text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
```
import numpy as np
import pandas as pd
import pyodbc
import time
import pickle
import operator
from operator import itemgetter
from joblib import Parallel, delayed
from sklearn import linear_model
from sklearn.linear_model import Ridge
from sklearn.tree import DecisionTreeRegressor
from sklearn.model_selection import cross_val_score
from sqlalchemy import create_engine
import matplotlib.pyplot as plt
%matplotlib inline
```
Before everything, please download a preprocessed version of the natality data from https://www.dropbox.com/s/395rrh0c826gw9r/Natality_small.csv?dl=0
```
conn = pyodbc.connect("Driver={SQL Server Native Client 11.0};"
"Server=localhost;"
"Database=master;"
"Trusted_Connection=yes;")
cur = conn.cursor()
engine = create_engine('mssql+pyodbc://localhost/master?driver=SQL+Server+Native+Client+11.0')
def construct_sec_order(arr):
# data generation function helper.
second_order_feature = []
num_cov_sec = len(arr[0])
for a in arr:
tmp = []
for i in range(num_cov_sec):
for j in range(i+1, num_cov_sec):
tmp.append( a[i] * a[j] )
second_order_feature.append(tmp)
return np.array(second_order_feature)
def data_generation_dense_2(num_control, num_treated, num_cov_dense, num_covs_unimportant,
control_m = 0.1, treated_m = 0.9):
# the data generation function that I'll use.
xc = np.random.binomial(1, 0.5, size=(num_control, num_cov_dense)) # data for conum_treatedrol group
xt = np.random.binomial(1, 0.5, size=(num_treated, num_cov_dense)) # data for treatmenum_treated group
errors1 = np.random.normal(0, 0.05, size=num_control) # some noise
errors2 = np.random.normal(0, 0.05, size=num_treated) # some noise
dense_bs_sign = np.random.choice([-1,1], num_cov_dense)
#dense_bs = [ np.random.normal(dense_bs_sign[i]* (i+2), 1) for i in range(len(dense_bs_sign)) ]
dense_bs = [ np.random.normal(s * 10, 1) for s in dense_bs_sign ]
yc = np.dot(xc, np.array(dense_bs)) + errors1 # y for conum_treatedrol group
treatment_eff_coef = np.random.normal( 1.5, 0.15, size=num_cov_dense)
treatment_effect = np.dot(xt, treatment_eff_coef)
second = construct_sec_order(xt[:,:5])
treatment_eff_sec = np.sum(second, axis=1)
yt = np.dot(xt, np.array(dense_bs)) + treatment_effect + treatment_eff_sec + errors2 # y for treated group
xc2 = np.random.binomial(1, control_m, size=(num_control, num_covs_unimportant)) #
xt2 = np.random.binomial(1, treated_m, size=(num_treated, num_covs_unimportant)) #
df1 = pd.DataFrame(np.hstack([xc, xc2]),
columns=['{0}'.format(i) for i in range(num_cov_dense + num_covs_unimportant)])
df1['outcome'] = yc
df1['treated'] = 0
df2 = pd.DataFrame(np.hstack([xt, xt2]),
columns=['{0}'.format(i) for i in range(num_cov_dense + num_covs_unimportant )] )
df2['outcome'] = yt
df2['treated'] = 1
df = pd.concat([df1,df2])
df['matched'] = 0
return df, dense_bs, treatment_eff_coef
# this function takes the current covariate list, the covariate we consider dropping, name of the data table,
# name of the holdout table, the threshold (below which we consider as no match), and balancing regularization
# as input; and outputs the matching quality
def score_tentative_drop_c(cov_l, c, db_name, holdout_df, thres = 0, tradeoff = 0.1):
covs_to_match_on = set(cov_l) - {c} # the covariates to match on
# the flowing query fetches the matched results (the variates, the outcome, the treatment indicator)
s = time.time()
##cur.execute('''with temp AS
## (SELECT
## {0}
## FROM {3}
## where "matched"=0
## group by {0}
## Having sum("treated")>'0' and sum("treated")<count(*)
## )
## (SELECT {1}, {3}."treated", {3}."outcome"
## FROM temp, {3}
## WHERE {2}
## )
## '''.format(','.join(['"C{0}"'.format(v) for v in covs_to_match_on ]),
## ','.join(['{1}."C{0}"'.format(v, db_name) for v in covs_to_match_on ]),
## ' AND '.join([ '{1}."C{0}"=temp."C{0}"'.format(v, db_name) for v in covs_to_match_on ]),
## db_name
## ) )
##res = np.array(cur.fetchall())
cur.execute('''with temp AS
(SELECT
{0}
FROM {3}
where "matched"=0
group by {0}
Having sum("treated")>'0' and sum("treated")<count(*)
)
(SELECT {1}, treated, outcome
FROM {3}
WHERE EXISTS
(SELECT 1
FROM temp
WHERE {2}
)
)
'''.format(','.join(['"{0}"'.format(v) for v in covs_to_match_on ]),
','.join(['{1}."{0}"'.format(v, db_name) for v in covs_to_match_on ]),
' AND '.join([ '{1}."{0}"=temp."{0}"'.format(v, db_name) for v in covs_to_match_on ]),
db_name
) )
res = np.array(cur.fetchall())
time_match = time.time() - s
s = time.time()
# the number of unmatched treated units
cur.execute('''select count(*) from {} where "matched" = 0 and "treated" = 0'''.format(db_name))
num_control = cur.fetchall()
# the number of unmatched control units
cur.execute('''select count(*) from {} where "matched" = 0 and "treated" = 1'''.format(db_name))
num_treated = cur.fetchall()
time_BF = time.time() - s
# fetch from database the holdout set
##s = time.time()
##cur.execute('''select {0}, "treated", "outcome"
## from {1}
## '''.format( ','.join( ['"C{0}"'.format(v) for v in covs_to_match_on ] ) , holdout))
##holdout = np.array(cur.fetchall())
s = time.time() # the time for fetching data into memory is not counted if use this
# below is the regression part for PE
#ridge_c = Ridge(alpha=0.1)
#ridge_t = Ridge(alpha=0.1)
tree_c = DecisionTreeRegressor(max_depth=8, random_state=0)
tree_t = DecisionTreeRegressor(max_depth=8, random_state=0)
holdout = holdout_df.copy()
holdout = holdout[ list(covs_to_match_on) + ['treated', 'outcome']]
mse_t = np.mean(cross_val_score(tree_t, holdout[holdout['treated'] == 1].iloc[:,:-2],
holdout[holdout['treated'] == 1]['outcome'] , scoring = 'neg_mean_squared_error' ) )
mse_c = np.mean(cross_val_score(tree_c, holdout[holdout['treated'] == 0].iloc[:,:-2],
holdout[holdout['treated'] == 0]['outcome'], scoring = 'neg_mean_squared_error' ) )
#mse_t = np.mean(cross_val_score(ridge_t, holdout[holdout['treated'] == 1].iloc[:,:-2],
# holdout[holdout['treated'] == 1]['outcome'] , scoring = 'neg_mean_squared_error' ) )
#mse_c = np.mean(cross_val_score(ridge_c, holdout[holdout['treated'] == 0].iloc[:,:-2],
# holdout[holdout['treated'] == 0]['outcome'], scoring = 'neg_mean_squared_error' ) )
# above is the regression part for BF
time_PE = time.time() - s
if len(res) == 0:
return (( mse_t + mse_c ), time_match, time_PE, time_BF)
##return mse_t + mse_c
else:
return (tradeoff * (float(len(res[res[:,-2]==0]))/num_control[0][0] + float(len(res[res[:,-2]==1]))/num_treated[0][0]) +\
( mse_t + mse_c ), time_match, time_PE, time_BF)
##return reg_param * (float(len(res[res[:,-2]==0]))/num_control[0][0] + float(len(res[res[:,-2]==1]))/num_treated[0][0]) +\
## ( mse_t + mse_c )
# update matched units
# this function takes the currcent set of covariates and the name of the database; and update the "matched"
# column of the newly mathced units to be "1"
def update_matched(covs_matched_on, db_name, level):
cur.execute('''with temp AS
(SELECT
{0}
FROM {3}
where "matched"=0
group by {0}
Having sum("treated")>'0' and sum("treated")<count(*)
)
update {3} set "matched"={4}
WHERE EXISTS
(SELECT {0}
FROM temp
WHERE {2} and {3}."matched" = 0
)
'''.format(','.join(['"{0}"'.format(v) for v in covs_matched_on]),
','.join(['{1}."{0}"'.format(v, db_name) for v in covs_matched_on]),
' AND '.join([ '{1}."{0}"=temp."{0}"'.format(v, db_name) for v in covs_matched_on ]),
db_name,
level
) )
conn.commit()
return
# get CATEs
# this function takes a list of covariates and the name of the data table as input and outputs a dataframe
# containing the combination of covariate values and the corresponding CATE
# and the corresponding effect (and the count and variance) as values
def get_CATE(cov_l, db_name, level):
cur.execute(''' select {0}, avg(outcome * 1.0), count(*)
from {1}
where matched = {2} and treated = 0
group by {0}
'''.format(','.join(['"{0}"'.format(v) for v in cov_l]),
db_name, level) )
res_c = cur.fetchall()
cur.execute(''' select {0}, avg(outcome * 1.0), count(*)
from {1}
where matched = {2} and treated = 1
group by {0}
'''.format(','.join(['"{0}"'.format(v) for v in cov_l]),
db_name, level) )
res_t = cur.fetchall()
if (len(res_c) == 0) | (len(res_t) == 0):
return None
cov_l = list(cov_l)
result = pd.merge(pd.DataFrame(np.array(res_c), columns=['{}'.format(i) for i in cov_l]+['effect_c', 'count_c']),
pd.DataFrame(np.array(res_t), columns=['{}'.format(i) for i in cov_l]+['effect_t', 'count_t']),
on = ['{}'.format(i) for i in cov_l], how = 'inner')
result_df = result[['{}'.format(i) for i in cov_l] + ['effect_c', 'effect_t', 'count_c', 'count_t']]
# -- the following section are moved to after getting the result
#d = {}
#for i, row in result.iterrows():
# k = ()
# for j in range(len(cov_l)):
# k = k + ((cov_l[j], row[j]),)
# d[k] = (row['effect_c'], row['effect_t'], row['std_t'], row['std_c'], row['count_c'], row['count_t'])
# -- the above section are moved to after getting the result
return result_df
def run(db_name, holdout_df, num_covs, reg_param = 0.1):
cur.execute('update {0} set matched = 0'.format(db_name)) # reset the matched indicator to 0
conn.commit()
covs_dropped = [] # covariate dropped
ds = []
level = 1
timings = [0]*5 # first entry - match (groupby and join),
# second entry - regression (compute PE),
# third entry - compute BF,
# fourth entry - keep track of CATE,
# fifth entry - update database table (mark matched units).
cur_covs = range(num_covs) # initialize the current covariates to be all covariates
# make predictions and save to disk
s = time.time()
update_matched(cur_covs, db_name, level) # match without dropping anything
timings[4] = timings[4] + time.time() - s
s = time.time()
d = get_CATE(cur_covs, db_name, level) # get CATE without dropping anything
timings[3] = timings[3] + time.time() - s
ds.append(d)
##s = time.time()
##cur.execute('''update {} set "matched"=2 WHERE "matched"=1 '''.format(db_name)) # mark the matched units as matched and
#they are no langer seen by the algorithm
##conn.commit()
##timings[4] = timings[4] + time.time() - s
while len(cur_covs)>1:
print(cur_covs) # print current set of covariates
level += 1
# the early stopping conditions
cur.execute('''select count(*) from {} where "matched"=0 and "treated"=0'''.format(db_name))
if cur.fetchall()[0][0] == 0:
break
cur.execute('''select count(*) from {} where "matched"=0 and "treated"=1'''.format(db_name))
if cur.fetchall()[0][0] == 0:
break
best_score = -np.inf
cov_to_drop = None
cur_covs = list(cur_covs)
for c in cur_covs:
score,time_match,time_PE,time_BF = score_tentative_drop_c(cur_covs, c, db_name,
holdout_df, tradeoff = 0.1)
timings[0] = timings[0] + time_match
timings[1] = timings[1] + time_PE
timings[2] = timings[2] + time_BF
if score > best_score:
best_score = score
cov_to_drop = c
cur_covs = set(cur_covs) - {cov_to_drop} # remove the dropped covariate from the current covariate set
s = time.time()
update_matched(cur_covs, db_name, level)
timings[4] = timings[4] + time.time() - s
s = time.time()
d = get_CATE(cur_covs, db_name, level)
timings[3] = timings[3] + time.time() - s
ds.append(d)
##s = time.time()
##cur.execute('''update {} set "matched"=2 WHERE "matched"=1 '''.format(db_name))
##conn.commit()
##timings[4] = timings[4] + time.time() - s
covs_dropped.append(cov_to_drop) # append the removed covariate at the end of the covariate
return timings, ds
```
PGARScore as outcome
```
df = pd.read_csv('Natality_small.csv')
df = df[df['ABAssistedVentilation'] != 'U']
df = df[df['ABAssistedVentilationMoreThan6Hrs'] != 'U']
df = df[df['ABAdmissionToNICU'] != 'U']
df = df[df['ABSurfactant'] != 'U']
df = df[df['ABAntibiotics'] != 'U']
df = df[df['ABSeizures'] != 'U']
df = df.drop(columns=['FiveMinuteAPGARScoreRecode', 'DeliveryMethodRecodeRevised'])
df['outcome'] = df['FiveMinAPGARScore']
df = df[df['outcome'] != 99]
df = df.drop(columns=['FiveMinAPGARScore'])
df.loc[df['CigaretteRecode'] == 'Y', 'CigaretteRecode'] = 1
df.loc[df['CigaretteRecode'] == 'N', 'CigaretteRecode'] = 0
df['matched'] = 0
cols = [c for c in df.columns if c.lower()[:4] != 'flag']
df = df[cols]
char_col = []
for c in df.columns:
if df[c].dtype != np.int64:
char_col.append(c)
for c in char_col:
df[c] = df[c].astype(str)
l = sorted(list(np.unique(df[c])))
print c, l
for i in range(len(l)):
df.loc[df[c] == l[i], c] = i
rename_dict = dict()
cols = list(df.columns)
cols.remove('matched')
cols.remove('outcome')
cols.remove('CigaretteRecode')
for i in range(len(cols)):
rename_dict[i] = cols[i]
rename_dict['treated'] = 'CigaretteRecode'
inv_rename_dict = {v:k for k,v in rename_dict.iteritems()}
import pickle
pickle.dump(rename_dict, open('natality_rename_dict_score', 'wb'))
df.rename(columns = inv_rename_dict, inplace = True)
from sklearn.model_selection import train_test_split
df, holdout = train_test_split(df, test_size = 0.1, random_state = 345)
df.to_csv('Natality_db_score.csv', index = False)
holdout.to_csv('Natality_holdout_score.csv', index = False)
#holdout = pd.read_csv('Natality_holdout.csv')
df.to_sql('natality_score', engine, chunksize=100)
del df
#df.to_sql('natality', engine, chunksize=100)
holdout = pd.read_csv('Natality_holdout_score.csv')
holdout.rename(columns = {str(i):i for i in range(166)}, inplace = True)
#del df
db_name = 'natality_score'
s = time.time()
res = run(db_name, holdout, 91)
print (time.time() - s)
#pickle.dump(res, open('natality_cigar_score_res', 'wb'))
```
abnormality as outcome, with prenatal care start time used to define sub-populations.
```
df = pd.read_csv('Natality_small.csv')
df = df[df['ABAssistedVentilation'] != 'U']
df = df[df['ABAssistedVentilationMoreThan6Hrs'] != 'U']
df = df[df['ABAdmissionToNICU'] != 'U']
df = df[df['ABSurfactant'] != 'U']
df = df[df['ABAntibiotics'] != 'U']
df = df[df['ABSeizures'] != 'U']
df = df.drop(columns=['FiveMinuteAPGARScoreRecode', 'DeliveryMethodRecodeRevised'])
df['outcome'] = np.array((df['ABAssistedVentilation'] == 'Y' ) |\
(df['ABAssistedVentilationMoreThan6Hrs'] == 'Y') |\
(df['ABAdmissionToNICU'] == 'Y') |\
(df['ABSurfactant'] == 'Y') |\
(df['ABAntibiotics'] == 'Y') |\
(df['ABSeizures'] == 'Y') )
df['outcome'] = df['outcome'].astype(int)
df = df.drop(columns = ['ABAssistedVentilation', 'ABAssistedVentilationMoreThan6Hrs',
'ABAdmissionToNICU', 'ABSurfactant', 'ABAntibiotics', 'ABSeizures'])
df.loc[df['CigaretteRecode'] == 'Y', 'CigaretteRecode'] = 1
df.loc[df['CigaretteRecode'] == 'N', 'CigaretteRecode'] = 0
df['matched'] = 0
cols = [c for c in df.columns if c.lower()[:4] != 'flag']
df = df[cols]
char_col = []
for c in df.columns:
if df[c].dtype != np.int64:
char_col.append(c)
for c in char_col:
df[c] = df[c].astype(str)
l = sorted(list(np.unique(df[c])))
print c, l
for i in range(len(l)):
df.loc[df[c] == l[i], c] = i
rename_dict = dict()
cols = list(df.columns)
cols.remove('matched')
cols.remove('outcome')
cols.remove('CigaretteRecode')
for i in range(len(cols)):
rename_dict[i] = cols[i]
rename_dict['treated'] = 'CigaretteRecode'
inv_rename_dict = {v:k for k,v in rename_dict.iteritems()}
import pickle
pickle.dump(rename_dict, open('natality_rename_dict_abnormality', 'wb'))
df.rename(columns = inv_rename_dict, inplace = True)
from sklearn.model_selection import train_test_split
df, holdout = train_test_split(df, test_size = 0.1, random_state = 345)
df.to_csv('Natality_db_abnormality.csv', index = False)
holdout.to_csv('Natality_holdout_abnormality.csv', index = False)
#holdout = pd.read_csv('Natality_holdout.csv')
df.to_sql('natality_abnormality', engine, chunksize=100)
holdout = pd.read_csv('Natality_holdout_abnormality.csv')
#df.to_sql('natality', engine, chunksize=100)
holdout = pd.read_csv('Natality_holdout_abnormality.csv')
holdout.rename(columns = {str(i):i for i in range(166)}, inplace = True)
#del df
db_name = 'natality_abnormality'
s = time.time()
res = run(db_name, holdout, 86)
print (time.time() - s)
pickle.dump(res, open('natality_cigar_abnormality_res', 'wb'))
res = pickle.load(open('natality_cigar_score_res', 'rb'))
rename_dict = pickle.load(open('natality_rename_dict_score'))
def weighted_avg_and_std(values, weights):
"""
Return the weighted average and standard deviation.
values, weights -- Numpy ndarrays with the same shape.
"""
ave = np.average(values, weights=weights)
var = np.average((values-ave)**2, weights=weights) # Fast and numerically precise
return (ave, np.sqrt(var))
g1_effect = []
g2_effect = []
g1_size = []
g2_size = []
for i in range(len(res[1])):
print i
r = res[1][i]
if r is None:
continue
if '11' not in list(r.columns):
break
r_small = r[(r['11'] <= 2) ]
r_large = r[(r['11'] >= 3) & (r['11'] <= 4) ]
g1_effect = g1_effect + list(r_small['effect_t'] - r_small['effect_c'] )
g1_size = g1_size + list(r_small['count_t'] + r_small['count_c'] )
g2_effect = g2_effect + list( r_large['effect_t'] - r_large['effect_c'] )
g2_size = g2_size + list( r_large['count_t'] + r_large['count_c'] )
g1_effect = [float(v) for v in g1_effect ]
g2_effect = [float(v) for v in g2_effect ]
g1_size = [float(v) for v in g1_size ]
g2_size = [float(v) for v in g2_size ]
g1_mean, g1_std = weighted_avg_and_std(np.array(g1_effect), np.array(g1_size) )
g2_mean, g2_std = weighted_avg_and_std(np.array(g2_effect), np.array(g2_size) )
plt.rcParams['font.size'] = 15
plt.figure(figsize = (5,5))
plt.scatter([0,1], [g1_mean, g2_mean])
plt.errorbar([0,1], [g1_mean, g2_mean], yerr = [g1_std, g2_std], linestyle = 'None')
plt.xticks([0,1], ['0', '1'])
plt.xlabel('Prenatal Care Beginning Time Code')
plt.ylabel('Estimated Treatment Effect')
plt.xlim([-1,2])
plt.ylim([-1,1])
plt.tight_layout()
plt.savefig('natality_score_prenatal.png', dpi = 150)
res = pickle.load(open('natality_cigar_score_res', 'rb'))
rename_dict = pickle.load(open('natality_rename_dict_score'))
effect = []
size = []
for i in range(len(res[1])):
print i
r = res[1][i]
if r is None:
continue
effect = effect + list(r['effect_t'] - r['effect_c'] )
size = size + list(r['count_t'] + r['count_c'] )
effect = [float(v) for v in effect ]
size = [float(v) for v in size ]
avr, std = weighted_avg_and_std(np.array(effect), np.array(size) )
g1_effect = []
g2_effect = []
g3_effect = []
g1_size = []
g2_size = []
g3_size = []
for i in range(len(res[1])):
print i
r = res[1][i]
if r is None:
continue
if '11' not in list(r.columns):
break
#r1 = r[(r['6'] == 0) ]
r2 = r[(r['7'] >=7) ]
r3 = r[(r['7'] < 7)]
#g1_effect = g1_effect + list(r1['effect_t'] - r1['effect_c'] )
#g1_size = g1_size + list(r1['count_t'] + r1['count_c'] )
g2_effect = g2_effect + list( r2['effect_t'] - r2['effect_c'] )
g2_size = g2_size + list( r2['count_t'] + r2['count_c'] )
g3_effect = g3_effect + list( r3['effect_t'] - r3['effect_c'] )
g3_size = g3_size + list( r3['count_t'] + r3['count_c'] )
g1_effect = [float(v) for v in g1_effect ]
g2_effect = [float(v) for v in g2_effect ]
g3_effect = [float(v) for v in g3_effect ]
g1_size = [float(v) for v in g1_size ]
g2_size = [float(v) for v in g2_size ]
g3_size = [float(v) for v in g3_size ]
#g1_mean, g1_std = weighted_avg_and_std(np.array(g1_effect), np.array(g1_size) )
g2_mean, g2_std = weighted_avg_and_std(np.array(g2_effect), np.array(g2_size) )
g3_mean, g3_std = weighted_avg_and_std(np.array(g3_effect), np.array(g3_size) )
plt.rcParams['font.size'] = 15
plt.figure(figsize = (5,5))
plt.scatter([0,1,2], [g3_mean, g2_mean, avr])
plt.errorbar([0,1,2], [g3_mean, g2_mean, avr], yerr = [g3_std, g2_std, std], linestyle = 'None')
plt.xticks([0,1,2], ['0', '1', 'whole'])
plt.xlabel("Gestational Hypertension Code")
plt.ylabel('Estimated Treatment Effect')
plt.xlim([-1,3])
#plt.ylim([-1,1])
plt.tight_layout()
#plt.savefig('natality_score_hypertensions_whole.png', dpi = 150)
```
| github_jupyter |
```
# Import some libraries
import torch
import torchvision
from torch import nn
from torch.utils.data import DataLoader
from torchvision import transforms
from torchvision.datasets import MNIST
from matplotlib import pyplot as plt
# Convert vector to image
def to_img(x):
x = 0.5 * (x + 1)
x = x.view(x.size(0), 28, 28)
return x
# Displaying routine
def display_images(in_, out, n=1):
for N in range(n):
if in_ is not None:
in_pic = to_img(in_.cpu().data)
plt.figure(figsize=(18, 6))
for i in range(4):
plt.subplot(1,4,i+1)
plt.imshow(in_pic[i+4*N])
plt.axis('off')
out_pic = to_img(out.cpu().data)
plt.figure(figsize=(18, 6))
for i in range(4):
plt.subplot(1,4,i+1)
plt.imshow(out_pic[i+4*N])
plt.axis('off')
# Define data loading step
batch_size = 256
img_transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
])
dataset = MNIST('./data', transform=img_transform, download=True)
dataloader = DataLoader(dataset, batch_size=batch_size, shuffle=True)
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# Define model architecture and reconstruction loss
# n = 28 x 28 = 784
d = 30 # for standard AE (under-complete hidden layer)
# d = 500 # for denoising AE (over-complete hidden layer)
class Autoencoder(nn.Module):
def __init__(self):
super().__init__()
self.encoder = nn.Sequential(
nn.Linear(28 * 28, d),
nn.Tanh(),
)
self.decoder = nn.Sequential(
nn.Linear(d, 28 * 28),
nn.Tanh(),
)
def forward(self, x):
x = self.encoder(x)
x = self.decoder(x)
return x
model = Autoencoder().to(device)
criterion = nn.MSELoss()
# Configure the optimiser
learning_rate = 1e-3
optimizer = torch.optim.Adam(
model.parameters(),
lr=learning_rate,
)
```
*Comment* or *un-comment out* a few lines of code to seamlessly switch between *standard AE* and *denoising one*.
Don't forget to **(1)** change the size of the hidden layer accordingly, **(2)** re-generate the model, and **(3)** re-pass the parameters to the optimiser.
```
# Train standard or denoising autoencoder (AE)
num_epochs = 1
# do = nn.Dropout() # comment out for standard AE
for epoch in range(num_epochs):
for data in dataloader:
img, _ = data
img.requires_grad_()
img = img.view(img.size(0), -1)
# img_bad = do(img).to(device) # comment out for standard AE
# ===================forward=====================
output = model(img) # feed <img> (for std AE) or <img_bad> (for denoising AE)
loss = criterion(output, img.data)
# ===================backward====================
optimizer.zero_grad()
loss.backward()
optimizer.step()
# ===================log========================
print(f'epoch [{epoch + 1}/{num_epochs}], loss:{loss.item():.4f}')
display_images(None, output) # pass (None, output) for std AE, (img_bad, output) for denoising AE
# Visualise a few kernels of the encoder
display_images(None, model.encoder[0].weight, 5)
! conda install -y --name codas-ml opencv
# Let's compare the autoencoder inpainting capabilities vs. OpenCV
from cv2 import inpaint, INPAINT_NS, INPAINT_TELEA
# Inpaint with Telea and Navier-Stokes methods
dst_TELEA = list()
dst_NS = list()
for i in range(3, 7):
corrupted_img = ((img_bad.data.cpu()[i].view(28, 28) / 4 + 0.5) * 255).byte().numpy()
mask = 2 - img_bad.grad_fn.noise.cpu()[i].view(28, 28).byte().numpy()
dst_TELEA.append(inpaint(corrupted_img, mask, 3, INPAINT_TELEA))
dst_NS.append(inpaint(corrupted_img, mask, 3, INPAINT_NS))
tns_TELEA = [torch.from_numpy(d) for d in dst_TELEA]
tns_NS = [torch.from_numpy(d) for d in dst_NS]
TELEA = torch.stack(tns_TELEA).float()
NS = torch.stack(tns_NS).float()
# Compare the results: [noise], [img + noise], [img], [AE, Telea, Navier-Stokes] inpainting
with torch.no_grad():
display_images(img_bad.grad_fn.noise[3:7], img_bad[3:7])
display_images(img[3:7], output[3:7])
display_images(TELEA, NS)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
#import matplotlib.pylab as plt
import matplotlib.pyplot as plt
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import silhouette_score
from sklearn import cluster
from sklearn.cluster import KMeans
from sklearn.datasets import make_blobs
import seaborn as sns
sns.set()
from sklearn.neighbors import NearestNeighbors
from yellowbrick.cluster import KElbowVisualizer
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
from sklearn.model_selection import train_test_split
from mpl_toolkits.mplot3d import Axes3D
from sklearn.metrics import accuracy_score
%matplotlib inline
plt.rcParams['figure.figsize'] = (16, 9)
plt.style.use('ggplot')
```
## Se visualiza los datos y se elimina las columnas que no son necesarias
```
dfRead = pd.read_csv('Interaccion_todasLasSesiones.csv')
df = dfRead.drop(['Sesion','Id'], axis=1)
#df = df[df['Fsm']!=0]
```
## Filtrado de datos
## Histograma de las notas
```
plt.rcParams['figure.figsize'] = (16, 9)
plt.style.use('ggplot')
datos = df.drop(['Nota'],1).hist()
plt.grid(True)
plt.show()
```
## Se crean los datos para el clusters y las categorias
```
clusters = df[['Nota']]
X = df.drop(['Nota'],1)
## Se reliza la normalización de los datos para que esten en un rango de (0,1)
scaler = MinMaxScaler(feature_range=(0, 1))
x = scaler.fit_transform(X)
```
## Se definen los metodos a emplear en el cluster
```
def clusterDBscan(x):
db = cluster.DBSCAN(eps=0.25, min_samples=5)
db.fit(x)
return db.labels_
def clusterKMeans(x, n_clusters):
return cluster.k_means(x, n_clusters=n_clusters)[1]
```
## Se crea funciones en caso de ser necesarias para poder reducir las dimensiones
```
def reducir_dim(x, ndim):
pca = PCA(n_components=ndim)
return pca.fit_transform(x)
def reducir_dim_tsne(x, ndim):
pca = TSNE(n_components=ndim)
return pca.fit_transform(x)
```
## Se grafica los valores de los posibles cluster en base a silohuette score
```
def calculaSilhoutter(x, clusters):
res=[]
fig, ax = plt.subplots(1,figsize=(20, 5))
for numCluster in range(2, 7):
res.append(silhouette_score(x, clusterKMeans(x,numCluster )))
ax.plot(range(2, 7), res)
ax.set_xlabel("n clusters")
ax.set_ylabel("silouhette score")
ax.set_title("K-Means")
calculaSilhoutter(x, clusters)
```
## Se grafica los valores de los posibles cluster en base a Elbow Method
```
model = KMeans()
visualizer = KElbowVisualizer(model, k=(2,7), metric='calinski_harabasz', timings=False)
visualizer.fit(x) # Fit the data to the visualizer
visualizer.show()
clus_km = clusterKMeans(x, 3)
clus_db = clusterDBscan(x)
def reducir_dataset(x, how):
if how == "pca":
res = reducir_dim(x, ndim=2)
elif how == "tsne":
res = reducir_dim_tsne(x, ndim=2)
else:
return x[:, :2]
return res
results = pd.DataFrame(np.column_stack([reducir_dataset(x, how="tsne"), clusters, clus_km, clus_db]), columns=["x", "y", "clusters", "clus_km", "clus_db"])
def mostrar_resultados(res):
"""Muestra los resultados de los algoritmos
"""
fig, ax = plt.subplots(1, 3, figsize=(20, 5))
sns.scatterplot(data=res, x="x", y="y", hue="clusters", ax=ax[0], legend="full")
ax[0].set_title('Ground Truth')
sns.scatterplot(data=res, x="x", y="y", hue="clus_km", ax=ax[1], legend="full")
ax[1].set_title('K-Means')
sns.scatterplot(data=res, x="x", y="y", hue="clus_db", ax=ax[2], legend="full")
ax[2].set_title('DBSCAN')
mostrar_resultados(results)
kmeans = KMeans(n_clusters=3,init = "k-means++")
kmeans.fit(x)
labels = kmeans.predict(x)
X['Cluster_Km']=labels
dfRead['Cluster_Km']=labels
X.groupby('Cluster_Km').mean()
```
## DBSCAN
```
neigh = NearestNeighbors(n_neighbors=2)
nbrs = neigh.fit(x)
distances, indices = nbrs.kneighbors(x)
distances = np.sort(distances, axis=0)
distancias = distances[:,1]
plt.plot(distancias)
plt.ylim(0, 0.4)
dbscan = cluster.DBSCAN(eps=0.25, min_samples=5)
dbscan.fit(x)
clusterDbscan = dbscan.labels_
X['Cluster_DB']=clusterDbscan
dfRead['Cluster_DB']=clusterDbscan
X.groupby('Cluster_DB').mean()
dfRead
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import treelib
from pathlib import Path
from treelib import Node, Tree
DATA_DIR = Path('../../data/retail-rocket')
EXPORT_DIR = Path('../../data/retail-rocket') / 'saved'
PATH_CATEGORY_TREE = DATA_DIR / 'category_tree.csv'
PATH_EVENTS = DATA_DIR /'events.csv'
PATH_ITEM_PROPS1 = DATA_DIR / 'item_properties_part1.csv'
PATH_ITEM_PROPS2 = DATA_DIR / 'item_properties_part2.csv'
```
# Creating Category Features via Tree
The category tree provided is given as a table of edges. We want to be able to get all the levels given a leaf node.
```
cat_tree_df = pd.read_csv(PATH_CATEGORY_TREE)
cat_tree_df.head()
tree = Tree()
ROOT = 'cat_tree'
tree.create_node(identifier=ROOT)
tree.create_node(identifier=-1, parent=ROOT) # temp
for _, row in cat_tree_df.iterrows():
categoryid, parentid = row
if np.isnan(parentid):
parentid = ROOT
else:
parentid = int(parentid)
categoryid = int(categoryid)
if not tree.contains(parentid):
tree.create_node(identifier=parentid, parent=-1)
if not tree.contains(categoryid):
tree.create_node(identifier=categoryid, parent=parentid)
else:
if tree.get_node(categoryid).bpointer == -1:
tree.move_node(categoryid, parentid)
tree.link_past_node(-1)
# Print the tree structure
# tree.show(line_type='ascii-em')
```
# Item Properties
We are provided with a bunch of item properties that can possibly change over time. But we will only be working with `categoryid` (and the latest record of it).
```
item_props_df = pd.concat([
pd.read_csv(PATH_ITEM_PROPS1, usecols=['itemid', 'property', 'value']),
pd.read_csv(PATH_ITEM_PROPS2, usecols=['itemid', 'property', 'value']),
])
item_props_df = item_props_df.loc[item_props_df['property']=='categoryid']\
.drop_duplicates().drop('property', axis=1).set_index('itemid')
item_props_df.columns = ['categoryid']
item_props_df['categoryid'] = item_props_df['categoryid'].astype(np.uint16)
# Could memoize if we wanted, meh
def get_cats(categoryid):
try:
return list(tree.rsearch(categoryid))[::-1][1:]
except treelib.exceptions.NodeIDAbsentError:
return []
item_categories_df = pd.DataFrame(item_props_df['categoryid'].map(get_cats).tolist())
item_categories_df.columns = [f'categoryid_lvl{i}' for i in range(6)]
item_categories_df.index = item_props_df.index
item_categories_df.reset_index(inplace=True)
# lvl3-5 are mostly NaN, probably want to chop them off
item_categories_df.to_msgpack(EXPORT_DIR / 'item_categories.msg')
item_categories_df.head()
```
# Events
Pre-split our event facts.
```
HOLDOUT_DATE = '2015-09-01'
events_df = pd.read_csv(PATH_EVENTS, usecols=['timestamp', 'visitorid', 'event', 'itemid'])
events_df['timestamp'] = pd.to_datetime(events_df['timestamp'], unit='ms')
events_df.to_msgpack(EXPORT_DIR / 'events.msg')
events_df.loc[events_df['timestamp'] < HOLDOUT_DATE].to_msgpack(EXPORT_DIR / 'events_tsplit.msg')
events_df.loc[events_df['timestamp'] >= HOLDOUT_DATE].to_msgpack(EXPORT_DIR / 'events_vsplit.msg')
```
| github_jupyter |
# VQGAN+CLIP Simplificado
```
# Licensed under the MIT License
# Copyright (c) 2021 Katherine Crowson
# Permission is hereby granted, free of charge, to any person obtaining a copy
# of this software and associated documentation files (the "Software"), to deal
# in the Software without restriction, including without limitation the rights
# to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
# copies of the Software, and to permit persons to whom the Software is
# furnished to do so, subject to the following conditions:
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
# AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
# OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN
# THE SOFTWARE.
```
## Docker Volumenes
```
from extra import *
MODELS_FOLDER = Path("/tf/models")
OUTPUTS_FOLDER= Path("/tf/outputs")
SRC_FOLDER= Path("/tf/outputs/src")
```
## Descargar modelos pre-entrenados:
```
from get_models import list_models, download_models
models = list_models(return_list=True)
print(models)
download_models(models, output=MODELS_FOLDER)
# Selecciona un modelo
model = models[1]
if (Path(MODELS_FOLDER)/ f"{model}.yaml").exists():
print(f"{model} ya existe.")
else:
print(f"Descargando: {model}")
download_models(model, output=MODELS_FOLDER)
```
## Configuración
**Definir parámetros principales:**
```
#!ls $MODELS_FOLDER
#Texto usado como Input para generar la imagen:
txt = 'Todo va a salir bien'
prompts = [txt]
# Tamaño de imagen
#size = (200, 200) #6129MiB de V-RAM
size = (340, 340) #8693MiB de V-RAM
#size = (450, 450) #10201MiB de V-RAM
# Numero de iteraciones en la creación de la imagen, a mayor el número de iteraciones mejor es el detalle en la imagen
iterations = 250
# Modelo a usar
# modelos descargados:
# !ls $MODELS_FOLDER
model = 'vqgan_imagenet_f16_1024'
```
**Definir parámetros secundarios:**
```
config = {
"init_image": None,
"seed": None,
"step_size": 0.2,
"cutn": 64,
"cut_pow": 1.,
"display_freq": 5,
"image_prompts": [],
"noise_prompt_seeds": [],
"noise_prompt_weights": [],
"init_weight": 0.,
"clip_model": 'ViT-B/32',
}
# SOBREESCRIBIR RESULTADOS
overwrite=True
```
## Generar frames del proceso resultante:
```
from generate_images import *
last_frame_index = generate_images(
prompts=prompts,
model=model,
outputs_folder=OUTPUTS_FOLDER,
models_folder=MODELS_FOLDER,
iterations=280,
**config,
overwrite=overwrite
)
experiment_name = to_experiment_name(prompts)
experiment_folder = Path(OUTPUTS_FOLDER) / experiment_name
create_video(last_frame_index, Path(experiment_folder))
!cp $experiment_folder/video.mp4 temp.mp4
from IPython.display import Video
video_file = "temp.mp4"
Video(video_file)
!nvidia-smi
# Liberar memoria de la GPU
reset_kernel()
```
---
Notebook Original: https://colab.research.google.com/drive/1L8oL-vLJXVcRzCFbPwOoMkPKJ8-aYdPN
| github_jupyter |
<a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_10_3_text_generation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# T81-558: Applications of Deep Neural Networks
**Module 10: Time Series in Keras**
* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
* For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# Module 10 Material
* Part 10.1: Time Series Data Encoding for Deep Learning [[Video]](https://www.youtube.com/watch?v=dMUmHsktl04&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_10_1_timeseries.ipynb)
* Part 10.2: Programming LSTM with Keras and TensorFlow [[Video]](https://www.youtube.com/watch?v=wY0dyFgNCgY&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_10_2_lstm.ipynb)
* **Part 10.3: Text Generation with Keras and TensorFlow** [[Video]](https://www.youtube.com/watch?v=6ORnRAz3gnA&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_10_3_text_generation.ipynb)
* Part 10.4: Image Captioning with Keras and TensorFlow [[Video]](https://www.youtube.com/watch?v=NmoW_AYWkb4&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_10_4_captioning.ipynb)
* Part 10.5: Temporal CNN in Keras and TensorFlow [[Video]](https://www.youtube.com/watch?v=i390g8acZwk&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_10_5_temporal_cnn.ipynb)
# Google CoLab Instructions
The following code ensures that Google CoLab is running the correct version of TensorFlow.
```
try:
%tensorflow_version 2.x
COLAB = True
print("Note: using Google CoLab")
except:
print("Note: not using Google CoLab")
COLAB = False
```
# Part 10.3: Text Generation with LSTM
Recurrent neural networks are also known for their ability to generate text. As a result, the output of the neural network can be free-form text. In this section, we will see how to train an LSTM can on a textual document, such as classic literature, and learn to output new text that appears to be of the same form as the training material. If you train your LSTM on [Shakespeare](https://en.wikipedia.org/wiki/William_Shakespeare), it will learn to crank out new prose similar to what Shakespeare had written.
Don't get your hopes up. You are not going to teach your deep neural network to write the next [Pulitzer Prize for Fiction](https://en.wikipedia.org/wiki/Pulitzer_Prize_for_Fiction). The prose generated by your neural network will be nonsensical. However, it will usually be nearly grammatically and of a similar style as the source training documents.
A neural network generating nonsensical text based on literature may not seem useful at first glance. However, this technology gets so much interest because it forms the foundation for many more advanced technologies. The fact that the LSTM will typically learn human grammar from the source document opens a wide range of possibilities. You can use similar technology to complete sentences when a user is entering text. Simply the ability to output free-form text becomes the foundation of many other technologies. In the next part, we will use this technique to create a neural network that can write captions for images to describe what is going on in the picture.
### Additional Information
The following are some of the articles that I found useful in putting this section together.
* [The Unreasonable Effectiveness of Recurrent Neural Networks](http://karpathy.github.io/2015/05/21/rnn-effectiveness/)
* [Keras LSTM Generation Example](https://keras.io/examples/lstm_text_generation/)
### Character-Level Text Generation
There are several different approaches to teaching a neural network to output free-form text. The most basic question is if you wish the neural network to learn at the word or character level. In many ways, learning at the character level is the more interesting of the two. The LSTM is learning to construct its own words without even being shown what a word is. We will begin with character-level text generation. In the next module, we will see how we can use nearly the same technique to operate at the word level. We will implement word-level automatic captioning in the next module.
We begin by importing the needed Python packages and defining the sequence length, named **maxlen**. Time-series neural networks always accept their input as a fixed-length array. Because you might not use all of the sequence elements, it is common to fill extra elements with zeros. You will divide the text into sequences of this length, and the neural network will train to predict what comes after this sequence.
```
from tensorflow.keras.callbacks import LambdaCallback
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import LSTM
from tensorflow.keras.optimizers import RMSprop
from tensorflow.keras.utils import get_file
import numpy as np
import random
import sys
import io
import requests
import re
```
For this simple example, we will train the neural network on the classic children's book [Treasure Island](https://en.wikipedia.org/wiki/Treasure_Island). We begin by loading this text into a Python string and displaying the first 1,000 characters.
```
r = requests.get("https://data.heatonresearch.com/data/t81-558/text/"\
"treasure_island.txt")
raw_text = r.text
print(raw_text[0:1000])
```
We will extract all unique characters from the text and sort them. This technique allows us to assign a unique ID to each character. Because we sorted the characters, these IDs should remain the same. If we add new characters to the original text, then the IDs would change. We build two dictionaries. The first **char2idx** is used to convert a character into its ID. The second **idx2char** converts an ID back into its character.
```
processed_text = raw_text.lower()
processed_text = re.sub(r'[^\x00-\x7f]',r'', processed_text)
print('corpus length:', len(processed_text))
chars = sorted(list(set(processed_text)))
print('total chars:', len(chars))
char_indices = dict((c, i) for i, c in enumerate(chars))
indices_char = dict((i, c) for i, c in enumerate(chars))
```
We are now ready to build the actual sequences. Just like previous neural networks, there will be an $x$ and $y$. However, for the LSTM, $x$ and $y$ will both be sequences. The $x$ input will specify the sequences where $y$ are the expected output. The following code generates all possible sequences.
```
# cut the text in semi-redundant sequences of maxlen characters
maxlen = 40
step = 3
sentences = []
next_chars = []
for i in range(0, len(processed_text) - maxlen, step):
sentences.append(processed_text[i: i + maxlen])
next_chars.append(processed_text[i + maxlen])
print('nb sequences:', len(sentences))
sentences
print('Vectorization...')
x = np.zeros((len(sentences), maxlen, len(chars)), dtype=np.bool)
y = np.zeros((len(sentences), len(chars)), dtype=np.bool)
for i, sentence in enumerate(sentences):
for t, char in enumerate(sentence):
x[i, t, char_indices[char]] = 1
y[i, char_indices[next_chars[i]]] = 1
x.shape
y.shape
```
The dummy variables for $y$ are shown below.
```
y[0:10]
```
Next, we create the neural network. This neural network's primary feature is the LSTM layer, which allows the sequences to be processed.
```
# build the model: a single LSTM
print('Build model...')
model = Sequential()
model.add(LSTM(128, input_shape=(maxlen, len(chars))))
model.add(Dense(len(chars), activation='softmax'))
optimizer = RMSprop(lr=0.01)
model.compile(loss='categorical_crossentropy', optimizer=optimizer)
model.summary()
```
The LSTM will produce new text character by character. We will need to sample the correct letter from the LSTM predictions each time. The **sample** function accepts the following two parameters:
* **preds** - The output neurons.
* **temperature** - 1.0 is the most conservative, 0.0 is the most confident (willing to make spelling and other errors).
The sample function below is essentially performing a [softmax]() on the neural network predictions. This causes each output neuron to become a probability of its particular letter.
```
def sample(preds, temperature=1.0):
# helper function to sample an index from a probability array
preds = np.asarray(preds).astype('float64')
preds = np.log(preds) / temperature
exp_preds = np.exp(preds)
preds = exp_preds / np.sum(exp_preds)
probas = np.random.multinomial(1, preds, 1)
return np.argmax(probas)
```
Keras calls the following function at the end of each training Epoch. The code generates sample text generations that visually demonstrate the neural network better at text generation. As the neural network trains, the generations should look more realistic.
```
def on_epoch_end(epoch, _):
# Function invoked at end of each epoch. Prints generated text.
print("******************************************************")
print('----- Generating text after Epoch: %d' % epoch)
start_index = random.randint(0, len(processed_text) - maxlen - 1)
for temperature in [0.2, 0.5, 1.0, 1.2]:
print('----- temperature:', temperature)
generated = ''
sentence = processed_text[start_index: start_index + maxlen]
generated += sentence
print('----- Generating with seed: "' + sentence + '"')
sys.stdout.write(generated)
for i in range(400):
x_pred = np.zeros((1, maxlen, len(chars)))
for t, char in enumerate(sentence):
x_pred[0, t, char_indices[char]] = 1.
preds = model.predict(x_pred, verbose=0)[0]
next_index = sample(preds, temperature)
next_char = indices_char[next_index]
generated += next_char
sentence = sentence[1:] + next_char
sys.stdout.write(next_char)
sys.stdout.flush()
print()
```
We are now ready to train. It can take up to an hour to train this network, depending on how fast your computer is. If you have a GPU available, please make sure to use it.
```
# Ignore useless W0819 warnings generated by TensorFlow 2.0. Hopefully can remove this ignore in the future.
# See https://github.com/tensorflow/tensorflow/issues/31308
import logging, os
logging.disable(logging.WARNING)
os.environ["TF_CPP_MIN_LOG_LEVEL"] = "3"
# Fit the model
print_callback = LambdaCallback(on_epoch_end=on_epoch_end)
model.fit(x, y,
batch_size=128,
epochs=60,
callbacks=[print_callback])
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import pickle
import json
import gensim
import os
import re
from sklearn.model_selection import train_test_split
from pandas.plotting import scatter_matrix
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.preprocessing import sequence
from keras.optimizers import RMSprop, SGD
from keras.models import Sequential, Model
from keras.layers.core import Dense, Dropout, Activation, Flatten, Reshape
from keras.layers import Input, Bidirectional, LSTM, regularizers
from keras.layers.embeddings import Embedding
from keras.layers.convolutional import Conv1D, MaxPooling1D, MaxPooling2D, Conv2D
from keras.layers.normalization import BatchNormalization
from keras.callbacks import EarlyStopping
%matplotlib inline
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
# Change these to match your file paths :)
filename = '../wyns/data/tweet_global_warming.csv' #64,706 reviews
model_path = "GoogleNews-vectors-negative300.bin"
word_vector_model = gensim.models.KeyedVectors.load_word2vec_format(model_path, binary=True)
def normalize(txt, vocab=None, replace_char=' ',
max_length=300, pad_out=False,
to_lower=True, reverse = False,
truncate_left=False, encoding=None,
letters_only=False):
txt = txt.split()
# Remove HTML
# This will keep characters and other symbols
txt = [re.sub(r'http:.*', '', r) for r in txt]
txt = [re.sub(r'https:.*', '', r) for r in txt]
txt = ( " ".join(txt))
# Remove non-emoticon punctuation and numbers
txt = re.sub("[.,!0-9]", " ", txt)
if letters_only:
txt = re.sub("[^a-zA-Z]", " ", txt)
txt = " ".join(txt.split())
# store length for multiple comparisons
txt_len = len(txt)
if truncate_left:
txt = txt[-max_length:]
else:
txt = txt[:max_length]
# change case
if to_lower:
txt = txt.lower()
# Reverse order
if reverse:
txt = txt[::-1]
# replace chars
if vocab is not None:
txt = ''.join([c if c in vocab else replace_char for c in txt])
# re-encode text
if encoding is not None:
txt = txt.encode(encoding, errors="ignore")
# pad out if needed
if pad_out and max_length>txt_len:
txt = txt + replace_char * (max_length - txt_len)
if txt.find('@') > -1:
print(len(txt.split('@'))-1)
print(txt.split('@'))
for i in range(len(txt.split('@'))-1):
try:
if str(txt.split('@')[1]).find(' ') > -1:
to_remove = '@' + str(txt.split('@')[1].split(' ')[0]) + " "
else:
to_remove = '@' + str(txt.split('@')[1])
print(to_remove)
txt = txt.replace(to_remove,'')
except:
pass
return txt
# What does this normalization function look like?
clean_text = normalize("This is A sentence. @sarahisabutthead with @sarah things! 123 :) and a link https://gitub.com @blah")
print(clean_text)
string = '@dez_blanchfield @SpEducatorCWSN @VolcanoScouting @USGSVolcanoes @Volcanoes_NPS @TmanSpeaks @DioFavatas @helene_wpli @ScheuerJo @martinfredras @HelenClarkNZ @dez_blanchfield, can you explain your answer? If oceans are rising, and getting heavier, why won�t this increased weight have consequences? See: https://t.co/DupaCkMnIE'
print(string)
print(normalize(string))
```
We want to balance the distrubtion of sentiment:
```
def balance(df):
print("Balancing the classes")
type_counts = df['Sentiment'].value_counts()
min_count = min(type_counts.values)
balanced_df = None
for key in type_counts.keys():
df_sub = df[df['Sentiment']==key].sample(n=min_count, replace=False)
if balanced_df is not None:
balanced_df = balanced_df.append(df_sub)
else:
balanced_df = df_sub
return balanced_df
def review_to_sentiment(review):
# Review is coming in as Y/N/NaN
# this then cleans the summary and review and gives it a positive or negative value
norm_text = normalize(review[0])
if review[1] in ('Yes', 'Y'):
return ['positive', norm_text]
elif review[1] in ('No', 'N'):
return ['negative', norm_text]
else:
return ['other', norm_text]
data = []
with open(filename, 'r', encoding='latin') as f:
for i,line in enumerate(f):
if i == 0: #skip header while i diagnose
continue
# as we read in, clean
line_data = line.split(",")
data.append(review_to_sentiment(line_data))
twitter = pd.DataFrame(data, columns=['Sentiment', 'clean_text'], dtype=str)
# print(twitter)
# For this demo lets just keep one and five stars the others are marked 'other
# twitter = twitter[twitter['Sentiment'].isin(['positive', 'negative'])]
# twitter.head()
# balanced_twitter = balance(twitter)
# len(balanced_twitter)
# Now go from the pandas into lists of text and labels
text = twitter['clean_text'].values
labels_0 = pd.get_dummies(twitter['Sentiment']) # mapping of the labels with dummies (has headers)
labels = labels_0.values # removes the headers
labels[:10] # negative, other, positive
labels = labels[:,[0,2]]
labels[:10] # negative, positive
# Perform the Train/test split
X_train_, X_test_, Y_train_, Y_test_ = train_test_split(text,labels, test_size = 0.2, random_state = 42)
max_fatures = 2000
max_len = 40
batch_size = 32
embed_dim = 300
lstm_out = 140
dense_out=len(labels[0])
tokenizer = Tokenizer(num_words=max_fatures, split=' ')
tokenizer.fit_on_texts(X_train_)
X_train = tokenizer.texts_to_sequences(X_train_)
X_train = pad_sequences(X_train, maxlen=max_len, padding='post')
X_test = tokenizer.texts_to_sequences(X_test_)
X_test = pad_sequences(X_test, maxlen=max_len, padding='post')
word_index = tokenizer.word_index
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.multioutput import MultiOutputClassifier
gb = GradientBoostingClassifier(n_estimators = 4000)
gb = MultiOutputClassifier(gb, n_jobs=2)
gb.fit(X_train,Y_train_)
print(gb.score(X_test,Y_test_))
blah = 'look \n at \n me \n go'
blah2 = blah.replace
print(blah)
# What does the data look like?
# It is a one-hot encoding of the label, either positive or negative
Y_train_[:5]
X_train_[42]
### Now for a simple bidirectional LSTM algorithm we set our feature sizes and train a tokenizer
# First we Tokenize and get the data into a form that the model can read - this is BoW
# In this cell we are also going to define some of our hyperparameters
max_fatures = 2000
max_len = 40
batch_size = 32
embed_dim = 300
lstm_out = 140
dense_out=len(labels[0]) #length of features
tokenizer = Tokenizer(num_words=max_fatures, split=' ')
tokenizer.fit_on_texts(X_train_)
X_train = tokenizer.texts_to_sequences(X_train_)
X_train = pad_sequences(X_train, maxlen=max_len, padding='post')
X_test = tokenizer.texts_to_sequences(X_test_)
X_test = pad_sequences(X_test, maxlen=max_len, padding='post')
word_index = tokenizer.word_index
# Now what does our data look like?
# Tokenizer creates a BOW encoding, which is then going to be fed into our Embedding matrix
# This will be used by the model to build up a word embedding
X_test[:,-1].mean()
# What does a word vector look like?
# Ahhhh, like a bunch of numbers
word_vector_model.word_vec('hello')
print('Prepare the embedding matrix')
# prepare embedding matrix
num_words = min(max_fatures, len(word_index))
embedding_matrix = np.zeros((num_words, embed_dim))
for word, i in word_index.items():
if i >= max_len:
continue
# words not found in embedding index will be all-zeros.
if word in word_vector_model.vocab:
embedding_matrix[i] = word_vector_model.word_vec(word)
# load pre-trained word embeddings into an Embedding layer
# note that we set trainable = True to fine tune the embeddings
embedding_layer = Embedding(num_words,
embed_dim,
weights=[embedding_matrix],
input_length=max_fatures,
trainable=True)
embedding_matrix[1]
# Define the model using the pre-trained embedding
# import tensorflow as tf
# with tf.device('/cpu:0'):
embedding_layer = Embedding(num_words,
embed_dim,
weights=[embedding_matrix],
input_length=max_fatures,
trainable=True)
sequence_input = Input(shape=(max_len,), dtype='int32')
embedded_sequences = embedding_layer(sequence_input)
x = Bidirectional(LSTM(lstm_out, recurrent_dropout=0.5, activation='tanh'))(embedded_sequences)
# preds = Dense(250, activation='softmax')(x)
preds = Dense(dense_out, activation='softmax')(x)
model = Model(sequence_input, preds)
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['acc'])
print(model.summary())
model_hist_embedding = model.fit(X_train, Y_train_, epochs = 80, batch_size=batch_size, verbose = 2,
validation_data=(X_test,Y_test_))
model_hist_embedding = model.fit(X_train, Y_train_, epochs = 80, batch_size=batch_size, verbose = 2,
validation_data=(X_test,Y_test_))
# Training Accuracy
x = np.arange(20)+1
plt.plot(x, model_hist_embedding.history['acc'])
plt.plot(x, model_hist_embedding.history['val_acc'])
plt.legend(['Training', 'Testing'], loc='lower right')
plt.ylabel("Accuracy")
axes = plt.gca()
axes.set_ylim([0.45,1.01])
plt.xlabel("Epoch")
plt.title("LSTM Accuracy")
plt.show()
#model_hist_embedding.model.save("../../wyns/data/climate_sentiment_m2.h5")
```
| github_jupyter |
# Statistical Data Modeling
Some or most of you have probably taken some undergraduate- or graduate-level statistics courses. Unfortunately, the curricula for most introductory statisics courses are mostly focused on conducting **statistical hypothesis tests** as the primary means for interest: t-tests, chi-squared tests, analysis of variance, etc. Such tests seek to esimate whether groups or effects are "statistically significant", a concept that is poorly understood, and hence often misused, by most practioners. Even when interpreted correctly, statistical significance is a questionable goal for statistical inference, as it is of limited utility.
A far more powerful approach to statistical analysis involves building flexible **models** with the overarching aim of *estimating* quantities of interest. This section of the tutorial illustrates how to use Python to build statistical models of low to moderate difficulty from scratch, and use them to extract estimates and associated measures of uncertainty.
```
%matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# Set some Pandas options
pd.set_option('display.notebook_repr_html', False)
pd.set_option('display.max_columns', 20)
pd.set_option('display.max_rows', 25)
```
Estimation
==========
An recurring statistical problem is finding estimates of the relevant parameters that correspond to the distribution that best represents our data.
In **parametric** inference, we specify *a priori* a suitable distribution, then choose the parameters that best fit the data.
* e.g. $\mu$ and $\sigma^2$ in the case of the normal distribution
```
x = np.array([ 1.00201077, 1.58251956, 0.94515919, 6.48778002, 1.47764604,
5.18847071, 4.21988095, 2.85971522, 3.40044437, 3.74907745,
1.18065796, 3.74748775, 3.27328568, 3.19374927, 8.0726155 ,
0.90326139, 2.34460034, 2.14199217, 3.27446744, 3.58872357,
1.20611533, 2.16594393, 5.56610242, 4.66479977, 2.3573932 ])
_ = plt.hist(x, bins=8)
```
### Fitting data to probability distributions
We start with the problem of finding values for the parameters that provide the best fit between the model and the data, called point estimates. First, we need to define what we mean by ‘best fit’. There are two commonly used criteria:
* **Method of moments** chooses the parameters so that the sample moments (typically the sample mean and variance) match the theoretical moments of our chosen distribution.
* **Maximum likelihood** chooses the parameters to maximize the likelihood, which measures how likely it is to observe our given sample.
### Discrete Random Variables
$$X = \{0,1\}$$
$$Y = \{\ldots,-2,-1,0,1,2,\ldots\}$$
**Probability Mass Function**:
For discrete $X$,
$$Pr(X=x) = f(x|\theta)$$

***e.g. Poisson distribution***
The Poisson distribution models unbounded counts:
<div style="font-size: 150%;">
$$Pr(X=x)=\frac{e^{-\lambda}\lambda^x}{x!}$$
</div>
* $X=\{0,1,2,\ldots\}$
* $\lambda > 0$
$$E(X) = \text{Var}(X) = \lambda$$
### Continuous Random Variables
$$X \in [0,1]$$
$$Y \in (-\infty, \infty)$$
**Probability Density Function**:
For continuous $X$,
$$Pr(x \le X \le x + dx) = f(x|\theta)dx \, \text{ as } \, dx \rightarrow 0$$

***e.g. normal distribution***
<div style="font-size: 150%;">
$$f(x) = \frac{1}{\sqrt{2\pi\sigma^2}}\exp\left[-\frac{(x-\mu)^2}{2\sigma^2}\right]$$
</div>
* $X \in \mathbf{R}$
* $\mu \in \mathbf{R}$
* $\sigma>0$
$$\begin{align}E(X) &= \mu \cr
\text{Var}(X) &= \sigma^2 \end{align}$$
### Example: Nashville Precipitation
The dataset `nashville_precip.txt` contains [NOAA precipitation data for Nashville measured since 1871](http://bit.ly/nasvhville_precip_data). The gamma distribution is often a good fit to aggregated rainfall data, and will be our candidate distribution in this case.
```
precip = pd.read_table("data/nashville_precip.txt", index_col=0, na_values='NA', delim_whitespace=True)
precip.head()
_ = precip.hist(sharex=True, sharey=True, grid=False)
plt.tight_layout()
```
The first step is recognixing what sort of distribution to fit our data to. A couple of observations:
1. The data are skewed, with a longer tail to the right than to the left
2. The data are positive-valued, since they are measuring rainfall
3. The data are continuous
There are a few possible choices, but one suitable alternative is the **gamma distribution**:
<div style="font-size: 150%;">
$$x \sim \text{Gamma}(\alpha, \beta) = \frac{\beta^{\alpha}x^{\alpha-1}e^{-\beta x}}{\Gamma(\alpha)}$$
</div>
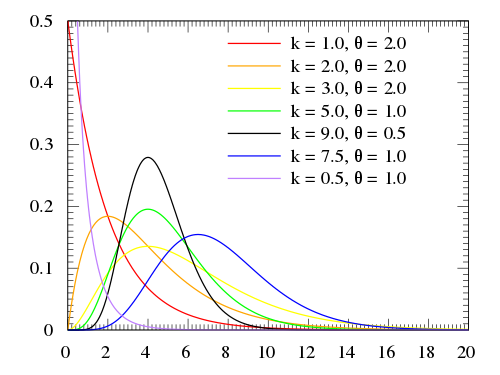
The ***method of moments*** simply assigns the empirical mean and variance to their theoretical counterparts, so that we can solve for the parameters.
So, for the gamma distribution, the mean and variance are:
<div style="font-size: 150%;">
$$ \hat{\mu} = \bar{X} = \alpha \beta $$
$$ \hat{\sigma}^2 = S^2 = \alpha \beta^2 $$
</div>
So, if we solve for these parameters, we can use a gamma distribution to describe our data:
<div style="font-size: 150%;">
$$ \alpha = \frac{\bar{X}^2}{S^2}, \, \beta = \frac{S^2}{\bar{X}} $$
</div>
Let's deal with the missing value in the October data. Given what we are trying to do, it is most sensible to fill in the missing value with the average of the available values.
```
precip.fillna(value={'Oct': precip.Oct.mean()}, inplace=True)
```
Now, let's calculate the sample moments of interest, the means and variances by month:
```
precip_mean = precip.mean()
precip_mean
precip_var = precip.var()
precip_var
```
We then use these moments to estimate $\alpha$ and $\beta$ for each month:
```
alpha_mom = precip_mean ** 2 / precip_var
beta_mom = precip_var / precip_mean
alpha_mom, beta_mom
```
We can use the `gamma.pdf` function in `scipy.stats.distributions` to plot the ditribtuions implied by the calculated alphas and betas. For example, here is January:
```
from scipy.stats.distributions import gamma
precip.Jan.hist(normed=True, bins=20)
plt.plot(np.linspace(0, 10), gamma.pdf(np.linspace(0, 10), alpha_mom[0], beta_mom[0]))
```
Looping over all months, we can create a grid of plots for the distribution of rainfall, using the gamma distribution:
```
axs = precip.hist(normed=True, figsize=(12, 8), sharex=True, sharey=True, bins=15, grid=False)
for ax in axs.ravel():
# Get month
m = ax.get_title()
# Plot fitted distribution
x = np.linspace(*ax.get_xlim())
ax.plot(x, gamma.pdf(x, alpha_mom[m], beta_mom[m]))
# Annotate with parameter estimates
label = 'alpha = {0:.2f}\nbeta = {1:.2f}'.format(alpha_mom[m], beta_mom[m])
ax.annotate(label, xy=(10, 0.2))
plt.tight_layout()
```
Maximum Likelihood
==================
**Maximum likelihood** (ML) fitting is usually more work than the method of moments, but it is preferred as the resulting estimator is known to have good theoretical properties.
There is a ton of theory regarding ML. We will restrict ourselves to the mechanics here.
Say we have some data $y = y_1,y_2,\ldots,y_n$ that is distributed according to some distribution:
<div style="font-size: 120%;">
$$Pr(Y_i=y_i | \theta)$$
</div>
Here, for example, is a **Poisson distribution** that describes the distribution of some discrete variables, typically *counts*:
```
y = np.random.poisson(5, size=100)
plt.hist(y, bins=12, normed=True)
plt.xlabel('y'); plt.ylabel('Pr(y)')
```
The product $\prod_{i=1}^n Pr(y_i | \theta)$ gives us a measure of how **likely** it is to observe values $y_1,\ldots,y_n$ given the parameters $\theta$. Maximum likelihood fitting consists of choosing the appropriate function $l= Pr(Y|\theta)$ to maximize for a given set of observations. We call this function the *likelihood function*, because it is a measure of how likely the observations are if the model is true.
> Given these data, how likely is this model?
In the above model, the data were drawn from a Poisson distribution with parameter $\lambda =5$.
$$L(y|\lambda=5) = \frac{e^{-5} 5^y}{y!}$$
So, for any given value of $y$, we can calculate its likelihood:
```
poisson_like = lambda x, lam: np.exp(-lam) * (lam**x) / (np.arange(x)+1).prod()
lam = 6
value = 10
poisson_like(value, lam)
np.sum(poisson_like(yi, lam) for yi in y)
lam = 8
np.sum(poisson_like(yi, lam) for yi in y)
```
We can plot the likelihood function for any value of the parameter(s):
```
lambdas = np.linspace(0,15)
x = 5
plt.plot(lambdas, [poisson_like(x, l) for l in lambdas])
plt.xlabel('$\lambda$')
plt.ylabel('L($\lambda$|x={0})'.format(x))
```
How is the likelihood function different than the probability distribution function (PDF)? The likelihood is a function of the parameter(s) *given the data*, whereas the PDF returns the probability of data given a particular parameter value. Here is the PDF of the Poisson for $\lambda=5$.
```
lam = 5
xvals = np.arange(15)
plt.bar(xvals, [poisson_like(x, lam) for x in xvals])
plt.xlabel('x')
plt.ylabel('Pr(X|$\lambda$=5)')
```
Why are we interested in the likelihood function?
A reasonable estimate of the true, unknown value for the parameter is one which **maximizes the likelihood function**. So, inference is reduced to an optimization problem.
Going back to the rainfall data, if we are using a gamma distribution we need to maximize:
$$\begin{align}l(\alpha,\beta) &= \sum_{i=1}^n \log[\beta^{\alpha} x^{\alpha-1} e^{-x/\beta}\Gamma(\alpha)^{-1}] \cr
&= n[(\alpha-1)\overline{\log(x)} - \bar{x}\beta + \alpha\log(\beta) - \log\Gamma(\alpha)]\end{align}$$
(*Its usually easier to work in the log scale*)
where $n = 2012 − 1871 = 141$ and the bar indicates an average over all *i*. We choose $\alpha$ and $\beta$ to maximize $l(\alpha,\beta)$.
Notice $l$ is infinite if any $x$ is zero. We do not have any zeros, but we do have an NA value for one of the October data, which we dealt with above.
### Finding the MLE
To find the maximum of any function, we typically take the *derivative* with respect to the variable to be maximized, set it to zero and solve for that variable.
$$\frac{\partial l(\alpha,\beta)}{\partial \beta} = n\left(\frac{\alpha}{\beta} - \bar{x}\right) = 0$$
Which can be solved as $\beta = \alpha/\bar{x}$. However, plugging this into the derivative with respect to $\alpha$ yields:
$$\frac{\partial l(\alpha,\beta)}{\partial \alpha} = \log(\alpha) + \overline{\log(x)} - \log(\bar{x}) - \frac{\Gamma(\alpha)'}{\Gamma(\alpha)} = 0$$
This has no closed form solution. We must use ***numerical optimization***!
Numerical optimization alogarithms take an initial "guess" at the solution, and iteratively improve the guess until it gets "close enough" to the answer.
Here, we will use Newton-Raphson algorithm:
<div style="font-size: 120%;">
$$x_{n+1} = x_n - \frac{f(x_n)}{f'(x_n)}$$
</div>
Which is available to us via SciPy:
```
from scipy.optimize import newton
```
Here is a graphical example of how Newtone-Raphson converges on a solution, using an arbitrary function:
```
# some function
func = lambda x: 3./(1 + 400*np.exp(-2*x)) - 1
xvals = np.linspace(0, 6)
plt.plot(xvals, func(xvals))
plt.text(5.3, 2.1, '$f(x)$', fontsize=16)
# zero line
plt.plot([0,6], [0,0], 'k-')
# value at step n
plt.plot([4,4], [0,func(4)], 'k:')
plt.text(4, -.2, '$x_n$', fontsize=16)
# tangent line
tanline = lambda x: -0.858 + 0.626*x
plt.plot(xvals, tanline(xvals), 'r--')
# point at step n+1
xprime = 0.858/0.626
plt.plot([xprime, xprime], [tanline(xprime), func(xprime)], 'k:')
plt.text(xprime+.1, -.2, '$x_{n+1}$', fontsize=16)
```
To apply the Newton-Raphson algorithm, we need a function that returns a vector containing the **first and second derivatives** of the function with respect to the variable of interest. In our case, this is:
```
from scipy.special import psi, polygamma
dlgamma = lambda m, log_mean, mean_log: np.log(m) - psi(m) - log_mean + mean_log
dl2gamma = lambda m, *args: 1./m - polygamma(1, m)
```
where `log_mean` and `mean_log` are $\log{\bar{x}}$ and $\overline{\log(x)}$, respectively. `psi` and `polygamma` are complex functions of the Gamma function that result when you take first and second derivatives of that function.
```
# Calculate statistics
log_mean = precip.mean().apply(np.log)
mean_log = precip.apply(np.log).mean()
```
Time to optimize!
```
# Alpha MLE for December
alpha_mle = newton(dlgamma, 2, dl2gamma, args=(log_mean[-1], mean_log[-1]))
alpha_mle
```
And now plug this back into the solution for beta:
<div style="font-size: 120%;">
$$ \beta = \frac{\alpha}{\bar{X}} $$
</div>
```
beta_mle = alpha_mle/precip.mean()[-1]
beta_mle
```
We can compare the fit of the estimates derived from MLE to those from the method of moments:
```
dec = precip.Dec
dec.hist(normed=True, bins=10, grid=False)
x = np.linspace(0, dec.max())
plt.plot(x, gamma.pdf(x, alpha_mom[-1], beta_mom[-1]), 'm-')
plt.plot(x, gamma.pdf(x, alpha_mle, beta_mle), 'r--')
```
For some common distributions, SciPy includes methods for fitting via MLE:
```
from scipy.stats import gamma
gamma.fit(precip.Dec)
```
This fit is not directly comparable to our estimates, however, because SciPy's `gamma.fit` method fits an odd 3-parameter version of the gamma distribution.
### Example: truncated distribution
Suppose that we observe $Y$ truncated below at $a$ (where $a$ is known). If $X$ is the distribution of our observation, then:
$$ P(X \le x) = P(Y \le x|Y \gt a) = \frac{P(a \lt Y \le x)}{P(Y \gt a)}$$
(so, $Y$ is the original variable and $X$ is the truncated variable)
Then X has the density:
$$f_X(x) = \frac{f_Y (x)}{1−F_Y (a)} \, \text{for} \, x \gt a$$
Suppose $Y \sim N(\mu, \sigma^2)$ and $x_1,\ldots,x_n$ are independent observations of $X$. We can use maximum likelihood to find $\mu$ and $\sigma$.
First, we can simulate a truncated distribution using a `while` statement to eliminate samples that are outside the support of the truncated distribution.
```
x = np.random.normal(size=10000)
a = -1
x_small = x < a
while x_small.sum():
x[x_small] = np.random.normal(size=x_small.sum())
x_small = x < a
_ = plt.hist(x, bins=100)
```
We can construct a log likelihood for this function using the conditional form:
$$f_X(x) = \frac{f_Y (x)}{1−F_Y (a)} \, \text{for} \, x \gt a$$
```
from scipy.stats.distributions import norm
trunc_norm = lambda theta, a, x: -(np.log(norm.pdf(x, theta[0], theta[1])) -
np.log(1 - norm.cdf(a, theta[0], theta[1]))).sum()
```
For this example, we will use another optimization algorithm, the **Nelder-Mead simplex algorithm**. It has a couple of advantages:
- it does not require derivatives
- it can optimize (minimize) a vector of parameters
SciPy implements this algorithm in its `fmin` function:
```
from scipy.optimize import fmin
fmin(trunc_norm, np.array([1,2]), args=(-1, x))
```
In general, simulating data is a terrific way of testing your model before using it with real data.
### Kernel density estimates
In some instances, we may not be interested in the parameters of a particular distribution of data, but just a smoothed representation of the data at hand. In this case, we can estimate the disribution *non-parametrically* (i.e. making no assumptions about the form of the underlying distribution) using kernel density estimation.
```
# Some random data
y = np.random.random(15) * 10
y
x = np.linspace(0, 10, 100)
# Smoothing parameter
s = 0.4
# Calculate the kernels
kernels = np.transpose([norm.pdf(x, yi, s) for yi in y])
plt.plot(x, kernels, 'k:')
plt.plot(x, kernels.sum(1))
plt.plot(y, np.zeros(len(y)), 'ro', ms=10)
```
SciPy implements a Gaussian KDE that automatically chooses an appropriate bandwidth. Let's create a bi-modal distribution of data that is not easily summarized by a parametric distribution:
```
# Create a bi-modal distribution with a mixture of Normals.
x1 = np.random.normal(0, 3, 50)
x2 = np.random.normal(4, 1, 50)
# Append by row
x = np.r_[x1, x2]
plt.hist(x, bins=8, normed=True)
from scipy.stats import kde
density = kde.gaussian_kde(x)
xgrid = np.linspace(x.min(), x.max(), 100)
plt.hist(x, bins=8, normed=True)
plt.plot(xgrid, density(xgrid), 'r-')
```
### Exercise: Cervical dystonia analysis
Recall the cervical dystonia database, which is a clinical trial of botulinum toxin type B (BotB) for patients with cervical dystonia from nine U.S. sites. The response variable is measurements on the Toronto Western Spasmodic Torticollis Rating Scale (TWSTRS), measuring severity, pain, and disability of cervical dystonia (high scores mean more impairment). One way to check the efficacy of the treatment is to compare the distribution of TWSTRS for control and treatment patients at the end of the study.
Use the method of moments or MLE to calculate the mean and variance of TWSTRS at week 16 for one of the treatments and the control group. Assume that the distribution of the `twstrs` variable is normal:
$$f(x \mid \mu, \sigma^2) = \sqrt{\frac{1}{2\pi\sigma^2}} \exp\left\{ -\frac{1}{2} \frac{(x-\mu)^2}{\sigma^2} \right\}$$
```
cdystonia = pd.read_csv("data/cdystonia.csv")
cdystonia[cdystonia.obs==6].hist(column='twstrs', by=cdystonia.treat, bins=8)
# Write your answer here
```
## Regression models
A general, primary goal of many statistical data analysis tasks is to relate the influence of one variable on another. For example, we may wish to know how different medical interventions influence the incidence or duration of disease, or perhaps a how baseball player's performance varies as a function of age.
```
x = np.array([2.2, 4.3, 5.1, 5.8, 6.4, 8.0])
y = np.array([0.4, 10.1, 14.0, 10.9, 15.4, 18.5])
plt.plot(x,y,'ro')
```
We can build a model to characterize the relationship between $X$ and $Y$, recognizing that additional factors other than $X$ (the ones we have measured or are interested in) may influence the response variable $Y$.
<div style="font-size: 150%;">
$y_i = f(x_i) + \epsilon_i$
</div>
where $f$ is some function, for example a linear function:
<div style="font-size: 150%;">
$y_i = \beta_0 + \beta_1 x_i + \epsilon_i$
</div>
and $\epsilon_i$ accounts for the difference between the observed response $y_i$ and its prediction from the model $\hat{y_i} = \beta_0 + \beta_1 x_i$. This is sometimes referred to as **process uncertainty**.
We would like to select $\beta_0, \beta_1$ so that the difference between the predictions and the observations is zero, but this is not usually possible. Instead, we choose a reasonable criterion: ***the smallest sum of the squared differences between $\hat{y}$ and $y$***.
<div style="font-size: 120%;">
$$R^2 = \sum_i (y_i - [\beta_0 + \beta_1 x_i])^2 = \sum_i \epsilon_i^2 $$
</div>
Squaring serves two purposes: (1) to prevent positive and negative values from cancelling each other out and (2) to strongly penalize large deviations. Whether the latter is a good thing or not depends on the goals of the analysis.
In other words, we will select the parameters that minimize the squared error of the model.
```
ss = lambda theta, x, y: np.sum((y - theta[0] - theta[1]*x) ** 2)
ss([0,1],x,y)
b0,b1 = fmin(ss, [0,1], args=(x,y))
b0,b1
plt.plot(x, y, 'ro')
plt.plot([0,10], [b0, b0+b1*10])
plt.plot(x, y, 'ro')
plt.plot([0,10], [b0, b0+b1*10])
for xi, yi in zip(x,y):
plt.plot([xi]*2, [yi, b0+b1*xi], 'k:')
plt.xlim(2, 9); plt.ylim(0, 20)
```
Minimizing the sum of squares is not the only criterion we can use; it is just a very popular (and successful) one. For example, we can try to minimize the sum of absolute differences:
```
sabs = lambda theta, x, y: np.sum(np.abs(y - theta[0] - theta[1]*x))
b0,b1 = fmin(sabs, [0,1], args=(x,y))
print b0,b1
plt.plot(x, y, 'ro')
plt.plot([0,10], [b0, b0+b1*10])
```
We are not restricted to a straight-line regression model; we can represent a curved relationship between our variables by introducing **polynomial** terms. For example, a cubic model:
<div style="font-size: 150%;">
$y_i = \beta_0 + \beta_1 x_i + \beta_2 x_i^2 + \epsilon_i$
</div>
```
ss2 = lambda theta, x, y: np.sum((y - theta[0] - theta[1]*x - theta[2]*(x**2)) ** 2)
b0,b1,b2 = fmin(ss2, [1,1,-1], args=(x,y))
print b0,b1,b2
plt.plot(x, y, 'ro')
xvals = np.linspace(0, 10, 100)
plt.plot(xvals, b0 + b1*xvals + b2*(xvals**2))
```
Although polynomial model characterizes a nonlinear relationship, it is a linear problem in terms of estimation. That is, the regression model $f(y | x)$ is linear in the parameters.
For some data, it may be reasonable to consider polynomials of order>2. For example, consider the relationship between the number of home runs a baseball player hits and the number of runs batted in (RBI) they accumulate; clearly, the relationship is positive, but we may not expect a linear relationship.
```
ss3 = lambda theta, x, y: np.sum((y - theta[0] - theta[1]*x - theta[2]*(x**2)
- theta[3]*(x**3)) ** 2)
bb = pd.read_csv("data/baseball.csv", index_col=0)
plt.plot(bb.hr, bb.rbi, 'r.')
b0,b1,b2,b3 = fmin(ss3, [0,1,-1,0], args=(bb.hr, bb.rbi))
xvals = np.arange(40)
plt.plot(xvals, b0 + b1*xvals + b2*(xvals**2) + b3*(xvals**3))
```
Of course, we need not fit least squares models by hand. The `statsmodels` package implements least squares models that allow for model fitting in a single line:
```
import statsmodels.api as sm
straight_line = sm.OLS(y, sm.add_constant(x)).fit()
straight_line.summary()
from statsmodels.formula.api import ols as OLS
data = pd.DataFrame(dict(x=x, y=y))
cubic_fit = OLS('y ~ x + I(x**2)', data).fit()
cubic_fit.summary()
```
### Exercise: Polynomial function
Write a function that specified a polynomial of arbitrary degree.
```
# Write your answer here
```
## Model Selection
How do we choose among competing models for a given dataset? More parameters are not necessarily better, from the standpoint of model fit. For example, fitting a 9-th order polynomial to the sample data from the above example certainly results in an overfit.
```
def calc_poly(params, data):
x = np.c_[[data**i for i in range(len(params))]]
return np.dot(params, x)
ssp = lambda theta, x, y: np.sum((y - calc_poly(theta, x)) ** 2)
betas = fmin(ssp, np.zeros(10), args=(x,y), maxiter=1e6)
plt.plot(x, y, 'ro')
xvals = np.linspace(0, max(x), 100)
plt.plot(xvals, calc_poly(betas, xvals))
```
One approach is to use an information-theoretic criterion to select the most appropriate model. For example **Akaike's Information Criterion (AIC)** balances the fit of the model (in terms of the likelihood) with the number of parameters required to achieve that fit. We can easily calculate AIC as:
$$AIC = n \log(\hat{\sigma}^2) + 2p$$
where $p$ is the number of parameters in the model and $\hat{\sigma}^2 = RSS/(n-p-1)$.
Notice that as the number of parameters increase, the residual sum of squares goes down, but the second term (a penalty) increases.
To apply AIC to model selection, we choose the model that has the **lowest** AIC value.
```
n = len(x)
aic = lambda rss, p, n: n * np.log(rss/(n-p-1)) + 2*p
RSS1 = ss(fmin(ss, [0,1], args=(x,y)), x, y)
RSS2 = ss2(fmin(ss2, [1,1,-1], args=(x,y)), x, y)
print aic(RSS1, 2, n), aic(RSS2, 3, n)
```
Hence, we would select the 2-parameter (linear) model.
## Logistic Regression
Fitting a line to the relationship between two variables using the least squares approach is sensible when the variable we are trying to predict is continuous, but what about when the data are dichotomous?
- male/female
- pass/fail
- died/survived
Let's consider the problem of predicting survival in the Titanic disaster, based on our available information. For example, lets say that we want to predict survival as a function of the fare paid for the journey.
```
titanic = pd.read_excel("data/titanic.xls", "titanic")
titanic.name
jitter = np.random.normal(scale=0.02, size=len(titanic))
plt.scatter(np.log(titanic.fare), titanic.survived + jitter, alpha=0.3)
plt.yticks([0,1])
plt.ylabel("survived")
plt.xlabel("log(fare)")
```
I have added random jitter on the y-axis to help visualize the density of the points, and have plotted fare on the log scale.
Clearly, fitting a line through this data makes little sense, for several reasons. First, for most values of the predictor variable, the line would predict values that are not zero or one. Second, it would seem odd to choose least squares (or similar) as a criterion for selecting the best line.
```
x = np.log(titanic.fare[titanic.fare>0])
y = titanic.survived[titanic.fare>0]
betas_titanic = fmin(ss, [1,1], args=(x,y))
jitter = np.random.normal(scale=0.02, size=len(titanic))
plt.scatter(np.log(titanic.fare), titanic.survived + jitter, alpha=0.3)
plt.yticks([0,1])
plt.ylabel("survived")
plt.xlabel("log(fare)")
plt.plot([0,7], [betas_titanic[0], betas_titanic[0] + betas_titanic[1]*7.])
```
If we look at this data, we can see that for most values of `fare`, there are some individuals that survived and some that did not. However, notice that the cloud of points is denser on the "survived" (y=1) side for larger values of fare than on the "died" (y=0) side.
### Stochastic model
Rather than model the binary outcome explicitly, it makes sense instead to model the *probability* of death or survival in a **stochastic** model. Probabilities are measured on a continuous [0,1] scale, which may be more amenable for prediction using a regression line. We need to consider a different probability model for this exerciese however; let's consider the **Bernoulli** distribution as a generative model for our data:
<div style="font-size: 120%;">
$$f(y|p) = p^{y} (1-p)^{1-y}$$
</div>
where $y = \{0,1\}$ and $p \in [0,1]$. So, this model predicts whether $y$ is zero or one as a function of the probability $p$. Notice that when $y=1$, the $1-p$ term disappears, and when $y=0$, the $p$ term disappears.
So, the model we want to fit should look something like this:
<div style="font-size: 120%;">
$$p_i = \beta_0 + \beta_1 x_i + \epsilon_i$$
</div>
However, since $p$ is constrained to be between zero and one, it is easy to see where a linear (or polynomial) model might predict values outside of this range. We can modify this model sligtly by using a **link function** to transform the probability to have an unbounded range on a new scale. Specifically, we can use a **logit transformation** as our link function:
<div style="font-size: 120%;">
$$\text{logit}(p) = \log\left[\frac{p}{1-p}\right] = x$$
</div>
Here's a plot of $p/(1-p)$
```
logit = lambda p: np.log(p/(1.-p))
unit_interval = np.linspace(0,1)
plt.plot(unit_interval/(1-unit_interval), unit_interval)
```
And here's the logit function:
```
plt.plot(logit(unit_interval), unit_interval)
```
The inverse of the logit transformation is:
<div style="font-size: 150%;">
$$p = \frac{1}{1 + \exp(-x)}$$
</div>
So, now our model is:
<div style="font-size: 120%;">
$$\text{logit}(p_i) = \beta_0 + \beta_1 x_i + \epsilon_i$$
</div>
We can fit this model using maximum likelihood. Our likelihood, again based on the Bernoulli model is:
<div style="font-size: 120%;">
$$L(y|p) = \prod_{i=1}^n p_i^{y_i} (1-p_i)^{1-y_i}$$
</div>
which, on the log scale is:
<div style="font-size: 120%;">
$$l(y|p) = \sum_{i=1}^n y_i \log(p_i) + (1-y_i)\log(1-p_i)$$
</div>
We can easily implement this in Python, keeping in mind that `fmin` minimizes, rather than maximizes functions:
```
invlogit = lambda x: 1. / (1 + np.exp(-x))
def logistic_like(theta, x, y):
p = invlogit(theta[0] + theta[1] * x)
# Return negative of log-likelihood
return -np.sum(y * np.log(p) + (1-y) * np.log(1 - p))
```
Remove null values from variables
```
x, y = titanic[titanic.fare.notnull()][['fare', 'survived']].values.T
```
... and fit the model.
```
b0,b1 = fmin(logistic_like, [0.5,0], args=(x,y))
b0, b1
jitter = np.random.normal(scale=0.01, size=len(x))
plt.plot(x, y+jitter, 'r.', alpha=0.3)
plt.yticks([0,.25,.5,.75,1])
xvals = np.linspace(0, 600)
plt.plot(xvals, invlogit(b0+b1*xvals))
```
As with our least squares model, we can easily fit logistic regression models in `statsmodels`, in this case using the `GLM` (generalized linear model) class with a binomial error distribution specified.
```
logistic = sm.GLM(y, sm.add_constant(x), family=sm.families.Binomial()).fit()
logistic.summary()
```
### Exercise: multivariate logistic regression
Which other variables might be relevant for predicting the probability of surviving the Titanic? Generalize the model likelihood to include 2 or 3 other covariates from the dataset.
```
# Write your answer here
```
## Bootstrapping
Parametric inference can be **non-robust**:
* inaccurate if parametric assumptions are violated
* if we rely on asymptotic results, we may not achieve an acceptable level of accuracy
Parmetric inference can be **difficult**:
* derivation of sampling distribution may not be possible
An alternative is to estimate the sampling distribution of a statistic *empirically* without making assumptions about the form of the population.
We have seen this already with the kernel density estimate.
### Non-parametric Bootstrap
The bootstrap is a resampling method discovered by [Brad Efron](http://www.jstor.org/discover/10.2307/2958830?uid=3739568&uid=2&uid=4&uid=3739256&sid=21102342537691) that allows one to approximate the true sampling distribution of a dataset, and thereby obtain estimates of the mean and variance of the distribution.
Bootstrap sample:
<div style="font-size: 120%;">
$$S_1^* = \{x_{11}^*, x_{12}^*, \ldots, x_{1n}^*\}$$
</div>
$S_i^*$ is a sample of size $n$, **with** replacement.
In Python, we have already seen the NumPy function `permutation` that can be used in conjunction with Pandas' `take` method to generate a random sample of some data without replacement:
```
np.random.permutation(titanic.name)[:5]
```
Similarly, we can use the `random.randint` method to generate a sample *with* replacement, which we can use when bootstrapping.
```
random_ind = np.random.randint(0, len(titanic), 5)
titanic.name[random_ind]
```
We regard S as an "estimate" of population P
> population : sample :: sample : bootstrap sample
The idea is to generate replicate bootstrap samples:
<div style="font-size: 120%;">
$$S^* = \{S_1^*, S_2^*, \ldots, S_R^*\}$$
</div>
Compute statistic $t$ (estimate) for each bootstrap sample:
<div style="font-size: 120%;">
$$T_i^* = t(S^*)$$
</div>
```
n = 10
R = 1000
# Original sample (n=10)
x = np.random.normal(size=n)
# 1000 bootstrap samples of size 10
s = [x[np.random.randint(0,n,n)].mean() for i in range(R)]
_ = plt.hist(s, bins=30)
```
### Bootstrap Estimates
From our bootstrapped samples, we can extract *estimates* of the expectation and its variance:
$$\bar{T}^* = \hat{E}(T^*) = \frac{\sum_i T_i^*}{R}$$
$$\hat{\text{Var}}(T^*) = \frac{\sum_i (T_i^* - \bar{T}^*)^2}{R-1}$$
```
boot_mean = np.sum(s)/R
boot_mean
boot_var = ((np.array(s) - boot_mean) ** 2).sum() / (R-1)
boot_var
```
Since we have estimated the expectation of the bootstrapped statistics, we can estimate the **bias** of T:
$$\hat{B}^* = \bar{T}^* - T$$
```
boot_mean - np.mean(x)
```
### Bootstrap error
There are two sources of error in bootstrap estimates:
1. **Sampling error** from the selection of $S$.
2. **Bootstrap error** from failing to enumerate all possible bootstrap samples.
For the sake of accuracy, it is prudent to choose at least R=1000
### Bootstrap Percentile Intervals
An attractive feature of bootstrap statistics is the ease with which you can obtain an estimate of *uncertainty* for a given statistic. We simply use the empirical quantiles of the bootstrapped statistics to obtain percentiles corresponding to a confidence interval of interest.
This employs the *ordered* bootstrap replicates:
$$T_{(1)}^*, T_{(2)}^*, \ldots, T_{(R)}^*$$
Simply extract the $100(\alpha/2)$ and $100(1-\alpha/2)$ percentiles:
$$T_{[(R+1)\alpha/2]}^* \lt \theta \lt T_{[(R+1)(1-\alpha/2)]}^*$$
```
s_sorted = np.sort(s)
s_sorted[:10]
s_sorted[-10:]
alpha = 0.05
s_sorted[[(R+1)*alpha/2, (R+1)*(1-alpha/2)]]
```
### Exercise: Cervical dystonia bootstrap estimates
Use bootstrapping to estimate the mean of one of the treatment groups, and calculate percentile intervals for the mean.
```
# Write your answer here
```
| github_jupyter |
# Tutorial
## Regime-Switching Model
`regime_switch_model` is a set of algorithms for learning and inference on regime-switching model. Let $y_t$ be a $p\times 1$ observed time series and $h_t$ be a homogenous and stationary hidden Markov
chain taking values in $\{1, 2, \dots, m\}$ with transition probabilities
\begin{equation}
w_{kj} = P(h_{t+1}=j\mid h_t=k), \quad k,j=1, \dots, m
\end{equation}
where the number of hidden states $m$ is known. It is assumed that the financial market in
each period can be realized as one of $m$ regime. Furthermore, the regimes are characterized
by a set of $J$ risk factors, which represent broad macro and micro economic indicators. Let $F_{tj}$ be the value of the $j$th risk factor $(j=1, \dots, J)$ in period $t$. Correspondingly, $F_t$ is the vector of risk factors in period $t$. We assumes that, for $t=1, \dots, n$, when the market is in regime $h_t$ in period $t$,
\begin{equation}
y_t = u_{h_t} + B_{h_t}F_t + \Gamma_{h_t}\epsilon_t,
\end{equation}
where $\epsilon_t \sim N(0,I)$. The model parameters $\{u_{h_t}, B_{h_t}, \Gamma_{h_t}\}$ depend on the regime $h_t$. $u_{h_t}$ is the state-dependent intercepts of the linear factor model. The matrix $B_{h_t}$ defines the sensitivities of asset returns to the common risk factors in state $h_t$ and is often called the loading matrix.
`regime_switch_model` solves the following fundamental problems:
* Given the observed data, estimate the model parameters
* Given the model parameters and observed data, estimate the optimal sequence of hidden states
The implementation of code is based on the well-known Baum-Welch algorithm and Viterbi algorithm that are widely used in hidden Markov model.
```
import numpy as np
import pandas as pd
from regime_switch_model.rshmm import *
```
## Generate samples based on the regime-switching model
```
model = HMMRS(n_components=2)
# startprob
model.startprob_ = np.array([0.9, 0.1])
# transition matrix
model.transmat_ = np.array([[0.9, 0.1], [0.6, 0.4]])
# risk factor matrix
# read file from Fama-French three-factor data
Fama_French = pd.read_csv('Global_ex_US_3_Factors_Daily.csv', skiprows=3)
Fama_French.rename(columns={'Unnamed: 0': 'TimeStamp'}, inplace=True)
Fama_French.replace(-99.99, np.nan);
Fama_French.replace(-999, np.nan);
# select data
#Fama_French_subset = Fama_French[(Fama_French['TimeStamp'] >= 20150101) & (Fama_French['TimeStamp'] <= 20171231)]
Fama_French_subset = Fama_French
Fama_French_subset.drop(['TimeStamp', 'RF'], axis=1, inplace=True)
F = np.hstack((np.atleast_2d(np.ones(Fama_French_subset.shape[0])).T, Fama_French_subset))
# loading matrix with intercept
loadingmat1 = np.array([[0.9, 0.052, -0.02],
[0.3, 0.27, 0.01],
[0.12, 0.1, -0.05],
[0.04, 0.01, -0.15],
[0.15, 0.04, -0.11]])
intercept1 = np.atleast_2d(np.array([-0.015, -0.01, 0.005, 00.1, 0.02])).T
model.loadingmat_ = np.stack((np.hstack((intercept1, loadingmat1)),
np.hstack((0.25*intercept1, -0.5* loadingmat1))), axis=0)
# covariance matrix
n_stocks = 5
rho = 0.2
Sigma1 = np.full((n_stocks, n_stocks), rho) + np.diag(np.repeat(1-rho, n_stocks))
model.covmat_ = np.stack((Sigma1, 10*Sigma1), axis=0)
save = True
# sample
Y, Z = model.sample(F)
```
## Split data into training and test
```
# Use the last 300 day as the test data
Y_train = Y[:-300,:]
Y_test = Y[-300:,:]
F_train = F[:-300,:]
F_test = F[-300:,:]
```
## Fitting Regime-Switch Model
```
remodel = HMMRS(n_components=2, verbose=True)
remodel.fit(Y_train, F_train)
Z2, logl, viterbi_lattice = remodel.predict(Y_train, F_train)
```
### Examine model parameters
```
np.set_printoptions(precision=2)
print("Number of data points = ", Y_train.shape[0])
print(" ")
print("Starting probability")
print(remodel.startprob_)
print(" ")
print("Transition matrix")
print(remodel.transmat_)
print(" ")
print("Means and vars of each hidden state")
for i in range(remodel.n_components):
print("{0}th hidden state".format(i))
print("loading matrix = ", remodel.loadingmat_[i])
print("covariance = ", remodel.covmat_[i])
print(" ")
```
### Examine the predicted hidden state
```
print("Prediction accuracy of the hidden states = ", np.mean(np.equal(Z[:-300], 1-Z2)))
```
| github_jupyter |
# RadarCOVID-Report
## Data Extraction
```
import datetime
import json
import logging
import os
import shutil
import tempfile
import textwrap
import uuid
import matplotlib.pyplot as plt
import matplotlib.ticker
import numpy as np
import pandas as pd
import pycountry
import retry
import seaborn as sns
%matplotlib inline
current_working_directory = os.environ.get("PWD")
if current_working_directory:
os.chdir(current_working_directory)
sns.set()
matplotlib.rcParams["figure.figsize"] = (15, 6)
extraction_datetime = datetime.datetime.utcnow()
extraction_date = extraction_datetime.strftime("%Y-%m-%d")
extraction_previous_datetime = extraction_datetime - datetime.timedelta(days=1)
extraction_previous_date = extraction_previous_datetime.strftime("%Y-%m-%d")
extraction_date_with_hour = datetime.datetime.utcnow().strftime("%Y-%m-%d@%H")
current_hour = datetime.datetime.utcnow().hour
are_today_results_partial = current_hour != 23
```
### Constants
```
from Modules.ExposureNotification import exposure_notification_io
spain_region_country_code = "ES"
germany_region_country_code = "DE"
default_backend_identifier = spain_region_country_code
backend_generation_days = 7 * 2
daily_summary_days = 7 * 4 * 3
daily_plot_days = 7 * 4
tek_dumps_load_limit = daily_summary_days + 1
```
### Parameters
```
environment_backend_identifier = os.environ.get("RADARCOVID_REPORT__BACKEND_IDENTIFIER")
if environment_backend_identifier:
report_backend_identifier = environment_backend_identifier
else:
report_backend_identifier = default_backend_identifier
report_backend_identifier
environment_enable_multi_backend_download = \
os.environ.get("RADARCOVID_REPORT__ENABLE_MULTI_BACKEND_DOWNLOAD")
if environment_enable_multi_backend_download:
report_backend_identifiers = None
else:
report_backend_identifiers = [report_backend_identifier]
report_backend_identifiers
environment_invalid_shared_diagnoses_dates = \
os.environ.get("RADARCOVID_REPORT__INVALID_SHARED_DIAGNOSES_DATES")
if environment_invalid_shared_diagnoses_dates:
invalid_shared_diagnoses_dates = environment_invalid_shared_diagnoses_dates.split(",")
else:
invalid_shared_diagnoses_dates = []
invalid_shared_diagnoses_dates
```
### COVID-19 Cases
```
report_backend_client = \
exposure_notification_io.get_backend_client_with_identifier(
backend_identifier=report_backend_identifier)
@retry.retry(tries=10, delay=10, backoff=1.1, jitter=(0, 10))
def download_cases_dataframe():
return pd.read_csv("https://raw.githubusercontent.com/owid/covid-19-data/master/public/data/owid-covid-data.csv")
confirmed_df_ = download_cases_dataframe()
confirmed_df_.iloc[0]
confirmed_df = confirmed_df_.copy()
confirmed_df = confirmed_df[["date", "new_cases", "iso_code"]]
confirmed_df.rename(
columns={
"date": "sample_date",
"iso_code": "country_code",
},
inplace=True)
def convert_iso_alpha_3_to_alpha_2(x):
try:
return pycountry.countries.get(alpha_3=x).alpha_2
except Exception as e:
logging.info(f"Error converting country ISO Alpha 3 code '{x}': {repr(e)}")
return None
confirmed_df["country_code"] = confirmed_df.country_code.apply(convert_iso_alpha_3_to_alpha_2)
confirmed_df.dropna(inplace=True)
confirmed_df["sample_date"] = pd.to_datetime(confirmed_df.sample_date, dayfirst=True)
confirmed_df["sample_date"] = confirmed_df.sample_date.dt.strftime("%Y-%m-%d")
confirmed_df.sort_values("sample_date", inplace=True)
confirmed_df.tail()
confirmed_days = pd.date_range(
start=confirmed_df.iloc[0].sample_date,
end=extraction_datetime)
confirmed_days_df = pd.DataFrame(data=confirmed_days, columns=["sample_date"])
confirmed_days_df["sample_date_string"] = \
confirmed_days_df.sample_date.dt.strftime("%Y-%m-%d")
confirmed_days_df.tail()
def sort_source_regions_for_display(source_regions: list) -> list:
if report_backend_identifier in source_regions:
source_regions = [report_backend_identifier] + \
list(sorted(set(source_regions).difference([report_backend_identifier])))
else:
source_regions = list(sorted(source_regions))
return source_regions
report_source_regions = report_backend_client.source_regions_for_date(
date=extraction_datetime.date())
report_source_regions = sort_source_regions_for_display(
source_regions=report_source_regions)
report_source_regions
def get_cases_dataframe(source_regions_for_date_function, columns_suffix=None):
source_regions_at_date_df = confirmed_days_df.copy()
source_regions_at_date_df["source_regions_at_date"] = \
source_regions_at_date_df.sample_date.apply(
lambda x: source_regions_for_date_function(date=x))
source_regions_at_date_df.sort_values("sample_date", inplace=True)
source_regions_at_date_df["_source_regions_group"] = source_regions_at_date_df. \
source_regions_at_date.apply(lambda x: ",".join(sort_source_regions_for_display(x)))
source_regions_at_date_df.tail()
#%%
source_regions_for_summary_df_ = \
source_regions_at_date_df[["sample_date", "_source_regions_group"]].copy()
source_regions_for_summary_df_.rename(columns={"_source_regions_group": "source_regions"}, inplace=True)
source_regions_for_summary_df_.tail()
#%%
confirmed_output_columns = ["sample_date", "new_cases", "covid_cases"]
confirmed_output_df = pd.DataFrame(columns=confirmed_output_columns)
for source_regions_group, source_regions_group_series in \
source_regions_at_date_df.groupby("_source_regions_group"):
source_regions_set = set(source_regions_group.split(","))
confirmed_source_regions_set_df = \
confirmed_df[confirmed_df.country_code.isin(source_regions_set)].copy()
confirmed_source_regions_group_df = \
confirmed_source_regions_set_df.groupby("sample_date").new_cases.sum() \
.reset_index().sort_values("sample_date")
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df.merge(
confirmed_days_df[["sample_date_string"]].rename(
columns={"sample_date_string": "sample_date"}),
how="right")
confirmed_source_regions_group_df["new_cases"] = \
confirmed_source_regions_group_df["new_cases"].clip(lower=0)
confirmed_source_regions_group_df["covid_cases"] = \
confirmed_source_regions_group_df.new_cases.rolling(7, min_periods=0).mean().round()
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df[confirmed_output_columns]
confirmed_source_regions_group_df = confirmed_source_regions_group_df.replace(0, np.nan)
confirmed_source_regions_group_df.fillna(method="ffill", inplace=True)
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df[
confirmed_source_regions_group_df.sample_date.isin(
source_regions_group_series.sample_date_string)]
confirmed_output_df = confirmed_output_df.append(confirmed_source_regions_group_df)
result_df = confirmed_output_df.copy()
result_df.tail()
#%%
result_df.rename(columns={"sample_date": "sample_date_string"}, inplace=True)
result_df = confirmed_days_df[["sample_date_string"]].merge(result_df, how="left")
result_df.sort_values("sample_date_string", inplace=True)
result_df.fillna(method="ffill", inplace=True)
result_df.tail()
#%%
result_df[["new_cases", "covid_cases"]].plot()
if columns_suffix:
result_df.rename(
columns={
"new_cases": "new_cases_" + columns_suffix,
"covid_cases": "covid_cases_" + columns_suffix},
inplace=True)
return result_df, source_regions_for_summary_df_
confirmed_eu_df, source_regions_for_summary_df = get_cases_dataframe(
report_backend_client.source_regions_for_date)
confirmed_es_df, _ = get_cases_dataframe(
lambda date: [spain_region_country_code],
columns_suffix=spain_region_country_code.lower())
```
### Extract API TEKs
```
raw_zip_path_prefix = "Data/TEKs/Raw/"
base_backend_identifiers = [report_backend_identifier]
multi_backend_exposure_keys_df = \
exposure_notification_io.download_exposure_keys_from_backends(
backend_identifiers=report_backend_identifiers,
generation_days=backend_generation_days,
fail_on_error_backend_identifiers=base_backend_identifiers,
save_raw_zip_path_prefix=raw_zip_path_prefix)
multi_backend_exposure_keys_df["region"] = multi_backend_exposure_keys_df["backend_identifier"]
multi_backend_exposure_keys_df.rename(
columns={
"generation_datetime": "sample_datetime",
"generation_date_string": "sample_date_string",
},
inplace=True)
multi_backend_exposure_keys_df.head()
early_teks_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.rolling_period < 144].copy()
early_teks_df["rolling_period_in_hours"] = early_teks_df.rolling_period / 6
early_teks_df[early_teks_df.sample_date_string != extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
early_teks_df[early_teks_df.sample_date_string == extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
multi_backend_exposure_keys_df = multi_backend_exposure_keys_df[[
"sample_date_string", "region", "key_data"]]
multi_backend_exposure_keys_df.head()
active_regions = \
multi_backend_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
active_regions
multi_backend_summary_df = multi_backend_exposure_keys_df.groupby(
["sample_date_string", "region"]).key_data.nunique().reset_index() \
.pivot(index="sample_date_string", columns="region") \
.sort_index(ascending=False)
multi_backend_summary_df.rename(
columns={"key_data": "shared_teks_by_generation_date"},
inplace=True)
multi_backend_summary_df.rename_axis("sample_date", inplace=True)
multi_backend_summary_df = multi_backend_summary_df.fillna(0).astype(int)
multi_backend_summary_df = multi_backend_summary_df.head(backend_generation_days)
multi_backend_summary_df.head()
def compute_keys_cross_sharing(x):
teks_x = x.key_data_x.item()
common_teks = set(teks_x).intersection(x.key_data_y.item())
common_teks_fraction = len(common_teks) / len(teks_x)
return pd.Series(dict(
common_teks=common_teks,
common_teks_fraction=common_teks_fraction,
))
multi_backend_exposure_keys_by_region_df = \
multi_backend_exposure_keys_df.groupby("region").key_data.unique().reset_index()
multi_backend_exposure_keys_by_region_df["_merge"] = True
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_df.merge(
multi_backend_exposure_keys_by_region_df, on="_merge")
multi_backend_exposure_keys_by_region_combination_df.drop(
columns=["_merge"], inplace=True)
if multi_backend_exposure_keys_by_region_combination_df.region_x.nunique() > 1:
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_combination_df[
multi_backend_exposure_keys_by_region_combination_df.region_x !=
multi_backend_exposure_keys_by_region_combination_df.region_y]
multi_backend_exposure_keys_cross_sharing_df = \
multi_backend_exposure_keys_by_region_combination_df \
.groupby(["region_x", "region_y"]) \
.apply(compute_keys_cross_sharing) \
.reset_index()
multi_backend_cross_sharing_summary_df = \
multi_backend_exposure_keys_cross_sharing_df.pivot_table(
values=["common_teks_fraction"],
columns="region_x",
index="region_y",
aggfunc=lambda x: x.item())
multi_backend_cross_sharing_summary_df
multi_backend_without_active_region_exposure_keys_df = \
multi_backend_exposure_keys_df[multi_backend_exposure_keys_df.region != report_backend_identifier]
multi_backend_without_active_region = \
multi_backend_without_active_region_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
multi_backend_without_active_region
exposure_keys_summary_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.region == report_backend_identifier]
exposure_keys_summary_df.drop(columns=["region"], inplace=True)
exposure_keys_summary_df = \
exposure_keys_summary_df.groupby(["sample_date_string"]).key_data.nunique().to_frame()
exposure_keys_summary_df = \
exposure_keys_summary_df.reset_index().set_index("sample_date_string")
exposure_keys_summary_df.sort_index(ascending=False, inplace=True)
exposure_keys_summary_df.rename(columns={"key_data": "shared_teks_by_generation_date"}, inplace=True)
exposure_keys_summary_df.head()
```
### Dump API TEKs
```
tek_list_df = multi_backend_exposure_keys_df[
["sample_date_string", "region", "key_data"]].copy()
tek_list_df["key_data"] = tek_list_df["key_data"].apply(str)
tek_list_df.rename(columns={
"sample_date_string": "sample_date",
"key_data": "tek_list"}, inplace=True)
tek_list_df = tek_list_df.groupby(
["sample_date", "region"]).tek_list.unique().reset_index()
tek_list_df["extraction_date"] = extraction_date
tek_list_df["extraction_date_with_hour"] = extraction_date_with_hour
tek_list_path_prefix = "Data/TEKs/"
tek_list_current_path = tek_list_path_prefix + f"/Current/RadarCOVID-TEKs.json"
tek_list_daily_path = tek_list_path_prefix + f"Daily/RadarCOVID-TEKs-{extraction_date}.json"
tek_list_hourly_path = tek_list_path_prefix + f"Hourly/RadarCOVID-TEKs-{extraction_date_with_hour}.json"
for path in [tek_list_current_path, tek_list_daily_path, tek_list_hourly_path]:
os.makedirs(os.path.dirname(path), exist_ok=True)
tek_list_base_df = tek_list_df[tek_list_df.region == report_backend_identifier]
tek_list_base_df.drop(columns=["extraction_date", "extraction_date_with_hour"]).to_json(
tek_list_current_path,
lines=True, orient="records")
tek_list_base_df.drop(columns=["extraction_date_with_hour"]).to_json(
tek_list_daily_path,
lines=True, orient="records")
tek_list_base_df.to_json(
tek_list_hourly_path,
lines=True, orient="records")
tek_list_base_df.head()
```
### Load TEK Dumps
```
import glob
def load_extracted_teks(mode, region=None, limit=None) -> pd.DataFrame:
extracted_teks_df = pd.DataFrame(columns=["region"])
file_paths = list(reversed(sorted(glob.glob(tek_list_path_prefix + mode + "/RadarCOVID-TEKs-*.json"))))
if limit:
file_paths = file_paths[:limit]
for file_path in file_paths:
logging.info(f"Loading TEKs from '{file_path}'...")
iteration_extracted_teks_df = pd.read_json(file_path, lines=True)
extracted_teks_df = extracted_teks_df.append(
iteration_extracted_teks_df, sort=False)
extracted_teks_df["region"] = \
extracted_teks_df.region.fillna(spain_region_country_code).copy()
if region:
extracted_teks_df = \
extracted_teks_df[extracted_teks_df.region == region]
return extracted_teks_df
daily_extracted_teks_df = load_extracted_teks(
mode="Daily",
region=report_backend_identifier,
limit=tek_dumps_load_limit)
daily_extracted_teks_df.head()
exposure_keys_summary_df_ = daily_extracted_teks_df \
.sort_values("extraction_date", ascending=False) \
.groupby("sample_date").tek_list.first() \
.to_frame()
exposure_keys_summary_df_.index.name = "sample_date_string"
exposure_keys_summary_df_["tek_list"] = \
exposure_keys_summary_df_.tek_list.apply(len)
exposure_keys_summary_df_ = exposure_keys_summary_df_ \
.rename(columns={"tek_list": "shared_teks_by_generation_date"}) \
.sort_index(ascending=False)
exposure_keys_summary_df = exposure_keys_summary_df_
exposure_keys_summary_df.head()
```
### Daily New TEKs
```
tek_list_df = daily_extracted_teks_df.groupby("extraction_date").tek_list.apply(
lambda x: set(sum(x, []))).reset_index()
tek_list_df = tek_list_df.set_index("extraction_date").sort_index(ascending=True)
tek_list_df.head()
def compute_teks_by_generation_and_upload_date(date):
day_new_teks_set_df = tek_list_df.copy().diff()
try:
day_new_teks_set = day_new_teks_set_df[
day_new_teks_set_df.index == date].tek_list.item()
except ValueError:
day_new_teks_set = None
if pd.isna(day_new_teks_set):
day_new_teks_set = set()
day_new_teks_df = daily_extracted_teks_df[
daily_extracted_teks_df.extraction_date == date].copy()
day_new_teks_df["shared_teks"] = \
day_new_teks_df.tek_list.apply(lambda x: set(x).intersection(day_new_teks_set))
day_new_teks_df["shared_teks"] = \
day_new_teks_df.shared_teks.apply(len)
day_new_teks_df["upload_date"] = date
day_new_teks_df.rename(columns={"sample_date": "generation_date"}, inplace=True)
day_new_teks_df = day_new_teks_df[
["upload_date", "generation_date", "shared_teks"]]
day_new_teks_df["generation_to_upload_days"] = \
(pd.to_datetime(day_new_teks_df.upload_date) -
pd.to_datetime(day_new_teks_df.generation_date)).dt.days
day_new_teks_df = day_new_teks_df[day_new_teks_df.shared_teks > 0]
return day_new_teks_df
shared_teks_generation_to_upload_df = pd.DataFrame()
for upload_date in daily_extracted_teks_df.extraction_date.unique():
shared_teks_generation_to_upload_df = \
shared_teks_generation_to_upload_df.append(
compute_teks_by_generation_and_upload_date(date=upload_date))
shared_teks_generation_to_upload_df \
.sort_values(["upload_date", "generation_date"], ascending=False, inplace=True)
shared_teks_generation_to_upload_df.tail()
today_new_teks_df = \
shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.upload_date == extraction_date].copy()
today_new_teks_df.tail()
if not today_new_teks_df.empty:
today_new_teks_df.set_index("generation_to_upload_days") \
.sort_index().shared_teks.plot.bar()
generation_to_upload_period_pivot_df = \
shared_teks_generation_to_upload_df[
["upload_date", "generation_to_upload_days", "shared_teks"]] \
.pivot(index="upload_date", columns="generation_to_upload_days") \
.sort_index(ascending=False).fillna(0).astype(int) \
.droplevel(level=0, axis=1)
generation_to_upload_period_pivot_df.head()
new_tek_df = tek_list_df.diff().tek_list.apply(
lambda x: len(x) if not pd.isna(x) else None).to_frame().reset_index()
new_tek_df.rename(columns={
"tek_list": "shared_teks_by_upload_date",
"extraction_date": "sample_date_string",}, inplace=True)
new_tek_df.tail()
shared_teks_uploaded_on_generation_date_df = shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.generation_to_upload_days == 0] \
[["upload_date", "shared_teks"]].rename(
columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_teks_uploaded_on_generation_date",
})
shared_teks_uploaded_on_generation_date_df.head()
estimated_shared_diagnoses_df = shared_teks_generation_to_upload_df \
.groupby(["upload_date"]).shared_teks.max().reset_index() \
.sort_values(["upload_date"], ascending=False) \
.rename(columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_diagnoses",
})
invalid_shared_diagnoses_dates_mask = \
estimated_shared_diagnoses_df.sample_date_string.isin(invalid_shared_diagnoses_dates)
estimated_shared_diagnoses_df[invalid_shared_diagnoses_dates_mask] = 0
estimated_shared_diagnoses_df.head()
```
### Hourly New TEKs
```
hourly_extracted_teks_df = load_extracted_teks(
mode="Hourly", region=report_backend_identifier, limit=25)
hourly_extracted_teks_df.head()
hourly_new_tek_count_df = hourly_extracted_teks_df \
.groupby("extraction_date_with_hour").tek_list. \
apply(lambda x: set(sum(x, []))).reset_index().copy()
hourly_new_tek_count_df = hourly_new_tek_count_df.set_index("extraction_date_with_hour") \
.sort_index(ascending=True)
hourly_new_tek_count_df["new_tek_list"] = hourly_new_tek_count_df.tek_list.diff()
hourly_new_tek_count_df["new_tek_count"] = hourly_new_tek_count_df.new_tek_list.apply(
lambda x: len(x) if not pd.isna(x) else 0)
hourly_new_tek_count_df.rename(columns={
"new_tek_count": "shared_teks_by_upload_date"}, inplace=True)
hourly_new_tek_count_df = hourly_new_tek_count_df.reset_index()[[
"extraction_date_with_hour", "shared_teks_by_upload_date"]]
hourly_new_tek_count_df.head()
hourly_summary_df = hourly_new_tek_count_df.copy()
hourly_summary_df.set_index("extraction_date_with_hour", inplace=True)
hourly_summary_df = hourly_summary_df.fillna(0).astype(int).reset_index()
hourly_summary_df["datetime_utc"] = pd.to_datetime(
hourly_summary_df.extraction_date_with_hour, format="%Y-%m-%d@%H")
hourly_summary_df.set_index("datetime_utc", inplace=True)
hourly_summary_df = hourly_summary_df.tail(-1)
hourly_summary_df.head()
```
### Official Statistics
```
import requests
import pandas.io.json
official_stats_response = requests.get("https://radarcovid.covid19.gob.es/kpi/statistics/basics")
official_stats_response.raise_for_status()
official_stats_df_ = pandas.io.json.json_normalize(official_stats_response.json())
official_stats_df = official_stats_df_.copy()
official_stats_df["date"] = pd.to_datetime(official_stats_df["date"], dayfirst=True)
official_stats_df.head()
official_stats_column_map = {
"date": "sample_date",
"applicationsDownloads.totalAcummulated": "app_downloads_es_accumulated",
"communicatedContagions.totalAcummulated": "shared_diagnoses_es_accumulated",
}
accumulated_suffix = "_accumulated"
accumulated_values_columns = \
list(filter(lambda x: x.endswith(accumulated_suffix), official_stats_column_map.values()))
interpolated_values_columns = \
list(map(lambda x: x[:-len(accumulated_suffix)], accumulated_values_columns))
official_stats_df = \
official_stats_df[official_stats_column_map.keys()] \
.rename(columns=official_stats_column_map)
official_stats_df["extraction_date"] = extraction_date
official_stats_df.head()
official_stats_path = "Data/Statistics/Current/RadarCOVID-Statistics.json"
previous_official_stats_df = pd.read_json(official_stats_path, orient="records", lines=True)
previous_official_stats_df["sample_date"] = pd.to_datetime(previous_official_stats_df["sample_date"], dayfirst=True)
official_stats_df = official_stats_df.append(previous_official_stats_df)
official_stats_df.head()
official_stats_df = official_stats_df[~(official_stats_df.shared_diagnoses_es_accumulated == 0)]
official_stats_df.sort_values("extraction_date", ascending=False, inplace=True)
official_stats_df.drop_duplicates(subset=["sample_date"], keep="first", inplace=True)
official_stats_df.head()
official_stats_stored_df = official_stats_df.copy()
official_stats_stored_df["sample_date"] = official_stats_stored_df.sample_date.dt.strftime("%Y-%m-%d")
official_stats_stored_df.to_json(official_stats_path, orient="records", lines=True)
official_stats_df.drop(columns=["extraction_date"], inplace=True)
official_stats_df = confirmed_days_df.merge(official_stats_df, how="left")
official_stats_df.sort_values("sample_date", ascending=False, inplace=True)
official_stats_df.head()
official_stats_df[accumulated_values_columns] = \
official_stats_df[accumulated_values_columns] \
.astype(float).interpolate(limit_area="inside")
official_stats_df[interpolated_values_columns] = \
official_stats_df[accumulated_values_columns].diff(periods=-1)
official_stats_df.drop(columns="sample_date", inplace=True)
official_stats_df.head()
```
### Data Merge
```
result_summary_df = exposure_keys_summary_df.merge(
new_tek_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
shared_teks_uploaded_on_generation_date_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
estimated_shared_diagnoses_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
official_stats_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = confirmed_eu_df.tail(daily_summary_days).merge(
result_summary_df, on=["sample_date_string"], how="left")
result_summary_df.head()
result_summary_df = confirmed_es_df.tail(daily_summary_days).merge(
result_summary_df, on=["sample_date_string"], how="left")
result_summary_df.head()
result_summary_df["sample_date"] = pd.to_datetime(result_summary_df.sample_date_string)
result_summary_df = result_summary_df.merge(source_regions_for_summary_df, how="left")
result_summary_df.set_index(["sample_date", "source_regions"], inplace=True)
result_summary_df.drop(columns=["sample_date_string"], inplace=True)
result_summary_df.sort_index(ascending=False, inplace=True)
result_summary_df.head()
with pd.option_context("mode.use_inf_as_na", True):
result_summary_df = result_summary_df.fillna(0).astype(int)
result_summary_df["teks_per_shared_diagnosis"] = \
(result_summary_df.shared_teks_by_upload_date / result_summary_df.shared_diagnoses).fillna(0)
result_summary_df["shared_diagnoses_per_covid_case"] = \
(result_summary_df.shared_diagnoses / result_summary_df.covid_cases).fillna(0)
result_summary_df["shared_diagnoses_per_covid_case_es"] = \
(result_summary_df.shared_diagnoses_es / result_summary_df.covid_cases_es).fillna(0)
result_summary_df.head(daily_plot_days)
def compute_aggregated_results_summary(days) -> pd.DataFrame:
aggregated_result_summary_df = result_summary_df.copy()
aggregated_result_summary_df["covid_cases_for_ratio"] = \
aggregated_result_summary_df.covid_cases.mask(
aggregated_result_summary_df.shared_diagnoses == 0, 0)
aggregated_result_summary_df["covid_cases_for_ratio_es"] = \
aggregated_result_summary_df.covid_cases_es.mask(
aggregated_result_summary_df.shared_diagnoses_es == 0, 0)
aggregated_result_summary_df = aggregated_result_summary_df \
.sort_index(ascending=True).fillna(0).rolling(days).agg({
"covid_cases": "sum",
"covid_cases_es": "sum",
"covid_cases_for_ratio": "sum",
"covid_cases_for_ratio_es": "sum",
"shared_teks_by_generation_date": "sum",
"shared_teks_by_upload_date": "sum",
"shared_diagnoses": "sum",
"shared_diagnoses_es": "sum",
}).sort_index(ascending=False)
with pd.option_context("mode.use_inf_as_na", True):
aggregated_result_summary_df = aggregated_result_summary_df.fillna(0).astype(int)
aggregated_result_summary_df["teks_per_shared_diagnosis"] = \
(aggregated_result_summary_df.shared_teks_by_upload_date /
aggregated_result_summary_df.covid_cases_for_ratio).fillna(0)
aggregated_result_summary_df["shared_diagnoses_per_covid_case"] = \
(aggregated_result_summary_df.shared_diagnoses /
aggregated_result_summary_df.covid_cases_for_ratio).fillna(0)
aggregated_result_summary_df["shared_diagnoses_per_covid_case_es"] = \
(aggregated_result_summary_df.shared_diagnoses_es /
aggregated_result_summary_df.covid_cases_for_ratio_es).fillna(0)
return aggregated_result_summary_df
aggregated_result_with_7_days_window_summary_df = compute_aggregated_results_summary(days=7)
aggregated_result_with_7_days_window_summary_df.head()
last_7_days_summary = aggregated_result_with_7_days_window_summary_df.to_dict(orient="records")[1]
last_7_days_summary
aggregated_result_with_14_days_window_summary_df = compute_aggregated_results_summary(days=13)
last_14_days_summary = aggregated_result_with_14_days_window_summary_df.to_dict(orient="records")[1]
last_14_days_summary
```
## Report Results
```
display_column_name_mapping = {
"sample_date": "Sample\u00A0Date\u00A0(UTC)",
"source_regions": "Source Countries",
"datetime_utc": "Timestamp (UTC)",
"upload_date": "Upload Date (UTC)",
"generation_to_upload_days": "Generation to Upload Period in Days",
"region": "Backend",
"region_x": "Backend\u00A0(A)",
"region_y": "Backend\u00A0(B)",
"common_teks": "Common TEKs Shared Between Backends",
"common_teks_fraction": "Fraction of TEKs in Backend (A) Available in Backend (B)",
"covid_cases": "COVID-19 Cases (Source Countries)",
"shared_teks_by_generation_date": "Shared TEKs by Generation Date (Source Countries)",
"shared_teks_by_upload_date": "Shared TEKs by Upload Date (Source Countries)",
"shared_teks_uploaded_on_generation_date": "Shared TEKs Uploaded on Generation Date (Source Countries)",
"shared_diagnoses": "Shared Diagnoses (Source Countries – Estimation)",
"teks_per_shared_diagnosis": "TEKs Uploaded per Shared Diagnosis (Source Countries)",
"shared_diagnoses_per_covid_case": "Usage Ratio (Source Countries)",
"covid_cases_es": "COVID-19 Cases (Spain)",
"app_downloads_es": "App Downloads (Spain – Official)",
"shared_diagnoses_es": "Shared Diagnoses (Spain – Official)",
"shared_diagnoses_per_covid_case_es": "Usage Ratio (Spain)",
}
summary_columns = [
"covid_cases",
"shared_teks_by_generation_date",
"shared_teks_by_upload_date",
"shared_teks_uploaded_on_generation_date",
"shared_diagnoses",
"teks_per_shared_diagnosis",
"shared_diagnoses_per_covid_case",
"covid_cases_es",
"app_downloads_es",
"shared_diagnoses_es",
"shared_diagnoses_per_covid_case_es",
]
summary_percentage_columns= [
"shared_diagnoses_per_covid_case_es",
"shared_diagnoses_per_covid_case",
]
```
### Daily Summary Table
```
result_summary_df_ = result_summary_df.copy()
result_summary_df = result_summary_df[summary_columns]
result_summary_with_display_names_df = result_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
result_summary_with_display_names_df
```
### Daily Summary Plots
```
result_plot_summary_df = result_summary_df.head(daily_plot_days)[summary_columns] \
.droplevel(level=["source_regions"]) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
summary_ax_list = result_plot_summary_df.sort_index(ascending=True).plot.bar(
title=f"Daily Summary",
rot=45, subplots=True, figsize=(15, 30), legend=False)
ax_ = summary_ax_list[0]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.95)
_ = ax_.set_xticklabels(sorted(result_plot_summary_df.index.strftime("%Y-%m-%d").tolist()))
for percentage_column in summary_percentage_columns:
percentage_column_index = summary_columns.index(percentage_column)
summary_ax_list[percentage_column_index].yaxis \
.set_major_formatter(matplotlib.ticker.PercentFormatter(1.0))
```
### Daily Generation to Upload Period Table
```
display_generation_to_upload_period_pivot_df = \
generation_to_upload_period_pivot_df \
.head(backend_generation_days)
display_generation_to_upload_period_pivot_df \
.head(backend_generation_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping)
fig, generation_to_upload_period_pivot_table_ax = plt.subplots(
figsize=(12, 1 + 0.6 * len(display_generation_to_upload_period_pivot_df)))
generation_to_upload_period_pivot_table_ax.set_title(
"Shared TEKs Generation to Upload Period Table")
sns.heatmap(
data=display_generation_to_upload_period_pivot_df
.rename_axis(columns=display_column_name_mapping)
.rename_axis(index=display_column_name_mapping),
fmt=".0f",
annot=True,
ax=generation_to_upload_period_pivot_table_ax)
generation_to_upload_period_pivot_table_ax.get_figure().tight_layout()
```
### Hourly Summary Plots
```
hourly_summary_ax_list = hourly_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.plot.bar(
title=f"Last 24h Summary",
rot=45, subplots=True, legend=False)
ax_ = hourly_summary_ax_list[-1]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.9)
_ = ax_.set_xticklabels(sorted(hourly_summary_df.index.strftime("%Y-%m-%d@%H").tolist()))
```
### Publish Results
```
github_repository = os.environ.get("GITHUB_REPOSITORY")
if github_repository is None:
github_repository = "pvieito/Radar-STATS"
github_project_base_url = "https://github.com/" + github_repository
display_formatters = {
display_column_name_mapping["teks_per_shared_diagnosis"]: lambda x: f"{x:.2f}" if x != 0 else "",
display_column_name_mapping["shared_diagnoses_per_covid_case"]: lambda x: f"{x:.2%}" if x != 0 else "",
display_column_name_mapping["shared_diagnoses_per_covid_case_es"]: lambda x: f"{x:.2%}" if x != 0 else "",
}
general_columns = \
list(filter(lambda x: x not in display_formatters, display_column_name_mapping.values()))
general_formatter = lambda x: f"{x}" if x != 0 else ""
display_formatters.update(dict(map(lambda x: (x, general_formatter), general_columns)))
daily_summary_table_html = result_summary_with_display_names_df \
.head(daily_plot_days) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.to_html(formatters=display_formatters)
multi_backend_summary_table_html = multi_backend_summary_df \
.head(daily_plot_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(formatters=display_formatters)
def format_multi_backend_cross_sharing_fraction(x):
if pd.isna(x):
return "-"
elif round(x * 100, 1) == 0:
return ""
else:
return f"{x:.1%}"
multi_backend_cross_sharing_summary_table_html = multi_backend_cross_sharing_summary_df \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(
classes="table-center",
formatters=display_formatters,
float_format=format_multi_backend_cross_sharing_fraction)
multi_backend_cross_sharing_summary_table_html = \
multi_backend_cross_sharing_summary_table_html \
.replace("<tr>","<tr style=\"text-align: center;\">")
extraction_date_result_summary_df = \
result_summary_df[result_summary_df.index.get_level_values("sample_date") == extraction_date]
extraction_date_result_hourly_summary_df = \
hourly_summary_df[hourly_summary_df.extraction_date_with_hour == extraction_date_with_hour]
covid_cases = \
extraction_date_result_summary_df.covid_cases.item()
shared_teks_by_generation_date = \
extraction_date_result_summary_df.shared_teks_by_generation_date.item()
shared_teks_by_upload_date = \
extraction_date_result_summary_df.shared_teks_by_upload_date.item()
shared_diagnoses = \
extraction_date_result_summary_df.shared_diagnoses.item()
teks_per_shared_diagnosis = \
extraction_date_result_summary_df.teks_per_shared_diagnosis.item()
shared_diagnoses_per_covid_case = \
extraction_date_result_summary_df.shared_diagnoses_per_covid_case.item()
shared_teks_by_upload_date_last_hour = \
extraction_date_result_hourly_summary_df.shared_teks_by_upload_date.sum().astype(int)
display_source_regions = ", ".join(report_source_regions)
if len(report_source_regions) == 1:
display_brief_source_regions = report_source_regions[0]
else:
display_brief_source_regions = f"{len(report_source_regions)} 🇪🇺"
def get_temporary_image_path() -> str:
return os.path.join(tempfile.gettempdir(), str(uuid.uuid4()) + ".png")
def save_temporary_plot_image(ax):
if isinstance(ax, np.ndarray):
ax = ax[0]
media_path = get_temporary_image_path()
ax.get_figure().savefig(media_path)
return media_path
def save_temporary_dataframe_image(df):
import dataframe_image as dfi
df = df.copy()
df_styler = df.style.format(display_formatters)
media_path = get_temporary_image_path()
dfi.export(df_styler, media_path)
return media_path
summary_plots_image_path = save_temporary_plot_image(
ax=summary_ax_list)
summary_table_image_path = save_temporary_dataframe_image(
df=result_summary_with_display_names_df)
hourly_summary_plots_image_path = save_temporary_plot_image(
ax=hourly_summary_ax_list)
multi_backend_summary_table_image_path = save_temporary_dataframe_image(
df=multi_backend_summary_df)
generation_to_upload_period_pivot_table_image_path = save_temporary_plot_image(
ax=generation_to_upload_period_pivot_table_ax)
```
### Save Results
```
report_resources_path_prefix = "Data/Resources/Current/RadarCOVID-Report-"
result_summary_df.to_csv(
report_resources_path_prefix + "Summary-Table.csv")
result_summary_df.to_html(
report_resources_path_prefix + "Summary-Table.html")
hourly_summary_df.to_csv(
report_resources_path_prefix + "Hourly-Summary-Table.csv")
multi_backend_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Summary-Table.csv")
multi_backend_cross_sharing_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Cross-Sharing-Summary-Table.csv")
generation_to_upload_period_pivot_df.to_csv(
report_resources_path_prefix + "Generation-Upload-Period-Table.csv")
_ = shutil.copyfile(
summary_plots_image_path,
report_resources_path_prefix + "Summary-Plots.png")
_ = shutil.copyfile(
summary_table_image_path,
report_resources_path_prefix + "Summary-Table.png")
_ = shutil.copyfile(
hourly_summary_plots_image_path,
report_resources_path_prefix + "Hourly-Summary-Plots.png")
_ = shutil.copyfile(
multi_backend_summary_table_image_path,
report_resources_path_prefix + "Multi-Backend-Summary-Table.png")
_ = shutil.copyfile(
generation_to_upload_period_pivot_table_image_path,
report_resources_path_prefix + "Generation-Upload-Period-Table.png")
```
### Publish Results as JSON
```
def generate_summary_api_results(df: pd.DataFrame) -> list:
api_df = df.reset_index().copy()
api_df["sample_date_string"] = \
api_df["sample_date"].dt.strftime("%Y-%m-%d")
api_df["source_regions"] = \
api_df["source_regions"].apply(lambda x: x.split(","))
return api_df.to_dict(orient="records")
summary_api_results = \
generate_summary_api_results(df=result_summary_df)
today_summary_api_results = \
generate_summary_api_results(df=extraction_date_result_summary_df)[0]
summary_results = dict(
backend_identifier=report_backend_identifier,
source_regions=report_source_regions,
extraction_datetime=extraction_datetime,
extraction_date=extraction_date,
extraction_date_with_hour=extraction_date_with_hour,
last_hour=dict(
shared_teks_by_upload_date=shared_teks_by_upload_date_last_hour,
shared_diagnoses=0,
),
today=today_summary_api_results,
last_7_days=last_7_days_summary,
last_14_days=last_14_days_summary,
daily_results=summary_api_results)
summary_results = \
json.loads(pd.Series([summary_results]).to_json(orient="records"))[0]
with open(report_resources_path_prefix + "Summary-Results.json", "w") as f:
json.dump(summary_results, f, indent=4)
```
### Publish on README
```
with open("Data/Templates/README.md", "r") as f:
readme_contents = f.read()
readme_contents = readme_contents.format(
extraction_date_with_hour=extraction_date_with_hour,
github_project_base_url=github_project_base_url,
daily_summary_table_html=daily_summary_table_html,
multi_backend_summary_table_html=multi_backend_summary_table_html,
multi_backend_cross_sharing_summary_table_html=multi_backend_cross_sharing_summary_table_html,
display_source_regions=display_source_regions)
with open("README.md", "w") as f:
f.write(readme_contents)
```
### Publish on Twitter
```
enable_share_to_twitter = os.environ.get("RADARCOVID_REPORT__ENABLE_PUBLISH_ON_TWITTER")
github_event_name = os.environ.get("GITHUB_EVENT_NAME")
if enable_share_to_twitter and github_event_name == "schedule" and \
(shared_teks_by_upload_date_last_hour or not are_today_results_partial):
import tweepy
twitter_api_auth_keys = os.environ["RADARCOVID_REPORT__TWITTER_API_AUTH_KEYS"]
twitter_api_auth_keys = twitter_api_auth_keys.split(":")
auth = tweepy.OAuthHandler(twitter_api_auth_keys[0], twitter_api_auth_keys[1])
auth.set_access_token(twitter_api_auth_keys[2], twitter_api_auth_keys[3])
api = tweepy.API(auth)
summary_plots_media = api.media_upload(summary_plots_image_path)
summary_table_media = api.media_upload(summary_table_image_path)
generation_to_upload_period_pivot_table_image_media = api.media_upload(generation_to_upload_period_pivot_table_image_path)
media_ids = [
summary_plots_media.media_id,
summary_table_media.media_id,
generation_to_upload_period_pivot_table_image_media.media_id,
]
if are_today_results_partial:
today_addendum = " (Partial)"
else:
today_addendum = ""
def format_shared_diagnoses_per_covid_case(value) -> str:
if value == 0:
return "–"
return f"≤{value:.2%}"
display_shared_diagnoses_per_covid_case = \
format_shared_diagnoses_per_covid_case(value=shared_diagnoses_per_covid_case)
display_last_14_days_shared_diagnoses_per_covid_case = \
format_shared_diagnoses_per_covid_case(value=last_14_days_summary["shared_diagnoses_per_covid_case"])
display_last_14_days_shared_diagnoses_per_covid_case_es = \
format_shared_diagnoses_per_covid_case(value=last_14_days_summary["shared_diagnoses_per_covid_case_es"])
status = textwrap.dedent(f"""
#RadarCOVID – {extraction_date_with_hour}
Today{today_addendum}:
- Uploaded TEKs: {shared_teks_by_upload_date:.0f} ({shared_teks_by_upload_date_last_hour:+d} last hour)
- Shared Diagnoses: ≤{shared_diagnoses:.0f}
- Usage Ratio: {display_shared_diagnoses_per_covid_case}
Last 14 Days:
- Usage Ratio (Estimation): {display_last_14_days_shared_diagnoses_per_covid_case}
- Usage Ratio (Official): {display_last_14_days_shared_diagnoses_per_covid_case_es}
Info: {github_project_base_url}#documentation
""")
status = status.encode(encoding="utf-8")
api.update_status(status=status, media_ids=media_ids)
```
| github_jupyter |
```
#load watermark
%load_ext watermark
%watermark -a 'Gopala KR' -u -d -v -p watermark,numpy,pandas,matplotlib,nltk,sklearn,tensorflow,theano,mxnet,chainer,seaborn,keras,tflearn,bokeh,gensim
```
# Cheatsheet for Decision Tree Classification
### Algorithm
1. Start at the root node as parent node
2. Split the parent node at the feature ***a*** to minimize the sum of the child node impurities (maximize information gain)
3. Assign training samples to new child nodes
3. Stop if leave nodes are pure or early stopping criteria is satisfied, else repeat steps 1 and 2 for each new child node
### Stopping Rules
- a maximal node depth is reached
- splitting a note does not lead to an information gain
### Criterion
Splitting criterion: Information Gain (IG), sum of node impurities
Objective function: Maximize IG at each split, eqiv. minimize the the impurity criterion
### Information Gain (IG)
***Examples below are given for binary splits.***
$$IG(D_{p}, a) = I(D_{p}) - \frac{N_{left}}{N_p} I(D_{left}) - \frac{N_{right}}{N_p} I(D_{right})$$
- $IG$: Information Gain
- $a$: feature to perform the split
- $N_p$: number of samples in the parent node
- $N_{left}$: number of samples in the left child node
- $N_{right}$: number of samples in the right child node
- $I$: impurity
- $D_{p}$: training subset of the parent node
- $D_{left}$: training subset of the left child node
- $D_{right}$: training subset of the right child node
# Impurity (I) Indices
### Entropy
The entropy is defined as
$$I_H(t) = - \sum_{i =1}^{C} p(i \mid t) \;log_2 \,p(i \mid t)$$
for all non-empty classes ($p(i \mid t) \neq 0$), where $p(i \mid t)$ is the proportion (or frequency or probability) of the samples that belong to class $i$ for a particular node $t$; $C$ is the number of unique class labels.
