text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Assigment 1
# Part One: Network Models
## 1. Watts-Strogatz Networks
* Use `nx.watts_strogatz_graph` to generate 3 graphs with 500 nodes each, average degree = 4, and rewiring probablity $p = 0, 0.1, \textrm{and} 1$. Calculate the average shortest path length $\langle d \rangle$ for each one. Describe what happens... | github_jupyter |
# Univariate time series classification with sktime
In this notebook, we will use sktime for univariate time series classification. Here, we have a single time series variable and an associated label for multiple instances. The goal is to find a classifier that can learn the relationship between time series and label ... | github_jupyter |
# End-to-End Example #1
1. [Introduction](#Introduction)
2. [Prerequisites and Preprocessing](#Prequisites-and-Preprocessing)
1. [Permissions and environment variables](#Permissions-and-environment-variables)
2. [Data ingestion](#Data-ingestion)
3. [Data inspection](#Data-inspection)
4. [Data conversion](#Data... | github_jupyter |
# Keras
Keras is fairly well-known in the Python deep learning community. It used to be a high-level API to make frameworks like CNTK, Theano and TensorFlow easier to use and was framework-agnostic (you only had to set the backend for processing, everything else was abstracted). A few years ago, Keras was migrated to t... | github_jupyter |
# Project 1: Trading with Momentum
## Instructions
Each problem consists of a function to implement and instructions on how to implement the function. The parts of the function that need to be implemented are marked with a `# TODO` comment. After implementing the function, run the cell to test it against the unit test... | github_jupyter |
# Introduction
## Guided Project - Visualizing The Gender Gap In College Degrees
In this guided project, we'll extend the work we did in the last two missions on visualizing the gender gap across college degrees. So far, we mostly focused on the STEM degrees but now we will generate line charts to compare across all... | github_jupyter |
```
import os
import sys
import networkx as nx
import pandas as pd
import community as community_louvain
import networkx.algorithms.community as nx_comm
nb_dir = os.path.split(os.getcwd())[0]
if nb_dir not in sys.path:
sys.path.append(nb_dir)
os.chdir('/home/tduricic/Development/workspace/structure-in-gnn')
from sr... | github_jupyter |
```
activations = [nn.ELU(),nn.LeakyReLU(),nn.PReLU(),nn.ReLU(),nn.ReLU6(),nn.RReLU(),nn.SELU(),nn.CELU(),nn.GELU(),nn.SiLU(),nn.Tanh()]
for activation in activations:
model = Test_Model(activation=activation)
optimizer = torch.optim.SGD(model.parameters(),lr=0.1)
criterion = nn.CrossEntropyLoss()
index... | github_jupyter |
# Rendezvous
Rendezvous problems involve the relative position, velocity, and acceleration of two objects in orbit around another (large) body—for example, two spacecraft in orbit around Earth.
```
import numpy as np
%matplotlib inline
from matplotlib import pyplot as plt
```
## Relative coordinate system
Given two... | github_jupyter |
```
%matplotlib inline
%load_ext autoreload
%autoreload 2
import sys
from pathlib import Path
sys.path.append(str(Path.cwd().parent))
from typing import Tuple
import numpy as np
import pandas as pd
from statsmodels.graphics import tsaplots
from load_dataset import Dataset
import matplotlib.pyplot as plt
import plott... | github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Tutorials/GlobalSurfaceWater/1_water_occurrence.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td>... | github_jupyter |
```
import math
import numpy as np
from joblib import load
from sklearn.ensemble import GradientBoostingClassifier
# Loading in final features test flows dicts
#
# Returns: all unknown test flows dict, mirror test flows dict, known test flows dict
def load_final_test_dicts(N):
if N == 100:
mirror_test_flo... | github_jupyter |
# Building DNN Models for Classification with TF core
Here we are using just a small subset of the data for demonstration pourposes. The complete dataset can be accessed here:
https://archive.ics.uci.edu/ml/machine-learning-databases/00280/HIGGS.csv.gz
```
import tensorflow as tf
import matplotlib.pyplot as plt
impor... | github_jupyter |
```
import os
import json
from datetime import datetime
import shutil
import subprocess
import pandas as pd
import seqeval
os.environ['MKL_THREADING_LAYER'] = 'GNU'
ROOT_DIR = !pwd
ROOT_DIR = "/".join(ROOT_DIR[0].split("/")[:-1])
ROOT_DIR
```
## Training Pipeline
```
downstream_dir = ROOT_DIR + "/token-classification... | github_jupyter |
##Tirmzi Analysis
n=1000 m+=1000 nm-=120 istep= 4 min=150 max=700
```
import sys
sys.path
import matplotlib.pyplot as plt
import numpy as np
import os
from scipy import signal
ls
import capsol.newanalyzecapsol as ac
ac.get_gridparameters
import glob
cd Output-Fortran
ls
folders = glob.glob("*NewTirmzi_large_range*/")
... | github_jupyter |
# Polynomial Regression
Polynomial Regression is a technique that is used for a nonlinear equation byt taking polynomial functions of indepedent variable.
