice.roll()
```
## Special display methods
The main idea of the first approach is that you have to implement special display methods when you define your class, one for each representation you want to use. Here is a list of the names of the special methods and the values they must return:
* `_repr_html_`: return raw HTML as a string
* `_repr_json_`: return a JSONable dict
* `_repr_jpeg_`: return raw JPEG data
* `_repr_png_`: return raw PNG data
* `_repr_svg_`: return raw SVG data as a string
* `_repr_latex_`: return LaTeX commands in a string surrounded by "`$`".
As an illustration, we build a class that holds data generated by sampling a Gaussian distribution with given mean and standard deviation. Here is the definition of the `Gaussian` class, which has a custom PNG and LaTeX representation, in addition to a standard `__repr__` representation.
```
from IPython.core.pylabtools import print_figure
from IPython.display import Image, SVG, Math
class Gaussian(object):
"""A simple object holding data sampled from a Gaussian distribution.
"""
def __init__(self, mean=0.0, std=1, size=1000):
self.data = np.random.normal(mean, std, size)
self.mean = mean
self.std = std
self.size = size
# For caching plots that may be expensive to compute
self._png_data = None
def __repr__(self):
return "A Gaussian process, mean %.2g, std %.2g, N %d" % (self.mean,
self.std,
self.size)
def _figure_data(self, format):
fig, ax = plt.subplots()
ax.hist(self.data, bins=50)
ax.set_title(self._repr_latex_())
ax.set_xlim(-10.0,10.0)
data = print_figure(fig, format)
# We MUST close the figure, otherwise IPython's display machinery
# will pick it up and send it as output, resulting in a double display
plt.close(fig)
return data
def _repr_png_(self):
if self._png_data is None:
self._png_data = self._figure_data('png')
return self._png_data
def _repr_latex_(self):
return r'$\mathcal{N}(\mu=%.2g, \sigma=%.2g),\ N=%d$' % (self.mean,
self.std, self.size)
```
Create an instance of the Gaussian distribution, `print` its standard representation, and return it to display the default representation:
```
x = Gaussian(2.0, 1.0)
print(x)
x
```
You can also pass the object to the `display` function to display the default representation:
```
display(x)
```
Use `display_png` to view the PNG representation:
```
display_png(x)
```
<div class="alert alert-success">
It is important to note a subtle different between <code>display</code> and <code>display_png</code>. The former computes <em>all</em> representations of the object, and lets the notebook UI decide which to display. The later only computes the PNG representation.
</div>
Create a new Gaussian with different parameters:
```
x2 = Gaussian(0, 2, 2000)
x2
```
You can then compare the two Gaussians by displaying their histograms:
```
display_png(x)
display_png(x2)
```
Note that like `print`, you can call any of the `display` functions multiple times in a cell.
## Adding IPython display support to existing objects
When you are directly writing your own classes, you can adapt them for display in IPython by following the above approach. But in practice, you often need to work with existing classes that you can't easily modify. We now illustrate how to add rich output capabilities to existing objects. We will use the NumPy polynomials and change their default representation to be a formatted LaTeX expression.
First, consider how a NumPy polynomial object renders by default:
```
p = np.polynomial.Polynomial([1,2,3], [-10, 10])
p
```
Next, define a function that pretty-prints a polynomial as a LaTeX string:
```
def poly_to_latex(p):
terms = ['%.2g' % p.coef[0]]
if len(p) > 1:
term = 'x'
c = p.coef[1]
if c!=1:
term = ('%.2g ' % c) + term
terms.append(term)
if len(p) > 2:
for i in range(2, len(p)):
term = 'x^%d' % i
c = p.coef[i]
if c!=1:
term = ('%.2g ' % c) + term
terms.append(term)
px = '$P(x)=%s$' % '+'.join(terms)
dom = r', $x \in [%.2g,\ %.2g]$' % tuple(p.domain)
return px+dom
```
This produces, on our polynomial ``p``, the following:
```
poly_to_latex(p)
```
You can render this string using the `Latex` class:
```
from IPython.display import Latex
Latex(poly_to_latex(p))
```
However, you can configure IPython to do this automatically by registering the `Polynomial` class and the `plot_to_latex` function with an IPython display formatter. Let's look at the default formatters provided by IPython:
```
ip = get_ipython()
for mime, formatter in ip.display_formatter.formatters.items():
print('%24s : %s' % (mime, formatter.__class__.__name__))
```
The `formatters` attribute is a dictionary keyed by MIME types. To define a custom LaTeX display function, you want a handle on the `text/latex` formatter:
```
ip = get_ipython()
latex_f = ip.display_formatter.formatters['text/latex']
```
The formatter object has a couple of methods for registering custom display functions for existing types.
```
help(latex_f.for_type)
help(latex_f.for_type_by_name)
```
In this case, we will use `for_type_by_name` to register `poly_to_latex` as the display function for the `Polynomial` type:
```
latex_f.for_type_by_name('numpy.polynomial.polynomial',
'Polynomial', poly_to_latex)
```
Once the custom display function has been registered, all NumPy `Polynomial` instances will be represented by their LaTeX form instead:
```
p
p2 = np.polynomial.Polynomial([-20, 71, -15, 1])
p2
```
## More complex display with `_ipython_display_`
Rich output special methods and functions can only display one object or MIME type at a time. Sometimes this is not enough if you want to display multiple objects or MIME types at once. An example of this would be to use an HTML representation to put some HTML elements in the DOM and then use a JavaScript representation to add events to those elements.
**IPython** recognizes another display method, `_ipython_display_`, which allows your objects to take complete control of displaying themselves. If this method is defined, IPython will call it, and make no effort to display the object using the above described `_repr_*_` methods for custom display functions. It's a way for you to say "Back off, IPython, I can display this myself." Most importantly, your `_ipython_display_` method can make multiple calls to the top-level `display` functions to accomplish its goals.
Here is an object that uses `display_html` and `display_javascript` to make a plot using the [Flot](http://www.flotcharts.org/) JavaScript plotting library:
```
import json
import uuid
from IPython.display import display_javascript, display_html, display
class FlotPlot(object):
def __init__(self, x, y):
self.x = x
self.y = y
self.uuid = str(uuid.uuid4())
def _ipython_display_(self):
json_data = json.dumps(list(zip(self.x, self.y)))
display_html('<div id="{}" style="height: 300px; width:80%;"></div>'.format(self.uuid),
raw=True
)
display_javascript("""
require(["https://cdnjs.cloudflare.com/ajax/libs/flot/0.8.2/jquery.flot.min.js"], function() {
var line = JSON.parse("%s");
console.log(line);
$.plot("#%s", [line]);
});
""" % (json_data, self.uuid), raw=True)
import numpy as np
x = np.linspace(0,10)
y = np.sin(x)
FlotPlot(x, np.sin(x))
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.