The entropy is therefore 0 if all samples at a node belong to the same class, and the entropy is maximal if we have an uniform class distribution. For example, in a binary class setting, the entropy is 0 if $p(i =1 \mid t) =1$ or $p(i =0 \mid t) =1$. And if the classes are distributed uniformly with $p(i =1 \mid t) = 0.5$ and $p(i =0 \mid t) =0.5$ the entropy is 1 (maximal), which we can visualize by plotting the entropy for binary class setting below.
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
def entropy(p):
return - p*np.log2(p) - (1 - p)*np.log2((1 - p))
x = np.arange(0.0, 1.0, 0.01)
ent = [entropy(p) if p != 0 else None for p in x]
plt.plot(x, ent)
plt.ylim([0,1.1])
plt.xlabel('p(i=1)')
plt.axhline(y=1.0, linewidth=1, color='k', linestyle='--')
plt.ylabel('Entropy')
plt.show()
```
### Gini Impurity
$$I_G(t) = \sum_{i =1}^{C}p(i \mid t) \big(1-p(i \mid t)\big)$$
```
def gini(p):
return (p)*(1 - (p)) + (1-p)*(1 - (1-p))
x = np.arange(0.0, 1.0, 0.01)
plt.plot(x, gini(x))
plt.ylim([0,1.1])
plt.xlabel('p(i=1)')
plt.axhline(y=0.5, linewidth=1, color='k', linestyle='--')
plt.ylabel('Gini Impurity')
plt.show()
```
### Misclassification Error
$$I_M(t) = 1 - max\{{p_i}\}$$
```
def error(p):
return 1 - np.max([p, 1-p])
x = np.arange(0.0, 1.0, 0.01)
err = [error(i) for i in x]
plt.plot(x, err)
plt.ylim([0,1.1])
plt.xlabel('p(i=1)')
plt.axhline(y=0.5, linewidth=1, color='k', linestyle='--')
plt.ylabel('Misclassification Error')
plt.show()
```
### Comparison
```
fig = plt.figure()
ax = plt.subplot(111)
for i, lab in zip([ent, gini(x), err],
['Entropy', 'Gini Impurity', 'Misclassification Error']):
line, = ax.plot(x, i, label=lab)
ax.legend(loc='upper center', bbox_to_anchor=(0.5, 1.15),
ncol=3, fancybox=True, shadow=False)
ax.axhline(y=0.5, linewidth=1, color='k', linestyle='--')
ax.axhline(y=1.0, linewidth=1, color='k', linestyle='--')
plt.ylim([0,1.1])
plt.xlabel('p(i=1)')
plt.ylabel('Impurity Index')
plt.tight_layout()
plt.show()
tested ; Gopal
```
| github_jupyter |
```
from orphics import sehgal, maps
import healpy as hp
from pixell import utils, enmap, curvedsky, enplot, wcsutils
import os
import numpy as np
import matplotlib.pyplot as plt
import lmdb
from cosmikyu import datasets, transforms, config
from pitas import modecoupling
import random
%matplotlib inline
%load_ext autoreload
%autoreload 2
data_dir = config.default_data_dir
sehgal_dir = os.path.join(data_dir, 'sehgal')
stat_dir = os.path.join(sehgal_dir, "stats")
#norm_info_file = "/home/dwhan89/workspace/cosmikyu/data/sehgal/201020_logzshrink_normalization_info_validation.npz"
#SDN = transforms.SehgalDataNormalizerScaledLogZShrink(norm_info_file)
#SDS_test = datasets.SehgalDataSet(sehgal_dir, "train141020", transforms=[SDN], dummy_label=False)
#SDS_input = datasets.SehgalDataSet(sehgal_dir, "train_tertiary191120",
# transforms=[SDN], dummy_label=False, dtype=np.float32)
norm_info_file = "/home/dwhan89/workspace/cosmikyu/data/sehgal/281220_logz_normalization_info_validation.npz"
SDN = transforms.SehgalDataNormalizerScaledLogZShrink(norm_info_file)
SDS_test = datasets.SehgalDataSet(sehgal_dir, "train281220_fromcat", transforms=[SDN], dummy_label=False,
dtype=np.float64)
SDS_input = datasets.SehgalDataSet(sehgal_dir, "train_tertiary281220_fromcat", transforms=[SDN],
dummy_label=False, dtype=np.float32)
plot_dir = "/home/dwhan89/scratch/outbox/cosmikyu"
def plot_path(x):
return os.path.join(plot_dir, x)
ps_file = "/home/dwhan89/workspace/cosmikyu/data/sehgal/ps_dist_train010121.npy"
nsample = len(SDS_test)
if not os.path.exists(ps_file):
storage = np.zeros((nsample, 5))
for i in range(len(SDS_test)):
if i % 5000 == 0: print(i)
storage[i,:] = np.var(SDS_test[i], axis=(-1,-2))
np.save(ps_file, storage)
else:
storage = np.load(ps_file)
bin_edges = np.linspace(0,storage[:,2].max(),200)
bin_center = (bin_edges[:-1]+ bin_edges[1:])/2
digitized = np.digitize(storage[:,2], bin_edges)
nbin = len(bin_center)
binned = np.zeros(nbin)
for i in range(nbin):
binned[i] = np.sum(digitized == i+1)
binned /= nsample
del digitized
#hist, binedges = np.histogram(storage[:,2], bins=100, density=False)
plt.plot(bin_center, binned)
plt.xlim(0,5)
plt.grid()
frac = 0.20
npart = int(nsample*frac)
ind = np.argpartition(storage[:,2], -npart)[-npart:]
print(np.min(storage[ind,2]))
print(npart/nsample*100)
for i in range(5):
cidx = random.choice(ind)
enplot.pshow(SDS_test[cidx][2])
highflux_idx = [i for i in range(nsample) if i in ind]
lowflux_idx = [i for i in range(nsample) if i not in ind]
np.save("/home/dwhan89/workspace/cosmikyu/data/sehgal/ps_dist_train010121_tszflux_sortedidx.npy", np.argsort(storage[:,2]))
ps_file = "/home/dwhan89/workspace/cosmikyu/data/sehgal/ps_dist_tertiary_train010121.npy"
nsample = len(SDS_input)
if not os.path.exists(ps_file):
storage = np.zeros((nsample, 5))
for i in range(len(SDS_input)):
if i % 5000 == 0: print(i)
storage[i,:] = np.var(SDS_input[i], axis=(-1,-2))
np.save(ps_file, storage)
else:
storage = np.load(ps_file)
bin_edges = np.linspace(0,storage[:,2].max(),200)
bin_center = (bin_edges[:-1]+ bin_edges[1:])/2
digitized = np.digitize(storage[:,2], bin_edges)
nbin = len(bin_center)
binned = np.zeros(nbin)
for i in range(nbin):
binned[i] = np.sum(digitized == i+1)
binned /= nsample
del digitized
#hist, binedges = np.histogram(storage[:,2], bins=100, density=False)
plt.plot(bin_center, binned)
plt.xlim(0,2)
plt.grid()
frac = 0.2
npart = int(nsample*frac)
ind = np.argpartition(storage[:,2], -npart)[-npart:]
print(np.min(storage[ind,2]))
print(npart/nsample*100)
for i in range(5):
cidx = random.choice(ind)
enplot.pshow(SDS_test[cidx][2])
np.save("/home/dwhan89/workspace/cosmikyu/data/sehgal/ps_dist_tertiary_train010121_tszflux_sortedidx.npy", np.argsort(storage[:,2]))
```
| github_jupyter |
```
#import necessary libraries
import torch
from transformers import *
import pandas as pd
import re
import collections
import numpy as np
import json
import time
from tqdm.notebook import tqdm
import torch.nn as nn
import pathlib
#output all items, not just last one
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
#set device
if torch.cuda.is_available():
device= "cuda"
else:
device = "cpu"
device
# define Articles dataset class for easy sampling, iteration, and weight creating
class Articles(torch.utils.data.Dataset):
def __init__(self, json_file):
super().__init__()
with open(json_file, "r") as data_file:
self.examples = json.loads(data_file.read())
def __getitem__(self, idx):
return self.examples[idx]
def __len__(self):
return len(self.examples)
def tokenize(self):
for idx, example in enumerate(self.examples):
self.examples[idx]['text'] = re.findall('[\w]+', self.examples[idx]['text'].lower())
def create_positive_sampler(self, target_publication):
prob = np.zeros(len(self))
for idx, example in enumerate(self.examples):
if example['model_publication'] == target_publication:
prob[idx] = 1
return torch.utils.data.WeightedRandomSampler(weights=prob, num_samples=len(self), replacement=True)
def create_negative_sampler(self, target_publication):
prob = np.zeros(len(self))
for idx, example in enumerate(self.examples):
if example['model_publication'] != target_publication:
prob[idx] = 1
return torch.utils.data.WeightedRandomSampler(weights=prob, num_samples=len(self), replacement=True)
def map_items(self, word_to_id, url_to_id, publication_to_id, filter=False, min_length=0):
min_length_articles = []
for idx, example in enumerate(self.examples):
self.examples[idx]['text'] = [word_to_id.get(word, len(word_to_id)) for word in example['text']]
self.examples[idx]['text'] = [word for word in example['text'] if word != len(word_to_id)]
if filter:
if len(self.examples[idx]['text']) > min_length:
min_length_articles.append(self.examples[idx])
self.examples[idx]['url'] = url_to_id.get(example['url'], url_to_id.get("miscellaneous"))
self.examples[idx]['model_publication'] = publication_to_id.get(example['model_publication'], publication_to_id.get("miscellaneous"))
return min_length_articles
from transformers import BertTokenizer
# Load the BERT tokenizer.
print('Loading BERT tokenizer...')
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased', do_lower_case=True)
sentences = "The White House medical unit and Secret Service will evaluate attendees before they're admitted. They'll be required to test negative for the virus on the day of the event, complete a health questionnaire and pass a temperature screening. The sites will be cleaned and sanitized before each event. Trump Victory, the president’s joint fundraising committee, will cover the testing costs. The move comes as Trump expresses his desire for the country to reopen even as medical professionals raise concerns about the potential dangers of doing so. The pandemic death toll passed 100,000 this week. Trump is also dead set on holding the GOP’s August national convention in Charlotte, even as North Carolina officials are raising concerns about safety. The Charlotte area has seen an uptick in coronavirus cases in recent days. Trump has also been itching to resume his trademark rallies, his primary method of connecting with supporters and broadcasting his message. The president’s campaign team has used the events to glean data from attendees which is used to turn out his voters. The president has left the confines of the White House over the past few weeks to hold ostensibly official events in swing states like Arizona and Michigan, where polls have shown him trailing presumptive Democratic nominee Joe Biden. The events have sometimes had the feel of a rally, complete with walk-out music. Trump’s advisers want him to be seen as eager to reopen the country while Democrats push for stay-at-home orders that keep the economy shuttered. Earlier this month, the reelection effort released a one-minute advertisement titled “American Comeback” highlighting the president’s desire to reignite the economy. The spot concluded with Trump’s mantra that he will “make America great again.” The president’s previously busy fundraising schedule came to a halt in March. A planned March 12 fundraiser with GOP megadonor Sheldon Adelson was scrapped, as were a pair of events that month to be headlined by first lady Melania Trump. Even with in-person fundraising events slashed, the president has maintained robust fundraising totals thanks to a massive small-donor operation. Trump’s political machine narrowly outraised Biden in April and has a $187 million cash-on-hand lead over Biden and the Democratic Party. Biden has sworn off in-person fundraisers during the pandemic, instead doing online events from his Delaware home. Couples will need to donate $580,600 to attend the Dallas fundraiser. A single attendee to the New Jersey fundraiser will need to give $250,000. The money will go to Trump Victory, a joint fundraising committee of the Trump campaign, Republican National Committee, and state parties."
print(' Original: ', sentences)
# Print the sentence split into tokens.
print('Tokenized: ', tokenizer.tokenize(sentences))
re.findall('[\w]+', sentences.lower())
# Print the sentence mapped to token ids.
print('Token IDs: ', tokenizer.convert_tokens_to_ids(tokenizer.tokenize(sentences[0])))
train_data = Articles("../data/final-data/debugdata/train_basic.json")
val_data = Articles("../data/final-data/debugdata/eval_basic.json")
for idx in range(len(train_data)):
train_data.examples[idx]['text'] = tokenizer.tokenize(train_data.examples[idx]['text'])
if len(train_data.examples[idx]['text']) > 512:
train_data.examples[idx]['text'] = train_data.examples[idx]['text'][:512]
train_data.examples[idx]['text'] = tokenizer.encode(
train_data.examples[idx]['text'],
add_special_tokens = True,
max_length = 512)
train_data.examples[idx]['model_publication'] = 1 if train_data.examples[idx]['model_publication'] == 'target' else 0
#Create batches with positive samples in first half and negative examples in second half
class BatchSamplerWithNegativeSamples(torch.utils.data.Sampler):
def __init__(self, pos_sampler, neg_sampler, batch_size, items):
self._pos_sampler = pos_sampler
self._neg_sampler = neg_sampler
self._items = items
assert batch_size % 2 == 0, 'Batch size must be divisible by two for negative samples.'
self._batch_size = batch_size
def __iter__(self):
batch, neg_batch = [], []
neg_sampler = iter(self._neg_sampler)
for pos_idx in self._pos_sampler:
batch.append(pos_idx)
neg_idx = pos_idx
# keep sampling until we get a true negative sample
while self._items[neg_idx] == self._items[pos_idx]:
try:
neg_idx = next(neg_sampler)
except StopIteration:
neg_sampler = iter(self._neg_sampler)
neg_idx = next(neg_sampler)
neg_batch.append(neg_idx)
if len(batch) == self._batch_size // 2:
batch.extend(neg_batch)
yield batch
batch, neg_batch = [], []
return
def __len__(self):
return len(self._pos_sampler) // self._batch_size
#define function to return necessary data for dataloader to pass into model
def collate_fn(examples):
words = []
articles = []
labels = []
publications = []
for example in examples:
words.append(example['text'])
articles.append(example['url'])
labels.append(example['model_publication'])
publications.append(example['publication'])
num_words = [len(x) for x in words]
words = np.concatenate(words, axis=0)
word_attributes = torch.tensor(words, dtype=torch.long)
articles = torch.tensor(articles, dtype=torch.long)
num_words.insert(0,0)
num_words.pop(-1)
attribute_offsets = torch.tensor(np.cumsum(num_words), dtype=torch.long)
publications = torch.tensor(publications, dtype=torch.long)
real_labels = torch.tensor(labels, dtype=torch.long)
return publications, articles, word_attributes, attribute_offsets, real_labels
from transformers import BertForSequenceClassification, AdamW, BertConfig
# Load BertForSequenceClassification, the pretrained BERT model with a single
# linear classification layer on top.
model = BertForSequenceClassification.from_pretrained(
"bert-base-uncased", # Use the 12-layer BERT model, with an uncased vocab.
num_labels = 2, # The number of output labels--2 for binary classification.
# You can increase this for multi-class tasks.
output_attentions = False, # Whether the model returns attentions weights.
output_hidden_states = False, # Whether the model returns all hidden-states.
)
# Tell pytorch to run this model on the GPU.
model.cuda()
```
| github_jupyter |
```
# Importing all necessary packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
retail = pd.read_csv('../Data/online_retail.csv')
retail
```
# Data Cleaning
```
retail.info()
```
### Figuring out number of missing values in each column
```
retail.isnull().sum().sort_values(ascending=False)
```
#### The total number of rows :
```
retail.shape[0]
```
** Hence, we find out that out of total 5 lakh rows, more than 1 lakh rows have no customerID specified and as many as 1454 rows have no product description mentioned. **
```
# Number of those rows with Description null
retail[retail.Description.isnull() & retail.CustomerID.isnull()].shape[0]
```
** This indicates that orders with no description mentioned have no customerID specified.
Hence all of those rows can be deleted. **
## Removing the rows with missing values
```
retail_new = retail.dropna()
retail_new
# checking for missing values
retail_new.isnull().sum().sort_values(ascending=False)
# changing the InvoiceDate format from string to Timestamp format
retail_new.InvoiceDate = pd.to_datetime(retail_new.InvoiceDate, format='%m/%d/%Y %H:%M')
# Total number of orders :
retail_new.InvoiceNo.nunique()
# number of unique products being ordered
retail_new.Description.nunique()
# total number of customers
retail_new.CustomerID.nunique()
# rows with negative quantity value
retail_new[retail_new.Quantity < 0 ].shape[0]
```
** There are 8905 rows with negative quantity ordered. **
## Removing the rows with negative quantity of product ordered
```
retail_new = retail_new[retail_new.Quantity >= 0]
retail_new[retail_new.Quantity <0].shape[0]
```
** All rows with negative quantity removed **
# Adding a column for the total price of the product ordered
```
retail_new['TotalPrice'] = retail_new.Quantity * retail_new.UnitPrice
retail_new
```
# Number of orders of different customers
```
orders = retail_new.groupby(by=['CustomerID', 'Country'], as_index=False).InvoiceNo.count()
orders
sorted_orders = orders.sort_values(by='InvoiceNo', ascending=False)
sorted_orders.head(n=20)
plt.subplots(figsize=(15,8))
plt.plot(orders.CustomerID, orders.InvoiceNo)
plt.xlabel('CustomerID')
plt.ylabel('Total orders')
plt.title('Number of orders for different customers')
plt.show()
orders[orders.InvoiceNo == orders.InvoiceNo.max()]
```
### Observation :
The customer that has placed the maximum orders is from UK.
# Number of customers from different countries
```
df = retail_new.groupby(['Country']).CustomerID.value_counts().unstack()
df.notnull().sum(axis=1)
```
### Analysing the top 20 customer countries
```
top20 = df.notnull().sum(axis=1)
top20 = top20.sort_values(ascending=False)
top20 = top20.head(n=20)
top20
# All the customers
df.notnull().sum(axis=1).plot(kind='bar', figsize=(15,8))
```
### Observation :
Maximum number(around 3921) of customers belong to UK.
```
retail_new.groupby(by=['CustomerID', 'Country']).TotalPrice.sum()
```
# Monthly revenue
```
retail_new['Month'] = retail_new.InvoiceDate.dt.month
retail_new.InvoiceDate
retail_new.groupby(['Month']).TotalPrice.sum()
plt.title('Monthly revenue')
retail_new.groupby(['Month']).TotalPrice.sum().plot(kind='bar')
```
### Observation:
November observes the maximum income for the company.
# Monthly sales
```
retail_new.Month.value_counts().sort_index()
# is same as :
# retail_new.groupby(['Month']).InvoiceNo.count()
plt.title('Monthly sales')
retail_new.Month.value_counts().sort_index().plot(kind='bar', figsize=(15,8))
```
### Observation :
The month of November observes the maximum sales.
# Analysis of discounted products
```
discounts = retail_new[retail_new.TotalPrice == 0]
discounts
discounts.Month.value_counts().sort_index()
discounts.Month.value_counts().sort_index().plot(kind='bar')
```
### Observation :
1. The firm gives away maximum products for free in the month of November.
2. Products are given for free throughout the year except for the month of June.
| github_jupyter |
# Interpreting Nodes and Edges by Saliency Maps in GAT
This demo shows how to use integrated gradients in graph attention networks to obtain accurate importance estimations for both the nodes and edges. The notebook consists of three parts:
setting up the node classification problem for Cora citation network
training and evaluating a GAT model for node classification
calculating node and edge importances for model's predictions of query ("target") nodes.
```
import networkx as nx
import pandas as pd
import numpy as np
from scipy import stats
import os
import time
import sys
import stellargraph as sg
from copy import deepcopy
from stellargraph.mapper import FullBatchNodeGenerator
from stellargraph.layer import GAT, GraphAttention
print(sg.__file__)
from keras import layers, optimizers, losses, metrics, models, Model
from sklearn import preprocessing, feature_extraction, model_selection
import keras.backend as K
import matplotlib.pyplot as plt
%matplotlib inline
```
### Loading the CORA network
**Downloading the CORA dataset:**
The dataset used in this demo can be downloaded from https://linqs-data.soe.ucsc.edu/public/lbc/cora.tgz
The following is the description of the dataset:
> The Cora dataset consists of 2708 scientific publications classified into one of seven classes.
> The citation network consists of 5429 links. Each publication in the dataset is described by a
> 0/1-valued word vector indicating the absence/presence of the corresponding word from the dictionary.
> The dictionary consists of 1433 unique words. The README file in the dataset provides more details.
Download and unzip the cora.tgz file to a location on your computer and set the `data_dir` variable to
point to the location of the dataset (the directory containing "cora.cites" and "cora.content").
```
data_dir = os.path.expanduser("~/data/cora")
```
Load the graph from edgelist
```
edgelist = pd.read_csv(os.path.join(data_dir, "cora.cites"), header=None, names=["source", "target"], sep='\t')
edgelist["label"] = "cites"
Gnx = nx.from_pandas_edgelist(edgelist, edge_attr="label")
nx.set_node_attributes(Gnx, "paper", "label")
```
Load the features and subject for the nodes
```
feature_names = ["w_{}".format(ii) for ii in range(1433)]
column_names = feature_names + ["subject"]
node_data = pd.read_csv(os.path.join(data_dir, "cora.content"), header=None, names=column_names, sep='\t')
```
### Splitting the data
For machine learning we want to take a subset of the nodes for training, and use the rest for validation and testing. We'll use scikit-learn again to do this.
Here we're taking 140 node labels for training, 500 for validation, and the rest for testing.
```
train_data, test_data = model_selection.train_test_split(node_data, train_size=140, test_size=None, stratify=node_data['subject'])
val_data, test_data = model_selection.train_test_split(test_data, train_size=500, test_size=None, stratify=test_data['subject'])
from collections import Counter
Counter(train_data['subject'])
```
### Converting to numeric arrays
For our categorical target, we will use one-hot vectors that will be fed into a soft-max Keras layer during training. To do this conversion ...
```
target_encoding = feature_extraction.DictVectorizer(sparse=False)
train_targets = target_encoding.fit_transform(train_data[["subject"]].to_dict('records'))
val_targets = target_encoding.transform(val_data[["subject"]].to_dict('records'))
test_targets = target_encoding.transform(test_data[["subject"]].to_dict('records'))
node_ids = node_data.index
all_targets = target_encoding.transform(
node_data[["subject"]].to_dict("records")
)
```
We now do the same for the node attributes we want to use to predict the subject. These are the feature vectors that the Keras model will use as input. The CORA dataset contains attributes 'w_x' that correspond to words found in that publication. If a word occurs more than once in a publication the relevant attribute will be set to one, otherwise it will be zero.
```
node_features = node_data[feature_names]
```
### Creating the GAT model in Keras
Now create a StellarGraph object from the NetworkX graph and the node features and targets. It is StellarGraph objects that we use in this library to perform machine learning tasks on.
```
G = sg.StellarGraph(Gnx, node_features=node_features)
print(G.info())
```
To feed data from the graph to the Keras model we need a generator. Since GAT is a full-batch model, we use the `FullBatchNodeGenerator` class to feed node features and graph adjacency matrix to the model.
```
generator = FullBatchNodeGenerator(G, method='gat',sparse=False)
```
For training we map only the training nodes returned from our splitter and the target values.
```
train_gen = generator.flow(train_data.index, train_targets)
```
Now we can specify our machine learning model, we need a few more parameters for this:
* the `layer_sizes` is a list of hidden feature sizes of each layer in the model. In this example we use two GAT layers with 8-dimensional hidden node features at each layer.
* `attn_heads` is the number of attention heads in all but the last GAT layer in the model
* `activations` is a list of activations applied to each layer's output
* Arguments such as `bias`, `in_dropout`, `attn_dropout` are internal parameters of the model, execute `?GAT` for details.
To follow the GAT model architecture used for Cora dataset in the original paper [Graph Attention Networks. P. Velickovic et al. ICLR 2018 https://arxiv.org/abs/1803.07294], let's build a 2-layer GAT model, with the 2nd layer being the classifier that predicts paper subject: it thus should have the output size of `train_targets.shape[1]` (7 subjects) and a softmax activation.
```
gat = GAT(
layer_sizes=[8, train_targets.shape[1]],
attn_heads=8,
generator=generator,
bias=True,
in_dropout=0,
attn_dropout=0,
activations=["elu","softmax"],
normalize=None,
saliency_map_support=True
)
# Expose the input and output tensors of the GAT model for node prediction, via GAT.node_model() method:
x_inp, predictions = gat.node_model()
```
### Training the model
Now let's create the actual Keras model with the input tensors `x_inp` and output tensors being the predictions `predictions` from the final dense layer
```
model = Model(inputs=x_inp, outputs=predictions)
model.compile(
optimizer=optimizers.Adam(lr=0.005),
loss=losses.categorical_crossentropy,
weighted_metrics=["acc"],
)
```
Train the model, keeping track of its loss and accuracy on the training set, and its generalisation performance on the validation set (we need to create another generator over the validation data for this)
```
val_gen = generator.flow(val_data.index, val_targets)
```
Train the model
```
N = len(node_ids)
history = model.fit_generator(train_gen, validation_data=val_gen, shuffle=False, epochs=10, verbose=2)
import matplotlib.pyplot as plt
%matplotlib inline
def remove_prefix(text, prefix):
return text[text.startswith(prefix) and len(prefix):]
def plot_history(history):
metrics = sorted(set([remove_prefix(m, "val_") for m in list(history.history.keys())]))
for m in metrics:
# summarize history for metric m
plt.plot(history.history[m])
plt.plot(history.history['val_' + m])
plt.title(m)
plt.ylabel(m)
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='best')
plt.show()
plot_history(history)
```
Evaluate the trained model on the test set
```
test_gen = generator.flow(test_data.index, test_targets)
test_metrics = model.evaluate_generator(test_gen)
print("\nTest Set Metrics:")
for name, val in zip(model.metrics_names, test_metrics):
print("\t{}: {:0.4f}".format(name, val))
```
Check serialization
```
# Save model
model_json = model.to_json()
model_weights = model.get_weights()
# Load model from json & set all weights
model2 = models.model_from_json(
model_json, custom_objects={"GraphAttention": GraphAttention}
)
model2.set_weights(model_weights)
model2_weights = model2.get_weights()
pred2 = model2.predict_generator(test_gen)
pred1 = model.predict_generator(test_gen)
print(np.allclose(pred1,pred2))
```
# Node and link importance via saliency maps
Now we define the importances of node features, nodes, and links in the target node's neighbourhood (ego-net), and evaluate them using our library.
Node feature importance: given a target node $t$ and the model's prediction of $t$'s class, for each node $v$ in its ego-net, feature importance of feature $f$ for node $v$ is defined as the change in the target node's predicted score $s(c)$ for the winning class $c$ if feature $f$ of node $v$ is perturbed.
The overall node importance for node $v$ is defined here as the sum of all feature importances for node $v$, i.e., it is the amount by which the target node's predicted score $s(c)$ would change if we set all features of node $v$ to zeros.
Link importance for link $e=(u, v)$ is defined as the change in target node $t$'s predicted score $s(c)$ if the link $e$ is removed from the graph. Links with high importance (positive or negative) affect the target node prediction more than links with low importance.
Node and link importances can be used to assess the role of neighbour nodes and links in model's predictions for the node(s) of interest (the target nodes). For datasets like CORA-ML, the features and edges are binary, vanilla gradients may not perform well so we use integrated gradients to compute them (https://arxiv.org/pdf/1703.01365.pdf).
```
from stellargraph.utils.saliency_maps import IntegratedGradientsGAT
from stellargraph.utils.saliency_maps import GradientSaliencyGAT
```
Select the target node whose prediction is to be interpreted.
```
graph_nodes = list(G.nodes())
all_gen = generator.flow(graph_nodes)
target_idx = 7
target_nid = graph_nodes[target_idx]
print(target_nid)
target_gen = generator.flow([target_nid])
```
Node id of the target node:
```
y_true = all_targets[target_idx] # true class of the target node
```
Extract adjacency matrix and feature matrix
```
y_pred = model.predict_generator(target_gen).squeeze()
class_of_interest = np.argmax(y_pred)
print("target node id: {}, \ntrue label: {}, \npredicted label: {}".format(target_nid, y_true, y_pred.round(2)))
```
Get the node feature importance by using integrated gradients
```
int_grad_saliency = IntegratedGradientsGAT(model, train_gen, generator.node_list)
saliency = GradientSaliencyGAT(model, train_gen)
```
Get the ego network of the target node.
```
G_ego = nx.ego_graph(G, target_nid, radius=len(gat.activations))
```
Compute the link importance by integrated gradients.
```
integrate_link_importance = int_grad_saliency.get_link_importance(target_nid, class_of_interest, steps=25)
print('integrated_link_mask.shape = {}'.format(integrate_link_importance.shape))
print(class_of_interest)
integrated_node_importance = int_grad_saliency.get_node_importance(target_nid, class_of_interest, steps=25)
print(integrated_node_importance.shape)
print('\nintegrated_node_importance', integrated_node_importance.round(2))
print('integrated self-importance of target node {}: {}'.format(target_nid, integrated_node_importance[target_idx].round(2)))
print("\nEgo net of target node {} has {} nodes".format(target_nid, G_ego.number_of_nodes()))
```
Get the ranks of the edge importance values.
```
sorted_indices = np.argsort(integrate_link_importance.flatten().reshape(-1))
sorted_indices = np.array(sorted_indices)
integrated_link_importance_rank = [(int(k/N), k%N) for k in sorted_indices[::-1]]
topk = 10
print('Top {} most important links by integrated gradients are {}'.format(topk, integrated_link_importance_rank[:topk]))
#print('Top {} most important links by integrated gradients (for potential edges) are {}'.format(topk, integrated_link_importance_rank_add[-topk:]))
```
In the following, we plot the link and node importance (computed by integrated gradients) of the nodes within the ego graph of the target node.
For nodes, the shape of the node indicates the positive/negative importance the node has. 'round' nodes have positive importance while 'diamond' nodes have negative importance. The size of the node indicates the value of the importance, e.g., a large diamond node has higher negative importance.
For links, the color of the link indicates the positive/negative importance the link has. 'red' links have positive importance while 'blue' links have negative importance. The width of the link indicates the value of the importance, e.g., a thicker blue link has higher negative importance.
```
nx.set_node_attributes(G_ego, values={x[0]:{'subject': x[1]} for x in node_data['subject'].items()})
node_size_factor = 1e2
link_width_factor = 4
nodes = list(G_ego.nodes())
colors = pd.DataFrame([v[1]['subject'] for v in G_ego.nodes(data=True)],
index=nodes, columns=['subject'])
colors = np.argmax(target_encoding.transform(colors.to_dict('records')), axis=1) + 1
fig, ax = plt.subplots(1, 1, figsize=(15, 10));
pos = nx.spring_layout(G_ego)
# Draw ego as large and red
node_sizes = [integrated_node_importance[graph_nodes.index(k)] for k in G_ego.nodes()]
node_shapes = ['o' if integrated_node_importance[graph_nodes.index(k)] > 0
else 'd' for k in G_ego.nodes()]
positive_colors, negative_colors = [], []
positive_node_sizes, negative_node_sizes = [], []
positive_nodes, negative_nodes = [], []
#node_size_sclae is used for better visualization of nodes
node_size_scale = node_size_factor/np.max(node_sizes)
for k in range(len(node_shapes)):
if list(nodes)[k] == target_nid:
continue
if node_shapes[k] == 'o':
positive_colors.append(colors[k])
positive_nodes.append(list(nodes)[k])
positive_node_sizes.append(node_size_scale*node_sizes[k])
else:
negative_colors.append(colors[k])
negative_nodes.append(list(nodes)[k])
negative_node_sizes.append(node_size_scale*abs(node_sizes[k]))
cmap = plt.get_cmap('jet', np.max(colors)-np.min(colors)+1)
nc = nx.draw_networkx_nodes(G_ego, pos, nodelist=positive_nodes,
node_color=positive_colors, cmap=cmap,
node_size=positive_node_sizes, with_labels=False,
vmin=np.min(colors)-0.5, vmax=np.max(colors)+0.5, node_shape='o')
nc = nx.draw_networkx_nodes(G_ego, pos, nodelist=negative_nodes,
node_color=negative_colors, cmap=cmap,
node_size=negative_node_sizes, with_labels=False,
vmin=np.min(colors)-0.5, vmax=np.max(colors)+0.5, node_shape='d')
# Draw the target node as a large star colored by its true subject
nx.draw_networkx_nodes(G_ego, pos, nodelist=[target_nid],
node_size=50*abs(node_sizes[nodes.index(target_nid)]), node_shape='*',
node_color=[colors[nodes.index(target_nid)]],
cmap=cmap, vmin=np.min(colors)-0.5, vmax=np.max(colors)+0.5, label="Target")
edges = G_ego.edges()
#link_width_scale is used for better visualization of links
weights = [integrate_link_importance[graph_nodes.index(u),list(G.node()).index(v)] for u,v in edges]
link_width_scale = link_width_factor/np.max(weights)
edge_colors = ['red' if integrate_link_importance[graph_nodes.index(u),list(G.node()).index(v)] > 0 else 'blue' for u,v in edges]
ec = nx.draw_networkx_edges(G_ego, pos, edge_color=edge_colors,
width = [link_width_scale*w for w in weights])
plt.legend()
plt.colorbar(nc, ticks=np.arange(np.min(colors),np.max(colors)+1))
plt.axis('off')
plt.show()
```
We then remove the node or edge in the ego graph one by one and check how the prediction changes. By doing so, we can obtain the ground truth importance of the nodes and edges. Comparing the following figure and the above one can show the effectiveness of integrated gradients as the importance approximations are relatively consistent with the ground truth.
```
[X,_,A], y_true_all = all_gen[0]
N = A.shape[-1]
X_bk = deepcopy(X)
edges = [(graph_nodes.index(u),graph_nodes.index(v)) for u,v in G_ego.edges()]
nodes = [list(G.node()).index(v) for v in G_ego.nodes()]
selected_nodes = np.array([[target_idx]], dtype='int32')
clean_prediction = model.predict([X, selected_nodes, A]).squeeze()
predict_label = np.argmax(clean_prediction)
groud_truth_edge_importance = np.zeros((N, N), dtype = 'float')
groud_truth_node_importance = []
for node in nodes:
if node == target_idx:
groud_truth_node_importance.append(0)
continue
X = deepcopy(X_bk)
#we set all the features of the node to zero to check the ground truth node importance.
X[0, node, :] = 0
predict_after_perturb = model.predict([X, selected_nodes, A]).squeeze()
prediction_change = clean_prediction[predict_label] - predict_after_perturb[predict_label]
groud_truth_node_importance.append(prediction_change)
node_shapes = ['o' if groud_truth_node_importance[k] > 0 else 'd' for k in range(len(nodes))]
positive_colors, negative_colors = [], []
positive_node_sizes, negative_node_sizes = [], []
positive_nodes, negative_nodes = [], []
#node_size_scale is used for better visulization of nodes
node_size_scale = node_size_factor/max(groud_truth_node_importance)
for k in range(len(node_shapes)):
if nodes[k] == target_idx:
continue
if node_shapes[k] == 'o':
positive_colors.append(colors[k])
positive_nodes.append(graph_nodes[nodes[k]])
positive_node_sizes.append(node_size_scale*groud_truth_node_importance[k])
else:
negative_colors.append(colors[k])
negative_nodes.append(graph_nodes[nodes[k]])
negative_node_sizes.append(node_size_scale*abs(groud_truth_node_importance[k]))
X = deepcopy(X_bk)
for edge in edges:
original_val = A[0, edge[0], edge[1]]
if original_val == 0:
continue
#we set the weight of a given edge to zero to check the ground truth link importance
A[0, edge[0], edge[1]] = 0
predict_after_perturb = model.predict([X, selected_nodes, A]).squeeze()
groud_truth_edge_importance[edge[0], edge[1]] = (predict_after_perturb[predict_label] - clean_prediction[predict_label])/(0 - 1)
A[0, edge[0], edge[1]] = original_val
# print(groud_truth_edge_importance[edge[0], edge[1]])
fig, ax = plt.subplots(1, 1, figsize=(15, 10));
cmap = plt.get_cmap('jet', np.max(colors)-np.min(colors)+1)
# Draw the target node as a large star colored by its true subject
nx.draw_networkx_nodes(G_ego, pos, nodelist=[target_nid], node_size=50*abs(node_sizes[nodes.index(target_idx)]), node_color=[colors[nodes.index(target_idx)]], cmap=cmap,
node_shape='*', vmin=np.min(colors)-0.5, vmax=np.max(colors)+0.5, label="Target")
# Draw the ego net
nc = nx.draw_networkx_nodes(G_ego, pos, nodelist=positive_nodes, node_color=positive_colors, cmap=cmap, node_size=positive_node_sizes, with_labels=False,
vmin=np.min(colors)-0.5, vmax=np.max(colors)+0.5, node_shape='o')
nc = nx.draw_networkx_nodes(G_ego, pos, nodelist=negative_nodes, node_color=negative_colors, cmap=cmap, node_size=negative_node_sizes, with_labels=False,
vmin=np.min(colors)-0.5, vmax=np.max(colors)+0.5, node_shape='d')
edges = G_ego.edges()
#link_width_scale is used for better visulization of links
link_width_scale = link_width_factor/np.max(groud_truth_edge_importance)
weights = [link_width_scale*groud_truth_edge_importance[graph_nodes.index(u),list(G.node()).index(v)] for u,v in edges]
edge_colors = ['red' if groud_truth_edge_importance[graph_nodes.index(u),list(G.node()).index(v)] > 0 else 'blue' for u,v in edges]
ec = nx.draw_networkx_edges(G_ego, pos, edge_color=edge_colors, width = weights)
plt.legend()
plt.colorbar(nc, ticks=np.arange(np.min(colors),np.max(colors)+1))
plt.axis('off')
plt.show()
```
| github_jupyter |
```
from sketching import settings
from sketching.datasets import Dataset, Covertype_Sklearn, KDDCup_Sklearn, Webspam_libsvm, Synthetic_Dataset, NoisyDataset
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
if not settings.PLOTS_DIR.exists():
settings.PLOTS_DIR.mkdir()
def get_results_df(dataset: Dataset, time_column):
df_list = []
for method in ["uniform", "l2s", "sketching"]:
df = (
pd.read_csv(settings.RESULTS_DIR / (dataset.get_name() + f"_{method}.csv"))
.filter(items=[time_column, "size", "ratio"])
.groupby(["size"], as_index=False)
.median()
.assign(method=method)
)
df_list.append(df)
return(pd.concat(df_list, ignore_index=True))
get_results_df(Covertype_Sklearn(), "sampling_time_s")
def make_plot(dataset, x_min, x_max, y_min, y_max, sampling_time=False, font_size=18, font_size_title=23):
if sampling_time:
time_column = "sampling_time_s"
else:
time_column = "total_time_s"
results_df = get_results_df(dataset, time_column=time_column)
# use TeX for typesetting
plt.rcParams["text.usetex"] = True
plt.rc("font", size=font_size)
fig, ax = plt.subplots()
colormap = matplotlib.cm.get_cmap(name="tab20")
colors = {
"uniform": colormap(0),
"sketching": colormap(2),
"l2s": colormap(4),
}
labels = {
"uniform": "Uniform",
"sketching": "Sketch",
"l2s": "L2S",
}
titles = {
"covertype_sklearn": "Covertype",
"covertype_sklearn_noisy": "Covertype, 1\% noisy",
"kddcup_sklearn": "Kddcup",
"kddcup_sklearn_noisy": "Kddcup, 1\% noisy",
"webspam_libsvm_desparsed": "Webspam",
"webspam_libsvm_desparsed_noisy": "Webspam, 1\% noisy",
"synthetic_n_100000": "Synthetic"
}
for cur_method in ["l2s", "uniform", "sketching"]:
cur_results = results_df.loc[results_df["method"] == cur_method]
ax.scatter(
cur_results[time_column],
cur_results["ratio"],
color=colors[cur_method],
label=labels[cur_method],
)
ax.set_xlim(left=x_min, right=x_max)
ax.set_ylim(bottom=y_min, top=y_max)
if sampling_time:
ax.set_xlabel("median sampling time (s)")
else:
ax.set_xlabel("median absolute running time (s)")
ax.set_ylabel("median approximation ratio")
ax.set_title(titles[dataset.get_name()], fontsize=font_size_title)
legend = ax.legend(loc="upper right", frameon=True)
fig.tight_layout()
if sampling_time:
plt.savefig(settings.PLOTS_DIR / f"{dataset.get_name()}_sampling_time_plot.pdf")
else:
plt.savefig(settings.PLOTS_DIR / f"{dataset.get_name()}_total_time_plot.pdf")
plt.show()
dataset = Covertype_Sklearn()
make_plot(dataset, x_min=None, x_max=None, y_min=None, y_max=None, sampling_time=False)
make_plot(dataset, x_min=None, x_max=None, y_min=None, y_max=None, sampling_time=True)
dataset = KDDCup_Sklearn()
make_plot(dataset, x_min=None, x_max=None, y_min=1, y_max=10, sampling_time=False)
make_plot(dataset, x_min=None, x_max=None, y_min=1, y_max=6, sampling_time=True)
dataset = Webspam_libsvm()
make_plot(dataset, x_min=None, x_max=None, y_min=1, y_max=2, sampling_time=False)
make_plot(dataset, x_min=None, x_max=None, y_min=1, y_max=2, sampling_time=True)
dataset = Synthetic_Dataset(n_rows=100000)
make_plot(dataset, x_min=None, x_max=None, y_min=None, y_max=None, sampling_time=False)
make_plot(dataset, x_min=None, x_max=None, y_min=None, y_max=None, sampling_time=True)
dataset = NoisyDataset(dataset=Webspam_libsvm(), percentage=0.01, std=10)
make_plot(dataset, x_min=None, x_max=None, y_min=1, y_max=2, sampling_time=False)
make_plot(dataset, x_min=None, x_max=None, y_min=1, y_max=2, sampling_time=True)
dataset = NoisyDataset(dataset=Covertype_Sklearn(), percentage=0.01, std=10)
make_plot(dataset, x_min=None, x_max=None, y_min=1, y_max=1.5, sampling_time=False)
make_plot(dataset, x_min=None, x_max=None, y_min=1, y_max=1.5, sampling_time=True)
dataset = NoisyDataset(dataset=KDDCup_Sklearn(), percentage=0.01, std=10)
make_plot(dataset, x_min=None, x_max=None, y_min=1, y_max=5, sampling_time=False)
make_plot(dataset, x_min=None, x_max=None, y_min=1, y_max=5, sampling_time=True)
```
| github_jupyter |
Introduction:
In the realm of sports betting, it is very difficult to make consistent profit. Sportsbooks intentionally create odds so that the general public is as close to a 50/50 split on a given game. Therefore, the sportsbooks try to predict the final outcome as accurately as possible. Each bet typically incurs a 10% fee that the sportsbooks keeps when the bettor loses. Therefore, in order to make any profit as a bettor you have to maintain higher than 53% accuracy. Simply relying on a coinflip will always lose money in the long-run. What if there was a way to beat the odds that the sportsbooks create, 53% of the time or better? This information could either be very valuable for professional sports bettors or sportsbooks themselves, if I can find a flaw in their algorithms.
Unfortunately, due to the high financial stakes involved in such scenarios, any other organizations who do work like this keep their work as proprietary information. Everyone claims to have the “winning” formula yet force you to pay for their services before seeing if the predictions are accurate or not. Additionally, once you sign up for one of these services, they provide suggestions of things to bet on. They do not reveal the data that are considered within their internal workings. Therefore, I will have to compare my work against that of the odds set by Vegas. Again, the algorithms they use are proprietary, so I cannot compare my means. Instead I will compare the ends, namely how often I can accurately predict the outcome of a game.
I will specifically look at the NBA. The most important variable that I hope to investigate is the number of minutes a starting lineup for a given NBA team, and its effects on their performance in future games. The fundamental question I will be asking is: Does the amount of minutes a starting lineup for a given NBA team have an effect on their individual performances in subsequent games, and how does this effect (if any) affect the overall performance of the team in terms of winning or losing the game. I will also compare the outcomes with the outcomes Against the Spread (ATS). ATS refers to how a team performs against the odds that sportsbooks set to try and level the playing field. If my hypothesis of minutes played having a role in determining the outcome ATS is correct, I will be able to predict the outcome more than 53% of the time using these data and determine that it is a variable not considered by sportsbooks when determining the spread.
Additionally, the information prior to the comparison of ATS could be useful to NBA teams themselves, as monitoring minutes could influence overall team performance. Some issues with the dataset arise with missing data, as not every player records every type of stat every time they play. Typically, it would make sense to place “zeros” for these events but for variables that are percentages (typically regarding shooting accuracy) it might make more sense to place their season average, in order to avoid creating synthetic outliers.
Clearly the most important variables that I will be looking at are minutes played, wins and losses, and wins and losses ATS. However, there are many other variables that will be considered when determining effect on individual player performance. Here is an explanation of each variable:
Age -- Age of the player at the start of February 1st of that season
Pos -- Position of a given player
Tm -- Team of the player
Opp -- Opponent for the given game
GS -- Confirms that the player started the game
MP -- Minutes played by the individual player for that specific game
FG -- Field Goals (how many shots the player made in the given game, excluding free throws)
FGA -- Field Goal Attempts (how many shots the player attempted in the given game, excluding free throws)
FG% -- Field Goal Percentage (the percentage of shots the player made in the given game, excluding free throws)
2P -- 2-Point Field Goals (how many 2-point shots the player made in the given game)
2PA -- 2-point Field Goal Attempts (how many 2-point shots the player attempted in the given game)
2P% -- 2-Point Field Goal Percentage (the percentage of 2-point shots the player made in the given game)
3P -- 3-Point Field Goals (how many 3-point shots the player made in the given game)
3PA -- 3-Point Field Goal Attempts (how many 3-point shots the player attempted in the given game)
3P% -- 3-Point Field Goal Percentage (the percentage of 3-point shots the player made in the given game)
FT -- Free Throws (how many free throws were made in a game)
FTA -- Free Throw Attempts (how many free throws were attemped in a game)
FT% -- Free Throw Percentage (the percentage of free throws were made in a game)
ORB -- Offensive Rebounds (amount of rebounds secured by a given player while his team is on offense)
DRB -- Defensive Rebounds (amount of rebounds secured by a given player while his team is on defense)
TRB -- Total Rebounds (combined total of offensive and defensive rebounds for a given player)
AST -- Assists (a pass that a player makes, leading to a teammate scoring)
STL -- Steals (taking the ball away from an opponent)
BLK -- Blocks (blocking the shot of an opponent)
TOV -- Turnovers (losing the ball to an opponent)
PF -- Personal Fouls
PTS -- Points
GmSc -- Game Score is a combination of many variable to determine a player’s overall positive impact on his team. The formula is as follows: (Points x 1.0) + (FGM x 0.4) + (FGA x -0.7) + ((FTA-FTM) x -0.4) + (OREB x 0.7) + (DREB x 0.3) + (STL x 1.0) + (AST x 0.7) + (BLK x 0.7) + (PF x -0.4) + (TO x -1.0).
Datasheet for NBA Fatigue Effects
Motivation for Dataset Creation
The main motivation behind the use of this dataset is to find a consistent determining factor of NBA games when comparing the outcome to Against the Spread (ATS) odds set up by sportsbooks. Most algorithms are proprietary due to the financial impacts they can have. However, there are some sharks (professional bettors) who have at least disclosed some of the metrics they consider. A common metric for the NBA is fatigue. This is based on the pace of previous games, determined by number of possessions, over a given number of days, with or without rest. However, only considering the number of possessions of a given team does not necessarily forecast fatigue, because it is not considering who is participating in these possessions. Therefore, this dataset looks at the minutes played by the starting five players of NBA teams, and the impact on future performances. The dataset focuses on starters, because they generally play at least twice as much as the bench players and should have a larger impact on the outcome of games.
Of course, the proposed use of this dataset is not exhaustive. It can also be used by the teams themselves. Coaches can use the data to determine each player’s threshold of minutes that cause the player to be less productive in subsequent games. This information can be invaluable when managing minutes throughout the season and within specific games. Also, fine tuning the rotation come time for playoffs can be very critical to maximize winning opportunities within important games.
The dataset was extracted from a much larger data source supplied by www.basketball-reference.com. This is a very common, public source of data for the NBA, so it has been used for many applications[1,2]. However, the specific dataset that I extracted was based on certain search parameters regarding players that started a game and played more than 15 minutes. I have not found any cited work using that specific parameter.
Basketball Reference is a part of a larger umbrella website, Sports Reference. This site prides itself on “providing both basic and sabermetric statistics and resources for sports fans everywhere”. Their mission is “to be the easiest-to-use, fastest, most complete sources for sports statistics anywhere”. In order to fund this project, Sports Reference uses lots of advertisements on its site. It also offers premium, paid data packages of the most up-to-date data collected while the games are occurring. It also offers a newsletter that one can sign up for that features more advertisements.
Dataset Composition
There are 18,478 instances, all of which represent an individual player’s performance during one game. These instances can be further categorized by all pertaining to a certain team, a certain game, or all games of a certain player. The amount for each of these types is variable based on how many times a given player was a starter for a specific team. This can be determined by performance, injury, or rest. The maximum for a given game is 5 (per team), for a given player is 82 (there are 82 games in a season), and for a given team is 410 (5 starters playing 82 games).
Most of the necessary data resides within this dataset. However, if interesting trends are determined, it might be useful to incorporate data collected for future seasons to create a predictive model. Additionally, it is necessary to import the spread for each game from an external website as this data only shows the raw data from games, but it fails to compare those results to the ATS outcome.
Data Collection Process
Initial NBA data collection is recorded by humans sitting courtside. Each professional team has at least one paid employee that records every stat for the five players on the court while the game is in progress, based on his/her direct observations. The average salary for these individuals is about $71,000, with a cap at about $85,000[3]. From there, the NBA releases the data to the public domain. Unfortunately, Basketball Reference does not disclose its specific collection process.
The final dataset includes all game logs of starters in the NBA who played at least 15 minutes. This means that some instances of stat lines for starters in the NBA this season were likely excluded. However, the typical starter in the NBA plays about 25-35 minutes per game. 15 is far below this average, and very uncommon considering a full game includes 48 minutes. The exclusion of instances with less than 15 minutes was intentional for a few reasons. First, and most obvious, it made the dataset a bit smaller and easier to manipulate. More importantly, if a starter is playing less than 15 minutes it is usually due to an injury. Even in the case where it is not an injury, playing less than 15 minutes does not have an impact on the initial thesis that extra fatigue could impact future performance.
Since the raw data is collected by humans and based on their direct observations in a chaotic and fast-paced environment, there are likely errors in the data. These errors would likely be random and nearly impossible to discover, unless one watched every second of every game to ensure the accuracy. However, this should not affect the goal of this dataset. The two main focuses, minutes and final outcome, will not be affected by such human error. The final outcome should be fairly intuitive as it is what everyone is extremely focused on. The minutes stat should be extremely accurate because in basketball players are only allowed to be substituted at stoppages in play. Additionally, there are only 5 players for each time on the court at a given time. With that said, it is very simple to keep track of this stat.
Data Preprocessing
It is unclear what data processing techniques may or may not have been used for this dataset. However, since it is data based directly on the events of basketball games, it is unlikely that much was done. Any future processing in relation to the scope of this project is still to be determined.
Dataset Distribution
As previously mentioned, the data is first published by the NBA and then collected second hand by Sports reference for ease of manipulation. As stated on the Sports Reference website, there are licensing agreements for some of their datasets to be used for paid services, however there is not a restriction on simply using the data. To more easily export the datasets on the site, Sports Reference will complete custom requests for a minimum of $1000. However, they encourage users to learn how to extract the data themselves.
Dataset Maintenance
The data set is maintained by the Basketball Reference team. Stats of games are posted within 24 hours of the conclusion, and often much quicker. The only way that the data would become obsolete is if the NBA disbanded, which seems extremely unlikely at this point in time, as the league is probably in the strongest position it has ever been. Basketball players have surpassed baseball players as the highest paid athletes in the country. Additionally, there is neither a repository to link use of the dataset, nor a feature to add to it.
Legal and Ethical Considerations
The component of sports betting certainly adds a legal layer to the use of this dataset. As of last year, 2018, sports betting is now federally legal. It is now up to each state to determine if they will permit this type of gambling. Currently only Nevada and New Jersey allow for mobile sports wagers. However, Pennsylvania, Rhode Island, Delaware, West Virginia, Mississippi, and New Mexico allow for in-person wagers[4]. Pennsylvania, Rhode Island, West Virginia, and Mississippi all have recently passed bills to allow for mobile wagers[5]. Many other states are expected to follow this trend. However, the actual legality of sports betting has little effect on the intended use of this data set. There are many apps and national television shows that provide gambling in such as: Action Network, Locked In, and The Sports Geek
```
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from scipy import stats
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.model_selection import RandomizedSearchCV
from sklearn.model_selection import ParameterGrid
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.svm import l1_min_c
from sklearn.metrics import roc_curve, auc
sns.set_context('talk')
sns.set_style('darkgrid')
sns.set_palette('colorblind')
NBAstats = pd.read_csv("NBA_stats_full.csv")
#import csv file
NBAstats.head()
#check accuracy
```
The dataset has strange player identifiers attached to the name. To clean it up I will need to split the string
```
NBAstats["Player"]= NBAstats["Player"].str.split("\\", n = 1, expand = True)
NBAstats.head()
#separate the \playerID from each Name
#NBAstats["Starter"] = NBAstats1[0]
#new column for name
NBAstats["Date"]= NBAstats["Date"].astype(str).str[:-3]
NBAstats.head()
```
The dataset that I wish to merge with this initial one (NBAstats) does not have years in the date column, so for a successful merge I needed to remove the end of end string starting with the final "/".
```
#NBAstats.drop(columns = ["Player"],inplace = True)
#delete initial column
NBAstats.head()
NBAstatsFinal= NBAstats.rename(columns= {'Tm': 'Team'})
#rename to match merge
NBAstatsFinal.head()
#NBAstatsFinal.count
#check rows for future reference
NBAodds = pd.read_csv("NBA_odds.csv")
NBAodds.head()
NBAoddsFix = pd.read_csv("NBA_odds.csv", index_col ="Team")
NBAoddsFix.rename(index = {"Atlanta": "ATL", "Brooklyn": "BKN", "Boston": "BOS", "Charlotte": "CHA", "Chicago": "CHI", "Cleveland": "CLE", "Dallas": "DAL", "Denver": "DEN", "Detroit": "DET", "Golden State": "GSW", "GoldenState": "GSW", "Houston": "HOU", "Indiana": "IND", "LAClippers": "LAC", "LA Clippers": "LAC", "LALakers": "LAL", "LA Lakers": "LAL", "Memphis": "MEM", "Miami": "MIA", "Milwaukee": "MIL", "Minnesota": "MIN", "NewOrleans": "NOP", "NewYork": "NYK", "New York": "NYK", "OklahomaCity": "OKC", "Oklahoma City": "OKC", "Orlando": "ORL", "Philadelphia": "PHI", "Phoenix": "PHO", "Portland": "POR", "Sacramento": "SAC", "San Antonio": "SAS", "SanAntonio": "SAS", "Toronto": "TOR", "Utah": "UTA", "Washington": "WAS"}, inplace = True)
#NBAoddsFix
#checked extra to make sure they all switched
```
This second dataset (NBAodds) has all of the team names spelled out. However, the first dataset has the abbreviations. I needed to go through and change them all. Additionally, teams like OklahomaCity and GoldenState had errors in the dataset as some were spelled with a space and some without. I noticed this after double checking more than just the .head(). So for every multi-word team I renamed both versions.
```
final_df = pd.merge(NBAstatsFinal, NBAoddsFix, how='left', left_on=['Team', 'Date'], right_on = ['Team', 'Date'])
final_df.head()
```
Team and date were the two consistent variables between the datasets so that is what I used to merge them.
```
final_df.rename( columns={'Unnamed: 6':'Location','Unnamed: 8':'Result'}, inplace=True )
#fix unnamed columns
final_df.head()
#final_df.count
#ensuring same instances
pd.DataFrame(final_df.MP.describe())
```
When tryoing to visualize with a bar graph there were clearly too many points to see, I will need segment into buckets with a histogram.
```
final_df['MP'].hist(bins=np.arange(15,60,3))
plt.xlabel('Minutes Played')
plt.ylabel('Number of Games')
#visualizing distribution of minutes being played
```
There's a clear dropoff at some point (perhaps around 37 that will be interesting to look at).
```
final_df['MP'].hist(bins=np.arange(33,57,3))
plt.xlabel('Minutes Played')
plt.ylabel('Number of Games')
#visualizing distribution of minutes being played for top 25%
final_df['MP'].hist(bins=np.arange(41,57,3))
plt.xlabel('Minutes Played')
plt.ylabel('Number of Games')
#visualizing distribution of minutes being played for over 2 std above mean
```
In order to sequence games in the future, which I will need if I wish to correlate minutes played in one game, with the future performance of the next, I need the dates to be numbers instead of strings.
```
Usagedata = {'Player': final_df['Player'], 'Date': final_df['Date'], 'High Usage' : final_df['MP'] >= 41, 'Normal' : final_df['MP'] < 41}
Usage_df= pd.DataFrame(Usagedata)
Usage_df.head()
pd.DataFrame(Usage_df.Player.describe())
final_df.head()
final_df = pd.merge(final_df, Usage_df, how='left', left_on=['Player', 'Date'], right_on = ['Player', 'Date'])
final_df.head()
```
I decided to split the date into months and days so that I could mathematically manipulate each datapoint and add it back together to give each day a unique number.
```
#pd.DataFrame(final_df.Player.describe())
#filter1 = final_df.Player == 'Jerami Grant'
#JG_df = final_df[filter1]
#JG_df
final_df['Date'].dtype.kind
#date split wasnt working, checked type to find its an object
#need to swith to string
#DateFix = {'': final_df['Date'], 'Date': final_df['Date']}
DateFix = {'Date': final_df['Date'], 'Player': final_df['Player']}
DateFix_df = pd.DataFrame(DateFix)
DateFix_df[['Month','Day']] = DateFix_df.Date.str.split("/",expand=True,)
#separated month and day to be able to do math and give each day differentiation
DateFix_df.head()
#pd.DataFrame(DateFix_df.Player.describe())
#DateFix_df['Month'].dtype.kind
#is obj, need int
DateFix_df['Month']= DateFix_df['Month'].astype(float)
DateFix_df['Month'] -= 1
DateFix_df['Month'] *= 31
DateFix_df['Day']= DateFix_df['Day'].astype(float)
DateFix_df.head()
```
I subtracted 1 from each month so that the starting point for January would be zero and the days could be added afterwards. Although all months do not have 31 days, using this as a multiple ensures that there are not any overlaps (ex: Jan 31 and Feb 1). Had I used 30, Jan 31 and Feb 1 would both have a value of 31. The actual value doesn't matter, because the games do not necessarily happen on consecutive days anyway. For future analysis it is only important that the days with correlating games are in order.
```
DateFix_df['Day'] = DateFix_df['Month'] + DateFix_df['Day']
DateFix_df.head()
final_df = pd.merge(final_df, DateFix_df, how='left', left_on=['Player', 'Date'], right_on = ['Player', 'Date'])
final_df.head()
#pd.DataFrame(final_df.Player.describe())
#realized that I do not really need all of the columns to look at relationship between
machine = {'Day': final_df['Day'], 'Team': final_df['Team'], 'Player': final_df['Player'], 'GmSc': final_df['GmSc'], 'Minutes': final_df['MP'], 'Fatigue': final_df['High Usage'], 'Rest': final_df['Normal'], 'Result': final_df['Result'] }
machine_df = pd.DataFrame(machine)
#creates new df for machine learning
pd.DataFrame(machine_df.Player.describe())
machine_df.head()
f, ax = plt.subplots(figsize=(10,4))
((machine_df.isnull().sum()/len(machine_df)) * 100).plot(kind='bar')
plt.xticks(rotation=45, horizontalalignment='right')
plt.title('Percent Missing by Variable')
#no missing data (:
filter1 = final_df.Player == 'Jerami Grant'
JG_df = final_df[filter1]
JG_df
final_df.groupby(['MP', 'GmSc'])['Result'].count()
```
A Pivot table is clearly not useful here, as there are far too many unique instances.
```
#final_df.plot.scatter('MP', 'GmSc')
plt.subplots(figsize=(6.5, 6.5))
sns.regplot('MP', 'GmSc', data=final_df, marker='.',
scatter_kws={'alpha':'0.2'}, line_kws={'color':'orange'}, truncate=True)
```
There appears to be a correlation between how long a player plays and how well they perform.
```
f, ax = plt.subplots(figsize=(6.5, 6.5))
sns.boxplot(x="Result", y="MP", data=final_df, fliersize=0.5, linewidth=0.75, ax=ax)
```
The minutes played by the startes does not seem to indicate the result of the game. However, this does not mean that the minutes do not have an affect on future performances. The effect can be delayed.
```
f, ax = plt.subplots(figsize=(6.5, 6.5))
sns.boxplot(x="Result", y="FG%", data=final_df, fliersize=0.5, linewidth=0.75, ax=ax)
```
The shooting efficiency of the starting line up seem to have a strong effect on the outcome of them game and might be good to consider in future analyses.
```
f, ax = plt.subplots(figsize=(6.5, 6.5))
sns.boxplot(x="Result", y="TOV", data=final_df, fliersize=0.5, linewidth=0.75, ax=ax)
```
While the median doesn't show a difference, there are clearly more turnovers associated with losses.
```
f, ax = plt.subplots(figsize=(6.5, 6.5))
sns.boxplot(x="TOV", y="MP", data=final_df, fliersize=0.5, linewidth=0.75, ax=ax)
```
The longer a starter plays, the more turnovers they tend to commit. This makes sense. As fatigue sets in, a player is likely to be more careless with the ball.
```
f, ax = plt.subplots(figsize=(6.5, 6.5))
sns.boxplot(x="Result", y="GmSc", data=final_df, fliersize=0.5, linewidth=0.75, ax=ax)
f, ax = plt.subplots(figsize=(6.5, 6.5))
sns.boxplot(x="Result", y="GmSc", data=final_df, fliersize=0.5, linewidth=0.75, ax=ax)
```
While the length of time each starter plays for did not have an effect on the Result, their efficiency determined by the Game Score does.
```
fig, ax = plt.subplots()
final_df['MP'].hist(bins=np.arange(41,57,3))
final_df['GmSc'].hist(bins=np.arange(41,57,3))
plt.xlabel('Minutes Played')
plt.ylabel('Number of Games')
```
As the starters play better it leads to more victories.
What I would like to look at with the ML, is what affects future games. For future discussion I would need a loop that finds the next game and associated that result with the minutes played in the previous instance. However, I was unable to accurately do this because there are multiple players for a given team and multiple teams playing games on a given day.
Every time I attempt to bring in the ML code it somehow ruins all of my previous code, and I then have to go back and fix it all over again. I'm not sure why this happens, but for that reason I have ommitted it to preserve my work above, since it did not work anyway.
References
Charles South, Ryan Elmore, Andrew Clarage, Rob Sickorez, Jing Cao. (2017) A Starting Point for Navigating the World of Daily Fantasy Basketball. The American Statistician 0:0, pages 1-7.
Balaban, Marc. “Exploring 13 Seasons (7,588,492 Plays) of The NBA in Real Time.” Exploring 13 Seasons (7,588,492 Plays) of The NBA in Real Time, 20 Sept. 2017, www.omnisci.com/blog/exploring-13-seasons-of-the-nba/.
Salary.com. “Salary for NBA Statistician in Greenwood, IN.” Salary.com, 2019, www.salary.com/research/salary/employer/nba/statistician-salary/greenwood-in.
Rodenberg, Ryan. “State-by-State Sports Betting Bill Tracker.” ESPN, ESPN Internet Ventures, 12 Mar. 2019, www.espn.com/chalk/story/_/id/19740480/gambling-sports-betting-bill-tracker-all-50-states.
Saville, Kendall. “US Sports Betting Sites - Legal US Online Sports Betting in 2019.” Legal Sports Report, Legal Sports Report, 19 Feb. 2019, www.legalsportsreport.com/us-betting-sites/.
| github_jupyter |
# Simple power spectrum estimation from an input dataset
This example shows how to estimate the power spectrum from a set of data files using an Optimal Quadratic Estimator (OQE) approach.
```
%matplotlib inline
from pyuvdata import UVData
import hera_pspec as hp
import numpy as np
import matplotlib.pyplot as plt
import copy, os, itertools, inspect
from hera_pspec.data import DATA_PATH
```
## Loading the input data: forming power spectra from adjacent time integrations
The input data are specified as a list of `UVData` objects, which are then packaged into a `PSpecData` class. This class is responsible for collecting the data and covariances together and performing the OQE power spectrum estimation.
At least two `UVData` objects must be specified, these could be different datasets, or just copies of a single dataset, given the use-case. In what follows, we will use only one data set and produce two copies of it, but will shift the second dataset by one time integration and interleave it relative to the first, such that we can form auto-baseline power spectra without noise-bias.
```
# select the data file to load
dfile = os.path.join(DATA_PATH, 'zen.all.xx.LST.1.06964.uvA')
# Load into UVData objects
uvd = UVData()
uvd.read_miriad(dfile)
# Check which baselines are available
print(uvd.get_antpairs())
```
## Define a cosmology
Here we will instantiate a `hera_pspec.conversions.Cosmo_Conversions` object that will define the cosmology we adopt throughout this notebook. **All calculations that depend on a cosmology will be tied to the one we adopt here: the various objects in `hera_pspec` will pass this specific object between themselves, which starts by attaching the cosmology to a `PSpecBeamUV` object.** Further on in the notebook, we will see how we can explicitly overwrite the adopted cosmology and appropriately re-calculate the necessary variables.
```
# Instantiate a Cosmo Conversions object
# we will need this cosmology to put the power spectra into cosmological units
cosmo = hp.conversions.Cosmo_Conversions()
print(cosmo)
```
Instantiate a beam object, and attach the cosmo conversions object onto it.
```
# List of beamfile to load. This is a healpix map.
beamfile = os.path.join(DATA_PATH, 'HERA_NF_dipole_power.beamfits')
# intantiate beam and pass cosmology, if not fed, a default Planck cosmology will be assumed
uvb = hp.pspecbeam.PSpecBeamUV(beamfile, cosmo=cosmo)
```
Next convert from Jy units to mK. This involves calculating the effective beam area (see HERA Memo #27 and #43), which can be done with the beam object we instantiated earlier.
```
# find conversion factor from Jy to mK
Jy_to_mK = uvb.Jy_to_mK(np.unique(uvd.freq_array), pol='XX')
# reshape to appropriately match a UVData.data_array object and multiply in!
uvd.data_array *= Jy_to_mK[None, None, :, None]
```
Configure data and instantiate a `PSpecData` object, while also feeding in the beam object.
```
# slide the time axis of uvd by one integration
uvd1 = uvd.select(times=np.unique(uvd.time_array)[:-1:2], inplace=False)
uvd2 = uvd.select(times=np.unique(uvd.time_array)[1::2], inplace=False)
# Create a new PSpecData object, and don't forget to feed the beam object
ds = hp.PSpecData(dsets=[uvd1, uvd2], wgts=[None, None], beam=uvb)
```
## Phase second `dset` to first `dset`
Because we are forming power spectra between datasets that are offset in LST there will be some level of decoherence (and therefore signal loss) of the EoR signal. For short baselines and small LST offsets this is typically negligible, but it is still good to try to recover what coherency we can, simply by phasing (i.e. fringe-stopping) the datasets before forming the power spectra. This can be done with the `rephase_to_dset` method, and can only be done once.
```
# Because the LST integrations are offset by more than ~15 seconds we will get a warning
# but this is okay b/c it is still **significantly** less than the beam-crossing time and we are using short
# baselines...
# here we phase all datasets in dsets to the zeroth dataset
ds.rephase_to_dset(0)
# change units of UVData objects
ds.dsets[0].vis_units = 'mK'
ds.dsets[1].vis_units = 'mK'
```
## Estimating the power spectrum for a handful of baseline pairs (auto-baseline pspec)
Estimate the power spectrum for a handful baseline pairs between the two datasets in `ds.dsets`. You can specify which baselines are included in the power spectrum estimate, which datasets to use, what freq channels to use, and how the estimate should be weighted.
```
# Specify which baselines to include
baselines = [(24,25), (37,38), (38,39)]
```
### Read the docstring! Here are the first few lines...
```
print("ds.pspec{}".format(inspect.signature(ds.pspec)))
print('\n'.join(ds.pspec.__doc__.split('\n')[:30]))
```
Call the `PSpecData.pspec` function to use the OQE framework. The result is a `UVPSpec` object that holds all of the power spectra and their meta-data.
```
# we will use the baselines list to produce 3 power spectra
# whose data will be drawn from the dsets[0] and dsets[1]
# across two spectral windows with identity weighting and a blackman-harris taper
uvp = ds.pspec(baselines, baselines, (0, 1), [('xx', 'xx')], spw_ranges=[(300, 400), (600,721)], input_data_weight='identity',
norm='I', taper='blackman-harris', verbose=True)
```
## Making sense of the meta-data
The fundamental unit in the `UVPSpec` object is a single delay spectrum, indexed by a spectral window selection (`spw`), a baseline-pair selection (`blpair`) and a polarization pair (`polpair`). Spectral windows are marked by their index (see `spw_array`), polarization pairs are marked by a pair of pol strings `('xx', 'xx')` or pol integers `(-5, -5)`, and a baseline-pair is marked by its blpair integer, which is simply the antenna numbers put into length-3 integers and concatenated. For example, the baseline-pair `((100, 200), (300, 400))` would have a blpair integer `100200300400`. Polpair tuples can be converted into integers in a similar way.
To access particular slices of the data in a `UVPSpec` object, the user should interface with the `get_data` method, which takes a selection of `spw`, `blpair` and `pol` as arguments. See the example below.
```
# let's get the delay spectra from spw 0, blpair ((24, 25), (24, 25)) and xx pol
key = (0, ((24, 25), (24, 25)), ('xx', 'xx'))
# output should be shape (Ntimes, Ndlys)
print(uvp.get_data(key).shape)
# we can also access data by feeding a dictionary
key = {'polpair':('xx','xx'), 'spw': 0, 'blpair':((24, 25), (24, 25))}
print(uvp.get_data(key).shape)
```
There is also metadata telling you things like the data weighting, normalization, tapering, units, telescope location, LST and JD time stamps, etc.
```
# get power spectrum units
print("pspec units: ", uvp.units)
# get weighting
print("pspec weighting: ", uvp.weighting)
```
Also, importantly, the cosmology we adopted originally and passed through to the beam and `PSpecData` objects was passed through to this `UVPSpec` object.
```
print(uvp.cosmo)
```
## Plotting
Here is an example of how to do some simple and quick plotting of power spectra, which includes getting the power spectra and the delay bins of their bandpowers. We can use the shorthand `'xx'` to denote the polarization pair `('xx', 'xx')`.
```
# plot power spectrum of spectral window 1
fig, ax = plt.subplots(figsize=(12,8))
spw = 1
blp =((24, 25), (24,25))
key = (spw, blp, 'xx')
dlys = uvp.get_dlys(spw) * 1e9
power = np.abs(np.real(uvp.get_data(key)))
p1 = ax.plot(dlys, power.T)
ax.set_yscale('log')
ax.grid()
ax.set_xlabel("delay [ns]", fontsize=14)
ax.set_ylabel(r"$P(k)\ \rm [mK^2\ h^{-3}\ Mpc^3]$", fontsize=14)
ax.set_title("spw : {}, blpair : {}, pol : {}".format(*key), fontsize=14)
```
## Form redundant baseline spectra (auto & cross baseline pspec)
Above we looked at auto-baseline power spectra. Here we will show how to form power spectra between groups of redundant baselines. To do this, we use the `pspecdata.construct_blpairs` helper function to construct the `baselines1` and `baselines2` lists that we then feed into `PSpecData.pspec`. This takes keywords `exclude_auto_bls`, which will remove all baselines paired with itself from the final `blpairs` list, as well as `exclude_permutations`, which will use a combination instead of a permutation to form the baseline-pairs from the input `baselines` list.
```
# baselines is a redundant baseline group
baselines = [(24,25), (37,38), (38,39)]
# calculate all baseline pairs from group
baselines1, baselines2, blpairs = hp.utils.construct_blpairs(baselines, exclude_auto_bls=True,
exclude_permutations=True)
# Inspect baseline pairs
print(blpairs)
```
Now we feed these lists through to `pspec`
```
uvp = ds.pspec(baselines1, baselines2, (0, 1), [('xx', 'xx')], spw_ranges=[(300, 400), (600,721)], input_data_weight='identity',
norm='I', taper='blackman-harris', verbose=True)
# plot power spectrum of a cross-baseline pspectra from spectral window 1
fig, ax = plt.subplots(figsize=(12,8))
spw = 1
blp =((24, 25), (37, 38))
key = (spw, blp, 'xx')
dlys = uvp.get_dlys(spw) * 1e9
power = np.abs(np.real(uvp.get_data(key)))
_ = ax.plot(dlys, power.T)
ax.set_yscale('log')
ax.grid()
ax.set_xlabel("delay [ns]", fontsize=14)
ax.set_ylabel(r"$P(k)\ \rm [mK^2\ h^{-3}\ Mpc^3]$", fontsize=14)
ax.set_title("spw : {}, blpair : {}, pol : {}".format(*key), fontsize=14)
```
## Convert to $\Delta^2(k)$
You can convert the power spectrum data from $P(k)$ format to "unitless" power spectrum $\Delta^2(k)$ by scaling by the cosmological wave-vectors cubed. This can be done by hand with the `get_kparas()` and `get_kperps()` methods, or can be done automatically using the `convert_to_deltasq()` method.
The cosmology used for this calculation will be the one stored in the `UVPSpec` object, which, recall, is the same cosmology we adopted originally.
```
# make a copy of the UVPSpec object
dsq = copy.deepcopy(uvp)
# how to get the wave vectors for spw = 1 by hand
k_perp, k_para = dsq.get_kperps(1, little_h=True), dsq.get_kparas(1, little_h=True)
# convert to deltasq automatically
dsq.convert_to_deltasq()
# see that units have automatically changed
print("dsq units: ",dsq.units)
# plot deltasq power spectrum of spw 1
fig, ax = plt.subplots(figsize=(12,8))
spw = 1
blp =((24, 25), (37, 38))
key = (spw, blp, 'xx')
power = np.abs(np.real(dsq.get_data(key)))
_ = ax.plot(k_para, power.T)
ax.set_yscale('log')
ax.grid()
ax.set_xlabel(r"$k_{\parallel}\ h\ Mpc^{-1}$", fontsize=14)
ax.set_ylabel(r"$\Delta^2(k)\ \rm [mK^2]$", fontsize=14)
ax.set_title("spw : {}, blpair : {}, pol : {}".format(*key), fontsize=14)
```
## Plotting Thermal Noise Curves
Using the `hera_pspec.noise` module, you can plot analytic thermal noise power spectra given a $T_{\rm sys}$ and a `UVPSpec` object, which carries with it the integration time and number of incoherent averages for **each delay spectrum** it contains.
To do this, all you need to do as the user is to interface with the `UVPSpec.generate_noise_spectra()` method. Given a selection of spectral window, polarization, system temperature and optionally which blpairs to compute for, it will return a dictionary of thermal noise curves (in either $P(k)$ or $\Delta^2(k)$ depending on `form` kwarg) for each baseline-pair in the `UVPSpec` object.
```
# generate noise curves for spw 0, pol 'xx' with a Tsys of 300 K
spw = 1
pol = 'xx'
Tsys = 300
P_N = uvp.generate_noise_spectra(spw, pol, Tsys)
```
Have generated the `Sense` object, we can now call `generate_noise_spectra` to produce noise curves. The result is a dictionary with baseline-pair integers as keys and ndarrays as values.
```
# plot power spectrum of spw 1 with noise curves
fig, ax = plt.subplots(figsize=(12,8))
blp = ((24, 25), (37, 38))
key = (spw, blp, pol)
power = np.abs(np.real(uvp.get_data(key)))
_ = ax.plot(k_para, power.T)
ax.set_yscale('log')
ax.grid()
ax.set_xlabel(r"$k_\parallel\ \rm h\ Mpc^{-1}$", fontsize=14)
ax.set_ylabel(r"$P(k)\ \rm [mK^2\ h^{-3}\ Mpc^3]$", fontsize=14)
ax.set_title("spw : {}, blpair : {}, pol : {}".format(*key), fontsize=14)
# plot noise curves for this baseline-pair
ax.set_prop_cycle(None)
_ = ax.plot(k_para, P_N[uvp.antnums_to_blpair(blp)].T, ls='--', lw=3)
```
## Incoherent Averaging
The `UVPSpec` object supports incoherent averaging across baseline-pairs and/or time using the `UVPSpec.average_spectra()` method. If the baseline-pairs are grouped by redundancy, this is equivalent to binning onto a single 3D $k_x, k_y, k_\parallel$ space. If the baseline-pairs are grouped by baseline separation, this is equivalent to cylindrical binning in $k_\perp$ and $k_\parallel$. Spherical binning to form a true 1D power spectrum is currently not supported.
Below we will incoherently average across all baseline-pairs in the `UVPSpec` object, knowing that we only constructed 14.6 m power spectra, so we are reasonably justified in doing so.
```
# form the baseline-pair group, which will be a single group
# consisting of all baseline-pairs in the object
blp_group = [sorted(np.unique(uvp.blpair_array))]
# average spectra with inplace = False and assign to a new "uvp2" object
uvp2 = uvp.average_spectra(blpair_groups=blp_group, time_avg=True, inplace=False)
```
We can then plot the averaged power spectrum, along with a new noise curve which will know to account for the averaging based on the changes to the `integration_array` and `nsample_array`. The colored lines show the same un-averaged pspectra from before, while the black line shows the averaged power spectrum, along with its new thermal noise estimate.
```
# plot power spectrum of spw 1
fig, ax = plt.subplots(figsize=(12,8))
spw = 1
blp = ((24, 25), (37, 38))
pol = 'xx'
key = (spw, blp, pol)
k_para = uvp.get_kparas(spw)
power = np.abs(np.real(uvp.get_data(key)))
P_N = uvp.generate_noise_spectra(spw, pol, 300)
P_N = P_N[uvp.antnums_to_blpair(blp)]
avg_power = np.abs(np.real(uvp2.get_data(key)))
avg_P_N = uvp2.generate_noise_spectra(spw, pol, 300)
avg_P_N = avg_P_N[uvp2.antnums_to_blpair(blp)]
_ = ax.plot(k_para, power.T)
ax.plot(k_para, avg_power.T, color='k')
ax.plot(k_para, avg_P_N.T, color='k', ls='--', lw=3)
ax.set_yscale('log')
ax.grid()
ax.set_xlabel(r"$k_\parallel\ \rm h\ Mpc^{-1}$", fontsize=14)
ax.set_ylabel(r"$P(k)\ \rm [mK^2\ h^{-3}\ Mpc^3]$", fontsize=14)
ax.set_title("spw : {}, blpair : {}, pol : {}".format(*key), fontsize=14)
ax.set_prop_cycle(None)
_ = ax.plot(k_para, P_N.T, ls='--', lw=3)
```
## Fold the power spectra
The power spectra are natively across positive and negative delays, but often we make plots with only positive delay bins. In addition, averaging bandpowers with identical negative with positive bandpowers gives us an extra factor of sqrt 2 sensitivity, which we can do with the `fold_spectra()` method.
```
uvp.fold_spectra()
uvp2.fold_spectra()
# plot power spectrum of spw 1
fig, ax = plt.subplots(figsize=(12,8))
spw = 1
blp =((24, 25), (37, 38))
pol = 'xx'
key = (spw, blp, pol)
k_para = uvp.get_kparas(spw)
power = np.abs(np.real(uvp.get_data(key)))
P_N = uvp.generate_noise_spectra(spw, pol, 300)
P_N = P_N[uvp.antnums_to_blpair(blp)]
avg_power = np.abs(np.real(uvp2.get_data(key)))
avg_P_N = uvp2.generate_noise_spectra(spw, pol, 300)
avg_P_N = avg_P_N[uvp2.antnums_to_blpair(blp)]
_ = ax.plot(k_para, power.T)
ax.plot(k_para, avg_power.T, color='k')
ax.plot(k_para, avg_P_N.T, color='k', ls='--', lw=3)
ax.set_yscale('log')
ax.grid()
ax.set_xlabel(r"$k_\parallel\ \rm h\ Mpc^{-1}$", fontsize=14)
ax.set_ylabel(r"$P(k)\ \rm [mK^2\ h^{-3}\ Mpc^3]$", fontsize=14)
ax.set_title("spw : {}, blpair : {}, pol : {}".format(*key), fontsize=14)
ax.set_prop_cycle(None)
_ = ax.plot(k_para, P_N.T, ls='--', lw=3)
```
## Re-defining the cosmology
We started by adopting a single cosmology, which was automatically passed between the various `hera_pspec` objects without user intervention in order to preserve self-consistency. If, however, we would like to adopt a *new cosmology*, we need to re-compute certain cosmology-dependent quantities, namely the normalization of the power spectrum scalar (see HERA Memos #27 and #43). We can do this with the `UVPSpec.set_cosmology` object.
```
# define a new cosmology with no dark energy
new_cosmo = hp.conversions.Cosmo_Conversions(Om_L = 0.0)
# attempt to overwrite the current UVPSpec cosmology:
# this will fail b/c one already exists and overwrite=False
uvp.set_cosmology(new_cosmo)
# now set overwrite = True
uvp.set_cosmology(new_cosmo, overwrite=True)
```
## Perform data selection on `UVPSpec`
We can select out parts of the data using the `select` function.
```
# select all baseline-pairs containing the baseline (24, 25)
# only_pairs_in_bls means to select only blpairs that have baseline1 _and_ baseline2 in the bls list
uvp2 = uvp.select(bls=[(24, 25)], inplace=False, only_pairs_in_bls=False)
# check than only baseline-pairs with a (24, 25) in them are kept
print("baseline pairs in uvp2: \n", map(lambda blp: uvp2.blpair_to_antnums(blp), np.unique(uvp2.blpair_array)))
```
## Write to HDF5
Write to disk using the `write_hdf5` method. Easy as that.
```
uvp.write_hdf5("pspec.hdf5", overwrite=True)
```
## Read from HDF5
Read the whole file, just the meta-data, or a data selection for partial I/O.
Frist, let's **read the whole file** and look at some meta-data.
```
# read the whole file
uvp2 = hp.uvpspec.UVPSpec()
uvp2.read_hdf5("pspec.hdf5")
# print some meta-data
print("polpairs: ", uvp2.polpair_array)
print("number of baseline-pairs: ", uvp2.Nblpairs)
# converting polpair integers to tuples
print("polpair tuples:", hp.uvpspec_utils.polpair_int2tuple(uvp2.polpair_array))
# Ensure data_array exists b/c we read the whole file
print("data_array in uvp2 : ", hasattr(uvp2, 'data_array'))
```
Now read **only the meta-data**.
```
# read only the meta-data
uvp2.read_hdf5("pspec.hdf5", just_meta=True)
# print some meta-data
print("polpairs : ", uvp2.polpair_array)
print("number of baseline-pairs: ",uvp2.Nblpairs)
# Ensure data_array doesn't exist b/c we only loaded the meta-data
print("data_array in uvp2 : ", hasattr(uvp2, 'data_array'))
```
Now do **partial loading of certain baseline pairs**.
```
# partial load on certain baseline-pairs
uvp2.read_hdf5("pspec.hdf5", just_meta=False, bls=[(37, 38), (38, 39)], only_pairs_in_bls=True)
# print baseline pairs
print("baseline pairs in uvp2: \n", map(lambda blp: uvp2.blpair_to_antnums(blp), np.unique(uvp2.blpair_array)))
```
Now do **partial loading of certain times**. To do this, we need to feed the exact float of the times we want, but how do we get these to float precision in the first place without loading the whole file? This is where the `just_meta` load comes in handy.
First load just the metadata, then do a partial load using the times you desire from the metadata.
```
# read only meta
uvp2.read_hdf5("pspec.hdf5", just_meta=True)
# now only load in one of the time integrations
uvp2.read_hdf5("pspec.hdf5", times=np.unique(uvp2.time_avg_array)[:1])
# check that only one time integration exists, but that all baseline pairs exist
print("number of unique times: ", len(np.unique(uvp2.time_avg_array)))
print("number of baseline-pairs: ", uvp2.Nblpairs)
if os.path.exists("pspec.hdf5"): os.remove("pspec.hdf5")
```
## Forming Pseudo-Stokes Power Spectra
Forming polarized pseudo-Stokes power spectra is identical to forming linear-dipole polarization power spectra, except that we need pseudo-Stokes visibilities and beam models. See the `Forming_PseudoStokes_Vis` notebook on details on how to do that. Here we will show how to form pseudo-I spectra.
```
# form pseudoI visibility
dset1 = UVData()
dset1.read_miriad(os.path.join(DATA_PATH, 'zen.all.xx.LST.1.06964.uvA')) # xx miriad file
dset2 = UVData()
dset2.read_miriad(os.path.join(DATA_PATH, 'zen.all.yy.LST.1.06964.uvA')) # yy miriad file
uvdI = hp.pstokes.construct_pstokes(dset1=dset1, dset2=dset2, pstokes='pI')
# List of beamfile to load. This is a healpix map for both XX and YY, and can therefore make an I map
beamfile = os.path.join(DATA_PATH, 'HERA_NF_pstokes_power.beamfits')
# intantiate beam and pass cosmology, if not fed, a default Planck cosmology will be assumed
uvb = hp.pspecbeam.PSpecBeamUV(beamfile, cosmo=cosmo)
# find conversion factor from Jy to mK for XX pol
Jy_to_mK = uvb.Jy_to_mK(np.unique(uvdI.freq_array), pol='pI')
# reshape to appropriately match a UVData.data_array object and multiply in!
uvdI.data_array *= Jy_to_mK[None, None, :, None]
# changeunits of UVData objects
uvdI.vis_units = 'mK'
# instantiate a PSpecData object
ds = hp.PSpecData(dsets=[uvdI, uvdI], wgts=[None, None], beam=uvb)
# baselines is a redundant baseline group
baselines = [(24,25), (37,38), (38,39)]
# calculate all baseline pairs from group
baselines1, baselines2, blpairs = hp.utils.construct_blpairs(baselines, exclude_auto_bls=True,
exclude_permutations=True)
# Inspect baseline pairs
print(blpairs)
uvp = ds.pspec(baselines1, baselines2, (0, 1), [('pI', 'pI')], spw_ranges=[(300, 400)],
input_data_weight='identity', norm='I', taper='blackman-harris', verbose=True)
# plot power spectrum of spectral window 0
fig, ax = plt.subplots(figsize=(12,8))
spw = 0
blp = ((24, 25), (37, 38))
key = (spw, blp, 'pI')
dlys = uvp.get_dlys(spw) * 1e9
power = np.abs(np.real(uvp.get_data(key)))
p1 = ax.plot(dlys, power.T)
ax.set_yscale('log')
ax.grid()
ax.set_xlabel("delay [ns]", fontsize=14)
ax.set_ylabel(r"$P(k)\ \rm [mK^2\ h^{-3}\ Mpc^3]$", fontsize=14)
ax.set_title("spw : {}, blpair : {}, pol : {}".format(*key), fontsize=14)
```
## Using Custom Polarized Beams
In the case that you have information on polarized beams that can't be put into a `UVBeam` object, you can use the custom beam capability of `hera_pspec.pspecbeam.PSpecBeamFromArray`, which will allow you to create a `PSpecBeam` object from custom $\Omega_P$ and $\Omega_{PP}$ arrays. See the `PSpecBeam` tutorial to see how to do this.
One can then use this object in the same way as above to create polarized power spectra with custom polarized beams.
| github_jupyter |
### Test to evaluate the use of global mass fraction in tracer solution
The discretized mass conservation equation of a component X can be written as
\begin{equation*}
\frac{m_T\phi-m_T^o\phi^o}{\Delta t}=\sum_{faces} \dot{m}_{face}\phi^{up}_{face}+\dot{m}_{comp}\phi_{comp}
\end{equation*}
where
\begin{equation*}
m_T = V_w\rho_w + V_o\rho_o+V_g\rho_g
\end{equation*}
\begin{equation*}
\dot{m} = F_w\rho_w^{up} + F_o\rho_o^{up} + F_g\rho_g^{up}
\end{equation*}
Phase densities and volumes are inputs from a reservoir simulation as well as the phase flow rates between grid blocks and the completion rates.
```
import numpy as np
# Main inputs
# 1D case in steady state and costant properties
nx = 20
number_of_time_steps = 100
saturation = np.array([1.0, 0.0, 0.0]) # [water, oil, gas]
time_step = 1.0 # d
injector_concentration = [0.1, 0.0, 0.0] # kg of X / kg of phase [water, oil, gas]
flow_rate = np.array([20.0, 0.0, 0.0]) # [water, oil, gas] in m3/d
# other inputs that change less frequently
L = 3000.0 # m
A = 1.0 # m2
total_volume = L * A # m3
bulk_volume = total_volume / nx # m3
porosity = 0.3 # m3 of pore space/ m3 of total volume
pore_volume = porosity * bulk_volume # m3
density = np.array([1000.0, 800.0, 200.0]) # [water, oil, gas] in kg/m3
injector_cell = 0
producer_cell = nx - 1
injector_rate = - flow_rate
producer_rate = flow_rate
A = np.zeros((nx, nx))
B = np.zeros(nx)
concentration = np.zeros(nx)
concentration_old = np.zeros(nx)
# steady state for now
density_old = density
saturation_old = saturation
pore_volume_old = pore_volume
def accumulation_term():
global A, B
for i in range(nx):
total_mass = np.sum(pore_volume * saturation * density)
total_mass_old = np.sum(pore_volume_old * saturation_old * density_old)
A[i,i] += total_mass
B[i] += total_mass_old * concentration_old[i]
def completions_term():
global A, B
# injection
B[injector_cell] += -np.sum(injector_rate * density * injector_concentration) * time_step
# production
A[producer_cell, producer_cell] += np.sum(producer_rate * density) * time_step
def flow_term():
global A
for i in range(nx-1):
cell_from = i
cell_to = i + 1
total_mass = np.sum(flow_rate * density) * time_step
A[cell_from, cell_from] += total_mass
A[cell_to, cell_from] += -total_mass
def assemble_linear_system():
global A, B
A = np.zeros((nx, nx))
B = np.zeros(nx)
accumulation_term()
completions_term()
flow_term()
def print_balance_error(cell):
injected_mass = 0.0
if cell == injector_cell:
injected_mass = abs(np.sum(injector_rate * density * injector_concentration) * time_step)
produced_mass = 0.0
if cell == producer_cell:
produced_mass = np.sum(producer_rate * density * injector_concentration) * time_step
flow_plus = 0.0
if cell < nx-1:
flow_plus = np.sum(flow_rate * density) * time_step * concentration[cell]
flow_minus = 0.0
if cell > 0:
flow_plus = np.sum(flow_rate * density) * time_step * concentration[cell-1]
total_mass = np.sum(pore_volume * saturation * density) * concentration[cell]
total_mass_old = np.sum(pore_volume_old * saturation_old * density_old) * concentration_old[cell]
change_in_time = total_mass - total_mass_old
balance_error = change_in_time - flow_minus - injected_mass + produced_mass + flow_plus
phase_mass = pore_volume * saturation * density
print(
"""
Phase mass: %s kg
Component X concentration: %f kg/kg
\tInjected mass: %f kg
\tProduced mass: %f kg
\tMass on I+ face: %f kg
\tMass on I- face: %f kg
\tChange of mass in time: %f kg (%f - %f)
\tBalance error: %f kg
"""
% (phase_mass, concentration[cell], injected_mass, produced_mass, flow_plus, flow_minus, change_in_time, total_mass, total_mass_old, balance_error))
%matplotlib notebook
import numpy as np
import matplotlib.pyplot as plt
# assemble the matrix just for visualization purposes, so we can see the coefficients in their places
assemble_linear_system()
plt.imshow(A)
plt.colorbar()
plt.show()
solutions = []
solutions.append(concentration)
def solve():
global concentration, concentration_old
for time in range(number_of_time_steps):
assemble_linear_system()
concentration = np.linalg.solve(A, B)
print_balance_error(0)
solutions.append(concentration)
concentration_old = np.copy(concentration)
solve()
%matplotlib notebook
import matplotlib.pyplot as plt
from ipywidgets import *
x = np.linspace(0, L, nx)
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
ax.set(ylim=(0, 1.1*np.max(injector_concentration)))
line, = ax.plot(x, solutions[0])
def update(x = 0):
line.set_ydata(solutions[x])
fig.canvas.draw_idle()
interact(update, x=widgets.IntSlider(min=0, max=len(solutions)-1, step=1, value=0))
```
| github_jupyter |
# parm@frosst-y to SMIRNOFF
This notebook provides examples/utility functionality to assist with conversion of parm@frosst or relatives to SMIRNOFF format. Particularly, Christopher Bayly is generating modified AMBER `frcmod` files where the first entry for each parameter (i.e. `CT-CT-CT`) is replaced by the relevant SMIRKS pattern, for conversion into SMIRNOFF FFXML format.
This notebook will:
1. Convert a specified smirks-frcmod file to SMIRNOFF FFXML
2. Generate (or take in) a set of molecules in OpenEye oemol format
3. Use the SMIRNOFF `ForceField` class to determine (a) which parameters are used in which molecules; (b) which molecules contain a specified parameter; and (c) which molecules do NOT contain a specified parameter.
Bayly has also updates the notebook with visualization for 3(b) and 3(c).
Bannan added printed current atom types to make looking up references easier
**Authors**:
* David L. Mobley (UCI)
* Contributions from Christopher I. Bayly (OpenEye Scientific Software Inc.) and Caitlin C. Bannan (UCI)
```
# Imports
from __future__ import print_function
from convert_frcmod import *
import openeye.oechem as oechem
import openeye.oeiupac as oeiupac
import openeye.oeomega as oeomega
import openeye.oedepict as oedepict
from IPython.display import display
from openff.toolkit.typing.engines.smirnoff.forcefield import *
from openff.toolkit.typing.engines.smirnoff.forcefield_utils import get_molecule_parameterIDs
from openff.toolkit.utils import *
% matplotlib inline
import matplotlib
import numpy as np
import pylab as pl
import matplotlib.pyplot as plt
import time
import IPython
import pickle
import glob
```
# Relevant methods
# Relevant methods
```
def depictAtomByIdx(mol_copy, atomIdxList, supH = True, width=900, height=500):
mol = oechem.OEMol(mol_copy)
OEGenerate2DCoordinates(mol)
atomBondSet = oechem.OEAtomBondSet()
for atom in mol.GetAtoms():
if atom.GetIdx() in atomIdxList:
atomBondSet.AddAtom( atom)
for bond in atom.GetBonds():
nbrAtom = bond.GetNbr(atom)
nbrIdx = nbrAtom.GetIdx()
if (nbrIdx in atomIdxList) and nbrIdx>atom.GetIdx():
atomBondSet.AddBond( bond)
from IPython.display import Image
dopt = oedepict.OEPrepareDepictionOptions()
dopt.SetDepictOrientation( oedepict.OEDepictOrientation_Horizontal)
dopt.SetSuppressHydrogens(supH)
oedepict.OEPrepareDepiction(mol, dopt)
opts = oedepict.OE2DMolDisplayOptions(width, height, oedepict.OEScale_AutoScale)
disp = oedepict.OE2DMolDisplay(mol, opts)
aroStyle = oedepict.OEHighlightStyle_Color
aroColor = oechem.OEColor(oechem.OEGrey)
oedepict.OEAddHighlighting(disp, aroColor, aroStyle,
oechem.OEIsAromaticAtom(), oechem.OEIsAromaticBond() )
hstyle = oedepict.OEHighlightStyle_BallAndStick
hcolor = oechem.OEColor(oechem.OELightGreen)
oedepict.OEAddHighlighting(disp, hcolor, hstyle, atomBondSet)
#ofs = oechem.oeosstream()
img = oedepict.OEImage(width, height)
oedepict.OERenderMolecule(img, disp)
#oedepict.OERenderMolecule(ofs, 'png', disp)
#ofs.flush()
#return Image(data = "".join(ofs.str()))
return Image(oedepict.OEWriteImageToString("png",img))
def getMolParamIDToAtomIndex( oemol, ff):
"""Take an OEMol and a SMIRNOFF force field object and return a dictionary,
keyed by parameter ID, where each entry is a tuple of
( smirks, [[atom1, ... atomN], [atom1, ... atomN]) giving the SMIRKS
corresponding to that parameter ID and a list of the atom groups in that
molecule that parameter is applied to.
Parameters
----------
oemol : OEMol
OpenEye OEMol with the molecule to investigate.
ff : ForceField
SMIRNOFF ForceField object (obtained from an ffxml via ForceField(ffxml)) containing FF of interest.
Returns
-------
param_usage : dictionary
Dictionary, keyed by parameter ID, where each entry is a tuple of
( smirks, [[atom1, ... atomN], [atom1, ... atomN]) giving the SMIRKS
corresponding to that parameter ID and a list of the atom groups in
that molecule that parameter is applied to.
"""
labels = ff.labelMolecules([oemol])
param_usage = {}
for mol_entry in range(len(labels)):
for force in labels[mol_entry].keys():
for (atom_indices, pid, smirks) in labels[mol_entry][force]:
if not pid in param_usage:
param_usage[pid] = (smirks, [atom_indices])
else:
param_usage[pid][1].append( atom_indices )
return param_usage
def GetAtomInfo(mol, indices, skip_atoms = []):
#print(indices)
atoms_by_index = dict()
charges_by_index = dict()
for atom in mol.GetAtoms():
idx = atom.GetIdx()
if idx in indices:
atoms_by_index[idx] = atom
charge = atom.GetFormalCharge()
if charge == 0:
charges_by_index[idx] = ''
elif charge > -1:
charges_by_index[idx] = '+%i' % charge
else:
charges_by_index[idx] = str(charge)
atoms = [(atoms_by_index[idx],charges_by_index[idx]) for idx in indices]
types = [atom.GetType() for (atom,charge) in atoms]
atom_smarts = ['[#%i%s]' % (atom.GetAtomicNum(),charge) for (atom,charge) in atoms]
smarts = '~'.join(atom_smarts)
types = '~'.join(types)
for (atom, charge) in atoms:
if atom.GetAtomicNum() in skip_atoms:
return (True, smarts, types)
return (False, smarts, types)
def DepictMolWithParam(mol, indice_list, supH = False, print_atoms = True, skip_atoms = []):
skip_count = 0
for IdxByOccurrence in indice_list:
skip_it, approx_smarts, types = GetAtomInfo(mol, IdxByOccurrence, skip_atoms)
if skip_it:
skip_count += 1
continue
if print_atoms:
print("Approximate SMARTS: %s" % approx_smarts)
print("Current Atom Types: %s" % types)
display(depictAtomByIdx(mol, IdxByOccurrence, supH = supH))
if skip_count > 0:
skips = ','.join([str(i) for i in skip_atoms])
print("This molecule contains %i fragment(s) with at least one atom in (%s)" % (skip_count, skips))
def make_param_histogram(param_id_counts, param_ids, letter, title):
# Graph occurrences of bond parameters
parm_ids = [ pid for pid in param_ids if pid[0]==letter]
parm_ids.sort()
counts_parms = [param_id_counts[parm_id] for parm_id in parm_ids]
#print( parm_ids)
#print( counts_parms)
split = int(len(parm_ids)/2)
indices = np.arange(len(parm_ids))
fix, ax = plt.subplots(2,1,figsize=(16,5))
ax[0].set_yscale('log', nonposy='clip')
ax[1].set_yscale('log', nonposy='clip')
rects2 = ax[0].bar(indices[0:split], counts_parms[0:split] )
ax[0].set_ylabel('Count')
ax[0].set_xticks( indices[0:split])
ax[0].set_xticklabels( parm_ids[0:split], rotation=-60, ha='left')
ax[0].set_xlim(indices[0], indices[split])
plt.yscale('log',nonposy='clip')
rects2 = ax[1].bar(indices[split:], counts_parms[split:])
ax[1].set_ylabel('Count')
ax[1].set_xticks( indices[split:])
ax[1].set_xticklabels( parm_ids[split:], rotation=-60, ha='left')
ax[1].set_xlim(indices[split], indices[-1]+1)
ax[0].set_title(title)
plt.show()
def check_valence(mol):
"""
Checks for hypervalency
Parameter
---------
mol - OEMol()
Return
------
boolean - True (no inappropriate valency)
False (an atom with atomic number < 10 has > 4 Valence)
"""
for atom in mol.GetAtoms():
atomNum = atom.GetAtomicNum()
# find number of neighbors to this atom
valence = atom.GetValence()
if atomNum <= 10: # first row elements
if valence > 4:
print("Found a #%i atom with valence %i in molecule %s" % (atomNum, valence, oechem.OECreateIsoSmiString(mol)))
return False
return True
```
## 1. Convert specified SMIRKS `frcmod` file to SMIRNOFF FFXML
```
# Input and output info
#infile = 'example.frcmod' # smirnoffish frcmod file to convert
infile = 'smirnoffishFrcmod.parm99Frosst.txt' # smirnoffish frcmod file to convert
ffxmlFile = 'smirnoff99FrosstFrcmod.offxml'
template = 'template.offxml' # Template FFXML file without parameters (but with remainder of contents)
# Convert
# Already converted
convert_frcmod_to_ffxml( infile, template, ffxmlFile)
# Load SMIRNOFF FFXML
ff = ForceField(ffxmlFile) # We will use this below to access details of parameters
```
## 2. Generate or take in a set of molecules in OpenEye OEMol format
Here we will take a set of molecules from openff-toolkit (or elsewhere), read in all molecules and then uncomment any filters you want.
Here are some examples of molecule sets in openff-toolkit (at /openff/toolkit/data/molecules/):
* `AlkEthOH_test_filt1_ff.mol2` - 42 alkanes, ethers, and alcohols with parm@frosst atom types
* `DrugBank_atyped.oeb` - DrugBank database with parm@frosst atom types (including "untypable" atoms)
* `zinc-subset-parm@frosst.mol2.gz` - ZINC parm@frosst subset from CCL
```
# Un-comment this section if you want to use a local directory with individual mol2 files
#DBpath = "path/to/molecules/*.mol2"
#DBpath = "/Users/bannanc/gitHub/FreeSolv/mol2files_sybyl/*mol2"
#DB_files = glob.glob(DBpath)
#molecules = list()
#for f in DB_files:
# molecules += read_molecules(f, verbose=False)
# These are atoms you don't want in your set, in this case metaloids or nobel gases
skip_atoms = [2, 5, 14,33, 34, 52, 54]
# Molecules file in openforcefield/data/molecules/
# OR any relative/absolute path
mol_File = 'DrugBank_atyped.oeb'
#mol_File = "zinc-subset-parm@frosst.mol2.gz"
#mol_File = "/Users/bannanc/Google Drive/RESEARCH/OFF_draftAndTestSpace/eMolecules_missingParameters/output.mol2"
molecules = read_molecules(mol_File)
# For use later, generate isomeric SMILES for these so we can easily look up molecules by smiles
isosmiles_to_mol = dict()
repeat = 0
skipped = 0
for mol in molecules:
c_mol = OEMol(mol)
oechem.OEAddExplicitHydrogens(c_mol)
# Get the smiles string for this molecule
smi = oechem.OECreateIsoSmiString(c_mol)
# uncomment to skip molecules with > n heavy atoms
n=200
if OECount(c_mol, OEIsHeavy()) > n:
continue
# uncomment to skip molecules with metals
#if OECount(c_mol, OEIsMetal()) > 0:
#skipped +=1
# continue
# uncomment to skip molecules containing the skip_atoms
has_skip_atom = False
for n in skip_atoms:
if OECount(c_mol, OEHasAtomicNum(n)) > 0:
has_skip_atom = True
#if has_skip_atom:
#skipped += 1
# continue
# uncomment to skip molecules with 5 bonds to atoms with atomic number < 10 (i.e. pentavalent nitrogen)
#if not check_valence(c_mol):
#skipped += 1
# continue
# uncomment to skip single molecules that contain > 1 molecule
#if '.' in smi:
# skipped +=1
# continue
if smi in isosmiles_to_mol:
repeat += 1
isosmiles_to_mol[smi] = c_mol
oemols = [mol for smi, mol in isosmiles_to_mol.items()]
print("\nAfter filtering %i molecules there were %i repeated SMILES.\nThe final set has %i/%i molecules"\
% (skipped, repeat, len(oemols), len(molecules)))
```
## 3. Determine parameter usage in molecules
Here we will use the SMIRNOFF ForceField class to determine (a) which parameters are used in which molecules; (b) which molecules contain a specified parameter; and (c) which molecules do NOT contain a specified parameter. We begin by just loading the SMIRNOFF force field we generated in section 1.
### 3(a). Determine which parameters are used in which molecules
Here we determine which parameters are actually used in which molecules, and make a couple example plots of the frequency of parameter occurrences for some of our example parameters.
```
# Track time
init_time = time.time()
# label molecules
labels = ff.labelMolecules(oemols, verbose = False)
elapsed = (time.time() - init_time) / 60.0
print("Assigned labels took %.2f minutes" % (elapsed))
# organize dictionaries to reference information
init_time = time.time()
parameters_by_molecule = dict()
parameters_by_ID = dict()
param_ids = set()
param_id_counts = dict()
for idx, mol_dict in enumerate(labels):
smi = OECreateIsoSmiString(oemols[idx])
parameters_by_molecule[smi] = dict()
for force_type, label_set in mol_dict.items():
for (indices, pid, smirks) in label_set:
if not pid in parameters_by_molecule[smi]:
parameters_by_molecule[smi][pid] = list()
parameters_by_molecule[smi][pid].append(indices)
if not pid in parameters_by_ID:
parameters_by_ID[pid] = set()
parameters_by_ID[pid].add(smi)
param_ids.add(pid)
for pid in param_ids:
param_id_counts[pid] = 0
for smi, pid_dict in parameters_by_molecule.items():
for pid, ind_list in pid_dict.items():
param_id_counts[pid] += len(ind_list)
elapsed = (time.time() - init_time) / 60.0
print("Organizing dictionaries took %.2f minutes" % (elapsed))
```
For fun/info, do a quick graph of frequency of occurrence of particular parameters. Here, let's just do bond parameters
```
# Create Hisogram for each type of parameter
# VdW
make_param_histogram(param_id_counts, param_ids, 'n', "VdW for DrugBank Molecules")
# Bonds
make_param_histogram(param_id_counts, param_ids, 'b', "Bonds for DrugBank Molecules")
# Angles
make_param_histogram(param_id_counts, param_ids, 'a', "Angles for DrugBank Molecules")
# Torsions
make_param_histogram(param_id_counts, param_ids, 't', "Torsions for DrugBank Molecules")
#make_param_histogram(param_id_counts, param_ids, 'i', "Impropers for DrugBank Molecules")
```
### 3(b)-3(c). Determine which molecules do/do not contain selected parameter
Determine which molecules do and do not contain a specified parameter; give access to isomeric smiles and OEMol for each molecule in each case.
```
# INPUT: Pick what parameter to look at
parameter_id = 'n1'
# For info, get details of that parameter
params = ff.getParameter(paramID=parameter_id)
print("For parameter %s, the relevant parameters are:" % parameter_id)
print(params)
# Find molecules which do/do not use that parameter
mols_with_param = []
mols_wo_param = []
for isosmi in parameters_by_molecule:
# Store a tuple of (isomeric smiles, oemol) for each
if parameter_id in parameters_by_molecule[isosmi].keys():
mols_with_param.append( (isosmi, isosmiles_to_mol[isosmi] ))
else:
mols_wo_param.append( (isosmi, isosmiles_to_mol[isosmi] ))
print("\nThere are %s molecules containing that parameter and %s which do not, out of %s.\n" %
(len(mols_with_param), len(mols_wo_param), len(isosmiles_to_mol)))
# Print first 10 molecules not containing parameter
not_containing = min(10, len(mols_wo_param))
if not_containing == 0:
print("\nAll molecules conatin parameter '%s'" % parameter_id)
elif not_containing <= 10:
print("\nThe %i molecule(s) that do(es) not contain parameter '%s'" % (not_containing, parameter_id))
else:
print("First 10 molecules not containing '%s' parameter:" % parameter_id)
for i in range(not_containing):
print(" %s" % mols_wo_param[i][0])
# Print first 10 molecules containing parameter
containing = min(10,len(mols_with_param))
if containing == 0:
print("\nNO molecules contain '%s' parameter" % parameter_id)
elif containing <= 10:
print("\nThe %i molecule(s) containing '%s' parameter:" % (containing, parameter_id))
else:
print("\nFirst 10 molecules containing '%s' parameter:" % parameter_id)
for i in range(containing):
print(" %s" % mols_with_param[i][0])
lowerlimit = 0
upperlimit = 100
# Skip showing molecule if the highlighted parameter
# includes an element we know we don't have parameters for
skip_atoms = [2, 5, 14,33, 34, 52]
# Correct with list of molecules is less than upper limit
if len(mols_with_param) < upperlimit:
upperlimit = len(mols_with_param)
print("\nDepicting molecules %i to %i with parameter '%s'\n\n" % (lowerlimit,upperlimit-1, parameter_id))
# Show molecules form lower to upper limit highlighting fragment with the parameter_id
for idx in range(lowerlimit, upperlimit):
smiles = mols_with_param[idx][0]
mol = mols_with_param[idx][1]
skip_it = False
OEAddExplicitHydrogens(mol)
indice_list = parameters_by_molecule[smiles][parameter_id]
print("looking at molecule %i" % idx)
print('Selected smiles is %s' % smiles)
print('Selected IUPAC name guessed: %s' % oeiupac.OECreateIUPACName(mol) )
print( 'mol title and NumAtoms: ', mol.GetTitle(), mol.NumAtoms() )
print( "Number of times '%s' appears: %i" % (parameter_id, len(indice_list)))
DepictMolWithParam( mol, indice_list, supH = False, skip_atoms=skip_atoms)
print()
print()
print()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/WeizmannML/course2020/blob/master/Tutorial6/Graph_Classification_DGL.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
# !pip install dgl
# !pip install networkx
import torch as th
from torch import nn
import torch.nn.functional as F
from torch.nn import init
import dgl.function as fn
from dgl.base import DGLError
import torch
import dgl
```
## The graph classifier will be built out of two graph-convolution layers
## The graph convolution : <br> $h_i^{(l+1)} = \sigma(b^{(l)} + \sum_{j\in\mathcal{N}(i)}\frac{1}{c_{ij}}h_j^{(l)}W^{(l)})$ <br> with $c_{ij} = \sqrt{|\mathcal{N}(i)|}\sqrt{|\mathcal{N}(j)|}$ ; <br> $H^{(l+1)} = \sigma ( \tilde{D}^{-\frac{1}{2}} \tilde{A} \tilde{D}^{-\frac{1}{2}} W^{l}h^{l} ) $
```
# implemented in https://docs.dgl.ai/_modules/dgl/nn/pytorch/conv/graphconv.html#GraphConv
class GraphConv(nn.Module):
r"""Apply graph convolution over an input signal.
Graph convolution is introduced in `GCN <https://arxiv.org/abs/1609.02907>`__
and can be described as below:
.. math::
h_i^{(l+1)} = \sigma(b^{(l)} + \sum_{j\in\mathcal{N}(i)}\frac{1}{c_{ij}}h_j^{(l)}W^{(l)})
where :math:`\mathcal{N}(i)` is the neighbor set of node :math:`i`. :math:`c_{ij}` is equal
to the product of the square root of node degrees:
:math:`\sqrt{|\mathcal{N}(i)|}\sqrt{|\mathcal{N}(j)|}`. :math:`\sigma` is an activation
function.
The model parameters are initialized as in the
`original implementation <https://github.com/tkipf/gcn/blob/master/gcn/layers.py>`__ where
the weight :math:`W^{(l)}` is initialized using Glorot uniform initialization
and the bias is initialized to be zero.
Notes
-----
Zero in degree nodes could lead to invalid normalizer. A common practice
to avoid this is to add a self-loop for each node in the graph, which
can be achieved by:
>>> g = ... # some DGLGraph
>>> g.add_edges(g.nodes(), g.nodes())
Parameters
----------
in_feats : int
Input feature size.
out_feats : int
Output feature size.
norm : str, optional
How to apply the normalizer. If is `'right'`, divide the aggregated messages
by each node's in-degrees, which is equivalent to averaging the received messages.
If is `'none'`, no normalization is applied. Default is `'both'`,
where the :math:`c_{ij}` in the paper is applied.
weight : bool, optional
If True, apply a linear layer. Otherwise, aggregating the messages
without a weight matrix.
bias : bool, optional
If True, adds a learnable bias to the output. Default: ``True``.
activation: callable activation function/layer or None, optional
If not None, applies an activation function to the updated node features.
Default: ``None``.
Attributes
----------
weight : torch.Tensor
The learnable weight tensor.
bias : torch.Tensor
The learnable bias tensor.
"""
def __init__(self,
in_feats,
out_feats,
norm='both',
weight=True,
bias=True,
activation=None):
super(GraphConv, self).__init__()
if norm not in ('none', 'both', 'right'):
raise DGLError('Invalid norm value. Must be either "none", "both" or "right".'
' But got "{}".'.format(norm))
self._in_feats = in_feats
self._out_feats = out_feats
self._norm = norm
if weight:
self.weight = nn.Parameter(th.Tensor(in_feats, out_feats))
else:
self.register_parameter('weight', None)
if bias:
self.bias = nn.Parameter(th.Tensor(out_feats))
else:
self.register_parameter('bias', None)
self.reset_parameters()
self._activation = activation
def reset_parameters(self):
"""Reinitialize learnable parameters."""
if self.weight is not None:
init.xavier_uniform_(self.weight)
if self.bias is not None:
init.zeros_(self.bias)
def forward(self, graph, feat, weight=None):
"""Compute graph convolution.
Notes
-----
* Input shape: :math:`(N, *, \text{in_feats})` where * means any number of additional
dimensions, :math:`N` is the number of nodes.
* Output shape: :math:`(N, *, \text{out_feats})` where all but the last dimension are
the same shape as the input.