Transform the data to polynomail. Polynomial regression is for special case of the general linear regression model. It is useful for describing curvilinear r... | github_jupyter |
# Assignment 1: Bandits and Exploration/Exploitation
Welcome to Assignment 1. This notebook will:
- Help you create your first bandit algorithm
- Help you understand the effect of epsilon on exploration and learn about the exploration/exploitation tradeoff
- Introduce you to some of the reinforcement learning software... | github_jupyter |
# NumPy and Pandas for 2D Data
This notebook contains the code assignments that are in the _NumPy and Pandas for 2D data_ lesson.
## Two-dimensional NumPy Arrays
In this section we will learn how to deal with numpy two-dimensinal arrays.
```
import numpy as np
# Subway ridership for 5 stations on 10 different days... | github_jupyter |
# Metadata management in kubeflow
```
!pip install kubeflow-metadata --user
from kubeflow.metadata import metadata
from datetime import datetime
from uuid import uuid4
METADATA_STORE_HOST = "metadata-grpc-service.kubeflow" # default DNS of Kubeflow Metadata gRPC serivce.
METADATA_STORE_PORT = 8080
#Define a workspace
... | github_jupyter |
To get started, let's import graphcat and create an empty computational graph:
```
import graphcat
graph = graphcat.StaticGraph()
```
The first step in our workflow will be to load an image from disk. We're going to use [Pillow](https://pillow.readthedocs.org) to do the heavy lifting, so you'll need to install it wi... | github_jupyter |
# Data analysis
This feature able the user to develop real-time data analysis, consist of the complete Python-powered environment, with a set of custom methods for agile development.
Extensions > New Extension > Data analysis
## Bare minimum
```
from bci_framework.extensions.data_analysis import DataAnalysis
... | github_jupyter |
```
# Initialize Otter
import otter
grader = otter.Notebook("hw07.ipynb")
```
# Homework 7 – Visualization Fundamentals 🐧
## Data 94, Spring 2021
This homework is due on **Thursday, April 8th at 11:59PM.** You must submit the assignment to Gradescope. Submission instructions can be found at the bottom of this noteb... | github_jupyter |
```
import tensorflow.keras
tensorflow.keras.__version__
```
# Understanding recurrent neural networks
This notebook contains the code samples found in Chapter 6, Section 2 of [Deep Learning with Python](https://www.manning.com/books/deep-learning-with-python?a_aid=keras&a_bid=76564dff). Note that the original text f... | github_jupyter |
# Validation of gens_eia860
This notebook runs sanity checks on the Generators data that are reported in EIA Form 860. These are the same tests which are run by the gens_eia860 validation tests by PyTest. The notebook and visualizations are meant to be used as a diagnostic tool, to help understand what's wrong when th... | github_jupyter |
# Intro to Hidden Markov Models (optional)
---
### Introduction
In this notebook, you'll use the [Pomegranate](http://pomegranate.readthedocs.io/en/latest/index.html) library to build a simple Hidden Markov Model and explore the Pomegranate API.
<div class="alert alert-block alert-info">
**Note:** You are not require... | github_jupyter |
<a id='ppd'></a>
<div id="qe-notebook-header" align="right" style="text-align:right;">
<a href="https://quantecon.org/" title="quantecon.org">
<img style="width:250px;display:inline;" width="250px" src="https://assets.quantecon.org/img/qe-menubar-logo.svg" alt="QuantEcon">
</a>
</div>
... | github_jupyter |
```
import math
import random
import gym
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torch.distributions import Normal,Beta
from sklearn import preprocessing
from IPython.display import clear_output
import matplotlib.pyplot as plt
%mat... | github_jupyter |
# Planar data classification with one hidden layer
Welcome to your week 3 programming assignment. It's time to build your first neural network, which will have a hidden layer. You will see a big difference between this model and the one you implemented using logistic regression.
**You will learn how to:**
- Implemen... | github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W3D2_HiddenDynamics/W3D2_Intro.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Intro
**Our 2021 Sponsors, including Presenting Sponsor Facebo... | github_jupyter |
```
# A script to calculate tolerance factors of ABX3 perovskites using bond valences from 2016
# Data from the International Union of Crystallography
# Author: Nick Wagner
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
bv = pd.read_csv("../Bond_valences2016.csv")
bv.head()
def calc_tol_factor(i... | github_jupyter |
# Variations on Binary Search
Now that you've gone through the work of building a binary search function, let's take some time to try out a few exercises that are variations (or extensions) of binary search. We'll provide the function for you to start:
```
def recursive_binary_search(target, source, left=0):
if ... | github_jupyter |
<a href="https://colab.research.google.com/github/phenix-project/Colabs/blob/main/CCTBX_Quickstart.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# CCTBX Quickstart
Get all dependencies installed and start coding using CCTBX
#Installation
### Inst... | github_jupyter |
# Conversations query
```
from rekall.interval_list import IntervalList, Interval
from rekall.temporal_predicates import overlaps
```
## Using Identity Labels
```
def conversationsq(video_name):
from query.models import FaceCharacterActor, Shot
from rekall.video_interval_collection import VideoIntervalCollec... | github_jupyter |
# The `Particle` Classes
The `Particle` class is the base class for all particles, whether introduced discretely one by one or as a distribution. In reality, the `Particle` class is based on two intermediate classes: `ParticleDistribution` and `ParticleInstances` to instantiate particle distributions and particles dir... | github_jupyter |
#blueqatのバックエンドを作る(簡易編)
今回はblueqatのバックエンドをqasmをベースに作る方法を確認します。今回はqiskitとcirqバックエンドを実装します。IBM社のQiskitとGoogle非公式のCirqをバックエンドとして利用してみます。
まずはインストールです。
```
pip install blueqat qiskit cirq
```
##まずQiskit
まずはQiskitです。ツールを読み込み、引数を設定してバックエンドが呼び出された時に返す値を設定すれば終わります。
```
import warnings
from collections import Counter
from bl... | github_jupyter |
```
import numpy as np
import pandas as pd
import os
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
from scipy import stats
from sklearn.linear_model import LogisticRegression
from imblearn.over_sampling import SMOTE
from co... | github_jupyter |
```
import gc
import os
import cv2
import sys
import json
import time
import timm
import torch
import random
import sklearn.metrics
from PIL import Image
from pathlib import Path
from functools import partial
from contextlib import contextmanager
import numpy as np
import scipy as sp
import pandas as pd
import torch.... | github_jupyter |
```
from random import randint, seed
import numpy as np
def random_sum_pairs(n_examples, n_numbers, largest):
X, y = [], []
for i in range(n_examples):
in_pattern = [randint(1, largest) for _ in range(n_numbers)]
out_pattern = sum(in_pattern)
X.append(in_pattern)
y.append(out_pat... | github_jupyter |
+ 目前来说就最后一点小问题:Y 比 Y_norm 跑的好,两份参考答案一份均值求的有问题应该是错的,另一份跟我遇到一样的问题。
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import scipy.io as scio
```
# 1 Anomaly detection
```
fpath = 'data/ex8data1.mat'
data = scio.loadmat(fpath)
data.keys()
X,Xval,yval = data['X'],data['Xval'],data['yval']
X.shap... | github_jupyter |
# SGDClassifier with RobustScaler & Quantile Transformer
This Code template is for classification analysis using the SGD Classifier where rescaling method used is RobustScaler and feature transformation is done via Quantile Transformer.