* Weight shape: "math:`(\text{in_feats}, \text{out_feats})`.
Parameters
----------
graph : DGLGraph
The graph.
feat : torch.Tensor
The input feature
weight : torch.Tensor, optional
Optional external weight tensor.
Returns
-------
torch.Tensor
The output feature
"""
graph = graph.local_var()
if self._norm == 'both':
degs = graph.out_degrees().to(feat.device).float().clamp(min=1)
norm = th.pow(degs, -0.5)
shp = norm.shape + (1,) * (feat.dim() - 1)
norm = th.reshape(norm, shp)
feat = feat * norm
if weight is not None:
if self.weight is not None:
raise DGLError('External weight is provided while at the same time the'
' module has defined its own weight parameter. Please'
' create the module with flag weight=False.')
else:
weight = self.weight
if self._in_feats > self._out_feats:
# mult W first to reduce the feature size for aggregation.
if weight is not None:
feat = th.matmul(feat, weight)
graph.srcdata['h'] = feat
graph.update_all(fn.copy_src(src='h', out='m'),
fn.sum(msg='m', out='h'))
rst = graph.dstdata['h']
else:
# aggregate first then mult W
graph.srcdata['h'] = feat
graph.update_all(fn.copy_src(src='h', out='m'),
fn.sum(msg='m', out='h'))
rst = graph.dstdata['h']
if weight is not None:
rst = th.matmul(rst, weight)
if self._norm != 'none':
degs = graph.in_degrees().to(feat.device).float().clamp(min=1)
if self._norm == 'both':
norm = th.pow(degs, -0.5)
else:
norm = 1.0 / degs
shp = norm.shape + (1,) * (feat.dim() - 1)
norm = th.reshape(norm, shp)
rst = rst * norm
if self.bias is not None:
rst = rst + self.bias
if self._activation is not None:
rst = self._activation(rst)
return rst
def extra_repr(self):
"""Set the extra representation of the module,
which will come into effect when printing the model.
"""
summary = 'in={_in_feats}, out={_out_feats}'
summary += ', normalization={_norm}'
if '_activation' in self.__dict__:
summary += ', activation={_activation}'
return summary.format(**self.__dict__)
from dgl.data import MiniGCDataset
import matplotlib.pyplot as plt
import networkx as nx
# A dataset with 80 samples, each graph is
# of size [10, 20]
dataset = MiniGCDataset(80, 10, 20)
graph, label = dataset[0]
fig, ax = plt.subplots()
nx.draw(graph.to_networkx(), ax=ax)
ax.set_title('Class: {:d}'.format(label))
plt.show()
dataset.graphs
dataset.max_num_v, dataset.num_classes
class_names ={
0 : 'cycle graph',
1 : 'star graph',
2 : 'wheel graph',
3 : 'lollipop graph',
4 : 'hypercube graph',
5 : 'grid graph',
6 : 'clique graph',
7 : 'circular ladder graph',
}
plt.rcParams["figure.figsize"]=18,12
fig, axes = plt.subplots(nrows=2, ncols=4)
iter = 0
for ax in axes.flatten() :
graph, label = dataset[10 * iter]
nx.draw(graph.to_networkx(), with_labels=True, ax=ax)
ax.set_title('Class: {:d}'.format(label.item() ) +', '+class_names[label.item()] )
iter += 1
plt.show()
```
# Forming a graph minibatch
## In general different elements of a batch can have different shapes (slide)
```
def collate(samples):
# The input `samples` is a list of pairs
# (graph, label).
graphs, labels = map(list, zip(*samples))
batched_graph = dgl.batch(graphs)
return batched_graph, torch.tensor(labels)
```
# Readout & Classification
###For this demonstration, consider initial node features to be their degrees. After two rounds of graph convolution, perform a graph readout by averaging over all node features for each graph in the batch. <br> $h_g = \frac{1}{\mathcal{|N|}} \sum_{\nu \in \mathcal{N}} h_{\nu} $
```
class Classifier(nn.Module):
def __init__(self, in_dim, hidden_dim, n_classes):
super(Classifier, self).__init__()
self.conv1 = GraphConv(in_dim, hidden_dim)
self.conv2 = GraphConv(hidden_dim, hidden_dim)
self.classify = nn.Linear(hidden_dim, n_classes)
def forward(self, g):
# Use node degree as the initial node feature. For undirected graphs, the in-degree
# is the same as the out_degree.
h = g.in_degrees().view(-1, 1).float()
# Perform graph convolution and activation function.
h = F.relu(self.conv1(g, h))
h = F.relu(self.conv2(g, h))
g.ndata['h'] = h
# Calculate graph representation by averaging all the node representations.
hg = dgl.mean_nodes(g, 'h')
return self.classify(hg)
import torch.optim as optim
from torch.utils.data import DataLoader
# Create training and test sets.
trainset = MiniGCDataset(1000, 10, 20)
testset = MiniGCDataset(300, 10, 20)
validset = MiniGCDataset(200, 10, 20)
# Use PyTorch's DataLoader and the collate function
# defined before.
data_loader = DataLoader(trainset, batch_size=32, shuffle=True,
collate_fn=collate)
val_data_loader = DataLoader(validset, batch_size=42, shuffle=True,
collate_fn=collate)
test_data_loader = DataLoader(testset, batch_size=42, shuffle=True,
collate_fn=collate)
# Create model
model = Classifier(1, 256, trainset.num_classes)
#model = model.cuda()
# if torch.cuda.is_available():
# model = model.cuda()
loss_func = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=0.001)
model
## --------- Run the training loop ------------------- ##
n_epoch = 200
epoch_losses, epoch_val_losses = [], []
for epoch in range(1, n_epoch+1):
epoch_loss = 0
iter_batch = 0
model.train()
for bg, label in data_loader:
# if torch.cuda.is_available():
# bg = bg.cuda()
# label = label.cuda()
prediction = model(bg)
loss = loss_func(prediction, label )
optimizer.zero_grad()
loss.backward()
optimizer.step()
epoch_loss += loss.detach().item()
iter_batch += 1
epoch_loss /= (iter_batch + 1)
#print('Epoch {}, loss {:.4f}'.format(epoch, epoch_loss))
epoch_losses.append(epoch_loss)
epoch_val_loss = 0
iter_batch_val = 0
model.eval()
for bg, label in val_data_loader:
prediction = model(bg)
loss = loss_func(prediction, label )
epoch_val_loss += loss.detach().item()
iter_batch_val += 1
epoch_val_loss /= (iter_batch_val + 1)
#print('Epoch {}, loss {:.4f}'.format(epoch, epoch_loss))
epoch_val_losses.append(epoch_val_loss)
print('Epoch {}, loss {:.4f}, val_loss {:.4f}'.format(epoch, epoch_loss, epoch_val_loss))
import numpy as np
plt.rcParams["figure.figsize"]=10,10
plt.plot( range(1, n_epoch+1), np.array(epoch_losses), label='Training Loss' )
plt.plot( range(1, n_epoch+1), np.array(epoch_val_losses), label='Validation Loss' )
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.legend()
plt.show()
model.eval()
# Convert a list of tuples to two lists
test_X, test_Y = map(list, zip(*testset))
test_bg = dgl.batch(test_X)
test_Y = torch.tensor(test_Y).float().view(-1, 1)
probs_Y = torch.softmax(model(test_bg), 1)
sampled_Y = torch.multinomial(probs_Y.cpu() , 1)
argmax_Y = torch.max(probs_Y.cpu(), 1)[1].view(-1, 1)
print('Accuracy of sampled predictions on the test set: {:.4f}%'.format(
(test_Y == sampled_Y.float()).sum().item() / len(test_Y) * 100))
# print('Accuracy of argmax predictions on the test set: {:4f}%'.format(
# (test_Y == argmax_Y.float()).sum().item() / len(test_Y) * 100))
```
| github_jupyter |
```
# author: Leonardo Filipe
# website: https://www.leonardofilipe.com
# contact: contact[at]leonardofilipe.com
import io
import re
import requests
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
plt.style.use('seaborn')
def getdata(tickers,start,end,frequency):
OHLC = {}
cookie = ''
crumb = ''
res = requests.get('https://finance.yahoo.com/quote/SPY/history')
cookie = res.cookies['B']
pattern = re.compile('.*"CrumbStore":\{"crumb":"(?P<crumb>[^"]+)"\}')
for line in res.text.splitlines():
m = pattern.match(line)
if m is not None:
crumb = m.groupdict()['crumb']
for ticker in tickers:
url_str = "https://query1.finance.yahoo.com/v7/finance/download/%s"
url_str += "?period1=%s&period2=%s&interval=%s&events=history&crumb=%s"
url = url_str % (ticker, start, end, frequency, crumb)
res = requests.get(url, cookies={'B': cookie}).text
OHLC[ticker] = pd.read_csv(io.StringIO(res), index_col=0,
error_bad_lines=False).replace('null', np.nan).dropna()
OHLC[ticker].index = pd.to_datetime(OHLC[ticker].index)
OHLC[ticker] = OHLC[ticker].apply(pd.to_numeric)
return OHLC
tickers = ['%5EGSPC','000001.SS']
data = None
while data is None:
try:
data = getdata(tickers,'946610000','1538960000','1d')
except:
pass
ReturnSP = data['%5EGSPC']['Adj Close']/data['%5EGSPC']['Adj Close'].shift(1)-1
BuyHoldSP = data['%5EGSPC']['Adj Close']/float(data['%5EGSPC']['Adj Close'][:1])-1
BuyHold1ySP = data['%5EGSPC']['Adj Close']/data['%5EGSPC']['Adj Close'].shift(252)-1
ReturnSSE = data['000001.SS']['Adj Close']/data['000001.SS']['Adj Close'].shift(1)-1
BuyHoldSSE = data['000001.SS']['Adj Close']/float(data['000001.SS']['Adj Close'][:1])-1
BuyHold1ySSE = data['000001.SS']['Adj Close']/data['000001.SS']['Adj Close'].shift(252)-1
BuyHoldAll = (BuyHoldSP*(0.8)+BuyHoldSSE*(0.2)).fillna(method='ffill')
BuyHold1yAll = (BuyHold1ySP*(0.8)+BuyHold1ySSE*(0.2)).fillna(method='ffill')
holdings = pd.DataFrame({'Asset 1':ReturnSP*0.8,'Asset 2':ReturnSSE*0.2}).fillna(0)
signal = pd.DataFrame({'Asset 1':BuyHold1ySP.shift(1),'Asset 2':BuyHold1ySSE.shift(1)}).fillna(method='ffill')
holdings['Asset 1'][signal['Asset 1'] > -0.17] = ReturnSP*1
holdings['Asset 2'][signal['Asset 1'] > -0.17] = ReturnSSE*0
holdings['Asset 1'][signal['Asset 2'] > 0.29] = ReturnSP*0
holdings['Asset 2'][signal['Asset 2'] > 0.29] = ReturnSSE*1
DynAssAll = (holdings['Asset 1']+holdings['Asset 2']).fillna(0).cumsum()
DynAssAll1y = (holdings['Asset 1']+holdings['Asset 2']).fillna(0).rolling(window=252).sum()
plt.figure(figsize=(15,6))
plt.plot(BuyHoldSP*100, label='Buy & Hold SP&500')
plt.plot(BuyHoldAll*100, label='Optimal Sharpe')
plt.plot(DynAssAll*100, label='Dynamic Asset Allocation')
plt.xlabel('Time')
plt.ylabel('Returns (in %)')
plt.margins(x=0.005,y=0.02)
plt.axhline(y=0, xmin=0, xmax=1, linestyle='--', color='k')
plt.legend()
plt.show()
print('Mean yearly excess return over SP500 =',round(float(DynAssAll1y.mean()-BuyHold1ySP.mean())*100,1),'%')
print('"" "" "" "" over Optimal Sharpe =',round(float(DynAssAll1y.mean()-BuyHold1yAll.mean())*100,1),'%')
print()
print('Strategy YTD Performance (at 05/10/2018) =',round(float(DynAssAll1y[-1:]*100),1),'%')
print('S&P500 Buy & Hold "" "" (at 05/10/2018) =',round(float(BuyHold1ySP[-1:]*100),1),'%')
```
| github_jupyter |
# Matplotlib
## [`matplotlib`](https://matplotlib.org/) is the most widely used scientific plotting library in Python.
* Commonly use a sub-library called [`matplotlib.pyplot`](https://matplotlib.org/api/pyplot_api.html).
* The Jupyter Notebook will render plots inline if we ask it to using a "magic" command.
```
%matplotlib inline
import matplotlib.pyplot as plt
```
* Simple plots are then (fairly) simple to create.
```
time = [0, 1, 2, 3]
position = [0, 100, 200, 300]
plt.plot(time, position)
plt.xlabel('Time (hr)')
plt.ylabel('Position (km)')
```
## Plot data directly from a [`Pandas dataframe`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.html).
* We can also plot [Pandas dataframes](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.html).
* This implicitly uses [`matplotlib.pyplot`](https://matplotlib.org/api/pyplot_api.html).
* Before plotting, we convert the column headings from a `string` to `integer` data type, since they represent numerical values
```
import pandas
data = pandas.read_csv('data/gapminder_gdp_oceania.csv', index_col='country')
# Extract year from last 4 characters of each column name
years = data.columns.str.strip('gdpPercap_')
# Convert year values to integers, saving results back to dataframe
data.columns = years.astype(int)
data.loc['Australia'].plot()
```
## Select and transform data, then plot it.
* By default, [`DataFrame.plot`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.plot.html#pandas.DataFrame.plot) plots with the rows as the X axis.
* We can transpose the data in order to plot multiple series.
```
data.T.plot()
plt.ylabel('GDP per capita')
```
## Many styles of plot are available.
* For example, do a bar plot using a fancier style.
```
plt.style.use('ggplot')
data.T.plot(kind='bar')
plt.ylabel('GDP per capita')
```
## Data can also be plotted by calling the `matplotlib` `plot` function directly.
* The command is `plt.plot(x, y)`
* The color / format of markers can also be specified as an optical argument: e.g. 'b-' is a blue line, 'g--' is a green dashed line.
## Get Australia data from dataframe
```
years = data.columns
gdp_australia = data.loc['Australia']
plt.plot(years, gdp_australia, 'g--')
```
## Can plot many sets of data together.
```
# Select two countries' worth of data.
gdp_australia = data.loc['Australia']
gdp_nz = data.loc['New Zealand']
# Plot with differently-colored markers.
plt.plot(years, gdp_australia, 'b-', label='Australia')
plt.plot(years, gdp_nz, 'g-', label='New Zealand')
# Create legend.
plt.legend(loc='upper left')
plt.xlabel('Year')
plt.ylabel('GDP per capita ($)')
```
* Plot a scatter plot correlating the GDP of Australia and New Zealand
* Use either `plt.scatter` or `DataFrame.plot.scatter`
```
plt.scatter(gdp_australia, gdp_nz)
data.T.plot.scatter(x = 'Australia', y = 'New Zealand')
```
## Adding a Legend
Often when plotting multiple datasets on the same figure it is desirable to have
a legend describing the data. This can be done in `matplotlib` in two stages:
Provide a label for each dataset in the figure:
```
plt.plot(years, gdp_australia, label='Australia')
plt.plot(years, gdp_nz, label='New Zealand')
```
Instruct `matplotlib` to create the legend.
```
plt.legend()
```
By default matplotlib will attempt to place the legend in a suitable position. If you would rather specify a position this can be done with the `loc=` argument, e.g to place the legend in the upper left corner of the plot, specify `loc='upper left'`
## Saving your plot to a file
If you are satisfied with the plot you see you may want to save it to a file, perhaps to include it in a publication. There is a function in the matplotlib.pyplot module that accomplishes this: [savefig](https://matplotlib.org/api/_as_gen/matplotlib.pyplot.savefig.html). Calling this function, e.g. with
`plt.savefig('my_figure.png')`
will save the current figure to the file `my_figure.png`. The file format will automatically be deduced from the file name extension (other formats are pdf, ps, eps and svg).
Note that functions in `plt` refer to a global figure variable and after a figure has been displayed to the screen (e.g. with `plt.show`) matplotlib will make this variable refer to a new empty figure. Therefore, make sure you call `plt.savefig` before the plot is displayed to the screen, otherwise you may find a file with an empty plot.
When using dataframes, data is often generated and plotted to screen in one line, and `plt.savefig` seems not to be a possible approach. One possibility to save the figure to file is then to
* save a reference to the current figure in a local variable (with `plt.gcf`)
* call the `savefig` class method from that varible.
`fig = plt.gcf() # get current figure
data.plot(kind='bar')
fig.savefig('my_figure.png')`
## Making your plots accessible
Whenever you are generating plots to go into a paper or a presentation, there are a few things you can do to make sure that everyone can understand your plots.
Always make sure your text is large enough to read. Use the `fontsize` parameter in `xlabel`, `ylabel`, `title`, and `legend`, and [`tick_params` with `labelsize`](https://matplotlib.org/2.1.1/api/_as_gen/matplotlib.pyplot.tick_params.html) to increase the text size of the numbers on your axes.
Similarly, you should make your graph elements easy to see. Use `s` to increase the size of your scatterplot markers and `linewidth` to increase the sizes of your plot lines.
Using color (and nothing else) to distinguish between different plot elements will make your plots unreadable to anyone who is colorblind, or who happens to have a black-and-white office printer. For lines, the `linestyle` parameter lets you use different types of lines. For scatterplots, `marker` lets you change the shape of your points. If you're unsure about your colors, you can use [Coblis](https://www.color-blindness.com/coblis-color-blindness-simulator/) or [Color Oracle](https://colororacle.org/) to simulate what your plots would look like to those with colorblindness.
## Questions
### Q1: Minima and Maxima
Fill in the blanks below to plot the minimum GDP per capita over time for all the countries in Europe. Modify it again to plot the maximum GDP per capita over time for Europe.
```
data_europe = pandas.read_csv('data/gapminder_gdp_europe.csv', index_col='country')
data_europe.____.plot(label='min')
data_europe.____
plt.legend(loc='best')
plt.xticks(rotation=90
```
**Solution**
Click on the '...' below to show the solution.
```
data_europe = pandas.read_csv('../data/gapminder_gdp_europe.csv', index_col='country')
data_europe.min().plot(label='min')
data_europe.max().plot(label='max')
plt.legend(loc='best')
plt.xticks(rotation=90)
```
### Q2: Correlations
Modify the example in the notes to create a scatter plot showing the relationship between the minimum and maximum GDP per capita among the countries in Asia for each year in the data set. What relationship do you see (if any)?
`data_asia = pandas.read_csv('data/gapminder_gdp_asia.csv, index_col='country')
data_asia.describe() .T.plot(kind='scatter', x='min', y='max')`
You might note that the variability in the maximum is much higher than that of the minimum. Take a look at the maximum and the max indexes:
```
data_asia = pandas.read_csv('data/gapminder_gdp_asia.csv', index_col='country')
data_asia.max().plot()
print(data_asia.idxmax())
print(data_asia.idxmin())
```
**Solution**
Click on the '...' below to show the solution.
```
data_asia = pandas.read_csv('../data/gapminder_gdp_asia.csv', index_col='country')
data_asia.describe().T.plot(kind='scatter', x='min', y='max')
# No particular correlations can be seen between the minimum and maximum gdp
# values year on year. It seems the fortunes of asian countries do not rise and
# fall together.
data_asia = pandas.read_csv('../data/gapminder_gdp_asia.csv', index_col='country')
plt.style.use('default')
data_asia.max().plot()
# Seems the variability in this value is due to a sharp drop after 1972. Some
# geopolitics at play perhaps? Given the dominance of oil producing countries,
# maybe the Brent crude index would make an interesting comparison? Whilst
# Myanmar consistently has the lowest gdp, the highest gdb nation has varied
# more notably.
```
### Q3: More Correlations
This short program creates a plot showing the correlation between GDP and life expectancy for 2007, normalizing marker size by population:
```
data_all - pandas.read.csv9'../data/gpaminder_all.csv', index-col -'country'
data_all.plot(kind='scatter', x='gdpPercap_2007', y='lifeExp_2007',
s=data_all['pop_2007']/1e6)
```
Using online help and other resources, explain what each argument to `plot` does.
**Solution**
Click on the '...' below to show the solution.
```
data_all = pandas.read_csv('../data/gapminder_all.csv', index_col='country')
data_all.plot(kind='scatter', x='gdpPercap_2007', y='lifeExp_2007',
s=data_all['pop_2007']/1e6)
# A good place to look is the documentation for the plot function -
# help(data_all.plot).
# kind - As seen already this determines the kind of plot to be drawn.
# x and y - A column name or index that determines what data will be placed on
# the x and y axes of the plot
# s - Details for this can be found in the documentation of plt.scatter. A
# single number or one value for each data point. Determines the size of the
# plotted points.
```
| github_jupyter |
Tornado 异步非阻塞浅析
===
## 先上代码演示
```
#!/usr/bin/python
# coding: utf-8
"""
File: demo.py
Author: zhangxu01 <zhangxu01@zhihu.com>
Date: 2017-08-28 22:59
Description: demo
"""
import random
import time
import urllib
import requests
import tornado
from tornado import gen, web
from tornado.httpclient import AsyncHTTPClient, HTTPRequest
@gen.coroutine
def async_fetch(url, method="GET", data=None, timeout=2 * 30,):
request = url
if isinstance(url, HTTPRequest):
url = request.url
if not isinstance(url, HTTPRequest):
kwargs = {
"connect_timeout": timeout,
"request_timeout": timeout,
}
if data:
if method == "GET":
url += '?' + urllib.urlencode(data)
else:
kwargs["body"] = urllib.urlencode(data)
request = HTTPRequest(url, method, **kwargs)
http_client = AsyncHTTPClient()
response = yield http_client.fetch(request)
raise gen.Return(response)
@gen.coroutine
def service_method():
url = "http://127.0.0.1:2345/api/time"
data = {
"time": str(time.time())
}
response = yield async_fetch(url, data=data)
raise gen.Return(response)
class NoBlockHandler(tornado.web.RequestHandler):
@web.asynchronous
@gen.coroutine
def get(self):
result = yield service_method()
self.write(result.body)
self.finish()
class BlockHandler(tornado.web.RequestHandler):
def get(self):
begin_time = time.time()
response = requests.get("http://127.0.0.1:2345/api/time", data={"time": str(begin_time)})
self.write(response.content)
self.finish()
class TimeHandler(tornado.web.RequestHandler):
def get(self):
req_time = self.get_argument("time", "")
sleep_time = random.randint(10, 500) * 0.001
time.sleep(sleep_time)
self.write("b:{},s:{} => e:{}".format(req_time, sleep_time, time.time()))
self.finish()
class Application(tornado.web.Application):
def __init__(self):
settings = {
"xsrf_cookies": False,
}
handlers = [
(r"/api/noblock", NoBlockHandler), # 非阻塞 IO 请求
(r"/api/block", BlockHandler), # 阻塞 IO 请求
(r"/api/time", TimeHandler), # 被请求接口
]
tornado.web.Application.__init__(self, handlers, **settings)
def main():
""" main"""
tornado.httpserver.HTTPServer(Application()).listen(2345)
tornado.ioloop.IOLoop.instance().start()
if __name__ == "__main__":
main()
```
http://python.jobbole.com/84188/
http://blog.csdn.net/wyx819/article/details/45420017
## IOLoop 模块
主要用来处理任务调度
```
def start(self):
# do something...
try:
while True:
# do something...
try:
event_pairs = self._impl.poll(poll_timeout)
except Exception as e:
# do something...
pass
self._events.update(event_pairs)
while self._events:
fd, events = self._events.popitem()
try:
fd_obj, handler_func = self._handlers[fd]
handler_func(fd_obj, events)
except (OSError, IOError) as e:
# do something...
pass
except Exception:
# do something...
pass
finally:
# do something...
```
`self._impl.poll(poll_timeout)` 会接收请求,等待期间阻塞状态,超时会进入下一次轮训继续等待。
`handler_func(fd_obj, events)` 会去处理接收的请求,通过HTTPServer -> TCPServer -> gen.coroutine
## gen.coroutine 装饰器
主要用来将一个 IOLoop 任务拆分成多个异步子任务
```
def coroutine(func, replace_callback=True):
@functools.wraps(wrapped)
def wrapper(*args, **kwargs):
future = TracebackFuture()
# do something...
try:
result = func(*args, **kwargs)
except Exception:
# do something...
pass
else:
if isinstance(result, GeneratorType):
try:
yielded = next(result)
# do something...
except Exception:
future.set_exc_info(sys.exc_info())
else:
Runner(result, future, yielded)
yielded = None
try:
return future
finally:
future = None
future.set_result(result)
return future
return wrapper
```
```
def am():
abc = 1234
try:
print "try"
return abc
finally:
print "finally"
abc = None
print "out"
return abc
am()
```
```
try
finally
Out[19]: 1234
```
高性能 IO 模型在 Tornado 中的应用
===
## 大纲
### 1. 基本概念
* 同步与异步
* 阻塞与非阻塞
* 用户空间和内核空间
* 缓存 I/O
### 2. 四种常见 IO 模型
* 同步阻塞 IO
* 同步非阻塞 IO
* 异步 IO
* IO 多路复用
### 3. Tornado 异步非阻塞 IO
* Tornado AsyncHTTPClient() 实例演示
* Tornado IOLoop 的调度方式
--------
## 1. 基本概念
### 同步和异步
#### 描述的是用户线程与内核的交互方式
*同步是指用户线程发起IO请求后需要等待或者轮询内核IO操作完成后才能继续执行;而异步是指用户线程发起IO请求后仍继续执行,当内核IO操作完成后会通知用户线程,或者调用用户线程注册的回调函数。*
### 阻塞和非阻塞
#### 描述的是用户线程调用内核IO的操作方式
*阻塞是指IO操作需要彻底完成后才返回到用户空间;而非阻塞是指IO操作被调用后立即返回给用户一个状态值,无需等到IO操作彻底完成。*
#### [同步异步 阻塞非阻塞的区别](https://www.zhihu.com/question/19732473)
同步和异步关注的是消息通信机制;阻塞和非阻塞关注的是程序在等待调用结果(消息,返回值)时的状态。
同步,就是在发出一个『调用』时,在没有得到结果之前,该『调用』就不返回。但是一旦调用返回,就得到返回值了。换句话说,就是由『调用者』主动等待这个『调用』的结果。异步则是相反,『调用』在发出之后,这个调用就直接返回了,所以没有返回结果。换句话说,当一个异步过程调用发出后,调用者不会立刻得到结果。而是在『调用』发出后,『被调用者』通过状态、通知来通知调用者,或通过回调函数处理这个调用。
阻塞调用是指调用结果返回之前,当前线程会被挂起,调用线程只有在得到结果之后才会返回。非阻塞调用指在不能立刻得到结果之前,该调用不会阻塞当前线程。
### 用户空间与内核空间
#### Linux 操作系统和驱动程序运行在内核空间,应用程序运行在用户空间
*现在操作系统都是采用虚拟存储器,那么对32位操作系统而言,它的寻址空间(虚拟存储空间)为4G(2的32次方)。操作系统的核心是内核,独立于普通的应用程序,可以访问受保护的内存空间,也有访问底层硬件设备的所有权限。为了保证用户进程不能直接操作内核(kernel),保证内核的安全,操心系统将虚拟空间划分为两部分,一部分为内核空间,一部分为用户空间。针对linux操作系统而言,将最高的1G字节(从虚拟地址0xC0000000到0xFFFFFFFF),供内核使用,称为内核空间,而将较低的3G字节(从虚拟地址0x00000000到0xBFFFFFFF),供各个进程使用,称为用户空间。*
### 缓存 I/O
*又被称作标准 I/O,大多数文件系统的默认 I/O 操作都是缓存 I/O。在 Linux 的缓存 I/O 机制中,操作系统会将 I/O 的数据缓存在文件系统的页缓存( page cache )中,也就是说,数据会先被拷贝到操作系统内核的缓冲区中,然后才会从操作系统内核的缓冲区拷贝到应用程序的地址空间。*
## 2. 四种常见 IO 模型
### 同步阻塞 IO

*用户线程通过系统调用read发起IO读操作,由用户空间转到内核空间。内核等到数据包到达后,然后将接收的数据拷贝到用户空间,完成read操作。用户需要等待read将socket中的数据读取到buffer后,才继续处理接收的数据。整个IO请求的过程中,用户线程是被阻塞的,这导致用户在发起IO请求时,不能做任何事情,对CPU的资源利用率不够。*
### 同步非阻塞 IO

*同步非阻塞IO是在同步阻塞IO的基础上,将socket设置为NONBLOCK。这样做用户线程可以在发起IO请求后可以立即返回。用户需要不断地调用read,尝试读取socket中的数据,直到读取成功后,才继续处理接收的数据。整个IO请求的过程中,虽然用户线程每次发起IO请求后可以立即返回,但是为了等到数据,仍需要不断地轮询、重复请求,消耗了大量的CPU的资源。一般很少直接使用这种模型,而是在其他IO模型中使用非阻塞IO这一特性。*
### 异步 IO

*异步IO模型中,用户线程直接使用内核提供的异步IO API发起read请求,且发起后立即返回,继续执行用户线程代码。不过此时用户线程已经将调用的AsynchronousOperation和CompletionHandler注册到内核,然后操作系统开启独立的内核线程去处理IO操作。当read请求的数据到达时,由内核负责读取socket中的数据,并写入用户指定的缓冲区中。最后内核将read的数据和用户线程注册的CompletionHandler分发给内部Proactor,Proactor将IO完成的信息通知给用户线程(一般通过调用用户线程注册的完成事件处理函数),完成异步IO。*
### IO 多路复用

*IO多路复用模型是建立在内核提供的多路分离函数select基础之上的,使用select函数可以避免同步非阻塞IO模型中轮询等待的问题。通过Reactor的方式,可以将用户线程轮询IO操作状态的工作统一交给handle_events事件循环进行处理。用户线程注册事件处理器之后可以继续执行做其他的工作(异步),而Reactor线程负责调用内核的select函数检查socket状态。当有socket被激活时,则通知相应的用户线程(或执行用户线程的回调函数),执行handle_event进行数据读取、处理的工作。由于select函数是阻塞的,因此多路IO复用模型也被称为异步阻塞IO模型。注意,这里的所说的阻塞是指select函数执行时线程被阻塞,而不是指socket。一般在使用IO多路复用模型时,socket都是设置为NONBLOCK的,不过这并不会产生影响,因为用户发起IO请求时,数据已经到达了,用户线程一定不会被阻塞。*
Linux支持IO多路复用的系统调用有select、poll、epoll,这些调用都是内核级别的,从 Reactor 和内核的交互方式来看它们都是同步I/O,先是block住等待就绪的socket,再是block住将数据从内核拷贝到用户内存。

#### Epoll
epoll是在2.6内核中提出的,相对于select和poll来说,epoll更加灵活。epoll使用事件的就绪通知方式,通过epoll_ctl注册fd,一旦该fd就绪,内核就会采用类似callback的回调机制来激活该fd,epoll_wait便可以收到通知。
#### epoll_create
用来创建一个 epoll 描述符( 就是创建了一个 epoll )
#### epoll_ctl
操作 epoll 中的 event;可用参数有:
EPOLL_CTL_ADD 添加一个新的epoll事件
EPOLL_CTL_DEL 删除一个epoll事件
EPOLL_CTL_MOD 改变一个事件的监听方式
#### epoll_wait
就是让 epoll 开始工作,里面有个参数 timeout,当设置为非 0 正整数时,会监听(阻塞) timeout 秒;设置为 0 时立即返回,设置为 -1 时一直监听。
在监听时有数据活跃的连接时其返回活跃的文件句柄列表。
## 3. Tornado 异步非阻塞 IO
### Tornado AsyncHTTPClient() 实例演示
```
#!/usr/bin/python
# coding: utf-8
"""
File: demo.py
Author: zhangxu01 <zhangxu01@zhihu.com>
Date: 2017-08-28 22:59
Description: demo
"""
import random
import time
import urllib
import requests
import tornado
from tornado import gen, web
from tornado.httpclient import AsyncHTTPClient, HTTPRequest
@gen.coroutine
def async_fetch(url, method="GET", data=None, timeout=2 * 30,):
request = url
if isinstance(url, HTTPRequest):
url = request.url
if not isinstance(url, HTTPRequest):
kwargs = {
"connect_timeout": timeout,
"request_timeout": timeout,
}
if data:
if method == "GET":
url += '?' + urllib.urlencode(data)
else:
kwargs["body"] = urllib.urlencode(data)
request = HTTPRequest(url, method, **kwargs)
http_client = AsyncHTTPClient()
response = yield http_client.fetch(request)
raise gen.Return(response)
@gen.coroutine
def service_method():
url = "http://127.0.0.1:2345/api/block2"
data = {
"time": str(time.time())
}
response = yield async_fetch(url, data=data)
raise gen.Return(response)
class NoBlockHandler(tornado.web.RequestHandler):
@web.asynchronous
@gen.coroutine
def get(self):
result = yield service_method()
self.write(result.body)
self.finish()
class BlockHandler(tornado.web.RequestHandler):
def get(self):
begin_time = time.time()
response = requests.get("http://127.0.0.1:2345/api/block2", data={"time": str(begin_time)})
self.write(response.content)
self.finish()
class BlockHandler2(tornado.web.RequestHandler):
def get(self):
req_time = self.get_argument("time", "")
sleep_time = random.randint(10, 500) * 0.001
time.sleep(sleep_time)
self.write("b:{},s:{} => e:{}".format(req_time, sleep_time, time.time()))
self.finish()
class Application(tornado.web.Application):
def __init__(self):
settings = {
"xsrf_cookies": False,
}
handlers = [
(r"/api/noblock", NoBlockHandler),
(r"/api/block", BlockHandler),
(r"/api/block2", BlockHandler2),
]
tornado.web.Application.__init__(self, handlers, **settings)
def main():
""" main"""
tornado.httpserver.HTTPServer(Application()).listen(2345)
tornado.ioloop.IOLoop.instance().start()
if __name__ == "__main__":
pass
# main()
```
### Tornado IOLoop 的调度方式
#### epoll 操作
epoll_ctl:这个三个方法分别对应 epoll_ctl 中的 add 、 modify 、 del 参数。 所以这三个方法实现了 epoll 的 epoll_ctl 。
epoll_create:然后 epoll 的生成在前文 EPollIOLoop 的初始化中就已经完成了:`super(EPollIOLoop, self).initialize(impl=select.epoll(), **kwargs)`。 这个相当于 epoll_create 。
epoll_wait:epoll_wait 操作则在 start() 中:`event_pairs = self._impl.poll(poll_timeout)`
epoll_close:而 epoll 的 close 则在 PollIOLoop 中的 close 方法内调用:`self._impl.close()`完成。
#### IOLoop 的核心调度集中在 start() 方法中。
start 方法中主要分三个部分:一个部分是对超时的相关处理;一部分是 epoll 事件通知阻塞、接收;一部分是对 epoll 返回I/O事件的处理。
```
class PollIOLoop(IOLoop):
def initialize(self, impl, time_func=None, **kwargs):
super(PollIOLoop, self).initialize(**kwargs)
self._impl = impl
if hasattr(self._impl, 'fileno'):
set_close_exec(self._impl.fileno())
self.time_func = time_func or time.time
self._handlers = {}
self._events = {}
self._callbacks = collections.deque() # 双端队列
self._timeouts = []
self._cancellations = 0
self._running = False
self._stopped = False
self._closing = False
self._thread_ident = None
self._pid = os.getpid()
self._blocking_signal_threshold = None
self._timeout_counter = itertools.count()
self._waker = Waker()
self.add_handler(self._waker.fileno(),
lambda fd, events: self._waker.consume(),
self.READ)
@classmethod
def configurable_base(cls):
return PollIOLoop
@classmethod
def configurable_default(cls):
"""根据系统环境选择"""
if hasattr(select, "epoll"):
from tornado.platform.epoll import EPollIOLoop
return EPollIOLoop
if hasattr(select, "kqueue"):
# Python 2.6+ on BSD or Mac
from tornado.platform.kqueue import KQueueIOLoop
return KQueueIOLoop
from tornado.platform.select import SelectIOLoop
return SelectIOLoop
def close(self, all_fds=False):
self._closing = True
self.remove_handler(self._waker.fileno())
if all_fds:
for fd, handler in list(self._handlers.values()):
self.close_fd(fd)
self._waker.close()
self._impl.close()
self._callbacks = None
self._timeouts = None
def add_handler(self, fd, handler, events):
"""注册事件"""
fd, obj = self.split_fd(fd)
self._handlers[fd] = (obj, stack_context.wrap(handler))
self._impl.register(fd, events | self.ERROR)
def update_handler(self, fd, events):
"""更新事件"""
fd, obj = self.split_fd(fd)
self._impl.modify(fd, events | self.ERROR)
def remove_handler(self, fd):
"""删除事件"""
fd, obj = self.split_fd(fd)
self._handlers.pop(fd, None)
self._events.pop(fd, None)
try:
self._impl.unregister(fd)
except Exception:
gen_log.debug("Error deleting fd from IOLoop", exc_info=True)
def start(self):
# 判断运行状态
if self._running:
raise RuntimeError("IOLoop is already running")
if os.getpid() != self._pid:
raise RuntimeError("Cannot share PollIOLoops across processes")
self._setup_logging()
if self._stopped:
self._stopped = False
return
# 启动当前实例
old_current = getattr(IOLoop._current, "instance", None)
IOLoop._current.instance = self
self._thread_ident = thread.get_ident()
self._running = True
try:
while True:
ncallbacks = len(self._callbacks) # 活动队列的长度
due_timeouts = [] # 存放这个周期已过期且有回调的任务
# 处理超时的任务
if self._timeouts:
now = self.time()
while self._timeouts: # 最小堆
# 处理掉超时的
if self._timeouts[0].callback is None: # todo ?
heapq.heappop(self._timeouts)
self._cancellations -= 1
elif self._timeouts[0].deadline <= now:
due_timeouts.append(heapq.heappop(self._timeouts))
else:
break
# 当超时计数器超过 512 并且大于超时队列一半的长度时,计数器归零,并弹出所有超时无 callback 任务
if (self._cancellations > 512 and
self._cancellations > (len(self._timeouts) >> 1)):
self._cancellations = 0
self._timeouts = [x for x in self._timeouts
if x.callback is not None]
heapq.heapify(self._timeouts) # 最小堆化处理
# 运行所有的可回调任务
for i in range(ncallbacks):
self._run_callback(self._callbacks.popleft())
# 运行所有已过期的任务
for timeout in due_timeouts:
if timeout.callback is not None:
self._run_callback(timeout.callback)
due_timeouts = timeout = None
# 这里设置等待时间
if self._callbacks:
poll_timeout = 0.0
elif self._timeouts:
poll_timeout = self._timeouts[0].deadline - self.time()
poll_timeout = max(0, min(poll_timeout, _POLL_TIMEOUT))
else:
poll_timeout = _POLL_TIMEOUT
# 检查中断状态
if not self._running:
break
if self._blocking_signal_threshold is not None:
signal.setitimer(signal.ITIMER_REAL, 0, 0)
try:
# 通过 self._impl.poll(poll_timeout) 进行事件阻塞,当有事件通知或超时时 poll 返回特定的 event_pairs。
event_pairs = self._impl.poll(poll_timeout) # 获取返回的活跃事件队列
except Exception as e:
if errno_from_exception(e) == errno.EINTR:
continue
else:
raise
if self._blocking_signal_threshold is not None:
signal.setitimer(signal.ITIMER_REAL,
self._blocking_signal_threshold, 0)
self._events.update(event_pairs) # 将活跃事件加入 _events
# 循环处理活跃事件
# epoll 返回通知事件后将新事件加入待处理队列,将就绪事件逐个弹出,通过stack_context.wrap(handler)保存的可执行对象调用事件处理。
while self._events:
fd, events = self._events.popitem()
try:
fd_obj, handler_func = self._handlers[fd]
handler_func(fd_obj, events)
except (OSError, IOError) as e:
if errno_from_exception(e) == errno.EPIPE:
# Happens when the client closes the connection
pass
else:
self.handle_callback_exception(self._handlers.get(fd))
except Exception:
self.handle_callback_exception(self._handlers.get(fd))
fd_obj = handler_func = None
finally:
# reset the stopped flag so another start/stop pair can be issued
self._stopped = False
if self._blocking_signal_threshold is not None:
signal.setitimer(signal.ITIMER_REAL, 0, 0)
IOLoop._current.instance = old_current
if old_wakeup_fd is not None:
signal.set_wakeup_fd(old_wakeup_fd)
def stop(self):
self._running = False
self._stopped = True
self._waker.wake()
def time(self):
return self.time_func()
def call_at(self, deadline, callback, *args, **kwargs):
timeout = _Timeout(
deadline,
functools.partial(stack_context.wrap(callback), *args, **kwargs),
self)
heapq.heappush(self._timeouts, timeout)
return timeout
def remove_timeout(self, timeout):
# Removing from a heap is complicated, so just leave the defunct
# timeout object in the queue (see discussion in
# http://docs.python.org/library/heapq.html).
# If this turns out to be a problem, we could add a garbage
# collection pass whenever there are too many dead timeouts.
timeout.callback = None
self._cancellations += 1
def add_callback(self, callback, *args, **kwargs):
if self._closing:
return
# Blindly insert into self._callbacks. This is safe even
# from signal handlers because deque.append is atomic.
self._callbacks.append(functools.partial(
stack_context.wrap(callback), *args, **kwargs))
if thread.get_ident() != self._thread_ident:
# This will write one byte but Waker.consume() reads many
# at once, so it's ok to write even when not strictly
# necessary.
self._waker.wake()
else:
# If we're on the IOLoop's thread, we don't need to wake anyone.
pass
def add_callback_from_signal(self, callback, *args, **kwargs):
with stack_context.NullContext():
self.add_callback(callback, *args, **kwargs)
```
### 参考资料
> https://segmentfault.com/a/1190000005659237
> http://www.cnblogs.com/fanzhidongyzby/p/4098546.html
> http://blog.csdn.net/iter_zc/article/details/39291647
> https://segmentfault.com/a/1190000003063859
> http://blog.csdn.net/feitianxuxue/article/details/8936802
> https://hexiangyu.me/posts/15
> http://blog.csdn.net/lisonglisonglisong/article/details/51328062
> http://blog.csdn.net/wyx819/article/details/45420017
> https://segmentfault.com/a/1190000002971992
> http://www.nowamagic.net/academy/detail/13321037
```
# todo future, reactor, promise, ioloop协作,callback,future,之间的关系;twisted,gevent关联区别,应用场景,优缺点。
```
沛然
陈思
张旭
```
%%latex
```
| github_jupyter |
```
import matplotlib.pyplot as plt
%matplotlib inline
import IPython.display
import librosa.display
import numpy as np
import librosa
import tensorflow as tf
import glob
c_drone_path = '../../../1m/*.wav'
m_drone_path = '../../../20m/*.wav'
f_drone_path = '../../../50m/*.wav'
background_path = '../../../40sec.wav'
drone_files = glob.glob(drone_path)
background_files = glob.glob(background_path)
CHUNK_SIZE = 8192
SR = 22050
N_MFCC = 16
def load(files, sr=22050):
[raw, sr] = librosa.load(files[0], sr=sr)
for f in files[1:]:
[array, sr] = librosa.load(f, sr=sr)
raw = np.hstack((raw, array))
print(raw.shape)
return raw
drone_raw = load(drone_files)
background_raw = load(background_files)
def mfcc4(raw, label, chunk_size=8192, window_size=4096, sr=22050, n_mfcc=16, n_frame=16):
mfcc = np.empty((0, n_mfcc, n_frame))
y = []
print(raw.shape)
for i in range(0, len(raw), chunk_size//2):
mfcc_slice = librosa.feature.mfcc(raw[i:i+chunk_size], sr=sr, n_mfcc=n_mfcc) #n_mfcc,17
if mfcc_slice.shape[1] < 17:
print("small end:", mfcc_slice.shape)
continue
mfcc_slice = mfcc_slice[:,:-1]
mfcc_slice = mfcc_slice.reshape((1, mfcc_slice.shape[0], mfcc_slice.shape[1]))
mfcc = np.vstack((mfcc, mfcc_slice))
y.append(label)
y = np.array(y)
return mfcc, y
mfcc_drone, y_drone = mfcc4(drone_raw, 1)
mfcc_background, y_background = mfcc4(background_raw, 0)
print(mfcc_drone.shape, y_drone.shape)
print(mfcc_background.shape, y_background.shape)
X = np.concatenate((mfcc_drone, mfcc_background), axis=0)
y = np.hstack((y_drone, y_background))
print(X.shape, y.shape)
n_labels = y.shape[0]
n_unique_labels = 2
y_encoded = np.zeros((n_labels, n_unique_labels))
y_encoded[np.arange(n_labels), y] = 1
print(y_encoded.shape)
X_train2 = X.reshape(X.shape[0], X.shape[1], X.shape[2], 1)
print(X_train2.shape)
n_mfcc = 16
n_frame = 16
n_classes = 2
n_channels = 1
learning_rate = 0.0002 ## 괜찮나용?
X = tf.placeholder(tf.float32, shape=[None,n_mfcc*n_frame*n_channels])
X = tf.reshape(X, [-1, n_mfcc, n_frame, n_channels])
Y = tf.placeholder(tf.float32, shape=[None,n_classes])
conv1 = tf.layers.conv2d(inputs=X, filters=1, kernel_size=[3, 3],
padding="SAME", activation=tf.nn.relu)
pool1 = tf.layers.max_pooling2d(inputs=conv1, pool_size=[2, 2],
padding="SAME", strides=1)
conv2 = tf.layers.conv2d(inputs=pool1, filters=1, kernel_size=[3, 3],
padding="SAME", activation=tf.nn.relu)
pool2 = tf.layers.max_pooling2d(inputs=conv2, pool_size=[2, 2],
padding="SAME", strides=1)
flat = tf.reshape(pool2, [-1, 16*16*1])
dense2 = tf.layers.dense(inputs=flat, units=625, activation=tf.nn.relu)
logits = tf.layers.dense(inputs=dense2, units=2)
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits_v2(logits=logits, labels=Y))
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(cost)
sess = tf.Session()
saver = tf.train.Saver()
# 모델 불러오는 path
saver.restore(sess, '../../model/CNN/cnn_model')
# Prediction
y_pred = sess.run(tf.argmax(logits,1), feed_dict={X:X_train2})
y_true = sess.run(tf.argmax(y_encoded,1))
# Print Result
from sklearn.metrics import precision_recall_fscore_support
p,r,f,s = precision_recall_fscore_support(y_true, y_pred, average='micro')
print("F-Score:", round(f,3))
from sklearn.metrics import accuracy_score
print("Accuracy: ", accuracy_score(y_true, y_pred))
#fig = plt.figure(figsize=(15,9))
#ax = fig.add_subplot(1,1,1)
#ax.plot(np.linspace(0,len(y_pred), len(y_pred)),y_true, 'x')
fig = plt.figure(figsize=(15,9))
ax = fig.add_subplot(1,1,1)
ax.plot(np.linspace(0,len(y_pred), len(y_pred)),y_pred, 'x')
from sklearn.metrics import classification_report
print(classification_report(y_true, y_pred))
from sklearn.metrics import confusion_matrix
print(confusion_matrix(y_true, y_pred))
```
| github_jupyter |
## 1. The most Nobel of Prizes
<p><img style="float: right;margin:5px 20px 5px 1px; max-width:250px" src="https://assets.datacamp.com/production/project_441/img/Nobel_Prize.png"></p>
<p>The Nobel Prize is perhaps the world's most well known scientific award. Except for the honor, prestige and substantial prize money the recipient also gets a gold medal showing Alfred Nobel (1833 - 1896) who established the prize. Every year it's given to scientists and scholars in the categories chemistry, literature, physics, physiology or medicine, economics, and peace. The first Nobel Prize was handed out in 1901, and at that time the Prize was very Eurocentric and male-focused, but nowadays it's not biased in any way whatsoever. Surely. Right?</p>
<p>Well, we're going to find out! The Nobel Foundation has made a dataset available of all prize winners from the start of the prize, in 1901, to 2016. Let's load it in and take a look.</p>
```
# Loading in required libraries
# ... YOUR CODE FOR TASK 1 ...
import pandas as pd
import seaborn as sns
import numpy as np
import matplotlib.pyplot as plt
# Reading in the Nobel Prize data
nobel = pd.read_csv("datasets/nobel.csv")
# Taking a look at the first several winners
nobel.head(6)
# ... YOUR CODE FOR TASK 1 ...
```
## 2. So, who gets the Nobel Prize?
<p>Just looking at the first couple of prize winners, or Nobel laureates as they are also called, we already see a celebrity: Wilhelm Conrad Röntgen, the guy who discovered X-rays. And actually, we see that all of the winners in 1901 were guys that came from Europe. But that was back in 1901, looking at all winners in the dataset, from 1901 to 2016, which sex and which country is the most commonly represented? </p>
<p>(For <em>country</em>, we will use the <code>birth_country</code> of the winner, as the <code>organization_country</code> is <code>NaN</code> for all shared Nobel Prizes.)</p>
```
# Display the number of (possibly shared) Nobel Prizes handed
# out between 1901 and 2016
result = len(nobel)
display(result)
# ... YOUR CODE FOR TASK 2 ...
# Display the number of prizes won by male and female recipients.
display(nobel['sex'].value_counts())
# ... YOUR CODE FOR TASK 2 ...
# Display the number of prizes won by the top 10 nationalities.
nobel['birth_country'].value_counts().head(10)
# ... YOUR CODE FOR TASK 2 ...
```
## 3. USA dominance
<p>Not so surprising perhaps: the most common Nobel laureate between 1901 and 2016 was a man born in the United States of America. But in 1901 all the winners were European. When did the USA start to dominate the Nobel Prize charts?</p>
```
# Calculating the proportion of USA born winners per decade
nobel['usa_born_winner']=nobel['birth_country']=="United States of America"
nobel['decade'] = (np.floor(nobel['year'] / 10) * 10).astype(int)
prop_usa_winners =nobel.groupby(nobel['decade'],as_index=False)[['decade','usa_born_winner']].mean()
# Display the proportions of USA born winners per decade
display(prop_usa_winners)
# ... YOUR CODE FOR TASK 3 ...
```
## 4. USA dominance, visualized
<p>A table is OK, but to <em>see</em> when the USA started to dominate the Nobel charts we need a plot!</p>
```
# Setting the plotting theme
sns.set()
# and setting the size of all plots.
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = [11, 7]
# Plotting USA born winners
ax = sns.lineplot(x='decade',y='usa_born_winner',data=nobel)
# Adding %-formatting to the y-axis
from matplotlib.ticker import PercentFormatter
ax.yaxis.set_major_formatter(PercentFormatter(1.0))
# ... YOUR CODE FOR TASK 4 ...
```
## 5. What is the gender of a typical Nobel Prize winner?
<p>So the USA became the dominating winner of the Nobel Prize first in the 1930s and had kept the leading position ever since. But one group that was in the lead from the start, and never seems to let go, are <em>men</em>. Maybe it shouldn't come as a shock that there is some imbalance between how many male and female prize winners there are, but how significant is this imbalance? And is it better or worse within specific prize categories like physics, medicine, literature, etc.?</p>
```
# Calculating the proportion of female laureates per decade
nobel['female_winner'] = nobel['sex']=='Female'
prop_female_winners = nobel.groupby(['decade','category'],as_index=False)['decade','female_winner'].mean()
# Plotting USA born winners with % winners on the y-axis
# Setting the plotting theme
sns.set()
# and setting the size of all plots.
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = [11, 7]
# Plotting USA born winners
ax = sns.lineplot(x='decade',y='female_winner',data=prop_female_winners,hue='category')
# Adding %-formatting to the y-axis
from matplotlib.ticker import PercentFormatter
ax.yaxis.set_major_formatter(PercentFormatter(1.0))
# ... YOUR CODE FOR TASK 4 ...
# ... YOUR CODE FOR TASK 5 ...
```
## 6. The first woman to win the Nobel Prize
<p>The plot above is a bit messy as the lines are overplotting. But it does show some interesting trends and patterns. Overall the imbalance is pretty large with physics, economics, and chemistry having the largest imbalance. Medicine has a somewhat positive trend, and since the 1990s the literature prize is also now more balanced. The big outlier is the peace prize during the 2010s, but keep in mind that this just covers the years 2010 to 2016.</p>
<p>Given this imbalance, who was the first woman to receive a Nobel Prize? And in what category?</p>
```
# Picking out the first woman to win a Nobel Prize
nobel[nobel['female_winner']==True].nsmallest(1,'year')['full_name']
# ... YOUR CODE FOR TASK 5 ...
```
## 7. Repeat laureates
<p>For most scientists/writers/activists a Nobel Prize would be the crowning achievement of a long career. But for some people, one is just not enough, and few have gotten it more than once. Who are these lucky few? (Having won no Nobel Prize myself, I'll assume it's just about luck.)</p>
```
# Selecting the laureates that have received 2 or more prizes.
nobel.groupby('full_name').filter(lambda group: len(group) >= 2)
# ... YOUR CODE FOR TASK 5 ...
```
## 8. How old are you when you get the prize?
<p>The list of repeat winners contains some illustrious names! We again meet Marie Curie, who got the prize in physics for discovering radiation and in chemistry for isolating radium and polonium. John Bardeen got it twice in physics for transistors and superconductivity, Frederick Sanger got it twice in chemistry, and Linus Carl Pauling got it first in chemistry and later in peace for his work in promoting nuclear disarmament. We also learn that organizations also get the prize as both the Red Cross and the UNHCR have gotten it twice.</p>
<p>But how old are you generally when you get the prize?</p>
```
# Converting birth_date from String to datetime
nobel['birth_date'] = pd.to_datetime(nobel['birth_date'])
# Calculating the age of Nobel Prize winners
nobel['age'] = nobel['year'] - pd.to_datetime(nobel['birth_date']).dt.year
# Plotting the age of Nobel Prize winners
sns.lmplot(x='year',y='age',data=nobel)
```
## 9. Age differences between prize categories
<p>The plot above shows us a lot! We see that people use to be around 55 when they received the price, but nowadays the average is closer to 65. But there is a large spread in the laureates' ages, and while most are 50+, some are very young.</p>
<p>We also see that the density of points is much high nowadays than in the early 1900s -- nowadays many more of the prizes are shared, and so there are many more winners. We also see that there was a disruption in awarded prizes around the Second World War (1939 - 1945). </p>
<p>Let's look at age trends within different prize categories.</p>
```
# Same plot as above, but separate plots for each type of Nobel Prize
sns.lmplot(x='year',y='age',data=nobel,row='category')
# ... YOUR CODE FOR TASK 9 ...
```
## 10. Oldest and youngest winners
<p>More plots with lots of exciting stuff going on! We see that both winners of the chemistry, medicine, and physics prize have gotten older over time. The trend is strongest for physics: the average age used to be below 50, and now it's almost 70. Literature and economics are more stable. We also see that economics is a newer category. But peace shows an opposite trend where winners are getting younger! </p>
<p>In the peace category we also a winner around 2010 that seems exceptionally young. This begs the questions, who are the oldest and youngest people ever to have won a Nobel Prize?</p>
```
# The oldest winner of a Nobel Prize as of 2016
# ... YOUR CODE FOR TASK 10 ...
display(nobel.nlargest(1,'age'))
# The youngest winner of a Nobel Prize as of 2016
nobel.nsmallest(1,'age')
# ... YOUR CODE FOR TASK 10 ...
```
## 11. You get a prize!
<p><img style="float: right;margin:20px 20px 20px 20px; max-width:200px" src="https://assets.datacamp.com/production/project_441/img/paint_nobel_prize.png"></p>
<p>Hey! You get a prize for making it to the very end of this notebook! It might not be a Nobel Prize, but I made it myself in paint so it should count for something. But don't despair, Leonid Hurwicz was 90 years old when he got his prize, so it might not be too late for you. Who knows.</p>
<p>Before you leave, what was again the name of the youngest winner ever who in 2014 got the prize for "[her] struggle against the suppression of children and young people and for the right of all children to education"?</p>
```
# The name of the youngest winner of the Nobel Prize as of 2016
youngest_winner ='Malala Yousafzai'
```
| github_jupyter |
# Distribution of Weights in a Network
Varun Nayyar, 2020-08-23
Let us consider the simplest possible neural network, 1 input $x$, 1 output $y$ with some non-linearity $f$. This is expressed as
$$
\begin{aligned}
y = f(wx + b)
\end{aligned}
$$
where $w$, $b$ are the weight and bias in the network. Putting this into a slightly different form
$$
\begin{aligned}
y = f(w(x + b/w))
\end{aligned}
$$
we know that the activation function is centered at $-b/w$.
For this experiment, we look at the distribution of $-b/w$ for a swarm fitting to a
- trig function: sin and cos have very obvious turning points.
- ReLU activation: as a very simple activation, the $-b/w$ will correspond exactly to the turning points
- Single hidden layer: this makes interpretability a bit clearer since we have a clearer understanding of the mix
```
import numpy as np
import torch
from torch import nn
from matplotlib import pyplot as plt
from IPython.display import Video
from swarm import core, animator, networks
import env
plt.rcParams["figure.figsize"] = (12.0, 12.0)
SEED = 20
if not env.FULL:
NUM_EPOCHS = 4
NUM_BEES = 5
else:
NUM_EPOCHS = 400
NUM_BEES = 500
def bee_trainer(xt, yt, width=2, num_epochs=200):
"""Define a simple training loop for use with swarm"""
net = networks.flat_net(1, width, activation=nn.ReLU)
optimiser = torch.optim.SGD(net.parameters(), lr=0.01, momentum=0.9)
loss_func = torch.nn.MSELoss()
for epoch in range(num_epochs):
optimiser.zero_grad()
ypred = net(xt)
loss = loss_func(ypred, yt)
if torch.isnan(loss):
raise RuntimeError("NaN loss, poorly configured experiment")
loss.backward()
optimiser.step()
weight, bias, *_ = net.parameters()
yield ypred, weight.detach().flatten().numpy().copy(), bias.detach().numpy().copy()
def main():
xt = torch.linspace(-3 * np.pi, 3 * np.pi, 101)
yt = torch.sin(xt)
bp = {"xt": xt, "yt": yt, "width": 20, "num_epochs": NUM_EPOCHS}
res = core.swarm_train(bee_trainer, bp, num_bees=NUM_BEES, fields="ypred,weights,biases", seed=SEED)
bw = -res["biases"] / res["weights"]
# reduce range to be safe
bw = bw.clip(-10, 10)
ls = animator.LineSwarm.standard(xt.detach().numpy(), yt.detach().numpy(), res["ypred"][::10], set_xlim=(-10,10))
hist = animator.HistogramSwarm.from_swarm(
bw, 100, set_title="- Biases/Weights", set_ylabel="Count", set_xlim=(-10,10)
)
animator.swarm_animate([ls, hist], "weight_distr.mp4")
main()
Video("weight_distr.mp4", embed=True)
```
## Weight Distributions
We can see that the biases and weights cluster around the places where the sin curve turns. As you'd expect with the starting conditions being quite close to 0, we see that most of the bends assigned by the network fit into the first curves and not the turning points at extremities.
| github_jupyter |
# Self-Driving Car Engineer Nanodegree
## Project: **Finding Lane Lines on the Road**
***
In this project, you will use the tools you learned about in the lesson to identify lane lines on the road. You can develop your pipeline on a series of individual images, and later apply the result to a video stream (really just a series of images). Check out the video clip "raw-lines-example.mp4" (also contained in this repository) to see what the output should look like after using the helper functions below.
Once you have a result that looks roughly like "raw-lines-example.mp4", you'll need to get creative and try to average and/or extrapolate the line segments you've detected to map out the full extent of the lane lines. You can see an example of the result you're going for in the video "P1_example.mp4". Ultimately, you would like to draw just one line for the left side of the lane, and one for the right.
In addition to implementing code, there is a brief writeup to complete. The writeup should be completed in a separate file, which can be either a markdown file or a pdf document. There is a [write up template](https://github.com/udacity/CarND-LaneLines-P1/blob/master/writeup_template.md) that can be used to guide the writing process. Completing both the code in the Ipython notebook and the writeup template will cover all of the [rubric points](https://review.udacity.com/#!/rubrics/322/view) for this project.
---
Let's have a look at our first image called 'test_images/solidWhiteRight.jpg'. Run the 2 cells below (hit Shift-Enter or the "play" button above) to display the image.
**Note: If, at any point, you encounter frozen display windows or other confounding issues, you can always start again with a clean slate by going to the "Kernel" menu above and selecting "Restart & Clear Output".**
---
**The tools you have are color selection, region of interest selection, grayscaling, Gaussian smoothing, Canny Edge Detection and Hough Tranform line detection. You are also free to explore and try other techniques that were not presented in the lesson. Your goal is piece together a pipeline to detect the line segments in the image, then average/extrapolate them and draw them onto the image for display (as below). Once you have a working pipeline, try it out on the video stream below.**
---
<figure>
<img src="line-segments-example.jpg" width="380" alt="Combined Image" />
<figcaption>
<p></p>
<p style="text-align: center;"> Your output should look something like this (above) after detecting line segments using the helper functions below </p>
</figcaption>
</figure>
<p></p>
<figure>
<img src="laneLines_thirdPass.jpg" width="380" alt="Combined Image" />
<figcaption>
<p></p>
<p style="text-align: center;"> Your goal is to connect/average/extrapolate line segments to get output like this</p>
</figcaption>
</figure>
**Run the cell below to import some packages. If you get an `import error` for a package you've already installed, try changing your kernel (select the Kernel menu above --> Change Kernel). Still have problems? Try relaunching Jupyter Notebook from the terminal prompt. Also, see [this forum post](https://carnd-forums.udacity.com/cq/viewquestion.action?spaceKey=CAR&id=29496372&questionTitle=finding-lanes---import-cv2-fails-even-though-python-in-the-terminal-window-has-no-problem-with-import-cv2) for more troubleshooting tips.**
## Import Packages
```
#importing some useful packages
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import numpy as np
import cv2
%matplotlib inline
```
## Read in an Image
```
#reading in an image
image = mpimg.imread('test_images/solidWhiteRight.jpg')
#printing out some stats and plotting
print('This image is:', type(image), 'with dimesions:', image.shape)
plt.imshow(image) # if you wanted to show a single color channel image called 'gray', for example, call as plt.imshow(gray, cmap='gray')
```
## Ideas for Lane Detection Pipeline
**Some OpenCV functions (beyond those introduced in the lesson) that might be useful for this project are:**
`cv2.inRange()` for color selection
`cv2.fillPoly()` for regions selection
`cv2.line()` to draw lines on an image given endpoints
`cv2.addWeighted()` to coadd / overlay two images
`cv2.cvtColor()` to grayscale or change color
`cv2.imwrite()` to output images to file
`cv2.bitwise_and()` to apply a mask to an image
**Check out the OpenCV documentation to learn about these and discover even more awesome functionality!**
## Helper Functions
Below are some helper functions to help get you started. They should look familiar from the lesson!
```
import math
def grayscale(img):
"""Applies the Grayscale transform
This will return an image with only one color channel
but NOTE: to see the returned image as grayscale
(assuming your grayscaled image is called 'gray')
you should call plt.imshow(gray, cmap='gray')"""
#return cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# Or use BGR2GRAY if you read an image with cv2.imread()
return cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
def canny(img, low_threshold, high_threshold):
"""Applies the Canny transform"""
return cv2.Canny(img, low_threshold, high_threshold)
def gaussian_blur(img, kernel_size):
"""Applies a Gaussian Noise kernel"""
return cv2.GaussianBlur(img, (kernel_size, kernel_size), 0)
def region_of_interest(img, vertices):
"""
Applies an image mask.
Only keeps the region of the image defined by the polygon
formed from `vertices`. The rest of the image is set to black.
"""
#defining a blank mask to start with
mask = np.zeros_like(img)
#defining a 3 channel or 1 channel color to fill the mask with depending on the input image
if len(img.shape) > 2:
channel_count = img.shape[2] # i.e. 3 or 4 depending on your image
ignore_mask_color = (255,) * channel_count
else:
ignore_mask_color = 255
#filling pixels inside the polygon defined by "vertices" with the fill color
cv2.fillPoly(mask, vertices, ignore_mask_color)
#returning the image only where mask pixels are nonzero
masked_image = cv2.bitwise_and(img, mask)
return masked_image
def draw_lines_old(img, lines, color=[255, 0, 0], thickness=6):
"""
NOTE: this is the function you might want to use as a starting point once you want to
average/extrapolate the line segments you detect to map out the full
extent of the lane (going from the result shown in raw-lines-example.mp4
to that shown in P1_example.mp4).
Think about things like separating line segments by their
slope ((y2-y1)/(x2-x1)) to decide which segments are part of the left
line vs. the right line. Then, you can average the position of each of
the lines and extrapolate to the top and bottom of the lane.
This function draws `lines` with `color` and `thickness`.
Lines are drawn on the image inplace (mutates the image).
If you want to make the lines semi-transparent, think about combining
this function with the weighted_img() function below
"""
for line in lines:
for x1,y1,x2,y2 in line:
cv2.line(img,(x1,y1),(x2,y2), color, thickness)
def draw_lines(img, lines, color=[255, 0, 0], thickness=5):
"""
NOTE: this is the function you might want to use as a starting point once you want to
average/extrapolate the line segments you detect to map out the full
extent of the lane (going from the result shown in raw-lines-example.mp4
to that shown in P1_example.mp4).
Think about things like separating line segments by their
slope ((y2-y1)/(x2-x1)) to decide which segments are part of the left
line vs. the right line. Then, you can average the position of each of
the lines and extrapolate to the top and bottom of the lane.
This function draws `lines` with `color` and `thickness`.
Lines are drawn on the image inplace (mutates the image).
If you want to make the lines semi-transparent, think about combining
this function with the weighted_img() function below
"""
#define parameters here
left_m = 0
left_bias = 0
right_m = 0
right_bias = 0
left_size = 0
right_size = 0
height = img.shape[0]
width = img.shape[1]
#in this function, we use y = m * x + b; m is slope, and b is bias.
for line in lines:
for x1,y1,x2,y2 in line:
#calculate the slope
slope = ((y2 - y1)/(x2 - x1))
#print(slope)
#because in lane line detection, if the slope is close to 0, this line maybe is noise
if(slope < 0.5 and slope > -0.5):
break
point = np.mean(line, axis=0)
#left line
if slope < -0.5:
#sum of m & bias
left_m += slope
left_bias += point[1] - point[0] * slope
left_size += 1
#print(slope)
if slope > 0.5:
#sum of m & bias
right_m += slope
right_bias += point[1] - point[0] * slope
right_size += 1
#print(slope)
if(right_size>0):
#draw single right line
cv2.line(img,(int(width*9/16), int(right_m/right_size * int(width*9/16) + right_bias/right_size)),(width, int(right_m/right_size * width +right_bias/right_size)), color, thickness )
#cv2.line(img, (int((539 - right_bias/right_size)/(right_m/right_size)), 539), (549, int(right_m/right_size * 549 + right_bias/right_size)), color, thickness)
if(left_size > 0):
pass
#draw single left line
cv2.line(img, (0, int(left_bias/left_size)), (int(width*7/16), int(left_m/left_size * int(width*7/16) + left_bias/left_size)), color, thickness)
def hough_lines(img, rho, theta, threshold, min_line_len, max_line_gap, new_draw_fun = True):
"""
`img` should be the output of a Canny transform.
Returns an image with hough lines drawn.
"""
lines = cv2.HoughLinesP(img, rho, theta, threshold, np.array([]), minLineLength=min_line_len, maxLineGap=max_line_gap)
line_img = np.zeros((img.shape[0], img.shape[1], 3), dtype=np.uint8)
if(new_draw_fun):
draw_lines(line_img, lines)
else:
draw_lines_old(line_img, lines)
return line_img
# Python 3 has support for cool math symbols.
def weighted_img(img, initial_img, α=0.8, β=1., λ=0.):
"""
`img` is the output of the hough_lines(), An image with lines drawn on it.
Should be a blank image (all black) with lines drawn on it.
`initial_img` should be the image before any processing.
The result image is computed as follows:
initial_img * α + img * β + λ
NOTE: initial_img and img must be the same shape!
"""
return cv2.addWeighted(initial_img, α, img, β, λ)
```
## Test Images
Build your pipeline to work on the images in the directory "test_images"
**You should make sure your pipeline works well on these images before you try the videos.**
```
import os
os.listdir("test_images/")
```
## Build a Lane Finding Pipeline
Build the pipeline and run your solution on all test_images. Make copies into the test_images directory, and you can use the images in your writeup report.
Try tuning the various parameters, especially the low and high Canny thresholds as well as the Hough lines parameters.
```
# TODO: Build your pipeline that will draw lane lines on the test_images
# then save them to the test_images directory.
def fine_lane_lines_function(image, new_draw_func):
# define parameters here
kernel_size = 3
low_threshold = 50
high_threshold = 150
imshape = image.shape
vertices = np.array([[(0,imshape[0]),(imshape[1]*7/16,imshape[0]*5/8), (imshape[1]*9/16, imshape[0]*5/8), (imshape[1],imshape[0])]], dtype=np.int32)
#convert the RGB image to grayscale
gray = grayscale(image)
#blur the gray image
blur_gray = gaussian_blur(gray, kernel_size)
#using canny edge function for the blur gray image
edges = canny(blur_gray, low_threshold, high_threshold)
#masked edges image by using region_of_interest function
masked_edges = region_of_interest(edges, vertices)
#here we use hough transform to process the masked edges
#define parameters here
rho = 2 # distance resolution in pixels of the Hough grid
theta = np.pi/180 # angular resolution in radians of the Hough grid
threshold = 100 # minimum number of votes (intersections in Hough grid cell)
min_line_len = 100 #minimum number of pixels making up a line
max_line_gap = 160 # maximum gap in pixels between connectable line segments
line_image = np.copy(image)*0 # creating a blank to draw lines on
#calling hough transform function and output color_edges
color_edges = hough_lines(masked_edges, rho, theta, threshold, min_line_len, max_line_gap, new_draw_fun = new_draw_func)
lines_edges = weighted_img(color_edges, image)
return lines_edges
for img_ in os.listdir("test_images/"):
image_address = "test_images/"+img_
#read image from input address
image = cv2.imread(image_address)
lines_edges = fine_lane_lines_function(image, False)
#debug-code for visiulization
fig = plt.figure()
plt.imshow(lines_edges)
#saving image to the test_images directory
cv2.imwrite("test_images/test_"+img_,lines_edges)
```
## Test on Videos
You know what's cooler than drawing lanes over images? Drawing lanes over video!
We can test our solution on two provided videos:
`solidWhiteRight.mp4`
`solidYellowLeft.mp4`
**Note: if you get an `import error` when you run the next cell, try changing your kernel (select the Kernel menu above --> Change Kernel). Still have problems? Try relaunching Jupyter Notebook from the terminal prompt. Also, check out [this forum post](https://carnd-forums.udacity.com/questions/22677062/answers/22677109) for more troubleshooting tips.**
**If you get an error that looks like this:**
```
NeedDownloadError: Need ffmpeg exe.
You can download it by calling:
imageio.plugins.ffmpeg.download()
```
**Follow the instructions in the error message and check out [this forum post](https://carnd-forums.udacity.com/display/CAR/questions/26218840/import-videofileclip-error) for more troubleshooting tips across operating systems.**
```
# Import everything needed to edit/save/watch video clips
from moviepy.editor import VideoFileClip
from IPython.display import HTML
def process_image(image):
# NOTE: The output you return should be a color image (3 channel) for processing video below
# TODO: put your pipeline here,
# you should return the final output (image where lines are drawn on lanes)
result = fine_lane_lines_function(image, False)
return result
```
Let's try the one with the solid white lane on the right first ...
```
white_output = 'white_1.mp4'
clip1 = VideoFileClip("solidWhiteRight.mp4")
white_clip = clip1.fl_image(process_image) #NOTE: this function expects color images!!
%time white_clip.write_videofile(white_output, audio=False)
```
Play the video inline, or if you prefer find the video in your filesystem (should be in the same directory) and play it in your video player of choice.
```
HTML("""
<video width="960" height="540" controls>
<source src="{0}">
</video>
""".format(white_output))
yello_output = 'yello_1.mp4'
clip1 = VideoFileClip("solidYellowLeft.mp4")
white_clip = clip1.fl_image(process_image) #NOTE: this function expects color images!!
%time white_clip.write_videofile(yello_output, audio=False)
HTML("""
<video width="960" height="540" controls>
<source src="{0}">
</video>
""".format(yello_output))
```
## Improve the draw_lines() function
**At this point, if you were successful with making the pipeline and tuning parameters, you probably have the Hough line segments drawn onto the road, but what about identifying the full extent of the lane and marking it clearly as in the example video (P1_example.mp4)? Think about defining a line to run the full length of the visible lane based on the line segments you identified with the Hough Transform. As mentioned previously, try to average and/or extrapolate the line segments you've detected to map out the full extent of the lane lines. You can see an example of the result you're going for in the video "P1_example.mp4".**
**Go back and modify your draw_lines function accordingly and try re-running your pipeline. The new output should draw a single, solid line over the left lane line and a single, solid line over the right lane line. The lines should start from the bottom of the image and extend out to the top of the region of interest.**
```
def process_image(image):
# NOTE: The output you return should be a color image (3 channel) for processing video below
# TODO: put your pipeline here,
# you should return the final output (image where lines are drawn on lanes)
result = fine_lane_lines_function(image, True)
return result
white_output = 'white_2.mp4'
clip1 = VideoFileClip("solidWhiteRight.mp4")
white_clip = clip1.fl_image(process_image) #NOTE: this function expects color images!!
%time white_clip.write_videofile(white_output, audio=False)
HTML("""
<video width="960" height="540" controls>
<source src="{0}">
</video>
""".format(white_output))
```
Now for the one with the solid yellow lane on the left. This one's more tricky!
```
yellow_output = 'yellow_2.mp4'
clip2 = VideoFileClip('solidYellowLeft.mp4')
yellow_clip = clip2.fl_image(process_image)
%time yellow_clip.write_videofile(yellow_output, audio=False)
HTML("""
<video width="960" height="540" controls>
<source src="{0}">
</video>
""".format(yellow_output))
```
## Writeup and Submission
If you're satisfied with your video outputs, it's time to make the report writeup in a pdf or markdown file. Once you have this Ipython notebook ready along with the writeup, it's time to submit for review! Here is a [link](https://github.com/udacity/CarND-LaneLines-P1/blob/master/writeup_template.md) to the writeup template file.
## Optional Challenge
Try your lane finding pipeline on the video below. Does it still work? Can you figure out a way to make it more robust? If you're up for the challenge, modify your pipeline so it works with this video and submit it along with the rest of your project!
```
def pipeline(image, new_draw_func):
# define parameters here
#print(image.shape)
kernel_size = 6
low_threshold = 50
high_threshold = 150
imshape = image.shape
vertices = np.array([[(0,imshape[0]),(imshape[1]*7/16,imshape[0]*5/8), (imshape[1]*9/16, imshape[0]*5/8), (imshape[1],imshape[0])]], dtype=np.int32)
#using hsv image
hsv = cv2.cvtColor(image, cv2.COLOR_RGB2HSV)
yellow = cv2.inRange(hsv, (20, 80, 80), (25, 255, 255))
white = cv2.inRange(hsv, (0, 0, 180), (255, 25, 255))
#convert to grayscale
gray = cv2.bitwise_or(yellow, white)
#using canny edge function for the blur gray image
edges = canny(gray, low_threshold, high_threshold)
#masked edges image by using region_of_interest function
masked_edges = region_of_interest(edges, vertices)
#here we use hough transform to process the masked edges
#define parameters here
rho = 4 # distance resolution in pixels of the Hough grid
theta = np.pi/180 # angular resolution in radians of the Hough grid
threshold = 100 # minimum number of votes (intersections in Hough grid cell)
min_line_len = 10 #minimum number of pixels making up a line
max_line_gap = 10 # maximum gap in pixels between connectable line segments
line_image = np.copy(image)*0 # creating a blank to draw lines on
#calling hough transform function and output color_edges
color_edges = hough_lines(masked_edges, rho, theta, threshold, min_line_len, max_line_gap, new_draw_fun = new_draw_func)
lines_edges = weighted_img(color_edges, image)
return lines_edges
def process_image_1(image):
# NOTE: The output you return should be a color image (3 channel) for processing video below
# TODO: put your pipeline here,
# you should return the final output (image where lines are drawn on lanes)
result = pipeline(image, True)
return result
challenge_output = 'extra.mp4'
clip2 = VideoFileClip('challenge.mp4')
challenge_clip = clip2.fl_image(process_image_1)
%time challenge_clip.write_videofile(challenge_output, audio=False)
HTML("""
<video width="960" height="540" controls>
<source src="{0}">
</video>
""".format(challenge_output))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/dheerajrathee/IADS-Summer-School-2021/blob/main/GradientBoostingClassifier_IADS_2021.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
from google.colab import drive
drive.mount('/gdrive')
%cd /gdrive/My Drive/IADS-Summer-School-2021
!pwd
```
# **Gradient Boosting for Classification**
## 1. IRIS Data
## 2. Mushroom Classification Data (Kaggle)
# 1. IRIS DATA
## Dataset for Classification
> **Dataset:** [Iris dataset](https://scikit-learn.org/stable/datasets/index.html#iris-dataset).
* **Number of Instances:**
* 150 (50 in each of three classes)
* **Number of Attributes:**
* 4 numeric, predictive attributes and the class
* **Attribute Information:**
* sepal length in cm
* sepal width in cm
* petal length in cm
* petal width in cm
* **Classes:**
* Setosa (0)
* Versicolour (1)
* Virginica (2)
```
# Add liberaries
from sklearn import datasets # DATA
from sklearn.model_selection import train_test_split # to Split Train-Test data
from sklearn import ensemble # To get Gradient Boosting classifier
from sklearn import metrics # To generate evaluation metrices
from sklearn.model_selection import cross_val_predict
from sklearn.model_selection import cross_val_score
from sklearn.tree import export_graphviz # exporting the tree structure as dot file
from pydotplus.graphviz import graph_from_dot_data # export png image from dot file
from IPython.display import Image, SVG # Show the image within colab notebook
from graphviz import Source
import matplotlib.pyplot as plt
import pandas as pd # for basic data manipulations
import numpy as np
import warnings
warnings.filterwarnings('ignore')
```
### 1. Load Data
```
#load data and see meta info
iris = datasets.load_iris()
dir(iris)
```
### 2. Explore Data
```
# print type and shape of data
print(type(iris.data))
print(type(iris.target))
print(iris.data.shape)
print(iris.target.shape)
```
### 3. Create Panda Dataframe and do data manipulations
```
dfCls = pd.DataFrame(iris.data, columns=iris.feature_names)
dfCls.head()
# Add target data to the panda dataframe
dfCls['target'] = iris.target
dfCls.head()
```
### 4. Split the data for Training and Testing
```
X_train, X_test, y_train, y_test = train_test_split(dfCls.drop(['target'],axis='columns'), iris.target, test_size=0.2,random_state=0, stratify=iris.target)
print(X_train.shape)
print(X_test.shape)
```
### 5. Initialise a Gradient Boosting Classifier
```
gbClassifier = ensemble.GradientBoostingClassifier(criterion='friedman_mse', init=None,
learning_rate=0.1, loss='deviance', max_depth=3,
max_features=None, max_leaf_nodes=None,
min_impurity_decrease=0.0, min_impurity_split=None,
min_samples_leaf=1, min_samples_split=2,
min_weight_fraction_leaf=0.0, n_estimators=500,
n_iter_no_change=None, presort='auto',
random_state=0, subsample=1.0, tol=0.0001,
validation_fraction=0.1, verbose=0,
warm_start=False)
```
> ***Let's dig into*** **[tree.GradientBoostingClassifier](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.GradientBoostingClassifier.html#sklearn.ensemble.GradientBoostingClassifier)**
### 6. Model Evaluation on Train data
```
#perform 10 fold cross validation and plot the CM
CV_predicted = cross_val_predict(gbClassifier, X_train, y_train, cv=10) #CV predicted values (training data)
CV_score = cross_val_score(gbClassifier, X_train, y_train, cv=10) #CV model score (training data)
print("Cross validation Score on train data: ",CV_score.mean())
print("\n")
print("Confusion matrix on CV predictions (train data)")
print(metrics.confusion_matrix(y_train, CV_predicted)) # confusion matrix on CV predictions (train data)
print("\n")
print("Classification report CV predictions (train data)")
print(metrics.classification_report(y_train, CV_predicted, target_names=['Setosa', 'Versicolor', 'Virginica'])) # classification report CV predictions (train data)
```
### 7. Let's fit the GB model on Training data and perform prediction with the Test data
```
gbClassMdl = gbClassifier.fit(X_train,y_train)
y_predicted = gbClassMdl.predict(X_test)
```
### 8. Model Evaluation on Test Data
```
mdl_score = gbClassMdl.score(X_test,y_test) #model score (test data)
print ("Model Score on test data:",mdl_score)
print("\n")
print("Confusion matrix (test data)")
print(metrics.confusion_matrix(y_test, y_predicted)) #confusion matrix (test data)
print("\n")
print("Classification report (test data)")
print(metrics.classification_report(y_test, y_predicted, target_names=['Setosa', 'Versicolor', 'Virginica'])) # classification report (test data)
```
# 2. Mushroom Classification Data (Kaggle)
[See further details](https://www.kaggle.com/uciml/mushroom-classification)
### 1. Load Data
```
#load data from local drive
mushData = pd.read_csv('mushrooms.csv')
```
### 2. Explore Data
```
#print first five rows of the data
mushData.head()
#print size of the data
mushData.shape
#print data attributes
mushData.describe()
#print key informations about the data
mushData.info()
#Check the class balance
mushData['class'].value_counts()
```
### 3. Perform data manipulations
```
from sklearn.preprocessing import LabelEncoder #print first five rows of the data
labelencoder=LabelEncoder()
for col in mushData.columns:
mushData[col] = labelencoder.fit_transform(mushData[col]) #Transform categrical data to numerical data
mushData.head()
mushData.describe()
target = mushData['class'] #get the labels as targets and convert to numpy array
np.array(target, dtype=pd.Series)
```
### 4. Split the data for Training and Testing
```
X_train, X_test, y_train, y_test = train_test_split(mushData.drop(['class'],axis='columns'), target, test_size=0.2,random_state=123, stratify=target)
print(X_train.shape)
print(X_test.shape)
```
### 5. Perform Grid search for getting the best parameters
```
from sklearn.model_selection import GridSearchCV # get gridsearch with cross validation
#provide GB hyperparameters
gb_hyperparameters = {
"n_estimators": [100, 200],
'learning_rate': [0.05, 0.1, 0.2],
'max_depth': [1, 3, 5]
}
nfolds = 10 #number of folds for CV
gbClassifier = ensemble.GradientBoostingClassifier(random_state=123) #initialise GB classifier
# create Grid search object
gs_gb_clf = GridSearchCV(gbClassifier, gb_hyperparameters,
n_jobs=-1, cv=nfolds,
scoring='accuracy')
gs_gb_clf.fit(X_train, y_train) #fit the grid search object
print(gs_gb_clf.best_score_)
print(gs_gb_clf.best_params_)
best_parameters_gs = gs_gb_clf.best_params_ #get the best parameters based on 10x CV grid search
```
### 6. Initialise a Gradient Boosting Classifier
```
gbClassifier_best = ensemble.GradientBoostingClassifier(**best_parameters_gs, random_state=123) #intialise GB classifier with best set of parameters
```
### 7. Model Evaluation on Train data
```
#perform 10 fold cross validation and plot the CM
CV_predicted = cross_val_predict(gbClassifier_best, X_train, y_train, cv=10) #CV predicted values (training data)
CV_score = cross_val_score(gbClassifier_best, X_train, y_train, cv=10) #CV model score (training data)
print("Cross validation Score on train data: ",CV_score.mean())
print("\n")
print("Confusion matrix on CV predictions (train data)")
print(metrics.confusion_matrix(y_train, CV_predicted)) # confusion matrix on CV predictions (train data)
print("\n")
print("Classification report CV predictions (train data)")
print(metrics.classification_report(y_train, CV_predicted, target_names=['Poisonous', 'Edigble'])) # classification report CV predictions (train data)
```
### 9. Model Evaluation on Test data
```
gbClassifier_best_mdl= gbClassifier_best.fit(X_train, y_train) #fit the best GB classifier with training data
y_predicted = gbClassifier_best_mdl.predict(X_test) #Predict the outcomes with best GB classifier for test data
mdl_score = gbClassifier_best_mdl.score(X_test,y_test) #model score (test data)
print ("Model Score on test data:",mdl_score)
print("\n")
print("Confusion matrix (test data)")
print(metrics.confusion_matrix(y_test, y_predicted)) #confusion matrix (test data)
print("\n")
print("Classification report (test data)")
print(metrics.classification_report(y_test, y_predicted, target_names=['Poisonous', 'Edigble'])) # classification report (test data)
```
| github_jupyter |
# Facial Expression Recognition Project
## Library Installations and Imports
```
!pip install -U -q PyDrive
!apt-get -qq install -y graphviz && pip install -q pydot
!pip install -q keras
from google.colab import files
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import pydot
import tensorflow as tf
from tensorflow.python.client import device_lib
from keras.models import Sequential
from keras.layers import Conv2D, LocallyConnected2D, MaxPooling2D, Dense
from keras.layers import Activation, Dropout, Flatten
from keras.callbacks import EarlyStopping
from keras.utils import plot_model, to_categorical
from keras import backend as K
```
### Confirm Tensorflow and GPU Support
```
K.tensorflow_backend._get_available_gpus()
device_lib.list_local_devices()
tf.test.gpu_device_name()
```
## Helper Functions
```
def uploadFiles():
uploaded = files.upload()
for fn in uploaded.keys():
print('User uploaded file "{name}" with length {length} bytes'.format(
name=fn, length=len(uploaded[fn])))
filenames = list(uploaded.keys())
for f in filenames:
data = str(uploaded[f], 'utf-8')
file = open(f, 'w')
file.write(data)
file.close()
def pullImage(frame, index: int):
"""
Takes in a pandas data frame object and an index and returns the 48 x 48 pixel
matrix as well as the label for the type of emotion.
"""
img = frame.loc[index]['pixels'].split(' ')
img = np.array([np.int(i) for i in img])
img.resize(48,48)
label = np.uint8(frame.loc[index]['emotion'])
return img, label
def splitImage_Labels(frame):
"""
Takes in a pandas data frame object filled with pixel field and label field
and returns two numpy arrays; one for images and one for labels.
"""
labels = np.empty(len(frame))
images = np.empty((len(frame), 48, 48, 1)) # using channel last notation.
for i in range(len(frame)):
img, lbl = pullImage(frame, i)
img = np.reshape(img, (48,48,1))
images[i], labels[i] = img, lbl
return images.astype(np.uint8), to_categorical(labels, 7).astype(np.uint8)
```
## Import FER2013 Dataset and Other Files
```
# Authenticate and create the PyDrive client.
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
fer2013 = drive.CreateFile({'id':'1Xdlvej7eXaVcfCf3CsQ1LcSFAiNx_63c'})
fer2013.GetContentFile('fer2013file.csv')
```
Save file as a pandas dataframe.
```
df = pd.read_csv('fer2013file.csv')
```
## Parse Data
Each image is a 48 x 48 grayscale photo.
The contents of pixel string are space-separated pixel values in row major order.
Emotional assignment convention:
* 0 = Angry
* 1 = Disgust
* 2 = Fear
* 3 = Happy
* 4 = Sad
* 5 = Surprise
* 6 = Neutral
```
df_Training = df[df.Usage == 'Training']
df_Testing = df[df.Usage == 'PrivateTest'].reset_index(drop = True)
img_train, lbl_train = splitImage_Labels(df_Training)
img_test, lbl_test = splitImage_Labels(df_Testing)
print('Type and Shape of Image Datasets: ' + '\n\tTraining: ' + '\t' +
str(type(img_train[0][0][0][0])) + '\t' + str(img_train.shape) +
'\n\tTesting: ' + '\t' + str(type(img_train[0][0][0][0])) + '\t' +
str(img_test.shape))
print('Type and Shape of Image Datasets: ' + '\n\tTraining: ' + '\t' +
str(type(lbl_train[0][0])) + '\t' + str(lbl_train.shape) +
'\n\tTesting: ' + '\t' + str(type(lbl_train[0][0])) + '\t' +
str(lbl_test.shape))
```
### Save Data to .npy Files
```
#np.save('img_train.npy', img_train)
#np.save('lbl_train.npy', lbl_train)
#np.save('img_test.npy', img_test)
#np.save('lbl_test.npy', img_test)
```
### Verify Image Import
```
plt.imshow(np.reshape(img_train[0], (48,48)))
plt.title('Training Image 1 (with label ' + str(lbl_train[0]) + ')')
plt.imshow(np.reshape(img_test[0], (48,48)))
plt.title('Training Image 1 (with label ' + str(lbl_test[0]) + ')')
```
## Build Convolutional Neural Network Model
```
model = Sequential()
```
### Phase 1
- Convolutional and Max Pooling Phase.
- **Kernal Size: 5x5**
- **Ouput Filters: 40**
- **Stride: 1 (default)**
- **Active Padding**
```
outputFilters = 40
kernelSize = 5
model.add(Conv2D(outputFilters, kernelSize, padding='same', activation='relu',
input_shape=img_train[0].shape))
model.add(Conv2D(outputFilters, kernelSize, padding='same', activation='relu'))
model.add(Conv2D(outputFilters, kernelSize, padding='same', activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
```
### Phase 2
- Convolutional and Max Pooling Phase.
- **Kernal Size: 5x5**
- **Ouput Filters: 60**
- **Stride: 1 (default)**
- **Active Padding**
```
outputFilters = 60
kernelSize = 5
model.add(Conv2D(outputFilters, kernelSize, padding='same', activation='relu'))
model.add(Conv2D(outputFilters, kernelSize, padding='same', activation='relu'))
model.add(Conv2D(outputFilters, kernelSize, padding='same', activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
```
### Phase 3
- Locally-Connected Convlutional Filtering Phase.
- The locally-connected layer works similarly to the traditional 2D convolutional layer, except that weights are unshared, that is, a different set of filters is applied at each different patch of the input.
- **Ouput Filters: 100**
- **Kernal Size: 3x3**
- **Stride: 1 (default)**
- **Non-Active Padding**
```
outputFilters = 100
kernelSize = 3
model.add(LocallyConnected2D(outputFilters, kernelSize, padding='valid', activation='relu'))
model.add(LocallyConnected2D(outputFilters, kernelSize, padding='valid', activation='relu'))
```
### Dense Layers
```
layerSize = 64
model.add(Flatten())
model.add(Dense(layerSize, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(layerSize, activation='relu'))
model.add(Dense(7, activation='softmax'))
model.summary()
```
## Compile, Train, and Evaluate the Model
```
batchSize = 200
trainingEpochs = 50
model.compile(optimizer='adam', loss='categorical_crossentropy',
metrics=['accuracy'])
early_stopping = EarlyStopping(monitor='val_loss', patience=5, verbose=1)
trainingHistory = model.fit(img_train, lbl_train, batch_size=batchSize,
epochs=trainingEpochs,
validation_split=0.3,
callbacks=[early_stopping],
shuffle=True,)
trainingAccuracy = trainingHistory.history['acc']
validationAccuracy = trainingHistory.history['val_acc']
print("Done Training: ")
print('Final Training Accuracy: ', trainingAccuracy[-1])
print('Final Validation Accuracy: ', validationAccuracy[-1])
print('Overfit Ratio: ', validationAccuracy[-1]/trainingAccuracy[-1])
metrics = model.evaluate(img_test, lbl_test, batch_size=batchSize, verbose=1)
print('Evaluation Loss: ', metrics[0])
print('Evaluation Accuracy: ', metrics[1])
```
## Predictions
```
def predict(mdl, img):
emotions = ['Anger', 'Disgust', 'Fear', 'Happy', 'Sad', 'Surprise', 'Neutral']
im = np.zeros((1, 48, 48, 1))
im[0] = img
p = mdl.predict(im)
i = np.argmax(p)
plt.imshow(np.reshape(img, (48,48)))
plt.title('Test Image (Predicted Label: ' + emotions[i] + ')')
return p
predict(model, img_test[12])
```
## Intermediate Layer Output
```
layer0 = K.function([model.layers[0].input, K.learning_phase()],
[model.layers[0].output])
layer1 = K.function([model.layers[0].input, K.learning_phase()],
[model.layers[1].output])
layer3 = K.function([model.layers[0].input, K.learning_phase()],
[model.layers[3].output])
layer7 = K.function([model.layers[0].input, K.learning_phase()],
[model.layers[7].output])
```
- Image being used for this example:
```
im = np.zeros((1, 48, 48, 1))
im[0] = img_test[21]
plt.imshow(np.reshape(img_test[21], (48,48)))
```
### Layer 0 (after first convolution)
```
layer_output = layer0([im, 0])[0]
ALL = layer_output[0,:,:,:]
ALL = np.reshape(ALL, (48*5, 48 * 8), order='F')
plt.imshow(ALL)
plt.imshow(layer_output[0, :, :,0])
plt.imshow(layer_output[0, :, :,1])
plt.imshow(layer_output[0, :, :,6])
plt.imshow(layer_output[0, :, :,7])
plt.imshow(layer_output[0, :, :,8])
plt.imshow(layer_output[0, :, :,9])
plt.imshow(layer_output[0, :, :,10])
plt.imshow(layer_output[0, :, :,11])
```
### Layer 1 (after second convolution)
```
layer_output = layer1([im, 0])[0]
ALL = layer_output[0,:,:,:]
ALL = np.reshape(ALL, (48*5, 48 * 8), order='F')
plt.imshow(ALL)
plt.imshow(layer_output[0, :, :,14])
```
### Layer 3 (after first pooling)
```
layer_output = layer3([im, 0])[0]
ALL = layer_output[0,:,:,:]
ALL = np.reshape(ALL, (24*5, 24 * 8), order='F')
plt.imshow(ALL)
plt.imshow(layer_output[0, :, :,2])
plt.imshow(layer_output[0, :, :,3])
```
### Layer 7 (after second pooling)
```
layer_output = layer7([im, 0])[0]
ALL = layer_output[0,:,:,:]
ALL = np.reshape(ALL, (12*6, 12 * 10), order='F')
plt.imshow(ALL)
```
## Look at the Weights
```
weights = model.get_weights()
```
### Layer 0
```
w0 = weights[0]
w0 = w0[:,:,0,:]
ALL = np.reshape(w0, (5*5, 5 * 8), order='F')
plt.imshow(ALL)
```
| github_jupyter |
```
%pylab inline
%pdb
import pandas as pd
from datetime import datetime as dt
import os
import glob
import statsmodels.api as sm
fname = "./data/date-hour-soo-dest-2015.csv"
bart_df = pd.read_csv(fname, names = ["Date", "Hour", "Origin", "Destination", "Count"],
parse_dates = ["Date"], index_col = "Date" )
#hour_df = bart_df.groupby("Hour")
#hour_df["Count"].mean().plot(kind="bar")
#piv = bart_df.pivot_table(index=bart_df.index.dayofweek, values="Count", aggfunc=np.sum, columns="Hour")
#piv/sum(piv)
#resamp = bart_df.resample("d").mean()
#resamp.dropna(how="any", inplace=True)
#resamp = resamp["Count"]
#resamp = resamp.diff().dropna()
#sm.graphics.tsa.plot_acf(resamp, lags=30)
#resamp
#pd.tools.plotting.autocorrelation_plot(resamp["Count"],)
#resamp
bart_df["weekday"].values[-20:-10]
piv.index = ["m", "t", "w", "tr", "f","s","su"]
piv = piv[piv["Hour"] < 10]
(piv.T/sum(piv.T)).plot(kind="bar")
t = bart_df.apply(lambda row: row["Hour"]+1, axis = 1 )
t.head()
bart_df["weekday"] = bart_df.index.dayofweek
weekday_filter = bart_df["weekday"] < 5
hour_filter = bart_df["Hour"] < 12
bart_df = bart_df[weekday_filter & hour_filter & (bart_df["Count"] > 5)]
#mon = bart_df.resample("M", how=np.mean)
weekdays = ["Mon", "Tues", "Wed", "Thurs", "Fri"]
bart_df["weekday"].unique = weekdays
mon = bart_df.pivot_table(aggfunc=np.sum, index="Hour", columns=bart_df.index.month, values="Count")
mon = mon.dropna(axis=0, how="any")
#mon = mon/sum(mon)
#mon = bart_df.groupby("Origin")
#mon["Count"].mean().plot(kind = "bar")
mon.boxplot()
csv_files = glob.glob("./data/*.csv")
my_list = []
for f_name in csv_files:
with open(f_name, 'r') as f:
f.read()
my_list.append()
bart_df = bart_df[(bart_df["Count"] > 2) & (bart_df["Hour"] > 6) & (bart_df["Hour"] < 10)]
bart_df["DateTime"] = bart_df.apply(lambda row: dt.strptime(row["Date"], "%Y-%m-%d") , axis=1)
bart_df["DayOfWeek"] = bart_df.apply(lambda row: datetime.date.weekday(row["DateTime"]), axis=1)
daily_df = bart_df[(bart_df["DayOfWeek"] < 5)].groupby("Date")
hourly_df = bart_df[(bart_df["Hour"] > 6) & (bart_df["Hour"] < 10)
& (bart_df["DayOfWeek"]) > 0 & (bart_df["DayOfWeek"] < 6)].groupby("Hour")
bart_df_pivot = bart_df.head(10000).pivot_table(values="Count", index="Origin", columns="Destination")
bart_df[bart_df["Count"] > 50].hist(column="Count", by="DayOfWeek", bins=30)
a = sm.OLS(bart_df.head(10000)["DayOfWeek"], bart_df.head(10000)["Count"])
res = a.fit()
res.summary()
plot(bart_df.head(10000)["DayOfWeek"], bart_df.head(10000)["Count"])
bart_df[0:1000][["DateTime","Count"]].set_index("DateTime").rolling(window=10).mean()
rm.plot()
```
| github_jupyter |
### Import libraries and read data
```
from __future__ import division
import pandas as pd
import numpy as np
from numpy import argmax
from scipy import constants
import random
import os
import sys
import re
import pdb
import glob
#import suftware
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import OneHotEncoder
from sklearn.model_selection import train_test_split
import keras
from keras.models import Sequential, Model
from keras.layers import Dense, Dropout, Activation, Input, Lambda, Concatenate
from keras.optimizers import Adam, SGD
from keras import metrics
from keras import regularizers
from keras import callbacks
import keras.backend as K
import tensorflow as tf
from tensorflow.python.framework import ops
from tensorflow.python.ops import math_ops
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.patches as patches
from matplotlib import cm
from matplotlib.colors import ListedColormap, LinearSegmentedColormap
%matplotlib inline
plt.ion()
input_data_ordered_raw = pd.read_csv('psi_9nt.csv')
#input_data_ordered_raw = pd.read_csv('ratios_9nt_ss_locus.txt',delim_whitespace=True)
input_data_ordered_raw.head()
len(input_data_ordered_raw)
```
## Remove nan rows from dataframe
```
input_data_ordered_raw = input_data_ordered_raw[['seq','brca2_9nt']].dropna()
# so I don't have to use pseudocounts
input_data_ordered_raw = input_data_ordered_raw[input_data_ordered_raw['brca2_9nt']>0]
#sequences_9nt = input_data_ordered_raw['seq'].values
sequences = input_data_ordered_raw['seq'].values
val = np.log10(input_data_ordered_raw['brca2_9nt'].values)
#val = input_data_ordered_raw['smn1_9nt']
len(input_data_ordered_raw)
# sequences = []
# # delete characters G and U at positions 3 and 4 in the sequences
# for _ in range(len(sequences_9nt)):
# sequences.append(sequences_9nt[_][0:3]+sequences_9nt[_][5:])
# sequences = np.array(sequences)
#val_norm = (val-min(val))/(max(val)-min(val))
val_norm = val
len(sequences)
```
## Split the data into test and train
```
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(sequences,val_norm,test_size=0.2,random_state=0)
#x_train, x_test, y_train, y_test = train_test_split(sequences,val,test_size=0.2)
# min_y_train = min(y_train)
# max_y_train = max(y_train)
# min_y_test = min(y_test)
# max_y_test = max(y_test)
# y_train = (y_train-min(y_train))/(max(y_train)-min(y_train))
# y_test = (y_test-min(y_test))/(max(y_test)-min(y_test))
x_train[0]
plt.hist(y_train,bins=50,color='r',density=True,alpha=0.5)
plt.hist(y_test,bins=50,color='b',density=True,alpha=0.5)
plt.show()
# le = LabelEncoder().fit(bases).transform(bases).reshape(len(bases), 1)
# onehot_encoder = OneHotEncoder(sparse = False)
# onehot_encoder.fit(le)
# # Fit a label encoder and a onehot encoder
# bases = ["A","C","G","U"]
# label_encoder = LabelEncoder()
# label_encoder.fit(bases)
# tmp = label_encoder.transform(bases)
# tmp = tmp.reshape(len(tmp), 1)
# onehot_encoder = OneHotEncoder(sparse = False)
# onehot_encoder.fit(tmp)
# # Encode sequence into onehot
# def onehot_sequence(sequence, lab_encoder = label_encoder, one_encoder = onehot_encoder):
# """Sequence as a string"""
# tmp = lab_encoder.transform(list(sequence))
# tmp = tmp.reshape(len(tmp),1)
# tmp = one_encoder.transform(tmp)
# return tmp
# Fit a label encoder and a onehot encoder
bases = ["A","C","G","U"]
label_encoder = LabelEncoder()
label_encoder.fit(bases)
tmp = label_encoder.transform(bases)
tmp = tmp.reshape(len(tmp), 1)
onehot_encoder = OneHotEncoder(sparse = False)
onehot_encoder.fit(tmp)
# Encode sequence into onehot
def onehot_sequence(sequence, lab_encoder = label_encoder, one_encoder = onehot_encoder):
"""Sequence as a string"""
tmp = lab_encoder.transform(list(sequence))
tmp = tmp.reshape(len(tmp),1)
tmp = one_encoder.transform(tmp)
return tmp
print(x_train[2])
onehot_sequence(x_train[2]).shape
```
### Form input from sequences for all-pairs model
```
# nbr_dinucleotides = ['AA', 'AC', 'AG', 'AU',
# 'CA', 'CC', 'CG', 'CU',
# 'GA', 'GC', 'GG', 'GU',
# 'UA', 'UC', 'UG', 'UU']
# # one-hot encode di-nucleotide training set
# dinuc_seqs_OHE = []
# for _ in range(len(X_train)):
# # take current raw training sequence
# raw_sequence = X_train[_]
# # split it into all nucleotide pairs
# all_nucl_pairs = []
# for i in range(len(raw_sequence)):
# for j in range(i+1,len(raw_sequence)):
# all_nucl_pairs.append(raw_sequence[i]+raw_sequence[j])
# # get indices of where pairs occur so that these indices could be used to one-hot encode.
# list_of_nbr_indices = [nbr_dinucleotides.index(dn) for dn in all_nucl_pairs]
# # do One-hot encoding. Every time a pair from list 'nbr_dinucleotides'
# # appears at a position, put 1 there, otherwise zeros.
# tmp_seq = np.array(list_of_nbr_indices)
# OHE_dinucl_seq = np.zeros((tmp_seq.size, len(nbr_dinucleotides)))
# OHE_dinucl_seq[np.arange(tmp_seq.size),tmp_seq] = 1
# dinuc_seqs_OHE.append(OHE_dinucl_seq)
# dinuc_seqs_OHE_test = []
# for _ in range(len(X_test)):
# # take current raw test sequence
# raw_sequence = X_test[_]
# # split it into all nucleotide pairs
# all_nucl_pairs = []
# for i in range(len(raw_sequence)):
# for j in range(i+1,len(raw_sequence)):
# all_nucl_pairs.append(raw_sequence[i]+raw_sequence[j])
# # get indices of where pairs occur so that these indices could be used to one-hot encode.
# list_of_nbr_indices = [nbr_dinucleotides.index(dn) for dn in all_nucl_pairs]
# # do One-hot encoding. Every time a pair from list 'nbr_dinucleotides'
# # appears at a position, put 1 there, otherwise zeros.
# tmp_seq = np.array(list_of_nbr_indices)
# OHE_dinucl_seq = np.zeros((tmp_seq.size, len(nbr_dinucleotides)))
# OHE_dinucl_seq[np.arange(tmp_seq.size),tmp_seq] = 1
# dinuc_seqs_OHE_test.append(OHE_dinucl_seq)
import time
start_time = time.time()
input_seqs_ohe = []
for _ in range(len(x_train)):
if _%5000==0:
print(_)
input_seqs_ohe.append(onehot_sequence(x_train[_]))
print("--- %s seconds ---" % (time.time() - start_time))
test_seqs_ohe = []
for _ in range(len(x_test)):
test_seqs_ohe.append(onehot_sequence(x_test[_]))
np.shape(input_seqs_ohe)
type(input_seqs_ohe[0])
type(input_seqs_ohe)
input_sequences_tensor2 = np.array(input_seqs_ohe)
test_input_sequences_tensor2 = np.array(test_seqs_ohe)
print(input_sequences_tensor2.shape)
print(test_input_sequences_tensor2.shape)
labels = np.array(y_train)
test_labels = np.array(y_test)
y_train = np.array(y_train).reshape(y_train.shape[0],1)
y_train.shape
x_train_tensor = input_seqs_ohe.copy()
x_test_tensor = test_seqs_ohe.copy()
input_sequences_tensor2 = []
test_input_sequences_tensor2 = []
for _ in range(len(x_train_tensor)):
if _%5000==0:
print(_)
temp = x_train_tensor[_].ravel()
input_sequences_tensor2.append(temp)
for _ in range(len(x_test_tensor)):
temp = x_test_tensor[_].ravel()
test_input_sequences_tensor2.append(temp)
input_sequences_tensor2 = np.array(input_sequences_tensor2)
test_input_sequences_tensor2 = np.array(test_input_sequences_tensor2)
input_sequences_tensor2.shape, labels.shape
```
## Custom error metric
```
# coefficient of determination (R^2) for regression
def r_square(y_true, y_pred):
from keras import backend as K
SS_res = K.sum(K.square(y_true - y_pred))
SS_tot = K.sum(K.square(y_true - K.mean(y_true)))
return (1 - SS_res/(SS_tot + K.epsilon()))
# import keras.backend as K
# import numpy as np
# def gaussian_nll(ytrue, ypreds):
# """
# TF implmementation of multivariate Gaussian negative loglikelihood loss function.
# This implementation implies diagonal covariance matrix.
# Parameters
# ----------
# ytrue: tf.tensor of shape [n_samples, n_dims]
# ground truth values
# ypreds: tf.tensor of shape [n_samples, n_dims*2]
# predicted mu and logsigma values (e.g. by your neural network)
# Returns
# -------
# neg_log_likelihood: float
# negative loglikelihood
# """
# # number of outputs
# #n_dims = int(int(ypreds.shape[1])/2)
# # output 1 coming out of observable node respenting mu
# mu = ypreds
# logsigma = model.layers[4].sigma
# print(logsigma)
# #squared_error_term = 0.5*K.sum(K.square((ytrue-mu)/K.exp(logsigma)),axis=1)
# negative_log_likelihood = 0.5*K.sum(K.square((ytrue-mu)/K.exp(logsigma))+logsigma,axis=1)
# #negative_log_likelihood = squared_error_term+logsigma
# #negative_log_likelihood = squared_error_term
# #return K.sum(negative_log_likelihood)
# return negative_log_likelihood
```
## Custom Loss
```
import keras.backend as K
import tensorflow as tf
#K.clear_session()
from keras.models import Sequential
from keras.layers import Dense, Activation, Flatten, Dropout
from keras import regularizers
from keras.optimizers import SGD, Adam, RMSprop
from keras.constraints import maxnorm, nonneg
from mavenn.src.utils import get_example_dataset
from sklearn.model_selection import train_test_split
from mavenn.src.utils import _generate_nbr_features_from_sequences
from mavenn.src.utils import _generate_all_pair_features_from_sequences
from mavenn.src.utils import onehot_encode_array
X, y = get_example_dataset('MPSA')
x_train, x_test, y_train, y_test = train_test_split(X, y)
x_train_nbr_OHE = _generate_all_pair_features_from_sequences(x_train,'rna')
x_test_nbr_OHE = _generate_all_pair_features_from_sequences(x_test,'rna')
# x_train_nbr_OHE = onehot_encode_array(x_train,['A','C','G','U'])
# x_test_nbr_OHE = onehot_encode_array(x_test,['A','C','G','U'])
class CustomDense(Dense):
def __init__(self, units=1,**kwargs):
self.units = units
super(CustomDense,self).__init__(units,**kwargs)
def build(self, input_shape):
self.logsigma = self.add_weight(name='logsigma',
shape=(1, 1),
initializer="random_normal",
trainable=True,
)
def call(self, inputs):
# compute per-datum ll here
# this is yhat
yhat = inputs[:,0:1]
# these are the labels
ytrue = inputs[:,1:]
# Gaussian likelihood
#ll =0.5*K.sum(K.square((ytrue-yhat)/K.exp(self.logsigma))+self.logsigma,axis=1)
# Cauchy likelihood
ll = K.sum(K.log(K.square((ytrue-yhat))+K.square(K.exp(self.logsigma)))-self.logsigma,axis=1)
return ll
def custom_loss(y_true, y_pred):
#return K.sum(y_pred)
return y_pred
number_input_layer_nodes = len(x_train_nbr_OHE[0]) +1
inputTensor = Input((number_input_layer_nodes,),name='Sequence')
sequence_input = Lambda(lambda x: x[:,0:len(x_train_nbr_OHE[0])], output_shape=((len(x_train_nbr_OHE[0]),)))(inputTensor)
labels_input = Lambda(lambda x: x[:,len(x_train_nbr_OHE[0]):len(x_train_nbr_OHE[0])+1], output_shape=((1,)), trainable=False)(inputTensor)
phi = Dense(1)(sequence_input)
intermediateTensor = Dense(50,activation='sigmoid',kernel_constraint=nonneg())(phi)
yhat = Dense(1,kernel_constraint=nonneg())(intermediateTensor)
concatenateLayer = Concatenate()([yhat,labels_input])
outputTensor = CustomDense()(concatenateLayer)
#create the model:
model = Model(inputTensor,outputTensor)
model.summary()
model.layers
```
### Define model
```
# number_input_layer_nodes = len(x_train_nbr_OHE[0])
# inputTensor = Input((number_input_layer_nodes,),name='Sequence')
# phi = Dense(1)(inputTensor)
# intermediateTensor = Dense(50,activation='sigmoid',kernel_constraint=nonneg())(phi)
# yhat = Dense(1,kernel_constraint=nonneg())(intermediateTensor)
# outputTensor = CustomDense(ytrue = K.constant(y_train.reshape(-1,1)))(yhat)
# #create the model:
# model = Model(inputTensor,outputTensor)
model.layers[7].get_weights()
model.summary()
#model.compile(loss='mean_squared_error',optimizer=sgd, metrics=['mean_absolute_error'])
#model.compile(loss='mean_squared_error',optimizer=Adam(lr=0.005), metrics=['mean_absolute_error'])
model.compile(loss=custom_loss,optimizer=Adam(lr=0.005))
train_sequences = []
test_sequences = []
for _ in range(len(x_train_nbr_OHE)):
temp = x_train_nbr_OHE[_].ravel()
temp = np.append(temp,y_train[_])
train_sequences.append(temp)
for _ in range(len(x_test_nbr_OHE)):
temp = x_test_nbr_OHE[_].ravel()
temp = np.append(temp,y_test[_])
test_sequences.append(temp)
train_sequences = np.array(train_sequences)
test_sequences = np.array(test_sequences)
#history = model.fit(x_train_flat, y_train, validation_split=0.2, epochs=25) # starts training
#history = model.fit(input_sequences_tensor2, labels, validation_split=0.2, epochs=25,verbose=1) # starts training
history = model.fit(train_sequences, y_train, validation_split=0.2, epochs=100,verbose=1) # starts training
plt.figure()
plt.plot(history.history['loss'],color='blue')
plt.plot(history.history['val_loss'],color='orange')
plt.title('Model loss',fontsize=12)
plt.ylabel('loss',fontsize=12)
plt.xlabel('epoch',fontsize=12)
plt.legend(['train', 'validation'])
#plt.savefig('Write_up/model_loss.png')
plt.show()
model.layers[3].output
#model.layers[1].output
# Note this gives liklelihood
model_prediction = model.predict(test_sequences)
model_prediction_train = model.predict(train_sequences)
plt.scatter(model_prediction_train.ravel(),y_train,color='black',s=1)
#plt.scatter(model_prediction[:,0],y_test,color='blue',s=1)
plt.show()
np.exp(model.layers[7].get_weights()[0][0][0])
get_yhat_out = K.function([model.layers[1].input],[model.layers[4].output])
yhat = get_yhat_out([x_test_nbr_OHE])
model.layers
get_1st_layer_output = K.function([model.layers[1].input],[model.layers[2].output])
layer_output = get_1st_layer_output([x_test_nbr_OHE])
get_1st_layer_output_train = K.function([model.layers[0].input],[model.layers[1].output])
layer_output_train = get_1st_layer_output_train([x_train_nbr_OHE])
sigma = np.exp(model.layers[7].get_weights()[0][0][0])
plt.figure(figsize=(7,5))
plt.scatter(layer_output[0],y_test,s=1,color='gray',alpha=0.5,label='')
plt.scatter(layer_output[0],yhat[0].ravel(),color='black',s=1,label='$\hat{y}$')
plt.scatter(layer_output[0],yhat[0].ravel()+sigma,color='blue',s=1,alpha=0.25,label='$\hat{y} \pm \gamma$')
plt.scatter(layer_output[0],yhat[0].ravel()-sigma,color='blue',s=1,alpha=0.25)
plt.tick_params(labelsize=14)
leg = plt.legend(fontsize=14,loc='upper left')
for lh in leg.legendHandles:
lh.set_alpha(1)
lh.set_sizes([5.0])
plt.ylabel('Observations',fontsize=14)
plt.xlabel('Latent phenotype ($\phi$)',fontsize=14)
plt.title('Cauchy Likelihood',fontsize=14)
plt.show()
plt.scatter(yhat[0].ravel(),y_test,s=1,color='black')
np.corrcoef(yhat[0].ravel(),y_test)[0][1]**2
```
| github_jupyter |
# Keras Benchmark
##### Importing libraries
```
import numpy as np
import matplotlib.pyplot as plt
from glob import glob
from PIL import Image
from time import time
from sklearn.model_selection import train_test_split
from keras.layers import Conv2D, MaxPooling2D, ZeroPadding2D,\
Dropout, Flatten, Dense
from keras.callbacks import EarlyStopping
from keras.models import Sequential
from keras.optimizers import Adam
from keras.utils import to_categorical
import os
os.chdir('C:/Users/Nicolas/Documents/Data/Faces')
```
##### Loading all file names
```
files = glob('combined/*.jpg')
files = np.random.permutation(files)
files[:5]
```
##### Keeping all correctly formatted labels
```
np.unique([i[-34] for i in files], return_counts=True)
```
##### Removing problematic target names
```
faces = [i for i in files if (i[-34] in ('0', '1')) and len(i[-37:-35].strip('\\').strip('d')) == 2 ]
y = [i[-34] for i in files if (i[-34] in ('0', '1')) and len(i[-37:-35].strip('\\').strip('d')) > 1 ]
sex = ['men', 'women']
assert len(y) == len(faces), 'The X and Y are not of the same length!'
```
#### This is the shape width/height
```
dim = 60
```
#### Cropping function
```
def crop(img):
if img.shape[0]<img.shape[1]:
x = img.shape[0]
y = img.shape[1]
crop_img = img[: , int(y/2-x/2):int(y/2+x/2)]
else:
x = img.shape[1]
y = img.shape[0]
crop_img = img[int(y/2-x/2):int(y/2+x/2) , :]
return crop_img
```
##### Loading and cropping images
```
print('Scaling...', end='')
start = time()
x = []
num_to_load = len(faces)
for ix, file in enumerate(faces[:num_to_load]):
image = plt.imread(file, 'jpg')
image = Image.fromarray(image).resize((dim, dim)).convert('L')
image = crop(np.array(image))
x.append(image)
print(f'\rDone. {int(time() - start)} seconds')
y = y[:num_to_load]
```
##### Turning the pictures into arrays
```
x = np.array(x, dtype=np.float32).reshape(-1, 60, 60, 1)
y = np.array(y, dtype=np.float32)
labels = y.copy()
```
##### Turning the targets into a 2D matrix
```
y = to_categorical(y, 2)
y
x.shape, y.shape
assert x.ndim == 4, 'The input is the wrong shape!'
yy, xx = y.nbytes, x.nbytes
print(f'The size of X is {xx:,} bytes and the size of Y is {yy:,} bytes.')
files, faces = None, None
```
##### Displaying the pictures
```
fig = plt.figure(figsize=(12, 12))
for i in range(1, 5):
plt.subplot(1, 5, i)
rand = np.random.randint(0, x.shape[0])
ax = plt.imshow(x[rand][:, :, 0], cmap='gray')
plt.title('<{}>'.format(sex[int(labels[rand])].capitalize()))
yticks = plt.xticks([])
yticks = plt.yticks([])
```
##### Cross-validation
```
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=2e-1)
x, y = None, None
trainsize, testsize = x_train.shape[0], x_test.shape[0]
print(f'The size of the training set is {trainsize:,} and the '\
f'size of the test set is {testsize:,}.')
```
##### Scaling, casting the arrays
```
print('Scaling...', end='')
image_size = x_train.shape[1] * x_train.shape[1]
x_train = x_train.astype('float32') / 255
x_test = x_test.astype('float32') / 255
print('\rDone. ')
```
##### Building the ConvNet
```
x_train.shape
model = Sequential([
Conv2D(16*4, (3, 3), input_shape=(60, 60, 1), activation='relu'),
MaxPooling2D(),
Conv2D(32*4, (3, 3), activation='relu'),
MaxPooling2D(),
Conv2D(64*4, (3, 3), activation='relu'),
MaxPooling2D(),
Conv2D(128*4, (3, 3), activation='relu'),
MaxPooling2D(),
Flatten(),
Dense(1024, activation='relu'),
Dense(2048, activation='relu'),
Dense(2, activation='sigmoid')
])
model.summary()
```
##### Compiling the model
```
model.compile(optimizer=Adam(lr=0.001),
loss='binary_crossentropy',
metrics=['accuracy'])
```
##### Making callbacks
```
e_s = EarlyStopping(monitor='val_loss', patience=10)
```
##### Training the model
```
hist = model.fit(x_train, y_train,
epochs=50,
validation_data=[x_test, y_test],
batch_size=32,
callbacks=[e_s])
```
##### Evaluating the model
```
test_loss, test_acc = model.evaluate(x_test, y_test)
print(f'Test loss: {np.round(test_loss, 4)} — Test accuracy: {int(test_acc*100)}%.')
```
##### Getting examples of misclassified samples
```
yy = np.argmax(y_test, axis=1)
xx = np.argmax(model.predict(x_test), axis=1)
wrong_input = x_test[xx != yy]
wrong_target = np.argmax(y_test[xx != yy], axis=1)
assert wrong_input.shape[0] == wrong_target.shape[0], 'Input shape error.'
rand = np.random.randint(wrong_target.shape[0])
fig = plt.figure(figsize=(12, 12))
for i in range(1, 5):
plt.subplot(1, 5, i)
rand = np.random.randint(wrong_target.shape[0])
ax = plt.imshow(wrong_input[rand][:, :, 0], cmap='gray')
plt.title('<{}>'.format(sex[int(wrong_target[rand])].capitalize()))
yticks = plt.xticks([])
yticks = plt.yticks([])
```
The above women were classified as men. It's unclear why.
##### Evolution of the Metrics per Epoch
```
len_epochs = len(hist.history['loss'])
fig, ax = plt.subplots()
ax.set_facecolor('#DEDEDE')
ax.plot(np.arange(1, len_epochs + 1), hist.history['loss'],
color='orange', lw=3)
ax.plot(np.arange(1, len_epochs + 1), hist.history['accuracy'],
color='blue', lw=3)
ax.plot(np.arange(1, len_epochs + 1), hist.history['val_loss'],
color='orange', ls=':', lw=3)
ax.plot(np.arange(1, len_epochs + 1), hist.history['val_accuracy'],
color='blue', ls=':', lw=3)
plt.ylim(0, 1)
plt.title('Metrics per Epochs')
plt.legend(['Loss', 'Accuracy', 'Val Loss', 'Val Accuracy'])
plt.grid(True, alpha=.3)
plt.show()
```
So we reached 90% accuracy. We will do some more work on the Pytorch CNN and get the same accuracy, now that we know that it is possible.
| github_jupyter |
<!-- dom:TITLE: Demo - Some fast transforms -->
# Demo - Some fast transforms
<!-- dom:AUTHOR: Mikael Mortensen Email:mikaem@math.uio.no at Department of Mathematics, University of Oslo. -->
<!-- Author: -->
**Mikael Mortensen** (email: `mikaem@math.uio.no`), Department of Mathematics, University of Oslo.
Date: **May 27, 2021**
**Summary.** This demo will show how to compute fast forward transforms for the three
different Dirichlet bases that are implemented for Chebyshev
polynomials in Shenfun.
## Forward and backward transforms
A function $u(x)$ can be approximated in a finite global spectral
expansion $u_N(x)$ as
<!-- Equation labels as ordinary links -->
<div id="eq:expansion"></div>
$$
\label{eq:expansion} \tag{1}
u_N(x) = \sum_{k=0}^{N-1} \hat{u}_k \phi_k(x), \quad \forall \, x \, \in [-1, 1],
$$
where $\phi_k(x)$ are the basis functions and $\boldsymbol{\hat{u}} = \{\hat{u}_k\}_{k=0}^{N-1}$
are the expansion coefficients. The function $u_N(x)$ is continuous
on the interval domain $[-1, 1]$. The span of the basis functions
$V_N = \text{span} \{\phi_k\}_{k=0}^{N-1}$ represents a functionspace.
Associated with this functionspace is a set of quadrature points
$\{x_k\}_{k=0}^{N-1}$ that, along with quadrature weights $\{\omega_k\}_{k=0}^{N-1}$, can be used
for efficient integration. We can also evaluate the function $u_N(x)$ at
these quadrature points to get the sequence
$\boldsymbol{u} = \{u_N(x_k)\}_{k=0}^{N-1}$. If $\boldsymbol{\hat{u}}=\{\hat{u}_k\}_{k=0}^{N-1}$ are *known*,
then $\boldsymbol{u}$ can be evaluated directly from
Eq. ([1](#eq:expansion))
<!-- Equation labels as ordinary links -->
<div id="eq:expansionQ"></div>
$$
\label{eq:expansionQ} \tag{2}
u_N(x_j) = \sum_{k=0}^{N-1} \hat{u}_k \phi_k(x_j), \quad \forall \, j=0,1, \ldots, N-1.
$$
This would correspond to a *backward* transform according to
the Shenfun terminology. A direct evaluation of the backward
([2](#eq:expansionQ)) transform takes $\mathcal{O}(N^2)$
operations since it requires a double sum (over both $j$
and $k$). A *fast* transform is
a transform that can be computed in $\mathcal{O}(N \log N)$ operations.
This is what the Fast Fourier Transform (FFT) does. It computes a double
sum, like ([2](#eq:expansionQ)), in $\mathcal{O}(N \log N)$ operations.
The other way around, computing $\{\hat{u}_k\}_{k=0}^{N-1}$ from the
*known* $\{u_N(x_k)\}_{k=0}^{N-1}$ corresponds to a *forward* transform.
The forward transform is computed using a projection of $u$
into $V_N$, which is formulated as: find $u_N \in V_N$ such that
<!-- Equation labels as ordinary links -->
<div id="eq:projection"></div>
$$
\label{eq:projection} \tag{3}
(u_N-u, v)_{\omega^{\sigma}} = 0, \quad \forall \, v \in V_{N},
$$
where $(a, b)_{\omega^{\sigma}} = \int_{I} a b \omega^{\sigma} dx$ is the
inner product in $L^2_{\omega^{\sigma}}(I)$, and $\omega^{\sigma}(x)=(1-x^2)^{\sigma}$ is a weight function.
For Chebyshev polynomials the weight function is usually $\omega^{-1/2}=(1-x^2)^{-1/2}$.
Inserting for $u_N$ and $v=\phi_k$, we get
<!-- Equation labels as ordinary links -->
<div id="_auto1"></div>
$$
\begin{equation}
\sum_{j=0}^{N-1}(\phi_j, \phi_k)_{\omega^{\sigma}} \hat{u}_{j} = (u, \phi_k)_{\omega^{\sigma}},
\label{_auto1} \tag{4}
\end{equation}
$$
<!-- Equation labels as ordinary links -->
<div id="_auto2"></div>
$$
\begin{equation}
B \boldsymbol{\hat{u}} = \boldsymbol{\tilde{u}},
\label{_auto2} \tag{5}
\end{equation}
$$
<!-- Equation labels as ordinary links -->
<div id="_auto3"></div>
$$
\begin{equation}
\boldsymbol{\hat{u}} = B^{-1} \boldsymbol{\tilde{u}},
\label{_auto3} \tag{6}
\end{equation}
$$
where
$\boldsymbol{\tilde{u}} = \{(u, \phi_k)_{\omega^{\sigma}}\}_{k=0}^{N-1}$ and the mass matrix
$B = (b_{kj})_{k,j=0}^{N-1}$, with $b_{kj}=(\phi_j, \phi_k)_{\omega^{\sigma}}$.
Note that the forward transform requires both an inner product
$\boldsymbol{\tilde{u}}$ and a matrix inversion. By a *fast* forward transform
we mean a transform that can be computed in $\mathcal{O}(N \log N)$
operations. If $B$ is a diagonal or banded matrix, the matrix inversion costs $\mathcal{O}(N)$,
and the limiting factor is then the inner product. Like for the backward transform,
the inner product, computed with quadrature, is a double sum
$$
(u, \phi_k)_{\omega^{\sigma}} = \sum_{j=0}^{N-1} u(x_j) \phi_k(x_j) \omega_j, \quad \forall \, k = 0, 1, \ldots, N-1,
$$
where $\{\omega_j\}_{j=0}^{N-1}$ are the quadrature weights.
A naive implementation of the inner product
takes $\mathcal{O}(N^2)$ operations. However,
for Chebyshev polynomials we can compute the double loop with
fast $\mathcal{O}(N \log N)$ discrete sine or cosine transforms,
that are versions of the FFT. To see this, assume that the basis functions are $\phi_k(x) =T_k(x)$, where
$T_k(x)$ is the $k$'th Chebyshev polynomial of the first kind,
and the weight function is $\omega^{-1/2}$.
We then choose Gauss-Chebyshev points $x_j = \cos(\theta_j)$,
where $\theta_j=\pi (2j+1)/(2N)$, and the associated quadrature weights
that are constant $\omega_j = \pi/N$. The Chebyshev polynomials evaluated
on the quadrature points can now
alternatively be written as $T_k(x_j) = \cos(k \theta_j)$,
and the inner product becomes
$$
(u, T_k)_{\omega^{-1/2}} = \sum_{j=0}^{N-1} u(x_j) \cos(k \theta_j) \pi/N, \quad \forall \, k = 0, 1, \ldots, N-1.
$$
From the [FFTW documentation](http://www.fftw.org/fftw3_doc/1d-Real_002deven-DFTs-_0028DCTs_0029.html#g_t1d-Real_002deven-DFTs-_0028DCTs_0029)
we recognise this sum as half a DCT-II (the FFTW DCT-II has a factor
2 in front of the sum) of $\boldsymbol{u}\pi/N$. Hence, we can compute the inner product as
$$
(u, T_k)_{\omega^{-1/2}} = \frac{\pi}{2N} \text{dct}^{II}(\boldsymbol{u})_k, \quad k = 0, 1, \ldots, N-1.
$$
## Dirichlet bases
The basis function $T_k$ satisfies $T_k(\pm 1) = (\pm 1)^k$ at the
boundaries of the domain, and the space $S_N=\text{span}\{T_k\}_{k=0}^{N-1}$,
of dimension $N$,
is thus not associated with any specific set of boundary conditions.
A functionspace for homogeneous Dirichlet boundary conditions is
given as $V_N=\{v\in S_N | v(\pm 1)=0 \}$. Because of the two restrictions
the space has dimension $N-2$.
There are several different choices of basis functions
for $V_N$.
The most interesting we name $\phi_k^n$, for integer $n$, and
define them as
<!-- Equation labels as ordinary links -->
<div id="_auto4"></div>
$$
\begin{equation}
\phi^n_k = \omega T^{(n)}_{k+n} = (1-x^2) T^{(n)}_{k+n},
\label{_auto4} \tag{7}
\end{equation}
$$
where $T^{(n)}_{k+n}$ is the $n$'th derivative of $T_{k+n}$. We have
for any integer $n$ that $V_N=\text{span}\{\phi^n_k\}_{k=0}^{N-3}$, and an
expansion in any of these basis functions is
<!-- Equation labels as ordinary links -->
<div id="eq:uNgeneric"></div>
$$
\begin{equation}
\label{eq:uNgeneric} \tag{8}
u_N = \sum_{k=0}^{N-3} \hat{u}^n_k \phi^n_k.
\end{equation}
$$
We can find the sequence $\{\hat{u}^n_{k}\}_{k=0}^{N-3}$ for any $n$
using a projection into the space $V_N$. The projection is computed
by using Eq. ([8](#eq:uNgeneric)) and $v=\phi^n_k$ in
Eq. ([3](#eq:projection))
<!-- Equation labels as ordinary links -->
<div id="eq:projortho"></div>
$$
\begin{equation}
\label{eq:projortho} \tag{9}
\sum_{j=0}^{N-3} ( T^{(n)}_{j+n}, T^{(n)}_{k+n})_{\omega^{\sigma+2}} \hat{u}^{n}_j = (u, T^{(n)}_{k+n})_{\omega^{\sigma+1}}.
\end{equation}
$$
Now how can this projection be computed as efficiently as possible?
The Chebyshev polynomials and their derivatives are known to satisfy
the following orthogonality relation
<!-- Equation labels as ordinary links -->
<div id="eq:orthon"></div>
$$
\begin{equation}
\label{eq:orthon} \tag{10}
(T^{(n)}_j, T^{(n)}_k)_{\omega^{n-1/2}} = \alpha^{n}_k \delta_{kj}, \quad \text{for}\, n \ge 0,
\end{equation}
$$
where $\delta_{kj}$ is the Kronecker delta function and
<!-- Equation labels as ordinary links -->
<div id="_auto5"></div>
$$
\begin{equation}
\alpha^n_k = \frac{c_{k+n}\pi k (k+n-1)!}{2(k-n)!},
\label{_auto5} \tag{11}
\end{equation}
$$
where $c_0=2$ and $c_k=1$ for $k>0$. This can be used in
computing ([9](#eq:projortho)), because we just
need to choose the $\sigma$ that leads to a diagonal mass matrix.
For $n=(0, 1, 2)$ this will be $\sigma=-5/2, -3/2$ and $-1/2$,
respectively. So, choosing $\sigma=-5/2, -3/2$ and $-1/2$
for $n=0, 1$ and 2, respectively, will lead to a diagonal
mass matrix $( T^{(n)}_{j+n}, T^{(n)}_{k+n})_{\omega^{\sigma+2}}$.
Using these $\sigma$'s we can invert the diagonal mass matrices
in Eq. ([9](#eq:projortho)) to get
<!-- Equation labels as ordinary links -->
<div id="_auto6"></div>
$$
\begin{equation}
\hat{u}^n_k = \frac{1}{\alpha^n_{k+n}}(u, T^{(n)}_{k+n})_{\omega^{\sigma+1}}, \quad k=0, 1, \ldots, N-3, \text{ for } n \in (0, 1, 2).
\label{_auto6} \tag{12}
\end{equation}
$$
Using now quadrature, $1-x^2_i=\sin^2 \theta_i$ and the
fast transforms $(u, T_k)_{\omega^{-1/2}} = \pi/2/N \text{dct}^{II}(\boldsymbol{u})_k$
and $(u, U_k)_{\omega^{-1/2}} = \pi/2/N \text{dst}^{II}(\boldsymbol{u}/\sin \boldsymbol{\theta})_k$,
where $\boldsymbol{u}/\sin \boldsymbol{\theta}$ implies element-wise division,
we get
<!-- Equation labels as ordinary links -->
<div id="eq:fast1"></div>
$$
\begin{equation}
\hat{u}^0_k = \frac{1}{c_k N} \text{dct}^{II}(\boldsymbol{u}/\sin^2 \boldsymbol{\theta})_k, \quad k = 0, 1, \ldots, N-3, \label{eq:fast1} \tag{13}
\end{equation}
$$
<!-- Equation labels as ordinary links -->
<div id="eq:fast2"></div>
$$
\begin{equation}
\hat{u}^1_k = \frac{1}{(k+1)N}\text{dst}^{II}(\boldsymbol{u}/\sin \boldsymbol{\theta})_k, \quad k = 0, 1, \ldots, N-3, \label{eq:fast2} \tag{14}
\end{equation}
$$
<!-- Equation labels as ordinary links -->
<div id="eq:fast3"></div>
$$
\begin{equation}
\hat{u}^2_k = \frac{1}{2(k+2)}\left(\hat{u}^1_k - \hat{u}^1_{k+2} \right), \quad k=0, 1, \ldots, N-3. \label{eq:fast3} \tag{15}
\end{equation}
$$
The last one requires some work, using the identity
$\phi^2_k=(1-x^2)T''_{k+2}=0.5(k+2)(k+3)(U_k - (k+1)/(k+3)U_{k+2})$.
## Verification
To validate all the fast methods we compute the projection first regularly
using the Shenfun function [project](https://github.com/spectralDNS/shenfun/blob/master/shenfun/forms/project.py),
which is using $\sigma=-1/2$, and then the fast methods above. The two
projections should be the same, but they will not give identical results.
In general, the fast transforms above should be both faster and more
accurate, because they only take a discrete transform and merely a diagonal
mass matrix inversion.
We start the implementation by importing necessary modules from Shenfun
and mpi4py-fft
```
from shenfun import *
from mpi4py_fft import fftw
```
The three bases $\{\phi^n_k\}_{k=0}^{N-3}$ are implemented
with slightly different scaling in shenfun.
The first, with $n=0$, is obtained with no special scaling using
```
N = 20
D0 = FunctionSpace(N, 'C', bc=(0, 0), basis='Heinrichs')
```
The second basis is implemented in Shenfun as $\phi_k = \frac{2}{k+1}\phi^1_k$,
which can be simplified as
<!-- Equation labels as ordinary links -->
<div id="eq:shen"></div>
$$
\label{eq:shen} \tag{16}
\phi_k(x) = T_k-T_{k+2}, \quad k=0,1, \ldots, N-3,
$$
and implemented as
```
D1 = FunctionSpace(N, 'C', bc=(0, 0)) # this is the default basis
```
Because of the scaling the expansion coefficients for $\phi_k$ are
$\hat{u}^{\phi}_k=\frac{k+1}{2}\hat{u}^1_k$. Using ([14](#eq:fast2)) we get
$$
\hat{u}^{\phi}_k = \frac{1}{2N}\text{dst}^{II}(\boldsymbol{u}/\sin \boldsymbol{\theta})_k, \quad k = 0, 1, \ldots, N-3.
$$
The third basis is also scaled and implemented in Shenfun as $\psi_k = \frac{2}{(k+3)(k+2)}\phi^2_k$,
which can be simplified using Chebyshev polynomials of the second
kind $U_k$
<!-- Equation labels as ordinary links -->
<div id="eq:dirichletU"></div>
$$
\label{eq:dirichletU} \tag{17}
\psi_k(x) = U_k-\frac{k+1}{k+3}U_{k+2}, \quad k=0,1, \ldots, N-3.
$$
We get the basis using
```
D2 = FunctionSpace(N, 'C', bc=(0, 0), basis='DirichletU')
```
and the expansion coefficients are found as
$\hat{u}^{\psi}_k = \frac{(k+3)(k+2)}{2} \hat{u}^2_k$.
For verification of all the fast transforms we first create a vector
consisting of random expansion coefficients, and then transform
it backwards to physical space
```
f = Function(D0, buffer=np.random.random(N))
f[-2:] = 0
fb = f.backward().copy()
```
Next, we perform the regular projections into the three spaces
`D0`, `D1` and `D2`, using the inner product
in $L^2_{\omega^{-1/2}}$. Now `u0`, `u1` and `u2` will be the
three solution vectors
$\boldsymbol{\hat{u}}^{\varphi}$, $\boldsymbol{\hat{u}}^{\phi}$
and $\boldsymbol{\hat{u}}^{\psi}$, respectively.
```
u0 = project(fb, D0)
u1 = project(fb, D1)
u2 = project(fb, D2)
```
Now compute the fast transforms and assert that they are equal to `u0`, `u1` and `u2`
```
theta = np.pi*(2*np.arange(N)+1)/(2*N)
# Test for n=0
dct = fftw.dctn(fb.copy(), type=2)
ck = np.ones(N); ck[0] = 2
d0 = dct(fb/np.sin(theta)**2)/(ck*N)
assert np.linalg.norm(d0-u0) < 1e-8, np.linalg.norm(d0-f0)
# Test for n=1
dst = fftw.dstn(fb.copy(), type=2)
d1 = dst(fb/np.sin(theta))/(2*N)
assert np.linalg.norm(d1-u1) < 1e-8
# Test for n=2
ut = d1
k = np.arange(N)
d2 = Function(D2)
d2[:-2] = (k[:-2]+3)/2/(k[:-2]+1)*ut[:-2]
d2[:-2] = d2[:-2] - 0.5*ut[2:]
assert np.linalg.norm(d2-u2) < 1e-8
```
That's it! If you make it to here with no errors, then the three tests pass, and the fast transforms are equal to the slow ones, at least within given precision.
Let's try some timings
```
%timeit project(fb, D1)
%timeit dst(fb/np.sin(theta))/(2*N)
```
We can precompute the sine term, because it does not change
```
dd = np.sin(theta)*2*N
%timeit dst(fb/dd)
```
The other two transforms are approximately the same speed.
```
%timeit dct(fb/np.sin(theta)**2)/(ck*N)
```
<!-- ======= Bibliography ======= -->
| github_jupyter |
```
import torch
import torch.nn as nn
from torch.autograd import Variable
import numpy as np
class Model(nn.Module):
def __init__(self):
super(Model,self).__init__()
self.conv0 = nn.Conv2d(1, 16, kernel_size=3, padding=5)
self.conv1 = nn.Conv2d(16, 32, kernel_size=3)
def forward(self, x):
h = self.conv0(x)
h = self.conv1(h)
return h
model = Model()
import torch
import torch.nn as nn
from torch.autograd import Variable
import numpy as np
class SizeEstimator(object):
def __init__(self, model, input_size=(1,1,32,32), bits=32):
'''
Estimates the size of PyTorch models in memory
for a given input size
'''
self.model = model
self.input_size = input_size
self.bits = 32
return
def get_parameter_sizes(self):
'''Get sizes of all parameters in `model`'''
mods = list(model.modules())
for i in range(1,len(mods)):
m = mods[i]
p = list(m.parameters())
sizes = []
for j in range(len(p)):
sizes.append(np.array(p[j].size()))
self.param_sizes = sizes
return
def get_output_sizes(self):
'''Run sample input through each layer to get output sizes'''
input_ = Variable(torch.FloatTensor(*self.input_size), volatile=True)
mods = list(model.modules())
out_sizes = []
for i in range(1, len(mods)):
m = mods[i]
out = m(input_)
out_sizes.append(np.array(out.size()))
input_ = out
self.out_sizes = out_sizes
return
def calc_param_bits(self):
'''Calculate total number of bits to store `model` parameters'''
total_bits = 0
for i in range(len(self.param_sizes)):
s = self.param_sizes[i]
bits = np.prod(np.array(s))*self.bits
total_bits += bits
self.param_bits = total_bits
return
def calc_forward_backward_bits(self):
'''Calculate bits to store forward and backward pass'''
total_bits = 0
for i in range(len(self.out_sizes)):
s = self.out_sizes[i]
bits = np.prod(np.array(s))*self.bits
total_bits += bits
# multiply by 2 for both forward AND backward
self.forward_backward_bits = (total_bits*2)
return
def calc_input_bits(self):
'''Calculate bits to store input'''
self.input_bits = np.prod(np.array(self.input_size))*self.bits
return
def estimate_size(self):
'''Estimate model size in memory in megabytes and bits'''
self.get_parameter_sizes()
self.get_output_sizes()
self.calc_param_bits()
self.calc_forward_backward_bits()
self.calc_input_bits()
total = self.param_bits + self.forward_backward_bits + self.input_bits
total_megabytes = (total/8)/(1024**2)
return total_megabytes, total
se = SizeEstimator(model)
print(se.estimate_size())
se = SizeEstimator(model, input_size=(16,1,256,256))
print(se.estimate_size())
```
| github_jupyter |
<img src="module-01/ScDo-Bandeau_Lingua_Technologies.png" style="width: 100%;float:center;"/>
<h1 style="font-size:200%;text-align:center">Survol des applications de la science des données</h1>
<h1 style="font-size:200%;text-align:center">et de l’intelligence artificielle</h1>
<h2 style="text-align:center">par</h2>
<h2 style="text-align:center">Claude COULOMBE</h2>
<h3 style="text-align:center">scientifique des données</h3>
# Syllabus
Dans cet atelier de type « prise en main » entièrement en français, nous ferons un survol de la science des données (en anglais, data science), de l’analyse intelligente des données (en anglais, data analysis), l’exploitation des données massives (en anglais, big data) et des principes de base du forage des données (en anglais, data mining) avec le langage Python.
À partir de données réelles en utilisant des carnets web (Jupyter iPython Notebooks), nous utiliserons les outils Pandas (manipulation de données), matplotlib (visualisation) et statsmodels (statistique de base) avec quelques incursions au niveau des bibliothèques Numpy (calcul numérique), Scipy (calcul scientifique) et Scikit-learn (apprentissage statistique).
Des activités pratiques seront choisies selon des référents culturels autour de la justice, du droit et du monde juridique. L'objectif ici est de motiver et de susciter des idées et des projets d'application.
Nous conclurons avec les perspectives offertes par et l’apprentissage automatique (en anglais, machine learning) en préparation du module suivant.
## Prérequis
* Une connaissance de base langage Python est requise.
* IMPORTANT: Vous devez avoir installé un environnement scientifique Python <a href="https://www.anaconda.com/distribution/">Anaconda</a> fonctionnel.
<hr style="height:1px">
NOTE: Ce tutoriel s’inspire du livre <a href="http://shop.oreilly.com/product/0636920023784.do">
Python for Data Analysis</a> de Wes McKinney, des <a href="https://github.com/ogrisel">carnets web de Olivier Grisel</a> de l’INRIA et des données du <a href="https://www.kaggle.com/c/titanic">« Titanic challenge »</a> de Kaggle.
<hr style="height:1px">
# Plan
<ul>
<li>Syllabus</li>
<li>Prérequis</li>
<li>Plan</li>
<li>Contexte</li>
<li>Motivation - exemples d'applications</li>
<ul>
<li>Un maraîcher japonais trie ses concombres avec l'IA </li>
<li>Identifier des parasites sur des saumons d'aquaculture</li>
<li>Surveillance automatisée de la durabilité des pêches</li>
<li>Utilisation de drones pour planter des arbres</li>
<li>Un assistant intelligent pour diagnostiquer les maladies du manioc</li>
</ul>
<li>L'écosystème de la science des données et de l'IA</li>
<li>La science des données</li>
<li>Définition d'intelligence et d'intelligence artificielle</li>
<li>Qu’est-ce que l’apprentissage automatique?</li>
<ul>
<li>Définition formelle de l'apprentissage automatique</li>
<li>Les composantes d’un algorithme d’apprentissage</li>
<li>Types d'apprentissage automatique</li>
<li>Deux grands types de modèles statistiques</li>
</ul>
<li>Outils</li>
<ul>
<li>Pourquoi Python?</li>
<li>Écosystème Python?</li>
<li>Autres outils</li>
</ul>
<li>Processus de forage des données selon la norme CRISP-DM</li>
<li>Configuration du tutoriel</li>
<ul>
<li>Que sont les données?</li>
<li>Introduction à NumPy</li>
<li>Premier contact avec Matplotlib</li>
</ul>
<li>Motivation - Exemples d'applications</li>
<ul>
</ul>
</ul>
# Contexte
Dans les dernières années, on note un intérêt croissant pour l'exploitation « intelligente » des données. Cette évolution ou révolution s’appuie sur l’informatisation croissante des activités humaines conséquente du développement de la Toile, de l’abondance des données, de l’augmentation des capacités de calcul et de l’émergence de la science des données, de l’apprentissage automatique et de l’intelligence artificielle.
Ceci s’inscrit dans un courant de société plus global qui vise à prendre de meilleures décisions par l’exploitation des données car « <b>les données sont le nouveau pétrole </b>».
Prendre des décisions à partir des données et des faits améliore la production et la productivité des organisations. Les entreprises, plus particulièrement les géants de la Toile, le fameux GAFAM (Google, Amazon, Facebook, Apple et Microsoft) ont été les premières à prendre le virage des données massives et de l'intelligence artificielle (IA) pour devenir des incontournables dans ces domaines avec les avantages et inconvénients que cela peut représenter.
# Motivation - exemples d'applications
Les applications de l'intelligence artificielle (plus précisément de l'apprentissage profond) les plus mûres et les plus faciles à réaliser concernent la reconnaissance visuelle. Il existe de gros modèles pré-entraînés et on peut recueillir facilement des images spécifiques avec une caméra numérique (ou un simple téléphone intelligent). Là où c'est plus compliqué c'est d'interfacer la reconnaissance visuelle avec un équipement électromécanique ou robotique.
## Un maraîcher japonais, M. Makoto Koike, trie ses concombres avec l'IA
L'application de tri de concombres vise à remplacer une petite installation où un convoyeur transporte des concombres qui pour être triés manuellement en fonction de différents attributs: taille, couleur, forme (plus ou moins courbée). Les usages et les prix des légumes varient en fonction de ces caractéristiques.
</br>
<figure>
<img src="module-01/ScDo-Un_maraicher_japonais_trie_ses_concombres_avec_TensorFlow-1.png" width='600' />
<figcaption style="text-align:center">M. Koike avec ses parents sur la ferme familiale - Source: Google Blog</figcaption>
</figure>
Il fallait normalement compter 2 à 3 heures dans cette petite ferme japonaise pour trier environ 50 kg de concombres en neuf catégories (ou classes) différentes. Mais l'emploi l'IA a permis d’augmenter le rendement de la ferme de 140%.
Bref, il faut environ 1h45 min pour trier 50 Kg au lieu de 2h30.
* <a href="https://cloud.google.com/blog/products/gcp/how-a-japanese-cucumber-farmer-is-using-deep-learning-and-tensorflow">How a cucumber farmer is using Deep Learning and TensorFlow</a>
* <a href="https://opensource.com/article/17/9/tensorflow">TensorFlow brings machine learning to the masses</a>
```
from IPython.display import Audio,Image, YouTubeVideo
id='pSqcwSGDPqc'
YouTubeVideo(id=id,width=600,height=300,hl='fr',cc_lang_pref='fr',title="Source:welcome.ai")
from IPython.display import Audio,Image, YouTubeVideo
id='4HCE1P-m1l8'
YouTubeVideo(id=id,width=600,height=300,hl='fr',cc_lang_pref='fr',title="Source:welcome.ai")
```
### Données
* 7000 images de différentes types de concombres classées / annotées manuellement (ici 9 catégories ou classes). Cette opération a pris 3 mois.
* Les photos furent converties en images de 80 par 80 pixels
### Algorithme
* Apprentissage supervisé
* Un réseau de neurones convolutif à plusieurs couches (CNN)
* 2 à 3 jours de calculs pour entraîner le modèle sur un PC roulant Windows
* Résultat: 70 % d'exactitude (accuracy)
Initialement M. Koike est parti du code d'un tutoriel de Google (<a href="https://chromium.googlesource.com/external/github.com/tensorflow/tensorflow/+/r0.10/tensorflow/g3doc/tutorials/mnist/pros/index.md">Deep MNIST for experts</a>)
## Identifier des parasites sur des saumons d'aquaculture
L'infestation par les poux de mer, de petis parasites qui s'accrochent aux flancs des poissons, est l'un des principal problème sanitaire des bassins d'aquaculture de saumons.
Le contrôle exige la prise de mesure régulière qui demande de pêcher régulièrement un certain nombre de poissons (10 à 20 poissons par semaine) pour compter le nombre de parasites qui les affectent.
L'idée est de faire un contrôle grâce à la vision artificielle. Un obstacle important est la mise au point d'une caméra submersible capable de répondre aux besoins.
```
from IPython.display import Audio,Image, YouTubeVideo
id='YZ_qJ5JFD3I'
YouTubeVideo(id=id,start=700,end=750,width=600,height=300,hl='fr',cc_lang_pref='fr',iv_load_policy=1)
```
#### Données
* Images de saumons prises par des caméras sous-marines, annotées manuellement pour les différentes parties du poisson et la détection des poux de mer
#### Algorithme
* Apprentissage supervisé
* Un réseau de neurones convolutif à plusieurs couches (CNN)
Note: Autre application: estimation de la taille et du poids de la population de poissons.
## Surveillance automatisée de la durabilité des pêches
IA et pêche durable - En Nouvelle-Angleterrre, on utilise des caméras et une vision par ordinateur (grâce à ConvNets / CNN) sur les bateaux de pêche pourrait être un moyen rentable de s’assurer que les bateaux de pêche ne capturent pas plus de poissons que les quotas de capture autorisés tout en fournissant des données pour la gestion de la pêche.
Des caméras de surveillance collectent des images sur le déroulement de la pêche qui sont stockées sur des disques durs à haute capacité. L'objectif est de déterminer si les pêcheurs respectents les règlements de la pêche et leurs quotas. Cela permet d'assurer un contrôle systématique et continu sans avoir à embarquer un inspecteur à bord.
* Article <a href="https://www.pri.org/stories/2018-05-10/future-fishing-big-data-and-artificial-intelligence">The future of fishing is big data and artificial intelligence</a>
```
from IPython.display import Audio,Image, YouTubeVideo
id='1LuvZlwy75Y'
YouTubeVideo(id=id,start=0,width=600,height=300,hl='fr',cc_lang_pref='fr',iv_load_policy=1)
```
### Données
* Images des prises de poissons et des remises à la mer par des caméras de surveillance. Les images incluent des régles et gabarits pour annoter la taille des prises. Des données de géolocalisation sont ajoutées automatiquement(GPS). Au début, des images sont probablement annotées manuellement pour identifier les espèces et calibrer le système de mesure de la taille des prises. On doit aussi pouvoir estimer le poids, le tonnage pêché.
### Algorithme
* Apprentissage supervisé
* Un réseau de neurones convolutif à plusieurs couches (CNN) pour identifier les espèces
* Un réseau de neurones convolutif à plusieurs couches (CNN) pour identifier la taille
* L'application compte également le nombre de prises de chaque espèce, les prises rejetées et le lieu de la prise
## Utilisation de drones pour planter des arbres
Susan Graham présente BioCarbon Engineering à Ecosummit Berlin 2019. Basé à Oxford Grande-Bretagne, BioCarbon Engineering reconstruit les écosystèmes en tant que service destiné aux sociétés minières et pétrolières. Pour y arriver, BioCarbon Engineering utilise combine l'intelligence artificielle et l'usage de drones pour planter des arbres.
* Article dans Wired <a href="https://www.wired.com/brandlab/2015/07/re-planting-forest-one-drone-time/">Re-Planting a Forest, One Drone at a Time</a>
* Site de <a href="https://www.biocarbonengineering.com/">BioCarbon Engineering</a>
```
from IPython.display import Audio,Image, YouTubeVideo
id='YqXL7tYOCWc'
YouTubeVideo(id=id,start=87,width=600,height=300,hl='fr',cc_lang_pref='fr',iv_load_policy=1)
from IPython.display import Audio,Image, YouTubeVideo
id='hxMsIb8_bDE'
YouTubeVideo(id=id,start=0,width=600,height=300,hl='fr',cc_lang_pref='fr',iv_load_policy=1)
from IPython.display import Audio,Image, YouTubeVideo
id='-j_6092U-m4'
YouTubeVideo(id=id,start=0,width=600,height=300,hl='fr',cc_lang_pref='fr',iv_load_policy=1)
```
#### Données
* Images aériennes des terrains à replanter. Des données de géolocalisation sont ajoutées automatiquement (GPS) comprenant l'altitude. Au début, des images sont probablement annotées manuellement pour identifier les particularités du terrain et les meilleurs sites de plantation.
#### Algorithme
* Cartographie 3 D du terrain
* Apprentissage supervisé (probable)
* Réseaux de neurones convolutifs à plusieurs couches (CNN) pour identifier des particularités du terrain (probable)
* Algorithme de prédiction des meilleurs sites de plantation (probable)
* Algorithme de planification des trajectoires de plantation d'arbres qui optimise la couverture de plantation, le temps de vol et la consommation d'énergie
## Un assistant intelligent pour diagnostiquer les maladies du manioc
Nuru qui signifie «lumière» en Swahili est un assistant intelligent sur téléphone mobile pour identifier les maladies du manioc.
```
from IPython.display import Audio,Image, YouTubeVideo
id='NlpS-DhayQA'
YouTubeVideo(id=id,start=0,width=600,height=300,hl='fr',cc_lang_pref='fr',iv_load_policy=1)
```
#### Données
* 5000 images annotées manuellement des différentes maladies du manioc.
#### Algorithme
* Apprentissage supervisé
* Réseaux de neurones convolutifs à plusieurs couches (CNN) pour identifier des particularités des images
* Algorithme de prédiction des maladies
https://plantvillage.psu.edu/solutions
## L'écosystème de la science des données et de l'IA
<img src="module-01/ScDo-Ecosysteme_ScDo_IA.png" style="float:center;"/>
# La science des données
La science des données couvre l'ensemble des domaines liés à l'exploitation, la gestion et l'analyse de grands volumes de données, structurées et non structurées dans le but d'en extraire de l'information pertinente à l'aide d'algorithmes.
La science des données touche aux métiers de scientifique de données (data scientist) ou d'analyste des données (data analyst), d'analyste d'affaires (business analyst), d'ingénieur statisticien, de statisticien, d'informaticien, d'administrateur de bases de données, de développeur d'applications innovantes et du chercheur en informatique spécialisé en apprentissage automatique, en gestion avancée des données, en fouille de données ou en mathématiques de l'apprentissage.
Des exemples d'applications sont l'aide au diagnostic (notamment médical), la prévision financière (dont l'analyse du marché boursier), la publicité sur Internet, la recommandation de produits / filtrage collaboratif, le forage de données (data mining), le forage de textes (text mining) (par exemple, l'extraction de connaissances à partir du web), l'exploration des réseaux sociaux (graph mining), le filtrage des pourriels, la détection de menaces (incluant l'espionnage), la détection de fraudes, l'indexation / la recherche / la classification de documents (dont les moteurs de recherche), l'analytique des données d'apprentissage (learning analytics), la bioinformatique (séquençage de l'ADN), etc.
Par extension, on peut y ajouter la reconnaissance de formes (pattern recognition) comme la vision artificielle (dont la reconnaissance de visages ou d'objets), la reconnaissance d'écriture (codes postaux manuscrits), et le traitement automatique du langage incluant la reconnaissance vocale et la traduction automatique. Enfin, on peut y inclure les mécanismes d'adaptation à l'environnement appliqués à la robotique.
La science des données touche aussi les aspects juridiques et éthiques en relation avec l’utilisation des données personnelles.
</br>
<figure>
<img src="module-01/ScDo-petit.png" width='350' />
<figcaption style="text-align:center">La science des données est un domaine multidisciplinaire - Source: Wikipédia</figcaption>
</figure>
# Définition d'intelligence et d'intelligence artificielle
La définition de l'intelligence et de l'intelligence artificielle a fait l'objet de nombreuses discussions philosophiques et scientifiques.
L'intelligence est l'<b>ensemble des processus</b> retrouvés dans des <b>systèmes, plus ou moins complexes, vivants ou non</b>, qui mettent en jeu la <b>mémoire</b>, le <b>langage</b>, le <b>raisonnement</b>, l'<b>apprentissage</b>, la <b>résolution de problèmes</b>, la <b>compréhension du réel</b>, l'<b>adaptation à l'environnement</b>, la <b>prise de décision</b>, la <b>perception</b> ou l'<b>attention</b>.
Le terme est dérivé du latin intelligentia, « faculté de comprendre », dont le préfixe intus (« entre »), et le radical legere (« choisir, cueillir ») ou ligare (« lier ») suggèrent essentiellement l'aptitude à faire des choix, à lier des faits par la logique, le raisonnement déductif et inductif.
# Qu’est-ce que l’apprentissage automatique?
L’apprentissage automatique (en anglais machine learning) est un sous-domaine de l’intelligence artificielle.
L’apprentissage automatique consiste à programmer un algorithme capable d’apprendre par lui-même, à partir de données sans avoir à coder explicitement son comportement.
Plus précisément, l’apprentissage automatique permet à un logiciel d’apprendre à partir d’exemples (qu’on appelle aussi les données d’entraînement) afin d’acquérir de nouvelles connaissances, d’améliorer son fonctionnement, de s'adapter à son environnement ou de simuler l'intelligence humaine ou animale.
La plus grande force d’un algorithme d’apprentissage vient de sa capacité de généralisation qui est de bien fonctionner avec des données qu’il n’a jamais traitées.
L'apprentissage automatique est capable d'accomplir des tâches difficiles voir impossibles à réaliser par des moyens algorithmiques classiques. On a pas à coder explicitement le comportement de l’algorithme car l’algorithme apprend par lui-même à partir des données.
Cela passe le plus souvent par la création et la mise au point de modèles statistiques.
*<i> On dit aussi apprentissage artificiel, apprentissage machine ou apprentissage statistique</i>
## Définition formelle de l'apprentissage automatique (notion avancée)
Ci-dessous une définition plus formelle de l’apprentissage automatique énoncée en 1997 par Tom Mitchell, chercheur en IA à l’université Carnegie Mellon (CMU).
<img src="Module-05/ScDo-Definition_Apprentissage_Tom_Mitchell.png" width=700 style="float:center;" />
## Les composantes d’un algorithme d’apprentissage (notion avancée)
Un algorithme d’apprentissage automatique comporte quatre (4) parties: données, modèle, fonction de coût et algorithme d’optimisation.
<img src="Module-05/ScDo-Les_composantes_dun_algorithme_dapprentissage.png" width=700 style="float:center;" />
Un algorithme d’optimisation mathématique (ou optimiseur) est utilisé dans l'apprentissage automatique pendant l’entraînement d’un modèle sur les données. L’algorithme d'optimisation cherche à minimiser le coût des erreurs (i.e. la fonction de coût) entre les prédictions du modèle et chacune des données d’entraînement. Grossièrement, entraîner consiste à optimiser en calculant itérativement la valeur des paramètres qui minimisent l’erreur commise par un modèle sur un jeu de données d’entraînement. L’entraînement consiste à réduire progressivement les erreurs que fait l’algorithme.
Note pédagogique: L'aspect itératif et graduel du processus d’optimisation est un concept clé des algorithmes d'apprentissage. En effet, le calcul d’optimisation est répété sans cesse en boucle jusqu’à l’atteinte d’un critère d’arrêt. Le critère d’arrêt peut être un nombre d’itérations fixé à l’avance ou la constatation que le coût ou l'erreur ne change plus ou que le changement est inférieur à un seuil minimal fixé à l’avance.
Note: Cette recette s'applique aussi bien à un algorithme d'apprentissage supervisé où les données sont des couples (attributs, étiquette) alors qu'un algorithme d'apprentissage non-supervisé on aura uniquement des attributs sans étiquette avec une fonction de coût et un modèle appropriés.
Note terminologique: En français, on utilise fonction de coût, fonction de perte, fonction d’erreur ou fonction objective. En anglais: cost function, loss function, error function objective function. Souvent en apprentissage automatique, on utilisera les termes erreur, coût, perte et objectif d'une manière interchangeable.
Inspiré de la section 5.10 du livre «Deep Learning» [Goodfellow, Bengio & Courville, 2016]
## Types d'apprentissage automatique
<ul>
<li><b>L'apprentissage supervisé</b></li>
<br/>
L’apprentissage est dit supervisé quand les exemples (ou données d’entraînement) sont accompagnés de la réponse ou de la classe que l’on désire apprendre.
<br/><br/>
L'apprentissage supervisé se déroule en trois phases:
<br/><br/>
<ol>
<li>Lors de la première phase, dite phase d’étiquetage (ou d’annotation), les classes sont prédéterminées et les données d'entraînement (ou exemples) sont préalablement étiquetées selon leur classe, le plus souvent par un expert humain.</li>
<br/><br/>
<li>Puis lors de la seconde phase, dite phase d'apprentissage, l'algorithme entraîne un modèle à partir des données étiquetées. À cette étape, l’algorithme cherche à minimiser le coût des erreurs entre la prédiction de classe du modèle et la vraie classe (i.e. classe étiquetée) pour chaque donnée d’entraînement.</li>
<br/><br/>
<li>Enfin, dans la troisième phase, dite de prédiction, l'algorithme prédit la classe d'une nouvelle donnée mais cette fois non-étiquetée, en utilisant le modèle appris à la phase précédente.</li>
<br/><br/>
</ol>
Le résultat est l'appartenance à une classe ou une probabilité d'appartenance à chacune des classes prédéterminées.
<br/><br/>
Un des plus beaux exemples d'apprentissage supervisé est la classification des courriels entre pourriels (spam) et bon courriels.
<br/><br/>
<b>Un exemple d’apprentissage supervisé</b>
<br/><br/>
Pour bien comprendre, nous allons prendre un exemple simple d'apprentissage supervisé qui est la détection des pourriels (en anglais spam). Plus précisément, on apprend à un algorithme à classer les courriels entre « pourriel » et « non pourriel ».
<br/><br/>
<ol>
<li>Dans un premier temps, chaque courriel servant à l’entraînement du modèle est examiné et étiqueté par un humain selon deux classes: « pourriel » et « non pourriel ».</li>
<br/><br/>
<li>Puis l’algorithme d’apprentissage supervisé apprend à associer les différents mots contenus dans le courriel à la classe « pourriel » ou à la classe « non pourriel ». On entraîne l’algorithme sur des centaines, voire des milliers de courriels. Petit à petit, en traitant chaque courriel des données d’entraînement, l’algorithme améliore ses prédictions en cherchant à réduire ses erreurs de prédictions.
</li>
<br/><br/>
<li>À la fin, on fournira le contenu d’un courriel jamais examiné (et non étiqueté) et à partir de son contenu, notre algorithme ou plutôt notre modèle entraîné sera capable de prédire s’il s’agit d’un pourriel ou non.</li>
</ol>
<br/><br/>
<b>Note:</b> Le plus beau de l'affaire est que l'utilisateur d'un outil de courriel comme le GMail de Google fait bénévolement l'étiquetage des pourriels par sa simple utilisation de l'outil.
<br/><br/>
<li><b>L'apprentissage non supervisé</b></li>
<br/>
L'algorithme ne dispose que d'exemples non étiquetés. De plus, le nombre de classes et leur nature n'ont pas été prédéterminés.
<br/><br/>
En situation d'apprentissage non supervisé, l'algorithme doit découvrir par lui-même la structure des données.
<br/><br/>
Par exemple, le groupage de données (en anglais clustering) est un algorithme d'apprentissage non supervisé. L’apprentissage non supervisé cherche à découvrir la distribution statistique des données d’entraînement afin d’en faire ressortir tendance et dispersion.
<br/><br/>
L'algorithme se base sur les données (exemples) disponibles, pour les classer en groupes homogènes correspondant à une classe selon une mesure de similarité qui calcule la distance entre paires d'exemples. Le résultat est l'appartenance à un groupe (ou classe) ou une probabilité d'appartenance à chacune des groupes (ou classes) formés par l'algorithme.
<br/><br/>
<b>Exemples d’apprentissage non-supervisé</b>
<br/><br/>
Un exemple typique d’apprentissage non-supervisé en marketing est de regrouper des clients en différentes catégories émergentes en fonction de leurs comportements d'achat.
<br/><br/>
Un autre exemple d’apprentissage non supervisé dans le domaine bancaire est la détection de fraude sur la base d’anomalies de comportement ou de données aberrantes (en anglais outliers).
<br/><br/>
<li><b>L'apprentissage par renforcement</b></li>
<br/>
L'algorithme apprend un comportement à partir d'expériences de façon à optimiser les récompenses reçues au cours du temps.
<br/><br/>
Typiquement, un agent autonome réalise une action en fonction de son état courant et de l'observation de son environnement. En retour de l'action de l'agent, l'environnement procure à l'agent une récompense ou une punition.
<br/><br/>
<b>Un exemple d’apprentissage par renforcement</b>
<br/><br/>
Une application spectaculaire de l'apprentissage par renforcement est AlphaGo de DeepMind, le premier logiciel d'intelligence artificielle capable de battre un champion au jeu de Go.
</ul>
## Deux grands types de modèles statistiques (notion avancée)
Sur le plan statistique, il existe deux grands types de modèles, les <b>modèles prédictifs</b> et les <b>modèles génératifs</b>. Comme son nom l'indique, un modèle prédictif est capable de prédire un résultat comme la classe d'un objet à partir des données le concernant. Plus général, un modèle génératif est capable de produire de nouvelles données selon la distribution statistique des données originales.
Notez que l'on peut créer un modèle prédictif à partir d'un modèle génératif en appliquant la loi statistique de Bayes.
# Outils
## Pourquoi Python?
<img src="Module-01/ScDo-Python_Logo.png" width='300' height='100' style="float:left;" />
Depuis sa création en 1991 par Guido van Rossum, Python est devenu un des langages dynamiques de programmation (ou langages à scripts, ou langages interprétés) les plus populaires avec Perl et Ruby. Python permet un développement rapide, interactif et incrémentiel, idéalement selon une approche de développement guidé par les tests.
Python se distingue par l’importance de son utilisation dans le domaine scientifique et plus particulièrement dans les sciences des données et l'apprentissage automatique. On ne peut éviter de comparer Python aux autres langages utilisés pour l’analyse des données comme R, Mathlab, S-plus et SAS.
Dans les dernières années, l’apparition de bibliothèques et d’outils spécialisés comme NumPy, SciPy, pandas, matplotlib, IPython et scikit-learn, et plus particulièrement l'émergence de l'apprentissage profond (deep learning) avec TensorFlow, Keras, PyTorch combinée aux forces de Python comme langage de programmation générique font de Python un excellent choix de langage unique pour toute application basée sur les données. Notez que la quasi totalité des bibliothèques scientifiques en Python sont des logiciels libres.
Une <a href="http://www.xmind.net/m/WvfC/">carte conceptuelle</a> (ou carte mentale) de l'écosystème Python pour la science des données. Source: <a href="http://www.xmind.net/">XMind</a>
## Écosystème Python
<ol>
<li><b>IPython</b></li><br>
<a href="https://ipython.org/">IPython</a> pour Iteractive Python, un outil Python de programmation interactive en logiciel libre, offre des carnets web (Notebooks) qui intègrent dans une même page web du code Python exécutable, des textes, des images, des graphiques, des hyperliens, en somme tout le travail des différents outils en Python sur les différentes données.
<br><br><li><b>NumPy</b></li><br>
Diminutif de Numerical Python, <a href="https://numpy.org/">NumPy</a> est une bibliothèque Python en logiciel libre utilisée pour les calculs scientifiques de base. Essentiellement basé sur la notion de matrices, NumPy est à la fois performant et super utile pour l'analyse des données et les calculs mathématiques.
<br><br><li><b>SciPy</b></li><br>
Diminutif de Scientific Python, <a href="https://www.scipy.org/">SciPy</a> est une bibliothèque Python en logiciel libre construite au-dessus de NumPy. SciPy offre des fonctions avancées sur le plan mathématique comme la résolution d'équations différentielles, les fonctions spéciales, l'optimisation, l'intégration, etc.
<br><br><li><b>Pandas</b></li><br>
Diminutif de (PANel DAta Structures), <a href="https://pandas.pydata.org/">Pandas</a> est une bibliothèque Python en logiciel libre qui touche la manipulation et l'analyse des données. Plus particulièrement, Pandas a été conçu pour manipuler des données contenues dans un tableau de type tableur (DataFRame) ou chaque ligne du tableau représente un exemplaire de la donnée et cahque colonne un attribut de la donnée.
<br><br><li><b>Matplotlib</b></li><br>
<a href="https://matplotlib.org/">Matplotlib</a> est une bibliothèque en logiciel libre de visualisation en Python. Matplotlib offre une gamme d'outils personnalisables principalement en 2D pour visualiser des ensembles de données complexes (nuages de points, histogrammes, diagrammes en bâtons, courbes, camemberts, etc.).
<br><br><li><b>Scikit-learn</b></li><br>
Développée à l’INRIA en France, <a href="https://scikit-learn.org/stable/">Scikit-Learn</a> (aussi appelée Sklearn) est une bibliothèque Python en logiciel libre spécialisée en apprentissage automatique classique (machine learning) qui se marie bien avec l'écosystème Python dont NumPy et SciPy.
<br><br><li><b>TensorFlow</b></li><br>
<a href="https://www.tensorflow.org/">TensorFlow</a> est une bibliothèque Python en logiciel libre pour l'apprentissage profond développée par Google et fortement inspiré de <a href="http://deeplearning.net/software/theano/">Theano</a> , (l'outil pionnier développé au MILA, Université de Montréal, [Bastien et al., 2012]). Le terme désigne également la couche API de base de la pile technologique TensorFlow qui soutient les graphes de calculs (dataflow). Bien que TensorFlow soit principalement utilisé pour l'apprentissage profond, il peut aussi être utilisé pour des tâches autres nécessitant de l'apprentissage automatique classique ou un calcul numérique à l'aide des graphes de calculsÀ.
<br><br><li><b>Keras</b></li><br>
<a href="https://keras.io/">Keras</a> est une bibliothèque Python de haut niveau en logiciel libre pour l'apprentissage profond bâtie au dessus de TensorFlow, Theano ou CNTK. Keras est publié en code source ouvert sous la licence MIT. Keras est développé par François Chollet, un ingénieur de Google, pour faciliter la création et la mise au point de modèles d'apprentissage profond.
</ol>
## Autres outils
En logiciel libre, il existe trois principales familles d’outils pour l’analyse statistique de données basées sur autant de langages de programmation.
<img src="Module-01/ScDo-Outils_Libres.png" />
Note: Julia est également un langage à surveiller...
## Processus de forage des données selon la norme CRISP-DM
CRISP-DM (Cross Industry Standard Process for Data Mining) est une norme industrielle qui décrit le processus de forage des données communément utilisée par les experts de la science des données (ou scientifiques de données).
Source: <a href="https://fr.wikipedia.org/wiki/Cross_Industry_Standard_Process_for_Data_Mining#/media/Fichier:Diagramme_du_Processus_CRISP-DM.png">Wikipédia - Diagramme du Processus CRISP-DM - Abdessamad DERRAZ<a/>
<img src="Module-01/ScDo-Diagramme_du_Processus_CRISP-DM.png" width=500 style="float:center;"/>
| github_jupyter |
# Deep Q-Network (DQN)
---
In this notebook, you will implement a DQN agent with OpenAI Gym's LunarLander-v2 environment.
### 1. Import the Necessary Packages
```
import gym
import random
import torch
import numpy as np
from collections import deque
import matplotlib.pyplot as plt
%matplotlib inline
```
### 2. Instantiate the Environment and Agent
Initialize the environment in the code cell below.
```
env = gym.make('LunarLander-v2')
env.seed(0)
print('State shape: ', env.observation_space.shape)
print('Number of actions: ', env.action_space.n)
```
Please refer to the instructions in `Deep_Q_Network.ipynb` if you would like to write your own DQN agent. Otherwise, run the code cell below to load the solution files.
```
from dqn_agent import Agent
agent = Agent(state_size=8, action_size=4, seed=0)
# watch an untrained agent
state = env.reset()
for j in range(200):
action = agent.act(state)
env.render()
state, reward, done, _ = env.step(action)
if done:
break
env.close()
```
### 3. Train the Agent with DQN
Run the code cell below to train the agent from scratch. You are welcome to amend the supplied values of the parameters in the function, to try to see if you can get better performance!
Alternatively, you can skip to the next step below (**4. Watch a Smart Agent!**), to load the saved model weights from a pre-trained agent.
```
def dqn(n_episodes=2000, max_t=1000, eps_start=1.0, eps_end=0.01, eps_decay=0.995):
"""Deep Q-Learning.
Params
======
n_episodes (int): maximum number of training episodes
max_t (int): maximum number of timesteps per episode
eps_start (float): starting value of epsilon, for epsilon-greedy action selection
eps_end (float): minimum value of epsilon
eps_decay (float): multiplicative factor (per episode) for decreasing epsilon
"""
scores = [] # list containing scores from each episode
scores_window = deque(maxlen=100) # last 100 scores
eps = eps_start # initialize epsilon
for i_episode in range(1, n_episodes+1):
state = env.reset()
score = 0
for t in range(max_t):
action = agent.act(state, eps)
next_state, reward, done, _ = env.step(action)
agent.step(state, action, reward, next_state, done)
state = next_state
score += reward
if done:
break
scores_window.append(score) # save most recent score
scores.append(score) # save most recent score
eps = max(eps_end, eps_decay*eps) # decrease epsilon
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_window)), end="")
if i_episode % 100 == 0:
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_window)))
if np.mean(scores_window)>=200.0:
print('\nEnvironment solved in {:d} episodes!\tAverage Score: {:.2f}'.format(i_episode-100, np.mean(scores_window)))
torch.save(agent.qnetwork_local.state_dict(), 'checkpoint.pth')
break
return scores
scores = dqn()
# plot the scores
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(len(scores)), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
```
### 4. Watch a Smart Agent!
In the next code cell, you will load the trained weights from file to watch a smart agent!
```
# load the weights from file
agent.qnetwork_local.load_state_dict(torch.load('checkpoint.pth'))
for i in range(3):
state = env.reset()
for j in range(200):
action = agent.act(state)
env.render()
state, reward, done, _ = env.step(action)
if done:
break
env.close()
```
### 5. Explore
In this exercise, you have implemented a DQN agent and demonstrated how to use it to solve an OpenAI Gym environment. To continue your learning, you are encouraged to complete any (or all!) of the following tasks:
- Amend the various hyperparameters and network architecture to see if you can get your agent to solve the environment faster. Once you build intuition for the hyperparameters that work well with this environment, try solving a different OpenAI Gym task with discrete actions!
- You may like to implement some improvements such as prioritized experience replay, Double DQN, or Dueling DQN!
- Write a blog post explaining the intuition behind the DQN algorithm and demonstrating how to use it to solve an RL environment of your choosing.
| github_jupyter |
### Note
* Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
```
# Dependencies and Setup
import pandas as pd
# File to Load (Remember to Change These)
file_to_load = "Resources/purchase_data.csv"
# Read Purchasing File and store into Pandas data frame
purchase_data = pd.read_csv(file_to_load)
```
## Player Count
* Display the total number of players
```
#Each players will have unique SN and can purchase multiple times,
#so one way to find out how many players without dups is to use SN
total_players = len(purchase_data["SN"].unique())
print(f'Total players is {total_players}')
```
## Purchasing Analysis (Total)
* Run basic calculations to obtain number of unique items, average price, etc.
* Create a summary data frame to hold the results
* Optional: give the displayed data cleaner formatting
* Display the summary data frame
```
#Setting variables
unique_item = len(purchase_data["Item Name"].unique())
average = round(purchase_data["Price"].mean(),2)
total_purchase = purchase_data["Purchase ID"].count()
total_revenue = purchase_data["Price"].sum()
#Put them in a dictionary DF
summary_data = pd.DataFrame({
"Number of Unique Items": [unique_item],
"Average Price": [average],
"Number of Purchases": [total_purchase],
"Total Revenue": [total_revenue],
})
summary_data["Average Price"] = summary_data["Average Price"].map('${:,.2f}'.format)
summary_data["Total Revenue"] = summary_data["Total Revenue"].map('${:,.2f}'.format)
summary_data
```
## Gender Demographics
* Percentage and Count of Male Players
* Percentage and Count of Female Players
* Percentage and Count of Other / Non-Disclosed
```
#Create a list of male player and remove duplicates
male_list = purchase_data.loc[purchase_data["Gender"] == "Male"]
male_count = len(male_list["SN"].unique())
percent_male = round((male_count/total_players)*100,2)
#Create a list of female player and remove duplicates
female_list = purchase_data.loc[purchase_data["Gender"] == "Female"]
female_count = len(female_list["SN"].unique())
percent_female = round((female_count/total_players)*100,2)
#Create a list of others and remove duplicates
others_list = purchase_data.loc[purchase_data["Gender"] == "Other / Non-Disclosed"]
others_count = len(others_list)
percent_others = round((others_count/total_players)*100,2)
gender_demo = {'Total Count': [male_count, female_count, others_count],
'Percentage of Players': [percent_male, percent_female, percent_others]}
gender_demographics_table = pd.DataFrame(gender_demo, columns = ['Total Count', 'Percentage of Players'], index=['Male', 'Female', 'Others/Non-Disclosed'])
gender_demographics_table["Percentage of Players"]=gender_demographics_table["Percentage of Players"].map("{0:.2f}%".format)
gender_demographics_table
```
## Purchasing Analysis (Gender)
* Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. by gender
* Create a summary data frame to hold the results
* Optional: give the displayed data cleaner formatting
* Display the summary data frame
```
#Use the male list to find total of purchase price for male only
male_total_pur_val = male_list["Price"].sum()
male_avg_pur_price = round((male_total_pur_val/male_count),2)
#Use total purchase price divide male(removed duplicates) to find Avg Total Purchase per Person
male_avg_total_pur_per = round((male_total_pur_val/male_count), 2)
#Use male list to find total purchase for male
total_male_count = male_list["Gender"].count()
#Use the female list to find total of purchase price for female only
female_total_pur_val = female_list["Price"].sum()
female_avg_pur_price = round((female_total_pur_val/male_count),2)
#Use total purchase price divide female(removed duplicates) to find Avg Total Purchase per Person
female_avg_total_pur_per = round((female_total_pur_val/female_count), 2)
#Use female list to find total purchase for female
total_female_count = female_list["Gender"].count()
#Use the other list to find total of purchase price for other only
others_total_pur_val = others_list["Price"].sum()
others_avg_pur_price = round((others_total_pur_val/others_count),2)
#Use total purchase price divide others(removed duplicates) to find Avg Total Purchase per Person
others_avg_total_pur_per = round((others_total_pur_val/others_count), 2)
#Use female list to find total purchase for others
total_others_count = others_list["Gender"].count()
#Put them in a dictionary
purchase_analysis = { 'Purchase Count': [total_female_count, total_male_count, total_others_count],
'Average Purchase Price': [female_avg_pur_price, male_avg_pur_price, others_avg_pur_price],
'Total Purchase Value': [female_total_pur_val, male_total_pur_val, others_total_pur_val],
'Avg Total Purchase per Person': [female_avg_total_pur_per, male_avg_total_pur_per, others_avg_total_pur_per]}
purchase_analysis_table = pd.DataFrame(purchase_analysis, columns = ['Purchase Count', 'Average Purchase Price',
'Total Purchase Value', 'Avg Total Purchase per Person'],
index=['Male', 'Female', 'Others/Non-Disclosed'])
index = purchase_analysis_table.index
index.name = "Gender"
purchase_analysis_table["Average Purchase Price"]=purchase_analysis_table["Average Purchase Price"].map('${:,.2f}'.format)
purchase_analysis_table["Total Purchase Value"]=purchase_analysis_table["Total Purchase Value"].map('${:,.2f}'.format)
purchase_analysis_table["Avg Total Purchase per Person"]=purchase_analysis_table["Avg Total Purchase per Person"].map('${:,.2f}'.format)
purchase_analysis_table
```
## Age Demographics
* Establish bins for ages
* Categorize the existing players using the age bins. Hint: use pd.cut()
* Calculate the numbers and percentages by age group
* Create a summary data frame to hold the results
* Optional: round the percentage column to two decimal points
* Display Age Demographics Table
```
#SCRAP
#Create bins where age will be held
bins = [0, 9, 14, 19, 24, 29, 34, 39, 100]
#Create group names for the bins
group_names = ["<10", "10-14", "15-19", "20-24", "25-29", "30-34", "35-39", "40+"]
purchase_data["Age Group"] = pd.cut(purchase_data["Age"], bins, labels=group_names, include_lowest=True)
#Remove duplicates player in the new list
purchase_data_nodup = purchase_data.drop_duplicates(subset=['SN'])
#Create lists of group age and store it as value
less_ten_list = purchase_data_nodup.loc[purchase_data_nodup["Age Group"] == "<10"]
less_ten = len(less_ten_list)
fourteen_list = purchase_data_nodup.loc[purchase_data_nodup["Age Group"] == "10-14"]
fourteen = len(fourteen_list)
nineteen_list = purchase_data_nodup.loc[purchase_data_nodup["Age Group"] == "15-19"]
nineteen = len(nineteen_list)
twenty_four_list = purchase_data_nodup.loc[purchase_data_nodup["Age Group"] == "20-24"]
twenty_four = len(twenty_four_list)
twenty_nine_list = purchase_data_nodup.loc[purchase_data_nodup["Age Group"] == "25-29"]
twenty_nine = len(twenty_nine_list)
thirty_four_list = purchase_data_nodup.loc[purchase_data_nodup["Age Group"] == "30-34"]
thirty_four = len(thirty_four_list)
thirty_nine_list = purchase_data_nodup.loc[purchase_data_nodup["Age Group"] == "35-39"]
thirty_nine = len(thirty_nine_list)
forty_list = purchase_data_nodup.loc[purchase_data_nodup["Age Group"] == "40+"]
forty = len(forty_list)
#Convert them into percentage for each group
per_less_ten = round(((less_ten/total_players)*100), 2)
per_fourteen = round(((fourteen/total_players)*100), 2)
per_nineteen = round(((nineteen/total_players)*100), 2)
per_twenty_four = round(((twenty_four/total_players)*100), 2)
per_twenty_nine = round(((twenty_nine/total_players)*100), 2)
per_thirty_four = round(((thirty_four/total_players)*100), 2)
per_thirty_nine = round(((thirty_nine/total_players)*100), 2)
per_forty = round(((forty/total_players)*100), 2)
#Put them in a dictionary
age_demo = { 'Total Count': [less_ten, fourteen, nineteen, twenty_four, twenty_nine, thirty_four, thirty_nine, forty],
'Percentage of Players':[per_less_ten, per_fourteen, per_nineteen, per_twenty_four, per_twenty_nine, per_thirty_four, per_thirty_nine, per_forty]}
age_demo_table = pd.DataFrame(age_demo, columns = ['Total Count', 'Percentage of Players'], index = ["<10", "10-14", "15-19", "20-24", "25-29", "30-34", "35-39", "40+"])
age_demo_table["Percentage of Players"]=age_demo_table["Percentage of Players"].map("{0:.2f}%".format)
age_demo_table
```
## Purchasing Analysis (Age)
* Bin the purchase_data data frame by age
* Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. in the table below
* Create a summary data frame to hold the results
* Optional: give the displayed data cleaner formatting
* Display the summary data frame
```
#Get value for Purchase Count for each group
count_ten = len(purchase_data.loc[purchase_data['Age Group'] == '<10'])
count_fourteen = len(purchase_data.loc[purchase_data['Age Group'] == '10-14'])
count_nineteen = len(purchase_data.loc[purchase_data['Age Group'] == '15-19'])
count_twentyfour = len(purchase_data.loc[purchase_data['Age Group'] == '20-24'])
count_twentynine = len(purchase_data.loc[purchase_data['Age Group'] == '25-29'])
count_thirtyfour = len(purchase_data.loc[purchase_data['Age Group'] == '30-34'])
count_thirtynine = len(purchase_data.loc[purchase_data['Age Group'] == '35-39'])
count_forty = len(purchase_data.loc[purchase_data['Age Group'] == '40+'])
#Get value for Total Purchase Value for each group
total_pur_val_ten = purchase_data.loc[purchase_data['Age Group'] == '<10', 'Price'].sum()
total_pur_val_fourteen = purchase_data.loc[purchase_data['Age Group'] == '10-14', 'Price'].sum()
total_pur_val_nineteen = purchase_data.loc[purchase_data['Age Group'] == '15-19', 'Price'].sum()
total_pur_val_twentyfour = purchase_data.loc[purchase_data['Age Group'] == '20-24', 'Price'].sum()
total_pur_val_twentynine = purchase_data.loc[purchase_data['Age Group'] == '25-29', 'Price'].sum()
total_pur_val_thirtyfour = purchase_data.loc[purchase_data['Age Group'] == '30-34', 'Price'].sum()
total_pur_val_thirtynine = purchase_data.loc[purchase_data['Age Group'] == '35-39', 'Price'].sum()
total_pur_val_forty = purchase_data.loc[purchase_data['Age Group'] == '40+', 'Price'].sum()
#Get value for Average Purchase Price
avg_ten = round((total_pur_val_ten/count_ten), 2)
avg_fourteen = round((total_pur_val_fourteen/count_ten), 2)
avg_nineteen = round((total_pur_val_nineteen/count_ten), 2)
avg_twentyfour = round((total_pur_val_twentyfour/count_ten), 2)
avg_twentynine = round((total_pur_val_twentynine/count_ten), 2)
avg_thirtyfour = round((total_pur_val_thirtyfour/count_ten), 2)
avg_thirtynine = round((total_pur_val_thirtynine/count_ten), 2)
avg_forty = round((total_pur_val_forty/count_ten), 2)
#Get value for Avg Total Purchase per Person
avg_per_ten = round((total_pur_val_ten/less_ten), 2)
avg_per_fourteen = round((total_pur_val_fourteen/fourteen), 2)
avg_per_nineteen = round((total_pur_val_nineteen/count_ten), 2)
avg_per_twentyfour = round((total_pur_val_twentyfour/twenty_four), 2)
avg_per_twentynine = round((total_pur_val_twentynine/twenty_nine), 2)
avg_per_thirtyfour = round((total_pur_val_thirtyfour/thirty_four), 2)
avg_per_thirtynine = round((total_pur_val_thirtynine/thirty_nine), 2)
avg_per_forty = round((total_pur_val_forty/forty), 2)
purchase_analysis_age = {'Purchase Count': [count_ten, count_fourteen, count_nineteen, count_twentyfour,
count_twentynine, count_thirtyfour, count_thirtynine, count_forty],
'Average Purchase Price': [avg_ten, avg_fourteen, avg_nineteen, avg_twentyfour,
avg_twentynine, avg_thirtyfour, avg_thirtynine, avg_forty],
'Total Purchase Value': [total_pur_val_ten, total_pur_val_fourteen, total_pur_val_nineteen, total_pur_val_twentyfour,
total_pur_val_twentynine, total_pur_val_thirtyfour, total_pur_val_thirtynine, total_pur_val_forty],
'Avg Total Purchase per Person': [avg_per_ten, avg_per_fourteen, avg_per_nineteen, avg_per_twentyfour,
avg_per_twentynine, avg_per_thirtyfour, avg_per_thirtynine, avg_per_forty]}
purchase_analysis_age_table = pd.DataFrame(purchase_analysis_age, columns = ['Purchase Count', 'Average Purchase Price',
'Total Purchase Value', 'Avg Total Purchase per Person'],
index = ["<10", "10-14", "15-19", "20-24", "25-29", "30-34", "35-39", "40+"])
index = purchase_analysis_age_table.index
index.name = "Age Ranges"
#setting format
purchase_analysis_age_table["Average Purchase Price"]=purchase_analysis_age_table["Average Purchase Price"].map('${:,.2f}'.format)
purchase_analysis_age_table["Total Purchase Value"]=purchase_analysis_age_table["Total Purchase Value"].map('${:,.2f}'.format)
purchase_analysis_age_table["Avg Total Purchase per Person"]=purchase_analysis_age_table["Avg Total Purchase per Person"].map('${:,.2f}'.format)
purchase_analysis_age_table
```
## Top Spenders
* Run basic calculations to obtain the results in the table below
* Create a summary data frame to hold the results
* Sort the total purchase value column in descending order
* Optional: give the displayed data cleaner formatting
* Display a preview of the summary data frame
```
#Count how many players occurs in the whole data
purchase_count = purchase_data["SN"].value_counts()
#Create a group by SN data frame
grouped_data = purchase_data.groupby(['SN'])
total_purchase_value = grouped_data["Price"].sum()
top_spenders_table = pd.DataFrame({'Purchase Count': purchase_count,
'Average Purchase Price': round((total_purchase_value/players_count),2),
'Total Purchase Value': total_purchase_value})
top_spenders_table = top_spenders_table.sort_values(["Total Purchase Value"], ascending=False)
#Setting format
top_spenders_table["Average Purchase Price"]=top_spenders_table["Average Purchase Price"].map('${:,.2f}'.format)
top_spenders_table["Total Purchase Value"]=top_spenders_table["Total Purchase Value"].map('${:,.2f}'.format)
top_spenders_table.head(5)
```
## Most Popular Items
* Retrieve the Item ID, Item Name, and Item Price columns
* Group by Item ID and Item Name. Perform calculations to obtain purchase count, average item price, and total purchase value
* Create a summary data frame to hold the results
* Sort the purchase count column in descending order
* Optional: give the displayed data cleaner formatting
* Display a preview of the summary data frame
```
#Retrieve the Item ID, Item Name, and Item Price columns
popular_list = purchase_data[["Item ID", "Item Name", "Price"]]
#Group by Item ID and Item Name
grouped_list = popular_list.groupby(["Item ID", "Item Name"])
popular_item_count = grouped_list["Item Name"].count()
popular_item_total_purchase = grouped_list["Price"].sum()
popular_item_avg_price = round((popular_item_total_purchase/popular_item_count),2)
#Put them in dictionary
popular_item = pd.DataFrame({'Purchase Count': popular_item_count,
'Item Price': popular_item_avg_price,
'Total Purchase Value': popular_item_total_purchase})
popular_item = popular_item.sort_values(["Purchase Count"], ascending=False)
#Setting format
popular_item["Item Price"]=popular_item["Item Price"].map('${:,.2f}'.format)
popular_item["Total Purchase Value"]=popular_item["Total Purchase Value"].map('${:,.2f}'.format)
popular_item.head(5)
```
## Most Profitable Items
* Sort the above table by total purchase value in descending order
* Optional: give the displayed data cleaner formatting
* Display a preview of the data frame
```
popular_item = popular_item.sort_values(["Purchase Count", "Total Purchase Value"], ascending=False)
popular_item.head(5)
```
| github_jupyter |
# Part 1 - Point source plotting
To perform a neutronics simulation a neutron source must also be defined.
This python notebook allows users to make a simple OpenMC point source and plot its energy, position and initial directions.
```
from IPython.display import HTML
HTML('<iframe width="560" height="315" src="https://www.youtube.com/embed/j9dT1Viqcu4" frameborder="0" allow="accelerometer; autoplay; clipboard-write; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>')
import openmc
from source_extraction_utils import * # imports plotting functions
```
This first code block creates an isotropic point source with 14 MeV monoenergetic neutrons.
```
# initialises a new source object
source = openmc.Source()
# sets the location of the source to x=0 y=0 z=0
source.space = openmc.stats.Point((0, 0, 0))
# sets the direction to isotropic
source.angle = openmc.stats.Isotropic()
# sets the energy distribution to 100% 14MeV neutrons
source.energy = openmc.stats.Discrete([14e6], [1])
create_inital_particles(source)
plot_energy_from_initial_source(input_filename='initial_source.h5')
```
The next code block creates an isotropic point source with a fission energy distribution.
```
source = openmc.Source()
source.space = openmc.stats.Point((0, 0, 0))
source.angle = openmc.stats.Isotropic()
# Documentation on the Watt distribution is here
# https://docs.openmc.org/en/stable/pythonapi/generated/openmc.data.WattEnergy.html
source.energy = openmc.stats.Watt(a=988000.0, b=2.249e-06)
create_inital_particles(source)
plot_energy_from_initial_source(input_filename='initial_source.h5')
```
This code block creates an isotropic point source with a fusion energy distribution.
```
source = openmc.Source()
source.space = openmc.stats.Point((0, 0, 0))
source.angle = openmc.stats.Isotropic()
# Documentation on the Muir distribution is here
# https://docs.openmc.org/en/stable/pythonapi/generated/openmc.stats.Muir.html
source.energy = openmc.stats.Muir(e0=14080000.0, m_rat=5.0, kt=20000.0)
create_inital_particles(source)
plot_energy_from_initial_source(input_filename='initial_source.h5')
```
The following code block plots the birth location of the neutrons from a 14 MeV monoenergetic point source.
```
# Creates an isotropic point source with monoenergetic 14MeV neutrons
source = openmc.Source()
source.space = openmc.stats.Point((0, 0, 0))
source.angle = openmc.stats.Isotropic()
source.energy = openmc.stats.Discrete([14e6], [1])
create_inital_particles(source)
# plots the position of neutrons created
plot_postion_from_initial_source()
```
Finally, the following code block plots the birth direction of the neutrons from the same source.
```
# Creates an isotropic point source with monoenergetic 14MeV neutrons
source = openmc.Source()
source.space = openmc.stats.Point((0, 0, 0))
source.angle = openmc.stats.Isotropic()
source.energy = openmc.stats.Discrete([14e6], [1])
create_inital_particles(source)
# plots the initial direction of neutrons created
plot_direction_from_initial_source()
```
**Learning Outcomes for Part 1:**
- OpenMC can be used to create neutron point sources with different energy distributions.
| github_jupyter |
# Calibrating qubits using Qiskit and OpenPulse
Qiskit is an open-source framework for programming quantum computers (Ref. [1](#refs)). Using Qiskit, quantum circuits can be built, simulated and executed on quantum devices.
OpenPulse provides a language for specifying pulse level control (i.e. control of the continuous time dynamics of input signals) of a general quantum device independent of the specific hardware implementation (Ref. [2](#refs)).
In this tutorial, we show how to implement typical single-qubit calibration and characterization experiments using Qiskit and OpenPulse. These are typically the first round of experiments that would be done in the lab immediately after a device has been fabricated and installed into a system. The presentation is pedagogical, and allows students to explore two-level-system dynamics experimentally. All units are returned as standard SI (ie Hz, sec, etc).
Each experiment gives us more information about the system, which is typically used in subsequent experiments. For this reason, this notebook has to be mostly executed in order.
## Contents
Part 0. [Getting started](#importing) <br/>
Part 1. [Finding the qubit frequency using a frequency sweep](#frequencysweep) <br/>
Part 2. Calibrating and using a $\pi$ pulse <br/>
A. [Calibrating $\pi$ pulses using a Rabi experiment](#rabi) <br/>
B. [Determining 0 vs 1](#zerovone) <br/>
C. [Measuring $T_1$ using inversion recovery](#T1)
Part 3. Determining qubit coherence <br/>
A. [Measuring the qubit frequency precisely using a Ramsey experiment](#ramsey) <br/>
B. [Measuring $T_2$ using Hahn echoes](#hahn) <br/>
Part 4. [References](#refs)
## 0. Getting started <a id="importing"></a>
We'll first get our basic dependencies set up and ready to go. Since we want to use real, noisy devices for our calibration experiments in this notebook, we need to load our IBMQ account and set the appropriate backend.
```
import warnings
warnings.filterwarnings('ignore')
from qiskit.tools.jupyter import *
from qiskit import IBMQ
IBMQ.load_account()
provider = IBMQ.get_provider(hub='ibm-q', group='open', project='main')
backend = provider.get_backend('ibmq_armonk')
```
We verify that the backend supports OpenPulse features by checking the backend configuration. The config provides us with general information about the structure of the backend setup.
```
backend_config = backend.configuration()
assert backend_config.open_pulse, "Backend doesn't support OpenPulse"
```
For instance, we can find the sampling time for the backend pulses within the backend configuration. This will be a very useful value to us as we build and execute our calibration routines.
```
dt = backend_config.dt
print(f"Sampling time: {dt*1e9} ns") # The configuration returns dt in seconds, so multiply by
# 1e9 to get nanoseconds
```
The backend defaults provide a starting point for how to use the backend. It contains estimates for qubit frequencies and default programs to enact basic quantum operators. We can access them with the following:
```
backend_defaults = backend.defaults()
```
# Part 1
## 1. Finding the qubit frequency using a frequency sweep <a id="frequencysweep"></a>
We begin by searching for the qubit frequency. The qubit frequency is the difference in energy between the ground and excited states, which we label the $\vert0\rangle$ and $\vert1\rangle$ states, respectively. This frequency will be crucial for creating pulses which enact particular quantum operators on the qubit -- the final goal of our calibration!
With superconducting qubits, higher energy levels are also available, but we fabricate the systems to be anharmonic so that we can control which transition we are exciting. That way, we are able to isolate two energy levels and treat each qubit as a basic two-level system, ignoring higher energy states.
In a typical lab setting, the qubit frequency can be found by sweeping a range of frequencies and looking for signs of absorption using a tool known as a Network Analyzer. This measurement gives a rough estimate of the qubit frequency. Later on, we will see how to do a more precise measurement using a Ramsey pulse sequence.
First, we define the frequency range that will be swept in search of the qubit. Since this can be arbitrarily broad, we restrict ourselves to a window of 40 MHz around the estimated qubit frequency in `backend_defaults`. We step the frequency in units of 1 MHz.
```
import numpy as np
# unit conversion factors -> all backend properties returned in SI (Hz, sec, etc)
GHz = 1.0e9 # Gigahertz
MHz = 1.0e6 # Megahertz
us = 1.0e-6 # Microseconds
ns = 1.0e-9 # Nanoseconds
# We will find the qubit frequency for the following qubit.
qubit = 0
# The sweep will be centered around the estimated qubit frequency.
center_frequency_Hz = backend_defaults.qubit_freq_est[qubit] # The default frequency is given in Hz
# warning: this will change in a future release
print(f"Qubit {qubit} has an estimated frequency of {center_frequency_Hz / GHz} GHz.")
# We will sweep 40 MHz around the estimated frequency
frequency_span_Hz = 40 * MHz
# in steps of 1 MHz.
frequency_step_Hz = 1 * MHz
# We will sweep 20 MHz above and 20 MHz below the estimated frequency
frequency_min = center_frequency_Hz - frequency_span_Hz / 2
frequency_max = center_frequency_Hz + frequency_span_Hz / 2
# Construct an np array of the frequencies for our experiment
frequencies_GHz = np.arange(frequency_min / GHz,
frequency_max / GHz,
frequency_step_Hz / GHz)
print(f"The sweep will go from {frequency_min / GHz} GHz to {frequency_max / GHz} GHz \
in steps of {frequency_step_Hz / MHz} MHz.")
```
Next, we define the pulses we will use for our experiment. We will start with the drive pulse, which is a Gaussian pulse.
Remember the value `dt` from earlier? All durations in pulse are given in terms of `dt`. In the next cell, we define the length of the drive pulse in terms of `dt`.
```
# samples need to be multiples of 16
def get_closest_multiple_of_16(num):
return (int(num) - (int(num)%16))
from qiskit import pulse # This is where we access all of our Pulse features!
from qiskit.pulse import pulse_lib # This Pulse module helps us build sampled pulses for common pulse shapes
# Drive pulse parameters (us = microseconds)
drive_sigma_us = 0.075 # This determines the actual width of the gaussian
drive_samples_us = drive_sigma_us*8 # This is a truncating parameter, because gaussians don't have
# a natural finite length
drive_sigma = get_closest_multiple_of_16(drive_sigma_us * us /dt) # The width of the gaussian in units of dt
drive_samples = get_closest_multiple_of_16(drive_samples_us * us /dt) # The truncating parameter in units of dt
drive_amp = 0.3
# Drive pulse samples
drive_pulse = pulse_lib.gaussian(duration=drive_samples,
sigma=drive_sigma,
amp=drive_amp,
name='freq_sweep_excitation_pulse')
```
Next, we will create the instructions we need to measure our qubit. This actually consists of two pulses: one stimulates the readout with a Gaussian-Square pulse applied at the readout resonator frequency, and the other triggers the data acquisition instrument to acquire data for the duration of the pulse.
```
### Construct the measurement pulse
# Measurement pulse parameters
meas_samples_us = 3.0
meas_sigma_us = 0.014 # The width of the gaussian part of the rise and fall
meas_risefall_us = 0.1 # and the truncating parameter: how many samples to dedicate to the risefall
meas_samples = get_closest_multiple_of_16(meas_samples_us * us/dt)
meas_sigma = get_closest_multiple_of_16(meas_sigma_us * us/dt) # The width of the gaussian part in units of dt
meas_risefall = get_closest_multiple_of_16(meas_risefall_us * us/dt) # The truncating parameter in units of dt
meas_amp = 0.25
# Measurement pulse samples
meas_pulse = pulse_lib.gaussian_square(duration=meas_samples,
sigma=meas_sigma,
amp=meas_amp,
risefall=meas_risefall,
name='measurement_pulse')
### Construct the acquire pulse to trigger the acquisition
# Acquire pulse samples
acq_cmd = pulse.Acquire(duration=meas_samples)
```
We have to check one additional thing in order to properly measure our qubits: the measurement map. This is a hardware constraint. When acquisition is done for one qubit, it is also done on other qubits. We have to respect this constraint when building our program in OpenPulse. Let's check which group of qubits our qubit is in:
```
# Find out which group of qubits need to be acquired with this qubit
meas_map_idx = None
for i, measure_group in enumerate(backend_config.meas_map):
if qubit in measure_group:
meas_map_idx = i
break
assert meas_map_idx is not None, f"Couldn't find qubit {qubit} in the meas_map!"
```
Lastly, we specify the channels on which we will apply our pulses. Drive, measure, and acquire channels are indexed by qubit index.
```
### Collect the necessary channels
drive_chan = pulse.DriveChannel(qubit)
meas_chan = pulse.MeasureChannel(qubit)
acq_chan = pulse.AcquireChannel(qubit)
```
Now that the pulse parameters have been defined, and we have created the pulse shapes for our experiments, we can proceed to creating the pulse schedules.
At each frequency, we will send a drive pulse of that frequency to the qubit and measure immediately after the pulse. The pulse envelopes are independent of frequency, so we will build a reusable `schedule`, and we will specify the drive pulse frequency with a frequency configuration array.
```
# Create the base schedule
# Start with drive pulse acting on the drive channel
schedule = pulse.Schedule(name='Frequency sweep')
schedule += drive_pulse(drive_chan)
# In a new schedule, which we will use again later, add a measurement stimulus on the
# measure channel pulse to trigger readout
measure_schedule = meas_pulse(meas_chan)
# Trigger data acquisition, and store measured values into respective memory slots
measure_schedule += acq_cmd([pulse.AcquireChannel(i) for i in backend_config.meas_map[meas_map_idx]],
[pulse.MemorySlot(i) for i in backend_config.meas_map[meas_map_idx]])
# The left shift `<<` is special syntax meaning to shift the start time of the schedule by some duration
schedule += measure_schedule << schedule.duration
# Create the frequency settings for the sweep (MUST BE IN HZ)
frequencies_Hz = frequencies_GHz*GHz
schedule_frequencies = [{drive_chan: freq} for freq in frequencies_Hz]
```
As a sanity check, it's always a good idea to look at the pulse schedule. This is done using `schedule.draw()` as shown below.
```
schedule.draw(channels_to_plot=[drive_chan, meas_chan], label=True, scaling=1.0)
```
We assemble the `schedules` and `schedule_frequencies` above into a program object, called a Qobj, that can be sent to the quantum device. We request that each schedule (each point in our frequency sweep) is repeated `num_shots_per_frequency` times in order to get a good estimate of the qubit response.
We also specify measurement settings. `meas_level=0` returns raw data (an array of complex values per shot), `meas_level=1` returns kerneled data (one complex value per shot), and `meas_level=2` returns classified data (a 0 or 1 bit per shot). We choose `meas_level=1` to replicate what we would be working with if we were in the lab, and hadn't yet calibrated the discriminator to classify 0s and 1s. We ask for the `'avg'` of the results, rather than each shot individually.
```
from qiskit import assemble
num_shots_per_frequency = 1024
frequency_sweep_program = assemble(schedule,
backend=backend,
meas_level=1,
meas_return='avg',
shots=num_shots_per_frequency,
schedule_los=schedule_frequencies)
```
Finally, we can run the assembled program on the backend using:
```
job = backend.run(frequency_sweep_program)
```
It is always a good idea to print the `job_id` for later retrieval, and to monitor the job status by using `job_monitor()`
```
# print(job.job_id())
from qiskit.tools.monitor import job_monitor
job_monitor(job)
```
Once the job is run, the results can be retrieved using:
```
frequency_sweep_results = job.result(timeout=120) # timeout parameter set to 120 seconds
```
We will extract the results and plot them using `matplotlib`:
```
%matplotlib inline
import matplotlib.pyplot as plt
sweep_values = []
for i in range(len(frequency_sweep_results.results)):
# Get the results from the ith experiment
res = frequency_sweep_results.get_memory(i)
# Get the results for `qubit` from this experiment
sweep_values.append(res[qubit])
plt.scatter(frequencies_GHz, sweep_values, color='black') # plot real part of sweep values
plt.xlim([min(frequencies_GHz), max(frequencies_GHz)])
plt.xlabel("Frequency [GHz]")
plt.ylabel("Measured signal [a.u.]")
plt.show()
```
As you can see above, the peak near the center corresponds to the location of the qubit frequency. The signal shows power-broadening, which is a signature that we are able to drive the qubit off-resonance as we get close to the center frequency. To get the value of the peak frequency, we will fit the values to a resonance response curve, which is typically a Lorentzian shape.
```
from scipy.optimize import curve_fit
def fit_function(x_values, y_values, function, init_params):
fitparams, conv = curve_fit(function, x_values, y_values, init_params)
y_fit = function(x_values, *fitparams)
return fitparams, y_fit
fit_params, y_fit = fit_function(frequencies_GHz,
sweep_values,
lambda x, A, q_freq, B, C: (A / np.pi) * (B / ((x - q_freq)**2 + B**2)) + C,
[-2e10, 4.975, 1, 3e10] # initial parameters for curve_fit
)
plt.scatter(frequencies_GHz, sweep_values, color='black')
plt.plot(frequencies_GHz, y_fit, color='red')
plt.xlim([min(frequencies_GHz), max(frequencies_GHz)])
plt.xlabel("Frequency [GHz]")
plt.ylabel("Measured Signal [a.u.]")
plt.show()
A, rough_qubit_frequency, B, C = fit_params
rough_qubit_frequency = rough_qubit_frequency*GHz # make sure qubit freq is in Hz
print(f"We've updated our qubit frequency estimate from "
f"{round(backend_defaults.qubit_freq_est[qubit] / GHz, 5)} GHz to {round(rough_qubit_frequency/GHz, 5)} GHz.")
```
# Part 2. Calibrating and using a $\pi$ pulse
## A. Calibrating $\pi$ pulses using a Rabi experiment <a id="rabi"></a>
Once we know the frequency of our qubit, the next step is to determine the strength of a $\pi$ pulse. Strictly speaking of the qubit as a two-level system, a $\pi$ pulse is one that takes the qubit from $\vert0\rangle$ to $\vert1\rangle$, and vice versa. This is also called the $X$ or $X180$ gate, or bit-flip operator. We already know the microwave frequency needed to drive this transition from the previous frequency sweep experiment, and we now seek the amplitude needed to achieve a $\pi$ rotation from $\vert0\rangle$ to $\vert1\rangle$. The desired rotation is shown on the Bloch sphere in the figure below -- you can see that the $\pi$ pulse gets its name from the angle it sweeps over on a Bloch sphere.
<img src="https://github.com/aasfaw/qiskit-intros/blob/master/zero_to_one_X180.png?raw=true">
We will change the drive amplitude in small increments and measuring the state of the qubit each time. We expect to see oscillations which are commonly named Rabi oscillations, as the qubit goes from $\vert0\rangle$ to $\vert1\rangle$ and back.
```
# This experiment uses these values from the previous experiment:
# `qubit`,
# `measure_schedule`, and
# `rough_qubit_frequency`.
# Rabi experiment parameters
num_rabi_points = 50
# Drive amplitude values to iterate over: 50 amplitudes evenly spaced from 0 to 0.75
drive_amp_min = 0
drive_amp_max = 0.75
drive_amps = np.linspace(drive_amp_min, drive_amp_max, num_rabi_points)
# Build the Rabi experiments:
# A drive pulse at the qubit frequency, followed by a measurement,
# where we vary the drive amplitude each time.
rabi_schedules = []
for drive_amp in drive_amps:
rabi_pulse = pulse_lib.gaussian(duration=drive_samples, amp=drive_amp,
sigma=drive_sigma, name=f"Rabi drive amplitude = {drive_amp}")
this_schedule = pulse.Schedule(name=f"Rabi drive amplitude = {drive_amp}")
this_schedule += rabi_pulse(drive_chan)
# Reuse the measure_schedule from the frequency sweep experiment
this_schedule += measure_schedule << this_schedule.duration
rabi_schedules.append(this_schedule)
```
The schedule will look essentially the same as the frequency sweep experiment. The only difference is that we are running a set of experiments which vary the amplitude of the drive pulse, rather than its modulation frequency.
```
rabi_schedules[-1].draw(channels_to_plot=[drive_chan, meas_chan], label=True, scaling=1.0)
# Assemble the schedules into a Qobj
num_shots_per_point = 1024
rabi_experiment_program = assemble(rabi_schedules,
backend=backend,
meas_level=1,
meas_return='avg',
shots=num_shots_per_point,
schedule_los=[{drive_chan: rough_qubit_frequency}]
* num_rabi_points)
# print(job.job_id())
job = backend.run(rabi_experiment_program)
job_monitor(job)
rabi_results = job.result(timeout=120)
```
Now that we have our results, we will extract them and fit them to a sinusoidal curve. For the range of drive amplitudes we selected, we expect that we will rotate the qubit several times completely around the Bloch sphere, starting from $|0\rangle$. The amplitude of this sinusoid tells us the fraction of the shots at that Rabi drive amplitude which yielded the $|1\rangle$ state. We want to find the drive amplitude needed for the signal to oscillate from a maximum (all $|0\rangle$ state) to a minimum (all $|1\rangle$ state) -- this gives the calibrated amplitude that enacts a $\pi$ pulse.
```
# center data around 0
def baseline_remove(values):
return np.array(values) - np.mean(values)
rabi_values = []
for i in range(num_rabi_points):
# Get the results for `qubit` from the ith experiment
rabi_values.append(rabi_results.get_memory(i)[qubit])
rabi_values = np.real(baseline_remove(rabi_values))
plt.xlabel("Drive amp [a.u.]")
plt.ylabel("Measured signal [a.u.]")
plt.scatter(drive_amps, rabi_values, color='black') # plot real part of Rabi values
plt.show()
fit_params, y_fit = fit_function(drive_amps,
rabi_values,
lambda x, A, B, drive_period, phi: (A*np.cos(2*np.pi*x/drive_period - phi) + B),
[1.5e10, 0.1e10, 0.5, 0])
plt.scatter(drive_amps, rabi_values, color='black')
plt.plot(drive_amps, y_fit, color='red')
drive_period = fit_params[2] # get period of rabi oscillation
plt.axvline(drive_period/2, color='red', linestyle='--')
plt.axvline(drive_period, color='red', linestyle='--')
plt.annotate("", xy=(drive_period, 0), xytext=(drive_period/2,0), arrowprops=dict(arrowstyle="<->", color='red'))
plt.annotate("$\pi$", xy=(drive_period/2-0.03, 0.1e10), color='red')
plt.xlabel("Drive amp [a.u.]", fontsize=15)
plt.ylabel("Measured signal [a.u.]", fontsize=15)
plt.show()
pi_amp = abs(drive_period / 2)
print(f"Pi Amplitude = {pi_amp}")
```
### Our $\pi$ pulse!
Let's define our pulse, with the amplitude we just found, so we can use it in later experiments.
```
pi_pulse = pulse_lib.gaussian(duration=drive_samples,
amp=pi_amp,
sigma=drive_sigma,
name='pi_pulse')
```
## B. Determining 0 vs 1 <a id="zerovone"></a>
Once our $\pi$ pulses have been calibrated, we can now create the state $\vert1\rangle$ with good probability. We can use this to find out what the states $\vert0\rangle$ and $\vert1\rangle$ look like in our measurements, by repeatedly preparing them and plotting the measured signal. This is what we use to build a discriminator, which is simply a function which takes a measured and kerneled complex value (`meas_level=1`) and classifies it as a 0 or a 1 (`meas_level=2`).
```
# Create two schedules
# Ground state schedule
gnd_schedule = pulse.Schedule(name="ground state")
gnd_schedule += measure_schedule
# Excited state schedule
exc_schedule = pulse.Schedule(name="excited state")
exc_schedule += pi_pulse(drive_chan) # We found this in Part 2A above
exc_schedule += measure_schedule << exc_schedule.duration
gnd_schedule.draw(channels_to_plot=[drive_chan, meas_chan], label=True, scaling=1.0)
exc_schedule.draw(channels_to_plot=[drive_chan, meas_chan], label=True, scaling=1.0)
```
We assemble the ground and excited state preparation schedules into one Qobj. Each of these will run `num_shots` times. We choose `meas_level=1` this time, because we do not want the results already classified for us as $|0\rangle$ or $|1\rangle$. Instead, we want kerneled data: raw acquired data that has gone through a kernel function to yield a single complex value for each shot. (You can think of a kernel as a dot product applied to the raw measurement data.)
We pass the same frequency for both schedules, although it is only used by the `exc_schedule`.
```
# Execution settings
num_shots = 1024
gnd_exc_program = assemble([gnd_schedule, exc_schedule],
backend=backend,
meas_level=1,
meas_return='single',
shots=num_shots,
schedule_los=[{drive_chan: rough_qubit_frequency}] * 2)
# print(job.job_id())
job = backend.run(gnd_exc_program)
job_monitor(job)
gnd_exc_results = job.result(timeout=120)
```
Now that we have the results, we can visualize the two populations which we have prepared on a simple scatter plot, showing results from the ground state program in blue and results from the excited state preparation program in red.
```
gnd_results = gnd_exc_results.get_memory(0)[:, qubit]
exc_results = gnd_exc_results.get_memory(1)[:, qubit]
plt.figure(figsize=[4,4])
# Plot all the results
# All results from the gnd_schedule are plotted in blue
plt.scatter(np.real(gnd_results), np.imag(gnd_results),
s=5, cmap='viridis', c='blue', alpha=0.5, label='state_0')
# All results from the exc_schedule are plotted in red
plt.scatter(np.real(exc_results), np.imag(exc_results),
s=5, cmap='viridis', c='red', alpha=0.5, label='state_1')
# Plot a large dot for the average result of the 0 and 1 states.
mean_gnd = np.mean(gnd_results) # takes mean of both real and imaginary parts
mean_exc = np.mean(exc_results)
plt.scatter(np.real(mean_gnd), np.imag(mean_gnd),
s=200, cmap='viridis', c='black',alpha=1.0, label='state_0_mean')
plt.scatter(np.real(mean_exc), np.imag(mean_exc),
s=200, cmap='viridis', c='black',alpha=1.0, label='state_1_mean')
plt.ylabel('I [a.u.]', fontsize=15)
plt.xlabel('Q [a.u.]', fontsize=15)
plt.title("0-1 discrimination", fontsize=15)
plt.show()
```
We can clearly see that the two populations of $|0\rangle$ and $|1\rangle$ form their own clusters. Kerneled measurement results (from `meas_level=1`) are classified (into `meas_level=2`) by applying a discriminator which optimally separates these two clusters. Optimal separation is simply a line in the IQ plane, equidistant from the average results we plotted above in the large dot, and normal to the line connecting the two dots.
We can set up a quick classifier function by returning 0 if a given point is closer to the mean of the ground state results, and returning 1 if the point is closer to the average excited state results.
```
import math
def classify(point: complex):
"""Classify the given state as |0> or |1>."""
def distance(a, b):
return math.sqrt((np.real(a) - np.real(b))**2 + (np.imag(a) - np.imag(b))**2)
return int(distance(point, mean_exc) < distance(point, mean_gnd))
```
## C. Measuring $T_1$ using inversion recovery <a id="T1"></a>
The $T_1$ time of a qubit is the time it takes for a qubit to decay from the excited state to the ground state. It is important because it limits the duration of meaningful programs we can run on the quantum computer.
Measuring $T_1$ is similar to our previous experiments, and uses the $\pi$ pulse we've calibrated. We again apply a single drive pulse, our $\pi$ pulse, then apply a measure pulse. However, this time we do not apply the measurement immediately. We insert a delay, and vary that delay between experiments. When we plot the measured signal against delay time, we will see a signal that decays exponentially as the qubit relaxes in energy. The decay time is the $T_1$, or relaxation time, of the qubit!
```
# T1 experiment parameters
time_max_us = 450
time_step_us = 6
times_us = np.arange(1, time_max_us, time_step_us)
# Convert to units of dt
delay_times_dt = times_us * us / dt
# We will use the same `pi_pulse` and qubit frequency that we calibrated and used before
# Create schedules for the experiment
t1_schedules = []
for delay in delay_times_dt:
this_schedule = pulse.Schedule(name=f"T1 delay = {delay * dt/us} us")
this_schedule += pi_pulse(drive_chan)
this_schedule |= measure_schedule << int(delay)
t1_schedules.append(this_schedule)
```
We can check out our $T_1$ schedule, too. To really get a sense of this experiment, try looking at a couple of the schedules by running the next cell multiple times, with different values of `sched_idx`. You will see the measurement pulse start later as you increase `sched_idx`.
```
sched_idx = 0
t1_schedules[sched_idx].draw(channels_to_plot=[drive_chan, meas_chan], label=True, scaling=1.0)
# Execution settings
num_shots = 256
t1_experiment = assemble(t1_schedules,
backend=backend,
meas_level=1,
meas_return='avg',
shots=num_shots,
schedule_los=[{drive_chan: rough_qubit_frequency}] * len(t1_schedules))
job = backend.run(t1_experiment)
# print(job.job_id())
job_monitor(job)
t1_results = job.result(timeout=120)
t1_values = []
for i in range(len(times_us)):
t1_values.append(t1_results.get_memory(i)[qubit])
plt.scatter(times_us, t1_values, color='black')
plt.title("$T_1$ Experiment", fontsize=15)
plt.xlabel('Delay before measurement [$\mu$s]', fontsize=15)
plt.ylabel('Signal [a.u.]', fontsize=15)
plt.show()
```
We can then fit the data to a decaying exponential, giving us T1!
```
# Fit the data
fit_params, y_fit = fit_function(times_us, t1_values,
lambda x, A, C, T1: (A * np.exp(-x / T1) + C),
[-3e10, 3e10, 100]
)
_, _, T1 = fit_params
plt.scatter(times_us, t1_values, color='black')
plt.plot(times_us, y_fit, color='red', label=f"T1 = {T1:.2f} us")
plt.xlim(0, np.max(times_us))
plt.title("$T_1$ Experiment", fontsize=15)
plt.xlabel('Delay before measurement [$\mu$s]', fontsize=15)
plt.ylabel('Signal [a.u.]', fontsize=15)
plt.legend()
plt.show()
```
# Part 3. Determining qubit coherence
## A. Measuring the qubit frequency precisely using a Ramsey experiment <a id="ramsey"></a>
Now, we determine the qubit frequency to better precision. This is done using a Ramsey pulse sequence. In this pulse sequence, we first apply a $\pi/2$ ("pi over two") pulse, wait some time $\Delta t$, and then apply another $\pi/2$ pulse. Since we are measuring the signal from the qubit at the same frequency as the pulses, we should observe oscillations at the difference in frequency between the applied pulses and the qubit.
<img src="https://github.com/aasfaw/qiskit-intros/blob/master/dephasing.png?raw=true">
```
# Ramsey experiment parameters
time_max_us = 1.8
time_step_us = 0.025
times_us = np.arange(0.1, time_max_us, time_step_us)
# Convert to units of dt
delay_times_dt = times_us * us / dt
# Drive parameters
# The drive amplitude for pi/2 is simply half the amplitude of the pi pulse
drive_amp = pi_amp / 2
# x_90 is a concise way to say pi_over_2; i.e., an X rotation of 90 degrees
x90_pulse = pulse_lib.gaussian(duration=drive_samples,
amp=drive_amp,
sigma=drive_sigma,
name='x90_pulse')
# create schedules for Ramsey experiment
ramsey_schedules = []
for delay in delay_times_dt:
this_schedule = pulse.Schedule(name=f"Ramsey delay = {delay * dt / us} us")
this_schedule |= x90_pulse(drive_chan)
this_schedule |= x90_pulse(drive_chan) << int(this_schedule.duration + delay)
this_schedule |= measure_schedule << int(this_schedule.duration)
ramsey_schedules.append(this_schedule)
```
Just like for $T_1$ schedules, it will be illuminating to execute the next cell multiple times to inspect a few of the schedules we've made. As you look at increasing indices of `ramsey_schedules`, the delay between the two $\pi/2$ pulses will increase.
```
ramsey_schedules[0].draw(channels_to_plot=[drive_chan, meas_chan], label=True, scaling=1.0)
```
Here, we will apply a commonly used experimental trick. We will drive the pulses off-resonance by a known amount, which we will call `detuning_MHz`. The measured Ramsey signal should show oscillations with frequency near `detuning_MHz`, with a small offset. This small offset is exactly how far away `rough_qubit_frequency` was from the qubit frequency.
```
# Execution settings
num_shots = 256
detuning_MHz = 2
ramsey_frequency = round(rough_qubit_frequency + detuning_MHz * MHz, 6) # need ramsey freq in Hz
ramsey_program = assemble(ramsey_schedules,
backend=backend,
meas_level=1,
meas_return='avg',
shots=num_shots,
schedule_los=[{drive_chan: ramsey_frequency}]*len(ramsey_schedules)
)
job = backend.run(ramsey_program)
# print(job.job_id())
job_monitor(job)
ramsey_results = job.result(timeout=120)
ramsey_values = []
for i in range(len(times_us)):
ramsey_values.append(ramsey_results.get_memory(i)[qubit])
plt.scatter(times_us, ramsey_values, color='black')
plt.xlim(0, np.max(times_us))
plt.title("Ramsey Experiment", fontsize=15)
plt.xlabel('Delay between X90 pulses [$\mu$s]', fontsize=15)
plt.ylabel('Measured Signal [a.u.]', fontsize=15)
plt.show()
```
We will fit the data to a sinusoid, and extract the information we are interested in -- namely, $\Delta f$.
```
fit_params, y_fit = fit_function(times_us, ramsey_values,
lambda x, A, del_f_MHz, C, B: (
A * np.cos(2*np.pi*del_f_MHz*x - C) + B
),
[2e10, 1./0.4, 0, 0.25e10]
)
# Off-resonance component
_, del_f_MHz, _, _, = fit_params # freq is MHz since times in us
plt.scatter(times_us, ramsey_values, color='black')
plt.plot(times_us, y_fit, color='red', label=f"df = {del_f_MHz:.2f} MHz")
plt.xlim(0, np.max(times_us))
plt.xlabel('Delay between X90 pulses [$\mu$s]', fontsize=15)
plt.ylabel('Measured Signal [a.u.]', fontsize=15)
plt.title('Ramsey Experiment', fontsize=15)
plt.legend()
plt.show()
```
Now that we know `del_f_MHz`, we can update our estimate of the qubit frequency.
```
precise_qubit_freq = rough_qubit_frequency + (del_f_MHz - detuning_MHz) * MHz # get new freq in Hz
print(f"Our updated qubit frequency is now {round(precise_qubit_freq/GHz, 6)} GHz. "
f"It used to be {round(rough_qubit_frequency / GHz, 6)} GHz")
```
## B. Measuring $T_2$ using Hahn echoes <a id="hahn"></a>
Next, we can measure the coherence time, $T_2$, of our qubit. The pulse sequence used to do this experiment is known as a Hahn echo, a term that comes from the NMR community. A Hahn echo experiment is very similar to the Ramsey experiment above, with an additional $\pi$ pulse between the two $\pi/2$ pulses. The $\pi$ pulse at time $\tau$ reverses the accumulation of phase, and results in an echo at time $2\tau$, where we apply the last $\pi/2$ pulse to do our measurement.
The decay time for the Hahn echo experiment gives us the coherence time, $T_2$.
```
# T2 experiment parameters
tau_max_us = 200
tau_step_us = 4
taus_us = np.arange(2, tau_max_us, tau_step_us)
# Convert to units of dt
delay_times_dt = taus_us * us / dt
# We will use the pi_pulse and x90_pulse from previous experiments
t2_schedules = []
for tau in delay_times_dt:
this_schedule = pulse.Schedule(name=f"T2 delay = {tau *dt/us} us")
this_schedule |= x90_pulse(drive_chan)
this_schedule |= pi_pulse(drive_chan) << int(this_schedule.duration + tau)
this_schedule |= x90_pulse(drive_chan) << int(this_schedule.duration + tau)
this_schedule |= measure_schedule << int(this_schedule.duration)
t2_schedules.append(this_schedule)
t2_schedules[0].draw(channels_to_plot=[drive_chan, meas_chan], label=True, scaling=1.0)
# Execution settings
num_shots_per_point = 512
t2_experiment = assemble(t2_schedules,
backend=backend,
meas_level=1,
meas_return='avg',
shots=num_shots_per_point,
schedule_los=[{drive_chan: precise_qubit_freq}]
* len(t2_schedules))
job = backend.run(t2_experiment)
# print(job.job_id())
job_monitor(job)
t2_results = job.result(timeout=120)
t2_values = []
for i in range(len(taus_us)):
t2_values.append(t2_results.get_memory(i)[qubit])
plt.scatter(2*taus_us, t2_values, color='black')
plt.xlabel('Delay between X90 pulse and $\pi$ pulse [$\mu$s]', fontsize=15)
plt.ylabel('Measured Signal [a.u.]', fontsize=15)
plt.title('Hahn Echo Experiment', fontsize=15)
plt.show()
fit_params, y_fit = fit_function(2*taus_us, t2_values,
lambda x, A, B, T2: (A * np.exp(-x / T2) + B),
[-1.2e15, -2.4e15, 20])
_, _, T2 = fit_params
print()
plt.scatter(2*taus_us, t2_values, color='black')
plt.plot(2*taus_us, y_fit, color='red', label=f"T2 = {T2:.2f} us")
plt.xlim(0, np.max(2*taus_us))
plt.xlabel('Delay between X90 pulse and $\pi$ pulse [$\mu$s]', fontsize=15)
plt.ylabel('Measured Signal [a.u.]', fontsize=15)
plt.title('Hahn Echo Experiment', fontsize=15)
plt.legend()
plt.show()
```
## C. Dynamical decoupling <a id="DD"></a>
A single $\pi$ pulse is able to eliminate quasi-static noise due to the reversal of phase accumulation. This concept can be extended to noise that cannot be approximated as quasi-static by applying several $\pi$ pulses in succession. This technique, commonly known as dynamical decoupling, allows us to cancel different frequencies of noise and is used to extract longer coherence times from qubits.
```
# DD experiment parameters
tau_us_min = 1
tau_us_max = 40
tau_step_us = 1.5
taus_us = np.arange(tau_us_min, tau_us_max, tau_step_us)
# Convert to units of dt
taus_dt = taus_us * us / dt
num_pi_pulses = 6 # apply two pi pulses
print(f"Total time ranges from {2.*num_pi_pulses*taus_us[0]} to {2.*num_pi_pulses*taus_us[-1]} us")
T2DD_schedules = []
for delay in taus_dt:
this_schedule = pulse.Schedule(name=f"T2DD delay = {delay * dt/us} us")
this_schedule |= x90_pulse(drive_chan)
this_schedule |= pi_pulse(drive_chan) << int(this_schedule.duration + delay)
for _ in range(num_pi_pulses - 1):
this_schedule |= pi_pulse(drive_chan) << int(this_schedule.duration + 2*delay)
this_schedule |= x90_pulse(drive_chan) << int(this_schedule.duration + delay)
this_schedule |= measure_schedule << int(this_schedule.duration)
T2DD_schedules.append(this_schedule)
T2DD_schedules[0].draw(channels_to_plot=[drive_chan, meas_chan], label=True, scaling=1.0)
num_shots_per_point = 1024
T2DD_experiment = assemble(T2DD_schedules,
backend=backend,
meas_level=1,
meas_return='avg',
shots=num_shots_per_point,
schedule_los=[{drive_chan: precise_qubit_freq}]
* len(T2DD_schedules))
job = backend.run(T2DD_experiment)
# print(job.job_id())
job_monitor(job)
T2DD_results = job.result(timeout=120)
times_us = 2.*num_pi_pulses*taus_us
DD_values = []
for i in range(len(taus_us)):
DD_values.append(T2DD_results.get_memory(i)[qubit])
plt.scatter(times_us, DD_values, color='black')
plt.xlim(0, np.max(times_us))
plt.xlabel('Total time before measurement [$\mu$s]', fontsize=15)
plt.ylabel('Measured Signal [a.u.]', fontsize=15)
plt.title('Dynamical Decoupling Experiment', fontsize=15)
plt.show()
# Fit the data
fit_func = lambda x, A, B, T2DD: (A * np.exp(-x / T2DD) + B)
fitparams, conv = curve_fit(fit_func, times_us, DD_values, [1.7e10, 0.8e10, 150])
_, _, T2DD = fitparams
plt.scatter(times_us, DD_values, color='black')
plt.plot(times_us, fit_func(times_us, *fitparams), color='red', label=f"T2DD = {T2DD:.2f} us")
plt.xlim([0, np.max(times_us)])
plt.xlabel('Total time before measurement [$\mu$s]', fontsize=15)
plt.ylabel('Measured Signal [a.u.]', fontsize=15)
plt.title('Dynamical Decoupling Experiment', fontsize=15)
plt.legend()
plt.show()
```
# Part 4. References <a id="refs"></a>
1. H. Abraham, I. Y. Akhalwaya, G. Aleksandrowicz, T. Alexander, G. Alexandrowics, E. Arbel, A. Asfaw, C. Azaustre, P. Barkoutsos, G. Barron, L. Bello, Y. Ben-Haim, L. S. Bishop, S. Bosch, D. Bucher, CZ, F. Cabrera, P. Calpin, L. Capelluto, J. Carballo, C.-F. Chen, A. Chen, R. Chen, J. M. Chow, C. Claus, A. W. Cross, A. J. Cross, J. Cruz- Benito, C. Culver, A. D. C ́orcoles-Gonzales, S. Dague, M. Dartiailh, A. R. Davila, D. Ding, E. Dumitrescu, K. Dumon, I. Duran, P. Eendebak, D. Egger, M. Everitt, P. M. Fern ́andez, A. Frisch, A. Fuhrer, J. Gacon, Gadi, B. G. Gago, J. M. Gambetta, L. Garcia, S. Garion, Gawel-Kus, L. Gil, J. Gomez-Mosquera, S. de la Puente Gonz ́alez, D. Green- berg,J.A.Gunnels,I.Haide,I.Hamamura,V.Havlicek,J.Hellmers,L.Herok,H.Horii, C. Howington, W. Hu, S. Hu, H. Imai, T. Imamichi, R. Iten, T. Itoko, A. Javadi-Abhari, Jessica, K. Johns, N. Kanazawa, A. Karazeev, P. Kassebaum, V. Krishnan, K. Kr- sulich, G. Kus, R. LaRose, R. Lambert, J. Latone, S. Lawrence, P. Liu, P. B. Z. Mac, Y. Maeng, A. Malyshev, J. Marecek, M. Marques, D. Mathews, A. Matsuo, D. T. Mc- Clure, C. McGarry, D. McKay, S. Meesala, A. Mezzacapo, R. Midha, Z. Minev, P. Mu- rali, J. Mu ̈ggenburg, D. Nadlinger, G. Nannicini, P. Nation, Y. Naveh, Nick-Singstock, P. Niroula, H. Norlen, L. J. O’Riordan, S. Oud, D. Padilha, H. Paik, S. Perriello, A. Phan, M. Pistoia, A. Pozas-iKerstjens, V. Prutyanov, J. P ́erez, Quintiii, R. Raymond, R. M.-C. Redondo, M. Reuter, D. M. Rodr ́ıguez, M. Ryu, M. Sandberg, N. Sathaye, B. Schmitt, C. Schnabel, T. L. Scholten, E. Schoute, I. F. Sertage, Y. Shi, A. Silva, Y. Siraichi, S. Sivarajah, J. A. Smolin, M. Soeken, D. Steenken, M. Stypulkoski, H. Takahashi, C. Taylor, P. Taylour, S. Thomas, M. Tillet, M. Tod, E. de la Torre, K. Trabing, M. Treinish, TrishaPe, W. Turner, Y. Vaknin, C. R. Valcarce, F. Varchon, D. Vogt- Lee, C. Vuillot, J. Weaver, R. Wieczorek, J. A. Wildstrom, R. Wille, E. Winston, J. J. Woehr, S. Woerner, R. Woo, C. J. Wood, R. Wood, S. Wood, J. Wootton, D. Yeralin, J. Yu, L. Zdanski, Zoufalc, azulehner, drholmie, fanizzamarco, kanejess, klinvill, merav aharoni, ordmoj, tigerjack, yang.luh, and yotamvakninibm, “Qiskit: An open-source framework for quantum computing,” 2019.
2. D. C. McKay, T. Alexander, L. Bello, M. J. Biercuk, L. Bishop, J. Chen, J. M. Chow, A. D. C ́orcoles, D. Egger, S. Filipp, J. Gomez, M. Hush, A. Javadi-Abhari, D. Moreda, P. Nation, B. Paulovicks, E. Winston, C. J. Wood, J. Wootton, and J. M. Gambetta, “Qiskit backend specifications for OpenQASM and OpenPulse experiments,” 2018.
```
import qiskit.tools.jupyter
%qiskit_version_table
```
| github_jupyter |
## Note-level dataset generation
This notebook uses raw data from the MusicNet dataset to set up sequential numpy arrays suitable for training deep neural networks.
**Before running:** Make sure to run the "Levels Computation" notebook to produce the numpy array files with global audio levels.
```
#### START HERE ####
dataFolder = 'data/' # make sure the path to data folder is correct
num_folds = 1
```
### Computing Note Information
```
import os
import json
import re
import numpy as np
import pandas as pd
from IPython.display import clear_output
import expression_modeling as m
def preprocess(labelsDir, csv, outfile=None, include_header=False, include_transp=True):
# load the symbolic information from the dataset
notearray = np.genfromtxt(os.path.join(labelsDir, csv + '.csv'), delimiter=',', names=True, dtype=['i', 'i', 'i', 'i', 'f', 'f', '|U40'])
# sort by score time first for correct parsing
notearray.sort(order=['start_beat', 'start_time'])
# load levels (generated by "Levels computation" notebook)
levels = np.load(os.path.join(dataFolder, 'levels', csv + '_global_lvls.npy'))
piece = m.Piece(name=csv)
piece.dynMean = np.mean(levels)
piece.dynStd = np.std(levels)
piece.startTime = notearray['start_time'][0]
piece.startBeat = notearray['start_beat'][0]
piece.endTime = notearray['end_time'][-1]
piece.endBeat = notearray['start_beat'][-1] + notearray['end_beat'][-1]
piece.part = m.buildPart(notearray, (levels - piece.dynMean)/piece.dynStd, 44100)
df = []
if include_transp:
for tr in range(-3,4):
di = m.buildSimpleNoteDataframe(piece, transpose=tr)
di['transposition'] = tr
df.append(di)
else:
df = m.buildSimpleNoteDataframe(piece, transpose=0)
df['transposition'] = 0
df = pd.concat(df, ignore_index=True)
df['pieceId'] = int(csv)
if outfile is None:
outfile = open(os.path.join(dataFolder, csv + '.csv'), 'w+')
df.to_csv(outfile)
else:
df.to_csv(outfile, mode='a', header=include_header)
return outfile
# select pieces containing violin
csvfolder = os.path.join(dataFolder, 'musicnet', 'train_labels')
dataset = [csv for csv in os.listdir(csvfolder) if re.search(r'^(.*?,){2}\s*?41\s*?,(.*?,){3}', open(os.path.join(csvfolder, csv), 'r').read(), re.MULTILINE)]
with open(os.path.join(dataFolder, 'musicnet_violin_IE1.csv'), 'w+') as outfile:
header = True
for i,mvt in enumerate(dataset):
clear_output(wait=True)
print("Processing piece " + str(i+1) + '/' + str(len(dataset)))
preprocess(csvfolder, mvt[:-4], outfile, include_header=header)
header = False
```
## Sequence Generation (Features)
The following cells use the CSV file produced above to format the data into sequences of notes containing a pitch vocabulary and a set of musicologically relevant features about the note.
### Loading Note Information
```
import os
import numpy as np
import pandas as pd
# read csv
with open(os.path.join(dataFolder, 'musicnet_violin_IE1.csv'), 'r') as path:
df = pd.read_csv(path)
# Input Encoding I --> minimal, i.e.: no musicological info
df.drop(['Unnamed: 0'], axis=1, inplace=True)
instrs = set(df.instrument)
df['instrument'] = df['instrument'].astype(pd.CategoricalDtype(instrs))
print('initial size: ' + str(len(df)))
```
### Mapping vocabulary
```
import pickle
pitches = list(df.loc[:,['pitch']].itertuples(index=False, name=None))
# using mapping from maestro dataset (88 keys + 4 ctrl. words)
with open(os.path.join(dataFolder, 'mF_pitch_dict.data'), 'rb') as filehandle:
lex_to_ix = pickle.load(filehandle)
pitches = list(df.loc[:,['pitch']].itertuples(index=False, name=None))
df['pitch'] = [lex_to_ix.get(m, len(lex_to_ix)+1) for m in pitches]
```
### Picking Training / Validation / Test sets:
```
import os
import json
import random
random.seed(777)
csvfolder = os.path.join(dataFolder, 'musicnet', 'train_labels')
all = [csv[0:-4] for csv in os.listdir(csvfolder) if re.search(r'^(.*?,){2}\s*?41\s*?,(.*?,){3}', open(os.path.join(csvfolder, csv), 'r').read(), re.MULTILINE)]
folds = []
test_pieces = []
# reserve 10% for test
test_sz = int(len(all) / 10)
for i in range(test_sz):
m = all[random.randint(0, len(all) - 1)]
test_pieces.append(m)
all.remove(m)
print('Test set pieces: ' + str(test_pieces))
for i in range(num_folds):
train = all.copy()
val = []
# another 10% of what remains for validation
val_sz = int(len(train) / 10)
for j in range(val_sz):
m = train[random.randint(0, len(train) - 1)]
val.append(m)
train.remove(m)
folds.append((train, val))
print('Val. set IDs: ' + str([f[1] for f in folds]))
```
### Arranging data for sequential training and saving dataset
```
import pickle
def sequencer(df, one_hot_cols=None, include_transp=True):
sequences = []
maxLen = 0
# list the pieces
pieces = set(df.pieceId)
for p in pieces:
dp = df.loc[df.pieceId == p, :].copy()
transps = range(-3,4) if include_transp else [0]
for tr in transps:
d = dp.loc[dp.transposition == tr, :].copy()
maxLen = len(d) if len(d) > maxLen else maxLen
d.drop(['pieceId', 'transposition'], axis=1, inplace=True)
# convert categories to one-hot
if one_hot_cols:
for attrib in one_hot_cols:
d = pd.concat([d, pd.get_dummies(d[attrib], prefix=attrib)], axis=1)
d.drop([attrib], axis=1, inplace=True)
# instance standardization for relevant features
feats = ['localTempo', 'peakLevel', 'ioiRatio']
aux = d.loc[:, feats]
moments = np.zeros((aux.shape[1], 2))
moments[:, 0] = aux.mean().to_numpy()
moments[:, 1] = aux.std().to_numpy()
d.loc[:, feats] = (aux - moments[:,0])/ moments[:,1]
# add <END> token to sequence
end = pd.DataFrame(np.zeros((1,d.shape[1])), columns=d.columns)
end["pitch"] = len(lex_to_ix) + 2
d = d.append(end)
# add <SOS> token to sequence
start = pd.DataFrame(np.zeros((1,d.shape[1])), columns=d.columns)
start["pitch"] = len(lex_to_ix) + 3
d = pd.concat([start, d])
# separate output features
outCols = ['ioiRatio', 'timingDev', 'timingDevLocal', 'localTempo', 'peakLevel', 'startTime', 'durationSecs']
y = d.loc[:, outCols].copy()
d.drop(outCols, axis=1, inplace=True)
sequences.append(((d, y, moments), p, tr))
return sequences
def standardize(df, moments=None, cols=None):
if cols is None:
cols = (df.dtypes == 'float64')
nums = df.loc[:,cols]
if moments is None:
moments = np.zeros((nums.shape[1],2)) # output mean and std for reverting predictions
moments[:,0] = nums.mean().to_numpy()
moments[:,1] = nums.std().to_numpy()
df.loc[:, cols] = (nums - moments[:,0]) / moments[:,1]
return moments, cols
# Separate Training / Validation / Test:
test = df.loc[df.pieceId.isin(test_pieces), :].copy()
moments = None
cols = None
for i, (training_pieces, val_pieces) in enumerate(folds):
train = df.loc[df.pieceId.isin(training_pieces), :].copy()
val = df.loc[df.pieceId.isin(val_pieces), :].copy()
# Standardization
moments, cols = standardize(train, cols=['beatDiff', 'duration', 'ioi', 'startTime', 'durationSecs', 'timingDev', 'timingDevLocal'])
standardize(val, moments=moments, cols=cols)
with open(os.path.join(dataFolder, 'MNv_I_normalizer_fold_' + str(i) + '.data'), 'wb') as filehandle:
pickle.dump((moments, cols), filehandle)
train_seq = sequencer(train, one_hot_cols=['instrument'])
val_seq = sequencer(val, one_hot_cols=['instrument'], include_transp=False)
# Save arrays
print('Saving fold ' + str(i))
with open(os.path.join(dataFolder, 'MNv_I_train_sequences_fold_' + str(i) + '.data'), 'wb') as filehandle:
pickle.dump(train_seq, filehandle)
with open(os.path.join(dataFolder, 'MNv_I_val_sequences_fold_' + str(i) + '.data'), 'wb') as filehandle:
pickle.dump(val_seq, filehandle)
# Prepare test sequences
print('Saving test data')
standardize(test, moments=moments, cols=cols) # using last fold moments (it's good enough)
test_seq = sequencer(test, one_hot_cols=['instrument'], include_transp=False)
with open(os.path.join(dataFolder, 'MNv_I_test_sequences.data'), 'wb') as filehandle:
pickle.dump(test_seq, filehandle)
print('Finished.')
```
| github_jupyter |
```
class FooClass:...
def test_sep():...
# local variable
var = "lowercase"
# internal use
_var = "_single_leading_underscore"
# avoid conflicts with Python keyword
var_ = "single_trailing_underscore_"
# a class attribute (private use in class)
__var = " __double_leading_underscore"
# "magic" objects or attributes, ex: __init__
__name__
# throwaway variable, ex: _, v = (1, 2)
_ = "throwaway"
print("Hello world!")
import __hello__
import __phello__
import __phello__.spam
import sys
print(sys.version)
sys.version_info >= (3, 7)
import platform
platform.python_version()
... == Ellipsis
type(...)
import random
num = random.randint(0, 10)
if num < 3:
print("less than 3")
elif num < 5:
print("less than 5")
else:
print(num)
a=[random.randint(1,50) for x in range(50)]
for _ in range(5):
pass
else:
print("no break")
sys.getsizeof(range(100000000))
sys.getsizeof(xrange(100000000))
n = 0
while n < 5:
if n == 3:
break
n += 1
else:
print("no break")
print ("test")
n=0
while True:
n += 1
if n == 5:
break
print(n)
try:
print("No exception")
except:
pass
else:
print("Success")
a = "hello"
#a[1]='t'
a=list(a)
a
a[0],a[4]= 'b', 'o'
''.join(a)
#lists
a = [1, 2, 3, 4, 5]
a[-1] # negative index
a[1:] # slicing
a[1:-1]
a[1:-1:2]
a[::-1] # reverse
a[0] = 0 # set an item
a.append(6) # append an item
del a[-1] # del an item
b = [x for x in range(3)] # list comprehension
a + b # add two lists
d = {'timmy': 'red', 'barry': 'green', 'guido': 'blue'}
d['timmy'] = "yellow" # set data
del d['guido'] # del data
'guido' in d # contain data
{k: v for k ,v in d.items()} # dict comprehension
d.keys() # list all keys
d.values() # list all values
def foo_with_doc():
"""Documentation String."""
def foo_with_arg(arg): ...
def foo_with_args(*arg): ...
def foo_with_kwarg(a, b="foo"): ...
def foo_with_args_kwargs(*args, **kwargs): ...
def foo_with_kwonly(a, b, *, k): ... # python3
def foo_with_annotations(a: int) -> int: ... # python3
def fib(n: int) -> int:
a,b = 0,1
for _ in range(n):
b,a = a+b,b
return a
fib(10)
def fib(n):
a,b =0,1
for _ in range(n):
yield a
b, a =a +b, b
for f in fib(20):
print(f)
def fibonacci(n):
yield from fib(n)
[f for f in fibonacci(10)]
class A: ...
class B: ...
class Foo(A, B):
"""A class document."""
foo = "class variable"
def __init__(self, v):
self.attr = v
self.__private = "private var"
@staticmethod
def bar_static_method(): ...
@classmethod
def bar_class_method(cls): ...
def bar(self):
"""A method document."""
def bar_with_arg(self, arg): ...
def bar_with_args(self, *args): ...
def bar_with_kwarg(self, kwarg="bar"): ...
def bar_with_args_kwargs(self, *args, **kwargs): ...
def bar_with_kwonly(self, *, k): ...
def bar_with_annotations(self, a: int): ...
'''import asyncio
async def http_ok(r, w):
head = b"HTTP/1.1 200 OK\r\n"
head += b"Content-Type: text/html\r\n"
head += b"\r\n"
body = b"<html>"
body += b"<body><h1>Hello world!</h1></body>"
body += b"</html>"
_ = await r.read(1024)
w.write(head + body)
await w.drain()
w.close()
async def main():
server = await asyncio.start_server(
http_ok, "127.0.0.1", 8888
)
async with server:
await server.serve_forever()
asyncio.run(main())
''''
py = '''
... def fib(n):
... a, b = 0, 1
... for _ in range(n):
... b, a = b + a, b
... return a
... print(fib(10))
... '''
>>> exec(py, globals(), locals())
```
| github_jupyter |
# Introduction to Python
## Introduction
### Why teach Python?
* In this first session, we will introduce [Python](http://www.python.org).
* This course is about programming for data analysis and visualisation in research.
* It's not mainly about Python.
* But we have to use some language.
### Why Python?
* Python is quick to program in
* Python is popular in research, and has lots of libraries for science
* Python interfaces well with faster languages
* Python is free, so you'll never have a problem getting hold of it, wherever you go.
### Why write programs for research?
* Not just labour saving
* Scripted research can be tested and reproduced
### Sensible Input - Reasonable Output
Programs are a rigorous way of describing data analysis for other researchers, as well as for computers.
Computational research suffers from people assuming each other's data manipulation is correct. By sharing codes,
which are much more easy for a non-author to understand than spreadsheets, we can avoid the "SIRO" problem. The old saw "Garbage in Garbage out" is not the real problem for science:
* Sensible input
* Reasonable output
## Many kinds of Python
### The Jupyter Notebook
The easiest way to get started using Python, and one of the best for research data work, is the Jupyter Notebook.
In the notebook, you can easily mix code with discussion and commentary, and mix code with the results of that code;
including graphs and other data visualisations.
```
### Make plot
%matplotlib inline
import math
import numpy as np
import matplotlib.pyplot as plt
theta = np.arange(0, 4 * math.pi, 0.1)
eight = plt.figure()
axes = eight.add_axes([0, 0, 1, 1])
axes.plot(0.5 * np.sin(theta), np.cos(theta / 2))
```
These notes are created using Jupyter notebooks and you may want to use it during the course. However, Jupyter notebooks won't be used for most of the activities and exercises done in class. To get hold of a copy of the notebook, follow the setup instructions shown on the course website, or use the installation in Desktop@UCL (available in the teaching cluster rooms or [anywhere](https://www.ucl.ac.uk/isd/services/computers/remote-access/desktopucl-anywhere)).
Jupyter notebooks consist of discussion cells, referred to as "markdown cells", and "code cells", which contain Python. This document has been created using Jupyter notebook, and this very cell is a **Markdown Cell**.
```
print("This cell is a code cell")
```
Code cell inputs are numbered, and show the output below.
Markdown cells contain text which uses a simple format to achive pretty layout,
for example, to obtain:
**bold**, *italic*
* Bullet
> Quote
We write:
**bold**, *italic*
* Bullet
> Quote
See the Markdown documentation at [This Hyperlink](http://daringfireball.net/projects/markdown/)
### Typing code in the notebook
When working with the notebook, you can either be in a cell, typing its contents, or outside cells, moving around the notebook.
* When in a cell, press escape to leave it. When moving around outside cells, press return to enter.
* Outside a cell:
* Use arrow keys to move around.
* Press `b` to add a new cell below the cursor.
* Press `m` to turn a cell from code mode to markdown mode.
* Press `shift`+`enter` to calculate the code in the block.
* Press `h` to see a list of useful keys in the notebook.
* Inside a cell:
* Press `tab` to suggest completions of variables. (Try it!)
*Supplementary material*: Learn more about [Jupyter notebooks](https://jupyter.org/).
### Python at the command line
Data science experts tend to use a "command line environment" to work. You'll be able to learn this at our ["Software Carpentry" workshops](http://rits.github-pages.ucl.ac.uk/software-carpentry/), which cover other skills for computationally based research.
```
%%bash
# Above line tells Python to execute this cell as *shell code*
# not Python, as if we were in a command line
# This is called a 'cell magic'
python -c "print(2 * 4)"
```
### Python scripts
Once you get good at programming, you'll want to be able to write your own full programs in Python, which work just
like any other program on your computer. Here are some examples:
```
%%bash
echo "print(2 * 4)" > eight.py
python eight.py
```
We can make the script directly executable (on Linux or Mac) by inserting a [hashbang](https://en.wikipedia.org/wiki/Shebang_(Unix%29) and [setting the permissions](http://v4.software-carpentry.org/shell/perm.html) to execute.
```
%%writefile fourteen.py
#! /usr/bin/env python
print(2 * 7)
%%bash
chmod u+x fourteen.py
./fourteen.py
```
### Python Libraries
We can write our own python libraries, called modules which we can import into the notebook and invoke:
```
%%writefile draw_eight.py
# Above line tells the notebook to treat the rest of this
# cell as content for a file on disk.
import math
import numpy as np
import matplotlib.pyplot as plt
def make_figure():
theta = np.arange(0, 4 * math.pi, 0.1)
eight = plt.figure()
axes = eight.add_axes([0, 0, 1, 1])
axes.plot(0.5 * np.sin(theta), np.cos(theta / 2))
return eight
```
In a real example, we could edit the file on disk
using a program such as [Atom](https://atom.io) or [VS code](https://code.visualstudio.com/).