### Required Packages
```
import numpy as np
import pandas as pd
import seabor... | github_jupyter |
#### New to Plotly?
Plotly's Python library is free and open source! [Get started](https://plotly.com/python/getting-started/) by downloading the client and [reading the primer](https://plotly.com/python/getting-started/).
<br>You can set up Plotly to work in [online](https://plotly.com/python/getting-started/#initiali... | github_jupyter |
# Data Science Fundamentals 5
Basic introduction on how to perform typical machine learning tasks with Python.
Prepared by Mykhailo Vladymyrov & Aris Marcolongo,
Science IT Support, University Of Bern, 2020
This work is licensed under <a href="https://creativecommons.org/share-your-work/public-domain/cc0/">CC0</a>.
... | github_jupyter |
```
import pandas as pd
from bicm import BipartiteGraph
import numpy as np
from tqdm import tqdm
import csv
import itertools
import matplotlib.pyplot as plt
from sklearn.metrics import confusion_matrix, f1_score, classification_report
from sklearn.metrics import roc_curve, roc_auc_score, precision_recall_curve, averag... | github_jupyter |
##### Copyright 2018 The TensorFlow Probability Authors.
Licensed under the Apache License, Version 2.0 (the "License");
```
#@title Licensed under the Apache License, Version 2.0 (the "License"); { display-mode: "form" }
# you may not use this file except in compliance with the License.
# You may obtain a copy of th... | github_jupyter |
```
from ciml import gather_results
from ciml import tf_trainer
from sklearn.cluster import KMeans
from sklearn.mixture import GaussianMixture
import tensorflow as tf
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.cm as cmx
import matplot... | github_jupyter |
# Prospect demo 2: inspect a set of targets
See companion notebook `Prospect_demo.ipynb` for more general informations
Note that standalone VI pages (html files) can also be created from a list of targets, see examples 6 and 10 in prospect/bin/examples_prospect_pages.sh
```
import os, sys
# If not using the desicond... | github_jupyter |
<a href="https://colab.research.google.com/github/google-research/tapas/blob/master/notebooks/tabfact_predictions.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
##### Copyright 2020 The Google AI Language Team Authors
Licensed under the Apache Lic... | github_jupyter |
# Parte de función
## Bibliografía
- Wai Kai Chen, capítulo 2
- Araujo, capítulo 2
- Schaumann & M.E. Van Valkenburg, capítulo 11
## Introducción
El estudio de las funciones de red es uno de los vectores principales de la materia y el análisis pormenorizado de sus partes deriva en aplicaciones particulares a cada u... | github_jupyter |
#### Copyright 2017 Google LLC.
```
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writin... | github_jupyter |
# 用DQN强化学习算法玩“合成大西瓜”!
<iframe src="//player.bilibili.com/player.html?aid=586526003&bvid=BV1Tz4y1U7HE&cid=293880206&page=1" scrolling="no" border="0" frameborder="no" framespacing="0" allowfullscreen="true"> </iframe>
<iframe src="//player.bilibili.com/player.html?aid=801504295&bvid=BV1Wy4y1n73E&cid=294254486&page=1" ... | github_jupyter |
```
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '3'
import numpy as np
import tensorflow as tf
import json
with open('dataset-bpe.json') as fopen:
data = json.load(fopen)
train_X = data['train_X']
train_Y = data['train_Y']
test_X = data['test_X']
test_Y = data['test_Y']
EOS = 2
GO = 1
vocab_size = 32000
train_Y ... | github_jupyter |
# Day 1, Part 6: Two body motion, analytical and numeric
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
```
Let's begin by defining the mass of the star we are interested in. We'll start with something that is the mass of the Sun.