```
import draw_eight # Load the library file we just wrote to disk
image = draw_eight.make_figure()
```
There is a huge variety of available packages to do pretty much anything. For instance, try `import antigravity`.
The `%%` at the beginning of a cell is called *magics*. There's a [large list of them available](https://ipython.readthedocs.io/en/stable/interactive/magics.html) and you can [create your own](http://ipython.readthedocs.io/en/stable/config/custommagics.html).
| github_jupyter |
```
## 9/12/17: this notebook subsets the relevant stuff from tf_sketchy.ipynb
## in order to compare triplet features to imagenet-only vgg
## on the image retrieval task
from __future__ import division
import numpy as np
from numpy import *
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn import svm
from sklearn import linear_model
import os
import PIL
from PIL import Image
import matplotlib.pyplot as plt
from matplotlib.path import Path
import matplotlib.patches as patches
from skimage import data, io, filters
import pandas as pd
# load in features for photos
path_to_features = 'sketchy/triplet_features'
photo_features = np.load(os.path.join(path_to_features,'photo_features.npy'))
F = photo_features
# read in filenames and generate pandas dataframe with object labels
_filenames = pd.read_csv(os.path.join(path_to_features,'photo_filenames.txt'),header=None,names=['filename'])
filenames = []
for i in range(len(_filenames)):
filenames.append(_filenames[_filenames.index==i].values[0][0])
filenames = ['sketchy' + f[1:] for f in filenames]
path = filenames
obj = [f.split('/')[3] for f in filenames]
img = [f.split('/')[4] for f in filenames]
data = {'path': path,
'object': obj,
'filename': img}
X = pd.DataFrame.from_dict(data)
# subset airplane features only
matches = X['object']=='airplane'
inds = np.where(matches==True)
X0 = X[matches]
F0 = F[inds]
# construct (11094,1024) version of photo feature matrix, called PF, that matches indexing of the sketch feature matrix
sketch_features = np.load('sketchy/airplane_features/airplane_sketch_features.npy')
_sketch_filenames = pd.read_csv('sketchy/airplane_features/airplane_sketch_filenames.txt',header=None,names=['filename'])
sketch_filenames = []
for i in range(len(_sketch_filenames)):
sketch_filenames.append(_sketch_filenames[_sketch_filenames.index==i].values[0][0])
PF = []
inds = []
for sf in sketch_filenames:
q = sf.split('/')[2]+'.jpg'
PF.append(F0[X0['filename']==q])
inds.append(np.where(X0['filename']==q)[0][0])
PF = np.squeeze(np.array(PF))
SF = sketch_features
inds = np.array(inds)
## zip together/concatenate the photo and sketch features
_F = np.hstack((PF,SF))
## get just complete sketches from each sketch folder
sketch_folders = np.unique([os.path.dirname(s) for s in sketch_filenames])
complete_paths = []
SF_complete = []
photos_complete = []
for (j,s) in enumerate(sketch_folders):
complete_sketch = str(max([int(i.split('.')[0]) for i \
in os.listdir(s)])) + '.png'
complete_paths.append(os.path.join(os.path.dirname(s),complete_sketch))
SF_complete.append(SF[j])
photos_complete.append(os.path.dirname(s).split('/')[-1])
SF_complete = np.array(SF_complete)
photos_complete = np.array(photos_complete)
## get image classification within airplane class
run_this = 1
FEAT = SF_complete
LABELS = photos_complete
if run_this:
# split sketch feature data for linear classification
X_train, X_test, y_train, y_test = train_test_split(
FEAT, LABELS, test_size=0.2, random_state=0)
# check dimensionality of split data
print 'dimensionality of train/test split'
print X_train.shape, y_train.shape
print X_test.shape, y_test.shape
print ' '
cval = True
if cval==False:
# compute linear classification accuracy (takes a minute or so to run)
clf = svm.SVC(kernel='linear', C=1).fit(X_train, y_train)
clf.score(X_test, y_test)
else:
# compute linear classification accuracy (takes several minutes to run)
# clf = svm.SVC(kernel='linear', C=1)
clf = linear_model.LogisticRegression(penalty='l2')
scores = cross_val_score(clf, FEAT, LABELS, cv=2)
print("Accuracy: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
## SVM Accuracy: 0.41 (+/- 0.08) with cv=5 measured on 6/26/17 on intermediate sketches
## softmax Accuracy: 0.43 (+/- 0.01) with cv=2 measured on 9/11/17
```
#### euclidean distances among sketches
```
from sklearn.metrics.pairwise import pairwise_distances
euc = pairwise_distances(SF_complete,metric='euclidean')
print euc.shape
p_ind = 4
fp = 20
fig = plt.figure(figsize=(9,9))
for (_i,p_ind) in enumerate(np.arange(fp,fp+9)):
unique_photos = np.unique(photos_complete)
inds = np.where(photos_complete==unique_photos[p_ind])[0]
start = inds[0]
stop = inds[-1]
# get within-photo sketch distances
within_block = euc[start:stop+1,start:stop+1]
assert len(within_block[np.triu_indices(len(within_block),k=1)])==(len(within_block)**2-len(within_block))/2
within_distances = within_block[np.triu_indices(len(within_block),k=1)]
# get between-photo sketch distances
all_inds = np.arange(len(photos_complete))
non_matches = [i for i in all_inds if i not in inds]
_non_matches_shuff = np.random.RandomState(seed=0).permutation(non_matches)
non_matches_shuff = _non_matches_shuff[:len(inds)]
btw_distances = euc[start:stop+1,non_matches_shuff].flatten()
# plot
plt.subplot(3,3,_i+1)
h = plt.hist(within_distances,bins=20,alpha=0.3)
h = plt.hist(btw_distances,bins=20,alpha=0.3)
plt.title(str(p_ind))
plt.show()
```
| github_jupyter |
# Training Image Classifier
We will use part of the training data provided to us, separated by high level [entity] clusters, to train the image classifier. Due to the scale of the full dataset, a random subsample is taken. See [this notebook block](http://localhost:8888/notebooks/02.train_tiered_classifiers.ipynb#Training-level-1---Image-classifier) for image classifier training.
# Compute evaluation image descriptors
When we get the evaluation image data, we must compute descriptors for that data. We will use the eval index (see the `common.descriptor_set.eval.json` config) and the common descriptor store (``common.descriptor_factory.json``). Using the ``common.cmd.eval.config.json`` with the ``compute_many_descriptors.py`` script should be used, which is set to these locations.
After descriptors are computed, we can proceed to scoring via the image classifier.
# Using image classifier for scoring
Here we will use the trained image classifer to score clustered ad images, pooling the maximum and average HT positive scores each ad and then cluster resulting in two score sets that we will "submit" for evaluation.
Output must be in the form of an **ordered** json-lines file with each line having the structure:
{"cluster_id": "...", "score": <float>}
Thus, we need the evaluation truth file in order to get the cluster ID ordering, which is also json-lines and of the form:
{"cluster_id": "...", "class": <int>}
...
The evaluation script (for plotting the ROC curve) can be [found here](https://github.com/istresearch/qpr-summer-2016-eval/tree/master/CP1).
The steps that need to be performed:
1. Get images + cluster/ad/sha CSV
- Compute descriptors for imagery provided
- Load cluster/ad/sha maps after knowing what images were successfully described
- Run classifier over descriptors computed
- Determine ad/cluster scores via max/avg pooling
- Output json-line files for scoring in evaluation script (linked above)
```
# Initialize logging
import logging
from smqtk.utils.bin_utils import initialize_logging
initialize_logging(logging.getLogger('smqtk'), logging.DEBUG)
initialize_logging(logging.getLogger('__name__'), logging.DEBUG)
# File path parameters
CMD_PROCESSED_CSV = 'eval.cmd.processed.csv'
CLUSTER_ADS_IMAGES_CSV = 'eval.clusters_ads_images.csv'
EVAL_IMAGE_CLASSIFICATIONS_CACHE = 'eval.image_classifications_cache.pickel'
OUTPUT_MAX_SCORE_JL = 'eval.cluster_scores.max_pool.jl'
OUTPUT_AVG_SCORE_JL = 'eval.cluster_scores.avg_pool.jl'
from smqtk.algorithms.classifier.libsvm import LibSvmClassifier
from smqtk.representation.classification_element.memory import MemoryClassificationElement
from smqtk.representation.classification_element.file import FileClassificationElement
from smqtk.representation import ClassificationElementFactory
image_classifier = LibSvmClassifier('image_classifier.train1.classifier.model',
'image_classifier.train1.classifier.label',
normalize=2)
c_file_factory = ClassificationElementFactory(FileClassificationElement,
{
"save_dir": "image_classifier.classifications",
"subdir_split": 10
})
from smqtk.representation import DescriptorSet
from smqtk.utils.plugin import from_plugin_config
with open('eval.test.cmd.json') as f:
descr_index = from_plugin_config(json.load(f)['descriptor_set'], DescriptorSet.get_impls())
descr_index.count() # should equal lines of eval.cmd .processed.csv
# TESTING
# Make up ground truth file from test-set clusters/ads/shas
test_pos_clusters = cPickle.load(open('test_pos_clusters.pickle'))
test_neg_clusters = cPickle.load(open('test_neg_clusters.pickle'))
pos_cluster2ads = cPickle.load(open('positive.cluster2ads.pickle'))
neg_cluster2ads = cPickle.load(open('negative.cluster2ads.pickle'))
pos_ad2shas = cPickle.load(open('positive.ad2shas.pickle'))
neg_ad2shas = cPickle.load(open('negative.ad2shas.pickle'))
with open('eval.test.clusters_ads_images.csv', 'w') as csv_out:
writer = csv.writer(csv_out)
writer.writerow(['cluster', 'ad', 'sha1'])
for c in test_pos_clusters:
for ad in pos_cluster2ads[c]:
for sha in pos_ad2shas[ad]:
writer.writerow([c, ad, sha])
for c in test_neg_clusters:
for ad in neg_cluster2ads[c]:
for sha in neg_ad2shas[ad]:
writer.writerow([c, ad, sha])
with open('eval.test.gt.jl', 'w') as f:
for c in sorted(test_pos_clusters | test_neg_clusters, key=lambda k: str(k)):
if c in test_pos_clusters:
f.write( json.dumps({'cluster_id': str(c), 'class': 1}) + '\n' )
elif c in test_neg_clusters:
f.write( json.dumps({'cluster_id': str(c), 'class': 0}) + '\n' )
else:
raise ValueError("Cluster %d not positive or negative?" % c)
# Step [3]
# Load in successfully processed image shas
# This is a result file from descriptor computation.
with open(CMD_PROCESSED_CSV) as f:
computed_shas = {r[1] for r in csv.reader(f)}
# Load cluster/ad/sha relationship maps, filtered by what was actually processed
import collections
cluster2ads = collections.defaultdict(set)
cluster2shas = collections.defaultdict(set)
ad2shas = collections.defaultdict(set)
sha2ads = collections.defaultdict(set)
with open(CLUSTER_ADS_IMAGES_CSV) as f:
reader = csv.reader(f)
for i, r in enumerate(reader):
if i == 0:
# skip header line
continue
c, ad, sha = r
if sha in computed_shas:
cluster2ads[c].add(ad)
cluster2shas[c].add(sha)
ad2shas[ad].add(sha)
sha2ads[sha].add(ad)
# Step [4]
# Classify eval set images
if os.path.isfile(EVAL_IMAGE_CLASSIFICATIONS_CACHE):
with open(EVAL_IMAGE_CLASSIFICATIONS_CACHE) as f:
image_descr2classifications = cPickle.load(f)
else:
img_descriptors = descr_index.get_many_descriptors(set(sha2ads))
image_descr2classifications = image_classifier.classify_async(img_descriptors,
c_file_factory,
use_multiprocessing=True,
ri=1.0)
with open(EVAL_IMAGE_CLASSIFICATIONS_CACHE, 'w') as f:
cPickle.dump(image_descr2classifications, f, -1)
# Step [5]
print "Collecting scores for SHA1s"
sha2score = {}
for c in image_descr2classifications.itervalues():
sha2score[c.uuid] = c['positive']
# select ads score from max and average of child image scores
print "Collecting scores for ads (MAX and AVG)"
import numpy
ad2score_max = {}
ad2score_avg = {}
for ad, child_shas in ad2shas.iteritems():
scores = [sha2score[sha] for sha in child_shas]
ad2score_max[ad] = numpy.max(scores)
ad2score_avg[ad] = numpy.average(scores)
# select cluster score from max and average of child ad scores
print "Collecting scores for ads (MAX and AVG)"
cluster2score_max = {}
cluster2score_avg = {}
for c, child_ads in cluster2ads.iteritems():
cluster2score_max[c] = numpy.max( [ad2score_max[ad] for ad in child_ads])
cluster2score_avg[c] = numpy.average([ad2score_avg[ad] for ad in child_ads])
len(cluster2score_max)
# Step [6]
# Write out json-lines file in same order as GT file
# The ordering we will save out json-lines (arbitrary?)
cluster_id_order = sorted(cluster2score_avg.iterkeys())
with open(OUTPUT_MAX_SCORE_JL, 'w') as f:
for c in cluster_id_order:
if c in cluster2score_max:
f.write( json.dumps({"cluster_id": c, "score": cluster2score_max[c]}) + '\n' )
else:
# Due to a cluster having no child ads with imagery
f.write( json.dumps({"cluster_id": c, "score": 0.5}) + '\n' )
with open(OUTPUT_AVG_SCORE_JL, 'w') as f:
for c in cluster_id_order:
if c in cluster2score_avg:
f.write( json.dumps({"cluster_id": c, "score": cluster2score_avg[c]}) + '\n' )
else:
# Due to a cluster having no child ads with imagery
f.write( json.dumps({"cluster_id": c, "score": 0.5}) + '\n' )
import numpy
numpy.average(sha2score.values()), numpy.min(sha2score.values()), numpy.max(sha2score.values())
```
| github_jupyter |
<h1><center><u>SAC -- 2D Navigation Robot(particle) Environment</u></center></h1>
```
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib notebook
# %matplotlib nbagg
%matplotlib qt
from robolearn.envs.simple_envs.goal_composition import GoalCompositionEnv
from robolearn.envs.normalized_box_env import NormalizedBoxEnv
from robolearn.torch.models import NNQFunction, NNVFunction
from robolearn.torch.policies import TanhGaussianPolicy
from robolearn.torch.rl_algos.sac.sac import SoftActorCritic
from robolearn.utils.data_management import SimpleReplayBuffer
from robolearn.utils.launchers.launcher_util import setup_logger
import robolearn.torch.pytorch_util as ptu
# Environment
env = GoalCompositionEnv(
goal_reward=50,
actuation_cost_coeff=0.5,
distance_cost_coeff=1.5,
log_distance_cost_coeff=10.0,
alpha=1e-3,
# Initial Condition
init_position=(-4., -4.),
init_sigma=1.50,
# Goal
goal_position=(5., 5.),
goal_threshold=0.05,
# Others
dynamics_sigma=0.1,
# horizon=PATH_LENGTH,
horizon=None,
)
# Normalize environment
env = NormalizedBoxEnv(
env,
normalize_obs=False,
online_normalization=False,
obs_mean=None,
obs_var=None,
obs_alpha=0.001,
)
# Visualize costs
env.reset()
env.render()
# env.close()
```
# IU-SAC
```
# Common parameters
n_unintentions = env.n_subgoals
obs_dim = np.prod(env.observation_space.shape)
action_dim = np.prod(env.action_space.shape)
# --------------------- #
# Value Function Models #
# --------------------- #
# Hyperparameters
net_size = 32
# Intentional Value function --> Environment goal (full task)
i_qf = NNQFunction(obs_dim=obs_dim,
action_dim=action_dim,
hidden_sizes=(net_size, net_size))
i_qf2 = NNQFunction(obs_dim=obs_dim,
action_dim=action_dim,
hidden_sizes=(net_size, net_size))
i_vf = NNVFunction(obs_dim=obs_dim,
hidden_sizes=(net_size, net_size))
# ------ #
# Policy #
# ------ #
# Hyperparameters
net_size = 32
n_unshared_layers = 2
policy = TanhGaussianPolicy(
obs_dim=obs_dim,
action_dim=action_dim,
hidden_sizes=[net_size for _ in range(n_unshared_layers)],
std=None,
reparameterize=True,
)
# --------- #
# Algorithm #
# --------- #
# Hypeparameters
render = False
reward_scale=1.0e-1
n_epochs = 100
replay_buffer_size = 1e3
batch_size=128
paths_per_epoch = 5
paths_per_eval = 3
path_length = 50
algo_hyperparams = dict(
# Common RL algorithm params
num_steps_per_epoch=paths_per_epoch * path_length,
num_epochs=n_epochs,
num_updates_per_train_call=1,
num_steps_per_eval=paths_per_eval * path_length,
# EnvSampler params
max_path_length=path_length,
render=render,
# SAC params
min_steps_start_train=batch_size,
min_start_eval=paths_per_epoch * path_length,
reparameterize=True,
action_prior='uniform',
entropy_scale=1.0e-0,
discount=0.99,
reward_scale=reward_scale,
)
# Logger
setup_logger('notebook_2d_nav',
variant=algo_hyperparams,
snapshot_mode='gap_and_last',
snapshot_gap=25,
log_dir=None,
log_stdout=False,
)
# Replay Buffer
replay_buffer = SimpleReplayBuffer(
max_replay_buffer_size=replay_buffer_size,
obs_dim=obs_dim,
action_dim=action_dim
)
algorithm = SoftActorCritic(
env=env,
policy=policy,
qf=i_qf,
vf=i_vf,
replay_buffer=replay_buffer,
batch_size=batch_size,
qf2=i_qf2,
eval_env=env,
save_environment=False,
**algo_hyperparams,
)
seed = 10
start_epoch = 0
env.seed(seed)
ptu.seed(seed)
algorithm.train(start_epoch=start_epoch)
# Test
%tb
deterministic = True
print('Max path length:', path_length)
env.close()
obs = env.reset()
for t in range(path_length):
env.render()
action, pol_info = policy.get_action(obs, deterministic=deterministic)
obs, reward, done, env_info = env.step(action)
# print('obs:', obs, '| goal:', env.wrapped_env.goal_position, ' | reward:', reward)
# print('---')
if done:
print('Environment done!')
break
```
| github_jupyter |
## Sensitivity analysis demonstration
This notebook contains an example of how to account for uncertainty in the parameters of the production process. The resulting variability is explored through a Monte Carlo-based sensitivity analysis, in which different values are used to run the facility and the outputs of interest are compiled.
First we set up the Facility to analyse, just as in previous demos.
```
import biopharma as bp
facility = bp.Facility(data_path='data')
# Define the steps needed to create our single product
from biopharma.process_steps import (
)
steps = [
]
product = bp.Product(facility, steps)
# Need to explicitly call this so that later functions can refer to quantities of interest
facility.load_parameters()
```
Once the facility is created, we can set up the sensitivity analysis. This requires two pieces of information:
* the aspects to be varied, and
* the outputs we are interested in.
The outputs are declared through selector functions, similar to how [optimisation targets are specified](Optimisation_demo.ipynb#optimisation_targets). In this example, we choose to track four outputs. Note that these outputs do not need to be related to the product: any parameter or output of a component can be tracked, by providing an appropriate selector function.
```
from biopharma import optimisation as opt
analyser = opt.SensitivityAnalyser(facility)
# Specify the variables whose sensitivity we are interested in.
analyser.add_output("CoG", component=opt.sel.product(0), item="cogs")
analyser.add_output("step_int_param", component=opt.sel.step('test_step'), item="int_param")
analyser.add_output("facility_info", component=opt.sel.facility(), item="param")
```
For each variable (uncertain aspect), we must say what distribution represents its possible values. There are several families of distributions available, each governed by appropriate parameters:
* Uniform (over a given domain)
* Triangular (over a given domain)
* Gaussian (with a given mean and variance)
Here, we choose two variables. Both are given uniform distributions.
```
# Specify which aspects to vary.
param1_mean = facility.products[0].parameters["param1"]
param1_width = 2 * param1_mean.units
analyser.add_variable(gen=opt.dist.Uniform(param1_mean - param1_width, param1_mean + param1_width),
component=opt.sel.product(), item="param")
param2_mean = facility.products[0].parameters["param2"]
param2_width = 100000 * param2_mean.units
analyser.add_variable(gen=opt.dist.Uniform(param2_mean - param2_width, param2_mean + param2_width),
component=opt.sel.product(), item="param2")
```
We are now ready to run the sensitivity analysis and collect the results.
The commented-out line (starting with a '# ') shows how to override the number of samples, which by default is set to 100 in the [SensitivityAnalyser.yaml](data/SensitivityAnalyser.yaml) file.
```
# analyser.parameters["numberOfSamples"] = 1000
analyser.run()
```
For each output, we can access the minimum and maximum values recorded (```min```, ```max```), the average value (```avg```) and the variance (```var```):
```
print("Minimum CoG: {:f}".format(analyser.outputs["CoG"]["min"]))
print("Maximum CoG: {:f}".format(analyser.outputs["CoG"]["max"]))
from numpy import sqrt
print("Average CoG: {:f} +/- {:f}".format(analyser.outputs["CoG"]["avg"], sqrt(analyser.outputs["CoG"]["var"])))
```
We can also directly access the list of all the values encountered (```all```), to examine their distribution in more detail:
```
# Plot a histogram of the CoG
import matplotlib.pyplot as plt
# The values to be plotted must first have their units removed
values = [value.magnitude for value in analyser.outputs["CoG"]["all"]]
units = analyser.outputs["CoG"]["all"][0].units
plt.hist(values)
plt.xlabel("Cost of goods ({})".format(units))
plt.ylabel("Frequency")
plt.show()
```
Some parameter values can lead to errors when evaluating the facility output (e.g. if a negative value is chosen for a quantity which must be positive). A careful choice of distributions for the variables can help avoid such problems. If, however, an error does occur, that particular run will be discarded and will not count towards the total number of runs requested. The number of failed runs is available as an output after the analysis is complete:
```
analyser.outputs["failed_runs"]
```
As with optimisation, it is possible to replicate a sensitivity analysis by specifying an initial random state. For more details, see the [relevant section in the optimisation demo](Optimisation_demo.ipynb#replication).
| github_jupyter |
<a href="https://colab.research.google.com/github/Tessellate-Imaging/monk_v1/blob/master/study_roadmaps/2_transfer_learning_roadmap/4_effect_of_training_epochs/2)%20Understand%20the%20effect%20of%20number%20of%20epochs%20in%20transfer%20learning%20-%20pytorch.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Goals
### Understand the role of number of epochs in transfer learning
### Till what point increasing epochs helps in imporving acuracy
### How overtraining can result in overfitting the data
### You will be using skin-cancer mnist to train the classifiers
# Table of Contents
## [0. Install](#0)
## [1. Train a resnet18 network for 5 epochs](#1)
## [2. Re-Train a new experiment for 10 epochs](#2)
## [3. Re-Train a third experiment for 20 epochs](#3)
## [4. Compare the experiments](#4)
<a id='0'></a>
# Install Monk
- git clone https://github.com/Tessellate-Imaging/monk_v1.git
- cd monk_v1/installation/Linux && pip install -r requirements_cu9.txt
- (Select the requirements file as per OS and CUDA version)
```
!git clone https://github.com/Tessellate-Imaging/monk_v1.git
# If using Colab install using the commands below
!cd monk_v1/installation/Misc && pip install -r requirements_colab.txt
# If using Kaggle uncomment the following command
#!cd monk_v1/installation/Misc && pip install -r requirements_kaggle.txt
# Select the requirements file as per OS and CUDA version when using a local system or cloud
#!cd monk_v1/installation/Linux && pip install -r requirements_cu9.txt
```
## Dataset Details
- Credits: https://www.kaggle.com/kmader/skin-cancer-mnist-ham10000
- Seven classes
- benign_keratosis_like_lesions
- melanocytic_nevi
- dermatofibroma
- melanoma
- vascular_lesions
- basal_cell_carcinoma
- Bowens_disease
### Download the dataset
```
! wget --load-cookies /tmp/cookies.txt "https://docs.google.com/uc?export=download&confirm=$(wget --save-cookies /tmp/cookies.txt --keep-session-cookies --no-check-certificate 'https://docs.google.com/uc?export=download&id=1MRC58-oCdR1agFTWreDFqevjEOIWDnYZ' -O- | sed -rn 's/.*confirm=([0-9A-Za-z_]+).*/\1\n/p')&id=1MRC58-oCdR1agFTWreDFqevjEOIWDnYZ" -O skin_cancer_mnist_dataset.zip && rm -rf /tmp/cookies.txt
! unzip -qq skin_cancer_mnist_dataset.zip
```
# Imports
```
# Monk
import os
import sys
sys.path.append("monk_v1/monk/");
#Using pytorch backend
from pytorch_prototype import prototype
```
<a id='1'></a>
# Train a resnet18 network for 5 epochs
## Creating and managing experiments
- Provide project name
- Provide experiment name
- For a specific data create a single project
- Inside each project multiple experiments can be created
- Every experiment can be have diferent hyper-parameters attached to it
```
gtf = prototype(verbose=1);
gtf.Prototype("Project", "Epochs-5");
```
### This creates files and directories as per the following structure
workspace
|
|--------Project
|
|
|-----Freeze_Base_Network
|
|-----experiment-state.json
|
|-----output
|
|------logs (All training logs and graphs saved here)
|
|------models (all trained models saved here)
## Set dataset and select the model
## Quick mode training
- Using Default Function
- dataset_path
- model_name
- freeze_base_network
- num_epochs
## Sample Dataset folder structure
parent_directory
|
|
|------cats
|
|------img1.jpg
|------img2.jpg
|------.... (and so on)
|------dogs
|
|------img1.jpg
|------img2.jpg
|------.... (and so on)
## Modifyable params
- dataset_path: path to data
- model_name: which pretrained model to use
- freeze_base_network: Retrain already trained network or not
- num_epochs: Number of epochs to train for
```
gtf.Default(dataset_path="skin_cancer_mnist_dataset/images",
path_to_csv="skin_cancer_mnist_dataset/train_labels.csv",
model_name="resnet18",
freeze_base_network=True,
num_epochs=5); #Set number of epochs here
#Read the summary generated once you run this cell.
```
## From summary above
Training params
Num Epochs: 5
## Train the classifier
```
#Start Training
gtf.Train();
#Read the training summary generated once you run the cell and training is completed
```
### Final training loss - 0.955
### Final validation loss - 0.811
(You may get a different result)
<a id='2'></a>
# Re-Train a new experiment for 10 epochs
## Creating and managing experiments
- Provide project name
- Provide experiment name
- For a specific data create a single project
- Inside each project multiple experiments can be created
- Every experiment can be have diferent hyper-parameters attached to it
```
gtf = prototype(verbose=1);
gtf.Prototype("Project", "Epochs-10");
```
### This creates files and directories as per the following structure
workspace
|
|--------Project
|
|
|-----Epochs-5 (Previously created)
|
|-----experiment-state.json
|
|-----output
|
|------logs (All training logs and graphs saved here)
|
|------models (all trained models saved here)
|
|
|-----Epochs-10 (Created Now)
|
|-----experiment-state.json
|
|-----output
|
|------logs (All training logs and graphs saved here)
|
|------models (all trained models saved here)
## Set dataset and select the model
## Quick mode training
- Using Default Function
- dataset_path
- model_name
- freeze_base_network
- num_epochs
## Sample Dataset folder structure
parent_directory
|
|
|------cats
|
|------img1.jpg
|------img2.jpg
|------.... (and so on)
|------dogs
|
|------img1.jpg
|------img2.jpg
|------.... (and so on)
## Modifyable params
- dataset_path: path to data
- model_name: which pretrained model to use
- freeze_base_network: Retrain already trained network or not
- num_epochs: Number of epochs to train for
```
gtf.Default(dataset_path="skin_cancer_mnist_dataset/images",
path_to_csv="skin_cancer_mnist_dataset/train_labels.csv",
model_name="resnet18",
freeze_base_network=True,
num_epochs=10); #Set number of epochs here
#Read the summary generated once you run this cell.
```
## From summary above
Training params
Num Epochs: 10
## Train the classifier
```
#Start Training
gtf.Train();
#Read the training summary generated once you run the cell and training is completed
```
### Final training loss - 0.747
### Final validation loss - 0.722
(You may get a different result)
```
```
<a id='3'></a>
# Re-Train a third experiment for 20 epochs
## Creating and managing experiments
- Provide project name
- Provide experiment name
- For a specific data create a single project
- Inside each project multiple experiments can be created
- Every experiment can be have diferent hyper-parameters attached to it
```
gtf = prototype(verbose=1);
gtf.Prototype("Project", "Epochs-20");
```
### This creates files and directories as per the following structure
workspace
|
|--------Project
|
|
|-----Epochs-5 (Previously created)
|
|-----experiment-state.json
|
|-----output
|
|------logs (All training logs and graphs saved here)
|
|------models (all trained models saved here)
|
|
|-----Epochs-10 (Previously Created)
|
|-----experiment-state.json
|
|-----output
|
|------logs (All training logs and graphs saved here)
|
|------models (all trained models saved here)
|
|
|-----Epochs-20 (Created Now)
|
|-----experiment-state.json
|
|-----output
|
|------logs (All training logs and graphs saved here)
|
|------models (all trained models saved here)
## Modifyable params
- dataset_path: path to data
- model_name: which pretrained model to use
- freeze_base_network: Retrain already trained network or not
- num_epochs: Number of epochs to train for
```
gtf.Default(dataset_path="skin_cancer_mnist_dataset/images",
path_to_csv="skin_cancer_mnist_dataset/train_labels.csv",
model_name="resnet18",
freeze_base_network=True,
num_epochs=20); #Set number of epochs here
#Read the summary generated once you run this cell.
```
## From summary above
Training params
Num Epochs: 20
## Train the classifier
```
#Start Training
gtf.Train();
#Read the training summary generated once you run the cell and training is completed
```
### Final training loss - 0.700
### Final validation loss - 0.785
(You may get a different result)
<a id='4'></a>
# Compare the experiments
```
# Invoke the comparison class
from compare_prototype import compare
```
### Creating and managing comparison experiments
- Provide project name
```
# Create a project
gtf = compare(verbose=1);
gtf.Comparison("Compare-effect-of-num-epochs");
```
### This creates files and directories as per the following structure
workspace
|
|--------comparison
|
|
|-----Compare-effect-of-num-epochs
|
|------stats_best_val_acc.png
|------stats_max_gpu_usage.png
|------stats_training_time.png
|------train_accuracy.png
|------train_loss.png
|------val_accuracy.png
|------val_loss.png
|
|-----comparison.csv (Contains necessary details of all experiments)
### Add the experiments
- First argument - Project name
- Second argument - Experiment name
```
gtf.Add_Experiment("Project", "Epochs-5");
gtf.Add_Experiment("Project", "Epochs-10");
gtf.Add_Experiment("Project", "Epochs-20");
```
### Run Analysis
```
gtf.Generate_Statistics();
```
## Visualize and study comparison metrics
### Training Accuracy Curves
```
from IPython.display import Image
Image(filename="workspace/comparison/Compare-effect-of-num-epochs/train_accuracy.png")
```
### Training Loss Curves
```
from IPython.display import Image
Image(filename="workspace/comparison/Compare-effect-of-num-epochs/train_loss.png")
```
### Validation Accuracy Curves
```
from IPython.display import Image
Image(filename="workspace/comparison/Compare-effect-of-num-epochs/val_accuracy.png")
```
### Validation loss curves
```
from IPython.display import Image
Image(filename="workspace/comparison/Compare-effect-of-num-epochs/val_loss.png")
```
## Training Accuracies achieved
### With 5 epochs - 68.2%
### With 10 epochs - 73.6%
### With 20 epochs - 74.5%
## Validation accuracies achieved
### With 5 epochs - 74.4%
### With 10 epochs - 75.5%
### With 20 epochs - 74.2%
#### Thing to note - After 7-8 epochs, accuracies and losses tend to saturate
(You may get a different result)
| github_jupyter |
#Bayesian Inference to predict water well functionality.
In this notebook, we train a model using Bayesian inference and then make predictions based on this model.
```
import pandas as pd
import numpy as np
from sklearn.neighbors import KNeighborsClassifier
import matplotlib.pyplot as plt
%matplotlib inline
```
Below we define the model, the prior distribution for parameters, likelihood function, and the posterior distribution.
```
def logprior(b, sigma2):
return -np.log(sigma2)
def logistic(x, b):
theta = linear(x,b)
return 1.0/(1+np.exp(theta))
def linear(x, b):
nFeat = x.shape[0]
y = np.zeros(x.shape[1])
for i in xrange(nFeat):
y += x[i]*b[i+1]
return b[0] + y
#return b[0] + x[0]*b[1] + x[1]*b[2] + x[2]*b[3] + x[3]*b[4]
def loglikelihood(b, sigma2, x, y, model):
n = len(x)
return -n*np.log(sigma2) - 0.5*((y-model(x, b))**2).sum()/sigma2
def logpost(b, sigma2, x, y, model):
return logprior(b, sigma2) + loglikelihood(b, sigma2, x, y, model)
```
We implement the MH algorithm using both blockwise updating and componentwise updating.
```
# mcmc algorithm
def mcmc(b_init, sig2, x, y, N, burnin, be, sig2e, model):
#MH with blockwise updating.
nFeat = len(b_init) - 1
B = np.zeros((N,nFeat + 1))
Sig2 = np.zeros(N)
b_prev = b_init
sig2_prev = sig2
count = 0
r = np.random.random(N)
for i in xrange(N):
b_star = np.random.normal(b_prev,be)
sig2_star = abs(np.random.normal(sig2_prev, sig2e))
p = logpost(b_star, sig2_star, x, y, model) - logpost(b_prev, sig2_prev, x, y, model)
if np.log(r[i]) < p:
b_prev = b_star
sig2_prev = sig2_star
count += 1
B[i] = b_prev
Sig2[i] = sig2_prev
print "The acceptance rate is "+ str(float(count)/N)+"."
return B, Sig2
def mcmcComp(b_init, sig2, x, y, N, burnin, be, sig2e, model):
#MCMC with componentwise updating instead of block updating
nParam = len(b_init)
B = np.zeros((N,nParam))
Sig2 = np.zeros(N)
b_prev = b_init
sig2_prev = sig2
count = 0
r = np.random.random((N,nParam+1))
for i in xrange(N):
#updating all the beta parameters
for j in xrange(nParam):
b_star = np.copy(b_prev)
b_star[j] = np.random.normal(b_prev[j],be[j])
p = logpost(b_star, sig2_prev, x, y, model) - logpost(b_prev, sig2_prev, x, y, model)
if np.log(r[i,j]) < p:
b_prev = b_star
count += 1
#updating sig2
sig2_star = abs(np.random.normal(sig2_prev, sig2e))
p = logpost(b_prev, sig2_star, x, y, model) - logpost(b_prev, sig2_prev, x, y, model)
if np.log(r[i,-1]) < p:
sig2_prev = sig2_star
count += 1
B[i] = b_prev
Sig2[i] = sig2_prev
print "The acceptance rate is "+ str(float(count)/(N*(nParam+1)))+"."
return B, Sig2
```
Below are functions to import the data and process them for use in our MH sampler.
```
def import_data():
"""Import data and labels and separate into training and testing data for cross-validation."""
#import data and labels
train_data = pd.read_csv('Waterpump-training-values.csv')
train_labels = pd.read_csv('Waterpump-training-labels.csv')
#separating dataset into training and testing for cross-validation - 90% into training
test_idx = np.random.uniform(0, 1, len(train_data)) <= 0.9
train = train_data[test_idx==True]
trainLabels = train_labels[test_idx==True]
test = train_data[test_idx==False]
testLabels = train_labels[test_idx==False]
return train, trainLabels, test, testLabels
def processAllData(train, trainLabels, test, testLabels):
"""Process data for use in training and testing."""
train, nFeatures = processData(train)
test, _ = processData(test)
trainLabelsVect = pd.get_dummies(trainLabels['status_group'])
trainLabelsVect['functionality'] = trainLabelsVect['functional'] + 0.5*trainLabelsVect['functional needs repair']
return train, trainLabelsVect, test, testLabels, nFeatures
def processData(data):
"""Pick out features we want to use in our model."""
features = ['longitude', 'latitude', 'age', 'gps_height', 'dry', 'population']
nFeatures = len(features)
data['age'] = 2015 - data['construction_year']
data['dry'] = data['quantity'] == 'dry'
return np.transpose(data[features].values), nFeatures
```
####Blockwise Updating
First we test the blockwise updating MH algorithm.
```
train, trainLabels, test, testLabels = import_data()
train, trainLabels, test, testLabels, nFeatures = processAllData(train, trainLabels, test, testLabels)
numBeta = nFeatures + 1 #1 more for the constant
#blockwise updating
model = logistic
b_init = [-0.5, 0, 0, 0, 0, 4, 0]
be = [0.01, 0.001, 0.001, 0.001, 0.001, 0.05, 0.001]
B, sig2= mcmc(b_init, 1, train, trainLabels['functionality'], 1000, 0, be, 0.1, model)
```
The blockwise update has an extremely low acceptance rate of 0.039, even when we start the parameters near where the componentwise updating converges. This is because we have 8 parameters!
```
plt.plot(B[:,0], label='b0')
plt.plot(B[:,1], label='b1')
plt.plot(B[:,2], label='b2')
plt.plot(B[:,3], label='b3')
plt.plot(B[:,4], label='b4')
plt.plot(B[:,5], label='b5')
plt.plot(B[:,6], label='b6')
plt.legend()
```
Now we make predictions on the withheld data with our trained parameters.
```
b_final = B[-1,:]
b_final
#predictions - these are the continuous values, need to convert to labels
yPredict = model(test, b_final)
n = len(testLabels)
correct = 0.0
upBound =0.6
lowBound = 0.4
for yPred, label in zip(yPredict, testLabels['status_group']):
if yPred >= upBound and label=='functional':
correct += 1
elif yPred <= lowBound and label == 'non functional':
correct += 1
elif yPred > lowBound and yPred < upBound and label == 'functional needs repair':
correct += 1
print correct/n
```
Extremely low prediction accuracy of 43%.
```
###plt.hist(yPredict)
```
######Componentwise Updating
Now we do the same process using component-wise updating.
```
#using componentwise updating
model = logistic
b_init = [0.5, 0, 0, 0, 0, 2, 0]
be = [0.05, 0.1, 0.1, 0.1, 0.1, 0.1, 0.1]
B, sig2= mcmcComp(b_init, 10, train, trainLabels['functionality'], 1000, 0, be, 0.1, model)
```
We get a much higher acceptance rate of 35%! The trace plot below looks like it has approximately converged.
```
plt.plot(B[:,0], label='b0')
plt.plot(B[:,1], label='b1')
plt.plot(B[:,2], label='b2')
plt.plot(B[:,3], label='b3')
plt.plot(B[:,4], label='b4')
plt.plot(B[:,5], label='b5')
plt.plot(B[:,6], label='b6')
plt.legend()
b_final = B[-1,:]
#predictions - these are the continuous values, need to convert to labels
yPredict = model(test, b_final)
n = len(testLabels)
correct = 0.0
upBound =0.6
lowBound = 0.4
for yPred, label in zip(yPredict, testLabels['status_group']):
if yPred >= upBound and label=='functional':
correct += 1
elif yPred <= lowBound and label == 'non functional':
correct += 1
elif yPred > lowBound and yPred < upBound and label == 'functional needs repair':
correct += 1
print correct/n
```
And we get 60% prediction accuracy rate with our model.
| github_jupyter |
# Regular expressions and word tokenization
> This chapter will introduce some basic NLP concepts, such as word tokenization and regular expressions to help parse text. You'll also learn how to handle non-English text and more difficult tokenization you might find. This is the Summary of lecture "Introduction to Natural Language Processing in Python", via datacamp.
- toc: true
- badges: true
- comments: true
- author: Chanseok Kang
- categories: [Python, Datacamp, Natural_Language_Processing]
- image: images/line_num_words.png
```
import re
from pprint import pprint
```
## Introduction to regular expressions
- Regular expressions
- Strings with a special syntax
- Allow us to match patterns in other strings
- Applications of regular expressions
- Find all web links in a document
- Parse email addresses, remove/replace unwanted characters
- Common Regex patterns
| pattern | matches | examples |
| ------- | ------- | -------- |
| \w+ | word | 'Magic' |
| \d | digit | 9 |
| \s | space | '' |
| .* | wildcard | 'username74' |
| + or * | greedy match | 'aaaaaaa' |
| \S | not space | 'no_spaces' |
| [a-z] | lowercase group | 'abcdfg' |
- Python's re Module
- `split`: split a string on regex
- `findall`: find all patterns in a string
- `search`: search for a pattern
- `match`: match an entire string or substring based on a pattern
- Pattern first, and the string second
- May return an iterator, string, or match object
```
my_string = "Let's write RegEx!"
PATTERN = r"\w+"
re.findall(PATTERN, my_string)
```
### Practicing regular expressions - re.split() and re.findall()
Now you'll get a chance to write some regular expressions to match digits, strings and non-alphanumeric characters. Take a look at `my_string` first by printing it in the IPython Shell, to determine how you might best match the different steps.
Note: It's important to prefix your regex patterns with `r` to ensure that your patterns are interpreted in the way you want them to. Else, you may encounter problems to do with escape sequences in strings. For example, `"\n"` in Python is used to indicate a new line, but if you use the `r` prefix, it will be interpreted as the raw string `"\n"` - that is, the character `"\"` followed by the character `"n"` - and not as a new line.
Remember from the video that the syntax for the regex library is to always to pass the **pattern first**, and then the **string second**.
```
my_string = "Let's write RegEx! Won't that be fun? I sure think so. Can you find 4 sentences? Or perhaps, all 19 words?"
# Write a pattern to match sentence endings: sentence_endings
sentence_endings = r"[.?!]"
# Split my_string on sentence endings and print the result
print(re.split(sentence_endings, my_string))
# Find all capicalized words in my_string and print the result
capitalized_words = r"[A-Z]\w+"
print(re.findall(capitalized_words, my_string))
# Split my_string on spaces and print the result
spaces = r"\s+"
print(re.split(spaces, my_string))
# Find all digits in my_string and print the result
digits = r"\d+"
print(re.findall(digits, my_string))
```
## Introduction to tokenization
- Tokenization
- Turning a string or document into **tokens** (smaller chunks)
- One step in preparing a text for NLP
- Many different theories and rules
- You can create your own rules using regular expressions
- Some examples:
- Breaking out words or sentences
- Separating punctuation
- Separating all hashtags in a tweet
- Why tokenize?
- Easier to map part of speech
- Matching common words
- Removing unwanted tokens
- Other `nltk` tokenizers
- `sent_tokenize`: tokenize a document into sentences
- `regexp_tokenize`: tokenize a string or document based on a regular expression pattern
- `TweetTokenizer`: special class just for tweet tokenization, allowing you to separate hashtags, mentions and lots of exclamation points
### Word tokenization with NLTK
Here, you'll be using the first scene of Monty Python's Holy Grail, which has been pre-loaded as `scene_one`.
Your job in this exercise is to utilize `word_tokenize` and `sent_tokenize` from `nltk.tokenize` to tokenize both words and sentences from Python strings - in this case, the first scene of Monty Python's Holy Grail.
> Note: Before using NLTK, you must install `punkt` package for tokenizer
```
with open('./dataset/grail.txt', 'r') as file:
holy_grail = file.read()
scene_one = re.split('SCENE 2:', holy_grail)[0]
scene_one
from nltk.tokenize import word_tokenize, sent_tokenize
# Split scene_one into sentences: sentences
sentences = sent_tokenize(scene_one)
# Use word_tokenize to tokenize the fourth sentence: tokenized_sent
tokenized_sent = word_tokenize(sentences[3])
# Make a set of unique tokens in the entire scene: unique_tokens
unique_tokens = set(word_tokenize(scene_one))
# Print the unique tokens result
print(unique_tokens)
```
### More regex with re.search()
In this exercise, you'll utilize `re.search()` and `re.match()` to find specific tokens. Both search and match expect regex patterns, similar to those you defined in an earlier exercise. You'll apply these regex library methods to the same Monty Python text from the `nltk` corpora.
```
# Search for the first occurrence of "coconuts" in scene_one: match
match = re.search("coconuts", scene_one)
# Print the start and end indexes of match
print(match.start(), match.end())
# Write a regular expression to search for anything in square brackets: pattern1
pattern1 = r"\[.*\]"
# Use re.search to find the first text in square brackets
print(re.search(pattern1, scene_one))
# Find the script notation at the beginning of the fourth sentence and print it
pattern2 = r"[\w\s]+:"
print(re.match(pattern2, sentences[3]))
```
## Advanced tokenization with NLTK and regex
- Regex groups using or `|`
- OR is represented using `|`
- You can define a group using `()`
- You can define explicit character ranges using `[]`
- Regex ranges and groups
| pattern | matches | example |
| ------- | ------- | ------- |
| [A-Za-z]+ | upper and lowercase English alphabet | 'ABCDEFghijk' |
| [0-9] | numbers from 0 to 9 | 9 |
| [A-Za-z\-\.]+ | upper and lowercase English alphabet, - and . | 'My-Website.com' |
| (a-z) | a, - and z | 'a-z' |
| (\s+|,) | spaces or a comma | ', ' |
### Choosing a tokenizer
Given the following string, which of the below patterns is the best tokenizer? If possible, you want to retain sentence punctuation as separate tokens, but have `'#1'` remain a single token.
```python
my_string = "SOLDIER #1: Found them? In Mercea? The coconut's tropical!"
```
Additionally, `regexp_tokenize` has been imported from `nltk.tokenize`. You can use `regexp_tokenize(string, pattern)` with `my_string` and one of the patterns as arguments to experiment for yourself and see which is the best tokenizer.
```
from nltk.tokenize import regexp_tokenize
my_string = "SOLDIER #1: Found them? In Mercea? The coconut's tropical!"
pattern1 = r'(\\w+|\\?|!)'
pattern2 = r"(\w+|#\d|\?|!)"
pattern3 = r'(#\\d\\w+\\?!)'
pattern4 = r'\\s+'
pprint(regexp_tokenize(my_string, pattern2))
```
### Regex with NLTK tokenization
Twitter is a frequently used source for NLP text and tasks. In this exercise, you'll build a more complex tokenizer for tweets with hashtags and mentions using nltk and regex. The `nltk.tokenize.TweetTokenizer` class gives you some extra methods and attributes for parsing tweets.
Here, you're given some example tweets to parse using both `TweetTokenizer` and `regexp_tokenize` from the `nltk.tokenize` module.
Unlike the syntax for the regex library, with `nltk_tokenize()` you pass the pattern as the second argument.
```
tweets = ['This is the best #nlp exercise ive found online! #python',
'#NLP is super fun! <3 #learning',
'Thanks @datacamp :) #nlp #python']
from nltk.tokenize import regexp_tokenize, TweetTokenizer
# Define a regex pattern to find hashtags: pattern1
pattern1 = r"#\w+"
# Use the pattern on the first tweet in the tweets list
hashtags = regexp_tokenize(tweets[0], pattern1)
print(hashtags)
# write a pattern that matches both mentions (@) and hashtags
pattern2 = r"[@|#]\w+"
# Use the pattern on the last tweet in the tweets list
mentions_hashtags = regexp_tokenize(tweets[-1], pattern2)
print(mentions_hashtags)
# Use the TweetTokenizer to tokenize all tweets into one list
tknzr = TweetTokenizer()
all_tokens = [tknzr.tokenize(t) for t in tweets]
print(all_tokens)
```
### Non-ascii tokenization
In this exercise, you'll practice advanced tokenization by tokenizing some non-ascii based text. You'll be using German with emoji!
Here, you have access to a string called `german_text`, which has been printed for you in the Shell. Notice the emoji and the German characters!
Unicode ranges for emoji are:
`('\U0001F300'-'\U0001F5FF')`, `('\U0001F600-\U0001F64F')`, `('\U0001F680-\U0001F6FF')`, and `('\u2600'-\u26FF-\u2700-\u27BF')`.
```
german_text = 'Wann gehen wir Pizza essen? 🍕 Und fährst du mit Über? 🚕'
# Tokenize and print all words in german_text
all_words = word_tokenize(german_text)
print(all_words)
# Tokenize and print only capital words
capital_words = r"[A-ZÜ]\w+"
print(regexp_tokenize(german_text, capital_words))
# Tokenize and print only emoji
emoji = "['\U0001F300-\U0001F5FF'|'\U0001F600-\U0001F64F'|'\U0001F680-\U0001F6FF'|'\u2600-\u26FF\u2700-\u27BF']"
print(regexp_tokenize(german_text, emoji))
```
## Charting word length with NLTK
### Charting practice
Try using your new skills to find and chart the number of words per line in the script using matplotlib. The Holy Grail script is loaded for you, and you need to use regex to find the words per line.
Using list comprehensions here will speed up your computations. For example: `my_lines = [tokenize(l) for l in lines]` will call a function tokenize on each line in the list lines. The new transformed list will be saved in the `my_lines` variable.
```
import matplotlib.pyplot as plt
# Split the script into lines: lines
lines = holy_grail.split('\n')
# Replace all script lines for speaker
pattern = "[A-Z]{2,}(\s)?(#\d)?([A-Z]{2,})?:"
lines = [re.sub(pattern, '', l) for l in lines]
# Tokenize each line: tokenized_lines
tokenized_lines = [regexp_tokenize(s, '\w+') for s in lines]
# Make a frequency list of lengths: line_num_words
line_num_words = [len(t_line) for t_line in tokenized_lines]
# Plot a histogram of the line lengths
plt.figure(figsize=(8,8))
plt.hist(line_num_words);
plt.title('# of words per line in holy_grail');
```
| github_jupyter |
# How to build a linear factor model
Algorithmic trading strategies use linear factor models to quantify the relationship between the return of an asset and the sources of risk that represent the main drivers of these returns. Each factor risk carries a premium, and the total asset return can be expected to correspond to a weighted average of these risk premia.
There are several practical applications of factor models across the portfolio management process from construction and asset selection to risk management and performance evaluation. The importance of factor models continues to grow as common risk factors are now tradeable:
- A summary of the returns of many assets by a much smaller number of factors reduces the amount of data required to estimate the covariance matrix when optimizing a portfolio
- An estimate of the exposure of an asset or a portfolio to these factors allows for the management of the resultant risk, for instance by entering suitable hedges when risk factors are themselves traded
- A factor model also permits the assessment of the incremental signal content of new alpha factors
- A factor model can also help assess whether a manager's performance relative to a benchmark is indeed due to skill in selecting assets and timing the market, or if instead, the performance can be explained by portfolio tilts towards known return drivers that can today be replicated as low-cost, passively managed funds without incurring active management fees
## Imports & Settings
```
from pprint import pprint
from pandas_datareader.famafrench import get_available_datasets
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from statsmodels.api import OLS, add_constant
from pathlib import Path
import warnings
from linearmodels.asset_pricing import TradedFactorModel, LinearFactorModel, LinearFactorModelGMM
# due to https://stackoverflow.com/questions/50394873/import-pandas-datareader-gives-importerror-cannot-import-name-is-list-like
# may become obsolete when fixed
pd.core.common.is_list_like = pd.api.types.is_list_like
import pandas_datareader.data as web
warnings.filterwarnings('ignore')
plt.style.use('fivethirtyeight')
```
## Get Data
Fama and French make updated risk factor and research portfolio data available through their [website](http://mba.tuck.dartmouth.edu/pages/faculty/ken.french/data_library.html), and you can use the `pandas_datareader` package to obtain the data.
### Risk Factors
In particular, we will be using the five Fama—French factors that result from sorting stocks first into three size groups and then into two for each of the remaining three firm-specific factors.
Hence, the factors involve three sets of value-weighted portfolios formed as 3 x 2 sorts on size and book-to-market, size and operating profitability, and size and investment. The risk factor values computed as the average returns of the portfolios (PF) as outlined in the following table:
| Label | Name | Description |
|-------|-------------------------------|-------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------|
| SMB | Small Minus Big | Average return on the nine small stock portfolios minus the average return on the nine big stock portfolios |
| HML | High Minus Low | Average return on the two value portfolios minus the average return on the two growth portfolios |
| RMW | Robust minus Weak | Average return on the two robust operating profitability portfolios minus the average return on the two weak operating profitability portfolios |
| CMA | Conservative Minus Aggressive | Average return on the two conservative investment portfolios minus the average return on the two aggressive investment portfolios |
| Rm-Rf | Excess return on the market | Value-weight return of all firms incorporated in the US and listed on the NYSE, AMEX, or NASDAQ at the beginning of month t with 'good' data for t minus the one-month Treasury bill rate |
The Fama-French 5 factors are based on the 6 value-weight portfolios formed on size and book-to-market, the 6 value-weight portfolios formed on size and operating profitability, and the 6 value-weight portfolios formed on size and investment.
We will use returns at a monthly frequency that we obtain for the period 2010 – 2017 as follows:
```
ff_factor = 'F-F_Research_Data_5_Factors_2x3'
ff_factor_data = web.DataReader(ff_factor, 'famafrench', start='2010', end='2017-12')[0]
ff_factor_data.info()
ff_factor_data.describe()
```
### Portfolios
Fama and French also make available numerous portfolios that we can illustrate the estimation of the factor exposures, as well as the value of the risk premia available in the market for a given time period. We will use a panel of the 17 industry portfolios at a monthly frequency.
We will subtract the risk-free rate from the returns because the factor model works with excess returns:
```
ff_portfolio = '17_Industry_Portfolios'
ff_portfolio_data = web.DataReader(ff_portfolio, 'famafrench', start='2010', end='2017-12')[0]
ff_portfolio_data = ff_portfolio_data.sub(ff_factor_data.RF, axis=0)
ff_portfolio_data.info()
ff_portfolio_data.describe()
```
### Equity Data
```
with pd.HDFStore('../../data/assets.h5') as store:
prices = store['/quandl/wiki/prices'].adj_close.unstack().loc['2010':'2017']
equities = store['/us_equities/stocks'].drop_duplicates()
sectors = equities.filter(prices.columns, axis=0).sector.to_dict()
prices = prices.filter(sectors.keys()).dropna(how='all', axis=1)
returns = prices.resample('M').last().pct_change().mul(100).to_period('M')
returns = returns.dropna(how='all').dropna(axis=1)
returns.info()
```
### Align data
```
ff_factor_data = ff_factor_data.loc[returns.index]
ff_portfolio_data = ff_portfolio_data.loc[returns.index]
ff_factor_data.describe()
```
### Compute excess Returns
```
excess_returns = returns.sub(ff_factor_data.RF, axis=0)
excess_returns.info()
excess_returns = excess_returns.clip(lower=np.percentile(excess_returns, 1),
upper=np.percentile(excess_returns, 99))
```
## Fama-Macbeth Regression
Given data on risk factors and portfolio returns, it is useful to estimate the portfolio's exposure, that is, how much the risk factors drive portfolio returns, as well as how much the exposure to a given factor is worth, that is, the what market's risk factor premium is. The risk premium then permits to estimate the return for any portfolio provided the factor exposure is known or can be assumed.
```
ff_portfolio_data.info()
ff_factor_data.info()
```
To address the inference problem caused by the correlation of the residuals, Fama and MacBeth proposed a two-step methodology for a cross-sectional regression of returns on factors. The two-stage Fama—Macbeth regression is designed to estimate the premium rewarded for the exposure to a particular risk factor by the market. The two stages consist of:
- First stage: N time-series regression, one for each asset or portfolio, of its excess returns on the factors to estimate the factor loadings.
- Second stage: T cross-sectional regression, one for each time period, to estimate the risk premium.
See corresponding section in Chapter 7 of [Machine Learning for Trading](https://www.amazon.com/Hands-Machine-Learning-Algorithmic-Trading-ebook/dp/B07JLFH7C5/ref=sr_1_2?ie=UTF8&qid=1548455634&sr=8-2&keywords=machine+learning+algorithmic+trading) for details.
Now we can compute the factor risk premia as the time average and get t-statistic to assess their individual significance, using the assumption that the risk premia estimates are independent over time.
If we had a very large and representative data sample on traded risk factors we could use the sample mean as a risk premium estimate. However, we typically do not have a sufficiently long history to and the margin of error around the sample mean could be quite large.
The Fama—Macbeth methodology leverages the covariance of the factors with other assets to determine the factor premia. The second moment of asset returns is easier to estimate than the first moment, and obtaining more granular data improves estimation considerably, which is not true of mean estimation.
### Step 1: Factor Exposures
We can implement the first stage to obtain the 17 factor loading estimates as follows:
```
betas = []
for industry in ff_portfolio_data:
step1 = OLS(endog=ff_portfolio_data.loc[ff_factor_data.index, industry],
exog=add_constant(ff_factor_data)).fit()
betas.append(step1.params.drop('const'))
betas = pd.DataFrame(betas,
columns=ff_factor_data.columns,
index=ff_portfolio_data.columns)
betas.info()
```
### Step 2: Risk Premia
For the second stage, we run 96 regressions of the period returns for the cross section of portfolios on the factor loadings
```
lambdas = []
for period in ff_portfolio_data.index:
step2 = OLS(endog=ff_portfolio_data.loc[period, betas.index],
exog=betas).fit()
lambdas.append(step2.params)
lambdas = pd.DataFrame(lambdas,
index=ff_portfolio_data.index,
columns=betas.columns.tolist())
lambdas.info()
lambdas.mean()
t = lambdas.mean().div(lambdas.std())
t
```
#### Results
```
ax1 = plt.subplot2grid((1, 3), (0, 0))
ax2 = plt.subplot2grid((1, 3), (0, 1), colspan=2)
lambdas.mean().plot.barh(ax=ax1)
lambdas.rolling(60).mean().plot(lw=2, figsize=(14,10), sharey=True, ax=ax2);
lambdas.rolling(60).mean().plot(lw=2, figsize=(14,10), subplots=True,sharey=True);
```
## Fama-Macbeth with the LinearModels library
The linear_models library extends statsmodels with various models for panel data and also implements the two-stage Fama—MacBeth procedure:
```
mod = LinearFactorModel(portfolios=ff_portfolio_data,
factors=ff_factor_data)
res = mod.fit()
print(res)
plt.rc('figure', figsize=(12, 7))
plt.text(0.01, 0.05, str(res), {'fontsize': 14}, fontproperties = 'monospace')
plt.axis('off')
plt.tight_layout()
plt.subplots_adjust(left=0.2, right=0.8, top=0.8, bottom=0.1)
plt.savefig('factor_model.png', bbox_inches='tight', dpi=300);
```
This provides us with the same result:
```
lambdas.mean()
```
| github_jupyter |
# Download the data
## Summary: Create lists with updated stocks yahoo finance codes to download the data
```
# Import required libraries
import os
import pickle
# Get current working directory
mycwd = os.getcwd()
print(mycwd)
# Change to data directory
os.chdir("..")
os.chdir(str(os.getcwd()) + "\\Models")
```
### Define Dictionary for NIFTY50
```
# Define NIFTY50 yahoo finance tickers
NIFTY50 = {"ADANIPORTS": "ADANIPORTS.NS", "ASIANPAINT": "ASIANPAINT.NS", "AXISBANK": "AXISBANK.NS",
"BAJAJ-AUTO": "BAJAJ-AUTO.NS", "BAJAJFINSV": "BAJAJFINSV.NS", "BAJFINANCE": "BAJFINANCE.NS",
"BHARTIARTL": "BHARTIARTL.NS", "BPCL": "BPCL.NS", "BRITANNIA": "BRITANNIA.NS", "CIPLA": "CIPLA.NS",
"COALINDIA": "COALINDIA.NS", "DIVISLAB": "DIVISLAB.NS", "DRREDDY": "DRREDDY.NS", "EICHERMOT": "EICHERMOT.NS",
"GAIL": "GAIL.NS", "GRASIM": "GRASIM.NS", "HCLTECH": "HCLTECH.NS", "HDFC": "HDFC.NS", "HDFCBANK": "HDFCBANK.NS",
"HDFCLIFE": "HDFCLIFE.NS", "HEROMOTOCO": "HEROMOTOCO.NS", "HINDALCO": "HINDALCO.NS",
"HINDUNILVR": "HINDUNILVR.NS", "ICICIBANK": "ICICIBANK.NS", "INDUSINDBK": "INDUSINDBK.NS", "INFY": "INFY.NS",
"IOC": "IOC.NS", "ITC": "ITC.NS", "JSWSTEEL": "JSWSTEEL.NS", "KOTAKBANK": "KOTAKBANK.NS", "LT": "LT.NS",
"M&M": "M&M.NS", "MARUTI": "MARUTI.NS", "NESTLEIND": "NESTLEIND.NS", "NTPC": "NTPC.NS", "ONGC": "ONGC.NS",
"POWERGRID": "POWERGRID.NS", "RELIANCE": "RELIANCE.NS", "SBILIFE": "SBILIFE.NS", "SBIN": "SBIN.NS",
"SHREECEM": "SHREECEM.NS", "SUNPHARMA": "SUNPHARMA.NS", "TATAMOTORS": "TATAMOTORS.NS",
"TATASTEEL": "TATASTEEL.NS", "TCS": "TCS.NS", "TECHM": "TECHM.NS", "TITAN": "TITAN.NS",
"ULTRACEMCO": "ULTRACEMCO.NS", "UPL": "UPL.NS", "WIPRO": "WIPRO.NS" }
# Save the NIFTY50 Tickers dictionary
with open('NIFTY50.pkl', 'wb') as f:
pickle.dump(NIFTY50, f, pickle.HIGHEST_PROTOCOL)
# Define NIFTYNEXT50 tickers
NIFTYNEXT50 = {"ABBOTINDIA": "ABBOTINDIA.NS", "ACC": "ACC.NS", "ADANIGREEN": "ADANIGREEN.NS",
"ADANITRANS": "ADANITRANS.NS", "ALKEM": "ALKEM.NS", "AMBUJACEM": "AMBUJACEM.NS",
"AUROPHARMA": "AUROPHARMA.NS", "BAJAJHLDNG": "BAJAJHLDNG.NS", "BANDHANBNK": "BANDHANBNK.NS",
"BANKBARODA": "BANKBARODA.NS", "BERGEPAINT": "BERGEPAINT.NS", "BIOCON": "BIOCON.NS",
"BOSCHLTD": "BOSCHLTD.NS", "CADILAHC": "CADILAHC.NS", "COLPAL": "COLPAL.NS", "CONCOR": "CONCOR.NS",
"DABUR": "DABUR.NS", "DLF": "DLF.NS", "DMART": "DMART.NS", "GICRE": "GICRE.NS", "GODREJCP": "GODREJCP.NS",
"HAVELLS": "HAVELLS.NS", "HDFCAMC": "HDFCAMC.NS", "HINDPETRO": "HINDPETRO.NS", "HINDZINC": "HINDZINC.NS",
"ICICIGI": "ICICIGI.NS", "ICICIPRULI": "ICICIPRULI.NS", "IGL": "IGL.NS", "INDIGO": "INDIGO.NS",
"INDUSTOWER": "INDUSTOWER.NS", "LTI": "LTI.NS", "LUPIN": "LUPIN.NS", "MARICO": "MARICO.NS",
"MCDOWELL-N": "MCDOWELL-N.NS", "MOTHERSUMI": "MOTHERSUMI.NS", "MUTHOOTFIN": "MUTHOOTFIN.NS",
"NAUKRI": "NAUKRI.NS", "NMDC": "NMDC.NS", "OFSS": "OFSS.NS", "PEL": "PEL.NS", "PETRONET": "PETRONET.NS",
"PFC": "PFC.NS", "PGHH": "PGHH.NS", "PIDILITIND": "PIDILITIND.NS", "PNB": "PNB.NS", "SBICARD": "SBICARD.NS",
"SIEMENS": "SIEMENS.NS", "TATACONSUM": "TATACONSUM.NS", "TORNTPHARM": "TORNTPHARM.NS", "UBL": "UBL.NS"}
# Save NIFTYNEXT50 tickers dictionary
with open('NIFTYNEXT50.pkl', 'wb') as f:
pickle.dump(NIFTYNEXT50, f, pickle.HIGHEST_PROTOCOL)
# Define NIFTYMIDCAP100 dictionary
NIFTYMIDCAP100 = {"AARTIIND": "AARTIIND.NS", "ABCAPITAL": "ABCAPITAL.NS", "ABFRL": "ABFRL.NS", "ADANIENT": "ADANIENT.NS",
"AJANTPHARM": "AJANTPHARM.NS", "AMARAJABAT": "AMARAJABAT.NS", "APLLTD": "APLLTD.NS",
"APOLLOHOSP": "APOLLOHOSP.NS", "APOLLOTYRE": "APOLLOTYRE.NS", "ASHOKLEY": "ASHOKLEY.NS",
"ATGL": "ATGL.NS", "AUBANK": "AUBANK.NS", "BALKRISIND": "BALKRISIND.NS", "BANKINDIA": "BANKINDIA.NS",
"BATAINDIA": "BATAINDIA.NS", "BBTC": "BBTC.NS", "BEL": "BEL.NS", "BHARATFORG": "BHARATFORG.NS",
"BHEL": "BHEL.NS", "CANBK": "CANBK.NS", "CASTROLIND": "CASTROLIND.NS", "CESC": "CESC.NS",
"CHOLAFIN": "CHOLAFIN.NS", "COFORGE": "COFORGE.NS", "COROMANDEL": "COROMANDEL.NS",
"CROMPTON": "CROMPTON.NS", "CUB": "CUB.NS", "CUMMINSIND": "CUMMINSIND.NS", "DALBHARAT": "DALBHARAT.NS",
"DHANI": "DHANI.NS", "EDELWEISS": "EDELWEISS.NS", "EMAMILTD": "EMAMILTD.NS", "ENDURANCE": "ENDURANCE.NS",
"ESCORTS": "ESCORTS.NS", "EXIDEIND": "EXIDEIND.NS", "FEDERALBNK": "FEDERALBNK.NS", "FORTIS": "FORTIS.NS",
"FRETAIL": "FRETAIL.NS", "GLENMARK": "GLENMARK.NS", "GMRINFRA": "GMRINFRA.NS",
"GODREJAGRO": "GODREJAGRO.NS", "GODREJIND": "GODREJIND.NS", "GODREJPROP": "GODREJPROP.NS",
"GSPL": "GSPL.NS", "GUJGASLTD": "GUJGASLTD.NS", "HUDCO": "HUDCO.NS", "IBULHSGFIN": "IBULHSGFIN.NS",
"IDEA": "IDEA.NS", "IDFCFIRSTB": "IDFCFIRSTB.NS", "INDHOTEL": "INDHOTEL.NS", "IPCALAB": "IPCALAB.NS",
"IRCTC": "IRCTC.NS", "ISEC": "ISEC.NS", "JINDALSTEL": "JINDALSTEL.NS", "JSWENERGY": "JSWENERGY.NS",
"JUBLFOOD": "JUBLFOOD.NS", "L&TFH": "L&TFH.NS", "LALPATHLAB": "LALPATHLAB.NS",
"LICHSGFIN": "LICHSGFIN.NS", "LTTS": "LTTS.NS", "M&MFIN": "M&MFIN.NS", "MANAPPURAM": "MANAPPURAM.NS",
"MFSL": "MFSL.NS", "MGL": "MGL.NS", "MINDTREE": "MINDTREE.NS", "MPHASIS": "MPHASIS.NS", "MRF": "MRF.NS",
"NAM-INDIA": "NAM-INDIA.NS", "NATCOPHARM": "NATCOPHARM.NS", "NATIONALUM": "NATIONALUM.NS",
"NAVINFLUOR": "NAVINFLUOR.NS", "OBEROIRLTY": "OBEROIRLTY.NS", "OIL": "OIL.NS", "PAGEIND": "PAGEIND.NS",
"PFIZER": "PFIZER.NS", "PIIND": "PIIND.NS", "POLYCAB": "POLYCAB.NS", "PRESTIGE": "PRESTIGE.NS",
"RAJESHEXPO": "RAJESHEXPO.NS", "RAMCOCEM": "RAMCOCEM.NS", "RBLBANK": "RBLBANK.NS", "RECLTD": "RECLTD.NS",
"SAIL": "SAIL.NS", "SANOFI": "SANOFI.NS", "SRF": "SRF.NS", "SRTRANSFIN": "SRTRANSFIN.NS",
"SUNTV": "SUNTV.NS", "SYNGENE": "SYNGENE.NS", "TATACHEM": "TATACHEM.NS", "TATAPOWER": "TATAPOWER.NS",
"TORNTPOWER": "TORNTPOWER.NS", "TRENT": "TRENT.NS", "TVSMOTOR": "TVSMOTOR.NS",
"UNIONBANK": "UNIONBANK.NS", "VBL": "VBL.NS", "VGUARD": "VGUARD.NS", "VOLTAS": "VOLTAS.NS",
"WHIRLPOOL": "WHIRLPOOL.NS", "YESBANK": "YESBANK.NS", "ZEEL": "ZEEL.NS"}
# Save NIFTYMIDCAP100 dictionary
with open('NIFTYMIDCAP100.pkl', 'wb') as f:
pickle.dump(NIFTYMIDCAP100, f, pickle.HIGHEST_PROTOCOL)
# Define dictionary for other indices
OTHERINDICES = {
"INDIAVIX": "^INDIAVIX"
}
# Save other indices dictionary
with open('OTHERINDICES.pkl', 'wb') as f:
pickle.dump(OTHERINDICES, f, pickle.HIGHEST_PROTOCOL)
```
__Comments:__ All dictionaries are saved in "//Data" folder and should be updated in case of any error. To change the strategy for other market economy, please redefine the dictionaries as per the market with "Yahoo finance Tickers"
| github_jupyter |
```
# default_exp series.preproc
```
# series.preproc
> Tools for preprocessing DICOM metadata imported using `dicomtools.core` into in a `pandas.DataFrame` in preparation for training RandomForest classifier to predict series type.
```
#hide
from nbdev.showdoc import *
#export
from dicomtools.imports import *
from dicomtools.core import *
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import MultiLabelBinarizer
#export
def exclude_other(df):
if 'BodyPartExamined' not in df.columns: return df
other = ['SPINE', 'CSPINE']
filt = df.BodyPartExamined.isin(other)
filt1 = df.SOPClassUID == "MR Image Storage"
df1 = df[~filt].copy()
return df1[filt1].reset_index(drop=True)
#export
def get_series_fp(fn): return Path(fn).parent
#export
def compute_plane(row):
'''
Computes the plane of imaging from the direction cosines provided in the `ImageOrientationPatient` field.
The format of the values in this field is: `[x1, y1, z1, x2, y2, z2]`,
which correspond to the direction cosines for the first row and column of the image pixel data.
'''
planes = ['sag', 'cor', 'ax']
if 'ImageOrientationPatient1' in row.keys():
dircos = [v for k,v in row.items() if 'ImageOrientationPatient' in k]
else:
dircos = row['ImageOrientationPatient']
dircos = np.array(dircos).reshape(2,3)
pnorm = abs(np.cross(dircos[0], dircos[1]))
return planes[np.argmax(pnorm)]
#export
_c = re.compile(r'(\+-?c|post)')
#export
def detect_contrast(row):
if _c.search(str(row['SeriesDescription']).lower()): return 1
c = row['ContrastBolusAgent']
if type(c) == str: return 1
return 0
row = {'SeriesDescription': 'ax t1 +c', 'ContrastBolusAgent': 'Gadavist'}
row1 = {'SeriesDescription': 'ax t1', 'ContrastBolusAgent': np.nan}
row2 = {'SeriesDescription': 'ax t1', 'ContrastBolusAgent': 'Gadavist'} # example of discordant SD
row3 = {'SeriesDescription': 'AX T1 POST', 'ContrastBolusAgent': np.nan} # also discordant
assert detect_contrast(row) == 1
assert detect_contrast(row1) == 0
assert detect_contrast(row2) == 1
assert detect_contrast(row3) == 1
#export
_re_extra_info = re.compile(r'[<\([].*?[\]\)>]')
#export
def rm_extra_info(t):
"Remove extraneous info in closures"
return _re_extra_info.sub('', t).strip()
assert rm_extra_info('ax t1 <mpr>') == 'ax t1'
assert rm_extra_info('adc (mm^2/s)') == 'adc'
assert rm_extra_info('ax t1 [date]') == 'ax t1'
#export
_t1 = re.compile(r't1')
_spgr = re.compile(r'spgr|mprage')
_t2 = re.compile(r't2')
_flair = re.compile(r'flair')
_swi = re.compile(r'swi|gre|susc|mag|pha|sw')
_adc = re.compile(r'adc|apparent')
_eadc = re.compile(r'exp|eadc')
_dwi = re.compile(r'diff|dwi|trace')
_mra = re.compile(r'mra|angio|cow|tof|mip')
_loc = re.compile(r'loc|scout')
#export
def _find_seq(sd):
if _t1.search(sd):
if _spgr.search(sd): return 'spgr'
else: return 't1'
if _spgr.search(sd): return 'spgr'
if _t2.search(sd):
if _flair.search(sd): return 'flair'
elif _swi.search(sd): return 'swi'
else: return 't2'
if _flair.search(sd): return 'flair'
if _swi.search(sd): return 'swi'
if _adc.search(sd):
if _eadc.search(sd): return 'other'
else: return 'adc'
if _dwi.search(sd):
if _eadc.search(sd): return 'other'
else: return 'dwi'
if _mra.search(sd): return 'mra'
if _loc.search(sd): return 'loc'
return 'unknown'
def test_find_seq(sd, targ): assert _find_seq(sd) == targ
test_find_seq('ax t1 +c', 't1')
test_find_seq('ax t1 flair +c', 't1')
test_find_seq('ax t2 +c', 't2')
test_find_seq('ax t2 flair', 'flair')
test_find_seq('ax t2 gre', 'swi')
test_find_seq('ax swi', 'swi')
test_find_seq('ax susc', 'swi')
test_find_seq('adc', 'adc')
test_find_seq('eadc', 'other')
test_find_seq('ax dwi', 'dwi')
test_find_seq('ax diffusion', 'dwi')
test_find_seq('ax spgr +c', 'spgr')
test_find_seq('localizer', 'loc')
#export
def _extract_label(sd):
t = rm_extra_info(str(sd).lower())
return _find_seq(t)
assert _extract_label('ax t1 +c') == 't1'
assert _extract_label('ax t1 +c [date]') == 't1'
assert _extract_label('<MPR Thick Range>') == 'unknown'
#export
def extract_labels(df):
"Extract candidate labels from Series Descriptions and computed plane"
df1 = df[['fname', 'SeriesDescription']].copy()
df1['fname'] = df1.fname.apply(get_series_fp)
print("Computing planes of imaging from `ImageOrientationPatient`.")
df1['plane'] = df.apply(compute_plane, axis=1)
print("Extracting candidate labels from `SeriesDescription`.")
df1['seq_label'] = df1['SeriesDescription'].apply(_extract_label)
print("Detecting contrast from `SeriesDescription` and `ContrastMediaAgent`.")
df1['contrast'] = df.apply(detect_contrast, axis=1)
return df1
#export
_keep = [
'fname',
# Patient info
'PatientID',
# Study info
'StudyInstanceUID',
'StudyID',
# Series info
'SeriesInstanceUID',
'SeriesNumber',
'SeriesDescription',
'AcquisitionNumber',
# Image info and features
'InstanceNumber',
'ImageOrientationPatient',
'ScanningSequence',
'SequenceVariant',
'ScanOptions',
'MRAcquisitionType',
'AngioFlag',
'SliceThickness',
'RepetitionTime',
'EchoTime',
'EchoTrainLength',
'PixelSpacing',
'ContrastBolusAgent',
'InversionTime',
'DiffusionBValue'
]
_dummies = [
'ScanningSequence',
'SequenceVariant',
'ScanOptions'
]
_d_prefixes = [
'seq',
'var',
'opt'
]
_binarize = [
'MRAcquisitionType',
'AngioFlag',
'ContrastBolusAgent',
'DiffusionBValue'
]
_rescale = [
'SliceThickness',
'RepetitionTime',
'EchoTime',
'EchoTrainLength',
'PixelSpacing',
'InversionTime'
]
#export
def _make_col_binary(df, col):
s = df[col].isna()
if any(s):
df[col] = s.apply(lambda x: 0 if x else 1)
else:
targ = df.loc[0, col]
df[col] = df[col].apply(lambda x: 0 if x == targ else 1)
#export
def make_binary_cols(df, cols):
df1 = df.copy()
for col in cols:
_make_col_binary(df1, col)
return df1
#export
def rescale_cols(df, cols):
df1 = df.copy()
scaler = MinMaxScaler()
df1[cols] = scaler.fit_transform(df1[cols])
return df1.fillna(0)
#export
def get_dummies(df, cols=_dummies, prefix=_d_prefixes):
df1 = df.copy()
for i, col in enumerate(cols):
df1[col] = df1[col].fillna('NONE')
mlb = MultiLabelBinarizer()
df1 = df1.join(
pd.DataFrame(mlb.fit_transform(df1.pop(col)), columns=mlb.classes_).add_prefix(f'{prefix[i]}_')
)
return df1
#export
_features = ['MRAcquisitionType', 'AngioFlag', 'SliceThickness', 'RepetitionTime',
'EchoTime', 'EchoTrainLength', 'PixelSpacing', 'ContrastBolusAgent',
'InversionTime', 'DiffusionBValue', 'seq_E', 'seq_EP', 'seq_G',
'seq_GR', 'seq_I', 'seq_IR', 'seq_M', 'seq_P', 'seq_R', 'seq_S',
'seq_SE', 'var_E', 'var_K', 'var_MP', 'var_MTC', 'var_N', 'var_O',
'var_OSP', 'var_P', 'var_S', 'var_SK', 'var_SP', 'var_SS', 'var_TOF',
'opt_1', 'opt_2', 'opt_A', 'opt_ACC_GEMS', 'opt_B', 'opt_C', 'opt_D',
'opt_E', 'opt_EDR_GEMS', 'opt_EPI_GEMS', 'opt_F', 'opt_FAST_GEMS',
'opt_FC', 'opt_FC_FREQ_AX_GEMS', 'opt_FC_SLICE_AX_GEMS',
'opt_FILTERED_GEMS', 'opt_FR_GEMS', 'opt_FS', 'opt_FSA_GEMS',
'opt_FSI_GEMS', 'opt_FSL_GEMS', 'opt_FSP_GEMS', 'opt_FSS_GEMS', 'opt_G',
'opt_I', 'opt_IFLOW_GEMS', 'opt_IR', 'opt_IR_GEMS', 'opt_L', 'opt_M',
'opt_MP_GEMS', 'opt_MT', 'opt_MT_GEMS', 'opt_NPW', 'opt_P', 'opt_PFF',
'opt_PFP', 'opt_PROP_GEMS', 'opt_R', 'opt_RAMP_IS_GEMS', 'opt_S',
'opt_SAT1', 'opt_SAT2', 'opt_SAT_GEMS', 'opt_SEQ_GEMS', 'opt_SP',
'opt_T', 'opt_T2FLAIR_GEMS', 'opt_TRF_GEMS', 'opt_VASCTOF_GEMS',
'opt_VB_GEMS', 'opt_W', 'opt_X', 'opt__']
#export
def preprocess(df, keepers=_keep, dummies=_dummies, d_prefixes=_d_prefixes, binarize=_binarize, rescale=_rescale):
"Preprocess metadata for Random Forest classifier to predict sequence type"
print("Preprocessing metadata for Random Forest classifier.")
df1 = exclude_other(df)
df1 = df1[keepers]
df1['PixelSpacing'] = df1['PixelSpacing'].apply(lambda x: x[0])
df1 = get_dummies(df1, dummies, d_prefixes)
df1 = make_binary_cols(df1, binarize)
df1 = rescale_cols(df1, rescale)
for f in _features:
if f not in df1.columns:
df1[f] = 0
return df1
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import scanpy as sc
import os
from sklearn.cluster import KMeans
from sklearn.cluster import AgglomerativeClustering
from sklearn.metrics.cluster import adjusted_rand_score
from sklearn.metrics.cluster import adjusted_mutual_info_score
from sklearn.metrics.cluster import homogeneity_score
import rpy2.robjects as robjects
from rpy2.robjects import pandas2ri
df_metrics = pd.DataFrame(columns=['ARI_Louvain','ARI_kmeans','ARI_HC',
'AMI_Louvain','AMI_kmeans','AMI_HC',
'Homogeneity_Louvain','Homogeneity_kmeans','Homogeneity_HC'])
workdir = './output/'
path_fm = os.path.join(workdir,'feature_matrices/')
path_clusters = os.path.join(workdir,'clusters/')
path_metrics = os.path.join(workdir,'metrics/')
os.system('mkdir -p '+path_clusters)
os.system('mkdir -p '+path_metrics)
metadata = pd.read_csv('./input/metadata.tsv',sep='\t',index_col=0)
num_clusters = len(np.unique(metadata['label']))
files = [x for x in os.listdir(path_fm) if x.startswith('FM')]
len(files)
files
def getNClusters(adata,n_cluster,range_min=0,range_max=3,max_steps=20):
this_step = 0
this_min = float(range_min)
this_max = float(range_max)
while this_step < max_steps:
print('step ' + str(this_step))
this_resolution = this_min + ((this_max-this_min)/2)
sc.tl.louvain(adata,resolution=this_resolution)
this_clusters = adata.obs['louvain'].nunique()
print('got ' + str(this_clusters) + ' at resolution ' + str(this_resolution))
if this_clusters > n_cluster:
this_max = this_resolution
elif this_clusters < n_cluster:
this_min = this_resolution
else:
return(this_resolution, adata)
this_step += 1
print('Cannot find the number of clusters')
print('Clustering solution from last iteration is used:' + str(this_clusters) + ' at resolution ' + str(this_resolution))
for file in files:
file_split = file.split('_')
method = file_split[1]
dataset = file_split[2].split('.')[0]
if(len(file_split)>3):
method = method + '_' + '_'.join(file_split[3:]).split('.')[0]
print(method)
pandas2ri.activate()
readRDS = robjects.r['readRDS']
df_rds = readRDS(os.path.join(path_fm,file))
fm_mat = pandas2ri.ri2py(robjects.r['data.frame'](robjects.r['as.matrix'](df_rds)))
fm_mat.columns = metadata.index
adata = sc.AnnData(fm_mat.T)
adata.var_names_make_unique()
adata.obs = metadata.loc[adata.obs.index,]
df_metrics.loc[method,] = ""
#Louvain
sc.pp.neighbors(adata, n_neighbors=15,use_rep='X')
# sc.tl.louvain(adata)
getNClusters(adata,n_cluster=num_clusters)
#kmeans
kmeans = KMeans(n_clusters=num_clusters, random_state=2019).fit(adata.X)
adata.obs['kmeans'] = pd.Series(kmeans.labels_,index=adata.obs.index).astype('category')
#hierachical clustering
hc = AgglomerativeClustering(n_clusters=num_clusters).fit(adata.X)
adata.obs['hc'] = pd.Series(hc.labels_,index=adata.obs.index).astype('category')
#clustering metrics
#adjusted rank index
ari_louvain = adjusted_rand_score(adata.obs['label'], adata.obs['louvain'])
ari_kmeans = adjusted_rand_score(adata.obs['label'], adata.obs['kmeans'])
ari_hc = adjusted_rand_score(adata.obs['label'], adata.obs['hc'])
#adjusted mutual information
ami_louvain = adjusted_mutual_info_score(adata.obs['label'], adata.obs['louvain'],average_method='arithmetic')
ami_kmeans = adjusted_mutual_info_score(adata.obs['label'], adata.obs['kmeans'],average_method='arithmetic')
ami_hc = adjusted_mutual_info_score(adata.obs['label'], adata.obs['hc'],average_method='arithmetic')
#homogeneity
homo_louvain = homogeneity_score(adata.obs['label'], adata.obs['louvain'])
homo_kmeans = homogeneity_score(adata.obs['label'], adata.obs['kmeans'])
homo_hc = homogeneity_score(adata.obs['label'], adata.obs['hc'])
df_metrics.loc[method,['ARI_Louvain','ARI_kmeans','ARI_HC']] = [ari_louvain,ari_kmeans,ari_hc]
df_metrics.loc[method,['AMI_Louvain','AMI_kmeans','AMI_HC']] = [ami_louvain,ami_kmeans,ami_hc]
df_metrics.loc[method,['Homogeneity_Louvain','Homogeneity_kmeans','Homogeneity_HC']] = [homo_louvain,homo_kmeans,homo_hc]
adata.obs[['louvain','kmeans','hc']].to_csv(os.path.join(path_clusters ,method + '_clusters.tsv'),sep='\t')
df_metrics.to_csv(path_metrics+'clustering_scores.csv')
df_metrics
```
| github_jupyter |
# week08: Self-critical Sequence Training
_Reference: based on Practical RL_ [week07](https://github.com/yandexdataschool/Practical_RL/blob/master/week07_seq2seq)
This time we'll solve a problem of transribing hebrew words in english, also known as g2p (grapheme2phoneme)
* word (sequence of letters in source language) -> translation (sequence of letters in target language)
Unlike what most deep learning practicioners do, we won't only train it to maximize likelihood of correct translation, but also employ reinforcement learning to actually teach it to translate with as few errors as possible.
### About the task
One notable property of Hebrew is that it's consonant language. That is, there are no wovels in the written language. One could represent wovels with diacritics above consonants, but you don't expect people to do that in everyay life.
Therefore, some hebrew characters will correspond to several english letters and others - to none, so we should use encoder-decoder architecture to figure that out.
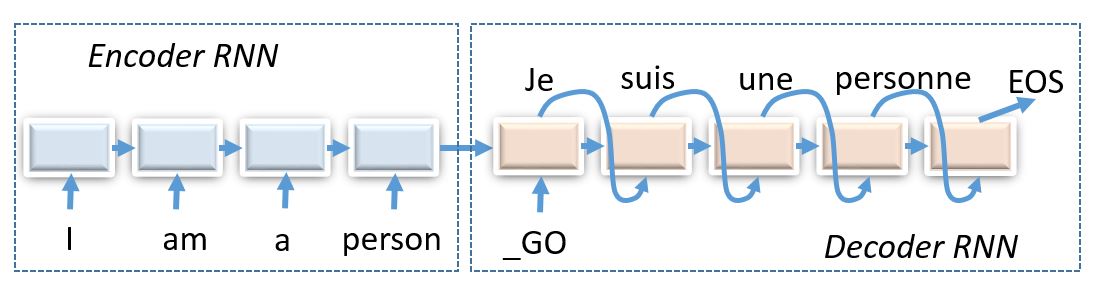
_(img: esciencegroup.files.wordpress.com)_
Encoder-decoder architectures are about converting anything to anything, including
* Machine translation and spoken dialogue systems
* [Image captioning](http://mscoco.org/dataset/#captions-challenge2015) and [image2latex](https://htmlpreview.github.io/?https://github.com/openai/requests-for-research/blob/master/_requests_for_research/im2latex.html) (convolutional encoder, recurrent decoder)
* Generating [images by captions](https://arxiv.org/abs/1511.02793) (recurrent encoder, convolutional decoder)
* Grapheme2phoneme - convert words to transcripts
We chose simplified __Hebrew->English__ machine translation for words and short phrases (character-level), as it is relatively quick to train even without a gpu cluster.
```
# # if running in colab, uncomment
!wget https://raw.githubusercontent.com/girafe-ai/ml-mipt/advanced/week11_RL_outside_games/basic_model_torch.py -O basic_model_torch.py -nc
!wget https://raw.githubusercontent.com/girafe-ai/ml-mipt/advanced/week11_RL_outside_games/main_dataset.txt -O main_dataset.txt -nc
!wget https://raw.githubusercontent.com/girafe-ai/ml-mipt/advanced/week11_RL_outside_games/voc.py -O voc.py -nc
!pip install nltk editdistance
# If True, only translates phrases shorter than 20 characters (way easier).
EASY_MODE = True
# Useful for initial coding.
# If false, works with all phrases (please switch to this mode for homework assignment)
# way we translate. Either "he-to-en" or "en-to-he"
MODE = "he-to-en"
# maximal length of _generated_ output, does not affect training
MAX_OUTPUT_LENGTH = 50 if not EASY_MODE else 20
REPORT_FREQ = 100 # how often to evaluate validation score
```
### Step 1: preprocessing
We shall store dataset as a dictionary
`{ word1:[translation1,translation2,...], word2:[...],...}`.
This is mostly due to the fact that many words have several correct translations.
We have implemented this thing for you so that you can focus on more interesting parts.
__Attention python2 users!__ You may want to cast everything to unicode later during homework phase, just make sure you do it _everywhere_.
```
import numpy as np
from collections import defaultdict
word_to_translation = defaultdict(list) # our dictionary
bos = '_'
eos = ';'
with open("main_dataset.txt", encoding="utf-8") as fin:
for line in fin:
en, he = line[:-1].lower().replace(bos, ' ').replace(eos,
' ').split('\t')
word, trans = (he, en) if MODE == 'he-to-en' else (en, he)
if len(word) < 3:
continue
if EASY_MODE:
if max(len(word), len(trans)) > 20:
continue
word_to_translation[word].append(trans)
print("size = ", len(word_to_translation))
# get all unique lines in source language
all_words = np.array(list(word_to_translation.keys()))
# get all unique lines in translation language
all_translations = np.array(
[ts for all_ts in word_to_translation.values() for ts in all_ts])
```
### split the dataset
We hold out 10% of all words to be used for validation.
```
from sklearn.model_selection import train_test_split
train_words, test_words = train_test_split(
all_words, test_size=0.1, random_state=42)
```
### Building vocabularies
We now need to build vocabularies that map strings to token ids and vice versa. We're gonna need these fellas when we feed training data into model or convert output matrices into english words.
```
from voc import Vocab
inp_voc = Vocab.from_lines(''.join(all_words), bos=bos, eos=eos, sep='')
out_voc = Vocab.from_lines(''.join(all_translations), bos=bos, eos=eos, sep='')
# Here's how you cast lines into ids and backwards.
batch_lines = all_words[:5]
batch_ids = inp_voc.to_matrix(batch_lines)
batch_lines_restored = inp_voc.to_lines(batch_ids)
print("lines")
print(batch_lines)
print("\nwords to ids (0 = bos, 1 = eos):")
print(batch_ids)
print("\nback to words")
print(batch_lines_restored)
```
Draw word/translation length distributions to estimate the scope of the task.
```
import matplotlib.pyplot as plt
%matplotlib inline
plt.figure(figsize=[8, 4])
plt.subplot(1, 2, 1)
plt.title("words")
plt.hist(list(map(len, all_words)), bins=20)
plt.subplot(1, 2, 2)
plt.title('translations')
plt.hist(list(map(len, all_translations)), bins=20)
```
### Step 3: deploy encoder-decoder
__assignment starts here__
Our architecture consists of two main blocks:
* Encoder reads words character by character and outputs code vector (usually a function of last RNN state)
* Decoder takes that code vector and produces translations character by character
Than it gets fed into a model that follows this simple interface:
* __`model(inp, out, **flags) -> logp`__ - takes symbolic int32 matrices of hebrew words and their english translations. Computes the log-probabilities of all possible english characters given english prefices and hebrew word.
* __`model.translate(inp, **flags) -> out, logp`__ - takes symbolic int32 matrix of hebrew words, produces output tokens sampled from the model and output log-probabilities for all possible tokens at each tick.
* if given flag __`greedy=True`__, takes most likely next token at each iteration. Otherwise samples with next token probabilities predicted by model.
That's all! It's as hard as it gets. With those two methods alone you can implement all kinds of prediction and training.
```
import torch
import torch.nn as nn
import torch.nn.functional as F
from basic_model_torch import BasicTranslationModel
model = BasicTranslationModel(inp_voc, out_voc,
emb_size=64, hid_size=256)
device = torch.device('cuda:0') if torch.cuda.is_available() else torch.device('cpu')
model.to(device)
# Play around with symbolic_translate and symbolic_score
inp = torch.tensor(np.random.randint(0, 10, [3, 5]), dtype=torch.int64).to(device)
out = torch.tensor(np.random.randint(0, 10, [3, 5]), dtype=torch.int64).to(device)
# translate inp (with untrained model)
sampled_out, logp = model.translate(inp, greedy=False)
print("Sample translations:\n", sampled_out)
print("Log-probabilities at each step:\n", logp)
# score logp(out | inp) with untrained input
logp = model(inp, out)
print("Symbolic_score output:\n", logp)
print("Log-probabilities of output tokens:\n",
torch.gather(logp, dim=2, index=out[:, :, None]))
def translate(lines, max_len=MAX_OUTPUT_LENGTH, device=device):
"""
You are given a list of input lines.
Make your neural network translate them.
:return: a list of output lines
"""
# Convert lines to a matrix of indices
lines_ix = inp_voc.to_matrix(lines)
lines_ix = torch.tensor(lines_ix, dtype=torch.int64).to(device)
# Compute translations in form of indices
trans_ix, _ = model.translate(lines_ix, greedy=True, max_len=max_len)
# Convert translations back into strings
return out_voc.to_lines(trans_ix.cpu().data.numpy())
print("Sample inputs:", all_words[:3])
print("Dummy translations:", translate(all_words[:3]))
trans = translate(all_words[:3])
assert translate(all_words[:3]) == translate(
all_words[:3]), "make sure translation is deterministic (use greedy=True and disable any noise layers)"
assert type(translate(all_words[:3])) is list and (type(translate(all_words[:1])[0]) is str or type(
translate(all_words[:1])[0]) is unicode), "translate(lines) must return a sequence of strings!"
# note: if translation freezes, make sure you used max_len parameter
print("Tests passed!")
```
### Scoring function
LogLikelihood is a poor estimator of model performance.
* If we predict zero probability once, it shouldn't ruin entire model.
* It is enough to learn just one translation if there are several correct ones.
* What matters is how many mistakes model's gonna make when it translates!
Therefore, we will use minimal Levenshtein distance. It measures how many characters do we need to add/remove/replace from model translation to make it perfect. Alternatively, one could use character-level BLEU/RougeL or other similar metrics.
The catch here is that Levenshtein distance is not differentiable: it isn't even continuous. We can't train our neural network to maximize it by gradient descent.
```
# !pip install editdistance
import editdistance
def get_distance(word, trans):
"""
A function that takes word and predicted translation
and evaluates (Levenshtein's) edit distance to closest correct translation
"""
references = word_to_translation[word]
assert len(references) != 0, "wrong/unknown word"
return min(editdistance.eval(trans, ref) for ref in references)
def score(words, bsize=100):
"""a function that computes levenshtein distance for bsize random samples"""
assert isinstance(words, np.ndarray)
batch_words = np.random.choice(words, size=bsize, replace=False)
batch_trans = translate(batch_words)
distances = list(map(get_distance, batch_words, batch_trans))
return np.array(distances, dtype='float32')
# should be around 5-50 and decrease rapidly after training :)
[score(test_words, 10).mean() for _ in range(5)]
```
### Supervised pre-training
Here we define a function that trains our model through maximizing log-likelihood a.k.a. minimizing crossentropy.
```
import random
def sample_batch(words, word_to_translation, batch_size):
"""
sample random batch of words and random correct translation for each word
example usage:
batch_x,batch_y = sample_batch(train_words, word_to_translations,10)
"""
# choose words
batch_words = np.random.choice(words, size=batch_size)
# choose translations
batch_trans_candidates = list(map(word_to_translation.get, batch_words))
batch_trans = list(map(random.choice, batch_trans_candidates))
return batch_words, batch_trans
bx, by = sample_batch(train_words, word_to_translation, batch_size=3)
print("Source:")
print(bx)
print("Target:")
print(by)
from basic_model_torch import infer_length, infer_mask, to_one_hot
def compute_loss_on_batch(input_sequence, reference_answers, device=device):
""" Compute crossentropy loss given a batch of sources and translations """
input_sequence = torch.tensor(inp_voc.to_matrix(input_sequence), dtype=torch.int64).to(device)
reference_answers = torch.tensor(out_voc.to_matrix(reference_answers), dtype=torch.int64).to(device)
# Compute log-probabilities of all possible tokens at each step. Use model interface.
logprobs_seq = model(input_sequence, reference_answers) # YOUR CODE
# compute elementwise crossentropy as negative log-probabilities of reference_answers.
crossentropy = -torch.sum(to_one_hot(reference_answers, len(out_voc)) * logprobs_seq, dim=-1)
# YOUR CODE
assert crossentropy.dim(
) == 2, "please return elementwise crossentropy, don't compute mean just yet"
# average with mask
mask = infer_mask(reference_answers, out_voc.eos_ix)
loss = torch.sum(crossentropy * mask) / torch.sum(mask)
return loss
# test it
loss = compute_loss_on_batch(*sample_batch(train_words, word_to_translation, 3))
print('loss = ', loss)
assert loss.item() > 0.0
loss.backward()
for w in model.parameters():
assert w.grad is not None and torch.max(torch.abs(w.grad)).item() != 0, \
"Loss is not differentiable w.r.t. a weight with shape %s. Check comput_loss_on_batch." % (
w.size(),)
print('Seems fine!')
```
##### Actually train the model
Minibatches and stuff...
```
from IPython.display import clear_output
from tqdm import tqdm, trange # or use tqdm_notebook,tnrange
loss_history = []
editdist_history = []
entropy_history = []
opt = torch.optim.Adam(model.parameters())
for i in trange(5000):
loss = compute_loss_on_batch(*sample_batch(train_words, word_to_translation, 32))
# train with backprop
loss.backward()
opt.step()
opt.zero_grad()
loss_history.append(loss.item())
if (i+1) % REPORT_FREQ == 0:
clear_output(True)
current_scores = score(test_words)
editdist_history.append(current_scores.mean())
print("llh=%.3f, mean score=%.3f" %
(np.mean(loss_history[-10:]), np.mean(editdist_history[-10:])))
plt.figure(figsize=(12, 4))
plt.subplot(131)
plt.title('train loss / traning time')
plt.plot(loss_history)
plt.grid()
plt.subplot(132)
plt.title('val score distribution')
plt.hist(current_scores, bins=20)
plt.subplot(133)
plt.title('val score / traning time (lower is better)')
plt.plot(editdist_history)
plt.grid()
plt.show()
```
__How to interpret the plots:__
* __Train loss__ - that's your model's crossentropy over minibatches. It should go down steadily. Most importantly, it shouldn't be NaN :)
* __Val score distribution__ - distribution of translation edit distance (score) within batch. It should move to the left over time.
* __Val score / training time__ - it's your current mean edit distance. This plot is much whimsier than loss, but make sure it goes below 8 by 2500 steps.
If it doesn't, first try to re-create both model and opt. You may have changed it's weight too much while debugging. If that doesn't help, it's debugging time.
```
for word in train_words[:10]:
print("%s -> %s" % (word, translate([word])[0]))
test_scores = []
for start_i in trange(0, len(test_words), 32):
batch_words = test_words[start_i:start_i+32]
batch_trans = translate(batch_words)
distances = list(map(get_distance, batch_words, batch_trans))
test_scores.extend(distances)
print("Supervised test score:", np.mean(test_scores))
```
### Self-critical policy gradient
In this section you'll implement algorithm called self-critical sequence training (here's an [article](https://arxiv.org/abs/1612.00563)).
The algorithm is a vanilla policy gradient with a special baseline.
$$ \nabla J = E_{x \sim p(s)} E_{y \sim \pi(y|x)} \nabla log \pi(y|x) \cdot (R(x,y) - b(x)) $$
Here reward R(x,y) is a __negative levenshtein distance__ (since we minimize it). The baseline __b(x)__ represents how well model fares on word __x__.
In practice, this means that we compute baseline as a score of greedy translation, $b(x) = R(x,y_{greedy}(x)) $.

Luckily, we already obtained the required outputs: `model.greedy_translations, model.greedy_mask` and we only need to compute levenshtein using `compute_levenshtein` function.
```
def compute_reward(input_sequence, translations):
""" computes sample-wise reward given token ids for inputs and translations """
distances = list(map(get_distance,
inp_voc.to_lines(input_sequence.data.numpy()),
out_voc.to_lines(translations.data.numpy())))
# use negative levenshtein distance so that larger reward means better policy
return - torch.tensor(distances, dtype=torch.int64)
def scst_objective_on_batch(input_sequence, max_len=MAX_OUTPUT_LENGTH, device=device):
""" Compute pseudo-loss for policy gradient given a batch of sources """
input_sequence = torch.tensor(inp_voc.to_matrix(input_sequence), dtype=torch.int64).to(device)
# use model to __sample__ symbolic translations given input_sequence
sample_translations, sample_logp = model.translate(input_sequence, greedy=False, max_len=max_len) # YOUR CODE
# use model to __greedy__ symbolic translations given input_sequence
greedy_translations, greedy_logp = model.translate(input_sequence, greedy=True, max_len=max_len) # YOUR CODE
# compute rewards and advantage
# be careful with the device, rewards require casting to numpy, so send everything to cpu
rewards = compute_reward(input_sequence.cpu(), sample_translations.cpu())
baseline = compute_reward(input_sequence.cpu(), greedy_translations.cpu()) # <compute __negative__ levenshtein for greedy mode >
# compute advantage using rewards and baseline
# be careful with the device, advantage is used to compute gradients, so send it to device
advantage = (rewards - baseline).float().to(device) # YOUR CODE
# compute log_pi(a_t|s_t), shape = [batch, seq_length]
logp_sample = torch.sum(to_one_hot(sample_translations, n_dims=len(out_voc)) * sample_logp, dim=-1)
# YOUR CODE
# ^-- hint: look at how crossentropy is implemented in supervised learning loss above
# mind the sign - this one should not be multiplied by -1 :)
# policy gradient pseudo-loss. Gradient of J is exactly policy gradient.
J = logp_sample * advantage[:, None]
assert J.dim() == 2, "please return elementwise objective, don't compute mean just yet"
# average with mask
mask = infer_mask(sample_translations, out_voc.eos_ix)
loss = - torch.sum(J * mask) / torch.sum(mask)
# regularize with negative entropy. Don't forget the sign!
# note: for entropy you need probabilities for all tokens (sample_logp), not just logp_sample
entropy = -torch.sum(torch.exp(sample_logp) * sample_logp, dim=-1)
# <compute entropy matrix of shape[batch, seq_length], H = -sum(p*log_p), don't forget the sign!>
# hint: you can get sample probabilities from sample_logp using math :)
assert entropy.dim(
) == 2, "please make sure elementwise entropy is of shape [batch,time]"
reg = - 0.01 * torch.sum(entropy * mask) / torch.sum(mask)
return loss + reg, torch.sum(entropy * mask) / torch.sum(mask)
```
### Policy gradient training
```
entropy_history = [np.nan] * len(loss_history)
opt = torch.optim.Adam(model.parameters(), lr=1e-5)
for i in trange(100000):
loss, ent = scst_objective_on_batch(
sample_batch(train_words, word_to_translation, 32)[0]) # [0] = only source sentence
# train with backprop
loss.backward()
opt.step()
opt.zero_grad()
loss_history.append(loss.item())
entropy_history.append(ent.item())
if (i+1) % REPORT_FREQ == 0:
clear_output(True)
current_scores = score(test_words)
editdist_history.append(current_scores.mean())
plt.figure(figsize=(12, 4))
plt.subplot(131)
plt.title('val score distribution')
plt.hist(current_scores, bins=20)
plt.subplot(132)
plt.title('val score / traning time')
plt.plot(editdist_history)
plt.grid()
plt.subplot(133)
plt.title('policy entropy / traning time')
plt.plot(entropy_history)
plt.grid()
plt.show()
print("J=%.3f, mean score=%.3f" %
(np.mean(loss_history[-10:]), np.mean(editdist_history[-10:])))
```
__Debugging tips:__

* As usual, don't expect improvements right away, but in general the model should be able to show some positive changes by 5k steps.
* Entropy is a good indicator of many problems.
* If it reaches zero, you may need greater entropy regularizer.
* If it has rapid changes time to time, you may need gradient clipping.
* If it oscillates up and down in an erratic manner... it's perfectly okay for entropy to do so. But it should decrease at the end.
* We don't show loss_history cuz it's uninformative for pseudo-losses in policy gradient. However, if something goes wrong you can check it to see if everything isn't a constant zero.
### Results
```
for word in train_words[:10]:
print("%s -> %s" % (word, translate([word])[0]))
test_scores = []
for start_i in trange(0, len(test_words), 32):
batch_words = test_words[start_i:start_i+32]
batch_trans = translate(batch_words)
distances = list(map(get_distance, batch_words, batch_trans))
test_scores.extend(distances)
print("Supervised test score:", np.mean(test_scores))
# ^^ If you get Out Of MemoryError, please replace this with batched computation
```
### Bonus area: Make it actually work
In this section we want you to finally __restart with EASY_MODE=False__ and experiment to find a good model/curriculum for that task.
We recommend you to start with the following architecture
```
encoder---decoder
P(y|h)
^
LSTM -> LSTM
^ ^
biLSTM -> LSTM
^ ^
input y_prev
```
__Note:__ you can fit all 4 state tensors of both LSTMs into a in a single state - just assume that it contains, for example, [h0, c0, h1, c1] - pack it in encode and update in decode.
Here are some cool ideas on what you can do then.
__General tips & tricks:__
* You will likely need to adjust pre-training time for such a network.
* Supervised pre-training may benefit from clipping gradients somehow.
* SCST may indulge a higher learning rate in some cases and changing entropy regularizer over time.
* It's often useful to save pre-trained model parameters to not re-train it every time you want new policy gradient parameters.
* When leaving training for nighttime, try setting REPORT_FREQ to a larger value (e.g. 500) not to waste time on it.
__Formal criteria:__
Build an architecture that:
* _doesn't consist of single GRU_
* _works better_ than single GRU baseline.
* We also want you to provide either learning curve or trained model, preferably both
* ... and write a brief report or experiment log describing what you did and how it fared.
### Attention
There's more than one way to connect decoder to encoder
* __Vanilla:__ layer_i of encoder last state goes to layer_i of decoder initial state
* __Every tick:__ feed encoder last state _on every iteration_ of decoder.
* __Attention:__ allow decoder to "peek" at one (or several) positions of encoded sequence on every tick.
The most effective (and cool) of those is, of course, attention.
You can read more about attention [in this nice blog post](https://distill.pub/2016/augmented-rnns/). The easiest way to begin is to use "soft" attention with "additive" or "dot-product" intermediate layers.
__Tips__
* Model usually generalizes better if you no longer allow decoder to see final encoder state
* Once your model made it through several epochs, it is a good idea to visualize attention maps to understand what your model has actually learned
* There's more stuff [here](https://github.com/yandexdataschool/Practical_RL/blob/master/week8_scst/bonus.ipynb)
* If you opted for hard attention, we recommend [gumbel-softmax](https://blog.evjang.com/2016/11/tutorial-categorical-variational.html) instead of sampling. Also please make sure soft attention works fine before you switch to hard.
### UREX
* This is a way to improve exploration in policy-based settings. The main idea is that you find and upweight under-appreciated actions.
* Here's [video](https://www.youtube.com/watch?v=fZNyHoXgV7M&feature=youtu.be&t=3444)
and an [article](https://arxiv.org/abs/1611.09321).
* You may want to reduce batch size 'cuz UREX requires you to sample multiple times per source sentence.
* Once you got it working, try using experience replay with importance sampling instead of (in addition to) basic UREX.
### Some additional ideas:
* (advanced deep learning) It may be a good idea to first train on small phrases and then adapt to larger ones (a.k.a. training curriculum).
* (advanced nlp) You may want to switch from raw utf8 to something like unicode or even syllables to make task easier.
* (advanced nlp) Since hebrew words are written __with vowels omitted__, you may want to use a small Hebrew vowel markup dataset at `he-pron-wiktionary.txt`.
```
assert not EASY_MODE, "make sure you set EASY_MODE = False at the top of the notebook."
```
`[your report/log here or anywhere you please]`
__Contributions:__ This notebook is brought to you by
* Yandex [MT team](https://tech.yandex.com/translate/)
* Denis Mazur ([DeniskaMazur](https://github.com/DeniskaMazur)), Oleg Vasilev ([Omrigan](https://github.com/Omrigan/)), Dmitry Emelyanenko ([TixFeniks](https://github.com/tixfeniks)) and Fedor Ratnikov ([justheuristic](https://github.com/justheuristic/))
* Dataset is parsed from [Wiktionary](https://en.wiktionary.org), which is under CC-BY-SA and GFDL licenses.
| github_jupyter |
# Relativistic kinematics
<h3>Learning goals</h3>
<ul>
<li>Relativistic kinematics.
<li>Standard model particles.
</ul>
<b>Background</b>
If you know the mass of a particle, most of the time you know <i>what that particle is</i>. However, there is no way to just build a single detector that gives you the mass. You need to be clever and make use of Special relativity, specifically <a href="http://en.wikipedia.org/wiki/Relativistic_mechanics">relativistic kinematics</a>.
To determine the mass ($m$) of a particle you need to know the 4-momenta of the particles ($\mathbf{P}$) that are detected after the collision: the energy ($E$), the momentum in the x direction ($p_x$), the momentum in the y direction ($p_y$), the momentum in the z direction ($p_z$).
$$\mathbf{P} = (E,p_x,p_y,p_z)$$
\begin{equation*} m = \sqrt{E^2-(p_x^2+p_y^2 + p_z^2)} \end{equation*}
<b>Let's code!</b>
Here is some sample code that reads in data from a small sample file from the <a href = "https://cms.cern">CMS experiment</a>. It loops over data from many different proton-proton collisions.
If you haven't already, you will want to go through the [data model](https://github.com/particle-physics-playground/playground/blob/master/activities/codebkg_DataInterfacing.ipynb) (also included when you cloned this directory) exercise so you know how to pull out the relevant information.
For each collision, you can get the 4-momenta of the jets, muons, electrons, and photons produced in these collisions.
```
%matplotlib notebook
import pandas as pd
import tools.cms as cms
import h5py
filename = './data/dimuons_1000_collisions.hdf5'
filehandler = h5py.File(filename, 'r')
print("Keys: ")
for i, key in enumerate(filehandler.keys(), 1):
print(f"{i:>2d}. {key}")
filehandler["datasets_and_counters"][()]
```
<h2><font color="red">Challenge!</font></h2>
Copy this sample code and use it to calculate the mass of the muons. Make a histogram of this quantity.
<i>Hint!</i>
Make sure you do this for all the muons! Each collision can produce differing numbers of muons, so take care when you code this up.
Your histogram should look something like the following sketch, though the peak will be at different values.
The value of the peak, should be the mass of the particle <a href="http://en.wikipedia.org/wiki/Muon">Check your answer!</a>
You should also make histograms of the energy and magnitude of momentum ($|p|$). You should see a pretty wide range of values for these, and yet the mass is a very specific number.
```
from IPython.display import Image
Image(filename='images/muons_sketch.jpeg')
# Your code here
```
Suppose we didn't know anything about special relativity and we tried calculating the mass from what we know about classical physics.
$$KE = \frac{1}{2}mv^2 \qquad KE = \frac{p^2}{2m} \qquad m = \frac{p^2}{2KE}$$
Let's interpret the energy from the CMS data as the kinetic energy ($KE$). Use classical mechanics then to calculate the mass of the muon, given the energy/KE and the momentum. What does <b>that</b> histogram look like?
*Your histogram should not look like the last one! We know that the Classical description of kinematics is not accurate for particle moving at high energies, so don't worry if the two histograms are different. That's the point! :)*
```
# Your code here
```
| github_jupyter |
# QuickSort
Like MergeSort, QuickSort is a divide-and-conquer algorithm. We need to pick a pivot, then sort both sublists that are created on either side of the pivot. Similar to the video, we'll follow the convention of picking the last element as the pivot.
Start with our unordered list of items:









```
items = [8, 3, 1, 7, 0, 10, 2]
```
Let's sketch out what a first iteration would look like.
We can use `len` to grab the pivot value, but in order to sort in-place we'll also want the index of the pivot.
```
pivot_index = len(items) - 1
pivot_value = items[pivot_index]
```
Because we plan on sorting in-place, we want to iterate through the items to the left of our pivot (`left_items`). When they're larger than `pivot_value` though, we will not increment our position through `left_items`, but instead change `pivot_index`. We'll know we're done with this pass when `pivot_index` and `left_items` index are equal.
```
left_index = 0
while (pivot_index != left_index):
item = items[left_index]
if item <= pivot_value:
left_index += 1
continue
# Place the item before the pivot at left_index
items[left_index] = items[pivot_index - 1]
# Shift pivot one to the left
items[pivot_index - 1] = pivot_value
# Place item at pivot's previous location
items[pivot_index] = item
# Update pivot_index
pivot_index -= 1
print(items)
```
You should see:
```
[0, 1, 2, 7, 3, 10, 8]
```
When our loop terminated, we knew everything to the left of our pivot was less than pivot, and everything to the right of our pivot was greater than pivot. Great! Now we need to do that again for the sublists that are left and right of pivot's final location.
We can do that by abstracting our above code to a function, just passing the list of items as a parameter.
```
def sort_a_little_bit(items):
left_index = 0
pivot_index = len(items) - 1
pivot_value = items[pivot_index]
while (pivot_index != left_index):
item = items[left_index]
if item <= pivot_value:
left_index += 1
continue
items[left_index] = items[pivot_index - 1]
items[pivot_index - 1] = pivot_value
items[pivot_index] = item
pivot_index -= 1
items = [8, 3, 1, 7, 0, 10, 2]
sort_a_little_bit(items)
print(items)
```
Now what would it require to recurse on this? We want to take the result of that iteration and act on it. So first off, we see that in order to call the function again, we need to communicate the final `pivot_index` value. And then with that, we can mark off segments of the list and have our function operate on less than the entire list. So let's change our function to accept the indices it should stay within, and return the pivot_index.
```
def sort_a_little_bit(items, begin_index, end_index):
left_index = begin_index
pivot_index = end_index
pivot_value = items[pivot_index]
while (pivot_index != left_index):
item = items[left_index]
if item <= pivot_value:
left_index += 1
continue
items[left_index] = items[pivot_index - 1]
items[pivot_index - 1] = pivot_value
items[pivot_index] = item
pivot_index -= 1
return pivot_index
items = [8, 3, 1, 7, 0, 10, 2]
pivot_index = sort_a_little_bit(items, 0, len(items) - 1)
print(items)
print('pivot index %d' % pivot_index)
```
Almost there! Let's create another function, the recursive function we want, that uses this. And then we'll have our top level definition of `quicksort` call it with our initial parameters. But we need a way to know if we're done! We'll use the indices to see if they demark a list of more than one item. If the demarked sublist is 0 or 1 item, we know it's already sorted.
```
def sort_all(items, begin_index, end_index):
if end_index <= begin_index:
return
pivot_index = sort_a_little_bit(items, begin_index, end_index)
sort_all(items, begin_index, pivot_index - 1)
sort_all(items, pivot_index + 1, end_index)
def quicksort(items):
sort_all(items, 0, len(items) - 1)
items = [8, 3, 1, 7, 0, 10, 2]
quicksort(items)
print(items)
```
It's a good idea to test a few more scenarios. Does it work with an even number of items? What if they're already sorted?
```
items = [1, 0]
quicksort(items)
print(items)
items = [96, 97, 98]
quicksort(items)
print(items)
```
### Mission Accomplished!
| github_jupyter |
# Introduction to NumPy
The learning objectives of this section are:
* Understand advantages of vectorized code using Numpy (over standard python ways)
* Create NumPy arrays
* Convert lists and tuples to numpy arrays
* Create (initialise) arrays
* Inspect the structure and content of arrays
* Subset, slice, index and iterate through arrays
* Compare computation times in NumPy and standard Python lists
### NumPy Basics
NumPy is a library written for scientific computing and data analysis. It stands for numerical python.
The most basic object in NumPy is the ```ndarray```, or simply an ```array```, which is an **n-dimensional, homogenous** array. By homogenous, we mean that all the elements in a numpy array have to be of the **same data type**, which is commonly numeric (float or integer).
Let's see some examples of arrays.
```
# Import the numpy library
# np is simply an alias, you may use any other alias, though np is quite standard
import numpy as np
# Creating a 1-D array using a list
# np.array() takes in a list or a tuple as argument, and converts into an array
array_1d = np.array([2, 4, 5, 6, 7, 9])
print(array_1d)
print(type(array_1d))
# Creating a 2-D array using two lists
array_2d = np.array([[2, 3, 4], [5, 8, 7]])
print(array_2d)
```
In NumPy, dimensions are called **axes**. In the 2-d array above, there are two axes, having two and three elements respectively.
In Numpy terminology, for 2-D arrays:
* ```axis = 0``` refers to the rows
* ```axis = 1``` refers to the columns
<img src="numpy_axes.jpg" style="width: 600px; height: 400px">
### Advantages of NumPy
What is the use of arrays over lists, specifically for data analysis? Putting crudely, it is **convenience and speed **:<br>
1. You can write **vectorised** code on numpy arrays, not on lists, which is **convenient to read and write, and concise**.
2. Numpy is **much faster** than the standard python ways to do computations.
Vectorised code typically does not contain explicit looping and indexing etc. (all of this happens behind the scenes, in precompiled C-code), and thus it is much more concise.
Let's see an example of convenience, we'll see one later for speed.
Say you have two lists of numbers, and want to calculate the element-wise product. The standard python list way would need you to map a lambda function (or worse - write a ```for``` loop), whereas with NumPy, you simply multiply the arrays.
```
list_1 = [3, 6, 7, 5]
list_2 = [4, 5, 1, 7]
# the list way to do it: map a function to the two lists
product_list = list(map(lambda x, y: x*y, list_1, list_2))
print(product_list)
# The numpy array way to do it: simply multiply the two arrays
array_1 = np.array(list_1)
array_2 = np.array(list_2)
array_3 = array_1*array_2
print(array_3)
print(type(array_3))
```
As you can see, the numpy way is clearly more concise.
Even simple mathematical operations on lists require for loops, unlike with arrays. For example, to calculate the square of every number in a list:
```
# Square a list
list_squared = [i**2 for i in list_1]
# Square a numpy array
array_squared = array_1**2
print(list_squared)
print(array_squared)
```
This was with 1-D arrays. You'll often work with 2-D arrays (matrices), where the difference would be even greater. With lists, you'll have to store matrices as lists of lists and loop through them. With NumPy, you simply multiply the matrices.
### Creating NumPy Arrays
There are multiple ways to create numpy arrays, the most commmon ones being:
* Convert lists or tuples to arrays using ```np.array()```, as done above
* Initialise arrays of fixed size (when the size is known)
```
# Convert lists or tuples to arrays using np.array()
# Note that np.array(2, 5, 6, 7) will throw an error - you need to pass a list or a tuple
array_from_list = np.array([2, 5, 6, 7])
array_from_tuple = np.array((4, 5, 8, 9))
print(array_from_list)
print(array_from_tuple)
```
The other common way is to initialise arrays. You do this when you know the size of the array beforehand.
The following ways are commonly used:
* ```np.ones()```: Create array of 1s
* ```np.zeros()```: Create array of 0s
* ```np.random.random()```: Create array of random numbers
* ```np.arange()```: Create array with increments of a fixed step size
* ```np.linspace()```: Create array of fixed length
```
# Tip: Use help to see the syntax when required
help(np.ones)
# Creating a 5 x 3 array of ones
np.ones((5, 3))
# Notice that, by default, numpy creates data type = float64
# Can provide dtype explicitly using dtype
np.ones((5, 3), dtype = np.int)
# Creating array of zeros
np.zeros(4, dtype = np.int)
# Array of random numbers
np.random.random([3, 4])
# np.arange()
# np.arange() is the numpy equivalent of range()
# Notice that 10 is included, 100 is not, as in standard python lists
# From 10 to 100 with a step of 5
numbers = np.arange(10, 100, 5)
print(numbers)
# np.linspace()
# Sometimes, you know the length of the array, not the step size
# Array of length 25 between 15 and 18
np.linspace(15, 18, 25)
```
### Inspect the Structure and Content of Arrays
It is helpful to inspect the structure of numpy arrays, especially while working with large arrays. Some attributes of numpy arrays are:
* ```shape```: Shape of array (n x m)
* ```dtype```: data type (int, float etc.)
* ```ndim```: Number of dimensions (or axes)
* ```itemsize```: Memory used by each array elememnt in bytes
Let's say you are working with a moderately large array of size 1000 x 300. First, you would want to wrap your head around the basic shape and size of the array.
```
# Initialising a random 1000 x 300 array
rand_array = np.random.random((1000, 300))
# Print the first row
print(rand_array[1, ])
# Inspecting shape, dtype, ndim and itemsize
print("Shape: {}".format(rand_array.shape))
print("dtype: {}".format(rand_array.dtype))
print("Dimensions: {}".format(rand_array.ndim))
print("Item size: {}".format(rand_array.itemsize))
```
Reading 3-D arrays is not very obvious, because we can only print maximum two dimensions on paper, and thus they are printed according to a specific convention. Printing higher dimensional arrays follows the following conventions:
* The last axis is printed from left to right
* The second-to-last axis is printed from top to bottom
* The other axes are also printed top-to-bottom, with each slice separated by another using an empty line
Let's see some examples.
```
# Creating a 3-D array
# reshape() simply reshapes a 1-D array
array_3d = np.arange(24).reshape(2, 3, 4)
print(array_3d)
```
* The last axis has 4 elements, and is printed from left to right.
* The second last has 3, and is printed top to bottom
* The other axis has 2, and is printed in the two separated blocks
### Subset, Slice, Index and Iterate through Arrays
For **one-dimensional arrays**, indexing, slicing etc. is **similar to python lists** - indexing starts at 0.
```
# Indexing and slicing one dimensional arrays
array_1d = np.arange(10)
print(array_1d)
# Third element
print(array_1d[2])
# Specific elements
# Notice that array[2, 5, 6] will throw an error, you need to provide the indices as a list
print(array_1d[[2, 5, 6]])
# Slice third element onwards
print(array_1d[2:])
# Slice first three elements
print(array_1d[:3])
# Slice third to seventh elements
print(array_1d[2:7])
# Subset starting 0 at increment of 2
print(array_1d[0::2])
# Iterations are also similar to lists
for i in array_1d:
print(i**2)
```
**Multidimensional arrays** are indexed using as many indices as the number of dimensions or axes. For instance, to index a 2-D array, you need two indices - ```array[x, y]```.
Each axes has an index starting at 0. The following figure shows the axes and their indices for a 2-D array.
<img src="2_d_array.png" style="width: 350px; height: 300px">
```
# Creating a 2-D array
array_2d = np.array([[2, 5, 7, 5], [4, 6, 8, 10], [10, 12, 15, 19]])
print(array_2d)
# Third row second column
print(array_2d[2, 1])
# Slicing the second row, and all columns
# Notice that the resultant is itself a 1-D array
print(array_2d[1, :])
print(type(array_2d[1, :]))
# Slicing all rows and the third column
print(array_2d[:, 2])
# Slicing all rows and the first three columns
print(array_2d[:, :3])
```
**Iterating on 2-D arrays** is done with respect to the first axis (which is row, the second axis is column).
```
# Iterating over 2-D arrays
for row in array_2d:
print(row)
# Iterating over 3-D arrays: Done with respect to the first axis
array_3d = np.arange(24).reshape(2, 3, 4)
print(array_3d)
# Prints the two blocks
for row in array_3d:
print(row)
```
### Compare Computation Times in NumPy and Standard Python Lists
We mentioned that the key advantages of numpy are convenience and speed of computation.
You'll often work with extremely large datasets, and thus it is important point for you to understand how much computation time (and memory) you can save using numpy, compared to standard python lists.
Let's compare the computation times of arrays and lists for a simple task of calculating the element-wise product of numbers.
```
## Comparing time taken for computation
list_1 = [i for i in range(1000000)]
list_2 = [j**2 for j in range(1000000)]
# list multiplication
import time
# store start time, time after computation, and take the difference
t0 = time.time()
product_list = list(map(lambda x, y: x*y, list_1, list_2))
t1 = time.time()
list_time = t1 - t0
print(t1-t0)
# numpy array
array_1 = np.array(list_1)
array_2 = np.array(list_2)
t0 = time.time()
array_3 = array_1*array_2
t1 = time.time()
numpy_time = t1 - t0
print(t1-t0)
print("The ratio of time taken is {}".format(list_time/numpy_time))
```
In this case, numpy is **an order of magnitude faster** than lists. This is with arrays of size in millions, but you may work on much larger arrays of sizes in order of billions. Then, the difference is even larger.
Some reasons for such difference in speed are:
* NumPy is written in C, which is basically being executed behind the scenes
* NumPy arrays are more compact than lists, i.e. they take much lesser storage space than lists
The following discussions demonstrate the differences in speeds of NumPy and standard python:
1. https://stackoverflow.com/questions/8385602/why-are-numpy-arrays-so-fast
2. https://stackoverflow.com/questions/993984/why-numpy-instead-of-python-lists
| github_jupyter |
# Starbucks Capstone Challenge: Customer Segmentation
### Introduction
This data set contains simulated data that mimics customer behavior on the Starbucks rewards mobile app. Once every few days, Starbucks sends out an offer to users of the mobile app. An offer can be merely an advertisement for a drink or an actual offer such as a discount or BOGO (buy one get one free). Some users might not receive any offer during certain weeks.
In this notebook we will attempt to divide our customers in segments and evaluate our decisions.
# Data Sets
The data is contained in three files:
* portfolio.json - containing offer ids and meta data about each offer (duration, type, etc.)