```
# mass of particle 1 in solar masses
mass_of_star = 1.... | github_jupyter |
# Tile Coding
---
Tile coding is an innovative way of discretizing a continuous space that enables better generalization compared to a single grid-based approach. The fundamental idea is to create several overlapping grids or _tilings_; then for any given sample value, you need only check which tiles it lies in. You c... | github_jupyter |
```
%matplotlib inline
```
# Violin plot basics
Violin plots are similar to histograms and box plots in that they show
an abstract representation of the probability distribution of the
sample. Rather than showing counts of data points that fall into bins
or order statistics, violin plots use kernel density estimati... | github_jupyter |
# Cart-pole Balancing Model with Amazon SageMaker and Coach library
---
## Introduction
In this notebook we'll start from the cart-pole balancing problem, where a pole is attached by an un-actuated joint to a cart, moving along a frictionless track. Instead of applying control theory to solve the problem, this exampl... | github_jupyter |
```
import os
import sys
import re
import json
import numpy as np
import pandas as pd
from collections import defaultdict
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
module_path = os.path.abspath(os.path.join('../onmt'))
if module_path not in sys.p... | github_jupyter |

[](https://colab.research.google.com/github/JohnSnowLabs/nlu/blob/master/examples/colab/Training/multi_class_text_classification/NLU_training_multi_class_text_classifier_demo_m... | github_jupyter |
```
import re
text_to_search = '''
abcdefghijklmnopqurtuvwxyz
ABCDEFGHIJKLMNOPQRSTUVWXYZ
1234567890
Ha HaHa
MetaCharacters (Need to be escaped):
. ^ $ * + ? { } [ ] \ | ( )
coreyms.com
321-555-4321
123.555.1234
123*555*1234
800-555-1234
900-555-1234
Mr. Schafer
Mr Smith
Ms Davis
Mrs. Robinson
Mr. T
cat
mat
bat
'... | github_jupyter |
This page explains the multiple layouts components and all the options to control the layout of the dashboard.
There are 4 main components in a jupyter-flex dashboard in this hierarchy:
1. Pages
2. Sections
3. Cards
4. Cells
Meaning that Pages contain one or more Sections, Sections contains one or multiple Cards an... | github_jupyter |
```
# Import libraries
import numpy as np
import matplotlib.pyplot as plt
# Import libraries
import keras
import keras.backend as K
from keras.models import Model
# Activation and Regularization
from keras.regularizers import l2
from keras.activations import softmax
# Keras layers
from keras.layers.convolutional import... | github_jupyter |
# 飞桨常规赛:PALM眼底彩照中黄斑中央凹定位 - 12月第3名方案
# (1)比赛介绍
## 赛题介绍
PALM黄斑定位常规赛的重点是研究和发展与患者眼底照片黄斑结构定位相关的算法。该常规赛的目标是评估和比较在一个常见的视网膜眼底图像数据集上定位黄斑的自动算法。具体目的是预测黄斑中央凹在图像中的坐标值。
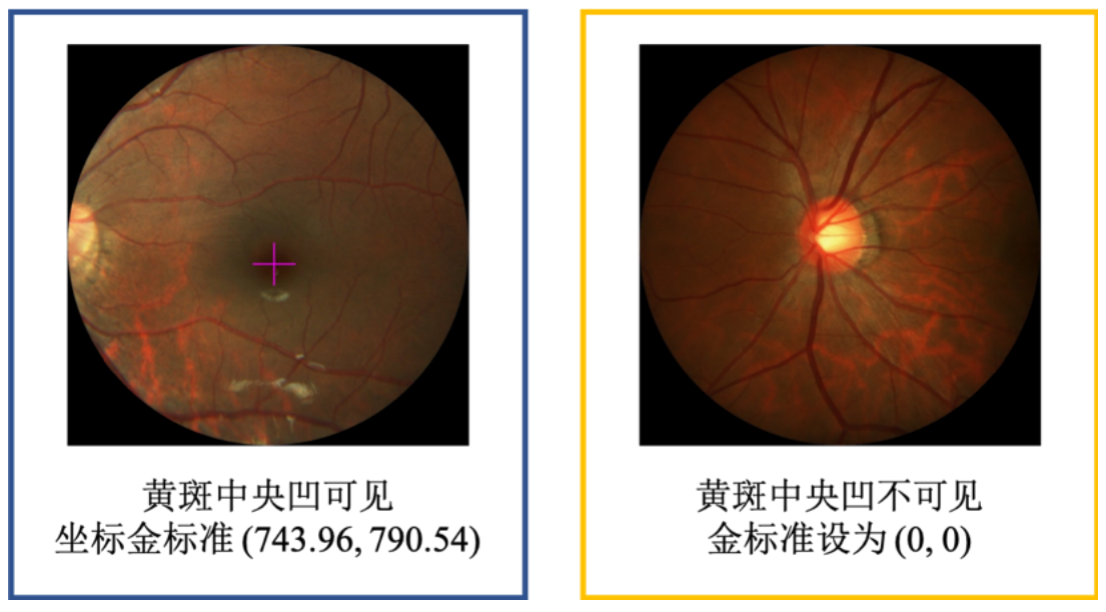
中央凹是视网膜中辨色力、分辨力最敏锐的区域。以人为例,在视盘颞侧约3.5mm处... | github_jupyter |
```
import pandas as pd
import re
import os
import time
import random
import numpy as np
try:
%tensorflow_version 2.x # enable TF 2.x in Colab
except Exception:
pass
import tensorflow as tf
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
from sklearn.model_selection import train_test_split
from... | github_jupyter |
[](https://github.com/labmlai/annotated_deep_learning_paper_implementations)
[](https://colab.research.google.com/github/labmlai/anno... | github_jupyter |
# 14. Image classification by machine learning: Optical text recognition
There are different types of machine learning. In some cases, like in the pixel classification task, the algorithm does the classification on its own by trying to optimize groups according to a given rule (unsupervised). In other cases one has to... | github_jupyter |
##### Copyright 2021 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or ... | github_jupyter |
# Part I: Set Up
- Import Packages
```
import seaborn as sns
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.pyplot as plt2
import pandas as pd
from pandas import datetime
import math, time
import itertools
from sklearn import preprocessing
import datetime
from sklearn.metrics import mean_squared... | github_jupyter |
```
import numpy as np
import pandas as pd
import torch
import torchvision.datasets as datasets
import torchvision.transforms as transforms
from torch.utils.data.sampler import SubsetRandomSampler
import torch.utils.data as DataUtils
import numpy as np
import time
import sys
import torch.nn as nn
import torch.nn.funct... | github_jupyter |
# Imports
```
%matplotlib inline
import pandas as pd
import numpy as np
from sklearn import model_selection, linear_model
import matplotlib.pyplot as plt
```
# Functions
```
def normalize(a):
return (a - np.min(a)) / (np.max(a) - np.min(a))
def linear_regression(x, y, iters, alpha):
m = len(x)
cost = np... | github_jupyter |
# OpenRXN Example: Membrane slab
### This notebook demostrates how a complicated system can be set up easily with the OpenRXN package
We are interested in setting up a 3D system with a membrane slab at the bottom (both lower and upper leaflets), and a bulk region on top. There will be three Species in our model (dru... | github_jupyter |
```
## A taste of things to come
# Print the list created using the Non-Pythonic approach
i = 0
new_list= []
while i < len(names):
if len(names[i]) >= 6:
new_list.append(names[i])
i += 1
print(new_list)
# Print the list created by looping over the contents of names
better_list = []
for name in names:
... | github_jupyter |
```
import numpy as np
import matplotlib.pyplot as pp
import pandas as pd
import seaborn
%matplotlib inline
import zipfile
zipfile.ZipFile('names.zip').extractall('.')
import os
os.listdir('names')
open('names/yob2011.txt','r').readlines()[:10]
names2011 = pd.read_csv('names/yob2011.txt')
names2011.head()
names2011 = p... | github_jupyter |
# Day 7 "The Treachery of Whales"
## Part 1
### Problem
A giant whale has decided your submarine is its next meal, and it's much faster than you are. There's nowhere to run!
Suddenly, a swarm of crabs (each in its own tiny submarine - it's too deep for them otherwise) zooms in to rescue you! They seem to be prepari... | github_jupyter |
# Simulation and comparison of dual pol and dual pol diagonal only
```
%matplotlib inline
import numpy as np
from osgeo import gdal
from osgeo.gdalconst import GDT_Float32, GA_ReadOnly
def make_simimage(fn,m=5,bands=9,sigma=1,alpha=0.2,beta=0.2):
simimage = np.zeros((100**2,9))
ReSigma = np.zeros((3,3))
... | github_jupyter |
```
import os
import pandas as pd
from pvoutput import *
```
* Uses PVOutput.org API search to try to get all systems in UK.