* profile.json - demographic data for each customer
* transcript.json - records for transactions, offers received, offers viewed, and offers completed
Here is the schema and explanation of each variable in the files:
**portfolio.json**
* id (string) - offer id
* offer_type (string) - type of offer ie BOGO, discount, informational
* difficulty (int) - minimum required spend to complete an offer
* reward (int) - reward given for completing an offer
* duration (int) - time for offer to be open, in days
* channels (list of strings)
**profile.json**
* age (int) - age of the customer
* became_member_on (int) - date when customer created an app account
* gender (str) - gender of the customer (note some entries contain 'O' for other rather than M or F)
* id (str) - customer id
* income (float) - customer's income
**transcript.json**
* event (str) - record description (ie transaction, offer received, offer viewed, etc.)
* person (str) - customer id
* time (int) - time in hours since start of test. The data begins at time t=0
* value - (dict of strings) - either an offer id or transaction amount depending on the record
# Import Packages
```
#import holy grail of data science
import pandas as pd
import numpy as np
#helper imports
from cleaner_help import transcript_cleaner
#more imports
import math
import json
import matplotlib.pyplot as plt
import pickle
import seaborn as sns
from scipy import stats
from sklearn.preprocessing import Imputer , MinMaxScaler
from sklearn.decomposition import PCA
from tqdm import tqdm
from sklearn.cluster import KMeans
from sklearn.cluster import MiniBatchKMeans
from sklearn.metrics import silhouette_score
#matplot magic
%matplotlib inline
```
# Load Data
```
user_df = pd.read_csv("data/user_df.csv")
user_df_median = pd.read_csv("data/user_df_median.csv")
user_df.head()
```
# Indentifying Customer Segments
## Reduce Dimensionality
```
# Apply feature scaling
scaler=MinMaxScaler()
#fit scaler
scaler.fit(user_df_median)
#transformed scaled data
user_scaled=scaler.transform(user_df_median)
# Apply PCA to the data
pca_pre= PCA()
pca_pre.fit(user_scaled)
# Investigate the variance accounted for by each principal component.
pca_pre.explained_variance_ratio_[:10]
#store cumulative sume of explained variance ratio
cumsum=np.cumsum(pca_pre.explained_variance_ratio_)
#find how many components account for 95% of the variance
pca_95= np.argmax(cumsum>0.95)
print(pca_95)
plt.figure(figsize=(8,6))
plt.plot(np.cumsum(pca_pre.explained_variance_ratio_))
plt.xlabel('Number of Components')
plt.ylabel('cumulative explained variance')
plt.title('Cumulative Explained Variance')
plt.xticks(np.arange(0,len(pca_pre.explained_variance_ratio_), 1))
plt.yticks(np.arange(0,1.1,0.1))
plt.grid(b=True)
plt.savefig('plots/pca_cumsum_pre.png')
plt.show()
# Re-apply PCA to the data while selecting for number of components to retain.
pca=PCA(n_components=pca_95)
pca.fit(user_scaled)
user_pca=pca.transform(user_scaled)
user_pca.shape
```
## Principal Components
```
# Map weights for the first principal component to corresponding feature names
# and then print the linked values, sorted by weight.
#index dimensions
dimensions = ['Dimension {}'.format(i) for i in range(1,len(pca.components_)+1)]
#PCA componenets
components= pd.DataFrame(np.round(pca.components_, 4), columns=user_df.keys().tolist())
components.index=dimensions
components=components.T
components.head()
def comp_weights(components, comp=0, topk=5, saveloc=None):
#get sorted weights for dimension
#note that comp 0 corresponds to dimension 1
dim=components.copy()
dim=dim.iloc[:,comp]
#dim_sort= np.sort(dim)
dim_sort=dim.sort_values(ascending=True)
#plot the feature weights as a function of the components
plt.figure(figsize=(20,10))
plt.bar(x=dim_sort.index, height=dim_sort, tick_label=dim_sort.index)
plt.xlabel("Features")
plt.xticks(rotation=90, fontsize=5)
plt.ylabel("Weights")
plt.title("Weights of Dimension {}".format(comp+1))
plt.savefig("plots/expl_var"+str(comp+1)+'.png')
if saveloc:
plt.savefig(saveloc)
plt.show()
print("{} most positive weights for the {} component: \n".format(topk, comp+1))
print(dim_sort[-5:], '\n')
print("{} most negative weights for the {} component: \n".format(topk, comp+1))
print(dim_sort[0:5])
comp_weights(components, 0, saveloc='plots/dim_1.png')
comp_weights(components, 1,saveloc='plots/dim_2.png')
comp_weights(components, 2,saveloc='plots/dim_3.png')
```
## Clustering
```
def minik_ssd(df, batch_size=100, K=range(1,30)):
'''
Peforms MiniBatchKMeans on dataframe with user specified batch size & range of clusters
Input:
- Dataframe
- Batch size
- Range of clusters
Output:
- Sum of squared distances over range
'''
#store evolution of ssd
ssd=[]
#put fancy progress bar on number of clusters
for k in tqdm(K):
# intantiate a mini kmeans
kmeans_mini=MiniBatchKMeans(n_clusters=k, batch_size=batch_size)
kmeans_mini.fit(df)
# compute the average within-cluster distances.
ssd.append(kmeans_mini.inertia_)
return ssd
K=range(1,41)
###get ssd for arbitrary batches
ssd_100=minik_ssd(user_pca, batch_size=100, K=K)
ssd_1000=minik_ssd(user_pca, batch_size=1000, K=K)
ssd_10000=minik_ssd(user_pca, batch_size=10000, K=K)
ssd_20000=minik_ssd(user_pca, batch_size=20000, K=K)
ssd_50000=minik_ssd(user_pca, batch_size=50000, K=K)
# Investigate the change in within-cluster distance across number of clusters.
# Use matplotlib's plot function to visualize this relationship.
plt.figure(figsize=(8,8))
plt.plot(K, np.log(ssd_100), marker='o') #apply natural log to refuce magnitude
plt.plot(K, np.log(ssd_1000), marker='+')
plt.plot(K, np.log(ssd_10000), marker='x')
plt.plot(K, np.log(ssd_20000), marker='.')
plt.plot(K, np.log(ssd_50000), marker='*')
plt.legend(['100', '1000', '10000', '20000', '50000'], loc='upper right')
plt.xlabel('Number of Clusters')
plt.ylabel('Log of Sum of Squared Distances')
plt.title('Elbow Method: Sum of Squared Distances Vs Number of Clusters')
plt.grid(b=True)
plt.savefig('plots/elbow_minik.png')
plt.show()
# Re-fit the k-means model with the selected number of clusters and obtain
# cluster predictions for the general population demographics data.
best_k=15
kmeans=KMeans(n_clusters=best_k)
kmeans.fit(user_pca)
with open('data/kmeans.pickle', 'wb') as f:
# Pickle the 'data' dictionary using the highest protocol available.
pickle.dump(kmeans, f, pickle.HIGHEST_PROTOCOL)
#get kmeans predictions
kmean_pred = kmeans.fit_predict(user_pca)
#get centroids
centers = kmeans.cluster_centers_
# get silhouette score
silh_score = silhouette_score (user_pca, kmean_pred, metric='euclidean')
print("For {} clusters our silhouette score is {:.3f}".format(best_k, silh_score))
```
## Visualize Clusters
```
#reduce data to 2 components
reduced_data=PCA(n_components=2).fit_transform(user_scaled)
clusters=KMeans(n_clusters=best_k, random_state=42).fit(reduced_data)
#set up meshgrid space
x_min = reduced_data[:, 0].min() -1
x_max = reduced_data[:, 0].max() +1
y_min = reduced_data[:, 1].min() - 1
y_max = reduced_data[:, 1].max() + 1
hx = (x_max-x_min)/1000.
hy = (y_max-y_min)/1000.
xx, yy = np.meshgrid(np.arange(x_min, x_max, hx), np.arange(y_min, y_max, hy))
#set up predictions and centroids
Z = clusters.predict(np.c_[xx.ravel(), yy.ravel()])
centroids = clusters.cluster_centers_
def PCA_plot(Z, centroids, saveloc=None):
Z = Z.reshape(xx.shape)
plt.figure(1, figsize=(20, 10))
plt.clf()
plt.imshow(Z, interpolation='nearest',
extent=(xx.min(), xx.max(), yy.min(), yy.max()),
cmap=plt.cm.Paired,
aspect='auto', origin='lower')
plt.plot(reduced_data[:, 0], reduced_data[:, 1], 'k.', markersize=2, alpha=0.1)
plt.scatter(centroids[:, 0], centroids[:, 1],
marker='x', s=169, linewidths=3,
color='w', zorder=10)
plt.title('Clustering on the dataset (PCA-reduced data)\n'
'Centroids are marked with white cross')
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
plt.xticks(())
plt.yticks(())
if saveloc:
plt.savefig(saveloc)
plt.show()
PCA_plot(Z, centroids, 'plots/pca_space.png')
# Compare the proportion of data in each cluster
user_df_kpredict =kmeans.predict(user_pca)
#get frequency of cluster counts for general pop
user_df_freq=np.bincount(user_df_kpredict)/len(user_df_kpredict)
sns.barplot(x=np.arange(0,len(user_df_freq)), y=user_df_freq)
plt.title("Clusters Frequency: Customers")
plt.savefig('plots/FrequencyClusters.png')
plt.show()
```
| github_jupyter |
# Rayleigh Scattering
**Scott Prahl**
**April 2021**
*If miepython is not installed, uncomment the following cell (i.e., delete the #) and run (shift-enter)*
```
#!pip install --user miepython
import numpy as np
import matplotlib.pyplot as plt
try:
import miepython
except ModuleNotFoundError:
print('miepython not installed. To install, uncomment and run the cell above.')
print('Once installation is successful, rerun this cell again.')
```
## Goals for this notebook:
* Plot Rayleigh scattering
* Compare total scattering between Rayleigh and Mie
* Compare scattering functions for unpolarized light
* Compare polarized results.
## Rayleigh Scattering Functions
```
def rayleigh(m,x):
"""
Calculate the efficiencies for a small sphere.
Based on equations 5.7 - 5.9 in Bohren and Huffman
Args:
m: the complex index of refraction of the sphere
x: the size parameter of the sphere
Returns:
qext: the total extinction efficiency
qsca: the scattering efficiency
qback: the backscatter efficiency
g: the average cosine of the scattering phase function
"""
ratio = (m**2-1)/(m**2+2)
qsca = 8/3*x**4*abs(ratio)**2
qext = 4*x*ratio*(1+x**2/15*ratio*(m**4+27*m**2+38)/(2*m**2+3))
qext = abs(qext.imag + qsca)
qback = 4*x**4*abs(ratio)**2
g = 0
return qext, qsca, qback, g
def rayleigh_S1_S2(m,x,mu):
"""
Calculate the scattering amplitude functions for small spheres.
Based on equation 5.4 in Bohren and Huffman
The amplitude functions are normalized so that when integrated
over all 4*pi solid angles, the integral will be qext*pi*x**2.
The units are weird, sr**(-0.5)
Args:
m: the complex index of refraction of the sphere
x: the size parameter of the sphere
mu: the angles, cos(theta), to calculate scattering amplitudes
Returns:
S1, S2: the scattering amplitudes at each angle mu [sr**(-0.5)]
"""
a1 = (2*x**3)/3 * (m**2-1)/(m**2+2)*1j
a1 += (2*x**5)/5 * (m**2-2)*(m**2-1)/(m**2+2)**2 *1j
s1 = (3/2)*a1*np.ones_like(mu)
s2 = (3/2)*a1*mu
## scale so integral over all angles is single scattering albedo
qext, qsca, qback, g = rayleigh(m,x)
factor = np.sqrt(np.pi*qext)*x
return s1/factor, s2/factor
def rayleigh_unpolarized(m,x,mu):
"""
Return the unpolarized scattered intensity for small spheres.
This is the average value for randomly polarized incident light.
The intensity is normalized so the integral of the unpolarized
intensity over 4pi steradians is equal to the single scattering albedo.
Args:
m: the complex index of refraction of the sphere
x: the size parameter
mu: the cos(theta) of each direction desired
Returns
The intensity at each angle in the array mu. Units [1/sr]
"""
s1, s2 = rayleigh_S1_S2(m,x,mu)
return (abs(s1)**2+abs(s2)**2)/2
```
Mie scattering describes the special case of the interaction of light passing through a non-absorbing medium with a single embedded spherical object. The sphere itself can be non-absorbing, moderately absorbing, or perfectly absorbing.
Rayleigh scattering is a simple closed-form solution for the scattering from small spheres.
## The Rayleigh scattering phase function
Rayleigh scattering describes the elastic scattering of light by spheres that are much smaller than the wavelength of light. The intensity $I$ of the scattered radiation is given by
$$
I=I_{0}\left(\frac {1+\cos ^{2}\theta }{2R^{2}}\right)
\left(\frac {2\pi }{\lambda }\right)^{4}
\left(\frac {n^{2}-1}{n^{2}+2}\right)^{2}
\left(\frac {d}{2}\right)^{6}
$$
where $I_0$ is the light intensity before the interaction with the particle, $R$ is the distance between the particle and the observer, $\theta$ is the scattering angle, $n$ is the refractive index of the particle, and $d$ is the diameter of the particle.
$$
x = \frac{\pi d}{\lambda} \qquad \rho=\frac{R}{\lambda}
$$
and thus
$$
I=\frac{I_0}{8\pi^2\rho^2}
\left(\frac{n^2-1}{n^2+2}\right)^{2}
x^{4}(1+\cos^2\theta)
$$
## Compare Efficiencies with Mie Code
```
for x in [0.1,0.2,0.3,0.4]:
m = 1.5-1j
theta = np.linspace(-180,180,180)
mu = np.cos(theta*np.pi/180)
rscat = rayleigh_unpolarized(m,x,mu)
mscat = miepython.i_unpolarized(m,x,mu)
plt.plot(theta,rscat,'--b')
plt.plot(theta,mscat,'r')
plt.annotate('x=%.1f '%x,(theta[-20],mscat[-20]),ha='right',va='bottom')
plt.xlim(-180,180)
plt.xlabel('Angle [degrees]')
plt.ylabel('Scattered Light [1/sr]')
plt.title('Solid Mie, Dashed Rayleigh')
plt.show()
```
## Polar plots for fun
```
m = 1.5
x = 0.1
theta = np.linspace(-180,180,180)
mu = np.cos(theta/180*np.pi)
unp = rayleigh_unpolarized(m,x,mu)
s1,s2 = rayleigh_S1_S2(m,x,mu)
par = abs(s1)**2
per = abs(s2)**2
fig,ax = plt.subplots(1,2,figsize=(12,5))
ax=plt.subplot(121, projection='polar')
ax.plot(theta/180*np.pi,unp)
ax.plot(theta/180*np.pi,par)
ax.plot(theta/180*np.pi,per)
ax.set_rticks([0.05, 0.1,0.15])
plt.subplot(122)
#plt.plot(theta,scat)
plt.plot(theta,unp)
plt.plot(theta,par)
plt.plot(theta,per)
plt.xlabel('Exit Angle [degrees]')
plt.ylabel('Unpolarized Scattered light [1/sr]')
plt.title("m=1.5, x = %.2f"%x)
plt.ylim(0.00,0.2)
plt.xlim(0,180)
plt.show()
```
## Compare Rayleigh and Mie efficiencies
```
m = 1.5
x = 0.1
qext, qsca, qback, g = miepython.mie(m,x)
rext, rsca, rback, rg = rayleigh(m,x)
print('Qext Qsca Qback g')
print("%.5e %.5e %.5e %.5f Mie"%(qext, qsca, qback, g))
print("%.5e %.5e %.5e %.5f Rayleigh"%(rext, rsca, rback, rg))
```
## Compare scattering amplitudes S1 and S2
```
m = 1.5
x = 0.1
theta = np.linspace(-180,180,19)
mu = np.cos(np.deg2rad(theta))
s1,s2 = miepython.mie_S1_S2(m,x,mu)
rs1, rs2 = rayleigh_S1_S2(m,x,mu)
# the real part of the Rayleigh scattering is always zero
print(" Mie Rayleigh | Mie Rayleigh")
print(" angle | S1.imag S1.imag | S2.imag S2.imag")
print("------------------------------------------------")
for i,angle in enumerate(theta):
print("%7.2f | %8.5f %8.5f | %8.5f %8.5f " % (angle,s1[i].imag,rs1[i].imag, s2[i].imag ,rs2[i].imag))
```
| github_jupyter |
```
# setup
from mlwpy import *
%matplotlib inline
iris = datasets.load_iris()
tts = skms.train_test_split(iris.data, iris.target,
test_size=.33, random_state=21)
(iris_train_ftrs, iris_test_ftrs,
iris_train_tgt, iris_test_tgt) = tts
# normal usage: build-fit-predict-evaluate
baseline = dummy.DummyClassifier(strategy="most_frequent")
baseline.fit(iris_train_ftrs, iris_train_tgt)
base_preds = baseline.predict(iris_test_ftrs)
base_acc = metrics.accuracy_score(base_preds, iris_test_tgt)
print(base_acc)
strategies = ['constant', 'uniform', 'stratified',
'prior', 'most_frequent']
# setup args to create diff. DummyClassifier strategies
baseline_args = [{'strategy':s} for s in strategies]
baseline_args[0]['constant'] = 0 # class 0 is setosa
accuracies = []
for bla in baseline_args:
baseline = dummy.DummyClassifier(**bla)
baseline.fit(iris_train_ftrs, iris_train_tgt)
base_preds = baseline.predict(iris_test_ftrs)
accuracies.append(metrics.accuracy_score(base_preds, iris_test_tgt))
display(pd.DataFrame({'accuracy':accuracies}, index=strategies))
# helpful stdlib tool for cleaning up printouts
import textwrap
print(textwrap.fill(str(sorted(metrics.SCORERS.keys())),
width=70))
knn = neighbors.KNeighborsClassifier()
# help(knn.score) # verbose, but complete
print(knn.score.__doc__.splitlines()[0])
print('\n---and---\n')
print("\n".join(knn.score.__doc__.splitlines()[-6:]))
tgt_preds = (neighbors.KNeighborsClassifier()
.fit(iris_train_ftrs, iris_train_tgt)
.predict(iris_test_ftrs))
print("accuracy:", metrics.accuracy_score(iris_test_tgt,
tgt_preds))
cm = metrics.confusion_matrix(iris_test_tgt,
tgt_preds)
print("confusion matrix:", cm, sep="\n")
fig, ax = plt.subplots(1,1,figsize=(4,4))
cm = metrics.confusion_matrix(iris_test_tgt, tgt_preds)
ax = sns.heatmap(cm, annot=True, square=True,
xticklabels=iris.target_names,
yticklabels=iris.target_names)
ax.set_xlabel('Predicted')
ax.set_ylabel('Actual');
macro_prec = metrics.precision_score(iris_test_tgt,
tgt_preds,
average='macro')
print("macro:", macro_prec)
cm = metrics.confusion_matrix(iris_test_tgt, tgt_preds)
n_labels = len(iris.target_names)
print("should equal 'macro avg':",
# correct column # columns
(np.diag(cm) / cm.sum(axis=0)).sum() / n_labels)
print("micro:", metrics.precision_score(iris_test_tgt,
tgt_preds,
average='micro'))
cm = metrics.confusion_matrix(iris_test_tgt, tgt_preds)
print("should equal avg='micro':",
# TP.sum() / (TP&FP).sum() -->
# all correct / all preds
np.diag(cm).sum() / cm.sum())
print(metrics.classification_report(iris_test_tgt,
tgt_preds))
# average is a weighted macro average (see text)
# verify sums-across-rows
cm = metrics.confusion_matrix(iris_test_tgt, tgt_preds)
print("row counts equal support:", cm.sum(axis=1))
# warning: this is 1 "one" not l "ell"
is_versicolor = iris.target == 1
tts_1c = skms.train_test_split(iris.data, is_versicolor,
test_size=.33, random_state = 21)
(iris_1c_train_ftrs, iris_1c_test_ftrs,
iris_1c_train_tgt, iris_1c_test_tgt) = tts_1c
# build, fit, predict (probability scores) for NB model
gnb = naive_bayes.GaussianNB()
prob_true = (gnb.fit(iris_1c_train_ftrs, iris_1c_train_tgt)
.predict_proba(iris_1c_test_ftrs)[:,1]) # [:,1]=="True"
fpr, tpr, thresh = metrics.roc_curve(iris_1c_test_tgt,
prob_true)
auc = metrics.auc(fpr, tpr)
print("FPR : {}".format(fpr),
"TPR : {}".format(tpr), sep='\n')
# create the main graph
fig, ax = plt.subplots(figsize=(8,4))
ax.plot(fpr, tpr, 'o--')
ax.set_title("1-Class Iris ROC Curve\nAUC:{:.3f}".format(auc))
ax.set_xlabel("FPR")
ax.set_ylabel("TPR");
# do a bit of work to label some points with their
# respective thresholds
investigate = np.array([1,3,5])
for idx in investigate:
th, f, t = thresh[idx], fpr[idx], tpr[idx]
ax.annotate('thresh = {:.3f}'.format(th),
xy=(f+.01, t-.01), xytext=(f+.1, t),
arrowprops = {'arrowstyle':'->'})
title_fmt = "Threshold {}\n~{:5.3f}\nTPR : {:.3f}\nFPR : {:.3f}"
pn = ['Positive', 'Negative']
add_args = {'xticklabels': pn,
'yticklabels': pn,
'square':True}
fig, axes = plt.subplots(1,3, sharey = True, figsize=(12,4))
for ax, thresh_idx in zip(axes.flat, investigate):
preds_at_th = prob_true < thresh[thresh_idx]
cm = metrics.confusion_matrix(1-iris_1c_test_tgt, preds_at_th)
sns.heatmap(cm, annot=True, cbar=False, ax=ax,
**add_args)
ax.set_xlabel('Predicted')
ax.set_title(title_fmt.format(thresh_idx,
thresh[thresh_idx],
tpr[thresh_idx],
fpr[thresh_idx]))
axes[0].set_ylabel('Actual');
# note: e.g. for threshold 3
# FPR = 1-spec = 1 - 31/(31+2) = 1 - 31/33 = 0.0606...
fig,ax = plt.subplots(1,1,figsize=(3,3))
model = neighbors.KNeighborsClassifier(3)
cv_auc = skms.cross_val_score(model, iris.data, iris.target==1,
scoring='roc_auc', cv=10)
ax = sns.swarmplot(cv_auc, orient='v')
ax.set_title('10-Fold AUCs');
checkout = [0,50,100]
print("Original Encoding")
print(iris.target[checkout])
print("'Multi-label' Encoding")
print(skpre.label_binarize(iris.target, [0,1,2])[checkout])
iris_multi_tgt = skpre.label_binarize(iris.target, [0,1,2])
# im --> "iris multi"
(im_train_ftrs, im_test_ftrs,
im_train_tgt, im_test_tgt) = skms.train_test_split(iris.data,
iris_multi_tgt,
test_size=.33,
random_state=21)
# knn wrapped up in one-versus-rest (3 classifiers)
knn = neighbors.KNeighborsClassifier(n_neighbors=5)
ovr_knn = skmulti.OneVsRestClassifier(knn)
pred_probs = (ovr_knn.fit(im_train_ftrs, im_train_tgt)
.predict_proba(im_test_ftrs))
# make ROC plots
lbl_fmt = "Class {} vs Rest (AUC = {:.2f})"
fig,ax = plt.subplots(figsize=(8,4))
for cls in [0,1,2]:
fpr, tpr, _ = metrics.roc_curve(im_test_tgt[:,cls],
pred_probs[:,cls])
label = lbl_fmt.format(cls, metrics.auc(fpr,tpr))
ax.plot(fpr, tpr, 'o--', label=label)
ax.legend()
ax.set_xlabel("FPR")
ax.set_ylabel("TPR");
knn = neighbors.KNeighborsClassifier(n_neighbors=5)
ovo_knn = skmulti.OneVsOneClassifier(knn)
pred_scores = (ovo_knn.fit(iris_train_ftrs, iris_train_tgt)
.decision_function(iris_test_ftrs))
df = pd.DataFrame(pred_scores)
df['class'] = df.values.argmax(axis=1)
display(df.head())
# note: ugly to make column headers
mi = pd.MultiIndex([['Class Indicator', 'Vote'], [0, 1, 2]],
[[0]*3+[1]*3,list(range(3)) * 2])
df = pd.DataFrame(np.c_[im_test_tgt, pred_scores],
columns=mi)
display(df.head())
def hand_and_till_M_statistic(test_tgt, test_probs, weighted=False):
def auc_helper(truth, probs):
fpr, tpr, _ = metrics.roc_curve(truth, probs)
return metrics.auc(fpr, tpr)
classes = np.unique(test_tgt)
n_classes = len(classes)
indicator = skpre.label_binarize(test_tgt, classes)
avg_auc_sum = 0.0
# comparing class i and class j
for ij in it.combinations(classes, 2):
# use use sum to act like a logical or
ij_indicator = indicator[:,ij].sum(axis=1,
dtype=np.bool)
# slightly ugly, can't broadcast these as indexes
# use .ix_ to save the day
ij_probs = test_probs[np.ix_(ij_indicator, ij)]
ij_test_tgt = test_tgt[ij_indicator]
i,j = ij
auc_ij = auc_helper(ij_test_tgt==i, ij_probs[:,0])
auc_ji = auc_helper(ij_test_tgt==j, ij_probs[:,1])
# compared to Hand & Till reference
# no / 2 ... factor it out since it will cancel
avg_auc_ij = (auc_ij + auc_ji)
if weighted:
avg_auc_ij *= ij_indicator.sum() / len(test_tgt)
avg_auc_sum += avg_auc_ij
# compared to Hand & Till reference
# no * 2 ... factored out above and they cancel
M = avg_auc_sum / (n_classes * (n_classes-1))
return M
knn = neighbors.KNeighborsClassifier()
knn.fit(iris_train_ftrs, iris_train_tgt)
test_probs = knn.predict_proba(iris_test_ftrs)
hand_and_till_M_statistic(iris_test_tgt, test_probs)
fig,ax = plt.subplots(1,1,figsize=(3,3))
htm_scorer = metrics.make_scorer(hand_and_till_M_statistic,
needs_proba=True)
cv_auc = skms.cross_val_score(model,
iris.data, iris.target,
scoring=htm_scorer, cv=10)
sns.swarmplot(cv_auc, orient='v')
ax.set_title('10-Fold H&T Ms');
fig,ax = plt.subplots(figsize=(6,3))
for cls in [0,1,2]:
prc = metrics.precision_recall_curve
precision, recall, _ = prc(im_test_tgt[:,cls],
pred_probs[:,cls])
prc_auc = metrics.auc(recall, precision)
label = "Class {} vs Rest (AUC) = {:.2f})".format(cls, prc_auc)
ax.plot(recall, precision, 'o--', label=label)
ax.legend()
ax.set_xlabel('Recall')
ax.set_ylabel('Precision');
# negate b/c we want big values first
myorder = np.argsort(-prob_true)
# cumulative sum then to percent (last value is total)
realpct_myorder = iris_1c_test_tgt[myorder].cumsum()
realpct_myorder = realpct_myorder / realpct_myorder[-1]
# convert counts of data into percents
N = iris_1c_test_tgt.size
xs = np.linspace(1/N,1,N)
print(myorder[:3])
fig, (ax1, ax2) = plt.subplots(1,2, figsize=(8,4))
fig.tight_layout()
# cumulative response
ax1.plot(xs, realpct_myorder, 'r.')
ax1.plot(xs, xs, 'b-')
ax1.axes.set_aspect('equal')
ax1.set_title("Cumulative Response")
ax1.set_ylabel("Percent of Actual Hits")
ax1.set_xlabel("Percent Of Population\n" +
"Starting with Highest Predicted Hits")
# lift
# replace divide by zero with 1.0
ax2.plot(xs, realpct_myorder / np.where(xs > 0, xs, 1))
ax2.set_title("Lift Versus Random")
ax2.set_ylabel("X-Fold Improvement") # not cross-fold!
ax2.set_xlabel("Percent Of Population\n" +
"Starting with Highest Predicted Hits")
ax2.yaxis.tick_right()
ax2.yaxis.set_label_position('right');
classifiers = {'base' : baseline,
'gnb' : naive_bayes.GaussianNB(),
'3-NN' : neighbors.KNeighborsClassifier(n_neighbors=10),
'10-NN' : neighbors.KNeighborsClassifier(n_neighbors=3)}
# define the one_class iris problem so we don't have random ==1 around
iris_onec_ftrs = iris.data
iris_onec_tgt = iris.target==1
msrs = ['accuracy', 'average_precision', 'roc_auc']
fig, axes = plt.subplots(len(msrs), 1, figsize=(6, 2*len(msrs)))
fig.tight_layout()
for mod_name, model in classifiers.items():
# abbreviate
cvs = skms.cross_val_score
cv_results = {msr:cvs(model, iris_onec_ftrs, iris_onec_tgt,
scoring=msr, cv=10) for msr in msrs}
for ax, msr in zip(axes, msrs):
msr_results = cv_results[msr]
my_lbl = "{:12s} {:.3f} {:.2f}".format(mod_name,
msr_results.mean(),
msr_results.std())
ax.plot(msr_results, 'o--', label=my_lbl)
ax.set_title(msr)
ax.legend(loc='lower center', ncol=2)
fig, axes = plt.subplots(2,2, figsize=(4,4), sharex=True, sharey=True)
fig.tight_layout()
for ax, (mod_name, model) in zip(axes.flat, classifiers.items()):
preds = skms.cross_val_predict(model,
iris_onec_ftrs, iris_onec_tgt,
cv=10)
cm = metrics.confusion_matrix(iris.target==1, preds)
sns.heatmap(cm, annot=True, ax=ax,
cbar=False, square=True, fmt="d")
ax.set_title(mod_name)
axes[1,0].set_xlabel('Predicted')
axes[1,1].set_xlabel('Predicted')
axes[0,0].set_ylabel('Actual')
axes[1,0].set_ylabel('Actual');
fig, ax = plt.subplots(1, 1, figsize=(6,4))
cv_prob_true = {}
for mod_name, model in classifiers.items():
cv_probs = skms.cross_val_predict(model,
iris_onec_ftrs, iris_onec_tgt,
cv=10, method='predict_proba')
cv_prob_true[mod_name] = cv_probs[:,1]
fpr, tpr, thresh = metrics.roc_curve(iris_onec_tgt,
cv_prob_true[mod_name])
auc = metrics.auc(fpr, tpr)
ax.plot(fpr, tpr, 'o--', label="{}:{}".format(mod_name, auc))
ax.set_title('ROC Curves')
ax.legend();
fig, (ax1,ax2) = plt.subplots(1, 2, figsize=(10,5))
N = len(iris_onec_tgt)
xs = np.linspace(1/N,1,N)
ax1.plot(xs, xs, 'b-')
for mod_name in classifiers:
# negate b/c we want big values first
myorder = np.argsort(-cv_prob_true[mod_name])
# cumulative sum then to percent (last value is total)
realpct_myorder = iris_onec_tgt[myorder].cumsum()
realpct_myorder = realpct_myorder / realpct_myorder[-1]
ax1.plot(xs, realpct_myorder, '.', label=mod_name)
ax2.plot(xs,
realpct_myorder / np.where(xs > 0, xs, 1),
label=mod_name)
ax1.legend()
ax2.legend()
ax1.set_title("Cumulative Response")
ax2.set_title("Lift versus Random");
student_df = pd.read_csv('data/portugese_student_numeric_discrete.csv')
student_df['grade'] = pd.Categorical(student_df['grade'],
categories=['low', 'mid', 'high'],
ordered=True)
student_ftrs = student_df[student_df.columns[:-1]]
student_tgt = student_df['grade'].cat.codes
fig,ax = plt.subplots(1,1,figsize=(3,3))
model = neighbors.KNeighborsClassifier(3)
cv_auc = skms.cross_val_score(model,
student_ftrs, student_tgt,
scoring='accuracy', cv=10)
ax = sns.swarmplot(cv_auc, orient='v')
ax.set_title('10-Fold Accuracy');
model = neighbors.KNeighborsClassifier(3)
my_scorer = metrics.make_scorer(metrics.precision_score,
average='macro')
cv_auc = skms.cross_val_score(model,
student_ftrs, student_tgt,
scoring=my_scorer, cv=10)
fig,ax = plt.subplots(1,1,figsize=(3,3))
sns.swarmplot(cv_auc, orient='v')
ax.set_title('10-Fold Macro Precision');
htm_scorer = metrics.make_scorer(hand_and_till_M_statistic,
needs_proba=True)
cv_auc = skms.cross_val_score(model,
student_ftrs, student_tgt,
scoring=htm_scorer, cv=10)
fig,ax = plt.subplots(1,1,figsize=(3,3))
sns.swarmplot(cv_auc, orient='v')
ax.set_title('10-Fold H&T Ms');
classifiers = {'base' : dummy.DummyClassifier(strategy="most_frequent"),
'gnb' : naive_bayes.GaussianNB(),
'3-NN' : neighbors.KNeighborsClassifier(n_neighbors=10),
'10-NN' : neighbors.KNeighborsClassifier(n_neighbors=3)}
macro_precision = metrics.make_scorer(metrics.precision_score,
average='macro')
macro_recall = metrics.make_scorer(metrics.recall_score,
average='macro')
htm_scorer = metrics.make_scorer(hand_and_till_M_statistic,
needs_proba=True)
msrs = ['accuracy', macro_precision,
macro_recall, htm_scorer]
fig, axes = plt.subplots(len(msrs), 1, figsize=(6, 2*len(msrs)))
fig.tight_layout()
for mod_name, model in classifiers.items():
# abbreviate
cvs = skms.cross_val_score
cv_results = {msr:cvs(model, student_ftrs, student_tgt,
scoring=msr, cv=10) for msr in msrs}
for ax, msr in zip(axes, msrs):
msr_results = cv_results[msr]
my_lbl = "{:12s} {:.3f} {:.2f}".format(mod_name,
msr_results.mean(),
msr_results.std())
ax.plot(msr_results, 'o--')
ax.set_title(msr)
# uncomment to see summary stats (clutters plots)
#ax.legend(loc='lower center')
fig, axes = plt.subplots(2,2, figsize=(5,5), sharex=True, sharey=True)
fig.tight_layout()
for ax, (mod_name, model) in zip(axes.flat,
classifiers.items()):
preds = skms.cross_val_predict(model,
student_ftrs, student_tgt,
cv=10)
cm = metrics.confusion_matrix(student_tgt, preds)
sns.heatmap(cm, annot=True, ax=ax,
cbar=False, square=True, fmt="d",
xticklabels=['low', 'med', 'high'],
yticklabels=['low', 'med', 'high'])
ax.set_title(mod_name)
axes[1,0].set_xlabel('Predicted')
axes[1,1].set_xlabel('Predicted')
axes[0,0].set_ylabel('Actual')
axes[1,0].set_ylabel('Actual');
student_url = ('https://archive.ics.uci.edu/' +
'ml/machine-learning-databases/00320/student.zip')
def grab_student_numeric_discrete():
# download zip file and unzip
# unzipping unknown files can be a security hazard
import urllib.request, zipfile
urllib.request.urlretrieve(student_url,
'port_student.zip')
zipfile.ZipFile('port_student.zip').extract('student-mat.csv')
# preprocessing
df = pd.read_csv('student-mat.csv', sep=';')
# g1 & g2 are highly correlated with g3;
# dropping them makes the problem sig. harder
# we also remove all non-numeric columns
# and discretize the final grade by 0-50-75-100 percentile
# which were determined by hand
df = df.drop(columns=['G1', 'G2']).select_dtypes(include=['number'])
df['grade'] = pd.cut(df['G3'], [0, 11, 14, 20],
labels=['low', 'mid', 'high'],
include_lowest=True)
df.drop(columns=['G3'], inplace=True)
# save as
df.to_csv('portugese_student_numeric_discrete.csv', index=False)
```
| github_jupyter |
```
from azureml.core import Workspace, Dataset, Datastore
from azureml.core import Environment, Model
from azureml.core.compute import ComputeTarget
from azureml.core.runconfig import RunConfiguration, CondaDependencies, DEFAULT_CPU_IMAGE
from azureml.pipeline.steps import PythonScriptStep
from azureml.pipeline.core import Pipeline, PublishedPipeline
from azureml.pipeline.core import StepSequence
from azureml.widgets import RunDetails
import pandas as pd
import numpy as np
import os
import random as r
ws = Workspace.from_config()
compute_name = 'compute-cluster'
compute_target = ComputeTarget(ws, compute_name)
datastore = Datastore.get_default(ws)
my_datastore_name = 'workspaceblobstore'
my_datastore = Datastore.get(ws, my_datastore_name)
os.makedirs('Scoring_Scripts', exist_ok=True)
%%writefile Scoring_Scripts/Iris_Scoring_Local.py
from azureml.core import Run, Workspace
from azureml.core import Dataset, Datastore, Model
import os
import joblib
import numpy as np
import pandas as pd
run = Run.get_context()
def main():
ws = run.experiment.workspace
datastore = Datastore.get_default(ws)
model_path = Model.get_model_path('Iris-Multi-Classification-AutoML')
model = joblib.load(model_path)
dataset = Dataset.get_by_name(ws,'Iris Scoring')
scoringDF = dataset.to_pandas_dataframe()
predictions = model.predict(scoringDF)
predSeries = pd.Series(predictions)
scoringDF['Prediction'] = predSeries
output_datastore_path = 'Output_Folder'
os.makedirs(output_datastore_path, exist_ok=True)
FileName = "Iris_Predictions.csv"
OutputPath = os.path.join(output_datastore_path, FileName)
scoringDF.to_csv(OutputPath, index = False, sep=',')
datastore.upload_files(files=[OutputPath], target_path=output_datastore_path, overwrite=True)
os.remove(OutputPath)
os.rmdir(output_datastore_path)
if __name__ == '__main__':
main()
# Create an Environment (collection of Python packages) you will use to run the script
Env = Environment(name='AutoML Environment')
# Create a CondaDepencies variable
conda_dep = CondaDependencies()
# Add conda packages
conda_dep.add_conda_package("numpy==1.18.5")
conda_dep.add_conda_package("joblib==0.14.1")
conda_dep.add_conda_package("pandas==0.25.3")
conda_dep.add_conda_package("packaging==20.7")
conda_dep.add_conda_package("xgboost==0.90")
# Add pip packages
conda_dep.add_pip_package("azureml-defaults==1.19.0")
conda_dep.add_pip_package("azureml-automl-core==1.19.0")
conda_dep.add_pip_package("azureml-automl-runtime==1.19.0")
# Adds dependencies to PythonSection of myenv
Env.python.conda_dependencies=conda_dep
# Register the environment to your workspace
RegisteredEnvironment = Env.register(workspace=ws)
run_config = RunConfiguration()
run_config.environment = Env
run_config.environment.docker.enabled = True
run_config.environment.docker.base_image = DEFAULT_CPU_IMAGE
# Create the training (Estimator) step by filling in all of the parameters
scoring_step = PythonScriptStep(name='iris-scoring-step',
script_name='Iris_Scoring_Local.py',
source_directory='Scoring_Scripts',
arguments=[],
inputs=[],
compute_target=compute_target,
runconfig=run_config,
allow_reuse=False)
step_sequence = StepSequence(steps=[scoring_step])
pipeline = Pipeline(workspace=ws, steps=step_sequence)
# Run your pipeline
pipeline_experiment = Experiment(ws, 'Iris-Scoring-Pipeline-Run')
pipeline_run = pipeline_experiment.submit(pipeline, show_output=True)
RunDetails(pipeline_run).show()
pipeline_run.wait_for_completion(show_output=True)
published_pipeline = pipeline_run.publish_pipeline(
name='Iris-Local-Scoring-Pipeline',\
description='Pipeline that Scores Iris Data', version= '1.0')
published_pipeline
```
| github_jupyter |
### Plotting Sine and Cosine Wave in Python
```
import numpy as np
import matplotlib.pyplot as plt
plt.plot()
%matplotlib inline
```
### Sine Wave
```
Time = np.arange(0,200,0.1)
Amplitude = np.sin(Time)
plt.plot(Time, Amplitude)
plt.title('Sine Wave')
plt.xlabel('Time')
plt.ylabel('Amplitude=sin(Time)')
plt.grid(True, which="both")
plt.axhline(y=0, color='b')
plt.figure(figsize=[206,106])
plt.show()
```
### Cosine Wave
```
time = np.arange(0,10,0.1)
amplitude = np.cos(time)
plt.plot(time, amplitude)
plt.title('Cosine Wave')
plt.xlabel('Time')
plt.ylabel('Amplitude=cos(time)')
plt.grid(True, which="both")
plt.axhline(y=0, color='b')
plt.show()
```
### Plotting Sine*Cosine wave
```
time = np.arange(0,10,0.1)
amplitude = np.cos(time)*np.sin(time)
plt.plot(time, amplitude)
plt.title('Cosine-Sine Wave')
plt.xlabel('Time')
plt.ylabel('Amplitude=cos(time)*sin(time)')
plt.grid(True, which="both")
plt.axhline(y=0, color='b')
plt.show()
x = np.arange(0,8*np.pi,0.1)
y = np.cos(x)*np.sin(x)
plt.plot(x,y)
plt.title('Cosine_Sine Wave')
plt.xlabel('Time')
plt.ylabel('Amplitude=cos(x)*sin(x)')
plt.grid(True, which="both")
plt.axhline(y=0, color='r')
plt.show()
```
### Plotting Damped Oscillation
```
time = np.arange(0,8000,4)
amplitude = 1008*np.sin(time/108)*np.exp(-time/1008)
exponential_decay = 1008*np.exp(-time/1008)
reciprocal_exponential =1/1008*np.exp(time/1008)
sinusoidal_wave=1008*np.sin(time/108)
plt.plot(time, sinusoidal_wave,color='cyan')
plt.plot(time, exponential_decay, color='red')
plt.plot(time, amplitude, color='indigo' )
plt.title('Damped Oscillation')
plt.xlabel('Time')
plt.ylabel('Amplitude')
plt.grid(True, which="both")
plt.axhline(y=0, color='yellow')
plt.show()
```
### Sine-Cosine wave together but seperate
```
x=np.arange(0,8*np.pi,0.1)
y=np.sin(x)
z=np.cos(x)
plt.plot(x,y,x,z)
plt.show()
```
### Soundwave graph: Wave Packet
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
x = np.linspace(0, 10.0, num=50000)
plt.plot(x, np.sin(x*4*np.pi), 'r')
plt.plot(x, -np.sin(x*1.1*2*np.pi), 'b')
plt.plot(x, np.sin(x*2*np.pi) - np.sin(x*1.1*2*np.pi), 'g')
plt.show()
x = np.linspace(0, 100.0, num=5000)
plt.plot(x, np.sin(x*4*np.pi), 'b')
plt.plot(x, -np.sin(x*1.1*2*np.pi), 'r')
plt.plot(x, np.sin(x*2*np.pi) - np.sin(x*1.1*2*np.pi), 'g')
plt.show()
x = np.linspace(0, 1000.0, num=500)
plt.plot(x, np.sin(x*2*np.pi), 'b')
plt.plot(x, -np.sin(x*1.1*2*np.pi), 'r')
plt.plot(x, np.sin(x*2*np.pi) - np.sin(x*1.1*2*np.pi), 'g')
plt.show()
```
### Plotting the result of a Fourier transform
### Let's try a Fourier transform of a sine wave signal with a pure frequency, namely,
$$f(t) = \sin(2 \pi \nu t)$$,
sampled for $t = j \Delta t$ with $j = 0, 1, \ldots{}, N-1$. To get a pure frequency, we set,
$$\nu = \nu_m = m/(N \Delta t)$$
so that, for frequency $\nu_m$, our discretized signal becomes
$$f_j = \sin(2 \pi m j/N)$$.
```
import numpy as np
import matplotlib.pyplot as plt
N=15
t=np.arange(N)
m=9
nu=float(m)/N
f=np.sin(8*np.pi*nu*t)
ft=np.fft.fft(f)
freq=np.fft.fftfreq(N)
plt.plot(freq, ft.real**2, ft.imag**2)
plt.show()
```
| github_jupyter |
SOP006 - az logout
==================
Use the az command line interface to logout of Azure.
Steps
-----
### Common functions
Define helper functions used in this notebook.
```
# Define `run` function for transient fault handling, suggestions on error, and scrolling updates on Windows
import sys
import os
import re
import platform
import shlex
import shutil
import datetime
from subprocess import Popen, PIPE
from IPython.display import Markdown
retry_hints = {} # Output in stderr known to be transient, therefore automatically retry
error_hints = {} # Output in stderr where a known SOP/TSG exists which will be HINTed for further help
install_hint = {} # The SOP to help install the executable if it cannot be found
def run(cmd, return_output=False, no_output=False, retry_count=0, base64_decode=False, return_as_json=False, regex_mask=None):
"""Run shell command, stream stdout, print stderr and optionally return output
NOTES:
1. Commands that need this kind of ' quoting on Windows e.g.:
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='data-pool')].metadata.name}
Need to actually pass in as '"':
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='"'data-pool'"')].metadata.name}
The ' quote approach, although correct when pasting into Windows cmd, will hang at the line:
`iter(p.stdout.readline, b'')`
The shlex.split call does the right thing for each platform, just use the '"' pattern for a '
"""
MAX_RETRIES = 5
output = ""
retry = False
# When running `azdata sql query` on Windows, replace any \n in """ strings, with " ", otherwise we see:
#
# ('HY090', '[HY090] [Microsoft][ODBC Driver Manager] Invalid string or buffer length (0) (SQLExecDirectW)')
#
if platform.system() == "Windows" and cmd.startswith("azdata sql query"):
cmd = cmd.replace("\n", " ")
# shlex.split is required on bash and for Windows paths with spaces
#
cmd_actual = shlex.split(cmd)
# Store this (i.e. kubectl, python etc.) to support binary context aware error_hints and retries
#
user_provided_exe_name = cmd_actual[0].lower()
# When running python, use the python in the ADS sandbox ({sys.executable})
#
if cmd.startswith("python "):
cmd_actual[0] = cmd_actual[0].replace("python", sys.executable)
# On Mac, when ADS is not launched from terminal, LC_ALL may not be set, which causes pip installs to fail
# with:
#
# UnicodeDecodeError: 'ascii' codec can't decode byte 0xc5 in position 4969: ordinal not in range(128)
#
# Setting it to a default value of "en_US.UTF-8" enables pip install to complete
#
if platform.system() == "Darwin" and "LC_ALL" not in os.environ:
os.environ["LC_ALL"] = "en_US.UTF-8"
# When running `kubectl`, if AZDATA_OPENSHIFT is set, use `oc`
#
if cmd.startswith("kubectl ") and "AZDATA_OPENSHIFT" in os.environ:
cmd_actual[0] = cmd_actual[0].replace("kubectl", "oc")
# To aid supportability, determine which binary file will actually be executed on the machine
#
which_binary = None
# Special case for CURL on Windows. The version of CURL in Windows System32 does not work to
# get JWT tokens, it returns "(56) Failure when receiving data from the peer". If another instance
# of CURL exists on the machine use that one. (Unfortunately the curl.exe in System32 is almost
# always the first curl.exe in the path, and it can't be uninstalled from System32, so here we
# look for the 2nd installation of CURL in the path)
if platform.system() == "Windows" and cmd.startswith("curl "):
path = os.getenv('PATH')
for p in path.split(os.path.pathsep):
p = os.path.join(p, "curl.exe")
if os.path.exists(p) and os.access(p, os.X_OK):
if p.lower().find("system32") == -1:
cmd_actual[0] = p
which_binary = p
break
# Find the path based location (shutil.which) of the executable that will be run (and display it to aid supportability), this
# seems to be required for .msi installs of azdata.cmd/az.cmd. (otherwise Popen returns FileNotFound)
#
# NOTE: Bash needs cmd to be the list of the space separated values hence shlex.split.
#
if which_binary == None:
which_binary = shutil.which(cmd_actual[0])
# Display an install HINT, so the user can click on a SOP to install the missing binary
#
if which_binary == None:
print(f"The path used to search for '{cmd_actual[0]}' was:")
print(sys.path)
if user_provided_exe_name in install_hint and install_hint[user_provided_exe_name] is not None:
display(Markdown(f'HINT: Use [{install_hint[user_provided_exe_name][0]}]({install_hint[user_provided_exe_name][1]}) to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)")
else:
cmd_actual[0] = which_binary
start_time = datetime.datetime.now().replace(microsecond=0)
cmd_display = cmd
if regex_mask is not None:
regex = re.compile(regex_mask)
cmd_display = re.sub(regex, '******', cmd)
print(f"START: {cmd_display} @ {start_time} ({datetime.datetime.utcnow().replace(microsecond=0)} UTC)")
print(f" using: {which_binary} ({platform.system()} {platform.release()} on {platform.machine()})")
print(f" cwd: {os.getcwd()}")
# Command-line tools such as CURL and AZDATA HDFS commands output
# scrolling progress bars, which causes Jupyter to hang forever, to
# workaround this, use no_output=True
#
# Work around a infinite hang when a notebook generates a non-zero return code, break out, and do not wait
#
wait = True
try:
if no_output:
p = Popen(cmd_actual)
else:
p = Popen(cmd_actual, stdout=PIPE, stderr=PIPE, bufsize=1)
with p.stdout:
for line in iter(p.stdout.readline, b''):
line = line.decode()
if return_output:
output = output + line
else:
if cmd.startswith("azdata notebook run"): # Hyperlink the .ipynb file
regex = re.compile(' "(.*)"\: "(.*)"')
match = regex.match(line)
if match:
if match.group(1).find("HTML") != -1:
display(Markdown(f' - "{match.group(1)}": "{match.group(2)}"'))
else:
display(Markdown(f' - "{match.group(1)}": "[{match.group(2)}]({match.group(2)})"'))
wait = False
break # otherwise infinite hang, have not worked out why yet.
else:
print(line, end='')
if wait:
p.wait()
except FileNotFoundError as e:
if install_hint is not None:
display(Markdown(f'HINT: Use {install_hint} to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)") from e
exit_code_workaround = 0 # WORKAROUND: azdata hangs on exception from notebook on p.wait()
if not no_output:
for line in iter(p.stderr.readline, b''):
try:
line_decoded = line.decode()
except UnicodeDecodeError:
# NOTE: Sometimes we get characters back that cannot be decoded(), e.g.
#
# \xa0
#
# For example see this in the response from `az group create`:
#
# ERROR: Get Token request returned http error: 400 and server
# response: {"error":"invalid_grant",# "error_description":"AADSTS700082:
# The refresh token has expired due to inactivity.\xa0The token was
# issued on 2018-10-25T23:35:11.9832872Z
#
# which generates the exception:
#
# UnicodeDecodeError: 'utf-8' codec can't decode byte 0xa0 in position 179: invalid start byte
#
print("WARNING: Unable to decode stderr line, printing raw bytes:")
print(line)
line_decoded = ""
pass
else:
# azdata emits a single empty line to stderr when doing an hdfs cp, don't
# print this empty "ERR:" as it confuses.
#
if line_decoded == "":
continue
print(f"STDERR: {line_decoded}", end='')
if line_decoded.startswith("An exception has occurred") or line_decoded.startswith("ERROR: An error occurred while executing the following cell"):
exit_code_workaround = 1
# inject HINTs to next TSG/SOP based on output in stderr
#
if user_provided_exe_name in error_hints:
for error_hint in error_hints[user_provided_exe_name]:
if line_decoded.find(error_hint[0]) != -1:
display(Markdown(f'HINT: Use [{error_hint[1]}]({error_hint[2]}) to resolve this issue.'))
# Verify if a transient error, if so automatically retry (recursive)
#
if user_provided_exe_name in retry_hints:
for retry_hint in retry_hints[user_provided_exe_name]:
if line_decoded.find(retry_hint) != -1:
if retry_count < MAX_RETRIES:
print(f"RETRY: {retry_count} (due to: {retry_hint})")
retry_count = retry_count + 1
output = run(cmd, return_output=return_output, retry_count=retry_count)
if return_output:
if base64_decode:
import base64
return base64.b64decode(output).decode('utf-8')
else:
return output
elapsed = datetime.datetime.now().replace(microsecond=0) - start_time
# WORKAROUND: We avoid infinite hang above in the `azdata notebook run` failure case, by inferring success (from stdout output), so
# don't wait here, if success known above
#
if wait:
if p.returncode != 0:
raise SystemExit(f'Shell command:\n\n\t{cmd_display} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(p.returncode)}.\n')
else:
if exit_code_workaround !=0 :
raise SystemExit(f'Shell command:\n\n\t{cmd_display} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(exit_code_workaround)}.\n')
print(f'\nSUCCESS: {elapsed}s elapsed.\n')
if return_output:
if base64_decode:
import base64
return base64.b64decode(output).decode('utf-8')
else:
return output
```
### Logout of azure
```
run("az logout")
print("Notebook execution is complete.")
```
Related
-------
- [SOP005 - az login](../common/sop005-az-login.ipynb)
| github_jupyter |
# Matplotlib
```
# Notebook Magic Line
%matplotlib inline # create visualizations in the notebook itself
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from google.colab import drive
drive.mount('/content/drive')
df = pd.read_csv("/content/drive/My Drive/Colab Notebooks/PA Projects/ML Class/Week 4/Top_AI_scientists_cleaned.csv")
df
```
## Basic Plotting
```
world_rank = np.asarray(df["World Rank"])
dblp = np.asarray(df["#DBLP"])
plt.plot(world_rank,dblp)
```
### Title, Labels, Legend, Ticks
```
plt.plot(world_rank,dblp)
# Title
plt.title("Relation between World Rank and Number of Publications")
# Axis Labels
plt.xlabel("World Rank")
plt.ylabel("Number of Publications")
# Adding more to a single plot
hidx = np.asarray(df["H-index"])
plt.plot(dblp)
plt.plot(hidx)
# Legends
plt.plot(dblp)
plt.plot(hidx)
plt.legend(labels=['#DBLP','H-Index'],loc='best')
```
**loc can be :**
best,
upper right,
upper left,
lower left,
lower right,
right,
center left,
center right,
lower center,
upper center,
center
```
world_rank = np.asarray(df["World Rank"]) - 1
names = np.asarray(df["Name"])
plt.plot(dblp[:100])
plt.plot(hidx[:100])
plt.legend(labels=['#DBLP','H-Index'],loc='best')
# Setting label for each person
plt.xticks(ticks=world_rank[:100],labels=names[:100])
plt.show()
# Vertical Rotation
world_rank = np.asarray(df["World Rank"]) - 1
names = np.asarray(df["Name"])
plt.plot(dblp[:100])
plt.plot(hidx[:100])
plt.legend(labels=['#DBLP','H-Index'],loc='best')
plt.xticks(ticks=world_rank[:100],labels=names[:100], rotation='vertical')
plt.show()
# Tuning the Size in inches
plt.rcParams["figure.figsize"] = (40,10)
world_rank = np.asarray(df["World Rank"]) - 1
names = np.asarray(df["Name"])
plt.plot(dblp[:100])
plt.plot(hidx[:100])
plt.legend(labels=['#DBLP','H-Index'],loc='best')
plt.xticks(ticks=world_rank[:100],labels=names[:100], rotation='vertical')
plt.show()
# Change back to default size
plt.rcParams["figure.figsize"] = plt.rcParamsDefault["figure.figsize"]
# Playing with colors
plt.rcParams["figure.figsize"] = (40,10)
world_rank = np.asarray(df["World Rank"]) - 1
names = np.asarray(df["Name"])
plt.plot(dblp[:100],'r-')
plt.plot(hidx[:100],'g--')
plt.legend(labels=['#DBLP','H-Index'],loc='best')
plt.xticks(ticks=world_rank[:100],labels=names[:100], rotation='vertical')
plt.show()
```
## Subplots
```
# National Rank, Citations, #DBLP, H-index
national_rank = np.asarray(df["National Rank"])[:50]
citations = np.asarray(df["Citations"])[:50]
dblp = np.asarray(df["#DBLP"])[:50]
hidx = np.asarray(df["H-index"])[:50]
world_rank = np.asarray(df["World Rank"])[:50] - 1
names = np.asarray(df["Name"])[:50]
plt.rcParams["figure.figsize"] = (50,30)
fig, ax = plt.subplots(nrows=2,ncols=2)
ax[0,0].plot(national_rank,'r*')
ax[0,1].plot(citations,'g^')
ax[1,0].plot(dblp,'yo')
ax[1,1].plot(hidx,'b--')
ax[0,0].set_title("National Rank")
ax[0,1].set_title("Citations")
ax[1,0].set_title("#DBLP")
ax[1,1].set_title("H-index")
for x in range(2):
for y in range(2):
ax[x,y].set_xticks(ticks=world_rank)
ax[x,y].set_xticklabels(labels=names, rotation='vertical')
plt.show()
```
| github_jupyter |
# Transfer Learning on TPUs
In the <a href="3_tf_hub_transfer_learning.ipynb">previous notebook</a>, we learned how to do transfer learning with [TensorFlow Hub](https://www.tensorflow.org/hub). In this notebook, we're going to kick up our training speed with [TPUs](https://www.tensorflow.org/guide/tpu).
## Learning Objectives
1. Know how to set up a [TPU strategy](https://www.tensorflow.org/api_docs/python/tf/distribute/experimental/TPUStrategy?version=nightly) for training
2. Know how to use a TensorFlow Hub Module when training on a TPU
3. Know how to create and specify a TPU for training
First things first. Configure the parameters below to match your own Google Cloud project details.
Each learning objective will correspond to a __#TODO__ in the notebook, where you will complete the notebook cell's code before running the cell. Refer to the [solution notebook](../solutions/4_tpu_training.ipynb))for reference.
```
import os
os.environ["BUCKET"] = "your-bucket-here"
```
## Packaging the Model
In order to train on a TPU, we'll need to set up a python module for training. The skeleton for this has already been built out in `tpu_models` with the data processing functions from the pevious lab copied into <a href="tpu_models/trainer/util.py">util.py</a>.
Similarly, the model building and training functions are pulled into <a href="tpu_models/trainer/model.py">model.py</a>. This is almost entirely the same as before, except the hub module path is now a variable to be provided by the user. We'll get into why in a bit, but first, let's take a look at the new `task.py` file.
We've added five command line arguments which are standard for cloud training of a TensorFlow model: `epochs`, `steps_per_epoch`, `train_path`, `eval_path`, and `job-dir`. There are two new arguments for TPU training: `tpu_address` and `hub_path`
`tpu_address` is going to be our TPU name as it appears in [Compute Engine Instances](https://console.cloud.google.com/compute/instances). We can specify this name with the [ctpu up](https://cloud.google.com/tpu/docs/ctpu-reference#up) command.
`hub_path` is going to be a Google Cloud Storage path to a downloaded TensorFlow Hub module.
The other big difference is some code to deploy our model on a TPU. To begin, we'll set up a [TPU Cluster Resolver](https://www.tensorflow.org/api_docs/python/tf/distribute/cluster_resolver/TPUClusterResolver), which will help tensorflow communicate with the hardware to set up workers for training ([more on TensorFlow Cluster Resolvers](https://www.tensorflow.org/api_docs/python/tf/distribute/cluster_resolver/ClusterResolver)). Once the resolver [connects to](https://www.tensorflow.org/api_docs/python/tf/config/experimental_connect_to_cluster) and [initializes](https://www.tensorflow.org/api_docs/python/tf/tpu/experimental/initialize_tpu_system) the TPU system, our Tensorflow Graphs can be initialized within a [TPU distribution strategy](https://www.tensorflow.org/api_docs/python/tf/distribute/experimental/TPUStrategy), allowing our TensorFlow code to take full advantage of the TPU hardware capabilities.
**TODO**: Complete the code below to setup the `resolver` and define the TPU training strategy.
```
%%writefile tpu_models/trainer/task.py
import argparse
import json
import os
import sys
import tensorflow as tf
from . import model
from . import util
def _parse_arguments(argv):
"""Parses command-line arguments."""
parser = argparse.ArgumentParser()
parser.add_argument(
'--epochs',
help='The number of epochs to train',
type=int, default=5)
parser.add_argument(
'--steps_per_epoch',
help='The number of steps per epoch to train',
type=int, default=500)
parser.add_argument(
'--train_path',
help='The path to the training data',
type=str, default="gs://cloud-ml-data/img/flower_photos/train_set.csv")
parser.add_argument(
'--eval_path',
help='The path to the evaluation data',
type=str, default="gs://cloud-ml-data/img/flower_photos/eval_set.csv")
parser.add_argument(
'--tpu_address',
help='The path to the evaluation data',
type=str, required=True)
parser.add_argument(
'--hub_path',
help='The path to TF Hub module to use in GCS',
type=str, required=True)
parser.add_argument(
'--job-dir',
help='Directory where to save the given model',
type=str, required=True)
return parser.parse_known_args(argv)
def main():
"""Parses command line arguments and kicks off model training."""
args = _parse_arguments(sys.argv[1:])[0]
# TODO: define a TPU strategy
resolver = # TODO: Your code goes here
tf.config.experimental_connect_to_cluster(resolver)
tf.tpu.experimental.initialize_tpu_system(resolver)
strategy = # TODO: Your code goes here
with strategy.scope():
train_data = util.load_dataset(args.train_path)
eval_data = util.load_dataset(args.eval_path, training=False)
image_model = model.build_model(args.job_dir, args.hub_path)
model_history = model.train_and_evaluate(
image_model, args.epochs, args.steps_per_epoch,
train_data, eval_data, args.job_dir)
if __name__ == '__main__':
main()
```
## The TPU server
Before we can start training with this code, we need a way to pull in [MobileNet](https://tfhub.dev/google/imagenet/mobilenet_v2_035_224/feature_vector/4). When working with TPUs in the cloud, the TPU will [not have access to the VM's local file directory](https://cloud.google.com/tpu/docs/troubleshooting#cannot_use_local_filesystem) since the TPU worker acts as a server. Because of this **all data used by our model must be hosted on an outside storage system** such as Google Cloud Storage. This makes [caching](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#cache) our dataset especially critical in order to speed up training time.
To access MobileNet with these restrictions, we can download a compressed [saved version](https://www.tensorflow.org/hub/tf2_saved_model) of the model by using the [wget](https://www.gnu.org/software/wget/manual/wget.html) command. Adding `?tf-hub-format=compressed` at the end of our module handle gives us a download URL.
```
!wget https://tfhub.dev/google/imagenet/mobilenet_v2_100_224/feature_vector/4?tf-hub-format=compressed
```
This model is still compressed, so lets uncompress it with the `tar` command below and place it in our `tpu_models` directory.
```
%%bash
rm -r tpu_models/hub
mkdir tpu_models/hub
tar xvzf 4?tf-hub-format=compressed -C tpu_models/hub/
```
Finally, we need to transfer our materials to the TPU. We'll use GCS as a go-between, using [gsutil cp](https://cloud.google.com/storage/docs/gsutil/commands/cp) to copy everything.
```
!gsutil rm -r gs://$BUCKET/tpu_models
!gsutil cp -r tpu_models gs://$BUCKET/tpu_models
```
## Spinning up a TPU
Time to wake up a TPU! Open the [Google Cloud Shell](https://console.cloud.google.com/home/dashboard?cloudshell=true) and copy the [gcloud compute](https://cloud.google.com/sdk/gcloud/reference/compute/tpus/execution-groups/create) command below. Say 'Yes' to the prompts to spin up the TPU.
`gcloud compute tpus execution-groups create \
--name=my-tpu \
--zone=us-central1-b \
--tf-version=2.3.2 \
--machine-type=n1-standard-1 \
--accelerator-type=v3-8`
It will take about five minutes to wake up. Then, it should automatically SSH into the TPU, but alternatively [Compute Engine Interface](https://console.cloud.google.com/compute/instances) can be used to SSH in. You'll know you're running on a TPU when the command line starts with `your-username@your-tpu-name`.
This is a fresh TPU and still needs our code. Run the below cell and copy the output into your TPU terminal to copy your model from your GCS bucket. Don't forget to include the `.` at the end as it tells gsutil to copy data into the currect directory.
```
!echo "gsutil cp -r gs://$BUCKET/tpu_models ."
```
Time to shine, TPU! Run the below cell and copy the output into your TPU terminal. Training will be slow at first, but it will pick up speed after a few minutes once the Tensorflow graph has been built out.
**TODO**: Complete the code below by adding flags for `tpu_address` and the `hub_path`. Have another look at `task.py` to see how these flags are used. The `tpu_address` denotes the TPU you created above and `hub_path` should denote the location of the TFHub module. (Note that the training code requires a TPU_NAME environment variable, set in the first two lines below -- you may reuse it in your code.)
```
%%bash
export TPU_NAME=my-tpu
echo "export TPU_NAME="$TPU_NAME
echo "python3 -m tpu_models.trainer.task \
# TODO: Your code goes here \
# TODO: Your code goes here \
--job-dir=gs://$BUCKET/flowers_tpu_$(date -u +%y%m%d_%H%M%S)"
```
How did it go? In the previous lab, it took about 2-3 minutes to get through 25 images. On the TPU, it took 5-6 minutes to get through 2500. That's more than 40x faster! And now our accuracy is over 90%! Congratulations!
Time to pack up shop. Run `exit` in the TPU terminal to close the SSH connection, and `gcloud compute tpus execution-groups delete my-tpu --zone=us-central1-b` in the [Cloud Shell](https://console.cloud.google.com/home/dashboard?cloudshell=true) to delete the Cloud TPU and Compute Engine instances. Alternatively, they can be deleted through the [Compute Engine Interface](https://console.cloud.google.com/compute/instances), but don't forget to separately delete the [TPU](https://console.cloud.google.com/compute/tpus) too!
Copyright 2022 Google Inc.
Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| github_jupyter |
```
import pandas as pd
import numpy as np
from tqdm import tqdm
from sklearn.neighbors import BallTree
import seaborn as sns
import geopandas as gpd
from shapely.geometry import Point, LineString
from pyproj import Proj, transform
from matplotlib import pyplot as plt
%matplotlib inline
from urbansim_templates import modelmanager as mm
from urbansim_templates.models import LargeMultinomialLogitStep
import orca
import os; os.chdir('../')
import warnings;warnings.simplefilter('ignore')
from scripts import datasources, models, variables
from choicemodels import MultinomialLogit
from choicemodels.tools import MergedChoiceTable
```
### Load data
```
orca.run(['initialize_network_small', 'initialize_network_walk'])
```
Load UrbanSim data
```
jobs = orca.get_table('jobs').to_frame()
buildings = orca.get_table('buildings').to_frame()
parcels = orca.get_table('parcels').to_frame()
parcels.head()
```
Load and format CHTS data for estimation
```
data_dir = '/home/data/fall_2018/'
chts_dir = 'CHTS_csv_format/'
chts_persons = pd.read_csv(data_dir + chts_dir + 'data/Deliv_PER.csv', low_memory=False)
chts_persons_lookup = pd.read_csv(data_dir + chts_dir + 'data/LookUp_PER.csv')
chts_households_lookup = pd.read_csv(data_dir + chts_dir + 'data/LookUp_Home.csv')
chts_persons = pd.merge(
chts_persons.set_index(['SAMPN','PERNO']),
chts_persons_lookup.set_index(['SAMPN','PERNO']),
left_index=True, right_index=True,
suffixes=('_persons', '_lookup')).reset_index()
chts_persons = pd.merge(
chts_persons.set_index(['SAMPN']),
chts_households_lookup.set_index(['SAMPN']),
left_index=True, right_index=True).reset_index()
chts_persons = chts_persons[chts_persons['HCTFIP'].isin([1, 13, 41, 55, 75, 81, 85, 95, 97])].reset_index()
```
### Identify in-region workers
```
chts_persons['work_in_region'] = chts_persons['WCTFIP_lookup'].isin([1, 13, 41, 55, 75, 81, 85, 95, 97])
chts_persons['work_in_region'].value_counts()
```
### Get job coords
```
buildings = pd.merge(buildings, parcels[['x', 'y']], left_on='parcel_id', right_index=True)
jobs = pd.merge(jobs, buildings[['x', 'y']], left_on='building_id', right_index=True)
jobs.rename(columns={'x': 'lng', 'y': 'lat'}, inplace=True)
```
### Prepare jobs table and CHTS persons table for job assignment
```
jobs.loc[:,'taken'] = False
chts_persons.loc[:, 'job_id'] = None
# haversine requires data in form of [lat, lng] and inputs/outputs in units of radians
persons_work_rad = np.deg2rad(chts_persons[['WYCORD_lookup', 'WXCORD_lookup']])
jobs_rad = np.deg2rad(jobs[['lat', 'lng']])
jobs.loc[:, 'x'] = jobs_rad['lng']
jobs.loc[:, 'y'] = jobs_rad['lat']
```
### Assign CHTS persons a job ID
```
%%time
dists = []
no_job_info = []
no_work_coords = []
matched_on_industry = 0
matched_on_occupation = 0
matched_on_both = 0
matched_on_none = 0
for i, person in tqdm(chts_persons.iterrows(), total=len(chts_persons)):
# only assign a job ID for employed persons with a fixed
# work location other than their home
if (person['EMPLY'] == 1) & (person['WLOC'] == 1) & (person['work_in_region'] == True):
# skip person if no CHTS industry or occupation
if (person['INDUS'] > 96) & (person['OCCUP'] > 96):
no_job_info.append(i)
continue
# skip person if no work location
elif pd.isnull(person[['WYCORD_lookup', 'WXCORD_lookup']]).any():
no_work_coords.append(i)
continue
# if CHTS industry is unknown, match jobs based on occupation only
elif person['INDUS'] > 96:
potential_jobs = jobs[
(jobs['occupation_id'] == person['OCCUP']) &
(jobs['taken'] == False)]
matched_on_occupation += 1
# if occupation is unknown, match jobs based on industry only
elif person['OCCUP'] > 96:
potential_jobs = jobs[
(jobs['sector_id'] == person['INDUS']) &
(jobs['taken'] == False)]
matched_on_industry += 1
elif (person['INDUS'] < 97) & (person['OCCUP'] < 97):
# define potential jobs based on industry and occupation
potential_jobs = jobs[
(jobs['sector_id'] == person['INDUS']) &
(jobs['occupation_id'] == person['OCCUP']) &
(jobs['taken'] == False)]
# if no such jobs exist, define jobs by industry
if len(potential_jobs) == 0:
potential_jobs = jobs[
(jobs['sector_id'] == person['INDUS']) &
(jobs['taken'] == False)]
matched_on_industry += 1
# if no such jobs exist, define jobs by occupation
if len(potential_jobs) == 0:
potential_jobs = jobs[
(jobs['occupation_id'] == person['OCCUP']) &
(jobs['taken'] == False)]
matched_on_occupation += 1
# otherwise, continue
if len(potential_jobs) == 0:
matched_on_none += 1
continue
else:
matched_on_both += 1
# build the tree of potential jobs for fast nearest-neighbor search
tree = BallTree(potential_jobs[['y','x']], metric='haversine')
# query the tree for nearest job to each workplace
dist, idx = tree.query(persons_work_rad.iloc[i].values.reshape(1,-1), return_distance=True)
# save results
job = potential_jobs.iloc[idx[0]]
dists.append(dist[0][0])
chts_persons.loc[i, 'job_id'] = job.index.values[0]
jobs.loc[jobs.index.values == job.index.values[0], 'taken'] = True
print('Assigned job IDs to {0}% of workers with a fixed work location.'.format(
np.round(chts_persons.job_id.count() / len(
chts_persons[(chts_persons['EMPLY'] == 1) & (chts_persons['WLOC'] == 1)]) * 100, 1)))
print('{0}% had no industry/occupation info.'.format(
np.round(len(no_job_info) / len(
chts_persons[(chts_persons['EMPLY'] == 1) & (chts_persons['WLOC'] == 1)]) * 100, 1)))
print('{0}% had no work coordinates.'.format(
np.round(len(no_work_coords) / len(
chts_persons[(chts_persons['EMPLY'] == 1) & (chts_persons['WLOC'] == 1)]) * 100, 1)))
print('{0} workers were not assigned to jobs based on industry OR occupation.'.format(
str(matched_on_none)))
print('{0} workers were assigned to jobs based on industry AND occupation.'.format(
str(matched_on_both)))
print('{0} workers were assigned to jobs based on industry only.'.format(
str(matched_on_industry)))
print('{0} workers were assigned to jobs based on occupation only.'.format(
str(matched_on_occupation)))
chts_persons.to_csv('data/chts_persons_w_jobs.csv', index=False)
```
Save the table so we don't have to re-run every time:
```
chts_persons = pd.read_csv('data/chts_persons_w_jobs.csv')
```
### Validate the work assignements
Compare the distance between the observed work location and the location of the job ID assigned in UrbanSim:
```
km_dists = [dist * 6371 for dist in dists]
pd.Series(km_dists).plot(kind='hist', bins=1000, xlim=(0, 2.5))
```
Compare the distribution of jobs per census tract for the observed work locations versus those we assigned:
```
chts_workers = chts_persons[~pd.isnull(chts_persons['job_id'])]
# chts data
perwgt_tract_count = chts_workers.groupby('WTRACT_lookup').PERWGT.sum()
perwgt_cumsum = chts_workers.groupby('WTRACT_lookup').PERWGT.sum().reset_index().sort_values('WTRACT_lookup')['PERWGT'].cumsum().values
wtract_sorted = chts_workers.groupby('WTRACT_lookup').PERWGT.sum().reset_index().sort_values('WTRACT_lookup')['WTRACT_lookup'].values
# assignment data
merged = pd.merge(chts_workers, jobs, left_on='job_id', right_index=True)
merged = pd.merge(merged, buildings, left_on='building_id', right_index=True)
merged = pd.merge(merged, parcels, left_on='parcel_id', right_index=True)
merged['WTRACT_assigned'] = merged['block_id'].str[5:11].astype(int)
perwgt_tract_count_assigned = merged.groupby('WTRACT_assigned').PERWGT.sum()
perwgt_cumsum_assigned = merged.groupby('WTRACT_assigned').PERWGT.sum().reset_index().sort_values('WTRACT_assigned')['PERWGT'].cumsum().values
wtract_sorted_assigned = merged.groupby('WTRACT_assigned').PERWGT.sum().reset_index().sort_values('WTRACT_assigned')['WTRACT_assigned'].values
perwgts = pd.concat([perwgt_tract_count, perwgt_tract_count_assigned], axis=1)
perwgts.columns = ['chts', 'assigned']
fig, ax = plt.subplots(figsize=(7,5))
ax.scatter(perwgts['chts'], perwgts['assigned'], alpha=0.2, s=10)
ax.set_xlim([0,15])
ax.set_ylim([0,15])
ax.set_xlabel('CHTS perwgt')
ax.set_ylabel('assigned perwgt')
fig, ax = plt.subplots(figsize=(7,5))
ax.plot(wtract_sorted, perwgt_cumsum, label='CHTS', lw=2, alpha=0.6)
ax.plot(wtract_sorted_assigned, perwgt_cumsum_assigned, label='assigned', lw=2, alpha=0.6)
ax.set_xlabel('work tract')
ax.set_ylabel('perwgt cumsum')
ax.legend()
```
### Assign CHTS persons to buildings
```
%%time
persons_home_rad = np.deg2rad(chts_persons[['HYCORD', 'HXCORD']])
bldg_rad = np.deg2rad(buildings[['y', 'x']])
buildings.loc[:, 'x_rad'] = bldg_rad['x']
buildings.loc[:, 'y_rad'] = bldg_rad['y']
tree = BallTree(buildings[['y_rad','x_rad']], metric='haversine')
dists, idxs = tree.query(persons_home_rad, return_distance=True)
km_dists = pd.Series([dist[0] * 6371 for dist in dists])
pd.Series(km_dists).plot(kind='hist', bins=100, xlim=(0,0.5))
```
Save the updated persons table so we don't have to re-run every time:
```
chts_persons['building_id'] = buildings.iloc[idxs[:,0]].index
chts_persons.to_csv('./data/chts_persons_w_jobs_and_res_bldgs.csv', index=False)
chts_persons = pd.read_csv('./data/chts_persons_w_jobs_and_res_bldgs.csv')
```
### Generate Accessibility Vars
```
%%time
orca.run(['initialize_network_small', 'network_aggregations_small'])
%%time
orca.run(['initialize_network_walk','network_aggregations_walk'])
walk_net_vars = orca.get_table('nodeswalk').to_frame()
drive_net_vars = orca.get_table('nodessmall').to_frame()
walk_net_vars.head()
drive_net_vars.head()
```
Save the network accessibility variables tables so we don't have to re-run every time:
```
walk_net_vars.to_csv('./data/walk_net_vars.csv')
drive_net_vars.to_csv('./data/drive_net_vars.csv')
```
### Generate the interaction terms
#### Travel Time
```
skims = pd.read_csv('./data/TimeSkimsDatabaseAM.csv')
interaction_terms = skims[['orig', 'dest', 'da', 'wTrnW']].rename(
columns={'orig': 'zone_id_home', 'dest': 'zone_id_work', 'da': 'tt_da', 'wTrnW': 'tt_wTrnW'})
interaction_terms.set_index(['zone_id_home', 'zone_id_work'], inplace=True)
interaction_terms.to_csv('./data/WLCM_interaction_terms_tt.csv')
```
#### Distance
```
skims = pd.read_csv('./data/DistanceSkimsDatabaseAM.csv')
interaction_terms = skims[['orig', 'dest', 'da', 'walk']].rename(
columns={'orig': 'zone_id_home', 'dest': 'zone_id_work', 'da': 'dist_da', 'walk': 'dist_walk'})
interaction_terms.set_index(['zone_id_home', 'zone_id_work'], inplace=True)
interaction_terms.to_csv('./data/WLCM_interaction_terms_dist.csv')
```
#### Cost
```
skims = pd.read_csv('./data/CostSkimsDatabaseAM.csv')
interaction_terms = skims[['orig', 'dest', 'daToll', 'wTrnW']].rename(
columns={'orig': 'zone_id_home', 'dest': 'zone_id_work', 'daToll': 'cost_da_toll', 'wTrnW': 'cost_wTrnW'})
interaction_terms.set_index(['zone_id_home', 'zone_id_work'], inplace=True)
interaction_terms.to_csv('./data/WLCM_interaction_terms_cost.csv')
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
from IPython.display import Image
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
import os
import json
import jax.numpy as np
import numpy as onp
import jax
import pickle
import matplotlib.pyplot as plt
import pandas as pd
from timecast.learners import AR
from timecast.learners._ar import _ar_predict, _ar_batch_window
from timecast.utils.numpy import ecdf
from timecast.utils.losses import MeanSquareError
import torch
import matplotlib
plt.rcParams['figure.figsize'] = [20, 10]
import tqdm.notebook as tqdm
from ealstm.gaip import FloodLSTM
from ealstm.gaip import FloodData
from ealstm.gaip.utils import MSE, NSE
from timecast.optim import SGD
from timecast.learners import AR
cfg_path = "/home/dsuo/src/toy_flood/ealstm/runs/run_2503_0429_seed283956/cfg.json"
flood_lstm = FloodLSTM(cfg_path)
flood_data = FloodData(cfg_path)
ea_data = pickle.load(open("../ealstm/runs/run_2503_0429_seed283956/lstm_seed283956.p", "rb"))
results = {}
date_range = pd.date_range(start=flood_data.cfg["val_start"], end=flood_data.cfg["val_end"])
for X, y, basin in tqdm.tqdm(flood_data.generator(), total=len(flood_data.basins)):
pred = flood_lstm.predict(X)
true = y[flood_data.cfg["seq_length"] - 1:]
df = pd.DataFrame(data={"qobs": true.ravel(), "qsim": pred.ravel()}, index=date_range)
results[basin] = df
for basin in tqdm.tqdm(flood_data.basins):
onp.testing.assert_array_almost_equal(results[basin], ea_data[basin], decimal=4)
from ealstm.gaip import FloodLSTM
from ealstm.gaip import FloodData
from ealstm.gaip.utils import MSE, NSE
from timecast.optim import SGD
from timecast.learners import AR
cfg_path = "/home/dsuo/src/toy_flood/ealstm/runs/run_2503_0429_seed283956/cfg.json"
flood_data = FloodData(cfg_path)
LR_AR = 1e-5
AR_INPUT_DIM=32
AR_OUTPUT_DIM=1
results = {}
mses = []
nses = []
for X, _, basin in tqdm.tqdm(flood_data.generator(), total=len(flood_data.basins)):
sgd = SGD(learning_rate=LR_AR, online=False)
ar = AR(input_dim=AR_INPUT_DIM,
output_dim=AR_OUTPUT_DIM,
window_size=flood_data.cfg["seq_length"],
optimizer=sgd,
history=X[:flood_data.cfg["seq_length"]],
fit_intercept=True,
constrain=False
)
# NOTE: difference in indexing convention, so need to pad one row
X = np.vstack((X[flood_data.cfg["seq_length"]:], np.ones((1, X.shape[1]))))
Y = np.array(ea_data[basin].qobs).reshape(-1, 1)
Y_lstm = np.array(ea_data[basin].qsim).reshape(-1, 1)
Y_target = Y - Y_lstm
Y_ar = ar.predict_and_update(X, Y_target)
Y_hat = Y_lstm + Y_ar
mse = MSE(Y, Y_hat)
nse = NSE(Y, Y_hat)
results[basin] = {
"mse": mse,
"nse": nse,
"count": X.shape[0],
"avg_mse": np.mean(np.array(mses)),
"avg_nse": np.mean(np.array(nses))
}
mses.append(mse)
nses.append(nse)
print(basin, mse, nse, np.mean(np.array(mses)), np.mean(np.array(nses)))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/john-s-butler-dit/Intro-to-Algorithms/blob/master/Chapter%201-%20Introduction_to_Algorithms/Analysis%20of%20an%20Algorithm.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Analysis of an Algorithm
## Cost Function
The cost function $C(n)$ is how many operations it takes an algorithm to perform a task of length $n$.
For example, to sort a list of $n$ numbers using the selection sort algorithm costs
$$T(n)=\frac{n(n-1)}{2}.$$
For example, to find a number in a list of sorted numbers of lenght $n$ using the binary search algorithm costs
$$T(n)=\log_2(n).$$
```
import numpy as np
import math
%matplotlib inline
import matplotlib.pyplot as plt # side-stepping mpl backend
import matplotlib.gridspec as gridspec # subplots
import warnings
import pandas as pd
warnings.filterwarnings("ignore")
```
## Linear Algorithm
A linear Algorithm is one that the cost function is linear
$$T(n)=an+b, $$
where $a$ and $b$ are real numbers.
```
## FUNCTION TO PLOT COST OF ALGORITHM AS A FUNCTION OF N
def Analysis_of_Algorithm(n,T):
fig = plt.figure(figsize=(12,3))
ax=fig.add_subplot(1,2,1) # PLOT OF T(N) AS A FUNCTION OF N
plt.plot(n,T,'o:',color='red')
plt.title('Cost as a function of n')
plt.xlabel('n')
plt.ylabel('T(n)')
# LOG TRANSFORM OF n AND T
log_n=np.log2(n)
log_T=np.log2(T)
ax=fig.add_subplot(1,2,2) # PLOT OF LOG TRANSFORM DATA
plt.plot(log_n,log_T,'o:',color='blue') # PLOT OF log(T(N)) AS A FUNCTION OF log(N)
plt.title('Log Transformed')
plt.xlabel(r'$\log_2(n)$')
plt.ylabel(r'$\log_2(T(n))$')
plt.plot();
## FUNCTION OUTPUT A TABLE COST OF ALGORITHM AS A FUNCTION OF N
def Analysis_of_Algorithm_Table(n,T):
slope=(np.diff(T)/np.diff(n))
log_n=np.log2(n)
log_T=np.log2(T)
log_slope=(np.diff(log_T)/np.diff(log_n))
# table.append(['n', 'T(n)',r'$\log_2(n)$',r'$\log_2(T(n))$'])
# for i in range(len(n)):
# table.append([round(n[i],5), round(T[i],5), round(log_n[i],7),round(log_T[i],7)])
# print(np.diff(log_T)/np.diff(log_n))
d = {'n ': np.round(n,5), 'T(n) ':np.round(T,5),'log2(n)':np.round(log_n,7),'log2(T(n))':np.round(log_T,7)}
df = pd.DataFrame(data=d)
d = {'Slope ': slope, 'log(Slope) ':log_slope}
df_slope = pd.DataFrame(data=d)
df_slope
return df, df_slope
```
### Example Linear Algorithm
The figure below show a plot of linear algorithm,
$$ T(n)=4n+10,$$
on the left.
```
n=2**np.arange(4,11,2)
T=4*n+10
print('The plots below shows the Cost as a function of n (left) and the log transformed (right).')
Analysis_of_Algorithm(n,T)
df,df_slope=Analysis_of_Algorithm_Table(n,T)
df.to_csv('Linear.csv')
df
df_slope.to_csv('Linear_slope.csv')
df_slope
```
## Quadratic Algorithm
A quadratic Algorithm is one that the cost function is of the form:
$$T(n)=an^2+bn+c, $$
where $a$, $b$ and $c$ are real numbers.
## Example Quadratic Algorithm
```
n=2**np.arange(10,18,2)
T=2*n**2++200*n+100
Analysis_of_Algorithm(n,T)
df, df_slope=Analysis_of_Algorithm_Table(n,T)
df.to_csv('Quad.csv')
df
df_slope.to_csv('Quad_slope.csv')
df_slope
```
## Cubic Algorithm
A quadratic Algorithm is one that the cost function is of the form:
$$T(n)=an^3+bn^2+cn+d, $$
where $a$, $b$, $c$ and $d$ are real numbers.
## Example Cubic Algorithm
```
T=0.1*n**3+n**2-n+1000
Analysis_of_Algorithm(n,T)
df, df_slope=Analysis_of_Algorithm_Table(n,T)
df.to_csv('Cubic.csv')
df
df_slope.to_csv('Cubic_slope.csv')
df_slope
```
## Different Powers Algorithm
A different power Algorithm is one that the cost function is of the form:
$$T(n)=an^{0.5}+b, $$
where $a$ and $b$ are real numbers.
```
T=2*n**0.5+12
Analysis_of_Algorithm(n,T)
df, df_slope=Analysis_of_Algorithm_Table(n,T)
df
df_slope
```
## It does not work so well for Log Algorithms
A log Algorithm is one that the cost function is of the form:
$$T(n)=an^0.5+b, $$
where $a$ and $b$ are real numbers.
```
T=np.log(n)+1
Analysis_of_Algorithm(n,T)
df, df_slope=Analysis_of_Algorithm_Table(n,T)
df
df_slope
```
## Tutorial Sheet 2
## Question 6
The order of growth of an algorithm, T (n), was measured experimentally for various values of n and the results are shown below:
n operations | 1 |2 | 3| 4|
Growth T (n)| 2| 3 |4 | 5|
```
n=[1,2,3,4]
T=[2,3,4,5]
d = {'n ': n, 'Cost (T(n)': T}
df = pd.DataFrame(data=d)
df
Analysis_of_Algorithm(n,T)
df, df_slope=Analysis_of_Algorithm_Table(n,T)
df
df_slope
```
## Tutorial Sheet 2
## Question 7
7. The order of growth of an algorithm, T (n), was measured experimentally for various values of n and the results are shown below:
n operations 1 2 3 4
Growth T (n) 10 17 36 73
```
n=[1,2,3,4]
T=[10,17,36,73]
d = {'n ': n, 'Cost (T(n)': T}
df = pd.DataFrame(data=d)
df
Analysis_of_Algorithm(n,T)
df, df_slope=Analysis_of_Algorithm_Table(n,T)
df
df_slope
```
## Question 8.
The order of growth of an algorithm, T (n), was measured experimentally for various values of n and the results are shown below:
n operations 16 64 256 1024
Growth T (n) 1512 9192 132072 2098152
```
n=[16, 64, 256, 1024]
T=[1512, 9192, 132072, 2098152]
d = {'n ': n, 'Cost (T(n)': T}
df = pd.DataFrame(data=d)
df
Analysis_of_Algorithm(n,T)
df, df_slope=Analysis_of_Algorithm_Table(n,T)
df
df_slope
```
## Question 9
The order of growth of an algorithm, T (n), was measured experimentally for various values of n and the results are shown below:
n operations 256 1024 4096 16384
Growth T (n) 44.0 76.0 140.0 268.0
```
n=[256, 1024, 4096, 16384]
T=[44.0, 76.0, 140.0, 268.0]
d = {'n ': n, 'Cost (T(n)': T}
df = pd.DataFrame(data=d)
df
Analysis_of_Algorithm(n,T)
df, df_slope=Analysis_of_Algorithm_Table(n,T)
df
df_slope
```
| github_jupyter |
## Importing and prepping data
```
import pandas as pd
import numpy as np
import diff_classifier.aws as aws
import diff_classifier.pca as pca
features = []
remote_folder = 'Gel_Studies/11_09_18_gel_experiment' #Folder in AWS S3 containing files to be analyzed
bucket = 'ccurtis.data'
vids = 20
gels = ['0_4', '0_6', '0_8', '1_0', '1_2']
nonnum = ['Percent Agarose', 'Well', 'Video Number', 'Track_ID', 'Mean Mean_Intensity', 'Std Mean_Intensity', 'X', 'Y',
'Mean X', 'Mean Y', 'Std X', 'Std Y']
featofvar = 'Percent Agarose'
counter = 0
for gel in gels:
for num in range(1, vids+1):
try:
filename = 'features_Gels_{}_XY{}.csv'.format(gel, '%02d' % num)
aws.download_s3('{}/{}'.format(remote_folder, filename), filename, bucket_name='ccurtis.data')
fstats = pd.read_csv(filename, encoding = "ISO-8859-1", index_col='Unnamed: 0')
print('{} size: {}'.format(filename, fstats.shape))
fstats['Percent Agarose'] = pd.Series(fstats.shape[0]*[gel], index=fstats.index)
fstats['Video Number'] = pd.Series(fstats.shape[0]*[num], index=fstats.index)
if num < 11:
fstats['Well'] = pd.Series(fstats.shape[0]*[1], index=fstats.index)
else:
fstats['Well'] = pd.Series(fstats.shape[0]*[2], index=fstats.index)
counter = counter + 1
if counter == 1:
fstats_tot = fstats
else:
fstats_tot = fstats_tot.append(fstats, ignore_index=True)
except:
print('skip filename {}'.format(filename))
#PCA analyses with too many datapoints fail. You get rows with lots of NAs. I'm going to try making a subset of the data first
#and then do a PCA analysis on that.
#include all in analysis
import random
subset = np.sort(np.array(random.sample(range(fstats_tot.shape[0]), 500000)))
fstats_sub = fstats_tot.loc[subset, :].reset_index(drop=True)
for typ in fstats_tot['Particle Size'].unique():
fstats_type = fstats_tot[fstats_tot['Particle Size']==typ].reset_index(drop=True)
print(fstats_type.shape)
#with equal sample sizes for each particle type
import random
counter = 0
for typ in fstats_tot['Particle Size'].unique():
fstats_type = fstats_tot[fstats_tot['Particle Size']==typ].reset_index(drop=True)
print(fstats_type.shape)
subset = np.sort(np.array(random.sample(range(fstats_type.shape[0]), 55000)))
if counter == 0:
fstats_sub = fstats_type.loc[subset, :].reset_index(drop=True)
else:
fstats_sub = fstats_sub.append(fstats_type.loc[subset, :].reset_index(drop=True), ignore_index=True)
counter = counter + 1
#fstats = pd.read_csv(filename, encoding = "ISO-8859-1", index_col='Unnamed: 0')
fstats_num = fstats_tot.drop(nonnum, axis=1)
fstats_raw = fstats_num.as_matrix()
#fstats
```
## PCA analysis
The pca.pca_analysis function provides a completely contained PCA analysis of the input trajectory features dataset. It includes options to impute NaN values (fill in with average values or drop them), and to scale features. Read the docstring for more information.
```
pcadataset = pca.pca_analysis(fstats_tot, dropcols=nonnum, n_components=16)
pcadataset.components.to_csv('components.csv')
aws.upload_s3('components.csv', '{}/components.csv'.format(remote_folder, filename), bucket_name=bucket)
fstats_num.columns
kmostat = pca.kmo(pcadataset.scaled)
```
The pca.kmo function calculates the Kaiser-Meyer-Olkin statistic, a measure of sampling adequacy. Check the docstring for more information.
```
import scipy.stats as stat
stat.bartlett(pcadataset.scaled[0, :], pcadataset.scaled[1, :], pcadataset.scaled[2, :], pcadataset.scaled[3, :])
newstr = ''
for i in range(pcadataset.scaled.shape[0]-1):
newstr = newstr + 'pcadataset.scaled[{}, :], '.format(i)
newstr = 'stat.bartlett(' + newstr + 'pcadataset.scaled[{}, :])'.format(i+1)
test = eval(newstr)
test
```
## Visualization
Users can then compare average principle component values between subgroups of the data. In this case, all particles were taken from the same sample, so there are no experimental subgroups. I chose to compare short trajectories to long trajectories, as I would expect differences between the two groups.
```
import numpy as np
ncomp = 16
dicti = {}
#test = np.exp(np.nanmean(np.log(pcadataset.final[pcadataset.final['Particle Size']==200].as_matrix()), axis=0))[-6:]
#test1 = np.exp(np.nanmean(np.log(pcadataset.final[pcadataset.final['Particle Size']==500].as_matrix()), axis=0))[-6:]
dicti[0] = np.nanmean(pcadataset.final[pcadataset.final['Percent Agarose']=='0_4'].values[:, -ncomp:], axis=0)
dicti[1] = np.nanmean(pcadataset.final[pcadataset.final['Percent Agarose']=='0_6'].values[:, -ncomp:], axis=0)
dicti[2] = np.nanmean(pcadataset.final[pcadataset.final['Percent Agarose']=='0_8'].values[:, -ncomp:], axis=0)
dicti[3] = np.nanmean(pcadataset.final[pcadataset.final['Percent Agarose']=='1_0'].values[:, -ncomp:], axis=0)
dicti[4] = np.nanmean(pcadataset.final[pcadataset.final['Percent Agarose']=='1_2'].values[:, -ncomp:], axis=0)
pca.plot_pca(dicti, savefig=True, labels=['0.4%', '0.6%', '0.8%', '1.0%', '1.2%'], rticks=np.linspace(-4, 4, 9))
```
The variable pcadataset.prcomps shows the user the major contributions to each of the new principle components. When observing the graph above, users can see that there are some differences between short trajectories and long trajectories in component 0 (asymmetry1 being the major contributor) and component 1 (elongation being the major contributor).
```
pcadataset.prcomps
labels=gels
feats = pca.feature_violin(pcadataset.final, label='Percent Agarose', lvals=labels, fsubset=ncomp, yrange=[-12, 12])
fstats1 = pca.feature_plot_3D(pcadataset.final, label='Percent Agarose', lvals=labels, randcount=400, ylim=[-12, 12],
xlim=[-12, 12], zlim=[-12, 12])
ncomp = 14
trainp = np.array([])
testp = np.array([])
for i in range(0, 20):
KNNmod, X, y = pca.build_model(pcadataset.final, 'Percent Agarose', labels, equal_sampling=True,
tsize=500, input_cols=ncomp, model='MLP', NNhidden_layer=(6, 2))
trainp = np.append(trainp, pca.predict_model(KNNmod, X, y))
X2 = pcadataset.final.values[:, -ncomp:]
y2 = pcadataset.final['Particle Size'].values
testp = np.append(testp, pca.predict_model(KNNmod, X2, y2))
print('Run {}: {}'.format(i, testp[i]))
print('{} +/ {}'.format(np.mean(trainp), np.std(trainp)))
print('{} +/ {}'.format(np.mean(testp), np.std(testp)))
fstats_num
```
## Neural Network
```
from sklearn.model_selection import GridSearchCV, cross_val_score, KFold, train_test_split
from sklearn.neural_network import MLPClassifier
from sklearn.metrics import classification_report
featofvar = 'Percent Agarose'
test = pcadataset.final.values[:, -ncomp:]
y = pcadataset.final[featofvar].values
for run in range(1):
X_train, X_test, y_train, y_test = train_test_split(test, y, test_size=0.4)
clf = MLPClassifier(hidden_layer_sizes=(900, ), solver='sgd', verbose=True, max_iter=500, tol=0.00001,
alpha=0.001, batch_size=50, learning_rate_init=0.005, learning_rate='adaptive',
early_stopping=True, validation_fraction=0.1)
clf.fit(X_train, y_train)
print('Training Results')
y_true1, y_pred1 = y_train, clf.predict(X_train)
print(classification_report(y_true1, y_pred1, digits=4))
print('Test Results')
print()
y_true, y_pred = y_test, clf.predict(X_test)
print(classification_report(y_true, y_pred, digits=4))
print('Training Results')
y_true1, y_pred1 = y_train, clf.predict(X_train)
print(classification_report(y_true1, y_pred1, digits=4))
print('Test Results')
print()
y_true, y_pred = y_test, clf.predict(X_test)
print(classification_report(y_true, y_pred, digits=4))
featofvar = 'Percent Agarose'
ncomp=16
test = pcadataset.final[pcadataset.final['Percent Agarose'].isin(['0_4', '0_8', '1_2'])].values[:, -ncomp:]
y = pcadataset.final[pcadataset.final['Percent Agarose'].isin(['0_4', '0_8', '1_2'])][featofvar].values
for run in range(1):
X_train, X_test, y_train, y_test = train_test_split(test, y, test_size=0.4)
clf = MLPClassifier(hidden_layer_sizes=(900, ), solver='sgd', verbose=True, max_iter=500, tol=0.00001,
alpha=0.001, batch_size=50, learning_rate_init=0.005, learning_rate='adaptive',
early_stopping=True, validation_fraction=0.1)
clf.fit(X_train, y_train)
print('Training Results')
y_true1, y_pred1 = y_train, clf.predict(X_train)
print(classification_report(y_true1, y_pred1, digits=4))
print('Test Results')
print()
y_true, y_pred = y_test, clf.predict(X_test)
print(classification_report(y_true, y_pred, digits=4))
fstats_play = fstats_tot.dropna(subset=['Deff1'])
kb = 1.381*10**-23
T = 313
nu = 0.000397
#size2 = 2*10**9*10**12*kb*T/(fstats_tot.Deff1.values*6*np.pi*nu)
Draw = kb*T*10**9*10**12/(6*np.pi*nu*140)
Drat = fstats_play.Deff1/Draw
from scipy.optimize import curve_fit
def pore_model(x, a, b):
return np.exp(-np.pi*((70+b)/(a*(x**-0.75)+2*b))**2)
means1 = np.log(pore_model(np.array([0.4, 0.6, 0.8, 1.0, 1.2]), a, b)*Draw)
means1
Dbins2 = np.log(pore_model(np.array([0.5, 0.7, 0.9, 1.1]), a, b)*Draw)
Dbins2
import matplotlib.pyplot as plt
Dbins = [-10, 0.46210, 0.665, 0.865, 1.0576, 10]
bins = np.linspace(-10, 10, 200)
fig, axes = plt.subplots(nrows=5, figsize=(12, 13))
counter = 0
means = []
for ax in axes:
means.append(fstats_tot[fstats_tot['Percent Agarose']==gels[counter]]['LogDeff1'].median())
for i in range(5):
fstats_tot[(fstats_tot['Percent Agarose']==gels[counter]) & (Dbins[i] < fstats_tot['LogDeff1']) & (fstats_tot['LogDeff1'] < Dbins[i+1])].hist(column='LogDeff1', bins=bins, figsize=(12,3), edgecolor='k', ax=ax, )
ax.set_xlim([-7.5, 3.5])
ax.set_ylim([0, 10000])
ax.axvline(means1[counter], color='k', linestyle='dashed', linewidth=3)
ax.set_title(gels[counter]+ 'nm')
if counter == 4:
ax.set_xlabel(r'$log(D_{eff})$')
counter = counter + 1
pore_model(ngels, 100, 40)
fstats_tot['Percent Agarose Int'] = fstats_tot['Percent Agarose']
fstats_tot['Percent Agarose Int'] = fstats_tot['Percent Agarose Int'].replace('0_4', 0.4)
fstats_tot['Percent Agarose Int'] = fstats_tot['Percent Agarose Int'].replace('0_6', 0.6)
fstats_tot['Percent Agarose Int'] = fstats_tot['Percent Agarose Int'].replace('0_8', 0.8)
fstats_tot['Percent Agarose Int'] = fstats_tot['Percent Agarose Int'].replace('1_0', 1.0)
fstats_tot['Percent Agarose Int'] = fstats_tot['Percent Agarose Int'].replace('1_2', 1.2)
fstats_tot['Percent Agarose Int'].mean()
nmeans = np.exp(np.array(means))/Draw
nmeans
ngels = np.array([0.4, 0.6, 0.8, 1.0, 1.2])
ngels
popt, pcov = curve_fit(pore_model, ngels, nmeans, p0=[100, 40])
a = popt[0]
b = popt[1]
gelpred = (((70+b)/np.sqrt(-np.log(Drat)/np.pi)) - 2*b)/a
gelpred
y_pred2 = pd.Series(list(pd.cut(gelpred.values, bins=[0, 0.5, 0.7, 0.9, 1.1, 100000], labels=['0_4', '0_6', '0_8', '1_0', '1_2']).astype(str)))
ypred2 = y_pred2.replace('nan', '0_4')
ytrue2 = fstats_play['Percent Agarose']
print(classification_report(ytrue2, ypred2, digits=4))
fstats_tot['LogDeff1'] = np.log(fstats_tot.Deff1).replace([np.inf, -np.inf], np.nan)
meanD = np.array(means)
meanD.sort()
Dbins = meanD[0:-1] + np.diff(meanD)/2
print(Dbins)
Drat1 = fstats_play[fstats_play['Percent Agarose'].isin(['0_4', '0_8', '1_2'])].Deff1/Draw
gelpred1 = (((70+b)/np.sqrt(-np.log(Drat1)/np.pi)) - 2*b)/a
y_pred3 = pd.Series(list(pd.cut(gelpred1.values, bins=[0, 0.6, 1.0, 100000], labels=['0_4', '0_8', '1_2']).astype(str)))
ypred3 = y_pred3.replace('nan', '0_4')
ytrue3 = fstats_play[fstats_play['Percent Agarose'].isin(['0_4', '0_8', '1_2'])]['Percent Agarose']
print(classification_report(ytrue3, ypred3, digits=4))
import matplotlib.pyplot as plt
Dbins = [-10, 0.520, 0.5686, 0.7651, 1.0865, 10]
bins = np.linspace(-10, 10, 200)
fig, axes = plt.subplots(nrows=5, figsize=(12, 13))
counter = 0
means = []
for ax in axes:
means.append(fstats_tot[fstats_tot['Percent Agarose']==gels[counter]]['LogDeff1'].median())
for i in range(5):
fstats_tot[(fstats_tot['Percent Agarose']==gels[counter]) & (Dbins[i] < fstats_tot['LogDeff1']) & (fstats_tot['LogDeff1'] < Dbins[i+1])].hist(column='LogDeff1', bins=bins, figsize=(12,3), edgecolor='k', ax=ax, )
ax.set_xlim([-7.5, 3.5])
ax.set_ylim([0, 10000])
ax.axvline(fstats_tot[fstats_tot['Percent Agarose']==gels[counter]]['LogDeff1'].median(), color='k', linestyle='dashed', linewidth=3)
ax.set_title(gels[counter]+ 'nm')
if counter == 4:
ax.set_xlabel(r'$log(D_{eff})$')
counter = counter + 1
y_true2 = fstats_tot['Percent Agarose'].values
y_pred2 = list(pd.cut(fstats_tot['LogDeff1'].values, bins=Dbins, labels=['1_2', '1_0', '0_8', '0_6', '0_4']).astype(str))
print(classification_report(y_true2, y_pred2, digits=4))
meanD = np.array(means)
meanD.sort()
Dbins = meanD[0:-1] + np.diff(meanD)/2
print(Dbins)
import matplotlib.pyplot as plt
Dbins = [-10, 0.56440, 0.9344, 10]
bins = np.linspace(-10, 10, 200)
fig, axes = plt.subplots(nrows=3, figsize=(12, 13))
gels1 = ['0_4', '0_8', '1_2']
counter = 0
means = []
for ax in axes:
means.append(fstats_tot[fstats_tot['Percent Agarose']==gels1[counter]]['LogDeff1'].median())
for i in range(3):
fstats_tot[(fstats_tot['Percent Agarose']==gels1[counter]) & (Dbins[i] < fstats_tot['LogDeff1']) & (fstats_tot['LogDeff1'] < Dbins[i+1])].hist(column='LogDeff1', bins=bins, figsize=(12,3), edgecolor='k', ax=ax, )
ax.set_xlim([-7.5, 3.5])
ax.set_ylim([0, 10000])
ax.axvline(fstats_tot[fstats_tot['Percent Agarose']==gels1[counter]]['LogDeff1'].median(), color='k', linestyle='dashed', linewidth=3)
ax.set_title(gels1[counter]+ 'nm')
if counter == 2:
ax.set_xlabel(r'$log(D_{eff})$')
counter = counter + 1
y_true2 = fstats_tot[fstats_tot['Percent Agarose'].isin(['0_4', '0_8', '1_2'])]['Percent Agarose'].values
y_pred2 = list(pd.cut(fstats_tot[fstats_tot['Percent Agarose'].isin(['0_4', '0_8', '1_2'])]['LogDeff1'].values, bins=Dbins, labels=['1_2', '0_8', '0_4']).astype(str))
print(classification_report(y_true2, y_pred2, digits=4))
fstats_tot['LogMeanDeff1'] = np.log(fstats_tot['Mean Deff1']).replace([np.inf, -np.inf], np.nan)
meanD = np.array(means)
meanD.sort()
Dbins = meanD[0:-1] + np.diff(meanD)/2
print(Dbins)
means
import matplotlib.pyplot as plt
Dbins = [-10, 0.6748, 0.7152, 0.8690, 1.172, 10]
bins = np.linspace(-10, 10, 200)
fig, axes = plt.subplots(nrows=5, figsize=(12, 11))
counter = 0
means = []
for ax in axes:
means.append(fstats_tot[fstats_tot['Percent Agarose']==gels[counter]]['LogMeanDeff1'].median())
for i in range(5):
fstats_tot[(fstats_tot['Percent Agarose']==gels[counter]) & (Dbins[i] < fstats_tot['LogMeanDeff1']) & (fstats_tot['LogMeanDeff1'] < Dbins[i+1])].hist(column='LogMeanDeff1', bins=bins, figsize=(12,3), edgecolor='k', ax=ax, )
ax.set_xlim([-6.5, 3.5])
ax.set_ylim([0, 12000])
ax.axvline(fstats_tot[fstats_tot['Percent Agarose']==gels[counter]]['LogMeanDeff1'].median(), color='k', linestyle='dashed', linewidth=3)
ax.set_title(gels[counter]+ 'nm')
if counter == 2:
ax.set_xlabel(r'$log(D_{eff})$')
counter = counter + 1
y_true2 = fstats_tot['Percent Agarose'].values
y_pred2 = list(pd.cut(fstats_tot['LogMeanDeff1'].values, bins=Dbins, labels=['1_2', '1_0', '0_8', '0_6', '0_4']).astype(str))
print(classification_report(y_true2, y_pred2, digits=4))
import matplotlib.pyplot as plt
fig, ax1 = plt.subplots()
ax1.plot(clf.loss_curve_, linewidth=4)
#ax1.set_xlim([0, 60])
#ax1.set_ylim([0.04, 0.18])
ax1.set_ylabel('Loss Curve')
ax2 = ax1.twinx()
ax2.plot(clf.validation_scores_, linewidth=4, c='g')
#ax2.set_ylim([0.94, 0.99])
ax2.set_ylabel('Validation Scores')
print('Training Results')
y_true1, y_pred1 = y_train, clf.predict(X_train)
print(classification_report(y_true1, y_pred1, digits=4))
print('Test Results')
print()
y_true, y_pred = y_test, clf.predict(X_test)
print(classification_report(y_true, y_pred, digits=4))
classification_report()
Deff1 = fstats_sub.Deff1.values
bins = [0, 1.872, 3.858, 10**20]
binned = np.histogram(Deff1, [0, 1.872, 3.858, 10**20])
binned
y_pred2 = list(pd.cut(fstats_tot.Deff1.values, bins=[0, 1.872, 3.858, 1000], labels=['500', '200', '100']).astype(str))
y_true2 = fstats_tot['Particle Size'].values
print(classification_report(y_true2, y_pred2, digits=4))
kb = 1.381*10**-23
T = 303
nu = 0.000797
size2 = 2*10**9*10**12*kb*T/(fstats_tot.Deff1.values*6*np.pi*nu)
y_pred2 = list(pd.cut(size2, bins=[-100, 150, 350, 1000000], labels=['100', '200', '500']).astype(str))
print(classification_report(y_true2, y_pred2, digits=4))
fstats_tot.hist(column='Deff1', by='Particle Size', sharex=True, bins=np.linspace(0, 10, 100),
figsize=(9, 7), grid=False, layout=(3,1), sharey=True, )
fstats_tot['LogDeff1'] = np.log(fstats_tot.Deff1).replace([np.inf, -np.inf], np.nan)
bins = np.linspace(-10, 10, 200)
axes = fstats_tot.hist(column='LogDeff1', by='Particle Size', layout=(3, 1), bins=bins, sharex=True, sharey=True,
figsize=(10, 8), edgecolor='k')
means = []
types2 = ['100', '200', '500']
for ax, typ in zip(axes, types2):
ax.set_ylim([0,10000])
#ax.set_xscale("log", nonposx='clip')
ax.set_xlim([-7.5,3.5])
means.append(fstats_tot[fstats_tot['Particle Size']==typ]['LogDeff1'].median())
ax.axvline(fstats_tot[fstats_tot['Particle Size']==typ]['LogDeff1'].median(), color='k', linestyle='dashed', linewidth=3)
means
meanD = np.array(means)
meanD.sort()
Dbins = meanD[0:-1] + np.diff(meanD)/2
print(Dbins)
import matplotlib.pyplot as plt
Dbins = [-10, -0.233, 0.895, 10]
bins = np.linspace(-10, 10, 200)
fig, axes = plt.subplots(nrows=3, figsize=(12, 9))
counter = 0
means = []
for ax in axes:
means.append(fstats_tot[fstats_tot['Particle Size']==sizes[counter]]['LogDeff1'].median())
for i in range(3):
fstats_tot[(fstats_tot['Particle Size']==sizes[counter]) & (Dbins[i] < fstats_tot['LogDeff1']) & (fstats_tot['LogDeff1'] < Dbins[i+1])].hist(column='LogDeff1', bins=bins, figsize=(12,3), edgecolor='k', ax=ax, )
ax.set_xlim([-7.5, 3.5])
ax.set_ylim([0, 10000])
ax.axvline(fstats_tot[fstats_tot['Particle Size']==sizes[counter]]['LogDeff1'].median(), color='k', linestyle='dashed', linewidth=3)
ax.set_title(sizes[counter]+ 'nm')
if counter == 2:
ax.set_xlabel(r'$log(D_{eff})$')
counter = counter + 1
y_true2 = fstats_tot['Particle Size'].values
y_pred2 = list(pd.cut(fstats_tot['LogDeff1'].values, bins=Dbins, labels=['500', '200', '100']).astype(str))
print(classification_report(y_true2, y_pred2, digits=4))
fstats_tot['LogMeanDeff1'] = np.log(fstats_tot['Mean Deff1']).replace([np.inf, -np.inf], np.nan)
meanD = np.array(means)
meanD.sort()
Dbins = meanD[0:-1] + np.diff(meanD)/2
print(Dbins)
Dbins = [-10, -0.0937, 0.9617, 10]
bins = np.linspace(-10, 10, 200)
fig, axes = plt.subplots(nrows=3, figsize=(12, 9))
counter = 0
means = []
for ax in axes:
means.append(fstats_tot[fstats_tot['Particle Size']==sizes[counter]]['LogMeanDeff1'].median())
for i in range(3):
fstats_tot[(fstats_tot['Particle Size']==sizes[counter]) & (Dbins[i] < fstats_tot['LogMeanDeff1']) & (fstats_tot['LogMeanDeff1'] < Dbins[i+1])].hist(column='LogMeanDeff1', bins=bins, figsize=(12,3), edgecolor='k', ax=ax, )
ax.set_xlim([-7.5, 3.5])
ax.set_ylim([0, 10000])
ax.axvline(fstats_tot[fstats_tot['Particle Size']==sizes[counter]]['LogMeanDeff1'].median(), color='k', linestyle='dashed', linewidth=3)
ax.set_title(sizes[counter]+ 'nm')
if counter == 2:
ax.set_xlabel(r'$log(D_{eff})$')
counter = counter + 1
y_true2 = fstats_tot['Particle Size'].values
y_pred2 = list(pd.cut(fstats_tot['LogMeanDeff1'].values, bins=Dbins, labels=['500', '200', '100']).astype(str))
print(classification_report(y_true2, y_pred2, digits=4))
y_pred2 = list(pd.cut(size2, bins=[-100, 150, 350, 1000000], labels=['100', '200', '500']).astype(str))
print(classification_report(y_true2, y_pred2, digits=4))
size3 = np.random.rand(len(size2))
y_pred2 = list(pd.cut(size3, bins=[0, 0.33333333, 0.666666666, 10], labels=['100', '200', '500']).astype(str))
print(classification_report(y_true2, y_pred2, digits=4))
size2 = 2*10**9*10**12*kb*T/(fstats_tot['Mean Deff1'].values*6*np.pi*nu)
y_pred2 = list(pd.cut(size2, bins=[-100, 150, 350, 1000000], labels=['100', '200', '500']).astype(str))
print(classification_report(y_true2, y_pred2, digits=4))
kb = 1.381*10**-23
T = 303
nu = 0.000797
sizes3 = np.array([150, 350])
Dbins2 = np.log(2*10**9*10**12*kb*T/(sizes3*6*np.pi*nu))
Dbins2.sort()
Dbins2
import matplotlib.pyplot as plt
Dbins = [-10, 0.4648, 1.3120, 10]
bins = np.linspace(-10, 10, 200)
fig, axes = plt.subplots(nrows=3, figsize=(12, 9))
counter = 0
means = []
Ds = np.log(2*10**9*10**12*kb*T/(np.array([100, 200, 500])*6*np.pi*nu))
for ax in axes:
means.append(fstats_tot[fstats_tot['Particle Size']==sizes[counter]]['LogDeff1'].median())
for i in range(3):
fstats_tot[(fstats_tot['Particle Size']==sizes[counter]) & (Dbins[i] < fstats_tot['LogDeff1']) & (fstats_tot['LogDeff1'] < Dbins[i+1])].hist(column='LogDeff1', bins=bins, figsize=(12,3), edgecolor='k', ax=ax, )
ax.set_xlim([-7.5, 3.5])
ax.set_ylim([0, 10000])
ax.axvline(Ds[counter], color='k', linestyle='dashed', linewidth=3)
ax.set_title(sizes[counter]+ 'nm')
if counter == 2:
ax.set_xlabel(r'$log(D_{eff})$')
counter = counter + 1
import matplotlib.pyplot as plt
Dbins = [-10, 0.4648, 1.3120, 10]
bins = np.linspace(-10, 10, 200)
fig, axes = plt.subplots(nrows=3, figsize=(12, 9))
counter = 0
means = []
Ds = np.log(2*10**9*10**12*kb*T/(np.array([100, 200, 500])*6*np.pi*nu))
for ax in axes:
means.append(fstats_tot[fstats_tot['Particle Size']==sizes[counter]]['LogMeanDeff1'].median())
for i in range(3):
fstats_tot[(fstats_tot['Particle Size']==sizes[counter]) & (Dbins[i] < fstats_tot['LogMeanDeff1']) & (fstats_tot['LogMeanDeff1'] < Dbins[i+1])].hist(column='LogMeanDeff1', bins=bins, figsize=(12,3), edgecolor='k', ax=ax, )
ax.set_xlim([-7.5, 3.5])
ax.set_ylim([0, 10000])
ax.axvline(Ds[counter], color='k', linestyle='dashed', linewidth=3)
ax.set_title(sizes[counter]+ 'nm')
if counter == 2:
ax.set_xlabel(r'$log(D_{eff})$')
counter = counter + 1
```
| github_jupyter |
# Streaming Algorithms in Machine Learning
In this notebook, we will use an extremely simple "machine learning" task to learn about streaming algorithms. We will try to find the median of some numbers in batch mode, random order streams, and arbitrary order streams.
The idea is to observe first hand the advantages of the streaming model as well as to appreciate some of the complexities involved in using it.
The task at hand will be to approximate the median (model) of a long sequence of numbers (the data). This might seem to have little to do with machine learning. We are used to thinking of a median, $m$, of number $x_1,\ldots,x_n$ in the context of statistics as the number, $m$, which is smaller than at most half the values $x_i$ and larger than at most half the values $x_i$.
Finding the median, however, also solves a proper machine learning optimization problem (albeit a simple one). The median minimizes the following clustering-like objective function
$$m = \min_x \frac1n\sum_i|x - x_i|.$$
In fact, the median is the solution to the well studied k-median clustering problem in one dimension and $k=1$. Moreover, the extension to finding all quantiles is common in feature transformations and an important ingedient in speeding up decission tree training.
## Batch Algorithms
Let's first import a few libraries and create some random data.
Our data will simple by $100,000$ equally spaced points between $0$ and $1$.
```
import numpy as np
import matplotlib.pyplot as plt
n = 100000
data = np.linspace(0,1,n)
np.random.shuffle(data)
def f(x, data):
return sum(abs(x - datum) for datum in data)/len(data)
```
Let's look at the data to make sure everything is correct.
```
%matplotlib inline
# Plotting every 10th point
plt.scatter(range(0,n,100),data[0:n:100],vmin=0,vmax=1.0)
plt.ylim((0.0,1.0))
plt.xlim((0,n))
plt.show()
```
Computing the median brute force is trivial.
```
from math import floor
def batchMedian(data):
n = len(data)
median = sorted(data)[floor(n/2)]
return(median)
median = batchMedian(data)
print('The median found if {}'.format(median))
print('The objective value is {}'.format(f(median,data)))
```
The result is, of course, correct ($0.5$).
To get the median we sorted the data in $O(n\log n)$ time even though QuickSelect would have been faster ($O(n)$). The algorithm speed is not the main issue here though. The main drawback of this algorithm is that it must store the entire dataset in memory. For either sorting or quickSelect the algorithm must also duplicate the array. Binary search is also a possible solution which doesn't require data duplication but does require $O(\log(n))$ passes over the data.
When the data is large this is either very expensive or simply impossible.
## Streaming Algorithms (Random Order, SGD)
In the streaming model, we assume only an iterator over the data is given. That is, we can only make one pass over the data. Moreover, the algorithm is limited in its memory footprint and the limit is much lower than the data size. Otherwise, we could "cheat" by storing all the data in memory and executing the batch mode algorithm.
Gradient Descent (GD) type solutions are extremely common in this setting and are, de facto, the only mechanism for optimizing neural networks. In gradient descent, a step is taken in the direction opposite of the gradient. In one dimension, this simply means going left if the derivative is positive or right if the derivative is negative.
```
%matplotlib inline
xs = list(np.linspace(-1.0,2.0,50))
ys = [f(x,data) for x in xs]
plt.plot(xs,ys)
plt.ylim((0.0,2.0))
plt.xlim((-1.0,2.0))
ax = plt.axes()
ax.arrow(-0.5, 1.1, 0.3, -0.3, head_width=0.05, head_length=0.1, fc='k', ec='k')
ax.arrow(1.5, 1.1, -0.3, -0.3, head_width=0.05, head_length=0.1, fc='k', ec='k')
plt.show()
```
In **Stochastic Gradience Descent**, one only has a stochastic (random) unbiased estimator of the gradient. So, instead of computing the gradient of $\frac1n\sum_i|x - x_i|$ we can compute the gradient of $|x - x_i|$ where $x_i$ is chosen **uniformly at random** from the data. Note that a) the derivative of $|x - x_i|$ is simply $1$ if $x > x_i$ and $-1$ otherwise and b) the *expectation* of the derivative is exactly equal to the derivative of the overall objective function.
Comment: the authors of the paper below suggest essentially this algorithm but do not mention the connection to SGD for some reason.
Frugal Streaming for Estimating Quantiles: One (or two) memory suffices: Qiang Ma, S. Muthukrishnan, Mark Sandler
```
from math import sqrt
def sgdMedian(data, learningRate=0.1, initMedianValue=0):
median = initMedianValue
for (t,x) in enumerate(data):
gradient = 1.0 if x < median else -1.0
median = median - learningRate*gradient/sqrt(t+1)
return(median)
median = sgdMedian(data, learningRate=0.1, initMedianValue=0)
print('The median found if {}'.format(median))
print('The objective value is {}'.format(f(median,data)))
```
The result isn't exactly $0.5$ but it is pretty close. If this was a real machine learning problem, matching the objective up to the 5th digit of the true global minimum would have been very good.
Why does this work? Let's plot our objective function to investigate further.
It should not come as a big surprise to you that the objective function is convex. After all, it is the sum of convex functions (absolute values). It is a piece-wise linear curve that approximates a parabole in the range $(0,1)$ and is linear outside that range. Therefore, gradient descent is guaranteed to converge.
SGD significantly more efficient than sorting or even QuickSelect. More importantly, its memory footprint is tiny, a handful of doubles, *regardless of the size of the data*!!!
This is a huge advantage when operating with large datasets or with limited hardware.
Alas, SGD has some subtleties that make it a little tricky to use sometimes.
```
# SGD needs to be initialized carfully
median = sgdMedian(data, learningRate=0.1, initMedianValue=100.0)
print('The median found if {}'.format(median))
print('The objective value is {}'.format(f(median,data)))
# SGD needs to set step sizes corectly (controled via the learing rate)
median = sgdMedian(data, learningRate=0.001, initMedianValue=0.0)
print('The median found if {}'.format(median))
print('The objective value is {}'.format(f(median,data)))
```
These issues are usually alleviated by adaptive versions of SGD. Enhancements to SGD such as second order (based) methods, adaptive learning rate, and momentum methods may help in these situations but still require tuning in many cases. A common approach is to use many epochs.
```
median=0.0
numEpochs = 100
for i in range(numEpochs):
median = sgdMedian(data, learningRate=0.001, initMedianValue=median)
print('The median found if {}'.format(median))
print('The objective value is {}'.format(f(median,data)))
```
While clearly much less efficient than a single pass, increasing the number of epochs seemed to have solved the problem. Machine learning practitioners can relate to this result. That is, SGD is a great algorithm IF one finds good parameters for initialization, learning rate, number of epochs etc.
One of the main challenges in designing fundamentally better SGD-based streaming algorithms is in adaptively controlling these parameters during the run of the algorithm.
It is important to mention that there are also fundamentally better algorithms than SGD for this problem. See for example:
_Sudipto Guha, Andrew McGregor_ <br>
Stream Order and Order Statistics: Quantile Estimation in Random-Order Streams. <br>
_SIAM J. Comput. 38(5): 2044-2059 (2009)_
Unfortunately, we don't have time to dive into that...
### Trending data poses a challenge...
SGD and the other above algorithm have a fundamental drawback. They inherently rely on the fact that the data is random. For SGD, the gradient of the loss on a single point (or minibatch) must be an estimator of the global gradient. This is not true if trends in data make its statistics change (even slightly) over time. Let's simulate this with our data.
```
%matplotlib inline
# SGD also depends on the data being reandomly suffeled
n,k = len(data),10
minibatches = [data[i:i+k] for i in range(0,n,k)]
minibatches.sort(key=sum)
trendyData = np.array(minibatches).reshape(n)
# Plotting every 10th point in the trending dataset
plt.scatter(range(0,n,100),trendyData[0:n:100],vmin=0,vmax=1.0)
plt.ylim((0.0,1.0))
plt.xlim((0,n))
plt.show()
median = sgdMedian(trendyData, learningRate=0.1, initMedianValue=0.0)
print('The median found if {}'.format(median))
print('The objective value is {}'.format(f(median,data)))
```
## Streaming Algorithms (single pass, arbitrary order)
One way not to be fooled by trends in data and to sample from it.
The algorithm uses Reservoir Sampling to obtain $k$ (in this case $k=1000$) uniformly chosen samples from the stream. Then, compute the batch median of the sample.
The main drawback of sampling is that we now use more memory. Roughly the sample size $k$ ($k=1000$ here). This much more than $O(1)$ needed for SGD. Yet, it has some very appealing properties. Sampling very efficient ($O(1)$ per update), it is very simple to implement, it doesn't have any numeric sensitivities or tunable input parameters, and it is provably correct.
_(For the sake of simplicity below we use python's builtin sample function rather than recode reservoir sampling)_
```
from random import sample
def sampleMedian(data):
k=300
samples = sample(list(data),k)
return batchMedian(samples)
median = sampleMedian(trendyData)
print('The median found if {}'.format(median))
print('The objective value is {}'.format(f(median,data)))
```
As you can see, sampling provides relatively good results.
Nevertheless, there is something deeply dissatisfying about it. The algorithm was given $100,000$ points and used on $1,000$ of them. I other words, it would have been just as accurate had we collected only $1\%$ of the data.
Can we do better? Can an algorithm simultaneously take advantage of all the data, have a fixed memory footprint, and not be sensitive to the order in which the data is consumed? The answer is _yes!_. These are known in the academic literature as Sketching (or simply streaming) algorithms.
Specifically for approximating the median (or any other quantile), there is a very recent result that shows how best to achieve that:
_Zohar S. Karnin, Kevin J. Lang, Edo Liberty_ <br>
Optimal Quantile Approximation in Streams. <br>
FOCS 2016: 71-78
The following code is a hacky version of the algorithm described in the paper above. Warning: this function will not work for streams much longer than $100,000$!
```
from kll300 import KLL300
from bisect import bisect
def sketchMedian(data):
sketch = KLL300()
for x in data:
sketch.update(x)
assert sketch.size <= 300 # making sure there is no cheating involved...
items, cdf = sketch.cdf()
i = bisect(cdf, 0.5)
median = items[i]
return median
median = sketchMedian(trendyData)
print('The median found if {}'.format(median))
print('The objective value is {}'.format(f(median,data)))
```
Note that sketchMedian and sampleMedian both retain at most 300 items from the stream.
Still, the sketching solution is significantly more accurate.
Note that both sampling and sketching are randomized algorithms.
It could be that sampling happens to be more accurate than sketching for any single run. But, as a whole, you should expect the sketching algorithm to be much more accurate.
If you are curious about what sketchMedian actually does, you should look here:
* Academic paper - https://arxiv.org/abs/1603.05346
* JAVA code as part of datasketches - https://github.com/DataSketches/sketches-core/tree/master/src/main/java/com/yahoo/sketches/kll
* Scala code by Zohar Karnin - https://github.com/zkarnin/quantiles-scala-kll
* Python experiments by Nikita Ivkin - https://github.com/nikitaivkin/quantilesExperiments
The point is, getting accurate and stable streaming algorithms is complex. This is true even for very simple problems (like the one above). But, if one can do that, the benefits are well worth it.
| github_jupyter |
```
import numpy as np
import pandas as pd
import pickle
import time
import itertools
import matplotlib
matplotlib.rcParams.update({'font.size': 17.5})
import matplotlib.pyplot as plt
%matplotlib inline
matplotlib.rc('axes.formatter', useoffset=False)
import sys
import os.path
sys.path.append( os.path.abspath(os.path.join( os.path.dirname('..') , os.path.pardir )) )
#import FLAME with parameter for the number of important covariates
from colFLAMEbit_imp import *
from FLAMEbit_imp import *
def construct_sec_order(arr):
''' an intermediate data generation function used
for generating second order information '''
second_order_feature = []
num_cov_sec = len(arr[0])
for a in arr:
tmp = []
for i in range(num_cov_sec):
for j in range(i+1, num_cov_sec):
tmp.append( a[i] * a[j] )
second_order_feature.append(tmp)
return np.array(second_order_feature)
def data_generation_dense_2(num_control, num_treated, num_cov_dense,
num_covs_unimportant, control_m = 0.1,
treated_m = 0.9):
''' the data generating function that we will use;
includes second order information '''
# generate data for control group
xc = np.random.binomial(1, 0.5, size=(num_control, num_cov_dense)) #bernouilli
# generate data for treated group
xt = np.random.binomial(1, 0.5, size=(num_treated, num_cov_dense)) #bernouilli
errors1 = np.random.normal(0, 0.1, size=num_control) # some noise
errors2 = np.random.normal(0, 0.1, size=num_treated) # some noise
dense_bs_sign = np.random.choice([-1,1], num_cov_dense)
dense_bs = [ np.random.normal(s * 10, 1) for s in dense_bs_sign ] #alpha in the paper
# y for control group
yc = np.dot(xc, np.array(dense_bs)) #+ errors1
# y for treated group
treatment_eff_coef = np.random.normal( 1.5, 0.15, size=num_cov_dense) #beta
treatment_effect = np.dot(xt, treatment_eff_coef)
second = construct_sec_order(xt[:,:(num_covs_unimportant -1)])
treatment_eff_sec = np.sum(second, axis=1)
yt = np.dot(xt,np.array(dense_bs))+treatment_effect+treatment_eff_sec
# + errors2
# generate unimportant covariates for control group
xc2 = np.random.binomial(1, control_m, size=(num_control,
num_covs_unimportant))
# generate unimportant covariates for treated group
xt2 = np.random.binomial(1, treated_m, size=(num_treated,
num_covs_unimportant))
df1 = pd.DataFrame(np.hstack([xc, xc2]),
columns=range(num_cov_dense + num_covs_unimportant))
df1['outcome'] = yc
df1['treated'] = 0
df2 = pd.DataFrame(np.hstack([xt, xt2]),
columns=range(num_cov_dense + num_covs_unimportant))
df2['outcome'] = yt
df2['treated'] = 1
df = pd.concat([df1,df2])
df['matched'] = 0
return df, dense_bs, treatment_eff_coef
# data generation: 5 important covariates, 10 unimportant covariates
d = data_generation_dense_2(15000, 15000, 5,10, control_m = 0.1, treated_m = 0.9)
df = d[0]
holdout,_,_ = data_generation_dense_2(15000, 15000, 5,10, control_m = 0.1, treated_m = 0.9)
d[0].to_csv('411datatest.csv')
#run generic flame til no more matches
res_gen = run_bit(df, holdout, range(15), [2]*15, covs_unimportant = -10, threshold = -10, tradeoff_param = 0.001)
# run collapsing flame til no more matches
res = run_mpbit(df, holdout, range(15), [2]*15, covs_unimportant=-10, threshold =-10, tradeoff_param = 0.001)
#save data and result from generic and collapsing FLAME to compare to other methods
pickle.dump(res_gen, open('411genendrestest', 'wb'))
pickle.dump(res, open('411colendrestest', 'wb'))
pickle.dump(d, open('411datatest', 'wb'))
d[0].to_csv('411data.csv')
# define a ground truth
def ground_truth( eff_coef, covs_ordered, num_covs_dense = 5, num_second_order = 10, second_order = True):
arr = np.array(list(itertools.product([0,1], repeat=num_covs_dense)))
effect = np.dot(arr, eff_coef)
if second_order:
second_effect = np.sum(construct_sec_order(arr[:,:num_second_order] ), axis=1)
effect = effect + second_effect
df = pd.DataFrame(arr, columns=covs_ordered)
df['effect'] = effect
return df
# generate ground truth data
ground_truth = ground_truth(d[2], list(range(5)), num_covs_dense = 5, num_second_order = 10, second_order = True)
ground_truth.to_csv('411groundtruthtest.csv')
# get true and estimated cates for matched units
truth_list = []
pred_list = []
count = 0
av_err_cate = []
aux_size = []
for r in res[1]:
count = count +1
tmp = pd.merge(r, ground_truth, on = list(set(range(5)) & set(r.columns) ), how = 'left')
truth_list = truth_list + list(tmp['effect_y'])
pred_list = pred_list + list(tmp['effect_x'])
aux_size = aux_size + list(tmp['size'])
truth_list_gen = []
pred_list_gen = []
aux_size_gen = []
av_err_cate_gen = []
for r_gen in res_gen[1]:
tmp_gen = pd.merge(r_gen, ground_truth, on = list(set(range(5)) & set(r_gen.columns) ), how = 'left')
truth_list_gen = truth_list_gen + list(tmp_gen['effect_y'])
pred_list_gen = pred_list_gen + list(tmp_gen['effect_x'])
aux_size_gen = aux_size_gen + list(tmp_gen['size'])
# create a dataframe with the true and estimated cates : data to be used in R for nice plots and compare other methods
effect_col = pd.DataFrame()
effect_col['pred'] = pred_list
effect_col['true'] = truth_list
effect_col['size'] = aux_size
effect_col['method'] = ['collapsing FLAME']*len(truth_list)
#effect_col.to_csv('effect411endcol.csv')
effect_gen = pd.DataFrame()
effect_gen['pred'] = pred_list_gen
effect_gen['true'] = truth_list_gen
effect_gen['method'] = ['generic FLAME']*len(truth_list_gen)
effect_gen['size'] = aux_size_gen
effect = pd.concat([effect_gen, effect_col])
effect_gen.to_csv('effect411endgen.csv')
effect.to_csv('effect411end.csv')
#-----PLOT generic FLAME and collapsing FLAME in same figure----------#
import seaborn as sns
sns.lmplot(x="true", y="pred",hue="method", data = effect, fit_reg=False)
#----------- early stopping----------------#
#run generic flame before dropping important covariates
res_genearly = run_bit(df, holdout, range(15), [2]*15, covs_unimportant = 10, threshold = -10, tradeoff_param = 0.001)
# run collapsing flame efore dropping important covariates
researly = run_mpbit(df, holdout, range(15), [2]*15, covs_unimportant=10, threshold =-10, tradeoff_param = 0.001)
pickle.dump(res_genearly, open('411genendrestestearly', 'wb'))
pickle.dump(researly, open('411colendrestestearly', 'wb'))
# get true and estimated cates for matched units
truth_liste = []
pred_liste = []
count = 0
av_err_catee = []
aux_sizee = []
for r in researly[1]:
count = count +1
tmp = pd.merge(r, ground_truth, on = list(set(range(5)) & set(r.columns) ), how = 'left')
truth_liste = truth_liste + list(tmp['effect_y'])
pred_liste = pred_liste + list(tmp['effect_x'])
aux_sizee = aux_sizee + list(tmp['size'])
truth_list_gene = []
pred_list_gene = []
aux_size_gene = []
for r_gen in res_genearly[1]:
tmp_gen = pd.merge(r_gen, ground_truth, on = list(set(range(5)) & set(r_gen.columns) ), how = 'left')
truth_list_gene = truth_list_gene + list(tmp_gen['effect_y'])
pred_list_gene = pred_list_gene + list(tmp_gen['effect_x'])
aux_size_gene = aux_size_gene + list(tmp_gen['size'])
# create a dataframe with the true and estimated cates : data to be used in R for plots
effect_col = pd.DataFrame()
effect_col['pred'] = pred_liste
effect_col['true'] = truth_liste
effect_col['size'] = aux_sizee
effect_col['method'] = ['collapsing FLAME']*len(truth_liste)
effect_col.to_csv('effect411earlycoltest.csv')
effect_gen = pd.DataFrame()
effect_gen['pred'] = pred_list_gene
effect_gen['true'] = truth_list_gene
effect_gen['method'] = ['generic FLAME']*len(truth_list_gene)
effect_gen['size'] = aux_size_gene
effect = pd.concat([effect_gen, effect_col])
effect_gen.to_csv('effect411earlygentest.csv')
effect.to_csv('effect411earlytest.csv')
#-----PLOT generic FLAME and collapsing FLAME in same figure----------#
import seaborn as sns
sns.lmplot(x="true", y="pred",hue="method", data = effect, fit_reg=False)
sns.lmplot(x="true", y="pred",hue="method", data = effect_col, fit_reg=False)
sns.lmplot(x="true", y="pred",hue="method", data = effect_gen, fit_reg=False)
```
| github_jupyter |
# Validating performance of regression models
This notebook explains how to use CNTK metric functions to validate the performance of a regression model.
We're using the [car MPG dataset](https://archive.ics.uci.edu/ml/datasets/Auto+MPG) from the UCI dataset library. This dataset is perfect for demonstrating how to build a regression model using CNTK.
In the dataset, you'll find 9 columns:
1. mpg: continuous
2. cylinders: multi-valued discrete
3. displacement: continuous
4. horsepower: continuous
5. weight: continuous
6. acceleration: continuous
7. model year: multi-valued discrete
8. origin: multi-valued discrete
9. car name: string (unique for each instance)
All columns in the dataset contain numeric values except for the origin column which is a categorical value.
We'll strip the `car name` column as it cannot be used in our model.
## The model
The model we're using features two hidden layers. Each with 64 neurons with a ReLU (Rectified Linear Unit) activation function. The output is a single neuron without an activation function. This is necessary to turn this neural network into a regression model.
We're using the 8 input features and the miles per gallon as target for our neural network.
```
from cntk import default_options, input_variable
from cntk.layers import Dense, Sequential
from cntk.ops import relu
with default_options(activation=relu):
model = Sequential([
Dense(64),
Dense(64),
Dense(1,activation=None)
])
features = input_variable(9)
target = input_variable(1)
z = model(features)
```
## Preprocessing
In this section we'll first preprocess the data so that it is compatible for use with our neural network.
We need to load the data and then clean it up.
```
import pandas as pd
import numpy as np
df_cars = pd.read_csv('auto-mpg.csv', na_values=['?'])
df_cars = df_cars.dropna()
```
The origin column contains three possible values, as is shown in the dictionary below. To use the origin in the neural network we need to split it into three separate columns. For this we'll first replace the numeric values with a string value. After we've done that, we ask pandas to generate dummy columns. This creates three columns: usa, europa, and japan. For each sample in the dataset, one of these columns will contain a value of 1 and the rest will contain a value of 0.
```
origin_mapping = {
1: 'usa',
2: 'europe',
3: 'japan'
}
df_cars.replace({'origin': origin_mapping}, inplace=True)
categorical_origin = pd.get_dummies(df_cars['origin'], prefix='origin')
df_cars = pd.concat([df_cars, categorical_origin], axis=1)
df_cars = df_cars.drop(columns=['origin', 'car name'])
```
The final result of this operation is the following dataset. It contains 9 columns. Of these columns the `mpg` column is used as the target output. The rest is used as a feature for the model.
```
df_cars.head()
```
Let's create a training and validation set for training.
First we extract the car properties by dropping the mpg column. This produces a copy of the dataset without the mpg column.
If we were to use the data as-is, we would quickly run into exploding gradients. To prevent this from happening we need to scale the data so that it has a mean of zero and a variance of 1. The result of this is that we get values in a range of -1 to +1.
Finally we use the `train_test_split` function to produce a training and validation set.
```
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
X = df_cars.drop(columns=['mpg']).values.astype(np.float32)
y = df_cars.iloc[:,0].values.reshape(-1,1).astype(np.float32)
scaler = StandardScaler()
X = scaler.fit_transform(X)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
```
## Training the neural network
Now that we have a neural network, let's train it using the training set.
We're using a squared error loss function which is a regular loss that you will find in almost any regression model. We'll train the model using a SGD learner, which is the most basic learner around for CNTK.
The criterion for training is the mean squared error function. Additionally we would like to measure the mean absolute error rate for the model. In order to do this, we need to create a CNTK function factory that produces a combination of these two.
When you create a new function in python marked with `cntk.Function` annotation CNTK will automatically convert it to a function object that has a `train` method for training and a `test` method for validation.
Since we don't have a mean absolute error function out-of-the-box we'll create it here as well using the standard CNTK operators.
```
import cntk
from cntk.losses import squared_error
def absolute_error(output, target):
return cntk.ops.reduce_mean(cntk.ops.abs(output - target))
@cntk.Function
def criterion_factory(output, target):
loss = squared_error(output, target)
metric = absolute_error(output, target)
return loss, metric
from cntk.logging import ProgressPrinter
from cntk.losses import squared_error
from cntk.learners import sgd
loss = criterion_factory(z, target)
learner = sgd(z.parameters, 0.001)
progress_printer = ProgressPrinter(0)
train_summary = loss.train((X_train,y_train),
parameter_learners=[learner],
callbacks=[progress_printer],
minibatch_size=16,
max_epochs=10)
```
The output of the training session is looking promising, you can see that the loss is going down quite nicely. It's not perfect, but not bad for a first attempt.
## Evaluating model performance
In order to measure the performance of our model we're going first going to use the squared error function from CNTK.
This gives us an rough idea of the error rate of the model. But this is in squares so it is quite hard to read depending on your background.
As an alternative we'll also use the mean absolute error metric. This gives a more understandable error rate.
This metric gives us a good idea of just how much we're off predicting the miles per gallon.
CNTK doesn't include a mean absolute error function, but you can easily create it yourself using the standard CNTK ops.
We're using the test method on the metric to determine how well our model is doing. This is different from the classification model where we had to do quite a bit more to measure the performance of our model.
The output of the `test` method tells us how many miles per gallon the model is off on average when predicting based on the test set we created earlier.
```
loss.test((X_test, y_test))
```
| github_jupyter |
<table align="left" width="100%"> <tr>
<td style="background-color:#ffffff;"><a href="https://qsoftware.lu.lv/index.php/qworld/" target="_blank"><img src="..\images\qworld.jpg" width="35%" align="left"></a></td>
<td align="right" style="background-color:#ffffff;vertical-align:bottom;horizontal-align:right">
prepared by Özlem Salehi (<a href="http://qworld.lu.lv/index.php/qturkey/" target="_blank">QTurkey</a>)
</td>
</tr></table>
<table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
$ \newcommand{\bra}[1]{\langle #1|} $
$ \newcommand{\ket}[1]{|#1\rangle} $
$ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
$ \newcommand{\dot}[2]{ #1 \cdot #2} $
$ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
$ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
$ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
$ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
$ \newcommand{\mypar}[1]{\left( #1 \right)} $
$ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
$ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
$ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
$ \newcommand{\onehalf}{\frac{1}{2}} $
$ \newcommand{\donehalf}{\dfrac{1}{2}} $
$ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
$ \newcommand{\vzero}{\myvector{1\\0}} $
$ \newcommand{\vone}{\myvector{0\\1}} $
$ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
$ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
$ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
$ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
$ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
$ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
$ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
$ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
$ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
<h1> Quantum Fourier Transform </h1>
What makes Fourier Transform special is that it can be computed faster on a quantum computer than on a classical computer. In this notebook, we will learn about the quantum analogue of Discrete Fourier Transform.
Let's recall the definition for $DFT$. $DFT$ of the vector $x=\myvector{x_0~x_1\dots~x_{N-1}}^T$ is the complex vector $y=\myvector{y_0~y_1\dots y_{N-1}}^T$ where
$$
y_k=\frac{1}{\sqrt{N}} \sum_{j=0}^{N-1}e^{\frac{2\pi i j k }{N}}x_j.
$$
Now suppose that we have an $N=2^n$-dimensional quantum state vector $x=\myvector{x_0~x_1\dots~x_{N-1}}^T$ representing the state
$$
\ket{\psi} = \sum_{j=0}^{N-1} x_j \ket{j}.
$$
Quantum Fourier Transform $QFT$ of state $\ket{\psi}$ is given by
$$
\ket{\phi}=\sum_{k=0}^{N-1}y_k \ket{k} = \frac{1}{\sqrt{N}} \sum_{k=0}^{N-1} \sum_{j=0}^{N-1}e^{\frac{2\pi i j k }{N}}x_j \ket{k},
$$
where $y_k$ is defined as above for $k=0,\dots,N-1$.
Hence for a basis state $\ket{j}$ represented by the vector $\myvector{x_0~x_1\dots~x_{N-1}}^T$ where $x_j=1$ and the rest of the entries are 0, its $QFT$ is given by
$$
\frac{1}{\sqrt{N}} \sum_{k=0}^{N-1}e^{\frac{2\pi i j k }{N}} \ket{k}.
$$
$e^{\frac{2\pi i}{N}}$ is often denoted by $\omega$. The above expression can be equivalently expressed as
$$
\frac{1}{\sqrt{N}} \sum_{k=0}^{N-1}{\omega^{ j k} } \ket{k}.
$$
<h3>Task 1 (on paper)</h3>
Apply $QFT$ to the basis state $\ket{10}$ and find the new quantum state.
<a href="D02_Quantum_Fourier_Transform_Solutions.ipynb#task1">click for our solution</a>
<h3>Task 2 (on paper)</h3>
Apply $QFT$ to the state $ \ket \psi=\frac{1}{\sqrt{2}} \ket {01}+\frac{1}{\sqrt{2}} \ket {10} $.
<a href="D02_Quantum_Fourier_Transform_Solutions.ipynb#task2">click for our solution</a>
<h3>Task 3 (on paper)</h3>
Apply $QFT$ to the quantum state $\ket{\psi}=\alpha \ket{0} + \beta \ket{1}$ and find the new quantum state. Conclude that applying 1 qubit $QFT$ is equivalent to applying Hadamard gate.
<a href="D02_Quantum_Fourier_Transform_Solutions.ipynb#task3">click for our solution</a>
<b>Remark:</b> Note that two vectors that differ by a relative phase can not be distinguished when measured. For instance the vectors $\frac{1}{2} \myvector{1 \\ -i \\ -1 \\ i}$ and $\frac{1}{2} \myvector{1 \\ -1 \\ 1 \\ -1}$ are not distinguishable. But when $QFT$ is applied, then we obtain the vectors $ \myvector{0 \\ 0 \\ 0 \\ 1}$ and $ \myvector{ 0\\ 0 \\ 1 \\ 0}$ respectively and now the two vectors are distinguishable.
<hr>
<h2>QFT Operator</h2>
Let's try to find out the matrix corresponding to $QFT$. Given $x$, the entries of the vector $y$ corresponding to $QFT$ of the vector $x$ is given by
$$
y_k=\frac{1}{\sqrt{N}} \sum_{j=0}^{N-1}e^{\frac{2\pi i j k }{N}}x_j = \frac{1}{\sqrt{N}} \sum_{j=0}^{N-1}\omega^{j k}x_j .
$$
Hence, we can equivalently write:
\begin{align*}
y_0 &= \frac{1}{\sqrt{N}} \left (x_0 + x_1 + x_2 + \cdots + x_{N-1} \right ) \\
y_1 &= \frac{1}{\sqrt{N}} \left (x_0 + x_1\omega + x_2 \omega^2+ \cdots + x_{N-1} \omega^{N-1} \right )\\
y_2 &= \frac{1}{\sqrt{N}} \left (x_0 + x_1\omega^2 + x_2 \omega^4+ \cdots + x_{N-1} \omega^{2N-2} \right )\\
\vdots \\
y_{N-1} &= \frac{1}{\sqrt{N}} \left (x_0 + x_1\omega^{N-1} + x_2 \omega^{2N-2}+ \cdots + x_{N-1} \omega^{(N-1)(N-1)} \right ).
\end{align*}
From above, the matrix representation can be given by the following matrix.
$$ \frac{1}{\sqrt{N}} \mymatrix{rrrrrr}{1 & 1 & 1 &1 & \cdots & 1\\
1 & \omega & \omega^2 & \omega^3 & \cdots& \omega^{N-1} \\
1 & \omega^2 &\omega^4 & \omega^6 & \cdots& \omega^{2N-2} \\
1 & \omega^3 & \omega^6 & \omega^9 & \cdots& \omega^{3N-3} \\
\vdots & \vdots & \vdots & \vdots & \vdots & \vdots \\
1 & \omega^{N-1} & \omega^{2N-2} & \omega^{3N-3} & \cdots& \omega^{(N-1)(N-1)} \\
} $$
<h4> Observation</h4>
Let $\ket{j}$ be a basis state where $j=0,\dots N-1$. Then after applying $QFT$, the vector that is obtained is exactly the $j+1$'st column of the $QFT$ opreator.
<h3>Task 4 (on paper)</h3>
Apply $QFT$ to the basis state $\ket{10}$ using the matrix representation and find the new quantum state.
<a href="D02_Quantum_Fourier_Transform_Solutions.ipynb#task4">click for our solution</a>
<hr>
<h2>Properties of QFT (Optional) </h2>
<ul>
<li> $QFT$ is unitary: You can check this from the matrix corresponding to $QFT$. </li>
<li> Linear shift: Linear shift of a state vector causes relative phase shift of its $QFT$. The converse is also true. </li>
</ul>
If $QFT$ $\myvector{\alpha_0 \\ \alpha_1 \\ \vdots \\ \alpha_{N-1} } = \myvector{\beta_0 \\ \beta_1 \\ \vdots \\ \beta_{N-1}}$, then $QFT$ $\myvector{\alpha_{N-1} \\ \alpha_0 \\ \vdots \\ \alpha_{N-2} } = \myvector{\beta_0 \\ \omega \beta_1 \\ \vdots \\ \omega^{N-1}\beta_{N-1}}$ and
$QFT$ $\myvector{\alpha_0 \\ \omega \alpha_1 \\ \vdots \\ \omega^{N-1}\alpha_{N-1} } = \myvector{\beta_1 \\ \beta_2 \\ \vdots \\ \beta_{0}}$.
<h3>Task 5 (on paper)</h3>
What is the quantum state obtained after applying $QFT$ to the state $\ket{11}$? Find using Task 4 and the linear shift property.
<a href="D02_Quantum_Fourier_Transform_Solutions.ipynb#task5">click for our solution</a>
<hr>
<h2> Circuit implementation for $QFT$ </h2>
Recall that after applying $QFT$ to the state $\ket{\psi}=\alpha\ket{0} + \beta \ket{1}$, we get $\frac{\alpha+\beta}{\sqrt{2}} \ket{0} + \frac{\alpha-\beta}{\sqrt{2}} \ket{1}$, which is the state obtained after applying Hadamard to the state $\ket{\psi}$. Hence, the effect of applying $QFT$ to a single qubit is equivalent to applying Hadamard.
Now let us look at the effect of applying $QFT$ to a $n$-qubit system. Let $N=2^n$.
Let $\ket{j}$ be a basis state where $j=0,\dots,N-1$. We will use the binary representation for $j$, that is $j=j_1j_2\cdots j_n$ for $j_i \in \{0,1\}$. More formally
\begin{equation} \label{eq: binary}
j=j_12^{n-1} + j_2 2^{n-2} + \cdots + j_n2^0 = 2^n \sum_{l=1}^n 2^{-l} j_l~~~~~~~~(1)
\end{equation}
We will stick to the expression above but equivalently you may encounter the following expression for $j$ in various places.
\begin{align*}
j&=2^n(j_1 2^{-1} + j_2 2^{-2} + \cdots j_n 2^{-n}) \\
&= 2^n (0.j_1j_2\cdots j_n)
\end{align*}
Now let's use this representation to express the state obtained after applying $QFT$ to $\ket{j}$.
\begin{align*}
\ket{j}=\ket{j_1j_2\cdots j_n} &\rightarrow \frac{1}{\sqrt{N}} \sum_{k=0}^{N-1}e^{\frac{2\pi i j k }{N}} \ket{k}\\
&= \frac{1}{2^{n/2}} \sum_{k_1=0}^{1} \cdots \sum_{k_n=0}^{1}e^{2\pi i j (\sum_{o=1}^n k_o2^{-o})} \ket{k_1k_2\cdots k_n} \mbox{ by replacing $k$ using Equation (1)} \\
&=\frac{1}{2^{n / 2}} \sum_{k_{1}=0}^{1} \sum_{k_2=0}^{1}\cdots \sum_{k_{n}=0}^{1} \prod_{o=1}^{n} e^{2 \pi i j k_o 2^{-o}}|k_{1} \ldots k_{n}\rangle\\
&= \frac{1}{2^{n/2}}\otimes_{o=1}^n \sum_{k_o=0}^{1} e^{2\pi i jk_o 2^{-o} } \ket{k_o} \mbox{ by writing the quantum state as the tensor product of $n$ qubits} \\
&= \frac{1}{2^{n/2}} \otimes_{o=1}^n \biggl (\ket{0} + e^{2\pi i j 2^{-o} } \ket{1} \biggr) \\
&= \frac{1}{2^{n/2}} \biggl ( \bigl(\ket{0} + e^{2\pi i j 2^{-1}} \ket{1} \bigr) \otimes \bigl(\ket{0} + e^{2\pi i j 2^{-2}}\ket{1} \bigr) \otimes \cdots \otimes \bigl(\ket{0} + e^{2\pi i j 2^{-n}}\ket{1} \bigr ) \biggr )~~~~~~ (2)
\end{align*}
Let's find the correct gates for applying $QFT$. We are going to use two gates: Hadamard and controlled rotation gate. Check the following unitary operator:
$$ CR_k = \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 1 & 0 \\ 0 & 0 & 0 & e^{\frac{2\pi i}{2^{k}}} } $$
$CR_k$ operator puts a relative phase of $e^{\frac{2\pi i}{2^k}}$ in front of the quantum state if both the controlled and the target qubits are in state $\ket{1}$. It corresponds to a rotation around $z$-axis.
\begin{align*}
CR_k \ket{00} &\rightarrow \ket{00} \\
CR_k \ket{01} &\rightarrow \ket{01} \\
CR_k \ket{10} &\rightarrow \ket{10} \\
CR_k \ket{11} &\rightarrow e^{\frac{2\pi i}{{2^k}}}\ket{11} \\
\end{align*}
Let $\ket{j_1j_2\cdots j_n}$ be the input state. Applying Hadamard to the first qubit, we get
$$
\frac{1}{2^{1/2}} \bigl (\ket{0} + e^{2\pi i j_1 2^{-1}} \ket{1} \bigr ) \ket{j_2\cdots j_n}
$$
since $e^{\pi i j_1}=1$ if $j_1=0$ and $e^{\pi i j_1} = -1$ if $j_1=1$.
Now let's apply the opreator $CR_2$ where second qubit is the control and the first qubit is the target. The resulting state is
$$
\frac{1}{2^{1/2}} \bigl (\ket{0} + e^{2\pi i (j_1 2^{-1}+{j_2}{2^{-2}} )} \ket{1} \bigr ) \ket{j_2\cdots j_n}
$$
After applying $CR_i$ where qubit $i$ is the controller and the first qubit is the target consecutively for $i=3\dots n$, we obtain the state
$$
\frac{1}{2^{1/2}} \bigl (\ket{0} + e^{2\pi i (j_1 2^{-1}+{j_2}{2^{-2}} +{j_3}{2^{-3}} \dots {j_n}{2^{-n}} )} \ket{1} \bigr ) \ket{j_2\cdots j_n}
$$
Noting that $j=2^n(j_1 2^{-1} + j_2 2^{-2} + \cdots j_n 2^{-n})$, we get
$$
\frac{1}{2^{1/2}} \bigl (\ket{0} + e^{2\pi i j 2^{-n}} \ket{1} \bigr ) \ket{j_2\cdots j_n}
$$
<h3>Task 6 (on paper)</h3>
Continue with the same procedure this time using the second qubit as the target and obtain the following state:
$$
\frac{1}{2^{1/{4}}} \bigl (\ket{0} + e^{2\pi i j 2^{-n}} \ket{1} \bigr ) \bigl (\ket{0} + e^{2\pi i j 2^{-n+1} } \ket{1} \bigr ) \ket{j_3\cdots j_n}
$$
- First apply Hadamard to second qubit
- Then apply $CR_i$ where qubit $i+1$ is the control and the second qubit is the target for $i= 2 \dots n-1$.
<a href="D02_Quantum_Fourier_Transform_Solutions.ipynb#task6">click for our solution</a>
The above procedure is repeated for each qubit and finally we get the state
$$
\frac{1}{2^{n/2}} \bigl (\ket{0} + e^{2\pi i j 2^{-n}} \ket{1} \bigr ) \otimes (\ket{0} + e^{2\pi i j 2^{-n+1}} \ket{1} \bigr ) \otimes \cdots \otimes (\ket{0} + e^{2\pi i j 2^{-1}} \ket{1} \bigr )
$$
Note that this is exactly Equation (2) except that the qubits are in the reverse order. This also proves that $QFT$ is unitary, as the gates we used are unitary gates. Overall, the $QFT$ circuit is given below:
<img src="../images/qft.png" width="80%" align="left">
<hr>
Now we are ready to implement our circuit in Cirq. To implement $CR_k$, we will use the $CZPowGate$. Let's start by importing Cirq.
```
import cirq
```
<h3>CZPowGate</h3>
In Cirq, $CZPowGate$ is given by the following unitary matrix:
$$ CZPowGate(t) = \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 1 & 0 \\ 0 & 0 & 0 & e^{\pi i t} } $$
To use $CZPowGate$, you should use the following command to create a gate with parameter $t$:
<pre>cirq.CZPowGate(exponent=t)
In our case, by letting $t=2/2^k$ we can create the gate $CR_k$.
<pre>CRk = cirq.CZPowGate(exponent=2/2^k)<pre>
Then we can use $CR_k$ in our circuit by providing the control and target gates.
<pre>circuit.append(CRk(q0,q1))<pre>
<h3>Task 7</h3>
Implement the circuit for applying $QFT$ to a 2 qubit system. Try it using $\ket{01}$ as your input state. Simulate your circuit and compare it with the solution of Task 1.
```
import cirq
from cirq import CZPowGate, X, H, SWAP
#Create two qubits
q1, q2 = cirq.LineQubit.range(2)
#Create and initialize circuit
circuit = cirq.Circuit(X(q1))
#Define CR_2 gate with t=2/2^2
#Apply Hadamard to first qubit
#Apply CR_2 where first qubit is target and second qubit is control
#Apply Hadamard to second qubit
#Swap the qubits
#Simulate the circuit and print the results
print('Simulate the circuit:')
s=cirq.Simulator()
results=s.simulate(circuit)
print(results)
```
<a href="D02_Quantum_Fourier_Transform_Solutions.ipynb#task7">click for our solution</a>
<i>In the following tasks, to impose the order of application of the gates you may need to use the parameter <pre>circuit.append(...,strategy=InsertStrategy.NEW)</pre>
when appending gates to your circuit.</i>
<h3>Task 8 </h3>
Implement the circuit which applies $QFT$ to any $n$-qubit system. Define a function named <i>qft</i> which takes the number of qubits, the qubits and the circuit as its parameter. Simulate the circuit for $n=3$ on all possible input states.
```
from cirq.circuits import InsertStrategy
from cirq import H, SWAP
import cirq
def qft(n,qubits,circuit):
#Your code here
n=3
inputs = ['000','001','010','011','100','101','110','111']
for input in inputs:
#Create n qubits
#
#Create circuit
#
#initialization
#
#Call qft function
#
#Printing circuit
#
#Simulating circuit
#
```
<a href="D02_Quantum_Fourier_Transform_Solutions.ipynb#task8">click for our solution</a>
<h2>Inverse Quantum Fourier Transform </h2>
Inverse Quantum Fourier Transform ($QFT^{\dagger}$) is the transformation which satisfies $QFT\cdot QFT^{\dagger}=I$. It is defined exactly the same with the exponents having negative sign.
$$
QFT^{\dagger} \ket{k}=\frac{1}{\sqrt{N}} \sum_{l=0}^{N-1}e^{-\frac{2\pi i j k }{N}}\ket{l}.
$$
To implement $QFT^{\dagger}$, one should apply all the operations in reverse order to undo the circuit.
<h3>Task 9</h3>
Implement the circuit which applies $QFT^{\dagger}$ to any $n$-qubit system. Define a function named <i>iqft</i> which takes the number of qubits, the qubits and the circuit as its parameter. Simulate the circuit for different $n$ values. Apply both $QFT$ and $QFT^{\dagger}$ and check whether the corresponding unitary matrix is the identity matrix. Write your function to a file so that you can use it later.
```
#%%writefile iqft.py
import cirq
from cirq.circuits import InsertStrategy
from cirq import H, SWAP, CZPowGate
def iqft(n,qubits,circuit):
#Your code here
n=4 #change n
#Create circuit
circuit = cirq.Circuit()
#Create n qubits
#
#Call qft function
#Call iqft function
#
#Printing circuit
#
#Simulating circuit
#
#Unitary matrix representation
print(cirq.unitary(circuit))
```
<a href="D02_Quantum_Fourier_Transform_Solutions.ipynb#task9">click for our solution</a>
<hr>
<h2> Discussion on the number of gates </h2>
For the first qubit, we apply a single Hadamard gate followed by $n-1$ controlled rotation gates, which makes $n$ gates in total.
For the second qubit we apply a single Hadamard gate followed by $n-2$ controlled rotation gates, which makes $n-1$ gates in total.
Overall, $n + (n-1) + \cdots + 1 = \frac{(n)(n+1)}{2}$ gates are required.
Furthermore, $\frac{n}{2}$ $SWAP$ gates are required, each of which can be implemented using three $CNOT$ gates.
To sum up, $\theta(n^2)$ gates are required for applying $QFT$ to an $n$-qubit system represented by a vector of size $N=2^n$.
The best known classical algorithm for computing the Discrete Fourier Transform of $2^n$ entries, such as Fast Fourier Transform (FFT) requires $\Theta(n2^n)$, equivalently $\Theta(N \log N)$ gates, which means that the classical algorithm requires exponentially many more operations to compute $DFT$ (Note that the obvious algorithm for computing $DFT$ requires ${2}^{2n}$ gates). An improved algorithm presented [here](https://ieeexplore.ieee.org/document/892139) provides a circuit for computing $QFT$ with $O(n \log n)$ gates.
Nevertheless, this does not mean that we can use $QFT$ directly to accelerate the classical computation process. There are two reasons for this: The first reason is that the amplitudes can not be accessed directly after applying $QFT$. The second reason is that we may not know how to efficiently prepare the input state to $QFT$.
Next, we will see some applications of $QFT$.
| github_jupyter |
```
import torch
from torch.autograd import Variable
import torch.nn.functional as F
import matplotlib.pyplot as plt
%matplotlib inline
torch.manual_seed(1) # reproducible
# make some fake data and display them
n_data = torch.ones(100, 2)
x0 = torch.normal(2*n_data, 1) # class0 x data (tensor), shape=(100, 2), with mean=2, variance=1
y0 = torch.zeros(100) # class0 y data (tensor), shape=(100, 1), label = 0
x1 = torch.normal(-2*n_data, 1) # class1 x data (tensor), shape=(100, 2), with mean=-2, variance=1
y1 = torch.ones(100) # class1 y data (tensor), shape=(100, 1), label = 1
x = torch.cat((x0, x1), 0).type(torch.FloatTensor) # shape (200, 2) FloatTensor = 32-bit floating
y = torch.cat((y0, y1), ).type(torch.LongTensor) # shape (200,) LongTensor = 64-bit integer
# torch can only train on Variable, so convert them to Variable
x, y = Variable(x), Variable(y)
plt.scatter(x.data.numpy()[:, 0], x.data.numpy()[:, 1], c=y.data.numpy(), s=100, lw=0, cmap='RdYlGn')
plt.show()
class Net(torch.nn.Module):
def __init__(self, n_feature, n_hidden, n_output):
super(Net, self).__init__()
self.hidden = torch.nn.Linear(n_feature, n_hidden) # hidden layer
self.out = torch.nn.Linear(n_hidden, n_output) # output layer
def forward(self, x):
x = F.relu(self.hidden(x)) # activation function for hidden layer
x = self.out(x)
return x
net = Net(n_feature=2, n_hidden=10, n_output=2) # instantiate the network
print(net) # print the net architecture
optimizer = torch.optim.SGD(net.parameters(), lr=0.2) # initialize the optimizer, SGD ==> 梯度下降
loss_func = torch.nn.CrossEntropyLoss() # this is for regression mean squared loss, 交叉熵损失函数
plt.ion() # something about plotting
for t in range(100):
out = net(x) # input x and predict based on x
loss = loss_func(out, y) # must be (1. nn output, 2. target)
optimizer.zero_grad() # must clear gradients for next train
loss.backward() # backpropagation, compute gradients
optimizer.step() # apply gradients to renew your params
if t % 10 == 0 or t in [3, 6]:
# plot and show learning process
plt.cla()
_, prediction = torch.max(F.softmax(out), 1)
pred_y = prediction.data.numpy().squeeze()
target_y = y.data.numpy()
plt.scatter(x.data.numpy()[:, 0], x.data.numpy()[:, 1], c=pred_y, s=100, lw=0, cmap='RdYlGn')
accuracy = sum(pred_y == target_y)/200.
plt.text(1.5, -4, 'Accuracy=%.2f' % accuracy, fontdict={'size': 20, 'color': 'red'})
plt.show()
plt.pause(0.1)
plt.ioff()
```
| github_jupyter |
```
##################################################################
#《Python机器学习及实践:从零开始通往Kaggle竞赛之路(2023年度版)》开源代码
#-----------------------------------------------------------------
# @章节号:6.6.2(注意力机制的TensorFlow实践)
# @作者:范淼
# @电子邮箱:fanmiao.cslt.thu@gmail.com
# @微博:https://weibo.com/fanmiaothu
# @官方交流QQ群号:561500762
##################################################################
from tensorflow.keras import models, layers, losses, optimizers, Model
#设置超参数。
INPUT_UNITS = 56
TIME_STEPS = 14
NUM_HEADS = 4
NUM_CLASSES = 10
EPOCHS = 5
BATCH_SIZE = 64
LEARNING_RATE = 1e-3
class Attention(Model):
'''
自定义注意力机制类,继承自keras.Model。
'''
def __init__(self, input_units, num_heads, num_classes):
super(Attention, self).__init__()
self.multihead_attn = layers.MultiHeadAttention(key_dim=input_units, num_heads=num_heads)
self.linear = layers.Dense(num_classes)
def call(self, input_tensor):
out = self.multihead_attn(input_tensor, input_tensor, input_tensor)
out = self.linear(out[:, -1, :])
return out
#初始化注意力机制的神经网络模型。
model = Attention(INPUT_UNITS, NUM_HEADS, NUM_CLASSES)
#设定神经网络的损失函数、优化方式,以及评估方法。
model.compile(optimizer=optimizers.Adam(LEARNING_RATE),
loss=losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
import pandas as pd
#使用pandas,读取fashion_mnist的训练和测试数据文件。
train_data = pd.read_csv('../datasets/fashion_mnist/fashion_mnist_train.csv')
test_data = pd.read_csv('../datasets/fashion_mnist/fashion_mnist_test.csv')
#从训练数据中,拆解出训练特征和类别标签。
X_train = train_data[train_data.columns[1:]]
y_train = train_data['label']
#从测试数据中,拆解出测试特征和类别标签。
X_test = test_data[train_data.columns[1:]]
y_test = test_data['label']
from sklearn.preprocessing import StandardScaler
#初始化数据标准化处理器。
ss = StandardScaler()
#标准化训练数据特征。
X_train = ss.fit_transform(X_train)
#标准化测试数据特征。
X_test = ss.transform(X_test)
X_train = X_train.reshape([-1, TIME_STEPS, INPUT_UNITS])
#使用fashion_mnist的训练集数据训练网络模型。
model.fit(X_train, y_train.values, batch_size=BATCH_SIZE, epochs=EPOCHS, verbose=1)
X_test = X_test.reshape([-1, TIME_STEPS, INPUT_UNITS])
#使用fashion_mnist的测试集数据评估网络模型的效果。
result = model.evaluate(X_test, y_test.values, verbose=0)
print('注意力机制(TensorFlow版本)在fashion_mnist测试集上的准确率为: %.2f%%。' %(result[1] * 100))
```
| github_jupyter |
<img src="images/strathsdr_banner.png" align="left">
# RFSoC QPSK Transceiver
----
<div class="alert alert-box alert-info">
Please use Jupyter Labs http://board_ip_address/lab for this notebook.
</div>
This design is a full QPSK transceiver, which transmits and receives randomly-generated pulse-shaped symbols with full carrier and timing synchronisation. PYNQ is used to visualise the data at both the DAC and ADC side of the RFSoC data converters, as well as visualising various DSP stages throughout the transmit and receive signal path.
## Contents
* [Introduction](#introduction)
* [Hardware Setup](#hardware-setup)
* [Software Setup](#software-setup)
* [RFSoC QPSK Transceiver](#RFSoC-QPSK-Transceiver)
* [Inspecting the transmit path](#Inspecting-the-transmit-path)
* [Inspecting the receive path](#Inspecting-the-receive-path)
* [Reconfigure the RF Data Path](#Reconfigure-the-RF-Data-Path)
* [Conclusion](#conclusion)
## References
* [Xilinx, Inc, "USP RF Data Converter: LogiCORE IP Product Guide", PG269, v2.3, June 2020](https://www.xilinx.com/support/documentation/ip_documentation/usp_rf_data_converter/v2_3/pg269-rf-data-converter.pdf)
## Revision History
* **v1.0** | 02/11/2020 | RFSoC QPSK demonstrator
* **v1.1** | 23/02/2021 | Reformatted notebook
----
## Introduction <a class="anchor" id="introduction"></a>
The ZCU111 platform and XM500 development board can be configured as a simple QPSK transceiver. The RFSoC QPSK demonstrator uses the RFSoC's RF Data Converters (RF DCs) to transmit and receive QPSK modulated waveforms. There are setup steps for hardware and software that you must follow.
### Hardware Setup <a class="anchor" id="hardware-setup"></a>
Your ZCU111 development board can be configured to host one QPSK transceiver channel. To setup your board for this demonstration, you can connect a channel in loopback as shown in [Figure 1](#fig-1).
The default loopback configuration is connected as follows:
* Channel 0: DAC6 (Tile 229 Block 2) to ADC0 (Tile 224 Block 0)
There are several XM500 board revisions, and some contain different silkscreen and labels for the ADCs and DACs. Use the image below for further guidance and pay attention to the associated Tile and Block.
<a class="anchor" id="fig-1"></a>
<figure>
<img src='images/zcu111_setup.png' height='50%' width='50%'/>
<figcaption><b>Figure 1: ZCU111 and XM500 development board setup in loopback mode.</b></figcaption>
</figure>
**Do not** attach an antenna to any SMA interfaces labelled DAC.
<div class="alert alert-box alert-danger">
<b>Caution:</b>
In this demonstration, we generate tones using the RFSoC development board. Your device should be setup in loopback mode. You should understand that the RFSoC platform can also transmit RF signals wirelessly. Remember that unlicensed wireless transmission of RF signals may be illegal in your geographical location. Radio signals may also interfere with nearby devices, such as pacemakers and emergency radio equipment. Note that it is also illegal to intercept and decode particular RF signals. If you are unsure, please seek professional support.
</div>
### Software Setup <a class="anchor" id="software-setup"></a>
Start by including the `xrfdc` drivers so we can configure the RF data converters, `ipywidgets` to make interactive controls, `numpy` for numerical analysis, and `rfsoc_qpsk` for the QPSK design.
```
import xrfdc
import ipywidgets as ipw
import numpy as np
from rfsoc_qpsk.qpsk_overlay import QpskOverlay
```
We can now initialise the overlay by downloading the bitstream and executing the drivers.
```
ol = QpskOverlay()
```
For a quick reference of all the things you can do with the QPSK overlay, ask the Python interpreter!
Pop open a new console (right click here and select "_New Console for Notebook_") and type `ol.plot_group?` to query a method of our new overlay. Tab completion works for discovery too.
----
## RFSoC QPSK Transceiver <a class="anchor" id="RFSoC-QPSK-Transceiver"></a>
We will now explore three interesting components of the QPSK demonstrator. Initially, the transmit path will be inspected and then the same inspection will also be carried out on the receive path. Finally, we will explore the control capabilities of our design and determine how these properties affect the transmit and receive signals.
### Inspecting the transmit path <a class="anchor" id="Inspecting-the-transmit-path"></a>
There are 3 main steps in the QPSK transmit IP signal path:
1. Random symbol generation
2. Pulse shaping
3. Interpolation
This design "taps off" this path after the first two stages so we can inspect the signals in Jupyter Lab.
The RF data converter can be reconfigured from Python too - we'll look at that [later](#Reconfigure-the-RF-Data-Path).

First we plot our raw QPSK symbols in the time domain.
```
ol.plot_group(
'tx_symbol', # Plot group's ID
['time-binary'], # List of plot types chosen from:
# ['time','time-binary','frequency','constellation']
ol.qpsk_tx.get_symbols, # Function to grab a buffer of samples
500 # Sample frequency (Hz)
)
```
We can stream new samples into this plot using the play/stop buttons. By default the samples are stored in a rolling buffer, so we can keep this running for a while without worrying too much about total memory usage. As you continue to work through this notebook though, you should stop any previous plot streams to keep your browser happy.
For the pulse shaped signal, let's have a look at the frequency domain too. This FFT is accelerated in the PL so we pass in an extra argument, `get_freq_data`, telling the plotting library how to grab the accelerated FFT data.
```
ol.plot_group('tx_shaped', ['time', 'frequency'], ol.qpsk_tx.get_shaped_time, 4000,
get_freq_data=ol.qpsk_tx.get_shaped_fft)
```
### Inspecting the receive path <a class="anchor" id="Inspecting-the-receive-path"></a>
The receive side is nearly the inverse of the transmit path (there's just some extra work for properly synchronising).
Again, there are taps off from a few places in the signal path:
1. After decimation
2. After coarse synchronisation
3. After root-raised-cosine filtering
4. and the data output

Because there are a few different intermediate stages, let's reuse the same cells to plot any of them on-demand.
First we describe how to generate plots for each of the intermediate taps.
```
rx_domains = ['time', 'frequency', 'constellation']
plot_rx_decimated = lambda : ol.plot_group(
'rx_decimated', rx_domains, ol.qpsk_rx.get_decimated, 4000
)
plot_rx_coarse_sync = lambda : ol.plot_group(
'rx_coarse_sync', rx_domains, ol.qpsk_rx.get_coarse_synced, 4000
)
plot_rx_rrced = lambda : ol.plot_group(
'rx_rrced', rx_domains, ol.qpsk_rx.get_rrced, 16000
)
```
Now we can just execute the function whichever tap you want. For example, let's look at the tap after decimation below.
```
plot_rx_decimated()
```
And for the final plot, let's look at the synchronised output data. To recover the bits we need to take our sampled, synchronised signal (seen in the constellation plot below) and decide which quadrant each symbol has fallen into.
```
def classify_bits(frame):
bit_quantise = lambda b: 1 if b>0 else 0
symbol_quantise = lambda i, q: bit_quantise(i) + 1j*bit_quantise(q)
return np.fromiter(
map(symbol_quantise, np.real(frame), np.imag(frame)),
dtype=np.complex
)
ol.plot_group(
'rx_data',
['constellation', 'time-binary'],
lambda : classify_bits(ol.qpsk_rx.get_data()),
500,
get_const_data=ol.qpsk_rx.get_data
)
```
Now is a good time to note that Jupyter Lab can manage multiple windows. Next we'll be playing with the RF settings, so you may want to make a new window for the constellation plot and leave it streaming. Make a new window for the plot by right clicking the plot and selecting "_Create New View for Output_". Feel free to snap this new window to the side by clicking the window's title ("Output View") and dragging it to the side of the web page. Now we can play with RF settings further down the notebook while still getting instant feedback about our received signal — pretty neat!
### Reconfigure the RF Data Path <a class="anchor" id="Reconfigure-the-RF-Data-Path"></a>
#### Transmit Power
The QPSK bitstream includes a digital attenuator on the transmit path. We can configure this via a memory-mapped register.
Let's use this as an example of interactive reconfiguration because the effects are quite clear in the constellation diagram. Try reducing the output power by setting a gain between 0 (off) and 1 (full scale).
```
ol.qpsk_tx.set_gain(0.6)
```
The constellation plot should shrink in a little towards the origin. Let's return to full power now.
```
ol.qpsk_tx.set_gain(1)
```
We can use some `ipywidgets` to make a more natural interface to control the gain too. Let's expose this as a slider with a callback to the `set_gain` function.
```
pow_slider = ipw.SelectionSlider(
options=[0.1, 0.3, 0.6, 1],
value=1,
description='',
)
accordion = ipw.Accordion(children=[pow_slider])
accordion.set_title(0, 'Transmitter power')
display(accordion)
def unwrap_slider_val(callback):
return lambda slider_val : callback(slider_val['new'])
pow_slider.observe(unwrap_slider_val(ol.qpsk_tx.set_gain), names='value')
```
#### Transmit and Receive Mixer Settings
So far the RF Data Converter settings have been controlled by `QpskOverlay` but we can also reconfigure these on the fly in python with the `xrfdc` driver.
First of all, consider the DAC block used for the transmit side.

There's a lot of scope for reconfiguration here — see the [IP product guide](https://www.xilinx.com/support/documentation/ip_documentation/usp_rf_data_converter/v2_1/pg269-rf-data-converter.pdf) or type `ol.dac_block?` for more details.
As an example, let's play with the mixer settings. Try changing the DAC's mixer frequency from the deafult 1000 MHz to 900 MHz.
```
def update_nco(rf_block, nco_freq):
mixer_cfg = rf_block.MixerSettings
mixer_cfg['Freq'] = nco_freq
rf_block.MixerSettings = mixer_cfg
rf_block.UpdateEvent(xrfdc.EVENT_MIXER)
update_nco(ol.dac_block, 900)
```
The received signal should disappear until we configure the receiver's ADC to match the new carrier frequency. Set the new carrier frequency for the ADC side mixer below.
```
update_nco(ol.adc_block, 900)
```
Again, we can use `ipywidgets` to make an interactive interface for these settings. Below we setup an RX and a TX slider and a TX slider that are linked together so we can scrub along the spectrum keeping both sides in near lock-step. If you've got any analog RF filters to hand, try them out with different mixer settings!
```
def new_nco_slider(title):
return ipw.FloatSlider(
value=1000,
min=620,
max=1220,
step=20,
description=title,
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.1f',
)
tx_nco_slider = new_nco_slider('TX (MHz)')
rx_nco_slider = new_nco_slider('RX (MHz)')
accordion = ipw.Accordion(children=[ipw.VBox([tx_nco_slider, rx_nco_slider])])
accordion.set_title(0, 'Carrier frequency')
display(accordion)
ipw.link((rx_nco_slider, 'value'), (tx_nco_slider, 'value'))
tx_nco_slider.observe(
unwrap_slider_val(lambda v: update_nco(ol.dac_block, v)),
names='value'
)
rx_nco_slider.observe(
unwrap_slider_val(lambda v: update_nco(ol.adc_block, v)),
names='value'
)
```
## Conclusion <a class="anchor" id="conclusion"></a>
We've now lead you through how we can interact with the RF data converters from PYNQ, using a QPSK transmit/receive loopback system as an example. More exhaustively, we've shown:
* Use of the programmable logic in the context of a real RF application
* Performing on-board introspection of an RF design:
* Leveraging existing plotting libraries from the Python ecosystem
* Interacting with a QPSK hardware design
* Configuring the signal path, using transmit power as an example
* Configuring the RF data converter, using TX/RX mixer frequencies as an example
| github_jupyter |
The notebook measures how well learned reward functions generalize to new environments.
It trains a reward function on a series of environments with different colors for the agent and background. It measures how well the reward function can generalize to colors it hasn't seen before.
```
import gym
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import random
import time
start = time.process_time()
# your code here
class TransferTestEnv(gym.Env):
"""Makes an environment with one color for the agent and another color for the background.
Generates the agent in a random position each reset.
Returns the reward based on how close the agent is to the top right.
"""
def __init__(self, bg_color, agent_color):
self.bg_color = bg_color
self.agent_color = agent_color
self.color_num = 1
self.env_size = 5
pass
def reset(self):
self.location = np.random.randint(0, self.env_size, [2])
color_extra_dim = np.reshape(self.bg_color, [1,1,self.color_num])
state = np.tile(color_extra_dim, [self.env_size, self.env_size, self.color_num])
state[self.location[0], self.location[1]] = self.agent_color
reward = self.location[0] + self.location[1]
return state, reward
state, reward = TransferTestEnv(5, 1).reset()
plt.imshow(np.squeeze(state))
def generate_data(dataset_size, env_params_list):
"""Generates data by randomly sampling from environments with the specified parameters."""
xs = []
ys = []
for _ in range(dataset_size):
env_params = random.choice(env_params_list)
state, reward = TransferTestEnv(*env_params).reset()
xs.append(state)
ys.append(reward)
xs = np.array(xs)
ys = np.array(ys)
return xs, ys
#------small problem (can be ignored): a not balanced amount of environemnts generated every type of setup
xs, ys = generate_data(10, [[1.111111, -.1111111],[1.111, -.111], [1,0]])
print(xs.shape)
print(ys.shape)
[m[1,1] for m in xs]
plt.imshow(np.squeeze(xs[0]))
# Creates parameters with different background and foreground colors. Make sure that the colors are different enough
max_num_envs = 40
while True:
half_max_num_envs = int(max_num_envs/2)
train_env_params = np.random.random(size=(half_max_num_envs, 2))
train_env_params = np.concatenate((np.array([[pair[0]+1.2, pair[1]-1.2] for pair in train_env_params]), np.random.random(size=(half_max_num_envs, 2))))
diff = np.abs(train_env_params[:, 0] - train_env_params[:, 1])
if np.min(diff) > .2:
break
print(train_env_params.shape)
print(train_env_params)
np.concatenate((np.array([1,23]),np.array([1,23])))
# np.array([1,23])
def get_generalization_amount(train_env_params):
xs, ys = generate_data(1000, train_env_params)
print("train_env_params", train_env_params)
#Model architecture
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(16, (3, 3), activation='relu', input_shape=xs.shape[1:]),
tf.keras.layers.Conv2D(16, (3, 3), activation='relu'),
tf.keras.layers.Dropout(0.3),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(64),
])
model.compile(optimizer='adam', loss='mse')
# Train the model on the data.
model.fit(xs, ys, epochs=400, verbose=0)
xs_val, ys_val = generate_data(1000, train_env_params)
xs_val_differnt, ys_val_different = generate_data(1000, [(1.1, -.1)])
model.evaluate(xs, ys)
# This is the loss from different data on the same environments.
val_loss = model.evaluate(xs_val, ys_val)
# This is the loss from different data on different environments.
different_loss = model.evaluate(xs_val_differnt, ys_val_different)
return val_loss, different_loss
# Get the val and generalization losses using different numbers of environments.
gaps_for_nums = []
num_envs_ar = list(range(1, max_num_envs))
val_loss_ar = []
different_loss_ar = []
for num_envs in num_envs_ar:
val_loss, different_loss = get_generalization_amount(train_env_params[:num_envs])
val_loss_ar.append(val_loss)
different_loss_ar.append(different_loss)
print([num_envs, val_loss, different_loss])
val_loss_ar = np.array(val_loss_ar)
different_loss_ar = np.array(different_loss_ar)
gaps = val_loss_ar / different_loss_ar
# Plot the ratio of loss on the data from the same environments vs different environments. Higher is better.
# X axis is the number of training environments.
plt.plot(num_envs_ar, gaps)
plt.plot(num_envs_ar, val_loss_ar)
plt.plot(num_envs_ar, different_loss_ar)
print("time spent (min):", (time.process_time() - start)/60)
```
| github_jupyter |
```
"""
@author: Ajay
"""
import torch
import torch.nn.functional as F
import torch.nn as nn
from torch.autograd import Variable
from torch.utils.data import Dataset, DataLoader
import torch.optim as optim
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
from sklearn.preprocessing import StandardScaler, MinMaxScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, precision_score, recall_score
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.linear_model import LinearRegression
from sklearn.ensemble import RandomForestRegressor
from xgboost import XGBRegressor
from tqdm.notebook import tqdm
from sklearn.datasets import load_iris
from sklearn.metrics import roc_curve, auc
from sklearn.metrics import confusion_matrix
from pyNM.cf_matrix import make_confusion_matrix
from plot_metric.functions import MultiClassClassification
import seaborn as sns
from random import *
get_ipython().run_line_magic('matplotlib', 'inline')
from pyNM.nonspiking_regressor import *
def run_regressor():
torch.multiprocessing.freeze_support()
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print(device)
# read house price dataset
df_house = pd.read_csv('../data/house_price_data.csv')
print(df_house.shape)
print(df_house.columns)
# peek into the data
print(df_house.head())
# final dataset
X = df_house.iloc[:,1:4] # independent attributes
y = df_house.iloc[:,0] # house price - target
#scaler = StandardScaler()
#scaler = MinMaxScaler()
#X_scaled = scaler.fit_transform(X)
# Create Train — Validation — Test split
# Train - Test
X_trainval, X_test, y_trainval, y_test = train_test_split(X, y, test_size=0.3, random_state=1)
# Split train into train-val
X_train, X_val, y_train, y_val = train_test_split(X_trainval, y_trainval, test_size=0.2, random_state=2)
print(X_train.shape)
print(y_train.shape)
print(X_val.shape)
print(y_val.shape)
print(X_test.shape)
print(y_test.shape)
# Scale data to have mean '0' and variance '1'
# which is importance for convergence of the neural network
scaler = MinMaxScaler()
X_train = scaler.fit_transform(X_train)
X_val = scaler.transform(X_val)
X_test = scaler.transform(X_test)
X_train, y_train = np.array(X_train), np.array(y_train)
X_val, y_val = np.array(X_val), np.array(y_val)
X_test, y_test = np.array(X_test), np.array(y_test)
# Create & Initialize dataset
class CreateRegressionDataset(Dataset):
def __init__(self, X_data, y_data):
self.X_data = X_data
self.y_data = y_data
def __getitem__(self, index):
return self.X_data[index], self.y_data[index]
def __len__(self):
return len(self.X_data)
# create train, val, test dataset
train_dataset = CreateRegressionDataset(torch.from_numpy(X_train).float(),
torch.from_numpy(y_train).float())
val_dataset = CreateRegressionDataset(torch.from_numpy(X_val).float(),
torch.from_numpy(y_val).float())
test_dataset = CreateRegressionDataset(torch.from_numpy(X_test).float(),
torch.from_numpy(y_test).float())
# define model parameters
EPOCHS = 150
BATCH_SIZE = 64
LEARNING_RATE = 0.001
NUM_OF_FEATURES = len(X.columns)
# load the training, validation, and test sets
print("Loading training, validation, and test data")
train_loader = DataLoader(dataset=train_dataset,
batch_size=BATCH_SIZE, shuffle=True)
val_loader = DataLoader(dataset=val_dataset,
batch_size=1)
test_loader = DataLoader(dataset=test_dataset,
batch_size=1)
print("Data loaded...")
# create spiking neural network architecture
'''
We will create a feed forward SNN with 3 hidden layers
'''
non_spiking_model_regressor = NonSpikingNeuralNetwork(input_dim=X_train.shape[1], hidden_dim_l1=16,
hidden_dim_l2=32, hidden_dim_l3=16)
print("Model created")
print(non_spiking_model_regressor)
# Initialize the model, optimizer, and loss function
'''
Transfer the model to GPU/CPU.
Here, we use Mean Squared Error (MSE) as loss function
'''
loss_func = nn.MSELoss()
optimizer = optim.SGD(non_spiking_model_regressor.parameters(), lr = LEARNING_RATE)
print("Optimizer done")
# dictionary to store loss each epoch for both train & val sets
loss_stats = {'train': [],
'val': []
}
# Train Model
print("Begin Training...")
for e in tqdm(range(1, EPOCHS+1)):
# TRAINING
train_epoch_loss = 0
non_spiking_model_regressor.train()
for X_train_batch, y_train_batch in train_loader:
X_train_batch, y_train_batch = X_train_batch.to(device), y_train_batch.to(device)
#feed forward
y_train_pred = non_spiking_model_regressor(X_train_batch.requires_grad_())
#calculate the loss
train_loss = loss_func(y_train_pred, y_train_batch.unsqueeze(1))
#backward propagation: calculate gradients
train_loss.backward()
#update the weights
optimizer.step()
#clear out the gradients from the last step loss.backward()
optimizer.zero_grad()
train_epoch_loss += train_loss.item()
# VALIDATION
with torch.no_grad():
val_epoch_loss = 0
non_spiking_model_regressor.eval()
for X_val_batch, y_val_batch in val_loader:
X_val_batch, y_val_batch = X_val_batch.to(device), y_val_batch.to(device)
y_val_pred = non_spiking_model_regressor(X_val_batch)
val_loss = loss_func(y_val_pred, y_val_batch.unsqueeze(1))
val_epoch_loss += val_loss.item()
loss_stats['train'].append(train_epoch_loss/len(train_loader))
loss_stats['val'].append(val_epoch_loss/len(val_loader))
print(f'Epoch {e+0:03}: | Train Loss: {train_epoch_loss/len(train_loader):.5f} | \
Val Loss: {val_epoch_loss/len(val_loader): .5f}')
# Visualize Loss and Accuracy
#print(loss_stats)
#To plot the loss line plots, we will create dataframe
#from the loss_stats dictionary
train_val_loss_df = pd.DataFrame.from_dict(loss_stats).reset_index().melt(id_vars=['index']).rename(columns={"index":"epochs"})
plt.figure(figsize=(15,8))
sns.lineplot(data=train_val_loss_df, x = "epochs", y="value", hue="variable").set_title('Train-Val Loss/Epoch')
# Test the performance of the model
y_pred_list = list()
with torch.no_grad():
non_spiking_model_regressor.eval()
for X_batch, _ in test_loader:
X_batch = X_batch.to(device)
y_test_pred = non_spiking_model_regressor(X_batch)
y_pred_list.append(y_test_pred.cpu().numpy())
y_pred_list = [b.squeeze().tolist() for a in y_pred_list for b in a]
#print(y_pred_list)
#print(y_test)
#print(len(y_pred_list))
#print(len(y_test))
# Lets check widely used regression metrics like MSE, R-squared, RMSE, RMSRE,
mse = mean_squared_error(y_test, y_pred_list) # MSE
r_square = r2_score(y_test, y_pred_list) # R^2
r = np.corrcoef(y_test, y_pred_list)[0, 1], # r
rmse = np.sqrt(mean_squared_error(y_test, y_pred_list)), # RMSE
rmsre = np.sqrt(np.mean(((y_test-y_pred_list)/y_test)**2)), # RMSRE
mape = np.mean(np.abs((y_test-y_pred_list) / y_test)) * 100 # MAPE
print("Mean Squared Error:", mse)
print("R^2 :",r_square)
print("pcc coefficient", r)
print("Root Mean Squared Error:", rmse)
print("Root Mean Squared Relative Error", rmsre)
print("Mean Absolute Percentage Error", mape)
# dictionary to store all the metrics
eval_metrics = {'Mean Squared Error': mse,
'R^2': r_square,
'pcc coefficient': r,
'Root Mean Squared Error':rmse,
'Root Mean Squared Relative Error': rmsre,
'Mean Absolute Percentage Error': mape
}
#print(eval_metrics)
# Visualize Metrics as bar plot with sns
fig, axes = plt.subplots(nrows=1, ncols=1, figsize=(25,7))
sns.barplot(data = pd.DataFrame.from_dict(eval_metrics).melt(), x = "variable", y="value",
hue="variable").set_title('Model Peformance on Test set')
#eval_metric_df = pd.DataFrame()
'''
eval_metric_df = pd.DataFrame((eval_metrics_list), \
columns=['MSE', 'R-Squared','PearsonCorrelationCoefficient',\
'RMSE', 'RMSRE', 'MAPE'])
print(eval_metric_df)
'''
# plot
#plt.figure(figsize=(6,3))
#sns.barplot(data=eval_metric_df, x="variable", y="value", hue="variable").set_title('Model Peformance on Test set')
'''
plt.plot(log['epoch'], log['loss'], label='Training')
plt.plot(log['epoch'], log['val_loss'], label='Validation')
plt.set_title('Loss Plot', fontsize=35)
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.legend()
'''
# Install regressormetricgraphplot package from terminal or notebook
'''
Terminal:
$ pip install regressormetricgraphplot
OR
$ git clone https://github.com/ajayarunachalam/RegressorMetricGraphPlot
$ cd RegressorMetricGraphPlot
$ python setup.py install
Notebook:
!git clone https://github.com/ajayarunachalam/RegressorMetricGraphPlot.git
cd RegressorMetricGraphPlot/
Just replace the line 'from CompareModels import *' with 'from regressormetricgraphplot import CompareModels'
'''
# Now, let us check how machine learning algorithms perform on this dataset in comparison to the build neural network
from regressioncomparemetricplot import CompareModels
# Linear Regression
# Fitting training set to linear regression model
lr = LinearRegression(n_jobs=-1)
lr.fit(X_train, y_train)
# Predicting the house price
y_pred = lr.predict(X_test)
# Metrics
print(f'R2_nd_RMSE LR MODEL: {CompareModels.R2AndRMSE(y_test=y_test, y_pred=y_pred)}')
plot = CompareModels()
plot.add(model_name='Linear Regression', y_test=y_test, y_pred=y_pred)
plot.show(figsize=(10, 5))
# Fitting Random Forest model to the dataset
rfr = RandomForestRegressor(n_estimators=10, random_state=10, n_jobs=-1)
rfr.fit(X_train, y_train)
# Predicting the house price
y_pred = rfr.predict(X_test)
print(f'R2_nd_RMSE RF MODEL: {CompareModels.R2AndRMSE(y_test=y_test, y_pred=y_pred)}')
plot.add('Random Forest', y_test, y_pred)
plot.show(figsize=(10, 5))
xgb = XGBRegressor(n_jobs=4, silent=False, objective='reg:linear',
max_depth=3, random_state=10, n_estimators=100,
learning_rate=0.3, verbose=True)
xgb.fit(X_train, y_train)
# Predicting the house price
y_pred = xgb.predict(X_test)
print(f'R2_nd_RMSE XGB MODEL: {CompareModels.R2AndRMSE(y_test=y_test, y_pred=y_pred)}')
plot.add('XGBoost', y_test, y_pred)
plot.show(figsize=(10, 5))
if (__name__ == '__main__'):
run_regressor()
```
| github_jupyter |
```
%matplotlib inline
```
# Wasserstein Discriminant Analysis
This example illustrate the use of WDA as proposed in [11].
[11] Flamary, R., Cuturi, M., Courty, N., & Rakotomamonjy, A. (2016).
Wasserstein Discriminant Analysis.
```
# Author: Remi Flamary <remi.flamary@unice.fr>
#
# License: MIT License
import numpy as np
import matplotlib.pylab as pl
from ot.dr import wda, fda
```
Generate data
-------------
```
#%% parameters
n = 1000 # nb samples in source and target datasets
nz = 0.2
# generate circle dataset
t = np.random.rand(n) * 2 * np.pi
ys = np.floor((np.arange(n) * 1.0 / n * 3)) + 1
xs = np.concatenate(
(np.cos(t).reshape((-1, 1)), np.sin(t).reshape((-1, 1))), 1)
xs = xs * ys.reshape(-1, 1) + nz * np.random.randn(n, 2)
t = np.random.rand(n) * 2 * np.pi
yt = np.floor((np.arange(n) * 1.0 / n * 3)) + 1
xt = np.concatenate(
(np.cos(t).reshape((-1, 1)), np.sin(t).reshape((-1, 1))), 1)
xt = xt * yt.reshape(-1, 1) + nz * np.random.randn(n, 2)
nbnoise = 8
xs = np.hstack((xs, np.random.randn(n, nbnoise)))
xt = np.hstack((xt, np.random.randn(n, nbnoise)))
```
Plot data
---------
```
#%% plot samples
pl.figure(1, figsize=(6.4, 3.5))
pl.subplot(1, 2, 1)
pl.scatter(xt[:, 0], xt[:, 1], c=ys, marker='+', label='Source samples')
pl.legend(loc=0)
pl.title('Discriminant dimensions')
pl.subplot(1, 2, 2)
pl.scatter(xt[:, 2], xt[:, 3], c=ys, marker='+', label='Source samples')
pl.legend(loc=0)
pl.title('Other dimensions')
pl.tight_layout()
```
Compute Fisher Discriminant Analysis
------------------------------------
```
#%% Compute FDA
p = 2
Pfda, projfda = fda(xs, ys, p)
```
Compute Wasserstein Discriminant Analysis
-----------------------------------------
```
#%% Compute WDA
p = 2
reg = 1e0
k = 10
maxiter = 100
Pwda, projwda = wda(xs, ys, p, reg, k, maxiter=maxiter)
```
Plot 2D projections
-------------------
```
#%% plot samples
xsp = projfda(xs)
xtp = projfda(xt)
xspw = projwda(xs)
xtpw = projwda(xt)
pl.figure(2)
pl.subplot(2, 2, 1)
pl.scatter(xsp[:, 0], xsp[:, 1], c=ys, marker='+', label='Projected samples')
pl.legend(loc=0)
pl.title('Projected training samples FDA')
pl.subplot(2, 2, 2)
pl.scatter(xtp[:, 0], xtp[:, 1], c=ys, marker='+', label='Projected samples')
pl.legend(loc=0)
pl.title('Projected test samples FDA')
pl.subplot(2, 2, 3)
pl.scatter(xspw[:, 0], xspw[:, 1], c=ys, marker='+', label='Projected samples')
pl.legend(loc=0)
pl.title('Projected training samples WDA')
pl.subplot(2, 2, 4)
pl.scatter(xtpw[:, 0], xtpw[:, 1], c=ys, marker='+', label='Projected samples')
pl.legend(loc=0)
pl.title('Projected test samples WDA')
pl.tight_layout()
pl.show()
```
| github_jupyter |
```
import json
import matplotlib.patches as mpatches
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
```
# Load Dataset
```
# image_val_path = 'bdd100k/images/100k/train/'
# label_path = 'bdd100k/labels/bdd100k_labels_images_train.json'
# save_label_path = 'bdd100k/labels/bdd100k_labels_images_train_highway_lane.json'
image_val_path = 'bdd100k/images/100k/val/'
label_path = 'bdd100k/labels/bdd100k_labels_images_val.json'
save_label_path = 'bdd100k/labels/bdd100k_labels_images_val_highway_lane.json'
json_file = open(label_path, 'r')
labels_json = json.load(json_file)
```
# Keys
```
labels_json[0].keys()
labels_json[0]['labels'][0].keys()
labels_json[0]['attributes'].keys()
labels_json[0]['attributes']
```
# Pandas Dataframe
```
df = pd.DataFrame(labels_json)
print(len(df))
df.head()
```
# Search highway seans
```
df_highway = df[df.apply(lambda x: x['attributes']['scene'], axis=1) == 'highway']
df_highway = df_highway.reset_index(drop=True)
df_highway.head()
df_highway['attributes'][0]
```
# Search parallel four lanes seans
```
df_highway_lane = pd.DataFrame()
for i in range(len(df_highway)):
tmp_se = df_highway.iloc[i]
tmp_label = df_highway.iloc[i]['labels']
lane_count = 0
except_flag = False
for j in range(len(tmp_label)):
if tmp_label[j]['category'] == 'lane':
if tmp_label[j]['attributes']['laneDirection'] == 'parallel' and \
(tmp_label[j]['attributes']['laneType'] == 'single other' or \
tmp_label[j]['attributes']['laneType'] == 'single white' or \
tmp_label[j]['attributes']['laneType'] == 'single yellow'):
lane_count += 1
elif tmp_label[j]['attributes']['laneDirection'] == 'vertical' or \
tmp_label[j]['attributes']['laneType'] == 'crosswalk':
except_flag = True
elif tmp_label[j]['category'] == 'bike' or \
tmp_label[j]['category'] == 'motor' or \
tmp_label[j]['category'] == 'person' or \
tmp_label[j]['category'] == 'rider' or \
tmp_label[j]['category'] == 'traffic light':
except_flag = True
if lane_count >= 4 and except_flag == False:
df_highway_lane = df_highway_lane.append(tmp_se)
df_highway_lane = df_highway_lane.reset_index(drop=True)
print(len(df_highway_lane))
df_highway_lane.head()
```
# Show
```
data_example_name = df_highway_lane.iloc[0]['name']
img_path = image_val_path + data_example_name
img = mpimg.imread(img_path)
img_path
fig = plt.figure(figsize=(10,10))
plt.plot()
plt.imshow(img)
```
# pandas dataframe -> dict
```
dict_highway_lane = df_highway_lane.to_dict(orient='index')
```
# dict -> list
```
list_highway_lane = list(dict_highway_lane.items())
```
# delete list index
```
list_highway_lane[0][0]
l = list_highway_lane
list_highway_lane_delete_index = list([v[1] for v in l])
```
# Save into a json file with indent
```
with open(save_label_path, 'w') as f:
json.dump(list_highway_lane_delete_index, f, indent=4)
```
# Result
```
print(len(df), '->', len(df_highway), '->', len(df_highway_lane))
```
| github_jupyter |
# Lesson 3 Demo 2: Focus on Primary Key
Cassandra logo
### In this demo we are going to walk through the basics of creating a table with a good Primary Key in Apache Cassandra, inserting rows of data, and doing a simple SQL query to validate the information.
#### We will use a python wrapper/ python driver called cassandra to run the Apache Cassandra queries. This library should be preinstalled but to install this library in the future you can run this command in a notebook to install locally:
! pip install cassandra-driver
#### More documentation can be found here: https://datastax.github.io/python-driver/
#### Import Apache Cassandra python package
```
import cassandra
```
### First let's create a connection to the database
```
from cassandra.cluster import Cluster
try:
cluster = Cluster(['127.0.0.1']) #If you have a locally installed Apache Cassandra instance
session = cluster.connect()
except Exception as e:
print(e)
```
### Let's create a keyspace to do our work in
```
try:
session.execute("""
CREATE KEYSPACE IF NOT EXISTS udacity
WITH REPLICATION =
{ 'class' : 'SimpleStrategy', 'replication_factor' : 1 }"""
)
except Exception as e:
print(e)
```
#### Connect to our Keyspace. Compare this to how we had to create a new session in PostgreSQL.
```
try:
session.set_keyspace('udacity')
except Exception as e:
print(e)
```
### Let's imagine we would like to start creating a new Music Library of albums. We are going to work with one of the queries from Exercise 1.
### We want to ask 1 question of our data
#### 1. Give me every album in my music library that was released in a given year
`select * from music_library WHERE YEAR=1970`
### Here is our Collection of Data
Please refer to table 3 image in the video
### How should we model this data? What should be our Primary Key and Partition Key? Since our data is looking for the YEAR let's start with that. Is Partitioning our data by year a good idea? In this case our data is very small, but if we had a larger data set of albums partitions by YEAR might be a find choice. We would need to validate from our dataset. We want an equal spread of the data.
`Table Name: music_library
column 1: Year
column 2: Artist Name
column 3: Album Name
Column 4: City
PRIMARY KEY(year)`
```
query = "CREATE TABLE IF NOT EXISTS music_library "
query = query + "(year int, artist_name text, album_name text, city text, PRIMARY KEY (year))"
try:
session.execute(query)
except Exception as e:
print(e)
```
### Let's insert our data into of table
```
query = "INSERT INTO music_library (year, artist_name, album_name, city)"
query = query + " VALUES (%s, %s, %s, %s)"
try:
session.execute(query, (1970, "The Beatles", "Let it Be", "Liverpool"))
except Exception as e:
print(e)
try:
session.execute(query, (1965, "The Beatles", "Rubber Soul", "Oxford"))
except Exception as e:
print(e)
try:
session.execute(query, (1965, "The Who", "My Generation", "London"))
except Exception as e:
print(e)
try:
session.execute(query, (1966, "The Monkees", "The Monkees", "Los Angeles"))
except Exception as e:
print(e)
try:
session.execute(query, (1970, "The Carpenters", "Close To You", "San Diego"))
except Exception as e:
print(e)
```
### Let's Validate our Data Model -- Did it work?? If we look for Albums from 1965 we should expect to see 2 rows.
`select * from music_library WHERE YEAR=1965`
```
query = "select * from music_library WHERE YEAR=1965"
try:
rows = session.execute(query)
except Exception as e:
print(e)
for row in rows:
print (row.year, row.artist_name, row.album_name, row.city)
```
### That didn't work out as planned! Why is that? Because we did not create a unique primary key.
### Let's Try Again. Let's focus on making the PRIMARY KEY unique. Look at our dataset do we have anything that is unique for each row? We have a couple of options (City and Album Name) but that will not get us the query we need which is looking for album's in a particular year. Let's make a composite key of the `YEAR` AND `ALBUM NAME`. This is assuming that an album name is unique to the year it was released (not a bad bet). --But remember this is just a demo, you will need to understand your dataset fully (no betting!)
```
query = "CREATE TABLE IF NOT EXISTS music_library1 "
query = query + "(year int, artist_name text, album_name text, city text, PRIMARY KEY (year, album_name))"
try:
session.execute(query)
except Exception as e:
print(e)
query = "INSERT INTO music_library1 (year, artist_name, album_name, city)"
query = query + " VALUES (%s, %s, %s, %s)"
try:
session.execute(query, (1970, "The Beatles", "Let it Be", "Liverpool"))
except Exception as e:
print(e)
try:
session.execute(query, (1965, "The Beatles", "Rubber Soul", "Oxford"))
except Exception as e:
print(e)
try:
session.execute(query, (1965, "The Who", "My Generation", "London"))
except Exception as e:
print(e)
try:
session.execute(query, (1966, "The Monkees", "The Monkees", "Los Angeles"))
except Exception as e:
print(e)
try:
session.execute(query, (1970, "The Carpenters", "Close To You", "San Diego"))
except Exception as e:
print(e)
```
### Let's Validate our Data Model -- Did it work?? If we look for Albums from 1965 we should expect to see 2 rows.
`select * from music_library WHERE YEAR=1965`
```
query = "select * from music_library1 WHERE YEAR=1965"
try:
rows = session.execute(query)
except Exception as e:
print(e)
for row in rows:
print (row.year, row.artist_name, row.album_name, row.city)
```
### Success it worked! We created a unique Primary key that evenly distributed our data.
### For the sake of the demo, I will drop the table.
```
query = "drop table music_library"
try:
rows = session.execute(query)
except Exception as e:
print(e)
query = "drop table music_library1"
try:
rows = session.execute(query)
except Exception as e:
print(e)
```
### And Finally close the session and cluster connection
```
session.shutdown()
cluster.shutdown()
```
| github_jupyter |
In this notebook, we preprocessed the data and feed the data to gradient boosting tree models, and got 1.39 on public leaderboard.
the workflow is as follows:
1. **Data preprocessing**. The purpose of data preprocessing is to achieve higher time/space efficiency. What we did includes round, constant features removal, duplicate features removal, insignificant features removal, etc. The key here is to ensure the preprocessing shall not hurt the accuracy.
2. **Feature transform**. The purpose of feature transform is to help the models to better grasp the information in the data, and fight overfitting. What we did includes dropping features which "live" on different distributions on training/testing set, adding statistical features, adding low-dimensional representation as features.
3. **Modeling**. We used 2 models: xgboost and lightgbm. We averaged the 2 models for the final prediction.
Stay tuned, more update will come.
references:
* [Distribution of Test vs. Training data](https://www.kaggle.com/nanomathias/distribution-of-test-vs-training-data)
* [Ensemble of LGBM and XGB](https://www.kaggle.com/lightsalsa/ensemble-of-lgbm-and-xgb)
* [predict house prices-model tuning & ensemble](https://www.kaggle.com/alexpengxiao/predict-house-prices-model-tuning-ensemble)
* [Stacked Regressions : Top 4% on LeaderBoard](https://www.kaggle.com/serigne/stacked-regressions-top-4-on-leaderboard)
**step 1**: load train & test data, drop duplicate columns, round the features to NUM_OF_DECIMALS decimals. here NUM_OF_DECIMALS is a experience value which can be tuned.
```
#coding:utf-8
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import os
import warnings
warnings.filterwarnings("ignore")
print(os.listdir("/Users/szkfzx/datasets/santander-value-prediction-challenge"))
train = pd.read_csv('/Users/szkfzx/datasets/santander-value-prediction-challenge/train.csv')
test = pd.read_csv('/Users/szkfzx/datasets/santander-value-prediction-challenge/test.csv')
#去掉ID,处理target列
test_ID = test['ID']
y_train = train['target']
y_train = np.log1p(y_train) # log1p=log(x+1)
train.drop("ID", axis = 1, inplace = True)
train.drop("target", axis = 1, inplace = True)
test.drop("ID", axis = 1, inplace = True)
#去掉只有一个值的列
cols_with_onlyone_val = train.columns[train.nunique() == 1]
train.drop(cols_with_onlyone_val.values, axis=1, inplace=True)
test.drop(cols_with_onlyone_val.values, axis=1, inplace=True)
#四舍五入,保留32位小数
NUM_OF_DECIMALS = 32
train = train.round(NUM_OF_DECIMALS)
test = test.round(NUM_OF_DECIMALS)
#删除重复的列
colsToRemove = []
columns = train.columns
for i in range(len(columns)-1):
v = train[columns[i]].values
dupCols = []
for j in range(i + 1,len(columns)):
if np.array_equal(v, train[columns[j]].values):
colsToRemove.append(columns[j])
train.drop(colsToRemove, axis=1, inplace=True)
test.drop(colsToRemove, axis=1, inplace=True)
print(train.shape)
print(test.shape)
```
**step 2**: Select features by importance. here we used a weak RandomForestRegressor to get the feature importance. here we select top NUM_OF_FEATURES important features. NUM_OF_FEATURES here is a hyper parameter that can be tuned.
```
# model_selection和ensemble通常一起使用
from sklearn import model_selection
from sklearn import ensemble
NUM_OF_FEATURES = 1000
def rmsle(y, pred):
return np.sqrt(np.mean(np.power(y - pred, 2)))
x1, x2, y1, y2 = model_selection.train_test_split(
train, y_train.values, test_size=0.20, random_state=5)
model = ensemble.RandomForestRegressor(n_jobs=-1, random_state=7)
model.fit(x1, y1)
print(rmsle(y2, model.predict(x2)))
col = pd.DataFrame({'importance': model.feature_importances_, 'feature': train.columns}).sort_values(
by=['importance'], ascending=[False])[:NUM_OF_FEATURES]['feature'].values
train = train[col]
test = test[col]
train.shape
col #最重要的1000个特征
```
**step 3**: we try to test the training data and testing data with Kolmogorov-Smirnov test. This is a two-sided test for the null hypothesis that whether 2 independent samples are drawn from the same continuous distribution([see more](https://docs.scipy.org/doc/scipy-0.14.0/reference/generated/scipy.stats.ks_2samp.html)). If a feature has different distributions in training set than in testing set, we should remove this feature since what we learned during training cannot generalize. THRESHOLD_P_VALUE and THRESHOLD_STATISTIC are hyper parameters.
```
#去掉训练数据集和测试数据集中分布差异很大的列
from scipy.stats import ks_2samp
THRESHOLD_P_VALUE = 0.01 #need tuned
THRESHOLD_STATISTIC = 0.3 #need tuned
diff_cols = []
for col in train.columns:
statistic, pvalue = ks_2samp(train[col].values, test[col].values)
if pvalue <= THRESHOLD_P_VALUE and np.abs(statistic) > THRESHOLD_STATISTIC:
diff_cols.append(col)
for col in diff_cols:
if col in train.columns:
train.drop(col, axis=1, inplace=True)
test.drop(col, axis=1, inplace=True)
print(train.shape)
print(test.shape)
```
**step 4**: We add some additional statistical features to the original features. Moreover, we also added low-dimensional representations as features. NUM_OF_COM is hyper parameter
```
#增加一些统计特征作为训练数据集合测试数据集的列
from sklearn import random_projection
ntrain = len(train)
ntest = len(test)
tmp = pd.concat([train,test])#RandomProjection
weight = ((train != 0).sum()/len(train)).values
tmp_train = train[train!=0]
tmp_test = test[test!=0]
#权重
train["weight_count"] = (tmp_train*weight).sum(axis=1)
test["weight_count"] = (tmp_test*weight).sum(axis=1)
#非0
train["count_not0"] = (train != 0).sum(axis=1)
test["count_not0"] = (test != 0).sum(axis=1)
#和
train["sum"] = train.sum(axis=1)
test["sum"] = test.sum(axis=1)
#方差
train["var"] = tmp_train.var(axis=1)
test["var"] = tmp_test.var(axis=1)
#中位数
train["median"] = tmp_train.median(axis=1)
test["median"] = tmp_test.median(axis=1)
#均值
train["mean"] = tmp_train.mean(axis=1)
test["mean"] = tmp_test.mean(axis=1)
#标准差
train["std"] = tmp_train.std(axis=1)
test["std"] = tmp_test.std(axis=1)
#最大值
train["max"] = tmp_train.max(axis=1)
test["max"] = tmp_test.max(axis=1)
#最小值
train["min"] = tmp_train.min(axis=1)
test["min"] = tmp_test.min(axis=1)
#偏度
train["skew"] = tmp_train.skew(axis=1)
test["skew"] = tmp_test.skew(axis=1)
#峰度
train["kurtosis"] = tmp_train.kurtosis(axis=1)
test["kurtosis"] = tmp_test.kurtosis(axis=1)
del(tmp_train)
del(tmp_test)
NUM_OF_COM = 100 #need tuned
transformer = random_projection.SparseRandomProjection(n_components = NUM_OF_COM)
RP = transformer.fit_transform(tmp)
rp = pd.DataFrame(RP)
columns = ["RandomProjection{}".format(i) for i in range(NUM_OF_COM)]
rp.columns = columns
rp_train = rp[:ntrain]
rp_test = rp[ntrain:]
rp_test.index = test.index
#concat RandomProjection and raw data
train = pd.concat([train,rp_train],axis=1)
test = pd.concat([test,rp_test],axis=1)
del(rp_train)
del(rp_test)
train.shape
test.shape
tmp.head()
weight
train.head()
test.head()
```
**step 5**: Define cross-validation methods and models. xgboost and lightgbm are used as base models. the hyper parameters are already tuned by grid search, here we use them directly. NUM_FOLDS can be treat as hyper parameter
```
from sklearn.model_selection import KFold, cross_val_score, train_test_split
from sklearn.base import BaseEstimator, TransformerMixin, RegressorMixin, clone #转换器基类、回归基类
from sklearn.metrics import mean_squared_error
import xgboost as xgb
import lightgbm as lgb
#define evaluation method for a given model. we use k-fold cross validation on the training set.
#the loss function is root mean square logarithm error between target and prediction
#note: train and y_train are feeded as global variables
NUM_FOLDS = 5 #need tuned
def rmsle_cv(model):
kf = KFold(NUM_FOLDS, shuffle=True, random_state=42).get_n_splits(train.values)
rmse= np.sqrt(-cross_val_score(model, train, y_train, scoring="neg_mean_squared_error", cv = kf))
return(rmse)
#ensemble method: model averaging
class AveragingModels(BaseEstimator, RegressorMixin, TransformerMixin):
def __init__(self, models):
self.models = models
# we define clones of the original models to fit the data in
# the reason of clone is avoiding affect the original base models
def fit(self, X, y):
self.models_ = [clone(x) for x in self.models]
# Train cloned base models
for model in self.models_:
model.fit(X, y)
return self
#Now we do the predictions for cloned models and average them
def predict(self, X):
predictions = np.column_stack([ model.predict(X) for model in self.models_ ])
return np.mean(predictions, axis=1)
model_xgb = xgb.XGBRegressor(colsample_bytree=0.055, colsample_bylevel =0.5,
gamma=1.5, learning_rate=0.02, max_depth=32,
objective='reg:linear',booster='gbtree',
min_child_weight=57, n_estimators=1000, reg_alpha=0,
reg_lambda = 0,eval_metric = 'rmse', subsample=0.7,
silent=1, n_jobs = -1, early_stopping_rounds = 14,
random_state =7, nthread = -1)
model_lgb = lgb.LGBMRegressor(objective='regression',num_leaves=144,
learning_rate=0.005, n_estimators=720, max_depth=13,
metric='rmse',is_training_metric=True,
max_bin = 55, bagging_fraction = 0.8,verbose=-1,
bagging_freq = 5, feature_fraction = 0.9)
score = rmsle_cv(model_xgb)
print("Xgboost score: {:.4f} ({:.4f})\n".format(score.mean(), score.std()))
score = rmsle_cv(model_lgb)
print("LGBM score: {:.4f} ({:.4f})\n" .format(score.mean(), score.std()))
averaged_models = AveragingModels(models = (model_xgb, model_lgb))
score = rmsle_cv(averaged_models)
print("averaged score: {:.4f} ({:.4f})\n" .format(score.mean(), score.std()))
```
**step 6**: average the two base models and submit the final predictions.
```
averaged_models.fit(train.values, y_train)
pred = np.expm1(averaged_models.predict(test.values))
ensemble = pred
sub = pd.DataFrame()
sub['ID'] = test_ID
sub['target'] = ensemble
sub.to_csv('submission.csv',index=False)
#Xgboost score: 1.3582 (0.0640)
#LGBM score: 1.3437 (0.0519)
#averaged score: 1.3431 (0.0586)
#Xgboost score: 1.3566 (0.0525)
#LGBM score: 1.3477 (0.0497)
#averaged score: 1.3438 (0.0516)
#Xgboost score: 1.3540 (0.0621)
#LGBM score: 1.3463 (0.0485)
#averaged score: 1.3423 (0.0556)
sub.T
```
| github_jupyter |
# Task: Decision Tree Classifier
# Author: Vibhuti Mayekar
```
import pandas as pd
import numpy as np
import os
import seaborn as sns
import matplotlib.pyplot as plt
%matplotlib inline
from sklearn.model_selection import train_test_split
from sklearn.cluster import KMeans
from sklearn.tree import DecisionTreeClassifier
from sklearn.linear_model import LinearRegression
os.chdir("F:\\Vibhuti\\business analytics and intelligence\\Internshala\\internshipwork_grip")
#
df = pd.read_csv('Iris.csv')
df.head(5)
sns.pairplot(df,hue='Species',palette='Set1')
###Visualizing the NULL or NA pr NAN values
fig, ax = plt.subplots(figsize = (15,5))
x = df.isna().sum().index
y = df.isna().sum()
ax.bar(x=x, height = y)
ax.set_xticklabels(x, rotation = 45)
plt.tight_layout()
plt.figure(figsize = (12,8))
sns.boxplot(data = df)
feature_cols = ['SepalLengthCm', 'SepalWidthCm', 'PetalLengthCm', 'PetalWidthCm']
x = df[feature_cols]
y= df['Species']
# Split dataset into training set and test set
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size=0.3, random_state=1) # 70% training and 30% test
# Create Decision Tree classifer object
clf = DecisionTreeClassifier()
# Train Decision Tree Classifer
clf = clf.fit(X_train,y_train)
#Predict the response for test dataset
y_pred = clf.predict(X_test)
from sklearn import metrics #Import scikit-learn metrics module for accuracy calculation
# Model Accuracy, how often is the classifier correct?
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
import graphviz
# Create Decision Tree classifer object
clf = DecisionTreeClassifier(criterion="entropy", max_depth=3)
# Train Decision Tree Classifer
clf = clf.fit(X_train,y_train)
#Predict the response for test dataset
y_pred = clf.predict(X_test)
# Model Accuracy, how often is the classifier correct?
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
import matplotlib.pyplot as plt
from sklearn.datasets import load_iris
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
import pandas as pd
import numpy as np
from sklearn import tree
tree.plot_tree(clf);
from matplotlib import pyplot as plt
from sklearn import datasets
from sklearn.tree import DecisionTreeClassifier
from sklearn import tree
# Prepare the data data
iris = datasets.load_iris()
X = iris.data
y = iris.target
# Fit the classifier with default hyper-parameters
clf = DecisionTreeClassifier(random_state=1234)
model = clf.fit(X, y)
text_representation = tree.export_text(clf)
print(text_representation)
with open("decistion_tree.log", "w") as fout:
fout.write(text_representation)
```
# Plotting the graph
```
fig = plt.figure(figsize=(25,20))
_ = tree.plot_tree(clf,
feature_names=iris.feature_names,
class_names=iris.target_names,
filled=True)
```
# Thank You..
| github_jupyter |
# Artificial Intelligence Nanodegree
## Convolutional Neural Networks
## Project: Write an Algorithm for a Dog Identification App
---
In this notebook, some template code has already been provided for you, and you will need to implement additional functionality to successfully complete this project. You will not need to modify the included code beyond what is requested. Sections that begin with **'(IMPLEMENTATION)'** in the header indicate that the following block of code will require additional functionality which you must provide. Instructions will be provided for each section, and the specifics of the implementation are marked in the code block with a 'TODO' statement. Please be sure to read the instructions carefully!
> **Note**: Once you have completed all of the code implementations, you need to finalize your work by exporting the iPython Notebook as an HTML document. Before exporting the notebook to html, all of the code cells need to have been run so that reviewers can see the final implementation and output. You can then export the notebook by using the menu above and navigating to \n",
"**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
In addition to implementing code, there will be questions that you must answer which relate to the project and your implementation. Each section where you will answer a question is preceded by a **'Question X'** header. Carefully read each question and provide thorough answers in the following text boxes that begin with **'Answer:'**. Your project submission will be evaluated based on your answers to each of the questions and the implementation you provide.
>**Note:** Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. Markdown cells can be edited by double-clicking the cell to enter edit mode.
The rubric contains _optional_ "Stand Out Suggestions" for enhancing the project beyond the minimum requirements. If you decide to pursue the "Stand Out Suggestions", you should include the code in this IPython notebook.
---
### Why We're Here
In this notebook, you will make the first steps towards developing an algorithm that could be used as part of a mobile or web app. At the end of this project, your code will accept any user-supplied image as input. If a dog is detected in the image, it will provide an estimate of the dog's breed. If a human is detected, it will provide an estimate of the dog breed that is most resembling. The image below displays potential sample output of your finished project (... but we expect that each student's algorithm will behave differently!).

In this real-world setting, you will need to piece together a series of models to perform different tasks; for instance, the algorithm that detects humans in an image will be different from the CNN that infers dog breed. There are many points of possible failure, and no perfect algorithm exists. Your imperfect solution will nonetheless create a fun user experience!
### The Road Ahead
We break the notebook into separate steps. Feel free to use the links below to navigate the notebook.
* [Step 0](#step0): Import Datasets
* [Step 1](#step1): Detect Humans
* [Step 2](#step2): Detect Dogs
* [Step 3](#step3): Create a CNN to Classify Dog Breeds (from Scratch)
* [Step 4](#step4): Use a CNN to Classify Dog Breeds (using Transfer Learning)
* [Step 5](#step5): Create a CNN to Classify Dog Breeds (using Transfer Learning)
* [Step 6](#step6): Write your Algorithm
* [Step 7](#step7): Test Your Algorithm
---
<a id='step0'></a>
## Step 0: Import Datasets
### Import Dog Dataset
In the code cell below, we import a dataset of dog images. We populate a few variables through the use of the `load_files` function from the scikit-learn library:
- `train_files`, `valid_files`, `test_files` - numpy arrays containing file paths to images
- `train_targets`, `valid_targets`, `test_targets` - numpy arrays containing onehot-encoded classification labels
- `dog_names` - list of string-valued dog breed names for translating labels
## Working with a Mounted Dataset
- In order to carry out this project on a GPU. I've mounted the datasets into the two directories below via FloydHub. The code will be modified so as to load_files from the mounted directories instead of the local directories
```
from sklearn.datasets import load_files
from keras.utils import np_utils
import numpy as np
from glob import glob
# define function to load train, test, and validation datasets
def load_dataset(path):
data = load_files(path)
dog_files = np.array(data['filenames'])
dog_targets = np_utils.to_categorical(np.array(data['target']), 133)
return dog_files, dog_targets
# load train, test, and validation datasets
train_files, train_targets = load_dataset('dogImages/train')
valid_files, valid_targets = load_dataset('dogImages/valid')
test_files, test_targets = load_dataset('dogImages/test')
# load list of dog names
dog_names = [item[20:-1] for item in sorted(glob("dogImages/train/*/"))]
# print statistics about the dataset
print('There are %d total dog categories.' % len(dog_names))
print('There are %s total dog images.\n' % len(np.hstack([train_files, valid_files, test_files])))
print('There are %d training dog images.' % len(train_files))
print('There are %d validation dog images.' % len(valid_files))
print('There are %d test dog images.'% len(test_files))
```
### Import Human Dataset
In the code cell below, we import a dataset of human images, where the file paths are stored in the numpy array `human_files`.
```
import random
random.seed(8675309)
# load filenames in shuffled human dataset
human_files = np.array(glob("lfw/*/*"))
random.shuffle(human_files)
# print statistics about the dataset
print('There are %d total human images.' % len(human_files))
```
---
<a id='step1'></a>
## Step 1: Detect Humans
We use OpenCV's implementation of [Haar feature-based cascade classifiers](http://docs.opencv.org/trunk/d7/d8b/tutorial_py_face_detection.html) to detect human faces in images. OpenCV provides many pre-trained face detectors, stored as XML files on [github](https://github.com/opencv/opencv/tree/master/data/haarcascades). We have downloaded one of these detectors and stored it in the `haarcascades` directory.
In the next code cell, we demonstrate how to use this detector to find human faces in a sample image.
```
import cv2
import matplotlib.pyplot as plt
%matplotlib inline
# extract pre-trained face detector
face_cascade = cv2.CascadeClassifier('haarcascades/haarcascade_frontalface_alt.xml')
# load color (BGR) image
img = cv2.imread(human_files[3])
# convert BGR image to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
# find faces in image
faces = face_cascade.detectMultiScale(gray)
# print number of faces detected in the image
print('Number of faces detected:', len(faces))
# get bounding box for each detected face
for (x,y,w,h) in faces:
# add bounding box to color image
cv2.rectangle(img,(x,y),(x+w,y+h),(255,0,0),2)
# convert BGR image to RGB for plotting
cv_rgb = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
# display the image, along with bounding box
plt.imshow(cv_rgb)
plt.show()
```
Before using any of the face detectors, it is standard procedure to convert the images to grayscale. The `detectMultiScale` function executes the classifier stored in `face_cascade` and takes the grayscale image as a parameter.
In the above code, `faces` is a numpy array of detected faces, where each row corresponds to a detected face. Each detected face is a 1D array with four entries that specifies the bounding box of the detected face. The first two entries in the array (extracted in the above code as `x` and `y`) specify the horizontal and vertical positions of the top left corner of the bounding box. The last two entries in the array (extracted here as `w` and `h`) specify the width and height of the box.
### Write a Human Face Detector
We can use this procedure to write a function that returns `True` if a human face is detected in an image and `False` otherwise. This function, aptly named `face_detector`, takes a string-valued file path to an image as input and appears in the code block below.
```
# returns "True" if face is detected in image stored at img_path
def face_detector(img_path):
img = cv2.imread(img_path)
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
faces = face_cascade.detectMultiScale(gray)
return len(faces) > 0
```
### (IMPLEMENTATION) Assess the Human Face Detector
__Question 1:__ Use the code cell below to test the performance of the `face_detector` function.
- What percentage of the first 100 images in `human_files` have a detected human face?
- What percentage of the first 100 images in `dog_files` have a detected human face?
Ideally, we would like 100% of human images with a detected face and 0% of dog images with a detected face. You will see that our algorithm falls short of this goal, but still gives acceptable performance. We extract the file paths for the first 100 images from each of the datasets and store them in the numpy arrays `human_files_short` and `dog_files_short`.
__Answer:__
Answers for both questions are in the following code cell
```
human_files_short = human_files[:100]
dog_files_short = train_files[:100]
# Do NOT modify the code above this line.
human_file = [face_detector(file) for file in human_files_short]
dog_file = [face_detector(file) for file in dog_files_short]
human_percentage = human_file.count(True) / len(human_file) * 100
human_in_dog_percentange = dog_file.count(True) / len(dog_file) * 100
## TODO: Test the performance of the face_detector algorithm
## on the images in human_files_short and dog_files_short.
print("% of human faces detected in human file : {}%".format(human_percentage))
print("% of human faces deteted in dog file : {}%".format(human_in_dog_percentange))
```
__Question 2:__ This algorithmic choice necessitates that we communicate to the user that we accept human images only when they provide a clear view of a face (otherwise, we risk having unneccessarily frustrated users!). In your opinion, is this a reasonable expectation to pose on the user? If not, can you think of a way to detect humans in images that does not necessitate an image with a clearly presented face?
__Answer:__
Yes I believe that this expectation is reasonable because I think that people have very high expectations for AI systems. AI is one of those technologies that has to work really well otherwise the output of the system loses all the magic to the user. For example, a good conversational interface that picks up on exactly what I'm asking feels to me like a magical experience. However, when I am grossly misunderstood it takes away the excitement of using the algorithm. It is definately frustrating. On the other hand if you wanted to be sure you could detect a human in a photo even though the face was unclear other body parts could be used such as hands, two arms, and feet which are very distinguishable when it comes to classifiying a human.
We suggest the face detector from OpenCV as a potential way to detect human images in your algorithm, but you are free to explore other approaches, especially approaches that make use of deep learning :). Please use the code cell below to design and test your own face detection algorithm. If you decide to pursue this _optional_ task, report performance on each of the datasets.
```
## (Optional) TODO: Report the performance of another
## face detection algorithm on the LFW dataset
### Feel free to use as many code cells as needed.
```
---
<a id='step2'></a>
## Step 2: Detect Dogs
In this section, we use a pre-trained [ResNet-50](http://ethereon.github.io/netscope/#/gist/db945b393d40bfa26006) model to detect dogs in images. Our first line of code downloads the ResNet-50 model, along with weights that have been trained on [ImageNet](http://www.image-net.org/), a very large, very popular dataset used for image classification and other vision tasks. ImageNet contains over 10 million URLs, each linking to an image containing an object from one of [1000 categories](https://gist.github.com/yrevar/942d3a0ac09ec9e5eb3a). Given an image, this pre-trained ResNet-50 model returns a prediction (derived from the available categories in ImageNet) for the object that is contained in the image.
```
from keras.applications.resnet50 import ResNet50
# define ResNet50 model
ResNet50_model = ResNet50(weights='imagenet')
```
### Pre-process the Data
When using TensorFlow as backend, Keras CNNs require a 4D array (which we'll also refer to as a 4D tensor) as input, with shape
$$
(\text{nb_samples}, \text{rows}, \text{columns}, \text{channels}),
$$
where `nb_samples` corresponds to the total number of images (or samples), and `rows`, `columns`, and `channels` correspond to the number of rows, columns, and channels for each image, respectively.
The `path_to_tensor` function below takes a string-valued file path to a color image as input and returns a 4D tensor suitable for supplying to a Keras CNN. The function first loads the image and resizes it to a square image that is $224 \times 224$ pixels. Next, the image is converted to an array, which is then resized to a 4D tensor. In this case, since we are working with color images, each image has three channels. Likewise, since we are processing a single image (or sample), the returned tensor will always have shape
$$
(1, 224, 224, 3).
$$
The `paths_to_tensor` function takes a numpy array of string-valued image paths as input and returns a 4D tensor with shape
$$
(\text{nb_samples}, 224, 224, 3).
$$
Here, `nb_samples` is the number of samples, or number of images, in the supplied array of image paths. It is best to think of `nb_samples` as the number of 3D tensors (where each 3D tensor corresponds to a different image) in your dataset!
```
from keras.preprocessing import image
from tqdm import tqdm
def path_to_tensor(img_path):
# loads RGB image as PIL.Image.Image type
img = image.load_img(img_path, target_size=(224, 224))
# convert PIL.Image.Image type to 3D tensor with shape (224, 224, 3)
x = image.img_to_array(img)
# convert 3D tensor to 4D tensor with shape (1, 224, 224, 3) and return 4D tensor
return np.expand_dims(x, axis=0)
def paths_to_tensor(img_paths):
list_of_tensors = [path_to_tensor(img_path) for img_path in tqdm(img_paths)]
return np.vstack(list_of_tensors)
```
### Making Predictions with ResNet-50
Getting the 4D tensor ready for ResNet-50, and for any other pre-trained model in Keras, requires some additional processing. First, the RGB image is converted to BGR by reordering the channels. All pre-trained models have the additional normalization step that the mean pixel (expressed in RGB as $[103.939, 116.779, 123.68]$ and calculated from all pixels in all images in ImageNet) must be subtracted from every pixel in each image. This is implemented in the imported function `preprocess_input`. If you're curious, you can check the code for `preprocess_input` [here](https://github.com/fchollet/keras/blob/master/keras/applications/imagenet_utils.py).
Now that we have a way to format our image for supplying to ResNet-50, we are now ready to use the model to extract the predictions. This is accomplished with the `predict` method, which returns an array whose $i$-th entry is the model's predicted probability that the image belongs to the $i$-th ImageNet category. This is implemented in the `ResNet50_predict_labels` function below.
By taking the argmax of the predicted probability vector, we obtain an integer corresponding to the model's predicted object class, which we can identify with an object category through the use of this [dictionary](https://gist.github.com/yrevar/942d3a0ac09ec9e5eb3a).
```
from keras.applications.resnet50 import preprocess_input, decode_predictions
def ResNet50_predict_labels(img_path):
# returns prediction vector for image located at img_path
img = preprocess_input(path_to_tensor(img_path))
return np.argmax(ResNet50_model.predict(img))
```
### Write a Dog Detector
While looking at the [dictionary](https://gist.github.com/yrevar/942d3a0ac09ec9e5eb3a), you will notice that the categories corresponding to dogs appear in an uninterrupted sequence and correspond to dictionary keys 151-268, inclusive, to include all categories from `'Chihuahua'` to `'Mexican hairless'`. Thus, in order to check to see if an image is predicted to contain a dog by the pre-trained ResNet-50 model, we need only check if the `ResNet50_predict_labels` function above returns a value between 151 and 268 (inclusive).
We use these ideas to complete the `dog_detector` function below, which returns `True` if a dog is detected in an image (and `False` if not).
```
### returns "True" if a dog is detected in the image stored at img_path
def dog_detector(img_path):
prediction = ResNet50_predict_labels(img_path)
return ((prediction <= 268) & (prediction >= 151))
```
### (IMPLEMENTATION) Assess the Dog Detector
__Question 3:__ Use the code cell below to test the performance of your `dog_detector` function.
- What percentage of the images in `human_files_short` have a detected dog?
- What percentage of the images in `dog_files_short` have a detected dog?
__Answer:__
Answer to both questions generated in the bottom cell
```
### TODO: Test the performance of the dog_detector function
### on the images in human_files_short and dog_files_short.
human_file = [dog_detector(file) for file in human_files_short]
dog_file = [dog_detector(file) for file in dog_files_short]
p_detected_dog_in_human_file = human_file.count(True) / len(human_file) * 100
p_detected_dog_in_dog_file = dog_file.count(True) / len(dog_file) * 100
print("Percentage of the images in human_files_short have a detected dog: {}%".format(p_detected_dog_in_human_file))
print("Percentage of the images in dog_files_short have a detected dog {}%".format(p_detected_dog_in_dog_file))
```
---
<a id='step3'></a>
## Step 3: Create a CNN to Classify Dog Breeds (from Scratch)
Now that we have functions for detecting humans and dogs in images, we need a way to predict breed from images. In this step, you will create a CNN that classifies dog breeds. You must create your CNN _from scratch_ (so, you can't use transfer learning _yet_!), and you must attain a test accuracy of at least 1%. In Step 5 of this notebook, you will have the opportunity to use transfer learning to create a CNN that attains greatly improved accuracy.
Be careful with adding too many trainable layers! More parameters means longer training, which means you are more likely to need a GPU to accelerate the training process. Thankfully, Keras provides a handy estimate of the time that each epoch is likely to take; you can extrapolate this estimate to figure out how long it will take for your algorithm to train.
We mention that the task of assigning breed to dogs from images is considered exceptionally challenging. To see why, consider that *even a human* would have great difficulty in distinguishing between a Brittany and a Welsh Springer Spaniel.
Brittany | Welsh Springer Spaniel
- | -
<img src="images/Brittany_02625.jpg" width="100"> | <img src="images/Welsh_springer_spaniel_08203.jpg" width="200">
It is not difficult to find other dog breed pairs with minimal inter-class variation (for instance, Curly-Coated Retrievers and American Water Spaniels).
Curly-Coated Retriever | American Water Spaniel
- | -
<img src="images/Curly-coated_retriever_03896.jpg" width="200"> | <img src="images/American_water_spaniel_00648.jpg" width="200">
Likewise, recall that labradors come in yellow, chocolate, and black. Your vision-based algorithm will have to conquer this high intra-class variation to determine how to classify all of these different shades as the same breed.
Yellow Labrador | Chocolate Labrador | Black Labrador
- | -
<img src="images/Labrador_retriever_06457.jpg" width="150"> | <img src="images/Labrador_retriever_06455.jpg" width="240"> | <img src="images/Labrador_retriever_06449.jpg" width="220">
We also mention that random chance presents an exceptionally low bar: setting aside the fact that the classes are slightly imabalanced, a random guess will provide a correct answer roughly 1 in 133 times, which corresponds to an accuracy of less than 1%.
Remember that the practice is far ahead of the theory in deep learning. Experiment with many different architectures, and trust your intuition. And, of course, have fun!
### Pre-process the Data
We rescale the images by dividing every pixel in every image by 255.
```
from PIL import ImageFile
ImageFile.LOAD_TRUNCATED_IMAGES = True
# pre-process the data for Keras
train_tensors = paths_to_tensor(train_files).astype('float32')/255
valid_tensors = paths_to_tensor(valid_files).astype('float32')/255
test_tensors = paths_to_tensor(test_files).astype('float32')/255
```
### (IMPLEMENTATION) Model Architecture
Create a CNN to classify dog breed. At the end of your code cell block, summarize the layers of your model by executing the line:
model.summary()
We have imported some Python modules to get you started, but feel free to import as many modules as you need. If you end up getting stuck, here's a hint that specifies a model that trains relatively fast on CPU and attains >1% test accuracy in 5 epochs:

__Question 4:__ Outline the steps you took to get to your final CNN architecture and your reasoning at each step. If you chose to use the hinted architecture above, describe why you think that CNN architecture should work well for the image classification task.
__Answer:__
Reasoning behind the CNN design from scratch listed after the next block
```
from keras.layers import Conv2D, MaxPooling2D, GlobalAveragePooling2D
from keras.layers import Dropout, Flatten, Dense
from keras.models import Sequential
model = Sequential()
### TODO: Define your architecture.
model.add(Conv2D(16, (3,3), input_shape=(224,224,3), activation = 'relu'))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Conv2D(16, (3,3), activation= 'relu'))
model.add(MaxPooling2D(pool_size =(2,2)))
model.add(Conv2D(32, (3,3), activation= 'relu'))
model.add(GlobalAveragePooling2D())
model.add(Dense(units=64, activation= 'relu'))
model.add(Dropout(0.3))
model.add(Dense(units= 133, activation = 'softmax'))
model.summary()
```
### Reasoning behind architecture
Before creating this network I had to consider the type of machine I was working on. I currently don't have access to a GPU enabled machine so my goal was to come up with a architecture that was small, but was good enough to predict better than 1%. The above architecture was what I came up with.
Initially, I chose a Convolutional layer of 16 filters utilizing a 3,3 window to handle the initial input of the Neural Network. This way the images could be examined in a much more compacted way. I also chose the activation function `relu`. I followed this layer with a max pooling layer of 2,2 which is standard.
Next, I Added another convolutional layer that is the same size as the first layer so as to preserve the data that was passed through from the prior layer. Nothing was changed here as it was designed purposely to be the exact same.
Following the first 2 layers I changed the third convolutional layer to include 32 filters as this allows for the network to start looking at more detail and open up room for more data. This way I could give the network more information to classify the dog breed successfully. The `kernel_size` remained the same, as well as the activation function. The major difference here is the 3 layer yielded a Global Average pooling layer that would then be fed into the fully connected layer.
The first fully connected hidden layer makes use of 64 units as this was a estimate by me to allow for a butter before the output layer to give the network more space to determine the weights. This layer is followed by a Dropout layer to help prevent overfitting before finally being passed to the output layer that makes a prediction of the specific dog breed.
The output layer has a `softmax` activation function which allows it maintain the probability distribution of all 133 species in between numbers 0,1.
### Compile the Model
```
model.compile(optimizer='rmsprop', loss='categorical_crossentropy', metrics=['accuracy'])
```
### (IMPLEMENTATION) Train the Model
Train your model in the code cell below. Use model checkpointing to save the model that attains the best validation loss.
You are welcome to [augment the training data](https://blog.keras.io/building-powerful-image-classification-models-using-very-little-data.html), but this is not a requirement.
```
from keras.callbacks import ModelCheckpoint
from keras.preprocessing.image import ImageDataGenerator
### TODO: specify the number of epochs that you would like to use to train the model.
## Data Augmentation (Took examples from keras blog)
train_datagen = ImageDataGenerator(rescale=1./255,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True)
test_datagen = ImageDataGenerator(rescale=1./255)
train_generator = train_datagen.flow_from_directory('dogImages/train',
target_size=(224, 224),
batch_size=20,
class_mode='categorical')
validation_generator = test_datagen.flow_from_directory('dogImages/valid',
target_size=(224,224),
batch_size=20,
class_mode='categorical')
# Have to be mindful here as This is really expensive Computationaly, so lowered the number of training epoch steps
model.fit_generator(train_generator,
steps_per_epoch=250,
epochs = 20,
validation_data=validation_generator,
validation_steps=600)
model.save_weights('saved_models/cnn_from_scratch.h5')
#### Training
epochs = 20
### Do NOT modify the code below this line.
checkpointer = ModelCheckpoint(filepath='saved_models/weights.best.from_scratch.hdf5',
verbose=1, save_best_only=True)
model.load_weights('saved_models/cnn_from_scratch.h5')
model.fit(train_tensors, train_targets,
validation_data=(valid_tensors, valid_targets),
epochs=epochs, batch_size=20, callbacks=[checkpointer], verbose=1)
```
### Load the Model with the Best Validation Loss
```
model.load_weights('saved_models/weights.best.from_scratch.hdf5')
```
### Test the Model
Try out your model on the test dataset of dog images. Ensure that your test accuracy is greater than 1%.
```
# get index of predicted dog breed for each image in test set
dog_breed_predictions = [np.argmax(model.predict(np.expand_dims(tensor, axis=0))) for tensor in test_tensors]
# report test accuracy
test_accuracy = 100*np.sum(np.array(dog_breed_predictions)==np.argmax(test_targets, axis=1))/len(dog_breed_predictions)
print('Test accuracy: %.4f%%' % test_accuracy)
```
---
<a id='step4'></a>
## Step 4: Use a CNN to Classify Dog Breeds
To reduce training time without sacrificing accuracy, we show you how to train a CNN using transfer learning. In the following step, you will get a chance to use transfer learning to train your own CNN.
### Obtain Bottleneck Features
```
bottleneck_features = np.load('bottleneck_features/DogVGG16Data.npz')
train_VGG16 = bottleneck_features['train']
valid_VGG16 = bottleneck_features['valid']
test_VGG16 = bottleneck_features['test']
```
### Model Architecture
The model uses the the pre-trained VGG-16 model as a fixed feature extractor, where the last convolutional output of VGG-16 is fed as input to our model. We only add a global average pooling layer and a fully connected layer, where the latter contains one node for each dog category and is equipped with a softmax.
```
VGG16_model = Sequential()
VGG16_model.add(GlobalAveragePooling2D(input_shape=train_VGG16.shape[1:]))
VGG16_model.add(Dense(133, activation='softmax'))
VGG16_model.summary()
```
### Compile the Model
```
VGG16_model.compile(loss='categorical_crossentropy', optimizer='rmsprop', metrics=['accuracy'])
```
### Train the Model
```
checkpointer = ModelCheckpoint(filepath='saved_models/weights.best.VGG16.hdf5',
verbose=1, save_best_only=True)
VGG16_model.fit(train_VGG16, train_targets,
validation_data=(valid_VGG16, valid_targets),
epochs=20, batch_size=20, callbacks=[checkpointer], verbose=1)
```
### Load the Model with the Best Validation Loss
```
VGG16_model.load_weights('saved_models/weights.best.VGG16.hdf5')
```
### Test the Model
Now, we can use the CNN to test how well it identifies breed within our test dataset of dog images. We print the test accuracy below.
```
# get index of predicted dog breed for each image in test set
VGG16_predictions = [np.argmax(VGG16_model.predict(np.expand_dims(feature, axis=0))) for feature in test_VGG16]
# report test accuracy
test_accuracy = 100*np.sum(np.array(VGG16_predictions)==np.argmax(test_targets, axis=1))/len(VGG16_predictions)
print('Test accuracy: %.4f%%' % test_accuracy)
```
### Predict Dog Breed with the Model
```
from extract_bottleneck_features import *
def VGG16_predict_breed(img_path):
# extract bottleneck features
bottleneck_feature = extract_VGG16(path_to_tensor(img_path))
# obtain predicted vector
predicted_vector = VGG16_model.predict(bottleneck_feature)
# return dog breed that is predicted by the model
return dog_names[np.argmax(predicted_vector)]
```
---
<a id='step5'></a>
## Step 5: Create a CNN to Classify Dog Breeds (using Transfer Learning)
You will now use transfer learning to create a CNN that can identify dog breed from images. Your CNN must attain at least 60% accuracy on the test set.
In Step 4, we used transfer learning to create a CNN using VGG-16 bottleneck features. In this section, you must use the bottleneck features from a different pre-trained model. To make things easier for you, we have pre-computed the features for all of the networks that are currently available in Keras:
- [VGG-19](https://s3-us-west-1.amazonaws.com/udacity-aind/dog-project/DogVGG19Data.npz) bottleneck features
- [ResNet-50](https://s3-us-west-1.amazonaws.com/udacity-aind/dog-project/DogResnet50Data.npz) bottleneck features
- [Inception](https://s3-us-west-1.amazonaws.com/udacity-aind/dog-project/DogInceptionV3Data.npz) bottleneck features
- [Xception](https://s3-us-west-1.amazonaws.com/udacity-aind/dog-project/DogXceptionData.npz) bottleneck features
The files are encoded as such:
Dog{network}Data.npz
where `{network}`, in the above filename, can be one of `VGG19`, `Resnet50`, `InceptionV3`, or `Xception`. Pick one of the above architectures, download the corresponding bottleneck features, and store the downloaded file in the `bottleneck_features/` folder in the repository.
### (IMPLEMENTATION) Obtain Bottleneck Features
In the code block below, extract the bottleneck features corresponding to the train, test, and validation sets by running the following:
bottleneck_features = np.load('bottleneck_features/Dog{network}Data.npz')
train_{network} = bottleneck_features['train']
valid_{network} = bottleneck_features['valid']
test_{network} = bottleneck_features['test']
```
### TODO: Obtain bottleneck features from another pre-trained CNN.
bottleneck_features = np.load('bottleneck_features/DogResnet50Data.npz')
train_Resnet50 = bottleneck_features['train']
valid_Resnet50 = bottleneck_features['valid']
test_Resnet50 = bottleneck_features['test']
```
### (IMPLEMENTATION) Model Architecture
Create a CNN to classify dog breed. At the end of your code cell block, summarize the layers of your model by executing the line:
<your model's name>.summary()
__Question 5:__ Outline the steps you took to get to your final CNN architecture and your reasoning at each step. Describe why you think the architecture is suitable for the current problem.
__Answer:__
This time I wanted to be certain I took full advantage of transfer learning so I decided to leverage the current resnet weights and make that an input into the global average pooling layer. This way, I could make use of what the network already understood from previous training. Then I finally added the fully connected layer with an output of 133 units because there are 133 dognames. I also continued to use the softmax function
```
### TODO: Define your architecture.
Resnet50_model = Sequential()
### TODO: Define your architecture.
Resnet50_model.add(GlobalAveragePooling2D(input_shape=train_Resnet50.shape[1:]))
Resnet50_model.add(Dense(units= 133, activation = 'softmax'))
Resnet50_model.summary()
```
### (IMPLEMENTATION) Compile the Model
```
### TODO: Compile the model.
Resnet50_model.compile(loss='categorical_crossentropy', optimizer='rmsprop', metrics=['accuracy'])
```
### (IMPLEMENTATION) Train the Model
Train your model in the code cell below. Use model checkpointing to save the model that attains the best validation loss.
You are welcome to [augment the training data](https://blog.keras.io/building-powerful-image-classification-models-using-very-little-data.html), but this is not a requirement.
```
### TODO: Train the model
from keras.preprocessing.image import ImageDataGenerator
datagen = ImageDataGenerator(
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
train_datagen = ImageDataGenerator(rescale=1./255,
shear_range=0.2,
zoom_range = 0.2,
horizontal_flip = True)
test_datagen = ImageDataGenerator(rescale=1./255)
train_generator = train_datagen.flow_from_directory('dogImages/train',
target_size=(224,224),
batch_size= 20,
class_mode='categorical')
validation_generator = test_datagen.flow_from_directory('dogImages/valid',
target_size=(224,224),
batch_size= 20,
class_mode='categorical')
Resnet50_model.fit_generator(train_generator,
steps_per_epoch=100,
epochs=10,
validation_data=validation_generator,
validation_steps=100)
Resnet50_model.save_weights('saved_models/transfer_cnn_image_train.hdf5')
checkpointer = ModelCheckpoint(filepath='saved_models/weights.best.Resnet50.hdf5',
verbose=1, save_best_only=True)
Resnet50_model.fit(train_Resnet50, train_targets,
validation_data=(valid_Resnet50, valid_targets),
epochs=20, batch_size=20, callbacks=[checkpointer], verbose=1)
```
### (IMPLEMENTATION) Load the Model with the Best Validation Loss
```
### TODO: Load the model weights with the best validation loss.
Resnet50_model.load_weights('saved_models/weights.best.Resnet50.hdf5')
```
### (IMPLEMENTATION) Test the Model
Try out your model on the test dataset of dog images. Ensure that your test accuracy is greater than 60%.
```
### TODO: Calculate classification accuracy on the test dataset.
# get index of predicted dog breed for each image in test set
Resnet50_predictions = [np.argmax(Resnet50_model.predict(np.expand_dims(feature, axis=0))) for feature in test_Resnet50]
# report test accuracy
test_accuracy = 100*np.sum(np.array(Resnet50_predictions)==np.argmax(test_targets, axis=1))/len(Resnet50_predictions)
print('Test accuracy: %.4f%%' % test_accuracy)
```
### (IMPLEMENTATION) Predict Dog Breed with the Model
Write a function that takes an image path as input and returns the dog breed (`Affenpinscher`, `Afghan_hound`, etc) that is predicted by your model.
Similar to the analogous function in Step 5, your function should have three steps:
1. Extract the bottleneck features corresponding to the chosen CNN model.
2. Supply the bottleneck features as input to the model to return the predicted vector. Note that the argmax of this prediction vector gives the index of the predicted dog breed.
3. Use the `dog_names` array defined in Step 0 of this notebook to return the corresponding breed.
The functions to extract the bottleneck features can be found in `extract_bottleneck_features.py`, and they have been imported in an earlier code cell. To obtain the bottleneck features corresponding to your chosen CNN architecture, you need to use the function
extract_{network}
where `{network}`, in the above filename, should be one of `VGG19`, `Resnet50`, `InceptionV3`, or `Xception`.
```
### TODO: Write a function that takes a path to an image as input
### and returns the dog breed that is predicted by the model.
def resnet50_predict_breed(img_path):
bottleneck_feature = extract_Resnet50(path_to_tensor(img_path))
predicted_vector = Resnet50_model.predict(bottleneck_feature)
return dog_names[np.argmax(predicted_vector)]
```
---
<a id='step6'></a>
## Step 6: Write your Algorithm
Write an algorithm that accepts a file path to an image and first determines whether the image contains a human, dog, or neither. Then,
- if a __dog__ is detected in the image, return the predicted breed.
- if a __human__ is detected in the image, return the resembling dog breed.
- if __neither__ is detected in the image, provide output that indicates an error.
You are welcome to write your own functions for detecting humans and dogs in images, but feel free to use the `face_detector` and `dog_detector` functions developed above. You are __required__ to use your CNN from Step 5 to predict dog breed.