* The API search only allows us to get all systems within a search radius of <= 25 km.
* This script loads the appropriately-spaced UK grid points (generated with `get_grid_points_for_UK.ipynb`)... | github_jupyter |
<img src="http://i67.tinypic.com/2jcbwcw.png" align="left"></img><br><br><br><br>
## Notebook: Web Scraping & Web Crawling
**Author List**: Alexander Fred Ojala
**Original Sources**: https://www.crummy.com/software/BeautifulSoup/bs4/doc/ & https://www.dataquest.io/blog/web-scraping-tutorial-python/
**License**: Fe... | github_jupyter |
```
import os
os.chdir(os.path.split(os.getcwd())[0])
import random
import numpy as np
import matplotlib.pyplot as plt
import gym
from agent import *
from optionpricing import *
import yaml
import torch
from collections import defaultdict
import matplotlib.style as style
style.use('seaborn-poster')
experiment_folder = ... | github_jupyter |
# 유사 이미지 검출 샘플모델 데모
DNN기반 이미지 유사도 검출 샘플 모델(BaseNet) 데모.
```
# load package
import tensorflow as tf
from functools import partial
import itertools
from tensorflow.keras.datasets import mnist
import numpy as np
import cv2
from matplotlib import pyplot as plt
import os.path as osp
from pathlib import Path
from model impo... | github_jupyter |
Cotton Diseases Prediction Detection Using Deep Learning
```
from tensorflow.compat.v1 import ConfigProto, InteractiveSession
config=ConfigProto()
config.gpu_options.per_process_gpu_memory_fraction=0.5
config.gpu_options.allow_growth=True
session = InteractiveSession(config=config)
from tensorflow.keras.layers import... | github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or ... | github_jupyter |
# Monitor a Model
When you've deployed a model into production as a service, you'll want to monitor it to track usage and explore the requests it processes. You can use Azure Application Insights to monitor activity for a model service endpoint.
## Install the Azure Machine Learning SDK
The Azure Machine Learning SD... | github_jupyter |
<a href="https://colab.research.google.com/github/PUC-RecSys-Class/RecSysPUC-2020/blob/master/practicos/pyRecLab_MostPopular.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
<a href="https://youtu.be/MEY4UK4QCP4" target="_parent"><img src="https://up... | github_jupyter |
Best Model : LSTM on Hist. of pixels ( 16 bin)
```
import math
from pandas import DataFrame
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import mean_squared_error
from numpy import array
from keras.layers import Convolution2D, MaxPooling2D, Flatten, Reshape,Conv2D
from keras.models import Seque... | github_jupyter |
# Artificial Intelligence Nanodegree
## Convolutional Neural Networks
---
In this notebook, we train an MLP to classify images from the MNIST database.
### 1. Load MNIST Database
```
from keras.datasets import mnist
# use Keras to import pre-shuffled MNIST database
(X_train, y_train), (X_test, y_test) = mnist.loa... | github_jupyter |
Deep Learning Models -- A collection of various deep learning architectures, models, and tips for TensorFlow and PyTorch in Jupyter Notebooks.
- Author: Sebastian Raschka
- GitHub Repository: https://github.com/rasbt/deeplearning-models
---
```
%load_ext watermark
%watermark -a 'Sebastian Raschka' -v -p torch
```
# ... | github_jupyter |
```
import numpy as np
from numba import jit
from scipy import ndimage
from osgeo import gdal, osr, ogr
import matplotlib.pyplot as plt
plt.style.use('default')
@jit(nopython=True)
def np_mean(neighborhood):
return np.nanmean(neighborhood)
def lonlat_to_utm(lon, lat):
if lat < 0:
return int(32700 + np.... | github_jupyter |
<a href="https://colab.research.google.com/github/Rajansharma05/A-mobile-based-photo-editing-app/blob/master/Copy_of_Copy_of_Welcome_to_Colaboratory.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
<p><img alt="Colaboratory logo" height="45px" src="/... | github_jupyter |
# Prepare Data for TPZ
* query GCR with the same cuts we used for BPZ
* deredden the magnitudes
* fill in missing values
* reformat to TPZ output
## Query GCR with the same cuts we used for BPZ
```
# everything we need for the whole notebook
import sys
import numpy as np
import pandas as pd
import matplotlib.pyplot... | github_jupyter |
# making an aggregate master dataframe for baseline model 10/22/18 -11/6/18
this notebook is going to make standardized longformat dataframes for each dataframe that i will adjust for each model to.
for my first pass, i will work to establish a baseline model by using:
the single "worst", or value that most indicate... | github_jupyter |
### Primitive Data Types: Booleans
These are the basic data types that constitute all of the more complex data structures in python. The basic data types are the following:
* Strings (for text)
* Numeric types (integers and decimals)
* Booleans
### Booleans
Booleans represent the truth or success of a statement, an... | github_jupyter |
Copyright Amazon.com, Inc. or its affiliates. All Rights Reserved.
SPDX-License-Identifier: Apache-2.0
# Using the AWS Batch Architecture for AlphaFold
This notebook allows you to predict protein structures using AlphaFold on AWS Batch.
**Differences to AlphaFold Notebooks**
In comparison to AlphaFold v2.1.0, this... | github_jupyter |
# Convolutional Neural Networks: Step by Step
Welcome to Course 4's first assignment! In this assignment, you will implement convolutional (CONV) and pooling (POOL) layers in numpy, including both forward propagation and (optionally) backward propagation.
**Notation**:
- Superscript $[l]$ denotes an object of the $l... | github_jupyter |
```
#Importing relevant libraries
import numpy as np
import pandas as pd
from pathlib import Path
import os.path
import matplotlib.pyplot as plt
from IPython.display import Image, display
import matplotlib.cm as cm
from sklearn.model_selection import train_test_split
import tensorflow as tf
#Set the image di... | github_jupyter |
```
%matplotlib inline
import gym
import itertools
import matplotlib
import numpy as np
import pandas as pd
import sys
if "../" not in sys.path:
sys.path.append("../")
from collections import defaultdict
from lib.envs.windy_gridworld import WindyGridworldEnv
from lib import plotting
matplotlib.style.use('ggplot'... | github_jupyter |
<a href="https://colab.research.google.com/github/sokrypton/ColabFold/blob/main/beta/alphafold_output_at_each_recycle.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
%%bash
if [ ! -d alphafold ]; then
pip -q install biopython dm-haiku ml-colle... | github_jupyter |
<small><small><i>
All the IPython Notebooks in this **Python Examples** series by Dr. Milaan Parmar are available @ **[GitHub](https://github.com/milaan9/90_Python_Examples)**
</i></small></small>
# Python Program to Differentiate Between `del`, `remove`, and `pop` on a List
In this example, you will learn to differe... | github_jupyter |
```
import scipy.io
import torch
import numpy as np
import torch.nn as nn
import torch.utils.data as Data
import matplotlib.pyplot as plt
import torch.nn.functional as F
#from tensorboardX import SummaryWriter
from sklearn.metrics import roc_auc_score,roc_curve,auc,average_precision_score,precision_recall_curve
torch.m... | github_jupyter |
# Python and Web Tutorial
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Python-and-Web-Tutorial" data-toc-modified-id="Python-and-Web-Tutorial-1">Python and Web Tutorial</a></span></li><li><span><a href="#웹(Web)" data-toc-modified-id="웹(Web)-2">웹(Web)... | github_jupyter |
```
import os
import os.path as op
import psutil
from pathlib import Path
from IPython.display import display
import findspark
findspark.init()
import pyspark.sql.functions as F
from pyspark import SparkConf
from pyspark.sql import SparkSession
from pyspark.sql.functions import col
if os.getenv('SLURM_TMPDIR'):
SP... | github_jupyter |
```
import sys, os, re, csv, codecs, numpy as np, pandas as pd
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.layers import Dense, Input, LSTM, Embedding, Dropout, Activation
from keras.layers import Bidirectional, GlobalMaxPool1D
from keras.models impo... | github_jupyter |
# Rust Crash Course - 01 - Variables and Data Types
In order to process data correctly and efficiently, Rust needs to know the data type of a variable.
In the following, variables and common data types of the Rust programming language are explained.
The contents represent a brief and compact introduction to the topi... | github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.