Some sample output for our algorithm is provided below, but feel free to design your own user experience!

### (IMPLEMENTATION) Write your Algorithm
```
### TODO: Write your algorithm.
### Feel free to use as many code cells as needed.
def test_with(file_name):
""" takes a file name and folder name and makes a prediction"""
if face_detector(file_name):
breed = resnet50_predict_breed(file_name)
print("| IMAGE | TYPE_IN_IMAGE | BREED |")
print("| {0} | Human | {1} |".format(file_name, breed))
elif dog_detector(file_name):
breed = resnet50_predict_breed(file_name)
print("| IMAGE | TYPE_IN_IMAGE | BREED |")
print("| {0} | DOG | {1} |".format(file_name, breed))
else:
print("| IMAGE | TYPE_IN_IMAGE | BREED |")
print("| {0} | Neither Human or Dog | {1} |".format(file_name, "N/A"))
```
---
<a id='step7'></a>
## Step 7: Test Your Algorithm
In this section, you will take your new algorithm for a spin! What kind of dog does the algorithm think that __you__ look like? If you have a dog, does it predict your dog's breed accurately? If you have a cat, does it mistakenly think that your cat is a dog?
### (IMPLEMENTATION) Test Your Algorithm on Sample Images!
Test your algorithm at least six images on your computer. Feel free to use any images you like. Use at least two human and two dog images.
__Question 6:__ Is the output better than you expected :) ? Or worse :( ? Provide at least three possible points of improvement for your algorithm.
__Answer:__
Answer to this question is in the following code cell
```
## TODO: Execute your algorithm from Step 6 on
## at least 6 images on your computer.
## Feel free to use as many code cells as needed.
file_names = ['test_1', 'test_2', 'test_3', 'test_4', 'test_5', 'test_6']
file_names = list(map(lambda f: "algo_test/" + f + ".jpg", file_names))
# image_tensors = [path_to_tensor(file).astype('float32')/255 for file in file_names]
# test_data = zip(file_names, image_tensors)
for file_name in file_names:
test_with(file_name)
```
### Explanation of Testing of the Algorithm
In order to get a good idea of how this algorithm was working I decided to add in some images that I thought would be kind of tricky for the algorithm. All the test images can be found in the `/algo_test` directory but a brief break down of the actual images and my analysis can be found below.
| Image | Expected | Actual |Description |
|:-------------:|:------------------:|-------------:|:------------------------------------: |
| test_1 | human | human | Was actually a picture of Lavar Ball |
| test_2 | human | human | Was a picture of me |
| test_3 | Dog | Dog | Was a picture of a puppy |
| test_4 | Neither | Neither | Was a picture of a basketball |
| test_5 | Dog | Dog | Was a picture of a tibetan_mastiff |
| test_6 | Dog | Dog & human | picture of a tibetan mastiff and its owner|
This was interesting to see the results at the end of the experiement. A brief description of my views are below...
- First, The algorithm correctly detected that the picture in `test_1.jpg` was a human. What was more interesting here was that it chose a `Can_corso` as the breed of dog that `Lavar Ball` who was depicted in the image, looked similar too. This was not at all expected.
- In `test_2.jpg` was an image of myself which was fed into the algorithm. The algorithm correctly detected my face as human, but what was rather offensive is the algorithm said that the breed that I looked like was a `Dogue_de_bordeaux`. These dogs are extremely unattractive, so I guess this algorithm shows what it thinks of my face.
- In `test_3.jpg` I found a random picture of a puppy which I believe is a Jack Russle tearrier puppy. The algorithm thought that this image was a `Beagle` which I could see how it chose that as both of those dog breeds look similar as puppies.
- The fourth image `test_4.jpg` was a picture of a basketball. I added this image into the test folder to see if the algorithm would figure out the difference. The output from the network was correct as it predicted that Neither dog nor human was found in the photo, and thus, no dog breed was selected. This is what I was expected from it.
- `test_5.jpg` was meant as tricky test as it was a picture of a tibetan Mastiff, which looked really close to a bear. I wanted to see if the algorithm was good enough to decipher this image as it could have fulled a person as it did me the first time I took a glance at it. Sure enough, the algorithm noticed that it was a dog, and classified it as a `Tibetan Mastiff` which is correct.
- `test_6.jpg` was designed to be a trick image. I found an image with a `Tibetan Mastiff` as well as a human in the photo together. I wanted to see what the algorithm would do. It located the dog in the photo and classified it correctly as a tibetan mastiff, however, It didn't seem to pay attention to the fact that a human was sitting right next to the dog in the photo. I found this really interesting as I didn't expect this to be the behavior. I thought that it would detect the human first and choose a dog breed the human was similar to.
| github_jupyter |
Copyright 2016 Google Inc. All Rights Reserved.
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software
distributed under the License is distributed on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and
limitations under the License.
<h1 id="tocheading">Table of Contents</h1>
<div id="toc"></div>
```
%%javascript
// From https://github.com/kmahelona/ipython_notebook_goodies
$.getScript('https://kmahelona.github.io/ipython_notebook_goodies/ipython_notebook_toc.js')
```
# Basics
There's lots of guides out there on decorators ([this](http://thecodeship.com/patterns/guide-to-python-function-decorators/) one is good), but I was never really sure when I would need to use decorators. Hopefully this will help motivate them a little more. Here I hope to show you:
* When decorators might come in handy
* How to write one
* How to generalize using `*args` and `**kwargs` sorcery.
You should read this if:
* You've heard of decorators and want to know more about them, and/or
* You want to know what `*args` and `**kwargs` mean.
If you're here just for `*args` and `**kwargs`, start reading [here](#args).
## Motivation
Let's say you're defining methods on numbers:
```
def add(n1, n2):
return n1 + n2
def multiply(n1, n2):
return n1 * n2
def exponentiate(n1, n2):
"""Raise n1 to the power of n2"""
import math
return math.pow(n1, n2)
```
Well, we only want these functions to work if both inputs are numbers. So we could do:
```
def is_number(n):
"""Return True iff n is a number."""
# A number can always be converted to a float
try:
float(n)
return True
except ValueError:
return False
def add(n1, n2):
if not (is_number(n1) and is_number(n2)):
print("Arguments must be numbers!")
return
return n1 + n2
def multiply(n1, n2):
if not (is_number(n1) and is_number(n2)):
print("Arguments must be numbers!")
return
return n1 * n2
def exponentiate(n1, n2):
"""Raise n1 to the power of n2"""
if not (is_number(n1) and is_number(n2)):
print("Arguments must be numbers!")
return
import math
return math.pow(n1, n2)
```
But this is yucky: we had to copy and paste code. This should always make you sad! For example, what if you wanted to change the message slightly? Or to return an error instead? You'd have to change it everywhere it appears...
We want the copy & pasted code to live in just one place, so any changes just go there (DRY code: Don't Repeat Yourself). So let's **refactor**.
```
def validate_two_arguments(n1, n2):
"""
Returns True if n1 and n2 are both numbers.
"""
if not (is_number(n1) and is_number(n2)):
return False
return True
def add(n1, n2):
if validate_two_arguments(n1, n2):
return n1 + n2
def multiply(n1, n2):
if validate_two_arguments(n1, n2):
return n1 * n2
def exponentiate(n1, n2):
"""Raise n1 to the power of n2"""
if validate_two_arguments(n1, n2):
import math
return math.pow(n1, n2)
```
This is definitely better. But there's still some repeated logic. Like, what if we want to return an error if we don't get numbers, or print something before running the code? We'd still have to make the changes in multiple places. The code isn't DRY.
## Basic decorators
We can refactor further with the **decorator pattern**.
We want to write something that looks like
@decorator
def add(n1, n2):
return n1 + n2
so that all the logic about validating `n1` and `n2` lives in one place, and the functions just do what we want them to do.
Since the @ syntax just means `add = decorator(add)`, we know the decorator needs to take a function as an argument, and it needs to return a function. (This should be confusing at first. Functions returning functions are scary, but think about it until that doesn't seem outlandish to you.)
This returned function should act the same way as `add`, so it should take two arguments. And within this returned function, we want to first check that the arguments are numbers. If they are, we want to call the original function that we decorated (in this case, `add`). If not, we don't want to do anything. Here's what that looks like (there's a lot here, so use the comments to understand what's happening):
```
# The decorator: takes a function.
def validate_arguments(func):
# The decorator will be returning wrapped_func, a function that has the
# same signature as add, multiply, etc.
def wrapped_func(n1, n2):
# If we don't have two numbers, we don't want to run the function.
# Best practice ("be explicit") is to raise an error here
# instead of just returning None.
if not validate_two_arguments(n1, n2):
raise Exception("Arguments must be numbers!")
# We've passed our checks, so we can call the function with the passed in arguments.
# If you like, think of this as
# result = func(n1, n2)
# return result
# to distinguish it from the outer return where we're returning a function.
return func(n1, n2)
# This is where we return the function that has the same signature.
return wrapped_func
@validate_arguments
def add(n1, n2):
return n1 + n2
# Don't forget, the @ syntax just means
# add = validate_decorator(add)
print(add(1, 3))
try:
add(2, 'hi')
except Exception as e:
print("Caught Exception: {}".format(e))
```
This pattern is nice because we've even refactored out all the validation logic (even the "if blah then blah" part) into the decorator.
## Generalizing with \*args and \**kwargs
What if we want to validate a function that has a different number of arguments?
```
@validate_arguments # Won't work!
def add3(n1, n2, n3):
return n1 + n2 + n3
add3(1, 2, 3)
```
We can't decorate this because the wrapped function expects 2 arguments.
Here's where we use the `*` symbol. I'll write out the code so you can see how it looks, and we'll look at what `*args` is doing below.
```
# The decorator: takes a function.
def validate_arguments(func):
# Note the *args! Think of this as representing "as many arguments as you want".
# So this function will take an arbitrary number of arguments.
def wrapped_func(*args):
# We just want to apply the check to each argument.
for arg in args:
if not is_number(arg):
raise Exception("Arguments must be numbers!")
# We also want to make sure there's at least two arguments.
if len(args) < 2:
raise Exception("Must specify at least 2 arguments!")
# We've passed our checks, so we can call the function with the
# passed-in arguments.
# Right now, args is a tuple of all the different arguments passed in
# (more explanation below), so we want to expand them back out when
# calling the function.
return func(*args)
return wrapped_func
@validate_arguments # This works
def add3(n1, n2, n3):
return n1 + n2 + n3
add3(1, 2, 3)
@validate_arguments # And so does this
def addn(*args):
"""Add an arbitrary number of numbers together"""
cumu = 0
for arg in args:
cumu += arg
return cumu
print(addn(1, 2, 3, 4, 5))
# range(n) gives a list, so we expand the list into positional arguments...
print(addn(*range(10)))
```
### <a id='args'>`*args`</a>
What is this `*` nonsense?
You've probably seen `*args` and `**kwargs` in documentation before. Here's what they mean:
* When calling a function, `*` **expands an iterable** into **positional arguments**.
* Terminology note: in a call like `bing(1, 'hi', name='fig')`, `1` is the first positional argument, `'hi'` is the second positional argument, and there's a keyword argument `'name'` with the value `'fig'`.
* When defining a signature, `*args` represents an **arbitrary number of positional arguments**.
```
def foo(*args):
print("foo args: {}".format(args))
print("foo args type: {}".format(type(args)))
# So foo can take an arbitrary number of arguments
print("First call:")
foo(1, 2, 'a', 3, True)
# Which can be written using the * syntax to expand an iterable
print("\nSecond call:")
l = [1, 2, 'a', 3, True]
foo(*l)
```
### Back to the decorator
(If you're just here for \*args and \*\*kwargs, skip down to [here](#kwargs))
So let's look at the decorator code again, minus the comments:
def validate_decorator(func):
def wrapped_func(*args):
for arg in args:
if not is_number(arg):
print("arguments must be numbers!")
return
return func(*args)
return wrapped_func
* `def wrapped_func(*args)` says that `wrapped_func` can take an arbitrary number of arguments.
* Within `wrapped_func`, we interact with `args` as a tuple containing all the (positional) arguments passed in.
* If all the arguments are numbers, we call `func`, the function we decorated, by **expanding** the `args` tuple back out into positional arguments: `func(*args)`.
* Finally the decorator needs to return a function (remember that the `@` syntax is just sugar for `add = decorator(add)`.
Congrats, you now understand decorators! You can do tons of other stuff with them, but hopefully now you're equipped to read the other guides online.
---
### <a id='kwargs'>As for `**kwargs`:</a>
* When calling a function, `**` **expands a dict** into **keyword arguments**.
* When defining a signature, `**kwargs` represents an **arbitrary number of keyword arguments**.
```
def bar(**kwargs):
print("bar kwargs: {}".format(kwargs))
# bar takes an arbitrary number of keyword arguments
print("First call:")
bar(location='US-PAO', ldap='awan', age=None)
# Which can also be written using the ** syntax to expand a dict
print("\nSecond call:")
d = {'location': 'US-PAO', 'ldap': 'awan', 'age': None}
bar(**d)
```
And in case your head doesn't hurt yet, we can do both together:
```
def baz(*args, **kwargs):
print("baz args: {}. kwargs: {}".format(args, kwargs))
# Calling baz with a mixture of positional and keyword arguments
print("First call:")
baz(1, 3, 'hi', name='Joe', age=37, occupation='Engineer')
# Which is the same as
print("\nSecond call:")
l = [1, 3, 'hi']
d = {'name': 'Joe', 'age': 37, 'occupation': 'Engineer'}
baz(*l, **d)
```
---
# Advanced decorators
This section will introduce some of the many other useful ways you can use decorators. We'll talk about
* Passing arguments into decorators
* `functools.wraps`
* Returning a different function
* Decorators and objects.
Use the [table of contents](#toc) at the top to make it easier to look around.
## Decorators with arguments
A common thing to want to do is to do some kind of configuration in a decorator. For example, let's say we want to define a `divide_n` method, and to make it easy to use we want to hide the existence of integer division. Let's define a decorator that converts arguments into floats.
```
def convert_arguments(func):
"""
Convert func arguments to floats.
"""
# Introducing the leading underscore: (weakly) marks a private
# method/property that should not be accessed outside the defining
# scope. Look up PEP 8 for more.
def _wrapped_func(*args):
new_args = [float(arg) for arg in args]
return func(*new_args)
return _wrapped_func
@convert_arguments
@validate_arguments
def divide_n(*args):
cumu = args[0]
for arg in args[1:]:
cumu = cumu / arg
return cumu
# The user doesn't need to think about integer division!
divide_n(103, 2, 8)
```
But now let's say we want to define a `divide_n_as_integers` function. We could write a new decorator, or we could alter our decorator so that we can specify what we want to convert the arguments to. Let's try the latter.
(For you smart alecks out there: yes you could use the `//` operator, but you'd still have to replicate the logic in `divide_n`. Nice try.)
```
def convert_arguments_to(to_type=float):
"""
Convert arguments to the given to_type by casting them.
"""
def _wrapper(func):
def _wrapped_func(*args):
new_args = [to_type(arg) for arg in args]
return func(*new_args)
return _wrapped_func
return _wrapper
@validate_arguments
def divide_n(*args):
cumu = args[0]
for arg in args[1:]:
cumu = cumu / arg
return cumu
@convert_arguments_to(to_type=int)
def divide_n_as_integers(*args):
return divide_n(*args)
@convert_arguments_to(to_type=float)
def divide_n_as_float(*args):
return divide_n(*args)
print(divide_n_as_float(7, 3))
print(divide_n_as_integers(7, 3))
```
Did you notice the tricky thing about creating a decorator that takes arguments? **We had to create a function to "return a decorator".** The outermost function, `convert_arguments_to`, returns a function that takes a function, which is what we've been calling a "decorator".
To think about why this is necessary, let's start from the form that we wanted to write, and unpack from there. We wanted to be able to do:
@decorator(decorator_arg)
def myfunc(*func_args):
pass
Unpacking the syntactic sugar gives us
def myfunc(*func_args):
pass
myfunc = decorator(decorator_arg)(myfunc)
Written this way, it should immediately be clear that `decorator(decorator_arg)` **returns a function that takes a function**.
So that's how you write a decorator that takes an argument: it actually has to be a function that takes your decorator arguments, and returns a function that takes a function.
## functools.wraps
If you've played around with the examples above, you might've seen that the name of the wrapped function changes after you apply a decorator... And perhaps more importantly, the docstring of the wrapped function changes too (this is important for when generating documentation, e.g. with Sphinx).
```
@validate_arguments
def foo(*args):
"""foo frobs bar"""
pass
print(foo.__name__)
print(foo.__doc__)
```
[`functools.wraps`](https://docs.python.org/2/library/functools.html#functools.wraps) solves this problem. Use it as follows:
```
from functools import wraps
def better_validate_arguments(func):
@wraps(func)
def wrapped_func(*args):
for arg in args:
if not is_number(arg):
raise Exception("Arguments must be numbers!")
if len(args) < 2:
raise Exception("Must specify at least 2 arguments!")
return func(*args)
return wrapped_func
@better_validate_arguments
def bar(*args):
"""bar frobs foo"""
pass
print(bar.__name__)
print(bar.__doc__)
```
Think of the `@wraps` decorator making it so that `wrapped_func` knows what function it originally wrapped.
## Returning a different function
Decorators don't even have to return the function that's passed in. You can have some fun with this...
```
def jedi_mind_trick(func):
def _jedi_func():
return "Not the droid you're looking for"
return _jedi_func
@jedi_mind_trick
def get_droid():
return "Found the droid!"
get_droid()
```
But more seriously, this can be useful for things like
* Authentication: you don't want to return the function if the user isn't recognized, instead redirecting to a login page (e.g. you could check an environment variable)
* Disabling test methods when deployed to a production environment
This is also how `@unittest.skip` works, if you've ever used it to skip functions that weren't ready for testing or couldn't be tested on a particular operating system.
## Objects
### Decorators that alter "self"
### Decorating a class
| github_jupyter |
(Source: http://www.scipy-lectures.org/packages/scikit-learn/index.html#basic-principles-of-machine-learning-with-scikit-learn)
```
%matplotlib notebook
```
## Estimators
Every algorithm is exposed in scikit-learn via an ‘’Estimator’’ object. For instance a linear regression is: `sklearn.linear_model.LinearRegression`
```
from sklearn.linear_model import LinearRegression
model = LinearRegression(normalize=True)
print(model.normalize)
print(model)
import numpy as np
x = np.array([0, 1, 2])
y = np.array([0, 1, 2])
X = x[:, np.newaxis] # The input data for sklearn is 2D: (samples == 3 x features == 1)
X
model.fit(X, y)
model.coef_
```
## Supervised Learning: Classifcation and Regression
```
from sklearn import neighbors, datasets
iris = datasets.load_iris()
X, y = iris.data, iris.target
knn = neighbors.KNeighborsClassifier(n_neighbors=1)
knn.fit(X, y)
```
> What kind of iris has 3cm x 5cm sepal and 4cm x 2cm petal?
```
print(iris.target_names[knn.predict([[3, 5, 4, 2]])])
from matplotlib import pyplot as plt
from matplotlib.colors import ListedColormap
# Create color maps for 3-class classification problem, as with iris
cmap_light = ListedColormap(['#FFAAAA', '#AAFFAA', '#AAAAFF'])
cmap_bold = ListedColormap(['#FF0000', '#00FF00', '#0000FF'])
iris = datasets.load_iris()
X = iris.data[:, :2] # we only take the first two features. We could avoid this by using a 2D dataset
y = iris.target
number_of_neighbors = [1, 3]
plt.figure(figsize=(9, 4))
for idx, neighbor_number in enumerate(number_of_neighbors):
knn = neighbors.KNeighborsClassifier(n_neighbors=neighbor_number)
knn.fit(X, y)
x_min, x_max = X[:, 0].min() - .1, X[:, 0].max() + .1
y_min, y_max = X[:, 1].min() - .1, X[:, 1].max() + .1
xx, yy = np.meshgrid(np.linspace(x_min, x_max, 100),
np.linspace(y_min, y_max, 100))
Z = knn.predict(np.c_[xx.ravel(), yy.ravel()])
# Put the result into a color plot
Z = Z.reshape(xx.shape)
plt.subplot(1,2,idx+1)
plt.pcolormesh(xx, yy, Z, cmap=cmap_light)
if idx == 0:
plt.xlabel('sepal length (cm)')
plt.ylabel('sepal width (cm)')
plt.axis('tight')
plt.title('k = {0}'.format(neighbor_number))
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=cmap_bold)
plt.show()
```
## Estimator interface summary
http://www.scipy-lectures.org/packages/scikit-learn/index.html#a-recap-on-scikit-learn-s-estimator-interface
## Regularization
Train errors Suppose you are using a 1-nearest neighbor estimator. How many errors do you expect on your train set?
- Train set error is not a good measurement of prediction performance. You need to leave out a test set.
- In general, we should accept errors on the train set.
**An example of regularization** The core idea behind regularization is that we are going to prefer models that are simpler, for a certain definition of ‘’simpler’‘, even if they lead to more errors on the train set.
As an example, let’s generate with a 9th order polynomial, with noise:
```
from sklearn import linear_model
# Create color maps for 3-class classification problem, as with iris
cmap_light = ListedColormap(['#FFAAAA', '#AAFFAA', '#AAAAFF'])
cmap_bold = ListedColormap(['#FF0000', '#00FF00', '#0000FF'])
rng = np.random.RandomState(0)
x = 2*rng.rand(100) - 1
f = lambda t: 1.2 * t**2 + .1 * t**3 - .4 * t **5 - .5 * t ** 9
y = f(x) + .4 * rng.normal(size=100)
x_test = np.linspace(-1, 1, 100)
plt.figure(figsize=(6, 4))
plt.scatter(x, y, s=4)
plt.figure(figsize=(6, 4))
plt.scatter(x, y, s=4)
X = np.array([x**i for i in range(5)]).T
X_test = np.array([x_test**i for i in range(5)]).T
regr = linear_model.LinearRegression()
regr.fit(X, y)
plt.plot(x_test, regr.predict(X_test), label='4th order')
X = np.array([x**i for i in range(10)]).T
X_test = np.array([x_test**i for i in range(10)]).T
regr = linear_model.LinearRegression()
regr.fit(X, y)
plt.plot(x_test, regr.predict(X_test), label='9th order')
plt.plot(x_test, f(x_test), label="truth")
plt.legend(loc='best')
plt.axis('tight')
plt.title('Fitting a 4th and a 9th order polynomial')
plt.show()
```
Similar example for classification: http://www.scipy-lectures.org/packages/scikit-learn/index.html#simple-versus-complex-models-for-classification
| github_jupyter |
# Building and training a mutli-layer network with Keras
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from keras.models import Sequential
from keras.layers import Dense, Activation
from keras.optimizers import SGD
%matplotlib inline
```
## Classifying Iris versicolor
Let us know try a slightly different problem: identifying the species *versicolor* instead of *setosa*.
This is a more challenging problem, because the species *versicolor* is very close to the related species *virginica*, as shown on the data below.
```
# Load data
df = pd.read_csv('./data/versicolor/train.csv')
X = df[['petal length (cm)', 'petal width (cm)']].values
y = df['versicolor'].values
def plot_keras_model( model=None ):
"Plot the Keras model, along with data"
plt.clf()
# Calculate the probability on a mesh
if model is not None:
petal_width_mesh, petal_length_mesh = \
np.meshgrid( np.linspace(0,3,100), np.linspace(0,8,100) )
petal_width_mesh = petal_width_mesh.flatten()
petal_length_mesh = petal_length_mesh.flatten()
p = model.predict( np.stack( (petal_length_mesh, petal_width_mesh), axis=1 ) )
p = p.reshape((100,100))
# Plot the probability on the mesh
plt.imshow( p.T, extent=[0,8,0,3], origin='lower',
vmin=0, vmax=1, cmap='RdBu', aspect='auto', alpha=0.7 )
# Plot the data points
plt.scatter( df['petal length (cm)'], df['petal width (cm)'], c=df['versicolor'], cmap='RdBu')
plt.xlabel('petal length (cm)')
plt.ylabel('petal width (cm)')
cb = plt.colorbar()
cb.set_label('versicolor')
plot_keras_model()
```
## Single layer network
Let us see how a single-layer neural network performs in this case. Here are building exactly the same kind of network as we did in the previous notebook.
```
# Build the model
single_layer_model = Sequential()
single_layer_model.add( Dense( output_dim=1, input_dim=2 ) )
single_layer_model.add( Activation( 'sigmoid' ) )
# Prepare the model for training
single_layer_model.compile(loss='binary_crossentropy', optimizer=SGD(lr=0.1), metrics=['accuracy'])
# Train the network
single_layer_model.fit( X, y, batch_size=16, nb_epoch=1000, verbose=0 )
plot_keras_model( model=single_layer_model )
```
The network is unable to make the correct prediction, even after 1000 epochs of training.
This because, as we saw when tuning the weights by hand, a single-layer network is only capable of producing a single linear boundary between two areas of the plane. For a more complicated model, we need several layers.
# Two layer network
A two layer network looks like this:

where the number of units in the intermediate layer (4 here) is a parameter that the user needs to choose.
```
# Build the model: pick 8 units in the intermediate layer
two_layer_model = Sequential()
two_layer_model.add( Dense( output_dim=8, input_dim=2 ) )
two_layer_model.add( Activation( 'sigmoid' ) )
two_layer_model.add( Dense( output_dim=1, input_dim=8 ) )
two_layer_model.add( Activation( 'sigmoid' ) )
# Compile the model
two_layer_model.compile(loss='binary_crossentropy', optimizer=SGD(lr=0.1), metrics=['accuracy'])
# Train it
two_layer_model.fit( X, y, batch_size=16, nb_epoch=1000, verbose=0 )
plot_keras_model( model=two_layer_model )
```
## A few more remarks on Keras and neural networks
Keras allows to build and train a number of neural network architectures:
- fully-connected networks
- convolutional networks
- recurrent networks
The corresponding code with the Keras interface is much less verbose than directly with the Tensorflow interfact (but also less flexible).
Keras still requires the user to make many educated guesses:
- Structure of the network (architecture, number of layers, number of nodes in each layers, activation functions)
- Training parameters (loss function, optimizer and learning rate, batch size, number of epochs)
| github_jupyter |
# Chicago crime dataset analysis
---
This notebook is a Spark and Python learner's to perform data analysis on some real-world data set.
In this notebook, I am capriciously using Spark, Pandas, Matplotlib, Seaborn without any meaningful distinction of purpose. The point is:
* Perform data reading, transforming, and querying using Apache Spark
* Visualize using existing Python libraries. Matplotlib will remain when I know to do with it all that I'm currenly using Seaborn for.
* Where interoperation between Spark and Matplotlib is a hindrance, I use Pandas and Numpy.
This will be evolutionary and I hope that a few weeks from now, it will look much better.
---
# How to run this and what to run it on?
---
I wrote this on Apache Spark 2.3.0. The entire notebook was executed on a single machine using the `pyspark` shell without problems.
Here are some important parameters:
* Executor count: 4
* Executor Memory: 4G
* Driver Memory: 8G
This may not be necessary, but some data frames are being cached and performance degrades remarkably when the percentage of cached RDDs drops.
So, the command with which the notebook was launched is:
```
pyspark --driver-memory 8g --executor-memory 4g --master local[4]
```
---
## Get Imports out of the way
Spark imports
```
## spark imports
from pyspark.sql import Row, SparkSession
from pyspark.sql.functions import *
```
Standard python data analysis imports
```
## standard imports
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
```
Jupyter visualization options
```
%matplotlib inline
#Not too sure the following 2 work. This is a TODO
sns.set_color_codes("pastel")
plt.rcParams["figure.figsize"] = [20, 8]
```
---
## Dataset information
First, clean the headers. This is just for nice presentation of headers. Having valid data frame field names that conform to Python identifier rules is good. I personally prefer that as it's always helpful to use the dot notation in many places.
These are the column names, as pulled from the Kaggle dataset page [here](https://www.kaggle.com/djonafegnem/chicago-crime-data-analysis)
```
content_cols = '''
ID - Unique identifier for the record.
Case Number - The Chicago Police Department RD Number (Records Division Number), which is unique to the incident.
Date - Date when the incident occurred. this is sometimes a best estimate.
Block - The partially redacted address where the incident occurred, placing it on the same block as the actual address.
IUCR - The Illinois Unifrom Crime Reporting code. This is directly linked to the Primary Type and Description. See the list of IUCR codes at https://data.cityofchicago.org/d/c7ck-438e.
Primary Type - The primary description of the IUCR code.
Description - The secondary description of the IUCR code, a subcategory of the primary description.
Location Description - Description of the location where the incident occurred.
Arrest - Indicates whether an arrest was made.
Domestic - Indicates whether the incident was domestic-related as defined by the Illinois Domestic Violence Act.
Beat - Indicates the beat where the incident occurred. A beat is the smallest police geographic area – each beat has a dedicated police beat car. Three to five beats make up a police sector, and three sectors make up a police district. The Chicago Police Department has 22 police districts. See the beats at https://data.cityofchicago.org/d/aerh-rz74.
District - Indicates the police district where the incident occurred. See the districts at https://data.cityofchicago.org/d/fthy-xz3r.
Ward - The ward (City Council district) where the incident occurred. See the wards at https://data.cityofchicago.org/d/sp34-6z76.
Community Area - Indicates the community area where the incident occurred. Chicago has 77 community areas. See the community areas at https://data.cityofchicago.org/d/cauq-8yn6.
FBI Code - Indicates the crime classification as outlined in the FBI's National Incident-Based Reporting System (NIBRS). See the Chicago Police Department listing of these classifications at http://gis.chicagopolice.org/clearmap_crime_sums/crime_types.html.
X Coordinate - The x coordinate of the location where the incident occurred in State Plane Illinois East NAD 1983 projection. This location is shifted from the actual location for partial redaction but falls on the same block.
Y Coordinate - The y coordinate of the location where the incident occurred in State Plane Illinois East NAD 1983 projection. This location is shifted from the actual location for partial redaction but falls on the same block.
Year - Year the incident occurred.
Updated On - Date and time the record was last updated.
Latitude - The latitude of the location where the incident occurred. This location is shifted from the actual location for partial redaction but falls on the same block.
Longitude - The longitude of the location where the incident occurred. This location is shifted from the actual location for partial redaction but falls on the same block.
Location - The location where the incident occurred in a format that allows for creation of maps and other geographic operations on this data portal. This location is shifted from the actual location for partial redaction but falls on the same block.'''
def entry_dic(line):
"""
Convert a header - description line into a single dictionary that holds the original header as 'title', a corresponding field name as 'header', and a description.
"""
pair = line.split(' - ')
return {'title': pair[0], 'description': pair[1], 'header': pair[0].lower().replace(' ', '_')}
```
Turn a list of headers to a list of dictionaries produced by the above function
```
header_dics = list(map(entry_dic, list(filter(lambda l: l != '', content_cols.split('\n')))))
header_dics[:2]
```
### Environment
The data set is expected to be located at `../data/from-kaggle/`. The dataset downloaded from the page mentioned above comes in multiple .csv files. I extracted those files into the directory (see below `ls` output).
It shouldn't matter where the data comes from or how many files it's broken up into. As long as the pattern matches, that should work.
```
!ls -lh ../data/from-kaggle/
```
---
## Reading the data
Using the Spark's csv reader to parse the files. It processes multiple files and returns a single data frame:
```
df = spark.read.csv('../data/from-kaggle/*.csv', inferSchema=True, header=True)
# Caching this data frame as it's going to be read over and over again
df = df.cache()
df.printSchema()
```
---
**Rename columns so that we can use the dot notation**
```
for h in header_dics:
df = df.withColumnRenamed(h['title'], h['header'])
```
There's a row that's been identified as having no value for `location_description`. We will just get rid of it right now...
Initially implemented using a `rdd.filter` call. Although, alternatively, one can use the Dataset.filter function
```
df = df.rdd.filter(lambda rec: rec.arrest.find('Location Description') < 0).toDF().cache()
```
Taking an initial look at the content of the data frame:
```
df.show(n=3, truncate=False)
```
---
## Looking at the data
```
# crime types
crime_type_groups = df.groupBy('primary_type').count()
crime_type_counts = crime_type_groups.orderBy('count', ascending=False)
```
Total record count:
```
print(df.count())
```
How many features do we have
```
df.columns
#Let's see the schema of the data frame:
df.printSchema()
```
### Crime types
These are the top 20 most frequent crime types:
```
crime_type_counts.show(truncate=False)
```
I like doing some things by hand, but it's clear that here, one can obtain a Pandas data frame by calling:
``
crime_type_counts.toPandas()
``
```
counts_pddf = pd.DataFrame(crime_type_counts.rdd.map(lambda l: l.asDict()).collect())
counts_pddf.head(10)
plt.rcParams["figure.figsize"] = [20, 8]
sns.set(style="whitegrid")
sns.set_color_codes("pastel")
#sns.despine(left=True, bottom=True)
type_graph = sns.barplot(x='count', y='primary_type', data=counts_pddf)
type_graph.set(ylabel="Primary Type", xlabel="Crimes Record Count")
```
### Recorded Date
```
import datetime
from pyspark.sql.functions import *
df.select(min('date').alias('first_record_date'), max('date').alias('latest_record_date')).show(truncate=False)
```
So it seems that the dataset we're dealing with comprises records from **2001-01-01** to **2016-12-31**
---
Converting dates to a timestamp type. As seen in the schema output above, the `date` field is of `string` type, which won't be very helpful
The format specifier that seems valid for date like '02/23/2006 09:06:22 PM' is **`'MM/dd/yyyy hh:mm:ss a'`**
```
df = df.withColumn('date_time', to_timestamp('date', 'MM/dd/yyyy hh:mm:ss a'))\
.withColumn('month', trunc('date_time', 'YYYY')) #adding a month column to be able to view stats on a monthly basis
df.select(['date','date_time', 'month'])\
.show(n=2, truncate=False)
```
### Primary type and arrest
```
# crime types and arrest over years:
type_arrest_date = df.groupBy(['arrest', 'month'])\
.count()\
.orderBy(['month', 'count'], ascending=[True, False])
print()
type_arrest_date.show(3, truncate=False)
# A small detour to learn/see how datetime works
import datetime
datetime.datetime.now()
datetime.datetime.strftime(datetime.datetime.now(), '%H')
# A pandas data frame of the collected dictionary version of the date-grouped DF above
type_arrest_pddf = pd.DataFrame(type_arrest_date.rdd.map(lambda l: l.asDict()).collect())
```
Convert date/time values to Pandas type
*A lot of this is unnecessary, but that's called fiddling...*
```
type_arrest_pddf['yearpd'] = type_arrest_pddf['month'].apply(lambda dt: datetime.datetime.strftime(pd.Timestamp(dt), '%Y'))
type_arrest_pddf['arrest'] = type_arrest_pddf['arrest'].apply(lambda l: l=='True')
type_arrest_pddf.head(5)
```
### How have arrests evolved over the 16 years?
```
# Data for plotting
t = type_arrest_pddf['count'] - 20 # np.arange(0.0, 2.0, 0.01)
s = type_arrest_pddf['month']
arrested = type_arrest_pddf[type_arrest_pddf['arrest'] == True]
not_arrested = type_arrest_pddf[type_arrest_pddf['arrest'] == False]
# Note that using plt.subplots below is equivalent to using
# fig = plt.figure() and then ax = fig.add_subplot(111)
fig, ax = plt.subplots()
ax.plot(arrested['month'], arrested['count'], label='Arrested')
ax.plot(not_arrested['month'], not_arrested['count'], label='Not Arrested')
ax.set(xlabel='Year - 2001-2017', ylabel='Total records',
title='Year-on-year crime records')
ax.grid(b=True, which='both', axis='y')
ax.legend()
```
It looks like the relative distance between arrests and non-arrests has remained constant..
### What time of the day are ciminal the busiest?
```
# Extract the "hour" field from the date into a separate column called "hour"
df_hour = df.withColumn('hour', hour(df['date_time']))
# Derive a data frame with crime counts per hour of the day:
hourly_count = df_hour.groupBy(['primary_type', 'hour']).count().cache()
hourly_total_count = hourly_count.groupBy('hour').sum('count')
hourly_count_pddf = pd.DataFrame(hourly_total_count.select(hourly_total_count['hour'], hourly_total_count['sum(count)'].alias('count'))\
.rdd.map(lambda l: l.asDict())\
.collect())
hourly_count_pddf = hourly_count_pddf.sort_values(by='hour')
```
Seems that 18-22 are the worst hours...
```
fig, ax = plt.subplots()
ax.plot(hourly_count_pddf['hour'], hourly_count_pddf['count'], label='Hourly Count')
ax.set(xlabel='Hour of Day', ylabel='Total records',
title='Overall hourly crime numbers')
ax.grid(b=True, which='both', axis='y')
ax.legend()
```
### Types of locations with crime entries
```
# Number of types of location recorded in the dataset
df.select('location_description').distinct().count()
```
What are the top 10 places where crime occurred?
```
df.groupBy(['location_description']).count().orderBy('count', ascending=False).show(10)
```
Can we see how the top crime locations are affected over the day?
```
#Being on the street and at home
street_home_hour = location_hour.where((location_hour['location_description'] == 'STREET') | (location_hour['location_description'] == 'RESIDENCE'))
#a data frame with location descriptions and counts of recorded crimes, and hours...
street_home_hour_pddf = pd.DataFrame(street_home_hour.rdd.map(lambda row: row.asDict()).collect())
street_home_hour_pddf = street_home_hour_pddf.sort_values(by='hour')
figure, axes = plt.subplots()
axes.plot(street_home_hour_pddf[street_home_hour_pddf['location_description'] == 'RESIDENCE']['hour'],
street_home_hour_pddf[street_home_hour_pddf['location_description'] == 'RESIDENCE']['count'],
label='Crimes at home')
axes.plot(street_home_hour_pddf[street_home_hour_pddf['location_description'] == 'STREET']['hour'],
street_home_hour_pddf[street_home_hour_pddf['location_description'] == 'STREET']['count'],
label='Crimes on the street')
axes.legend()
ax.grid(b=True, which='both', axis='y')
```
#### This may be just me, but it seems that crimes at home increase when crimes on the street decrease. Could it be that it's the same kind of people that are in charge of both home and streets?
### Domestic crimes:
```
domestic_hour = pd.DataFrame(df_hour.groupBy(['domestic', 'hour']).count().orderBy('hour').rdd.map(lambda row: row.asDict()).collect())
dom = domestic_hour[domestic_hour['domestic'] == 'True']['count']
non_dom = domestic_hour[domestic_hour['domestic'] == 'False']['count']
either_dom = domestic_hour.groupby(by=['hour']).sum()['count']
dom_keys = domestic_hour[domestic_hour['domestic'] == 'False']['hour']
```
#### How do domestic crimes compare the other crimes?
```
figure, axes = plt.subplots()
axes.plot(dom_keys, either_dom, label='Total hourly count')
axes.plot(dom_keys, dom, label='Domestic crime count')
axes.plot(dom_keys, non_dom, label='Non-Domestic hourly count')
axes.legend()
axes.grid(which='b', b=True)
```
### A closer look at crime date and time
The information given here indicates when the crime is perpetrated. The date/time field may be able to draw a meaningful trend that can be used to predict crime. However, I believe that this leads much more to external factors, such as policy changes, law inforcement-related factors, and so on.
It's much more likely that time-related features that are more closely relatable to crime occurence be better predictors than the date and time. I mean, knowing the month of the year, the day of the week, and the hour of the day that the crime occurred can enable better chances of predicting accurately than simply knowing "when" AD crimes occurred.
Adding predictors (so far fields) that read time information:
* hour of day (already added the 'hour' field)
* day of week
* month of year
* day in a range. Instead of using the entire date-time, we'll use a "day sequence" that is counted from 2001-01-01.
```
df_dates = df_hour.withColumn('week_day', dayofweek(df_hour['date_time']))\
.withColumn('year_month', month(df_hour['date_time']))\
.withColumn('month_day', dayofmonth(df_hour['date_time']))\
.withColumn('date_number', datediff(df['date_time'], to_date(lit('2001-01-01'), format='yyyy-MM-dd')))\
.cache()
df_dates.select(['date', 'month', 'hour', 'week_day', 'year', 'year_month', 'month_day', 'date_number']).show(20, truncate=False)
```
## Day of week crime levels
It seems intriguing that the're little variance... But bad people are bad all week, aren't they...
```
week_day_crime_counts = df_dates.groupBy('week_day').count()
week_day_crime_counts_pddf = pd.DataFrame(week_day_crime_counts.orderBy('week_day').rdd.map(lambda e: e.asDict()).collect())
sns.barplot(data=week_day_crime_counts_pddf, x='week_day', y='count')
```
## Month of year
It seems that May-August are the busiest months for criminals... What could be the reason?
```
year_month_crime_counts = df_dates.groupBy('year_month').count()
year_month_crime_counts_pddf = pd.DataFrame(year_month_crime_counts.orderBy('year_month').rdd.map(lambda e: e.asDict()).collect())
year_month_crime_counts_pddf
```
What happens in MAY through AUGUST?
* Salary increase season?
* Weather?
I think these don't form the Christmas season in Chicago...
```
sns.barplot(data=year_month_crime_counts_pddf, y='count', x='year_month')
```
## Day of month
```
month_day_crime_counts = df_dates.groupBy('month_day').count()
month_day_crime_counts_pddf = pd.DataFrame(month_day_crime_counts.orderBy('month_day').rdd.map(lambda e: e.asDict()).collect())
```
#### Top 10 worst days of the month
```
month_day_crime_counts_pddf.sort_values(by='count', ascending=False).head(10)
month_day_crime_counts_pddf = month_day_crime_counts_pddf.sort_values(by='month_day', ascending=True)
fg, ax = plt.subplots()
ax.plot(month_day_crime_counts_pddf['month_day'], month_day_crime_counts_pddf['count'], label='Crimes over the month')
ax.grid(b=True, which='both')
ax.legend()
```
### Community areas?
Chicago has 77 community areas. How do they appear next to one another in a count plot?
```
df_dates_community_areas = df_dates.na.drop(subset=['community_area']).groupBy('community_area').count()
```
What are the to 10 areas with recorded crime?
```
df_dates_community_areas.orderBy('count', ascending=False).show(10)
## Is it possible to see how the top crime types occur in the top crime-affected areas?
top_crime_types = df_dates.select('primary_type').groupBy('primary_type').count().rdd.map(lambda row: row.asDict()).takeOrdered(10, key=lambda l: 1/l['count'])
top_busy_areas = df_dates_community_areas.rdd.map(lambda row: row.asDict()).takeOrdered(10, key=lambda l: 1/l['count'])
top_crime_types_lst = [dc['primary_type'] for dc in top_crime_types]
top_busy_areas_lst = [dc['community_area'] for dc in top_busy_areas]
top_crime_types_lst
top_busy_areas_lst
```
Find crimes of the top 10 types, that were perpetrated in the 10 10 busiest cities...
```
q1 = "instr('" + ' '.join(top_busy_areas_lst) + "', community_area) > 0"
q2 = "instr('" + ' '.join(top_crime_types_lst) + "', primary_type) > 0"
print(q1)
## Construct a data frame filtered on these top community areas and top crime types:
df_dates_tops = df_dates.filter(q1).filter(q2)
df_dates_tops.count()
tops_of_tops = df_dates_tops.groupBy(['primary_type', 'community_area']).count().orderBy(['primary_type', 'count', 'community_area'], ascending=[True, False, True]).cache()
tops_of_tops.show(20)
```
#### Community area names
Let's link names of these community areas. These float numbers are having a hard time making any sense...
Sourced from Chicago Tribune at http://www.chicagotribune.com/chi-community-areas-htmlstory.html
```
area_names = """
01 Rogers Park
40 Washington Park
02 West Ridge
41 Hyde Park
03 Uptown
42 Woodlawn
04 Lincoln Square
43 South Shore
05 North Center
44 Chatham
06 Lakeview
45 Avalon Park
07 Lincoln Park
46 South Chicago
08 Near North Side
47 Burnside
09 Edison Park
48 Calumet Heights
10 Norwood Park
49 Roseland
11 Jefferson Park
50 Pullman
12 Forest Glen
51 South Deering
13 North Park
52 East Side
14 Albany Park
53 West Pullman
15 Portage Park
54 Riverdale
16 Irving Park
55 Hegewisch
17 Dunning
56 Garfield Ridge
18 Montclare
57 Archer Heights
19 Belmont Cragin
58 Brighton Park
20 Hermosa
59 McKinley Park
21 Avondale
60 Bridgeport
22 Logan Square
61 New City
23 Humboldt Park
62 West Elsdon
24 West Town
63 Gage Park
25 Austin
64 Clearing
26 West Garfield Park
65 West Lawn
27 East Garfield Park
66 Chicago Lawn
28 Near West Side
67 West Englewood
29 North Lawndale
68 Englewood
30 South Lawndale
69 Greater Grand Crossing
31 Lower West Side
70 Ashburn
32 Loop
71 Auburn Gresham
33 Near South Side
72 Beverly
34 Armour Square
73 Washington Heights
35 Douglas
74 Mount Greenwood
36 Oakland
75 Morgan Park
37 Fuller Park
76 O'Hare
38 Grand Boulevard
77 Edgewater
39 Kenwood
"""
code_pairs = [[float(p[0]), p[1]] for p in [pair.strip().split('\t') for pair in area_names.strip().split('\n')]]
code_pairs[:5]
```
#### A view of crime count per community area
```
community_area_counts = pd.DataFrame(df_dates_community_areas.rdd.map(lambda row: row.asDict()).collect())
# Create a dictionary of area code to names
area_name_dic = {float(k[0]):k[1] for k in code_pairs}
community_area_counts['community_area_name'] = community_area_counts['community_area'].apply(lambda area: area_name_dic.get(float(area), 'unknown_%s'%area))
community_area_counts = community_area_counts.sort_values(by='count')
community_area_counts.head(5)
```
**What is happening in Austin?**
plt.rcParams["figure.figsize"] = [32, 32]
sns.set(style="whitegrid")
sns.set_color_codes("pastel")
#sns.despine(left=True, bottom=True)
area_chart = sns.barplot(x='count', y='community_area_name', data=community_area_counts)
area_chart.set(ylabel="Community Area Name", xlabel="Overall Crimes Record Count")
** *Join these community areas on the main data frame* **
```
code_pairs_df = spark.createDataFrame(code_pairs, ['community_area', 'area_name'])
named_tops_of_tops = code_pairs_df.join(tops_of_tops, on='community_area', how='right')
named_tops_of_tops.show(10)
tops_of_tops_dff = pd.DataFrame(named_tops_of_tops.rdd.map(lambda l: l.asDict()).collect() )
plt.rcParams["figure.figsize"] = [64, 16]
sns.barplot(data=tops_of_tops_dff, x='area_name', y='count', hue='primary_type', palette='pastel')
```
---
# Can we try to predict the primary type of crime?
Now on to another objective of this exercise. We're going to try to predict what kind of crime is going to occur given the selected features.
Speaking about features, here's the list of predictors that I chose to go for:
### Excluded variables:
* 'id' - Random information that isn't a predictor of crime type
* 'case_number' - Random information that isn't a predictor of crime type
* 'date' - Removed because it's been re-featurized in other features generated above
* 'block' - Excluded as this may simply mean noise
* 'iucr' - Excluded as correlated with crime type. No point.
* 'x_coordinate' - Not included
* 'y_coordinate' - Not included
* 'year' - Not included (already otherwise featurized)
* 'updated_on' - not included
* 'latitude' - not included
* 'longitude' - not included
* 'location' - not included
* 'date_time' - Taken into account in other time-related features
* 'description' - Excluded. I want to see this as associated with the response (primary type)
### Selected predictors:
* 'location_description'
* 'arrest'
* 'domestic'
* 'beat'
* 'district'
* 'ward'
* 'community_area'
* 'fbi_code'
* 'hour'
* 'week_day'
* 'year_month'
* 'month_day'
* 'date_number'
```
selected_features = [
'location_description',
'arrest',
'domestic',
'beat',
'district',
'ward',
'community_area',
'fbi_code',
'hour',
'week_day',
'year_month',
'month_day',
'date_number']
#Let's see the schema of these selected features:
features_df = df_dates.select(selected_features)
features_df.printSchema()
```
Let's try to see all level counts in all the selected features...
```
feature_level_count_dic = []
for feature in selected_features:
print('Analysing %s' % feature)
levels_list_df = features_df.select(feature).distinct()
feature_level_count_dic.append({'feature': feature, 'level_count': levels_list_df.count()})
pd.DataFrame(feature_level_count_dic).sort_values(by='level_count', ascending=False)
```
### Preparing model
```
from pyspark.ml.feature import StringIndexer, VectorAssembler
df_dates_features = df_dates.na.drop(subset=selected_features)
```
Let us use Spark's string indexer to index selected features
```
for feature in feature_level_count_dic:
indexer = StringIndexer(inputCol=feature['feature'], outputCol='%s_indexed' % feature['feature'])
print('Fitting feature "%s"' % feature['feature'])
model = indexer.fit(df_dates_features)
print('Transforming "%s"' % feature['feature'])
df_dates_features = model.transform(df_dates_features)
```
Now let's do the same with the label
```
## String-index the response variable:
response_indexer = StringIndexer(inputCol='primary_type', outputCol='primary_type_indexed')
response_model = response_indexer.fit(df_dates_features)
df_dates_features = response_model.transform(df_dates_features)
#What does it look like now...
df_dates_features.show(1)
```
Let's vectorize the features
We use a vector assembler to vectorize all predictors into a `features` column
```
indexed_features = ['%s_indexed' % fc['feature'] for fc in feature_level_count_dic]
indexed_features
assembler = VectorAssembler(inputCols=indexed_features, outputCol='features')
vectorized_df_dates = assembler.transform(df_dates_features)
vectorized_df_dates.select('features').take(1)
```
### Now let's train the model.
We're using **60%** to **40%** split between the train and the test datasets
Let's try to train a logistic regression.
```
train, test = vectorized_df_dates.randomSplit([0.6, 0.4])
from pyspark.ml.classification import LogisticRegression
logisticRegression = LogisticRegression(labelCol='primary_type_indexed', featuresCol='features', maxIter=10, family='multinomial')
fittedModel = logisticRegression.fit(train)
```
## Model performance?
```
fittedModel.summary.accuracy
model_summary = fittedModel.summary
fittedModel.coefficientMatrix
```
#### Why the 34X13 shape?
That's because the multinomial logistic regression is fitted on **each class** of the label. It computes the probability of each class and then predicts based on these probabilities.
```
print(fittedModel.coefficientMatrix)
print('Coefficient matrix:\nRow count = %s\nCol count = %s' % (fittedModel.coefficientMatrix.numRows, fittedModel.coefficientMatrix.numCols))
print('Model:\nNum Classes = %s\nNum Features = %s' % (fittedModel.numClasses, fittedModel.numFeatures))
print('Training "primary_type" factor level count = %s' % train.select('primary_type_indexed').distinct().count())
```
So our `randomSplit` call to split train/test data sets took some primary types to only either of the two parts
```
vectorized_df_dates.select('features').show(2, truncate=False)
fittedModel.numClasses
fittedModel.numFeatures
train.select('primary_type_indexed').distinct().count()
df_dates.select('primary_type').distinct().count()
fittedModel.interceptVector.values.size
print(model_summary.objectiveHistory)
print()
print('Objective history size ', len(model_summary.objectiveHistory))
sns.barplot(y=model_summary.objectiveHistory, x=list(range(len(model_summary.objectiveHistory))))
label_stats = {float(i):{'index': float(i)} for i in range(34)}
print(label_stats)
print("False positive rate by label:")
for i, rate in enumerate(model_summary.falsePositiveRateByLabel):
label_stats[i]['false_positive_rate'] = rate
for i, rate in enumerate(model_summary.truePositiveRateByLabel):
label_stats[i]['true_positive_rate'] = rate
for i, rate in enumerate(model_summary.precisionByLabel):
label_stats[i]['precision_rate'] = rate
for i, rate in enumerate(model_summary.recallByLabel):
label_stats[i]['recall_rate'] = rate
for i, rate in enumerate(model_summary.fMeasureByLabel()):
label_stats[i]['f_measure'] = rate
train_rdd = train.select(['primary_type', 'primary_type_indexed']).distinct().orderBy('primary_type_indexed').rdd.map(lambda l: l.asDict()).collect()
for l in train_rdd:
print(l)
label_stats[l['primary_type_indexed']]['primary_type'] = l['primary_type']
rates_pddf = pd.DataFrame(list(label_stats.values()))
rates_pddf = rates_pddf.sort_values(by='precision_rate', ascending=False)
```
#### Why does this model seem to be so good at predicting theft? (TODO)
```
rates_pddf
```
## TODO:
* Run the model on the test dataset
| github_jupyter |
```
##Author: Gene Burinskiy
!pip install plinkio
#!pip install h5py --for some reason, h5py doesn't install :/
#!pip install tables --since h5py can't be installed, neither can tables
import os
import re
import numpy as np
import pandas as pd
from plinkio import plinkfile
os.getcwd()
#working with original dataset
data_path = '/home/jovyan/work/LEAP/leap/regression/dataset1'
os.chdir(data_path)
os.listdir()
bed = plinkfile.open("dataset1")
loci = bed.get_loci()
len(loci)
locus = loci[0]
locus.name
locus.chromosome
np.unique([x.chromosome for x in loci])
samples = bed.get_samples()
print("Object of type: ",type(samples), "and length:" ,len(samples))
sample = samples[500]
print(sample.fid, sample.father_iid, sample.iid, sample.phenotype, sample.sex)
h = [row for row in bed][0]
phenos = pd.read_csv("dataset1.phe", sep=' ', header=0, names=['fam', 'person', 'pheno'], skiprows=0)
phenos.iloc[:5]
len([x for x in h])
##each row in bed is of length 10000 and there are 10,499 rows thus to make a matrix:
mat = np.zeros((10499,1000), dtype='int16') #1/4 of the taken up space
i=0
for row in bed:
mat[i,:] = np.array([snp for snp in row])
i+=1
#this matrix is equivalent to transposed bed.val
print("Data type:", mat.dtype)
print(mat[:2,:5])
print("Size of bed matrix: %4.0fmb" %(mat.nbytes/(1024**2)))
df = pd.DataFrame(mat.transpose()) #normally, it reads them in as floats which is a huge waste of space
df.columns = [x.name for x in loci]
df.index = [x.iid for x in bed.get_samples()] #could also double index on chromosomes
df.iloc[:5,:5]
np.unique(df.dtypes)
df = df.astype('float32')-df.astype('float32').mean() #this gets us pretty close to their normalization stuff
df.iloc[:5,:5]
##Save the file to sql db - not feasible for this data
#from sqlalchemy import create_engine
#engine = create_engine('sqlite:///dataset1.db', echo=False)
#df.transpose().to_sql(name='dataset1', con=engine, if_exists = 'replace', index=True)
%%timeit
np.cov(df)
cov = np.cov(df)
print("Shave of covariance matrix:", cov.shape)
cov[:5,:5]
"""
A straightforward solution of the optimization problem presented above is difficult owing to high dimensionality, which is equal to the number of genotyped
variants. Fortunately, the problem can be reformulated as a lower-dimensional problem, with dimensionality equal to the number of individuals.
The equivalence stems from the fact that the genotypes
matrix X can be represented in terms of the eigenvectors of its covariance
"""
%%timeit
U,s,V = np.linalg.svd(df,full_matrices=False )
import scipy
%%timeit
U,s,V, = scipy.linalg.svd(df, full_matrices=False)
%%timeit
U,s,V = scipy.linalg.svd(df, full_matrices=False)
"""
Dependencies don't quite exist.
from pandas import HDFStore
hdf = HDFStore('dataset1.h5')
# put the dataset in the storage
hdf.put('dataset1', df, format='table', data_columns=True)
hdf.append('d1', DataFrame(np.random.rand(5,3),
columns=('A','B','C')),
format='table', data_columns=True)
hdf.close() # closes the file
hdf = read_hdf('storage.h5', 'd1',
where=['A>.5'], columns=['A','B'])
"""
U,s,V = scipy.linalg.svd(df, full_matrices=False)
print("Shapes of U,s,and V respectively")
print(U.shape, s.shape, V.shape)
```
| github_jupyter |
# Transforming Images
```
#@ImageJ ij
image = ij.io().open("http://imagej.net/images/clown.png")
```
Image transformations such as rotation, scaling and cropping are accomplished using ops of the `transform` namespace.
Most ops of this namespace have the nice property of being _views_: they do not actually copy image samples, but rather wrap the image, offering a modified "window" into the original data. Using views helps to greatly reduce computer memory usage, at a very minimal time performance cost. If you need a deep copy of the image for some reason (e.g., if time performance is paramount, or if you want to modify the transformed image samples in-place without affecting other transformed versions of the image), you can still copy it using the `copy` namespace; see "Generating images notebook" for details.
```
ij.op().ops().findAll{op ->
op.startsWith("transform.")
}.collect{op -> op[10..-1]}
```
### Rotating an image
The `transform.rotateView` op rotates the image 90 degrees from one dimensional axis to another.
```
ij.op().help("rotateView")
```
Here is an example usage of the `transform.rotateView` op.
```
// set parameter for rotate
x = 0; y = 1; c = 2
// define functions to see the bounds of an image
bounds = {interval ->
return "(" + interval.min(0) + ", " + interval.min(1) + ") - " +
"(" + interval.max(0) + ", " + interval.max(1) + ")"
}
// Rotate the image (image, fromAxis, toAxis)
rotated = ij.op().run("rotateView", image, x, y) // 90 degrees clockwise
//rotated = ij.op().run("rotateView", image, y, x)// 90 degrees counter-clockwise
//rotated = ij.op().run("rotateView", image, x, c) // rotate through channels! WAT
// The interval bounds have automatically changed!
println("Old bounds: " + bounds(image))
println("New bounds: " + bounds(rotated))
rotated
```
### Cropping an image
The `transform.crop` op crops an image N-dimensionally. E.g., you can use it to create a substack of a 3D dataset, cut out irrelevant channels, or crop the XY planes as with 2D image processing software.
```
ij.op().help("crop")
```
Below, we show two ways to crop an image: 1) with `transform.crop`; and 2) using `transform.intervalView`. The former translates the image back to the origin, while the latter does not.
```
import net.imglib2.FinalInterval
region = FinalInterval.createMinSize(75, 27, 0, 40, 28, 3)
eye = ij.op().run("crop", image, region)
eyeView = ij.op().run("intervalView", image, region)
[["eye":eye, "view":eyeView]]
```
### Scaling an image
To perform [image scaling](https://en.wikipedia.org/wiki/Image_scaling), use the `transform.scaleView` op. You already saw it in action in the "Getting started" section, but here it is again—this time enlarging an image rather than reducing it.
Just for fun, we compare three different interpolation strategies: [nearest neighbor](https://en.wikipedia.org/wiki/Nearest-neighbor_interpolation), [N-linear](https://en.wikipedia.org/wiki/Linear_interpolation), and [Lanczos](https://en.wikipedia.org/wiki/Lanczos_resampling).
```
import net.imglib2.interpolation.randomaccess.NearestNeighborInterpolatorFactory
import net.imglib2.interpolation.randomaccess.NLinearInterpolatorFactory
import net.imglib2.interpolation.randomaccess.LanczosInterpolatorFactory
scaleFactors = [4, 4, 1] // Enlarge X and Y by 4x; leave channel count the same.
nearestNeighborEye = ij.op().run("scaleView", eye, scaleFactors, new NearestNeighborInterpolatorFactory())
nLinearEye = ij.op().run("scaleView", eye, scaleFactors, new NLinearInterpolatorFactory())
lanczosEye = ij.op().run("scaleView", eye, scaleFactors, new LanczosInterpolatorFactory())
[["nearest neighbor":nearestNeighborEye, "N-linear":nLinearEye, "Lanczos":lanczosEye]]
```
Of course, some detail from the original image has been lost, since we scaled down and then back up again.
### Padding an image
The `transform.intervalView` can also be used to expand the bounding box of an image. However, there is one catch: you must first decide what the out-of-bounds sample values will be. The `transform.extend` ops achieve this goal. If you forget to extend the image before padding it via `intervalView`, you will receive an exception when attempting to query any out-of-bounds samples.
Note that the various `transform.extend` ops explicitly remove the bounding box of an image, expanding the defined sample values to infinity in all directions. In most circumstances, you will want to use `transform.intervalView` to rebound the image after extending it.
```
ij.op().ops().findAll{op ->
op.startsWith("transform.extend")
}.collect{op -> op[10..-1]}
```
Here are some side-by-side examples of what happens when you pad an image with these various approaches:
```
def pad(image, extended, t, r, b, l) {
min = new long[image.numDimensions()]
max = new long[image.numDimensions()]
image.min(min)
image.max(max)
min[0] -= l; min[1] -= t; max[0] += r; max[1] += b
return ij.op().run("intervalView", extended, min, max)
}
// Define the top, right, bottom and left padding amounts.
t = r = b = l = 20
// Pad the image with different out-of-bounds strategies.
extendedBorder = ij.op().run("transform.extendBorderView", eye)
paddedBorder = pad(eye, extendedBorder, t, r, b, l)
extendedMirrorDouble = ij.op().run("transform.extendMirrorDoubleView", eye)
paddedMirrorDouble = pad(eye, extendedMirrorDouble, t, r, b, l)
extendedMirrorSingle = ij.op().run("transform.extendMirrorSingleView", eye)
paddedMirrorSingle = pad(eye, extendedMirrorSingle, t, r, b, l)
extendedPeriodic = ij.op().run("transform.extendPeriodicView", eye)
paddedPeriodic = pad(eye, extendedPeriodic, t, r, b, l)
minValue = eye.firstElement().getMinValue().doubleValue()
maxValue = eye.firstElement().getMaxValue().doubleValue()
extendedRandom = ij.op().run("transform.extendRandomView", eye, minValue, maxValue)
paddedRandom = pad(eye, extendedRandom, t, r, b, l)
value = eye.firstElement().copy(); value.set(100)
extendedValue = ij.op().run("transform.extendValueView", eye, value)
paddedValue = pad(eye, extendedValue, t, r, b, l)
extendedZero = ij.op().run("transform.extendZeroView", eye)
paddedZero = pad(eye, extendedZero, t, r, b, l)
[["border":paddedBorder, "mirror double":paddedMirrorDouble,
"mirror single":paddedMirrorSingle, "periodic":paddedPeriodic,
"random":paddedRandom, "value":paddedValue, "zero":paddedZero]]
```
| github_jupyter |
# Cross-Entropy Method
---
In this notebook, we will train the Cross-Entropy Method with OpenAI Gym's MountainCarContinuous environment.
### 1. Import the Necessary Packages
```
import gym
import math
import numpy as np
from collections import deque
import matplotlib.pyplot as plt
%matplotlib inline
import time
import torch
import torch.nn as nn
#import torch.nn.tanh as tanh
from torch.autograd import Variable
```
### 2. Instantiate the Environment and Agent
```
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
env = gym.make('MountainCarContinuous-v0')
env.seed(101)
np.random.seed(101)
print('observation space:', env.observation_space)
print('action space:', env.action_space)
print(' - low:', env.action_space.low)
print(' - high:', env.action_space.high)
class Agent(nn.Module):
def __init__(self, env, h_size=16):
super(Agent, self).__init__()
self.env = env
# state, hidden layer, action sizes
self.s_size = env.observation_space.shape[0]
self.h_size = h_size
self.a_size = env.action_space.shape[0]
# define layers
self.fc1 = nn.Linear(self.s_size, self.h_size)
self.fc2 = nn.Linear(self.h_size, self.a_size)
def set_weights(self, weights):
s_size = self.s_size
h_size = self.h_size
a_size = self.a_size
# separate the weights for each layer
fc1_end = (s_size*h_size)+h_size
fc1_W = torch.from_numpy(weights[:s_size*h_size].reshape(s_size, h_size))
fc1_b = torch.from_numpy(weights[s_size*h_size:fc1_end])
fc2_W = torch.from_numpy(weights[fc1_end:fc1_end+(h_size*a_size)].reshape(h_size, a_size))
fc2_b = torch.from_numpy(weights[fc1_end+(h_size*a_size):])
# set the weights for each layer
self.fc1.weight.data.copy_(fc1_W.view_as(self.fc1.weight.data))
self.fc1.bias.data.copy_(fc1_b.view_as(self.fc1.bias.data))
self.fc2.weight.data.copy_(fc2_W.view_as(self.fc2.weight.data))
self.fc2.bias.data.copy_(fc2_b.view_as(self.fc2.bias.data))
def get_weights_dim(self):
return (self.s_size+1)*self.h_size + (self.h_size+1)*self.a_size
def forward(self, x):
x = torch.relu(self.fc1(x))
x = torch.tanh(self.fc2(x))
return x.cpu().data
def evaluate(self, weights, gamma=1.0, max_t=5000):
self.set_weights(weights)
episode_return = 0.0
state = self.env.reset()
for t in range(max_t):
state = torch.from_numpy(state).float().to(device)
action = self.forward(state)
state, reward, done, _ = self.env.step(action)
episode_return += reward * math.pow(gamma, t)
if done:
break
return episode_return
agent = Agent(env).to(device)
```
### 3. Train the Agent with the Cross-Entropy Method
Run the code cell below to train the agent from scratch. Alternatively, you can skip to the next code cell to load the pre-trained weights from file.
```
def cem(n_iterations=5000, max_t=1000, gamma=0.999, print_every=10, pop_size=50, elite_frac=0.2, sigma=0.5):
"""PyTorch implementation of the cross-entropy method.
Params
======
n_iterations (int): maximum number of training iterations
max_t (int): maximum number of timesteps per episode
gamma (float): discount rate
print_every (int): how often to print average score (over last 100 episodes)
pop_size (int): size of population at each iteration
elite_frac (float): percentage of top performers to use in update
sigma (float): standard deviation of additive noise
"""
n_elite=int(pop_size*elite_frac)
scores_deque = deque(maxlen=100)
scores = []
best_weight = sigma*np.random.randn(agent.get_weights_dim())
for i_iteration in range(1, n_iterations+1):
weights_pop = [best_weight + (sigma*np.random.randn(agent.get_weights_dim())) for i in range(pop_size)]
rewards = np.array([agent.evaluate(weights, gamma, max_t) for weights in weights_pop])
elite_idxs = rewards.argsort()[-n_elite:]
elite_weights = [weights_pop[i] for i in elite_idxs]
best_weight = np.array(elite_weights).mean(axis=0)
reward = agent.evaluate(best_weight, gamma=1.0)
scores_deque.append(reward)
scores.append(reward)
torch.save(agent.state_dict(), 'checkpoint.pth')
if i_iteration % print_every == 0:
print('Episode {}\tAverage Score: {:.2f}'.format(i_iteration, np.mean(scores_deque)))
# if np.mean(scores_deque)>=200.0:
# print('\nEnvironment solved in {:d} iterations!\tAverage Score: {:.2f}'.format(i_iteration, np.mean(scores_deque)))
# break
return scores
print(f"start_time: {time.ctime()}\n")
scores = cem()
print(f"\nend time_time: {time.ctime()}")
# plot the scores
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(1, len(scores)+1), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
```
### 4. Watch a Smart Agent!
In the next code cell, you will load the trained weights from file to watch a smart agent!
```
# load the weights from file
agent.load_state_dict(torch.load('checkpoint.pth'))
state = env.reset()
env.render()
time.sleep(5)
while True:
state = torch.from_numpy(state).float().to(device)
with torch.no_grad():
action = agent(state)
action = env.action_space.sample()
env.render()
next_state, reward, done, _ = env.step(action)
state = next_state
if done:
break
env.close()
```
| github_jupyter |
```
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib import pyplot
import numpy as np
plt.rcParams.update({'figure.max_open_warning': 0})
df = pd.read_csv('\\Results_CSV\\Results_Transfer_Learning_Sim_to_Physical\\result_transfer_Sim_to_Phy_dijkstra_and_sha.csv')
df
ls = df['model_name']
ls
def convert_to_num(arr, ind):
# print(arr[ind])
arr = arr[ind]
arr = [float(s) for s in arr[1:-1].split(',')]
# print(arr)
return arr
def convert_to_num_mape(arr, ind):
# print(arr[ind])
arr = arr[ind]
arr = [float(s) for s in arr[1:-1].split(',')]
arr = [x*100 for x in arr]
# print(arr)
return arr
r2_dijkstra = []
r2_sha = []
ape_dijkstra = []
ape_sha = []
for i in range(11):
temp = df.iloc[i]['r2_dijkstra']
r2_dijkstra.append(temp)
for i in range(11):
temp = df.iloc[i]['r2_sha']
r2_sha.append(temp)
for i in range(11):
temp = df.iloc[i]['ape_dijkstra']
ape_dijkstra.append(temp)
for i in range(11):
temp = df.iloc[i]['ape_sha']
ape_sha.append(temp)
ape_sha = [x*100 for x in ape_sha]
ape_dijkstra = [x*100 for x in ape_dijkstra]
print(ls)
ape_dijkstra, ape_sha, np.mean(ape_dijkstra), np.mean(ape_sha)
np.mean(r2_sha[5:10]), np.mean(r2_dijkstra[5:10]), np.mean(ape_sha[5:10]), np.mean(ape_dijkstra[5:10])
def find_y_r2(r2):
y = []
for i in range(len(r2)):
y.append(np.mean(r2[i]))
return y
def find_y_mape(mape):
y = []
for i in range(len(mape)):
y_temp = [x * 100 for x in mape[i]]
y.append(np.median(y_temp))
return y
import matplotlib as mpl
font = {'family' : 'normal',
'weight' : 'bold',
'size' : 18}
mpl.rc('font', **font)
```
# SHA
```
ls = ['svr', 'lr', 'rr', 'knn', 'gpr','dt','rf','etr',
'gbr', 'xgb','dnn']
fig, ax = plt.subplots(1,1, figsize = (11, 8)) # create 2x2 array of subplots
# color = 'tab:red'
# y_r2 = r2_qsort_s
y_mape = ape_sha
# px = ax.bar(ls,y_r2,color = 'mediumorchid', label = 'R2')
# ax.legend(frameon=True, loc='upper left', prop={'size': 20})
# ax[i,j].bar(ls, y,color = 'blue')
# ax.set_xticklabels(ls, rotation = 90, fontsize = 35, fontweight = 'bold')
# if j == 4:
# ax[i,j].set_ylabel('MedAPE score')
# ax.set_title(dataset_names[cb[j]] + ' (' + dataset[cb[j]] + ')', fontsize = 25, fontweight = 'bold')
# ax.set_ylim(-0.5,1.5)
# ax.grid()
ax_temp = ax
# create data
x = ls
values = y_mape
# stem function: first way
ax_temp.stem(x, values)
# plt.ylim(0, 1.2)
#plt.show()
# stem function: If no X provided, a sequence of numbers is created by python:
# plt.stem(values)
#plt.show()
# stem function: second way
(markerline, stemlines, baseline) = ax_temp.stem(x, values, linefmt='-*', markerfmt='bs', basefmt='k-',label='APE score')
plt.setp(baseline, visible = False)
plt.setp(stemlines, 'linewidth', 12)
#plt.show()
# ax_temp = ax.twinx() # instantiate a second axes that shares the same x-axis
# line2 = ax_temp.bar(ls,y_mape, color = 'mediumorchid',label='APE score') # add boxplot to 1st subplot
# line, = ax.plot([1, 2, 3], label='Inline label')
# ax_temp.legend(frameon= True, loc='upper right', prop={'size': 20})
plt.title('Absolute Percentage Error', fontsize = 30, fontweight = 'bold')
# ax_temp.boxplot(y_mape,labels = ls)
# ax[i,j].bar(ls, y,color = 'blue')
ax_temp.set_xticklabels(ls, rotation = 90, fontsize = 35, fontweight = 'bold')
# ax_temp.set_yticklabels(fontsize = 15)
ax_temp.ticklabel_format(style='sci', axis='y', scilimits=(0,0))
# ax_temp.legend(['MedAPE score'])
fig.tight_layout() # otherwise the right y-label is slightly clipped
ax_temp.set_ylim(0,30)
# plt.show()
# extent = ax.get_tightbbox(fig.canvas.get_renderer()).transformed(fig.dpi_scale_trans.inverted())
# fig.savefig('cb_1.pdf', bbox_inches = extent)
ax_temp.grid()
name = 'transfer_learning_sha'
plt.rc('font', size=25)
fig.savefig(name+ '.eps')
fig.show()
```
# Dijkstra
```
ls = ['svr', 'lr', 'rr', 'knn', 'gpr','dt','rf','etr',
'gbr', 'xgb','dnn']
fig, ax = plt.subplots(1,1, figsize = (11, 8)) # create 2x2 array of subplots
# color = 'tab:red'
# y_r2 = r2_qsort_s
y_mape = ape_dijkstra
# px = ax.bar(ls,y_r2,color = 'mediumorchid', label = 'R2')
# ax.legend(frameon=True, loc='upper left', prop={'size': 20})
# ax[i,j].bar(ls, y,color = 'blue')
# ax.set_xticklabels(ls, rotation = 90, fontsize = 35, fontweight = 'bold')
# if j == 4:
# ax[i,j].set_ylabel('MedAPE score')
# ax.set_title(dataset_names[cb[j]] + ' (' + dataset[cb[j]] + ')', fontsize = 25, fontweight = 'bold')
# ax.set_ylim(-0.5,1.5)
# ax.grid()
ax_temp = ax
# create data
x = ls
values = y_mape
# stem function: first way
ax_temp.stem(x, values)
# plt.ylim(0, 1.2)
#plt.show()
# stem function: If no X provided, a sequence of numbers is created by python:
# plt.stem(values)
#plt.show()
# stem function: second way
(markerline, stemlines, baseline) = ax_temp.stem(x, values, linefmt='-*', markerfmt='bs', basefmt='k-',label='APE score')
plt.setp(baseline, visible = False)
plt.setp(stemlines, 'linewidth', 12)
#plt.show()
# ax_temp = ax.twinx() # instantiate a second axes that shares the same x-axis
# line2 = ax_temp.bar(ls,y_mape, color = 'mediumorchid',label='APE score') # add boxplot to 1st subplot
# line, = ax.plot([1, 2, 3], label='Inline label')
# ax_temp.legend(frameon= True, loc='upper right', prop={'size': 20})
plt.title('Absolute Percentage Error', fontsize = 30, fontweight = 'bold')
# ax_temp.boxplot(y_mape,labels = ls)
# ax[i,j].bar(ls, y,color = 'blue')
ax_temp.set_xticklabels(ls, rotation = 90, fontsize = 35, fontweight = 'bold')
# ax_temp.set_yticklabels(fontsize = 15)
ax_temp.ticklabel_format(style='sci', axis='y', scilimits=(0,0))
# ax_temp.legend(['MedAPE score'])
fig.tight_layout() # otherwise the right y-label is slightly clipped
ax_temp.set_ylim(0,30)
# plt.show()
# extent = ax.get_tightbbox(fig.canvas.get_renderer()).transformed(fig.dpi_scale_trans.inverted())
# fig.savefig('cb_1.pdf', bbox_inches = extent)
ax_temp.grid()
name = 'transfer_learning_dijkstra'
plt.rc('font', size=25)
fig.savefig(name+ '.eps')
fig.show()
```
| github_jupyter |
### HomeWork 8
#### Mouselinos Spyridon
#### February 2020
***
### Exersize 2
***
```
### Imports
import numpy as np
import matplotlib.pyplot as plt
### Let's define the sigmoid function with scale a
def sigmoid(t,a):
return 1/(1+ np.exp(-a*t))
### a) Plot the function for different parameters of a
datapoints = np.arange(-6.0,6.0,0.5)
a_values =[0, 0.01, 0.1, 1, 10, 100]
subplot_values = [231,232,233,234,235,236]
fig = plt.figure()
for i in range(6):
plt.subplot(subplot_values[i])
plt.plot(datapoints, sigmoid(datapoints,a_values[i]))
plt.title('Value of a= {}'.format(a_values[i]))
plt.subplots_adjust(hspace=0.75, wspace=0.95)
plt.show()
```
It is shown that as a increases the sigmoid function approaches the unit step function.
Also as a goes to 0, the sigmoid becomes constant approaching the value of 0.5.
### Exersize 4
***
```
import scipy.io as sio
import numpy as np
import numpy.matlib as nm
import matplotlib.pyplot as plt
from scipy.stats import multivariate_normal, norm
Dataset = sio.loadmat('HW8.mat')
train_x = Dataset['train_x']
train_y= Dataset['train_y']
test_x = Dataset['test_x']
test_y = Dataset['test_y']
### First let's separate the class 1 and class 2 points
class_1_points = train_x[np.where(train_y==1),:][0]
class_2_points = train_x[np.where(train_y==2),:][0]
```
#### a) Bayes classifier
***
```
### Count and get statistics for each class
N1 = class_1_points.shape[0]
N2 = class_2_points.shape[0]
N = len(train_y)
N1_mean = np.mean(class_1_points,axis=0)
N2_mean = np.mean(class_2_points,axis=0)
N1_cov = 1/ N1*((class_1_points - nm.repmat(N1_mean,N1,1) ).T ).dot((class_1_points - nm.repmat(N1_mean,N1,1)))
N2_cov = 1/ N2*((class_2_points - nm.repmat(N2_mean,N2,1)).T).dot((class_2_points - nm.repmat(N2_mean,N2,1)))
P1 = N1/N
P2 = N2/N
N_test = len(test_y)
class1 = multivariate_normal(N1_mean,N1_cov)
class1_scores = class1.pdf(test_x)*P1
class2 = multivariate_normal(N2_mean,N2_cov)
class2_scores = class2.pdf(test_x)*P2
Bayes_test_y = np.zeros((N,1))
Bayes_test_y[class1_scores > class2_scores] = 1
Bayes_test_y[class2_scores > class1_scores] = 2
error_bayes = np.sum(np.fromiter((i==1 for i in Bayes_test_y != test_y),np.float)) /N_test
print(error_bayes)
```
#### b) Naive Bayes CLassifier
***
```
### Here we need to recalculate the S matrices assuming independence
Cov_11 = np.sum(np.power(class_1_points[:,0].reshape(N1,1) - np.matlib.repmat(N1_mean[0],N1,1),2)) / N1
Cov_12 = np.sum(np.power(class_1_points[:,1].reshape(N1,1) - np.matlib.repmat(N1_mean[1],N1,1),2)) / N1
Cov_21 = np.sum(np.power(class_2_points[:,0].reshape(N1,1) - np.matlib.repmat(N2_mean[0],N2,1),2)) / N2
Cov_22 = np.sum(np.power(class_2_points[:,1].reshape(N1,1) - np.matlib.repmat(N2_mean[1],N2,1),2)) / N2
class1_scores = np.multiply(norm(N1_mean[0],np.sqrt(Cov_11)).pdf(test_x[:,0]), norm(N1_mean[1],np.sqrt(Cov_12)).pdf(test_x[:,1]))
class2_scores = np.multiply(norm(N2_mean[0],np.sqrt(Cov_21)).pdf(test_x[:,0]), norm(N2_mean[1],np.sqrt(Cov_22)).pdf(test_x[:,1]))
Naive_Bayes_test_y = np.zeros((N,1))
Naive_Bayes_test_y[class1_scores > class2_scores] = 1
Naive_Bayes_test_y[class2_scores > class1_scores] = 2
error_naive_bayes = np.sum(np.fromiter((i==1 for i in Naive_Bayes_test_y!=test_y),np.float)) /N_test
print(error_naive_bayes)
```
#### c) knn classifier
***
```
from sklearn.neighbors import KNeighborsClassifier
clf = KNeighborsClassifier(n_neighbors=5)
clf.fit(train_x, train_y.ravel())
kNN_test_y = clf.predict(test_x).reshape(N_test,1)
error_kNN = np.sum(np.fromiter((i==1 for i in kNN_test_y!=test_y),np.float)) /N_test
print(error_kNN)
```
#### d) logistic regression classifier
***
```
class LogisticRegressionClassifier:
def __init__(self, lr=0.01, tolerance=0.0001, maxIterations=500, extend_with_ones=True):
self.lr = lr
self.tolerance = tolerance
self.maxIterations = maxIterations
self.extend_with_ones = extend_with_ones
self.final_theta = None
self._isfit = False
def sigmoid(self,t):
return 1/(1+np.exp(-t))
def fit(self, X, y):
### Rectify y to 0-1
y = y-1
### Find the Input shape
input_length = X.shape[0]
input_shape = X.shape[1]
### If bias is required extend the input with ones
if self.extend_with_ones:
X = np.hstack((np.ones((input_length,1)),X))
input_shape += 1
### Initialize the theta parameter table
theta = np.random.normal(0,1,size=(input_shape,1))
### If there is not significant update of theta stop
### The magnitude of the gradient update can be calculated
### By the L1 norm of the gradient at each step
gradient_magnitude = 10000
step = 0
while step < self.maxIterations:
step += 1
### We can use the sigmoid we created at Exersize 2 with alpha 1
s = self.sigmoid((theta.T).dot(X.T))
gradient = (X.T).dot((s.T - y))
gradient_magnitude = np.linalg.norm(gradient)
if gradient_magnitude < self.tolerance:
self.final_theta = theta
break
old_theta = theta
theta = old_theta - self.lr * gradient
return
def transform(self, X, y_test, threshold=0.5):
### Rectify y
y_test = y_test-1
### Find the Input shape
input_shape = X.shape[0]
### If bias is required extend the input with ones
if self.extend_with_ones:
X = np.hstack((np.ones((input_shape,1)),X))
input_shape += 1
predictions = (self.sigmoid((self.final_theta.T).dot(X.T))).T
class_predictions = np.zeros_like(predictions)
class_predictions[np.where(predictions>0.5)] = 1
error_Logistic_Regression = np.sum(np.fromiter((i==1 for i in class_predictions!=y_test),np.float)) / len(y_test)
return error_Logistic_Regression, class_predictions+1
clf = LogisticRegressionClassifier(lr=0.01, tolerance=0.0001, maxIterations=500)
clf.fit(X=train_x,y=train_y)
error = clf.transform(X=test_x,y_test=test_y)[0]
lry = clf.transform(X=test_x,y_test=test_y)[1]
print("Error of Logistic Regression",error)
```
#### e) Classification results on the test set
***
```
### Let's Plot the Training Set and the Test set
def plot_dataset(name, points, labels):
plt.figure(figsize=(10,10))
plt.title(name)
class_1 = points[np.where(labels==1),:][0]
class_2 = points[np.where(labels==2),:][0]
plt.scatter(x=class_1[:,0], y=class_1[:,1], c='m', label='class 1')
plt.scatter(x=class_2[:,0], y=class_2[:,1], c='y', label='class 2')
plt.xlabel("1st Dimention of the Dataset")
plt.ylabel("2nd Dimention of the Dataset")
plt.legend(loc=1)
plt.show()
plt.close()
return
def plot_predictions(name, points, real_labels, predicted_labels):
plt.figure(figsize=(10,10))
plt.title(name)
class_1 = points[np.where(real_labels==1),:][0]
class_2 = points[np.where(real_labels==2),:][0]
class_1_correct = points[np.where(predicted_labels==1),:][0]
class_2_correct = points[np.where(predicted_labels==2),:][0]
plt.scatter(x=class_1[:,0], y=class_1[:,1], c='b',marker='o',label='Real class 1')
plt.scatter(x=class_1_correct[:,0], y=class_1_correct[:,1], c='b', marker='+',label='Predicted class 1')
plt.scatter(x=class_2[:,0], y=class_2[:,1], c='y', marker='o',label='Real class 2')
plt.scatter(x=class_2_correct[:,0], y=class_2_correct[:,1], c='y', marker='+',label='Predicted class 2')
plt.xlabel("1st Dimention of the Dataset")
plt.ylabel("2nd Dimention of the Dataset")
plt.legend(loc=1)
plt.show()
plt.close()
return
### Let's show the Train Set Points
plot_dataset(name='Train Set',points=train_x,labels=train_y)
### Let's show the Test Set Points
plot_dataset(name='Test Set',points=test_x,labels=test_y)
### Now let's plot for each classifier the predictions on the test set
plot_predictions(name='Bayesian', points=test_x, real_labels=test_y, predicted_labels=Bayes_test_y)
plot_predictions(name='Naive Bayes',points=test_x, real_labels=test_y, predicted_labels=Naive_Bayes_test_y)
plot_predictions(name='Knn Classifier',points=test_x,real_labels=test_y, predicted_labels=kNN_test_y)
plot_predictions(name='Logistic Regression Classifier',points=test_x,real_labels=test_y, predicted_labels=lry)
```
From the results it is shown that the Bayes classifier outperforms the other 3 classifiers.
This is expected due to the fact that the Bayes classifier is better that the Naive Bayes since the two classes have non-diagonal covariance matrices.
The same applies for the knn Classifier that we know from theory that has worse performance at small k and approaches the Bayes Classifier at high k.
The Bayes classifier would exhibit the same performance with Naive Bayes if the covariance matrices of normal distribution that generated the samples of the two classes was diagonal.
It should be also noted that the results obtained by Logistic regression on the test set are quite close to the ones obtained by the Bayes classifier who is optimal.The assumption here is of linear separability of the 2 classes that is strongly indicated visually.
| github_jupyter |
```
import os
import importlib.machinery
import importlib.util
loader = importlib.machinery.SourceFileLoader('baltic','/Users/evogytis/Documents/baltic/baltic.py')
spec = importlib.util.spec_from_loader(loader.name, loader)
bt = importlib.util.module_from_spec(spec)
loader.exec_module(bt)
base_path='/Users/evogytis/Documents/manuscripts/skeeters/data/narnavirus/'
RdRp=bt.loadNexus(os.path.join(base_path,"Batman.rooted.nexus"),absoluteTime=False,treestring_regex='tree tree_1 ',verbose=True).collapseBranches(lambda k: k.length==0,verbose=True)
Robin=bt.loadNexus(os.path.join(base_path,"Robin.rooted.nexus"),absoluteTime=False,treestring_regex='tree tree_1 ').collapseBranches(lambda k: k.length==0,verbose=True)
RdRp.treeStats()
Robin.treeStats()
RdRp.tipMap={k.numName: k.numName for k in RdRp.getExternal()}
Robin.tipMap={k.numName: k.numName for k in Robin.getExternal()}
for k in RdRp.getExternal():
k.name=k.numName
for k in Robin.getExternal():
k.name=k.numName
trees={'RdRp':RdRp,
'Robin':Robin}
segments=['RdRp','Robin']
%matplotlib inline
import matplotlib as mpl
from matplotlib import pyplot as plt
from matplotlib import gridspec
import numpy as np
typeface='Helvetica Neue'
# mpl.rcParams['font.weight']=400
mpl.rcParams['mathtext.fontset']='custom'
mpl.rcParams['font.sans-serif']=typeface
mpl.rcParams['mathtext.default']='sf'
mpl.rcParams['axes.labelweight']=300
mpl.rcParams['font.family']=typeface
mpl.rcParams['font.size']=22
tip_positions={x:{} for x in segments} ## remember the position of each tip in each tree
for t,tr in enumerate(trees.keys()): ## iterate over trees
cur_tree=trees[tr] ## fetch tree object
for k in cur_tree.Objects:
if k.branchType=='leaf':
tip_positions[tr][k.name]=(k.height,k.y) ## remember (X, Y) position of tip
cmap=mpl.cm.Spectral
tip_positions={x:{} for x in trees} ## remember the position of each tip in each tree
for t,tr in enumerate(trees.keys()): ## iterate over trees
cur_tree=trees[tr] ## fetch tree object
for k in cur_tree.Objects:
if k.branchType=='leaf':
tip_positions[tr][k.name]=(k.height,k.y) ## remember (X, Y) position of tip
cmap=mpl.cm.Spectral
for X in range(10): ## 10 untangling iterations
print('iteration %d'%(X+1))
for t,tr in enumerate(segments): ## iterate over each tree
print(tr)
ptr=segments[t-1] ## previous tree
ntr=segments[t] ## next tree
seg=trees[ptr] ## fetch appropriate tree
nex_seg=trees[ntr]
for k in sorted(nex_seg.Objects,key=lambda q:q.height): ## iterate over branches from most recent to oldest
if k.branchType=='node': ## can only sort nodes
leaves=[[seg.tipMap[tip] for tip in w.leaves if tip in seg.tipMap] if w.branchType=='node' else [w.name] for w in k.children] ## descendent tips in current order
# leaves=[[seg.tipMap[tip] for tip in w.leaves] if w.branchType=='node' else [w.name] for w in k.children] ## descendent tips in current order
for c in range(len(leaves)):
# leaves[c]=sorted(leaves[c],key=lambda x:tip_positions[ntr][x][1]) ## sort leaves according to their positions in the next tree
leaves[c]=sorted(leaves[c],key=lambda x:tip_positions[ntr][x][1] if x in tip_positions[ntr] else 0.0) ## sort leaves according to their positions in the next tree
ys=[sorted([tip_positions[ntr][w][1] for w in cl if w in tip_positions[ntr]]) for cl in leaves] ## extract y positions of descendents
merge_ys=sum(ys,[]) ## flatten list of tip y coordinates
m,M=int(min(merge_ys)+0.5),int(max(merge_ys)+0.5)
ypos=range(m,M+1) ## get y positions of tips in current order
order={i:x for i,x in enumerate(leaves)} ## dict of tip order: tip name
new_order=sorted(order.keys(),key=lambda x:-np.mean([(tip_positions[ptr][order[x][w]][1]-ypos[w]) for w in range(min([len(order[x]),len(ypos)])) if order[x][w] in tip_positions[ptr]])) ## get new order by sorting existing order based on y position differences
# new_order=sorted(order.keys(),key=lambda x:-np.mean([(tip_positions[ptr][order[x][w]][1]-ypos[w]) for w in range(len(order[x]))])) ## get new order by sorting existing order based on y position differences
if new_order!=range(len(leaves)): ## if new order is not current order
k.children=[k.children[i] for i in new_order] ## assign new order of child branches
nex_seg.drawTree() ## update y positions
for w in nex_seg.Objects: ## iterate over objects in next tree
if w.branchType=='leaf':
tip_positions[ntr][w.name]=(w.height,w.y) ## remember new positions
if t==0: ## if first tree
trees[segments[t]].drawTree() ## update positions
lvs=sorted([w for w in trees[segments[t]].Objects if w.branchType=='leaf'],key=lambda x:x.y) ## get leaves in y position order
norm=mpl.colors.Normalize(0,len(lvs))
pos_colours={w.name:cmap(norm(w.y)) for w in lvs} ## assign colour
fig = plt.subplots(figsize=(15,10),facecolor='w')
gs = gridspec.GridSpec(1, 1,hspace=0.0,wspace=0.0)
ax = plt.subplot(gs[0])
# traitName='HA'
tree1=RdRp
tree2=Robin
# cmap=mpl.cm.Spectral
cmap=mpl.cm.viridis
# for k in tree1.getInternal():
# if k.length>0.012:
# print(k.index,k.length,len(k.leaves))
tip_colours={}
colour_cycle=['steelblue','seagreen','indianred','purple','seagreen']
x=0
for k in sorted(tree2.getInternal()+tree2.getExternal(),key=lambda w: w.height):
if k.length>0.05:
c=colour_cycle[x%len(colour_cycle)]
if k.branchType=='node':
for w in k.leaves:
tip_colours[w]=c
else:
tip_colours[k.name]=c
x+=1
# print('2',k.index,k.length)
x_attr=lambda k: k.height ## branch x position is determined by height
y_attr=lambda k: k.y
b_func=lambda k: 4 ## branch width function
s_func=lambda k: 50 ## size of circle at tips
su_func=lambda k: 100 ## size of circle that sits underneath and acts as an outline
c_func=lambda k: 'k'
# ct_func=lambda k: cmap(k.y/float(tree2.ySpan)) ## call colour map with fraction that represents the y position of a tip (returns colour)
cu_func=lambda k: 'k' ## colour of circle sitting underneath the tip one is always black
z_func=lambda k: 100
zu_func=lambda k: 99
tree1.plotTree(ax,x_attr=x_attr,y_attr=y_attr,branchWidth=b_func,colour_function=c_func) ## plot tree
# tree1.plotPoints(ax,x_attr=x_attr,y_attr=y_attr,size_function=s_func,colour_function=ct_func,zorder_function=z_func) ## plot tip circle
# tree1.plotPoints(ax,x_attr=x_attr,y_attr=y_attr,size_function=su_func,colour_function=cu_func,zorder_function=zu_func) ## plot tip circle outline (another circle underneath)
cutoff=60
supported=lambda k: 'label' in k.traits and k.traits['label']>=cutoff
sp_func=lambda k: 30
spu_func=lambda k: 90
cp_func=lambda k: 'w'
tree1.plotPoints(ax,target=supported,x_attr=x_attr,y_attr=y_attr,size_function=sp_func,colour_function=cp_func,zorder_function=z_func) ## plot tip circle
tree1.plotPoints(ax,target=supported,x_attr=x_attr,y_attr=y_attr,size_function=spu_func,colour_function=cu_func,zorder_function=zu_func) ## plot tip circle
# skip=tree1.treeHeight*0.2 ## skip this many units between trees
skip=0.025
x_attr=lambda k: tree1.treeHeight+skip+tree2.treeHeight-k.height ## for tree2 we'll offset x coordinates by the height of the tree and invert branches
tree2.plotTree(ax,x_attr=x_attr,branchWidth=b_func,colour_function=c_func) ## plot black tree
# tree2.plotPoints(ax,x_attr=x_attr,size_function=s_func,colour_function=ct_func,zorder_function=z_func) ## plot circles at tips
# tree2.plotPoints(ax,x_attr=x_attr,size_function=su_func,colour_function=cu_func,zorder_function=zu_func) ## plot circles underneath tip circles as an outline
tree2.plotPoints(ax,target=supported,x_attr=x_attr,y_attr=y_attr,size_function=sp_func,colour_function=cp_func,zorder_function=z_func) ## plot tip circle
tree2.plotPoints(ax,target=supported,x_attr=x_attr,y_attr=y_attr,size_function=spu_func,colour_function=cu_func,zorder_function=zu_func) ## plot tip circle
ax.scatter(0.65,0.2,s=30,facecolor='w',edgecolor='none',zorder=99,transform=ax.transAxes)
ax.scatter(0.65,0.2,s=90,facecolor='k',edgecolor='none',zorder=98,transform=ax.transAxes)
ax.text(0.66,0.2,'bootstrap support >%s/100'%(cutoff),size=24,ha='left',va='center',transform=ax.transAxes)
for k in filter(lambda x: x.branchType=='leaf',tree1.Objects): ## grab leaf objects in tree1
x=k.height ## get height
y=k.y ## get y position
try:
matching_tip=tree2.getBranches(lambda x: x.branchType=='leaf' and x.name==k.name) ## fetch corresponding branch in tree2
match_y=matching_tip.y
xs=[x,tree1.treeHeight+0.1*skip,tree1.treeHeight+skip-0.1*skip,x_attr(matching_tip)] ## x coordinates for tangleline
ys=[y_attr(k),y_attr(k),match_y,match_y] ## y coordinates for tangleline
fc=tip_colours[matching_tip.name]
ec='k'
if 'CMS' not in k.numName:
# ec='w'
fc='slategrey'
# fc=cmap(match_y/float(tree2.ySpan))
ax.scatter(x,y_attr(k),s=s_func(k),facecolor=fc,edgecolor='none',zorder=z_func(k))
ax.scatter(x,y_attr(k),s=su_func(k),facecolor=ec,edgecolor='none',zorder=zu_func(k))
ax.scatter(x_attr(matching_tip),match_y,s=s_func(k),facecolor=fc,edgecolor='none',zorder=z_func(k))
ax.scatter(x_attr(matching_tip),match_y,s=su_func(k),facecolor=ec,edgecolor='none',zorder=zu_func(k))
ax.plot(xs,ys,color=fc,lw=6,solid_capstyle='round',alpha=0.7) ## plot tangleline
except:
pass
cumulative_displace=0
for cur_tree in [RdRp,Robin]:
tr='RdRp' if cur_tree==RdRp else 'Robin'
ax.text(cumulative_displace+cur_tree.treeHeight/2,max([T.ySpan for T in [RdRp,Robin]])+1,tr,size=26,va='bottom',ha='center')
xsm=np.arange(cumulative_displace,cumulative_displace+cur_tree.treeHeight,1e-2)
xsM=np.arange(cumulative_displace,cumulative_displace+cur_tree.treeHeight,5e-2)
if tr=='Robin':
xsm=np.arange(cumulative_displace+cur_tree.treeHeight,cumulative_displace,-1e-2)
xsM=np.arange(cumulative_displace+cur_tree.treeHeight,cumulative_displace,-5e-2)
ax.eventplot(xsm,linelengths=0.6,colors=['k'],lineoffsets=[-1])
ax.eventplot(xsM,linelengths=1.6,colors=['k'],lineoffsets=[-1])
ax.plot([cumulative_displace,cumulative_displace+cur_tree.treeHeight],[-1,-1],lw=2,color='k')
for t in xsM:
text=t
if tr=='Robin':
text=abs(t-cumulative_displace-cur_tree.treeHeight)
ax.text(t,-1.5,'%.2f'%(text),size=22,va='top',ha='center',weight=300,clip_on=False)
ax.text(cumulative_displace+cur_tree.treeHeight/2.0,-4.5,'subs/site',
size=28,va='top',ha='center',weight=300,clip_on=False)
cumulative_displace+=cur_tree.treeHeight
cumulative_displace+=skip
[ax.spines[loc].set_visible(False) for loc in ['top','right','left','bottom']]
ax.tick_params(axis='x',size=0)
ax.tick_params(axis='y',size=0)
ax.set_xticklabels([])
ax.set_yticklabels([])
ax.set_ylim(-1,max(tree1.ySpan,tree2.ySpan)+1) ## set y limits
ax.set_xlim(-0.001,tree1.treeHeight+skip+tree2.treeHeight+0.001)
plt.savefig('/Users/evogytis/Documents/manuscripts/skeeters/figures/figSX_narna_tanglegram.pdf',dpi=300,bbox_inches='tight')
plt.savefig('/Users/evogytis/Documents/manuscripts/skeeters/figures/figSX_narna_tanglegram.png',dpi=300,bbox_inches='tight')
plt.show()
```
| github_jupyter |
# Anacycliques
## Définition
Pour cet exercice, nous nous focaliserons sur une catégorie de mots qui conservent un sens lorsqu’on les lit de droite à gauche : les anacycliques. De la famille des anagrammes, ils se distinguent des palindromes en ce que leur sens n’est pas forcément identique dans les deux sens de lecture :
- *amuser* et *résuma*
- *super* et *repus*
- *trot* et *tort*
- *été* et *été* (anacyclique et palindrome)
- …
Pour reconnaître un anacylique, il ne suffit donc pas seulement de modifier le sens de lecture d’une chaîne de caractères puis de la comparer avec la version originale, mais il faut également s’assurer que le résultat produit ait un sens. Le mot *tabernacle*, par exemple, produit à l’envers la chaîne *elcanrebat*, qui n’a aucun sens.
## Conception
L’objectif de cet exercice est de fournir un programme qui repère dans un texte tous les anacycliques et dénombre leurs occurrences. Nous travaillerons sur un extrait du *Ventre de Paris* (1873) de Zola. Vous comprendrez qu’analyser le texte mobiliserait trop de ressources machines.
Recensons à présent les besoins du programme :
- ouvrir le fichier
- découper en mots
- (boucle) renverser le sens de lecture de chaque mot
- (test) si le mot est présent dans un lexique de référence :
- le sauvegarder et incrémenter le compteur
Comme on s’attend à ce qu’un anacyclique soit présent plus d’une fois dans le texte, l’enregistrer dans un dictionnaire avec le nombre de ses occurrences serait plus opportun.
## Astuces
### Découper un texte en mots
Vous utiliserez la méthode `split()` sans vous soucier du bruit engendré.
### Renverser le sens de lecture d’un mot
La technique la plus simple parmi toutes celles imaginables, recourt aux *slices* :
```
mot = "bazinga"
envers = mot[::-1]
print(envers)
```
La syntaxe `mot[::-1]` affiche chaque caractère du mot avec un pas négatif de -1, comprenez : en partant de la fin.
### Accéder à une valeur dans un dictionnaire
Pensez à la méthode `get()` qui permet, lorsqu’une clé n’est pas trouvée, de retourner une valeur au lieu de lever l’exception `KeyError` :
```
houses = {
'lannister': ['Jaime', 'Cersei'],
'bolton': ['Roose', 'Ramsey']
}
# If the key 'stark' is not in the dict,
# the second parameter will be returned.
houses.get('stark', 'Désolé, aucun Stark ici.')
```
## Code de départ
Pour démarrer, vous disposez des 100 000 premiers caractères du roman dans un objet `text`, ainsi que d’un lexique de référence dans un objet `lexicon`. À vous de produire tout le reste !
**Remarque :** le lexique de référence est un extrait de [*Lexique 3*](http://www.lexique.org/) où nous n’avons conservé que la colonne *ortho* à des fins de performance.
```
import csv
# Loading with the data
with open('../data/le-ventre-de-paris.txt') as file:
# Reads only the first 100.000 characters
text = file.read(100000)
# Loading a french lexicon
with open('../data/lexique383-ortho.tsv') as csvfile:
reader = csv.DictReader(csvfile, delimiter="\t")
lexicon = [ line["ortho"] for line in reader ]
# Your code here
# Text into words
words = text.split()
# A dictionary of ananyms
ananyms = dict()
"""If a word in its reversed form is
in the lexicon: put it in the dictionary
with a counter initialized at 0. The next
occurrence found will increment the counter.
"""
for word in words:
reverse = word[::-1]
if reverse in lexicon:
ananyms.update({
word: ananyms.get(word, 0) + 1
})
```
| github_jupyter |
This is an supervised classification example taken from the KDD 2009 cup. A copy of the data and details can be found here: [https://github.com/WinVector/PDSwR2/tree/master/KDD2009](https://github.com/WinVector/PDSwR2/tree/master/KDD2009). The problem was to predict account cancellation ("churn") from very messy data (column names not given, numeric and categorical variables, many missing values, some categorical variables with a large number of possible levels). In this example we show how to quickly use `vtreat` to prepare the data for modeling. `vtreat` takes in `Pandas` `DataFrame`s and returns both a treatment plan and a clean `Pandas` `DataFrame` ready for modeling.
Load our packages/modules.
```
import pandas
import xgboost
import vtreat
import vtreat.cross_plan
import numpy.random
import wvpy.util
import scipy.sparse
vtreat.__version__
```
Read in explanitory variables.
```
# data from https://github.com/WinVector/PDSwR2/tree/master/KDD2009
dir = "../../../PracticalDataScienceWithR2nd/PDSwR2/KDD2009/"
d = pandas.read_csv(dir + 'orange_small_train.data.gz', sep='\t', header=0)
vars = [c for c in d.columns]
d.shape
```
Read in dependent variable we are trying to predict.
```
churn = pandas.read_csv(dir + 'orange_small_train_churn.labels.txt', header=None)
churn.columns = ["churn"]
churn.shape
churn["churn"].value_counts()
```
Arrange test/train split.
```
numpy.random.seed(2020)
n = d.shape[0]
# https://github.com/WinVector/pyvtreat/blob/master/Examples/CustomizedCrossPlan/CustomizedCrossPlan.md
split1 = vtreat.cross_plan.KWayCrossPlanYStratified().split_plan(n_rows=n, k_folds=10, y=churn.iloc[:, 0])
train_idx = set(split1[0]['train'])
is_train = [i in train_idx for i in range(n)]
is_test = numpy.logical_not(is_train)
```
(The reported performance runs of this example were sensitive to the prevalance of the churn variable in the test set, we are cutting down on this source of evaluation variarance by using the stratified split.)
```
d_train = d.loc[is_train, :].copy()
churn_train = numpy.asarray(churn.loc[is_train, :]["churn"]==1)
d_test = d.loc[is_test, :].copy()
churn_test = numpy.asarray(churn.loc[is_test, :]["churn"]==1)
```
Take a look at the dependent variables. They are a mess, many missing values. Categorical variables that can not be directly used without some re-encoding.
```
d_train.head()
d_train.shape
```
Try building a model directly off this data (this will fail).
```
fitter = xgboost.XGBClassifier(n_estimators=10, max_depth=3, objective='binary:logistic')
try:
fitter.fit(d_train, churn_train)
except Exception as ex:
print(ex)
```
Let's quickly prepare a data frame with none of these issues.
We start by building our treatment plan, this has the `sklearn.pipeline.Pipeline` interfaces.
```
plan = vtreat.BinomialOutcomeTreatment(
outcome_target=True,
params=vtreat.vtreat_parameters({
'filter_to_recommended': True,
'sparse_indicators': True,
}))
```
Use `.fit_transform()` to get a special copy of the treated training data that has cross-validated mitigations againsst nested model bias. We call this a "cross frame." `.fit_transform()` is deliberately a different `DataFrame` than what would be returned by `.fit().transform()` (the `.fit().transform()` would damage the modeling effort due nested model bias, the `.fit_transform()` "cross frame" uses cross-validation techniques similar to "stacking" to mitigate these issues).
```
cross_frame = plan.fit_transform(d_train, churn_train)
```
Take a look at the new data. This frame is guaranteed to be all numeric with no missing values, with the rows in the same order as the training data.
```
cross_frame.head()
cross_frame.shape
```
Pick a recommended subset of the new derived variables.
```
plan.score_frame_.head()
model_vars = numpy.asarray(plan.score_frame_["variable"][plan.score_frame_["recommended"]])
len(model_vars)
```
Fit the model
```
cross_frame.dtypes
# fails due to sparse columns
# can also work around this by setting the vtreat parameter 'sparse_indicators' to False
try:
cross_sparse = xgboost.DMatrix(data=cross_frame.loc[:, model_vars], label=churn_train)
except Exception as ex:
print(ex)
# also fails
try:
cross_sparse = scipy.sparse.csc_matrix(cross_frame[model_vars])
except Exception as ex:
print(ex)
# works
cross_sparse = scipy.sparse.hstack([scipy.sparse.csc_matrix(cross_frame[[vi]]) for vi in model_vars])
# https://xgboost.readthedocs.io/en/latest/python/python_intro.html
fd = xgboost.DMatrix(
data=cross_sparse,
label=churn_train)
x_parameters = {"max_depth":3, "objective":'binary:logistic', "eval_metric": 'logloss'}
cv = xgboost.cv(x_parameters, fd, num_boost_round=100, verbose_eval=False)
cv.head()
best = cv.loc[cv["test-logloss-mean"]<= min(cv["test-logloss-mean"] + 1.0e-9), :]
best
ntree = best.index.values[0]
ntree
fitter = xgboost.XGBClassifier(n_estimators=ntree, max_depth=3, objective='binary:logistic')
fitter
model = fitter.fit(cross_sparse, churn_train)
```
Apply the data transform to our held-out data.
```
test_processed = plan.transform(d_test)
```
Plot the quality of the model on training data (a biased measure of performance).
```
pf_train = pandas.DataFrame({"churn":churn_train})
pf_train["pred"] = model.predict_proba(cross_sparse)[:, 1]
wvpy.util.plot_roc(pf_train["pred"], pf_train["churn"], title="Model on Train")
```
Plot the quality of the model score on the held-out data. This AUC is not great, but in the ballpark of the original contest winners.
```
test_sparse = scipy.sparse.hstack([scipy.sparse.csc_matrix(test_processed[[vi]]) for vi in model_vars])
pf = pandas.DataFrame({"churn":churn_test})
pf["pred"] = model.predict_proba(test_sparse)[:, 1]
wvpy.util.plot_roc(pf["pred"], pf["churn"], title="Model on Test")
```
Notice we dealt with many problem columns at once, and in a statistically sound manner. More on the `vtreat` package for Python can be found here: [https://github.com/WinVector/pyvtreat](https://github.com/WinVector/pyvtreat). Details on the `R` version can be found here: [https://github.com/WinVector/vtreat](https://github.com/WinVector/vtreat).
We can compare this to the [R solution (link)](https://github.com/WinVector/PDSwR2/blob/master/KDD2009/KDD2009vtreat.md).
We can compare the above cross-frame solution to a naive "design transform and model on the same data set" solution as we show below. Note we are leaveing filter to recommended on, to show the non-cross validated methodology still fails in an "easy case."
```
plan_naive = vtreat.BinomialOutcomeTreatment(
outcome_target=True,
params=vtreat.vtreat_parameters({'filter_to_recommended':True}))
plan_naive.fit(d_train, churn_train)
naive_frame = plan_naive.transform(d_train)
model_vars = numpy.asarray(plan_naive.score_frame_["variable"][plan_naive.score_frame_["recommended"]])
len(model_vars)
naive_sparse = scipy.sparse.hstack([scipy.sparse.csc_matrix(naive_frame[[vi]]) for vi in model_vars])
fd_naive = xgboost.DMatrix(data=naive_sparse, label=churn_train)
x_parameters = {"max_depth":3, "objective":'binary:logistic'}
cvn = xgboost.cv(x_parameters, fd_naive, num_boost_round=100, verbose_eval=False)
bestn = cvn.loc[cvn["test-logloss-mean"] <= min(cvn["test-logloss-mean"] + 1.0e-9), :]
bestn
ntreen = bestn.index.values[0]
ntreen
fittern = xgboost.XGBClassifier(n_estimators=ntreen, max_depth=3, objective='binary:logistic')
fittern
modeln = fittern.fit(naive_sparse, churn_train)
test_processedn = plan_naive.transform(d_test)
test_processedn = scipy.sparse.hstack([scipy.sparse.csc_matrix(test_processedn[[vi]]) for vi in model_vars])
pfn_train = pandas.DataFrame({"churn":churn_train})
pfn_train["pred_naive"] = modeln.predict_proba(naive_sparse)[:, 1]
wvpy.util.plot_roc(pfn_train["pred_naive"], pfn_train["churn"], title="Overfit Model on Train")
pfn = pandas.DataFrame({"churn":churn_test})
pfn["pred_naive"] = modeln.predict_proba(test_processedn)[:, 1]
wvpy.util.plot_roc(pfn["pred_naive"], pfn["churn"], title="Overfit Model on Test")
```
Note the naive test performance is worse, despite its far better training performance. This is over-fit due to the nested model bias of using the same data to build the treatment plan and model without any cross-frame mitigations.
| github_jupyter |
# Chapter 10: Sound Sharing and Retreival
## a) Create Audio Database
```
import os
import pandas as pd
import numpy as np
import freesound
from whoosh.fields import Schema, ID, TEXT, KEYWORD, NUMERIC
from whoosh.index import create_in
try:
from freesound_apikey import FREESOUND_API_KEY
except ImportError:
print('Can\'t load your Freesound API key!\nPlease request an API key at http://freesound.org/apiv2/apply/ and paste it in a file named \'freesound_apikey.py\'.')
def create_text_index():
schema = Schema(
name = TEXT,
description = TEXT,
username = TEXT,
tags = KEYWORD,
duration = NUMERIC,
license = TEXT,
pandas_index = ID(stored=True),
)
text_index = create_in("text_index", schema)
return text_index
def replace_license_label(record):
short_name = '' # Required for using as whoosh facets
if 'publicdomain' in record['license']: short_name = 'CC_0'
elif 'by/3.0' in record['license']: short_name = 'CC_BY'
elif 'by-nc/3.0' in record['license']: short_name = 'CC_BY_NC'
elif 'sampling+' in record['license']: short_name = 'S_Plus'
record['license'] = short_name
return record
def query_freesound(q):
freesound_client = freesound.FreesoundClient()
freesound_client.set_token(FREESOUND_API_KEY)
pager = freesound_client.text_search(
query = q,
fields = "id,name,tags,username,analysis,duration," \
+ "description,license,previews",
descriptors = "lowlevel.mfcc.mean,lowlevel.mfcc.var," \
+ "lowlevel.spectral_centroid.mean",
group_by_pack = 1,
page_size = 50
)
return [sound for sound in pager if sound.analysis]
def make_pandas_record(fs_object):
record = {key: fs_object.as_dict()[key]
for key in metadata_fields}
record["path"] = "files/" + \
fs_object.previews.preview_lq_mp3.split("/")[-1]
lowlevel = fs_object.analysis.lowlevel
for index, val in enumerate(lowlevel.mfcc.mean):
record["mfcc_mean_%i"%index] = val
for index, val in enumerate(lowlevel.mfcc.var):
record["mfcc_var_%i"%index] = val
record["spectral_centroid"] = lowlevel.spectral_centroid.mean
return replace_license_label(record)
def make_whoosh_record(fs_object, pandas_index):
record = {key: fs_object.as_dict()[key]
for key in metadata_fields}
record["pandas_index"] = str(pandas_index)
return replace_license_label(record)
def make_db():
sounds = sum([query_freesound(animal)
for animal in animal_sounds],[])
for sound in sounds:
sound.retrieve_preview("files/")
data_frame = pd.DataFrame(
[make_pandas_record(s) for s in sounds])
text_index = create_text_index()
writer = text_index.writer()
for index, sound in enumerate(sounds):
text_document = make_whoosh_record(sound, index)
writer.add_document(**text_document)
writer.commit()
data_frame.to_csv('database.csv')
print('Created dataset with %i sounds!' % len(sounds))
# Make the db with animal sounds
animal_sounds = ["dog bark", "cat meow", "lion roar", "nightingale"]
metadata_fields = ["name", "tags", "username",
"description","duration", "license"]
if not os.path.exists("text_index"): os.mkdir("text_index")
if not os.path.exists("files"): os.mkdir("files")
make_db()
```
| github_jupyter |
```
from datetime import date, timedelta
from Stock import *
s = date(2020,1,1)
e = date(2021,12,20)
tesla = Stock("tsla")
tesla.load_data()
#tesla.add_data_range(s,e,stockpath='pricedata/tsla.csv')
#tesla.save_data()
# adding individual days
#tesla.add_data(s)
len(tesla.df)
# Analysis
import seaborn as sns
import matplotlib.pyplot as plt
import re
df = tesla.df.dropna()
df['domain'] = df.apply(
lambda row: re.sub('www3.|www.','',row.link.split('//')[1].split('/')[0]), axis=1)
df1 = df.iloc[0:len(df)//2] #df.sample(frac=0.5)
df2 = df.drop(df1.index)
print(len(df))
print(len(df1))
print(len(df2))
count = {}
for p, tx in df1[['percent_change','text']].values:
tx = re.sub('[^a-zA-Z ]',' ',tx)
for word in tx.split():
if word.lower() not in count:
count[word.lower()] = [0,0]
count[word.lower()][p < 0] += 1
prob = {k:list(np.array(count[k])/(sum(count[k]))) for k in count}
len(prob)
print('--------- positive ---------')
for k in sorted(prob,key=prob.get,reverse=True):
if prob[k][0] >= 0.70 and sum(count[k]) > 25:
print(k,prob[k],count[k])
print('--------- negative ---------')
for k in sorted(prob,key=prob.get,reverse=False):
if prob[k][1] >= 0.70 and sum(count[k]) > 25:
print(k,prob[k],count[k])
rel = {}
for k in sorted(prob,key=prob.get,reverse=True):
if prob[k][0] >= 0.60 and sum(count[k]) > 75:
rel[k] = prob[k]
for k in sorted(prob,key=prob.get,reverse=False):
if prob[k][1] >= 0.60 and sum(count[k]) > 25:
rel[k] = prob[k]
len(rel)
accuracy = 0
for num in range(len(df2)):
prediction = [0,0]
count = [0, 0]
tx = df2['text'].iloc[num]
tx = re.sub('[^a-zA-Z ]',' ',tx)
for word in tx.split():
if word.lower() in prob:
prediction[prob[word.lower()][0] < 0.5] += prob[word.lower()][prob[word.lower()][0] < 0.5]
count[prob[word.lower()][0] < 0.5] += 1
if count[0] > 0:
prediction[0] /= count[0]
if count[1] > 0:
prediction[1] /= count[1]
prediction[0] /= sum(prediction)
prediction[1] = 1 - prediction[0]
#print(prediction,df2['percent_change'].iloc[num])
accuracy += int((df2['percent_change'].iloc[num] < 0) == (prediction[0] < 0))
accuracy /= len(df2)
print(accuracy)
prob['germany']
def encode_text(tx):
enc = np.zeros(len(rel))
tx = re.sub('[^a-zA-Z ]',' ',tx)
tx = tx.split()
#for i, k in enumerate(prob):
# if k in tx:
# enc[i] = 1
enc = [int(k in tx) for k in rel]
return enc
dataset = df2.apply(
lambda row: [row.percent_change,encode_text(row.text)],
axis=1,result_type='expand')
dataset.columns = ['label','data']
labels = dataset['label'].values.astype(np.float64)
inputs = np.array(dataset['data'].values.tolist())
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(inputs, labels, test_size=0.3, shuffle=True)
#from sklearn.ensemble import AdaBoostRegressor
#clf = AdaBoostRegressor(loss='square')
#clf.fit(X_train, y_train)
#print(clf.score(X_test,y_test))
from sklearn.ensemble import AdaBoostClassifier
clf = AdaBoostClassifier()
clf.fit(X_train, (y_train < 0).astype(int))
print(clf.score(X_test,(y_test < 0).astype(int)))
#from sklearn.ensemble import RandomForestRegressor
#clf = RandomForestRegressor()
#clf.fit(X_train, y_train)
#print(clf.score(X_test,y_test))
from sklearn.ensemble import RandomForestClassifier
clf = RandomForestClassifier()
clf.fit(X_train, (y_train < 0).astype(int))
print(clf.score(X_test,(y_test < 0).astype(int)))
from sklearn.linear_model import SGDClassifier
clf = SGDClassifier(max_iter=1000, tol=1e-10,shuffle=True)
clf.fit(X_train, (y_train < 0).astype(int))
print(clf.score(X_train,(y_train < 0).astype(int)))
print(clf.score(X_test,(y_test < 0).astype(int)))
#https://www.geeksforgeeks.org/python-word-embedding-using-word2vec/
time_res = np.array([int(r == (y_test[i] < 0)) for i,r in enumerate(clf.predict(X_test))])
# average length of correct guesses in a row
length = 0
count = 0
avg = 0
for x in time_res:
if x == 0:
count+=1
avg += length
length = 0
elif x == 1:
length+=1
avg /= count
avg
```
| github_jupyter |
# About: scpによるリストア
---
Moodle構築環境のデータ、設定ファイルなどのバックアップをscpを利用してリストアします。
## 概要
scpを利用してMoodle環境のリストアを行います。
### 前提条件
この Notebook を実行するには事前に以下のものを準備する必要があります。
* リストア対象のホストからバックアップ保存先のホストにSSH公開鍵認証でログインできること
* リストア先となるVCノード/EC2インスタンス/Azure仮想マシンが作成済であること
リストア先となる環境は「011-VCノードの作成」、「012-EC2インスタンスの作成」、「013-Azure仮想マシンの作成」のいずれかのNotebookで作成することができます。
また、リストアの操作により既存の環境を損ねることを避けるため、リストア先となるディレクトリ `/srv/moodle` が存在していないことを前提としています。
## パラメータ設定
### リストア先のMoodle環境を指定する
リストア先となるノードをAnsibleのグループ名で指定してください。
```
# (例)
# target_group = 'Moodle'
target_group =
```
リストア対象のノードにアクセスできることを確認します。
```
!ansible {target_group} -m ping
```
リストア先となるディレクトリ `/srv/moodle` が存在していないことを確認します。
```
!ansible {target_group} -a 'test ! -e /srv/moodle'
```
### SSH公開鍵認証
バックアップの保存先にSSHでアクセスするためのパラメータを指定します。
バックアップ先のホストにログインする際のユーザ名を指定してください。
```
# (例)
# backup_user = 'user01'
backup_user =
```
バックアップ先のホスト名を指定してください。
```
# (例)
# backup_host = 'backup.example.org'
backup_host =
```
バックアップ先のホストにログインする際のSSHの秘密鍵のパスを指定してください。ここで指定するパスはMoodle構築環境におけるパスを指定する必要があります。
```
# (例)
# backup_ssh_identity = '~/.ssh/id_rsa'
backup_ssh_identity =
```
指定されたパスに秘密鍵のファイルが存在していることをチェックします。次のセルを実行してエラーにならないことを確認してください。
```
!ansible {target_group} -m shell -a 'test -f {backup_ssh_identity}'
```
SSHの公開鍵ペアファイルをまだ作成していない場合は、次のセルのコメント `#` を外し実行することで公開鍵認証のファイルを作成することができます。
```
# !ansible {target_group} -m shell -a \
# 'test -f {backup_ssh_identity} || \
# ssh-keygen -q -t rsa -N "" -f {backup_ssh_identity} \
# && cat {backup_ssh_identity}.pub'
```
バックアップ先のホストにログインする前に `~/.ssh/known_hosts` を更新しておきます。
> 既に `~/.ssh/known_hosts` にバックアップ先のホストを登録してある場合は次のセルの実行をスキップしてください。
```
!ansible {target_group} -m shell -a \
'ssh-keyscan {backup_host} >> ~/.ssh/known_hosts'
```
バックアップ先のホストにSSHでログインしてコマンド `ls -la` を実行してみます。
```
ssh_command = f'ssh -i {backup_ssh_identity} {backup_user}@{backup_host}'
!ansible {target_group} -m shell -a \
'{ssh_command} ls -la'
```
### リストアを行うバックアップファイルを指定する
バックアップファイルの保存先であるディレクトリを指定してください。
```
# (例)
# backup_dir = 'moodle-simple/Moodle/2020-XX-XXTXX:XX:XX.XXXXXX'
backup_dir =
```
指定したバケット名、ディレクトリにバックアップファイルがあることを確認します。次のセルを実行してMoodle環境のバックアップファイル `db.sql.gz`, `moodle.tar.gz` が存在していることを確認してください。
```
!ansible {target_group} -m shell -a \
'{ssh_command} ls -la {backup_dir}'
```
## リストア
Moodle環境のリストアを行います。
### ディレクトリの作成
リストア先となるディレクトリを作成します。
```
!ansible {target_group} -b -m file -a \
'path=/srv/moodle state=directory owner={{{{ansible_ssh_user}}}}'
```
デフォルトのコンテナログの出力先ディレクトリを作成します。
```
!ansible {target_group} -b -m file -a 'path=/var/log/httpd state=directory'
!ansible {target_group} -b -m file -a 'path=/var/log/mysql owner=999 group=adm state=directory'
```
### コンテナ構成、Moodleデータなどのリストア
設定ファイルなどのバックアップファイル `moodle.tar.gz` をリストアします。
リストア対象のバックアップファイルを確認します。
```
moodle_backup = backup_dir + '/moodle.tar.gz'
print(moodle_backup)
```
リストアを行います。
```
!ansible {target_group} -m shell -a \
'{ssh_command} cat {moodle_backup} \
| bash -c "sudo tar xzpf - -C /srv/moodle"'
```
リストアされたことを確認します。
```
!ansible {target_group} -a 'chdir=/srv/moodle \
tree -L 3 -F'
```
### DBデータ
DBデータを復元するためのSQLファイルを取得します。
リストア対象のバックアップファイルを確認します。
```
db_backup = backup_dir + '/db.sql.gz'
print(db_backup)
```
リストア先となるディレクトリを作成します。
```
!ansible {target_group} -b -m file -a \
'path=/srv/moodle/db/data state=directory owner={{{{ansible_ssh_user}}}}'
!ansible {target_group} -b -m file -a \
'path=/srv/moodle/db/sql state=directory owner={{{{ansible_ssh_user}}}}'
```
DBデータをリストアするためのSQLファイルファイルを配置します。配置したSQLファイルはDBコンテナ起動時に自動的に実行されます。
```
!ansible {target_group} -m shell -a \
'{ssh_command} cat {db_backup} > /srv/moodle/db/sql/backup.sql.gz'
```
### コンテナイメージ
Dockerコンテナイメージのリストアを行います。
> コンテナイメージのバックアップを作成していない、あるいはレポジトリのコンテナイメージを利用する場合はこの節を実行せずにスキップしてください。
コンテナイメージをリストアする前の状態を確認しておきます。コンテナイメージの一覧を表示します。
```
!ansible {target_group} -a 'docker images'
```
リストア対象のバックアップファイルを確認します。
```
img_backup = backup_dir + '/container-image.tar.gz'
print(img_backup)
```
コンテナイメージをリストアします。
> バックアップファイルが存在していない場合はエラーになります。
```
!ansible {target_group} -m shell -a \
'{ssh_command} cat {img_backup} | \
gzip -cd | docker load'
```
リストアを行った後の状態を確認します。コンテナイメージの一覧を表示します。
```
!ansible {target_group} -a 'docker images'
```
### コンテナの起動
リストア環境のコンテナを起動します。
```
!ansible {target_group} -a 'chdir=/srv/moodle \
docker-compose up -d'
```
コンテナの状態を確認します。`State`が`Up`となっていることを確認してください。
```
!ansible {target_group} -a 'chdir=/srv/moodle \
docker-compose ps'
```
### crontab
crontabの設定を復元します。
現在の crontab の設定を確認します。
```
try:
!ansible {target_group} -a 'crontab -l'
except:
pass
```
バックアップファイルからcrontabの復元を行います。
```
!ansible {target_group} -a 'crontab /srv/moodle/misc/crontab'
```
復元後の設定を確認します。
```
!ansible {target_group} -a 'crontab -l'
```
### logrotate
コンテナログのログローテーションの設定を復元します。
```
!ansible {target_group} -b -m shell -a \
'cp /srv/moodle/misc/logrotate.d/* /etc/logrotate.d/'
```
### メンテナンスモードの解除
メンテナンスモードに変更してからバックアップを作成した場合はメンテナンスモードの解除が必要となります。
```
!ansible {target_group} -a 'chdir=/srv/moodle docker-compose exec -T moodle \
/usr/bin/php /var/www/html/admin/cli/maintenance.php --disable'
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.